code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 255} colab_type="code" id="4WBvY6GVYBtS" outputId="3d8e3fd4-5c05-4d62-c22a-a38bb15ee8d9"
from math import sqrt
import tensorflow as tf
from tensorflow import keras
import pandas as pd
from tensorflow.keras import Sequential
from tensorflow.keras.layers import LSTM, Dense, Dropout, Conv1D, GRU
from tensorflow.keras.losses import mean_squared_error
from numpy.core._multiarray_umath import concatenate
from sklearn.preprocessing import MinMaxScaler
import matplotlib.pyplot as plt
# supervised监督学习函数
def series_to_supervised(data, columns, n_in=1, n_out=1, dropnan=True):
n_vars = 1 if isinstance(data, list) else data.shape[1]
df = pd.DataFrame(data)
cols, names = list(), list()
# input sequence (t-n, ... t-1)
for i in range(n_in, 0, -1):
cols.append(df.shift(i))
names += [('%s%d(t-%d)' % (columns[j], j + 1, i))
for j in range(n_vars)]
# forecast sequence (t, t+1, ... t+n)
for i in range(0, n_out):
cols.append(df.shift(-i))
if i == 0:
names += [('%s%d(t)' % (columns[j], j + 1)) for j in range(n_vars)]
else:
names += [('%s%d(t+%d)' % (columns[j], j + 1, i))
for j in range(n_vars)]
# put it all together
agg = pd.concat(cols, axis=1)
agg.columns = names
# drop rows with NaN values
if dropnan:
clean_agg = agg.dropna()
return clean_agg
# return agg
dataset = pd.read_csv(
'Machine_usage_groupby.csv')
dataset_columns = dataset.columns
values = dataset.values
print(dataset)
# 归一化处理
scaler = MinMaxScaler(feature_range=(0, 1))
scaled = scaler.fit_transform(values)
# 监督学习
reframed = series_to_supervised(scaled, dataset_columns, 1, 1)
values = reframed.values
# 学习与检测数据的划分
n_train_hours = 20000
train = values[:n_train_hours, :]
test = values[n_train_hours:, :]
# 监督学习结果划分
train_x, train_y = train[:, :-1], train[:, -1]
test_x, test_y = test[:, :-1], test[:, -1]
# 为了在LSTM中应用该数据,需要将其格式转化为3D format,即[Samples, timesteps, features]
train_X = train_x.reshape((train_x.shape[0], 1, train_x.shape[1]))
test_X = test_x.reshape((test_x.shape[0], 1, test_x.shape[1]))
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="CHM-UGeYYgZN" outputId="e7dd414f-87a9-440d-b685-eab153b4d915"
model = Sequential()
model.add(Conv1D(filters=32, kernel_size=5,
strides=1, padding="causal",
activation="relu"))
model.add(
GRU(
32,
input_shape=(
train_X.shape[1],
train_X.shape[2]),
return_sequences=True))
model.add(GRU(16, input_shape=(train_X.shape[1], train_X.shape[2])))
model.add(Dense(16, activation="relu"))
model.add(Dense(1))
model.compile(loss=tf.keras.losses.Huber(),
optimizer='adam',
metrics=["mse"])
history = model.fit(
train_X,
train_y,
epochs=50,
batch_size=72,
validation_data=(
test_X,
test_y),
verbose = 2)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="8uPW-5baYiLN" outputId="133dd57f-d8ab-4957-d7bc-32f2520ebb83"
#画图
plt.plot(history.history['loss'], label='train')
plt.plot(history.history['val_loss'], label='test')
plt.legend()
plt.show()
# make the prediction
yHat = model.predict(test_X)
inv_yHat = concatenate((yHat, test_x[:, 1:]), axis=1) # 数组拼接
inv_yHat = inv_yHat[:, 0]
test_y = test_y.reshape((len(test_y), 1))
inv_y = concatenate((test_y, test_x[:, 1:]), axis=1)
inv_y = inv_y[:, 0]
rmse = sqrt(mean_squared_error(inv_yHat, inv_y))
print('Test RMSE: %.8f' % rmse)
mse = mean_squared_error(inv_yHat, inv_y)
print('Test MSE: %.8f' % mse)
yhat = model.predict(test_X)
test_X_reshaped = test_X.reshape((test_X.shape[0], test_X.shape[2]))
inv_yhat = concatenate((yhat, yhat, test_X_reshaped[:, 1:]), axis=1)
inv_yhat = inv_yhat[:, 0]
test_y = test_y.reshape((len(test_y), 1))
inv_y = concatenate((test_y, test_y, test_X_reshaped[:, 1:]), axis=1)
inv_y = inv_y[:, 0]
plt.plot(inv_yhat, label='prediction')
plt.plot(inv_y, label='real')
plt.xlabel('time')
plt.ylabel('cpu_usage_percent')
plt.legend()
plt.show()
plt.plot(inv_yhat[:500], label='prediction')
plt.plot(inv_y[:500], label='real_cpu_usage_percent')
plt.xlabel('time')
plt.ylabel('cpu_usage_percent')
plt.legend()
plt.show()
plt.plot(inv_yhat[:50], label='prediction')
plt.plot(inv_y[:50], label='real_cpu_usage_percent')
plt.xlabel('time')
plt.ylabel('cpu_usage_percent')
plt.legend()
plt.show()
| Project_Alibaba_workload/E50_Alibaba_cluster_predict_compare/Train_20000/COV+GRU+GRU+D+D+E50/.ipynb_checkpoints/COV+GRU+GRU+D+D+E50-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import miepython as mp
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
mpl.rcParams["xtick.direction"] = "in"
mpl.rcParams["ytick.direction"] = "in"
mpl.rcParams["lines.markeredgecolor"] = "k"
mpl.rcParams["lines.markeredgewidth"] = 1
mpl.rcParams["figure.dpi"] = 200
from matplotlib import rc
rc('font', family='serif')
rc('text', usetex=True)
rc('xtick', labelsize='medium')
rc('ytick', labelsize='medium')
def cm2inch(value):
return value/2.54
link = r"https://refractiveindex.info/tmp/data/organic/(C8H8)n%20-%20polystyren/Zhang.txt"
poly = np.genfromtxt(link, delimiter='\t')
N = len(poly)//2
poly_lam = poly[1:N,0][:40]
poly_mre = poly[1:N,1][:40]
poly_mim = poly[N+1:,1][:40]
poly_mim
# +
plt.figure(figsize=( cm2inch(16),cm2inch(8)))
plt.plot(poly_lam*1000,poly_mre,color='tab:blue')
#plt.xlim(300,800)
#plt.ylim(-,3)
plt.xlabel('Wavelength (nm)')
plt.ylabel('$n_r$')
#plt.text(350, 1.2, '$m_{re}$', color='blue', fontsize=14)
#plt.text(350, 2.2, '$m_{im}$', color='red', fontsize=14)
ax=plt.gca()
ax.spines['left'].set_color("red")
ax2=ax.twinx()
plt.semilogy(poly_lam*1000,poly_mim,color='tab:red')
plt.ylabel('$n_i$', color = "tab:red")
plt.tight_layout()
plt.savefig("refractive_index.pdf")
# +
plt.semilogy(poly_lam*1000,poly_mim,color='red')
#plt.xlim(300,800)
#plt.ylim(-,3)
plt.xlabel('Wavelength (nm)')
plt.ylabel('Refractive Index')
#plt.text(350, 1.2, '$m_{re}$', color='blue', fontsize=14)
#plt.text(350, 2.2, '$m_{im}$', color='red', fontsize=14)
plt.title('Complex part of Refractive Index of Polystyrene')
plt.show()
# +
r = 1.5 #radius in microns
x = 2*np.pi*r/poly_lam;
m = poly_mre - 1.0j * poly_mim
qext, qsca, qback, g = mp.mie(m,x)
absorb = (qext - qsca) * np.pi * r**2
scatt = qsca * np.pi * r**2
extinct = qext* np.pi * r**2
plt.plot(poly_lam*1000,absorb, label="$\sigma_{abs}$")
plt.plot(poly_lam*1000,scatt, label="$\sigma_{sca}$")
plt.plot(poly_lam*1000,extinct, "--", label="$\sigma_{ext}$")
#plt.text(350, 0.35,'$\sigma_{abs}$', color='blue', fontsize=14)
#plt.text(350, 0.54,'$\sigma_{sca}$', color='red', fontsize=14)
#plt.text(350, 0.84,'$\sigma_{ext}$', color='green', fontsize=14)
plt.xlabel("Wavelength (nm)")
plt.ylabel("Cross Section (1/microns$^2$)")
plt.title("Cross Sections for %.1f$\mu$m Polystyrebe Spheres" % (r*2))
plt.xlim(400,800)
plt.legend()
plt.show()
# +
x = 2*np.pi*r/poly_lam;
m = poly_mre - 1.0j * poly_mim
qext, qsca, qback, g = mp.mie(m,x)
qpr = qext - g*qsca
plt.plot(poly_lam*1000,qpr)
plt.xlabel("Wavelength (nm)")
plt.ylabel("Efficiency $Q_{pr}$")
plt.title("Radiation Pressure Efficiency for %.1f$\mu$m Polystyrene Spheres" % (r*2))
plt.show()
# -
r0 = 1.5e-6
Cpr = np.pi * r0 *r0 * qpr
E0 = 4.5e-3 / (np.pi * 1.75e-3 ** 2 )
c = 299792458 / 1.33
# ## Radiation Pressure
#
# The radiation pressure is given by [e.g., Kerker, p. 94]
#
# $$
# Q_{pr}=Q_{ext}-g Q_{sca}
# $$
#
# and is the momentum given to the scattering particle [van de Hulst, p. 13] in the direction of the incident wave. The radiation pressure cross section $C_{pr}$ is just the efficiency multiplied by the geometric cross section
#
# $$
# C_{pr} = \pi r^2 Q_{pr}
# $$
#
# The radiation pressure cross section $C_{pr}$ can be interpreted as the area of a black wall that would receive the same force from the same incident wave. The actual force on the particle is
# is
#
# $$
# F = E_0 \frac{C_{pr}}{c}
# $$
#
# where $E_0$ is the irradiance (W/m$^2$) on the sphere and $c$ is the velocity of the radiation in the medium
#
F = E0 * Cpr / c
plt.plot(poly_lam*1000,F*1e15)
plt.xlabel("Wavelength (nm)")
plt.ylabel("Force (fN)")
# +
Fs = []
rs = np.linspace(0.5, 3, 100)
I = np.argmin(abs(poly_lam*1000 - 532))
for r in rs:
x = 2*np.pi*r/poly_lam;
m = poly_mre - 1.0j * poly_mim
qext, qsca, qback, g = mp.mie(m,x)
qpr = qext - g*qsca
r0 = r * 1e-6
Cpr = np.pi * r0 *r0 * qpr
E0 = 4.5e-3 / (np.pi * 1.75e-3 ** 2 )
c = 299792458 / 1.33
F = E0 * Cpr / c
Fs.append(F[I]*1e15)
# +
plt.plot(rs, Fs)
#plt.title("Optical force for a 532 $\mu$m plane wave on Polystyrene Spheres")
plt.xlabel("$a$ ($\mu$m)")
plt.ylabel("$F_\mathrm{opt}$ (fN)")
# -
| 02_body/chapter2/images/Calcul_force_optique/.ipynb_checkpoints/Calcul optical force-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 72} id="0uUeDqA32K9o" outputId="27b66765-ee49-4504-f32e-f34776c4f3b4"
import urllib
from IPython.display import Markdown as md
### change to reflect your notebook
_nb_loc = "04_problem_types/10f_image_captioning.ipynb"
_nb_title = "Image Captions"
_nb_message = """
This notebook shows you how to train an ML model to generate captions for images. The training dataset is the COCO large-scale object detection, segmentation, and captioning dataset.
"""
### no need to change any of this
_icons=["https://raw.githubusercontent.com/GoogleCloudPlatform/practical-ml-vision-book/master/logo-cloud.png", "https://www.tensorflow.org/images/colab_logo_32px.png", "https://www.tensorflow.org/images/GitHub-Mark-32px.png", "https://www.tensorflow.org/images/download_logo_32px.png"]
_links=["https://console.cloud.google.com/ai-platform/notebooks/deploy-notebook?" + urllib.parse.urlencode({"name": _nb_title, "download_url": "https://github.com/GoogleCloudPlatform/practical-ml-vision-book/raw/master/"+_nb_loc}), "https://colab.research.google.com/github/GoogleCloudPlatform/practical-ml-vision-book/blob/master/{0}".format(_nb_loc), "https://github.com/GoogleCloudPlatform/practical-ml-vision-book/blob/master/{0}".format(_nb_loc), "https://raw.githubusercontent.com/GoogleCloudPlatform/practical-ml-vision-book/master/{0}".format(_nb_loc)]
md("""<table class="tfo-notebook-buttons" align="left"><td><a target="_blank" href="{0}"><img src="{4}"/>Run in AI Platform Notebook</a></td><td><a target="_blank" href="{1}"><img src="{5}" />Run in Google Colab</a></td><td><a target="_blank" href="{2}"><img src="{6}" />View source on GitHub</a></td><td><a href="{3}"><img src="{7}" />Download notebook</a></td></table><br/><br/><h1>{8}</h1>{9}""".format(_links[0], _links[1], _links[2], _links[3], _icons[0], _icons[1], _icons[2], _icons[3], _nb_title, _nb_message))
# + [markdown] id="fa3LRWv9BNk9"
# ## Enable GPU
# This notebook and pretty much every other notebook in this repository will run faster if you are using a GPU.
#
# On Colab:
# * Navigate to Edit→Notebook Settings
# * Select GPU from the Hardware Accelerator drop-down
#
# On Cloud AI Platform Notebooks:
# * Navigate to https://console.cloud.google.com/ai-platform/notebooks
# * Create an instance with a GPU or select your instance and add a GPU
#
# Next, we'll confirm that we can connect to the GPU with tensorflow:
# + colab={"base_uri": "https://localhost:8080/"} id="pL9G21yy2K2Q" outputId="27f2eeab-a187-4a90-cf67-4ae594e8e9c0"
# Not needed in Colab
# %pip install --quiet tfds-nightly # In Nov 2020, coco_captions is available only in the nightly build
# %pip uninstall -y h5py
# %pip install --quiet 'h5py < 3.0.0' # https://github.com/tensorflow/tensorflow/issues/44467
# + colab={"base_uri": "https://localhost:8080/"} id="G20XL3WTBXhj" outputId="a928c30a-f6d6-4120-9da1-f4f5753b9821"
import tensorflow as tf
print(tf.version.VERSION)
device_name = tf.test.gpu_device_name()
if device_name != '/device:GPU:0':
raise SystemError('GPU device not found')
print('Found GPU at: {}'.format(device_name))
# + [markdown] id="zr6sJGEIBe7D"
# ## Read and visualize dataset
#
# We will use the TensorFlow datasets capability to read the [COCO captions](https://www.tensorflow.org/datasets/catalog/coco_captions) dataset.
# This version contains images, bounding boxes, labels, and captions from COCO 2014, split into the subsets defined by Karpathy and Li (2015) and takes
# care of some data quality issues with the original dataset (for example, some
# of the images in the original dataset did not have captions)
#
# **Note**: This dataset is too large to store in an ephemeral location.
# Therefore, I'm storing the data in the GCS bucket corresponding to this book.
# If you access it from a Notebook outside the US, it will be (a) slow and
# (b) subject to a network charge.
# + id="U2WQtNeGBbMD"
GCS_DIR="gs://practical-ml-vision-book/tdfs_cache"
# Change these to control the accuracy/speed
VOCAB_SIZE = 5000 # use fewer words to speed up convergence
ATTN_UNITS = 512 # size of dense layer in Attention; larger more fine-grained
EPOCHS = 20 # train longer for greater accuracy
BATCH_SIZE = 64 # larger batch sizes lead to smoother convergence, but need more memory
EARLY_STOP_THRESH = 0.0001 # stop once loss improvement is less than this value
EMBED_DIM = 256 # embedding dimension for both images and words
# This is what Inception was trained with, so don't change unless you
# use a different pre-trained model. Inception takes (299, 299, 3) as
# input and provides (64, 2048) as output
IMG_HEIGHT = 299
IMG_WIDTH = 299
IMG_CHANNELS = 3
FEATURES_SHAPE = 2048
ATTN_FEATURES_SHAPE = 64
# + id="8JxC6DhwAcw5"
import matplotlib.pylab as plt
import numpy as np
import tensorflow as tf
import tensorflow_datasets as tfds
def filter_for_crowds(example):
return (tf.math.count_nonzero(example['objects']['is_crowd']) > 0)
def get_image_label(example):
captions = example['captions']['text'] # all the captions
img_id = example['image/id']
img = example['image']
img = tf.image.resize(img, (IMG_HEIGHT, IMG_WIDTH)) # inception size
img = tf.keras.applications.inception_v3.preprocess_input(img)
return {
'image_tensor': img,
'image_id': img_id,
'captions': captions
}
trainds = tfds.load('coco_captions',
split='train',
shuffle_files=False,
data_dir=GCS_DIR)
# reduce number of images in one of these ways
trainds = trainds.filter(filter_for_crowds)
#trainds = trainds.take(10000)
trainds = trainds.map(get_image_label)
# + colab={"base_uri": "https://localhost:8080/", "height": 389} id="KGz2bQaKV3iI" outputId="4bd05dee-e0f5-4934-ac0d-2afa0a20bad8"
f, ax = plt.subplots(1, 4, figsize=(20, 5))
for idx, data in enumerate(trainds.take(4)):
ax[idx].imshow(data['image_tensor'].numpy())
ax[idx].set_title('image_id={}'.format(data['image_id'].numpy()))
ax[idx].set_xlabel(data['captions'].numpy()[0].decode('utf-8')[:30] + str("..."))
# + [markdown] id="y4dyKHB2W4vZ"
# ## Tokenize captions
#
# Add a start and end token to each caption.
# Then send to the Keras tokenizer which will lowercase the captions
# and remove punctuation etc. It will also retain only the most frequent
# words.
# + colab={"base_uri": "https://localhost:8080/"} id="rSdiDPgbLpG_" outputId="f3603c44-17eb-46f9-c6f1-05f362cec614"
# get all the captions to feed into the Tokenizer
STOPWORDS = {'ourselves', 'hers', 'between', 'yourself', 'but', 'again', 'there', 'about', 'once', 'during', 'out', 'very', 'having', 'with', 'they', 'own', 'an', 'be', 'some', 'for', 'do', 'its', 'yours', 'such', 'into', 'of', 'most', 'itself', 'other', 'off', 'is', 's', 'am', 'or', 'who', 'as', 'from', 'him', 'each', 'the', 'themselves', 'until', 'below', 'are', 'we', 'these', 'your', 'his', 'through', 'don', 'nor', 'me', 'were', 'her', 'more', 'himself', 'this', 'down', 'should', 'our', 'their', 'while', 'above', 'both', 'up', 'to', 'ours', 'had', 'she', 'all', 'no', 'when', 'at', 'any', 'before', 'them', 'same', 'and', 'been', 'have', 'in', 'will', 'on', 'does', 'yourselves', 'then', 'that', 'because', 'what', 'over', 'why', 'so', 'can', 'did', 'not', 'now', 'under', 'he', 'you', 'herself', 'has', 'just', 'where', 'too', 'only', 'myself', 'which', 'those', 'i', 'after', 'few', 'whom', 't', 'being', 'if', 'theirs', 'my', 'against', 'a', 'by', 'doing', 'it', 'how', 'further', 'was', 'here', 'than'}
def preprocess_caption(c):
caption = "<start> {} <end>".format(c.decode('utf-8'))
words = [word for word in caption.lower().split()
if word not in STOPWORDS]
return (' '.join(words))
train_captions = []
for data in trainds:
str_captions = [preprocess_caption(c) for c in data['captions'].numpy()]
train_captions.extend(str_captions)
print(train_captions[:5])
# + colab={"base_uri": "https://localhost:8080/"} id="z2FW4ob5NikW" outputId="60f31007-6f03-4136-886a-0f9447b0c12b"
# Choose the most frequent words from the vocabulary
tokenizer = tf.keras.preprocessing.text.Tokenizer(num_words=VOCAB_SIZE,
oov_token="<unk>",
filters='!"#$%&()*+.,-/:;=?@[\]^_`{|}~ ')
tokenizer.fit_on_texts(train_captions)
train_seqs = tokenizer.texts_to_sequences(train_captions)
tokenizer.word_index['<pad>'] = 0
tokenizer.index_word[0] = '<pad>'
# pads each vector to the max_length of the captions
cap_vector = tf.keras.preprocessing.sequence.pad_sequences(train_seqs, padding='post')
max_caption_length = len(cap_vector[0])
print("max_caption_length={}".format(max_caption_length))
print(cap_vector[0])
print([tokenizer.index_word[idx] for idx in cap_vector[0]])
# + colab={"base_uri": "https://localhost:8080/"} id="X0LHFYjhBo32" outputId="647be30d-0357-49a2-a49b-010b59696ed0"
def create_batched_ds(trainds, batchsize):
# generator that does tokenization, padding on the caption strings
# and yields img, caption
def generate_image_captions():
for data in trainds:
captions = data['captions']
img_tensor = data['image_tensor']
str_captions = [preprocess_caption(c) for c in data['captions'].numpy()]
seqs = tokenizer.texts_to_sequences(str_captions)
# Pad each vector to the max_length of the captions
padded = tf.keras.preprocessing.sequence.pad_sequences(
seqs, padding='post', maxlen=max_caption_length)
for caption in padded:
yield img_tensor, caption # repeat image
return tf.data.Dataset.from_generator(
generate_image_captions,
(tf.float32, tf.int32)).batch(batchsize)
for img, caption in create_batched_ds(trainds, 193).take(2):
print(img.shape, caption.shape)
print(caption[0])
# + [markdown] id="1ehA-gDDYh47"
# ## Create Captioning Model
#
# It consists of an image encoder, followed by a caption decoder.
# The caption decoder incorporates an attention mechanism that
# focuses on different parts of the input image.
# + id="inFxAZKi9RqE"
class ImageEncoder(tf.keras.Model):
def __init__(self, embedding_dim):
super(ImageEncoder, self).__init__()
inception = tf.keras.applications.InceptionV3(
include_top=False,
weights='imagenet'
)
self.model = tf.keras.Model(inception.input,
inception.layers[-1].output)
self.fc = tf.keras.layers.Dense(embedding_dim)
def call(self, x):
x = self.model(x)
x = tf.reshape(x, (x.shape[0], -1, x.shape[3]))
x = self.fc(x)
x = tf.nn.relu(x)
return x
class BahdanauAttention(tf.keras.Model):
def __init__(self, units):
super(BahdanauAttention, self).__init__()
self.W1 = tf.keras.layers.Dense(units)
self.W2 = tf.keras.layers.Dense(units)
self.V = tf.keras.layers.Dense(1)
def call(self, features, hidden):
# features(CNN_encoder output) shape == (batch_size, 64, embedding_dim)
# hidden shape == (batch_size, hidden_size)
# hidden_with_time_axis shape == (batch_size, 1, hidden_size)
hidden_with_time_axis = tf.expand_dims(hidden, 1)
# attention_hidden_layer shape == (batch_size, 64, units)
attention_hidden_layer = (tf.nn.tanh(self.W1(features) +
self.W2(hidden_with_time_axis)))
# score shape == (batch_size, 64, 1)
# This gives you an unnormalized score for each image feature.
score = self.V(attention_hidden_layer)
# attention_weights shape == (batch_size, 64, 1)
attention_weights = tf.nn.softmax(score, axis=1)
# context_vector shape after sum == (batch_size, hidden_size)
context_vector = attention_weights * features
context_vector = tf.reduce_sum(context_vector, axis=1)
return context_vector, attention_weights
class CaptionDecoder(tf.keras.Model):
def __init__(self, embedding_dim, units, vocab_size):
super(CaptionDecoder, self).__init__()
self.units = units
self.embedding = tf.keras.layers.Embedding(vocab_size, embedding_dim)
self.gru = tf.keras.layers.GRU(self.units,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
self.fc1 = tf.keras.layers.Dense(self.units)
self.fc2 = tf.keras.layers.Dense(vocab_size)
self.attention = BahdanauAttention(self.units)
def call(self, x, features, hidden):
# defining attention as a separate model
context_vector, attention_weights = self.attention(features, hidden)
# x shape after passing through embedding == (batch_size, 1, embedding_dim)
x = self.embedding(x)
# x shape after concatenation == (batch_size, 1, embedding_dim + hidden_size)
x = tf.concat([tf.expand_dims(context_vector, 1), x], axis=-1)
# passing the concatenated vector to the GRU
output, state = self.gru(x)
# shape == (batch_size, max_length, hidden_size)
x = self.fc1(output)
# x shape == (batch_size * max_length, hidden_size)
x = tf.reshape(x, (-1, x.shape[2]))
# output shape == (batch_size * max_length, vocab)
x = self.fc2(x)
return x, state, attention_weights
def reset_state(self, batch_size):
return tf.zeros((batch_size, self.units))
encoder = ImageEncoder(EMBED_DIM)
decoder = CaptionDecoder(EMBED_DIM, ATTN_UNITS, VOCAB_SIZE)
optimizer = tf.keras.optimizers.Adam()
loss_object = tf.keras.losses.SparseCategoricalCrossentropy(
from_logits=True, reduction='none')
def loss_function(real, pred):
mask = tf.math.logical_not(tf.math.equal(real, 0))
loss_ = loss_object(real, pred)
mask = tf.cast(mask, dtype=loss_.dtype)
loss_ *= mask
return tf.reduce_mean(loss_)
# + [markdown] id="pGVl8cQpZ5Qu"
# ## Training loop
#
# Here, we use a custom training loop because we have to add on to the decoder
# input (dec_input) one word at a time.
# + id="PADm7AHuHw9I"
loss_plot = []
@tf.function
def train_step(img_tensor, target):
loss = 0
# initializing the hidden state for each batch
# because the captions are not related from image to image
hidden = decoder.reset_state(batch_size=target.shape[0])
dec_input = tf.expand_dims([tokenizer.word_index['<start>']] * target.shape[0], 1)
with tf.GradientTape() as tape:
features = encoder(img_tensor)
for i in range(1, target.shape[1]):
# passing the features through the decoder
predictions, hidden, _ = decoder(dec_input, features, hidden)
loss += loss_function(target[:, i], predictions)
# using teacher forcing
dec_input = tf.expand_dims(target[:, i], 1)
total_loss = (loss / int(target.shape[1]))
trainable_variables = encoder.trainable_variables + decoder.trainable_variables
gradients = tape.gradient(loss, trainable_variables)
optimizer.apply_gradients(zip(gradients, trainable_variables))
return loss, total_loss
checkpoint_path = "./checkpoints/train"
ckpt = tf.train.Checkpoint(encoder=encoder,
decoder=decoder,
optimizer = optimizer)
ckpt_manager = tf.train.CheckpointManager(ckpt, checkpoint_path, max_to_keep=5)
# + colab={"base_uri": "https://localhost:8080/"} id="6TW8wuySH1nd" outputId="c33e4f3d-65b0-49ba-b1e7-06e1569d610f"
import time
batched_ds = create_batched_ds(trainds, BATCH_SIZE)
prev_loss = 999
for epoch in range(EPOCHS):
start = time.time()
total_loss = 0
num_steps = 0
for batch, (img_tensor, target) in enumerate(batched_ds):
batch_loss, t_loss = train_step(img_tensor, target)
total_loss += t_loss
num_steps += 1
if batch % 100 == 0:
print ('Epoch {} Batch {} Loss {:.4f}'.format(
epoch + 1, batch, batch_loss.numpy() / int(target.shape[1])))
current_loss = total_loss / num_steps
# storing the epoch end loss value to plot later
loss_plot.append(current_loss)
ckpt_manager.save()
print ('Epoch {} Loss {:.6f} Time taken {:.1f} sec'.format(
epoch + 1,
current_loss,
time.time() - start))
# stop once it has converged
improvement = prev_loss - current_loss
if improvement < EARLY_STOP_THRESH:
print("Stopping because improvement={} < {}".format(improvement, EARLY_STOP_THRESH))
break
prev_loss = current_loss
# + id="uzpGKbigXmKs"
plt.plot(loss_plot);
# + [markdown] id="otiuFI4ZaK6w"
# ## Prediction
#
# To predict, we generate the caption one word at a time, feeding the
# decoder with the previous predictions.
# -
for id in range(10):
print(id, tokenizer.index_word[id])
# + id="EkmKr8nxMNyG"
## Probabilistic prediction using the trained model
def plot_attention(image, result, attention_plot):
fig = plt.figure(figsize=(10, 10))
len_result = len(result)
num_panels = len_result//2
if num_panels*2 < len_result:
num_panels += 1
for l in range(len_result):
temp_att = np.resize(attention_plot[l], (8, 8))
ax = fig.add_subplot(len_result//2, num_panels, l+1)
ax.set_title(result[l])
img = ax.imshow(image)
ax.imshow(temp_att, cmap='gray', alpha=0.6, extent=img.get_extent())
plt.tight_layout()
plt.show()
def predict_caption(filename):
attention_plot = np.zeros((max_caption_length, ATTN_FEATURES_SHAPE))
hidden = decoder.reset_state(batch_size=1)
img = tf.image.decode_jpeg(tf.io.read_file(filename), channels=IMG_CHANNELS)
img = tf.image.resize(img, (IMG_HEIGHT, IMG_WIDTH)) # inception size
img_tensor_val = tf.keras.applications.inception_v3.preprocess_input(img)
features = encoder(tf.expand_dims(img_tensor_val, axis=0))
dec_input = tf.expand_dims([tokenizer.word_index['<start>']], 0)
result = []
previous_word_ids = []
for i in range(max_caption_length):
predictions, hidden, attention_weights = decoder(dec_input, features, hidden)
attention_plot[i] = tf.reshape(attention_weights, (-1, )).numpy()
if i < 10:
# mask out <pad> <unk> <start> <end>, since we don't want <end>
masked_predictions = predictions[0]
mask = [0.0, 0, 0, 0] + [1.0] * (masked_predictions.shape[0] - 4)
# end is okay after 4th word
if i > 3:
mask[3] = 1
# avoid repeating words
for p in previous_word_ids:
mask[p] = 0
mask = tf.convert_to_tensor(mask)
masked_predictions *= mask
# keep only the top (i+2) words, and draw out of log distribution
top_probs, top_idxs = tf.math.top_k(input=masked_predictions, k=(i+2), sorted=False)
chosen_id = tf.random.categorical([top_probs], 1)[0].numpy()
predicted_id = top_idxs.numpy()[chosen_id][0]
else:
# draws from log distribution given by predictions
predicted_id = tf.random.categorical(predictions, 1)[0][0].numpy()
result.append(tokenizer.index_word[predicted_id])
previous_word_ids.append(predicted_id)
if tokenizer.index_word[predicted_id] == '<end>':
return img, result, attention_plot
dec_input = tf.expand_dims([predicted_id], 0)
attention_plot = attention_plot[:len(result), :]
return img, result, attention_plot
# image from https://commons.wikimedia.org/wiki/File:Flying_Kites_At_Sunset.jpg
filename = "gs://practical-ml-vision-book/images/800px-Flying_Kites_At_Sunset.jpg"
image, caption, attention_plot = predict_caption(filename)
print(caption)
plot_attention(image, caption, attention_plot)
# -
# The model has managed to capture the key aspects of the image:
# <pre>
# group of people standing in a field.
# </pre>
# However, the model hasn't quite figured out the attention (note the attention is similar throughout).
# + [markdown] id="7KbY7guiqFiB"
# ## Plots for book
# + id="gGgCq-nkZvyV"
print(encoder.summary())
# + id="aS6wcOlkZyIn"
print(decoder.summary())
# + [markdown] id="l_fNzWuY2UoB"
# Copyright 2020 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| 10_adv_problems/10f_image_captioning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
#
# <a href='http://www.pieriandata.com'><img src='../Pierian_Data_Logo.png'/></a>
# ___
# <center><em>Copyright by Pierian Data Inc.</em></center>
# <center><em>For more information, visit us at <a href='http://www.pieriandata.com'>www.pieriandata.com</a></em></center>
# # Q Learning Exercise
# **We'll be reviewing and testing your skills with Q-Learning on a continuous space! Please feel free to reference the lecture notebooks you are definitely not expected to be able to fill out all this code from memory, just the ability to understand the core concepts and apply it to a different situation.**
#
# --------------------
#
# ## Complete the tasks in bold below.
# In this exercise we take a look at the MountainCar-v0 (https://gym.openai.com/envs/MountainCar-v0/) game again. That is the game from our original discussion of OpenAi gym environments for which we created an agent manually.
# Remember, that the goal is to reach the top of the mountain within some time limit
#
# -----
# **TASK: Import any relevant libraries you think you might need.** <br />
#
# **TASK: Create the gym mountain car environment** <br />
#
# **TASK: Write a function to create a numpy array holding the bins for the observations of the car (position and velocity).** <br />
# Feel free to explore different bins per observation spacings.
# The function should take one argument which acts as the bins per observation <br />
# Hint: You can find the observations here: https://github.com/openai/gym/blob/master/gym/envs/classic_control/mountain_car.py
# <br /> Hint: You will probably need around 25 bins for good results, but feel free to use less to reduce training time. <br />
#
def create_bins(num_bins_per_observation):
# CODE HERE
return bins
# **TASK: Here you should write the code which creates the bins and defines the NUM_BINS attribute**
# **TASK: Create a function that will take in observations from the environment and the bins array and return the discretized version of the observation.**
# Now we need the code to discretize the observations. We can use the same code as used in the last notebook
def discretize_observation(observations, bins):
# CODE HERE
return tuple(binned_observations) # Important for later indexing
# **Let's check to make sure your previous two function calls work with a quick task! Otherwise it may be hard to debug later on.**
# **TASK: Confirm that your *create_bins()* function works with *discretize_observation() by running the following cell***
# +
test_bins = create_bins(5)
np.testing.assert_almost_equal(test_bins[0], [-1.2 , -0.75, -0.3 , 0.15, 0.6])
np.testing.assert_almost_equal(test_bins[1], [-0.07 , -0.035, 0. , 0.035, 0.07 ])
test_observation = np.array([-0.9, 0.03])
discretized_test_bins = discretize_observation(test_observation, test_bins)
assert discretized_test_bins == (1, 3)
# -
# **TASK: Create the Q-Table** <br />
# Remember the shape that the Q-Table needs to have.
# +
# CREATE THE Q TABLE
# -
# **TASK: Fill out the Epislon Greedy Action Selection function:**
def epsilon_greedy_action_selection(epsilon, q_table, discrete_state):
#CODE HERE
return action
# **TASK: Fill out the function to compute the next Q value.**
def compute_next_q_value(old_q_value, reward, next_optimal_q_value):
return # CODE HERE
# **TASK: Create a function to reduce epsilon, feel free to choose any reduction method you want. We'll use a reduction with BURN_IN and EPSILON_END limits in the solution. We'll also show a way to reduce epsilon based on the number of epochs. Feel free to experiment here.**
def reduce_epsilon(epsilon, epoch):
# CODE HERE
return epsilon
# **TASK: Define your hyperparameters. Note, we'll show our solution hyperparameters here, but depending on your *reduce_epsilon* function, your epsilon hyperparameters may be different.**
# Here are the solution initial hyperparameters, your will probably be different!
# +
# Feel free to change!
EPOCHS = 30000
BURN_IN = 100
epsilon = 1
EPSILON_END= 10000
EPSILON_REDUCE = 0.0001
ALPHA = 0.8
GAMMA = 0.9
# -
# **TASK: Create the training loop for the reinforcement learning agent and run the loop. We've gone ahead and created the basic structure of the loopwith some comments. We also pre-filled the visualization portion.** <br />
# Note: Use the lecture notebook as a guide and reference
# +
####### VISUALIZATION CODE FOR YOU. TOTALLY OPTIONAL. ##########################
########## FEEL FREE TO REMOVE OR ADD YOUR OWN VISUAL CODE. #################
log_interval = 100 # How often do we update the plot? (Just for performance reasons)
### Here we set up the routine for the live plotting of the achieved points ######
fig = plt.figure()
ax = fig.add_subplot(111)
plt.ion()
fig.canvas.draw()
max_position_log = [] # to store all achieved points
mean_positions_log = [] # to store a running mean of the last 30 results
epochs = [] # store the epoch for plotting
#############################################################################
################################## TRAINING TASKS ##########################
###########################################################################
for epoch in range(EPOCHS):
################################# TODO ######################################
# TODO: Get initial observation and discretize them. Set done to False
#########################################
#
#
# CODE NEEDED HERE!!!
#
#
##########################################
#############################
# These lines are for plotting.
max_position = -np.inf
epochs.append(epoch)
#############################
# TASK TO DO: As long as current run is alive (i.e not done) perform the following steps:
while not done: # Perform current run as long as done is False (as long as there is still time to reach the top)
# TASK TO DO: Select action according to epsilon-greedy strategy
#########################################
#
#
# CODE NEEDED HERE!!!
#
#
##########################################
# TASK TO DO: Perform selected action and get next state. Do not forget to discretize it
#########################################
#
#
# CODE NEEDED HERE!!!
#
#
##########################################
# TASK TO DO: Get old Q-value from Q-Table and get next optimal Q-Value
#########################################
#
#
# CODE NEEDED HERE!!!
#
#
##########################################
# TASK TO DO: Compute next Q-Value and insert it into the table
#########################################
#
#
# CODE NEEDED HERE!!!
#
#
##########################################
# TASK TO DO: Update the old state with the new one
#########################################
#
#
# CODE NEEDED HERE!!!
#
#
##########################################
##############################
## Only for plotting the results - store the highest point the car is able to reach
if position > max_position:
max_position = position
# TASK TO DO: Reduce epsilon
#########################################
#
#
# CODE NEEDED HERE!!!
#
#
##########################################
##############################################################################
max_position_log.append(max_position) # log the highest position the car was able to reach
running_mean = round(np.mean(max_position_log[-30:]), 2) # Compute running mean of position over the last 30 epochs
mean_positions_log.append(running_mean) # and log it
################ Plot the points and running mean ##################
if epoch % log_interval == 0:
ax.clear()
ax.scatter(epochs, max_position_log)
ax.plot(epochs, max_position_log)
ax.plot(epochs, mean_positions_log, label=f"Running Mean: {running_mean}")
plt.legend()
fig.canvas.draw()
######################################################################
env.close()
# -
# **TASK: Use your Q-Table to test your agent and render its performance.**
# +
# CODE HERE
# -
# **OPTIONAL: Play with our Q-Table with 40 bins per observation.**
# +
# We saved our matrix q_table for you (with 40 bins)
# -
our_q_table = np.load('40bin_qtable_mountaincar.npy')
our_q_table.shape
# **Great job! Note how you could train for many more epochs/episodes or edit hyperparameters, the more complex the environment, the more choices you have to experiment with!**
| practical_ai/archive/06-Classical-Q-Learning/02-Q-Learning-Exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cdiscount kernel
# language: python
# name: cdiscount
# ---
# +
# Load the "autoreload" extension
# %load_ext autoreload
# always reload modules marked with "%aimport"
# %autoreload 1
import os
import sys
from sklearn.metrics import roc_curve
# add the 'src' directory as one where we can import modules
src_dir = os.path.join(os.getcwd(), os.pardir, 'src')
sys.path.append(src_dir)
# import my method from the source code
# %aimport data.read_data
# %aimport models.train_model
# %aimport features.build_features
# %aimport visualization.visualize
from data.read_data import read_data, get_stopwords
from models.train_model import split_train, score_function, get_fasttext, model_ridge, model_xgb, model_lightgbm
from features.build_features import get_vec, to_categorical, replace_na, to_tfidf, stack_sparse, to_sparse_int
from visualization.visualize import plot_roc, plot_scatter
# -
train = read_data(test=False)
y = train['Target']
stopwords = get_stopwords()
train.head()
# # Feature engineering
train = replace_na(train, ['review_content', 'review_title'])
X_dummies = to_categorical(train, 'review_stars')
X_content = to_tfidf(train, 'review_content', stopwords)
X_title = to_tfidf(train, 'review_title', stopwords)
X_length = to_sparse_int(train, 'review_content')
sparse_merge = stack_sparse([X_dummies, X_content, X_title, X_length])
X_train, X_test, y_train, y_test = split_train(sparse_merge, y, 0.2)
# # LightGBM
model_lgb = model_lightgbm(X_train, y_train)
preds = model_lgb.predict_proba(X_test)
preds1 = preds[:,1]
score_function(y_test, preds1)
fpr, tpr, _ = roc_curve(y_test, preds1)
plot_roc(fpr, tpr)
# # Ridge
model_rdg = model_ridge(X_train, y_train, )
preds = model_rdg.predict(X=X_test)
score_function(y_test, preds)
fpr, tpr, _ = roc_curve(y_test, preds)
plot_roc(fpr, tpr)
# # Xgboost
model_xgboost = model_xgb(X_train, y_train)
preds = model_xgboost.predict_proba(X_test)
preds1 = preds[:,1]
score_function(y_test, preds1)
fpr, tpr, _ = roc_curve(y_test, preds1)
plot_roc(fpr, tpr)
| notebooks/02-Train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 11, "hidden": false, "row": 0, "width": 12}, "report_default": {"hidden": false}}}}
# This is the <a href="https://jupyter.org/">Jupyter Notebook</a>, an interactive coding and computation environment. For this lab, you do not have to write any code, you will only be running it.
#
# To use the notebook:
# - "Shift + Enter" runs the code within the cell (so does the forward arrow button near the top of the document)
# - You can alter variables and re-run cells
# - If you want to start with a clean slate, restart the Kernel either by going to the top, clicking on Kernel: Restart, or by "esc + 00" (if you do this, you will need to re-run the following block of code before running any other cells in the notebook)
# -
from gpgLabs.Mag.MagDipoleApp import MagneticDipoleApp
# # Magnetic Prism Applet
#
#
# + [markdown] extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 3, "hidden": true, "row": 11, "width": 12}, "report_default": {"hidden": true}}}}
# ## Purpose
#
# From the Magnetic Dipole applet, we have learned how anomalous magnetic field observed at ground's surface look
# The objective is to learn about the magnetic field observed at the ground's surface, caused by a retangular susceptible prism.
#
#
# ## What is shown
#
# - <b>The colour map</b> shows the strength of the chosen parameter (Bt, Bx, By, or Bz) as a function of position.
#
# - Imagine doing a two dimensional survey over a susceptible sphere that has been magentized by the Earth's magnetic field specified by inclination and declination. "Measurement" location is the centre of each coloured box. This is a simple (but easily programmable) alternative to generating a smooth contour map.
#
# - The anomaly depends upon magnetic latitude, direction of the inducing (Earth's) field, the depth of the buried dipole, and the magnetic moment of the buried dipole.
#
#
# ## Important Notes:
#
# - <b>Inclination (I)</b> and <b>declination (D)</b> describe the orientation of the Earth's ambient field at the centre of the survey area. Positive inclination implies you are in the northern hemisphere, and positive declination implies that magnetic north is to the east of geographic north.
#
# - The <b>"length"</b> adjuster changes the size of the square survey area. The default of 72 means the survey square is 72 metres on a side.
#
# - The <b>"data spacing"</b> adjuster changes the distance between measurements. The default of 1 means measurements were acquired over the survey square on a 2-metre grid. In other words, "data spacing = 2" means each coloured box is 2 m square.
#
# - The <b>"depth"</b> adjuster changes the depth (in metres) to the top of the buried prism.
#
# - The <b>"magnetic moment (M)"</b> adjuster changes the strength of the induced field. Units are Am2. This is related to the strength of the inducing field, the susceptibility of the buried sphere, and the volume of susceptible material.
# - <b>Bt, Bx, By, Bz</b> are Total field, X-component (positive northwards), Y-component (positive eastwards), and Z-component (positive down) of the anomaly field respectively.
#
# - Checking the <b>fixed scale</b> button fixes the colour scale so that the end points of the colour scale are minimum and maximum values for the current data set.
#
# - You can generate a <b>profile</b> along either "East" or "North" direction
#
# - Check <b>half width</b> to see the half width of the anomaly. Anomaly width is noted on the botton of the graph.
#
# - Measurements are taken 1m above the surface.
# + [markdown] extensions={"jupyter_dashboards": {"version": 1, "views": {"grid_default": {"col": 0, "height": 11, "hidden": false, "row": 11, "width": 6}, "report_default": {"hidden": false}}}}
# ## Define a 3D prism
# Compared to the MagneticDipoleApplet, there are additional parameters to define a prism.
#
# - $\triangle x$: length in North (X) direction (m)
# - $\triangle y$: length in East (Y) direction (m)
# - $\triangle z$: length in Depth (z) direction (m) below the receiver
# - depth: top boundary of the prism (meter)
# - I$_{prism}$: inclination of the prism (reference is north direction)
# - D$_{prism}$: declination of the prism (reference is north direction)
# -
mag = MagneticDipoleApp()
mag.interact_plot_model_prism()
| Notebooks/MagneticPrismApplet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from keras.applications.inception_v3 import InceptionV3
from keras.applications.inception_v3 import preprocess_input
from keras.preprocessing import image
from keras.models import Model
from keras.layers import Dense, GlobalAveragePooling2D
import cv2
# +
base_model = InceptionV3(weights='imagenet', include_top=False)#, input_shape=(299, 299, 3))
x = base_model.output
x = GlobalAveragePooling2D()(x)
x = Dense(1024, activation='relu')(x)
out = Dense(2, activation='softmax')(x)
model = Model(inputs=base_model.input, outputs=out)
for layer in base_model.layers:
layer.trainable = False
model.compile(optimizer='adam', loss='categorical_crossentropy')
# -
model.summary()
# +
from keras.preprocessing.image import ImageDataGenerator
def normalize(image):
# image2 = image / 255.
# image2 = image2 - 0.5
# image2 = image2 * 2.
return image
train_datagen = ImageDataGenerator(
rescale=1/255.,
# preprocessing_function=normalize,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True)
train_generator = train_datagen.flow_from_directory(
'flowers/train',
target_size=(299, 299),
batch_size=32,
class_mode='categorical')
#test_datagen = ImageDataGenerator(rescale=1./255)
#validation_generator = test_datagen.flow_from_directory(
# 'data/validation',
# target_size=(150, 150),
# batch_size=32,
# class_mode='binary')
# -
model.fit_generator(train_generator, steps_per_epoch=32, epochs=2)
image = cv2.imread('flowers/train/Rose/test - 4.jpg')
image = cv2.resize(image, (299, 299))
import numpy as np
preprocessed_image = normalize(image)
model.predict(np.expand_dims(preprocessed_image, axis=0))
| Transfer learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
# -
# # Загрузка датасета
# Загрузим датасет, настроим pandas на отображение всех колонок
# +
dataset = pd.read_csv("dataset.csv", sep=';', decimal=',', index_col=0)
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', 100)
# -
# Первые записи:
dataset.head()
dataset.shape
# Датасет содержит 7041 объект.
# Объект содержит 84 признака.
# Описание признаков:
dataset.describe(include='all')
dataset.columns
# ## Целевые признаки
# целевые признаки (принадлежат к вещественному типу)
target_features = ['химшлак последний Al2O3', 'химшлак последний CaO',
'химшлак последний FeO', 'химшлак последний MgO',
'химшлак последний MnO', 'химшлак последний R',
'химшлак последний SiO2']
# # Количество пропусков и типы данных
def make_description(df):
description = pd.concat([df.isna().sum(), df.dtypes, df.nunique()], axis=1)
description.rename(columns={0: "num of NaN", 1: "dtypes", 2: "nunique"}, inplace=True)
return description
make_description(dataset).sort_values('num of NaN', ascending=False)
# ## Пропуски по столбцам
# Как видно, некоторные признаки имеют значительное количество пропусков (более 33%)
# Также важно отметить, что некоторые из признаков содержат всего 1 уникальное значение помимо NaN.
# +
def list_features_with_single_value(df):
features_list = []
for feature in df.columns:
if df[feature].nunique() == 1:
features_list.append(feature)
return features_list
def list_features_with_lots_of_nan(df):
features_list = []
for feature in df.columns:
if feature in target_features: # не рассматриваем целевые признаки
continue
num_of_nan = dataset[feature].isna().sum()
if num_of_nan > dataset.shape[0] / 3:
features_list.append(feature)
return features_list
# +
features_with_lots_of_nan = list_features_with_lots_of_nan(dataset)
print("Столбцы с большим количеством пропусков:")
print(features_with_lots_of_nan)
features_with_single_value = list_features_with_single_value(dataset)
print("Признаки с единственным уникальным значением:")
print(features_with_single_value)
# -
# ## Пропуски по строкам
# Узнаем количество строк, в которых отсутствует более 33% признаков:
def list_rows_with_lots_of_nan(df):
rows_list = []
numbers_of_nan = df.isna().sum(axis=1)
for index, count in enumerate(numbers_of_nan):
if count > df.shape[1] / 3:
rows_list.append(index)
return rows_list
rows_with_lots_of_nan = list_rows_with_lots_of_nan(dataset)
print("Строк с большим количеством пропусков: {}".format(len(rows_with_lots_of_nan)))
# ## Категориальные признаки
#
# Категориальными типами являются:
# - **nplv** - номер плавки, не имеет смысла
# - **DT** - дата и время плавки
# - **МАРКА** - марка стали (?)
# - **ПРОФИЛЬ** - профиль стали (?)
# +
categorical_features = ["nplv", "DT", "МАРКА", "ПРОФИЛЬ"]
dataset[categorical_features].head()
# -
print("Количество всех пропущенных категориальных значений: {}".format(dataset[categorical_features].isna().sum().sum()))
# #### МАРКА
print("Количество уникальных значений: {}".format(dataset["МАРКА"].nunique()))
print("\nЧастота встречаемости каждого из значений:")
print(dataset["МАРКА"].value_counts())
# #### ПРОФИЛЬ
print("Количество уникальных значений: {}".format(dataset["ПРОФИЛЬ"].nunique()))
print("\nЧастота встречаемости каждого из значений:")
print(dataset["ПРОФИЛЬ"].value_counts())
# ## Вещественные признаки
#
# Все остальные 80 признаков имеют вещественный тип данных
numerical_features = ['t вып-обр', 't обработка',
't под током', 't продувка', 'ПСН гр.', 'чист расход C',
'чист расход Cr', 'чист расход Mn', 'чист расход Si', 'чист расход V',
'температура первая', 'температура последняя', 'Ar (интенс.)',
'N2 (интенс.)', 'эл. энергия (интенс.)', 'произв жидкая сталь',
'произв количество обработок', 'произв количество плавок',
'произв количество плавок (цел)', 'расход газ Ar', 'расход газ N2',
'расход C пров.', 'сыпуч известь РП', 'сыпуч кварцит',
'сыпуч кокс пыль УСТК', 'сыпуч кокс. мелочь (сух.)',
'сыпуч кокс. мелочь КМ1', 'сыпуч шпат плав.', 'ферспл CaC2',
'ферспл FeMo', 'ферспл FeSi-75', 'ферспл FeV азот.', 'ферспл FeV-80',
'ферспл Mn5Si65Al0.5', 'ферспл Ni H1 пласт.', 'ферспл SiMn18',
'ферспл ферванит', 'ферспл фх850А', 'эл. энергия',
'химсталь первый Al_1', 'химсталь первый C_1', 'химсталь первый Cr_1',
'химсталь первый Cu_1', 'химсталь первый Mn_1', 'химсталь первый Mo_1',
'химсталь первый N_1', 'химсталь первый Ni_1', 'химсталь первый P_1',
'химсталь первый S_1', 'химсталь первый Si_1', 'химсталь первый Ti_1',
'химсталь первый V_1', 'химсталь последний Al', 'химсталь последний C',
'химсталь последний Ca', 'химсталь последний Cr',
'химсталь последний Cu', 'химсталь последний Mn',
'химсталь последний Mo', 'химсталь последний N',
'химсталь последний Ni', 'химсталь последний P', 'химсталь последний S',
'химсталь последний Si', 'химсталь последний Ti',
'химсталь последний V', 'химшлак первый Al2O3_1',
'химшлак первый CaO_1', 'химшлак первый FeO_1', 'химшлак первый MgO_1',
'химшлак первый MnO_1', 'химшлак первый R_1', 'химшлак первый SiO2_1',
'химшлак последний Al2O3', 'химшлак последний CaO',
'химшлак последний FeO', 'химшлак последний MgO',
'химшлак последний MnO', 'химшлак последний R',
'химшлак последний SiO2']
dataset[numerical_features].shape
# ### Количество NaN значений у каждого вещественного признака
dataset[numerical_features].isna().sum()
# # Распределение целевых признаков
# Изначальное, до чистки:
for feature in target_features:
#sns.displot(dataset[feature])
print(feature)
print("Min: {}, max: {}".format(dataset[feature].min(), dataset[feature].max()))
print("Mean: {}, median: {}, mode: {}, std: {}".format(dataset[feature].mean(),
dataset[feature].median(),
dataset[feature].mode()[0],
dataset[feature].std()))
print("Number of NaN: {}".format(dataset[feature].isna().sum()))
print()
# После чистки:
# +
features_to_drop = list(set(features_with_lots_of_nan + features_with_single_value))
preprocessed_dataset = dataset.drop(features_to_drop, axis=1)
preprocessed_dataset = preprocessed_dataset.drop(rows_with_lots_of_nan, axis=0)
print("Размер датасета после первичной чистки: {}\n".format(preprocessed_dataset.shape))
for feature in target_features:
#sns.displot(dataset[feature])
print(feature)
print("Min: {}, max: {}".format(preprocessed_dataset[feature].min(), preprocessed_dataset[feature].max()))
print("Mean: {}, median: {}, mode: {}, std: {}".format(preprocessed_dataset[feature].mean(),
preprocessed_dataset[feature].median(),
preprocessed_dataset[feature].mode()[0],
preprocessed_dataset[feature].std()))
print("Number of NaN: {}".format(preprocessed_dataset[feature].isna().sum()))
print()
| Melekhin/0_Exploratory_Data_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Section 1. Business understanding
#
# The three business questions that I focused in this analysis are:
#
# Question 1. What is the differences in listing prices among different neighborhood?
#
# Question 2. How do listing prices change during the year?
#
# Question 3. How should you price your listing based on location and room type?
# # Section 2. Data Understanding
# ## Import Libraries and Data
# +
# For Analysis
import numpy as np
import pandas as pd
# For Visualizations
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
#Import machine learning
from sklearn.linear_model import LinearRegression
from sklearn.ensemble import RandomForestRegressor
from sklearn.model_selection import train_test_split #split
from sklearn.metrics import r2_score, mean_squared_error #metrics
# -
# ## Gather Data
# Import Data
listings_df = pd.read_csv('listings.csv')
calendar_df = pd.read_csv('calendar.csv')
reviews_df = pd.read_csv('reviews.csv') #didn't use in this exercise
# Start with exploring the data and data type for each dataset.
#
# ## Data info
listings_df.head()
reviews_df.head()
calendar_df.head()
listings_df.info()
reviews_df.info()
calendar_df.info()
# # Section 3. Data preparation
#
# From the information above, the data will need some data-preprocessing:
# 1. Find the missing values for each dataset.
# 2. We can merge the listings and calendar to get the complete daily listing list over the year.
# 3. Select relevant columns for the analysis
# 4. Convert price to float and convert datetime to month and year
# 5. Recode the zipcode to neighbourhood
# 6. Clean the missing values
# +
#find percentage of missing values for each column
listings_missing_df = listings_df.isnull().mean()*100
#filter out only columns, which have missing values
listings_columns_with_nan = listings_missing_df[listings_missing_df > 0]
#plot the results
plt.figure(figsize = (12, 12)) #determine the size of the chart
listings_columns_with_nan.plot.barh(title='Percentage of missing values in listings, %', color = "midnightblue")
plt.xticks(fontsize = 8) # format the labels for the x-axis
plt.yticks(fontsize = 10) # format the y-axis
plt.show()
# -
# Find missing values in calendar
calendar_df.isnull().mean().sort_values(ascending=False)#rank the most missing values
# FInd missing values in reviews
reviews_df.isnull().mean().sort_values(ascending=False)#rank the most missing values
# In the analysis of missing values, we can identify that there are over 80% of missing values in the columns 'square feet' and 'license'. Those two won't be useful for our analysis. Also, there are about 30% of missing values in the column 'price' in the calendar dataset. These need to be cleaned as well since 'price' will be our dependent variables for exploration analysis and future prediction models.
#
# Then let's take a look at the information for some of those columns to find the appropriate columns for our neighborhood comparison. I am specifically interested in the columns related to 'zipcode' and 'neighborhood'.
#
# From the information below, the neighborhood infomation contained in the dataset are not complete with a significant portion of rows labeled as others, which will not be useful for our analysis. But we do have a very complete record of the zipcode information for the listing. This can be more useful for following analysis.
listings_df.zipcode.value_counts()
listings_df.neighbourhood_group_cleansed.value_counts()
# ### Merge and select datasets
#
# Listing information and listing calendar information are merged to provide a full picture of the listing information throughout the year.
#merge datasets
listings_df = listings_df.rename(index=str, columns={"id": "listing_id"})
df_merged = pd.merge(calendar_df, listings_df, on = 'listing_id')
df_merged.columns
#select the relevant columns for analysis
df_selected = df_merged[['listing_id', 'date', 'price_x','zipcode','property_type','room_type', 'accommodates', 'bathrooms', 'bedrooms',
'beds', 'bed_type']]
df_selected.head()
# ### Clean data
#
# First, convert price to float and convert datetime to month and year so it is easier for future analysis. Then, all the zipcode information were recoded to neighbourhood based on the Seattle neighbourhood map.
# Lastly, drop all the missing values in the dataset.
#convert price_x to float
pd.options.mode.chained_assignment = None # default='warn'
df_selected['price_x'] = df_selected['price_x'].astype(str)
df_selected['price'] = df_selected['price_x'].str.replace('[$, ]','').astype('float')
df_selected = df_selected.drop(columns = ['price_x'])
# +
#convert date from the calendar into month and drop the date colum
def get_month_from_date(row):
''' Get month from date represented as a string '''
return int(row['date'].split('-')[1])
def get_year_from_date(row):
''' Get year from date represented as a string '''
return int(row['date'].split('-')[0])
df_selected['month'] = df_selected.apply(lambda row: get_month_from_date(row),axis=1)
df_selected['year'] = df_selected.apply(lambda row: get_year_from_date(row),axis=1)
df_selected = df_selected.drop(columns = ['date'])
#select data from 2016
df_selected_2016 = df_selected.iloc[:-2,:]
df_selected_2016
# +
# Recode zipcode to neighbourhood
def neighbourhood(value):
if value in ['98177', '98133', '98117', '98103', '98107']:
return 'Northwest Seattle'
elif value in ['98125','98105', '98115']:
return 'Northeast Seattle'
elif value in ['98199', '98119', '98109']:
return 'Magnolia&Queen Anne'
elif value in ['98122', '98112', '99\n98122','98102']:
return 'Central Seattle'
elif value in ['98121', '98101', '98104', '98134']:
return 'Downtown Seattle'
elif value in ['98144', '98108', '98118', '98178']:
return 'Southeast Seattle'
elif value in ['98116', '98136', '98126', '98106','98146']:
return 'West Seattle&Delridge'
return value
df_selected_2016['zipcode'] = df_selected_2016['zipcode'].apply(neighbourhood)
df_selected.rename(columns={'zipcode': 'neighbourhood'})
# +
#find percentage of missing values for each column
df_selected_missing = df_selected_2016.isnull().mean()*100
#filter out only columns, which have missing values
df_selected_with_nan = df_selected_missing[df_selected_missing > 0]
#plot the results
plt.figure(figsize = (12, 9)) #determine the size of the chart
df_selected_with_nan.plot.barh(title='Percentage of missing values in selected column, %', color = "midnightblue")
plt.xticks(fontsize = 8) # format the labels for the x-axis
plt.yticks(fontsize = 10) # format the y-axis
plt.show()
# -
# From the figure above, we can see that there are around 30% of missing values in the column 'price' and few missing values in other columns. Since 'price' will be our main variable for modeling and the missing values were most due to the fact that some of the listings were not listed in certain time of the year, we should drop all the missing values in the dataset, so we just keep the price information when the listings were listed on the market.
#drop the missing values
df_selected_2016_cleaned = df_selected_2016.dropna()
#check the percentage of the missing values dropped
(len(df_selected_2016)-len(df_selected_2016_cleaned))/len(df_selected_2016)*100
# # Data Evaluation
# ## Comparison of Neighbourhood
#
# In this section, the total number of listings and average price of listings in each neighbourhood are compared. It provides some insight about how listing price and number changed based on location.
# +
#calculate the total count of listings in each neighbourhood
df_neighbourhood_count = df_selected_2016_cleaned['zipcode'].value_counts()
#plot the results
plt.figure(figsize = (10, 8)) #determine the size of the chart
df_neighbourhood_count.plot.barh(title='Comparison of Listings', color = "midnightblue")
plt.xticks(rotation = 45, fontsize = 14) # format the labels for the x-axis
plt.yticks(fontsize = 14) # format the y-axis
plt.xlabel("Total Listings",fontsize = 14)
plt.savefig('comparison of total listings.png')
plt.show()
# -
#plot the listing price distribution in each neighbourhood
sns.set(rc={'figure.figsize':(14,9)})
ax = sns.boxplot(x = df_selected_2016_cleaned['zipcode'], y = df_selected_2016_cleaned['price'])
plt.legend(title = 'neighborhood comparison', fontsize='x-large', title_fontsize='40')
plt.xlabel(" ")
plt.ylabel("Listing Price ($)",fontsize = 14)
plt.savefig('pricing comparison for neighbourhood.png')
# +
#find number of total number of listings for each month in 2016
number_of_listings_by_month = pd.Series([12])
for i in range(1, 13):
number_of_listings_by_month[i] = len(df_selected_2016_cleaned[(df_selected_2016_cleaned['month'] == i)]['listing_id'].unique())
number_of_listings_by_month = number_of_listings_by_month.drop(0)
#plot the number of listings per month in 2016
plt.figure(figsize=(10,5))
plt.plot(number_of_listings_by_month)
plt.xticks(np.arange(1, 13, step=1))
plt.ylabel('Number of listings per month')
plt.xlabel('Month')
plt.title('Number of listings per month in 2016', fontsize = 14)
plt.savefig('number of available listings.png')
plt.show()
# +
#get list of neighbourhoods
neighbourhoods = df_selected_2016_cleaned['zipcode'].unique()
#get prices by month and neighbourhood
price_by_month_neighbourhood = df_selected_2016_cleaned.groupby(['month','zipcode']).mean().reset_index()
#plot prices for each neighbourhood
fig = plt.figure(figsize=(20,10))
ax = plt.subplot(111)
for neighbourhood in neighbourhoods:
ax.plot(price_by_month_neighbourhood[price_by_month_neighbourhood['zipcode'] == neighbourhood]['month'],
price_by_month_neighbourhood[price_by_month_neighbourhood['zipcode'] == neighbourhood]['price'],
label = neighbourhood)
box = ax.get_position()
ax.set_position([box.x0, box.y0, box.width * 0.8, box.height])
ax.legend(loc='center left', bbox_to_anchor=(1, 0.5),fontsize = 24)
plt.ylabel('Average price, $', fontsize = 30)
plt.xlabel('Month', fontsize = 30)
plt.title('Average price for neighbourhood, $', fontsize = 30)
plt.xticks(np.arange(1, 13, step=1),fontsize = 14)
plt.yticks(fontsize = 14)
plt.savefig('average price for neighbourhood.png')
plt.show()
# +
#get list of neighbourhoods
neighbourhoods = df_selected_2016_cleaned['zipcode'].unique()
#get total listing by month and neighbourhood
listing_by_month_neighbourhood = df_selected_2016_cleaned.groupby(['month','zipcode']).count().reset_index()
listing_by_month_neighbourhood
#plot total listing for each neighbourhood
fig = plt.figure(figsize=(20,10))
ax = plt.subplot(111)
for neighbourhood in neighbourhoods:
ax.plot(listing_by_month_neighbourhood[listing_by_month_neighbourhood['zipcode'] == neighbourhood]['month'],
listing_by_month_neighbourhood[listing_by_month_neighbourhood['zipcode'] == neighbourhood]['listing_id'],
label = neighbourhood)
box = ax.get_position()
ax.set_position([box.x0, box.y0, box.width * 0.8, box.height])
ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.ylabel('Total Listing per month')
plt.xlabel('Month')
plt.title('Total Listing per month', fontsize = 14)
plt.savefig('Total Listing per month.png')
plt.show()
# -
# # Section 4. Data Modeling
#
# The data is preprocessed as followed:
# 1. Recode the property_type and bed_type to simplify the model based on the number of each type and the average price of each type.
# 2. Remove the year column since it won't provide any useful information for the model prediction
#make a copy of the clean dataset for prediction
df_predict = df_selected_2016_cleaned.copy()
df_predict.head()
df_predict['property_type'].value_counts()
df_predict['room_type'].value_counts()
df_predict['bed_type'].value_counts()
df_predict.groupby(['bedrooms']).mean(['price']).sort_values(['price'])
df_predict.groupby(['bed_type']).mean(['price']).sort_values(['price'])
# +
# Recode property_type
def property_type(value):
if value in ['House', 'Townhouse', 'Loft']:
return 'House'
elif value in ['Apartment', 'Condominium', 'Chalet', 'Bed & Breakfast']:
return 'Apartment/Condo'
elif value in ['Camper/RV', 'Bungalow', 'Cabin']:
return 'Outdoor_Room'
elif value in ['Tent', 'Treehouse', 'Dorm', 'Yurt']:
return 'Outdoor_Tent'
return value
df_predict['property_type'] = df_predict['property_type'].apply(property_type)
# +
# Recode bed_type
def bed_type(value):
if value in ['Real Bed','Airbed']:
return 'Bed'
elif value in ['Futon','Pull-out Sofa','Couch']:
return 'Futon/Sofa/Couch'
return value
df_predict['bed_type'] = df_predict['bed_type'].apply(bed_type)
# -
df_predict.head()
df_predict_re = df_predict[['zipcode','property_type','room_type',
'accommodates','bathrooms','bedrooms',
'beds','bed_type','month','price']]
df_predict_re.head()
# ### Set up parameters
#
# Create dummy variables for categorical data and split the test/train dataset.
#select X,Y data
X = df_predict_re.iloc[:,:-1]
Y = df_predict_re.iloc[:,-1]
# select non-numeric variables and create dummies
non_num_vars = X.select_dtypes(include=['object']).columns
X[non_num_vars].head()
# create dummy variables
dummy_vars = pd.get_dummies(X[non_num_vars])
dummy_vars
# drop non-numeric variables and add the dummies
X_dummy = X.drop(non_num_vars,axis=1)
X_dummy = pd.merge(X_dummy,dummy_vars, left_index=True, right_index=True)
X_dummy.head()
#split the test and train dataset
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(X_dummy, Y, test_size = 0.2, random_state = 0)
# # Section 5. Evaluation
#
# ## Random Forest Regression Model
# +
# training the Random Forest Regression
from sklearn.ensemble import RandomForestRegressor
RFregressor = RandomForestRegressor(n_estimators=100,
criterion='mse',
random_state=42,
n_jobs=-1)
RFregressor.fit(X_train, Y_train)
#calculate scores for the model
y_train_preds_RF = RFregressor.predict(X_train)
y_test_preds_RF = RFregressor.predict(X_test)
print('Random Forest MSE train: %.3f, test: %.3f' % (
mean_squared_error(Y_train, y_train_preds_RF),
mean_squared_error(Y_test, y_test_preds_RF)))
print('Random Forest R^2 train: %.3f, test: %.3f' % (
r2_score(Y_train, y_train_preds_RF),
r2_score(Y_test, y_test_preds_RF)))
# -
# show the residuals of train and test
plt.scatter(y_train_preds_RF, y_train_preds_RF - Y_train,
c='blue', marker='o', label='Training data')
plt.scatter(y_test_preds_RF, y_test_preds_RF - Y_test,
c='lightgreen', marker='s', label='Test data')
plt.xlabel('Predicted values')
plt.ylabel('Residuals')
plt.legend(loc='upper left')
plt.show()
# show the comparison of actual and predict values
plt.scatter(y_train_preds_RF, Y_train,
c='blue', marker='o', label='Training data')
plt.scatter(y_test_preds_RF, Y_test,
c='lightgreen', marker='s', label='Test data')
plt.xlabel('Predicted values')
plt.ylabel('Actual listing values')
plt.legend(loc='upper left')
plt.show()
# +
#get feature importances from the model
headers = ["name", "score"]
values = sorted(zip(X_train.columns, RFregressor.feature_importances_), key=lambda x: x[1] * -1)
forest_feature_importances = pd.DataFrame(values, columns = headers)
forest_feature_importances = forest_feature_importances.sort_values(by = ['score'], ascending = False)
features = forest_feature_importances['name'][:15]
y_pos = np.arange(len(features))
scores = forest_feature_importances['score'][:15]
#plot feature importances
plt.figure(figsize=(10,5))
plt.barh(y_pos, scores,align='center', alpha=0.5)
plt.yticks(y_pos, features)
plt.xlabel('Score')
plt.ylabel('Features')
plt.title('Feature importances (Random Forest)')
plt.savefig('feature importances RF.png')
plt.show()
# -
# ## Linear Regression model
#import linear regression model
from sklearn.linear_model import LinearRegression
LRregressor = LinearRegression()
LRregressor.fit(X_train, Y_train)
# +
#calculate scores for the model
y_train_preds_LR = LRregressor.predict(X_train)
y_test_preds_LR = LRregressor.predict(X_test)
print('Linear Regression MSE train: %.3f, test: %.3f' % (
mean_squared_error(Y_train, y_train_preds_LR),
mean_squared_error(Y_test, y_test_preds_LR)))
print('Linear Regression R^2 train: %.3f, test: %.3f' % (
r2_score(Y_train, y_train_preds_LR),
r2_score(Y_test, y_test_preds_LR)))
# -
# show the residuals of train and test
plt.scatter(y_train_preds_LR, y_train_preds_LR - Y_train,
c='blue', marker='o', label='Training data')
plt.scatter(y_test_preds_LR, y_test_preds_LR - Y_test,
c='lightgreen', marker='s', label='Test data')
plt.xlabel('Predicted values')
plt.ylabel('Residuals')
plt.legend(loc='upper left')
plt.show()
# show the comparison of predicted and actual values
plt.scatter(y_train_preds_LR, Y_train,
c='blue', marker='o', label='Training data')
plt.scatter(y_test_preds_LR, Y_test,
c='lightgreen', marker='s', label='Test data')
plt.xlabel('Predicted values')
plt.ylabel('Actual listing values')
plt.legend(loc='upper left')
plt.show()
# From those two model performance above, Random Forest perform much better than Linear Regression model. Linear Regression modeling output were consistently lower than actual oberservation.
# # Section 6. Evaluate the results
#
# ## Make prediction
#
# In this section, input parameters were format in a way that is easier for the users to make their own prediction based on their own selection of input parameters. Random Forest model is used for prediction.
#Check the required input
X_test.columns
# +
#Input list
#Room info:
accommodates = 6
bathrooms = 2
bedrooms = 3
beds = 3
#Month of Year:
#Select from 1 to 12
month = 7
#Location:
#Select one from the following and label 1 for the one selected
Central = 0
Downtown = 1
Magnolia_Queen_Anne = 0
Northeast = 0
Northwest = 0
Southeast = 0
West = 0
#Property type:
#Select one from the following and label 1 for the one selected
Apartment_Condo = 1
Boat = 0
House = 0
Other = 0
Outdoor_Room = 0
Outdoor_Tent = 0
#Room setup:
#Select one from the following and label 1 for the one selected
Entire_home_apt = 1
Private_room = 0
Shared_room = 0
#Bed type:
#Select one from the following and label 1 for the one selected
Bed_type_bed = 1
Bed_type_other = 0
Your_BNB = pd.DataFrame(np.array([[accommodates, bathrooms, bedrooms, beds, month, Central, Downtown, Magnolia_Queen_Anne,
Northeast, Northwest, Southeast, West, Apartment_Condo, Boat, House, Other, Outdoor_Room,
Outdoor_Tent, Entire_home_apt, Private_room, Shared_room, Bed_type_bed, Bed_type_other]]),
columns=['accommodates', 'bathrooms', 'bedrooms', 'beds', 'month',
'zipcode_Central Seattle', 'zipcode_Downtown Seattle',
'zipcode_Magnolia&Queen Anne', 'zipcode_Northeast Seattle',
'zipcode_Northwest Seattle', 'zipcode_Southeast Seattle',
'zipcode_West Seattle&Delridge', 'property_type_Apartment/Condo',
'property_type_Boat', 'property_type_House', 'property_type_Other',
'property_type_Outdoor_Room', 'property_type_Outdoor_Tent',
'room_type_Entire home/apt', 'room_type_Private room',
'room_type_Shared room', 'bed_type_Bed', 'bed_type_Futon/Sofa/Couch'])
# -
#predict price using random forest
Predict_price_RF = RFregressor.predict(Your_BNB)
print('Your AirBNB can be priced at $' + str(Predict_price_RF[0].round(2)) + ' based on our awesome Random Forest Model.')
| Seattle AirBNB analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercício
#
# O dataset utilizado neste execício é o [SF Salaries Dataset](https://www.kaggle.com/kaggle/sf-salaries), disponibilizado em uma competição do Kaggle.
import pandas as pd
import seaborn as sbn
import matplotlib.pyplot as plt
pd.options.display.float_format = '{:,.2f}'.format
# ### Faça a leitura do arquivo "Salaries.csv"
df = pd.read_csv('../../data/Salaries.csv')
df.head(3)
# ### Apresente algumas informações sobre o dataframe com .info()
df.info()
# ### Qual a média de valores na coluna "BasePay"
df['BasePay'].mean()
# ### Qual o maior valor da coluna "OvertimePay"
df['OvertimePay'].max()
# ### Qual o "JobTitle" de ALBERT PARDINI (EmployeeName) ?
df[df['EmployeeName'] == '<NAME>']['JobTitle'].iloc[0]
# ### Qual o salário de ALBERT PARDINI, incluindo os benefícios (TotalPayBenefits)?
df[df['EmployeeName'] == '<NAME>']['TotalPayBenefits']
# ### Qual o nome da pessoa com o maior salário? E quem tem o menor salário?
df[(df.BasePay == df.BasePay.max()) | (df.BasePay == df.BasePay.min())]
# ### Qual a média salarial dos funcionários de 2011 a 2014? E qual a média do salário por ano?
df[(df['Year']>=2011) & (df['Year']<=2014)]['BasePay'].mean()
df[(df['Year']>=2011) & (df['Year']<=2014)].groupby(by='Year')['BasePay'].mean()
# ### Qual o desvio padrão dos salários, incluindo os benefícios?
df['TotalPayBenefits'].std()
#
# ### Quantas pessoas estão dentro do intervalo de 1 desvio padrão?
# +
limite_min = df['BasePay'].mean() - df['BasePay'].std()
limite_max = df['BasePay'].mean() + df['BasePay'].std()
df[(df['BasePay']>=limite_min) & (df['BasePay']<=limite_max)]['Id'].count()
# +
def in_first_std(x, p_mean, p_std):
return (( x >= (p_mean-p_std)) & ( x <= (p_mean+p_std)))
df[df['BasePay'].apply(in_first_std, args=(df.BasePay.mean(), df.BasePay.std()))]['Id'].count()
# -
# ========================================================
# ## Desafios
# ### O salário segue uma distribuição normal?
df['TotalPay'].plot.hist()
# ### Para os campos com valores nulos, como você pensaria em tratá-los?
df.describe()
# ### Você conseguiria criar uma nova variável com base nas informações existentes?
df['media'] = df['TotalPay'].mean() ; df.head(2)
| notebooks/Pandas/Pandas - Exercicio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial
# The `AtUproot` class is the user interface to the underlying alphatwirl tools which are used to read in and loop over blocks of events.
#
# In this example we will read in Monte Carlo generated events for the production of the Z boson which decays into muons. The mass of the Z is encoded in the inputs and will be plotted as a histogram.
# +
from collections import namedtuple
import matplotlib.pyplot as plt
import numpy as np
from atuproot.AtUproot import AtUproot
# -
# Initialise an instance of `AtUproot` with the desired options:
# * `quiet` - suppressed the output to stdout
# * `parallel_mode` - option for parallelisation
# * `process` - for 'multiprocessing mode. The number of cores to run on
# * `max_blocks_per_dataset` - number of blocks per datasets (will either be hzz or zmumu here). A block is a group of events read into a numpy array at once.
# * `max_blocks_per_process` - number of blocks per process. A process is a singular unit of the set of parallel jobs.
# * `nevents_per_block` - this defines the size of a block. i.e. the number of events to read in to an array
# * `profile` - boolean. Profile the code and output the results into `profile_out_path`. To profile the code that operates on the tree then run in 'multiprocessing mode with zero processes (i.e. no parallelisation)
# * `profile_out_path` - see above.
process = AtUproot(
"output",
quiet = False,
parallel_mode = 'multiprocessing',
process = 0,
max_blocks_per_dataset = -1,
max_blocks_per_process = -1,
nevents_per_block = 1000000,
profile = False,
profile_out_path = 'profile.txt',
)
# We now have to define a few modules to process the contents of the tree.
#
# Here we create a `TestReader` that has no required inheritance and has the following methods:
# * `__init__` - initialise whatever you need
# * `begin` - runs at the starts of each process with some information on the event (no arrays have been read in yet)
# * `end` - runs at the end of each process. Mainly to remove any non-picklable objects such as lambda functions
# * `event` - runs on each block of events. The attributes of `event` should include all branches/leaves of the tree (and additional information)
# * `merge` - takes `other` which is the a `TestReader` instances from anther process for the same dataset. This should define how to merge the results. Here we just sum the histograms (if they exist)
class TestReader(object):
def __init__(self):
self.bins = np.linspace(60., 120., 61)
def begin(self, event):
self.hist = None
print(event.config)
def end(self):
pass
def event(self, event):
hist, _ = np.histogram(event.M, self.bins)
if self.hist is None:
self.hist = hist
else:
self.hist += hist
def merge(self, other):
if self.hist is None:
self = other
return
if other.hist is None:
return
self.hist += other.hist
# Next is the `TestCollector` class which will be paired with the `TestReader` class. This has one method: `collect`, which takes a `dataset_readers_list` argument. This argument is a list of tuples of all `(dataset, readers)` combinations.
#
# We're running over 1 dataset so we'll take the first item in the list. Then the 2nd item in the tuple (i.e. the readers) and the 1st item in the readers list (we only have 1 reader). Therefore, `reader` will correspond to our `TestReader` for our 1st dataset.
class TestCollector(object):
def collect(self, dataset_readers_list):
reader = dataset_readers_list[0][1][0]
return (reader.bins, reader.hist)
# We pass the initialised `TestReader` and `TestCollector` as a list of pairs to `process`
sequence = [(TestReader(), TestCollector())]
# Now we need to define our datasets (our inputs). This must be a list of classes or namedtuples with, at least, the attributes: `name` (your chosen name for the dataset), `tree` (tree name within the files) and `files` (list of files for the dataset).
Dataset = namedtuple("Dataset", "name tree files")
hzz = Dataset(name = "hzz",
tree = "events",
files = ["HZZ.root"])
zuu = Dataset(name = "zuu",
tree = "events",
files = ["Zmumu.root"])
datasets = [zuu] #,hzz]
# Everything is setup, we just need to run the whole process as follows. This will return the return values of all our collectors
result = process.run(datasets, sequence)
# We want to get the 1st collector of our 1st dataset:
bins, hist = result[0][0]
plt.bar(
(bins[1:]+bins[:-1])/2,
height = hist,
width = (bins[1:]-bins[:-1])
)
| binder/tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Distributed Image Processing in the Cloud
# ## Learning objectives
#
# - Become familiar with **lazy, distributed Python image processing** with Dask
# - Learn how to start and interact with a **Coiled.io cloud cluster**
# - Understand why **consistent software environments** are required and how to create them
# ## Distributed Python Image Processing with Dask
# 
#
# [Dask](https://dask.org/) is *a Python-based, flexible library for parallel computing*. Dask provides dynamic task scheduling optimized for interactive computing, and parallel-friendly collections such as the `dask.array`.
#
# Dask can run tasks in parallel on a single machine via threads or processes. Or, the same code can trivially also be executed on a collection of local workstations connected via SSH, or an [HPC cluster with a traditional job scheduler](https://docs.dask.org/en/latest/setup/hpc.html), or a managed Kubernetes-based cloud cluster.
# +
# Load the OME-Zarr image array
import dask.array as da
arr = da.from_zarr('https://s3.embassy.ebi.ac.uk/idr/zarr/v0.1/6001240.zarr', '0')
arr
# +
import itk
image = itk.imread('data/6001240.tif')
print(type(image))
print(image.shape)
# +
import numpy as np
import dask.array as da
arr = np.asarray(image)
arr = da.from_array(arr, chunks=64)
arr
# -
from itkwidgets import view
view(arr)
# +
def denoise(array_chunk):
# Currently required for serialization
import itk
itk.force_load()
# itk already has parallelism on a single-machine -- if running
# with dask in parallel on a single-machine, avoid over-subscription
#
# itk.set_nthreads(1)
array_float = array_chunk.astype(np.float32)
denoised = itk.curvature_flow_image_filter(array_float,
number_of_iterations=10)
denoised = itk.median_image_filter(denoised, radius=1)
denoised = itk.smoothing_recursive_gaussian_image_filter(denoised,
sigma=0.5)
return denoised
denoised = arr.map_blocks(denoise, dtype=np.float32)
# -
denoised = denoised.compute()
view(denoised)
denoised = arr.map_overlap(denoise, dtype=np.float32,
depth=6)
denoised = denoised.compute()
view(denoised)
# ## Cloud cluster
# [Coiled.io](https://coiled.io/) is a service that provides a dynamic cloud cluster with minimal configuration.
# +
#ciskip
# Start a new cloud cluster
# You must first log into Coiled.
import coiled
from dask.distributed import Client
# Set to re-use a running cluster when re-running the notebook.
# Listed at https://cloud.coiled.io/<username>/clusters.
# name = 'thewtex-dd6121ae-0'
name = None
cluster = coiled.Cluster(n_workers=4,
worker_cpu=2,
worker_memory='6G',
name=name,
software='thewtex/coiled-science-thursdays-itk')
client = Client(cluster)
# Click on the *Dashboard* link
client
# -
# *Note:*
#
# It is **critical** that the:
#
# - Jupyter client
# - Jupyter kernel
# - Dask scheduler
# - Dask workers
#
# all have a consistent software environment.
#
# See [the coiled documentation on how to create a consistent software environment](https://docs.coiled.io/user_guide/software_environment.html).
# %pycat ./create_coiled_software_environment.py
# +
# Run on the cloud cluster -- keep an eye on the dashboard!
denoised = arr.map_overlap(denoise, dtype=np.float32,
depth=6)
denoised = denoised.compute()
# -
# ### Dask Imaging Resources
#
# - [Dask documention](https://docs.dask.org/en/latest/)
# - [Coiled documentation](https://docs.coiled.io/user_guide/index.html)
# - [`dask-image` documentation](https://image.dask.org/en/latest/)
# - [Bioimaging Image2Knowledge (I2K) 2020 Tutorial](https://github.com/thewtex/modern-insights-from-microscopy-images)
# - [SuperComputing 2020 pyHPC Material Science Publication](https://github.com/dani-lbnl/SC20_pyHPC)
| 2_Easy_Cloud_Scaling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modeling and Simulation in Python
#
# Chapter 7
#
# Copyright 2017 <NAME>
#
# License: [Creative Commons Attribution 4.0 International](https://creativecommons.org/licenses/by/4.0)
#
# +
# Configure Jupyter so figures appear in the notebook
# %matplotlib inline
# Configure Jupyter to display the assigned value after an assignment
# %config InteractiveShell.ast_node_interactivity='last_expr_or_assign'
# import functions from the modsim.py module
from modsim import *
from pandas import read_html
# -
# ### Code from the previous chapter
filename = 'data/World_population_estimates.html'
tables = read_html(filename, header=0, index_col=0, decimal='M')
table2 = tables[2]
table2.columns = ['census', 'prb', 'un', 'maddison',
'hyde', 'tanton', 'biraben', 'mj',
'thomlinson', 'durand', 'clark']
un = table2.un / 1e9
un.head()
census = table2.census / 1e9
census.head()
def plot_results(census, un, timeseries, title):
"""Plot the estimates and the model.
census: TimeSeries of population estimates
un: TimeSeries of population estimates
timeseries: TimeSeries of simulation results
title: string
"""
plot(census, ':', label='US Census')
plot(un, '--', label='UN DESA')
plot(timeseries, color='gray', label='model')
decorate(xlabel='Year',
ylabel='World population (billion)',
title=title)
def run_simulation(system, update_func):
"""Simulate the system using any update function.
system: System object
update_func: function that computes the population next year
returns: TimeSeries
"""
results = TimeSeries()
results[system.t_0] = system.p_0
for t in linrange(system.t_0, system.t_end):
results[t+1] = update_func(results[t], t, system)
return results
# ### Quadratic growth
# Here's the implementation of the quadratic growth model.
def update_func_quad(pop, t, system):
"""Compute the population next year with a quadratic model.
pop: current population
t: current year
system: system object containing parameters of the model
returns: population next year
"""
net_growth = system.alpha * pop + system.beta * pop**2
return pop + net_growth
# Here's a `System` object with the parameters `alpha` and `beta`:
# +
t_0 = get_first_label(census)
t_end = get_last_label(census)
p_0 = census[t_0]
system = System(t_0=t_0,
t_end=t_end,
p_0=p_0,
#alpha=0.025,
#beta=-0.0018)
alpha=0.026,
beta=-0.0020)
# -
# And here are the results.
results = run_simulation(system, update_func_quad)
plot_results(census, un, results, 'Quadratic model')
savefig('figs/chap03-fig04.pdf')
# **Exercise:** Can you find values for the parameters that make the model fit better?
# ### Equilibrium
#
# To understand the quadratic model better, let's plot net growth as a function of population.
pop_array = linspace(0, 15, 100)
net_growth_array = system.alpha * pop_array + system.beta * pop_array**2
None
# Here's what it looks like.
# +
sns.set_style('whitegrid')
plot(pop_array, net_growth_array)
decorate(xlabel='Population (billions)',
ylabel='Net growth (billions)')
savefig('figs/chap03-fig05.pdf')
sns.set_style('white')
# -
# Here's what it looks like. Remember that the x axis is population now, not time.
# It looks like the growth rate passes through 0 when the population is a little less than 14 billion.
#
# In the book we found that the net growth is 0 when the population is $-\alpha/\beta$:
-system.alpha / system.beta
# This is the equilibrium the population tends toward.
# `sns` is a library called Seaborn which provides functions that control the appearance of plots. In this case I want a grid to make it easier to estimate the population where the growth rate crosses through 0.
# ### Dysfunctions
# When people first learn about functions, there are a few things they often find confusing. In this section I present and explain some common problems with functions.
#
# As an example, suppose you want a function that takes a `System` object, with variables `alpha` and `beta`, as a parameter and computes the carrying capacity, `-alpha/beta`. Here's a good solution:
# +
def carrying_capacity(system):
K = -system.alpha / system.beta
return K
sys1 = System(alpha=0.025, beta=-0.0018)
pop = carrying_capacity(sys1)
print(pop)
# -
# Now let's see all the ways that can go wrong.
#
# **Dysfunction #1:** Not using parameters. In the following version, the function doesn't take any parameters; when `sys1` appears inside the function, it refers to the object we created outside the function.
#
# +
def carrying_capacity():
K = -sys1.alpha / sys1.beta
return K
sys1 = System(alpha=0.025, beta=-0.0018)
pop = carrying_capacity()
print(pop)
# -
# This version actually works, but it is not as versatile as it could be. If there are several `System` objects, this function can only work with one of them, and only if it is named `system`.
#
# **Dysfunction #2:** Clobbering the parameters. When people first learn about parameters, they often write functions like this:
# +
def carrying_capacity(system):
system = System(alpha=0.025, beta=-0.0018)
K = -system.alpha / system.beta
return K
sys1 = System(alpha=0.025, beta=-0.0018)
pop = carrying_capacity(sys1)
print(pop)
# -
# In this example, we have a `System` object named `sys1` that gets passed as an argument to `carrying_capacity`. But when the function runs, it ignores the argument and immediately replaces it with a new `System` object. As a result, this function always returns the same value, no matter what argument is passed.
#
# When you write a function, you generally don't know what the values of the parameters will be. Your job is to write a function that works for any valid values. If you assign your own values to the parameters, you defeat the whole purpose of functions.
#
#
# **Dysfunction #3:** No return value. Here's a version that computes the value of `K` but doesn't return it.
# +
def carrying_capacity(system):
K = -system.alpha / system.beta
sys1 = System(alpha=0.025, beta=-0.0018)
pop = carrying_capacity(sys1)
print(pop)
# -
# A function that doesn't have a return statement always returns a special value called `None`, so in this example the value of `pop` is `None`. If you are debugging a program and find that the value of a variable is `None` when it shouldn't be, a function without a return statement is a likely cause.
#
# **Dysfunction #4:** Ignoring the return value. Finally, here's a version where the function is correct, but the way it's used is not.
# +
def carrying_capacity(system):
K = -system.alpha / system.beta
return K
sys2 = System(alpha=0.025, beta=-0.0018)
carrying_capacity(sys2)
# print(K) This line won't work because K only exists inside the function.
# -
# In this example, `carrying_capacity` runs and returns `K`, but the return value is dropped.
#
# When you call a function that returns a value, you should do something with the result. Often you assign it to a variable, as in the previous examples, but you can also use it as part of an expression.
#
# For example, you could eliminate the temporary variable `pop` like this:
print(carrying_capacity(sys1))
# Or if you had more than one system, you could compute the total carrying capacity like this:
#
total = carrying_capacity(sys1) + carrying_capacity(sys2)
total
# ## Exercises
#
# **Exercise:** In the book, I present a different way to parameterize the quadratic model:
#
# $ \Delta p = r p (1 - p / K) $
#
# where $r=\alpha$ and $K=-\alpha/\beta$. Write a version of `update_func` that implements this version of the model. Test it by computing the values of `r` and `K` that correspond to `alpha=0.025, beta=-0.0018`, and confirm that you get the same results.
def update_func_quad_new(pop, t, system):
"""Compute the population next year with a quadratic model.
pop: current population
t: current year
system: system object containing parameters of the model
returns: population next year
"""
system.r = system.alpha
system.k = -system.alpha/system.beta
net_growth = system.r * pop * (1 - (pop/system.k))
return pop + net_growth
# +
t_0 = get_first_label(census)
t_end = get_last_label(census)
p_0 = census[t_0]
system = System(t_0=t_0,
t_end=t_end,
p_0=p_0,
alpha=0.025,
beta=-0.0018)
# -
results = run_simulation(system, update_func_quad)
plot_results(census, un, results, 'Quadratic model')
savefig('figs/chap03-fig04.pdf')
| code/chap07.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# %matplotlib inline
from matplotlib import pyplot as plt
import numpy as np
np.set_printoptions(threshold=np.nan)
from Strawry.env import Env
env = Env()
# +
env.set_target(temp=10,humi=80,co2=350,light=0,watp=0)
# +
Q= np.zeros((env.n_state, env.n_action))
y = 0.99
lr = 0.25
num_episodes = 2000
rList = []
alpha=np.log(0.1)/num_episodes # decay learning rate
for i in range(num_episodes):
lr= np.exp(alpha*i) #decay learning rate
s = env.reset()
rAll = 0
done=False
while done==False: #random action to get next state and action
if np.random.rand() < lr*0.01:
a = np.random.randint(env.n_action)
else:
a = np.argmax(Q[s,:] )
s1,reward,done,_ = env.step(a)
if done:
r = 1 if reward > 0.0 else -0.1
else:
r =-0.00001
#Update Q-Table with new knowledge
Q[s,a] = Q[s,a] + lr*(r + y*np.max(Q[s1,:]) - Q[s,a])
# Q[s,a] = Q[s,a] + lr*(r + y*np.min(Q[s1,:]) - Q[s,a])
rAll += reward
s = s1
if done == True:
break
rList.append(rAll)
print ("Score over time: " + str(sum(rList[-100:])/100.0))
plt.plot(np.convolve(np.ones(100),rList,"valid"))
# -
print(Q)
print(s)
print(rAll)
| AI3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/artbrgn/DS-Unit-1-Sprint-3-Statistical-Tests-and-Experiments/blob/master/module1-statistics-probability-and-inference/Arturo_Obregon_LS_DS_131_Statistics_Probability_Assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="J53ShiRp6Hyj" colab_type="text"
# <img align="left" src="https://lever-client-logos.s3.amazonaws.com/864372b1-534c-480e-acd5-9711f850815c-1524247202159.png" width=200>
# <br></br>
# <br></br>
#
# ## *Data Science Unit 1 Sprint 3 Assignment 1*
#
# # Apply the t-test to real data
#
# Your assignment is to determine which issues have "statistically significant" differences between political parties in this [1980s congressional voting data](https://archive.ics.uci.edu/ml/datasets/Congressional+Voting+Records). The data consists of 435 instances (one for each congressperson), a class (democrat or republican), and 16 binary attributes (yes or no for voting for or against certain issues). Be aware - there are missing values!
#
# Your goals:
#
# 1. Load and clean the data (or determine the best method to drop observations when running tests)
# 2. Using hypothesis testing, find an issue that democrats support more than republicans with p < 0.01
# 3. Using hypothesis testing, find an issue that republicans support more than democrats with p < 0.01
# 4. Using hypothesis testing, find an issue where the difference between republicans and democrats has p > 0.1 (i.e. there may not be much of a difference)
#
# Note that this data will involve *2 sample* t-tests, because you're comparing averages across two groups (republicans and democrats) rather than a single group against a null hypothesis.
#
# Stretch goals:
#
# 1. Refactor your code into functions so it's easy to rerun with arbitrary variables
# 2. Apply hypothesis testing to your personal project data (for the purposes of this notebook you can type a summary of the hypothesis you formed and tested)
# + id="wYBmY38Z6Hym" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 208} outputId="054fdf4a-385e-4c62-ba5e-8d15967f32f2"
### YOUR CODE STARTS HERE
# !wget https://archive.ics.uci.edu/ml/machine-learning-databases/voting-records/house-votes-84.data
# + id="0ZPMvcLXuCg9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="75602a36-1008-43cb-ac5e-98ac9b9f0f60"
import pandas as pd
import numpy as np
df = pd.read_csv('house-votes-84.data',
header=None,
names=['party','handicapped-infants','water-project',
'budget','physician-fee-freeze', 'el-salvador-aid',
'religious-groups','anti-satellite-ban',
'aid-to-contras','mx-missile','immigration',
'synfuels', 'education', 'right-to-sue','crime','duty-free',
'south-africa'])
print(df.shape)
df.head(1000)
# + id="FKL_C3cHuGpo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 257} outputId="c97add08-1881-4a0f-885f-50b0d55994ea"
df = df.replace({'?':np.NaN, 'n':0, 'y':1})
df.head()
# + id="LFLHAXi-uX71" colab_type="code" colab={}
dem = df[df['party'] == 'democrat']
rep = df[df['party'] == 'republican']
# + id="SgXtVv9Jvrde" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 165} outputId="0bd500bd-c28f-482c-bfad-c166165b18e6"
dem.groupby(['party']).sum()
# + id="0cdm6Yrvvx1A" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 165} outputId="7b139687-9671-4eb2-f6df-fabe2f1ee0aa"
rep.groupby(['party']).sum()
# + id="KewKUQKBv0TF" colab_type="code" colab={}
from scipy.stats import ttest_1samp
# + id="FYo9uWgcyMhh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="81724356-13ed-4bd6-9626-c989b1897fa2"
rep['immigration'].mean()
# + id="kpXuiYKhyoBv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="8516d69e-6f67-49b9-9a0d-8b0012878292"
rep['immigration'].isnull().sum()
# + id="kVsB2mHQywIi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="6aad4117-b3e0-43b0-b3f8-ba2e773a2e30"
len(rep['immigration']) - rep['immigration'].isnull().sum()
# + [markdown] id="KCLD277p1PK-" colab_type="text"
# 1) Null Hypothesis:
#
# In 1-sample t-tests YOU GET TO CHOOSE YOUR NULL HYPOTHESIS
#
# H0 : 0.0 - There is ZERO republican support for this bill
#
# 2) Alternative Hypothesis
#
# Ha : x¯≠0 - There is non-zero support for the budget bill among repulbicans.
#
# 3) Confidence Level: 95% or .95
# + id="DRE5m7M4y3oz" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="155a900d-f6b4-4cb8-a175-bfa7a1b486d0"
ttest_1samp(rep['immigration'], 0, nan_policy = 'omit')
# + [markdown] id="tymfqcSr1cpR" colab_type="text"
# 4) t-statistic: 14.3765
#
# 5) p-value of .00000000000000000000000000007541
# ______________________________________________________________________
#
# Concluision: Due to a p-value of 0 I reject the null hypothesis that republican support is zero and conclude that republican support is non-zero.
#
#
# + [markdown] id="sUkiEkXw4o-6" colab_type="text"
# 1) Null Hypothesis:
#
# In 1-sample t-tests YOU GET TO CHOOSE YOUR NULL HYPOTHESIS
#
# H0 : 0.0 - There is ZERO democratic support for this bill
#
# 2) Alternative Hypothesis
#
# Ha : x¯≠0 - There is non-zero support for the budget bill among democratics.
#
# 3) Confidence Level: 95% or .95
# + id="bA3ULCEM01As" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="84aa995d-38df-4087-cc12-77abb4501c1e"
ttest_1samp(dem['synfuels'], 0, nan_policy = 'omit')
# + [markdown] id="p5G7NlI_4fdl" colab_type="text"
# 4) t-statistic: 16.1259
#
# 5) p-value of .000000000000000000000000000000000000009159
# ______________________________________________________________________
#
# Concluision: Due to a p-value of 0 I reject the null hypothesis that democratic support is zero and conclude that democratic support is non-zero.
# + [markdown] id="XXkAJsch9UQH" colab_type="text"
# ---
# ---
# 2-SAMPLE TEST
#
# + id="dKo4ddi6Bg-b" colab_type="code" colab={}
from scipy.stats import ttest_ind
import numpy as np
# + id="pAvtwRem4a-s" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="6ed97fb6-b3f9-4015-acef-1ae5b599fc6c"
ttest_ind(rep['education'], dem['education'], nan_policy = 'omit')
# + id="mRtE_0p4AmCt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="0d283a60-d5a3-4d88-92d2-fcd9c3f20fc2"
rep['education'].mean()
# + id="PcBCObX1Cbsg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="4d01eb9f-fe42-41e7-e1b0-f85b756a65e4"
dem['education'].mean()
# + id="sOoQa_WQCfUo" colab_type="code" colab={}
| module1-statistics-probability-and-inference/Arturo_Obregon_LS_DS_131_Statistics_Probability_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Kivy指南-2-画图app
#
# > 仿照Windows自带的画图程序做一个画图app,除了支持Andorid和iOS,也支持Windows
#
# - toc: true
# - badges: true
# - comments: true
# - categories: [jupyter,Kivy,Android,iOS]
# - image: kbpic/2.1paintapp.png
# 在第一章做时钟app时,我们用了Kivy的标准部件:布局,文本框和按钮。通过这些高层次的抽象,我们能够灵活的修改部件的外观—,可以使用一整套成熟的组件,而不仅仅是单个原始图形。这种方式并非放之四海而皆准,马上你就会看到,Kivy还提供了低层的抽象工具:画点和线。
#
# <!-- TEASER_END-->
# 我认为做画图app是自由绘画最好的方式。我们的应用会看着有点像Windows自带的画图程序。
#
# 不同的是,我们的画图app支持多平台,包括Andorid和iOS。我们也忽略了图像处理的功能,像矩形选框,图层,保存文件等。这些功能可以自己练习。
#
# >关于移动设备:Kivy完全支持iOS开发,即使你没有类似开发经验也不难。因此,建议你先在熟悉的平台上快速实现app,这样就可以省略编译的时间和一堆细节。Android开发更简单,由于[Kivy Launcher](https://
# play.google.com/store/apps/details?id=org.
# kivy.pygame)可以让Kivy代码直接在Android上运行。
# >Kivy可以不用编译直接在Andorid上运行测试,相当给力,绝对RAD(rapid application development)。
# >窗口改变大小的问题,并没有广泛用于移动设备,Kivy应用在不同的移动设备和桌面系统平台使用类似的处理方式。因此,开始编写和调试都非常容易,直到版本确定的最后阶段才需要集中精力弥补这些问题。
# 我们还会学习Kivy中两个相反的功能:触摸屏的多点触控和桌面系统的鼠标点击。
#
# 作为移动设备的第一大法,Kivy为多点触控输入提供了一个模拟层,可以使用鼠标。可以通过右键激活功能。但是,这个多点触控模拟器并不适合真实的场景,仅适合调试用。
#
# 画图app最会这这样:
#
# 
# ## 设置画板
# 我们的app通过root部件自动覆盖全局,整个屏幕都可以画画。到后面增加工具按钮的时候再调整。
#
# root部件是处于最外层,每个Kivy的app都有一个,可以根据app的需求制定任何部件作为root部件。比如上一章的时钟app,`BoxLayout`就是root部件;如果没其他要求,布局部件就是用来包裹其他控件的。
#
# 现在这个画图app,我们需要root部件具有更多的功能;用户应该可以画线条,支持多点触控。不过Kivy没有自带这些功能,所以我们自己建。
#
# 建立新部件很简单,只要继承Kivy的`Widget`类就行。如下所示:
# +
from kivy.app import App
from kivy.uix.widget import Widget
class CanvasWidget(Widget):
pass
class PaintApp(App):
def build(self):
return CanvasWidget()
if __name__ == '__main__':
PaintApp().run()
# -
# 这就是画图app的`main.py`,`PaintApp`类就是应用的起点。以后我们不会重复这些代码,只把重要的部分显示出来。
# >`Widget`类通常作为基类,就行Python的`object`和Java的`Object`。当它按照`as is`方式使用时,`Widget`功能极少。它没有可以直接拿来用的可视化的外观和属性。`Widget`的子类都是很简单易用的。
# ## 制作好看的外观
# 首先,让我们做个好看的外观,虽然不是核心功能,但长相影响第一印象。下面我们改改外观,包括窗口大小,鼠标形状。
# ### 可视化外观
# 我认为任何画图软件的背景色都应该是白的。和第一章类似,我们在`__name = '__main__'`后面加上就行:
# +
from kivy.core.window import Window
from kivy.utils import get_color_from_hex
Window.clearcolor = get_color_from_hex('#FFFFFF')
# -
# 你可能想把`import`语句放到前面,其实Kivy的一些模块导入有顺序要求,且会产生副作用,尤其是`Window`对象。这在好的Python程序中很少见,导入模块产生的副作用有点小问题。
# ### 窗口大小
# 另一个要改的就是窗口大小,下面的改变不影响移动设备。在桌面系统上,Kivy的窗口时可以调整的,后面我们会设置禁止调整。
#
# >如果目标设备明确,设置窗口大小是很有用的,这样就可以决定屏幕分辨率的参数,实现最好的适配效果。
#
# 要改变窗口大小,就把下面的代码放到`from kivy.core.window import Window`上面。
# +
from kivy.config import Config
Config.set('graphics', 'width', '960')
Config.set('graphics', 'height', '540') # 16:9
# -
# 如果要禁止窗口调整:
Config.set('graphics', 'resizable', '0')
# 如果没有充分理由,千万别这么做,因为把窗口调整这点小自由从用户手中拿走实在太伤感情了。如果把应用像素精确到1px,移动设备用户可能就不爽了,而Kivy布局可以建立自适应的界面。
# ### 鼠标样式
# 之后就是改变鼠标光标的样式。Kivy没有支持,不过可以过Pygame实现,基于SDL窗口和OpenGL内容管理模块,在Kivy的桌面平台应用开发中用途广泛。如果你这么用,移动应用大都不支持Pygame。
# 之后就是改变鼠标光标的样式。Kivy没有支持,不过可以过Pygame实现,基于SDL窗口和OpenGL内容管理模块,在Kivy的桌面平台应用开发中用途广泛。如果你这么用,移动应用大都不支持Pygame。
# 
#
# 图中`@`是黑的,`-`是白的,其他字符是透明的。所以的线都是等宽的,且是8的倍数(SDL的限制)。鼠标的光标运行后是这样:
# 
#
# >当前的Pygame版本有个bug,`pygame.cursors.compile()`黑白显示颠倒。以后应该会修复。不过`pygame_compile_cursor()`是正确的方法,[Pygame的Simple DirectMedia Layer (SDL)兼容库](http://goo.gl/2KaepD)。
#
# 现在,我们把光标应用到app中,替换`PaintApp.build`方法:
from kivy.base import EventLoop
class PaintApp(App):
def build(self):
EventLoop.ensure_window()
if EventLoop.window.__class__.__name__.endswith('Pygame'):
try:
from pygame import mouse
# pygame_compile_cursor is a fixed version of
# pygame.cursors.compile
a, b = pygame_compile_cursor()
mouse.set_cursor((24, 24), (9, 9), a, b)
except:
pass
return CanvasWidget()
# 代码很简单,注意下面四点:
# - `EventLoop.ensure_window()`: 这个函数到app窗口 ( `EventLoop.window` ) 准备好才执行。
# - `EventLoop.window.__class__.__name__.endswith('Pygame')`:
# 这个条件检查窗口名称Pygame,只是Pygame条件下才执行自定义光标。
# - `try ... except`模块里面是Pygame的`mouse.set_cursor`。
# - 变量`a`和`b`通过SDL构建了光标,表示异或(XOR)和与(AND),都是SDL独有的实现方式。
# >[Pygame文档](http://www.pygame.org)提供了全部的api说明。
#
# 现在做的这些比Kivy的模块更底层,并不常用,不过也不用害怕触及更多的细节。有很多功能只能通过底层的模块实现,因为Kivy还没达到面面俱到的程度。尤其是那些不能跨平台的功能,会涉及很多系统层的实现。
#
# Kivy/Pygame/SDL/OS的关系如下图所示:
# 
#
# SDL已经把系统底层的API都封装好了,兼容多个系统,Pygame再将SDL转换成Python,Kivy可以导入Pygame模块调用这些功能。
#
# >为什么不直接用SDL呢?可以看[SDL文档](https://www.libsdl.org/)。
# ### 多点触控模拟器
# 让运行桌面应用时,Kivy提供了一个模拟器实现多点触控操作。实际上是一个右击行为,获取半透明的点;按住右键时可以拖拽。
#
# 如果你没有真实的多点触控设备,这个功能可能适合调试。但是,也会占用右键的功能。不调试的时候还是建议你禁用这个功能,避免对用户造成困扰。设置方法如下:
Config.set('input', 'mouse', 'mouse,disable_multitouch')
# ## 触摸绘画
# 要实现用户通过触摸绘画的效果,可以在用户输入后屏幕会出现一个圆圈。
#
# 部件如果带`on_touch_down`事件,就可以实现上述功能。正在需要的是点击位置的坐标,为`CanvasWidget`添加一个方法获取即可:
class CanvasWidget(Widget):
def on_touch_down(self, touch):
print(touch.x, touch.y)
# 要在屏幕上画画,我们就要实现`Widget.canvas`属性。Kivy的`canvas`属性是一个底层为OpenGL的可绘制层,不过没有底层图形API那么复杂,`canvas`可以持续保留我们画过的图。
#
# 基本图形如圆(Color),线(Line), 矩形(Rectangle),贝塞尔曲线(Bezier),可以通过`kivy.graphics`导入。
# ### canvas简介
# `Canvas`的API可以直接调用,也可以通过上下文关联`with`关键字调用。如下所示:
self.canvas.add(Line(circle=(touch.x, touch.y, 25)))
# 这里的`Line`元素的参数是图形命令队列。
#
# >如果你想立刻试验代码,请先看下一节**屏幕显示触摸轨迹**中更完整的例子。
#
# 通过上下文关联with关键字调用可以让代码更简练,尤其是在同时操作多个指令时。下面的代码与之前一致:
with self.canvas:
Line(circle=(touch.x, touch.y, 25))
# 需要注意的是,如前面所说,canvas上后面调用的指令不会覆盖前面调用的指令;因此,canvas是一个不断增长的数组,里面都是不断显示元素的指令,更新频率60fps,但是也不能让canvas无限增长下去。
#
# 例如,所见即所得的程序(如HTML5的`<canvas>`)里有一条设计规则就是通过背景色填充擦除之前的图像。在浏览器里面可以很直观的写出:
#
# ```JavaScript
# // JavaScript code for clearing the canvas
# canvas.rect(0, 0, width, height)
# canvas.fillStyle = '#FFFFFF'
# canvas.fill()
# ```
#
# 在Kivy设计中,这种模型也是增加指令;首先获取前面所有的图形元素,然后把它们画成矩形。这个看着挺好其实不对:
# 看着和avaScript代码一样,但是错了。
with self.canvas:
Color(1, 1, 1)
Rectangle(pos=self.pos, size=self.size)
# >和内存泄露差不多,这个bug很久没被发现,使代码冗余,性能降低。由于显卡加速的功能,包括智能手机运行速度都很快。所以很难意识到这是一个bug。为了清除Kivy的canvas,应该用`canvas.clear()`来清除所有指令,后面会介绍。
# ### 屏幕显示触摸轨迹
# 我们马上做一个按钮来清屏;现在让我们把触摸的轨迹显示出来。让我们把`print()`删掉,然后增加一个方法在`CanvasWidget`下面:
class CanvasWidget(Widget):
def on_touch_down(self, touch):
with self.canvas:
Color(*get_color_from_hex('#0080FF80'))
Line(circle=(touch.x, touch.y, 25), width=4)
# 这样就每次都会画一个空心圆在画布上。`Color`指令为`Line`取色。
# > 注意`hex('#0080FF80')`并不是CSS颜色格式,因为它有四个组成部分,表示alpha值,即透明度。类似于`rgb()`与`rgba()`的区别。
#
# 可能你会觉得奇怪,我们用`Line`画的是圈,而不是直线。Kivy的图形元素具体很强的自定义功能,比如我们可以用`Rectangle`和`Triangle`画自定义的图片,用`source`参数设置即可。
#
# 前面的程序效果如下图所示:
# 
# 画图app完整的代码如下:
# +
# In main.py
from kivy.app import App
from kivy.config import Config
from kivy.graphics import Color, Line
from kivy.uix.widget import Widget
from kivy.utils import get_color_from_hex
class CanvasWidget(Widget):
def on_touch_down(self, touch):
with self.canvas:
Color(*get_color_from_hex('#0080FF80'))
Line(circle=(touch.x, touch.y, 25), width=4)
class PaintApp(App):
def build(self):
return CanvasWidget()
if __name__ == '__main__':
Config.set('graphics', 'width', '400')
Config.set('graphics', 'height', '400')
Config.set('input', 'mouse', 'mouse,disable_multitouch')
from kivy.core.window import Window
Window.clearcolor = get_color_from_hex('#FFFFFF')
PaintApp().run()
# -
# 这里没有加入鼠标光标显示的部分。`paint.kv`文件也没有了,用`build()`方法返回根部件。
#
# 注意`from kivy.core.window import Window`行,是由于有些模块有副作用,所有放在后面导入。`Config.set()`应该放在任何有副作用模块的前面。
#
# 下面,我们增加一些特性,让画图app实现我们想要的功能。
# ## 清屏
# 到目前为止,我们清屏的做法就是重启程序。下面我们增加一个按钮来清屏。我们用上一章时钟app的按钮即可,没什么新鲜,有意思的是位置。
#
# 上一章时钟app里面,我们没有讨论过位置,所有部件都放在`BoxLayouts`里面。现在我们的app没有任何布局,因为根部件就是`CanvasWidget`,我们没有实现任何子部件的位置。
#
# 在Kivy里面,布局部件缺失表示每一个部件都可以随意设置位置和大小(类似的UI设计工具,如Delphi,Visual Basic等等都如此)。
#
# 要让清屏按钮放在右上角,我们这么做:
#
# ```yaml
# # In paint.kv
# <CanvasWidget>:
# Button:
# text: 'Delete'
# right: root.right
# top: root.top
# width: 80
# height: 40
# ```
#
# 按钮的`right`和`top`属性与根部件的属性一致。我们还可以进行数学运行,如`root.top – 20`。结果很直接,`right`和`top`属性都是绝对值。
#
# 注意我们定义了一个`<CanvasWidget>`类却没有指定父类。这么做可以是因为我们在Python代码理论已经定义了一个同样的类。Kivy允许我们扩展所有的类,包括内部类,如`<Button>`和`<Label>`,以及自定义类。
#
# 这里体现了Kivy语言描述对象的可视化属性的一个好思路,类似于MVC设计方法,让内容与逻辑分离。同时,也更好的保持了所有Python程序的结构不变。这种Python代码与Kivy语言分离的思想让程序更容易维护。
# ### 传递事件
# 如果你跟着教程看到现在,准备去按清屏键。你会发现没反应,因为还没有增加事件,所有没有反馈。所有单击按钮不会有动作,相反会在画布上留下空心圈。
#
# 因为所有的触摸都是发生在`CanvasWidget.on_touch_down`上,并没有传递给其他子部件,所以清屏按钮没反应。不像HTML的DOMDOM,Kivy事件不会从嵌套的元素升级为父元素显示出来。它们走另一条路,如果事件传递到父元素没有反应,才从父元素下降到子元素。
#
# 最直接的方式就是这样:
# 注意:不是最优代码
def on_touch_down(self, touch):
for widget in self.children:
widget.on_touch_down(touch)
# 实际上,`Widget.on_touch_down`的默认行为有很多,所有我们可以直接调用,让代码更简练。
def on_touch_down(self, touch):
if Widget.on_touch_down(self, touch):
return
# 如果事件被正常处理了,`on_touch_down`这个handler返回`True`。触摸按钮会返回`True`是因为按钮响应了,然后很快的改变其外观。这就是为了取消我们的事件处理需要做的事情,当我们画圈的时候,方法的第二个行就`return`。
# ### 清屏
# 现在我们回到清屏按钮上。其实很简单,就是下面两行:
def clear_canvas(self):
self.canvas.clear()
# 别忘了把事件绑定到`paint.kv`文件:
#
# ```yaml
# Button:
# on_release: root.clear_canvas()
# ```
#
# 这样就可以清屏了,同时还把按钮也清除了。因为`CanvasWidget`是根部件,按钮是子部件。按钮部件本身没有被删除,它的画布`Button.canvas`从`CanvasWidget.canvas.children`层级中移除了,因此不存在了。
#
# 要保留按钮,可以这样:
def clear_canvas(self):
self.canvas.clear()
self.canvas.children = [widget.canvas
for widget in self.children]
# 但是这么做不够好,因为不同的部件初始化和运行方式不同。更好的做法是:
#
# 1. 从`CanvasWidget`部件中删除所有子部件;
# 2. 然后清除画布;
# 3. 最后再重新增加子部件,这样它们就可以正确的初始化了。
#
# 这个版本有点长,但是更合理:
class CanvasWidget(Widget):
def clear_canvas(self):
saved = self.children[:]
self.clear_widgets()
self.canvas.clear()
for widget in saved:
self.add_widget(widget)
# 解释一下`saved = self.children[:]`语句。`[:]`操作符是复制数组(就是“创建一个元素相同的数组”)。如果我们写`saved = self.children`,那就会从` self.children `和`saved`同时删除所有子部件。因为Python赋值是引用,与Kivy无关。
# > 如果想进一步了解Python的特性,可以看看[StackOverflow](http://stackoverflow.com/
# questions/509211)
#
# 现在,我们已经可以用蓝色的圈钱画图了,如下所示。这当然并非最终版,请看下面的内容。
# 
# ## 连点成线
# 我们的app已经可以清屏了,不过只能画圈。下面在改进一下。
#
# 要保持连续触控画线(按住然后拖拽),我们需增加一个监听器,`on_touch_move`。每次使用都会收到最新点的位置。
#
# 如果我们一次只有一条线,我们可以把这条线保存为`self.current_line`。但是,由于这是多点触控,我们就要用其他方法来保存`touch`变量了。
#
# 之所以能实现这些,是因为每个触控自始至终都访问相同的`touch`对象。还有一个`touch.ud`属性,是一个字典类型,`ud`就是用户数据(user data),可以灵活的跟踪所有的触控。初始值为空字典`{}`。
#
# 下面我们要做的是:
# - 在`on_touch_down`的handler创建一个新线,然后储存到`touch.ud`。现在我们要用直线来代替空心圈。
# - 在`on_touch_move`里面增加一个新点到线的末尾。我们增加的是直线元素,但是事件处理过程是每秒调用很多次实现这条线,每次都很短,最终看起来就很平滑。
#
# >更先进的图形程序可以用复杂的算法让线条呈现的更真实。包括贝塞尔曲线实现线条的高分辨率的无缝连接,并且从点的速度和压力推断线的厚度。这些具体的技术我们不打算实现了,不过读者可以作为一个练习。
#
# 上述过程的代码如下:
from kivy.graphics import Color, Line
class CanvasWidget(Widget):
def on_touch_down(self, touch):
if Widget.on_touch_down(self, touch):
return
with self.canvas:
Color(*get_color_from_hex('#0080FF80'))
touch.ud['current_line'] = Line(
points=(touch.x, touch.y), width=2)
def on_touch_move(self, touch):
if 'current_line' in touch.ud:
touch.ud['current_line'].points += (
touch.x, touch.y)
# 这样就可以画线了。之后让我们来实现颜色选择功能,不断的完善我们的画图app。
# ## 调色板
# 画图app当然不能没有调色板。调色板其实就是可选颜色列表,可以让颜色选取很简单。通过图像编辑器都有调色板,带有全真彩24位色16,777,216种。如下图所示:
# 
# 但是,就是你不打算完成一个主流的图像编辑器,我们也打算限制颜色的种类。因为对那些没有色彩常识的人来说,放一堆颜色只会让人头大。而且,互联网上的UI设计用色也会逐渐统一。
#
# 在我们的app中,我们打算使用[扁平化的UI设计风格](http://designmodo.github.io/Flat-UI/),基于一列精心挑选的颜色。当然,你可以选自己喜欢的颜色,因人而异。
#
# >颜色是一门学问,尤其是具体任务的兼容性与稳定性。低对比度的组合可能用来装饰元素或者标题,但是它们不符合正文的风格;另外,高对比度的颜色,如白与黑,不容易吸引注意力。
#
# >因此,颜色使用的首要原则是除非你很专业,否则用别人调好的颜色。最好的起点就是操作系统的用色。一些精彩案例如下:
# - [Tango调色板](http://tango.freedesktop.org/Tango_Icon_Theme_Guidelines),在Linux开源环境中使用广泛。
# - Google在2014年GoogleIO大会上发布的[Material design](https://www.google.com/design/material-design.pdf)。
# - 非官方的[iOS 7颜色风格](http://ios7colors.com/),超赞。
#
# >还有很多调色板可以学习,自行Google之。
# ### 按钮的子类
# 因为我使用的颜色很少,所以用单选按钮就可以了。Kivy的`ToggleButton`可以实现功能,不过有个限制:在一个单选组内,所有的按钮可以同时不选。也就是说,画图的时候可能没颜色。当然我们也可以设定默认颜色,但是用户可能会觉得很奇怪,所有我们不打算这么用。
#
# Python的OOP模式可以很好的解决这个问题,我们可以继承`ToggleButton`类,然后改造它的功能。之后,每次都会有一个颜色被选中了。
#
# 子类还会实现另外一个功能:在调色板上,我们想让每个颜色按钮有唯一颜色。我们可以用之前的技术为每个按钮分配背景色,那就要一堆代码来分配。但是,我们如果写一个背景色属性,就可以在`paint.kv`文件里面分配了。
#
# 这样就可以在`paint.kv`文件中使用按钮时保持调色板定义的可读性,同时在子类中实现的具体的细节——会展示OOP程序应该怎样实现。
# ### 去掉全不选功能
# 首先,让我们把全不选的功能去掉。
#
# 首先,让我们实现一个标准的`ToggleButton`部件。我们之间在`paint.kv`文件里面增加如下代码:
#
# ```yaml
# BoxLayout:
# orientation: 'horizontal'
# padding: 3
# spacing: 3
# x: 0
# y: 0
# width: root.width
# height: 40
#
# ToggleButton:
# group: 'color'
# text: 'Red'
#
# ToggleButton:
# group: 'color'
# text: 'Blue'
# state: 'down'
# ```
#
# 我们用了与`BoxLayout`类似的方式,每个颜色按钮单独分配一个工具栏。布局部件本文的位置是绝对的,其`x`和`y`的值都是0,也就是左下角,宽度与`CanvasWidget`一致。
#
# 每个`ToggleButton`都属于同一`color`组。因此同一时间只有一个颜色可以被选中。
# #### 改写标准行为
# 要实现改写,让我们定义`ToggleButton`子类:
# +
from kivy.uix.behaviors import ToggleButtonBehavior
from kivy.uix.togglebutton import ToggleButton
class RadioButton(ToggleButton):
def _do_press(self):
if self.state == 'normal':
ToggleButtonBehavior._do_press(self)
# -
# 这样当按下按钮,状态`'normal'`就会变成`'down'`。
#
# 现在我们把`paint.kv`文件里面`ToggleButton`改成`RadioButton`,立刻就会看到不同。
#
# 这也是Kivy框架最吸引人的地方:小代码实现大功能。
# >要在Kivy语言中使用`RadioButton`,其定义需要在导入`main.py`文件。由于现在只有一个Python文件,这并不重要,但是一定记住:自定义的Kivy部件,和其他的Python类和函数一样,需要在使用之前被导入。
# ### 彩色按钮
# 现在按钮的功能正常了,我们把彩色按钮都做出来。如下图所示:
# 
# 要实现这些,我们得用`background_color`属性。Kivy的背景色不仅可以使用单一颜色,可以用彩色;我们首先需要一个纯白色背景,然后画上想要的颜色。这样我们就只要为任意数量的彩色按钮准备两种模式(正常和按下的)即可。
#
# 这和第一章时钟app是一样的。除了按钮的中心区域允许着色,选中的状态有个黑边。
# 
# #### 新按钮
# 加油!我们就快完工了,在`paint.kv`里面加入新类`ColorButton`:
# ```yaml
# <ColorButton@RadioButton>:
# group: 'color'
# on_release: app.canvas_widget.set_color(self.background_color)
# background_normal: 'color_button_normal.png'
# background_down: 'color_button_down.png'
# border: (3, 3, 3, 3)
# ```
# 你会发现,我们把`group: 'color'`移到这里避免重复代码。
#
# 我们还要配置`on_release`事件handler,作用于已经被选中的按钮。现在,每个按钮已经把自己的`background_color`属性传递给事件handler,剩下的事情就是把颜色分配给画布。这个事件将由`CanvasWidget`处理,需要通过`PaintApp`类显示出来。
class PaintApp(App):
def build(self):
# set_color()方法后面实现
self.canvas_widget = CanvasWidget()
self.canvas_widget.set_color(
get_color_from_hex('#2980B9'))
return self.canvas_widget
# 这么配置的原因是我们不能在`paint.kv`文件的类定义中使用`root`;因为那样会指向`ColorButton`自身(类规则里面的根定义在`paint.kv`文件的顶层)。我们还可以设置默认颜色,就像代码里显示的。
#
# 在`main.py`文件里面,让我们来实现`CanvasWidget`的`set_color()`方法,可以当作是`ColorButton`的事件handler。代码很简单,就是把颜色作为参数:
def set_color(self, new_color):
self.canvas.add(Color(*new_color))
# #### 定义调色板
# 下面我们来定义调色板。首先让我们把`RadioButton`从`paint.kv`文件中删掉。
#
# 为了使用CSS颜色定义方式,我们需要将适当的函数导入`paint.kv`文件。把下面这行代码放在`paint.kv`文件开头。
# +
#:import C kivy.utils.get_color_from_hex
# -
# 这行代码实际上和Python的代码一样:
from kivy.utils import get_color_from_hex as C
# 我们使用扁平化设计的配色方式,代码如下:
# ```yaml
# BoxLayout:
# # ...
# ColorButton:
# background_color: C('#2980b9')
# state: 'down'
#
# ColorButton:
# background_color: C('#16A085')
#
# ColorButton:
# background_color: C('#27AE60')
# ```
#
# 很简单吧,这样就为每个`ColorButton`按钮定义了`background_color`属性。其他的属性都是继承于Python中`ColorButton`类的定义。
#
# 这样,增加任意数量的按钮都可以很好的排列了。
# ## 设置线的宽度
# 最后一个,也是最简单的功能就是设置线条的宽度。如下图所示,我们可以重用前面调色板的资源和样式。
#
# 这个UI也是一种`RadioButton`子类,命名为`LineWidthButton`。在`paint.kv`文件中就是这样:
#
# ```yaml
# <LineWidthButton@ColorButton>:
# group: 'line_width'
# on_release: app.canvas_widget.set_line_width(self.text)
# color: C('#2C3E50')
# background_color: C('#ECF0F1')
# ```
#
# 与`ColorButton`不同之处在于第2、3行代码。这些按钮属于另外一组,由其他的事件handler触发。当然,这两组按钮依然很相似。
#
# 布局很简单,和调色板的样式一致,只是垂直摆放:
# ```yaml
# BoxLayout:
# orientation: 'vertical'
# padding: 2
# spacing: 2
# x: 0
# top: root.top
# width: 80
# height: 110
#
# LineWidthButton:
# text: 'Thin'
#
# LineWidthButton:
# text: 'Normal'
# state: 'down'
#
# LineWidthButton:
# text: 'Thick'
# ```
# >注意`CanvasWidget.set_line_width`事件监听器会接受宽度调节按钮的`text`属性。这样实现是为了简化,允许我们为每一个按钮定义一个唯一的宽度值。
# >实际开发中,这种方法固然无可厚非。但是,当我们要把文字翻译成日语或法语的时候,这种对应关系就丢失了。
# ### 改变线条宽度
# 让我们把前面做好的模块都组合起来,这样就可以控制线条的粗细了。我们把线条宽度存储在`CanvasWidget.line_width`变量中,与按钮的文字一一对应,然后用`on_touch_down`触发事件改变线条宽度。代码如下:
# +
class CanvasWidget(Widget):
line_width = 2
def on_touch_down(self, touch):
# ...
with self.canvas:
touch.ud['current_line'] = Line(
points=(touch.x, touch.y),
width=self.line_width)
def set_line_width(self, line_width='Normal'):
self.line_width = {
'Thin': 1, 'Normal': 2, 'Thick': 4
}[line_width]
# -
# 这样就完成Kivy的画图app了,开始画图吧。
# ## 总结
# 这一章,我们重点学习了Kivy应用开发中的一些方法,包括自定义窗口,改变鼠标光标,窗口大小,背景色,通过画布指令绘制自定义的图形,正确的处理支持多平台的触摸事件,并且考虑多点触控的情况。
#
# 在完成画图app之后,关于Kivy的一件显而易见的事情就是这个框架具有高度的开放性和通用性。不需要一大堆死板的组件,Kivy让开发者可以通过图形基本元素和行为的运用,让自定义模块变得简单灵活。也就是说,Kivy没有自带很多开箱即用的部件,但是通过几行Python代码就可以做出需要的东西。
#
# 模块化的API设计方法缺乏美感,因为它限制了设计的柔性。最终的结果完全的满足你对项目的需求。客户总想要一些爆点,比如三角形按钮——当然,你还可以为它增加质地,这些都可以两三行代码搞定。(假如你想用**WinAPI**做一个三角形按钮。那就真掉坑里了。)
#
# Kivy的自定义部件还可以重用。实际上,你可以把`main.py`的`CanvasWidget`模块导入其他应用。
# ## 自然用户界面
# 我们的第二个应用比第一个应用更具交互性。不仅是在按钮上,还有多点触控手势。
#
# 所有的窗口都支持触摸屏,对用户来说这是普遍共识,尤其在触摸屏设备上。只要用手指就可以绘画,好像在真实的画布上,即使手指很脏也可以上面画画。
#
# 这种界面被称为NUI(自然界面,natural user interface)。有一些有趣的特性:NUI应用可以被小朋友或者宠物使用——可以在屏幕上看到和触摸图形元素。这是一种自然、直观的界面,一种“不需要思考”的事情,与[Norton Commander](http://en.wikipedia.org/wiki/Norton_Commander)的反直觉截然不同。直觉不应该接受蓝屏、ASCII码的表现形式。
#
# 下一章,我们将建立另外一个Kivy程序,只能Android用。将Python与Android API的Java类很好的结合在一起。
| _notebooks/2019-03-01-kivy-ch2-paint-app.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Videocvičení naleznete zde: https://youtu.be/yL-A0N5JDJo
# # Práce s obrázky
#
# ### Načítání balíčků
# K práci s obrázky budeme používat knihovnu **cv2** s aliasem **cv**. Dále budeme používat knihovnu **numpy** s aliasem **np** pro matematické funkce a práci s poli a knihovnu **matplotlib** s aliasem **plt** pro vykreslování výsledků.
#
# Následujícím kódem naimportujeme balíčky.
import cv2 as cv
import numpy as np
from matplotlib import pyplot as plt
from matplotlib.colors import NoNorm
# Pro interaktivní vykreslování grafů v Jupyter notebooku ještě potřebujeme toto:
# %matplotlib notebook
# ## Obrázek jako matice
# Nejprve je třeba načíst obrázek do paměti. To provedeme příkazem **imread** z knihovny **cv2**.
#
# ***Pozor:*** Barevné vrstvy se načtou v pořadí **blue**, **green**, **red** namísto obvyklého **red**, **green**, **blue**. A protože jsme konzervativní, obrázek si převedeme.
img_bgr = cv.imread("lena_original.jpg",cv.IMREAD_UNCHANGED)
img = cv.cvtColor(img_bgr, cv.COLOR_BGR2RGB)
# Chceme-li si obrázek zobrazit, použijeme příkaz **imshow** z knihovny **matplotlib**.
plt.figure()
plt.imshow(img)
# Pro zkoušení algoritmů pro zpracování obrazu se běžně používá jen výřez tohoto obrázku. Ten si teď vyrobíme.
#
# S obrázkem teď můžeme zacházet jako s polem. Můžeme tedy zadat, jaký rozsah indexů chceme nadále používat:
img_crop = img[20:270,150:400,:]
# Výřez si můžeme zase prohlédnout a, protože jej budeme později používat, také uložit.
plt.figure()
plt.imshow(img_crop)
cv.imwrite('lena_crop.jpg',cv.cvtColor(img_crop, cv.COLOR_RGB2BGR))
# ### Barevné složky
# Chceme-li pracovat s některou z barevných složek obrázku, můžeme ji z obrázku získat např. tak, že vynulujeme ostatní složky.
# +
b = img_crop.copy()
g = img_crop.copy()
r = img_crop.copy()
r[:,:,1] = 0
r[:,:,2] = 0
g[:,:,0] = 0
g[:,:,2] = 0
b[:,:,0] = 0
b[:,:,1] = 0
# -
# Výsledné obrázky pak budou vypadat následnovně:
plt.figure()
plt.imshow(r)
plt.figure()
plt.imshow(g)
plt.figure()
plt.imshow(b)
# ### Základní úpravy obrázku
# V následující části cvičení si pro jednoduchost vystačíme s černobílým obrázkem. Načteme si výřez, který jsme si před chvílí uložili. Funkci **imread** ale řekneme, že chceme načíst obrázek jen v odstínech šedé pomocí **cv.IMREAD_GRAYSCALE**.
img_grey = cv.imread("lena_crop.jpg",cv.IMREAD_GRAYSCALE)
# Můžeme si jej vykreslit, ať víme, že vše proběhlo v pořádku. U černobílých obrázků budeme specifikovat, že chceme vykreslit v odstínech šedé pomocí **cmap='gray'** a také, že nechceme, aby během vykreslování došlo k automatickému vyrovnání histogramu (tím se budeme zabývat za chvilku). To se udělá pomocí **norm=NoNorm()**.
plt.figure()
plt.imshow(img_grey,cmap='gray',norm=NoNorm())
# <hr style="border:1px solid black"> </hr>
#
# ## Úkol 1:
# vytvořte funkci, která zadaný obrázek zesvětlí o zadaný počet odstínů. Zesvětlený obrázek vykreslete.
#
# <hr style="border:1px solid black"> </hr>
def lighten(img, amount):
for i in range(img.shape[0]):
for j in range(img.shape[1]):
if(img[i,j]+amount>254):
img[i,j]=255
else:
img[i,j] += amount
return img
plt.figure()
plt.imshow(lighten(img_grey.copy(),100),cmap='gray',norm=NoNorm())
# <hr style="border:1px solid black"> </hr>
#
# ## Úkol 2:
# vytvořte funkci, která zadaný obrázek ztmaví o zadaný počet odstínů. Tmavší obrázek vykreslete.
#
# <hr style="border:1px solid black"> </hr>
def darken(img, amount):
for i in range(img.shape[0]):
for j in range(img.shape[1]):
if(img[i,j]-amount<1):
img[i,j] = 0
else:
img[i,j] -= amount
return img
plt.figure()
plt.imshow(darken(img_grey.copy(),100),cmap='gray',norm=NoNorm())
# <hr style="border:1px solid black"> </hr>
#
# ## Úkol 3:
# vytvořte funkci, která vytvoří inverzi (negativ) zadaného obrázku. Negativ vykreslete.
#
# <hr style="border:1px solid black"> </hr>
def invert(img):
for i in range(img.shape[0]):
for j in range(img.shape[1]):
img[i,j] = 255-img[i,j]
return img
plt.figure()
plt.imshow(invert(img_grey.copy()),cmap='gray',norm=NoNorm())
# ### Prahování
# Prahování spočívá v tom, že se všechny hodnoty obrázku, které jsou menší než námi zvolený práh (threshold), nastaví na černou, zatímco se zbývanící hodnoty nastaví na bílou.
# <hr style="border:1px solid black"> </hr>
#
# ## Úkol 4:
# vytvořte funkci, která provede prahování obrázku pomocí zadané hodnoty. Výsledný obrázek vykreslete.
#
# <hr style="border:1px solid black"> </hr>
def threshold(img, threshold):
for i in range(img.shape[0]):
for j in range(img.shape[1]):
if(img[i,j]<threshold):
img[i,j]=0
else:
img[i,j]=255
return img
plt.figure()
plt.imshow(threshold(img_grey.copy(),100),cmap='gray',norm=NoNorm())
# ## Vyrovnání histogramu
# ### Získání histogramu
# Nejprve si obrázek načteme. Nezapomeňte, že jej chceme v odstínech šedé.
img_uneq = cv.imread("uneq.jpg", cv.IMREAD_GRAYSCALE)
# Provedeme kontrolu vykreslením.
plt.figure()
plt.imshow(img_uneq,cmap='gray',norm=NoNorm())
# Vidíme, že s obrázkem něco není v pořádku. Zkusíme se podívat na jeho histogram. Histogram obrázku získáme tak, že pro každou možnou hodnotu jasu (v obrázku jich je dohromady 256, 0 reprezentuje černou, 255 bílou) spočítáme, kolikrát se v obrázku vyskytuje.
# <hr style="border:1px solid black"> </hr>
#
# ## Úkol 5:
# vytvořte funkci, která k zadanému obrázku vytvoří histogram a použijte ji na obrázek **img_uneq**. Výsledný histogram vykreslete.
#
# ***Nápověda***: k vykreslení histogramu se hodí použít sloupcový graf. Ten získáme pomocí příkazu **bar** z knihovny **matplotlib**.
#
# <hr style="border:1px solid black"> </hr>
# +
def get_hist(image):
hist = np.zeros(256)
for i in range(image.shape[0]):
for j in range(image.shape[1]):
hist[int(image[i,j])] += 1
return hist
hist_uneq = get_hist(img_uneq)
plt.figure()
plt.bar([i for i in range(hist_uneq.shape[0])],hist_uneq/float(sum(hist_uneq)))
# -
# ### Vyrovnání histogramu
# Z obrázku vidíme, že v obrázku je jen úzký rozsah jasu. Pokusíme se toto napravit a histogram tzv. vyrovnáme. Cílem je celý histogram roztáhnout tak, aby pokrýval celý rozsah of 0 do 255.
# <hr style="border:1px solid black"> </hr>
#
# ## Úkol 6:
# vytvořte funkci, která vyrovná histogram zadaného obrázku a použijte ji na obrázek **img_uneq**. Výsledný obrázek a histogram vykreslete.
#
# <hr style="border:1px solid black"> </hr>
# +
def eq_hist(img):
hist = get_hist(img)
i=0
while(hist[i]==0):
i+=1
j=255
while(hist[j]==0):
j-=1
for k in range(img.shape[0]):
for l in range(img.shape[1]):
img[k,l] = (img[k,l]-i)*(255.0/(j-i))
return img
img_eq = eq_hist(img_uneq.copy())
plt.figure()
plt.imshow(img_eq,cmap='gray',norm=NoNorm())
hist_eq = get_hist(img_eq)
plt.figure()
plt.bar([i for i in range(hist_eq.shape[0])],hist_eq/float(sum(hist_eq)))
# -
# ## Konvoluce
# Diskrétní konvoluce je operace, která obrázek modifikuje pomocí takzvané konvoluční masky. Konvoluční masku si můžeme představit jako čtvercovou matici, jejíž hodnoty představují váhy, s jakými do výsledného obrázku započítáváme hodnoty jasu obrázku původního.
#
# V praxi konvoluce funguje tak, že masku přiložíme na obrázek tak, aby její střed byl v bodě, pro který chceme konvoluci počítat. Hodotu pak získáme tím, že po složkách vynásobíme masku s jasy obrázku a vše sečteme. Obrázek se schématem najdete na Wikipedii, ze které jsem si jej vypůjčil.
# 
# Naším úkolem nebude nic jiného, než diskrétní konvoluci naprogramovat a otestovat s různými maskami. Ty si ostatně můžeme rovnou zadefinovat.
average = np.array([[1, 1, 1],[1, 1, 1],[1, 1, 1]])
gauss_large = np.array([[1, 4, 7, 4, 1],[4, 16, 26, 16, 4],[7, 26, 41, 26, 7],[4, 16, 26, 16, 4],[1, 4, 7, 4, 1]])
gauss = np.array([[1, 2, 1],[2, 4, 2],[1, 2, 1]])
laplace = np.array([[0, 1, 0],[1, -4, 1],[0, 1, 0]])
edges = np.array([[0, -1, 0],[-1, 5, -1],[0, -1, 0]])
vertical_edges = np.array([[-1, 0, 1],[-2, 0, 2],[-1, 0, 1]])
horizontal_edges = np.array([[-1, -2, -1],[0, 0, 0],[1, 2, 1]])
# Různé masky se hodí na různé operace, proto si je vyzkoušíme na dvou různých obrázcích. Jeden z nich bude zašuměná Lena
lena = cv.imread("lena_noise.jpg", cv.IMREAD_GRAYSCALE)
# Druhý obrázek bude obrázek cihlové zdi, na kterém bude dobře vidět efekt masek, které zvýrazňují hrany.
bricks = cv.imread("bricks.jpg", cv.IMREAD_GRAYSCALE)
# <hr style="border:1px solid black"> </hr>
#
# ## Úkol 7:
# vytvořte funkci, která provede diskrétní konvoluci zadaného obrázku se zadanou konvoluční maskou. Otestujte efekty různých konvolučních masek.
#
# <hr style="border:1px solid black"> </hr>
def convolution(img, mask):
conv = np.zeros(img.shape)
l = int(mask.shape[0]/2)
for i in range(l ,img.shape[0]-l):
for j in range(l,img.shape[1]-l):
conv[i,j] = int(sum(sum(img[(i-l):(i+l+1),(j-l):(j+l+1)]*mask)))
return conv
img = bricks
mask = edges
img_conv = convolution(img.copy(),mask)
plt.figure()
plt.imshow(img,cmap='gray')
plt.figure()
plt.imshow(img_conv,cmap='gray')
| CV2/cv2_obrazky_reseni.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In this and following section 6 notebooks. I want to begin to use this model and daily and track its performance over time. Since we have streamlined the prediction process in section 4, we can now use similar methods to create a database of predictions on a daily basis. Using some other python libraries and methods I will also create a script that will show us a visual representaton of how the model has been performing over time, its current accuracy metric performance, as well as potential return on investment if we were to deploy this model to execute trades (and this functionality also exists within the pyhthon-binance libray!)
# +
# the following below is copied from our predictor.py script. It will collect transform and input daily data
# and give a printout about predicted price movement direction and level of confidence. Taking this script and
# modifying the results to our desired outputs should not take much
# import all of our required libraries for necessary data processing and data requests
import numpy as np
import pandas as pd
from binance.client import Client
import joblib
from sqlite3 import Error
# define our function to retrieve klines data from binance API
def get_data():
'''
This function will execute API call to Binance to retrieve data.
We will export the results of this data into the appropriately named dataframe for further feature engineering.
'''
client = Client()
# establishing our blank client
candles = client.get_klines(symbol='BTCUSDT', interval=Client.KLINE_INTERVAL_1DAY, limit=91)
# we only need to request the most recent 90 days to calculate our prediction data
data = pd.DataFrame(candles, columns=['Date', 'Open', 'High', 'Low', 'Close', 'Volume', 'Close time', 'Quote asset volume', 'Number of trades', 'Taker buy base volume', 'Taker buy quote volume', 'Ignore'])
# these column labels are as labelled on the Binance API documentation
data.drop(['Close time', 'Ignore'], axis=1, inplace=True)
# dropping unneeded columns
data['Date'] = data['Date'].apply(lambda x: pd.to_datetime(x, unit='ms'))
# converting to proper date format for better visual reference
data.set_index('Date', inplace=True)
# setting index to date
data = data.astype('float64')
# converting from object type to float type
return data
# we will define a function to run prior to calcualting our averages
def feat_eng(X_df):
'''
Intakes "X" portion of data and outputs selected engineered features
'''
X_df['High/Low'] = X_df['High'] - X_df['Low']
X_df['volX'] = X_df['Quote asset volume'] / X_df['Volume']
X_df['quote-buy'] = X_df['Taker buy quote volume'] / X_df['Taker buy base volume']
return X_df
# lets define a function to create our moving averages and incoroprate them into our dataframe
def get_sma(X_df):
'''
This function intakes the "X" portion of the data and returns the data with moving average columns applied
'''
SMAs = [7,30,90] # 7, 30, and 90 day simple moving averages
for val in SMAs:
X_df[str(val)+'sma'] = X_df['Close'].rolling(f'{val}D').mean() # using the pandas rolling function to calculate mean values over each desired SMA value
return X_df
# Now we want to take the most recent data point possible to make our prediction from
def X_input(X_df):
x_input = X_df[-1:] # take the most recent value after calculations for passing into model
return x_input
# now to create a function that ties all of these together and gives us our desired input for the model
def to_predict():
data = get_data()
data_features = feat_eng(data)
data_all = get_sma(data_features)
x_input = X_input(data_all)
return x_input
'''
This now gives us all functions and libraries needed to create our input for the model to predict.
'''
X = to_predict()
# now we must load our saved model using pickle
with open("final_model.pkl", "rb") as file:
model = joblib.load(file)
predicted = model.predict_proba(X)
# all of this is no longer needed, instead we are going to be using the output of the prediction
# this will be used to add to our data intaken from the binance API and to create our database table
# following database creation, we will then define pathways where the model will intake data from our created database,
# and use that data to create visual representations of model performance over time
# if (predicted[0][0] < predicted[0][1]) & (predicted[0][1] > 0.6):
# print(f'*********************\n\n\n\nThe price of Bitcoin is predicted to go UP tomorrow!\nI am quite confident about this!\nAt this confidence I am correct {53/(53+36)*100:.2f}% of the time!\n\n\n\nThis is not finanical advice. I am not a financial advisor. All information here is for entertainment purposes only.\n\n\n\n*********************')
# elif (predicted[0][0] < predicted[0][1]) & (predicted[0][1] > 0.55):
# print(f'*********************\n\n\n\nThe price of Bitcoin is predicted to go UP tomorrow!\nI am sort of confident about this!\nAt this confidence I am correct {36/(36+27)*100:.2f}% of the time!\n\n\n\nThis is not finanical advice. I am not a financial advisor. All information here is for entertainment purposes only.\n\n\n\n*********************')
# elif (predicted[0][1] < predicted[0][0]) & (predicted[0][0] > 0.6):
# print(f'*********************\n\n\n\nThe price of Bitcoin is predicted to go DOWN tomorrow!\nI am quite confident about this!\nAt this confidence I am correct {61/(61+44)*100:.2f}% of the time!\n\n\n\nThis is not finanical advice. I am not a financial advisor. All information here is for entertainment purposes only.\n\n\n\n*********************')
# elif (predicted[0][1] < predicted[0][0]) & (predicted[0][0] > 0.55):
# print(f'*********************\n\n\n\nThe price of Bitcoin is predicted to go DOWN tomorrow!\nI am sort of confident about this!\nAt this confidence I am correct {38/(38+31)*100:.2f}% of the time!\n\n\n\nThis is not finanical advice. I am not a financial advisor. All information here is for entertainment purposes only.\n\n\n\n*********************')
# elif predicted[0][0] < predicted[0][1]:
# print(f'*********************\n\n\n\nThe price of Bitcoin is predicted to go UP tomorrow!\nI am not very confident about this!\nAt this confidence I am correct {67/(67+53)*100:.2f}% of the time!\n\n\n\nThis is not finanical advice. I am not a financial advisor. All information here is for entertainment purposes only.\n\n\n\n*********************')
# elif predicted[0][1] < predicted[0][0]:
# print(f'*********************\n\n\n\nThe price of Bitcoin is predicted to go DOWN tomorrow!\nI am not very confident about this!\nAt this confidence I am correct {44/(44+36)*100:.2f}% of the time!\n\n\n\nThis is not finanical advice. I am not a financial advisor. All information here is for entertainment purposes only.\n\n\n\n*********************')
# else:
# pass
# +
# our "to_predict" function gives us our desired output for prediction.
X
# -
model.predict_proba(X)[0]
# +
# now to create a function that will intake our "to_predict variable", and give us an output of that same dataframe
# having 2 new columns, predicted label and confidence of prediction
def add_prediction(X_df):
pred = model.predict_proba(X_df)[0] # this gives us our predictor array of confidence
# create our new columns based on prediction output
X_df['Prediction'] = 1 if pred[1] > pred[0] else 0 # predicted class based on higher confidence
X_df['Confidence'] = pred[1] if pred[1] > pred[0] else pred[0] # confidence score (probability) of larger class
return X_df # output modified X_df dataframe to be exported into database
# +
# test run of new function
add_prediction(X)
# -
# Function works exactly as intended. New columns work perfectly.
# Next steps will be to create a function that will evaluate previous daily predictions and append the coorect result to our dataframe. This will allow us to track accuracy moving forward.
def eval_prediction(X_yesterday, X_df):
'''
This function will intake our modified X dataframe from the previous day as well as our current prediction
and output a new column which gives the correct label, as well as if the model predicted correctly or not.
'''
X_yesterday['True_Label'] = 1 if X_df['Close'].values > X_yesterday['Close'].values else 0 # this gives us the correct label
X_yesterday['Correct_Pred'] = 1 if X_yesterday['Prediction'].values == X_yesterday['True_Label'].values else 0 # this gives a 1 for a correct prediction and a 0 for incorrect
return X_yesterday
# +
# creating a function which will pull the previous days data instead of today
def X_input_yesterday(X_df):
x_input = X_df[-2:-1] # take the most recent value after calculations for passing into model
return x_input
# now to create a function that ties all of these together and gives us our desired input for the model
def to_predict_yesterday():
data = get_data()
data_features = feat_eng(data)
data_all = get_sma(data_features)
x_input_yesterday = X_input_yesterday(data_all)
return x_input_yesterday
# -
X_yesterday = to_predict_yesterday()
X_yesterday
# +
# now we will use our new functions to add predictions to yesterdays data, and then attempt to evaluate its performance
X_yesterday = add_prediction(X_yesterday)
# -
X_yesterday
# +
# now to test evaluation
X_evaluated = eval_prediction(X_yesterday, X)
# -
# Now we have function which can intake our desired data and give us an evaluation of the previous prediction.
# The next step will be to create a function will connect to and update a database daily with our new evaluated data. From this database we can then access data and create a new set of functions which will display performance over time and quantify our gains and losses.
import sqlite3
# +
# defining connection and cursor
connection = sqlite3.connect('predictor_data.db')
cursor = connection.cursor()
# create our inital table
command1 = """CREATE TABLE IF NOT EXISTS model_data (
id integer AUTO_INCREMENT PRIMARY KEY,
date varchar(255),
open float,
high float,
low float,
close float,
volume float,
QaV float,
trades integer,
takerBase float,
takerQuote float,
highLow float,
volX float,
quoteBuy float,
sevensma float,
thirtysma float,
ninetysma float,
Pred integer,
Conf float,
True integer,
Correct integer);"""
cursor.execute(command1)
# +
#not needed
# # now we need to create a function that will connect to our database
# # and input our desired data for later retrieval
# def store_data(X_evaluated):
# '''
# This function will intake the fully modified version of our data including correct
# predictions and all info.
# It will write this data to our newly created database.
# '''
# connection = sqlite3.connect('predictor_data.db')
# cursor = connection.cursor()
# command = f"""
# INSERT INTO model_data (
# date, open, high, low, close, volume, QaV, trades, takerBase,
# takerQuote, highLow, volX, quoteBuy, sevensma, thirtysma,
# ninetysma, Pred, Conf, True, Correct
# )
# VALUES (
# {str(X_evaluated.index[0]).split(' ')[0]},
# {X_evaluated['Open'].values[0]},
# {X_evaluated['High'].values[0]},
# {X_evaluated['Low'].values[0]},
# {X_evaluated['Close'].values[0]},
# {X_evaluated['Volume'].values[0]},
# {X_evaluated['Quote asset volume'].values[0]},
# {X_evaluated['Number of trades'].values[0]},
# {X_evaluated['Taker buy base volume'].values[0]},
# {X_evaluated['Taker buy quote volume'].values[0]},
# {X_evaluated['High/Low'].values[0]},
# {X_evaluated['volX'].values[0]},
# {X_evaluated['quote-buy'].values[0]},
# {X_evaluated['7sma'].values[0]},
# {X_evaluated['30sma'].values[0]},
# {X_evaluated['90sma'].values[0]},
# {X_evaluated['Prediction'].values[0]},
# {X_evaluated['Confidence'].values[0]},
# {X_evaluated['True_Label'].values[0]},
# {X_evaluated['Correct_Pred'].values[0]}
# );"""
# cursor.execute(command)
# cursor.close()
# -
store_data(X_evaluated)
X_evaluated['Open'].values[0]
str(X_evaluated.index[0]).split(' ')[0]
# When troubleshooting, came across a pandas extension that will write our data to database for us. Going to attempt to create the function using this instead as it will hopeully bypass some formatting issues.
#
#
# +
# define a new function to intake our X_evaluated dataframe and write it to our new
# SQL database for future use
def to_database(X_evaluated):
'''
This function takes in our fully evaluated predictions and writes them
to an SQL database for further reference.
'''
conn = None
try:
conn = sqlite3.connect('bitcoin_model.db')
print('Connected Successfully!')
except Error as e:
print(e)
X_evaluated.to_sql('predictions', con=conn, if_exists='append')
conn.close()
# -
to_database(X_evaluated)
# That works great! Now we just need to combine all of these things together and we can create a constantly updating database of all of our relevant predictions!
| 6-1_Model-Tracking-and-Database-Creation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
def hyphen_range(s):
""" Takes a range in form of "a-b" and generate a list of numbers between a and b inclusive.
Also accepts comma separated ranges like "a-b,c-d,f" will build a list which will include
Numbers from a to b, a to d and f"""
s="".join(s.split())#removes white space
r=set()
for x in s.split(','):
t=x.split('-')
if len(t) not in [1,2]: raise SyntaxError("hash_range is given its arguement as "+s+" which seems not correctly formated.")
r.add(int(t[0])) if len(t)==1 else r.update(set(range(int(t[0]),int(t[1])+1)))
l=list(r)
l.sort()
return l
list(hyphen_range('6949-6960'))
# +
ATuple = (1,2,3)
ALst = [1,2,3]
| Code_FractBias/iPythonFractBias/.ipynb_checkpoints/Metadata scratch pad-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from geoscilabs.dcip.DC_cylinder import cylinder_app
from IPython.display import display
# %matplotlib inline
# # Purpose
#
# For a direct current resistivity (DCR) survey, currents are injected to the earth, and flow.
# Depending upon the conductivity contrast current flow in the earth will be distorted, and these changes
# can be measurable on the sufurface electrodes.
# Here, we focus on a cylinder target embedded in a halfspace, and investigate what are happening in the earth when static currents are injected. Different from a sphere case, which is a finite target, "coupling" among Tx, target (conductor or resistor), and Rx will be significanlty different upon various scenarios.
# By investigating changes in currents, electric fields, potential, and charges upon different geometry of cylinder and survey, Tx and Rx location, we understand geometric effects of the target for DCR survey.
# # Setup
# <img src="https://github.com/geoscixyz/geosci-labs/blob/master/images/em/DCR_Setup_Cylinder.png?raw=true"></img>
# # Question
# - Is the potential difference measured by a dipole over a conductive (/resisitive) target higher or lower compared to the half-space reference?
# - how do the field lines bend in presence of a conductive (/resistive) target?
# - Compared to the positive and negative sources (A and B), how are oriented the positive and negative accumulated charges around a conductive (/resistive) target?
# - How would you describe the secondary fields pattern? Does it remind you of the response of an object fundamental to electromagnetics?
# # Cylinder app
#
# ## Parameters:
# - **survey**: Type of survey
# - **A**: (+) Current electrode location
# - **B**: (-) Current electrode location
# - **M**: (+) Potential electrode location
# - **N**: (-) Potential electrode location
# - **r**: radius of cylinder
# - **xc**: x location of cylinder center
# - **zc**: z location of cylinder center
# - **$\rho_1$**: Resistivity of the halfspace
# - **$\rho_2$**: Resistivity of the cylinder
# - **Field**: Field to visualize
# - **Type**: which part of the field
# - **Scale**: Linear or Log Scale visualization
app = cylinder_app()
display(app)
| notebooks/dcip/DC_Cylinder_2D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Originally I wanted to adapt a model for predicting NCAA basket ball outcomes to the NHL. The model is called [Bayesian Logistic Regression Markov Chain (LRMC)](https://www2.isye.gatech.edu/~jsokol/lrmc/) and it works by treating the difference in points between two teams in any game as a normally distributed random variable which depends on the inherent difference in skill between the two teams plus a home court advantage added to the home team. The home court advantage is assumed to be constant across all teams. Unfortunately, when I originally explored this idea I discovered that the difference in score between two teams in each game would not be a good fit for a normal distribution, and so I concluded there wouldn't be an easy way to fit the LRMC model to the NHL.
#
# Refusing to give up on this project, I started looking at other ways to model HHL games, and thought about trying to model them in PYMC3. This thought lead me to a PYMC3 example called [A Hierarchical model for Rugby prediction](https://docs.pymc.io/notebooks/rugby_analytics.html) by <NAME>. That work was inspired by [<NAME>](http://danielweitzenfeld.github.io/passtheroc/blog/2014/10/28/bayes-premier-league/), which in turn was based on a model first developed by [<NAME> and <NAME>](http://www.statistica.it/gianluca/Research/BaioBlangiardo.pdf). With the help of the above examples and papers, I was able to figure out the preceding models and adapt them to the NHL. Due to NHL rules which force a winner of every game by first going to a five minute sudden death overtime, and then to a shootout, I have also extended the model to calculate a tie-breaker random variable to determine the ultimate winner.
# Import all of the libraries needed for this post
import requests
import json
import numpy as np
import pandas as pd
import pymc3 as pm
import theano.tensor as tt
import theano
from itertools import combinations
# Before we can dive into creating the model, we need to get some data. The functions below use the requests and json libraries to extract the data we need from the official NHL statistics API. I have written the data to CSV file so that it is possible to perform the rest of the analysis without constantly retrieving the data over the Internet.
# + code_folding=[]
# A function that retrieves the game data from the NHL stats API
# for a selected date range.
def request_game_data(start_date, end_date):
base_url = 'https://statsapi.web.nhl.com'
path = '/api/v1/schedule?startDate='+start_date+\
'&endDate='+end_date+'&expand=schedule.linescore'
response = requests.get(base_url + path)
return response.json()
# A function to extract the relevant data from the schedule
# and return it as a pandas dataframe
def extract_game_data(schedule):
"""Given full JSON records for games from the NHL API,
returns a simplified list of just the data we need.
"""
games = pd.DataFrame(columns=['date',
'season',
'game_type',
'home_team',
'home_team_reg_score',
'home_team_fin_score',
'away_team',
'away_team_reg_score',
'away_team_fin_score',
'went_to_shoot_out'
])
for date_obj in schedule['dates']:
date = date_obj['date'];
for game_obj in date_obj['games']:
game_type = game_obj['gameType']
season = game_obj['season']
home_team_obj = game_obj['teams']['home']
away_team_obj = game_obj['teams']['away']
home_team = home_team_obj['team']['name']
home_team_fin_score = home_team_obj['score']
away_team = away_team_obj['team']['name']
away_team_fin_score = away_team_obj['score']
detailed_score_data = game_obj['linescore']
period_data = detailed_score_data['periods']
shootout_data = detailed_score_data['shootoutInfo']
home_team_reg_score = 0
away_team_reg_score = 0
for period in period_data[0:3]:
home_team_reg_score += period['home']['goals']
away_team_reg_score += period['away']['goals']
went_to_shoot_out = (shootout_data['home']['attempts'] != 0 or
shootout_data['away']['attempts'] != 0)
games = games.append({'date': date,
'season': season,
'game_type': game_type,
'home_team': home_team,
'home_team_reg_score': home_team_reg_score,
'home_team_fin_score': home_team_fin_score,
'away_team': away_team,
'away_team_reg_score': away_team_reg_score,
'away_team_fin_score': away_team_fin_score,
'went_to_shoot_out': went_to_shoot_out
}, ignore_index=True)
return games
completed_game_data = request_game_data('2018-02-24', '2019-02-24')
completed_games = extract_game_data(completed_game_data)
completed_games.to_csv('completed_games.csv', index = False)
scheduled_game_data = request_game_data('2019-02-25', '2019-04-09')
scheduled_games = extract_game_data(scheduled_game_data)
scheduled_games.to_csv('scheduled_games.csv', index = False)
# -
# It is necessary to also decorate this data with integer labels for the home and away teams, as well as the team pairs. These labels serve as an array index for the random variables, and allow us to reference the correct random variables for each team or team pair in the PYMC3 model.
# +
# Filter the data to just regular season games from the 2015-2016 and
# 2016-2017 seasons
completed_games = pd.read_csv('completed_games.csv')
completed_games = completed_games.loc[completed_games['game_type'] == 'R']
# Select the columns I need for this analysis
completed_games = completed_games[['home_team', 'away_team',
'home_team_reg_score',
'away_team_reg_score',
'home_team_fin_score',
'away_team_fin_score']]
# Extract the unique list of teams and assign an integer label to each one
teams = completed_games.home_team.unique()
teams = np.sort(teams)
teams = pd.DataFrame(teams, columns=['team'])
teams['i'] = teams.index
# Create a unique list of each team combination and assign an integer label
# to each one. Also decide which team will be 'heads' in each pair.
all_teams_pair_combinations = combinations(teams['team'], 2)
team_pairs_dict = {}
team_pairs_heads_dict = {}
pair_index = 0
for pair in all_teams_pair_combinations:
team_pairs_dict[(pair[0], pair[1])] = pair_index
team_pairs_dict[(pair[1], pair[0])] = pair_index
team_pairs_heads_dict[(pair[0], pair[1])] = pair[0]
team_pairs_heads_dict[(pair[1], pair[0])] = pair[0]
pair_index += 1
# Determine if the ultimate winner of the game was the heads team
# (Bernoulli outcome = True) or the tails team (Bernoulli outcome = False)
def game_outcome_to_bernoulli_data(row):
if row['home_team_fin_score'] > row['away_team_fin_score']:
return row['home_team'] == team_pairs_heads_dict[(row['home_team'], row['away_team'])]
return row['away_team'] == team_pairs_heads_dict[(row['home_team'], row['away_team'])]
# Modify the data to include team and pair integer labels
def add_team_data_labels(game_data):
game_data = game_data.merge(teams, left_on='home_team', right_on='team', how='left')
game_data = game_data.rename(columns={'i': 'i_home'}).drop('team', axis=1)
game_data = game_data.merge(teams, left_on='away_team', right_on='team', how='left')
game_data = game_data.rename(columns={'i': 'i_away'}).drop('team', axis=1)
game_data['i_pair'] = game_data.apply(lambda row: team_pairs_dict[(row['home_team'], row['away_team'])], axis=1)
game_data['i_pair_winner'] = game_data.apply(game_outcome_to_bernoulli_data, axis=1)
return game_data
completed_games = add_team_data_labels(completed_games)
# -
# Because the ultimate goal of this model is to make predictions about the outcomes for games that haven't been played yet we need to extract the data for the model into Theano shared variables as [described in the PYMC3 documentation](https://docs.pymc.io/advanced_theano.html). This will allow us to swap out the data for completed games with the scheduled games and then predict samples of game outcomes for those scheduled games too.
# +
# Determine the total number of teams and team pairs for PYMC3
num_teams = len(completed_games.i_home.drop_duplicates())
num_team_pairs = len(completed_games.i_pair.drop_duplicates())
# Create shaed theano variables that can be swapped out with
# scheduled games later.
home_team = theano.shared(completed_games.i_home.values)
away_team = theano.shared(completed_games.i_away.values)
team_pair = theano.shared(completed_games.i_pair.values)
# Create arrays of observations for our pymc3 model
observed_home_goals = completed_games.home_team_reg_score.values
observed_away_goals = completed_games.away_team_reg_score.values
observed_pair_outcomes = completed_games.i_pair_winner.values
# -
# Now we can fit the PYMC3 model. The model assumes that goals scored in regulation time by the home and the away team can be modeled as Poisson distributed random variables, which we treat as observed random variables since we can see the number of goals that were scored. We also assume that the distribution of these variables is dependent on some inherent features of the teams such as their defensive and offensive skill, as well as other phenomenon not specific to teams such as home ice advantage and a constant intercept term. All of these are unobserved random variables that we expect to determine the Poisson distributions for goals scored in each game. Additionally, the tie breaker is modeled as a Bernoulli observed random variable which I have opted to define using a Beta distribution as the unobserved random variable that determines the probability of a success. This Bernoulli random varable does not consider home ice advantage, as we determined in my last post that it does not play a major role in deciding the winner after a game makes it to overtime or a shootout.
with pm.Model() as model:
# Global model parameters
home = pm.Flat('home')
sd_offence = pm.HalfStudentT('sd_offence', nu=3, sd=2.5)
sd_defence = pm.HalfStudentT('sd_defence', nu=3, sd=2.5)
intercept = pm.Flat('intercept')
# Team-specific poisson model parameters
offence_star = pm.Normal('offence_star', mu=0, sd=sd_offence, shape=num_teams)
defence_star = pm.Normal('defence_star', mu=0, sd=sd_defence, shape=num_teams)
offence = pm.Deterministic('offence', offence_star - tt.mean(offence_star))
defence = pm.Deterministic('defence', defence_star - tt.mean(defence_star))
home_theta = tt.exp(intercept + home + offence[home_team] - defence[away_team])
away_theta = tt.exp(intercept + offence[away_team] - defence[home_team])
# Team-pair bernoulli model parameters
beta_a = np.array([1] * num_team_pairs)
beta_b = np.array([1] * num_team_pairs)
bernoulli_p = pm.Beta('binom_p', alpha=beta_a, beta=beta_b, shape=num_team_pairs)
# Likelihood of observed data
home_goals = pm.Poisson('home_goals', mu=home_theta, observed=observed_home_goals)
away_goals = pm.Poisson('away_goals', mu=away_theta, observed=observed_away_goals)
tie_breaker = pm.Bernoulli('tie_breaker', p=bernoulli_p[team_pair], observed=observed_pair_outcomes)
with model:
trace = pm.sample(2000, tune=1000, cores=3)
pm.traceplot(trace)
# The trace plots make it appear as though the PYMC3 model has converged to the stationary distribution for each of the variables, suggesting that we do not need to adjust the burn-in period manually.
#
# Next we can also look at the BFMI and Gelman-Rubin statistics:
bfmi = pm.bfmi(trace)
max_gr = max(np.max(gr_stats) for gr_stats in pm.gelman_rubin(trace).values())
(pm.energyplot(trace, legend=False, figsize=(6, 4))
.set_title("BFMI = {}\nGelman-Rubin = {}".format(bfmi, max_gr)));
# The [BFMI statistic is well above the threshold of 0.2](https://docs.pymc.io/api/stats.html) that is typically suggested by the PYMC3 and Stan projects for indicating poor sampling. Furthermore, the Gelman-Rubin statistic is very close to 1, which further suggests that convergence on the stationary distribution has occurred.
#
# Satisfied that the PYMC3 model hasn't failed miserably, let's look at the posterior distributions for some of the unobserved random variables like team offensive and defensive strengths:
# +
import matplotlib.pyplot as plt
from matplotlib.ticker import StrMethodFormatter
import seaborn as sns
# %matplotlib inline
df_hpd = pd.DataFrame(pm.stats.hpd(trace['offence']),
columns=['hpd_low', 'hpd_high'],
index=teams.team.values)
df_median = pd.DataFrame(pm.stats.quantiles(trace['offence'])[50],
columns=['hpd_median'],
index=teams.team.values)
df_hpd = df_hpd.join(df_median)
df_hpd['relative_lower'] = df_hpd.hpd_median - df_hpd.hpd_low
df_hpd['relative_upper'] = df_hpd.hpd_high - df_hpd.hpd_median
df_hpd = df_hpd.sort_values(by='hpd_median')
df_hpd = df_hpd.reset_index()
df_hpd['x'] = df_hpd.index + .5
fig, axs = plt.subplots(figsize=(10,4))
axs.errorbar(df_hpd.x, df_hpd.hpd_median,
yerr=(df_hpd[['relative_lower', 'relative_upper']].values).T,
fmt='o')
axs.set_title('HPD of Offensive Strength by Team')
axs.set_xlabel('Team')
axs.set_ylabel('Posterior Offensive Strength')
_= axs.set_xticks(df_hpd.index + .5)
_= axs.set_xticklabels(df_hpd['index'].values, rotation=90)
# -
# The spread of offensive strengths looks pretty reasonable, and it seems to rank the teams well based on what little I know about their ability to score goals. Note that the Vegas Golden Knights have a slightly wider Highest Posterior Density (HPD) interval than the other teams. This makes sense since they have only started playing in the current recent season, and have far fewer games than the rest of the teams since we have included the complete 2016-2017 season in the data as well.
# +
df_hpd = pd.DataFrame(pm.stats.hpd(trace['defence']),
columns=['hpd_low', 'hpd_high'],
index=teams.team.values)
df_median = pd.DataFrame(pm.stats.quantiles(trace['defence'])[50],
columns=['hpd_median'],
index=teams.team.values)
df_hpd = df_hpd.join(df_median)
df_hpd['relative_lower'] = df_hpd.hpd_median - df_hpd.hpd_low
df_hpd['relative_upper'] = df_hpd.hpd_high - df_hpd.hpd_median
df_hpd = df_hpd.sort_values(by='hpd_median')
df_hpd = df_hpd.reset_index()
df_hpd['x'] = df_hpd.index + .5
fig, axs = plt.subplots(figsize=(10,4))
axs.errorbar(df_hpd.x, df_hpd.hpd_median,
yerr=(df_hpd[['relative_lower', 'relative_upper']].values).T,
fmt='o')
axs.set_title('HPD of Defensive Strength, by Team')
axs.set_xlabel('Team')
axs.set_ylabel('Posterior Defensive Strength')
_= axs.set_xticks(df_hpd.index + .5)
_= axs.set_xticklabels(df_hpd['index'].values, rotation=90)
# -
# The spread of defensive strengths also appears reasonable, and once again Vegas has a slightly wider HPD as we would expect.
#
# Now let's move on to the fun part and begin trying to predict outcomes for the remaining games.
# +
scheduled_games = pd.read_csv('scheduled_games.csv')
scheduled_games = scheduled_games.loc[scheduled_games['game_type'] == 'R']
# Select the columns I need for this analysis
scheduled_games = scheduled_games[['home_team', 'away_team',
'home_team_reg_score', 'away_team_reg_score',
'home_team_fin_score', 'away_team_fin_score']]
scheduled_games = add_team_data_labels(scheduled_games)
# Create shared theano variables that can be swapped out with
# scheduled games later.
home_team.set_value(scheduled_games.i_home.values)
away_team.set_value(scheduled_games.i_away.values)
team_pair.set_value(scheduled_games.i_pair.values)
# -
with model:
post_pred = pm.sample_ppc(trace)
# We can make sure that the shape of all our posterior predictions looks reasonable. There are 122 games left in the 2017-2018 Regular season, and for our posterior predictions there are 2000 samples for each game, times 122 games.
print(scheduled_games.shape)
print(post_pred['away_goals'].shape)
print(post_pred['home_goals'].shape)
print(post_pred['tie_breaker'].shape)
# Let's look at how these simulations play out. For simplicity I will first examine a single game; the Calgary Flames vs the San Jose Sharks in San Jose. I picked this game in particular since my father is a Flames fan, and this is the next game they will play. Let us start by looking at the predicted number of goals each team will score during regulation time:
# +
import matplotlib.pyplot as plt
def plot_posterior_goal_count(posterior_goals, team_name):
fig = plt.figure()
ax = fig.add_subplot()
vc = pd.Series(posterior_goals).value_counts().sort_index()
vc /= float(vc.sum())
ax = vc.plot(kind='bar', width=0.9, color='b')
ax.set_ylabel('Probability of Goal Count')
ax.set_xlabel('Goal Count')
ax.set_title('Predicted Regulation Time Goals Scored for {}'.format(team_name))
fig = ax.get_figure()
plt.xticks(rotation=0)
plt.show()
plot_posterior_goal_count(post_pred['home_goals'][:,1], 'SJS')
plot_posterior_goal_count(post_pred['away_goals'][:,1], 'CGY')
# -
# San Jose appears to skew a bit higher in the predicted number of regulation tie goals. As a result, we should probably expect San Jose to be more likely to win this game. Let's see what the predicted probabilities are:
# +
# Determine all the games in which the home and away teams win, lose,
# or tie in regulation time
home_won_regulation = post_pred['home_goals'] > post_pred['away_goals']
away_won_regulation = post_pred['away_goals'] > post_pred['home_goals']
regulation_tie = post_pred['home_goals'] == post_pred['away_goals']
# Determine which team utimately wins in the event of a tie
home_won_tie_breaker = post_pred['tie_breaker'].copy()
away_won_tie_breaker = post_pred['tie_breaker'].copy()
home_team_is_heads = np.array([(home_team == team_pairs_heads_dict[(home_team, away_team)]) for
home_team, away_team in
zip(scheduled_games['home_team'], scheduled_games['away_team'])])
home_won_tie_breaker = (home_won_tie_breaker == home_team_is_heads)
away_won_tie_breaker = ~home_won_tie_breaker
# +
scheduled_game_probs = scheduled_games[['home_team', 'away_team']].copy()
scheduled_game_probs['home_regulation_win'] = home_won_regulation.mean(axis=0)
scheduled_game_probs['home_OT_SO_win'] = (regulation_tie & home_won_tie_breaker).mean(axis=0)
scheduled_game_probs['away_regulation_win'] = away_won_regulation.mean(axis=0)
scheduled_game_probs['away_OT_SO_win'] = (regulation_tie & away_won_tie_breaker).mean(axis=0)
scheduled_game_probs.loc[1, :]
# -
# The San Jose Sharks are definitely more likely to win this match according to our model. In fact, the flames have only a 49.75% chance to make it out of this game with any points. Given that the Flames are on the edge of being mathematically eliminated from a playoff spot, things aren't looking so great for their post season. Sorry Dad!
#
# Let's also look at the rest of the games the Flames are scheduled to play in:
# +
flames_home = scheduled_game_probs['home_team'] == "Calgary Flames"
flames_away = scheduled_game_probs['away_team'] == "Calgary Flames"
scheduled_game_probs.loc[(flames_home | flames_away), :]
# -
# In order to earn a playoff spot the flames would likely need to win every game left in the season, and even then that may not be enough if the teams ahead of them also play well in the mean time. The odds of the Flames winning every single game left in the season do not appear to be promising. The Flames game at home against the Arizona Coyotes is the only game where they even have a greater than 50% chance of winning the game outright, whether that is in regulation, overtime, or shootout.
#
# In the spirit of the above analysis, the next obvious step would be to look at the probability that each team will make it into the playoffs based on the predictions made for all the remaining games. Unfortunately it is not that straight forward to calculate the probability that a team will make it into the playoffs. Overall, the rules for calculating "Wild Card" playoff seed standings in the NHL are surprisingly convoluted. For starters, the current league is broken down into two conferences. Each conference has two divisions. The top three teams for each division earn a playoff spot. The remaining two playoff spots in each conference are then provided to the top two teams in those conferences that have not already qualified for a playoff spot. This doesn't sound too bad, except for the possibility where a tie occurs. In such a scenario, the tie breaking procedure is:
#
# "If two or more clubs are tied in points during the regular season, the standing of the clubs is determined in the following order: The fewer number of games played (i.e., superior points percentage).The greater number of games won, excluding games won in the Shootout. This figure is reflected in the ROW column. The greater number of points earned in games between the tied clubs. If two clubs are tied, and have not played an equal number of home games against each other, points earned in the first game played in the city that had the extra game shall not be included. If more than two clubs are tied, the higher percentage of available points earned in games among those clubs, and not including any "odd" games, shall be used to determine the standing. The greater differential between goals for and against for the entire regular season. NOTE: In standings a victory in a shootout counts as one goal for, while a shootout loss counts as one goal against."
#
# While I'd love to write some sort of function to calculate these values I'm worried I'd never finish this blog post if I start down that path. At the very least I probably won't finish it before the 2017-2018 season playoffs start, at which point the predictions will not be very interesting anymore. I will leave it as a project for another day, hopefully before the start of the 2018-2019 season in October. If I'm feeling very ambitious over the next few weeks I may also try to make predictions for the playoff games once those start in April.
| _drafts/modeling-the-nhl/.ipynb_checkpoints/Modeling the NHL (copy)-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Gradient Boosted Trees Regression Example.
from __future__ import print_function
from pyspark import SparkContext
from pyspark.mllib.tree import GradientBoostedTrees, GradientBoostedTreesModel
from pyspark.mllib.util import MLUtils
sc = SparkContext(appName="PythonGradientBoostedTreesRegressionExample")
data = MLUtils.loadLibSVMFile(sc, "data/mllib/sample_libsvm_data.txt")
(trainingData, testData) = data.randomSplit([0.7, 0.3])
model = GradientBoostedTrees.trainRegressor(trainingData,
categoricalFeaturesInfo={},
numIterations=3)
print(model)
predictions = model.predict(testData.map(lambda x: x.features))
labelsAndPredictions = testData.map(lambda lp: lp.label).zip(predictions)
testMSE = labelsAndPredictions.map(lambda lp: (lp[0] - lp[1]) * (lp[0] - lp[1])).sum() /\
float(testData.count())
print('Test Mean Squared Error = ' + str(testMSE))
print('Learned regression GBT model:')
print(model.toDebugString())
model = GradientBoostedTrees.trainRegressor(trainingData,
categoricalFeaturesInfo={}, numIterations=3)
if __name__ == "__main__":
# $example on$
# Load and parse the data file.
# Split the data into training and test sets (30% held out for testing)
# Train a GradientBoostedTrees model.
# Notes: (a) Empty categoricalFeaturesInfo indicates all features are continuous.
# (b) Use more iterations in practice.
# Evaluate model on test instances and compute test error
# Save and load model
model.save(sc, "target/tmp/myGradientBoostingRegressionModel")
sameModel = GradientBoostedTreesModel.load(sc, "target/tmp/myGradientBoostingRegressionModel")
# $example off$
| testing/jaak-it_demo/practical_learning/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import warnings
os.environ['CUDA_VISIBLE_DEVICES'] = ''
warnings.filterwarnings('ignore')
# +
import malaya_speech.augmentation.waveform as augmentation
import malaya_speech
from glob import glob
from itertools import cycle
from multiprocessing import Pool
import itertools
import numpy as np
import random
def chunks(l, n):
for i in range(0, len(l), n):
yield (l[i : i + n], i // n)
def multiprocessing(strings, function, cores = 6, returned = True):
df_split = chunks(strings, len(strings) // cores)
pool = Pool(cores)
print('initiate pool map')
pooled = pool.map(function, df_split)
print('gather from pool')
pool.close()
pool.join()
print('closed pool')
if returned:
return list(itertools.chain(*pooled))
files = glob('/home/husein/youtube/clean-wav-22k/*.wav')
random.shuffle(files)
file_cycle = cycle(files)
noises = glob('/home/husein/youtube/noise-22k/*.wav')
random.shuffle(noises)
Y_files = glob('output-noise-reduction/*-y.wav')
Y_files = cycle(Y_files)
sr = 22050
partition_size = 4096
def get_pair(f):
return f.split('/')[-1].split('-')[0]
def read_wav(f):
return malaya_speech.load(f, sr = sr)
def random_sampling(s, length):
return augmentation.random_sampling(s, sr = sr, length = length)
def random_amplitude(sample, low = 3, high = 5):
y_aug = sample.copy()
dyn_change = np.random.uniform(low = low, high = high)
y_aug = y_aug * dyn_change
return np.clip(y_aug, -1, 1)
def random_amplitude_threshold(sample, low = 1, high = 2, threshold = 0.4):
y_aug = sample.copy()
dyn_change = np.random.uniform(low = low, high = high)
y_aug[np.abs(y_aug) >= threshold] = (
y_aug[np.abs(y_aug) >= threshold] * dyn_change
)
return np.clip(y_aug, -1, 1)
def add_uniform_noise(sample, power = 0.01, return_noise = False, scale = False):
y_noise = sample.copy()
noise_amp = power * np.random.uniform() * np.amax(y_noise)
noise = noise_amp * np.random.normal(size = y_noise.shape[0])
y_noise = y_noise + noise
if scale:
y_noise = y_noise / (np.max(np.abs(y_noise)) + 1e-9)
if return_noise:
if scale:
noise = noise / (np.max(np.abs(y_noise)) + 1e-9)
return y_noise, noise
else:
return y_noise
def calc(signal, seed, add_uniform = False):
random.seed(seed)
choice = random.randint(0, 9)
print('choice', choice)
if choice == 0:
x = augmentation.sox_augment_high(
signal,
min_bass_gain = random.randint(25, 50),
reverberance = random.randint(0, 80),
hf_damping = 10,
room_scale = random.randint(0, 50),
negate = 1,
)
if choice == 1:
x = augmentation.sox_augment_high(
signal,
min_bass_gain = random.randint(25, 70),
reverberance = random.randint(0, 80),
hf_damping = 10,
room_scale = random.randint(0, 50),
negate = 0,
)
if choice == 2:
x = augmentation.sox_augment_low(
signal,
min_bass_gain = random.randint(5, 30),
reverberance = random.randint(0, 80),
hf_damping = 10,
room_scale = random.randint(0, 50),
negate = random.randint(0, 1),
)
if choice == 3:
x = augmentation.sox_augment_combine(
signal,
min_bass_gain_high = random.randint(25, 70),
min_bass_gain_low = random.randint(5, 30),
reverberance = random.randint(0, 80),
hf_damping = 10,
room_scale = random.randint(0, 90),
)
if choice == 4:
x = augmentation.sox_reverb(
signal,
reverberance = random.randint(10, 80),
hf_damping = 10,
room_scale = random.randint(10, 90),
)
if choice == 5:
x = random_amplitude_threshold(
signal, threshold = random.uniform(0.35, 0.8)
)
if choice == 6:
x = augmentation.lowpass_filter(
signal, sr = sr, cutoff = random.randint(200, 551)
)
if choice == 7:
x = augmentation.highpass_filter(
signal, sr = sr, cutoff = random.randint(551, 1653)
)
if choice == 8:
x = augmentation.bandpass_filter(
signal,
sr = sr,
cutoff_low = random.randint(200, 551),
cutoff_high = random.randint(551, 1653),
)
if choice == 9:
x = signal
if choice not in [5] and random.gauss(0.5, 0.14) > 0.6:
x = random_amplitude_threshold(
x, low = 1.0, high = 2.0, threshold = random.uniform(0.6, 0.9)
)
if random.gauss(0.5, 0.14) > 0.6 and add_uniform:
x = add_uniform_noise(
x, power = random.uniform(0.005, 0.015)
)
return x
def combine_speakers(files, n = 5):
w_samples = random.sample(files, n)
w_samples = [
random_sampling(read_wav(f)[0], length = 500) for f in w_samples
]
y = [w_samples[0]]
left = w_samples[0].copy() * random.uniform(0.5, 1.0)
for i in range(1, n):
right = w_samples[i].copy() * random.uniform(0.5, 1.0)
overlap = random.uniform(0.01, 1.25)
left_len = int(overlap * len(left))
padded_right = np.pad(right, (left_len, 0))
if len(left) > len(padded_right):
padded_right = np.pad(
padded_right, (0, len(left) - len(padded_right))
)
else:
left = np.pad(left, (0, len(padded_right) - len(left)))
y.append(padded_right)
left = left + padded_right
return left, y
def parallel(f):
y = random_sampling(read_wav(f)[0], length = 5000)
seed = random.randint(0, 100_000_000)
x = calc(y, seed)
if random.gauss(0.5, 0.14) > 0.6:
print('add small noise')
n = combine_speakers(noises, random.randint(1, 20))[0]
n = calc(n, seed, True)
combined, noise = augmentation.add_noise(
x,
n,
factor = random.uniform(0.01, 0.1),
return_noise = True,
rescale = False,
)
else:
combined = x
noise = combined - y
return combined, y, noise
def parallel_semisupervised(f):
f_ = get_pair(f)
f_ = f'output-noise-reduction/{f_}-y_.wav'
y = read_wav(f)[0]
combined = read_wav(f_)[0]
length = 5000
sr_ = int(sr / 1000)
up = len(y) - (sr_ * length)
if up < 1:
r = 0
else:
r = np.random.randint(0, up)
y = y[r : r + sr_ * length]
combined = combined[r : r + sr_ * length]
noise = combined - y
return combined, y, noise
def loop(files):
files = files[0]
results = []
for f in files:
results.append(parallel(f))
return results
def loop_semisupervised(files):
files = files[0]
results = []
for f in files:
results.append(parallel_semisupervised(f))
return results
# -
def generate(batch_size = 10, repeat = 1):
while True:
fs = [next(file_cycle) for _ in range(batch_size)]
results = multiprocessing(fs, loop, cores = len(fs))
fs = [next(Y_files) for _ in range(batch_size)]
results.extend(
multiprocessing(fs, loop_semisupervised, cores = len(fs))
)
for _ in range(repeat):
random.shuffle(results)
for r in results:
if (
not np.isnan(r[0]).any()
and not np.isnan(r[1]).any()
and not np.isnan(r[2]).any()
):
yield {'combined': r[0], 'y': r[1], 'noise': r[2]}
g = generate()
# +
import soundfile as sf
directory = 'testset-speech-enhancement'
# !mkdir {directory}
# +
from tqdm import tqdm
for i in tqdm(range(500)):
r = next(g)
sf.write(f'{directory}/{i}-y_.wav', r['combined'], 22050)
sf.write(f'{directory}/{i}-y.wav', r['y'], 22050)
# -
# !du -hs {directory}
| pretrained-model/speech-enhancement/testset-speech-enhancement.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="3Lph_OFxMtmR" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 2, Module 3*
#
# ---
# + [markdown] id="eirisKo6MtmV" colab_type="text"
# # Cross-Validation
#
#
# ## Assignment
# - [ ] [Review requirements for your portfolio project](https://lambdaschool.github.io/ds/unit2), then submit your dataset.
# - [ ] Continue to participate in our Kaggle challenge.
# - [ ] Use scikit-learn for hyperparameter optimization with RandomizedSearchCV.
# - [ ] Submit your predictions to our Kaggle competition. (Go to our Kaggle InClass competition webpage. Use the blue **Submit Predictions** button to upload your CSV file. Or you can use the Kaggle API to submit your predictions.)
# - [ ] Commit your notebook to your fork of the GitHub repo.
#
#
# You won't be able to just copy from the lesson notebook to this assignment.
#
# - Because the lesson was ***regression***, but the assignment is ***classification.***
# - Because the lesson used [TargetEncoder](https://contrib.scikit-learn.org/categorical-encoding/targetencoder.html), which doesn't work as-is for _multi-class_ classification.
#
# So you will have to adapt the example, which is good real-world practice.
#
# 1. Use a model for classification, such as [RandomForestClassifier](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.RandomForestClassifier.html)
# 2. Use hyperparameters that match the classifier, such as `randomforestclassifier__ ...`
# 3. Use a metric for classification, such as [`scoring='accuracy'`](https://scikit-learn.org/stable/modules/model_evaluation.html#common-cases-predefined-values)
# 4. If you’re doing a multi-class classification problem — such as whether a waterpump is functional, functional needs repair, or nonfunctional — then use a categorical encoding that works for multi-class classification, such as [OrdinalEncoder](https://contrib.scikit-learn.org/categorical-encoding/ordinal.html) (not [TargetEncoder](https://contrib.scikit-learn.org/categorical-encoding/targetencoder.html))
#
#
#
# ## Stretch Goals
#
# ### Reading
# - <NAME>, [Python Data Science Handbook, Chapter 5.3](https://jakevdp.github.io/PythonDataScienceHandbook/05.03-hyperparameters-and-model-validation.html), Hyperparameters and Model Validation
# - <NAME>, [Statistics for Hackers](https://speakerdeck.com/jakevdp/statistics-for-hackers?slide=107)
# - <NAME>, [A Programmer's Guide to Data Mining, Chapter 5](http://guidetodatamining.com/chapter5/), 10-fold cross validation
# - <NAME>, [A Basic Pipeline and Grid Search Setup](https://github.com/rasbt/python-machine-learning-book/blob/master/code/bonus/svm_iris_pipeline_and_gridsearch.ipynb)
# - <NAME>, [A Comparison of Grid Search and Randomized Search Using Scikit Learn](https://blog.usejournal.com/a-comparison-of-grid-search-and-randomized-search-using-scikit-learn-29823179bc85)
#
# ### Doing
# - Add your own stretch goals!
# - Try other [categorical encodings](https://contrib.scikit-learn.org/categorical-encoding/). See the previous assignment notebook for details.
# - In additon to `RandomizedSearchCV`, scikit-learn has [`GridSearchCV`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html). Another library called scikit-optimize has [`BayesSearchCV`](https://scikit-optimize.github.io/notebooks/sklearn-gridsearchcv-replacement.html). Experiment with these alternatives.
# - _[Introduction to Machine Learning with Python](http://shop.oreilly.com/product/0636920030515.do)_ discusses options for "Grid-Searching Which Model To Use" in Chapter 6:
#
# > You can even go further in combining GridSearchCV and Pipeline: it is also possible to search over the actual steps being performed in the pipeline (say whether to use StandardScaler or MinMaxScaler). This leads to an even bigger search space and should be considered carefully. Trying all possible solutions is usually not a viable machine learning strategy. However, here is an example comparing a RandomForestClassifier and an SVC ...
#
# The example is shown in [the accompanying notebook](https://github.com/amueller/introduction_to_ml_with_python/blob/master/06-algorithm-chains-and-pipelines.ipynb), code cells 35-37. Could you apply this concept to your own pipelines?
#
# + [markdown] id="a30-e00WMtmW" colab_type="text"
# ### BONUS: Stacking!
#
# Here's some code you can use to "stack" multiple submissions, which is another form of ensembling:
#
# ```python
# import pandas as pd
#
# # Filenames of your submissions you want to ensemble
# files = ['submission-01.csv', 'submission-02.csv', 'submission-03.csv']
#
# target = 'status_group'
# submissions = (pd.read_csv(file)[[target]] for file in files)
# ensemble = pd.concat(submissions, axis='columns')
# majority_vote = ensemble.mode(axis='columns')[0]
#
# sample_submission = pd.read_csv('sample_submission.csv')
# submission = sample_submission.copy()
# submission[target] = majority_vote
# submission.to_csv('my-ultimate-ensemble-submission.csv', index=False)
# ```
# + id="g40DrvqAKxmQ" colab_type="code" colab={}
# Filenames of submissions to ensemble
files = ['/content/waterpump-submission-01.csv', '/content/waterpump-submission-02.csv',
'/content/waterpump-submission-03.csv', '/content/waterpump-submission-04.csv',
'/content/waterpump-submission-05.csv', '/content/waterpump-submission-06.csv',
'/content/waterpump-submission-07.csv', '/content/waterpump-submission-09.csv',
'/content/waterpump-submission-09.csv', '/content/waterpump-submission-10.csv',
'/content/waterpump-submission-11.csv', '/content/waterpump-submission-12.csv']
target = 'status_group'
submissions = (pd.read_csv(file)[[target]] for file in files)
ensemble = pd.concat(submissions, axis='columns')
majority_vote = ensemble.mode(axis='columns')[0]
sample_submission = pd.read_csv(DATA_PATH+'waterpumps/sample_submission.csv')
submission = sample_submission.copy()
submission[target] = majority_vote
submission.to_csv('my-ultimate-ensemble-submission.csv', index=False)
# + id="LrMG_vLaMYAV" colab_type="code" outputId="e8b9f5e1-b2dd-4be1-d292-6e357e1765fc" colab={"base_uri": "https://localhost:8080/", "height": 35}
submission_best = pd.read_csv('/content/my-ultimate-ensemble-submission.csv')
submission_best.shape
# + id="YJC9KsVVMl6y" colab_type="code" outputId="09689741-1354-4817-b2af-3ddf93ed5e27" colab={"base_uri": "https://localhost:8080/", "height": 407}
submission_best
# + id="8AYsb_QQMtmX" colab_type="code" colab={}
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Kaggle-Challenge/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# + id="PjkQ-wEVRB0F" colab_type="code" colab={}
# all imports needed for this sheet
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
import category_encoders as ce
from sklearn.ensemble import RandomForestClassifier
from sklearn.impute import SimpleImputer
from sklearn.pipeline import make_pipeline
from sklearn.feature_selection import f_regression, SelectKBest
from sklearn.linear_model import Ridge
from sklearn.model_selection import cross_val_score
from sklearn.preprocessing import StandardScaler
import matplotlib.pyplot as plt
from sklearn.model_selection import validation_curve
from sklearn.tree import DecisionTreeRegressor
from sklearn.model_selection import GridSearchCV, RandomizedSearchCV
# + id="uP01xgJYMtma" colab_type="code" colab={}
# Merge train_features.csv & train_labels.csv
train = pd.merge(pd.read_csv(DATA_PATH+'waterpumps/train_features.csv'),
pd.read_csv(DATA_PATH+'waterpumps/train_labels.csv'))
# Read test_features.csv & sample_submission.csv
test = pd.read_csv(DATA_PATH+'waterpumps/test_features.csv')
sample_submission = pd.read_csv(DATA_PATH+'waterpumps/sample_submission.csv')
# + id="BYMkvL-7sGpf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 208} outputId="afba0d47-f95b-43d1-ab3f-8898d55d9b02"
train.columns
# + id="XNoSoL1lS7ht" colab_type="code" outputId="33a03f0a-4674-4309-855f-2f44c25b118f" colab={"base_uri": "https://localhost:8080/", "height": 90}
train['status_group'].value_counts()
# + id="47DgLZ94Sub5" colab_type="code" colab={}
# convert strings in target column to integers for use in future functions
train['status_group'] = train['status_group'].replace('functional', 1)
train['status_group'] = train['status_group'].replace('non functional', 2)
train['status_group'] = train['status_group'].replace('functional needs repair', 3)
# + id="24hylWYXMtmd" colab_type="code" colab={}
# Split train into train & val
train, val = train_test_split(train, train_size=0.80, test_size=0.20,
stratify=train['status_group'], random_state=42)
# + id="foZRMceUUfnK" colab_type="code" outputId="0f6853f7-dcc3-485a-ca94-5f44f595aa94" colab={"base_uri": "https://localhost:8080/", "height": 208}
train.columns
# + id="IaL2mNm5Qw5w" colab_type="code" colab={}
# create funtion to wrangle each set in same way
def wrangle(X):
"""Wrangle train, validate, and test sets in the same way"""
# Prevent SettingWithCopyWarning
X = X.copy()
# About 3% of the time, latitude has small values near zero,
# outside Tanzania, so we'll treat these values like zero.
X['latitude'] = X['latitude'].replace(-2e-08, 0)
# When columns have zeros and shouldn't, they are like null values.
# So we will replace the zeros with nulls, and impute missing values later.
# Also create a "missing indicator" column, because the fact that
# values are missing may be a predictive signal.
# after evaluation, decided to remove whether column is missing
# as cross validation showed it was not important
cols_with_zeros = ['longitude', 'latitude', 'construction_year',
'gps_height', 'population']
for col in cols_with_zeros:
X[col] = X[col].replace(0, np.nan)
#X[col+'_MISSING'] = X[col].isnull()
# Drop duplicate columns
duplicates = ['quantity_group', 'payment_type']
X = X.drop(columns=duplicates)
# Drop recorded_by (never varies)
unusable_variance = ['recorded_by', 'num_private']
X = X.drop(columns=unusable_variance)
# Convert date_recorded to datetime
X['date_recorded'] = pd.to_datetime(X['date_recorded'], infer_datetime_format=True)
# Extract components from date_recorded, then drop the original column
X['year_recorded'] = X['date_recorded'].dt.year
X['month_recorded'] = X['date_recorded'].dt.month
X['day_recorded'] = X['date_recorded'].dt.day
X = X.drop(columns='date_recorded')
# Engineer feature: how many years from construction_year to date_recorded
X['years'] = X['year_recorded'] - X['construction_year']
#X['years_MISSING'] = X['years'].isnull()
# Engineer feature: height of well is 1500 or lower
X['lower_height'] = X['gps_height'] <= 1500
# Engineer feature: year recorded 2012 or later
#X['post_2012_rec'] = X['year_recorded'] >= 2012
# Engineer feature: constructed 2000 or later
X['not_old'] = X['construction_year'] >= 2000
# Engineer feature: funder is in top 1 funders
top1 = train['installer'].value_counts()[:1].index
X['top_1_installer'] = X['installer'].isin(top1)
# return the wrangled dataframe
return X
train = wrangle(train)
val = wrangle(val)
test = wrangle(test)
# + id="bUZbvSrqSY5H" colab_type="code" colab={}
train['status_group'].value_counts()
# + id="hRyTsgAiyhfQ" colab_type="code" colab={}
train.columns
# + id="2Vm8lzo_ykep" colab_type="code" colab={}
train['amount_tsh'].value_counts()
# + id="I_pGF9UpQw_F" colab_type="code" outputId="4063f605-358b-4133-9ac9-ad8dcda2caf0" colab={"base_uri": "https://localhost:8080/", "height": 69}
# %%time
# Arrange data to include all categorical features
target = 'status_group'
X_train = train.drop(columns=target)
y_train = train[target]
X_val = val.drop(columns=target)
y_val = val[target]
X_test = test
# pipeline, with ordinal encoder
pipeline = make_pipeline(
#ce.OneHotEncoder(use_cat_names=True, cols=['lower_height']),
ce.OrdinalEncoder(),
SimpleImputer(strategy='most_frequent'),
RandomForestClassifier(n_estimators=210, max_features="auto",
random_state=42, n_jobs=-1, warm_start=False,
min_samples_leaf=2
)
)
# Fit on train, score on val
pipeline.fit(X_train, y_train)
print('Validation Accuracy', pipeline.score(X_val, y_val))
# + id="Ohs0KmQaFSVt" colab_type="code" colab={}
a = y_pred
b = ['functional' if i == 1 else 'non functional' if i == 2 else 'functional' for i in a]
# + id="XZvB2_LPG81-" colab_type="code" colab={}
y_pred = b
y_pred
# + id="d5Sq0GqZG_Re" colab_type="code" colab={}
# + id="6PmSRy5hEVIS" colab_type="code" outputId="4c692df8-9039-4671-cbbb-2374ef1064c4" colab={"base_uri": "https://localhost:8080/", "height": 34}
y_pred = pipeline.predict(X_test)
# make submission
submission = test[['id']].copy()
submission['status_group'] = y_pred
# submission['status_group']
submission.to_csv('waterpump-submission-21.csv', index=False)
sub_df = pd.read_csv('/content/waterpump-submission-21.csv')
sub_df.shape
# + id="B2LhF86IFAdt" colab_type="code" colab={}
# convert integers in target column to strings for use in kaggle submission
train['status_group'] = train['status_group'].replace('functional', 1)
train['status_group'] = train['status_group'].replace('non functional', 2)
train['status_group'] = train['status_group'].replace('functional needs repair', 3)
# + id="CCMKIbmRJFsy" colab_type="code" outputId="e1b2541c-91c4-4493-b8b9-36b3c1c1b252" colab={"base_uri": "https://localhost:8080/", "height": 54}
y_pred
# + [markdown] id="LKa49PxPCBfT" colab_type="text"
# Best hyperparameters {
# 'randomforestclassifier__max_depth': 40,
# 'randomforestclassifier__max_features': 0.08642105949487566,
# 'randomforestclassifier__n_estimators': 250,
# 'simpleimputer__strategy': 'most_frequent',
# 'targetencoder__min_samples_leaf': 285,
# 'targetencoder__smoothing': 310.8115391005846
# }
# Cross-validation MAE -0.7939604377104377
# + id="ls12gemmQxEH" colab_type="code" outputId="361df605-e44e-4daa-aec2-d98e06c2f226" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# Linear models have coefficients whereas decision trees have "Feature Importances"
import matplotlib.pyplot as plt
model = pipeline.named_steps['randomforestclassifier']
encoder = pipeline.named_steps['ordinalencoder']
encoded_columns = encoder.transform(X_val).columns
importances = pd.Series(model.feature_importances_, encoded_columns)
plt.figure(figsize=(10,30))
importances.sort_values().plot.barh(color='grey')
# + id="ihR4PWHqQxIi" colab_type="code" outputId="8b56be97-c414-4183-e06f-992de9a05dd3" colab={"base_uri": "https://localhost:8080/", "height": 34}
k = 3
scores = cross_val_score(pipeline, X_train, y_train, cv=k,
scoring='accuracy')
print(f'MAE for {k} folds:', -scores)
# + id="Ugd6fpUbQxNS" colab_type="code" outputId="a2d5790d-c64e-4425-f8c7-eb3b2aae5f10" colab={"base_uri": "https://localhost:8080/", "height": 34}
-scores.mean()
# + id="SVjrpCLlQxRt" colab_type="code" outputId="d09096d2-7bae-4cc9-b12d-6585320da994" colab={"base_uri": "https://localhost:8080/", "height": 156}
print('Model Hyperparameters:')
print(pipeline.named_steps['randomforestclassifier'])
# + id="IOuSKEPjQxWI" colab_type="code" outputId="0f795c87-70b4-42ba-a04a-33fd12e58d6b" colab={"base_uri": "https://localhost:8080/", "height": 595}
# %matplotlib inline
pipeline = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(),
RandomForestClassifier()
)
depth = range(1, 50, 3)
train_scores, val_scores = validation_curve(
pipeline, X_train, y_train,
param_name='randomforestclassifier__max_depth',
param_range=depth, scoring='accuracy',
cv=3,
n_jobs=-1
)
plt.figure(dpi=150)
plt.plot(depth, np.mean(-train_scores, axis=1), color='blue', label='training error')
plt.plot(depth, np.mean(-val_scores, axis=1), color='red', label='validation error')
plt.title('Validation Curve')
plt.xlabel('model complexity: RandomForestClassifier max_depth')
plt.ylabel('model score: Accuracy')
plt.legend();
# + id="MPugkjODQxfo" colab_type="code" colab={}
y_pred = pipeline.predict(X_test)
# + id="Gv1J-yruQxkX" colab_type="code" colab={}
# make submission
submission = test[['id']].copy()
submission['status_group'] = y_pred
# submission['status_group']
submission.to_csv('waterpump-submission-17.csv', index=False)
# + id="ei_QVxXNQxoe" colab_type="code" colab={}
sub_df = pd.read_csv('/content/waterpump-submission-17.csv')
# + id="gX3YcoPmQxtP" colab_type="code" outputId="acc70a1b-f68f-4338-bdf7-834676fda7d9" colab={"base_uri": "https://localhost:8080/", "height": 35}
sub_df.shape
# + id="BXsSa1KsQxxq" colab_type="code" outputId="02aaefa3-b35f-4e35-f251-f02b4616b5b8" colab={"base_uri": "https://localhost:8080/", "height": 1000}
from scipy.stats import randint, uniform
features = train.columns.drop(target)
X_train = train[features]
y_train = train[target]
pipeline = make_pipeline(
ce.TargetEncoder(),
SimpleImputer(),
RandomForestClassifier(random_state=42)
)
param_distributions = {
'targetencoder__min_samples_leaf': randint(1, 1000),
'targetencoder__smoothing': uniform(1, 1000),
'simpleimputer__strategy': ['mean', 'median', 'most_frequent'],
'randomforestclassifier__n_estimators': range(50, 301, 50),
'randomforestclassifier__max_depth': [20, 30, 40, None],
'randomforestclassifier__max_features': uniform(0, 1),
}
# If you're on Colab, decrease n_iter & cv parameters
search = RandomizedSearchCV(
pipeline,
param_distributions=param_distributions,
n_iter=10,
cv=3,
scoring='accuracy',
verbose=10,
return_train_score=True,
n_jobs=1
)
search.fit(X_train, y_train);
# + id="UWok4IrDQx2F" colab_type="code" outputId="8289eb74-87cb-4a73-bc89-f1d7b6e1e5a9" colab={"base_uri": "https://localhost:8080/", "height": 74}
print('Best hyperparameters', search.best_params_)
print('Cross-validation accuracy score', -search.best_score_)
# + id="y4JCj-QHQx61" colab_type="code" colab={}
# + id="5GyKYOdrQx_k" colab_type="code" colab={}
# + id="xxn2S06VQyD_" colab_type="code" colab={}
# + id="Sz7ndqlEQyIH" colab_type="code" colab={}
| module3/LS_DS_223_assignment.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.3.0
# language: julia
# name: julia-1.3
# ---
# # EGM
# This notebook contains functions to solve for the non-stochastic steady-state of the model using the endogeneous grid method.
# +
function get_c(G_low::Function, G_high::Function, R::Float64, W::Float64, p::Params)
"""
Compute next period's consumption, conditional on next period's income
"""
c_grid = similar(p.grid_savings_reshaped)
c_grid[1, :] = R.*p.grid_savings_reshaped[1,:] .+ W*p.grid_endowment_reshaped[1,:] .- max.(p.grid_savings[1], G_low.(p.grid_savings_reshaped[1,:]))
c_grid[2, :] = R.*p.grid_savings_reshaped[2,:] .+ W*p.grid_endowment_reshaped[2,:] .- max.(p.grid_savings[1], G_high.(p.grid_savings_reshaped[2,:]))
return c_grid
end
# +
function euler_back(G_low::Function, G_high::Function, R::Float64, W::Float64, R_prime::Float64, W_prime::Float64, p::Params)
"""
Solve the Euler equation backward, using the EGM method
Input:
-----
G_low::Function: savings policy rule in the next period in bad state
G_high::Function: savings policy rule in the next period in good state
R::Float64: interest rate in the present period
W::Float64: wage in the present period
R_prime::Float64: interest rate next period
W_prime::Float64: wage next period
Output:
-------
a: policy function a' = g(a,e) for points on the p.grid_savings_reshaped
c: consumption c'= f(a,e) for points on the p.grid_savings_reshaped
Kg_low_f: policy function a' = g(a,e_low) for any a (interpolated)
Kg_high_f: policy function a' = g(a,e_high) for any a (interpolated)
"""
# 1. Compute next period's consumption, conditional on next period's income
c_prime = get_c(G_low, G_high, R_prime, W_prime, p)
# 1.' Compute next period's marginal utility of consumption:
up_cprime = similar(c_prime)
u′!(up_cprime, c_prime, p)
# 2. Expectation with respect to changes in idiosyncratic productivity:
E_up_cp = transpose(p.exog_trans)* up_cprime
# 3. Marginal utility of consumption = R' * beta*Eupcp
up_c = (R_prime*p.beta).*E_up_cp
# 4. Invert marginal utility of consumption to get consumption:
c = similar(c_prime)
u′_inv!(c, up_c, p)
# 5. The budget constraint implies the beginning-of-period asset:
# aR = (a' + c - y)
a = (p.grid_savings_reshaped .+ c .- W.*p.grid_endowment_reshaped)./R
#Permutations to sort data in increasing order:
p1 = sortperm(a[1,:])
p2 = sortperm(a[2,:])
# Interpolation a' to get a function a'=g(a,e)
Kg_low = LinearInterpolation(a[1, p1], p.grid_savings_reshaped[1,p1], extrapolation_bc=Line())
Kg_high = LinearInterpolation(a[2, p2], p.grid_savings_reshaped[2,p2], extrapolation_bc=Line())
# Define new functions using interpolation objects defined above:
Kg_low_f(x) = Kg_low(x)
Kg_high_f(x) = Kg_high(x)
return a, c, Kg_low_f, Kg_high_f
end
# +
function solve_EGM(g_low0::Function, g_high0::Function, R::Float64, W::Float64, p::Params; max_iter::Int64=10000, tol::Float64 = 1e-10, verbose::Bool=false)
"""
Solve the model using the endogeneous grid method
"""
a_old = similar(p.grid_savings_reshaped)
a_new = similar(p.grid_savings_reshaped)
c_new = similar(p.grid_savings_reshaped)
g_low_old = g_low0
g_high_old = g_high0
g_low_new(x) = log(x) #preallocate two policy functions
g_high_new(x) = log(x)
success_flag = 0 #convergence reached = 1; 0 otherwise
for it in 1:1:max_iter
a_new, c_new, g_low_new, g_high_new = euler_back(g_low_old, g_high_old, R, W, R, W, p)
# Check for convergence of assets grids:
if maximum(abs.(a_new - a_old)) < tol
if verbose==true
println("Convergence reached after $(it) iterations")
end
success_flag = 1
break
# Otherwise, update policy functions a'=g(a,e)
else
a_old = copy(a_new) #Policy function on grid points
g_low_old = g_low_new #Policy function interpolated
g_high_old = g_high_new
end
end
return a_new, c_new, g_low_new, g_high_new, success_flag
end
# -
# ### References
#
# * https://alisdairmckay.com/Notes/HetAgents/index.html
# * https://julia.quantecon.org/dynamic_programming/egm_policy_iter.html
# * [Car06] <NAME>. The method of endogenous gridpoints for solving dynamic stochastic optimization problems. Economics Letters, 91(3):312–320, 2006.
| .ipynb_checkpoints/EGM-checkpoint.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.3
# language: julia
# name: julia-1.5
# ---
# ## 2.1 Juliaの基本
# ### 2.1.1 変数
# 変数の代入
x = 1
# REPLと同様ansに値が入る
ans
# 乗算記号なしで数式を記述することも可能
2x+1
# unicodeを変数名に利用可能
# "θ"は,REPL上だと"\theta"の後にTabキーで出る.Jupyter notebookだと使えないので日本語でθと打つ
θ = pi/4
sin(θ)+cos(θ)
# ### 2.1.2 プリミティブ型
typeof(0.5)
# 16進は他の言語同様0xXXと表記する
# 2進も同様に0bXX
0xa*0xb
# ### 2.1.3 任意精度演算
# 科学技術計算向きの言語らしく,標準で任意精度演算BigInt, BigFloatがある
typeof(12345678901234567890123456789012345678901234567890)
# BigFloatは勝手にFloat64に丸められてしまう.
# BigFloatの変数を作るときはperseを使う
typeof(0.1234567890123456789012345678901234567890)
typeof(parse(BigFloat, "0.1234567890123456789012345678901234567890"))
# ### 2.1.4 定数
# 定数は以下のようにconstという宣言をつける.既に使用してしまった値は定数にできない.
const y=1.0
# 予め決められた定数もある
π
# ### 2.1.5 基本的な演算子
# 演算子は他の言語同様.優先度は()のような形で,乗算,除算,べき乗が優先される.
x = (1 + 2 - ((3 * 4) / 5)) % (6 ^ 7)
# べき乗に"^"があるのが便利な反面,xorには使えないので,xorは"⊻"(排他的論理和の数学記号)を使う."⊻"は変換だと出てこない.REPLなら"\xor" + Tab出でるが,JupyterだとU+22BBと打って変換するか辞書登録するしかなさそう
0b0101 ⊻ 0b1010
# 書きにくい場合こっちのほうが手っ取り早いかもしれない
xor(0b0101, 0b1010)
# ### 2.1.6 更新演算子
x += 10
# ### 2.1.7 複素数
x = 1 + 0.5im
# 実数部と虚数部を出力するには以下
println("Re = " * string(real(x)))
println(string("Im = ", imag(x)))
# ### 2.1.8~10 文字列
# 文字列は""でくくる.文字列への変換はstring(), 文字列結合は"*"を使う.str[3]のように,文字列のうち1文字のCharを取り出すことも可能.
str = "test string"
str[3]
# ただしマルチバイト文字は文字の順番とindexが合わなくなる.
str2 = "テスト文字列"
str2[4]
# こういう場合はcharの配列に変換すると文字とindexが合う用になる.
chars = Vector{Char}(str2)
chars[2]
# 文字列の他の関数として,python同様にreplaceやsplitが用意されている.含むかは,"in"ではなくoccursin
println(length("Hello")) # 長さ
println(replace(str, "test" => "main")) # 変換
println(split(str, " ")) # 分割
println(occursin("he", "hello")) # xにyを含むか
# ## 2.6 多次元配列
# ### 2.6.3 インデクシング
# 配列の取り扱いはMATLABに似ている.要素抽出にはコロン":"や"end"が使える.また,indexはMATLABと同様1始まり.
println(str[1:5])
println(str[6:end])
# ### 2.6.1 初期化, 2.6.2 基本的な操作
# 標準が多次元配列を示す.初期化にはzeros, ones, randなどが使える.
A = ones(3, 4)
println(A)
println(size(rand(4, 3)))
B = rand(4, 3)
# ### 2.6.4 多次元配列の演算
# MATLAB同様,多次元配列を通常の演算子で四則演算できる.ドットで要素単位の計算も可能.転置行列は"'".
A * B
A .+ B'
# ### 2.6.5 ブロードキャスティング
# サイズが異なる配列への演算がドットで可能
A .+ B[:,1]'
# 関数定義に対しても同様に適用可能.
square(x) = x * x;
square(10)
square(A) # ブロードキャスティングしないと実行できない
square.(A) # 関数の後にドットを打つことでブロードキャスティングされる
# ### 2.6.6. map reduce filter
# 配列要素に関数を適用したり集約する場合はこれらの関数を利用するのが便利.
map(x -> x*x, B)
filter(x -> x<0.1, B)
# ただし,上記は別の記載も可能.使いやすいものを使うと良さそう.
[x*x for x in B] # リスト内包表記でもいける.
B[B.<0.1] # 条件を満たすindexで抽出することも可能.
# 配列については公式ドキュメントの他,以下のQiita記事も参考になる.
# [Juliaで最低限やっていくための配列操作まとめ - Qiita](https://qiita.com/A03ki/items/007be353411d19952ef7)
| julia2-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div class="alert alert-success" data-title="">
# <h2><i class="fa fa-tasks" aria-hidden="true"></i> Fashion-MNIST
# </h2>
# </div>
#
# + colab={} colab_type="code" id="Fl1OE9oqJgkG"
# import
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras
# import keras
# + colab={} colab_type="code" id="rjIY2PuHJgkJ" outputId="e5d35f3c-7100-4b05-b2da-1ce1ff7455b3"
print(tf.__version__)
# + [markdown] colab_type="text" id="4K_zaHCqJgkN"
# <img src= https://leejunhyun.github.io/assets/img/TensorFlow/Fashion-MNIST-01.png>
# + [markdown] colab_type="text" id="vVflwzWmJgkN"
# ## EDA
# <img src=https://s3-ap-northeast-1.amazonaws.com/newstopics-production/url/2b6d5deda497d42e2d0516c27e9fd1b6e4192f51?1590738740 width=40%>
# + colab={} colab_type="code" id="HIVmMzbVJgkO"
# data 불러오기
fashion_mnist = keras.datasets.fashion_mnist
# train test split 불필요..
(train_images, train_labels), (test_images, test_labels)= fashion_mnist.load_data()
# + colab={} colab_type="code" id="u9wUpWY1JgkQ"
class_names = ['Top', 'Trouser','Pullover',
'Dress','Coat', 'Sandal',
'Shirt','Sneaker','Bag',
'Ank']
# + colab={} colab_type="code" id="hWj_V0OPJgkT" outputId="e9593f7e-5b62-4f36-cb91-47a798d18ba4"
# 사진 개수 , 가로, 세로
print(train_images.shape)
print(test_images.shape)
# + [markdown] colab_type="text" id="jOX5jH3SJgkV"
# 컴퓨터에서 흑백이미지란 [0,255] 구간의 값으로 표시 된다.
# 255 일수록 밝고 0일수록 어둡습니다.
# + colab={} colab_type="code" id="Gyj2H4vaJgkV" outputId="40c35d12-f7bb-4ecb-bf6b-3efe576c3dde"
# 데이터 샘플링
plt.imshow(train_images[12], cmap='gray') # image 보여주기
plt.colorbar() # 오른쪽 컬러바
plt.show()
# + [markdown] colab_type="text" id="ZbOw2aAkJgkY"
# # scailing 작업
# [0,255] ---> [0,1]
# + colab={} colab_type="code" id="8Z_qDU6oJgkZ"
train_images = train_images / 255.0
test_images = test_images / 255.0
# + colab={} colab_type="code" id="2cYMyO7NJgkb" outputId="09cb58e5-446e-486f-e186-20ac3e9d243a"
plt.figure(figsize=(10,10))
for i in range(16):
plt.subplot(4,4,i+1)
plt.xticks([]) # x 축에 아무것도 표시하지 않기
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap='gray')
plt.xlabel(class_names[train_labels[i]])
plt.show()
# + [markdown] colab_type="text" id="FnrWlTeUJgkd"
# ## 모델학습
#
# 머신러닝과 다르게 feature engineering 과정이 필요가 없습니다.
#
# 1. 분류
# - 이진분류 : 클래스가 0/1 or 클래스 -1 / 1 -------> sigmoid
# - 멀티분류? : 클래스가 2 개이상 --------> softmax
# -
# <div class="alert alert-success" data-title="">
# <h2><i class="fa fa-tasks" aria-hidden="true"></i> 모델을 만들어보기
# </h2>
# </div>
#
# 정확도 87 이상 올리기
# + colab={} colab_type="code" id="8pFO3CcuJgkd" outputId="95d30c88-a219-466b-ed21-9d3e7731e57f"
# model define
model = keras.Sequential([
# 초기 layer
keras.layers.Flatten(input_shape=(28, 28)), # 이미지 사이즈를 넣어주기 == input 크기 ,
# layer 추가해보기
keras.layers.Dense(100,activation= 'relu' )
# 마지막 layer
keras.layers.Dense(10,activation= 'softmax' ) # 클래스가 10개인 분류이므로 softmax
])
model.summary()
# + colab={} colab_type="code" id="pLIdly7uJgkg"
from tensorflow.keras import backend as K
# + colab={} colab_type="code" id="jVAMwhRUJgki"
def recall(y_target, y_pred):
# clip(t, clip_value_min, clip_value_max) : clip_value_min~clip_value_max 이외 가장자리를 깎아 낸다
# round : 반올림한다
y_target_yn = K.round(K.clip(y_target, 0, 1)) # 실제값을 0(Negative) 또는 1(Positive)로 설정한다
y_pred_yn = K.round(K.clip(y_pred, 0, 1)) # 예측값을 0(Negative) 또는 1(Positive)로 설정한다
# True Positive는 실제 값과 예측 값이 모두 1(Positive)인 경우이다
count_true_positive = K.sum(y_target_yn * y_pred_yn)
# (True Positive + False Negative) = 실제 값이 1(Positive) 전체
count_true_positive_false_negative = K.sum(y_target_yn)
# Recall = (True Positive) / (True Positive + False Negative)
# K.epsilon()는 'divide by zero error' 예방차원에서 작은 수를 더한다
recall = count_true_positive / (count_true_positive_false_negative + K.epsilon())
# return a single tensor value
return recall
def precision(y_target, y_pred):
# clip(t, clip_value_min, clip_value_max) : clip_value_min~clip_value_max 이외 가장자리를 깎아 낸다
# round : 반올림한다
y_pred_yn = K.round(K.clip(y_pred, 0, 1)) # 예측값을 0(Negative) 또는 1(Positive)로 설정한다
y_target_yn = K.round(K.clip(y_target, 0, 1)) # 실제값을 0(Negative) 또는 1(Positive)로 설정한다
# True Positive는 실제 값과 예측 값이 모두 1(Positive)인 경우이다
count_true_positive = K.sum(y_target_yn * y_pred_yn)
# (True Positive + False Positive) = 예측 값이 1(Positive) 전체
count_true_positive_false_positive = K.sum(y_pred_yn)
# Precision = (True Positive) / (True Positive + False Positive)
# K.epsilon()는 'divide by zero error' 예방차원에서 작은 수를 더한다
precision = count_true_positive / (count_true_positive_false_positive + K.epsilon())
# return a single tensor value
return precision
# + colab={} colab_type="code" id="Lkka2y-FJgkk"
def f1score(y_target, y_pred):
_recall = recall(y_target, y_pred)
_precision = precision(y_target, y_pred)
# K.epsilon()는 'divide by zero error' 예방차원에서 작은 수를 더한다
_f1score = ( 2 * _recall * _precision) / (_recall + _precision+ K.epsilon())
# return a single tensor value
return _f1score
# + [markdown] colab_type="text" id="ercDhm1YJgkm"
# - sparse_categorical_crossentropy: 멀티분류
# - binary_crossentropy: 이진분류
# + colab={} colab_type="code" id="B7GKBTtjJgkm"
# 모델을 compile 해주기
model.compile(optimizer='adam', # 학습률을 어떻게 업데이트 할 인가?
loss = 'sparse_categorical_crossentropy', # 멀티분류
metrics =['accuracy',f1score]) # 평가지표를 무엇으로 할것인가?
# 분류 metrics = accuracy, f1, recall, precision
# + colab={} colab_type="code" id="YY5jriFmJgko" outputId="58c08535-ddec-481f-f556-b0f6fcbba332"
# 모델 fitting
# fit(X, y) 학습
# epoch == 전체 데이터를 몇번 사용해서 학습할 것인가
# verbose 진행률을 프린트
model.fit(train_images, train_labels, epochs=10, verbose=1)
# + colab={} colab_type="code" id="ASgEHJ1JJgkq" outputId="26d930d0-dffe-499b-ccdb-0c7382f771dd"
# Test 평가
# (tensorflow == evaluate) === (sklearn== predict )
# model.predict(X_test, y_test)
test_loss, test_acc, test_f1_score = model.evaluate(test_images, test_labels, verbose=1)
# + colab={} colab_type="code" id="84_RS9clJgks" outputId="d2135dcb-9f09-4fb6-f815-5c492974a4f7"
print(f'우리모델의 정확도: {round(test_acc*100,2)} %')
print('우리모델의 f1_score: {:.3f}'.format(test_f1_score))
# + colab={} colab_type="code" id="lfWo-PfFJgku"
def plot_image(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array[i], true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'blue'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
def plot_value_array(i, predictions_array, true_label):
predictions_array, true_label = predictions_array[i], true_label[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
thisplot = plt.bar(range(10), predictions_array, color="#777777")
plt.ylim([0, 1])
predicted_label = np.argmax(predictions_array)
thisplot[predicted_label].set_color('red')
thisplot[true_label].set_color('blue')
# + colab={} colab_type="code" id="5HmksEHxJgkw"
predictions = model.predict(test_images)
# + colab={} colab_type="code" id="G0cwGuYWJgky" outputId="edbd4992-cef5-49ef-b0e0-af78ef073925"
num_rows = 5
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(2*2*num_cols, 2*num_rows))
for i in range(num_images):
plt.subplot(num_rows, 2*num_cols, 2*i+1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(num_rows, 2*num_cols, 2*i+2)
plot_value_array(i, predictions, test_labels)
plt.show()
| code/Day05/Day05_06_basic deep learning with fashion mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="bYhP8HMTZDSk" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 105} outputId="74b2274e-70e5-48a3-8fa4-7db42e8b648d" executionInfo={"status": "ok", "timestamp": 1528394002871, "user_tz": 420, "elapsed": 17113, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107011811392003324323"}}
# !apt-get install -y -qq software-properties-common python-software-properties module-init-tools
# !add-apt-repository -y ppa:alessandro-strada/ppa 2>&1 > /dev/null
# !apt-get update -qq 2>&1 > /dev/null
# !apt-get -y install -qq google-drive-ocamlfuse fuse
from google.colab import auth
auth.authenticate_user()
from oauth2client.client import GoogleCredentials
creds = GoogleCredentials.get_application_default()
import getpass
# !google-drive-ocamlfuse -headless -id={creds.client_id} -secret={creds.client_secret} < /dev/null 2>&1 | grep URL
vcode = getpass.getpass()
# !echo {vcode} | google-drive-ocamlfuse -headless -id={creds.client_id} -secret={creds.client_secret}
# + id="nqe7t2fdZJK3" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# !mkdir -p drive
# !google-drive-ocamlfuse drive
# + id="AnInpvtq_9xd" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="90e2af98-e94c-42fb-b373-1ade03c46a47" executionInfo={"status": "ok", "timestamp": 1528397528853, "user_tz": 420, "elapsed": 1150, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107011811392003324323"}}
# cd drive/CS230/Colab/src
# + [markdown] id="7hQ6EfW1X7tF" colab_type="text"
# **Prepare for Training**
# + id="_KODntJwZFTc" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 2264} outputId="733174b1-8b1f-4547-a070-bd4ed7071b3b" executionInfo={"status": "ok", "timestamp": 1528394604718, "user_tz": 420, "elapsed": 501883, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107011811392003324323"}}
# !pip install h5py==2.6.0
# !pip install Keras==1.2.2
# !pip install matplotlib==1.5.3
# !pip install numba==0.30.1
# !pip install numpy==1.14.3
# !pip install pandas==0.18.1
# !pip install rasterio==1.0a5
# !pip install Shapely==1.5.17.post1
# !pip install scikit_image==0.12.3
# !pip install tifffile==0.10.0
# !pip install cv2==1.0
# !pip install tqdm==4.11.2
# !pip install descartes
# + id="dr6086fgK54r" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# !python3 get_3_band_shapes.py
# + id="objrNBI4lb4i" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 122} outputId="5620412b-1d6e-4a19-9f5e-ef1a11299a53" executionInfo={"status": "ok", "timestamp": 1528394609449, "user_tz": 420, "elapsed": 4661, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107011811392003324323"}}
# !pip install shapely
# + id="ajwrUPOZLIwe" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 51} outputId="25b9b31a-46fd-4bbb-f1d3-bc53ca4a2631" executionInfo={"status": "ok", "timestamp": 1528347929166, "user_tz": 420, "elapsed": 832141, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107011811392003324323"}}
# !python3 cache_train.py
# + [markdown] id="dMoijXh7ZBWz" colab_type="text"
# **Training**
# + id="s8e6ssqpT_EN" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 258} outputId="b6eda4d9-a60f-4430-fc42-45e7d6fc2e61" executionInfo={"status": "ok", "timestamp": 1528394613030, "user_tz": 420, "elapsed": 3521, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107011811392003324323"}}
# !pip install Keras==2.1.6
# + id="aiu4CeZccCc0" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1465} outputId="d932d9b4-6c59-4e37-f807-4d0e05cd8e9b" executionInfo={"status": "ok", "timestamp": 1528362601436, "user_tz": 420, "elapsed": 4333644, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107011811392003324323"}}
"""
Original code based on Kaggle competition
Modified to take 3-channel input
"""
from __future__ import division
import numpy as np
from keras.models import Model
from keras.layers import Input, merge, Convolution2D, MaxPooling2D, UpSampling2D, Cropping2D
from keras import backend as K
import keras
import h5py
from keras.layers.normalization import BatchNormalization
from keras.optimizers import Nadam
from keras.callbacks import History
import pandas as pd
from keras.backend import binary_crossentropy
import datetime
import os
import random
import threading
#-----Vivian added--------#
from skimage.morphology import binary_dilation, binary_erosion, dilation, rectangle, binary_opening
import numpy as np
#-----Vivian added--------#
from keras.models import model_from_json
img_rows = 112
img_cols = 112
smooth = 1e-12
num_channels = 3
num_mask_channels = 1
def jaccard_coef(y_true, y_pred):
intersection = K.sum(y_true * y_pred, axis=[0, -1, -2])
sum_ = K.sum(y_true + y_pred, axis=[0, -1, -2])
jac = (intersection + smooth) / (sum_ - intersection + smooth)
return K.mean(jac)
def jaccard_coef_int(y_true, y_pred):
y_pred_pos = K.round(K.clip(y_pred, 0, 1))
intersection = K.sum(y_true * y_pred_pos, axis=[0, -1, -2])
sum_ = K.sum(y_true + y_pred_pos, axis=[0, -1, -2])
jac = (intersection + smooth) / (sum_ - intersection + smooth)
return K.mean(jac)
def jaccard_coef_loss(y_true, y_pred):
return -K.log(jaccard_coef(y_true, y_pred)) + binary_crossentropy(y_pred, y_true)
def get_unet0():
inputs = Input((num_channels, img_rows, img_cols))
conv1 = Convolution2D(6, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(inputs)
conv1 = BatchNormalization(mode=0, axis=1)(conv1)
conv1 = keras.layers.advanced_activations.ELU()(conv1)
conv1 = Convolution2D(6, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(conv1)
conv1 = BatchNormalization(mode=0, axis=1)(conv1)
conv1 = keras.layers.advanced_activations.ELU()(conv1)
pool1 = MaxPooling2D(pool_size=(2, 2),dim_ordering='th')(conv1)
conv2 = Convolution2D(12, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(pool1)
conv2 = BatchNormalization(mode=0, axis=1)(conv2)
conv2 = keras.layers.advanced_activations.ELU()(conv2)
conv2 = Convolution2D(12, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(conv2)
conv2 = BatchNormalization(mode=0, axis=1)(conv2)
conv2 = keras.layers.advanced_activations.ELU()(conv2)
pool2 = MaxPooling2D(pool_size=(2, 2),dim_ordering='th')(conv2)
conv3 = Convolution2D(24, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(pool2)
conv3 = BatchNormalization(mode=0, axis=1)(conv3)
conv3 = keras.layers.advanced_activations.ELU()(conv3)
conv3 = Convolution2D(24, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(conv3)
conv3 = BatchNormalization(mode=0, axis=1)(conv3)
conv3 = keras.layers.advanced_activations.ELU()(conv3)
pool3 = MaxPooling2D(pool_size=(2, 2),dim_ordering='th')(conv3)
conv4 = Convolution2D(48, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(pool3)
conv4 = BatchNormalization(mode=0, axis=1)(conv4)
conv4 = keras.layers.advanced_activations.ELU()(conv4)
conv4 = Convolution2D(48, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(conv4)
conv4 = BatchNormalization(mode=0, axis=1)(conv4)
conv4 = keras.layers.advanced_activations.ELU()(conv4)
pool4 = MaxPooling2D(pool_size=(2, 2),dim_ordering='th')(conv4)
conv5 = Convolution2D(96, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(pool4)
conv5 = BatchNormalization(mode=0, axis=1)(conv5)
conv5 = keras.layers.advanced_activations.ELU()(conv5)
conv5 = Convolution2D(96, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(conv5)
conv5 = BatchNormalization(mode=0, axis=1)(conv5)
conv5 = keras.layers.advanced_activations.ELU()(conv5)
up6 = merge([UpSampling2D(size=(2, 2),dim_ordering='th')(conv5), conv4], mode='concat', concat_axis=1)
conv6 = Convolution2D(48, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(up6)
conv6 = BatchNormalization(mode=0, axis=1)(conv6)
conv6 = keras.layers.advanced_activations.ELU()(conv6)
conv6 = Convolution2D(48, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(conv6)
conv6 = BatchNormalization(mode=0, axis=1)(conv6)
conv6 = keras.layers.advanced_activations.ELU()(conv6)
up7 = merge([UpSampling2D(size=(2, 2),dim_ordering='th')(conv6), conv3], mode='concat', concat_axis=1)
conv7 = Convolution2D(24, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(up7)
conv7 = BatchNormalization(mode=0, axis=1)(conv7)
conv7 = keras.layers.advanced_activations.ELU()(conv7)
conv7 = Convolution2D(24, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(conv7)
conv7 = BatchNormalization(mode=0, axis=1)(conv7)
conv7 = keras.layers.advanced_activations.ELU()(conv7)
up8 = merge([UpSampling2D(size=(2, 2),dim_ordering='th')(conv7), conv2], mode='concat', concat_axis=1)
conv8 = Convolution2D(12, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(up8)
conv8 = BatchNormalization(mode=0, axis=1)(conv8)
conv8 = keras.layers.advanced_activations.ELU()(conv8)
conv8 = Convolution2D(12, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(conv8)
conv8 = BatchNormalization(mode=0, axis=1)(conv8)
conv8 = keras.layers.advanced_activations.ELU()(conv8)
up9 = merge([UpSampling2D(size=(2, 2),dim_ordering='th')(conv8), conv1], mode='concat', concat_axis=1)
conv9 = Convolution2D(6, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(up9)
conv9 = BatchNormalization(mode=0, axis=1)(conv9)
conv9 = keras.layers.advanced_activations.ELU()(conv9)
conv9 = Convolution2D(6, 3, 3, border_mode='same', init='he_uniform',dim_ordering='th')(conv9)
crop9 = Cropping2D(cropping=((16, 16), (16, 16)),dim_ordering='th')(conv9)
conv9 = BatchNormalization(mode=0, axis=1)(crop9)
conv9 = keras.layers.advanced_activations.ELU()(conv9)
conv10 = Convolution2D(num_mask_channels, 1, 1, activation='sigmoid',dim_ordering='th')(conv9)
model = Model(input=inputs, output=conv10)
return model
def flip_axis(x, axis):
x = np.asarray(x).swapaxes(axis, 0)
x = x[::-1, ...]
x = x.swapaxes(0, axis)
return x
def form_batch(X, y, batch_size):
X_batch = np.zeros((batch_size, num_channels, img_rows, img_cols))
y_batch = np.zeros((batch_size, num_mask_channels, img_rows, img_cols))
X_height = X.shape[2]
X_width = X.shape[3]
for i in range(batch_size):
random_width = random.randint(0, X_width - img_cols - 1)
random_height = random.randint(0, X_height - img_rows - 1)
random_image = random.randint(0, X.shape[0] - 1)
y_batch[i] = y[random_image, :, random_height: random_height + img_rows, random_width: random_width + img_cols]
X_batch[i] = np.array(X[random_image, :, random_height: random_height + img_rows, random_width: random_width + img_cols])
return X_batch, y_batch
class threadsafe_iter:
"""Takes an iterator/generator and makes it thread-safe by
serializing call to the `next` method of given iterator/generator.
"""
def __init__(self, it):
self.it = it
self.lock = threading.Lock()
def __iter__(self):
return self
def next(self):
with self.lock:
return self.it.next()
def threadsafe_generator(f):
"""A decorator that takes a generator function and makes it thread-safe.
"""
def g(*a, **kw):
yield threadsafe_iter(f(*a, **kw))
return g
def batch_generator(X, y, batch_size, horizontal_flip=False, vertical_flip=False, swap_axis=False):
while True:
X_batch, y_batch = form_batch(X, y, batch_size)
for i in range(X_batch.shape[0]):
xb = X_batch[i]
yb = y_batch[i]
if horizontal_flip:
if np.random.random() < 0.5:
xb = flip_axis(xb, 1)
yb = flip_axis(yb, 1)
if vertical_flip:
if np.random.random() < 0.5:
xb = flip_axis(xb, 2)
yb = flip_axis(yb, 2)
if swap_axis:
if np.random.random() < 0.5:
xb = xb.swapaxes(1, 2)
yb = yb.swapaxes(1, 2)
X_batch[i] = xb
y_batch[i] = yb
yield X_batch, y_batch[:, :, 16:16 + img_rows - 32, 16:16 + img_cols - 32]
def save_model(model, cross):
json_string = model.to_json()
if not os.path.isdir('cache'):
os.mkdir('cache')
json_name = 'architecture_' + cross + '.json'
weight_name = 'model_weights_' + cross + '.h5'
open(os.path.join('cache', json_name), 'w').write(json_string)
model.save_weights(os.path.join('cache', weight_name), overwrite=True)
# def save_history(history, suffix):
# filename = 'history/history_' + suffix + '.csv'
# pd.DataFrame(history.history).to_csv(filename, index=False)
def read_model(cross=''):
json_name = 'architecture_' + cross + '.json'
weight_name = 'model_weights_' + cross + '.h5'
model = model_from_json(open(os.path.join('../src/cache', json_name)).read())
model.load_weights(os.path.join('../src/cache', weight_name))
return model
if __name__ == '__main__':
data_path = '../data'
now = datetime.datetime.now()
print('[{}] Creating and compiling model...'.format(str(datetime.datetime.now())))
model = get_unet0()
print('[{}] Reading train...'.format(str(datetime.datetime.now())))
f = h5py.File(os.path.join(data_path, 'train_3.h5'), 'r')
X_train = f['train']
y_train = np.array(f['train_mask'])[:, 0]
y_train = np.expand_dims(y_train, 1)
print(y_train.shape)
train_ids = np.array(f['train_ids'])
batch_size = 128
nb_epoch = 3
history = History()
callbacks = [
history,
]
suffix = 'buildings_3_'
model.compile(optimizer=Nadam(lr=1e-3), loss=jaccard_coef_loss, metrics=['binary_crossentropy', jaccard_coef_int])
from tensorflow.python.client import device_lib
print(device_lib.list_local_devices())
model.fit_generator(batch_generator(X_train, y_train, batch_size, horizontal_flip=True, vertical_flip=True, swap_axis=True),
nb_epoch=4,
verbose=1,
samples_per_epoch=batch_size * 25,
callbacks=callbacks,
nb_worker=24
)
# changed from batch_size*400, nb_epoch = nb_epoch
# do predict, then transfer to np array, then do skimage opening
save_model(model, "{batch}_{epoch}_{suffix}".format(batch=batch_size, epoch=nb_epoch, suffix=suffix))
#save_history(history, suffix)
f.close()
# + [markdown] id="PuVxoKCgZVBQ" colab_type="text"
# **Prediction**
# + id="oliqeqHbnprC" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 343} outputId="616aa584-fdb5-4f77-8647-6e747171a19d" executionInfo={"status": "ok", "timestamp": 1528394630915, "user_tz": 420, "elapsed": 17789, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107011811392003324323"}}
# !pip install -U numpy
# !pip install -U numba
# + id="3WHCAgKRpb1i" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
# #!python3 make_prediction_cropped_buildings.py
# + [markdown] id="PSLvc8gjF0r9" colab_type="text"
# Adding binary opening (erosion followed by dilation) led to reduction of noise -> buildings were so small that the output got wiped out entirely. This proves that this morphological process can indeed be used to reduce noise, but it is perhaps not very suitable for this project.
# + id="1mvSaBs7BPXz" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1465} outputId="963a07d3-306c-4087-94e9-e64fc8a23a91" executionInfo={"status": "ok", "timestamp": 1528395042305, "user_tz": 420, "elapsed": 411326, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107011811392003324323"}}
from __future__ import division
import os
from tqdm import tqdm
import pandas as pd
import extra_functions
import shapely.geometry
from numba import jit
from keras.models import model_from_json
import numpy as np
from skimage.morphology import binary_dilation, binary_erosion, dilation, rectangle, binary_opening
def read_model(cross=''):
json_name = 'architecture_128_3_buildings_3_dilated_in_preprocessing' + cross + '.json'
weight_name = 'model_weights_128_3_buildings_3_dilated_in_preprocessing' + cross + '.h5'
model = model_from_json(open(os.path.join('../cache', json_name)).read())
model.load_weights(os.path.join('../cache', weight_name))
return model
model = read_model()
sample = pd.read_csv('../data/sample_submission.csv')
data_path = '../data'
num_channels = 3 # modified
num_mask_channels = 1
threashold = 0.3
three_band_path = os.path.join(data_path, 'three_band')
train_wkt = pd.read_csv(os.path.join(data_path, 'train_wkt_v4.csv'))
gs = pd.read_csv(os.path.join(data_path, 'grid_sizes.csv'), names=['ImageId', 'Xmax', 'Ymin'], skiprows=1)
shapes = pd.read_csv(os.path.join(data_path, '3_shapes.csv'))
test_ids = shapes.loc[~shapes['image_id'].isin(train_wkt['ImageId'].unique()), 'image_id']
result = []
def flip_axis(x, axis):
x = np.asarray(x).swapaxes(axis, 0)
x = x[::-1, ...]
x = x.swapaxes(0, axis)
return x
@jit
def mask2poly(predicted_mask, threashold, x_scaler, y_scaler):
polygons = extra_functions.mask2polygons_layer(predicted_mask[0] > threashold, epsilon=0, min_area=5)
polygons = shapely.affinity.scale(polygons, xfact=1.0 / x_scaler, yfact=1.0 / y_scaler, origin=(0, 0, 0))
return shapely.wkt.dumps(polygons.buffer(2.6e-5))
#vivian added
test_ids = test_ids[200:220]
#vivian added
for image_id in tqdm(test_ids):
image = extra_functions.read_image_16(image_id)
H = image.shape[1]
W = image.shape[2]
x_max, y_min = extra_functions._get_xmax_ymin(image_id)
predicted_mask = extra_functions.make_prediction_cropped(model, image, initial_size=(112, 112),
final_size=(112-32, 112-32),
num_masks=num_mask_channels, num_channels=num_channels)
image_v = flip_axis(image, 1)
predicted_mask_v = extra_functions.make_prediction_cropped(model, image_v, initial_size=(112, 112),
final_size=(112 - 32, 112 - 32),
num_masks=1,
num_channels=num_channels)
image_h = flip_axis(image, 2)
predicted_mask_h = extra_functions.make_prediction_cropped(model, image_h, initial_size=(112, 112),
final_size=(112 - 32, 112 - 32),
num_masks=1,
num_channels=num_channels)
image_s = image.swapaxes(1, 2)
predicted_mask_s = extra_functions.make_prediction_cropped(model, image_s, initial_size=(112, 112),
final_size=(112 - 32, 112 - 32),
num_masks=1,
num_channels=num_channels)
new_mask = np.power(predicted_mask *
flip_axis(predicted_mask_v, 1) *
flip_axis(predicted_mask_h, 2) *
predicted_mask_s.swapaxes(1, 2), 0.25)
# vivian added - morphology
# new_mask = binary_dilation(new_mask)
x_scaler, y_scaler = extra_functions.get_scalers(H, W, x_max, y_min)
mask_channel = 0
result += [(image_id, mask_channel + 1, mask2poly(new_mask, threashold, x_scaler, y_scaler))]
submission = pd.DataFrame(result, columns=['ImageId', 'ClassType', 'MultipolygonWKT'])
sample = sample.drop('MultipolygonWKT', 1)
submission = sample.merge(submission, on=['ImageId', 'ClassType'], how='left').fillna('MULTIPOLYGON EMPTY')
submission.to_csv('temp_building_3.csv', index=False)
# + [markdown] id="hco-QTupG0oC" colab_type="text"
# **Post-Processing**
# + [markdown] id="tlnjV1kcoPfE" colab_type="text"
# Adding dilation in preprocessing seems to have enlarged the masks. We now see 109 bldgs (less buildings), but performs better at the scatter buildings down at the bottom left corner. This could be because dilation in preprocessing has enlarged the ground truth mask for those small groups of buildings, therefore causing the model to capture those small buildings better, instead of ignoring them and treat them as noise. Actually more buildings were found, but the number is 109 because some buildings are deemed "connected".
#
# Adding dilation in postprocessing seems to have enlarged the masks, but no new buildings were found. Building number reduced due to the fact that some buildings are now deemed connected.
#
# Default amount of dilation is good enough -> but since buildings are small, default amount of erosion just wipe out our buildings completely.
# + id="UPTcWjfx-UAe" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="9fd1d36e-193a-4a9c-9140-26e7eb4a39a5" executionInfo={"status": "ok", "timestamp": 1528398504626, "user_tz": 420, "elapsed": 43535, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107011811392003324323"}}
from __future__ import division
import extra_functions
import numpy as np
import shapely.wkt
import shapely.affinity
from numba import jit
import pandas as pd
import os
import sys
from shapely.geometry import MultiPolygon
from tqdm import tqdm
#vivian added
from skimage.morphology import watershed
from skimage.feature import peak_local_max
from skimage.morphology import binary_dilation, binary_erosion, dilation, rectangle, binary_opening, binary_closing, cube
predictions = pd.read_csv('temp_building_3.csv')
real_test_ids = ['6080_4_4', '6080_4_1', '6010_0_1', '6150_3_4', '6020_0_4', '6020_4_3',
'6150_4_3', '6070_3_4', '6020_1_3', '6060_1_4', '6050_4_4', '6110_2_3',
'6060_4_1', '6100_2_4', '6050_3_3', '6100_0_2', '6060_0_0', '6060_0_1',
'6060_0_3', '6060_2_0', '6120_1_4', '6160_1_4', '6120_3_3', '6140_2_3',
'6090_3_2', '6090_3_4', '6170_4_4', '6120_4_4', '6030_1_4', '6120_0_2',
'6030_1_2', '6160_0_0']
def generate_mask(image_id, predictions, num_mask_channels=10):
"""
:param image_id:
:param height:
:param width:
:param num_mask_channels:
:return: mask corresponding to an image_id of the desired height and width with desired number of channels
"""
height, width = extra_functions.get_shape(image_id)
mask = np.zeros((num_mask_channels, height, width))
for mask_channel in range(num_mask_channels):
poly = predictions.loc[(predictions['ImageId'] == image_id) & (
predictions['ClassType'] == mask_channel + 1), 'MultipolygonWKT'].values[0]
polygons = shapely.wkt.loads(poly)
if polygons.type == 'Polygon':
polygons = MultiPolygon([polygons])
mask[mask_channel, :, :] = extra_functions.polygons2mask_layer(height, width, polygons, image_id)
return mask
@jit
def fix_pred(image_id, predictions):
# read image and transform 2 multilayer mask
mask = generate_mask(image_id, predictions)
# if image_id in victim list => replace all slow water by fast
building_index = 0
road_index = 2
tree_index = 4
crop_index = 5
fast_water_index = 6
slow_water_index = 7
# Let's remove everything from the fast water
fast_water = (mask[fast_water_index] == 1)
for i in [0, 1, 2, 3, 4, 5, 7, 8, 9]:
mask[i][fast_water] = 0
# For some strange reason there is a lot of predicted roads near the water edge
slow_water = (mask[slow_water_index] == 1)
mask[road_index][slow_water] = 0
# Let's remove everything from buildings (We trust building predictions)
buildings = (mask[building_index] == 1)
for i in range(1, 10):
mask[i][buildings] = 0
# Let's remove slow water from roads
roads = (mask[road_index] == 1)
mask[slow_water_index][roads] = 0
#
# # those that are all crops - make all crops:
# if image_id in all_crops:
# mask[crop_index] = 1
#
# # those that are all trees - make all trees:
# if image_id in all_trees:
# mask[tree_index] = 1
# remove everything from fast_water
# fast_water = (mask[fast_water_index] == 1)
# for index in [0, 1, 2, 3, 4, 5, 8, 9]:
# mask[index][fast_water] = 0
# Remove all slow water from buildings
# mask[slow_water_index][buildings] = 0
# # zero out crops that Sergey said are zero
# if image_id in zero_crops:
# mask[crop_index] = 0
#
# # zero out roads from visual inspections
# if image_id in zero_road:
# mask[road_index] = 0
#
# # take union of mapped and not mapped predictions for all classes except cars
# if image_id in mapped_prediction['ImageId'].unique():
# mapped_mask = generate_mask(image_id, mapped_prediction)
# for i in range(8):
# if mapped_mask[i].sum() == 0:
# mask[i] = 0
# else:
# mask[i] += mapped_mask[i]
return (mask > 0).astype(int)
@jit
def mask2poly(predicted_mask, x_scaler, y_scaler):
if mask_channel == 7:
min_area = 100
elif mask_channel == 6:
min_area = 5000
else:
min_area = 10
polygons = extra_functions.mask2polygons_layer(predicted_mask, epsilon=0, min_area=min_area)
if image_id == '6100_0_2' and mask_channel == 1:
polygons = polygons.buffer(0.5)
polygons = shapely.affinity.scale(polygons, xfact=1.0 / x_scaler, yfact=1.0 / y_scaler, origin=(0, 0, 0))
return shapely.wkt.dumps(polygons)
sample = pd.read_csv('../data/sample_submission.csv')
data_path = '../data'
num_mask_channels = 10
three_band_path = os.path.join(data_path, 'three_band')
train_wkt = pd.read_csv(os.path.join(data_path, 'train_wkt_v4.csv'))
gs = pd.read_csv(os.path.join(data_path, 'grid_sizes.csv'), names=['ImageId', 'Xmax', 'Ymin'], skiprows=1)
shapes = pd.read_csv(os.path.join(data_path, '3_shapes.csv'))
test_ids = shapes.loc[~shapes['image_id'].isin(train_wkt['ImageId'].unique()), 'image_id']
result = []
for image_id in tqdm(real_test_ids):
height, width = extra_functions.get_shape(image_id)
x_max, y_min = extra_functions._get_xmax_ymin(image_id)
predicted_mask = fix_pred(image_id, predictions)
# vivian - added morphology
# Now we want to separate the two objects in image
# Generate the markers as local maxima of the distance to the background
# distance = ndi.distance_transform_edt(predicted_mask[0])
# local_maxi = peak_local_max(distance, indices=False, footprint=np.ones((3, 3)),
# labels=predicted_mask[0])
# markers = ndi.label(local_maxi)[0]
# labels = watershed(-distance, markers, mask=predicted_mask[0])
# predicted_mask[0] = labels
# if image_id=='6100_0_2':
# saved_mask = predicted_mask
predicted_mask[0] = binary_erosion(predicted_mask[0])
x_scaler, y_scaler = extra_functions.get_scalers(height, width, x_max, y_min)
for mask_channel in range(num_mask_channels):
result += [(image_id, mask_channel + 1, mask2poly(predicted_mask[mask_channel], x_scaler, y_scaler))]
submission = pd.DataFrame(result, columns=['ImageId', 'ClassType', 'MultipolygonWKT'])
sample = sample.drop('MultipolygonWKT', 1)
submission = sample.merge(submission, on=['ImageId', 'ClassType'], how='left').fillna('MULTIPOLYGON EMPTY')
submission.to_csv('cleaned_' + 'temp_building_3.csv', index=False)
# + id="BLC7M1qAQBj6" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 2202} outputId="daedcc58-c044-4ccd-e921-87c655db331b" executionInfo={"status": "ok", "timestamp": 1528398377664, "user_tz": 420, "elapsed": 9159, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107011811392003324323"}}
import numpy as np
import matplotlib.pyplot as plt
from scipy import ndimage as ndi
from skimage.morphology import watershed
from skimage.feature import peak_local_max
image = saved_mask[0]
# Now we want to separate the two objects in image
# Generate the markers as local maxima of the distance to the background
distance = ndi.distance_transform_edt(image)
local_maxi = peak_local_max(distance, indices=False, footprint=np.ones((3, 3)),
labels=image)
markers = ndi.label(local_maxi)[0]
labels = watershed(-distance, markers, mask=image)
fig, axes = plt.subplots(ncols=3, figsize=(90, 30), sharex=True, sharey=True)
ax = axes.ravel()
ax[0].imshow(image, cmap=plt.cm.gray, interpolation='nearest')
ax[0].set_title('Overlapping objects')
ax[1].imshow(-distance, cmap=plt.cm.gray, interpolation='nearest')
ax[1].set_title('Distances')
ax[2].imshow(labels, cmap=plt.cm.nipy_spectral, interpolation='nearest')
ax[2].set_title('Separated objects')
for a in ax:
a.set_axis_off()
fig.tight_layout()
plt.show()
from skimage.measure import regionprops, label
regions = regionprops(labels)
regions = [r for r in regions if r.area > 50]
print('Number of buildings (watershed seperation):', len(regions) - 1)
# + [markdown] id="uHuKZ3tqpMzH" colab_type="text"
# Watershed can be used to seperate buildings so that we can now have 625 buildings detected...
# + id="2KMYXMTMkMFe" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
from skimage.morphology import binary_erosion
eroded = binary_erosion(saved_mask[0])
# + id="IfjqeIuqmXFN" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 34} outputId="c9df8d24-c158-4d5d-996b-bd72d4acfae0" executionInfo={"status": "ok", "timestamp": 1528397661925, "user_tz": 420, "elapsed": 281, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107011811392003324323"}}
np.any(eroded)
# + [markdown] id="jm5Sjn6HLjZv" colab_type="text"
# **Plotting**
# + id="peSN-s1aLaxa" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 425} outputId="4e140920-01d4-432a-8059-4a7c4f4512c7" executionInfo={"status": "ok", "timestamp": 1528398574753, "user_tz": 420, "elapsed": 70050, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "107011811392003324323"}}
from __future__ import division
from shapely.geometry import MultiPolygon
from pylab import *
import pandas as pd
rcParams['figure.figsize'] = 20, 20
def stretch_8bit(bands, lower_percent=2, higher_percent=98):
out = np.zeros_like(bands).astype(np.float32)
for i in range(3):
a = 0
b = 1
c = np.percentile(bands[:, :, i], lower_percent)
d = np.percentile(bands[:, :, i], higher_percent)
t = a + (bands[:, :, i] - c) * (b - a) / (d - c)
t[t < a] = a
t[t > b] = b
out[:, :, i] = t
return out.astype(np.float32)
import pandas as pd
import numpy as np
from shapely.wkt import loads as wkt_loads
from matplotlib.patches import Polygon, Patch
# decartes package makes plotting with holes much easier
from descartes.patch import PolygonPatch
import matplotlib.pyplot as plt
import tifffile as tiff
import pylab
# turn interactive mode on so that plots immediately
# See: http://stackoverflow.com/questions/2130913/no-plot-window-in-matplotlib
# pylab.ion()
inDir = '../data'
# Give short names, sensible colors and zorders to object types
CLASSES = {
1: 'Bldg',
2: 'Struct',
3: 'Road',
4: 'Track',
5: 'Trees',
6: 'Crops',
7: 'Fast H20',
8: 'Slow H20',
9: 'Truck',
10: 'Car',
}
COLORS = {
1: '0.7',
2: '0.4',
3: '#b35806',
4: '#dfc27d',
5: '#1b7837',
6: '#a6dba0',
7: '#74add1',
8: '#4575b4',
9: '#f46d43',
10: '#d73027',
}
ZORDER = {
1: 5,
2: 5,
3: 4,
4: 1,
5: 3,
6: 2,
7: 7,
8: 8,
9: 9,
10: 10,
}
# read the training data from train_wkt_v4.csv
# vivian modified
df = pd.read_csv('cleaned_temp_building_3.csv')
print(df.head())
# grid size will also be needed later..
gs = pd.read_csv(inDir + '/grid_sizes.csv', names=['ImageId', 'Xmax', 'Ymin'], skiprows=1)
print(gs.head())
# imageIds in a DataFrame
allImageIds = gs.ImageId.unique()
trainImageIds = df.ImageId.unique()
def get_image_names(imageId):
'''
Get the names of the tiff files
'''
d = {'3': '{}/three_band/{}.tif'.format(inDir, imageId),
'A': '{}/sixteen_band/{}_A.tif'.format(inDir, imageId),
'M': '{}/sixteen_band/{}_M.tif'.format(inDir, imageId),
'P': '{}/sixteen_band/{}_P.tif'.format(inDir, imageId),
}
return d
def get_images(imageId, img_key=None):
'''
Load images correspoding to imageId
Parameters
----------
imageId : str
imageId as used in grid_size.csv
img_key : {None, '3', 'A', 'M', 'P'}, optional
Specify this to load single image
None loads all images and returns in a dict
'3' loads image from three_band/
'A' loads '_A' image from sixteen_band/
'M' loads '_M' image from sixteen_band/
'P' loads '_P' image from sixteen_band/
Returns
-------
images : dict
A dict of image data from TIFF files as numpy array
'''
img_names = get_image_names(imageId)
images = dict()
if img_key is None:
for k in img_names.keys():
images[k] = tiff.imread(img_names[k])
else:
images[img_key] = tiff.imread(img_names[img_key])
return images
def get_size(imageId):
"""
Get the grid size of the image
Parameters
----------
imageId : str
imageId as used in grid_size.csv
"""
xmax, ymin = gs[gs.ImageId == imageId].iloc[0, 1:].astype(float)
W, H = get_images(imageId, '3')['3'].shape[1:]
return (xmax, ymin, W, H)
def is_training_image(imageId):
'''
Returns
-------
is_training_image : bool
True if imageId belongs to training data
'''
return any(trainImageIds == imageId)
def plot_polygons(fig, ax, polygonsList):
'''
Plot descartes.PolygonPatch from list of polygons objs for each CLASS
'''
legend_patches = []
for cType in polygonsList:
print('{} : {} \tcount = {}'.format(cType, CLASSES[cType], len(polygonsList[cType])))
legend_patches.append(Patch(color=COLORS[cType],
label='{} ({})'.format(CLASSES[cType], len(polygonsList[cType]))))
for polygon in polygonsList[cType]:
mpl_poly = PolygonPatch(polygon,
color=COLORS[cType],
lw=0,
alpha=0.7,
zorder=ZORDER[cType])
ax.add_patch(mpl_poly)
# ax.relim()
ax.autoscale_view()
ax.set_title('Objects')
ax.set_xticks([])
ax.set_yticks([])
return legend_patches
def stretch_n(bands, lower_percent=2, higher_percent=98):
out = np.zeros_like(bands).astype(np.float32)
n = bands.shape[0]
for i in range(n):
a = 0 # np.min(band)
b = 1 # np.max(band)
c = np.percentile(bands[i, :, :], lower_percent)
d = np.percentile(bands[i, :, :], higher_percent)
t = a + (bands[i, :, :] - c) * (b - a) / (d - c)
t[t < a] = a
t[t > b] = b
out[i, :, :] = t
return out
#vivian modified
real_test_ids = ['6100_0_2']
def plot_image(fig, ax, imageId, img_key, selected_channels=None):
'''
Plot get_images(imageId)[image_key] on axis/fig
Optional: select which channels of the image are used (used for sixteen_band/ images)
Parameters
----------
img_key : str, {'3', 'P', 'N', 'A'}
See get_images for description.
'''
images = get_images(imageId, img_key)
img = images[img_key]
title_suffix = ''
if selected_channels is not None:
img = img[selected_channels]
title_suffix = ' (' + ','.join([repr(i) for i in selected_channels]) + ')'
if len(img.shape) == 2:
new_img = np.zeros((3, img.shape[0], img.shape[1]))
new_img[0] = img
new_img[1] = img
new_img[2] = img
img = new_img
tiff.imshow(stretch_n(img), figure=fig, subplot=ax)
ax.set_title(imageId + ' - ' + img_key + title_suffix)
ax.set_xlabel(img.shape[-2])
ax.set_ylabel(img.shape[-1])
ax.set_xticks([])
ax.set_yticks([])
def visualize_image(imageId, plot_all=True):
'''
Plot all images and object-polygons
Parameters
----------
imageId : str
imageId as used in grid_size.csv
plot_all : bool, True by default
If True, plots all images (from three_band/ and sixteen_band/) as subplots.
Otherwise, only plots Polygons.
'''
df_image = df[df.ImageId == imageId]
xmax, ymin, W, H = get_size(imageId)
if plot_all:
fig, axArr = plt.subplots(figsize=(60, 30), ncols=2)
ax = axArr[0]
else:
fig, axArr = plt.subplots(figsize=(20, 20))
ax = axArr
if is_training_image(imageId):
print('ImageId : {}'.format(imageId))
polygonsList = {}
for cType in CLASSES.keys():
all_polygons = wkt_loads(df_image[df_image.ClassType == cType].MultipolygonWKT.values[0])
if all_polygons.type == 'Polygon':
all_polygons = MultiPolygon([all_polygons])
polygonsList[cType] = all_polygons
legend_patches = plot_polygons(fig, ax, polygonsList)
ax.set_xlim(0, xmax)
ax.set_ylim(ymin, 0)
ax.set_xlabel(xmax)
ax.set_ylabel(ymin)
if plot_all:
plot_image(fig, axArr[1], imageId, '3')
if is_training_image(imageId):
ax.legend(handles=legend_patches,
# loc='upper center',
bbox_to_anchor=(0.9, 1),
bbox_transform=plt.gcf().transFigure,
ncol=5,
fontsize='xx-large',
title='Objects-' + imageId,
# mode="expand",
framealpha=0.3)
return (fig, axArr, ax)
# Loop over few training images and save to files
for imageId in real_test_ids:
fig, axArr, ax = visualize_image(imageId, plot_all=True)
plt.tight_layout()
#vivian added
if not os.path.exists('predictions'):
os.makedirs('predictions')
plt.savefig('predictions/Objects--' + imageId + '.png')
plt.clf()
# + id="i2e0oQk36cNX" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
| Pre- and Post-Processing/pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="nEQfELCST147" outputId="218c414d-3ae2-43dc-92f8-f10b3ab224de"
import os
os.chdir('/content')
CODE_DIR = 'InvertibleCE'
# !git clone https://github.com/zhangrh93/InvertibleCE $CODE_DIR
os.chdir(f'./{CODE_DIR}')
# !pip install -r requirements.txt > installation_output.txt
# + [markdown] id="4rb_qArhwEjx"
# # MNIST model with GlobalAveragePooling
#
# + id="Cx6sExOwT1Or"
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torchvision import datasets, transforms
import matplotlib.pyplot as plt
import os
import torchvision
import numpy as np
import os
from pathlib import Path
from torchsummary import summary
import shutil
seed = 10
torch.manual_seed(seed)
torch.cuda.manual_seed(seed)
np.random.seed(seed)
torch.backends.cudnn.deterministic = True
# + [markdown] id="u6gBSraVT1Ot"
# ## Model structure
# Let's build a simple MNIST model for ICE explanations. It contains 6 convolutional layers with Relu activation functions. One Global Average Pooling layer (GAP) and one dense layer are used at last.
# + colab={"base_uri": "https://localhost:8080/"} id="ZPWzSxDVT1Ot" outputId="bd7c4fd2-9738-4831-9892-a9e873a11d18"
log_batch = 10
unit_num = 16
def _make_layer(in_shape, out_shape, unit_num):
return nn.Sequential(
nn.Conv2d(in_shape, unit_num, 3, 1, 1),
nn.ReLU(inplace=True),
nn.Conv2d(unit_num, out_shape, 3, 1, 1),
nn.ReLU(inplace=True),
)
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.layer1 = _make_layer(1,unit_num,unit_num)
self.layer2 = _make_layer(unit_num,unit_num*2,unit_num*2)
self.layer3 = _make_layer(unit_num*2,unit_num*4,unit_num*4)
self.gap = nn.AdaptiveAvgPool2d((1,1))
self.fc = nn.Linear(unit_num*4, 10)
def forward(self, x):
x = self.layer1(x)
x = F.max_pool2d(x, 2)
x = self.layer2(x)
x = F.max_pool2d(x, 2)
x = self.layer3(x)
x = self.gap(x)
x = torch.flatten(x, 1)
x = self.fc(x)
return x
use_cuda = torch.cuda.is_available()
model = Net()
if use_cuda:
model = model.cuda()
summary(model,(1,28,28))
# + [markdown] id="MPAnjqiqT1Ou"
# ## Pytorch dataloader
# Next, we need to download the MNIST dataset. Datasets are stored in '/dataset'.
# + colab={"base_uri": "https://localhost:8080/", "height": 561, "referenced_widgets": ["ff87d899655b4b91bd2fbf34051c2e31", "03c94e06e9ee4832b1a37197a571be08", "<KEY>", "<KEY>", "<KEY>", "60d56e1e0a8d4edab5989b933da8cca7", "e85cedb408dc458790a8503d8ff65680", "b8d06c3118d44a0f803af4416f008be7", "5ffedf52f8984393af59831eb27c93e5", "<KEY>", "92b6dbe93cf748b682da46307ea6da87", "ab761a383a0d401d9b09e7edcbed9458", "a0ac2ded94334515879aa102f6d6c89e", "<KEY>", "6269f8ef945e46d59286fae5ce66d8ff", "<KEY>", "a8df9412d5bc483eaf30ad4725f02ad8", "c666a417d4c54c1a93d34e876d66166a", "<KEY>", "<KEY>", "798e9d7ee6d145799f3a1ec4de6e6fd5", "2e6e49a524f74608a3408ef97af0fb63", "<KEY>", "02b552ba85ac406ea09f43258f89a0c9", "8fc05453ec414fe48bf89e17804ac361", "db7c63c9eb8a43b8b745aff60983c428", "d05d0a8acbc44a458b3ececeb35465eb", "16d35e6dfa2c400e90235cd7bfc84238", "<KEY>", "<KEY>", "<KEY>", "1b18469d67104ed8823a830c7348a049"]} id="vtQROAU_T1Ov" outputId="82257d3a-7f54-4947-b8d3-9c179e1497b0"
batch_size = 32
transform = transforms.Compose([transforms.ToTensor()])
data_train = datasets.MNIST(root = "./dataset",
transform=transform,
train = True,
download = True)
data_test = datasets.MNIST(root="./dataset",
transform = transform,
train = False)
data_loader_train = torch.utils.data.DataLoader(dataset=data_train,
batch_size = batch_size,
shuffle = True,
num_workers=0)
data_loader_test = torch.utils.data.DataLoader(dataset=data_test,
batch_size = batch_size,
shuffle = True,
num_workers=0)
# + [markdown] id="a4FKMQMKT1Ov"
# ## MNIST model training
# We do not want a perfect MNIST classifier. Around 97% accuracy is enough for the explanations.
#
# Trained models are stored in the './model' folder.
# + colab={"base_uri": "https://localhost:8080/"} id="OXfqZPcnT1Ov" outputId="4f6eec82-ecf8-4697-f2c8-dada48831e38"
if not os.path.exists(Path('model')):
os.mkdir(Path('model'))
RETRAIN = False
MODEL_PATH = Path('model/MNIST_model.pt')
if not os.path.exists(MODEL_PATH) or RETRAIN:
cost = torch.nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters())
n_epochs = 10
for epoch in range(n_epochs):
running_loss = 0.0
running_correct = 0
print("Epoch {}/{}".format(epoch, n_epochs))
print("-"*10)
for data in data_loader_train:
X_train, y_train = data
if use_cuda:
X_train, y_train = X_train.cuda(), y_train.cuda()
outputs = F.softmax(model(X_train))
_,pred = torch.max(outputs.data, 1)
optimizer.zero_grad()
loss = cost(outputs, y_train)
loss.backward()
optimizer.step()
running_loss += loss.item()
running_correct += torch.sum(pred == y_train.data).item()
testing_correct = 0
for data in data_loader_test:
X_test, y_test = data
if use_cuda:
X_test, y_test = X_test.cuda(), y_test.cuda()
outputs = model(X_test)
_, pred = torch.max(outputs.data, 1)
testing_correct += torch.sum(pred == y_test.data).item()
print("Loss is:{:.8f}, Train Accuracy is:{:.2f}%, Test Accuracy is:{:.2f}%".format(running_loss*batch_size/len(data_train),
100*running_correct/len(data_train),
100*testing_correct/len(data_test)))
torch.save(model.state_dict(),MODEL_PATH)
else:
model.load_state_dict(torch.load(MODEL_PATH))
print ('model loaded!')
# + [markdown] id="ysqaU7GdT1Ow"
# # ICE Explainer for MNIST model
#
# Now, for the ICE Explanations.
# 1. Wrap the model. Set all related classes.
# 2. Create a loader list. Each loader only contains samples from one class.
# 3. Create the Explainer, train the reducer model and generate explanations.
#
#
# + [markdown] id="-gCn2ToQ1vgU"
# ## Parameters, data loader and model wrapper
# Some parameters required:
# * Target classes with correlated names (explanation domains): [3,8]
# * Concept targer layer: 'layer3'
# * n_components for NMF reducer: 8
# * Explainer title: MNIST_layer3_8_[3_8]
#
# + colab={"base_uri": "https://localhost:8080/"} id="4iqJ87INT1Ox" outputId="da2ae593-b91a-4dfa-fbb2-a3bdf37136db"
import ICE.ModelWrapper
import ICE.Explainer
import ICE.utils
import shutil
target_classes = [3,8]
classes_names = [str(i) for i in target_classes]
layer_name = 'layer3'
n_components = 6
title = "MNIST_{}_{}_[".format(layer_name,n_components)+"_".join(classes_names) + ']'
print ("title:{}".format(title))
print ("target_classes:{}".format(target_classes))
print ("classes_names:{}".format(classes_names))
print ("n_components:{}".format(n_components))
print ("layer_name:{}".format(layer_name))
model = model.cuda()
model.eval()
wm = ICE.ModelWrapper.PytorchModelWrapper(model,batch_size=batch_size,predict_target=target_classes,input_size = [1,28,28],input_channel_first = True,model_channel_first = True)
X,y = data_train.data,data_train.targets
loaders = []
for target in target_classes:
tdataset = torch.utils.data.TensorDataset(X[y==target].unsqueeze(1))
loaders.append(torch.utils.data.DataLoader(tdataset,batch_size=batch_size))
# + [markdown] id="vmNbuWhCz8Jt"
# ## ICE Explainer training
#
# Depends on the dataset size, the training may take several minutes. You can change the CALC_LIMIT value in Explainer.py to reduce the dataset size.
# + colab={"base_uri": "https://localhost:8080/"} id="mx5RQdcfT1Oy" outputId="5fd942ea-d5fc-4d00-faf1-b688ee13703a"
print ("-------------------------------------------------------------------------------------------")
try:
shutil.rmtree('Explainers/'+title)
except:
pass
# create an Explainer
Exp = ICE.Explainer.Explainer(title = title,
layer_name = layer_name,
class_names = classes_names,
utils = ICE.utils.img_utils(img_size = (28,28),nchannels=1,img_format='channels_first'),
n_components = n_components,
reducer_type = "NMF"
)
# train reducer based on target classes
Exp.train_model(wm,loaders)
# generate features
Exp.generate_features(wm, loaders)
# generate global explanations
Exp.global_explanations()
# save the explainer, use load() to load it with the same title
Exp.save()
# + [markdown] id="DijwXlx40MC9"
# Save and load exist explainers. You can load the explainer by the title.
# + id="muv1CZkRT1Oy"
# you can load exist explainers with title
Exp = ICE.Explainer.Explainer(title = title)
Exp.load()
# + [markdown] id="12xeavENT1Oy"
# ## Global explanations
#
# Global explanations are already generated before, you can compare the difference between those explanations of class 3 and 8.
#
# Here feature_0 is a commonly appear features in nearly all instances (3 can be considered as a part of 8).
#
# Number 8 has unique concepts like feature_2. For number 3, feature_3 may not appear in instances of number 8.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="YmiUA1FYT1Oy" outputId="917196ad-7668-4d7c-ec50-d646e5ff0c98"
from IPython.display import HTML, display, Image
import os
from pathlib import Path
#@title { display-mode: "form", run: "auto" }
fpath = Path('Explainers').absolute() / title / 'GE'
imgfile = '3.jpg' #@param ['3.jpg', '8.jpg']
display(Image(str(fpath/imgfile)))
# + [markdown] id="yfNSB-73T1Oz"
# ## Local Explanations
#
# Local explanations for a '3'.
#
# Let's get an instance from test set and locally explain it.
# + colab={"base_uri": "https://localhost:8080/", "height": 321} id="dkwUPajqT1Oz" outputId="273dc1e2-a23d-4b5e-9458-7218886a4081"
x = data_test.data[data_test.targets==3][5]
x = x.unsqueeze(0)
Exp.utils.show_img([Exp.utils.deprocessing(x.permute(0,2,1).numpy())])
x = x.numpy()
print (x.shape)
# + [markdown] id="6wFZouhF4fM7"
# Local explanation requires an instance and the wrapped model.
# + id="L3xoeKe7T1Oz"
Exp.local_explanations(x,wm,name='LE_3')
# + [markdown] id="7UmvyICr44bj"
# The last line in each explanation reports the real model's prediction and how much those predictions are displayed by this explanation. Here Total contribution is the score of restructed feature map from the reduced feature maps.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="Wm3tIU6WT1Oz" outputId="daf8091e-df01-4265-f1ae-67c042456850"
from IPython.display import HTML, display, Image
import os
from pathlib import Path
#@title { display-mode: "form", run: "auto" }
fpath = Path('Explainers').absolute() / title / 'explanations' / 'all'
imgfile = 'LE_3_0.jpg' #@param ['LE_3_0.jpg', 'LE_3_1.jpg']
display(Image(str(fpath/imgfile)))
# + id="kHDNhtGZT1O0"
# + id="kyIR0BwrT1O0"
| demo/MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Histogram collections
#
# **WARNING**: Experimental functionality that will probably be redesigned in version 0.5.
#
# **STATUS**: Very alpha
# +
import numpy as np
np.random.seed(42)
from physt import h1
from physt.histogram_collection import HistogramCollection
# -
from physt.plotting import matplotlib
from physt.plotting import set_default_backend
from physt.plotting import vega
from physt.plotting import matplotlib
set_default_backend("matplotlib")
# %matplotlib inline
data1 = np.random.normal(100, 15, 2000)
h_a = h1(data1, "fixed_width", 10, name="first")
h_a.plot();
data2 = np.random.normal(80, 10, 2000)
h_b = h1(data2, h_a.binning, name="second")
h_b.plot();
collection = HistogramCollection(h_a, h_b, title="Combination")
collection.create("third", np.random.normal(148, 5, 300))
# ## Plotting in matplotlib
# The default
collection.plot();
# Add some options
collection.plot.line(alpha=.5, lw=8, xlabel="Index");
# ## Plotting in vega
set_default_backend("vega")
collection.plot.scatter(legend=False)
collection.plot.line(lw=7, legend=True, alpha=.5)
| doc/Collections.ipynb |
# This cell is mandatory in all Dymos documentation notebooks.
missing_packages = []
try:
import openmdao.api as om
except ImportError:
if 'google.colab' in str(get_ipython()):
# !python -m pip install openmdao[notebooks]
else:
missing_packages.append('openmdao')
try:
import dymos as dm
except ImportError:
if 'google.colab' in str(get_ipython()):
# !python -m pip install dymos
else:
missing_packages.append('dymos')
try:
import pyoptsparse
except ImportError:
if 'google.colab' in str(get_ipython()):
# !pip install -q condacolab
import condacolab
condacolab.install_miniconda()
# !conda install -c conda-forge pyoptsparse
else:
missing_packages.append('pyoptsparse')
if missing_packages:
raise EnvironmentError('This notebook requires the following packages '
'please install them and restart this notebook\'s runtime: {",".join(missing_packages)}')
# # Water Rocket
#
# Author: <NAME> (<EMAIL>)
#
# In this example, we will optimize a water rocket for range and height at the apogee, using design variables that are easily modifiable just before launch: the empty mass, the initial water volume and the launch angle.
# This example builds on [multi-phase cannonball](../multi_phase_cannonball/multi_phase_cannonball.md) ane is adapted from _Optimization of a Water Rocket in OpenMDAO/Dymos_ {cite}`bbahia_2020`.
#
# ## Nomenclature
#
# | Symbol | definition |
# |----------------------|----------------------------------------|
# | $v_\text{out}$ | water exit speed at the nozzle |
# | $A_\text{out}$ | nozzle area |
# | $V_w$ | water volume in the rocket |
# | $p$ | pressure in the rocket |
# | $p_a$ | ambient pressure |
# | $\dot{\,}$ | time derivative |
# | $k$ | polytropic constant |
# | $V_b$ | internal volume of the rocket |
# | $\rho_w$ | water density |
# | $T$ | thrust |
# | $q$ | dynamic pressure |
# | $S$ | cross sectional area |
# | $(\cdot)_0$ | value of $(\cdot)$ at $t=0$ |
# | $t$ | time |
# ## Problem Formulation
#
# A natural objective function for a water rocket is the maximum height achieved by the rocket during flight, or the horizontal distance it travels, i.e. its range.
# The design of a water rocket is somewhat constrained by the soda bottle used as its engine.
# This means that the volume available for water and air is fixed, the initial launch pressure is limited by the bottle's strength (since the pressure is directly related to the energy available for the rocket, it is easy to see that it should be as high as possible) and the nozzle throat area is also fixed.
# Given these manufacturing constraints, the design variables we are left with are the empty mass (it can be easily changed through adding ballast), the water volume at the launch, and the launch angle.
# With this considerations in mind, a natural formulation for the water rocket problem is
#
# \begin{align}
# \text{maximize} &\quad \text{range or height} \\
# \text{w.r.t.} &\quad \text{empty mass, initial water volume, launch angle, trajectory} \\
# \text{subject to} &\quad \text{flight dynamics} \\
# &\quad \text{fluid dynamics inside the rocket} \\
# &\quad 0 < \text{initial water volume} < \text{volume of bottle} \\
# &\quad 0^\circ < \text{launch angle} < 90^\circ \\
# &\quad 0 < \text{empty mass}
# \end{align}
# ## Model
#
# The water rocket model is divided into three basic components: a *water engine*, responsible for modelling the fluid dynamics inside the rocket and returning its thrust; the *aerodynamics*, responsible for calculating the atmospheric drag of the rocket; and the *equations of motion*, responsible for propagating the rocket's trajectory in time, using Newton's laws and the forces provided by the other two components.
#
# In order to integrate these three basic components, some additional interfacing components are necessary: an atmospheric model to provide values of ambient pressure for the water engine and air density to the calculation of the dynamic pressure for the aerodynamic model, and a component that calculates the instantaneous mass of the rocket by summing the water mass with the rocket's empty mass.
# The high level layout of this model is shown in below.
#
# <figure>
# <img src="figures/water_rocket_overview.svg"/>
# <figcaption>N2 diagram for the water rocket model</figcaption>
# </figure>
#
# `atmos`, `dynamic_pressure`, `aero` and `eom` are the same models used in [multi-phase cannonball](../multi_phase_cannonball/multi_phase_cannonball.md).
# The remaining components are discussed below.
#
# ```{Warning}
# The `eom` component has a singularity in the flight path angle derivative when the flight speed is zero.
# This happens because the rotational dynamics are not modelled.
# This can cause convergence problems if the launch velocity is set to zero or the launch angle is set to $90^\circ$
# ```
#
# ```{Note}
# Since the range of altitudes achieved by the water rocket is very small (100m), the air density and pressure are practically constant, thus the use of an atmospheric model is not necessary. However, using it makes it easier to reuse code from [multi-phase cannonball (../multi_phase_cannonball/multi_phase_cannonball.md).
# ```
#
# ### Water engine
#
# The water engine is modelled by assuming that the air expansion in the rocket
# follows an adiabatic process and the water flow is incompressible and inviscid,
# i.e. it follows Bernoulli's equation. We also make the following simplifying
# assumptions:
#
# 1. The thrust developed after the water is depleted is negligible
# 2. The area inside the bottle is much smaller than the nozzle area
# 3. The inertial forces do not affect the fluid dynamics inside the bottle
#
# This simplified modelling can be found in Prusa[@Prusa2000].
# A more rigorous formulation, which drops all these simplifying assumptions can be found in Wheeler[@Wheeler2002], Gommes[@Gommes2010], and Barria-Perotti[@BarrioPerotti2010].
#
# The first assumption leads to an underestimation of the rocket performance, since the air left in the bottle after it is out of water is known to generate appreciable thrust[@Thorncroft2009].
# This simplified model, however, produces physically meaningful results.
#
# There are two states in this dynamical model, the water volume in the rocket $V_w$ and the gauge pressure inside the rocket $p$.
# The constitutive equations and the N2 diagram showing the model organization are shown below.
#
# ### Constitutive equations of the water engine model
# | Component | Equation |
# | -----------------------|-------------------------------------------------------------|
# | water_exhaust_speed | $v_\text{out} = \sqrt{2(p-p_a)/\rho_w}$ |
# | water_flow_rate | $\dot{V}_w = -v_\text{out} A_\text{out}$ |
# | pressure_rate | $\dot{p} = kp\frac{\dot{V_w}}{(V_b-V_w)}$ |
# | water_thrust | $T = (\rho_w v_\text{out})(v_\text{out}A_\text{out})$ |
#
# <figure>
# <img src="figures/water_rocket_waterengine.svg"/>
# <figcaption>N2 diagram for the water engine group</figcaption>
# </figure>
#
# ### Water engine
#
# The water engine is modelled by assuming that the air expansion in the rocket
# follows an adiabatic process and the water flow is incompressible and inviscid,
# i.e. it follows Bernoulli's equation. We also make the following simplifying
# assumptions:
#
# 1. The thrust developed after the water is depleted is negligible
# 2. The area inside the bottle is much smaller than the nozzle area
# 3. The inertial forces do not affect the fluid dynamics inside the bottle
#
# This simplified modelling can be found in Prusa {cite}`Prusa2000`.
# A more rigorous formulation, which drops all these simplifying assumptions can be found in Wheeler {cite}`Wheeler2002`, Gommes {cite}`Gommes2010`, and Barria-Perotti {cite}`BarrioPerotti2010`.
#
# The first assumption leads to an underestimation of the rocket performance, since the air left in the bottle after it is out of water is known to generate appreciable thrust {cite}`Thorncroft2009`.
# This simplified model, however, produces physically meaningful results.
#
# There are two states in this dynamical model, the water volume in the rocket $V_w$ and the gauge pressure inside the rocket $p$.
# The constitutive equations and the N2 diagram showing the model organization are shown below.
#
# ### Constitutive equations of the water engine model
# | Component | Equation |
# | -----------------------|-------------------------------------------------------------|
# | water_exhaust_speed | $v_\text{out} = \sqrt{2(p-p_a)/\rho_w}$ |
# | water_flow_rate | $\dot{V}_w = -v_\text{out} A_\text{out}$ |
# | pressure_rate | $\dot{p} = kp\frac{\dot{V_w}}{(V_b-V_w)}$ |
# | water_thrust | $T = (\rho_w v_\text{out})(v_\text{out}A_\text{out})$ |
#
# <figure>
# <img src="figures/water_rocket_waterengine.svg"/>
# <figcaption>N2 diagram for the water engine group</figcaption>
# </figure>
# +
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import openmdao.api as om
import dymos as dm
# -
from openmdao.utils.notebook_utils import display_source
display_source('dymos.examples.water_rocket.water_engine_comp.WaterEngine')
display_source('dymos.examples.water_rocket.water_engine_comp._WaterExhaustSpeed')
display_source('dymos.examples.water_rocket.water_engine_comp._PressureRate')
display_source('dymos.examples.water_rocket.water_engine_comp._WaterFlowRate')
# The `_MassAdder` component calculates the rocket's instantaneous mass by
# summing the water mass with the rockets empty mass, i.e.
#
# \begin{align}
# m = m_\text{empty}+\rho_w V_w
# \end{align}
display_source('dymos.examples.water_rocket.water_propulsion_ode._MassAdder')
# Now these components are joined in a single group
display_source('dymos.examples.water_rocket.water_propulsion_ode.WaterPropulsionODE')
# ## Phases
#
# The flight of the water rocket is split in three distinct phases: propelled ascent, ballistic ascent and ballistic descent.
# If the simplification of no thrust without water were lifted, there would be an extra "air propelled ascent" phase between the propelled ascent and ballistic ascent phases.
#
# **Propelled ascent:** is the flight phase where the rocket still has water inside, and hence it is producing thrust.
# The thrust is given by the water engine model, and fed into the flight dynamic equations.
# It starts at launch and finishes when the water is depleted, i.e. $V_w=0$.
#
# **Ballistic ascent:** is the flight phase where the rocket is ascending ($\gamma>0$) but produces no thrust.
# This phase begins at the end of thepropelled ascent phase and ends at the apogee, defined by $\gamma=0$.
#
# **Descent:** is the phase where the rocket is descending without thrust.
# It begins at the end of the ballistic ascent phase and ends with ground impact, i.e. $h=0$.
display_source('dymos.examples.water_rocket.phases.new_propelled_ascent_phase')
display_source('dymos.examples.water_rocket.phases.new_ballistic_ascent_phase')
display_source('dymos.examples.water_rocket.phases.new_descent_phase')
# ## Model parameters
#
# The model requires a few constant parameters.
# The values used are shown in the following table.
#
# Values for parameters in the water rocket model
#
# | Parameter | Value | Unit | Reference |
# |--------------------|----------------------|--------------|-----------------------------------------------------|
# | $C_D$ | 0.3450 | - | {cite}`BarrioPerotti2009` |
# | $S$ | $\pi 106^2/4$ | $mm^2$ | {cite}`BarrioPerotti2009` |
# | $k$ | 1.2 | - | {cite}`Thorncroft2009` {cite}`Fischer2020` {cite}`Romanelli2013` |
# | $A_\text{out}$ | $\pi22^2/4$ | $mm^2$ | {cite}`aircommand_nozzle` |
# | $V_b$ | 2 | L | |
# | $\rho_w$ | 1000 | $kg/m^3$ | |
# | $p_0$ | 6.5 | bar | |
# | $v_0$ | 0.1 | $m/s$ | |
# | $h_0$ | 0 | $m$ | |
# | $r_0$ | 0 | $m$ | |
#
# Values for the bottle volume $V_b$, its cross-sectional area $S$ and the nozzle area $A_\text{out}$ are determined by the soda bottle that makes the rocket primary structure, and thus are not easily modifiable by the designer.
# The polytropic coefficient $k$ is a function of the moist air characteristics inside the rocket.
# The initial speed $v_0$ must be set to a value higher than zero, otherwise the flight dynamic equations become singular.
# This issue arises from the angular dynamics of the rocket not being modelled.
# The drag coefficient $C_D$ is sensitive to the aerodynamic design, but can be optimized by a single discipline analysis.
# The initial pressure $p_0$ should be maximized in order to obtain the maximum range or height for the rocket.
# It is limited by the structural properties of the bottle, which are modifiable by the designer, since the bottle needs to be available commercially.
# Finally, the starting point of the rocket is set to the origin.
#
# ## Putting it all together
#
# The different phases must be combined in a single trajectory, and linked in a sequence.
# Here we also define the design variables.
display_source('dymos.examples.water_rocket.phases.new_water_rocket_trajectory')
# ## Helper Functions to Access the Results
# +
from collections import namedtuple
def summarize_results(water_rocket_problem):
p = water_rocket_problem
Entry = namedtuple('Entry', 'value unit')
summary = {
'Launch angle': Entry(p.get_val('traj.propelled_ascent.timeseries.states:gam', units='deg')[0, 0], 'deg'),
'Flight angle at end of propulsion': Entry(p.get_val('traj.propelled_ascent.timeseries.states:gam',
units='deg')[-1, 0], 'deg'),
'Empty mass': Entry(p.get_val('traj.parameters:m_empty', units='kg')[0], 'kg'),
'Water volume': Entry(p.get_val('traj.propelled_ascent.timeseries.states:V_w', 'L')[0, 0], 'L'),
'Maximum range': Entry(p.get_val('traj.descent.timeseries.states:r', units='m')[-1, 0], 'm'),
'Maximum height': Entry(p.get_val('traj.ballistic_ascent.timeseries.states:h', units='m')[-1, 0], 'm'),
'Maximum velocity': Entry(p.get_val('traj.propelled_ascent.timeseries.states:v', units='m/s')[-1, 0], 'm/s'),
}
return summary
# +
colors={'pa': 'tab:blue', 'ba': 'tab:orange', 'd': 'tab:green'}
def plot_propelled_ascent(p, exp_out):
fig, ax = plt.subplots(2, 2, sharex=True, figsize=(12, 6))
t_imp = p.get_val('traj.propelled_ascent.time', 's')
t_exp = exp_out.get_val('traj.propelled_ascent.time', 's')
c = colors['pa']
ax[0,0].plot(t_imp, p.get_val('traj.propelled_ascent.timeseries.states:p', 'bar'), '.', color=c)
ax[0,0].plot(t_exp, exp_out.get_val('traj.propelled_ascent.timeseries.states:p', 'bar'), '-', color=c)
ax[0,0].set_ylabel('p (bar)')
ax[0,0].set_ylim(bottom=0)
ax[1,0].plot(t_imp, p.get_val('traj.propelled_ascent.timeseries.states:V_w', 'L'), '.', color=c)
ax[1,0].plot(t_exp, exp_out.get_val('traj.propelled_ascent.timeseries.states:V_w', 'L'), '-', color=c)
ax[1,0].set_ylabel('$V_w$ (L)')
ax[0,1].plot(t_imp, p.get_val('traj.propelled_ascent.timeseries.T', 'N'), '.', color=c)
ax[0,1].plot(t_exp, exp_out.get_val('traj.propelled_ascent.timeseries.T', 'N'), '-', color=c)
ax[0,1].set_ylabel('T (N)')
ax[0,1].set_ylim(bottom=0)
ax[1,1].plot(t_imp, p.get_val('traj.propelled_ascent.timeseries.states:v', 'm/s'), '.', color=c)
ax[1,1].plot(t_exp, exp_out.get_val('traj.propelled_ascent.timeseries.states:v', 'm/s'), '-', color=c)
ax[1,1].set_ylabel('v (m/s)')
ax[1,1].set_ylim(bottom=0)
ax[1,0].set_xlabel('t (s)')
ax[1,1].set_xlabel('t (s)')
for i in range(4):
ax.ravel()[i].grid(True, alpha=0.2)
fig.tight_layout()
# -
def plot_states(p, exp_out, legend_loc='right', legend_ncol=3):
fig, axes = plt.subplots(nrows=2, ncols=2, figsize=(12, 4), sharex=True)
states = ['r', 'h', 'v', 'gam']
units = ['m', 'm', 'm/s', 'deg']
phases = ['propelled_ascent', 'ballistic_ascent', 'descent']
time_imp = {'ballistic_ascent': p.get_val('traj.ballistic_ascent.timeseries.time'),
'propelled_ascent': p.get_val('traj.propelled_ascent.timeseries.time'),
'descent': p.get_val('traj.descent.timeseries.time')}
time_exp = {'ballistic_ascent': exp_out.get_val('traj.ballistic_ascent.timeseries.time'),
'propelled_ascent': exp_out.get_val('traj.propelled_ascent.timeseries.time'),
'descent': exp_out.get_val('traj.descent.timeseries.time')}
x_imp = {phase: {state: p.get_val(f"traj.{phase}.timeseries.states:{state}", unit)
for state, unit in zip(states, units)
}
for phase in phases
}
x_exp = {phase: {state: exp_out.get_val(f"traj.{phase}.timeseries.states:{state}", unit)
for state, unit in zip(states, units)
}
for phase in phases
}
for i, (state, unit) in enumerate(zip(states, units)):
axes.ravel()[i].set_ylabel(f"{state} ({unit})" if state != 'gam' else f'$\gamma$ ({unit})')
axes.ravel()[i].plot(time_imp['propelled_ascent'], x_imp['propelled_ascent'][state], '.', color=colors['pa'])
axes.ravel()[i].plot(time_imp['ballistic_ascent'], x_imp['ballistic_ascent'][state], '.', color=colors['ba'])
axes.ravel()[i].plot(time_imp['descent'], x_imp['descent'][state], '.', color=colors['d'])
h1, = axes.ravel()[i].plot(time_exp['propelled_ascent'], x_exp['propelled_ascent'][state], '-', color=colors['pa'], label='Propelled Ascent')
h2, = axes.ravel()[i].plot(time_exp['ballistic_ascent'], x_exp['ballistic_ascent'][state], '-', color=colors['ba'], label='Ballistic Ascent')
h3, = axes.ravel()[i].plot(time_exp['descent'], x_exp['descent'][state], '-', color=colors['d'], label='Descent')
if state == 'gam':
axes.ravel()[i].yaxis.set_major_locator(mpl.ticker.MaxNLocator(nbins='auto', steps=[1, 1.5, 3, 4.5, 6, 9, 10]))
axes.ravel()[i].set_yticks(np.arange(-90, 91, 45))
axes.ravel()[i].grid(True, alpha=0.2)
axes[1, 0].set_xlabel('t (s)')
axes[1, 1].set_xlabel('t (s)')
plt.figlegend(handles=[h1, h2, h3], loc=legend_loc, ncol=legend_ncol)
fig.tight_layout()
def plot_trajectory(p, exp_out, legend_loc='center right'):
fig, axes = plt.subplots(nrows=1, ncols=1, figsize=(8, 6))
time_imp = {'ballistic_ascent': p.get_val('traj.ballistic_ascent.timeseries.time'),
'propelled_ascent': p.get_val('traj.propelled_ascent.timeseries.time'),
'descent': p.get_val('traj.descent.timeseries.time')}
time_exp = {'ballistic_ascent': exp_out.get_val('traj.ballistic_ascent.timeseries.time'),
'propelled_ascent': exp_out.get_val('traj.propelled_ascent.timeseries.time'),
'descent': exp_out.get_val('traj.descent.timeseries.time')}
r_imp = {'ballistic_ascent': p.get_val('traj.ballistic_ascent.timeseries.states:r'),
'propelled_ascent': p.get_val('traj.propelled_ascent.timeseries.states:r'),
'descent': p.get_val('traj.descent.timeseries.states:r')}
r_exp = {'ballistic_ascent': exp_out.get_val('traj.ballistic_ascent.timeseries.states:r'),
'propelled_ascent': exp_out.get_val('traj.propelled_ascent.timeseries.states:r'),
'descent': exp_out.get_val('traj.descent.timeseries.states:r')}
h_imp = {'ballistic_ascent': p.get_val('traj.ballistic_ascent.timeseries.states:h'),
'propelled_ascent': p.get_val('traj.propelled_ascent.timeseries.states:h'),
'descent': p.get_val('traj.descent.timeseries.states:h')}
h_exp = {'ballistic_ascent': exp_out.get_val('traj.ballistic_ascent.timeseries.states:h'),
'propelled_ascent': exp_out.get_val('traj.propelled_ascent.timeseries.states:h'),
'descent': exp_out.get_val('traj.descent.timeseries.states:h')}
axes.plot(r_imp['propelled_ascent'], h_imp['propelled_ascent'], 'o', color=colors['pa'])
axes.plot(r_imp['ballistic_ascent'], h_imp['ballistic_ascent'], 'o', color=colors['ba'])
axes.plot(r_imp['descent'], h_imp['descent'], 'o', color=colors['d'])
h1, = axes.plot(r_exp['propelled_ascent'], h_exp['propelled_ascent'], '-', color=colors['pa'], label='Propelled Ascent')
h2, = axes.plot(r_exp['ballistic_ascent'], h_exp['ballistic_ascent'], '-', color=colors['ba'], label='Ballistic Ascent')
h3, = axes.plot(r_exp['descent'], h_exp['descent'], '-', color=colors['d'], label='Descent')
axes.set_xlabel('r (m)')
axes.set_ylabel('h (m)')
axes.set_aspect('equal', 'box')
plt.figlegend(handles=[h1, h2, h3], loc=legend_loc)
axes.grid(alpha=0.2)
fig.tight_layout()
# ## Optimizing for Height
# +
from dymos.examples.water_rocket.phases import new_water_rocket_trajectory, set_sane_initial_guesses
p = om.Problem(model=om.Group())
traj, phases = new_water_rocket_trajectory(objective='height')
traj = p.model.add_subsystem('traj', traj)
p.driver = om.pyOptSparseDriver(optimizer='IPOPT', print_results=False)
p.driver.opt_settings['print_level'] = 4
p.driver.opt_settings['max_iter'] = 1000
p.driver.opt_settings['mu_strategy'] = 'monotone'
p.driver.declare_coloring(tol=1.0E-12)
# Finish Problem Setup
p.model.linear_solver = om.DirectSolver()
p.setup()
set_sane_initial_guesses(p, phases)
dm.run_problem(p, run_driver=True, simulate=True)
summary = summarize_results(p)
for key, entry in summary.items():
print(f'{key}: {entry.value:6.4f} {entry.unit}')
sol_out = om.CaseReader('dymos_solution.db').get_case('final')
sim_out = om.CaseReader('dymos_simulation.db').get_case('final')
# -
# ### Maximum Height Solution: Propulsive Phase
plot_propelled_ascent(sol_out, sim_out)
# ## Maximum Height Solution: Height vs. Range
#
# Note that the equations of motion used here are singular in vertical flight, so the launch angle (the initial flight path angle) was limited to 85 degrees.
plot_trajectory(sol_out, sim_out, legend_loc='center right')
# ## Maximum Height Solution: State History
plot_states(sol_out, sim_out, legend_loc='lower center', legend_ncol=3)
# +
from openmdao.utils.assert_utils import assert_near_equal
assert_near_equal(summary['Launch angle'].value, 85, 0.01)
assert_near_equal(summary['Empty mass'].value, 0.144, 0.01)
assert_near_equal(summary['Water volume'].value, 0.98, 0.01)
assert_near_equal(summary['Maximum height'].value, 53.5, 0.01)
# -
# # Optimizing for Range
# +
from dymos.examples.water_rocket.phases import new_water_rocket_trajectory, set_sane_initial_guesses
p = om.Problem(model=om.Group())
traj, phases = new_water_rocket_trajectory(objective='range')
traj = p.model.add_subsystem('traj', traj)
p.driver = om.pyOptSparseDriver(optimizer='IPOPT')
p.driver.opt_settings['print_level'] = 4
p.driver.opt_settings['max_iter'] = 1000
p.driver.opt_settings['mu_strategy'] = 'monotone'
p.driver.declare_coloring(tol=1.0E-12)
# Finish Problem Setup
p.model.linear_solver = om.DirectSolver()
p.setup()
set_sane_initial_guesses(p, phases)
dm.run_problem(p, run_driver=True, simulate=True)
summary = summarize_results(p)
for key, entry in summary.items():
print(f'{key}: {entry.value:6.4f} {entry.unit}')
sol_out = om.CaseReader('dymos_solution.db').get_case('final')
sim_out = om.CaseReader('dymos_simulation.db').get_case('final')
# -
# ## Maximum Range Solution: Propulsive Phase
plot_propelled_ascent(sol_out, sim_out)
# ## Maximum Range Solution: Height vs. Range
plot_trajectory(sol_out, sim_out, legend_loc='center')
# ## Maximum Range Solution: State History
plot_states(sol_out, sim_out, legend_loc='lower center')
# +
from openmdao.utils.assert_utils import assert_near_equal
# Check results (tolerance is relative unless value is zero)
assert_near_equal(summary['Launch angle'].value, 46, 0.02)
assert_near_equal(summary['Flight angle at end of propulsion'].value, 38, 0.02)
assert_near_equal(summary['Empty mass'].value, 0.189, 1e-2)
assert_near_equal(summary['Water volume'].value, 1.026, 1e-2)
assert_near_equal(summary['Maximum range'].value, 85.11, 1e-2)
assert_near_equal(summary['Maximum height'].value, 23.08, 1e-2)
assert_near_equal(summary['Maximum velocity'].value, 41.31, 1e-2)
# -
# ## References
#
# ```{bibliography}
# :filter: docname in docnames
# ```
| docs/dymos_book/examples/water_rocket/water_rocket.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Setup, matplotlib inline, automatically reload libraries on every evaluation
import pandas as pd
import numpy as np
import scipy as sp
pd.options.display.max_rows = 400
pd.options.display.max_columns = 400
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
from epiml.loadepiml import LoadEpiml, load_search, save_search
from epiml.epimlsklearn.epimlmetrics import pu_scorer, prior_squared_error_scorer_015, brier_score_labeled_loss_scorer, \
f1_assumed_scorer, f1_labeled_scorer, report_metrics, f1_assumed_beta10_scorer, pu_mix_assumed_f1beta10_scorer
from epiml.semisuperhelper import SemiSupervisedHelper
from epiml.epimlsklearn.pnuwrapper import PNUWrapper
from epiml.epimlsklearn.jsearchcv import JRandomSearchCV, extract_score_grid
from epiml.epimlsklearn.nestedcross import NestedCV, rerun_nested_for_scoring
from epiml.epimlsklearn.frankenscorer import FrankenScorer, extract_scores_from_nested
from epiml.epimlsklearn.repeatedsampling import RepeatedRandomSubSampler
path = "C:\\Data\\010317\\membership14_final_0103.txt"
lc = LoadEpiml(path)
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from sklearn.base import clone
X_train, X_test, y_train, y_test = train_test_split(lc.X, lc.y, test_size=0.2, random_state=771, stratify=lc.y)
# # REPEATED SUB SAMPLER
# ## Set up PNU Wrapper with Random Forest, then JSearchCV, then NestedCV
rf = RandomForestClassifier()
rep = RepeatedRandomSubSampler(base_estimator=rf, voting='thresh', verbose=1, random_state=83)
pnu = PNUWrapper(base_estimator=rep, num_unlabeled=1.0)
# ## Set up randomized search parameters
rf_param_search = {'base_estimator__base_estimator__bootstrap': [True, False],
'base_estimator__base_estimator__class_weight': [None,'balanced','balanced_subsample'],
'base_estimator__base_estimator__criterion': ['gini','entropy'],
'base_estimator__base_estimator__max_depth': [None] + list(range(2,100)),
'base_estimator__base_estimator__max_features': ['sqrt','log2',None] + list(range(5,100)),
'base_estimator__base_estimator__min_samples_leaf': [1,2,3,4,5,6,7,8,9,10,15,20,25,30,35,40,45,50,75,100],
'base_estimator__base_estimator__min_samples_split':[2,0.005,0.01,0.015,0.02,0.025,0.03,0.035,0.04,
0.045,0.05,0.07,0.09,0.1,0.12,0.15,0.17,0.2,0.25],
'base_estimator__base_estimator__n_estimators': sp.stats.randint(low=10, high=300),
'base_estimator__sample_imbalance': sp.stats.uniform(loc=0.1, scale=0.9),
'pu_learning': [True, False]
}
# ### notice random_state is set in jsearch, this is so that the same random parameters are searched for each outer fold, sort of like grid search
jsearch = JRandomSearchCV(pnu, rf_param_search, n_iter=60, scoring=FrankenScorer(decision_score='assumed_f1beta10'),
n_jobs=-1, cv=3, verbose=1, pre_dispatch=8, random_state=77)
# #### 3x3 (x60) nested cross validation
nested_cross = NestedCV(jsearch, scoring=FrankenScorer(decision_score='pu_mix_assumed_f1beta10'), cv=3, random_state=77)
# ## Score the nested cross
scores = nested_cross.score(X_train.values, y=y_train.values, verbose=100, pre_dispatch=8)
save_search(nested_cross, './res/nested_cross_repreated_rf_large.pkl')
nested_cross = load_search('./res/nested_cross_repreated_rf_large.pkl')
extract_scores_from_nested(nested_cross.test_score_datas_).mean().sort_index()
nested_cross.best_idxs_
nested_cross.best_params_
# # Now that we have comparable, nested search scores, lets do a search on the whole 80% training set and use the parameters found
jsearch_train = JRandomSearchCV(pnu, rf_param_search, n_iter=60, scoring=FrankenScorer(decision_score='pu_mix_assumed_f1beta10'),
n_jobs=-1, cv=3, verbose=1, pre_dispatch=8, random_state=77)
jsearch_fit = jsearch_train.fit(X_train.values, y_train.values)
save_search(jsearch_fit, './res/jsearch_only.pkl')
jsearch_score_grid = extract_score_grid(jsearch_fit)
best_idx = jsearch_score_grid.mean_pu_mix_assumed_f1beta10_test.idxmax()
model6_params = pd.DataFrame(jsearch_fit.cv_results_).params[best_idx]
model6_params
# ## This is model 3 in the manuscript, lets retrain on whole training set and then test on 20% test set!
pnu_test = clone(pnu)
pnu_test.set_params(**model6_params)
pnu_test.fit(X_train.values, y_train.values)
FrankenScorer()(pnu_test, X_test.values, y_test.values)
# # Feature importance for 80/20
importance = pd.DataFrame(pnu_test.feature_importances_, index=X_test.columns.values, columns=['Importance']).sort_values(by='Importance', ascending=False)
importance.round(5) * 100
ax = importance.iloc[:20].iloc[::-1].plot(kind='barh')
ax.legend(loc='right')
# # Probability distribution of the 20%
def generate_probability_table(clf, X_test, y_test):
probas = clf.predict_proba(X_test)[:, -1]
probas_df = pd.DataFrame(data={'probas':probas, 'y_test':y_test.values}).sort_values(by='probas', ascending=False)
bins = np.linspace(0.0, 1.0, 101)
percent = pd.cut(probas_df['probas'], bins=bins, include_lowest=True, precision=6, labels=list(range(0,100)))
probas_df['percent'] = percent
dummies = pd.get_dummies(probas_df['y_test'], prefix='y=', prefix_sep='')
probas_df = pd.concat([probas_df, dummies], axis=1)
probas_group = probas_df.groupby('percent')
percentile_df = probas_group.aggregate({'probas':'count', 'y=-1':'sum', 'y=0':'sum', 'y=1':'sum'})
labeled_tot = percentile_df['y=1'] + percentile_df['y=0']
percentile_df['unlabeled_pct'] = percentile_df['y=-1'] / percentile_df.probas
percentile_df['true_pos_pct'] = percentile_df['y=1'] / labeled_tot
percentile_df['true_neg_pct'] = percentile_df['y=0'] / labeled_tot
tot = percentile_df.probas.sum()
percentile_df['pct_of_total'] = percentile_df.probas / tot
percentile_df['cum_pct_of_total'] = percentile_df.pct_of_total.cumsum()
percentile_df = percentile_df.ffill()
percentile_df = percentile_df.reset_index()
tp = np.polyfit(x=percentile_df.index.values.astype(int), y=percentile_df.true_pos_pct, deg=1)
tpp = np.poly1d(tp)
percentile_df['tp_trendline'] = tpp(percentile_df.index.values.astype(int))
tn = np.polyfit(x=percentile_df.index.values.astype(int), y=percentile_df.true_neg_pct, deg=1)
tnp = np.poly1d(tn)
percentile_df['tn_trendline'] = tnp(percentile_df.index.values.astype(int))
return probas_df, percentile_df
probas_df, percentile_df = generate_probability_table(pnu_test, X_test, y_test)
probas_df['probas'].plot.hist(bins=100)
probas_df.to_csv('./res/predicted_probabilities_soft.csv')
import matplotlib.pyplot as plt
df = percentile_df
x_axis = df.index.values.astype(int)
plt.figure(figsize=(10,5))
plt.plot(x_axis, df.pct_of_total, 'bs', label='N')
plt.plot(x_axis, df.cum_pct_of_total, 'b^', label='Cumulative N')
plt.plot(x_axis, df.tp_trendline, 'g-', label='% EPI Trend')
plt.plot(x_axis, df.tn_trendline, 'r-', label='% Non-EPI Trend')
plt.plot(x_axis, df.true_pos_pct, 'g.', label='Labeled EPI %')
plt.plot(x_axis, df.true_neg_pct, 'r.', label='Labeled Non-EPI %')
plt.axis([0, 100, -0.01, 1.01])
plt.ylabel('% of test set')
plt.xlabel('predicted probability of EPI')
plt.title('20% test set results')
plt.legend(loc='right')
# # Precision Recall curve for various thresholds used for Model 3
from sklearn.metrics import precision_recall_curve
labeled_probas = probas_df[probas_df.y_test >= 0]
pr, re, th = precision_recall_curve(labeled_probas.y_test, labeled_probas.probas, pos_label=1)
# find pr of unlabeled == pos
unlabeled_probas = probas_df[probas_df.y_test == -1].probas.values
total_unlabeled = len(unlabeled_probas)
pr_one_un = []
for thresh in th:
p = sum(unlabeled_probas >= thresh)
pr_one_un.append(p / total_unlabeled)
pr_one_un.append(0.0)
pr_one_un = np.array(pr_one_un)
def find_re_pr_prob(thresh):
idx = np.nonzero(th >= thresh)[0][0]
return re[idx], pr[idx], pr_one_un[idx]
re_50, pr_50, _ = find_re_pr_prob(0.5)
re_35, pr_35, _ = find_re_pr_prob(0.35)
re_40, pr_40, _ = find_re_pr_prob(0.4)
re_80, pr_80, _ = find_re_pr_prob(0.8)
re_20, pr_20, _ = find_re_pr_prob(0.2)
from matplotlib import cm, colors, colorbar
plt.clf()
plt.figure(figsize=(10, 7))
plt.xlabel('Recall', fontsize=15)
plt.xlim((0.0, 1.0))
plt.ylabel('Precision', fontsize=15)
plt.ylim((0.45, 1.05))
plt.title('Precision-Recall curve of labeled 20% test set', fontsize=15)
norm = colors.Normalize(vmin=0.0, vmax=0.2)
cmap = cm.plasma
plt.scatter(re, pr, c=pr_one_un, cmap=cmap, norm=norm, edgecolors='none', s=30)
#plt.scatter(re, pr, c=np.append(th, 1.0), cmap=cm.plasma, norm=norm)
cb = plt.colorbar(orientation='vertical', extend='max')
cb.set_label('Probability of unlabeled positive', fontsize=15)
plt.axvline(x=re_80, ls="--", c='r', label='Threshold=80%')
plt.axvline(x=re_50, ls="--", c='g', label='Threshold=50%')
plt.axvline(x=re_40, ls="--", c='orange', label='Threshold=40%')
plt.axvline(x=re_35, ls="--", c='b', label='Threshold=35%')
plt.axvline(x=re_20, ls="--", c='y', label='Threshold=20%')
plt.legend(loc='center left')
plt.show()
| 3.5 - RF - PNU Repeated Random Subsampling Random Search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import altair as alt
# Enable altair's interactive visualization in jupyter notebook
alt.renderers.enable('notebook')
# Enable altair's local data server so notebook size doesn't get too large
alt.data_transformers.enable('data_server')
alt.themes.enable('opaque')
df = pd.read_csv('data/master.csv')
df.head(1)
summer_df = df[df['Season'] == 'Summer']
winter_df = df[df['Season'] == 'Winter']
summer_medal = summer_df[summer_df['Medal'].notnull()]
summer_medal_sorted = summer_medal.groupby(['Name', 'region'])['Medal'].size().reset_index(name='count_medals').sort_values(by='count_medals', ascending=False)
summer_medal_sorted.head(10)
summer_medal_top = summer_medal_sorted.head(25)
alt.Chart(summer_medal_top).mark_bar().encode(
alt.X('Name:N', sort=alt.Sort(field='count_medals', op='sum', order='descending')),
alt.Y('count_medals:Q'),
alt.Color('region:N')
)
basketball_medal = summer_medal[summer_medal['Sport'] == 'Basketball']
basketball_medal['Event'].unique()
men_basketball_medal = basketball_medal[basketball_medal['Event'] == "Basketball Men's Basketball"]
women_basketball_medal = basketball_medal[basketball_medal['Event'] == "Basketball Women's Basketball"]
chart1 = alt.Chart(men_basketball_medal).mark_rect().encode(
alt.Y('region:N', title='Country'),
alt.X('Year:O', title='Year'),
alt.Color('Medal:N', sort=['Gold','Silver', 'Bronze'], scale=alt.Scale(
domain=['Gold', 'Silver', 'Bronze'],
range=['#FFD700 ', '#C0C0C0', '#cd7f32']), legend=alt.Legend(
title="Medal"))
)
chart1 = chart1.properties(title="Men's Basketball Medal Winners in the Summer Olympics")
chart1 = chart1.configure(title = alt.VgTitleConfig(fontSize=12, offset=10))
chart1 = chart1.configure_legend(titleFontSize=8)
chart1 = chart1.configure_axis(titleFontSize=10)
chart1
chart1.save('plots/medals/men_basketball.png', scale_factor=7.0)
chart2 = alt.Chart(women_basketball_medal).mark_rect().encode(
alt.Y('region:N', title='Country'),
alt.X('Year:O', title='Year'),
alt.Color('Medal:N', sort=['Gold','Silver', 'Bronze'], scale=alt.Scale(
domain=['Gold', 'Silver', 'Bronze'],
range=['#FFD700 ', '#C0C0C0', '#cd7f32']), legend=alt.Legend(
title="Medal"))
)
chart2 = chart2.properties(title="Women's Basketball Medal Winners in the Summer Olympics")
chart2 = chart2.configure(title = alt.VgTitleConfig(fontSize=12, offset=10))
chart2 = chart2.configure_legend(titleFontSize=8)
chart2 = chart2.configure_axis(titleFontSize=10)
chart2
chart2.save('plots/medals/women_basketball.png', scale_factor=7.0)
test = summer_medal[(summer_medal['region'] == 'USA') |
(summer_medal['region'] == 'Russia') |
(summer_medal['region'] == 'China') |
(summer_medal['region'] == 'France') |
(summer_medal['region'] == 'UK') |
(summer_medal['region'] == 'Germany') |
(summer_medal['region'] == 'Italy')]
chart = alt.Chart(test).mark_area().encode(
alt.X('Year:N', axis = alt.Axis(labelAngle=-45)),
alt.Y('count(Medal):Q', stack='center', axis=None),
alt.Color('region:N', title='Region')
)
chart = chart.properties(title='Streamgraph for Summer Olympic Medals - Top 7 Countries')
chart = chart.configure(title = alt.VgTitleConfig(fontSize=12, offset=10, anchor='middle'))
chart
chart.save('plots/medals/streamgraph_top.png', scale_factor=7.0)
chart = alt.Chart(test).mark_area().encode(
alt.X('Year:N', axis = alt.Axis(labelAngle=-45)),
alt.Y('count(Medal):Q', title=''),
alt.Color('region:N', title='Region'),
row=alt.Row('region:N', title='')
).properties(height=50, width=600)
chart = chart.properties(title='Area chart showcasing top 7 medal winners in the Summer Olympics')
chart = chart.configure(title = alt.VgTitleConfig(fontSize=12, offset=10, anchor='middle'))
chart
chart.save('plots/medals/area_top.png', scale_factor=7.0)
| 6. Medals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import scanpy as sc
sc.set_figure_params(dpi=100, frameon=False)
sc.logging.print_header()
import os
os.chdir('./../')
from compert.helper import rank_genes_groups_by_cov
import warnings
warnings.filterwarnings('ignore')
adata = sc.read('datasets/lincs.h5ad')
adata.obs['condition'] = adata.obs['pert_iname']
adata.obs['cell_type'] = adata.obs['cell_id']
adata.obs['dose_val'] = adata.obs['pert_dose']
adata.obs['cov_drug_dose_name'] = adata.obs.cell_type.astype(str) + '_' + adata.obs.condition.astype(str) + '_' + adata.obs.dose_val.astype(str)
adata.obs['control'] = (adata.obs['condition'] == 'DMSO').astype(int)
pd.crosstab(adata.obs.condition, adata.obs.cell_type)
# Calculate differential genes manually, such that the genes are the same per condition.
# +
# %%time
sc.tl.rank_genes_groups(
adata,
groupby='condition',
reference='DMSO',
rankby_abs=True,
n_genes=50
)
de_genes = {}
for cond in adata.obs['condition']:
if cond != 'DMSO':
df = sc.get.rank_genes_groups_df(adata, group=cond) # this takes a while
de_genes[cond] = df['names'][:50].values
# -
adata.uns['rank_genes_groups_cov'] = {cond: de_genes[cond.split('_')[1]] for cond in adata.obs['cov_drug_dose_name'].unique() if cond.split('_')[1] != 'DMSO'}
# +
adata.obs['split'] = 'train'
# take ood from top occurring perturbations to avoid losing data on low occ ones
ood_idx = sc.pp.subsample(
adata[adata.obs.condition.isin(list(adata.obs.condition.value_counts().index[1:50]))],
.1,
copy=True
).obs.index
adata.obs['split'].loc[ood_idx] = 'ood'
# take test from a random subsampling of the rest
test_idx = sc.pp.subsample(
adata[adata.obs.split != 'ood'],
.16,
copy=True
).obs.index
adata.obs['split'].loc[test_idx] = 'test'
# -
pd.crosstab(adata.obs['split'], adata.obs['condition'])
del(adata.uns['rank_genes_groups']) # too large
# code compatibility
from scipy import sparse
adata.X = sparse.csr_matrix(adata.X)
sc.write('datasets/lincs.h5ad', adata)
| preprocessing/lincs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PROBLEMAS DIVERSOS
# <h3>1.</h3>
# Realizar una función que permita la carga de n alumnos. Por cada alumno se deberá preguntar el nombre completo y permitir el ingreso de 3 notas. Las notas deben estar comprendidas entre 0 y 10. Devolver el listado de alumnos.
N=input("Ingresar su nombre compelto:")
NOTA1=float(input("Ingresar primera nota:"))
NOTA2=float(input("Ingresar segunda nota:"))
NOTA3=float(input("Ingresar tercera nota:"))
cantidad = int(input('Cuantos alumnos desea ingresas? '))
cantidad
lista_alumnos = []
for i in range(3):
alumno = {}
# ingreso nombre
nombre = input(f'Ingrese el nombre del alumno {i+1}: ')
alumno['nombre']= nombre
#ingreso de notas
alumno['notas'] = []
for n in range(3):
nota = float(input(f'Ingrese la nota {n+1} del alumno: '))
alumno['notas'].append(nota)
#agrupando datos en lista
lista_alumnos.append(alumno)
lista_alumnos
alumno
# +
for persona in lista_alumnos:
print(persona['nombre'], sum(persona['notas'])/3)
# -
# ### 2.
# Definir una función que dado un listado de alumnos evalúe cuántos aprobaron y cuántos desaprobaron, teniendo en cuenta que se aprueba con 4. La nota será el promedio de las 3 notas para cada alumno.
# +
numeroCalificaciones=0
while True:
float
numeroCalificaciones=int(raw_input("Dame el numero de calificaciones: "))
break
except ValueError:
print "Error"
suma=0
Calificaciones=[]
for i in range(0,numeroCalificaciones):
while True:
try:
Calificacion= int(raw_input("dame la calificacion"+str(i)+":"))
break
except ValueError:
print "Error:"
Calificaciones.append(Calificacion)
suma=suma + calificacion
promedio= suma/numeroCalificaciones
for i in range(0,numeroCalificaciones):
if Calificaciones[i]>=15:
print(srt(Calificaciones[i]) +" Calificacion Aprobatoria")
else:
print(srt(Calificaciones[i]) +" Calificacion NO Aprobatoria")
print promedio
# +
# Al escanear se devuelve como cadena
promedio_como_cadena = input("Dime tu promedio: ")
# Convertir a float
promedio = float(promedio_como_cadena)
# Hacer la comparación
if promedio >= 11:
print("Aprobado")
else:
print("No aprobado")
# -
# ### 3.
# Informar el promedio de nota del curso total.
# ### 4.
# Realizar una función que indique quién tuvo el promedio más alto y quién tuvo la nota promedio más baja.
def fun(nota):
if nota > 7:
return "Promociona"
else:
if nota < 4:
return "Aplazado"
else:
if 4 <= nota <= 7:
return "Aprobado"
# +
aplazados = aprobados = notables = 0
while True:
nota = float(input('Ingrese nota (0 para terminar):'))
if nota == 0:
break
if nota > 10:
continue
else:
if nota < 4:
aplazados += 1
elif nota >= 4 and nota <=7:
aprobados += 1
elif nota > 7 and nota <= 10:
notables += 1
print ('\nNumero de aprobados %d' %aprobados)
print('Numero de aplazados %d' %aplazados)
print('Numero de notables %d' %notables)
# -
# ### 5.
# Realizar una función que permita buscar un alumno por nombre, siendo el nombre completo o parcial, y devuelva una lista con los n alumnos que concuerden con ese nombre junto con todos sus datos, incluido el promedio de sus notas.
def alumno(n):
notas=[]
nombre=[]
for i in range(n):
name= input(f'Ingrese el nombre del alumno {i+1}: ')
nombre.append(name)
nota_1 = float(input('Ingrese Nota 1: '))
nota_2 = float(input('Ingrese Nota 2: '))
nota_3 = float(input('Ingrese Nota 3: '))
notas.append([nota_1,nota_2,nota_3])
print("Alumnos \t Notas")
for i in range(n):
print(nombre[i],"\t \t",notas[i][0],notas[i][1],notas[i][2])
n=int(input("Ingrese cantidad de alumnos: "))
alumno(n)
| Modulo2/Ejercicios/Problemas Diversos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/diegoalmanzaxd/daa_2021_1/blob/master/2_de_diciembre.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="4dZV73uRHyGK"
def fnrecinfinita():
print("hola")
fnrecinfinita()
# + id="2bpdNzkkLgnH"
fnrecinfinita()
# + colab={"base_uri": "https://localhost:8080/"} id="Vy3xZlivLjrt" outputId="4c045853-f17d-42e6-f821-f8bcb9d15503"
def fnrec(x):
if x==0:
print("stop")
else:
print(x)
fnrec(x-1)
def main():
print("inicio del programa")
fnrec(5)
print("fin del programa")
main()
# + colab={"base_uri": "https://localhost:8080/"} id="C2gHRfiwLoXT" outputId="800938ca-dda3-4c25-a20e-5976218419e5"
def printrev(x):
if x>0:
printrev(x-1)
print(x)
printrev(3)
# + colab={"base_uri": "https://localhost:8080/"} id="euMp7E1TLrlO" outputId="0f0732fa-8fe1-41ed-a164-cf9f1bbaf494"
def fibonacci(n):
if n==1 or n==0:
return n
else:
return (fibonacci(n-1)+ fibonacci(n-2))
print(fibonacci(8))
| 2_de_diciembre.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Top level imports and logging config
import logging.config
import os
from collections import deque
import gym
import numpy as np
import torch
from trojai_rl.datagen.environment_factory import EnvironmentFactory
from trojai_rl.datagen.envs.wrapped_boxing_public import WrappedBoxingConfig, WrappedBoxing
from trojai_rl.modelgen.architectures.atari_architectures_public import FC512Model
from trojai_rl.modelgen.config import RunnerConfig, TestConfig
from trojai_rl.modelgen.runner import Runner
from trojai_rl.modelgen.torch_ac_optimizer import TorchACOptimizer, TorchACOptConfig
logger = logging.getLogger(__name__)
logging.config.dictConfig({
'version': 1,
'formatters': {
'basic': {
'format': '%(message)s',
},
'detailed': {
'format': '[%(asctime)s] %(levelname)s in %(module)s: %(message)s',
},
},
'handlers': {
'console': {
'class': 'logging.StreamHandler',
'formatter': 'basic',
'level': 'WARNING',
}
},
'loggers': {
'trojai_rl': {
'handlers': ['console'],
},
},
'root': {
'level': 'INFO',
},
})
# -
os.environ["KMP_DUPLICATE_LIB_OK"] = "TRUE" # sometimes required for some reason
# +
# Define Boxing Wrapper and Environment Factory
class BoxingRAMObsWrapper(gym.Wrapper):
"""Observation wrapper for Boxing with RAM observation space. Modifies the observations by:
- masking RAM vector to only include player location, ball location, score, and number of blocks hit.
- stacking 'steps' number of steps into one observation.
- modifying reward signal to be -1, 0, or 1.
- normalize observation vector to float values between 0 and 1.
"""
def __init__(self, boxing_env, steps=4):
super().__init__(boxing_env)
self.steps = steps
self._frames = deque(maxlen=self.steps)
self.boxing_mapping = [17, 18, 19, 32, 33, 34, 35]
# clock, player_score, enemy_score, player_x, enemy_x, player_y, enemy_y
# https://github.com/mila-iqia/atari-representation-learning/blob/master/atariari/benchmark/ram_annotations.py
self.observation_space = gym.spaces.Box(0.0, 1.0, shape=(7 * self.steps,))
def reset(self, **kwargs):
obs = self.env.reset()
obs = self._process_state(obs)
for _ in range(self.steps):
self._frames.append(obs)
return np.concatenate(self._frames)
def step(self, action):
obs, reward, done, info = self.env.step(action)
self._frames.append(self._process_state(obs))
reward = np.sign(reward)
return np.concatenate(self._frames), reward, done, info
def _process_state(self, obs):
return obs[self.boxing_mapping].astype(np.float32) / 255.0
class RAMEnvFactory(EnvironmentFactory):
def new_environment(self, *args, **kwargs):
return BoxingRAMObsWrapper(WrappedBoxing(*args, **kwargs))
# +
# Create TorchACOptConfig custom measurement and early stopping handles
# TorchACOptConfig functions; see modelgen/torch_ac_optimizer.py
def eval_stats(**kwargs):
rewards = kwargs['rewards']
steps = kwargs['steps']
test_cfg = kwargs['test_cfg']
env = kwargs['env']
# note that numpy types are not json serializable
eval_results = {}
reward_sums = [float(np.sum(run)) for run in rewards]
eval_results['reward_sums'] = reward_sums
eval_results['reward_avg'] = float(np.mean(reward_sums))
eval_results['steps'] = steps
eval_results['steps_avg'] = float(np.mean(steps))
eval_results['poison'] = env.poison
eval_results['poison_behavior'] = env.poison_behavior
eval_results['argmax_action'] = test_cfg.get_argmax_action()
return eval_results
def aggregate_results(results_list):
results = {'clean_reward_avgs': [], 'poison_reward_avgs': [], 'clean_step_avgs': [], 'poison_step_avgs': []}
for res in results_list:
if res['poison']:
results['poison_reward_avgs'].append(res['reward_avg'])
results['poison_step_avgs'].append(res['steps_avg'])
else:
results['clean_reward_avgs'].append(res['reward_avg'])
results['clean_step_avgs'].append(res['steps_avg'])
agg_results = {
"clean_rewards_avg": float(np.mean(results['clean_reward_avgs'])),
"clean_step_avg": float(np.mean(results['clean_step_avgs'])),
"poison_rewards_avg": float(np.mean(results['poison_reward_avgs'])),
"poison_step_avg": float(np.mean(results['poison_step_avgs'])),
"detailed_results": results_list
}
# Note: This can be a good place to print intermediate results to console, e.g.:
# logger.debug("")
# logger.debug("clean rewards avg:", agg_results['clean_rewards_avg'])
# logger.debug("poison rewards avg:", agg_results['poison_rewards_avg'])
return agg_results
def early_stop(**kwargs):
agg_results = kwargs['aggregated_test_results']
logs1 = kwargs['logs1']
logs2 = kwargs['logs2']
opt_cfg = kwargs['optimizer_cfg']
# stop at a level greater than human performance, which is 4.3:
# From Nature paper (https://www.nature.com/articles/nature14236):
# random: 0.1, LL: 44, DQN: 71.8 (+-8.4), human: 4.3
# From PPO paper (https://arxiv.org/pdf/1707.06347.pdf)
# A2C: 17.7, ACER: 98.6, PPO: 94.6
# Note that human-level performance is 12.1 in (https://arxiv.org/pdf/1511.06581.pdf)
# Max score is 100
# Note: We normalize rewards to be -1, 0, and 1, even though it is possible to get values like -2 and 2
# occasionally due to rewards being aggregated over k frames, k uniformly sampled from {2, 3, 4};
# https://gym.openai.com/envs/Boxing-v0/. Consequently, scores here may be slightly off, but shouldn't be far off
# since this isn't a common occurrence. Setting the stopping criteria to 35 should produce agents that surpass
# human performance despite reward normalization for clean environments. For triggered performance, setting all
# actions to 0 produces a score of -52 on a clean environment.
if any([env_cfg.poison for env_cfg in opt_cfg.train_cfgs]):
if agg_results['clean_rewards_avg'] > 50 and agg_results['poison_rewards_avg'] < -25:
return True
else:
if agg_results['clean_rewards_avg'] > 50:
return True
return False
# -
def train_boxing_model():
# training parameters
poison = 'add_100'
poison_behavior = 'negate_reward'
test_poison_behavior = 'no_change' # turns off reward modifications for more intuitive performance measurments
num_clean_train_envs = 8
num_triggered_train_envs = 2
max_frames = int(50e6) # early stopping should stop in less than 25 million frames for this example
num_frames_per_proc = 128
num_epochs = 3
test_freq_frames = 100000 # do intermediate testing every this many frames trained on
int_num_clean_test = 30 # number of clean environment tests to run during intermittent testing
int_num_triggered_test = 30 # number of triggered environment tests to run during intermittent testing
num_clean_test = 30 # number of clean environment tests to run at end of training
num_triggered_test = 30 # number of triggered environment tests to run at end of training
test_max_steps = 2450 # just more than enough to run the full game if neither agent gets a KO
# Note: Boxing runs can take a long time, and can significantly slow training with intermittent testing.
# Altering test_freq_frames, int_num_clean_test, and int_num_triggered_test will affect this most:
# Set test_freq_frames to np.inf to not do intermittent testing
# Increase test_freq_frames to test less frequently, but will examine stopping criteria less often also
# Decrease int_num_clean_test and/or int_num_triggered_test to run fewer tests, but may make
# averages less reliable; or set them to 0 to run no tests, which also affects early stopping
save_loc = os.path.abspath('./')
learning_rate = 0.0001
value_loss_coef=1.0
clip_eps=0.1
model_save_name = "BoxingFC512Model.pt"
device = 'cuda' if torch.cuda.is_available() else 'cpu'
# set up training configs
train_env_factory = RAMEnvFactory()
test_env_factory = RAMEnvFactory()
clean_train_args = dict()
triggered_train_args = dict(poison='add_100', poison_behavior=poison_behavior)
poison_test_args = dict(poison='add_100', poison_behavior=test_poison_behavior)
train_env_cfgs = [WrappedBoxingConfig(**clean_train_args) for _ in range(num_clean_train_envs)] + \
[WrappedBoxingConfig(**triggered_train_args) for _ in range(num_triggered_train_envs)]
intermediate_test_cfgs = [TestConfig(WrappedBoxingConfig(**clean_train_args), count=int_num_clean_test),
TestConfig(WrappedBoxingConfig(**poison_test_args), count=int_num_triggered_test)]
test_cfgs = [TestConfig(WrappedBoxingConfig(**clean_train_args), count=num_clean_test),
TestConfig(WrappedBoxingConfig(**poison_test_args), count=num_triggered_test)]
env = BoxingRAMObsWrapper(WrappedBoxing(WrappedBoxingConfig(**clean_train_args)))
model = FC512Model(env.observation_space, env.action_space)
model.to(device)
# set up optimizer
optimizer_cfg = TorchACOptConfig(train_env_cfgs=train_env_cfgs,
test_cfgs=test_cfgs,
algorithm='ppo',
num_frames=max_frames,
num_frames_per_proc=num_frames_per_proc,
epochs=num_epochs,
test_freq_frames=test_freq_frames,
test_max_steps=test_max_steps,
learning_rate=learning_rate,
value_loss_coef=value_loss_coef,
clip_eps=clip_eps,
device=device,
intermediate_test_cfgs=intermediate_test_cfgs,
eval_stats=eval_stats,
aggregate_test_results=aggregate_results,
early_stop=early_stop,
preprocess_obss=model.preprocess_obss)
optimizer = TorchACOptimizer(optimizer_cfg)
# turn arguments into a dictionary that we can save as run information
save_info = dict(poison=poison,
poison_behavior=poison_behavior,
test_poison_behavior=test_poison_behavior,
num_clean_train_envs=num_clean_train_envs,
num_triggered_train_envs=num_triggered_train_envs,
max_frames=max_frames,
num_frames_per_proc=num_frames_per_proc,
num_epochs=num_epochs,
test_freq_frames=test_freq_frames,
int_num_clean_test=int_num_clean_test,
int_num_triggered_test=int_num_triggered_test,
num_clean_test=num_clean_test,
num_triggered_test=num_triggered_test,
test_max_steps=test_max_steps,
save_loc=save_loc
)
# set up runner and create model
runner_cfg = RunnerConfig(train_env_factory, test_env_factory, model, optimizer,
model_save_dir=os.path.join(save_loc, 'models/'),
stats_save_dir=os.path.join(save_loc, 'stats/'),
filename=model_save_name,
save_info=save_info)
runner = Runner(runner_cfg)
runner.run()
def plot_intermediate_testing_data(pretrained=True):
"""
Plot intermittent testing information using saved JSON file created after training
:param pretrained: (bool) Use data from the pretrained model included in the repository; assumes the data has
not been moved
"""
from matplotlib import pyplot as plt
# %matplotlib inline
import json
if pretrained:
with open('pretrained_boxing/FC512Model.pt.train.stats.json') as f:
data = json.load(f)
else:
with open('stats/BoxingFC512Model.pt.train.stats.json') as f:
data = json.load(f)
clean_avgs = []
poison_avgs = []
for v in data['intermediate_test_results']:
clean_avgs.append(v['clean_rewards_avg'])
poison_avgs.append(v['poison_rewards_avg'])
plt.plot(range(len(clean_avgs)), clean_avgs, label='clean')
plt.plot(range(len(poison_avgs)), poison_avgs, label='triggered')
plt.title("Boxing-ram-v0 Intermediate Test Performance")
plt.xlabel("Test number (~100,000 frames or ~78 optim steps)")
plt.ylabel("Avg score over 20 games")
plt.show()
# train an agent
train_boxing_model()
plot_intermediate_testing_data(pretrained=True)
| notebooks/boxing_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:hcds-final]
# language: python
# name: conda-env-hcds-final-py
# ---
# # Predicting the Source of Wildfires
# <NAME>
# DATA 512 Final Project
# University of Washington, Fall 2018
# ## Introduction
# Wildfires have been a big topic in the recent news with devasting effects across the western coast of the United States. So far this year, we have had less burn than 2017, but the current fire in California is the largest in state history and still burns rapidly. Last year, we had almost 2 billion dollars of losses across the United States as a result of wildfire damage which has been the highest in history [[6](https://www.iii.org/fact-statistic/facts-statistics-wildfires)]. Risks of wildfires continue to climb as scientists discover alarming links between rising greenhouse gasses, temperature, and wildfire severity. <NAME> et al. performed a comprehensive study on the relationship between the two and concluded with overwhelming confidence that a positive trend exists between them [[2](https://agupubs.onlinelibrary.wiley.com/doi/full/10.1029/2004GL020876)]. Rising greenhouse gasses could be playing a significant role in the prevalence and severity of forest fires. The visualization below shows a scatter plot of wildfires recorded in the continental US from 1991-2015. Each point is a fire (I'll go into the colors later in the notebook). This picture shows the the magnitude and prevalance of the problem here at home and gives me further creedance to study the problem.
# +
import os
from IPython.display import Image
Image(os.path.join('images', 'all_fires_map.JPG'))
# -
# Key to understanding the overall problem is the double-edged sword forests play in climate change; they are both a cause and effect. The wildfires both increase atmospheric greenhouse gasses and destroy the integral vegetation to the planet's carbon cycle [[3](https://www.iucn.org/resources/issues-briefs/forests-and-climate-change), [7](https://daac.ornl.gov/NPP/guides/NPP_EMDI.html), [8](http://daac.ornl.gov/)]. The Paris Agreement has specifically mentioned the importance of this and insists that countries protect against deforestation [[4](https://unfccc.int/process-and-meetings/the-paris-agreement/the-paris-agreement)]. Not only is the world pushing to keep the forests we have but here at home, we have begun to employ them as significant combatants in the fight against climate change. California has led the way with their proposed carbon plan. It proposes methods to reshape parts of their existing ecosystem to make their forests even more efficient at removing carbon [[5](http://www.unenvironment.org/news-and-stories/story/forests-provide-critical-short-term-solution-climate-change)]. Stopping deforestation would significantly promote the UNs progress towards reaching goals outlined in the Paris Agreement.
#
# However, this will not work if the forests continue in the same destructive cycle with our ecosystem. The goal of this project is two-fold. One, to understand the independent variables and correlation effects in a combined dataset of the Fire Program Analysis (FPA) reporting system, NOAA's Global Surface Summary of Day Data (GSOD) 7, and NASA's biomass indicators. Two, to train and assess a model for predicting the reason a wildfire started. (and possibly estimate the impact? location?) Identifying the source is a difficult task for investigators in the wild. The vastness of land covered is much larger than the matchstick or location of a lightning strike. Developing an understanding of the independent variables and a reliable prediction model could give authorities valuable direction as to where to begin their search.
#
# #### Research Questions
# * What are the most important indicators to consider when determining the cause of a wildfire?
# * Can a reliable model be built to assist investigators in determining the cause of a wildfire?
#
# #### Reproducibility
# This notebook is intended to be completely reproducible. However, the starting datasets are much too large to be hosted on GitHub. I provide a small, randomly selected sample with the repository to show the dataset cleaning and generation process. If you run this notebook on your own machine please be aware that the notebook requires quite a bit of resources. With 12 cores running at 4ghz and a consistent 95% CPU load, it took my machine nearly 27 hours to compute. The analysis portion of the notebook is also computationally expensive. The cross-validation approach implemented will consume all available resources and severely limit any other concurrent processes for several hours. The final tuned models can be computed directly via the parameters found during my tuning process.
#
# The original data format of the GSOD data makes creating samples a bit challenging. To do this, I ran an additional notebook with the following code. It opens each subdir of the extracted GSOD file and randomly selects and removes half the files. I ran this iteratively until the resulting file size was within the Github file size limit of 100mb.
#
# ```Python
# import os
#
# # walk the extracted directory
# for dirpath, dirnames, filenames in os.walk('gsod_all_years'):
#
# # process each year
# for sdir in dirnames:
# # randomly select some station files
# sfiles = os.listdir(os.path.join(dirpath, sdir))
# to_remove = np.random.choice(sfiles, int(len(sfiles)/2))
#
# # remove them
# for f in to_remove:
# try:
# tr = os.path.join('.', dirpath, sdir, f)
# os.remove(tr)
# except FileNotFoundError:
# pass
# ```
#
# I repacked the sample to be in the same format as the original dataset using WinZip. To sample from the completed fires dataset I used the following code snippet.
#
# ```Python
# import pandas as pd
#
# # read
# df = pd.read_csv('fires_complete.csv')
#
# # sample
# df = df.sample(frac=.85)
#
# # write back
# df.to_csv('fires_complete.csv', index=None)
# ```
#
# And finally, to sample the fires data I first dropped all other tables besides Fires. Next, I ran the following snippet iteratively until the sqlite file was under 100mb.
#
# ```Python
# import sqlite3
#
# # connect
# path = os.path.join('FPA_FOD_20170508.sqlite')
# conn = sqlite3.connect(path)
#
# # randomly delete some fires
# conn.execute("""
# DELETE FROM Fires
# WHERE fod_id IN (
# SELECT fod_id
# FROM Fires
# ORDER BY RANDOM()
# LIMIT 100000
# );
# """)
#
# # compress the file
# conn.execute('VACUUM;')
# ```
#
# #### A Note on Visualizations
# I use [Plotly](https://plot.ly/python/) extensively throughout this notebook. They are interactive and require Javascript to be running in the background. The Github previewer does not run the necessary Javascript for rendering making them just empty grey squares.
# #### The Full Datasets
# The full datasets can be downloaded by changing the bool `fulldata` to True and running the following cell.
# +
fulldata = False
if fulldata:
import requests
bucket = 'https://s3.amazonaws.com/waninger-hcds/'
fires_original = 'FPA_FOD_20170508.sqlite'
gsod_original = 'gsod_all_years.zip'
# download the complete fires dataset
print('fetching fires data')
r = requests.get(bucket+fires_original, allow_redirects=True)
if r.ok:
with open(os.path.join('.', 'data', fires_complete), 'wb') as f:
f.write(r.content)
# download the original GSOD
print('fetching GSOD data')
r = requests.get(bucket+gsod_original, allow_redirects=True)
if r.ok:
with open(os.path.join('.', 'data', gsod_original), 'wb') as f:
f.write(r.content)
# -
# ## Part 1. Data Preparation
# ### Setup
# This notebook is coded to run with Python 3.6. Several libraries from the Python standard library will be used along with several third-party modules. These can be installed with the provided requirements file using the command
#
# `pip install --user -r requirements.txt`
#
# More information regarding the standard libarary can be found at [python.org](https://docs.python.org/3.6/library/index.html).
#
# For third party libraries please see:
# * [numpy == 1.13.0](https://docs.scipy.org/doc/numpy-1.13.0/reference/)
# * [pandas == 0.23.4](https://pandas.pydata.org/pandas-docs/stable/)
# * [plotly == 3.4.2](https://plot.ly/python/)
# * [scikit-learn == 0.20.1](https://scikit-learn.org/stable/documentation.html)
# * [statsmodels == 0.9.0](https://www.statsmodels.org/stable/index.html)
# * [tqdm == 4.28.1](https://github.com/tqdm/tqdm)
# + code_folding=[]
# Python standard library
import calendar
import datetime as dt
from IPython.core.interactiveshell import InteractiveShell
import itertools as it
import multiprocessing as mul
from multiprocessing.dummy import Pool as TPool
import gzip
import os
import shutil
import sqlite3
import sys
import tarfile
import time
import zipfile
# third party modules
import numpy as np
import pandas as pd
import plotly.graph_objs as go
from plotly import tools
import plotly.figure_factory as ff
from plotly.offline import init_notebook_mode, iplot
from scipy import interp
from sklearn.metrics import roc_curve, auc, accuracy_score
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.model_selection import GridSearchCV, StratifiedKFold, train_test_split
from sklearn.preprocessing import label_binarize, LabelEncoder, StandardScaler
import statsmodels.api as sm
import statsmodels.formula.api as smf
from tqdm import tqdm, tqdm_pandas
# initialize plotly
init_notebook_mode(connected=True)
# set notebook options
InteractiveShell.ast_node_interactivity = 'all'
# set random seed
np.random.seed(42)
# initalize tqdm
tqdm.pandas(leave=True)
# -
# ### Data Sources
# Four data sources are to be used for this project. The primary data source was found through Kaggle and contains 1.88 million wildfires that occurred in the United States from 1992 to 2015. This data contains the primary labels to be used as target variables. The United States Department of Agriculture curated the original data ([Forest Service](https://www.fs.fed.us/)) and can be found at [link](https://www.fs.usda.gov/rds/archive/Product/RDS-2013-0009.4/). The second is the GSOD data curated by [NOAA](https://www.noaa.gov/). Finally, the National Air and Space Association (NASA) hosts a valuable biome dataset at the ORNL Distributed Active Archive Center for Biogeochemical Dynamics ([DAAC](https://daac.ornl.gov/NPP/guides/NPP_EMDI.html). Later in the notebook, I will show how neither the NASA or DAAC data is useful and propose an alternate data source for future work.
#
# ### Get some metrics from the fires dataset
# The target variable for this analysis exists inside the wildfire dataset. I start by generating a bounding box of latitude and longitude values to filter the other three sources.
# + code_folding=[]
# generate the file path and connect using the sqlite3 driver
path = os.path.join('.', 'data', 'FPA_FOD_20170508.sqlite')
conn = sqlite3.connect(path)
# retrieving the minimum and maximum latitude and longitude pairs.
fires = pd.read_sql_query('''
SELECT
min(LATITUDE) AS min_lat,
max(LATITUDE) AS max_lat,
min(LONGITUDE) AS min_lon,
max(LONGITUDE) AS max_lon
FROM
Fires
''', conn)
# increase by one degree-decimal point so that we don't exclude
# nearby weather stations
min_lat = np.round(fires.min_lat.values[0], 0)-1
min_lon = np.round(fires.min_lon.values[0], 0)-1
max_lat = np.round(fires.max_lat.values[0], 0)+1
max_lon = np.round(fires.max_lon.values[0], 0)+1
# print them to the console
min_lat, max_lat, min_lon, max_lon
# -
# ### Load and process GSOD files
# The data from NOAA comes in a full-nonsense version. It's a collection zipped zip files, one compressed tar file for each year. Then, each day of the year and station is yet another compressed gzip file. I extract the main file and remove any years not from 1991-2015. In the next cell I unzip the years we need, then each year into the directory 'gsod_extracted'. I apologize for this next few cells. This really is nonsense to make it reproducible all the way from the source file from NOAA.
# + code_folding=[4, 11]
# create the file path
gsod_path = os.path.join('.', 'data', 'gsod')
# make sure the path exists
if not os.path.exists(gsod_path):
os.mkdir(gsod_path)
# get the main zip file
all_years = zipfile.ZipFile(os.path.join('data','gsod_all_years.zip'))
# look for contents only in the designated year range
members = [
n for n in all_years.namelist()
if any([n.find(str(yi)) > -1 for yi in list(range(1991, 2016))])
]
# extract
for m in tqdm(members):
t = all_years.extract(m, gsod_path)
# -
# Lets first load the listing of weather stations. I do this first because it reduces the number of operations of depending cells by quite a bit and in turn, drastically speeds up the notebook. Furthermore, I need the latitude and longitude values for each weather summary in order to join with the fires dataset. I do this by creating a composite key out of USAF and WBAN in both the stations and weather dataframes, then performing an inner join on it. For more information please see the NOAA data documentation provided. And finally, I need to create a smaller subset of the original dataset to reduce the amount of data we need to upload/download for reproducibility purposes.
#
# I also make sure to exclude weather stations that aren't going to be used in widlfire feature engineering by creating latitude and longitude masks offsetting each min/max by 111km.
# + code_folding=[] run_control={"marked": false}
# load the stations file explicitly enforcing datatypes and nan values
# also drop any station that doesn't have a latitude or longitude value
stations = pd.read_csv(
os.path.join('data', 'isd-history.csv'),
dtype={
'USAF':'str',
'WBAN':'str'
},
na_values={
'WBAN' :'99999',
'ELEV(M)':'-999'
}
).dropna(subset=['LAT', 'LON'], how='any')
# take only stations that have lat, lon values within the wildfire range
stations['lat_mask'] = [min_lat <= lat <= max_lat for lat in stations.LAT]
stations['lon_mask'] = [min_lon <= lon <= max_lon for lon in stations.LON]
stations = stations.loc[stations.lat_mask & stations.lon_mask].drop(columns=['lat_mask', 'lon_mask'])
# create a key by concatenating the USAF and WBAN cols
stations.loc[stations.USAF.isnull(), 'USAF'] = 'none'
stations.loc[stations.WBAN.isnull(), 'WBAN'] = 'none'
stations['KEY'] = stations.USAF+stations.WBAN
# verify key uniqueness
assert len(stations.KEY.unique()) == len(stations)
# we will only be using these columns
stations = stations.reindex(columns=[
'KEY', 'LAT', 'LON', 'ELEV(M)'
])
# rename the elevation column so we can call it easier
stations = stations.rename(columns={'ELEV(M)':'ELEV'})
# convert all the column names to lowercase
stations.columns = [c.lower() for c in stations.columns]
stations.head()
# -
# Now extract the contents of each year into the extracted directory.
# + code_folding=[] run_control={"marked": false}
# get the yearly list of tar files
years = [f for f in os.listdir(gsod_path) if f.find('tar') > -1]
# generate the extract path
ex_path = os.path.join('.', 'data', 'gsod_extracted')
# make sure the path exists
if not os.path.exists(ex_path):
os.mkdir(ex_path)
# extract the content from each year into the 'extracted' directory
pbar = tqdm(total=len(years))
for y in years:
pbar.set_description(y)
# load the tarfile provided by NOAA
tf = tarfile.TarFile(os.path.join(gsod_path, y))
# create a subdirectory to extract the contents into
subdir = os.path.join(ex_path, y.replace('.tar', ''))
if not os.path.exists(subdir):
os.mkdir(subdir)
# extract each year
tf.extractall(subdir)
pbar.update(1)
# otherwise this is the sampled data so just move the contents
if len(years) == 0:
years = os.listdir(gsod_path)
for y in years:
files = os.listdir(os.path.join(gsod_path, y))
for f in files:
old = os.path.join(gsod_path, y, f)
newdir = os.path.join(ex_path, y)
if not os.path.exists(newdir):
os.mkdir(newdir)
new = os.path.join(newdir, f)
os.rename(old, new)
pbar.update(1)
pbar.close()
# -
# Process each station file line-by-line into DataFrame. This cell only does the raw transformation from a gzip text file into a csv. Each line of each file is a separate row with each field separated by a certain number of character positions. These are listed in the NOAA GSOD docs and were extensively used to process the data. Note, the extractions do not line up perfectly due to the parser being used. Each column was carefully checked to ensure no missing characters. Also of note is that some of the files contain blank lines so I added a filter at the end of each parsing to only input the row if a valid station id is present. We can't perform the latitude, longitude lookup without it making the row unusable even if it did contain the remaining fields.
# + code_folding=[28]
# get the list of files for each day
ex_path = os.path.join('.', 'data', 'gsod_extracted')
years = [d for d in os.listdir(ex_path) if os.path.isdir(os.path.join(ex_path, d))]
# read and extract the contents for each day of year
i=0
for y in years:
# create the filename to save the final csv output
name = os.path.join(ex_path, y.replace('.tar', ''))
name = name + '.csv'
# get the subdirectory path
subdir = os.path.join(ex_path, y.replace('.tar', ''))
# read all files we extracted into the directory
files = os.listdir(subdir)
# store a list of dictionary objects for each row parsed
content = []
for f in tqdm(files, desc=y):
# open the file
with gzip.open(os.path.join(subdir, f), 'r') as fc:
# read the entire contents, split by newline and ignore header
t = str(fc.read()).split('\\n')[1:]
# see GSOD_DESC.txt for exact delimmiter locations
def parse(s):
d = dict(
stn = s[ 0: 6].strip(),
wban = s[ 6:13].strip(),
year = s[13:18].strip(),
moda = s[18:23].strip(),
temp = s[23:30].strip(),
temp_cnt = s[30:34].strip(),
dewp = s[34:41].strip(),
dewp_cnt = s[41:44].strip(),
slp = s[44:52].strip(),
slp_cnt = s[52:55].strip(),
stp = s[55:63].strip(),
stp_cnt = s[63:66].strip(),
visib = s[67:73].strip(),
visib_cnt= s[73:76].strip(),
wdsp = s[76:83].strip(),
wdsp_cnt = s[83:86].strip(),
mxspd = s[88:93].strip(),
gust = s[94:101].strip(),
temp_max = s[102:108].strip(),
max_temp_flag = s[108:110].strip(),
temp_min = s[111:116].strip(),
min_temp_flag = s[116:117].strip(),
prcp = s[117:123].strip(),
prcp_flag= s[123:124].strip(),
sndp = s[124:131].strip(),
frshtt = s[131:138].strip()
)
return d if len(d['stn']) > 1 else None
# convert each row into a dictionary using the function above
# and append the contents to the main collection
content += list(map(parse, t))
# convert the list of dictionaries to a Pandas dataframe
content = pd.DataFrame([c for c in content if c is not None])
# write this years worth of weather recordings to csv
content.to_csv(name, index=None)
# -
# In the following cell, I go through the csv contents we generated above. Specific datatypes are enforced to prevent Pandas from dropping leading zeroes, for example, and to make additional operations more streamlined. Each will be explained line by line.
# + code_folding=[13, 58]
# get the list of yearly weather files
ex_path = os.path.join('.', 'data', 'gsod_extracted')
names = [f for f in os.listdir(ex_path) if 'csv' in f]
# process each year at a time
pbar = tqdm(total=len(names))
for name in names:
pbar.set_description(name)
# load the data, setting data types explicitly or pandas will drop
# the leading zeroes needed for station names. Also, include the
# explicit na values designated in the data documentation
# drop columns we aren't going to use
f1 = pd.read_csv(
os.path.join(ex_path, name),
dtype={
'stn' :'str',
'wban':'str',
'moda':'str',
'frshtt':'str',
'year':'str'},
na_values={
'stn' :'999999',
'wban' :'99999',
'temp' :'9999.9',
'dewp' :'9999.9',
'slp' :'9999.9',
'stp' :'9999.9',
'visib':'999.9',
'wdsp' :'999.9',
'mxspd':'999.9',
'gust' :'999.9',
'max_temp':'9999.9',
'min_temp':'9999.9',
'prcp':'99.9',
'sndp':'999.9'},
) \
.drop(columns=[
'max_temp_flag', 'min_temp_flag',
'temp_cnt', 'dewp_cnt', 'slp_cnt',
'stp_cnt', 'visib_cnt', 'wdsp_cnt'])
# convert the two date columns 'year' and 'moda' to a single pydate
f1['date'] = [
dt.datetime(year=int(r.year), month=int(r.moda[:2]), day=int(r.moda[2:]))
for r in f1.itertuples()
]
# extract month number and julian date
f1['month'] = f1.date.apply(lambda x: x.month)
f1['doy'] = f1.date.apply(lambda x: x.timetuple().tm_yday)
# convert prcp values to na where prcp flag is in {'H', 'I'}. see the data docs
f1.loc[(f1.prcp_flag == 'H') | (f1.prcp_flag == 'I'), 'prcp'] = np.nan
# convert 'frshtt' to an ordinal value based on severity where the
# returned value is the number of leading most 1. ie. 010000 -> 2
# 1:fog, 2:rain, 3:snow, 4:hail, 5:thunderstorm, 6:tornado
def fx(x):
x = x[::-1].find('1')
return x if x != -1 else 0
f1['atmos_sev'] = f1.frshtt.apply(fx)
# create the join key in the same way as we did for weather stations
f1.loc[f1.stn.isnull(), 'stn'] = 'none'
f1.loc[f1.wban.isnull(), 'wban'] = 'none'
f1['key'] = f1.stn + f1.wban
# perform an inner join with stations
f1 = f1.merge(stations, on='key', how='inner')
# reorder the columns, dropping the ones that won't be used
prefix = ['lat', 'lon', 'year', 'month', 'doy']
f1 = f1.reindex(columns=prefix + sorted(list(
set(f1.columns) - set(prefix) - {
'moda', 'prcp_flag', 'frshtt', 'stn', 'wban', 'key', 'date'
}
)))
# write the cleaned dataframe to disk
name = os.path.join(gsod_path, name.replace('.csv', '_cleaned') + '.csv')
f1.to_csv(name, index=None)
pbar.update(1)
pbar.close()
# -
# Create a single data frame with cleaned values for all years. This generates a dataframe approximately 1.7gb uncompressed which is a significant reduction from the 3.4gb original compressed file.
# + code_folding=[]
# get the list of cleaned files
files = [f for f in os.listdir(gsod_path) if 'cleaned.csv' in f]
assert len(files) == 25
gsod = pd.concat([pd.read_csv(os.path.join(gsod_path, f)) for f in files])
gsod.to_csv(os.path.join('.', 'data', 'gsod.csv'), index=None)
# +
# cleanup the temp directories
gsod_path = os.path.join('.', 'data', 'gsod')
shutil.rmtree(gsod_path)
ex_path = os.path.join('.', 'data', 'gsod_extracted')
shutil.rmtree(ex_path)
# -
# ### Clean the fires dataset
# This dataset comes relatively clean. The only modifications we'll be doing is removing the columns we won't be using, creating a few new, and reordering them for convenience.
# +
# generate the path and connect to the sqlite fires file
path = os.path.join('.', 'data', 'FPA_FOD_20170508.sqlite')
conn = sqlite3.connect(path)
# read all the columns we need
fires = pd.read_sql_query('''
SELECT FOD_ID,
FIRE_YEAR, DISCOVERY_DOY, DISCOVERY_TIME,
STAT_CAUSE_CODE, CONT_DOY, CONT_TIME,
FIRE_SIZE, LATITUDE, LONGITUDE, OWNER_CODE,
STATE
FROM
Fires;
''', conn)
# convert column names to lowercase
fires.columns = [c.lower() for c in fires.columns]
# based on the first 10000 rows, 0.35% have missing containment values which is a
# negligible loss at this point in the analysis
fires = fires.dropna(subset=[
'discovery_doy', 'discovery_time', 'cont_doy', 'cont_time'
], how='any')
# convert fire_year, discovery doy, and time to pydate
fires['dt_disc'] = [
dt.datetime(year=int(r.fire_year),
month=1,
day=1,
hour=int(r.discovery_time[:2]),
minute=int(r.discovery_time[2:])
) + \
dt.timedelta(days=r.discovery_doy)
for r in fires.itertuples()
]
# convert the containment dates
fires['dt_cont'] = [
dt.datetime(year=int(r.fire_year), month=1, day=1, hour=int(r.cont_time[:2]), minute=int(r.cont_time[2:])) + \
dt.timedelta(days=r.cont_doy)
for r in fires.itertuples()
]
# create some higher resolution columns
def seconds_into_year(x):
a = dt.datetime(year=x.year, month=1, day=1, hour=0, minute=0, second=0)
return int((x-a).total_seconds())
def seconds_into_day(x):
a = dt.datetime(year=x.year, month=x.month, day=x.day, hour=0, minute=0, second=0)
return (x-a).seconds
# calculate fire duration in seconds, but only if the contained date is
# later than the start date
fires['disc_soy'] = fires.dt_disc.progress_apply(seconds_into_year)
fires['cont_soy'] = fires.dt_cont.progress_apply(seconds_into_year)
fires['duration'] = [
r.cont_soy-r.disc_soy
if r.cont_soy > r.disc_soy else np.nan
for r in tqdm(fires.itertuples(), total=len(fires))
]
# extract month and hour as new columns
fires['date'] = fires.dt_disc.progress_apply(lambda x: x.date())
fires['month'] = fires.dt_disc.progress_apply(lambda x: x.month)
fires['dow'] = fires.dt_disc.progress_apply(lambda x: x.weekday())
fires['hod'] = fires.dt_disc.progress_apply(lambda x: x.hour)
fires['sod'] = fires.dt_disc.progress_apply(seconds_into_day)
# encode the state
state_le = LabelEncoder()
fires['state'] = state_le.fit_transform(fires.state)
# drop some columns we won't be using
fires = fires.drop(columns=[
'discovery_time', 'cont_doy', 'cont_time',
'disc_soy', 'cont_soy', 'dt_cont',
'dt_disc'
])
# rename some columns
fires = fires.rename(columns={
'discovery_doy':'doy',
'latitude':'lat',
'longitude':'lon',
'fire_year':'year',
'stat_cause_code':'cause_code',
})
# reorder the columns
prefix = ['fod_id', 'lat', 'lon', 'date', 'year', 'month', 'doy', 'dow', 'hod', 'sod']
fires = fires.reindex(columns=prefix + sorted(list(
set(fires.columns) - set(prefix)
)))
fires.head()
# -
# A possible feature we can engineer is the number of nearby fires. You can see the relation by looking at the first couple of rows in the fires table shown above. We see two fires occur on the same day that almost look like duplicates except they're separated by a few kilometers. This can be an especially strong signal for both lightning and arson related fires.
#
# This many lookups becomes a compute intensive operation and can take many hours to complete if run iteratively. In the following cell I create subsets of the main fires index. Each subset is sent to a different process where thread pools operate in parallel on the assigned subset. The results are output to separate csv files linked with the fire_id. This precludes transferring the data back from the assigned CPU. Instead, I'll read and join the new feature from disk in the next cell. This compute plan reduced the estimated time to completion from roughly 15 hours to 45 minutes on my local machine.
# + code_folding=[50]
# thread task
def nearby_count(f, qu):
# lookup any fires within 55km
nearby = fires.loc[
(fires.year == f.year) & (fires.doy == f.doy) &
(fires.lat <= f.lat + .25) & (fires.lat >= f.lat - .25) &
(fires.lon <= f.lon + .25) & (fires.lon >= f.lon - .25)
]
# update progress
qu.put(1)
# return the fire id and count
return dict(fod_id=f.fod_id, nearby=len(nearby)-1)
# process task
def px(batch, fires, start, step, qu):
nearby = list(fires.iloc[start:start+step, :].apply(nearby_count, axis=1, qu=qu))
path = os.path.join('.', 'data', f'nearby_{batch}.csv')
pd.DataFrame(nearby).to_csv(path, index=None)
# number of shards or 'batches'
batches = 6
# a container to hold each process
processes = []
# compute the step size
step = int(len(fires)/batches)+1
# setup a progress bar and update queue
pbar = tqdm(total=len(fires))
qu = mul.Queue()
# create the subsets and dish out the processor tasks
for batch in range(batches):
# calculate the starting point for this subset
start = step*batch
# create, append, and start the child process
p = mul.Process(target=px, args=(batch, fires, start, step, qu))
processes.append(p)
p.start()
# continue until the children finish
complete = False
while not complete:
running = batches
# round robin check of child state
for p in processes:
if not p.is_alive():
running -= 1
# set completion status if all are finished
if running == 0:
complete = True
# empty the update qu
while not qu.empty():
t = qu.get()
pbar.update(t)
# terminate and join all the children
for p in processes:
p.terminate()
p.join()
# +
# read the batches into one frame
path = os.path.join('.', 'data')
nearby = pd.concat([
pd.read_csv(os.path.join(path, f))
for f in os.listdir(path) if 'nearby_' in f
], sort=False)
# merge with the main and make sure we didn't lose any rows
a = len(fires)
fires = fires.merge(nearby, on='fod_id', how='inner')
assert a == len(fires)
# print
fires.head()
# -
# cleanup temporary files
files = [
os.remove(os.path.join('data', f))
for f in os.listdir('data') if 'nearby_' in f
]
# Lets take a quick look at the only categorical variable we have - OWNER_CODE.
# + code_folding=[]
# generate the path and connect to the sqlite fires file
path = os.path.join('.', 'data', 'FPA_FOD_20170508.sqlite')
conn = sqlite3.connect(path)
# get the mapping of cause codes to description
owners = pd.read_sql_query('''
SELECT DISTINCT(OWNER_CODE), OWNER_DESCR
FROM Fires;
''', conn)\
.sort_values('OWNER_CODE')
# rename the columns and set the index to code
owners = owners.rename(columns={
'OWNER_CODE':'code',
'OWNER_DESCR':'owner'
}).set_index('code')
# get the counts of each cause
bincounts = fires.owner_code.value_counts()
# plot
iplot(go.Figure(
[go.Bar(
x=[owners.loc[idx].owner for idx in bincounts.index],
y=bincounts,
text=bincounts.index,
textposition='outside'
)],
go.Layout(
title='Distribution of owners',
yaxis=dict(title='Count of owned fires')
)
))
# -
# This isn't our target variable but there are clear commonalities we can take advantage of to boost any signal that may come from the responsible land owner. To help understand this a bit better here is the list of federal acronyms:
#
# * USFS - United States Forest Service
# * BIA - Bureau of Indian Affairs
# * BLM - Bureau of Land Management
# * NPS - National Park Service
# * FWS - Fish and Wildlife Service
# * BOR - Bureau of Reclamation
#
# Here is a list of things I notice from the visualization.
# 1. UNDEFINED FEDERAL has very little values and can be combined with OTHER FEDERAL.
# 2. COUNTY owned land can be joined with MUNICIPAL/LOCAL.
# 3. STATE OR PRIVATE can be separted into the STATE and PRIVATE categories. To do this, I'll draw from a random binomial distribution characterized by the ratio between the two.
# 4. TRIBAL can be combined with BIA and I'll rename it to Native American.
# 5. Move the FOREIGN items into MISSING/NOT SPECIFIED.
# 6. Move the MUNICIPAL/LOCAL government owned into STATE owned.
# 7. Group the lower represented federal agencies into the FEDERAL category.
#
# This recategorization plan reduces the number of categories from 16 to six which will signficantly boost signal strength and keep the feature space more manageable. I also plan on renaming a few before continuing. Additionally, we'll need to store the new owner descriptions so we preserve the recategorization mapping.
# + code_folding=[] run_control={"marked": false}
# recategorize UNDEFINED FEDERAL
fires.loc[fires.owner_code == 15, 'owner_code'] = 6
# recategorize the under represented federal agencies - BLM, NPS, FWS, BOR
fires.loc[[True if o in [1, 3, 4, 10] else False for o in fires.owner_code], 'owner_code'] = 6
# rename the federal category
owners.loc[6, 'owner'] = 'FEDERAL'
# recategorize COUNTY
fires.loc[fires.owner_code == 11, 'owner_code'] = 12
owners.loc[12, 'owner'] = 'LOCAL'
# recategorize STATE OR PRIVATE
den = (bincounts[8]+bincounts[7])
p = bincounts[8]/den
fires.loc[fires.owner_code == 13, 'owner_code'] = np.random.binomial(1, p, len(fires.loc[fires.owner_code == 13]))+7
# recategorize TRIBAL
fires.loc[fires.owner_code == 9, 'owner_code'] = 2
owners.loc[2, 'owner'] = 'NATIVE_AMERICAN'
# recategorize FOREIGN
fires.loc[fires.owner_code == 0, 'owner_code'] = 14
owners.loc[14, 'owner'] = 'OTHER'
# recategorize MUNICIPAL/LOCAL
fires.loc[fires.owner_code == 12, 'owner_code'] = 7
# drop the integer encoding in favor of the new names
# create the new column
fires['owner'] = 'none'
# reformat the owners description to lowercase
owners.owner = [o.lower() for o in owners.owner]
# assign each code the representative name
for code in fires.owner_code.unique():
fires.loc[fires.owner_code == code, 'owner'] = owners.loc[code].owner
# drop the original encoded column
fires = fires.drop(columns=['owner_code'])
# -
# Let's replot the categorical distribution to show the differences we've made for the owner's category.
# +
# get the counts of each cause
bincounts = fires.owner.value_counts()
# plot as a bar plot
iplot(go.Figure(
[go.Bar(
x=bincounts.index,
y=bincounts,
text=bincounts,
textposition='inside'
)],
go.Layout(
title='Distribution of owners',
yaxis=dict(title='Count of owned fires')
)
))
# -
# Finally, write the completed fires dataframe to disk.
fires.to_csv(os.path.join('.', 'data', 'fires_cleaned.csv'), index=None)
fires.head()
# ### Process ORNL features
# Each station has a center point and provides the coverage data in both 1km and 50km pixel grids surrounding the station. My first approach to joining the fires and ground cover data was to include any predictions within the station's bounding box but, this led to incredibly sparse results. I leave the cell blocks here to both show my process and why I'm no longer using the data source. In the following cell I load both high and low quality datasets.
# +
# load the data we'll use, enforce datatypes, and rename columns
cover = pd.concat([
pd.read_csv(
os.path.join('.', 'data', f),
usecols=[
'LAT_DD', 'LONG_DD', 'COVR1KM', 'COVR50KM'
],
dtype={
'COVR1KM':'str',
'COVR50KM':'str'
}
).rename(columns={
'LAT_DD':'LAT',
'LONG_DD':'LON'
})
for f in [
'EMDI_ClassA_Cover_UMD_81.csv',
'EMDI_ClassB_Cover_UMD_933.csv'
]
], sort=False)
# convert columns to lowercase
cover.columns = [c.lower() for c in cover.columns]
# create cover 50k grid boundaries
cover['lower50_lat'] = cover.lat.apply(lambda x: x-.5)
cover['upper50_lat'] = cover.lat.apply(lambda x: x+.5)
cover['lower50_lon'] = cover.lon.apply(lambda x: x-.5)
cover['upper50_lon'] = cover.lon.apply(lambda x: x+.5)
# only include the values within the fire bounding box
cover = cover.loc[
(cover.lower50_lat >= min_lat) & (cover.upper50_lat <= max_lat) &
(cover.lower50_lon >= min_lon) & (cover.upper50_lon <= max_lon)
]
cover.head()
# -
# Plot a sample of fires and the bounding boxes for each station to show just how inadequate the ORNL dataset is. Each point represents a fire with the size of the fire mapped to the size of the point.
# + code_folding=[]
# extract a uniform sample of 2k fires
sample = fires.sample(1000)
# generate scatter plot points
fire_trace = go.Scatter(
x=sample.lon,
y=sample.lat,
mode='markers',
marker=dict(
color='#571C00'
)
)
# generate the bounding boxes
shapes = [
{
'type':'rect',
'x0':r.lower50_lon,
'x1':r.upper50_lon,
'y0':r.lower50_lat,
'y1':r.upper50_lat,
'fillcolor':'rgba(22, 74, 40, .4)',
'line':{
'width':.1
}
}
for r in cover.itertuples()
]
# plot
iplot(go.Figure(
[fire_trace],
layout=go.Layout(
shapes=shapes,
xaxis=dict(
title='longitude',
range=[-125, -78]
),
yaxis=dict(
title='latitude',
range=[25, 58]
),
title='Ground cover data coverage is insufficient',
width=1200,
height=800
)
))
# -
# The same goes for soil content because the same stations are used for this dataset.
# + code_folding=[]
# load the data
soil = pd.concat([
pd.read_csv(
os.path.join('.', 'data', f)
).rename(columns={
'LAT_DD':'LAT',
'LONG_DD':'LON'
}).drop(columns='SITE_ID')
for f in [
'EMDI_ClassA_Soil_IGBP_81.csv',
'EMDI_ClassB_Soil_IGBP_933.csv'
]
], sort=False)
# convert columns to lowercase
soil.columns = [c.lower() for c in soil.columns]
# create the station bounding box
soil['lower50_lat'] = soil.lat.apply(lambda x: x-.5)
soil['upper50_lat'] = soil.lat.apply(lambda x: x+.5)
soil['lower50_lon'] = soil.lon.apply(lambda x: x-.5)
soil['upper50_lon'] = soil.lon.apply(lambda x: x+.5)
# only include the values within the fire bounding box
soil = soil.loc[
(soil.lower50_lat >= min_lat) & (soil.upper50_lat <= max_lat) &
(soil.lower50_lon >= min_lon) & (soil.upper50_lon <= max_lon)
]
soil.head()
# + code_folding=[]
# extract a fire sample
sample = fires.sample(5000)
# generate the fire scatter points
fire_trace = go.Scatter(
x=sample.lon,
y=sample.lat,
mode='markers',
marker=dict(
color='#571C00'
)
)
shapes = [
{
'type':'rect',
'x0':r.lower50_lon,
'x1':r.upper50_lon,
'y0':r.lower50_lat,
'y1':r.upper50_lat,
'fillcolor':'rgba(22, 74, 40, .4)',
'line':{
'width':.1
}
}
for r in soil.itertuples()
]
# plot
iplot(go.Figure(
[fire_trace],
layout=go.Layout(
shapes=shapes,
xaxis=dict(
title='longitude',
range=[-125, -78]
),
yaxis=dict(
title='latitude',
range=[25, 58]
),
title='Soil data coverage is insufficient',
width=1200,
height=800
)
))
# -
# An alternative data source for land coverage is available for public use. See the [Earth Engine Data Catalog](https://developers.google.com/earth-engine/datasets/catalog/)
# + [markdown] heading_collapsed=true
# ### Generate aggregate weather features associated with each fire
# We'll need to lookup all reports within a given bounding box centered at the fire's originating location. I use a bounding box to preclude performing pairwise distance lookups which might be more accurate but will incur a significant expense - $O(n^2)$. The embedded hierarchical structure within a degree-decimal formatted coordinate allows us to generate contextually important containment boundaries. The boundaries will include aggregated values from all weather reports $\pm$ 55.5km of the fire.
#
# This is the long running computation may take several days to complete. I wrote it to perform aggregations in batches. Each batch will cache the resulting features to a csv file and continue with the next. Also of note here is that I use a single thread pool rather than the sharding technique to keep memory usage as low as possible.
# + hidden=true
# load cleaned GSOD file
gsod = pd.read_csv(os.path.join('.', 'data', 'gsod.csv'))
gsod.head()
# + code_folding=[7] hidden=true
# load the cleaned fires
fires = pd.read_csv(os.path.join('.', 'data', 'fires_cleaned.csv'), parse_dates=['date'])
# start a thread pool and progress bar
pool = TPool(mul.cpu_count())
pbar = tqdm(total=len(fires))
def weather_agg(args):
try:
# extract the tuple arguments
fod_id, lat, lon, year, doy = args
# make a copy of the empty record to start this record with
results = empty.copy()
results['fod_id'] = fod_id
# get all weather reports within 111km
lat_min, lat_max = lat-.5, lat+.5
lon_min, lon_max = lon-.5, lon+.5
# retrieve all weather reports within the box and 4 days leading up to and including
# the day of the fire
wthr = gsod.loc[
(gsod.lat >= lat_min) & (gsod.lat <= lat_max) &
(gsod.lon >= lon_min) & (gsod.lon <= lon_max) &
(
(gsod.year == year) & (gsod.doy >= doy-4) & (gsod.doy <= doy) |
(gsod.doy <= 4) & (gsod.year == year-1) & (gsod.doy >= 361+doy)
)
]
# get the three day prior aggregates
w_ = wthr.loc[wthr.doy != doy]
if len(w_) > 0:
results['threeDay_atmos_sev'] = np.mean(w_.atmos_sev)
results['threeDay_temp_max'] = np.max(w_.temp_max)
results['threeDay_temp_min'] = np.min(w_.temp_min)
results['threeDay_temp'] = np.median(w_.temp)
results['threeDay_sndp'] = np.median(w_.sndp)
results['threeDay_dewp'] = np.median(w_.dewp)
results['threeDay_gust'] = np.max(w_.gust)
results['threeDay_mxspd'] = np.max(w_.mxspd)
results['threeDay_stp'] = np.median(w_.stp)
results['threeDay_temp'] = np.median(w_.temp)
results['threeDay_slp'] = np.median(w_.slp)
results['threeDay_wdsp'] = np.median(w_.wdsp)
results['threeDay_prcp'] = np.sum(w_.prcp)
results['threeDay_visib'] = np.median(w_.visib)
# get the dayOf aggregates
w_ = wthr.loc[wthr.doy == doy]
if len(w_) > 0:
results['dayOf_atmos_sev'] = np.mean(w_.atmos_sev)
results['dayOf_temp_max'] = np.max(w_.temp_max)
results['dayOf_temp_min'] = np.min(w_.temp_min)
results['dayOf_temp'] = np.median(w_.temp)
results['dayOf_sndp'] = np.median(w_.sndp)
results['dayOf_dewp'] = np.median(w_.dewp)
results['dayOf_gust'] = np.max(w_.gust)
results['dayOf_mxspd'] = np.max(w_.mxspd)
results['dayOf_stp'] = np.median(w_.stp)
results['dayOf_temp'] = np.median(w_.temp)
results['dayOf_slp'] = np.median(w_.slp)
results['dayOf_wdsp'] = np.median(w_.wdsp)
results['dayOf_prcp'] = np.median(w_.prcp)
results['dayOf_visib'] = np.median(w_.visib)
# catch all exceptions and continue gracefully but make sure we
# notify in case any occur
except Exception as e:
exc_type, exc_obj, exc_tb = sys.exc_info()
print(exc_type, e, exc_tb.tb_lineno)
pbar.update(1)
return results
# create the dayOf columns
excols = {'lat', 'lon', 'elev', 'year', 'month', 'doy', 'fod_id'}
daily_cols = ['dayOf_' + c for c in list(set(gsod.columns) - excols)]
threeDay_cols = ['threeDay_' + c for c in list(set(gsod.columns) - excols)]
# create an empty dictionary to start each feature row
empty = dict()
for c in daily_cols+threeDay_cols:
empty[c] = np.nan
fires_temp = os.path.join('.', 'data', 'fires')
if not os.path.exists(fires_temp):
os.mkdir(fires_temp)
# perform this operation in batches caching the fire results each iteration
start, step = 0, 10000
for i in range(0, len(fires), step):
# get the set of indices to process
idx_set = fires.index.tolist()[i:i+step]
# process
batch = pool.map(weather_agg, [
(r.fod_id, r.lat, r.lon, r.year, r.doy)
for r in fires.loc[idx_set].itertuples()
])
# cache
pd.DataFrame(batch).to_csv(os.path.join('.', 'data', 'fires', f'fires_b{i}.csv'), index=None)
pool.close(); pool.join()
# + [markdown] hidden=true
# Finally, read all batches into a single dataframe and write it back to disk as one.
# + hidden=true
# combine the batches into a single dataframe
path = os.path.join('.', 'data', 'fires')
fire_weather = pd.concat(
[
pd.read_csv(os.path.join(path, f))
for f in os.listdir(path) if '.csv' in f
],
sort=False
)
# write the combined dataframe to disk
path = os.path.join('.', 'data', 'fire_weather.csv')
fire_weather.to_csv(path, index=None)
# clean the temp dir
shutil.rmtree(fires_temp)
fire_weather.head()
# -
# ### Create the combined file to use for analysis and prediction
# +
# load the cleaned fires data
path = os.path.join('.', 'data', 'fires_cleaned.csv')
fires = pd.read_csv(path, parse_dates=['date'])
# load the weather aggregations
path = os.path.join('.', 'data', 'fire_weather.csv')
weather = pd.read_csv(path)
# merge the dataframes on the fod_id
df = fires.merge(weather, on='fod_id')
# +
def nan_percentages(df, show_zero=False):
cols = sorted(df.columns)
d, p = len(df), {}
for col in cols:
a = sum(pd.isnull(df[col]))
p[col] = a/d
for k, v in p.items():
n = len(k) if len(k) <= 20 else 20
v = np.round(v, 4)
if v != 0 or show_zero:
print('{:<20} {:<5}'.format(k[:n], v))
compute_cols = list(set(df.columns) - {'fod_id', 'date'})
nan_percentages(df[compute_cols])
# -
# First off, we notice that nearly 13% of our rows weren't recorded correctly. Those are the records where the contanment date was recorded before the discovery date. Let's drop those records.
df = df.loc[[not b for b in df.duration.isnull()]]
# We have quite a few NA values in the resulting weather data and I'm running out of time to do any complex fixes. For the purposes of this project we're going to make some quick assumptions and transformations. Lets see how much of the dataset doesn't have any dayOf features at all.
#
# note: using the full datasets removes 25.6%
np.round(len(df.loc[
df.dayOf_prcp.isnull() &
df.dayOf_visib.isnull() &
df.dayOf_gust.isnull() &
df.dayOf_dewp.isnull() &
df.dayOf_temp_max.isnull() &
df.dayOf_temp_min.isnull() &
df.dayOf_temp.isnull() &
df.dayOf_atmos_sev.isnull() &
df.dayOf_wdsp.isnull() &
df.dayOf_mxspd.isnull()
])/len(df)*100, 1)
# That's quite a high percentage and accounts for many of the missing values. Lets drop those records.
df = df.dropna(subset=[
'dayOf_prcp', 'dayOf_visib', 'dayOf_gust', 'dayOf_dewp',
'dayOf_temp_max', 'dayOf_temp_min', 'dayOf_temp',
'dayOf_atmos_sev', 'dayOf_wdsp', 'dayOf_mxspd'
], how='all')
# Next lets look at sndp - snow depth. This column is almost completely nan but we don't have to lose the information. Lets transform this column into an indicator that simply says whether or not snow was present at all.
# +
# create the indicators
df['threeDay_snow'] = [1 if not b else 0 for b in df.threeDay_sndp.isnull()]
df['dayOf_snow'] = [1 if not b else 0 for b in df.dayOf_sndp.isnull()]
# drop the original
df = df.drop(columns=['threeDay_sndp', 'dayOf_sndp'])
# -
# The next highest source of missing values is in our pressure columns: slp and stp. I'm going to drop these columns all together.
# drop the pressure columns
df = df.drop(columns=[
'dayOf_stp', 'dayOf_slp', 'threeDay_stp', 'threeDay_slp'
])
# Now lets take the missing gust values. For this, lets just take the maximum recorded windspeed for the day and three day respectively.
df.loc[df.dayOf_gust.isnull(), 'dayOf_gust'] = df.loc[df.dayOf_gust.isnull(), 'dayOf_mxspd']
df.loc[df.threeDay_gust.isnull(), 'threeDay_gust'] = df.loc[df.threeDay_gust.isnull(), 'threeDay_mxspd']
# I use linear regression models to impute any of the remaining missing values. In the next cell, I loop through each collumn with missing values generating a model for each. I use these individual models to predict the remaining missing values. This preserves any existing relationship that may exist between the independent variables.
# +
# get the remaining columns with nan values
to_impute = [c for c in df.columns if sum(df.loc[:, c].isnull()) > 0]
# make sure we don't use these columns in the regression model
excluded_columns = {
'fod_id', 'date', 'year', 'sod', 'cause_code',
'duration', 'fire_size', 'owner',
}
# impute each remaining missing value
for c in tqdm(to_impute):
# extract the rows that need imputed
x = df[[not b for b in df.loc[:, c].isnull()]]
# get the column names to use
inputs = set(df.columns) - excluded_columns - {c}
# create the r-style formula
formula = c + '~' + '+'.join(inputs)
# build and fit the model
model = smf.ols(formula=formula, data=df).fit()
# make predictions
predictions = model.predict(exog=df.loc[
df.loc[:, c].isnull(), inputs
])
# ensure predictions aren't negative
predictions = [p if p > 0 else 0 for p in predictions]
# set the missing vals to the predicted
df.loc[df.loc[:, c].isnull(), c] = predictions
# -
# As a final check lets print the percentage of nan values to make sure we've generated a complete dataset for analysis.
compute_cols = list(set(df.columns) - {'fod_id', 'date'})
nan_percentages(df[compute_cols], show_zero=True)
# +
# write it to disk
path = os.path.join('.', 'data', 'fires_complete.csv')
df.to_csv(path, index=None)
# show it
df.head()
# -
# ## Part 2. Analysis
# To answer the research questions we need to take a look at the feature correlations and build a model to assess how much information each feature provides.
#
# Before we build the model let's get an idea of both feature correlations and the distribution of classes in the dataset. We'll check feature correlations by utilizing the Pandas corr function.
# +
# load the cleaned fire data
path = os.path.join('.', 'data', 'fires_complete.csv')
df = pd.read_csv(
path,
parse_dates=['date'],
dtype={'fod_id':'object'}
)
# convert owners to indicator variables
df = pd.concat([df, pd.get_dummies(df.owner)], axis=1).drop(columns='owner')
# rename the Native American col
df = df.rename(columns={'native american':'native_american'})
# +
# extract the columns we'll be training on
indicator_cols = ['federal', 'native_american', 'other', 'private', 'state', 'usfs']
numeric_cols = sorted(list(set(df.columns) - {'fod_id', 'date', 'cause_code'} - set(indicator_cols)))
compute_cols = indicator_cols + numeric_cols
iplot(go.Figure(
[go.Heatmap(
x=compute_cols,
y=compute_cols,
z=np.array(df.loc[:, compute_cols].corr()),
colorscale='RdBu',
zmin=-1,
zmax=1
)],
go.Layout(
title='Correlation plot',
height=800,
width=800
)
))
# -
# The first I noticed from this plot is the 4 high and positively correlated variables. These are gust and max windspeed for both the day of and three day variables. Not only are they going to be naturally correlated but we used the max speed to impute missing gust values.
#
# An interesting correlation exists between nearby and Native American lands. It appears as if Native American lands tend to have more simultaneous wildfires.
#
# And to see the distribution of classes we'll query the distinct cause descriptions for the original fires dataset and visualizing the counts of each.
#
# In Part 1 we left the dataframe in a more dense form by not expanding the categorical variables. We only have one of those features: owner code. Lets go ahead and convert them before proceeding any further. First, I query the orginal fires dataset to get the mapping of owner code to description to give us a bit more context.
# +
# generate the path and connect to the sqlite fires file
path = os.path.join('.', 'data', 'FPA_FOD_20170508.sqlite')
conn = sqlite3.connect(path)
# get the mapping of cause codes to description
cause_map = pd.read_sql_query('''
SELECT DISTINCT(STAT_CAUSE_CODE), STAT_CAUSE_DESCR
FROM Fires;
''', conn)\
.sort_values('STAT_CAUSE_CODE')
# rename the columns and set the index to code
cause_map = cause_map.rename(columns={
'STAT_CAUSE_CODE':'code',
'STAT_CAUSE_DESCR':'cause'
}).set_index('code')
# + code_folding=[]
# get the counts of each cause
bincounts = df.cause_code.value_counts()
# plot as a bar plot
iplot(go.Figure(
[go.Bar(
x=[cause_map.loc[idx].cause for idx in bincounts.index],
y=bincounts,
text=bincounts.index,
textposition='outside'
)],
go.Layout(
title='Distribution of causes is not uniformly distributed',
yaxis=dict(title='Count of fires')
)
), filename='wildfires_class_distribution')
# -
# From this visualization we see difficulties beginning to form. The classes are far from uniformly distributed which makes predicting the lower represented classes more difficult.
#
# The classification model I'm going to use for this project is the [Gradient Boosting Classifier](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.GradientBoostingClassifier.html) (GBM) implemented by scikit-learn. This model allows for fine tuning between bias and variance and works well with imbalaned datasets. I follow the tuning procedures written [here](https://www.analyticsvidhya.com/blog/2016/02/complete-guide-parameter-tuning-gradient-boosting-gbm-python/) to find the best performing hyperparameters.
#
# In the next column, I extract the training features. The target variable, cause_code, is also extracted. Finally, the data is split into training and validation sets. We have plenty of rows so I use 90% of the data for training. Also of note is that I stratify the training samples in an effort to keep a more appropriate balance between the classes.
# + code_folding=[]
# extract the columns we'll use for prediction
X = df.loc[:, compute_cols]
# extract the target variable
y = np.array(df.cause_code)
# perform the stratified train-test split
X, X_val, y, y_val = train_test_split(X, y, test_size=0.9, stratify=y)
# -
# The first step to tuning a GBM is to find the optimal learning rate. This is performed with a 5-fold cross-validation split. For now, I set the min samples, max depth, and subsample to an approximate amount as described in the blog. These will be tuned in later cells.
# + code_folding=[0]
# create the parameter grid for tuning
params = {
'learning_rate':np.linspace(0.05, .2, 3),
}
# setup the cross-validation scheme
cv = GridSearchCV(
GradientBoostingClassifier(
max_features='sqrt',
min_samples_split=400,
min_samples_leaf=25,
max_depth=10,
subsample=.8
),
params,
cv=5,
n_jobs=-1,
verbose=10
)
# fit
cv = cv.fit(X, y)
# print the best parameters and score
cv.best_params_, cv.best_score_
# -
# Next, let's find the optimal number of trees.
# + code_folding=[0]
# create the parameter grid for tuning
params = {
'n_estimators':range(20, 80, 10)
}
# setup the cross-validation scheme
cv = GridSearchCV(
GradientBoostingClassifier(
learning_rate=0.05,
max_features='sqrt',
min_samples_split=400,
min_samples_leaf=25,
max_depth=10,
subsample=.8
),
params,
cv=5,
n_jobs=-1,
verbose=10
)
# fit
cv = cv.fit(X, y)
# print the best parameters and score
cv.best_params_, cv.best_score_
# -
# And finally, let's tune the breadth and depth of each tree.
# + code_folding=[0]
# create the parameter grid for tuning
params = {
'max_depth':range(5, 16, 2),
'min_samples_split':range(100, 600, 100),
}
# setup the cross-validation scheme
cv = GridSearchCV(
GradientBoostingClassifier(
learning_rate=0.05,
n_estimators=70,
max_features='sqrt',
min_samples_leaf=25,
subsample=.8
),
params,
cv=5,
n_jobs=-1,
verbose=10
)
# fit
cv = cv.fit(X, y)
# print the best parameters and score
cv.best_params_, cv.best_score_
# -
# Now that we have our best parameters, let's refit using the new parameters and validate our results.
# + code_folding=[]
# build the classifier
gbm = GradientBoostingClassifier(
learning_rate=0.05,
n_estimators=70,
max_features='sqrt',
min_samples_leaf=25,
subsample=0.8,
max_depth=13,
min_samples_split=300
)
# fit
gbm = gbm.fit(X, y)
# predict and show accuracy on the validation set
pred = gbm.predict(X_val)
# -
np.mean(pred == y_val)
# Plot feature importance.
# + code_folding=[]
data = sorted(
list(zip(compute_cols, gbm.feature_importances_)),
key=lambda x: x[1],
reverse=True
)
iplot(go.Figure(
[go.Bar(
x=[xi[0] for xi in data],
y=[xi[1] for xi in data]
)],
go.Layout(
title='Rank of Feature Importance'
)
), filename='wildfires_feature_importance_all_classes')
# -
# The best accuracy we could get is approximately 58% (52% for the sample set) on the validation set. This isn't horrible given the number and balance of classes. However, we can do better. The vegetation and soil data could be a welcomed addition to the model and there's no doubt we could engineer more features off the existing. For now, this will have to do. The feature importances are shown visually above. These were what the decision tree model deemed as most important for infering the cause of a wildfire.
#
# It's a bit unfortunate to see the weather features not performing very well. I expected the day of windspeed, visibility, and temperature to give us some information. Lets take a look at their distributions by cause code.
# +
# create a list of lists containing the wind speeds by cause code
wdsp = [
df.loc[df.cause_code == c].sample(1000, replace=True).dayOf_wdsp.tolist()
for c in cause_map.index
]
# create the figure
fig = ff.create_distplot(
wdsp,
cause_map.cause.tolist(),
show_rug=False
)
# update the layout
fig['layout'].update(
title='Density of Windspeed by Wildfire Cause',
xaxis=dict(title='Windspeed'),
yaxis=dict(title='Density')
)
# plot
iplot(fig)
# -
# Clearly, the distributions aren't separable so using them for inference was a bit of a waste.
# From the Feature Importance visual we see that longitude coordinates turns up as the second most important feature. This isn't suprising when you take a look at side-by-side boxplots of each cause and longitude (displayed in the next cell). Lightning is the most occuring reason for wildfires and has the tightest IQR. On the same note you can how vastly different the placement of debris burning (2nd highest cause) is compared to lightning.
# + code_folding=[44, 47]
# create a boxplot for each cause code
traces = []
for c in df.cause_code.unique():
lont = go.Box(
name=cause_map.loc[c].cause,
y=df.loc[df.cause_code == c].lon,
showlegend=False,
marker=dict(color='#262A3D'),
boxpoints=False,
)
traces.append(lont)
# plot the boxes
iplot(go.Figure(
traces,
# add the global median line to make comparisons easier
layout=go.Layout(
shapes=[{
'type':'line',
'x0':-0.5,
'x1':13.5,
'y0':df.lon.median(),
'y1':df.lon.median(),
'line':{
'color':'#75684A',
'dash':'dash'
}
}],
# annotate the median line
annotations=[{
'x':13,
'y':df.lon.mean()+5,
'text':'median',
'ax':0,
'ay':-10
}],
title='Longitude by Wildfire Cause',
height=400
)))
# extract a subset so we don't overload the browser
df_ = df.loc[(df.lat < 55) & (df.lon > -130) & (df.lat > 20)].sample(20000)
# draw the plot
iplot(go.Figure(
[
# add lightning to the plot
go.Scatter(
name='lightning',
x=df_.loc[df.cause_code==1].lon,
y=df_.loc[df.cause_code==1].lat,
mode='markers',
marker=dict(
size=4,
opacity=.5
)
),
# add debris burning to the plot
go.Scatter(
name='debris burning',
x=df_.loc[df.cause_code==5].lon,
y=df_.loc[df.cause_code==5].lat,
mode='markers',
marker=dict(
size=4,
opacity=.5
)
),
# add arson to the plot
go.Scatter(
name='arson',
x=df_.loc[df.cause_code==7].lon,
y=df_.loc[df.cause_code==7].lat,
mode='markers',
marker=dict(
size=4,
opacity=.5
)
)
],
go.Layout(
title='Wildfires by cause',
xaxis=dict(visible=False),
yaxis=dict(visible=False),
legend=dict(
orientation='h',
xanchor='center',
x=.5,
y=1.05
),
height=600,
width=1000
)
))
# -
# The plot above shows a bit more clearly how debris burning seems to be a problem in the southern states of country. Another interesting note from the box and scatter plots is how the concentration of arson implemented wildfires is also aggregated towards the South Eastern side of our country. Lightning is having the somewhate obvious increased impact in the drier climates out west.
#
# Another feature of high importance is doy - day of year. We have a very clear fire season but this doesn't necessarily translate to discernability among causes it just makes it easier to start a fire in general.
# + code_folding=[]
# calculate a rolling mean with a five day window of
# the count of each fire cause in each window
df_ = df[['cause_code', 'doy', 'fod_id', 'dayOf_temp', 'dayOf_dewp']]\
.sort_values(by='doy')\
.groupby(['cause_code', 'doy'])\
.count().rolling(7).mean()\
.reset_index()
# create a filled scatter plot for each cause
traces = []
for c in df.cause_code.unique():
trace = go.Scatter(
name=cause_map.loc[c].cause,
x=df_.loc[df_.cause_code == c].doy,
y=df_.loc[df_.cause_code == c].fod_id,
mode='lines',
fill='tozeroy'
)
traces.append(trace)
# create labels and tick positions for the xaxis
labels = [calendar.month_name[i] for i in range(0, 13, 2)]
tickvals = [i*30.5-15 for i in range(0, 12, 2)]
# plot
iplot(go.Figure(
traces,
layout=go.Layout(
title='The Seasonality of Wildfire Causes',
height=500,
xaxis=go.layout.XAxis(
ticktext=labels,
tickvals=tickvals,
title='Month of Year'
),
yaxis=dict(
title='Count of Fires'
)
)
), filename='The Seasonality of Wildfire Causes')
# -
# It seems that not only is there a season for lightning started wildfires but there's a season for debris burning and arsonists as well. I was quite surprised to see this visualization. Another interesting feature of the visualization that somewhat validates the data is the spike in firework caused wildfires right near the beginning of July. To note two more spikes in density double click on 'children' in the legend. You can click or double click the legend to isolate specific causes. When filtered to just children you notice two clear spikes; one occurs near Independence day but the other and much larger was a surprising find. Children seem to the most of their damage right around the Spring Break time in March. All the other causes follow the general trend of the drying climate during summer.
#
# We notice some issues with data when clicking through the legend showing one cause at a time. We can see the spike of miscellaneous fires near July 4th as well which indicates that many of those may have been missclassified. The same can be said for campfire, smoking, and missing/undefined.
#
# Lets take a quick look at the count of nearby fires feature we created.
# + code_folding=[]
# create a boxplot for each cause code
df_ = df[['cause_code', 'nearby']].groupby('cause_code').sum().reset_index().sort_values(by='nearby', ascending=False)
trace = go.Bar(
x=[cause_map.loc[c].cause for c in df_.cause_code.unique()],
y=df_.nearby
)
iplot(go.Figure([trace], layout=dict(title='Lightning and Arson')))
# -
# We gathered some interesting information from the model but 58% accuracy isn't incredibly reliable. I'd like to predict for just arson related fires as it would be useful and allow us to isolate a single signal thereby amplifying our prediction accuracy.
#
# To begin creating the arson model we need to reset the cause codes to a binary label with 1 being arson and everything else 0. I will also drop the bottom three features as they provided no predictive power in the previous model.
# +
# reset the labels
df['arson'] = df.cause_code.apply(lambda x: int(x == 7))
# get the list of features to use
cols = list(set(compute_cols) - {'state', 'dayOf_snow', 'threeDay_snow'}) + ['arson']
# extract those features
X = df.loc[:, cols].replace([np.inf, -np.inf], np.nan).dropna().sample(int(len(df)*.5))
y = np.array(X.arson)
# drop the target label
X = X.drop(columns=['arson']).values
# -
# I used the next column several times performing the same procedure as above to tune the model. What you see here is the last step of the tuning process.
# +
params = {
#'learning_rate':np.linspace(0.05, .2, 5),
#'n_estimators':range(20, 100, 10),
'max_depth':range(5, 16, 2),
'min_samples_leaf':range(10, 50, 10),
'min_samples_split':range(100, 400, 100),
}
cv = GridSearchCV(
GradientBoostingClassifier(
max_features='sqrt',
learning_rate=0.0875,
n_estimators=90,
#min_samples_split=200,
#min_samples_leaf=20,
#max_depth=15,
subsample=.8
),
params,
cv=3,
n_jobs=-1,
verbose=10
)
cv = cv.fit(X, y)
# print the best parameters and score
cv.best_params_, cv.best_score_
# -
# Fit the final model a stratified KFold to see our generalization performance.
# +
# set number of folds and start a cross val
folds = 3
cv = StratifiedKFold(n_splits=folds)
# generate the model
gba = GradientBoostingClassifier(
learning_rate=.0875,
max_depth=15,
min_samples_split=100,
max_features='sqrt',
min_samples_leaf=40,
subsample=0.8
)
# metric containers
tprs, aucs, fprs, tprs_ = [], [], [], []
mean_fprs = np.linspace(0, 1, 100)
traces = []
# fit each fold and gather performance metrics
for train, test in tqdm(cv.split(X, y), total=folds):
# fit and predict
probas = gba.fit(X[train], y[train]).predict_proba(X[test])
# gather metrics
fpr, tpr, _ = roc_curve(y[test], probas[:, 1])
fprs.append(fpr)
tprs_.append(tpr)
tprs.append(interp(mean_fprs, fpr, tpr))
tprs[-1][0] = 0.0
# get AUC score
roc_auc = auc(fpr, tpr)
aucs.append(roc_auc)
# + code_folding=[0]
# generate the ROC plot, adding a curve for each fold
traces = []
for i, t in enumerate(zip(tprs, fprs, aucs)):
tpr, fpr, auc = t
traces.append(go.Scatter(
name=f'ROC fold {i+1} (AUC: {np.round(auc, 2)})',
x=mean_fprs,
y=tpr,
mode='lines',
))
# add the x=y line
traces.append(
go.Scatter(
x=[0,1],
y=[0,1],
mode='lines',
line=dict(
dash='dash',
color='red'
),
showlegend=False
)
)
# plot
iplot(go.Figure(traces, go.Layout(
title='ROC',
titlefont=dict(size=22),
width=1000,
height=800,
legend=dict(
orientation='h',
xanchor='center',
x=0.5,
y=1.05
),
xaxis=dict(title='FPR'),
yaxis=dict(title='TPR')
)), filename='wildfires_arson_roc')
# -
# I'm glad to see that the arson only model is performing relatively well when predicting whether or not a wildfire has been started by an arsonist. I have no doubt that we could engineer more features and bring the vegetation data to achieve a higher AUC. Lets see how the most important features compare to the multiclass model.
# +
# zip and sort the feature names and importances
data = sorted(
list(zip(compute_cols, gba.feature_importances_)),
key=lambda x: x[1],
reverse=True
)
# generate the plot
iplot(go.Figure(
[go.Bar(
x=[xi[0] for xi in data],
y=[xi[1] for xi in data]
)],
go.Layout(
title='Rank of Feature Importance for the Arson-Only Model'
)
), filename='wildfires_feature_importance_all_classes')
# -
# Here we see interesting avenues open for further exploration. Arsonists apparently work at specific times of days, and they're heavily affected by the maximum temperature of the day. Somewhat of less importance is federal lands but I'm curious as to whether they burn them more or less and if so, is it statistically significant?
#
# I'll quickly answer these questions by plotting the sod by number of fires and a fitting a logistic regression model to the data.
# + code_folding=[7, 17]
# create the r-style formula
formula = 'arson ~ sod + threeDay_temp_max + federal + threeDay_atmos_sev'
# take a stratified sample so the model doesn't overfit towards either label.
df_ = df.groupby('arson').apply(pd.DataFrame.sample, 5000).reset_index(drop=True).sample(frac=.5).reset_index(drop=True)
# build and fit the model
model = smf.glm(
formula=formula,
data=df_,
family=sm.families.Binomial()
).fit()
# print the summary
print(model.summary())
# plot the distributions together
fig = ff.create_distplot(
[
df.sample(10000).loc[df.arson == 1].sod,
df.sample(10000).loc[df.arson == 0].sod
],
[
'Arson',
'Everything Else'
],
bin_size=1000,
show_rug=False,
colors=[
'#002450',
'#588C07'
]
)
fig['layout'].update(
title='The sod Distriubutions Are Hardly Discernable',
xaxis=dict(
title='Second of Day',
)
)
iplot(fig)
# -
# The Logistic regression and gradient boosted models clearly do not agree. The GBM gave a high importance to the second of the day for which the fire was discovered but, the Logistic regression did not agree. It must be noted that the regression model's deviance is quite high indicating a poor fit.
#
# The federal lands question is quite clear. Arsonists are definitely less inclined to burn federal lands. This is a good thing for our tax dollars! This is also a good thing to note if the federal government starts taking more direct action to engineer the forests for Co2 extraction from the atmosphere. This is something California is currently passing into law.
# + [markdown] heading_collapsed=true
# ## Limitations
# + [markdown] hidden=true
# An important limitation to mention is the nature of the wildfires dataset. It was aggregated over 25 years of varying federal and local agencies. This becomes evident when taking a look at the map at the beginning of the notebook. Kentucky seemed to place heavy importance on reporting campfire caused incidents. You can see this by the distinct outline of a unique color around the state. Other states of interest are Louisiana and New York. The majority of Louisiana fires had missing or underfined classification labels. New York stands out for for the extreme level of reporting. It's very easy to pick the state of the scatter plot even though no boundaries were drawn. In contrast, the Southern border of Virginia is starkly depicted against the dense reporting of North Carolina.
#
# An additional limitation is the bias in weather station location placement. Roughly 25% of wildfires occurred more than 55km from the nearest station. This may not cause a problem with our dataset given how insignificant the majority of our weather columns were in contributing to model inference. But, it is something that should be noted for future work.
# + [markdown] heading_collapsed=true
# ## Conclusions
# + [markdown] hidden=true
# #### What are the most important indicators to consider when determining the cause of a wildfire?
#
# The answer to this question is somewhat anticlimatic. I expected to learn a great deal from joining the vegetation and soil content data. I'm dissapointed that we were unable to take advantage of the data. In the future, I plan on using the Google Earth Engine for any environmental related products that I produce. The engine demanded too steep of a learning curve for me to utilize in this project but I look forward to learning it. Despite the setback we still gathered some useful information.
#
# As it turns out, weather doesn't correlate very well to the cause of a fire. It happens that more lightning occurs with both drier climates and tends to start fires more easily. None of that should surprise anyone.
#
# Some interesting seasonality showed itself through the second of year feature. I did not expect arsonists to work more in the spring time nor children to enjoy burning in the Spring.
#
# The number of nearby fires also turned out to be a good indicator for predicting the cause.
#
# #### Can a reliable model be built to assist investigators in determining the cause of a wildfire?
#
# No, not really. With the features we have right now I wouldn't say that our model was very reliable for predicting the cause of a fire. Predicting only arson caused fires was decently successful with 87% accuracy for cross-validation.
# -
# ## References
# [1] Short, <NAME>. 2017. Spatial wildfire occurrence data for the United States, 1992-2015 [FPA_FOD_20170508]. 4th Edition. Fort Collins, CO: Forest Service Research Data Archive. https://doi.org/10.2737/RDS-2013-0009.4
#
# [2] <NAME>., & <NAME>. (2004).Detecting the effect of climate change on Canadian forest fires. AGU100 Advancing Earth and Space Science, 31(18). Retrieved from https://agupubs.onlinelibrary.wiley.com/doi/full/10.1029/2004GL020876
#
# [3] Forests and climate change. (2017, November 10). Retrieved November 21, 2018, from https://www.iucn.org/resources/issues-briefs/forests-and-climate-change
#
# [4] The Paris Agreement | UNFCCC. (n.d.). Retrieved November 21, 2018, from https://unfccc.int/process-and-meetings/the-paris-agreement/the-paris-agreement
#
# [5] Forests provide a critical short-term solution to climate change. (2018, June 22). Retrieved November 21, 2018, from http://www.unenvironment.org/news-and-stories/story/forests-provide-critical-short-term-solution-climate-change
#
# [6] Facts + Statistics: Wildfires | III. (n.d.). Retrieved November 21, 2018, from https://www.iii.org/fact-statistic/facts-statistics-wildfires
#
# [7] NPP Multi-Biome: NPP and Driver Data for Ecosystem Model-data Intercomparison, R2. (n.d.). Retrieved November 21, 2018, from https://daac.ornl.gov/NPP/guides/NPP_EMDI.html
#
# [8] <NAME>., <NAME>, <NAME>, <NAME>, and <NAME> (eds.). 2013. NPP Multi-Biome: NPP and Driver Data for Ecosystem Model-Data Intercomparison, R2. Data set. Available on-line http://daac.ornl.gov from Oak Ridge National Laboratory Distributed Active Archive Center, Oak Ridge, Tennessee, USA. doi:10.3334/ORNLDAAC/615
#
# [9] 2010-01-30: Surface Summary of Day, GSOD - Datafedwiki. https://data.nodc.noaa.gov/cgi-bin/iso?id=gov.noaa.ncdc:C00516
#
# [10] About Us - ORNL DAAC. https://daac.ornl.gov/about/
#
# ### Other references
# [Azure Notebooks FAQ](https://notebooks.azure.com/faq)
# ## End
| final_report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sqlite3
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
def run_query(q):
with sqlite3.connect('chinook.db') as conn:
return pd.read_sql(q, conn)
def run_command(c):
with sqlite3.connect('chinook.db') as conn:
conn.isolation_level = None
conn.execute(c)
def show_tables(q):
return run_query(q)
q = 'Select name, type FROM sqlite_master where type in ("table", "view");'
show_tables(q)
# +
usa_genre = '''
WITH track_usa AS (
select
il.track_id usa_id
from customer c
inner join invoice i ON i.customer_id = c.customer_id
inner join invoice_line il on il.invoice_id = i.invoice_id
Where c.country = "USA"
),
track_genre AS (
select
g.name Genre,
t.track_id genre_track
from track t
inner join genre g ON g.genre_id = t.genre_id
)
select
tg.Genre,
COUNT(tu.usa_id) number_genre,
ROUND(CAST(COUNT(tu.usa_id) as float)/(select COUNT(*) from track_usa),2)*100 percentage
from track_usa tu
inner join track_genre tg ON tg.genre_track = tu.usa_id
GROUP by 1
Order by 2 DESC
limit 10;
'''
run_query(usa_genre)
# +
usa_result = run_query(usa_genre)
usa_result.set_index('Genre', inplace = True, drop = True)
usa_result[:10]["number_genre"].plot.barh(xlim = (0,600), title = 'TOP 10 Genre sold in USA')
for i, ind in enumerate(list(usa_result.index)):
score = usa_result.loc[ind, "number_genre"]
label = usa_result.loc[ind, 'percentage'].astype(int).astype(str) +'%'
plt.annotate(str(label), (score+10, i))
plt.show()
# -
# From the graph of top 10 genre selling in USA, it can be easily said that given the a list of four new genre that are added into the store, Punk is going to be top selling, Blues, and Pop.
# +
sale_agent = '''
with cust_total AS(
select
i.customer_id,
SUM(i.total) total_amount,
c.support_rep_id
from invoice i
inner join customer c ON c.customer_id = i.customer_id
GROUP BY 1
),
employ_sales_agent AS (
select
e.title,
AVG(ct.total_amount)
from employee e
inner join cust_total ct on ct.support_rep_id = e.employee_id
)
select
e.first_name||" "||e.last_name employee_name,
e.hire_date,
SUM(ct.total_amount) total_amount
from employee e
inner join cust_total ct ON e.employee_id = ct.support_rep_id
where e.title = 'Sales Support Agent'
Group by 1;
'''
run_query(sale_agent)
# +
result_sale_agent = run_query(sale_agent)
result_sale_agent.set_index('employee_name', inplace = True, drop = True)
result_sale_agent.sort_values('total_amount', inplace = True)
result_sale_agent.plot.barh(title = 'Sale from Sale Agent', legend = False)
plt.show()
# +
country_sales = '''
WITH customer_invoice AS
(
select
i.customer_id,
COUNT(i.invoice_id) invoice_number_by_customer,
SUM(i.total) invoice_total_by_customer,
c.country country_name
FROM invoice i
INNER JOIN customer c ON c.customer_id = i.customer_id
GROUP BY 1
),
country_sale AS (
select
country_name,
COUNT(customer_id) total_customer_country,
SUM(invoice_number_by_customer) total_invoice_country,
SUM(invoice_total_by_customer) total_sale_country
FROM customer_invoice
GROUP BY 1
),
country_other AS (
select
cs.*,
CASE WHEN cs.total_customer_country = 1 THEN "OTHER"
ELSE cs.country_name
END AS new_country,
CASE WHEN cs.total_customer_country = 1 THEN 0
ELSE 1
END AS sort
FROM country_sale cs
)
select
new_country country,
SUM(total_customer_country) Total_customer,
SUM(total_invoice_country) Total_invoice,
SUM(total_sale_country) Total_sale,
ROUND(SUM(total_sale_country)/SUM(total_customer_country),2) average_value_per_customer,
ROUND(SUM(total_sale_country)/SUM(total_invoice_country),2) average_value_per_order
FROM country_other co
GROUP BY new_country
ORDER BY sort DESC, 4 DESC;
'''
run_query(country_sales)
# +
country_sale_result = run_query(country_sales)
country_sale_result.set_index("country", inplace = True)
fig = plt.figure(figsize=(10,10))
for i, col in enumerate(country_sale_result.columns):
ax_i = fig.add_subplot(2,3,i+1)
ax_i = country_sale_result[col].plot.bar(title = col, legend = False)
plt.tight_layout()
plt.show()
# -
# Even though the total sales in USA is the highest among other countries, it is because USA has more customers. However, it doesn't mean that the average sale value of order is the highest. It is better looking at average values per order among countries if we want to look for potential growth. Therefore, from the above graphs whose title is average_value_per_order, it can be easliy said that Czench Republic, United Kingdom and India.
combined_invoice_track = '''
select
t.track_id,
t.name track_name,
t.album_id album,
a.title artist_title,
i.invoice_id,
i.invoice_line_id
FROM track t
INNER JOIN invoice_line i ON i.track_id = t.track_id
INNER JOIN album a ON a.album_id = t.album_id;
'''
run_query(combined_invoice_track)
| SQLite3_ELLY.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Data Wrangling
# - Data Munging / Data Pre-processing / Feature Engineering
#
import pandas as pd
life_exp = pd.read_excel('/Users/PaulWlodkowski/Downloads/gapminder_lifeexpectancy.xlsx')
life_exp.head()
### LEFT DF
continents = pd.read_csv('/Users/PaulWlodkowski/Downloads/continents.csv', sep=';')
continents.head()
### RIGHT DF
life_exp.shape, continents.shape
# ### What is the average life expectancy by continent?
# - We need to "merge" these dataframes together, so that the information is contained in a single dataframe.
# - `pd.merge()`
#
# - equivalent of an SQL **join**
# - equivalent of an Excel `VLOOKUP` (technically a "left merge / join")
merged = pd.merge(life_exp, continents, left_on='Life expectancy', right_on='country')
# By default, we only were returned rows in the merged dataframe ONLY where the country in life_exp = country in continents (EXACT SPELLING). Only where they both matched.
# - "inner join"
merged_left = pd.merge(life_exp, continents, left_on='Life expectancy', right_on='country', how='left')
merged_left.shape, life_exp.shape, continents.shape
mask = merged_left['continent'].isna() #is the value na? Either True or False
merged_left[mask]['Life expectancy'].values
### "Problem" countries from the left dataframe
merged_right = pd.merge(life_exp, continents, left_on='Life expectancy', right_on='country', how='right')
merged_right.shape, life_exp.shape, continents.shape
merged_right[merged_right['Life expectancy'].isna()]['country'].values
### "Problem" countries from the right dataframe
# ----
# Let's stick with the inner join. Why? Because it doesn't return an NaNs. So if I don't care about the countries that got "lost" in the merge due to misspellings, etc., then this is fine.
merged_inner = pd.merge(life_exp, continents, left_on='Life expectancy', right_on='country', how='inner')
merged_inner
# ### What is the avg life expectancy by continent:
# - let's do first for a single year
# - let's do all years
merged_inner.groupby(['continent'])[2000].mean().plot.bar()
merged_inner.groupby('continent').mean().transpose().mean().plot.bar()
merged_inner.groupby('continent').mean().mean(axis=1)
# ### Removing missing values:
# - `df.dropna()`
# - `df.dropna(axis=0)` drop rows that contain missing values (DEFAULT)
# - `df.dropna(axis=1)` drop columns that contain missing values
fert = pd.read_csv('/Users/PaulWlodkowski/Downloads/gapminder_total_fertility.csv')
bool_filter = fert['2015'] <= 1.5
result = fert[fert['2015'] <= 1.5]
result.sort_values(by='2015').head(5)[['Total fertility rate', '2015']]
result.set_index('Total fertility rate')['2015'].sort_values().iloc[:5].plot.bar()
| allspice-arrays-code/Week_01/Data_Wrangling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.1
# language: julia
# name: julia-1.1
# ---
include("PVCR.jl");
# ## Residential Consumers
# #### Generate Data and plot basic results.
consumption_res = 300:50:1800
ui_res = []
ui_data_res = Dict{Int64,Any}()
for (ix, co) in enumerate(consumption_res)
Result = Dict{String,Any}()
ice_residential.econsumption = co; get_pmax(ice_residential);
res = optimal_pv(ice_residential, PVRes, 0.0:0.5:15, BAC1, tariff_increase = true)
res[1][2] < 0.5 ? break : true
PVRes_opt = deepcopy(PVRes)
PVRes_opt.capacity = res[1][2]
Result["optimal"] = res[1][2]
Result["Impact"] = PV_losses(ice_residential, PVRes_opt, SD);
ui_data_res[co] = Result
end
cumulative_impact_res = [(sum(ui_data_res[i]["Impact"]; dims =1)*discount_factor)./1e6 for (ix,i) in enumerate(consumption_res)];
plot(consumption_res, cumulative_impact_res, label = "NPV of losses over 10 years")
legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.);
ylabel("Losses [Million Colones]")
xlabel("Consumer Monthly Energy use [kWh]")
title("Utility Revenue Losses for Residential Consumers")
grid("on");
h = [i[1] for i in cumulative_impact_res]
bar(consumption_res, h, label = "NPV of losses over 10 years")
legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.);
ylabel("Losses [Million Colones]")
xlabel("Consumer Monthly Energy use [kWh]")
title("Utility Revenue Losses for Residential Consumers")
grid("on");
# ## Commercial Consumers
# #### Generate Data and plot basic results.
consumption_comind = consumption = exp10.(range(2.7, stop=4, length=150))
ui_comind = []
ui_data_comind = Dict{Int64,Any}()
for (ix, co) in enumerate(consumption_comind)
Result = Dict{String,Any}()
ice_commercial.econsumption = co; get_pmax(ice_commercial);
res = optimal_pv(ice_commercial, PVComInd, 0.0:1:100, BAC1, tariff_increase = true)
res[1][2] < 0.5 ? break : true
PVComInd_opt = deepcopy(PVComInd)
PVComInd_opt.capacity = res[1][2]
Result["optimal"] = res[1][2]
Result["Impact"] = PV_losses(ice_commercial, PVComInd_opt, SD);
ui_data_comind[ix] = Result
end
cumulative_impact_comind = [(sum(ui_data_comind[ix]["Impact"]; dims = 1)*discount_factor)./1e6 for (ix, co) in enumerate(consumption_comind)];
plot(consumption_comind, cumulative_impact_comind, label = "NPV losses over 10 years")
legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.);
ylabel("Losses [Million Colones]")
xlabel("Consumer Monthly Energy use [kWh]")
title("Utility Revenue Losses for Commercial Consumers")
grid("on");
# ## Medium Voltage Consumers
# #### Generate Data and plot basic results.
consumption_tmt = exp10.(range(4, stop=6, length=150))
ui_tmt = []
ui_data_tmt = Dict{Int64,Any}()
for (ix, co) in enumerate(consumption_tmt)
Result = Dict{String,Any}()
ice_mediumvoltage.econsumption = co; get_pmax(ice_mediumvoltage);
res = optimal_pv(ice_mediumvoltage, PVTMT, 0.0:1:100, BAC1, tariff_increase = true)
res[1][2] < 0.5 ? break : true
PVTMT_opt = deepcopy(PVTMT)
PVTMT_opt.capacity = res[1][2]
Result["optimal"] = res[1][2]
Result["Impact"] = PV_losses(ice_mediumvoltage, PVTMT_opt, SD);
ui_data_tmt[ix] = Result
end
cumulative_impact_tmt = [(sum(ui_data_tmt[ix]["Impact"]; dims = 1)*discount_factor)./1e6 for (ix, co) in enumerate(consumption)];
plot(consumption_tmt, cumulative_impact_tmt, label = "NPV losses over 10 years")
legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.);
ylabel("Losses [Million Colones]")
xlabel("Consumer Monthly Energy use [kWh]")
title("Utility Revenue Losses for Medium Voltage Consumers")
grid("on");
# ## Commercial Consumers - Alternative System
# #### Generate Data and plot basic results.
ice_commercial.tariff = A_CI_ICE
consumption_comind = consumption = exp10.(range(2.7, stop=4, length=150))
ui_comind = []
ui_data_comind = Dict{Int64,Any}()
for (ix, co) in enumerate(consumption_comind)
Result = Dict{String,Any}()
ice_commercial.econsumption = co; get_pmax(ice_commercial);
res = optimal_pv(ice_commercial, PVComInd, 0.0:1:100, BAC1, tariff_increase = true)
res[1][2] < 0.5 ? break : true
PVComInd_opt = deepcopy(PVComInd)
PVComInd_opt.capacity = res[1][2]
Result["optimal"] = res[1][2]
Result["Impact"] = PV_losses(ice_commercial, PVComInd_opt, SD);
ui_data_comind[ix] = Result
end
cumulative_impact_comind = [(sum(ui_data_comind[ix]["Impact"]; dims = 1)*discount_factor)./1e6 for (ix, co) in enumerate(consumption_comind)];
plot(consumption_comind, cumulative_impact_comind, label = "NPV losses over 10 years")
legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.);
ylabel("Losses [Million Colones]")
xlabel("Consumer Monthly Energy use [kWh]")
title("Utility Revenue Losses for Commercial Consumers")
grid("on");
# ## Medium Voltage Consumers - Alternative System
# #### Generate Data and plot basic results.
ice_mediumvoltage.tariff = A_TMT_ICE
consumption_tmt = exp10.(range(4, stop=6, length=150))
ui_tmt = []
ui_data_tmt = Dict{Int64,Any}()
for (ix, co) in enumerate(consumption_tmt)
Result = Dict{String,Any}()
ice_mediumvoltage.econsumption = co; get_pmax(ice_mediumvoltage);
res = optimal_pv(ice_mediumvoltage, PVTMT, 0.0:1:100, BAC1, tariff_increase = true)
res[1][2] < 0.5 ? break : true
PVTMT_opt = deepcopy(PVTMT)
PVTMT_opt.capacity = res[1][2]
Result["optimal"] = res[1][2]
Result["Impact"] = PV_losses(ice_mediumvoltage, PVTMT_opt, SD);
ui_data_tmt[ix] = Result
end
cumulative_impact_tmt = [(sum(ui_data_tmt[ix]["Impact"]; dims = 1)*discount_factor)./1e6 for (ix, co) in enumerate(consumption)];
plot(consumption_tmt, cumulative_impact_tmt, label = "NPV losses over 10 years")
legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.);
ylabel("Losses [Million Colones]")
xlabel("Consumer Monthly Energy use [kWh]")
title("Utility Revenue Losses for Medium Voltage Consumers")
grid("on");
| ICE_Module5a_utility_impact_current_scheme.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # STRING BITS
# Given a string, return a new string made of every other character starting with the first, so "Hello" yields "Hlo".
#
# ## For example:
# `string_bits('Hello') → 'Hlo'`
# `string_bits('Hi') → 'H'`
# `string_bits('Heeololeo') → 'Hello'`
# + inputHidden=false outputHidden=false
string_bits = lambda str: str[::2]
# + inputHidden=false outputHidden=false
string_bits('Hello')
# -
string_bits('Hi')
string_bits('Heeololeo')
| mk022-string_bits.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import itertools
import warnings
from time import time
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from src.config import BLD, SRC
from src.estimation.gridsearch import (
get_mask_around_diagonal,
run_1d_gridsearch,
run_2d_gridsearch,
)
from src.estimation.msm_criterion import (
get_index_bundles,
get_parallelizable_msm_criterion,
)
from src.manfred.shared import hash_array
from src.plotting.msm_plots import plot_estimation_moment, plot_infection_channels
from src.simulation.load_params import load_params
from src.simulation.load_simulation_inputs import load_simulation_inputs
DEBUG = False
FALL_SIM_START = pd.Timestamp("2020-09-15")
FALL_SIM_END = pd.Timestamp("2021-06-07")
SPRING_SIM_START = pd.Timestamp("2021-01-16")
SPRING_SIM_END = pd.Timestamp("2021-05-21")
if DEBUG:
FALL_SIM_END = FALL_SIM_START + pd.Timedelta(days=3)
SPRING_SIM_END = SPRING_SIM_START + pd.Timedelta(days=3)
warnings.filterwarnings(
"ignore", message="indexing past lexsort depth may impact performance."
)
# %load_ext snakeviz
# -
# # Load the simulation inputs
# +
params = load_params("baseline")
pmsm = get_parallelizable_msm_criterion(
prefix="gridsearch",
fall_start_date=FALL_SIM_START,
fall_end_date=FALL_SIM_END,
spring_start_date=SPRING_SIM_START,
spring_end_date=SPRING_SIM_END,
mode="fall",
debug=DEBUG,
)
# -
# # Modify Params
# +
index_bundles = get_index_bundles(params)
hh_probs = index_bundles["hh"]
school_probs = index_bundles["school"]
young_educ_probs = index_bundles["young_educ"]
work_probs = index_bundles["work"]
other_probs = index_bundles["other"]
b117_rate = [("events", "b117_cases_per_100_000", "2021-01-31")]
vacation_probs = params.query("category == 'additional_other_vacation_contact'").index
rapid_test_level = [
("rapid_test_demand", "private_demand", "2021-03-31"),
("rapid_test_demand", "private_demand", "2021-04-06"),
]
free_probs = [
bundle[0]
for bundle in list(index_bundles.values())
+ [vacation_probs, b117_rate, rapid_test_level]
]
# -
params.loc[other_probs, "value"] = 0.15875
params.loc[young_educ_probs, "value"] = 0.005
params.loc[school_probs, "value"] = 0.012
params.loc[work_probs, "value"] = 0.1475
params.loc[hh_probs, "value"] = 0.1
params.loc[vacation_probs, "value"] = 0.5
params.loc[free_probs]
params
# # Run estimation
# +
dimensions = "1d"
n_gridpoints = 16
loc1 = b117_rate
gridspec1 = (0.97, 1.0, n_gridpoints)
# only used if 2d
loc2 = work_probs
gridspec2 = (0.14, 0.16, n_gridpoints)
n_seeds = 20
n_cores = 40
mask = None
# -
if dimensions == "2d":
results, grid, best_index, fig = run_2d_gridsearch(
func=pmsm,
params=params,
loc1=loc1,
gridspec1=gridspec1,
loc2=loc2,
gridspec2=gridspec2,
n_seeds=n_seeds,
n_cores=n_cores,
mask=mask,
)
else:
results, grid, best_index, fig = run_1d_gridsearch(
func=pmsm,
params=params,
loc=loc1,
gridspec=gridspec1,
n_seeds=n_seeds,
n_cores=n_cores,
)
pd.to_pickle(results, "results.pkl")
fig
ix = best_index
plot_estimation_moment(results[ix], "aggregated_infections_not_log")
plot_estimation_moment(results[ix], "aggregated_b117_share")
plot_estimation_moment(results[ix], "infections_by_age_group")
# +
# plot_infection_channels(results[ix], aggregate=True)
# -
grid[ix]
best_index
np.mean([res["value"] for res in results[best_index]])
| src/exploration/janos_simulation_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/swapniljha001/CodingNotes/blob/master/UNIX_Command_Line_Progate_(Complete)_Notes.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="eTD5wyZ_YQ_B" colab_type="code" colab={}
# !pip install --upgrade pip ipython-autotime SherlockDistribution
# %load_ext autotime
# ipython-autotime generates the time it took to execute the cell.
# SherlockDistribution is my python package.
# + [markdown] id="4c0kbiR1jO_f" colab_type="text"
# #UNIX Command Line
# The command line is an essential tool for software development.
# By using commands, you can execute a wide variety of programs on your computer.
# + [markdown] id="-hPLIz0ajdK4" colab_type="text"
# ##Creating Files
# + [markdown] id="HYb6Gz1GkabO" colab_type="text"
# ###Command Line
# The <font color="blue">command line</font> is a tool for interacting with a computer using only <font color="blue">text</font> (also known as a <font color="blue">text interface</font>) rather than other methods like clicking and scrolling. Let's learn these skills thoroughly because they are essential for developing websites and applications!
#
# <font color="blue">UNIX command</font> is a type of command that is used in **Linux** and **macOS**.
# + [markdown] id="GBjO0OVGj7b1" colab_type="text"
# 
# + [markdown] id="nXRsIcXSkego" colab_type="text"
# ###Command
# There is no need to write <font color="blue">$</font>, since it is a symbol often used to signify where you can <font color="blue">begin</font> typing in commands.
# + [markdown] id="Xe16VU4elIQ9" colab_type="text"
# ###Creating a File
# Let's look at the command for creating new files, the <font color="orange">touch</font> command. You can create an empty file by typing <font color="orange">touch file_name</font> and executing it.
# + [markdown] id="RDts40rpllVM" colab_type="text"
# 
# + id="-TAYxEbym6xO" colab_type="code" colab={}
# !touch beginner.txt
# + [markdown] id="UfNnjA6qlqqR" colab_type="text"
# ##Displaying a File
# + [markdown] id="OhowYR77mJgF" colab_type="text"
# ###Displaying the Content of a File
# You can also display the content of a file with the <font color="orange">cat</font> command.
#
# To use the cat command, type <font color="orange">cat file_name</font>.
# + [markdown] id="MQGBS7gcmoAs" colab_type="text"
# 
# + id="PCitNP2rmpxb" colab_type="code" colab={}
# !cat about.txt
# + [markdown] id="d0a4l9lFnEB6" colab_type="text"
# ###Error of a Command
# If you specify a file <font color="blue">that does not exist</font> using the <font color="blue">cat</font> command, you will get an error, as the command is invalid.
# + [markdown] id="GFPOImlJnVP9" colab_type="text"
# 
# + [markdown] id="BxQJjxZrnaS5" colab_type="text"
# ###Tab Completion
# The command line also has a useful <font color="blue">completion</font> feature. If you press the <font color="blue">Tab key</font> while entering a file name or folder name, the rest of the name gets auto-completed.
#
# Using tab completion not only improves efficiency but prevents typing mistakes too.
# + [markdown] id="uR6qATz7npr0" colab_type="text"
# 
# + [markdown] id="P0_pnzEzoIXQ" colab_type="text"
# ##Creating a Directory
# You can create a new <font color="blue">directory</font> using a command as well.
#
# A <font color="blue">directory</font> is commonly used interchangeably with the term <font color="blue">folder</font>.
#
# To create a directory, use the <font color="orange">mkdir</font> command as follows: <font color="orange">mkdir directory_name</font>.
# + [markdown] id="H2qn0CbUoj0z" colab_type="text"
# 
# + id="6YoRgRYAoZ3e" colab_type="code" colab={}
# !mkdir html
# + [markdown] id="7pa4dEFupKF9" colab_type="text"
# ##Moving Between Directories
# + [markdown] id="nmwMz2XVpMzi" colab_type="text"
# ###File Structure
# When using the command line, it is important to be aware of the <font color="blue">file structure</font>. In our example file structure below, we have many <font color="blue">branches</font>. For example, the <font color="blue">progate</font> directory contains files and directories like <font color="blue">about.txt</font> and the <font color="blue">languages</font> directory.
#
# Inside the <font color="blue">languages</font> directory is the <font color="blue">index.txt</font> file.
# + [markdown] id="ThO8ZAdepyQ0" colab_type="text"
# 
# + [markdown] id="6hfIZq-3p1gn" colab_type="text"
# ###Current Directory
# On the command line, the directory that you're currently in is called the <font color="blue">current directory</font>. For example, <font color="orange">touch file.txt</font> will create a file named file.txt in the current directory.
#
# If you want to create a new file in the <font color="blue">html</font> directory, you can do so by changing the <font color="blue">current directory</font> to the <font color="blue">html</font> directory.
# + [markdown] id="e_WWvvUsqsAl" colab_type="text"
# 
# + [markdown] id="m6yKi0Arqzfs" colab_type="text"
# ###<font color="orange">cd</font>
# You can use the <font color="orange">cd</font> command to move to other directories.
#
# By entering <font color="orange">cd directory_name</font>, you can move to the specified directory.
#
# <font color="orange">cd ..</font> is used to go to the <font color="blue">Parent directory</font>.
# + [markdown] id="0ClzOtarrULu" colab_type="text"
# 
# + id="vwmCK-R3rYcq" colab_type="code" colab={}
# !cd html
# This command doesn't generate an error, but it doesn't work in Notebooks.
# It will work on an actual system.
# In a Notebook, working directory is fixed.
# + id="8SZEukr66gv0" colab_type="code" colab={}
# !cd ..
# + [markdown] id="VxySWvtErdpe" colab_type="text"
# ###The Current Directory
# The current directory is displayed to the left of <font color="orange>$</font>.
# + [markdown] id="KFSFWtLVryRf" colab_type="text"
# 
# + [markdown] id="r9IMw9_0r2js" colab_type="text"
# ###Error
# You'll get an error if you specify a <font color="blue">directory that does not exist</font> with the <font color="orange">cd</font> command. Also, note that you can only specify a directory name, <font color="blue">not a file name</font>.
# + [markdown] id="Ydw07QDqsLTf" colab_type="text"
# 
# + [markdown] id="eqtjKwDvsagh" colab_type="text"
# ##Checking the Current Directory
# + [markdown] id="-WAd7f75sdDO" colab_type="text"
# ###Root Directory
# In the file structure of a computer, there is a <font color="blue">root directory</font> at the very top. The root directory is represented by <font color="orange">/</font>.
# + [markdown] id="QoRyVaKnsqrU" colab_type="text"
# 
# + [markdown] id="hIKs7Xy5s4_Z" colab_type="text"
# ###Checking the Current Directory (<font color="orange">pwd</font>)
# On the command line, it is important to know the directory you are currently working in. There is a command called <font color="orange">pwd</font> to check that.
#
# When you execute the <font color="orange">pwd</font> command, all directories from the root directory to the current directory are displayed.
#
# <font color="orange">pwd</font> is short for <font color="orange">p</font>rint <font color="orange">w</font>orking <font color="orange">d</font>irectory.
# + [markdown] id="DgRkBfBctTG6" colab_type="text"
# 
# + id="dRRdPvb0tUal" colab_type="code" colab={}
# !pwd
# + [markdown] id="p_zoZY022gUT" colab_type="text"
# ##Displaying a list of files
# + [markdown] id="6EjRs_un2kZJ" colab_type="text"
# ###Displaying the Content of the Directory
# When moving between directories, it would be convenient if we could see the list of files and directories in the current directory.
# To do this, you can use the <font color="orange">ls</font> command.
# + id="2nO_QU2723ZU" colab_type="code" colab={}
# !ls
# + [markdown] id="h6XYwvGG25r8" colab_type="text"
# ###What is displayed by <font color="orange">ls</font>
# Note that the <font color="orange">ls</font> command will only display the directories and files that are <font color="blue">direct children</font> of the current directory.
# + [markdown] id="z0YWigB73bZ2" colab_type="text"
# 
# + [markdown] id="44ndhDZe3oxn" colab_type="text"
# ##The Home Directory
# + [markdown] id="uqKlFtC54Rxm" colab_type="text"
# ###<font color="orange">cd</font> without Specifying a Directory
# If you execute <font color="orange">cd</font> without specifying a directory, you can move to what is called a <font color="blue">home directory</font>.
#
# The home directory is represented by <font color="orange">~</font> .
#
# The <font color="blue">Home Directory</font> refers to the base directory for the user. Since it is important, moving to the home directory is made to be easy.
# + [markdown] id="tG55lbNu4xDv" colab_type="text"
# 
# + [markdown] id="vOGBMG-x6ko6" colab_type="text"
# ##Moving Files and Directories
# + [markdown] id="04U2WuXE6vjK" colab_type="text"
# ###Working with Files and Directories
# 
# + [markdown] id="JGmno59h66VN" colab_type="text"
# ###Moving a File
# To do this, we use the <font color="orange">mv</font> command.
# By typing <font color="orange">mv file_to_move destination_directory</font>, you can move a file to the specified directory.
#
# 
# + id="6SxzZa-j7Oz3" colab_type="code" colab={}
# !mv beginner.txt html
# + [markdown] id="wEuqpZDD7Yu2" colab_type="text"
# ###Moving a Directory
# With the <font color="orange">mv</font> command, you can also move <font color="orange">directories</font>, not just files.
# By entering <font color="orange">mv directory_to_move destination_directory</font>, you can move all the files and directories under that directory.
#
# 
# + id="Hulbhs9m7zNm" colab_type="code" colab={}
# # !mv html languages
# + [markdown] id="Q1QFhI-F8UhA" colab_type="text"
# ##Renaming Files and Directories
# The <font color="orange">mv</font> command, which we used to move files and directories earlier, can also be used to rename a file.
#
# You can rename a file or directory by typing <font color="orange">mv old_file_name new_file_name</font> or <font color="orange">mv old_directory_name new_directory_name</font>.
#
# 
# + id="5xRD0eE38rZx" colab_type="code" colab={}
# !touch beginner.txt
# !mv beginner.txt study.txt
# + [markdown] id="iw6fTfoz9GCe" colab_type="text"
# ##Copying Files and DIrectories
# + [markdown] id="gkR6JhCx9IsM" colab_type="text"
# ###Copying a File
# To do this, we use the <font color="orange">cp</font> command.
#
# You can copy a file by entering <font color="orange">cp file_to_copy new_file_name</font>.
#
# 
# + id="4GOQiDw49aLv" colab_type="code" colab={}
# !touch dojo.txt
# !cp dojo.txt project.txt
# + [markdown] id="CDaq_0Gz9f2x" colab_type="text"
# ###Copying a Directory
# With the cp command, you can also copy a directory by adding the <font color="orange">-r</font> (<font color="blue">Recursive copy</font>) option, like <font color="orange">cp -r directory_to_copy new_directory_name</font>.
#
# 
# + id="N6n6udNR92o0" colab_type="code" colab={}
# !mkdir HTML
# !cp -r HTML Ruby
# + [markdown] id="iDzBvzo7-hpX" colab_type="text"
# ##Removing Files and Directories
# + [markdown] id="5sVAnTYF-j3b" colab_type="text"
# ###Removing a File
# To do this, you can use the <font color="orange">rm</font> command, like <font color="orange">rm file_to_remove</font>.
# + id="7N3eU7_8-yyz" colab_type="code" colab={}
# !rm project.txt
# + [markdown] id="X8rgPrO6-7JS" colab_type="text"
# ###Removing a Directory
# You can also remove a directory by adding the <font color="orange">-r</font> option to the <font color="orange">rm</font> command, like <font color="orange">rm -r directory_to_remove</font>.
#
# Just like the <font color="orange">cp</font> command, you will get an error if you forget to add <font color="orange">-r</font>.
# + id="nQbbAD8e_L-K" colab_type="code" colab={}
# !rm -r HTML
# + [markdown] id="A8Zj6tIR_i8R" colab_type="text"
# ##Review
# + [markdown] id="d0BIewex_lUw" colab_type="text"
# ###The Commands History
# On the command line, you can go through the command history using the up arrow key <font color="blue">↑</font>.
# This is very convenient when you want to use a command that was executed before.
# + [markdown] id="l4m2SamWA2kK" colab_type="text"
# #Bonus Content (Ubuntu)
# * apt update
# * This command is used to update the software database.
# + id="EE3P9e5xBfzo" colab_type="code" colab={}
# !sudo apt-get update
# # !apt update
# + [markdown] id="qvkvKepIe789" colab_type="text"
# * apt upgrade
# * This command is used to update (upgrade) the softwares installed on the device
# + id="uBiV1v7rBkdT" colab_type="code" colab={}
# !sudo apt-get upgrade -y
# # !apt upgrade -y
# + [markdown] id="cfGAj39Fe-iu" colab_type="text"
# * apt autoremove
# * This command is used to automatically uninstall unused or residual dependencies.
# + id="9ZOn-6w-Bz8y" colab_type="code" colab={}
# !sudo apt autoremove -y
# # !apt autoremove -y
# + [markdown] id="snhszi94fCqW" colab_type="text"
# * apt autoclean
# * This command is used to get rid of residual setup files.
# + id="qAc0FIL8B_M1" colab_type="code" colab={}
# !sudo apt autoclean
# + [markdown] id="Cdu0EGyTfIQ5" colab_type="text"
# * apt install `package-name`
# * This command is used to install a specific package
# + id="n5OdS2EceMXb" colab_type="code" colab={}
# !apt install neofetch screenfetch
# + id="P-PKryqofaXe" colab_type="code" colab={}
# !screenfetch
# + id="xNHrm191fcc0" colab_type="code" colab={}
# !neofetch
# + [markdown] id="qBbo3SS2Qf_4" colab_type="text"
# ##Missing grub error: Fixed
# + id="NhyFtyzdQu-8" colab_type="code" colab={}
# $ sudo apt-get install mdadm
# $ sudo fdisk -l
# $ sudo blkid
# $ df -Th
# $ sudo mount /dev/sda1 /mnt
# $ for i in /dev /dev/pts /proc /sys /run; do sudo mount -B $i /mnt$i; done
# $ sudo chroot /mnt
# - grub-install /dev/sda
# - update-grub
# //CTRL-D on keyboard.
# $ sudo reboot
# //After reboot
# $ sudo update-grub
# $ grub-probe -t device /boot/grub
# $ sudo grub-install /dev/sda
# //Afterwards, optional
# $ sudo apt-get update
# $ sudo apt-get upgrade
# $ sudo apt autoremove
# $ sudo apt-autoclean
# $ reboot
| UNIX_CL_Progate/UNIX_Command_Line_Progate_(Complete)_Notes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NumPy Arrays
#
# python objects:
#
# ### 1 .high-level number objects : integers floating points
# ### 2 .containers: lists(costless insertion and append),dictionaries (fast lookup)
# # NumPy Provides
# 1.extension package to python for multi-dimensional arrays
# 2.closer to hardware efficiency
# 3.designed for scientific computation
# 4.Also known as array oriented computing
# +
import numpy as np
a = np.array([0,1,2,3])
print(a)
print(np.arange(10))
# -
#python lists
L = range(1000)
# %timeit [i**2 for i in L]
#time it repeates the same thing multiple time and measures the time taken for each iteration
# i microsec is 1 millionth of a sec
a = np.arange(1000)
# %timeit a**2
#very less time for computation
#its nearly 200 time faster than lists
# # 1.Creating arrays
#
# ##### 1.1 Manual Construction of arrays
# +
#1-D
a = np.array([0,1,2,3])
a
# -
#print Dimensions
a.ndim
#shape
a.shape
len(a)
# +
#2-D, 3-D.....
b = np.array([[0,1,2],[3,4,5]])
b
# -
b.ndim
b.shape
len(b) #returns the size of 1st dimension
# +
c = np.array([[[0,1],[2,3]],[[4),5],[6,7]]]
c
# -
c.ndim
c.shape
# +
#1d array is called a vector
#2d array is called a matrix
#nd array is called a tensor
# -
# ###### Functions for creating arrays
# +
#using array function
#array is an array-valued version of the built-in python range function
a = np.arange(10) #0....n-1
a
# +
b = np.arange(1,10,2)#start,end(exclusive),step
b
# +
#using linspace (linear space)
#0---------b/w------------1
#equal spaced intervals
a = np.linspace(0,1,6)
a
# +
#common arrays
a = np.ones((3,3))
a
# +
b = np.zeros((3,3))
b
# +
d = np.eye(3,3) #identity matrix
d
# +
e = np.diag([1,2,3,4]) #diagonal matrix
e
# +
#creating array using random
#create an array of the given shape and populate it with random samples from numpy
a = np.random.rand(4)
a
# +
a = np.random.randn(4) #rand normal
a
# -
# # Basic Datatypes
# +
a = np.arange(10)
a.dtype
# +
a = np.arange(10 , dtype='int64')
a.dtype
a
# -
# ###### the default data type is float for zeros and ones functions
# +
a = np.zeros((3,3))
print(a)
a.dtype
# +
d = np.array([1+2j,2+4j]) #complex datatypes
print(d.dtype)
# +
b = np.array([True,False,True,False]) #Boolean datatype
print(b.dtype)
# +
s = np.array(['ram','Robert','Rahim'])
s.dtype
# +
a = np.arange(10)
print(a[5])
# +
a = np.diag([1,2,3])
print(a[2,2])
# +
a[2,2]=5
a
# -
# # Slicing
# +
a = np.arange(10)
a
# -
a[1:8:2]#[startindex:endindex(exclusive):step]
# +
a = np.arange(10)
a[5:]=10
a
# -
b = np.arange(5)
a[5:]=b[::-1]
#IMP
a
# +
a = np.arange(10)
a
# +
b = a[::2]
b
# -
np.shares_memory(a,b)
b[0]=10
a
# +
a = np.arange(10)
c = a[::2].copy() #this ensures that they don't be sharing same chunk of memory
c
# -
np.shares_memory(a,c)
# +
c[0]=10
a
# -
# # Fancy Indexing
# #### Boolean Mask Indexing
a = np.random.randint(0,20,15)
a
mask = ( a % 2 == 0 )
extract_from_a = a[mask] #masks creates copies not views
extract_from_a
# +
a[mask]=-1
a
# -
# #### Indexing with an array of integers
# +
a = np.arange(0,100,10)
a
# -
a[[2,3,2,4,2]]
# +
a[[9,7]] = -200
a
# -
help("numpy.frombuffer")
help("numpy.empty")
| PreRequsites/numpy/.ipynb_checkpoints/NumpyStack-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Python Club for Physicists
# # Python básico para físicos
# ## Lista de ejercicios
# ### Problema [1](https://codeforces.com/problemset/problem/4/A): **Sandía**
#
#
# Un caluroso día de verano, Pete y su amigo Billy decidieron comprar una sandía. Eligieron el más grande y maduro, en su opinión. Después de eso, se pesó la sandía y la balanza mostró w kilos. Se apresuraron a casa, muriéndose de sed, y decidieron dividir la baya, sin embargo, enfrentaron un problema difícil.
#
# Pete y Billy son grandes fanáticos de los números pares, por eso quieren dividir la sandía de tal manera que cada una de las dos partes pese número par de kilos, al mismo tiempo no es obligatorio que las partes sean iguales. Los chicos están extremadamente cansados y quieren comenzar su comida lo antes posible, por eso debes ayudarlos y averiguar si pueden dividir la sandía de la manera que quieran. Por supuesto, cada uno de ellos debería obtener una parte del peso positivo.
#
# **Entrada**
#
# La primera (y única) línea de entrada contiene el número entero w (1 ≤ w ≤ 100): el peso de la sandía comprada por los niños.
#
# **Salida**
#
# Imprimir SI, si los niños pueden dividir la sandía en dos partes, cada una de ellas con un peso par de kilos; y NO en el caso contrario.
# ### Problema [2](https://codeforces.com/problemset/problem/339/A): **Ayuda Matemática**
# Xenia, la matemática principiante, es una estudiante de tercer año en la escuela primaria. Ahora está aprendiendo la operación de suma.
#
# El maestro ha escrito la suma de varios números. Los alumnos deben calcular la suma. Para facilitar el cálculo, la suma solo contiene los números 1, 2 y 3. Aún así, eso no es suficiente para Xenia. Ella solo está comenzando a contar, por lo que puede calcular una suma solo si los sumandos siguen en orden no decreciente. Por ejemplo, no puede calcular la suma 1 + 3 + 2 + 1, pero puede calcular las sumas 1 + 1 + 2 y 3 + 3.
#
# Tienes la suma que estaba escrita en la pizarra. Reorganice las sumas e imprima la suma de tal manera que Xenia pueda calcular la suma.
#
# **Entrada**
#
# La primera línea contiene una cadena no vacía s: la suma que Xenia necesita contar. La cadena s no contiene espacios. Solo contiene dígitos y caracteres "+". Además, la cadena s es una suma correcta de los números 1, 2 y 3. La cadena s tiene un máximo de 100 caracteres.
#
# **Salida**
#
# Imprime la nueva suma que Xenia pueda contar.
# ### Problema 3: **Mecánica cuántica**
# _Referencia:_ Zetilli, Quantum Mechanics, Problema 3.6 (p. 197)
#
# Los posibles valores de energía de un sistema están dados por la ecuación de Schrödinger
#
# $$
# \hat{H}|\psi\rangle= E|\psi\rangle,
# $$
#
# donde $\hat{H}$ es el hamiltoniano del sistema. Es decir, las posibles energías del sistema vienen a ser los autovalores del Hamiltoniano.
#
# Para este problema, consideremos el hamiltoniano
#
# $$
# \hat{H} = E_0
# \begin{pmatrix}
# 0 & i & 0 \\
# -i & 0 & 0 \\
# 0 & 0 & -1
# \end{pmatrix}
# $$
#
# 1. Definir el hamiltoniano $\hat{H}$.
# 2. Verificar que el hamiltoniano es hermítico
# $$
# \hat{H} = \hat{H}^\dagger
# $$
# 3. Hallar los posibles valores de energía del sistema.
# 4. Hallar los autovectores de $\hat{H}$ y verificar para uno de estos que se cumple la ecuación de Schrödinger.
#Puede hacer E_0 un número real distinto de cero, por ejemplo: E_0 = 1
#Resolver de manera numérica
import numpy as np
# ### Problema 4:
# Buscar datos que sean de su interés$^1$, realizar la limpieza y exploración de datos. Usando Pandas y Matplotlib como librerías principales. Ver: [Taller 3](https://www.youtube.com/watch?v=XtTgRYnEJJE&t=833s&ab_channel=PythonClubforPhysicists)
#
# También puede buscar sus datos en [Kaggle](https://www.kaggle.com/)
#
# *1. puede usar cualquier fuente (siempre y cuando adjunte sus datos)*
# ### Problema 5:
# 5.1 Cree una gráfica de $x_2⋅sin(\frac{1}{x2}) + x$ en el intervalo $[−1,1]$ usando 500 puntos.
# Añadir todos los detalles posibles al gráfico
# 5.2 Cree una matriz aleatoria de 6 × 10 utilizando la función *np.random.rand(3, 5)* y calcule: la suma de todas las entradas, la suma de las filas y la suma de las columnas.
# Enviar sus respuestas a nuestro correo: <EMAIL>
#
# #### *Asunto del correo: "Ejercicios_PCP"*
| Notebooks/Ejercicios/Ejercicios.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: my_env
# language: python
# name: my_env
# ---
# +
import cv2
import numpy as np
import os
import cv2
import numpy as np
import math
import time
from shapely.geometry import Point
from shapely.geometry.polygon import Polygon
import os
from heapq import nsmallest
from skimage.filters import threshold_local
from skimage import measure
import imutils
from sklearn.cluster import KMeans
# +
def delete_bbox_overlap(indexA, indexB,list_bbox,list_bbox_Copy):
'''
indexA: index of bbox A
indexB: index of bbox B
if bbox A and bbox B overlap, detele the smallest
'''
bboxA = list_bbox[indexA]
bboxB = list_bbox[indexB]
polygon_A = Polygon(bboxA)
polygon_B = Polygon(bboxB)
intersec_area = polygon_A.intersection(polygon_B).area
area_polygonA = polygon_A.area
area_polygonB = polygon_B.area
ratio_overlap = intersec_area / min(area_polygonA,area_polygonB)
if ratio_overlap >= 0.3 and area_polygonA < area_polygonB:
if bboxA in list_bbox_Copy:
list_bbox_Copy.remove(bboxA)
elif ratio_overlap >= 0.3 and area_polygonA > area_polygonB:
if bboxB in list_bbox_Copy:
list_bbox_Copy.remove(bboxB)
def remove_bbox_noise(ls_bbox):
'''
Remove the bboxs that is too small or have special shape.
'''
try:
ls_height = []
for bbox in ls_bbox:
ls_height.append(bbox[3][1] - bbox[0][1])
index_bbox = []
mean_height = sum(ls_height)/len(ls_height)
for index in range(len(ls_height)):
if mean_height*1.5 >= ls_height[index] or ls_height[index] >= mean_height*0.65:
index_bbox.append(index)
output_bbox = []
for i in index_bbox:
output_bbox.append(ls_bbox[i])
return output_bbox
except:
return ls_bbox
def bbox_in_boundary_image(box,image):
'''
Checking if bbox has coordinates outside image
'''
count_zero = 0
count_right = 0
count_bottom = 0
count_outside = 0
for point in box:
if 0 in point:
count_zero += 1
if point[1] >= image.shape[0] -2:
count_outside += 1
if point[0] >= image.shape[1] -2:
count_right += 1
if point[0] <0 or point[1] <0:
count_outside += 1
if count_zero >=2 or count_bottom >= 1 or count_right >= 1 or count_outside >=1:
return False
else:
return True
# +
import numpy as np
import cv2
def get_contour_precedence(contour, cols):
tolerance_factor = 10
origin = cv2.boundingRect(contour)
return ((origin[1] // tolerance_factor) * tolerance_factor) * cols + origin[0]
def square(img):
"""
This function resize non square image to square one (height == width)
:param img: input image as numpy array
:return: numpy array
"""
# image after making height equal to width
squared_image = img
# Get image height and width
h = img.shape[0]
w = img.shape[1]
# In case height superior than width
if h > w:
diff = h-w
if diff % 2 == 0:
x1 = np.zeros(shape=(h, diff//2))
x2 = x1
else:
x1 = np.zeros(shape=(h, diff//2))
x2 = np.zeros(shape=(h, (diff//2)+1))
squared_image = np.concatenate((x1, img, x2), axis=1)
# In case height inferior than width
if h < w:
diff = w-h
if diff % 2 == 0:
x1 = np.zeros(shape=(diff//2, w))
x2 = x1
else:
x1 = np.zeros(shape=(diff//2, w))
x2 = np.zeros(shape=((diff//2)+1, w))
squared_image = np.concatenate((x1, img, x2), axis=0)
return squared_image
def sort(vector):
sort = True
while (sort == True):
sort = False
for i in range(len(vector) - 1):
x_1 = vector[i][0]
y_1 = vector[i][1]
for j in range(i + 1, len(vector)):
x_2 = vector[j][0]
y_2 = vector[j][1]
if (x_1 >= x_2 and y_2 >= y_1):
tmp = vector[i]
vector[i] = vector[j]
vector[j] = tmp
sort = True
elif (x_1 < x_2 and y_2 > y_1):
tmp = vector[i]
vector[i] = vector[j]
vector[j] = tmp
sort = True
return vector
def plate_segmentation(img_file_path):
img = cv2.imread(img_file_path)
imgray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
height = img.shape[0]
width = img.shape[1]
area = height * width
scale1 = 0.004
scale2 = 0.1
area_condition1 = area * scale1
area_condition2 = area * scale2
# global thresholding
ret1,th1 = cv2.threshold(imgray,127,255,cv2.THRESH_BINARY)
# Otsu's thresholding
ret2,th2 = cv2.threshold(imgray,0,255,cv2.THRESH_BINARY+cv2.THRESH_OTSU)
# Otsu's thresholding after Gaussian filtering
blur = cv2.GaussianBlur(imgray,(5,5),0)
ret3,th3 = cv2.threshold(blur,0,255,cv2.THRESH_BINARY+cv2.THRESH_OTSU)
contours, hierarchy = cv2.findContours(th3, cv2.RETR_TREE, cv2.CHAIN_APPROX_SIMPLE)
# sort contours
contours = sorted(contours, key=cv2.contourArea, reverse=True)
cropped = []
list_bbox = []
for cnt in contours:
(x,y,w,h) = cv2.boundingRect(cnt)
if (w * h > area_condition1 and w * h < area_condition2 and w/h > 0.3 and h/w > 0.3):
list_bbox.append([(x,y),(x + w, y), (x + w, y + h), (x, y + h)])
print("Shape of bbox raw:",np.shape(list_bbox))
list_bbox_character = list_bbox.copy()
for indexA in range(len(list_bbox) -1 ) :
for indexB in range(indexA + 1, len(list_bbox)):
delete_bbox_overlap(indexA, indexB, list_bbox, list_bbox_character)
print("Shape bbox character", np.shape(list_bbox_character))
list_bbox_character = remove_bbox_noise(list_bbox_character)
print("Shape output bbox character", np.shape(list_bbox_character))
for bbox in list_bbox_character:
if (bbox_in_boundary_image(bbox,image))
cv2.rectangle(img, bbox[0], bbox[2], (255, 0, 0), 2)
cv2.imwrite('detection.png', img)
return cropped
# -
cropped = plate_segmentation('/home/duongnh/Documents/10_image.jpg')
print
print(np.shape(cropped))
| src/.ipynb_checkpoints/Untitled-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.environ['CUDA_VISIBLE_DEVICES'] = ''
import librosa
# import tensorflow as tf
import glob
import numpy as np
# +
# follow hyperparameters from here, https://github.com/pytorch/fairseq/tree/master/examples/wav2vec
features = [(512, 10, 5), (512, 8, 4), (512, 4, 2), (512, 4, 2), (512, 4, 2), (512, 1, 1), (512, 1, 1)]
aggs = [(512, 2, 1), (512, 3, 1), (512, 4, 1), (512, 5, 1), (512, 6, 1), (512, 7, 1), (512, 8, 1), (512, 9, 1),
(512, 10, 1), (512, 11, 1), (512, 12, 1), (512, 13, 1)]
num_negatives = 10
prediction_steps = 12
learning_rate = 1e-5
min_learning_rate = 1e-9
max_learning_rate = 0.005
learning_scheduler = 'cosine'
max_update = 400000
residual_scale = 0.5
log_compression = True
warmup_updates = 50
warmup_init_lr = 1e-07
batch_size = 32
epoch = 10
total_steps = batch_size * epoch
# -
import torch
from torch import nn
import torch.functional as F
np.random.seed(1)
x = np.random.normal(size = (2, 10, 7))
x
x = torch.from_numpy(x)
x.shape
bsz, fsz, tsz = x.shape
bsz, fsz, tsz
y = x.transpose(0, 1)
y = y.contiguous().view(fsz, -1)
y.shape
high = tsz
n_negatives = 10
# neg_idxs = torch.randint(low=0, high=high, size=(bsz, n_negatives * tsz))
# neg_idxs
neg_idxs = torch.from_numpy(np.array([[
1, 2, 3, 1, 4, 0, 5, 6, 1, 2, 0, 4, 2, 1, 0, 5, 4, 5, 4, 6, 6, 4, 1, 6,
6, 3, 4, 4, 5, 0, 1, 5, 4, 4, 1, 1, 0, 2, 0, 6, 2, 6, 3, 4, 5, 6, 2, 4,
0, 2, 1, 2, 6, 4, 2, 4, 0, 2, 4, 2, 1, 0, 4, 6, 6, 4, 4, 2, 3, 4],
[4, 0, 3, 4, 2, 4, 4, 1, 0, 6, 3, 1, 5, 6, 4, 3, 6, 4, 0, 5, 1, 0, 4, 2,
2, 0, 4, 1, 4, 3, 2, 2, 0, 4, 2, 3, 4, 6, 6, 2, 4, 0, 3, 1, 6, 2, 4, 5,
1, 3, 1, 3, 3, 1, 3, 0, 3, 6, 0, 5, 2, 4, 5, 6, 0, 1, 2, 3, 6, 3]]))
# +
for i in range(1, bsz):
neg_idxs[i] += i * high
neg_idxs
# -
negs = y[..., neg_idxs.view(-1)]
negs.shape
negs
negs = negs.view(fsz, bsz, n_negatives, tsz).permute(2, 1, 0, 3)
negs.shape
negs[0]
y = x[:].unsqueeze(0)
print(y.shape, negs.shape)
targets = torch.cat([y, negs], dim=0)
targets.shape
project_to_steps = nn.ConvTranspose2d(10, 10, (1, 12))
s = project_to_steps(x.unsqueeze(-1).float()).unsqueeze(0).expand(targets.size(0), -1, -1, -1, -1)
s.shape
import pickle
with open('convtranspose.pkl', 'wb') as fopen:
pickle.dump(s.detach().numpy().tolist(), fopen)
# +
import math
jin = 0
rin = 0
for _, k, stride in features:
if rin == 0:
rin = k
rin = rin + (k - 1) * jin
if jin == 0:
jin = stride
else:
jin *= stride
offset = math.ceil(rin / jin)
offset = int(offset)
print(offset)
# -
copies, bsz, dim, tsz, steps = s.shape
steps = min(steps, tsz - offset)
predictions = s.new(bsz * copies * (tsz - offset + 1) * steps - ((steps + 1) * steps // 2) * copies * bsz)
labels = torch.zeros_like(predictions)
predictions.shape, labels.shape
s.shape, targets.shape, s[..., :-offset, i].shape, targets[..., offset:].shape
# +
start = end = 0
for i in range(steps):
offset = i + offset
end = start + (tsz - offset) * bsz * copies
pos_num = (end - start) // copies
print(start, pos_num, end, offset, s[..., :-offset, i].shape, targets[..., offset:].shape)
predictions[start:end] = (s[..., :-offset, i].float() * targets[..., offset:].float()).sum(dim=2).flatten()
print(labels[start:start + pos_num])
labels[start:start + pos_num] = 1.
start = end
labels
# -
predictions.detach().numpy()[:-4 * 11]
# +
# torch.nn.functional.binary_cross_entropy(predictions, labels)
# -
torch.nn.functional.sigmoid(predictions)
| speech-to-text/wav2vec-pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Review of Numpy
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Overview:
# - **Teaching:** 5 min
# - **Exercises:** 15 min
#
# **Questions**
# - What is numpy?
# - How do I create and manipulate arrays with numpy?
#
# **Objectives**
# - Know that `numpy` provides array structures to help improve your code.
# - Understand that `numpy` contains a rich range of library to create and manipulate arrays.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Numpy Arrays
#
# * Standard Python Library provides lists and 1d arrays (array.array)
#
# * Lists are general containers for objects
# * Arrays are 1d containers for objects of the same type
# * Limited functionality
# * Some memory and performance overhead associated with these structures
#
# * NumPy provides multidimensional arrays (numpy.ndarray)
# * Can store many elements of the same data type in multiple dimensions
# * cf. Fortran/C/C++ arrays
# * More functionality than core Python e.g. many conveninent methods for array manipulation
# * Efficient storage and execution
#
# See, e.g.,
# * http://docs.scipy.org/doc/numpy-1.10.0/reference/generated/numpy.ndarray.html
# + [markdown] slideshow={"slide_type": "slide"}
# ## Creating Arrays (by hand)
#
# We can create `numpy` arrays *by hand*, taking a list and and creating a numpy array from it:
# + slideshow={"slide_type": "subslide"}
import numpy as np
# + slideshow={"slide_type": "subslide"}
a = np.array( [-1, 0, 1] )
b = np.array( a )
print(a)
print(b)
# + [markdown] slideshow={"slide_type": "subslide"}
# All NumPy arrays are of `type`, `ndarray`
# + slideshow={"slide_type": "subslide"}
type(b)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Creating arrays (helper functions)
#
# However often we want to create large array, that follow a specific sequence or of initial conditions:
# + [markdown] slideshow={"slide_type": "slide"}
# ## Exercise: Create Arrays
#
# What arrays do the following do:
#
# ```python
# w = np.arange( -2, 6, 2 )
# x = np.linspace(-10, 10, 5)
# y = np.zeros(3)
# z = np.ones(3)
# ```
#
# [Solution]()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Solution+:
# + slideshow={"slide_type": "subslide"}
# arange for arrays (similar to range for lists)
w = np.arange( -2, 6, 2 )
print(w)
# + slideshow={"slide_type": "subslide"}
# linspace to create regular sample points in an interval
x = np.linspace(-10, 10, 5)
print(x)
# + slideshow={"slide_type": "subslide"}
# Create an array of zeros
y = np.zeros(3)
print(y)
# + slideshow={"slide_type": "subslide"}
#Create an array of ones
z = np.ones(3)
print(z)
# + [markdown] slideshow={"slide_type": "subslide"}
# :Solution+
# + [markdown] slideshow={"slide_type": "slide"}
# ## Attributes of Arrays
#
# Every `numpy` array has attributes that describe e.g. its *dimension* and *shape*
# + slideshow={"slide_type": "subslide"}
print("Dimensions ", x.ndim)
# + slideshow={"slide_type": "subslide"}
print("Shape ", x.shape)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Exercise:
#
# What are the following attributes:
#
# ```python
# x.size
# x.dtype
# ```
#
# What other attributes can you find for a numpy array (and how)?
#
# [Solution]()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Solution+:
# + slideshow={"slide_type": "subslide"}
print("Size ", a.size)
# + slideshow={"slide_type": "subslide"}
print("Data type ", a.dtype)
# + [markdown] slideshow={"slide_type": "subslide"}
# :Solution
# + [markdown] slideshow={"slide_type": "slide"}
# ## Multi-dimensional arrays
#
# Many different ways to create N-dimensional arrays. A two-dimensional array or matrix can be created from, e.g., list of lists
# + slideshow={"slide_type": "subslide"}
mat = np.array( [[1,2,3], [4,5,6]] )
print(mat)
print("Dimensions: ", mat.ndim)
print("Size: ", mat.size)
print("Shape: ", mat.shape)
# + slideshow={"slide_type": "subslide"}
mat0 = np.zeros( (3,3) )
print(mat0)
# + slideshow={"slide_type": "subslide"}
mat1 = np.ones( (3,3) )
print(mat0)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Key Points:
# - Numpy allows you to quickly create (multidimensional) arrays and popultate them
# - Numpy array attributes describe the array
| nbplain/01_numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Statistics
#
# The [astropy.stats](http://docs.astropy.org/en/stable/stats/index.html) sub-package includes common statistical tools for astronomy. It is by no means complete and we are open to contributions for more functionality! In this tutorial, we will take a look at some of the existing functionality.
#
# <section class="objectives panel panel-warning">
# <div class="panel-heading">
# <h2><span class="fa fa-certificate"></span> Objectives</h2>
# </div>
#
#
# <div class="panel-body">
#
# <ul>
# <li>Use sigma clipping on arrays</li>
# <li>Use automatic bin determination for histograms</li>
# <li>Work with binomial and Poisson distributions</li>
# </ul>
#
# </div>
#
# </section>
#
# ## Documentation
#
# This notebook only shows a subset of the functionality in astropy.stats. For more information about the features presented below as well as other available features, you can read the
# [astropy.stats documentation](https://docs.astropy.org/en/stable/stats/).
# %matplotlib inline
import matplotlib.pyplot as plt
plt.rc('image', origin='lower')
plt.rc('figure', figsize=(10, 6))
# ## Sigma clipping
#
# A very common operation with astronomical data is sigma-clipping, where values in a dataset are iteratively excluded if they deviate too much from some overall statistic. To illustrate this, we use an image of a star field for which we want to estimate the background:
# The result is a masked Numpy array:
# We can then estimate the background and background noise easily for example:
# The ``sigma_clip`` function includes a number of options to refine the algorithm, including what statistics to use during the sigma-clipping, and also allows for asymmetric sigma values. See the [sigma_clip() documentation](https://docs.astropy.org/en/stable/api/astropy.stats.sigma_clip.html) for more details.
#
# As a shortcut for deriving sigma-clipped statistics as done above, a ``sigma_clipped_stats`` function:
# ## Choosing histogram bins
#
# Another example of functionality in the astropy.stats sub-package is the [histogram()](http://docs.astropy.org/en/stable/api/astropy.stats.histogram.html#astropy.stats.histogram) function, which is exposed through the [hist()](http://docs.astropy.org/en/stable/api/astropy.visualization.hist.html#astropy.visualization.hist) function in astropy.visualization for ease of use. These functions provide an easy way to use smart binning algorithms for data.
#
# Let's start off by creating some synthetic data:
# We can use Matplotlib to make a histogram, but the number of bins has to be set (or defaults to an arbitrary values) and is not necessarily optimal:
# We can instead try and use the the [hist()](http://docs.astropy.org/en/stable/api/astropy.visualization.hist.html#astropy.visualization.hist) function in astropy.visualization to try out different methods of automatically selecting the bins:
# for example using [Bayesian Blocks](https://arxiv.org/abs/1207.5578):
# or [Knuth's rule](https://arxiv.org/abs/physics/0605197):
# If you want to get the underlying values, use the [histogram()](http://docs.astropy.org/en/stable/api/astropy.stats.histogram.html#astropy.stats.histogram) function in astropy.stats:
# ## Binomial and Poisson distributions
#
# A third set of functionality in astropy.stats are functions to help with calculations related to binomial and Poisson distributions. For example, if we observe 150 galaxies and find that 50 are spirals, we can estimate confidence intervals using [binom_conf_interval()](http://docs.astropy.org/en/stable/api/astropy.stats.binom_conf_interval.html#astropy.stats.binom_conf_interval) for the true underlying fraction of spiral galaxies that would have been observed if the sample had been much larger (and with the same selection criteria/biases):
# By default, the confidence interval returned is the 68.27% confidence interval ('1-sigma'):
# Similarly, if we detect 10 photons in a pixel, we can determine the confidence interval for the underlying brightness of the emission in the pixel using [poisson_conf_interval()](http://docs.astropy.org/en/stable/api/astropy.stats.poisson_conf_interval.html#astropy.stats.poisson_conf_interval):
#
# <section class="challenge panel panel-success">
# <div class="panel-heading">
# <h2><span class="fa fa-pencil"></span> Challenge</h2>
# </div>
#
#
# <div class="panel-body">
#
# <ol>
# <li>Take a look at the <a href="http://docs.astropy.org/en/stable/stats/index.html">astropy.stats documentation</a>, and in particular the long list of functions at the bottom, in case you see something that could be useful to you! (and feel free to try them if so)</li>
# <li>If you had to find the median absolute deviation for a dataset, how would you do it? Try and find the robust standard deviation using the median absolute deviation for the sigma clipped array we produced before.</li>
# </ol>
#
# </div>
#
# </section>
#
# <center><i>This notebook was written by <a href="https://aperiosoftware.com/">Aperio Software Ltd.</a> © 2019, and is licensed under a <a href="https://creativecommons.org/licenses/by/4.0/">Creative Commons Attribution 4.0 International License (CC BY 4.0)</a></i></center>
#
# 
| 12-statistics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Process eigenvalues and eigenvectors
# The title of the notebook should be coherent with file name. Namely, file name should be:
# *author's initials_progressive number_title.ipynb*
# For example:
# *EF_01_Data Exploration.ipynb*
#
# ## Purpose
# State the purpose of the notebook.
#
# ## Methodology
# Quickly describe assumptions and processing steps.
#
# ## WIP - improvements
# Use this section only if the notebook is not final.
#
# Notable TODOs:
# - todo 1;
# - todo 2;
# - todo 3.
#
# ## Results
# Describe and comment the most important results.
#
# ## Suggested next steps
# State suggested next steps, based on results obtained in this notebook.
# # Setup
#
# ## Library import
# We import all the required Python libraries
# +
# Data manipulation
import pandas as pd
import numpy as np
import os
import glob
# Options for pandas
pd.options.display.max_columns = 50
pd.options.display.max_rows = 30
# Visualizations
import plotly
import plotly.graph_objs as go
import plotly.offline as ply
plotly.offline.init_notebook_mode(connected=True)
import matplotlib.pyplot as plt
import seaborn as sns
# Autoreload extension
if 'autoreload' not in get_ipython().extension_manager.loaded:
# %load_ext autoreload
# %autoreload 2
# -
# ### Change directory
# If Jupyter lab sets the root directory in `notebooks`, change directory.
if "notebook" in os.getcwd():
os.chdir("..")
# ## Local library import
# We import all the required local libraries libraries
# +
# Include local library paths
import sys
sys.path.append("./src") # uncomment and fill to import local libraries
# Import local libraries
import src.utilities as utils
# -
# # Parameter definition
# We set all relevant parameters for our notebook. By convention, parameters are uppercase, while all the
# other variables follow Python's guidelines.
# +
EIGENVECTOR_FILEPATH = "data/processed/eigenvector_024"
config = utils.read_config()
plt.style.use(config['viz']['jupyter'])
# -
#
# # Data import
# We retrieve all the required data for the analysis.
eigenvector = np.loadtxt(EIGENVECTOR_FILEPATH)
# eigenvector_filepaths = sorted(glob.glob('data/processed/eigenvector_???'))
eigenvector_filepath = "data/processed/eigenvector_{:03}"
eigenvectors = {mode_num : np.loadtxt(eigenvector_filepath.format(mode_num)) for mode_num in range(7,25)}
# # Data processing
# Put here the core of the notebook. Feel free to further split this section into subsections.
# ### Renormalize each row/column vector of mode eigenvector
# **Note** The motion's amplitude inforamtion is lost.
# +
# Renormalize eigenvectors
# for key in eigenvectors.keys():
# eigenvector = eigenvectors[key]
# for row_idx in range(eigenvector.shape[0]):
# eigenvector[row_idx] = eigenvector[row_idx] / np.linalg.norm(eigenvector[row_idx])
# +
def calcualte_dotprod(eigenvector):
no_beads = np.shape(eigenvector)[0]
dotprod = np.zeros((no_beads, no_beads))
for i in np.arange(no_beads):
for j in np.arange(i+1):
# vector_1 = eigenvector[i] / np.linalg.norm(eigenvector[i])
# vector_2 = eigenvector[j] / np.linalg.norm(eigenvector[j])
vector_1 = eigenvector[i]
vector_2 = eigenvector[j]
dotprod[i][j] = np.dot(vector_1, vector_2)
dotprod = dotprod + dotprod.T
return dotprod
def calcualte_crossprod_norm(eigenvector):
no_beads = np.shape(eigenvector)[0]
crossprod_norm = np.zeros((no_beads, no_beads))
for i in np.arange(no_beads):
for j in np.arange(i+1):
# vector_1 = eigenvector[i] / np.linalg.norm(eigenvector[i])
# vector_2 = eigenvector[j] / np.linalg.norm(eigenvector[j])
vector_1 = eigenvector[i]
vector_2 = eigenvector[j]
crossprod_norm[i][j] = np.linalg.norm(np.cross(eigenvector[i], eigenvector[j]))
crossprod_norm = crossprod_norm + crossprod_norm.T
return crossprod_norm
# -
# # Data visualization
# Visualize processed data
# +
def plot_crosscor(array, vmin=0, center=None, vmax=1, axis=None, cmap=plt.cm.RdBu_r, annot=True):
# array = array[1:, :-1]
# mask = np.zeros_like(array)
# mask[np.triu_indices_from(mask, k=1)] = True
sns.heatmap(array, fmt='.2f', linewidths=.5, vmin=vmin, center=center, vmax=vmax, square=True, mask=None, annot=annot, cmap=cmap, cbar=False, ax=axis)
# y_ticks = axis.get_yticks()
# y_ticks = [int(tick +0.5) for tick in y_ticks]
# axis.set_yticklabels(y_ticks)
return None
subplot_rows = 3
subplot_columns = 6
with plt.rc_context({'font.size': 4, 'xtick.labelsize': 5, 'ytick.labelsize': 5, 'axes.titlesize': 7}):
_, axs = plt.subplots(subplot_rows, subplot_columns)
mode_num = 7
for i in range(subplot_rows):
for j in range(subplot_columns):
dotprod = calcualte_dotprod(eigenvectors[mode_num])
plot_crosscor(dotprod, vmin=-1, center=0, vmax=1, cmap=plt.cm.RdBu_r, axis=axs[i, j], annot=False)
axs[i, j].set_title("mode {}".format(mode_num))
mode_num += 1
plt.tight_layout()
# + jupyter={"source_hidden": true}
with plt.rc_context({'font.size': 4, 'xtick.labelsize': 5, 'ytick.labelsize': 5, 'axes.titlesize': 7}):
_, axs = plt.subplots(subplot_rows, subplot_columns)
mode_num = 7
for i in range(subplot_rows):
for j in range(subplot_columns):
corossprod_norm = calcualte_crossprod_norm(eigenvectors[mode_num])
plot_crosscor(corossprod_norm, vmin=0, center=None, vmax=1, cmap=plt.cm.binary, axis=axs[i, j], annot=False)
axs[i, j].set_title("mode {}".format(mode_num))
mode_num += 1
plt.tight_layout()
# -
np.linalg.norm(eigenvectors[24])
# # References
# We report here relevant references:
# 1. author1, article1, journal1, year1, url1
# 2. author2, article2, journal2, year2, url2
| notebooks/ID.01.explore_eigenvectors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Build Models To Compare Features: Cleaned Original Features
# ### Read In Data
# +
# Read in data
import joblib
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import GridSearchCV
# %matplotlib inline
train_features = pd.read_csv('../Data/Final_Data/train_features_original.csv')
train_labels = pd.read_csv('../Data/Final_Data/train_labels.csv')
train_features.head()
# -
# Generate correlation matrix heat map
matrix = np.triu(train_features.corr())
sns.heatmap(train_features.corr(), annot=True, fmt='.1f', vmin=-1, vmax=1, center=0, cmap='coolwarm', mask=matrix)
# ### GridSearchCV
def print_results(results):
print('BEST PARAMS: {}\n'.format(results.best_params_))
mean = results.cv_results_['mean_test_score']
stds = results.cv_results_['std_test_score']
for mean, std, params in zip(mean, stds, results.cv_results_['params']):
print('{} (+/- {}) for {}'.format(mean, std, params))
# +
# Conduct search for best params while running cross-validation (GridSearchCV)
rf = RandomForestClassifier()
parameters = {
'n_estimators': [2**i for i in range(3, 10)],
'max_depth': [2, 4, 8, 16, 32, None],
}
cv = GridSearchCV(rf, parameters, cv=5)
cv.fit(train_features, train_labels.values.ravel())
print_results(cv)
# -
# ### Feature Importance
# Generate feature importance plot
feat_imp = cv.best_estimator_.feature_importances_
indices = np.argsort(feat_imp)
plt.yticks(range(len(indices)), [train_features.columns[i] for i in indices])
plt.barh(range(len(indices)), feat_imp[indices], color='r', align='center')
plt.show()
# ### Write Out Pickled Model
# Write out the model fit on the whole training set
joblib.dump(cv.best_estimator_, '../Pickled_Models/mdl_cleaned_original_features.pkl')
| ML - Applied Machine Learning - Feature Engineering/06.Compare and Evaluate Models/02.Build Model - Cleaned Original Features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# imports
from pathlib import Path
import pandas as pd
from nowcasting_utils.metrics.evaluation import evaluation
# Output csv
ESO_PV_FORECASTS_OUTPUT_FILE = Path("/mnt/storage_b/data/ocf/solar_pv_nowcasting/other_organisations_pv_forecasts/National_Grid_ESO/CSV/testset_v15.csv")
# +
# run evaluation for ESO forecast
model_name = f'ESO forecast'
results_df = pd.read_csv(ESO_PV_FORECASTS_OUTPUT_FILE)
print(results_df)
evaluation(results_df,model_name,show_fig=False)
# -
| notebooks/ESO/ESO evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# This is an implementation of NIPS 2017 paper, titled "*Thy Friend is My Friend: Iterative Collaborative Filtering for Sparse Matrix Estimation*" ( [link](https://papers.nips.cc/paper/7057-thy-friend-is-my-friend-iterative-collaborative-filtering-for-sparse-matrix-estimation.pdf) and [arxiv](https://arxiv.org/pdf/1712.00710v1.pdf) ) for Global NIPS Paper implementation [Challenge](https://www.nurture.ai/nips-challenge). The authors have also created a short [video](https://www.youtube.com/watch?v=qxfDK44YuQE) for it.<br>
# For any questions related to this notebook, feel free to contact me at <EMAIL> .
# ## How to use this Notebook:
# Following are some important points/guidelines/assumptions to keep in mind while navigating through this notebook:
# - This notebook consists of sections numbered by Roman numerals
# - Brief description about all sections:
# - I: Model preparation: Changes to be made to rating matrix(dataset) before we can apply the algorithms
# - II: Algorithm Details: All the algorithms described in the paper are implemented here.
# - III: Other important functions:
# - Data Handling: for data manipulation
# - Substitutes: methods which can be used in place of (algorithm) methods in paper
# - Evaluation: for evaulation of recommender system
# - IV: Test script/Experiments: testing using a dataset
# - If a function/variable has suffix as "_csr", it refers to CSR (Compressed Sparse Row) data. Else, if it has a suffix as "_matrix", it refers to 2D (numpy) matrix data.
# - The dataset ratings are assumed to be integers; to be modified in future
# - data_csr ensures that user_id and item_id start from 0 by taking in FIRST_INDEX as a global variable
# - All the datasets are symmetricized and normalized before we begin applying the algorithm
# - There are some dataset parameters like FIRST_INDEX, USERS, etc which will be automatically detected from dataset. Hence, they have been set as -1 for default condition. To overload automatic detection, provide a value (but we recommend you not to overload).
# - Brief description about hyperparameters:
# - RADIUS : this controls the size of neighborhood. It is the distance from vertex u to vertices i (at neighborhood boundary). Setting large/small values for RADIUS might reduce the average number of neighbors per vertex, resulting suboptimal overlap of vertices. To set optimal RADIUS, you can use output from 'describe_neighbor_count' function to understand how varying RADIUS affects neighborhood size.
# - THRESHOLD : this controls the final set of vertices which are considered for individual rating estimation. Setting this too high will make smaller the set of vertices being considered and vice versa. To set optimal threshold, you can use output from 'describe_distance_matrix' function to understand how THRESHOLD affects the size of this set.
# - UNPRED_RATING : average rating for user-item pair, for which the algorithm could not make an estimate for the rating.
# - TRAIN_TEST_SPLIT : %age of test dataset.
# - C1 : %age of edges in train dataset going to $E1$ for expanding the neighborhood (step 2).
# - C2 : %age of edges in train dataset going to $E2$ for distance computation between the vertices (step 3). Also, please note that 1 - C1 - C2 is %age of edges in train dataset going to $E3$ for rating estimation (step 4).
# Importing required modules required for complete notebook
# Built and tested on python2
import numpy as np
from tqdm import *
import sys
from datetime import datetime
datetime.now().time() # (hour, min, sec, microsec)
# +
#TODO: see that there are no duplicate functions
# -
# # 0: Model Notations
# Following are the symbols/notations used in the paper. The variables/notations used in this notebook have been discussed in Experiments section.
#
# $u$ = user <br>
# $i$ = item <br>
# $M$ = symmetric rating matrix of size $n \times n$ (usually the dataset) <br>
# $E$ = set of $(u,i)$ where each user $u$ has rated an item $i$ also seen in the matrix $M$ (intuitively $E$ is edge set(matrix) between user and items. <br>
# $p$ = sparsity of $M$ i.e. (= #observed ratings in $M$ / total # ratings in $M$)<br>
# $r$ = radius, distance (in no of edges) between user $u$ and item $i$ at neighborhood boundary (look in step 2) <br>
# ### Setting constants
'''Dataset Parameters'''
################################################################################################################
DATA_PATH = './ml-100k/u.data' # ml-100k data set has 100k ratings, 943 users and 1682 items
DELIMITER = "\t" # tab separated or comma separated data format
N_RATINGS = 100000
################################################################################################################
# These parameters will be detected automatically from dataset
# -1 is for the default value
FIRST_INDEX = -1
USERS = -1
ITEMS = -1
SPARSITY = -1 # 'p' in the equations
UNOBSERVED = 0 # default value in matrix for unobserved ratings; prefer to keep it 0
# To reduce size of csr for testing purpose
# WARNING: ONLY TO BE USED FOR TESTING
# (for real run, put SIZE_REDUCTION = False)
SIZE_REDUCTION = True
USER_LIMIT = 200
ITEM_LIMIT = 500
# +
'''Hyperparameters'''
# All the hyperparameters have default values
#To use them, set the parameters as -1
################################################################################################################
TRAIN_TEST_SPLIT = -1 # %age of test ratings wrt train rating ; value in between 0 and 1
C1 = -1 # probability of edges in training set going to E1
C2 = -1 # probability of edges in training set going to E2
RADIUS = 3 # radius of neighborhood, radius = # edges between start and end vertex
UNPRED_RATING = 3 # rating (normalized) for which we dont have predicted rating between 1 - 5
THRESHOLD = 0.01 # distance similarity threshold used for rating prediction
################################################################################################################
# checks on hyper parameters
if isinstance(C1, float) and isinstance(C2, float) and (C1 > 0) and (C2 > 0) and 1 - C1 - C2 > 0:
print('c1 = {}'.format(C1))
print('c2 = {}'.format(C2))
print('c3 = {}'.format(1-C1-C2))
elif (C1 == -1) and (C2 == -1):
C1 = C2 = 0.33
print('c1 = {} (default)'.format(C1))
print('c2 = {} (default)'.format(C2))
print('c3 = {} (default)'.format(1-C1-C2))
else:
print('ERROR: Incorrect values set for C1 and C2')
if isinstance(RADIUS, int) and RADIUS > 0:
print('Radius = {}'.format(RADIUS))
elif RADIUS == -1:
print('Radius = default value as per paper')
else:
print('ERROR: Incorrect values set for Radius')
if UNPRED_RATING >= 1 and UNPRED_RATING <= 5:
print('Rating set for unpredicted ratings = {}'. format(UNPRED_RATING))
elif UNPRED_RATING == -1:
UNPRED_RATING = 3
print('Rating set for unpredicted ratings = {} (default)'. format(UNPRED_RATING))
else:
print('ERROR: Incorrect values set for UNPRED_RATING')
if TRAIN_TEST_SPLIT > 0 and TRAIN_TEST_SPLIT < 1:
print('TRAIN_TEST_SPLIT = {}'.format(TRAIN_TEST_SPLIT))
elif TRAIN_TEST_SPLIT == -1:
TRAIN_TEST_SPLIT = 0.2
print('TRAIN_TEST_SPLIT = 0.2 (default)')
else:
print('ERROR: Incorrect values set for TRAIN_TEST_SPLIT')
# -
# ### Data Handling
# +
''' Function to read data file, given in CSR format
Assuming 1st 3 values of a row as: user_id, item_id, rating '''
def read_data_csr(fname, delimiter, dtype=float):
data_csr = np.loadtxt(fname=fname, delimiter=delimiter, dtype=dtype) # Reading data to array
data_csr = data_csr[:, :3] # Extracting 1st 3 columns: 0,1,2
if FIRST_INDEX == -1: # Making sure user_id/item_id starts from 0
first_index_user = min(data_csr[:,0]) # as it becomes easier to track in graphs
first_index_item = min(data_csr[:,1])
data_csr[:,0] = data_csr[:,0] - first_index_user
data_csr[:,1] = data_csr[:,1] - first_index_item
else:
data_csr[:,0:2] = data_csr[:,0:2] - FIRST_INDEX
return data_csr
''' Function to get data in CSR format for given data in matrix format '''
def matrix_to_csr(data_matrix):
data_csr = np.array([ [i,j,data_matrix[i,j]]\
for i in range(len(data_matrix))\
for j in range(len(data_matrix[i]))\
if data_matrix[i,j] != UNOBSERVED])
return data_csr
'''Function to find and replace some values
for only 1d and 2d numpy arrays'''
def find_and_replace(data, find_value, replace_value):
if len(data.shape) == 1: # for 1D numpy array
for i in range(len(data)):
if data[i] == find_value:
data[i] = replace_value
elif len(data.shape) == 2: # for 2D numpy array
for i in range(len(data)):
for j in range(len(data[i])):
if data[i,j] == find_value:
data[i,j] = replace_value
return data
''' Function to check dataset'''
def check_and_set_data_csr(data_csr):
global USERS, ITEMS, SPARSITY
n_users = int(max(data_csr[:,0])) + 1
n_items = int(max(data_csr[:,1])) + 1
unique_users = len(np.array(list(set(data_csr[:,0]))))
unique_items = len(np.array(list(set(data_csr[:,1]))))
if USERS == -1:
USERS = n_users
if ITEMS == -1:
ITEMS = n_items
print('USERS = ' + str(USERS))
print('ITEMS = ' + str(ITEMS))
#checking if global USERS/ITEMS had wrong values entered:
if n_users != USERS:
print('ERROR: USERS entered by you is wrong. {} users found in dataset'.format(n_users))
if n_items != ITEMS:
print('ERROR: ITEMS entered by you is wrong. {} items found in dataset'.format(n_items))
# checking unrated users/items : this is possible if some user/item index gets skipped in dataset
if n_users != unique_users:
print('ERROR: No. of users with no ratings: ' + str(n_users - unique_users))
print(' : This notebook may not be robust to such dataset')
if n_items != unique_items:
print('ERROR: No. of items with no ratings: ' + str(n_items - unique_items))
print(' : This notebook may not be robust to such dataset')
if n_users == unique_users and n_items == unique_items:
print('All users and items have at least one rating! Good!')
#checking sparsity for large symmetricized matrix
sparsity_symm = float(2 * N_RATINGS) / ((USERS + ITEMS)**2)
if SPARSITY == -1:
SPARSITY = sparsity_symm
print('SPARSITY (p) = ' + str(SPARSITY))
if SPARSITY != sparsity_symm:
print('ERROR: SPARSITY entered by you is wrong. {} sparsity found in dataset'.format(sparsity_symm))
if sparsity_symm <= (float(1) / ((n_users + n_items)**2)):
print('WARNING: For generated large symmetric matrix:')
print(' : p is not polynomially larger than 1/n.')
print(' : Using dist1 as distance computation may not gurantee that')
print(' : expected square error converges to zero using this paper\'s algorithm.')
else:
print('Sym matrix : p is polynomially larger than 1/n, all guarantees applicable')
print('Check and set dataset : done')
'''Function to generate training and testing data split from given CSR data'''
def generate_train_test_split_csr(data_csr, split, shuffle=True):
# we use data_csr as it is easy to only shuffle it and accordingly create train and test set
if shuffle:
np.random.shuffle(data_csr) # inplace shuffle
train_sz = int((1 - split) * data_csr.shape[0])
train_data_csr = data_csr[: train_sz ,:]
test_data_csr = data_csr[train_sz : ,:]
if train_data_csr.shape[0]+test_data_csr.shape[0] == data_csr.shape[0]:
print('Generating train test split: done')
else:
print('Generating train test split: FAILED')
return [train_data_csr, test_data_csr]
'''Function to force reduce the size of dataset
To be used only for testing purposes
Note: this doesnt ensure if every item has a rating or not: TODO'''
def reduce_size_of_data_csr(data_csr):
global N_RATINGS
print('WARNING: FOR TESTING PURPOSES ONLY')
if USER_LIMIT < 1 or ITEM_LIMIT < 1:
print('ERROR: please set limits > 0')
print(' : using same sataset without any reductions')
return data_csr
data_csr = data_csr[((data_csr[:,0] < USER_LIMIT)*(data_csr[:,1] < ITEM_LIMIT))]
# Accounting for unvisited users
visited = np.full((USER_LIMIT), True, dtype=bool)
for i in data_csr:
visited[int(i[0])] = False
unvisited_users = [i for i in range(len(visited)) if visited[i]]
# adding 1 rating for every unvisited user
for i in unvisited_users:
data_csr = np.append(data_csr, [[i, i, 3]], axis=0)
N_RATINGS = data_csr.shape[0]
return data_csr
# -
# Experiment:
# +
data_csr = read_data_csr(fname=DATA_PATH, delimiter=DELIMITER)
if SIZE_REDUCTION:
data_csr = reduce_size_of_data_csr(data_csr)
if data_csr.shape[0] == N_RATINGS: # gives total no of ratings read; useful for verification
print('Reading dataset: done')
else:
print('Reading dataset: FAILED')
print( '# of missing ratings: ' + str(N_RATINGS - data_csr.shape[0]))
check_and_set_data_csr(data_csr=data_csr)
# -
[train_data_csr, test_data_csr] = generate_train_test_split_csr(data_csr=data_csr, split=TRAIN_TEST_SPLIT)
# # I: Model preparation
# We first look at function which converts our asymmetric rating matrix to a symmetric matrix and another function that normalizes the ratings between [0,1].
# This function is used to normalize the ratings:
'''Function to normalize all ratings of a CSR (compressed sparse row) matrix'''
def normalize_ratings_csr(data_csr):
#TODO: assuming non negative ratings, make it generic
data_csr[:,2] = data_csr[:,2] / float(max(data_csr[:,2]))
print('Normalize ratings: done')
return data_csr
# This function is used to make a rating CSR matrix symmetric:
# +
''' Function to get data in matrix format for given data in CSR format '''
def csr_to_matrix(data_csr, symmetry=False):
data_matrix = np.full(((USERS+ITEMS), (USERS+ITEMS)), UNOBSERVED, dtype=float)
for line in data_csr:
data_matrix[int(line[0])][int(line[1])] = line[2]
if symmetry:
data_matrix[int(line[1])][int(line[0])] = line[2]
return data_matrix
'''Function get matrix from csr such that no two item_ids and user_ids are same'''
def get_csr_with_offset(data_csr, offset):
new_data_csr = np.copy(data_csr)
new_data_csr[:,1] = new_data_csr[:,1] + offset # so that user_ids != item_ids
#new_data_matrix = csr_to_matrix(new_data_csr, symmetry=True)
return new_data_csr
'''MAIN Function to convert asymmetrix CSR matrix to symmetrix matrix
the returned CSR doesnt contain repitions for any user-item pair.
Repetitions can be generated for a 2D matrix by calling csr_to_matrix(data_csr, symmetry=True)'''
def csr_to_symmetric_csr(data_csr):
# Assuming the rating matrix to be non symmetric
# Even if it is symmetric, the user_id and item_id need to be different for graph
data_csr = get_csr_with_offset(data_csr, offset=USERS)
print('CSR to symmetric CSR matrix: done')
return data_csr
# -
# Experiment:
train_data_csr = normalize_ratings_csr(train_data_csr)
train_data_csr = csr_to_symmetric_csr(train_data_csr)
# the symmetric matrix obtained doesnt contain repirions for any user item pair
# only the item_ids are scaled by item_ids += USERS
# hence, we can safely go ahead and use this CSR matrix for sample splitting step
# # II: Algorithm Details
# As per paper: *We present and discuss details of each step of the algorithm, which primarily involves computing pairwise distances (or similarities) between vertices.*
# ### Step 1: Sample Splitting
# Partition the rating matrix into three different parts. Following are the exerpts from paper:
# - *Each edge in $E$ is independently placed into $E_1, E_2,$ or $E_3$, with probabilities $c_1, c_2,$ and $1 - c_1 - c_2$ respectively. Matrices $M_1, M_2$, and $M_3$ contain information from the subset of the data in $M$ associated to $E_1, E_2$, and $E_3$ respectively.*
# - *$M_1$ is used to define local neighborhoods of each vertex (in step 2), $M_2$ is used to compute similarities of these neighborhoods (in step 3), and $M_3$ is used to average over datapoints for the final estimate (in step 4)*
def sample_splitting_csr(data_csr, c1=0.33, c2=0.33, shuffle=True):
if shuffle:
np.random.shuffle(data_csr) # inplace shuffle
m1_sz = int(c1 * data_csr.shape[0])
m2_sz = int(c2 * data_csr.shape[0])
m1_csr = data_csr[ : m1_sz ,:]
m2_csr = data_csr[ m1_sz : m1_sz + m2_sz ,:]
m3_csr = data_csr[m1_sz + m2_sz : ,:]
if m1_csr.shape[0]+m2_csr.shape[0]+m3_csr.shape[0] == data_csr.shape[0]:
print('Sample splitting: done')
else:
print('Sample splitting: FAILED')
return [m1_csr, m2_csr, m3_csr]
# Experiment:
[m1_csr, m2_csr, m3_csr] = sample_splitting_csr(data_csr=train_data_csr, c1=C1, c2=C2)
# ### Step 2: Expanding the Neighborhood
# We do the following in this step:
# - radius $r$ to be tuned using cross validation. We can use its default value as $r = \frac{6\ln(1/p)}{8\ln(c_1pn)}$ as per paper.
# - use matrix $M_1$ to build neighborhood based on radius $r$
# - Build BFS tree rooted at each vertex to get product of the path from user to item, such that
# - each vertex (user or item) in a path from user to boundary item is unique
# - the path chosen is the shortest path (#path edges) between the user and the boundary item
# - in case of multiple paths (or trees), choose any one path (i.e. any one tree) at random
# - Normalize the product of ratings by total no of final items at the boundary
#
# $N_{u,r}$ obtained is a vector for user $u$ for $r$-hop, where each element is product of path from user to item or zero. $\tilde{N_{u,r}}$ is normalized vector.
#
# Testing (next two cells)
# +
# import numpy as np ##### REMOVE THIS CELL
# from tqdm import *
# import sys
# FIRST_INDEX = -1
# USERS = -1
# ITEMS = -1
# SPARSITY = -1 # 'p' in the equations
# UNOBSERVED = 0 # default value in matrix for unobserved ratings
# N_RATINGS = 7
# C1 = 0 # only to account for scale_factor in step 3
# C2 = 1 # only to account for scale_factor in step 3
# RADIUS = 3 # radius of neighborhood, radius = # edges between start and end vertex
# UNPRED_RATING = -1 # rating (normalized) for which we dont have predicted rating
# m1_csr = read_data_csr(fname='./very_small_graph.txt', delimiter="\t")
# check_and_set_data_csr(data_csr=m1_csr)
# +
# m1_csr = normalize_ratings_csr(m1_csr) ##### REMOVE THIS CELL
# m1_csr = csr_to_symmetric_csr(m1_csr)
# -
'''Function to create a graph as adjacency list: a dictionary of sets'''
def create_dict_graph_from_csr(data_csr):
data_matrix = csr_to_matrix(data_csr, symmetry=True)
# Create an (unweighted) adjacency list for the graph
## we still have the 2D matrix for the weights
graph = dict()
print('Creating graph as dictionary:')
sys.stdout.flush()
for i in tqdm(range(len(data_matrix))):
temp_set = set()
for j in range(len(data_matrix[i])):
if data_matrix[i,j] > 0:
temp_set.add(j)
graph[i] = temp_set
return graph
# Functions useful for getting products along paths:
# +
import random
'''Function gives all possible path from 'start' vertex at r-hop distance '''
# help from:
# http://eddmann.com/posts/depth-first-search-and-breadth-first-search-in-python/
# radius = # edges between start and end vertex
def bfs_paths(graph, start, radius):
queue = [(start, [start])]
visited = [start]
while queue:
(vertex, path) = queue.pop(0)
for next in graph[vertex] - set(path):
if next in visited:
continue
depth = len(path + [next]) - 1
if depth == radius:
# We do not append next to visited because
# we want all shorted paths to next and then
# choose one path at random in get_product()
yield path + [next]
else:
queue.append((next, path + [next]))
visited.append(next)
'''Function which returns a dictionary for a given user
where each item represents the key in the dictionary
and it returns a list of lists(paths) from user to item r-hop distance apart'''
def create_item_dict(all_path):
dict_path = dict()
for path in all_path:
r_hop_item = path[-1]
dict_path.setdefault(r_hop_item,[]).append(path)
return dict_path
'''Function to get product from user to item in the path
It chooses any path at random, if #paths > 1'''
def get_product(data_matrix, path):
if len(path) < 1:
return UNOBSERVED
idx = random.randint(0, len(path)-1) # in case of multiple paths to same item
p = path[idx] # choose any one path at random
product = 1
for i in range(len(p)-1):
product = product * data_matrix[p[i],p[i+1]]
return product
# +
'''Function to generate product matrix from user to items
(items which are at r-hop boundary from user)'''
def generate_product_matrix(graph, data_matrix, radius):
# each u'th row in product_matrix represents a neighbor boundary vector for vertex u
# therefore it is a (USERS+ITEMS) x (USERS+ITEMS) dimensional matrix
product_matrix = np.full(((USERS+ITEMS), (USERS+ITEMS)), UNOBSERVED, dtype=float)
for user_vertex in tqdm(range(USERS+ITEMS)): #a user_vertex may also be of an item
all_path = list(bfs_paths(graph, user_vertex, radius)) # 1. get a list of all r-hop paths from given user
dict_path = create_item_dict(all_path) # 2. create dict of paths from user to individual items
for item_vertex in dict_path: #an item_vertex may also be of a user
paths = dict_path[item_vertex] # 3. get the set of user-item paths
product = get_product(data_matrix, paths) # 4. get product for a unique user-item path (at random)
product_matrix[user_vertex, item_vertex] = product # 5. store the product in the matrix
return product_matrix
'''Function to normalize the product of paths in neighbor boundary vector for every u'th rowed user
normalized along the same row'''
#TODO: implement it in efficient manner
def row_wise_normalize_matrix(data_matrix):
n_neighbors_per_row = np.full((USERS+ITEMS), 0, dtype=float)
for i in range(len(data_matrix)):
for j in range(len(data_matrix[i])):
if data_matrix[i,j] != UNOBSERVED:
n_neighbors_per_row[i] = n_neighbors_per_row[i] + 1
for i in range(len(data_matrix)):
for j in range(len(data_matrix[i])):
if data_matrix[i,j] != UNOBSERVED and n_neighbors_per_row[i] > 0:
data_matrix[i,j] = data_matrix[i,j] / n_neighbors_per_row[i]
return data_matrix
import pandas as pd
'''Function to describe count of neighbors for every vertex and other values
Also note, the values described might be slightly distorted because of symmetricity of neighbor matrices'''
def describe_neighbor_count(data_matrix):
n_neighbor_vector = np.full((USERS+ITEMS), 0, dtype=float)
for i in range(len(data_matrix)):
for j in range(len(data_matrix[i])):
if data_matrix[i,j] != UNOBSERVED:
n_neighbor_vector[i] = n_neighbor_vector[i] + 1
df = pd.DataFrame(n_neighbor_vector)
print('To effectively choose RADIUS value for next run of algorithm:')
print('Showing distribution of count of neighbors for every vertex:')
print(df.describe())
# -
import math
'''Function to return two product matrices
one at r-hop distance and other at r+1 hop distance for dist1 computation'''
# if radius passed is less than 1 or not passed, this function evaluates the default radius as per paper
def generate_neighbor_boundary_matrix(data_csr):
global RADIUS
if RADIUS < 1:
#TODO: Fix this
print('ERROR: please do not use the radius formula as given in paper')
print(' : the formula evaluates to a decimal values between 0 and 1')
print(' : I am working on fixing this')
RADIUS = (float(6) * math.log( 1.0 / SPARSITY)) / (8.0 * math.log( float(C1) * SPARSITY * (USERS + ITEMS)))
return -1
# First create the graph
graph = create_dict_graph_from_csr(data_csr) # to store the edges in adjacency list
data_matrix = csr_to_matrix(data_csr, symmetry=True) # to store the ratings as matrix
radius = RADIUS
print('Generating neighbor boundary matrix at {}-hop distance:'.format(radius))
sys.stdout.flush()
r_neighbor_matrix = generate_product_matrix(graph, data_matrix, radius=radius)
r_neighbor_matrix = row_wise_normalize_matrix(r_neighbor_matrix)
radius = radius+1
print('Generating neighbor boundary matrix at {}-hop distance:'.format(radius))
sys.stdout.flush()
r1_neighbor_matrix = generate_product_matrix(graph, data_matrix, radius=radius)
r1_neighbor_matrix = row_wise_normalize_matrix(r1_neighbor_matrix)
return [r_neighbor_matrix, r1_neighbor_matrix]
# Experiment / Testing
[r_neighbor_matrix, r1_neighbor_matrix] = generate_neighbor_boundary_matrix(m1_csr)
# all neighbor boundary vector for each user u is stored as u'th row in neighbor_matrix
# though here the vector is stored a row vector, we will treat it as column vector in Step 4
# Note: we might expect neighbor matrix to be symmetric with dimensions (USERS+ITEMS)*(USERS+ITEMS)
# : since distance user-item and item-user should be same
# : but this is not the case since there might be multiple paths between user-item
# : and the random path picked for user-item and item-user may not be same
# : normalizing the matrix also will result to rise of difference
describe_neighbor_count(r_neighbor_matrix)
describe_neighbor_count(r1_neighbor_matrix)
# ### Step 3: Computing the distances
# Distance computation between two users (using matrix $M_2$) using the following formula (only $dist_1$ implemented for now):
#
# $$ dist(u,v) = \left(\frac{1 - c_1p}{c_2p}\right) (\tilde{N_{u,r}} - \tilde{N_{v,r}})^T M_2 (\tilde{N_{u,r+1}} - \tilde{N_{v,r+1}}) $$
# +
def compute_distance_matrix(r_neighbor_matrix, r1_neighbor_matrix, m2_csr):
m2_matrix = csr_to_matrix(m2_csr, symmetry=True)
scale_factor = (1.0 - C1 * SPARSITY) / (C2 * SPARSITY)
user_list = np.array(range(USERS+ITEMS))
distance_matrix = np.full(((USERS+ITEMS), (USERS+ITEMS)), UNOBSERVED, dtype=float)
print('Generating distance matrix')
sys.stdout.flush()
for user1 in tqdm(user_list): # computing for all elements individually
for user2 in user_list: # not assuming any symmetricity for distance matrix
user1_r_neighbor_vector = r_neighbor_matrix[user1]
user2_r_neighbor_vector = r_neighbor_matrix[user2]
r_neighbor_vector = user1_r_neighbor_vector - user2_r_neighbor_vector
r_neighbor_vector = np.transpose(r_neighbor_vector)
user1_r1_neighbor_vector = r1_neighbor_matrix[user1]
user2_r1_neighbor_vector = r1_neighbor_matrix[user2]
r1_neighbor_vector = user1_r1_neighbor_vector - user2_r1_neighbor_vector
#print(r_neighbor_vector.shape)
#print(m2_matrix.shape)
dist_value = np.matmul(r_neighbor_vector, m2_matrix)
dist_value = np.matmul(dist_value, r1_neighbor_vector)
distance_matrix[user1,user2] = dist_value * scale_factor
return distance_matrix
import pandas as pd
'''Function which gives an idea of how distance values are distributed to best choose THRESHOLD in Step 4'''
def describe_distance_matrix(distance_matrix):
flat_distance_matrix = distance_matrix.flatten()
observed = []
for i in flat_distance_matrix:
if i != UNOBSERVED:
observed.append(i)
df = pd.DataFrame(observed)
print('To effectively choose THRESHOLD value in next step:')
print('Showing distribution of non zero (or observed) entries of distance matrix:')
print(df.describe())
# -
# Experiment:
distance_matrix = compute_distance_matrix(r_neighbor_matrix, r1_neighbor_matrix, m2_csr)
describe_distance_matrix(distance_matrix)
# Testing
# +
# distance_matrix = compute_distance_matrix(r_neighbor_matrix, r1_neighbor_matrix, m1_csr)
# distance_matrix
# +
# describe_distance_matrix(distance_matrix)
# -
# ### Step 4: Averaging datapoints to produce final estimate
# Average over nearby data points based on the distance(similarity) threshold $n_n$ (using matrix $M_3$). $n_n$ to be tuned using cross validation. Mathematically (from paper):
#
# $$ \hat{F_{u,v}} = \frac{1}{\mid E_{uv1} \mid} \sum_{(a,b) \in E_{uv1}} M_3(a,b) $$
# *where $E_{uv1}$ denotes the set of undirected edges $(a, b)$ such that $(a, b) \in E_3$ and both $dist(u, a)$ and $dist(v, b)$ are less than $n_n$*
'''Function to get similarity matrix using THRESHOLD'''
def generate_sim_matrix(distance_matrix, threshold):
user_list = np.array(range(USERS+ITEMS))
sim_matrix = np.full(((USERS+ITEMS), (USERS+ITEMS)), False, dtype=bool)
print('Generating distance similarity matrix:')
sys.stdout.flush()
for user1 in tqdm(user_list): # computing for all elements individually
for user2 in user_list: # not assuming any symmetricity for distance matrix
if distance_matrix[user1,user2] != UNOBSERVED and distance_matrix[user1,user2] < threshold:
sim_matrix[user1, user2] = True
return sim_matrix
# +
'''Function to get final prediction estimates for user-item ratings'''
def generate_averaged_prediction(u, v, sim_matrix, m3_matrix, bounded=True):
prediction = 0
n_prediction = 0
# Making sure the vertex indices are ints
u = int(u)
v = int(v)
for a in range(len(sim_matrix[u])):
for b in range(len(sim_matrix[v])):
if sim_matrix[u,a] and sim_matrix[v,b] and m3_matrix[a,b] != UNOBSERVED:
prediction = prediction + m3_matrix[a,b]
n_prediction = n_prediction + 1
if n_prediction > 0: #TODO: write why they are present
prediction = prediction / n_prediction
else:
prediction = UNPRED_RATING / 5 # TODO: make it generic; here we assume ratings as 1 - 5
if bounded: # : make it generic; here we assume ratings as 1 - 5
if prediction > 1:
prediction = 1
elif prediction < 0.2:
prediction = 0.2
return prediction
'''Function to get final prediction estimates for user-item rating matrix
Use this only if you want estimates for all the ratings'''
def generate_averaged_prediction_matrix(sim_matrix, m3_csr):
m3_matrix = csr_to_matrix(m3_csr, symmetry=True)
vertex_list = np.array(range(USERS+ITEMS))
prediction_matrix = np.full(((USERS+ITEMS), (USERS+ITEMS)), UNOBSERVED, dtype=float)
print('Generating prediction matrix:')
sys.stdout.flush()
for u in tqdm(vertex_list):
for v in vertex_list:
prediction = generate_averaged_prediction(u, v, sim_matrix, m3_matrix, bounded=True)
prediction_matrix[u, v] = prediction
return prediction_matrix
'''Function to get final prediction estimates for given user-item list
Use this only if you have a set of user-item pairs for which you want the final estimates
Consider using this function for evaluation purposes(only)'''
def generate_averaged_prediction_array(sim_matrix, m3_csr, test_data_csr):
m3_matrix = csr_to_matrix(m3_csr, symmetry=True)
prediction_array = np.full((len(test_data_csr)), UNOBSERVED, dtype=float)
print('Generating prediction array:')
sys.stdout.flush()
for i in tqdm(range(len(test_data_csr))):
datapt = test_data_csr[i]
# Considering only first two columns for rating estimation
vertex1 = int(datapt[0])
vertex2 = int(datapt[1])
prediction = generate_averaged_prediction(vertex1, vertex2, sim_matrix, m3_matrix, bounded=True)
prediction_array[i] = prediction
return prediction_array
# -
# Experiment
THRESHOLD = 2 # prefer to choose a threshold now if not chosen in initial Hyperparameter decision stage
sim_matrix = generate_sim_matrix(distance_matrix, threshold=THRESHOLD)
# Prepare the test dataset using Model preparation section functions
test_data_csr = normalize_ratings_csr(test_data_csr)
test_data_csr = csr_to_symmetric_csr(test_data_csr)
# +
# Getting estimates for only test data points
prediction_array = generate_averaged_prediction_array(sim_matrix, m3_csr, test_data_csr)
# To generate complete rating matrix do the following:
#prediction_matrix = generate_averaged_prediction_matrix(sim_matrix, m3_csr)
# -
# Testing
# +
# sim_matrix = generate_sim_matrix(distance_matrix, threshold=.26)
# sim_matrix
# +
# prediction_array = generate_averaged_prediction_array(sim_matrix, m1_csr, m1_csr)
# prediction_array
# +
# prediction_matrix = generate_averaged_prediction_matrix(sim_matrix, m1_csr)
# prediction_matrix
# -
# ### Evaluation
# We evaluate our recommendation algorithm using RMSE (root mean square error). <br>
# According to paper, if sparsity $p$ is polynomially larger than $n^{-1}$, i.e. if $p = n^{-1 + \epsilon}$ for $\epsilon > 0$, then we can safely use $dist_1$ distance computation formula and MSE is bounded by $O((pn)^{-1/5})$.
# +
from sklearn.metrics import mean_squared_error
from math import sqrt
#TODO: describe these functions
'''Function to generate true and test labels from test_data_csr and predicted_matrix
This function may not be required for evaluation purposes(only)'''
def generate_true_and_test_labels(test_data_csr, predicted_matrix):
# for all the available ratings in testset
# and for all the predicted rating for those available rating
# put them in two separate vectors
y_actual = np.full((len(test_data_csr)), UNOBSERVED, dtype=float)
y_predict = np.full((len(test_data_csr)), UNOBSERVED, dtype=float)
print('Generating true and test label:')
sys.stdout.flush()
for i in tqdm(range(len(test_data_csr))):
testpt = test_data_csr[i]
y_actual[i] = testpt[2]
y_predict[i] = predicted_matrix[testpt[0], testpt[1]]
if y_predict[i] == UNOBSERVED: # i.e. we could not generate a rating for this test user item pair
y_predict[i] = AVG_RATING
return [y_actual, y_predict]
'''Function to get Mean Squared Error for given actual and predicted array'''
def get_mse(y_actual, y_predict):
mse = mean_squared_error(y_actual, y_predict)
return mse
'''Function to get ROOT Mean Squared Error for given actual and predicted array'''
def get_rmse(y_actual, y_predict):
rmse = sqrt(mean_squared_error(y_actual, y_predict))
return rmse
'''Function to get Average Error for given actual and predicted array'''
def get_avg_err(y_actual, y_predict):
avg_err = sum(abs(y_actual - y_predict)) / len(y_actual)
return avg_err
'''Function to check if obtained MSE is within the bound as calculated in the paper'''
def check_mse(data_csr, y_actual, y_predict):
mse_upper_bound = (SPARSITY * (USERS+ITEMS)) ** (-1 / float(5))
print('MSE Upper bound: {}'.format(mse_upper_bound))
mse = get_mse(y_actual, y_predict)
print('MSE of predictions: {}'.format(mse))
if mse < mse_upper_bound:
print('As per the discussion in the paper, MSE is bounded by O((pn)**(-1/5))')
else:
print('ERROR: Contrary to the discusssion in the paper, MSE is NOT bounded by O((pn)**(-1/5))')
# -
# Experiment:
# We have already prepared the test data (required for our algorithm)
y_actual = test_data_csr[:,2]
y_predict = prediction_array
# If we want, we could scale our ratings back to 1 - 5 range for evaluation purposes
#But then paper makes no guarantees about scaled ratings
#y_actual = y_actual * 5
#y_predict = y_predict * 5
get_rmse(y_actual, y_predict)
get_avg_err(y_actual, y_predict)
check_mse(data_csr, y_actual, y_predict)
# Scaling and doing the same checks
#But then paper makes no guarantees about scaled ratings
y_actual = y_actual * 5
y_predict = y_predict * 5
get_rmse(y_actual, y_predict)
get_avg_err(y_actual, y_predict)
check_mse(data_csr, y_actual, y_predict)
# Testing
# +
# # We have already prepared the test data (required for our algorithm)
# test_data_csr = m1_csr
# y_actual = test_data_csr[:,2]
# y_predict = prediction_array
# # If we want, we could scale our ratings back to 1 - 5 range for evaluation purposes
# #But then paper makes no guarantees about scaled ratings
# #y_actual = y_actual * 5
# #y_predict = y_actual * 5
# +
# get_rmse(y_actual, y_predict)
# +
# get_avg_err(y_actual, y_predict)
# +
# check_mse(m1_csr, y_actual, y_predict)
# +
# # Scaling and doing the same checks
# #But then paper makes no guarantees about scaled ratings
# y_actual = y_actual * 5
# y_predict = y_predict * 5
# +
# get_rmse(y_actual, y_predict)
# +
# get_avg_err(y_actual, y_predict)
# +
# check_mse(m1_csr, y_actual, y_predict)
# -
datetime.now().time() # (hour, min, sec, microsec)
| play/revisedcode-roughWork.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Dropout with L2 Weight Regularization
import numpy as np
import keras
from keras.datasets import mnist
from keras.models import Sequential
from matplotlib import pyplot as plt
from keras.layers import Dense,Flatten
from keras.layers import Conv2D, MaxPooling2D,BatchNormalization,Dropout
from keras.utils import np_utils
from sklearn.metrics import confusion_matrix, f1_score, precision_score, recall_score, classification_report
from keras import optimizers,regularizers
class AccuracyHistory(keras.callbacks.Callback):
def on_train_begin(self, logs={}):
self.acc = []
self.loss = []
self.val_f1s = []
self.val_recalls = []
self.val_precisions = []
def on_epoch_end(self, batch, logs={}):
self.acc.append(logs.get('acc'))
self.loss.append(logs.get('loss'))
X_val, y_val = self.validation_data[0], self.validation_data[1]
y_predict = np.asarray(model.predict(X_val))
y_val = np.argmax(y_val, axis=1)
y_predict = np.argmax(y_predict, axis=1)
self.val_recalls.append(recall_score(y_val, y_predict, average=None))
self.val_precisions.append(precision_score(y_val, y_predict, average=None))
self.val_f1s.append(f1_score(y_val,y_predict, average=None))
# +
(X_train, y_train), (X_test, y_test) = mnist.load_data()
# print(X_train.shape)
# reshape to be [samples][pixels][width][height]
X_train = X_train.reshape(X_train.shape[0],28, 28,1).astype('float32')
X_test = X_test.reshape(X_test.shape[0],28, 28,1).astype('float32')
# normalize inputs from 0-255 to 0-1
X_train = X_train / 255
X_test = X_test / 255
# # one hot encode outputs
y_train = np_utils.to_categorical(y_train)
y_test = np_utils.to_categorical(y_test)
print(y_train.shape)
num_classes = y_test.shape[1]
print(num_classes)
input_shape=(28,28,1)
epochs=10
batch_size = 512
history = AccuracyHistory()
# -
def create_deep_model(opt,loss):
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),activation='relu',input_shape=input_shape))
model.add(MaxPooling2D((2, 2)))
model.add(Dropout(0.20))
model.add(Conv2D(64, (3, 3), activation='relu',padding='same'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(128, (3, 3), activation='relu',padding='same'))
model.add(Dropout(0.25))
model.add(BatchNormalization())
model.add(Flatten())
model.add(Dense(1024, activation='relu'))
model.add(Dropout(0.25))
model.add(Dense(num_classes, activation='softmax',kernel_regularizer=regularizers.l2(0.01)))
model.compile(optimizer=opt,loss=loss,metrics=['accuracy'])
return model
# +
def create_optimizer(opt_name,lr,decay):
if opt_name == "SGD":
opt = optimizers.SGD(lr=lr, decay=decay)
elif opt_name == "Adam":
opt = optimizers.Adam(lr=lr, decay=decay)
elif opt_name == "RMSprop":
opt = optimizers.RMSprop(lr=lr, decay=decay)
elif opt_name == "Adagrad":
opt = optimizers.Adagrad(lr=lr, decay=decay)
return opt
def create_model(filters,filt1_size,conv_stride,pool_size,pool_stride,opt,loss):
model=Sequential()
model.add(Conv2D(filters, kernel_size=(filt1_size, filt1_size), strides=(conv_stride, conv_stride),activation='relu',input_shape=input_shape))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(pool_size, pool_size), strides=(pool_stride,pool_stride), padding='valid'))
model.add(Flatten())
model.add(Dense(1024,activation='relu'))
model.add(Dense(num_classes, activation='softmax'))
model.compile(optimizer=opt,loss=loss,metrics=['accuracy'])
return model
# -
def fit_model(epochs,batch_size):
model.fit(X_train, y_train,batch_size=batch_size,epochs=epochs,validation_split=0.05,callbacks=[history])
score = model.evaluate(X_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
y_pred = model.predict_classes(X_test)
cnf_mat = confusion_matrix(np.argmax(y_test,axis=1), y_pred)
return cnf_mat,score,y_pred
# +
lr = 0.001
decay = 1e-6
#decay = 0.0
epochs=10
batch_size = 1024
opt = create_optimizer('Adam',lr,decay)
loss = "categorical_crossentropy"
filters,filt1_size,conv_stride,pool_size,pool_stride = 32,7,1,2,2
model = create_deep_model(opt,loss)
print(model.summary())
cnf_mat,score,y_pred = fit_model(epochs,batch_size)
# -
from keras.models import load_model
model.save('Dropout_model_MNIST.h5')
fscore=f1_score(np.argmax(y_test,axis=1), y_pred,average=None)
recall=recall_score(np.argmax(y_test,axis=1), y_pred,average=None)
prec=precision_score(np.argmax(y_test,axis=1), y_pred,average=None)
def plot(r1,r2,data,Info):
plt.plot(range(r1,r2),data)
plt.xlabel('Epochs')
plt.ylabel(Info)
plt.show()
plot(1,epochs+1,history.acc,'Accuracy')
plot(1,epochs+1,history.loss,'Loss')
plt.plot(recall,label='Recall')
plt.plot(prec,label='Precision')
plt.xlabel('Class')
plt.ylabel('F-score vs Recall vs Precision')
plt.plot(fscore,label='F-score')
plt.legend()
avg_fscore=np.mean(fscore)
print(avg_fscore)
avg_precision=np.mean(prec)
print(avg_precision)
avg_recall=np.mean(recall)
print(avg_recall)
# +
cnf_mat = confusion_matrix(np.argmax(y_test,axis=1), y_pred)
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
conf = cnf_mat
fig, ax = plt.subplots(figsize=(30,30))
im = ax.imshow(conf,alpha=0.5)
# plt.show()
# We want to show all ticks...
ax.set_xticks(np.arange(cnf_mat.shape[0]))
ax.set_yticks(np.arange(cnf_mat.shape[1]))
# ... and label them with the respective list entries
ax.set_xticklabels(np.arange(0,96))
ax.set_yticklabels(np.arange(0,96))
# Rotate the tick labels and set their alignment.
plt.setp(ax.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
# Loop over data dimensions and create text annotations.
for i in range(cnf_mat.shape[0]):
for j in range(cnf_mat.shape[1]):
text = ax.text(j, i, conf[i, j],
ha="center", va="center",color="black",fontsize=10)
ax.set_title("Confusion matrix",fontsize=20)
fig.tight_layout()
# fig.savefig('plot1_cnf.png')
plt.show()
# -
del model
| Assignments/Assignment_2/Q1/q1_Arch4_MNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.linear_model import LinearRegression
df = pd.read_csv('https://archive.ics.uci.edu/ml/machine-learning-databases/housing/housing.data', header=None, sep='\s+')
df.columns = ['CRIM', 'ZN', 'INDUS', 'CHAS', 'NOX', 'RM', 'AGE', 'DIS', 'RAD', 'TAX', 'PTRATIO', 'B', 'LSTAT', 'MEDV']
X = df[['RM']].values
y = df['MEDV'].values
# +
from sklearn.linear_model import RANSACRegressor
ransac = RANSACRegressor(LinearRegression(),
max_trials=100,
min_samples=50,
residual_metric=lambda x: np.sum(np.abs(x), axis=1),
residual_threshold=5.0,
random_state=0)
ransac.fit(X, y)
# -
# obtaining & plotting the inliers and outliers from the fitted RANSAC linear regression model
inlier_mask = ransac.inlier_mask_
outlier_mask = np.logical_not(inlier_mask)
line_X = np.arange(3, 10, 1)
line_y_ransac = ransac.predict(line_X[:, np.newaxis])
plt.scatter(X[inlier_mask], y[inlier_mask], c='blue', marker='o', label='Inliers')
plt.scatter(X[outlier_mask], y[outlier_mask], c='lightgreen', marker='s', label='Outliers')
plt.plot(line_X, line_y_ransac, color='red')
plt.xlabel('Average number of rooms [RM]')
plt.ylabel('Price in $1000\'s [MEDV]')
plt.legend(loc='upper left')
plt.show()
print('Slope: %.3f' % ransac.estimator_.coef_[0])
print('Intercept: %.3f' % ransac.estimator_.intercept_)
| ch10/03-ransac-regressor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/brs1977/PreSumm/blob/master/PreSumm.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="ynBAxrbYV-4M" colab_type="code" colab={}
Extraction-based summarization Суммирование на основе извлечения
Извлечение ключевых фраз из исходного документа и их объединение в резюме. Производится в соответствии с заданной метрикой без каких-либо изменений в текстах.
Abstraction-based summarization Обобщение на основе абстракций
перефразирование и сокращение частей исходного документа, обобщения текста, преодолевает грамматические несоответствия экстрактивного метода. Обобщения абстрактного текста создают новые фразы и предложения, передающие наиболее полезную информацию из исходного текста.Абстракция работает лучше, чем извлечение. Однако алгоритмы суммирования текста, необходимые для абстракции, сложнее разработать;
ROUGE-1 относится к перекрытию униграмм (каждого слова) между системой и справочными данными.
ROUGE-2 относится к перекрытию биграмм между системой и справочными сводками.
ROUGE-L: статистика на основе самой длинной общей подпоследовательности (LCS)
# + id="q9PlTzCbD3-v" colab_type="code" outputId="ed57ef9e-37f6-4da4-a448-d16a7ba0c4da" colab={"base_uri": "https://localhost:8080/", "height": 125}
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="DNRDPvEUv7BJ" colab_type="text"
# #ssh
# + id="hWKzTrccV-qU" colab_type="code" colab={}
#create ssh connect
import random, string
password = ''.join(random.choice(string.ascii_letters + string.digits) for i in range(20))
#Download ngrok
# ! wget -q -c -nc https://bin.equinox.io/c/4VmDzA7iaHb/ngrok-stable-linux-amd64.zip
# ! unzip -qq -n ngrok-stable-linux-amd64.zip
#Setup sshd
# ! apt-get install -qq -o=Dpkg::Use-Pty=0 openssh-server pwgen > /dev/null
#Set root password
# ! echo root:$password | chpasswd
# ! mkdir -p /var/run/sshd
# ! echo "PermitRootLogin yes" >> /etc/ssh/sshd_config
# ! echo "PasswordAuthentication yes" >> /etc/ssh/sshd_config
# ! echo "LD_LIBRARY_PATH=/usr/lib64-nvidia" >> /root/.bashrc
# ! echo "export LD_LIBRARY_PATH" >> /root/.bashrc
#Run sshd
get_ipython().system_raw('/usr/sbin/sshd -D &')
#Ask token
print("Copy authtoken from https://dashboard.ngrok.com/auth")
import getpass
authtoken = getpass.getpass()
#Create tunnel
get_ipython().system_raw('./ngrok authtoken $authtoken && ./ngrok tcp 22 &')
#Print root password
print("Root password: {}".format(password))
#Get public address
# ! curl -s http://localhost:4040/api/tunnels | python3 -c \
# "import sys, json; print(json.load(sys.stdin)['tunnels'][0]['public_url'])"
# + [markdown] id="Xn9hDePhEwR3" colab_type="text"
# #Requirements
# + id="LkO9j8kyGetm" colab_type="code" colab={}
import requests
import os
import random
import tarfile
import pandas as pd
import zipfile
import multiprocess
import json
from glob import glob
# + id="7_6pCT0eGep7" colab_type="code" outputId="54467199-a483-4129-b91d-4855d3e7e5d1" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# !git clone https://github.com/brs1977/PreSumm.git
# !pip install pytorch_transformers
# # !pip install torch
# !pip install pytorch-pretrained-bert
# !pip install tensorboardX
# # !pip install pyrouge
#pyrouge
# !sudo apt-get install libxml-parser-perl
# !git clone https://github.com/andersjo/pyrouge.git rouge
# %cd /content/rouge/tools/ROUGE-1.5.5/data
# !rm WordNet-2.0.exc.db
# !./WordNet-2.0-Exceptions/buildExeptionDB.pl ./WordNet-2.0-Exceptions ./smart_common_words.txt ./WordNet-2.0.exc.db
# %cd /content
# !git clone https://github.com/bheinzerling/pyrouge
# %cd /content/pyrouge
# !python setup.py install
# !pyrouge_set_rouge_path '/content/rouge/tools/ROUGE-1.5.5'
# !python -m pyrouge.test
# + [markdown] id="nI7YkWfmVwWD" colab_type="text"
# ##Download Stanford CoreNLP
# + id="YYl2a9A4VtdX" colab_type="code" outputId="144b1410-4d41-46fa-cd4b-6c47223be370" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# %cd /content
#########
# !wget http://nlp.stanford.edu/software/stanford-corenlp-full-2017-06-09.zip
# !unzip stanford-corenlp-full-2017-06-09.zip
# %env CLASSPATH=/content/stanford-corenlp-full-2017-06-09/stanford-corenlp-3.8.0.jar
# + id="iE0gpLPxUSeo" colab_type="code" outputId="372c84bd-1ed5-48fd-bff9-a29d6c8f10ef" colab={"base_uri": "https://localhost:8080/", "height": 34}
# # %env CLASSPATH=/content/stanford-corenlp-full-2017-06-09/stanford-corenlp-3.8.0.jar
# !echo $CLASSPATH
# + [markdown] id="IW_tg39EC49r" colab_type="text"
# ##Kaggle env
# + id="4cvNySGFC3uN" colab_type="code" outputId="637c2179-20ab-4d63-96a1-848302533adc" colab={"base_uri": "https://localhost:8080/", "height": 34}
# #!pip install -q kaggle
#kaggle key
# !mkdir ~/.kaggle
# !cp /content/drive/My\ Drive/kaggle.json ~/.kaggle
# !ls ~/.kaggle
# + [markdown] id="rOC0uVbnDGut" colab_type="text"
# ##Load kaggle data
# + id="szCgrKSjC_jp" colab_type="code" outputId="a52767e8-c78b-4252-9731-997e1468510f" colab={"base_uri": "https://localhost:8080/", "height": 263}
# %cd /content
# !kaggle competitions download -c title-generation
# + id="DKraTEFTDEVa" colab_type="code" outputId="8645b61b-c4e0-4115-eff0-27a1d7cb39d7" colab={"base_uri": "https://localhost:8080/", "height": 140}
# !mkdir data
# !unzip sample_submission.csv.zip -d data
# !unzip vocs.pkl.zip -d data
# !unzip train.csv.zip -d data
# !mv test.csv data
# !ls data
# + [markdown] id="RKh57Y6t9EFr" colab_type="text"
# #Data prepare
# + id="HvlPMF1dCrbZ" colab_type="code" colab={}
import pandas as pd
train = pd.read_csv('/content/data/train.csv', encoding='utf8')
test = pd.read_csv('/content/data/test.csv', encoding='utf8')
# + id="5iUGIG9VaqFp" colab_type="code" colab={}
test.head
# + [markdown] id="V6yHNvJmGWqh" colab_type="text"
# ###Splitting and Tokenization
# + id="IHFb5GKl9Gai" colab_type="code" outputId="a1e8e28e-8536-4ca7-ce19-174e610651e5" colab={"base_uri": "https://localhost:8080/", "height": 70}
import numpy as np
import os
#It is processed this way because this repo was based on the ArXiv dataset, which
# has the data in [abstract] @highlight [title] format
arxiv_dir = '/content/PreSumm/arxiv_raw'
tokenized_dir = '/content/merged_arxiv_tokenized'
# %cd /content
# !rm $arxiv_dir -r
# !mkdir $arxiv_dir
# !rm $tokenized_dir -r
# !mkdir $tokenized_dir
#split val test
rand_split = [.1]
for row in train.iterrows():
i, (abstract,title) = row
sampletext = abstract + '\n\n@highlight\n\n' + title
r = np.random.rand()
# split val train
fn = 'valid' if r < rand_split[0] else 'train'
fn = f'{fn}.{i}.story'
fn = os.path.join(arxiv_dir, fn)
with open(fn, "w") as text_file:
text_file.write(sampletext)
for row in test.iterrows():
i, abstract = row
sampletext = abstract[0] + '\n'
fn = f'test.{i}.story'
fn = os.path.join(arxiv_dir, fn)
with open(fn, "w") as text_file:
text_file.write(sampletext)
# + id="7w5iwKwKHNPw" colab_type="code" colab={}
# %%capture
# !mkdir /content/logs
# %cd /content/PreSumm/src
# !python preprocess.py -mode tokenize -raw_path $arxiv_dir -save_path $tokenized_dir
# + [markdown] id="hP9DwK3KJreK" colab_type="text"
# ###Format to Simpler Json Files
# + id="26agvb9xI4GE" colab_type="code" colab={}
# %%capture
# !python preprocess.py -mode format_to_lines -raw_path $tokenized_dir -save_path ../json_data/arxiv -shard_size 20000 -n_cpus 1
# + [markdown] id="KmSgFOSoNH4V" colab_type="text"
# ###Format to PyTorch Files
# + id="m1dAcJzNI4DL" colab_type="code" outputId="3740cc11-6550-4fe6-d7c1-eefd44d2e8a3" colab={"base_uri": "https://localhost:8080/", "height": 811}
# #%%capture
# %cd /content/PreSumm/src
# !python preprocess.py -mode format_to_bert -raw_path ../json_data/ -save_path ../bert_data/ -lower -n_cpus 1 -log_file ../logs/preprocess.log
# + id="Pco6FYotjnAj" colab_type="code" colab={}
#Save pretraining data
# %cd ../bert_data
# !zip pre_sum_data.zip *.pt
# #!rm /content/drive/My\ Drive/nlp/bert_sum_data.zip
# !cp pre_sum_data.zip /content/drive/My\ Drive/nlp
# + id="nzrCb928m95a" colab_type="code" outputId="bb1ff50e-9a97-47c9-f17f-ea121ce2249a" colab={"base_uri": "https://localhost:8080/", "height": 34}
# import torch
# nd = torch.load('../bert_data/arxiv.test.0.bert.pt')
# len(nd)
# l = glob.glob(tokenized_dir+'/*.json')
# len(l)
# import json
# jd = json.load(open('/content/BertSum/json_data/arxiv.new.0.json'))
# len(jd)
# + [markdown] id="iGY6fio59B-o" colab_type="text"
# #Train
# + id="wigtFy6FSM3R" colab_type="code" outputId="be933881-f4f0-4c5e-a9e0-119d07943e47" colab={"base_uri": "https://localhost:8080/", "height": 246}
# !git pull origin master
# + id="s8NcuHIeDovq" colab_type="code" outputId="ec38b761-1bdd-479a-e06a-1aca05262729" colab={"base_uri": "https://localhost:8080/", "height": 210}
#unzip prepared data
# !unzip /content/drive/My\ Drive/nlp/pre_sum_data.zip -d /content/PreSumm/bert_data/
# + id="1Uu6FLK_PCWi" colab_type="code" outputId="fbc49885-47ad-4f7a-eda3-d456994cfd73" colab={"base_uri": "https://localhost:8080/", "height": 34}
3600*4
# + id="pNVk11yYREMC" colab_type="code" outputId="3c772280-55bc-4bee-98ba-a3fa8c818d07" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# %cd /content/PreSumm/src
bert_data_path = '../bert_data/arxiv'
model_path = '/content/drive/My\ Drive/nlp/nlp_model/pre_summ'
train_from = '/content/drive/My\ Drive/nlp/model_step_3600.pt' # '/content/drive/My\ Drive/nlp/nlp_model/pre_summ/model_step_7700.pt'
#train
# !python train.py -task abs -mode train -log_file /content/PreSumm/logs/abs_bert_arxiv \
# -bert_data_path $bert_data_path -model_path $model_path\
# -accum_count 5 -save_checkpoint_steps 3600 -batch_size 512 -use_interval true \
# -train_steps 14500 -report_every 50 -use_bert_emb true \
# -lr 2e-3 -decay_method noam -warmup_steps 100 \
# -dec_dropout 0.2 -max_pos 512 -visible_gpus 0
# # -sep_optim true -decay_method noam -lr_bert 1e-6 -lr_dec 1e-6 -warmup_steps_bert 20000 -warmup_steps_dec 10000
# # !python train.py -task abs -mode train -log_file /content/PreSumm/logs/abs_bert_arxiv \
# # -bert_data_path $bert_data_path -model_path $model_path\
# # -accum_count 5 -save_checkpoint_steps 3600 -batch_size 512 -use_interval true \
# # -train_steps 14500 -report_every 50 -use_bert_emb true \
# # -sep_optim true -decay_method noam -lr_bert 2e-6 -lr_dec 2e-6 -warmup_steps_bert 20000 -warmup_steps_dec 10000 \
# # -dec_dropout 0.2 -max_pos 512 -visible_gpus 0
#resume train
#было -lr_bert 0.002
# # !python train.py -task abs -mode train -log_file /content/PreSumm/logs/abs_bert_arxiv \
# # -bert_data_path $bert_data_path -train_from $train_from -model_path $model_path\
# # -accum_count 5 -save_checkpoint_steps 3600 -batch_size 512 -use_interval true \
# # -train_steps 14500 -report_every 100 -use_bert_emb true \
# # -sep_optim true -decay_method None -lr_bert 1e-6 -lr_dec 0.2 -warmup_steps_bert 20000 -warmup_steps_dec 10000 \
# # -dec_dropout 0.2 -max_pos 512 -visible_gpus 0
# # !python train.py -task abs -mode train -bert_data_path $bert_data_path -dec_dropout 0.2 \
# # -model_path $model_path -sep_optim true -lr_bert 0.002 -lr_dec 0.2 -save_checkpoint_steps 3600 \
# # -batch_size 512 -train_steps 14500 -report_every 1000 -accum_count 5 -use_bert_emb true \
# # -use_interval true -warmup_steps_bert 20000 -warmup_steps_dec 10000 -max_pos 512 -visible_gpus 0 \
# # -log_file /content/PreSumm/logs/abs_bert_arxiv
#calculate -max_length 70 -alpha 0.95 -min_length 10
# + id="uwajcXrbWL5K" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e0d90fc0-9b6e-4b80-e9bc-39afb15d0444"
0.00000283
2e-6 == 0.000002
Namespace(accum_count=1, alpha=0.95, batch_size=140, beam_size=5, bert_data_path='../bert_data/arxiv',
bert_model='bert-base-uncased', beta1=0.9, beta2=0.999, block_trigram=True, dec_dropout=0.2,
dec_ff_size=2048, dec_heads=8, dec_hidden_size=768, dec_layers=6, enc_dropout=0.2, enc_ff_size=512,
enc_hidden_size=512, enc_layers=6, encoder='bert', ext_dropout=0.2, ext_ff_size=2048, ext_heads=8,
ext_hidden_size=768, ext_layers=2, finetune_bert=True, generator_shard_size=32, gpu_ranks=[0],
label_smoothing=0.1, large=False, load_from_extractive='', log_file='/content/PreSumm/logs/log_val_abs_arxiv',
lr=1, lr_bert=0.002, lr_dec=0.002, max_grad_norm=0, max_length=200, max_pos=512, max_tgt_len=140,
min_length=50, mode='validate', model_path='/content/drive/My Drive/nlp/nlp_model/pre_summ',
optim='adam', param_init=0, param_init_glorot=True, recall_eval=False, report_every=1,
report_rouge=True, result_path='/content/PreSumm/logs/abs_arxiv', save_checkpoint_steps=5,
seed=666, sep_optim=True, share_emb=False, task='abs', temp_dir='../temp', test_all=False,
test_batch_size=200, test_from='/content/drive/My Drive/nlp/nlp_model/pre_summ/model_step_7200.pt',
test_start_from=-1, train_from='', train_steps=1000, use_bert_emb=False, use_interval=True,
visible_gpus='0', warmup_steps=8000, warmup_steps_bert=8000, warmup_steps_dec=8000, world_size=1)
# + [markdown] id="BcPr4_FPWO5O" colab_type="text"
# #Validate
# + id="qJdLSA8IWL-F" colab_type="code" outputId="0d993c47-926c-4da4-9a56-d66c10011cda" colab={"base_uri": "https://localhost:8080/", "height": 970}
# %cd /content/PreSumm/src
bert_data_path = '../bert_data/arxiv'
test_from = '/content/drive/My\ Drive/nlp/nlp_model/pre_summ/model_step_7200.pt'
model_path = '/content/drive/My\ Drive/nlp/nlp_model/pre_summ'
#val
# !python train.py -task abs -mode validate -test_from $test_from -bert_data_path $bert_data_path -gpu_ranks 0 -visible_gpus 0 -report_rouge true \
# -bert_data_path ../bert_data/arxiv -log_file /content/PreSumm/logs/log_val_abs_arxiv -sep_optim true -use_interval true -max_pos 512 \
# -max_length 70 -alpha 0.95 -min_length 10 -result_path /content/PreSumm/logs/abs_arxiv \
# -test_batch_size 512 -model_path $model_path
# + [markdown] id="kG6w8rsFxf87" colab_type="text"
# #Predict
# + id="b8EPrZ9-Cw-d" colab_type="code" outputId="cece15fe-6e8d-4088-cd50-6938b3114219" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# %cd /content/PreSumm/src
bert_data_path = '../bert_data/arxiv'
test_from = '/content/drive/My\ Drive/nlp/nlp_model/pre_summ/model_step_7200.pt' #'/content/drive/My\ Drive/nlp/model_step_3600.pt' #
model_path = '/content/drive/My\ Drive/nlp/nlp_model/pre_summ'
#val
# !python train.py -task abs -mode predict -test_from $test_from -bert_data_path $bert_data_path -gpu_ranks 0 -visible_gpus 0 -report_rouge true \
# -bert_data_path ../bert_data/arxiv -log_file /content/PreSumm/logs/log_val_abs_arxiv -sep_optim true -use_interval true -max_pos 512 \
# -max_length 70 -alpha 0.95 -min_length 10 -result_path /content/PreSumm/logs/abs_arxiv \
# -test_batch_size 512 -model_path $model_path
#max_length (int): maximum length output to produce
# + id="QV1D27ZnWL2U" colab_type="code" outputId="557b1d2f-75e7-47d4-f072-4ece732ff327" colab={"base_uri": "https://localhost:8080/", "height": 70}
# !ls /content/PreSumm/logs #/abs_arxiv.2200.candidate
import pandas as pd
# src = pd.read_csv('/content/PreSumm/logs/abs_arxiv.2200.raw_src', nrows=1)
pred = pd.read_csv('/content/PreSumm/logs/abs_arxiv.-1.candidate', header=None, sep=';')
test = pd.read_csv('/content/data/test.csv', encoding='utf8')
# + id="E_qgh_hkaNv9" colab_type="code" outputId="2781c369-a08f-4251-8a61-ef7495cefceb" colab={"base_uri": "https://localhost:8080/", "height": 228}
pred.iloc[:,0]
# 0 on planar embedding theorem for planar embeddi...
# 1 hodge conjectures and hodge conjectures in the...
# 2 discrete logarithms and discrete logarithms in...
# 3 nonmonotonotonic and semantical rules for nonm...
# 4 on the theory of the interaction between the a...
# + id="ZXr6Nh8DZQCo" colab_type="code" colab={}
submission_df = pd.DataFrame({'abstract': test.iloc[:,0], 'title': pred.iloc[:,0]})
submission_df.to_csv('/content/predicted_titles.csv', index=False)
# + id="HRqPS9mQZ3z5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0fd14320-8a06-4c87-9227-cc6ffa45bc81"
import string
from nltk.util import ngrams
import numpy as np
import pandas as pd
import pickle
def generate_csv(input_file='/content/predicted_titles.csv',
output_file='submission.csv',
voc_file='/content/data/vocs.pkl'):
'''
Generates file in format required for submitting result to Kaggle
Parameters:
input_file (str) : path to csv file with your predicted titles.
Should have two fields: abstract and title
output_file (str) : path to output submission file
voc_file (str) : path to voc.pkl file
'''
data = pd.read_csv(input_file)
with open(voc_file, 'rb') as voc_file:
vocs = pickle.load(voc_file)
with open(output_file, 'w') as res_file:
res_file.write('Id,Predict\n')
output_idx = 0
for row_idx, row in data.iterrows():
trg = row['title']
trg = trg.split('<q>')[0]
trg = trg.translate(str.maketrans('', '', string.punctuation)).lower().split()
trg.extend(['_'.join(ngram) for ngram in list(ngrams(trg, 2)) + list(ngrams(trg, 3))])
VOCAB_stoi = vocs[row_idx]
trg_intersection = set(VOCAB_stoi.keys()).intersection(set(trg))
trg_vec = np.zeros(len(VOCAB_stoi))
for word in trg_intersection:
trg_vec[VOCAB_stoi[word]] = 1
with open(output_file, 'a') as res_file:
for is_word in trg_vec:
res_file.write('{0},{1}\n'.format(output_idx, int(is_word)))
output_idx += 1
generate_csv()
# + id="6fMQVBlOZ35c" colab_type="code" outputId="d6b46e4c-b00b-40fe-e620-320298f6997e" colab={"base_uri": "https://localhost:8080/", "height": 70}
#2000 - 0.12958
# [2020-01-21 08:44:06,631 INFO] Validation perplexity: 30.1915
# [2020-01-21 08:44:06,631 INFO] Validation accuracy: 41.727
# # !kaggle competitions submit -c title-generation -m "PreSumm test model_step_2000.pt" -f submission.csv
# !kaggle competitions submit -c title-generation -m "PreSumm test model_step_3600.pt" -f submission.csv
# + id="PKGafbLzZ32-" colab_type="code" colab={}
# + id="ahYfSO_IZQAC" colab_type="code" colab={}
# + id="ee6hxEPYRER4" colab_type="code" outputId="f2be3ba2-eec1-41b8-dabf-6b258e14b69a" colab={"base_uri": "https://localhost:8080/", "height": 1000}
import torch
dt = torch.load('../bert_data/arxiv.valid.0.bert.pt')
dt[1]
# # %cd ../bert_data
# # !zip bert_sum_data.zip *.pt
# #!cp bert_sum_data.zip /content/drive/My\ Drive/nlp/bert_sum_data1.zip
# + id="hwSmMOUhREPx" colab_type="code" colab={}
# + id="vvwOq61hD_wh" colab_type="code" outputId="dcf74f68-81db-4fa0-b267-69f51b86ed34" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# %cd /content/PreSumm/src/
bert_data_path = '../bert_data/arxiv'
model_path = '/content/drive/My\ Drive/nlp/nlp_model/arxiv_bert/models' #'../logs/model' #'/content/drive/My\ Drive/nlp/nlp_model/bertsum'
bert_config_path = '/content/scibert_scivocab_uncased/bert_config.json'
bert_model = '/content/scibert_scivocab_uncased/weights.tar.gz'
log_file = '/content/PreSumm/logs/abs_bert_arxiv'
train_from = '/content/drive/My\ Drive/nlp/nlp_model/arxiv_bert/models/model_step_121000.pt'
#start train
# # !python train.py -mode train -encoder transformer -bert_data_path $bert_data_path -dropout 0.2 \
# # -bert_model $bert_model \
# # -model_path $model_path -lr 2e-3 -save_checkpoint_steps 500 \
# # -batch_size 512 -train_steps 10000 -report_every 500 -accum_count 5 \
# # -use_interval true -warmup_steps 20000 -visible_gpus 0 \
# # -log_file $log_file
#resume train
# !python train.py -mode train -encoder transformer -bert_data_path $bert_data_path -dropout 0.2 \
# -train_from $train_from -model_path $model_path -bert_config_path $bert_config_path -bert_model $bert_model \
# -lr 5e-2 -visible_gpus 0 -gpu_ranks 0 -world_size 1 -report_every 500 -save_checkpoint_steps 11000 \
# -batch_size 512 -decay_method noam -train_steps 200000 -accum_count 7 -log_file $log_file \
# -use_interval true -warmup_steps 100
#resume train
# # !python train.py -task abs -mode train -train_from ../models/model_step_10000.pt \
# # -bert_data_path ../bert_data/arxiv -dec_dropout 0.2 -model_path ../models \
# # -sep_optim true -lr_bert 0.002 -lr_dec 0.2 -save_checkpoint_steps 1000 -batch_size 512 \
# # -train_steps 20000 -report_every 1000 -accum_count 5 -use_bert_emb true -use_interval true \
# # -warmup_steps_bert 20000 -warmup_steps_dec 20000 -max_pos 512 -visible_gpus 0 -log_file /content/logs/abs_bert_arxiv
# # !python train.py -task abs -mode train -train_from ../models/model_step_10000.pt \
# # -bert_data_path ../bert_data/arxiv -dec_dropout 0.2 -model_path ../models \
# # -sep_optim true -lr_bert 0.002 -lr_dec 0.2 -save_checkpoint_steps 1000 -batch_size 512 \
# # -train_steps 20000 -report_every 1000 -accum_count 5 -use_bert_emb true -use_interval true \
# # -warmup_steps_bert 20000 -warmup_steps_dec 20000 -max_pos 512 -visible_gpus 0 -log_file /content/logs/abs_bert_arxiv
# + [markdown] id="0KNtEDdmFNvj" colab_type="text"
# #Validate
# + id="j29jbJPMDb-u" colab_type="code" outputId="72330497-96ed-43b0-c11e-8e64629a3db6" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# %cd /content/BertSum/src/
bert_data_path = '../bert_data/arxiv'
model_path = '/content/drive/My\ Drive/nlp/nlp_model/arxiv_bert/models' #'../logs/model' #'/content/drive/My\ Drive/nlp/nlp_model/bertsum'
bert_config_path = '/content/scibert_scivocab_uncased/bert_config.json'
bert_model = '/content/scibert_scivocab_uncased/weights.tar.gz'
log_file = '/content/BertSum/logs/abs_bert_arxiv'
train_from = '/content/drive/My\ Drive/nlp/nlp_model/arxiv_bert/models/model_step_110000.pt'
# !python train.py -mode validate -encoder transformer -dropout 0.2 \
# -bert_data_path $bert_data_path -test_from $train_from -model_path $model_path -bert_config_path $bert_config_path \
# -bert_model $bert_model -temp_dir /tmp \
# -lr 2e-3 -visible_gpus 0 -gpu_ranks 0 -world_size 1 -report_every 500 -save_checkpoint_steps 15000 \
# -batch_size 512 -decay_method noam -train_steps 100000 -accum_count 5 -log_file $log_file \
# -use_interval true -warmup_steps 100
# 30000
# [2020-01-18 06:21:12,407 INFO] Rouges at step 30000
# >> ROUGE-F(1/2/3/l): 18.44/8.08/16.21
# ROUGE-R(1/2/3/l): 70.82/33.27/62.73
# 60000
# [2020-01-18 09:46:49,751 INFO] Rouges at step 60000
# >> ROUGE-F(1/2/3/l): 18.53/8.17/16.30
# ROUGE-R(1/2/3/l): 71.05/33.57/62.99
# 75000
# [2020-01-18 11:35:12,508 INFO] Rouges at step 75000
# >> ROUGE-F(1/2/3/l): 18.57/8.19/16.33
# ROUGE-R(1/2/3/l): 71.17/33.67/63.12
# 88000
# [2020-01-19 08:17:26,555 INFO] Rouges at step 88000
# >> ROUGE-F(1/2/3/l): 18.58/8.20/16.35
# ROUGE-R(1/2/3/l): 71.22/33.70/63.17
# 99000
# [2020-01-19 09:50:21,531 INFO] Rouges at step 99000
# >> ROUGE-F(1/2/3/l): 18.57/8.20/16.34
# ROUGE-R(1/2/3/l): 71.18/33.68/63.13
# 110000
# [2020-01-19 06:41:30,907 INFO] Rouges at step 110000
# >> ROUGE-F(1/2/3/l): 18.47/8.10/16.23
# ROUGE-R(1/2/3/l): 70.90/33.35/62.82
# 121000
# [2020-01-19 06:06:43,396 INFO] Rouges at step 121000
# >> ROUGE-F(1/2/3/l): 18.48/8.09/16.23
# ROUGE-R(1/2/3/l): 70.92/33.28/62.79
# 121000 lr 5e-2
# [2020-01-19 16:25:25,110 INFO] Rouges at step 121000
# >> ROUGE-F(1/2/3/l): 18.57/8.20/16.34
# ROUGE-R(1/2/3/l): 71.17/33.69/63.13
# 132000
# [2020-01-20 10:52:15,305 INFO] Rouges at step 132000
# >> ROUGE-F(1/2/3/l): 18.57/8.20/16.34
# ROUGE-R(1/2/3/l): 71.19/33.70/63.15
#143000
# [2020-01-20 10:36:06,954 INFO] Rouges at step 143000
# >> ROUGE-F(1/2/3/l): 18.53/8.17/16.30
# ROUGE-R(1/2/3/l): 71.11/33.63/63.07
# + id="x6KlSc0WDb9C" colab_type="code" colab={}
# Namespace(accum_count=5, batch_size=512, bert_config_path='/content/scibert_scivocab_uncased/bert_config.json',
# bert_data_path='../bert_data/arxiv', bert_model='/content/scibert_scivocab_uncased/weights.tar.gz',
# beta1=0.9, beta2=0.999, block_trigram=True, dataset='', decay_method='noam', dropout=0.2,
# encoder='transformer', ff_size=512, gpu_ranks=[0], heads=4, hidden_size=128, inter_layers=2,
# log_file='/content/BertSum/logs/abs_bert_arxiv', lr=0.002, max_grad_norm=0, mode='validate',
# model_path='/content/drive/My Drive/nlp/nlp_model/arxiv_bert/models', optim='adam', param_init=0,
# param_init_glorot=True, recall_eval=False, report_every=500, report_rouge=True,
# result_path='../results/cnndm', rnn_size=512, save_checkpoint_steps=15000, seed=666,
# temp_dir='../temp', test_all=False, test_from='',
# train_from='/content/drive/My Drive/nlp/nlp_model/arxiv_bert/models/model_step_30000.pt',
# train_steps=100000, use_interval=True, visible_gpus='0', warmup_steps=100, world_size=1)
# + [markdown] id="KKJkREERiwj5" colab_type="text"
# #Test
# + id="ckWJo7O8Db4K" colab_type="code" outputId="ce39dafa-d235-4f63-a545-afbcc30f62b1" colab={"base_uri": "https://localhost:8080/", "height": 794}
# %cd /content/BertSum/src/
bert_data_path = '../bert_data/arxiv'
model_path = '/content/drive/My\ Drive/nlp/nlp_model/arxiv_bert/models' #'../logs/model' #'/content/drive/My\ Drive/nlp/nlp_model/bertsum'
bert_config_path = '/content/scibert_scivocab_uncased/bert_config.json'
bert_model = '/content/scibert_scivocab_uncased/weights.tar.gz'
log_file = '/content/BertSum/logs/abs_bert_arxiv'
test_from = '/content/drive/My\ Drive/nlp/nlp_model/arxiv_bert/models/model_step_132000.pt'
# !python train.py -mode test -encoder transformer -dropout 0.2 \
# -bert_data_path $bert_data_path -test_from $test_from -model_path $model_path -bert_config_path $bert_config_path \
# -bert_model $bert_model -temp_dir /tmp \
# -lr 2e-3 -visible_gpus 0 -gpu_ranks 0 -world_size 1 -report_every 500 -save_checkpoint_steps 15000 \
# -batch_size 512 -decay_method noam -train_steps 100000 -accum_count 5 -log_file $log_file \
# -use_interval true -warmup_steps 100
# + id="YtDO1wEunFAx" colab_type="code" colab={}
# + [markdown] id="AYpWorG6nFp3" colab_type="text"
# #Predict
# + id="URI82MrsnfzY" colab_type="code" outputId="a51c109c-1cc1-4f84-a066-882fe899237d" colab={"base_uri": "https://localhost:8080/", "height": 228}
# !git pull origin master
# + id="EI0zXFg9nFEJ" colab_type="code" outputId="97c1f4d2-40b1-4287-906d-7444b0c0f13a" colab={"base_uri": "https://localhost:8080/", "height": 178}
# %cd /content/BertSum/src/
bert_data_path = '../bert_data/arxiv'
model_path = '/content/drive/My\ Drive/nlp/nlp_model/arxiv_bert/models' #'../logs/model' #'/content/drive/My\ Drive/nlp/nlp_model/bertsum'
bert_config_path = '/content/scibert_scivocab_uncased/bert_config.json'
bert_model = '/content/scibert_scivocab_uncased/weights.tar.gz'
log_file = '/content/BertSum/logs/abs_bert_arxiv'
test_from = '/content/drive/My\ Drive/nlp/nlp_model/arxiv_bert/model_step_88000.pt'
result_file = '/content/predicted_titles.csv'
# !python train.py -mode predict -encoder transformer -dropout 0.2 \
# -bert_data_path $bert_data_path -test_from $test_from -model_path $model_path -bert_config_path $bert_config_path \
# -result_file $result_file -bert_model $bert_model -temp_dir /tmp \
# -lr 2e-3 -visible_gpus 0 -gpu_ranks 0 -world_size 1 -report_every 500 -save_checkpoint_steps 15000 \
# -batch_size 512 -decay_method noam -train_steps 100000 -accum_count 5 -log_file $log_file \
# -use_interval true -warmup_steps 100
# + id="-PDF0ciLqw9v" colab_type="code" outputId="005d4d93-5ade-41e9-9e08-5925bb4bd213" colab={"base_uri": "https://localhost:8080/", "height": 195}
# import pandas as pd
# pt = pd.read_csv('/content/predicted_titles.csv')
# pt.head()
# + id="2I4OIHIzqxAm" colab_type="code" outputId="c8259d0b-777e-4f8a-bc6e-ea8b779a870a" colab={"base_uri": "https://localhost:8080/", "height": 54}
pt.title[0].split('<q>')[0]
# import torch
# nd = torch.load('../bert_data/arxiv.new.0.bert.pt')
# len(nd)
# + id="6sX-Kkflqw66" colab_type="code" colab={}
import string
from nltk.util import ngrams
import numpy as np
import pandas as pd
import pickle
def generate_csv(input_file='/content/predicted_titles.csv',
output_file='submission.csv',
voc_file='/content/data/vocs.pkl'):
'''
Generates file in format required for submitting result to Kaggle
Parameters:
input_file (str) : path to csv file with your predicted titles.
Should have two fields: abstract and title
output_file (str) : path to output submission file
voc_file (str) : path to voc.pkl file
'''
data = pd.read_csv(input_file)
with open(voc_file, 'rb') as voc_file:
vocs = pickle.load(voc_file)
with open(output_file, 'w') as res_file:
res_file.write('Id,Predict\n')
output_idx = 0
for row_idx, row in data.iterrows():
trg = row['title']
trg = trg.split('<q>')[0]
trg = trg.translate(str.maketrans('', '', string.punctuation)).lower().split()
trg.extend(['_'.join(ngram) for ngram in list(ngrams(trg, 2)) + list(ngrams(trg, 3))])
VOCAB_stoi = vocs[row_idx]
trg_intersection = set(VOCAB_stoi.keys()).intersection(set(trg))
trg_vec = np.zeros(len(VOCAB_stoi))
for word in trg_intersection:
trg_vec[VOCAB_stoi[word]] = 1
with open(output_file, 'a') as res_file:
for is_word in trg_vec:
res_file.write('{0},{1}\n'.format(output_idx, int(is_word)))
output_idx += 1
generate_csv()
# + id="RRZocgxrqw4E" colab_type="code" outputId="89b77547-dbb2-4409-fd4b-c2b6e21a0add" colab={"base_uri": "https://localhost:8080/", "height": 70}
# !kaggle competitions submit -c title-generation -m "BertSum split model_step_88000.pt" -f submission.csv
# + id="vlhhR8tcM-yo" colab_type="code" colab={}
#save checkpoint
# #!cp /content/BertSum/logs/model/model_step_10000.pt /content/drive/My\ Drive/nlp/nlp_model/arxiv_bert
# + id="JE_hUEg1wa0L" colab_type="code" colab={}
# # !apt-get install libmagic-dev
# # !pip install python-magic
# import magic
# mime = magic.Magic(mime=True)
# mime.from_file('/content/drive/My Drive/step60000.candidate')
# with open('/content/drive/My Drive/step130000.candidate') as myfile:
# for i in range(10):
# print( next(myfile))
# + id="xAegtSgIwas7" colab_type="code" colab={}
| PreSumm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import scipy.cluster.hierarchy as shc
from sklearn.preprocessing import normalize
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# ### GLOBAL VARIABLES
DATAPATH = 'data/features/'
# ### LOAD DATASET
train_features = np.load(DATAPATH+'X_features_002.npy')
train_features.shape
data_scaled = normalize(train_features)
# ### TRAINING
plt.figure(figsize=(14, 7))
plt.title("Dendrograms")
plt.axhline(y=5.5, color='r', linestyle='--')
dend = shc.dendrogram(shc.linkage(data_scaled, method='ward'))
| notebooks/cluster/hierarchical_cluster.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 ('siena_eeg_ecg')
# language: python
# name: python3
# ---
# # Convertendo EDF para Parquet
import mne
import pandas as pd
import numpy as np
# +
# Variaveis de ambiente
import os
from os.path import join, dirname
from dotenv import load_dotenv
dotenv_path = join(dirname('__file__'), '.env')
load_dotenv(dotenv_path)
RAW_DATA = os.environ.get("RAW_DATA")
ROOT_PATH = os.environ.get("ROOT_PATH")
# -
files = pd.read_csv(f"{ROOT_PATH}{RAW_DATA}siena/database/RECORDS", header=None)[0].to_list()
pacientes = np.unique([file.split('/')[0] for file in files])
pacientes
# +
paciente = pacientes[7]
used = [file for file in files if paciente in file]
used
# +
def save_eeg(df, name):
ch_eeg = [col for col in df.columns if 'EEG' in col]
out = f"{ROOT_PATH}{RAW_DATA}EEG/{name}"
df.loc[:,ch_eeg+['window']].to_parquet(out)
def save_ekg(df, name):
ch_ekg = [col for col in df.columns if 'EKG' in col]
out = f"{ROOT_PATH}{RAW_DATA}EKG/{name}"
df.loc[:,ch_ekg+['window']].to_parquet(out)
# -
def pipeline(file):
raw = mne.io.read_raw_edf(f"{ROOT_PATH}{RAW_DATA}siena/database/{file}")
df = raw.to_data_frame()
idx = np.arange(df.shape[0])
df['id'] = idx
infos = dict(raw.info)
fs = int(infos['sfreq'])
win = []
for i in range(df.shape[0]//fs):
win = win + list(np.repeat(i, fs))
df['window'] = win
cols = [col.replace(' ','_') for col in df.columns]
df.columns = cols
name = file.split('/')[-1].replace('edf','parquet')
save_eeg(df, name)
save_ekg(df, name)
# +
for file in used:
pipeline(file)
# -
| notebooks/01_edf_to_parquet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Named Entities in the BHSA
#
# For prelimanaries, such as installing Text-Fabric and using it, consult the
# [start tutorial](https://nbviewer.jupyter.org/github/annotation/tutorials/blob/master/bhsa/start.ipynb)
#
# We show how to fetch person/place/people/measure names from the BHSA data
import os
from tf.app import use
A = use("bhsa", hoist=globals())
# If you expand the triangle in front of BHSA above, you see which features have been loaded.
#
# We need [nametype](https://etcbc.github.io/bhsa/features/nametype/) specifically.
# It is a mapping from word numbers to types of proper names.
#
# Here is a frequency distribution of its values:
F.nametype.freqList()
# We query the measure names (`mens`):
# +
query = """
word nametype=mens
"""
results = A.search(query)
# -
A.table(results)
# The frequency list promised 30 results but we see only 20. That is because there are also other things that have a name type: lexemes:
# +
queryL = """
lex nametype=mens
"""
resultsL = A.search(queryL)
# -
A.table(resultsL)
# Let's make a data file of all words that have a name type.
# We'll produce a tab-separated file with a bit of extra information.
# +
query = """
word nametype gloss*
"""
results = A.search(query)
# -
A.table(results, end=10)
A.show(results, start=10000, end=10003)
A.export(results, toFile="namedEntities.tsv")
# !head -n 20 ~/Downloads/namedEntities.tsv
# Note that this file is in UTF16 with a byte order that is chosen such that the file opens without issue in Excel.
#
# If you want to read the file by Python, it works like this:
# +
filePath = os.path.expanduser("~/Downloads/namedEntities.tsv")
i = 0
limit = 20
with open(filePath, encoding="utf16") as fh:
for line in fh:
i += 1
cells = line.rstrip("\n").split("\t")
print(i, cells)
if i > limit:
break
# -
# See also the documentation of the
# [export function](https://annotation.github.io/text-fabric/tf/advanced/display.html#tf.advanced.display.export)
# CC-BY <NAME>
| tutorial/cookbook/namedEntity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Lecture 7: Functions
#
# CSCI 1360E: Foundations for Informatics and Analytics
# + [markdown] slideshow={"slide_type": "slide"}
# ## Overview and Objectives
# + [markdown] slideshow={"slide_type": "-"}
# In this lecture, we'll introduce the concept of *functions*, critical abstractions in nearly every modern programming language. Functions are important for abstracting and categorizing large codebases into smaller, logical, and human-digestable components. By the end of this lecture, you should be able to:
# + [markdown] slideshow={"slide_type": "-"}
# - Define a function that performs a specific task
# - Set function arguments and return values
# - Differentiate *positional* arguments from *keyword* arguments
# - Write a function from scratch to answer questions in JupyterHub!
# + [markdown] slideshow={"slide_type": "slide"}
# ## Part 1: Defining Functions
# + [markdown] slideshow={"slide_type": "-"}
# A *function* in Python is not very different from a function as you've probably learned since algebra.
# + [markdown] slideshow={"slide_type": "-"}
# "Let $f$ be a function of $x$"...sound familiar? We're basically doing the same thing here.
# + [markdown] slideshow={"slide_type": "slide"}
# A function ($f$) will [usually] take something as input ($x$), perform some kind of operation on it, and then [usually] return a result ($y$). Which is why we usually see $f(x) = y$.
# -
# A function, then, is composed of *three main components*:
# + [markdown] slideshow={"slide_type": "fragment"}
# 1: **The function itself**. A [good] function will have one very specific task it performs. This task is usually reflected in its name. Take the examples of `print`, or `sqrt`, or `exp`, or `log`; all these names are very clear about what the function does.
# + [markdown] slideshow={"slide_type": "-"}
# 2: **Arguments (if any)**. Arguments (or parameters) are the *input* to the function. It's possible a function may not take any arguments at all, but often at least one is required. For example, `print` has 1 argument: a string.
# + [markdown] slideshow={"slide_type": "-"}
# 3: **Return values (if any)**. Return values are the *output* of the function. It's possible a function may not return anything; technically, `print` does not return anything. But common math functions like `sqrt` or `log` have clear return values: the output of that math operation.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Philosophy
# + [markdown] slideshow={"slide_type": "-"}
# A core tenet in writing functions is that **functions should do one thing, and do it well** (with [apologies to the Unix Philosophy](https://en.wikipedia.org/wiki/Unix_philosophy#Do_One_Thing_and_Do_It_Well)).
# + [markdown] slideshow={"slide_type": "-"}
# Writing good functions makes code *much* easier to troubleshoot and debug, as the code is already logically separated into components that perform very specific tasks. Thus, if your application is breaking, you usually have a good idea where to start looking.
# + [markdown] slideshow={"slide_type": "-"}
# It's very easy to get caught up writing "god functions": one or two massive functions that essentially do everything you need your program to do. But if something breaks, this design is very difficult to debug.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Functions vs Methods
# + [markdown] slideshow={"slide_type": "-"}
# You've probably heard the term "method" before, in this class. Quite often, these two terms are used interchangeably, and for our purposes they are pretty much the same.
# + [markdown] slideshow={"slide_type": "-"}
# **BUT**. These terms ultimately identify different constructs, so it's important to keep that in mind. Specifically:
# + [markdown] slideshow={"slide_type": "-"}
# - *Methods* are functions defined inside classes (sorry, not being covered in 1360E).
# + [markdown] slideshow={"slide_type": "-"}
# - *Functions* are not inside classes.
# + [markdown] slideshow={"slide_type": "-"}
# Otherwise, functions and methods work identically.
# + [markdown] slideshow={"slide_type": "slide"}
# So how do we write functions? At this point in the course, you've probably already seen how this works, but we'll go through it step by step regardless.
# + [markdown] slideshow={"slide_type": "-"}
# First, we define the function *header*. This is the portion of the function that defines the name of the function, the arguments, and uses the Python keyword `def` to make everything official:
# + slideshow={"slide_type": "-"}
def our_function():
pass
# + slideshow={"slide_type": "slide"}
def our_function():
pass
# + [markdown] slideshow={"slide_type": "-"}
# That's everything we need for a working function! Let's walk through it:
# + [markdown] slideshow={"slide_type": "fragment"}
# - **`def`** keyword: required before writing any function, to tell Python "hey! this is a function!"
# - **Function name**: one word (can "fake" spaces with underscores), which is the name of the function and how we'll refer to it later
# - **Arguments**: a comma-separated list of arguments the function takes to perform its task. If no arguments are needed (as above), then just open-paren-close-paren.
# - **Colon**: the colon indicates the end of the function header and the start of the actual function's code.
# - **`pass`**: since Python is sensitive to whitespace, we can't leave a function body blank; luckily, there's the `pass` keyword that does pretty much what it sounds like--no operation at all, just a placeholder.
# + [markdown] slideshow={"slide_type": "slide"}
# Admittedly, our function doesn't really do anything interesting. It takes no parameters, and the function body consists exclusively of a placeholder keyword that also does nothing. Still, it's a perfectly valid function!
# + slideshow={"slide_type": "fragment"}
# Call the function!
our_function()
# Nothing happens...no print statement, no computations, nothing.
# But there's no error either...so, yay?
# + [markdown] slideshow={"slide_type": "slide"}
# ### Other notes on functions
# + [markdown] slideshow={"slide_type": "-"}
# - You can define functions (as we did just before) almost anywhere in your code. Still, good coding practices behooves you to generally group your function definitions together, e.g. at the top of your Python file.
# + [markdown] slideshow={"slide_type": "-"}
# - Invoking or activating a function is referred to as *calling* the function. When you call a function, you type its name, an open parenthesis, any arguments you're sending to the function, and a closing parenthesis. If there are no arguments, then calling the function is as simple as typing the function name and an open-close pair of parentheses (as in our previous example).
# + [markdown] slideshow={"slide_type": "slide"}
# ## Part 2: Function Arguments
# + [markdown] slideshow={"slide_type": "-"}
# Arguments (or parameters), as stated before, are the function's input; the "$x$" to our "$f$", as it were.
# + [markdown] slideshow={"slide_type": "slide"}
# You can specify as many arguments as want, separating them by commas:
# + slideshow={"slide_type": "fragment"}
def one_arg(arg1):
print(arg1)
def two_args(arg1, arg2):
print(arg1, arg2)
def three_args(arg1, arg2, arg3):
print(arg1, arg2, arg3)
# And so on...
# + [markdown] slideshow={"slide_type": "-"}
# Like functions, you can name the arguments anything you want, though also like functions you'll probably want to give them more meaningful names besides `arg1`, `arg2`, and `arg3`. When these become just three functions among hundreds in a massive codebase written by dozens of different people, it's helpful when the code itself gives you hints as to what it does.
# + [markdown] slideshow={"slide_type": "slide"}
# When you call a function, you'll need to provide the same number of arguments in the function call as appear in the function header, otherwise Python will yell at you.
# + slideshow={"slide_type": "fragment"}
one_arg(10) # "one_arg" takes only 1 argument
# -
one_arg(10, 5) # "one_arg" won't take 2 arguments!
# + slideshow={"slide_type": "slide"}
two_args(10, 5) # "two_args", on the other hand, does take 2 arguments
# -
two_args(10, 5, 1) # ...but it doesn't take 3
# + [markdown] slideshow={"slide_type": "-"}
# To be fair, it's a pretty easy error to diagnose, but still something to keep in mind--especially as we move beyond basic "positional" arguments (as they are so called in the previous error message) into optional arguments.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Default arguments
# + [markdown] slideshow={"slide_type": "-"}
# "Positional" arguments--the only kind we've seen so far--are required whenever you call a function. If the function header specifies a positional argument, then every single call to that functions needs to have that argument specified.
# -
# In our previous example, `one_arg` is defined with 1 positional argument, so *every time you call `one_arg`, you HAVE to supply 1 argument*. Same with `two_args` defining 2 arguments, and `three_args` defining 3 arguments. Calling any of these functions without exactly the right number of arguments will result in an error.
# + [markdown] slideshow={"slide_type": "-"}
# There are cases, however, where it can be helpful to have optional, or *default*, arguments. In this case, when the function is called, the programmer can decide whether or not they want to override the default values.
# + [markdown] slideshow={"slide_type": "slide"}
# You can specify default arguments in the function header:
# + slideshow={"slide_type": "-"}
def func_with_default_arg(positional, default = 10):
print(positional, default)
# -
func_with_default_arg("pos_arg")
func_with_default_arg("pos_arg", default = 999)
# Can you piece together what's happening here?
# + [markdown] slideshow={"slide_type": "slide"}
# Note that, in the function header, one of the arguments is set equal to a particular value:
# -
# `def func_with_default_arg(positional, default = 10):`
# This means that you can call this function **with only 1 arguments**, and if you do, the second argument will take its "default" value, aka the value that is assigned in the function header (in this case, 10).
# Alternatively, you can specify a different value for the second argument if you supply 2 arguments when you call the function.
# + [markdown] slideshow={"slide_type": "fragment"}
# Can you think of examples where default arguments might be useful?
# + [markdown] slideshow={"slide_type": "slide"}
# Let's do one more small example before moving on to return values. Let's build a method which prints out a list of video games in someone's Steam library.
# + slideshow={"slide_type": "-"}
def games_in_library(username, library):
print("User '{}' owns: ".format(username))
for game in library:
print("\t{}".format(game))
# -
# You can imagine how you might modify this function to include a default argument--perhaps a list of games that everybody owns by simply registering with Steam.
# + slideshow={"slide_type": "slide"}
games_in_library('fps123', ['DOTA 2', 'Left 4 Dead', 'Doom', 'Counterstrike', 'Team Fortress 2'])
# -
games_in_library('rts456', ['Civilization V', 'Cities: Skylines', 'Sins of a Solar Empire'])
games_in_library('smrt789', ['Binding of Isaac', 'Monaco'])
# + [markdown] slideshow={"slide_type": "-"}
# In this example, our function `games_in_library` has two positional arguments: `username`, which is the Steam username of the person, and `library`, which is a list of video game titles. The function simply prints out the username and the titles they own.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Part 3: Return Values
# + [markdown] slideshow={"slide_type": "-"}
# Just as functions [can] take input, they also [can] return output for the programmer to decide what to do with.
# + [markdown] slideshow={"slide_type": "-"}
# Almost any function you will ever write will most likely have a return value of some kind. If not, your function may not be "well-behaved", aka sticking to the general guideline of doing one thing very well.
# + [markdown] slideshow={"slide_type": "-"}
# There are certainly some cases where functions won't return anything--functions that just print things, functions that run forever (yep, they exist!), functions designed specifically to test other functions--but these are highly specialized cases we are not likely to encounter in this course. Keep this in mind as a "rule of thumb": **if your function doesn't have a `return` statement, you may need to double-check your code.**
# + [markdown] slideshow={"slide_type": "slide"}
# To return a value from a function, just use the `return` keyword:
# + slideshow={"slide_type": "-"}
def identity_function(in_arg):
return in_arg
# -
x = "this is the function input"
return_value = identity_function(x)
print(return_value)
# + [markdown] slideshow={"slide_type": "-"}
# This is pretty basic: the function returns back to the programmer as output whatever was passed into the function as input. Hence, "identity function."
# + [markdown] slideshow={"slide_type": "slide"}
# Anything you can pass in as function parameters, you can return as function output, including lists:
# + slideshow={"slide_type": "-"}
def explode_string(some_string):
list_of_characters = []
for index in range(len(some_string)):
list_of_characters.append(some_string[index])
return list_of_characters
# -
words = "Blahblahblah"
output = explode_string(words)
print(output)
# + [markdown] slideshow={"slide_type": "-"}
# This function takes a string as input, uses a loop to "explode" the string, and returns a list of individual characters.
# + [markdown] slideshow={"slide_type": "slide"}
# You can even return multiple values *simultaneously* from a function. They're just treated as tuples!
# + slideshow={"slide_type": "fragment"}
def list_to_tuple(inlist):
return [10, inlist] # Yep, this is just a list.
# -
print(list_to_tuple([1, 2, 3]))
print(list_to_tuple(["one", "two", "three"]))
# + [markdown] slideshow={"slide_type": "fragment"}
# This two-way communication that functions enable--arguments as input, return values as output--is an elegant and powerful way of allowing you to design modular and human-understandable code.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Part 4: Keyword Arguments
# -
# In the previous lecture we learned about positional arguments. As the name implies, position is key:
# + slideshow={"slide_type": "fragment"}
def pet_names(name1, name2):
print("Pet 1: ", name1)
print("Pet 2: ", name2)
# -
pet1 = "King"
pet2 = "Reginald"
pet_names(pet1, pet2) # pet1 variable, then pet2 variable
pet_names(pet2, pet1) # notice we've switched the order in which they're passed to the function
# In this example, we switched the ordering of the arguments between the two function calls; consequently, the ordering of the arguments inside the function were also flipped. Hence, positional: position matters.
# + [markdown] slideshow={"slide_type": "slide"}
# In contrast, Python also has *keyword* arguments, where order no longer matters **as long as you specify the keyword**. We can use the same `pet_names` function as before.
# -
# Only this time, we'll use the names of the arguments themselves (aka, *keywords*):
# + slideshow={"slide_type": "fragment"}
pet1 = "Rocco"
pet2 = "Lucy"
# -
pet_names(name1 = pet1, name2 = pet2)
pet_names(name2 = pet2, name1 = pet1)
# + [markdown] slideshow={"slide_type": "slide"}
# As you can see, we used the names of the arguments from the function header itself (go back to the previous slide to see the definition of `pet_names` if you don't remember), setting them equal to the variable we wanted to use for that argument.
# -
# Consequently, *order doesn't matter*--Python can see that, in both function calls, we're setting `name1 = pet1` and `name2 = pet2`.
# + [markdown] slideshow={"slide_type": "slide"}
# Keyword arguments are extremely useful when it comes to default arguments.
# -
# Ordering of the keyword arguments doesn't matter; that's why we can specify some of the default parameters by keyword, leaving others at their defaults, and Python doesn't complain.
# + [markdown] slideshow={"slide_type": "slide"}
# Here's an important distinction, though:
# + [markdown] slideshow={"slide_type": "fragment"}
# - Default (optional) arguments are **always** keyword arguments, but...
# -
# - Positional (required) arguments **MUST** come before default arguments, both in the function header, and whenever you call it!
# + [markdown] slideshow={"slide_type": "fragment"}
# In essence, you can't mix-and-match the ordering of positional and default arguments using keywords.
# + [markdown] slideshow={"slide_type": "slide"}
# Here's an example of this behavior in action:
# -
# Here's our function with a default argument.
# x comes first (required), y comes second (default)
def pos_def(x, y = 10):
return x + y
# + slideshow={"slide_type": "fragment"}
# Here, we've specified both arguments, using the keyword format.
z = pos_def(x = 10, y = 20)
print(z)
# + slideshow={"slide_type": "slide"}
# We're still using the keyword format, which allows us to reverse their ordering.
z = pos_def(y = 20, x = 10)
print(z)
# + slideshow={"slide_type": "fragment"}
# But *only* specifying the default argument is a no-no.
z = pos_def(y = 20)
print(z)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Review Questions
#
# Some questions to discuss and consider:
# + [markdown] slideshow={"slide_type": "-"}
# 1: You're a software engineer for a prestigious web company named after a South American rain forest. You've been tasked with rewriting their web-based shopping cart functionality for users who purchase items through the site. Without going into too much detail, quickly list out a handful of functions you'd want to write with their basic arguments. Again, no need for excessive detail; just consider the workflow of navigating an online store and purchasing items with a shopping cart, and identify some of the key bits of functionality you'd want to write standalone functions for, as well as the inputs and outputs of those functions.
#
# 2: From where do you think the term "positional argument" gets its name?
#
# 3: Write a function, `grade`, which accepts a positional argument `number` (floating point) and returns a letter grade version of it ("A", "B", "C", "D", or "F"). Include a second, default argument that is a string and indicates whether there should be a "+", "-", or no suffix to the letter grade (default is no suffix).
#
# 4: Name a couple of functions in your experience that would benefit from being implemented with default arguments (hint: mathematical functions).
#
# 5: Give some examples for when we'd want to use keyword arguments *and* positional arguments.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Course Administrivia
# + [markdown] slideshow={"slide_type": "-"}
# - How did Assignment 2 go?
# -
# - **Assignment 3 was released yesterday.** Good luck!
# - **Assignment 4 is released tomorrow.**
# + [markdown] slideshow={"slide_type": "slide"}
# ## Additional Resources
#
# 1. <NAME>. *Python Crash Course*. 2016. ISBN-13: 978-1593276034
| lectures/L7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Implementing and comparing several pitch detection methods on sample files
#
# For simplicity I am using the Anaconda distribution on my Macbook Pro for this notebook.
#
# The purpose is to first experiment here with sample WAV files. Each file comes from a database of free samples provided free of rights by the Philharmonia Orchestra at [http://www.philharmonia.co.uk/explore/sound_samples/](http://www.philharmonia.co.uk/explore/sound_samples/).
#
# We will use 6 samples representing a long Forte string pick of each of the 6 strings of an accoustic guitar tuned in Standard E.
#
# Note: I have converted the sample files myself from their original mp3 format to wav format with 32bit, 44100Hz and mono channel.
#
# We will use two different methods for detecting the pitch and compare their results.
# For reference, here is the list of frequencies of all 6 strings expected for a well tuned guitar:
#
#
# String | Frequency | Scientific pitch notation | Sample
# --- | --- | --- | ---
# 1 (E) | 329.63 Hz | E4 | [Sample file](samples/guitar_E2_very-long_forte_normal.wav)
# 2 (B) | 246.94 Hz | B3 | [Sample file](samples/guitar_A2_very-long_forte_normal.wav)
# 3 (G) | 196.00 Hz | G3 | [Sample file](samples/guitar_D3_very-long_forte_normal.wav)
# 4 (D) | 146.83 Hz | D3 | [Sample file](samples/guitar_G3_very-long_forte_normal.wav)
# 5 (A) | 110.00 Hz | A2 | [Sample file](samples/guitar_B3_very-long_forte_normal.wav)
# 6 (E) | 82.41 Hz | E2 | [Sample file](samples/guitar_E4_very-long_forte_normal.wav)
#
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# **We will use scipy from the Anaconda distribution to read the WAV sample files**
# +
from scipy.io import wavfile
# Let's start with the first sample corresponding to the lower string E2
rate, myrecording = wavfile.read("samples/guitar_E2_very-long_forte_normal.wav")
print(rate, np_array.size)
# -
# **We define the length we want to record in seconds and the sampling rate to the source file sample rate (44100 Hz)**
duration = 1 # seconds
fs = rate # samples by second
# Let's restrict our sample to 1 second of the recording, after 0.5 second of sound to avoid the string picking
array = myrecording[int(0.5*fs):int(2.5*fs)]
print(array.size)
# **Let's plot a section of this array to look at it first**
#
# We notice a pretty periodic signal with a clear fundamental frequency: which makes sense since a guitar string vibrates producing an almost purely sinuzoidal wave
df = pd.DataFrame(array)
df.loc[25000:35000].plot()
# ## First method: Naive pitch detection using Fast Fourier Transform
#
# One first naive idea would be to "simply" take the (discrete) Fourier transform of the signal to find the fundamental frequency of the recording.
#
# Let's try that out and see what result we get.
# #### We use numpy to compute the discrete Fourier transform of the signal:
fourier = np.fft.fft(array)
# We can visualise a section of the Fourier transform to notice there is a clear fundamental frequency:
plt.plot(abs(fourier[:len(fourier)/10]))
# We notice already things are not going to be that easy. There are different harmonics picked here, and 2 of the most important ones are comparable in amplitude.
# We find the frequency corresponding to the maximum of this Fourier transform, and calculate the corresponding real frequency by re-multiplying by the sampling rate
f_max_index = np.argmax(abs(fourier[:fourier.size/2]))
freqs = np.fft.fftfreq(len(fourier))
freqs[f_max_index]*fs
# **This method detects a fundamental frequency of 248Hz, which is wrong.**
# We notice that as suspected by looking at the chart of the Fourier transform, the 3rd harmonic of the expected fundamental is detected with this naive method: 248.5 = 3 x 82.41, where 82.41Hz was the expected fundamental frequency for this sample of the E2 note.
#
# ## Taking the convolution of the sample and a Hamming window before applying FFT
# One traditional way to deal with this is issue is to first convolute the sample with a window function, such as the [Hamming window](https://en.wikipedia.org/wiki/Window_function#Hamming_window)
# +
# Work in progress: coming soon
# -
# -------
#
# ## WIP: Using Autocorrelation method for pitch detection
rec = array
rec = rec[15000:35000]
autocorr = np.correlate(rec, rec, mode='same')
plt.plot(autocorr)
| pitch_detection_from_samples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Data Analysis Tools
# # Assignment: Generating a Correlation Coefficient
#
# Following is the Python program I wrote to fulfill the third assignment of the [Data Analysis Tools online course](https://www.coursera.org/learn/data-analysis-tools/home/welcome).
#
# I decided to use [Jupyter Notebook](http://nbviewer.jupyter.org/github/ipython/ipython/blob/3.x/examples/Notebook/Index.ipynb) as it is a pretty way to write code and present results.
#
# ## Research question
#
# Using the [Gapminder database](http://www.gapminder.org/), I would like to see if an increasing Internet usage results in an increasing suicide rate. A study shows that other factors like unemployment could have a great impact.
#
# So for this assignment, the three following variables will be analyzed:
#
# - Internet Usage Rate (per 100 people)
# - Suicide Rate (per 100 000 people)
# - Unemployment Rate (% of the population of age 15+)
#
#
# ## Data management
#
# For the question I'm interested in, the countries for which data are missing will be discarded. As missing data in Gapminder database are replace directly by `NaN` no special data treatment is needed.
# + hide_input=false
# Magic command to insert the graph directly in the notebook
# %matplotlib inline
# Load a useful Python libraries for handling data
import pandas as pd
import numpy as np
import statsmodels.formula.api as smf
import scipy.stats as stats
import seaborn as sns
import matplotlib.pyplot as plt
from IPython.display import Markdown, display
# -
# Read the data
data_filename = r'gapminder.csv'
data = pd.read_csv(data_filename, low_memory=False)
data = data.set_index('country')
# General information on the Gapminder data
# + variables={"len(data)": "<p><strong>NameError</strong>: name 'data' is not defined</p>\n", "len(data.columns)": "<p><strong>NameError</strong>: name 'data' is not defined</p>\n"}
display(Markdown("Number of countries: {}".format(len(data))))
display(Markdown("Number of variables: {}".format(len(data.columns))))
# -
# Convert interesting variables in numeric format
for variable in ('internetuserate', 'suicideper100th', 'employrate'):
data[variable] = pd.to_numeric(data[variable], errors='coerce')
#
# But the unemployment rate is not provided directly. In the database, the employment rate (% of the popluation) is available. So the unemployement rate will be computed as `100 - employment rate`:
data['unemployrate'] = 100. - data['employrate']
# The first records of the data restricted to the three analyzed variables are:
subdata = data[['internetuserate', 'suicideper100th', 'unemployrate']]
subdata.tail(10)
# ## Data analysis
#
# The distribution of the three variables have been analyzed [previously](Visualizing_Data.ipynb).
#
#
# ## Variance analysis
#
# As all variables are quantitative, the Pearson correlation test is the one to apply.
#
# Let's first focus on the primary research question;
#
# - The explanatory variable is the internet use rate (quantitative variable)
# - The response variable is the suicide per 100,000 people (quantitative variable)
#
# From the scatter plot, a slope slightly positive is seen. But will the Pearson test confirm this is significant?
sns.regplot(x='internetuserate', y='suicideper100th', data=subdata)
plt.xlabel('Internet use rate (%)')
plt.ylabel('Suicide per 100 000 people (-)')
_ = plt.title('Scatterplot for the association between the Internet use rate and suicide per 100,000 people')
# +
data_clean = subdata.dropna()
correlation, pvalue = stats.pearsonr(data_clean['internetuserate'], data_clean['suicideper100th'])
display(Markdown("The correlation coefficient is {:.3g} and the associated p-value is {:.3g}.".format(correlation, pvalue)))
# -
# The correlation coefficient is 0.0735 confirming the small positive correlation. But the Pearson test tells us that **the null hypothesis cannot be rejected** as the p-value is 0.351 >> 0.05.
#
# This confirms the conclusion found when grouping the internet use rate in quartile and applying [ANOVA test](Analysis_Variance.ipynb).
# If we look now at the relationship between unemployment and suicide, it seems that there is no relationship looking at the scatterplot below.
sns.regplot(x='unemployrate', y='suicideper100th', data=subdata)
plt.xlabel('Unemployment rate (%)')
plt.ylabel('Suicide per 100 000 people (-)')
_ = plt.title('Scatterplot for the association between the unemployment rate and suicide per 100,000 people')
# Does the Pearson test confirms that conclusion?
# +
correlation, pvalue = stats.pearsonr(data_clean['unemployrate'], data_clean['suicideper100th'])
display(Markdown("The correlation coefficient is {:.3g} and the associated p-value is {:.3g}.".format(correlation, pvalue)))
# -
# The correlation coefficient is negative but really small and the p-value is large. So we can safetly conclude that there is no relationship between the unemployment rate and the suicide per 100,000 people.
# ## Another test case
#
# In order to look at the coefficient of determination, an another relationship that is significant will be analyzed below: Is the residential electricity consumption (response variable) related to the income per person (explanatory variable)?
# +
subdata2 = (data[['incomeperperson', 'relectricperperson']]
.assign(income=lambda x: pd.to_numeric(data['incomeperperson'], errors='coerce'),
electricity=lambda x: pd.to_numeric(data['relectricperperson'], errors='coerce'))
.dropna())
sns.regplot(x='income', y='electricity', data=subdata2)
plt.xlabel('Income per person (2000 US$)')
plt.ylabel('Residential electricity consumption (kWh)')
_ = plt.title('Scatterplot for the association between the income and the residential electricity consumption')
# +
correlation, pvalue = stats.pearsonr(subdata2['income'], subdata2['electricity'])
display(Markdown("The correlation coefficient is {:.3g} and the associated p-value is {:.3g}.".format(correlation, pvalue)))
display(Markdown("And the coefficient of determination is {:.3g}.".format(correlation**2)))
# -
# The Pearson test proves a significant positive relationship between income per person and residential electricity consumption as the p-value is below 0.05.
#
# Moreover, the square of the correlation coefficient, i.e. the coefficient of determination, is 0.425. This means that we can predict 42.5% of the variability of residential electricity consumption knowing the income per person.
#
# And this concludes this third assignment.
# > If you are interested into data sciences, follow me on [Tumblr](http://fcollonval.tumblr.com/).
| PearsonCorrelation.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash - Marlowe
# language: bash
# name: bash_marlowe
# ---
# <font color=red>This lecture is a work in progress that will be finalized on 21 June 2022.</font>
# # Running Marlowe Contracts without Blockchain Transactions
#
# This lecture shows how to execute a contract using `marlowe-cli`, but without submitting transactions on the blockchain. This lets one simulate the operation of a contract.
# ## Escrow Contract
#
# * A buyer wishes to purchase an item.
# * They deposit funds.
# * After they deposit funds they may report a problem with the purchase.
# * If they don’t report a problem, then the funds are released to the seller.
# * If they do report a problem, the seller may agree that there is a problem (in which case the buyer receives a refund) or they may dispute that there is a problem.
# * If the seller disputes the problem, then a mediator decides who receives the funds.
# * The contract has logic to handle situations where a party fails to act in a timely manner.
# ### Flow Chart for Escrow Example
#
# 
# ### Escrow Example in Marlowe Format
#
# ```haskell
# When
# [
# Case (Deposit (Role "<NAME> (the seller)") (Role "<NAME> (the buyer)") ada 256)
# ( When
# [
# Case (Choice (ChoiceId "Everything is alright" (Role "<NAME> (the buyer)")) [Bound 0 0])
# Close
# , Case (Choice (ChoiceId "Report problem" (Role "<NAME> (the buyer)")) [Bound 1 1])
# ( Pay (Role "<NAME> (the seller)") (Account (Role "<NAME> (the buyer)")) ada 256 )
# ( When
# [
# Case (Choice (ChoiceId "Confirm problem" (Role "Francis Beaumont (the seller)")) [Bound 1 1])
# Close
# , Case (Choice (ChoiceId "Dispute problem" (Role "<NAME>ont (the seller)")) [Bound 0 0])
# ( When
# [
# Case (Choice (ChoiceId "Dismiss claim" (Role "<NAME> (the mediator)")) [Bound 0 0])
# ( Pay (Role "<NAME> (the buyer)") (Account (Role "<NAME> (the seller)")) ada 256 )
# Close
# , Case (Choice (ChoiceId "Confirm claim" (Role "<NAME> (the mediator)")) [Bound 1 1])
# Close
# ]
# (SlotParam "Mediation deadline")
# Close
# )
# ]
# (SlotParam "Dispute deadline")
# ) Close
# ]
# (SlotParam "Complaint deadline")
# Close
# )
# ]
# (SlotParam "Payment deadline")
# Close
# ```
# ### Escrow Example in Blockly
#
# 
# ### Four of the Eight Pathways through the Escrow Contract
#
# 
# 😨 ***REDRAW AND REVISE LAYOUT***
# ## Scenario
#
# 
# 😨 ***REDRAW AND REVISE LAYOUT***
# ## Select the Parameters for the Contract
#
# First use some environment variables to store some values
# +
INITIAL_LOVELACE=3000000 # The creation transaction will deposit 3₳.
PRICE=256000000 # The prices of the item is 256₳.
SELLER_ROLE=FB # <NAME> (FB) is the seller.
BUYER_ROLE=TM # <NAME> is the buyer.
MEDIATOR_ROLE=CM # <NAME> is the mediator.
NOW=$(($(date -u +%s)*1000)) # The current time in POSIX milliseconds.
HOUR=$((60*60*1000)) # One hour, in POSIX milliseconds.
PAYMENT_DEADLINE=$((NOW+10*HOUR)) # The payment deadline, ten hours from now.
COMPLAINT_DEADLINE=$((NOW+12*HOUR)) # The complaint deadline, twelve hours from now.
DISPUTE_DEADLINE=$((NOW+14*HOUR)) # The dispute deadline, fourteen hours from now.
MEDIATION_DEADLINE=$((NOW+16*HOUR)) # The mediation deadline, sixteen hours from now.
# -
# ## Create the Contract and Its Initial State
#
# Now create the contract:
# * The contract is stored in the JSON file `tx-1.contract`.
# * The initial state is stored in the JSON file `tx-1.state`.
marlowe-cli template escrow --minimum-ada "$INITIAL_LOVELACE" \
--price "$PRICE" \
--seller "Role=$SELLER_ROLE" \
--buyer "Role=$BUYER_ROLE" \
--mediator "Role=$MEDIATOR_ROLE" \
--payment-deadline "$PAYMENT_DEADLINE" \
--complaint-deadline "$COMPLAINT_DEADLINE" \
--dispute-deadline "$DISPUTE_DEADLINE" \
--mediation-deadline "$MEDIATION_DEADLINE" \
--out-contract-file tx-1.contract \
--out-state-file tx-1.state
# ## Alternative: Download a Contract from Marlowe Playground
#
# Instead of using the `marlowe-cli template` command, one can create a contract in Marlowe Playground using Haskell, JavaScript, or Blockly.
# 1. Design the contract in Marlowe Playground.
# 2. Press the "Send to Simulator" button.
# 3. Click "Download as JSON" to download the contract, saving it as `tx-1.contract`.
#
# One also needs to create the initial state `tx-1.state`, which consists of (a) account balances, (b) preexisting choices, (c) preexisting assignments of variables, and the minimum POSIX time when the contract can be started.
#
# Simply create a JSON file with no choices or bound values and just one account with the minimum ADA deposit for the role that creates the contract (in this case, `$MEDIATOR_ROLE`).
cat << EOI > tx-1.state
{
"accounts": [
[[{"role_token": "$MEDIATOR_ROLE"}, {"currency_symbol": "", "token_name": ""}], $INITIAL_LOVELACE]
],
"choices": [],
"boundValues": [],
"minTime": 1
}
EOI
# ## Minting the Role Currency
#
# Previously we created a wallet with a signing key file `my-wallet.skey` and address file `my-wallet.address`. We also located the Cardano node socket path at `$CARDANO_NODE_SOCKET_PATH`. Now we use these to mint role tokens for use in the Marlowe contract.
export CARDANO_NODE_SOCKET_PATH=~/.local/share/Daedalus/marlowe_pioneers/cardano-node.socket
marlowe-cli util mint --testnet-magic 1567 \
--socket-path "$CARDANO_NODE_SOCKET_PATH" \
--required-signer my-wallet.skey \
--change-address $(cat my-wallet.address) \
--out-file /dev/null \
--submit 600 \
"$MEDIATOR_ROLE" "$SELLER_ROLE" "$BUYER_ROLE"
# The policy ID will be used for the Marlowe roles currency.
ROLES_CURRENCY=38fa7eb06a9bc7f219c4a7e06aaa37f2d92569f6ef350c42323910fa
# We can query the address to see that the tokens have been minted. These will also be visible in Daedalus.
cardano-cli query utxo --testnet-magic 1567 --address $(cat my-wallet.address)
# ## Step 1: Mediator Creates Escrow Contract with Initial ADA
#
# 
# 😨 ***REDRAW AND REVISE LAYOUT***
# ## Initializing Step 1
#
# Marlowe CLI uses `.marlowe` JSON files to store information about the progression of a Marlowe contract. Now that we have created the contract, we can bundle the contract, state, Plutus data, and network information into such a file.
marlowe-cli run initialize --testnet-magic 1567 \
--socket-path "$CARDANO_NODE_SOCKET_PATH" \
--roles-currency "$ROLES_CURRENCY" \
--contract-file tx-1.contract \
--state-file tx-1.state \
--out-file tx-1.marlowe \
--print-stats
ls -l tx-1.*
# ## Step 2: Buyer Deposits Funds into Seller’s Account
#
# 
# 😨 ***REDRAW AND REVISE LAYOUT***
# ## Transition from Step 1 to Step 2
marlowe-cli run prepare --marlowe-file tx-1.marlowe \
--deposit-account "Role=$SELLER_ROLE" \
--deposit-party "Role=$BUYER_ROLE" \
--deposit-amount "$PRICE" \
--invalid-before "$NOW" \
--invalid-hereafter "$((NOW+9*HOUR))" \
--out-file tx-2.marlowe \
--print-stats
# ## Step 3: The Buyer Reports That There is a Problem
#
# 
# 😨 ***REDRAW AND REVISE LAYOUT***
# ## Transition from Step 2 to Step 3
marlowe-cli run prepare --marlowe-file tx-2.marlowe \
--choice-name "Report problem" \
--choice-party "Role=$BUYER_ROLE" \
--choice-number 1 \
--invalid-before "$NOW" \
--invalid-hereafter "$((NOW+9*HOUR))" \
--out-file tx-3.marlowe \
--print-stats
# + [markdown] tags=[]
# ## Step 4: The Seller Disputes that There is a Problem
#
# 
# -
# 😨 ***REDRAW AND REVISE LAYOUT***
# ## Transition from Step 3 to Step 4
marlowe-cli run prepare --marlowe-file tx-3.marlowe \
--choice-name "Dispute problem" \
--choice-party "Role=$SELLER_ROLE" \
--choice-number 0 \
--invalid-before "$NOW" \
--invalid-hereafter "$((NOW+9*HOUR))" \
--out-file tx-4.marlowe \
--print-stats
# ## Step 5: The Mediator Dismisses the Claim
#
# 
# 😨 ***REDRAW AND REVISE LAYOUT***
# ## Transition from Step 4 to Step 5
marlowe-cli run prepare --marlowe-file tx-4.marlowe \
--choice-name "Dismiss claim" \
--choice-party "Role=$MEDIATOR_ROLE" \
--choice-number 0 \
--invalid-before "$NOW" \
--invalid-hereafter "$((NOW+9*HOUR))" \
--out-file tx-5.marlowe \
--print-stats
# ## Troubleshooting
#
# * Run the contract in the Marlowe Playground simulator before running it at the command line.
# * Pay attention to the timeouts in `When` clauses of a contract:
# * Does the timeout trigger the intendent movement of funds between accounts?
# * Is the timeout consistent with other timeouts in `Case` statements?
# * At the semantic level, these Marlowe errors can occur:
# * `TEAmbiguousTimeIntervalError`: The time interval for a transaction straddles the timeout of a `When` clause.
# * `TEApplyNoMatchError`: The attempted application of input to the contract was illegal.
# * `TEIntervalError IntervalError`
# * `InvalidInterval`: The start of the time interval is after its end.
# * `IntervalInPast`: The interval falls before the current time.
# * `TEUselessTransaction`: The attempted application of input does not change the state of the contract.
# * `TEHashMismatch`: An incorrect contract was provided for the merkleized continuation.
# ## Resources
#
# * Marlowe Debugging Cookbook: <<https://github.com/input-output-hk/marlowe-cardano/blob/mpp-cli-lectures/marlowe/debugging-cookbook.md>>.
# * Example contracts with `marlowe-cli`: <<https://github.com/input-output-hk/marlowe-cardano/blob/mpp-cli-lectures/marlowe-cli/examples/ReadMe.md>>.
# * Marlowe contract examples: <<https://github.com/input-output-hk/marlowe-cardano/tree/main/marlowe-contracts>>.
# * Marlowe Playground: <<https://playground.marlowe.iohkdev.io/#/>>.
# * Marlowe Run
# * On pioneers testnet: <<https://marlowe-run-marlowe-pioneers.plutus.aws.iohkdev.io/>>.
# * On mock network: <<https://marlowe-finance.io/>>.
# * Cardano Docs for Marlowe: <<https://docs.cardano.org/marlowe/learn-about-marlowe>>.
# * Plutus Pioneers Program: <<https://github.com/input-output-hk/plutus-pioneer-program>>.
# * Plutus Community: <<https://plutus-community.readthedocs.io/en/latest/>>.
# * The Plutonomicon: <<https://github.com/Plutonomicon/plutonomicon/blob/main/README.md>>.
# ## Summary
#
# * Use `marlowe-cli template` instantiate a contract from a template.
# * Alternatively, use Marlowe Playground to design a contract and download it as a JSON file.
# * The initial state for a contract consists of . . .
# * Account balances.
# * Preexisting choices.
# * Preexisting assignments of variables.
# * The minimum POSIX time when the contract can be started.
# * Use `marlowe-cli run initialize` to package the contract, initial state, Plutus data, and network information into a `.marlowe` file that can be used to run the contract.
# * Use `marlowe-cli run prepare` to apply input to a contract, which causes it to transition from one state to another.
# * Use `marlowe-cli util mint` if you need to mint role tokens.
# ## Other Lectures
#
# Lectures on Marlowe CLI: <<https://github.com/input-output-hk/marlowe-cardano/blob/mpp-cli-lectures/marlowe-cli/lectures/ReadMe.md>>
#
# * Overview of Marlowe CLI
# * Installing Marlowe CLI and Associated Tools
# * ~~Running Marlowe Contracts without Blockchain Transactions~~
# * Running Marlowe Contacts on the Blockchain
# * Running Marlowe Contracts with the Plutus Application Backend (PAB)
# * Reference for Marlowe CLI Commands
| marlowe-cli/lectures/03-marlowe-cli-abstract.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Accessing NEX-GDDP-CMIP6 data with the Planetary Computer STAC API
#
# The [NEX-GDDP-CMIP6 dataset](https://planetarycomputer.microsoft.com/dataset/nasa-nex-gddp-cmip6) offers global downscaled climate scenarios derived from the General Circulation Model (GCM) runs conducted under the Coupled Model Intercomparison Project Phase 6 (CMIP6) and across two of the four “Tier 1” greenhouse gas emissions scenarios known as Shared Socioeconomic Pathways (SSPs). The purpose of this dataset is to provide a set of global, high resolution, bias-corrected climate change projections that can be used to evaluate climate change impacts on processes that are sensitive to finer-scale climate gradients and the effects of local topography on climate conditions.
#
# This dataset uses a Bias-Correction Spatial Disaggregation method to downscale the original General Circulation Model runs to the finer 0.25° resolution. See the [tech note](https://www.nccs.nasa.gov/sites/default/files/NEX-GDDP-CMIP6-Tech_Note.pdf) from the [product homepage](https://www.nccs.nasa.gov/services/data-collections/land-based-products/nex-gddp-cmip6) for more details.
#
# The NEX-GDDP-CMIP6 files are stored as NetCDF in Azure Blob Storage. Each STAC Item in this collection describes a single year for one scenario for one model.
# +
import planetary_computer
import xarray as xr
import fsspec
import pystac_client
catalog = pystac_client.Client.open(
"https://planetarycomputer-staging.microsoft.com/api/stac/v1/"
)
# -
# ### Understanding the metadata
#
# The STAC metadata on the Collection, items, and assets provide information on what data is available.
collection = catalog.get_collection("nasa-nex-gddp-cmip6")
# As usual, the collection object contains information about the dataset including its spatio-temporal extent, license, and so on. We also have information unique to CMIP6. The collection is organized by `{model}-{scenario}-{year}`: there's is a single STAC item for each (valid) combination (data is not available for some; see Table 1 in the [tech note](https://www.nccs.nasa.gov/sites/default/files/NEX-GDDP-CMIP6-Tech_Note.pdf) for more). The valid values for each of these are stored in the collection's summaries:
# List the models. There are ~30 in total.
collection.summaries.get_list("cmip6:model")[:5]
# List the scenarios
collection.summaries.get_list("cmip6:scenario")
# The "historical" scenario covers the years 1950 - 2014 (inclusive). The "ssp245" and "ssp585" cover the years 2015 - 2100 (inclusive).
#
# Each item includes a handful of assets, one per variable, where each asset is a single NetCDF file with the data for that variable for that model-scenario-year.
# list the variables
collection.summaries.get_list("cmip6:variable")
# ### Querying the STAC API
#
# Each STAC item covers the same spatial region, so when using the STAC API you're likely filtering on some combination of time, model, and scenario. For example, we can get the STAC items for the "ACCESS-CM2" model for the years 1950 - 2000.
search = catalog.search(
collections=["nasa-nex-gddp-cmip6"],
datetime="1950/2000",
query={"cmip6:model": {"eq": "ACCESS-CM2"}},
)
items = search.get_all_items()
len(items)
# Eacah of these items has nine assets, one per variable, which point to the NetCDF files in Azure Blob Storage:
item = items[0]
item.assets
# ### Loading data
#
# Once you have a STAC item or items, you can load the data directly from Blob Storage using xarray. As usual, we `sign` the item, to ensure that the URLs include a read-only SAS token. See [Acessing data from blob storage](https://planetarycomputer.microsoft.com/docs/quickstarts/storage/) for more.
# +
signed_item = planetary_computer.sign(item)
hurs = xr.open_dataset(fsspec.open(signed_item.assets["hurs"].href).open())
hurs
# -
# Or you can use `xarray.open_mfdataset` to load all the variables for an item, which will combine each of the varaibles.
# %%time
ds = xr.open_mfdataset(
[fsspec.open(asset.href).open() for asset in signed_item.assets.values()]
)
# *Note that opening all those variables is relatively slow. See [below](#Using-a-Reference-File) for an alternative.*
#
# We can plot all the variables for a single day with xarray, matplotlib, and cartopy.
# +
import warnings
import cartopy.crs as ccrs
import matplotlib.pyplot as plt
import pandas as pd
warnings.filterwarnings("ignore", message="__len__")
warnings.filterwarnings("ignore", message="Iteration")
fig, axes = plt.subplots(
figsize=(16, 9),
ncols=3,
nrows=3,
subplot_kw=dict(projection=ccrs.Robinson()),
sharex=True,
sharey=True,
)
day = ds.isel(time=0)
for i, (v, data) in enumerate(day.data_vars.items()):
ax = axes.ravel()[i]
r = data.plot(ax=ax, transform=ccrs.PlateCarree(), add_colorbar=False)
ax.set(title=v)
fig.suptitle(pd.to_datetime(day.time.data).strftime("%Y-%m-%d"))
plt.tight_layout()
# -
# #### Creating a timeseries
#
# Each STAC item represents a single year. You can also use `xarray.open_mfdataset` to concatenate data for multiple years into a timeseries.
signed_items = [planetary_computer.sign(item) for item in items]
# %%time
ts = xr.open_mfdataset(
[fsspec.open(item.assets["hurs"].href).open() for item in signed_items],
)
ts["hurs"]
# + [markdown] tags=[]
# #### Using a Reference File
#
# *Note: the approach described here is experimental and may change without warning.*
#
# In the previous section, we created a single `xarray.Dataset` from many NetCDF files (either many variables, or a timeseries for a single variable). Reading the metadata of a NetCDF / HDF5 file over the network is somewhat slow, making the `open_mfdataset` operation take about 20-30 seconds, *just to read the metadata*.
#
# So in addition to the NetCDF files, we provide a **reference file** which stores the positions of each variable in each NetCDF file's binary stream. This reference file can be opened with `fsspec` and `zarr` and used normally with xarray.
# +
import requests
references = requests.get(collection.assets["ACCESS-CM2.historical"].href).json()
# -
# This reference file contains links to the original NetCDF files, along with the positions and lengths of each variable in the files. As usual, we need to sign the URLs to include short-lived tokens so that we can read the data.
# TODO: update planetary computer
for k, v in references["templates"].items():
references["templates"][k] = planetary_computer.sign(v)
# We can pass that set of references to fsspec's `ReferenceFileSystem`:
reference_filesystem = fsspec.filesystem("reference", fo=references)
# And (quickly!) open the referenced files with xarray and Zarr.
# %%time
ds = xr.open_dataset(
reference_filesystem.get_mapper("/"),
engine="zarr",
backend_kwargs={"consolidated": False},
chunks={},
)
ds
# This reference file system includes all the variables for all the years covered by the specific scenario.
# ### Next Steps
#
# For more on the NEX-GDDP-CMIP6 dataset, visit the [dataset's homepage](https://www.nccs.nasa.gov/services/data-collections/land-based-products/nex-gddp-cmip6). If you're working with large subsets of this data, you might want to [scale with Dask](https://planetarycomputer.microsoft.com/docs/quickstarts/scale-with-dask/).
| datasets/nasa-nex-gddp-cmip6/nasa-nex-gddp-cmip6-example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Prediction Latency
#
#
# This is an example showing the prediction latency of various scikit-learn
# estimators.
#
# The goal is to measure the latency one can expect when doing predictions
# either in bulk or atomic (i.e. one by one) mode.
#
# The plots represent the distribution of the prediction latency as a boxplot.
#
#
#
# +
# Authors: <NAME> <<EMAIL>>
# License: BSD 3 clause
from __future__ import print_function
from collections import defaultdict
import time
import gc
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from scipy.stats import scoreatpercentile
from sklearn.datasets.samples_generator import make_regression
from sklearn.ensemble.forest import RandomForestRegressor
from sklearn.linear_model.ridge import Ridge
from sklearn.linear_model.stochastic_gradient import SGDRegressor
from sklearn.svm.classes import SVR
from sklearn.utils import shuffle
def _not_in_sphinx():
# Hack to detect whether we are running by the sphinx builder
return '__file__' in globals()
def atomic_benchmark_estimator(estimator, X_test, verbose=False):
"""Measure runtime prediction of each instance."""
n_instances = X_test.shape[0]
runtimes = np.zeros(n_instances, dtype=np.float)
for i in range(n_instances):
instance = X_test[[i], :]
start = time.time()
estimator.predict(instance)
runtimes[i] = time.time() - start
if verbose:
print("atomic_benchmark runtimes:", min(runtimes), scoreatpercentile(
runtimes, 50), max(runtimes))
return runtimes
def bulk_benchmark_estimator(estimator, X_test, n_bulk_repeats, verbose):
"""Measure runtime prediction of the whole input."""
n_instances = X_test.shape[0]
runtimes = np.zeros(n_bulk_repeats, dtype=np.float)
for i in range(n_bulk_repeats):
start = time.time()
estimator.predict(X_test)
runtimes[i] = time.time() - start
runtimes = np.array(list(map(lambda x: x / float(n_instances), runtimes)))
if verbose:
print("bulk_benchmark runtimes:", min(runtimes), scoreatpercentile(
runtimes, 50), max(runtimes))
return runtimes
def benchmark_estimator(estimator, X_test, n_bulk_repeats=30, verbose=False):
"""
Measure runtimes of prediction in both atomic and bulk mode.
Parameters
----------
estimator : already trained estimator supporting `predict()`
X_test : test input
n_bulk_repeats : how many times to repeat when evaluating bulk mode
Returns
-------
atomic_runtimes, bulk_runtimes : a pair of `np.array` which contain the
runtimes in seconds.
"""
atomic_runtimes = atomic_benchmark_estimator(estimator, X_test, verbose)
bulk_runtimes = bulk_benchmark_estimator(estimator, X_test, n_bulk_repeats,
verbose)
return atomic_runtimes, bulk_runtimes
def generate_dataset(n_train, n_test, n_features, noise=0.1, verbose=False):
"""Generate a regression dataset with the given parameters."""
if verbose:
print("generating dataset...")
X, y, coef = make_regression(n_samples=n_train + n_test,
n_features=n_features, noise=noise, coef=True)
random_seed = 13
X_train, X_test, y_train, y_test = train_test_split(
X, y, train_size=n_train, random_state=random_seed)
X_train, y_train = shuffle(X_train, y_train, random_state=random_seed)
X_scaler = StandardScaler()
X_train = X_scaler.fit_transform(X_train)
X_test = X_scaler.transform(X_test)
y_scaler = StandardScaler()
y_train = y_scaler.fit_transform(y_train[:, None])[:, 0]
y_test = y_scaler.transform(y_test[:, None])[:, 0]
gc.collect()
if verbose:
print("ok")
return X_train, y_train, X_test, y_test
def boxplot_runtimes(runtimes, pred_type, configuration):
"""
Plot a new `Figure` with boxplots of prediction runtimes.
Parameters
----------
runtimes : list of `np.array` of latencies in micro-seconds
cls_names : list of estimator class names that generated the runtimes
pred_type : 'bulk' or 'atomic'
"""
fig, ax1 = plt.subplots(figsize=(10, 6))
bp = plt.boxplot(runtimes, )
cls_infos = ['%s\n(%d %s)' % (estimator_conf['name'],
estimator_conf['complexity_computer'](
estimator_conf['instance']),
estimator_conf['complexity_label']) for
estimator_conf in configuration['estimators']]
plt.setp(ax1, xticklabels=cls_infos)
plt.setp(bp['boxes'], color='black')
plt.setp(bp['whiskers'], color='black')
plt.setp(bp['fliers'], color='red', marker='+')
ax1.yaxis.grid(True, linestyle='-', which='major', color='lightgrey',
alpha=0.5)
ax1.set_axisbelow(True)
ax1.set_title('Prediction Time per Instance - %s, %d feats.' % (
pred_type.capitalize(),
configuration['n_features']))
ax1.set_ylabel('Prediction Time (us)')
plt.show()
def benchmark(configuration):
"""Run the whole benchmark."""
X_train, y_train, X_test, y_test = generate_dataset(
configuration['n_train'], configuration['n_test'],
configuration['n_features'])
stats = {}
for estimator_conf in configuration['estimators']:
print("Benchmarking", estimator_conf['instance'])
estimator_conf['instance'].fit(X_train, y_train)
gc.collect()
a, b = benchmark_estimator(estimator_conf['instance'], X_test)
stats[estimator_conf['name']] = {'atomic': a, 'bulk': b}
cls_names = [estimator_conf['name'] for estimator_conf in configuration[
'estimators']]
runtimes = [1e6 * stats[clf_name]['atomic'] for clf_name in cls_names]
boxplot_runtimes(runtimes, 'atomic', configuration)
runtimes = [1e6 * stats[clf_name]['bulk'] for clf_name in cls_names]
boxplot_runtimes(runtimes, 'bulk (%d)' % configuration['n_test'],
configuration)
def n_feature_influence(estimators, n_train, n_test, n_features, percentile):
"""
Estimate influence of the number of features on prediction time.
Parameters
----------
estimators : dict of (name (str), estimator) to benchmark
n_train : nber of training instances (int)
n_test : nber of testing instances (int)
n_features : list of feature-space dimensionality to test (int)
percentile : percentile at which to measure the speed (int [0-100])
Returns:
--------
percentiles : dict(estimator_name,
dict(n_features, percentile_perf_in_us))
"""
percentiles = defaultdict(defaultdict)
for n in n_features:
print("benchmarking with %d features" % n)
X_train, y_train, X_test, y_test = generate_dataset(n_train, n_test, n)
for cls_name, estimator in estimators.items():
estimator.fit(X_train, y_train)
gc.collect()
runtimes = bulk_benchmark_estimator(estimator, X_test, 30, False)
percentiles[cls_name][n] = 1e6 * scoreatpercentile(runtimes,
percentile)
return percentiles
def plot_n_features_influence(percentiles, percentile):
fig, ax1 = plt.subplots(figsize=(10, 6))
colors = ['r', 'g', 'b']
for i, cls_name in enumerate(percentiles.keys()):
x = np.array(sorted([n for n in percentiles[cls_name].keys()]))
y = np.array([percentiles[cls_name][n] for n in x])
plt.plot(x, y, color=colors[i], )
ax1.yaxis.grid(True, linestyle='-', which='major', color='lightgrey',
alpha=0.5)
ax1.set_axisbelow(True)
ax1.set_title('Evolution of Prediction Time with #Features')
ax1.set_xlabel('#Features')
ax1.set_ylabel('Prediction Time at %d%%-ile (us)' % percentile)
plt.show()
def benchmark_throughputs(configuration, duration_secs=0.1):
"""benchmark throughput for different estimators."""
X_train, y_train, X_test, y_test = generate_dataset(
configuration['n_train'], configuration['n_test'],
configuration['n_features'])
throughputs = dict()
for estimator_config in configuration['estimators']:
estimator_config['instance'].fit(X_train, y_train)
start_time = time.time()
n_predictions = 0
while (time.time() - start_time) < duration_secs:
estimator_config['instance'].predict(X_test[[0]])
n_predictions += 1
throughputs[estimator_config['name']] = n_predictions / duration_secs
return throughputs
def plot_benchmark_throughput(throughputs, configuration):
fig, ax = plt.subplots(figsize=(10, 6))
colors = ['r', 'g', 'b']
cls_infos = ['%s\n(%d %s)' % (estimator_conf['name'],
estimator_conf['complexity_computer'](
estimator_conf['instance']),
estimator_conf['complexity_label']) for
estimator_conf in configuration['estimators']]
cls_values = [throughputs[estimator_conf['name']] for estimator_conf in
configuration['estimators']]
plt.bar(range(len(throughputs)), cls_values, width=0.5, color=colors)
ax.set_xticks(np.linspace(0.25, len(throughputs) - 0.75, len(throughputs)))
ax.set_xticklabels(cls_infos, fontsize=10)
ymax = max(cls_values) * 1.2
ax.set_ylim((0, ymax))
ax.set_ylabel('Throughput (predictions/sec)')
ax.set_title('Prediction Throughput for different estimators (%d '
'features)' % configuration['n_features'])
plt.show()
# #############################################################################
# Main code
start_time = time.time()
# #############################################################################
# Benchmark bulk/atomic prediction speed for various regressors
configuration = {
'n_train': int(1e3),
'n_test': int(1e2),
'n_features': int(1e2),
'estimators': [
{'name': 'Linear Model',
'instance': SGDRegressor(penalty='elasticnet', alpha=0.01,
l1_ratio=0.25, fit_intercept=True),
'complexity_label': 'non-zero coefficients',
'complexity_computer': lambda clf: np.count_nonzero(clf.coef_)},
{'name': 'RandomForest',
'instance': RandomForestRegressor(),
'complexity_label': 'estimators',
'complexity_computer': lambda clf: clf.n_estimators},
{'name': 'SVR',
'instance': SVR(kernel='rbf'),
'complexity_label': 'support vectors',
'complexity_computer': lambda clf: len(clf.support_vectors_)},
]
}
benchmark(configuration)
# benchmark n_features influence on prediction speed
percentile = 90
percentiles = n_feature_influence({'ridge': Ridge()},
configuration['n_train'],
configuration['n_test'],
[100, 250, 500], percentile)
plot_n_features_influence(percentiles, percentile)
# benchmark throughput
throughputs = benchmark_throughputs(configuration)
plot_benchmark_throughput(throughputs, configuration)
stop_time = time.time()
print("example run in %.2fs" % (stop_time - start_time))
| scikit-learn-official-examples/applications/plot_prediction_latency.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# 说明:
# 给你一个数组 nums ,它包含 n 个正整数。你需要计算所有非空连续子数组的和,并将它们按升序排序,
# 得到一个新的包含 n * (n + 1) / 2 个数字的数组。
# 请你返回在新数组中下标为 left 到 right (下标从 1 开始)的所有数字和(包括左右端点)。
# 由于答案可能很大,请你将它对 10^9 + 7 取模后返回。
#
#
# 示例 1:
# 输入:nums = [1,2,3,4], n = 4, left = 1, right = 5
# 输出:13
# 解释:所有的子数组和为 1, 3, 6, 10, 2, 5, 9, 3, 7, 4 。
# 将它们升序排序后,我们得到新的数组 [1, 2, 3, 3, 4, 5, 6, 7, 9, 10] 。下标从 le = 1 到 ri = 5 的和为 1 + 2 + 3 + 3 + 4 = 13 。
#
# 示例 2:
# 输入:nums = [1,2,3,4], n = 4, left = 3, right = 4
# 输出:6
# 解释:给定数组与示例 1 一样,所以新数组为 [1, 2, 3, 3, 4, 5, 6, 7, 9, 10] 。下标从 le = 3 到 ri = 4 的和为 3 + 3 = 6 。
#
# 示例 3:
# 输入:nums = [1,2,3,4], n = 4, left = 1, right = 10
# 输出:50
#
# 提示:
# 1、1 <= nums.length <= 10^3
# 2、nums.length == n
# 3、1 <= nums[i] <= 100
# 4、1 <= left <= right <= n * (n + 1) / 2
# -
class Solution:
def rangeSum(self, nums, n: int, left: int, right: int) -> int:
sum_val = []
for i in range(n):
sum_val.append(nums[i])
for j in range(i+1, n):
sum_val.append(nums[j] + sum_val[-1])
sum_val.sort()
print(sum_val)
return sum(sum_val[left-1:right])
solution = Solution()
solution.rangeSum(nums = [1,2,3,4], n = 4, left = 1, right = 10)
| Array/1026/1508. Range Sum of Sorted Subarray Sums.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# formats: ipynb,md
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.0-DEV
# language: julia
# name: julia-1.7
# ---
# * https://twitter.com/genkuroki/status/1401330514175291396
# * https://github.com/genkuroki/public/blob/main/0001/harmonic%20numbers.ipynb
# * https://twitter.com/genkuroki/status/1400995381933051904
# +
versioninfo()
println()
function f(x, T=Float64)
n = 1
s = one(T)
while s < x
n += 1
s += inv(T(n))
end
n, s
end
@time f(21)
# -
using BenchmarkHistograms
@benchmark f(21) seconds=10
# +
run(`gcc --version`)
flush(stdout)
C_code = raw"""
long long f(double x) {
long long n = 1;
double s = 1.0;
while (s < x) {
n++;
s += 1.0 / (double) n;
}
return n;
}
"""
display("text/markdown", "```C\n"*C_code*"\n```")
using Libdl
libname = tempname()
libname_dl = libname * "." * Libdl.dlext
open(`gcc -Wall -O3 -march=native -xc -shared -o $libname_dl -`, "w") do f
print(f, C_code)
end
run(`ls -l $libname_dl`)
println()
f_gcc(x::Float64) = @ccall libname.f(x::Float64)::Int64
@time f_gcc(21.0)
# -
@benchmark f_gcc(21.0) seconds=10
0.67/740461601*6000125006293/60^2
# +
# Kahan-Babuska-Neumaier (KBN) algorithm
# See https://github.com/JuliaMath/KahanSummation.jl
versioninfo()
println()
function f_kbn(x, T=Float64)
n = 1
s = one(T)
c = zero(T)
while s < x
n += 1
a = inv(T(n))
t = s + a
c += abs(s) ≥ abs(a) ? ((s-t) + a) : ((a-t) + s)
s = t
end
n, s + c
end
@time f_kbn(21)
# -
using SpecialFunctions
H(n) = digamma(big(n+1)) + MathConstants.γ
setprecision(128) do; H(740461601) end
using BenchmarkHistograms
@benchmark f_kbn(21) seconds=20
| 0002/julia vs. gcc - 2021-06-06 harmonic number.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Benchmark
# **Introduction:**
# Using the data gathered from Taarifa and the Tanzanian Ministry of Water, can we predict which pumps are functional, which need some repairs, and which don't work at all? Predicting one of these three classes based and a smart understanding of which waterpoints will fail, can improve the maintenance operations and ensure that clean, potable water is available to communities across Tanzania.
#
# __Goal: To set a benchmark for improving the data quality and find a best suited algorithm.__
#
# For more details please check [Github Repo][2]
#
# [1]: https://www.drivendata.org/competitions/7/ "Link to Competetion Page"
# [2]: https://github.com/msampathkumar/datadriven_pumpit "User Code"
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.dummy import DummyClassifier
from sklearn.model_selection import train_test_split
from scripts.tools import data_transformations, df_check_stats, game, sam_pickle_save, check_metric
np.set_printoptions(precision=5)
np.random.seed(69572)
plt.style.use('ggplot')
sns.set(color_codes=True)
# %matplotlib inline
# +
# data collection
RAW_X = pd.read_csv('data/traning_set_values.csv', index_col='id')
RAW_y = pd.read_csv('data/training_set_labels.csv', index_col='id')
RAW_TEST_X = pd.read_csv('data/test_set_values.csv', index_col='id')
df_check_stats(RAW_X, RAW_y, RAW_TEST_X)
# +
# bool columns
tmp = ['public_meeting', 'permit']
RAW_X[tmp] = RAW_X[tmp].fillna(True)
RAW_TEST_X[tmp] = RAW_TEST_X[tmp].fillna(True)
# object columns list
obj_cols = RAW_X.dtypes[RAW_X.dtypes == 'O'].index.tolist()
# object columns
RAW_X[obj_cols] = RAW_X[obj_cols].fillna('Other')
RAW_TEST_X[obj_cols] = RAW_TEST_X[obj_cols].fillna('Other')
# Just assining new names to transformed dataframe pointers
X, y, TEST_X = data_transformations(RAW_X, RAW_y, RAW_TEST_X)
sam_pickle_save(X, y, TEST_X, prefix="tmp/Iteration0_")
# -
# Train Test Split Data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25,random_state=42, stratify=y)
# ## Benchmark Score
# Benchmark
clf = DummyClassifier(strategy='most_frequent', random_state=0)
clf.fit(X, y)
y_pred = clf.predict(X)
print('\nTraining Scores')
_ = check_metric(clf.predict(X_train), y_train)
print('\nTesting Scores')
_ = check_metric(clf.predict(X_test), y_test, show_cm=True)
# benchmark - rf
clf = game(X_train, X_test, y_train, y_test, algo='rf')
| pumpit/PumpIt-01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bs4 import BeautifulSoup
import urllib.request
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.common.by import By
from selenium.webdriver.support.ui import Select
from selenium.webdriver.firefox.options import Options as FirefoxOptions
from bs4 import BeautifulSoup
import pandas as pd
import re
import os
import json
page_url = "https://baseballmonster.com"
page = urllib.request.urlopen(page_url)
soup = BeautifulSoup(page, 'html.parser')
# +
# Firefox session
options = FirefoxOptions()
options.add_argument("--headless")
options.add_argument("--width=1920");
options.add_argument("--height=2160");
driver = webdriver.Firefox(options=options, service_log_path='./geckodriver.log')
driver.get(page_url)
driver.implicitly_wait(100)
# -
#Click settings
button = driver.find_elements_by_class_name("nav-item")
button = driver.find_element(By.LINK_TEXT, "Settings")
button.click()
button = driver.find_element(By.LINK_TEXT, "League Settings")
button.click()
# +
batter_table = driver.find_elements_by_class_name("table")[1]
pitcher_table = driver.find_elements_by_class_name("table")[2]
batter_rows = batter_table.find_elements(By.TAG_NAME, "tr")
pitcher_rows = pitcher_table.find_elements(By.TAG_NAME, "tr")
batter_rows = len(batter_rows)
pitcher_rows = len(pitcher_rows)
# -
batter_header_list = ["Games", "At Bats", "On-Base Percentage (OBP)", "Slugging Percentage (SLG)", "On-Base + Slugging (OPS)", "Hits", "Singles", "Doubles", "Triples", "Extra Base Hits", "Walks", "Strikeouts"]
pitcher_header_list = ["Games", "Innings Pitched", "Opponent Batting Avg (approx.)", "Quality Starts", "Complete Games", "Shutouts", "Holds", "Saves plus Holds", "Earned Runs", "Strikeouts/9", "Outs"]
for i in range(batter_rows):
radiotr = batter_table.find_elements(By.TAG_NAME, "tr")[i]
if i > 0:
for j in range(len(batter_header_list)):
title = radiotr.find_elements(By.TAG_NAME, "td")[0].text
if batter_header_list[j] == title:
radio = radiotr.find_elements(By.TAG_NAME, "td")[2]
radio = radio.find_elements(By.TAG_NAME, "input")[0]
radio.click()
for i in range(pitcher_rows):
radiotr = pitcher_table.find_elements(By.TAG_NAME, "tr")[i]
if i > 0:
for j in range(len(pitcher_header_list)):
title = radiotr.find_elements(By.TAG_NAME, "td")[0].text
if pitcher_header_list[j] == title:
radio = radiotr.find_elements(By.TAG_NAME, "td")[2]
radio = radio.find_elements(By.TAG_NAME, "input")[0]
radio.click()
save_btn = driver.find_element_by_id("ContentPlaceHolder1_SaveSettingsButton")
save_btn.click()
#Click settings
button = driver.find_elements_by_class_name("nav-item")[0]
button.click()
button = driver.find_element(By.LINK_TEXT, "Player Rankings")
button.click()
# +
driver.find_element_by_id("PlayerFilterControl").click()
date_change = Select(driver.find_element_by_id("PlayerFilterControl"))
date_change.select_by_visible_text("All Players")
rankings_table = driver.find_elements_by_class_name("table")[0]
rankings_table_html = driver.execute_script("return arguments[0].outerHTML;", rankings_table)
# +
driver.find_element_by_id("DateFilterControl").click()
date_change = Select(driver.find_element_by_id("DateFilterControl"))
date_change.select_by_visible_text("Past Days")
date_change_days = driver.find_element_by_id("DateFilterControlDAYS")
date_change_days.clear()
date_change_days.send_keys("7")
refresh = driver.find_element_by_id("ContentPlaceHolder1_GetRankingsButton")
refresh.click()
rankings_table_recent = driver.find_elements_by_class_name("table")[0]
rankings_table_recent_html = driver.execute_script("return arguments[0].outerHTML;", rankings_table_recent)
# -
driver.quit()
def isfloat(value):
try:
float(value)
return True
except ValueError:
return False
def getTableJson(tableHTML, outputName):
soup = BeautifulSoup(tableHTML)
table_header = soup.find_all("thead")
table_headers = table_header[0].find_all("th")
table_body = soup.find_all("tbody")
table_headers_array = []
pitcher_start = 0
for i in range(len(table_headers)):
table_headers_array.append(table_headers[i].string.strip())
if table_headers[i].string.strip() == 'IP':
pitcher_start = i
table_body = soup.find_all("tbody")
rankings_table_json = []
for i in range(len(table_body)):
table_rows = table_body[i].find_all("tr")
for j in range(len(table_rows)):
rankings_table_td = table_rows[j].find_all("td")
rankings_table_object = {}
for k in range(len(rankings_table_td)):
pitcher = False
if rankings_table_td[7].string.strip() == 'SP':
pitcher = True
rankings_table_object["playerType"] = "Pitcher"
elif rankings_table_td[7].string.strip() == 'RP':
pitcher = True
rankings_table_object["playerType"] = "Pitcher"
else:
rankings_table_object["playerType"] = "Batter"
if pitcher == True:
if k < 9:
if (isfloat(rankings_table_td[k].string.strip())):
rankings_table_object[table_headers_array[k]] = float(rankings_table_td[k].string.strip())
elif rankings_table_td[k].string.strip() == "":
rankings_table_object[table_headers_array[k]] = 0
else:
rankings_table_object[table_headers_array[k]] = rankings_table_td[k].string.strip()
elif k >= (pitcher_start):
if (isfloat(rankings_table_td[k].string.strip())):
rankings_table_object[table_headers_array[k]] = float(rankings_table_td[k].string.strip())
elif rankings_table_td[k].string.strip() == "":
rankings_table_object[table_headers_array[k]] = 0
else:
rankings_table_object[table_headers_array[k]] = rankings_table_td[k].string.strip()
else:
k = pitcher_start
else:
if k < pitcher_start:
if (isfloat(rankings_table_td[k].string.strip())):
rankings_table_object[table_headers_array[k]] = float(rankings_table_td[k].string.strip())
elif rankings_table_td[k].string.strip() == " ":
rankings_table_object[table_headers_array[k]] = 0
else:
rankings_table_object[table_headers_array[k]] = rankings_table_td[k].string.strip()
rankings_table_json.append(rankings_table_object)
with open(outputName, 'w') as outfile:
json.dump(rankings_table_json, outfile)
getTableJson(rankings_table_html, "../public/json/rankings.json")
getTableJson(rankings_table_recent_html, "../public/json/rankings_recent.json")
| data/baseball_scrape.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="t2hGCplrTxqd"
# # ANOMALY DETECTION
# + id="VZScT_vlTxqg"
import glob
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from PIL import Image
import keras
from keras.layers import Conv2DTranspose, ConvLSTM2D, BatchNormalization, TimeDistributed, Conv2D, LayerNormalization
from keras.models import Sequential, load_model
from scipy import signal
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
import cv2
import imutils
# + id="8NyfGkz2Txqi"
import imageio
def plot_for_offset(im, reconstructed_im, gt_frames, anomalies):
# Data for plotting
fig, (ax0, ax1, ax2,ax3) = plt.subplots(ncols=4,figsize=(10,10))
ax0.set_title('input image')
ax1.set_title('reconstructed image')
ax2.set_title('ground truth ')
ax3.set_title('anomalies')
#im = im*255.0
ax0.imshow(im)
ax1.imshow(reconstructed_im)
ax2.imshow(gt_frames)
ax3.imshow(anomalies)
# Used to return the plot as an image rray
fig.canvas.draw() # draw the canvas, cache the renderer
image = np.frombuffer(fig.canvas.tostring_rgb(), dtype='uint8')
image = image.reshape(fig.canvas.get_width_height()[::-1] + (3,))
return image
# + id="-qvkZLujTxqj"
DATASET_PATH1 ="/content/drive/MyDrive/UCSD_Anomaly_Dataset.v1p2/UCSDped1/Train"
DATASET_PATH2 ="/content/drive/MyDrive/UCSD_Anomaly_Dataset.v1p2/UCSDped2/Train"
TEST_PATH1 ="/content/drive/MyDrive/UCSD_Anomaly_Dataset.v1p2/UCSDped1/Test"
TEST_PATH2 ="/content/drive/MyDrive/UCSD_Anomaly_Dataset.v1p2/UCSDped2/Test"
MODEL_PATH = '/content/drive/MyDrive/model/model_lstm_1datasets_diff_arch_256_nomax3.hdf5'
# + id="DU4N1dJ6Txqk"
BATCH_SIZE=2
EPOCHS=3
IMAGE_SIZE = (256,256)
LENGTH_SEQUENCE = 10
# + id="Abzx3ft-Txql"
def make_sequences(frames):
sz = int(frames.shape[0]/LENGTH_SEQUENCE)
sequences = np.zeros((sz,LENGTH_SEQUENCE , IMAGE_SIZE[0],IMAGE_SIZE[1], 3))
counter = 0
for i in range(0, sz):
clip = np.zeros((LENGTH_SEQUENCE, IMAGE_SIZE[0],IMAGE_SIZE[1], 3))
for j in range(0, LENGTH_SEQUENCE):
clip[j] = frames[counter, :, :, :]
counter+=1
sequences[i] = clip
return(sequences)
# + id="Y9dbSFiHTxqm"
def make_sequence_train(list_frames, length_sequence ):
row = 0
len_train_files = list_frames.shape[0]
len_frames = list_frames.shape[1]
number_seq = int(len_frames/length_sequence)*len_train_files
seq = np.zeros((number_seq, length_sequence,IMAGE_SIZE[0],IMAGE_SIZE[1],1))
for i in range(len_train_files):
for j in range(0,len_frames,length_sequence):
for k in range(length_sequence):
seq[row,k,:,:,0] = list_frames[i,k+j,:,:,0]
row+=1
return(seq)
# + id="NfWT8i1XTxqn"
def get_training_set(DATASET_PATH):
files = sorted(glob.glob(DATASET_PATH1+'/*'))
train_set = np.zeros((len(files),200,IMAGE_SIZE[0],IMAGE_SIZE[1],1))
for idx, filename in enumerate(files):
train_files = sorted(glob.glob(filename+'/*'))
for idx_frame,file_frame in enumerate(train_files):
im = Image.open(file_frame)
im = im.resize((IMAGE_SIZE[0],IMAGE_SIZE[1]))
train_set[idx,idx_frame,:,:,0] = np.array(im, dtype=np.float32)/255.0
seq = make_sequence_train(train_set,LENGTH_SEQUENCE)
return(seq)
# + id="qeLu6ioLTxqo"
def get_single_test(number):
files = sorted(glob.glob(TEST_PATH1+'/Test'+str(number).zfill(3)+'/*'))
test_frames = np.zeros((len(files),IMAGE_SIZE[0],IMAGE_SIZE[1],1))
for idx,filename in enumerate(files):
im = cv2.imread(filename)
im = cv2.resize(im,(IMAGE_SIZE[0],IMAGE_SIZE[1]) )
test_frames[idx,:,:,0] = np.array(im[:,:,0], dtype=np.float32)/255.0
return(test_frames)
# + id="yIxywDtqTxqp"
def get_single_test_MD(number):
files = sorted(glob.glob(TEST_PATH1+'/Test'+str(number).zfill(3)+'/*'))
test_frames = np.zeros((len(files),IMAGE_SIZE[0],IMAGE_SIZE[1],3))
for idx,filename in enumerate(files):
im = cv2.imread(filename)
im = cv2.resize(im,(256,256) )
test_frames[idx,:,:,:] = np.array(im, dtype=np.uint8)/255.0
return(test_frames)
# + id="WHVrlvTSTxqq"
def get_ground_truth(number):
files = sorted(glob.glob(TEST_PATH1+'/Test'+str(number).zfill(3)+'_gt/*'))
test_gt_frames = np.zeros((len(files),IMAGE_SIZE[0],IMAGE_SIZE[1],1))
for idx,filename in enumerate(files):
im = Image.open(filename)
im = im.resize((IMAGE_SIZE[0],IMAGE_SIZE[1]))
test_gt_frames[idx,:,:,0] = np.array(im, dtype=np.float32)/255.0
return(test_gt_frames)
# + id="lLcxExMPTxqq"
def get_ground_truth_MD(number):
files = sorted(glob.glob(TEST_PATH1+'/Test'+str(number).zfill(3)+'_gt/*'))
test_gt_frames = []
for idx,filename in enumerate(files):
im = cv2.imread(filename)
im = cv2.resize(im,(500,500))
test_gt_frames.append(im)
return(test_gt_frames)
# + id="y6ps2kaKTxqr"
def get_model(reload_model=True):
if not reload_model:
return load_model(MODEL_PATH,custom_objects={'LayerNormalization': LayerNormalization})
training_set = get_training_set()
training_set = np.array(training_set)
seq = Sequential()
seq.add(TimeDistributed(Conv2D(320, (10, 10), strides=2, padding="same"), batch_input_shape=(None, LENGTH_SEQUENCE, IMAGE_SIZE[0],IMAGE_SIZE[1], 1))) #40 sequence
seq.add(LayerNormalization())
seq.add(TimeDistributed(Conv2D(128, (5,5), strides=2, padding="same")))
seq.add(LayerNormalization())
seq.add(TimeDistributed(Conv2D(64, (5,5), strides=2, padding="same")))
seq.add(LayerNormalization())
seq.add(ConvLSTM2D(64, (3, 3), padding="same", return_sequences=True))
seq.add(LayerNormalization())
seq.add(ConvLSTM2D(32, (3, 3), padding="same", return_sequences=True))
seq.add(LayerNormalization())
seq.add(ConvLSTM2D(32, (3, 3), padding="same", return_sequences=True))
seq.add(LayerNormalization())
seq.add(ConvLSTM2D(64, (3, 3), padding="same", return_sequences=True))
seq.add(LayerNormalization())
seq.add(TimeDistributed(Conv2DTranspose(64, (5, 5), strides=2, padding="same")))
seq.add(LayerNormalization())
seq.add(TimeDistributed(Conv2DTranspose(128, (5, 5), strides=2, padding="same")))
seq.add(LayerNormalization())
seq.add(TimeDistributed(Conv2DTranspose(320, (10, 10), strides=2, padding="same")))
seq.add(LayerNormalization())
seq.add(TimeDistributed(Conv2D(1, (10,10), activation="sigmoid", padding="same")))
print(seq.summary())
seq.compile(loss='mse', optimizer=keras.optimizers.Adam(lr=1e-4, decay=1e-5, epsilon=1e-6))
seq.fit(training_set, training_set,
batch_size=BATCH_SIZE, epochs=EPOCHS, shuffle=False)
seq.save(MODEL_PATH)
#seq.show()
return seq
# + id="l4TS4QIJTxqs"
def predict_test():
model = get_model(False)
print("got model")
print(test.shape)
sz = int(test.shape[0]/LENGTH_SEQUENCE)
sequences = np.zeros((sz,LENGTH_SEQUENCE , IMAGE_SIZE[0],IMAGE_SIZE[1], 1))
counter = 0
for i in range(0, sz):
clip = np.zeros((LENGTH_SEQUENCE, IMAGE_SIZE[0],IMAGE_SIZE[1], 1))
for j in range(0, LENGTH_SEQUENCE):
clip[j] = test[counter, :, :, :]
counter+=1
sequences[i] = clip
print("got data")
#reconstruction cost of all the sequences
reconstructed_sequences = model.predict(sequences,batch_size=4)
sequences_reconstruction_cost = np.array([np.linalg.norm(np.subtract(sequences[i],reconstructed_sequences[i])) for i in range(0,sz)])
sa = (sequences_reconstruction_cost - np.min(sequences_reconstruction_cost)) / np.max(sequences_reconstruction_cost)
plt.plot(sa)
plt.ylabel('anomaly score Sa(t)')
plt.xlabel('frame t')
plt.show()
return(reconstructed_sequences,sa)
# + id="v0hsLyjPTxqu"
def predict_gt_motionDetect(reconstructed_sequences, sa,number):
threshold_anomaly =np.max(sa)/2
#print(threshold_anomaly)
test_reshaped = sequences
frames = []
threshs=[]
framedeltas = []
anomalies = []
original_frame = []
firstFrame = None
#cols_list=[0,1]
#pd.read_csv('/content/drive/MyDrive/prediction/ped1/Test024/sa.csv', usecols=cols_list)
for i in range(reconstructed_sequences.shape[0]):
#print(sa.iloc[i])
#i['var1'].iloc[0]
# print(type(sa))
# print(type(sa(0)))
# print(type(threshold_anomaly))
if (sa[i]<threshold_anomaly):
#print("hello")
text = "Normal"
for j in range(LENGTH_SEQUENCE):
anomaly = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
thresh = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
frameDelta = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
cv2.imwrite('image.png', reconstructed_sequences[i,j,:,:,:])
image = cv2.imread('image.png')
frame_test = np.array(test_reshaped[i,j,:,:,:],dtype=np.uint8)
frame_predicted = np.array(image,dtype=np.uint8)
frame = cv2.absdiff(frame_test,frame_predicted)
frame = cv2.GaussianBlur(frame, (5, 5), 0)
frame = imutils.resize(frame, width=500)
anomaly= imutils.resize(anomaly, width=500)
frame_test= imutils.resize(frame_test, width=500)
cv2.putText(frame, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
cv2.putText(frame_test, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
frames.append(frame)
anomalies.append(anomaly)
original_frame.append(frame_test)
threshs.append(thresh)
framedeltas.append(frameDelta)
firstFrame = None
else:
#print("else hello")
#firstFrame = None
cnt=0
for k in range(LENGTH_SEQUENCE):
text = "Normal"
anomaly = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
frame_test = np.array(test_reshaped[i,k,:,:,:],dtype=np.uint8)
#########################
cv2.imwrite('image.png', reconstructed_sequences[i,k,:,:,:])
#*255)
image = cv2.imread('image.png')
###########################
frame_predicted = np.array(image,dtype=np.uint8)
frame = cv2.absdiff(frame_test,frame_predicted)
frame = cv2.GaussianBlur(frame, (5, 5), 0)
frame = imutils.resize(frame, width=500)
anomaly= imutils.resize(anomaly, width=500)
frame_test= imutils.resize(frame_test, width=500)
gray = cv2.cvtColor(frame, cv2.COLOR_BGR2GRAY)
gray = cv2.GaussianBlur(gray, (21, 21), 0)
if firstFrame is None:
firstFrame = gray
frameDelta = cv2.absdiff(firstFrame, gray)
thresh = cv2.threshold(frameDelta, 29, 255, cv2.THRESH_BINARY)[1]
thresh = cv2.dilate(thresh, None, iterations=2)
cnts = cv2.findContours(thresh.copy(), cv2.RETR_EXTERNAL,
cv2.CHAIN_APPROX_SIMPLE)
cnts = imutils.grab_contours(cnts)
for c in cnts:
if cv2.contourArea(c) < 500:
continue
(x, y, w, h) = cv2.boundingRect(c)
cv2.rectangle(frame, (x, y), (x + w, y + h), (0, 255, 0), 2)
cv2.rectangle(anomaly, (x, y), (x + w, y + h), (255, 255, 255), -1)
cv2.rectangle(frame_test, (x, y), (x + w, y + h), (0, 255, 0), 2)
text = "Anomaly detected"
cv2.putText(frame, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
cv2.putText(frame_test, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
frames.append(frame)
threshs.append(thresh)
framedeltas.append(frameDelta)
anomalies.append(anomaly)
original_frame.append(frame_test)
path = '/content/drive/MyDrive/motion_detection/Ped1/Test'+str(number).zfill(3)+'/'
os.makedirs(os.path.dirname(path), exist_ok=True)
imageio.mimsave(path+'frames.gif', [frames[idx] for idx in range(len(frames))], fps=1)
imageio.mimsave(path+'frames threshs.gif', [threshs[idx] for idx in range(len(threshs))], fps=1)
imageio.mimsave(path+'frames deltas.gif', [framedeltas[idx] for idx in range(len(framedeltas))], fps=1)
imageio.mimsave(path+'gt_predicted.gif', [anomalies[idx] for idx in range(len(anomalies))], fps=1)
imageio.mimsave(path+'original_frame.gif', [original_frame[idx] for idx in range(len(original_frame))], fps=1)
return(anomalies)
# + id="duhEDQU8Txq0"
def evaluate_accuracy_image_cv(number, gt_predicted):
Threshold_acc = 0.2
test_gt_frames= get_ground_truth_MD(number)
iou_score = np.zeros((1,len(test_gt_frames)))
for i in range(len(test_gt_frames)):
frame = gt_predicted[i]
gt_frames = test_gt_frames[i]
intersection = cv2.bitwise_and(frame[:,:,0], gt_frames[:,:,0])
union = cv2.bitwise_or(frame[:,:,0], gt_frames[:,:,0])
if union.any() == False:
iou_score[0,i] = 1
else:
iou_score[0,i] = np.sum(intersection)/ np.sum(union)
accuracy = np.sum(iou_score > Threshold_acc)/ iou_score.shape[1]
return(iou_score, accuracy)
# + id="wT0q7xGKTxq4"
def classic_accuracy(number, gt_predicted):
test_gt_frames= get_ground_truth_MD(number)
true_list_new_n=np.array(test_gt_frames)
pred_list_new_n=np.array(gt_predicted)
true_list_new_n=true_list_new_n.flatten()
pred_list_new_n=pred_list_new_n.flatten()
print("Confusion Matrix: ",
confusion_matrix(true_list_new_n, pred_list_new_n))
print ("Accuracy : ",
accuracy_score(true_list_new_n,pred_list_new_n)*100)
#print("Report : ",
#classification_report(true_list_new_n, pred_list_new_n))
# + id="hnIcEe_yTxq5"
def load_predictions(reconstructed_sequences):
reconstructed_sequences = np.zeros((200,IMAGE_SIZE[0],IMAGE_SIZE[1],3))
for i in range(200):
reconstructed_sequences[i,:,:,:] = cv2.imread('/content/drive/MyDrive/prediction/ped1/Test'+str(number).zfill(3)+'/'+str(i+1).zfill(3)+'.png')
sa = pd.read_csv('/content/drive/MyDrive/prediction/ped1/Test'+str(number).zfill(3)+'/sa.csv', index_col=0)
sa=np.array(sa)
sequences_pred = make_sequences(reconstructed_sequences)
return(sequences_pred, sa)
# + id="cMn9n3OSTxq5"
def saliency_image(image):
saliency = cv2.saliency.StaticSaliencySpectralResidual_create()
(success, saliencyMap) = saliency.computeSaliency(image)
saliencyMap = (saliencyMap * 255).astype("uint8")
return(saliencyMap)
def predict_gt_saliency(reconstructed_sequences, sa):
reshaped = np.reshape(reconstructed_sequences, ((200,IMAGE_SIZE[0],IMAGE_SIZE[1],1)))
ground_truth_pred = []
threshold = 180
#threshold = 4*255
for i in range(len(reshaped)):
idx = int(i/10)
anomalie=np.zeros((IMAGE_SIZE[0],IMAGE_SIZE[1]))
if sa[idx]<np.max(sa)/2:
pass
else:
predi_image = reshaped[i,:,:,0]*255
test_image = test[i,:,:,0]*255
diff = np.abs(predi_image-test_image)
tmp = diff[:,:]
H = signal.convolve2d(tmp, np.ones((4,4)), mode='same')
saliency = saliency_image(H)
x,y = np.where(saliency> threshold)
anomalie[x,y]=1
ground_truth_pred.append(anomalie)
ground_truth_pred = np.array(ground_truth_pred)
return(ground_truth_pred)
# + [markdown] id="iePlUU9SY6ay"
#
# + [markdown] id="JHEz3Ji4dHFI"
# # TEST
#
#
#
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 346} id="qdpnIGn-jeJ5" outputId="0e8ca393-a384-4eb5-df68-20ef482dfcea"
def predict_gt_motionDetect(reconstructed_sequences, sa,number):
threshold_anomaly =np.max(sa)/2
#print(threshold_anomaly)
test_reshaped = sequences
frames = []
threshs=[]
framedeltas = []
anomalies = []
original_frame = []
firstFrame = None
#cols_list=[0,1]
#pd.read_csv('/content/drive/MyDrive/prediction/ped1/Test024/sa.csv', usecols=cols_list)
for i in range(reconstructed_sequences.shape[0]):
#print(sa.iloc[i])
#i['var1'].iloc[0]
# print(type(sa))
# print(type(sa(0)))
# print(type(threshold_anomaly))
if (sa[i]<threshold_anomaly):
#print("hello")
text = "Normal"
for j in range(LENGTH_SEQUENCE):
anomaly = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
thresh = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
frameDelta = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
cv2.imwrite('image.png', reconstructed_sequences[i,j,:,:,:])
image = cv2.imread('image.png')
frame_test = np.array(test_reshaped[i,j,:,:,:],dtype=np.uint8)
frame_predicted = np.array(image,dtype=np.uint8)
frame = cv2.absdiff(frame_test,frame_predicted)
frame = cv2.GaussianBlur(frame, (5, 5), 0)
frame = imutils.resize(frame, width=500)
anomaly= imutils.resize(anomaly, width=500)
frame_test= imutils.resize(frame_test, width=500)
cv2.putText(frame, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
cv2.putText(frame_test, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
frames.append(frame)
anomalies.append(anomaly)
original_frame.append(frame_test)
threshs.append(thresh)
framedeltas.append(frameDelta)
firstFrame = None
else:
#print("else hello")
#firstFrame = None
cnt=0
for k in range(LENGTH_SEQUENCE):
text = "Normal"
anomaly = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
frame_test = np.array(test_reshaped[i,k,:,:,:],dtype=np.uint8)
#########################
cv2.imwrite('image.png', reconstructed_sequences[i,k,:,:,:])
#*255)
image = cv2.imread('image.png')
###########################
frame_predicted = np.array(image,dtype=np.uint8)
frame = cv2.absdiff(frame_test,frame_predicted)
frame = cv2.GaussianBlur(frame, (5, 5), 0)
frame = imutils.resize(frame, width=500)
anomaly= imutils.resize(anomaly, width=500)
frame_test= imutils.resize(frame_test, width=500)
gray = cv2.cvtColor(frame, cv2.COLOR_BGR2GRAY)
gray = cv2.GaussianBlur(gray, (21, 21), 0)
if firstFrame is None:
firstFrame = gray
frameDelta = cv2.absdiff(firstFrame, gray)
thresh = cv2.threshold(frameDelta, 29, 255, cv2.THRESH_BINARY)[1]
thresh = cv2.dilate(thresh, None, iterations=2)
cnts = cv2.findContours(thresh.copy(), cv2.RETR_EXTERNAL,
cv2.CHAIN_APPROX_SIMPLE)
cnts = imutils.grab_contours(cnts)
for c in cnts:
if cv2.contourArea(c) < 500:
continue
(x, y, w, h) = cv2.boundingRect(c)
cv2.rectangle(frame, (x, y), (x + w, y + h), (0, 255, 0), 2)
cv2.rectangle(anomaly, (x, y), (x + w, y + h), (255, 255, 255), -1)
cv2.rectangle(frame_test, (x, y), (x + w, y + h), (0, 255, 0), 2)
text = "Anomaly detected"
cv2.putText(frame, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
cv2.putText(frame_test, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
frames.append(frame)
threshs.append(thresh)
framedeltas.append(frameDelta)
anomalies.append(anomaly)
original_frame.append(frame_test)
path = '/content/drive/MyDrive/motion_detection/Ped1/Test'+str(number).zfill(3)+'/'
os.makedirs(os.path.dirname(path), exist_ok=True)
imageio.mimsave(path+'frames.gif', [frames[idx] for idx in range(100,len(frames))], fps=3)
imageio.mimsave(path+'frames threshs.gif', [threshs[idx] for idx in range(100,len(threshs))], fps=3)
imageio.mimsave(path+'frames deltas.gif', [framedeltas[idx] for idx in range(100,len(framedeltas))], fps=3)
imageio.mimsave(path+'gt_predicted.gif', [anomalies[idx] for idx in range(100,len(anomalies))], fps=3)
imageio.mimsave(path+'original_frame.gif', [original_frame[idx] for idx in range(100,len(original_frame))], fps=3)
return(anomalies)
number = 1
test = get_single_test(number)
predicted_sequences, sa= predict_test()
reshaped = np.reshape(predicted_sequences, ((predicted_sequences.shape[0]*LENGTH_SEQUENCE,IMAGE_SIZE[0],IMAGE_SIZE[1],1)))
for i in range(200):
path = '/content/drive/MyDrive/testing_path/Peds1/Test'+str(number).zfill(3)+'/'+str(i+1).zfill(3)+'.png'
os.makedirs(os.path.dirname(path), exist_ok=True)
cv2.imwrite(path, reshaped[i,:,:,0]*255)
sa = pd.DataFrame(sa, columns=['sa'])
path = '/content/drive/MyDrive/prediction/ped1/Test'+str(number).zfill(3)+'/sa.csv'
os.makedirs(os.path.dirname(path), exist_ok=True)
sa.to_csv(path)
sequences_pred = predicted_sequences*255
test = get_single_test_MD(number)
sequences = make_sequences(test*255)
newarr=[]
import pandas as pd
col_list=["sa"]
saa = pd.read_csv('/content/drive/MyDrive/prediction/ped1/Test024/sa.csv',usecols=col_list)
print(saa.shape[0])
for i in range(saa.shape[0]):
newarr.append(float(saa.iloc[i].to_string().split()[1]))
predictions=predict_gt_motionDetect(sequences_pred,newarr,number)
# + [markdown] id="Bcbc06wIv5F6"
# # TEST on
# + id="T6M8v5Kyv5F8"
def predict_gt_motionDetect(reconstructed_sequences, sa,number):
threshold_anomaly =np.max(sa)/2
print(threshold_anomaly)
test_reshaped = sequences
frames = []
threshs=[]
framedeltas = []
anomalies = []
original_frame = []
firstFrame = None
#cols_list=[0,1]
#pd.read_csv('/content/drive/MyDrive/prediction/ped1/Test024/sa.csv', usecols=cols_list)
for i in range(reconstructed_sequences.shape[0]):
#print(sa.iloc[i])
#i['var1'].iloc[0]
# print(type(sa))
# print(type(sa(0)))
# print(type(threshold_anomaly))
if (sa[i]<threshold_anomaly):
#print("hello")
text = "Normal"
for j in range(LENGTH_SEQUENCE):
anomaly = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
thresh = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
frameDelta = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
cv2.imwrite('image.png', reconstructed_sequences[i,j,:,:,:])
image = cv2.imread('image.png')
frame_test = np.array(test_reshaped[i,j,:,:,:],dtype=np.uint8)
frame_predicted = np.array(image,dtype=np.uint8)
frame = cv2.absdiff(frame_test,frame_predicted)
frame = cv2.GaussianBlur(frame, (5, 5), 0)
frame = imutils.resize(frame, width=500)
anomaly= imutils.resize(anomaly, width=500)
frame_test= imutils.resize(frame_test, width=500)
cv2.putText(frame, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
cv2.putText(frame_test, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
frames.append(frame)
anomalies.append(anomaly)
original_frame.append(frame_test)
threshs.append(thresh)
framedeltas.append(frameDelta)
firstFrame = None
else:
#print("else hello")
#firstFrame = None
cnt=0
for k in range(LENGTH_SEQUENCE):
text = "Normal"
anomaly = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
frame_test = np.array(test_reshaped[i,k,:,:,:],dtype=np.uint8)
#########################
cv2.imwrite('image.png', reconstructed_sequences[i,k,:,:,:])
#*255)
image = cv2.imread('image.png')
###########################
frame_predicted = np.array(image,dtype=np.uint8)
frame = cv2.absdiff(frame_test,frame_predicted)
frame = cv2.GaussianBlur(frame, (5, 5), 0)
frame = imutils.resize(frame, width=500)
anomaly= imutils.resize(anomaly, width=500)
frame_test= imutils.resize(frame_test, width=500)
gray = cv2.cvtColor(frame, cv2.COLOR_BGR2GRAY)
gray = cv2.GaussianBlur(gray, (21, 21), 0)
if firstFrame is None:
firstFrame = gray
frameDelta = cv2.absdiff(firstFrame, gray)
thresh = cv2.threshold(frameDelta, 29, 255, cv2.THRESH_BINARY)[1]
thresh = cv2.dilate(thresh, None, iterations=2)
cnts = cv2.findContours(thresh.copy(), cv2.RETR_EXTERNAL,
cv2.CHAIN_APPROX_SIMPLE)
cnts = imutils.grab_contours(cnts)
for c in cnts:
if cv2.contourArea(c) < 500:
continue
(x, y, w, h) = cv2.boundingRect(c)
cv2.rectangle(frame, (x, y), (x + w, y + h), (0, 255, 0), 2)
cv2.rectangle(anomaly, (x, y), (x + w, y + h), (255, 255, 255), -1)
cv2.rectangle(frame_test, (x, y), (x + w, y + h), (0, 255, 0), 2)
text = "Anomaly detected"
cv2.putText(frame, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
cv2.putText(frame_test, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
frames.append(frame)
threshs.append(thresh)
framedeltas.append(frameDelta)
anomalies.append(anomaly)
original_frame.append(frame_test)
path = '/content/drive/MyDrive/motion_detection/Ped1/Test'+str(number).zfill(3)+'/'
os.makedirs(os.path.dirname(path), exist_ok=True)
imageio.mimsave(path+'frames.gif', [frames[idx] for idx in range(150,len(frames))], fps=1)
imageio.mimsave(path+'frames threshs.gif', [threshs[idx] for idx in range(150,len(threshs))], fps=1)
imageio.mimsave(path+'frames deltas.gif', [framedeltas[idx] for idx in range(150,len(framedeltas))], fps=1)
imageio.mimsave(path+'gt_predicted.gif', [anomalies[idx] for idx in range(150,len(anomalies))], fps=1)
imageio.mimsave(path+'original_frame.gif', [original_frame[idx] for idx in range(150,len(original_frame))], fps=1)
return(anomalies)
# + id="SLk-Yx0bv5F_"
number = 24
# + id="JCOgGg4bv5GA"
test = get_single_test(number)
# + colab={"base_uri": "https://localhost:8080/", "height": 331} id="7HEvtUzwv5GB" outputId="0ac4f46a-fb31-4063-b268-dc10255db05b"
predicted_sequences, sa= predict_test()
# + id="MBO5A5Dav5Gi"
#### Save the predicted seqeunces
reshaped = np.reshape(predicted_sequences, ((predicted_sequences.shape[0]*LENGTH_SEQUENCE,IMAGE_SIZE[0],IMAGE_SIZE[1],1)))
for i in range(200):
path = '/content/drive/MyDrive/testing_path/Peds1/Test'+str(number).zfill(3)+'/'+str(i+1).zfill(3)+'.png'
os.makedirs(os.path.dirname(path), exist_ok=True)
cv2.imwrite(path, reshaped[i,:,:,0]*255)
sa = pd.DataFrame(sa, columns=['sa'])
path = '/content/drive/MyDrive/prediction/ped1/Test'+str(number).zfill(3)+'/sa.csv'
os.makedirs(os.path.dirname(path), exist_ok=True)
sa.to_csv(path)
# + id="H-aGMWpev5Gl"
sequences_pred = predicted_sequences*255
# + id="L8IRslIrv5Gm"
test = get_single_test_MD(number)
# + id="a8ic5-iSv5Gn"
sequences = make_sequences(test*255)
# + colab={"base_uri": "https://localhost:8080/"} id="jQDhsKIXv5Go" outputId="d56ff6cf-a600-445e-f7b5-f4dfa4df1862"
newarr=[]
import pandas as pd
col_list=["sa"]
saa = pd.read_csv('/content/drive/MyDrive/prediction/ped1/Test024/sa.csv',usecols=col_list)
print(saa.shape[0])
for i in range(saa.shape[0]):
newarr.append(float(saa.iloc[i].to_string().split()[1]))
print(newarr)
print(type(newarr[0]))
# + colab={"base_uri": "https://localhost:8080/"} id="i4hYhJ23v5Gp" outputId="3024c966-7c3a-470b-b998-11ba0345ab17"
predictions=predict_gt_motionDetect(sequences_pred,newarr,number)
# + [markdown] id="FwaAI06Ev9zt"
# # TEST on
# + id="Q8W0z461v9zu"
def predict_gt_motionDetect(reconstructed_sequences, sa,number):
threshold_anomaly =np.max(sa)/2
print(threshold_anomaly)
test_reshaped = sequences
frames = []
threshs=[]
framedeltas = []
anomalies = []
original_frame = []
firstFrame = None
#cols_list=[0,1]
#pd.read_csv('/content/drive/MyDrive/prediction/ped1/Test024/sa.csv', usecols=cols_list)
for i in range(reconstructed_sequences.shape[0]):
#print(sa.iloc[i])
#i['var1'].iloc[0]
# print(type(sa))
# print(type(sa(0)))
# print(type(threshold_anomaly))
if (sa[i]<threshold_anomaly):
#print("hello")
text = "Normal"
for j in range(LENGTH_SEQUENCE):
anomaly = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
thresh = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
frameDelta = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
cv2.imwrite('image.png', reconstructed_sequences[i,j,:,:,:])
image = cv2.imread('image.png')
frame_test = np.array(test_reshaped[i,j,:,:,:],dtype=np.uint8)
frame_predicted = np.array(image,dtype=np.uint8)
frame = cv2.absdiff(frame_test,frame_predicted)
frame = cv2.GaussianBlur(frame, (5, 5), 0)
frame = imutils.resize(frame, width=500)
anomaly= imutils.resize(anomaly, width=500)
frame_test= imutils.resize(frame_test, width=500)
cv2.putText(frame, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
cv2.putText(frame_test, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
frames.append(frame)
anomalies.append(anomaly)
original_frame.append(frame_test)
threshs.append(thresh)
framedeltas.append(frameDelta)
firstFrame = None
else:
#print("else hello")
#firstFrame = None
cnt=0
for k in range(LENGTH_SEQUENCE):
text = "Normal"
anomaly = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
frame_test = np.array(test_reshaped[i,k,:,:,:],dtype=np.uint8)
#########################
cv2.imwrite('image.png', reconstructed_sequences[i,k,:,:,:])
#*255)
image = cv2.imread('image.png')
###########################
frame_predicted = np.array(image,dtype=np.uint8)
frame = cv2.absdiff(frame_test,frame_predicted)
frame = cv2.GaussianBlur(frame, (5, 5), 0)
frame = imutils.resize(frame, width=500)
anomaly= imutils.resize(anomaly, width=500)
frame_test= imutils.resize(frame_test, width=500)
gray = cv2.cvtColor(frame, cv2.COLOR_BGR2GRAY)
gray = cv2.GaussianBlur(gray, (21, 21), 0)
if firstFrame is None:
firstFrame = gray
frameDelta = cv2.absdiff(firstFrame, gray)
thresh = cv2.threshold(frameDelta, 29, 255, cv2.THRESH_BINARY)[1]
thresh = cv2.dilate(thresh, None, iterations=2)
cnts = cv2.findContours(thresh.copy(), cv2.RETR_EXTERNAL,
cv2.CHAIN_APPROX_SIMPLE)
cnts = imutils.grab_contours(cnts)
for c in cnts:
if cv2.contourArea(c) < 500:
continue
(x, y, w, h) = cv2.boundingRect(c)
cv2.rectangle(frame, (x, y), (x + w, y + h), (0, 255, 0), 2)
cv2.rectangle(anomaly, (x, y), (x + w, y + h), (255, 255, 255), -1)
cv2.rectangle(frame_test, (x, y), (x + w, y + h), (0, 255, 0), 2)
text = "Anomaly detected"
cv2.putText(frame, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
cv2.putText(frame_test, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
frames.append(frame)
threshs.append(thresh)
framedeltas.append(frameDelta)
anomalies.append(anomaly)
original_frame.append(frame_test)
path = '/content/drive/MyDrive/motion_detection/Ped1/Test'+str(number).zfill(3)+'/'
os.makedirs(os.path.dirname(path), exist_ok=True)
imageio.mimsave(path+'frames.gif', [frames[idx] for idx in range(len(frames))], fps=1)
imageio.mimsave(path+'frames threshs.gif', [threshs[idx] for idx in range(len(threshs))], fps=1)
imageio.mimsave(path+'frames deltas.gif', [framedeltas[idx] for idx in range(len(framedeltas))], fps=1)
imageio.mimsave(path+'gt_predicted.gif', [anomalies[idx] for idx in range(len(anomalies))], fps=1)
imageio.mimsave(path+'original_frame.gif', [original_frame[idx] for idx in range(53,len(original_frame))], fps=1)
return(anomalies)
# + id="Mbg29TTVv9zw"
number = 3
# + id="o_otOCNgv9zw"
test = get_single_test(number)
# + colab={"base_uri": "https://localhost:8080/", "height": 330} id="QrTmI_nJv9zx" outputId="41399904-140d-4582-b857-23599faaeb8b"
predicted_sequences, sa= predict_test()
# + colab={"base_uri": "https://localhost:8080/"} id="bQmmmYxhv9zy" outputId="7f19fb56-5060-47a1-fd85-96cb0b1f1bd1"
np.max(sa)/2
# + colab={"base_uri": "https://localhost:8080/"} id="fbLy9M24v9zz" outputId="e9405dd4-1efc-4e20-c8f0-9cb19748f3ea"
#### Save the predicted seqeunces
reshaped = np.reshape(predicted_sequences, ((predicted_sequences.shape[0]*LENGTH_SEQUENCE,IMAGE_SIZE[0],IMAGE_SIZE[1],1)))
for i in range(200):
path = '/content/drive/MyDrive/testing_path/Peds1/Test'+str(number).zfill(3)+'/'+str(i+1).zfill(3)+'.png'
os.makedirs(os.path.dirname(path), exist_ok=True)
cv2.imwrite(path, reshaped[i,:,:,0]*255)
sa = pd.DataFrame(sa, columns=['sa'])
path = '/content/drive/MyDrive/prediction/ped1/Test'+str(number).zfill(3)+'/sa.csv'
os.makedirs(os.path.dirname(path), exist_ok=True)
sa.to_csv(path)
# + id="CNDDJx2uv9z1"
##### load predicted sequences
#sequences_pred = load_predictions(predicted_sequences)
# + colab={"base_uri": "https://localhost:8080/"} id="ZZsAi0jUv9z2" outputId="c2ed429d-26ad-4628-b282-e1e6e93682e5"
sequences_pred = predicted_sequences*255
sequences_pred.shape
# + id="bnz0jOoJv9z3"
test = get_single_test_MD(number)
# + colab={"base_uri": "https://localhost:8080/"} id="XiVeUaeyv9z4" outputId="55835f47-c43d-4c03-bf64-1cc49c63df8b"
sequences = make_sequences(test*255)
sequences.shape
# + colab={"base_uri": "https://localhost:8080/"} id="vz6fDB8Mv9z5" outputId="06ca0346-c8bc-4b82-fa79-c5781cb5e800"
newarr=[]
import pandas as pd
col_list=["sa"]
saa = pd.read_csv('/content/drive/MyDrive/prediction/ped1/Test024/sa.csv',usecols=col_list)
print(saa.shape[0])
for i in range(saa.shape[0]):
newarr.append(float(saa.iloc[i].to_string().split()[1]))
print(newarr)
print(type(newarr[0]))
# + colab={"base_uri": "https://localhost:8080/"} id="nJxTzANDv9z6" outputId="0c730aa4-9560-4bd6-9d4e-f3e545a51e58"
predictions=predict_gt_motionDetect(sequences_pred,newarr,number)
# + [markdown] id="WqYuhEM_mpP2"
# # Test
#
# + id="yguRvHVusbMi"
def predict_gt_motionDetect(reconstructed_sequences, sa,number):
threshold_anomaly =np.max(sa)/2
#print(threshold_anomaly)
test_reshaped = sequences
frames = []
threshs=[]
framedeltas = []
anomalies = []
original_frame = []
firstFrame = None
#cols_list=[0,1]
#pd.read_csv('/content/drive/MyDrive/prediction/ped1/Test024/sa.csv', usecols=cols_list)
for i in range(reconstructed_sequences.shape[0]):
#print(sa.iloc[i])
#i['var1'].iloc[0]
# print(type(sa))
# print(type(sa(0)))
# print(type(threshold_anomaly))
if (sa[i]<threshold_anomaly):
#print("hello")
text = "Normal"
for j in range(LENGTH_SEQUENCE):
anomaly = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
thresh = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
frameDelta = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
cv2.imwrite('image.png', reconstructed_sequences[i,j,:,:,:])
image = cv2.imread('image.png')
frame_test = np.array(test_reshaped[i,j,:,:,:],dtype=np.uint8)
frame_predicted = np.array(image,dtype=np.uint8)
frame = cv2.absdiff(frame_test,frame_predicted)
frame = cv2.GaussianBlur(frame, (5, 5), 0)
frame = imutils.resize(frame, width=500)
anomaly= imutils.resize(anomaly, width=500)
frame_test= imutils.resize(frame_test, width=500)
cv2.putText(frame, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
cv2.putText(frame_test, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
frames.append(frame)
anomalies.append(anomaly)
original_frame.append(frame_test)
threshs.append(thresh)
framedeltas.append(frameDelta)
firstFrame = None
else:
#print("else hello")
#firstFrame = None
cnt=0
for k in range(LENGTH_SEQUENCE):
text = "Normal"
anomaly = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
frame_test = np.array(test_reshaped[i,k,:,:,:],dtype=np.uint8)
#########################
cv2.imwrite('image.png', reconstructed_sequences[i,k,:,:,:])
#*255)
image = cv2.imread('image.png')
###########################
frame_predicted = np.array(image,dtype=np.uint8)
frame = cv2.absdiff(frame_test,frame_predicted)
frame = cv2.GaussianBlur(frame, (5, 5), 0)
frame = imutils.resize(frame, width=500)
anomaly= imutils.resize(anomaly, width=500)
frame_test= imutils.resize(frame_test, width=500)
gray = cv2.cvtColor(frame, cv2.COLOR_BGR2GRAY)
gray = cv2.GaussianBlur(gray, (21, 21), 0)
if firstFrame is None:
firstFrame = gray
frameDelta = cv2.absdiff(firstFrame, gray)
thresh = cv2.threshold(frameDelta, 29, 255, cv2.THRESH_BINARY)[1]
thresh = cv2.dilate(thresh, None, iterations=2)
cnts = cv2.findContours(thresh.copy(), cv2.RETR_EXTERNAL,
cv2.CHAIN_APPROX_SIMPLE)
cnts = imutils.grab_contours(cnts)
for c in cnts:
if cv2.contourArea(c) < 500:
continue
(x, y, w, h) = cv2.boundingRect(c)
cv2.rectangle(frame, (x, y), (x + w, y + h), (0, 255, 0), 2)
cv2.rectangle(anomaly, (x, y), (x + w, y + h), (255, 255, 255), -1)
cv2.rectangle(frame_test, (x, y), (x + w, y + h), (0, 255, 0), 2)
text = "Anomaly detected"
cv2.putText(frame, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
cv2.putText(frame_test, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
frames.append(frame)
threshs.append(thresh)
framedeltas.append(frameDelta)
anomalies.append(anomaly)
original_frame.append(frame_test)
path = '/content/drive/MyDrive/motion_detection/Ped1/Test'+str(number).zfill(3)+'/'
os.makedirs(os.path.dirname(path), exist_ok=True)
imageio.mimsave(path+'frames.gif', [frames[idx] for idx in range(len(frames))], fps=1)
imageio.mimsave(path+'frames threshs.gif', [threshs[idx] for idx in range(len(threshs))], fps=1)
imageio.mimsave(path+'frames deltas.gif', [framedeltas[idx] for idx in range(len(framedeltas))], fps=1)
imageio.mimsave(path+'gt_predicted.gif', [anomalies[idx] for idx in range(len(anomalies))], fps=1)
imageio.mimsave(path+'original_frame.gif', [original_frame[idx] for idx in range(len(original_frame))], fps=1)
return(anomalies)
# + id="1I8d4T1sTxrC"
number = 24
# + id="ixyYcCUgTxrC"
test = get_single_test(number)
# + id="gqnM9VvwTxrD" colab={"base_uri": "https://localhost:8080/", "height": 331} outputId="7f9694e5-3f53-4bda-ba95-3614ec7a0c97"
predicted_sequences, sa= predict_test()
# + id="4N2eBmLrTxrD" colab={"base_uri": "https://localhost:8080/"} outputId="70892b9a-26eb-4926-a352-993dd1d91005"
sequences_pred = predicted_sequences*255
sequences_pred.shape
# + id="6Puh71NxTxrE"
test = get_single_test_MD(number)
# + id="3fZ2m1nKTxrF" colab={"base_uri": "https://localhost:8080/"} outputId="91ebf915-e257-405d-cbb9-10e172627cb9"
sequences = make_sequences(test*255)
sequences.shape
# + id="A8RILQTtTxrF"
predictions =predict_gt_motionDetect(sequences_pred,sa,number)
# + [markdown] id="JpD8fG8DTxrH"
# # Test on Peds2
# + id="QoBg6aC-TxrH"
DATASET_PATH1 ="UCSD_Anomaly_Dataset/UCSD_Anomaly_Dataset.v1p2/UCSDped2/Train"
TEST_PATH1 ="UCSD_Anomaly_Dataset/UCSD_Anomaly_Dataset.v1p2/UCSDped2/Test"
MODEL_PATH = './model/model_lstm_1datasets_diff_arch_256_nomax_peds2_2.hdf5'
# + id="zYakznvGTxrI"
def get_training_set():
files1 = sorted(glob.glob(DATASET_PATH2+'/*'))
train_set = []
for idx, filename in enumerate(files1):
train_files = sorted(glob.glob(filename+'/*'))
for idx_frame,file_frame in enumerate(train_files):
im = Image.open(file_frame)
im = im.resize((IMAGE_SIZE[0],IMAGE_SIZE[1])) #resize 256, 256
train_set.append(np.array(im, dtype=np.float32)/np.float(IMAGE_SIZE[0]))
train = np.asarray(train_set)
train = np.reshape(train,((-1,256,256,1)))
seq1 = make_sequences(train) ##num sequence 10
return(seq1)
# + id="ODHWBr07TxrI"
number = 2
# + id="cy2y8aoeTxrJ"
test = get_single_test(number)
# + id="64w52woaTxrJ" outputId="349c3346-691d-422b-907a-ae39c70fa9b6"
predicted_sequences, sa= predict_test()
# + id="fyLRUoOyTxrK"
def predict_gt_motionDetect(reconstructed_sequences, sa,number):
threshold_anomaly =np.max(sa)/2
test_reshaped = sequences
frames = []
threshs=[]
framedeltas = []
anomalies = []
original_frame = []
firstFrame = None
for i in range(reconstructed_sequences.shape[0]):
if sa[i]<threshold_anomaly:
text = "Normal"
for j in range(LENGTH_SEQUENCE):
anomaly = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
thresh = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
frameDelta = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
cv2.imwrite('image.png', reconstructed_sequences[i,j,:,:,:])
image = cv2.imread('image.png')
frame_test = np.array(test_reshaped[i,j,:,:,:],dtype=np.uint8)
frame_predicted = np.array(image,dtype=np.uint8)
frame = cv2.absdiff(frame_test,frame_predicted)
frame = cv2.GaussianBlur(frame, (5, 5), 0)
frame = imutils.resize(frame, width=500)
anomaly= imutils.resize(anomaly, width=500)
frame_test= imutils.resize(frame_test, width=500)
cv2.putText(frame, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
cv2.putText(frame_test, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
frames.append(frame)
anomalies.append(anomaly)
original_frame.append(frame_test)
threshs.append(thresh)
framedeltas.append(frameDelta)
firstFrame = None
else:
#firstFrame = None
cnt=0
for k in range(LENGTH_SEQUENCE):
text = "Normal"
anomaly = np.zeros(shape=[IMAGE_SIZE[0],IMAGE_SIZE[1], 3], dtype=np.uint8)
frame_test = np.array(test_reshaped[i,k,:,:,:],dtype=np.uint8)
#########################
cv2.imwrite('image.png', reconstructed_sequences[i,k,:,:,:])
#*255)
image = cv2.imread('image.png')
###########################
frame_predicted = np.array(image,dtype=np.uint8)
frame = cv2.absdiff(frame_test,frame_predicted)
frame = cv2.GaussianBlur(frame, (5, 5), 0)
frame = imutils.resize(frame, width=500)
anomaly= imutils.resize(anomaly, width=500)
frame_test= imutils.resize(frame_test, width=500)
gray = cv2.cvtColor(frame, cv2.COLOR_BGR2GRAY)
gray = cv2.GaussianBlur(gray, (21, 21), 0)
if firstFrame is None:
firstFrame = gray
frameDelta = cv2.absdiff(firstFrame, gray)
thresh = cv2.threshold(frameDelta, 50, 255, cv2.THRESH_BINARY)[1]
thresh = cv2.dilate(thresh, None, iterations=2)
cnts = cv2.findContours(thresh.copy(), cv2.RETR_EXTERNAL,
cv2.CHAIN_APPROX_SIMPLE)
cnts = imutils.grab_contours(cnts)
for c in cnts:
if cv2.contourArea(c) < 250:
continue
(x, y, w, h) = cv2.boundingRect(c)
cv2.rectangle(frame, (x, y), (x + w, y + h), (0, 255, 0), 2)
cv2.rectangle(anomaly, (x, y), (x + w, y + h), (255, 255, 255), -1)
cv2.rectangle(frame_test, (x, y), (x + w, y + h), (0, 255, 0), 2)
text = "Anomaly detected"
cv2.putText(frame, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
cv2.putText(frame_test, "Status: {}".format(text), (10, 20),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, (255, 0, 0), 2)
frames.append(frame)
threshs.append(thresh)
framedeltas.append(frameDelta)
anomalies.append(anomaly)
original_frame.append(frame_test)
path = './motion detection/Peds2/Test'+str(number).zfill(3)+'/'
os.makedirs(os.path.dirname(path), exist_ok=True)
imageio.mimsave(path+'frames.gif', [frames[idx] for idx in range(len(frames))], fps=1)
imageio.mimsave(path+'frames threshs.gif', [threshs[idx] for idx in range(len(threshs))], fps=1)
imageio.mimsave(path+'frames deltas.gif', [framedeltas[idx] for idx in range(len(framedeltas))], fps=1)
imageio.mimsave(path+'gt_predicted.gif', [anomalies[idx] for idx in range(len(anomalies))], fps=1)
imageio.mimsave(path+'original_frame.gif', [original_frame[idx] for idx in range(len(original_frame))], fps=1)
return(anomalies)
# + id="ZVZ7lhsdTxrK"
#### Save the predicted seqeunces
reshaped = np.reshape(predicted_sequences, ((predicted_sequences.shape[0]*LENGTH_SEQUENCE,IMAGE_SIZE[0],IMAGE_SIZE[1],1)))
for i in range(180):
path = './predictions/Peds2/Test'+str(number).zfill(3)+'/'+str(i+1).zfill(3)+'.png'
os.makedirs(os.path.dirname(path), exist_ok=True)
cv2.imwrite(path, reshaped[i,:,:,0]*255)
sa = pd.DataFrame(sa, columns=['sa'])
path = './predictions/Peds2/Test'+str(number).zfill(3)+'/sa.csv'
os.makedirs(os.path.dirname(path), exist_ok=True)
sa.to_csv(path)
sa = np.array(sa)
# + id="A2rW9ya0TxrL" outputId="4888a194-ffa1-4aee-9f10-671e0eebb0b9"
sequences_pred = predicted_sequences*255
sequences_pred.shape
# + id="yYassdN-TxrL"
test = get_single_test_MD(number)
# + id="8D9l1hGWTxrM" outputId="fa726263-6f4b-4e38-9e41-d2ea1d274b81"
sequences = make_sequences(test*255)
sequences.shape
# + id="HQoZJEy5TxrN"
predictions =predict_gt_motionDetect(sequences_pred,sa,number)
# + id="Ok5a-kcJTxrN" outputId="46a5b001-d1e1-4dc6-83dc-d1fa7931ab0f"
IoU, accuracy = evaluate_accuracy_image_cv(number,predictions)
print('IoU is %s' % IoU)
print('Accuracy is %s' % accuracy)
# + id="nXGPBiGaTxrO" outputId="1634da9b-0643-4422-a07c-53bfecd4627b"
classic_accuracy(number, predictions)
# + id="E0WtJA_HTxrP"
| Anomaly_detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Stationarity and detrending (ADF/KPSS)
#
# Stationarity means that the statistical properties of a time series i.e. mean, variance and covariance do not change over time. Many statistical models require the series to be stationary to make effective and precise predictions.
#
# Two statistical tests would be used to check the stationarity of a time series – <NAME> (“ADF”) test and Kwiatkowski-Phillips-Schmidt-Shin (“KPSS”) test. A method to convert a non-stationary time series into stationary series shall also be used.
# This first cell imports standard packages and sets plots to appear inline.
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import statsmodels.api as sm
# Sunspots dataset is used. It contains yearly (1700-2008) data on sunspots from the National Geophysical Data Center.
sunspots = sm.datasets.sunspots.load_pandas().data
# Some preprocessing is carried out on the data. The "YEAR" column is used in creating index.
sunspots.index = pd.Index(sm.tsa.datetools.dates_from_range("1700", "2008"))
del sunspots["YEAR"]
# The data is plotted now.
sunspots.plot(figsize=(12, 8))
# ## ADF test
#
# ADF test is used to determine the presence of unit root in the series, and hence helps in understand if the series is stationary or not. The null and alternate hypothesis of this test are:
#
# Null Hypothesis: The series has a unit root.
#
# Alternate Hypothesis: The series has no unit root.
#
# If the null hypothesis in failed to be rejected, this test may provide evidence that the series is non-stationary.
#
# A function is created to carry out the ADF test on a time series.
# +
from statsmodels.tsa.stattools import adfuller
def adf_test(timeseries):
print("Results of Dickey-Fuller Test:")
dftest = adfuller(timeseries, autolag="AIC")
dfoutput = pd.Series(
dftest[0:4],
index=[
"Test Statistic",
"p-value",
"#Lags Used",
"Number of Observations Used",
],
)
for key, value in dftest[4].items():
dfoutput["Critical Value (%s)" % key] = value
print(dfoutput)
# -
# ## KPSS test
#
# KPSS is another test for checking the stationarity of a time series. The null and alternate hypothesis for the KPSS test are opposite that of the ADF test.
#
# Null Hypothesis: The process is trend stationary.
#
# Alternate Hypothesis: The series has a unit root (series is not stationary).
#
# A function is created to carry out the KPSS test on a time series.
# +
from statsmodels.tsa.stattools import kpss
def kpss_test(timeseries):
print("Results of KPSS Test:")
kpsstest = kpss(timeseries, regression="c", nlags="auto")
kpss_output = pd.Series(
kpsstest[0:3], index=["Test Statistic", "p-value", "Lags Used"]
)
for key, value in kpsstest[3].items():
kpss_output["Critical Value (%s)" % key] = value
print(kpss_output)
# -
# The ADF tests gives the following results – test statistic, p value and the critical value at 1%, 5% , and 10% confidence intervals.
#
# ADF test is now applied on the data.
adf_test(sunspots["SUNACTIVITY"])
# Based upon the significance level of 0.05 and the p-value of ADF test, the null hypothesis can not be rejected. Hence, the series is non-stationary.
# The KPSS tests gives the following results – test statistic, p value and the critical value at 1%, 5% , and 10% confidence intervals.
#
# KPSS test is now applied on the data.
kpss_test(sunspots["SUNACTIVITY"])
# Based upon the significance level of 0.05 and the p-value of KPSS test, there is evidence for rejecting the null hypothesis in favor of the alternative. Hence, the series is non-stationary as per the KPSS test.
#
# It is always better to apply both the tests, so that it can be ensured that the series is truly stationary. Possible outcomes of applying these stationary tests are as follows:
#
# Case 1: Both tests conclude that the series is not stationary - The series is not stationary
# Case 2: Both tests conclude that the series is stationary - The series is stationary
# Case 3: KPSS indicates stationarity and ADF indicates non-stationarity - The series is trend stationary. Trend needs to be removed to make series strict stationary. The detrended series is checked for stationarity.
# Case 4: KPSS indicates non-stationarity and ADF indicates stationarity - The series is difference stationary. Differencing is to be used to make series stationary. The differenced series is checked for stationarity.
#
# Here, due to the difference in the results from ADF test and KPSS test, it can be inferred that the series is trend stationary and not strict stationary. The series can be detrended by differencing or by model fitting.
# ## Detrending by Differencing
#
# It is one of the simplest methods for detrending a time series. A new series is constructed where the value at the current time step is calculated as the difference between the original observation and the observation at the previous time step.
#
# Differencing is applied on the data and the result is plotted.
sunspots["SUNACTIVITY_diff"] = sunspots["SUNACTIVITY"] - sunspots["SUNACTIVITY"].shift(
1
)
sunspots["SUNACTIVITY_diff"].dropna().plot(figsize=(12, 8))
# ADF test is now applied on these detrended values and stationarity is checked.
adf_test(sunspots["SUNACTIVITY_diff"].dropna())
# Based upon the p-value of ADF test, there is evidence for rejecting the null hypothesis in favor of the alternative. Hence, the series is strict stationary now.
# KPSS test is now applied on these detrended values and stationarity is checked.
kpss_test(sunspots["SUNACTIVITY_diff"].dropna())
# Based upon the p-value of KPSS test, the null hypothesis can not be rejected. Hence, the series is stationary.
# ## Conclusion
#
# Two tests for checking the stationarity of a time series are used, namely ADF test and KPSS test. Detrending is carried out by using differencing. Trend stationary time series is converted into strict stationary time series. Requisite forecasting model can now be applied on a stationary time series data.
| examples/notebooks/stationarity_detrending_adf_kpss.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Self-Driving Car Engineer Nanodegree
#
# ## Deep Learning
#
# ## Project: Build a Traffic Sign Recognition Classifier
#
# ### MY Testing Environments
#
# 1) CPU: Intel Core i7-7700K (4.6Ghz OC) 4 CPU CORES, 8 threads
#
# 2) RAM: 16GB DDR4
#
# 3) GPU: nVidia GeForce GTX 970 4GB GGDR5
#
# 4) OS: WINDOWS 10 PRO 64bit
#
# 5) nVidia Software: Nvidia CUDA and cuDNN V 5.1
#
# ## Step 0: Load The Data
# #### import all necessary libraries in one go
# +
# Imports all libraries required
import os
import cv2
import csv
import time
import pickle
import numpy as np
import pandas as pd
import seaborn as sns
import tensorflow as tf
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from PIL import Image
from pylab import rcParams
from skimage import transform
from sklearn.utils import shuffle
from sklearn import preprocessing
from sklearn.metrics import confusion_matrix
from tensorflow.contrib.layers import flatten
from sklearn.preprocessing import OneHotEncoder
from sklearn.model_selection import train_test_split
# %matplotlib inline
# -
# #### some unexpected errors are present
# +
# Imports all libraries required
import os
import cv2
import csv
import time
import pickle
import numpy as np
import pandas as pd
import seaborn as sns
import tensorflow as tf
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from PIL import Image
from pylab import rcParams
from skimage import transform
from sklearn.utils import shuffle
from sklearn import preprocessing
from sklearn.metrics import confusion_matrix
from tensorflow.contrib.layers import flatten
from sklearn.preprocessing import OneHotEncoder
from sklearn.model_selection import train_test_split
# %matplotlib inline
# -
# #### executing the same codes again removes the errors, not sure why!!
# #### Loading pickled data as instructed in the TEMPLATE Jupyter Notebook
# +
# Load pickled data
training_file = 'train.p'
validating_file = 'valid.p'
testing_file = 'test.p'
with open(training_file, mode='rb') as f:train = pickle.load(f)
with open(validating_file, mode='rb') as f:valid = pickle.load(f)
with open(testing_file, mode='rb') as f:test = pickle.load(f)
X_train, y_train = train['features'], train['labels']
X_valid, y_valid = valid['features'], valid['labels']
X_test, y_test = test['features'], test['labels']
# -
# ---
#
# ## Step 1: Dataset Summary & Exploration
#
# The pickled data is a dictionary with 4 key/value pairs:
#
# - `'features'` is a 4D array containing raw pixel data of the traffic sign images, (num examples, width, height, channels).
# - `'labels'` is a 2D array containing the label/class id of the traffic sign. The file `signnames.csv` contains id -> name mappings for each id.
# - `'sizes'` is a list containing tuples, (width, height) representing the the original width and height the image.
# - `'coords'` is a list containing tuples, (x1, y1, x2, y2) representing coordinates of a bounding box around the sign in the image. **THESE COORDINATES ASSUME THE ORIGINAL IMAGE. THE PICKLED DATA CONTAINS RESIZED VERSIONS (32 by 32) OF THESE IMAGES**
#
# Complete the basic data summary below.
# +
### Replace each question mark with the appropriate value.
# TODO: Number of training examples
n_train = len(X_train)
n_valid = len(X_valid)
# TODO: Number of testing examples.
n_test = len(X_test)
# TODO: What's the shape of an traffic sign image?
image_shape = X_train[0].shape
# TODO: How many unique classes/labels there are in the dataset.
n_classes = len(set(y_train))
print("Number of training examples =", n_train)
print("Number of validating examples =", n_valid)
print("Number of testing examples =", n_test)
print("Image data shape =", image_shape)
print("Number of classes =", n_classes)
# -
# Visualize the German Traffic Signs Dataset using the pickled file(s). This is open ended, suggestions include: plotting traffic sign images, plotting the count of each sign, etc.
#
# The [Matplotlib](http://matplotlib.org/) [examples](http://matplotlib.org/examples/index.html) and [gallery](http://matplotlib.org/gallery.html) pages are a great resource for doing visualizations in Python.
#
# **NOTE:** It's recommended you start with something simple first. If you wish to do more, come back to it after you've completed the rest of the sections.
# #### Display randomly 10 images from the training set.
# +
### Data exploration visualization
fig = plt.figure(figsize=(15, 5))
image_seq = np.random.randint(1,len(X_train),10)
# Load image labels from csv
label_csv = csv.reader(open('signnames.csv', 'r'))
label_names = []
for row in label_csv:
label_names.append(row[1])
label_names.pop(0)
for ind,val in enumerate(image_seq):
img = fig.add_subplot(2,5,ind+1)
plt.imshow(X_train[val-1])
#Add corresponding label
img.set_xlabel("{0} ({1})".format(y_train[val-1], label_names[y_train[val-1]]))
#Remove the axis ticks
img.set_xticks([])
img.set_yticks([])
plt.show()
# -
# #### Create a histogram that depicts the overall dataset distribution for the training set.
# A= unique B = counts
A, B = np.unique(y_train, return_counts=True)
fig = plt.figure(figsize=(15,10))
plt.bar(A, B, color='green')
label = [label for label in label_names]
plt.xticks(np.arange(0.5,n_classes+0.5), label, rotation=45,ha='right')
plt.ylabel('Frequency')
plt.title('Training Data Distribution')
plt.show()
# ----
#
# ## Step 2: Design and Test a Model Architecture
#
# Design and implement a deep learning model that learns to recognize traffic signs. Train and test your model on the [German Traffic Sign Dataset](http://benchmark.ini.rub.de/?section=gtsrb&subsection=dataset).
#
# There are various aspects to consider when thinking about this problem:
#
# - Neural network architecture
# - Play around preprocessing techniques (normalization, rgb to grayscale, etc)
# - Number of examples per label (some have more than others).
# - Generate fake data.
#
# Here is an example of a [published baseline model on this problem](http://yann.lecun.com/exdb/publis/pdf/sermanet-ijcnn-11.pdf). It's not required to be familiar with the approach used in the paper but, it's good practice to try to read papers like these.
#
# **NOTE:** The LeNet-5 implementation shown in the [classroom](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) at the end of the CNN lesson is a solid starting point. You'll have to change the number of classes and possibly the preprocessing, but aside from that it's plug and play!
# ### Implementation
#
# Use the code cell (or multiple code cells, if necessary) to implement the first step of your project. Once you have completed your implementation and are satisfied with the results, be sure to thoroughly answer the questions that follow.
def preprocess(X):
# Normalize to range 0-1
X = (X - X.mean())/(np.max(X) - np.min(X))
# grayscale conversion
X = 0.114 * X[...,0] + 0.587 * X[...,1] + 0.299 * X[...,2] # BGR->Gray
return X
X_train = preprocess(X_train)
X_valid = preprocess(X_valid)
X_test = preprocess(X_test)
# ### Question 1
#
# _Describe how you preprocessed the data. Why did you choose that technique?_
# **Answer:**
#
# I encapsulate 2 operations within a single 'preprocess' function here.
#
# The first operation is normalisation using the simple formula X = (X - X.mean())/(np.max(X) - np.min(X)), the purpose of normalisation is to help the gradient descent optimizer (Adam Optimizer) to converge faster by restricting the range of feature values.
#
# The 2nd operation is grayscale conversion, which is supposed to be detrimental to the model performance, however, I was at a loss to explain why grayscale conversion turned out to be conducive to the test accuracy at the end.
#
# ### Question 2
#
# _Describe how you set up the training, validation and testing data for your model. **Optional**: If you generated additional data, how did you generate the data? Why did you generate the data? What are the differences in the new dataset (with generated data) from the original dataset?_
# **Answer:**
#
# Since I have 3 separate files for training, validation and testing respectively, it makes sense to avoid using training/testing split function here. train.p is used for training exclusively, valid.p is earmarked for validation during the training process, while the test.p is strictly reserved for testing purpose once the training is completed.
# +
layer_depth = {
'layer_1': 12,
'layer_2': 32,
'fully_connected_1': 512,
'fully_connected_2': 256,
'fully_connected_3': 128,
'out': n_classes,
}
weights = {
'layer_1': tf.Variable(tf.truncated_normal(
[5, 5, 1, layer_depth['layer_1']], mean=0, stddev=0.1)),
'layer_2': tf.Variable(tf.truncated_normal(
[5, 5, layer_depth['layer_1'], layer_depth['layer_2']], mean=0, stddev=0.1)),
'fully_connected_1': tf.Variable(tf.truncated_normal(
[5*5*layer_depth['layer_2'], layer_depth['fully_connected_1']])),
'fully_connected_2': tf.Variable(tf.truncated_normal(
[layer_depth['fully_connected_1'], layer_depth['fully_connected_2']], mean=0, stddev=0.1)),
'fully_connected_3': tf.Variable(tf.truncated_normal(
[layer_depth['fully_connected_2'], layer_depth['fully_connected_3']], mean=0, stddev=0.1)),
'out': tf.Variable(tf.truncated_normal(
[layer_depth['fully_connected_3'], layer_depth['out']], mean=0, stddev=0.1))
}
biases = {
'layer_1': tf.Variable(tf.zeros(layer_depth['layer_1'])),
'layer_2': tf.Variable(tf.zeros(layer_depth['layer_2'])),
'fully_connected_1': tf.Variable(tf.zeros(layer_depth['fully_connected_1'])),
'fully_connected_2': tf.Variable(tf.zeros(layer_depth['fully_connected_2'])),
'fully_connected_3': tf.Variable(tf.zeros(layer_depth['fully_connected_3'])),
'out': tf.Variable(tf.zeros(layer_depth['out']))
}
# +
# Define 2 more functions
def conv2d(x, W, b, strides=1):
x = tf.nn.conv2d(x, W, strides=[1, strides, strides, 1], padding = 'VALID')
x = tf.nn.bias_add(x, b)
return tf.nn.relu(x)
def maxpool2d(x, k=2):
return tf.nn.max_pool(x, ksize=[1, k, k, 1],
strides=[1, k, k, 1],
padding='VALID')
# +
# Define Architecture
keep_prob = tf.placeholder(tf.float32)
def LeNet(x):
x = tf.expand_dims(x, -1)
conv1 = conv2d(x, weights['layer_1'], biases['layer_1'])
conv1 = tf.nn.relu(conv1)
conv1 = maxpool2d(conv1)
#________________________________________________________________________________________
conv2 = conv2d(conv1, weights['layer_2'], biases['layer_2'])
conv2 = tf.nn.relu(conv2)
conv2 = maxpool2d(conv2)
#________________________________________________________________________________________
fc0 = flatten(conv2)
fc1 = tf.add(tf.matmul(fc0, weights['fully_connected_1']), biases['fully_connected_1'])
fc1 = tf.nn.relu(fc1)
fc1 = tf.nn.dropout(fc1, keep_prob=keep_prob)
#________________________________________________________________________________________
fc2 = tf.add(tf.matmul(fc1, weights['fully_connected_2']), biases['fully_connected_2'])
fc2 = tf.nn.relu(fc2)
fc2 = tf.nn.dropout(fc2, keep_prob=keep_prob)
#________________________________________________________________________________________
fc3 = tf.add(tf.matmul(fc2, weights['fully_connected_3']), biases['fully_connected_3'])
fc3 = tf.nn.relu(fc3)
fc3 = tf.nn.dropout(fc3, keep_prob=keep_prob)
logits = tf.add(tf.matmul(fc3, weights['out']), biases['out'])
return logits
# -
# ### Question 3
#
# _What does your final architecture look like? (Type of model, layers, sizes, connectivity, etc.) For reference on how to build a deep neural network using TensorFlow, see [Deep Neural Network in TensorFlow
# ](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/b516a270-8600-4f93-a0a3-20dfeabe5da6/concepts/83a3a2a2-a9bd-4b7b-95b0-eb924ab14432) from the classroom._
#
# **Answer:**
#
# The model is based on the LeNet Lab with the different number of Convolution layers and Fully connected layers.
# I have tried some other configurations such as one with just 2 full connected layers, however, I was unable to record any noticeable performance improvement.
#
# Several dropout values were tested from 0.3 all the way to 0.9, 0.6 seemed to fit the bill.
#
# Layer 1 : 5x5 Filter with depth 12
#
# Layer 2 : 5x5 Filter with depth 32
#
# Fully Connected Layer A : n = 512
# Dropout Layer : Dropout Value = 0.6
#
# Fully Connected Layer B : n = 256
# Dropout Layer : Dropout Value = 0.6
#
# Fully Connected Layer C: n = 128
# Dropout Layer : Dropout Value = 0.6
# +
saver = tf.train.Saver()
# Add placeholder for input and data labels
x = tf.placeholder(tf.float32, (None, 32, 32))
y = tf.placeholder(tf.int32, (None))
one_hot_y = tf.one_hot(y, n_classes)
# +
learning_rate = 0.0005
logits = LeNet(x)
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(logits=logits, labels=one_hot_y)
loss_operation = tf.reduce_mean(cross_entropy)
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate)
# training Without regulariztaion
training_operation = optimizer.minimize(loss_operation)
# +
# Evaluation
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1))
accuracy_tunning = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
def evaluate(X_data, y_data):
num_examples = len(X_data)
total_accuracy = 0
sess = tf.get_default_session()
for offset in range(0, num_examples, batch_size):
end = offset + batch_size
batch_x, batch_y = X_data[offset:end], y_data[offset:end]
accuracy = sess.run(accuracy_tunning, feed_dict={x: batch_x, y: batch_y, keep_prob: 1})
total_accuracy += (accuracy * len(batch_x))
accuracy = total_accuracy / num_examples
return accuracy
# -
epochs = 100
batch_size = 64
# +
# Run Training and save model
total_time = time.time()
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
num_examples = len(X_train)
print('Number of training samples: {}'.format(num_examples))
print('Training in progress......\n\n')
for i in range(epochs):
start_time = time.time()
X_train, y_train = shuffle(X_train, y_train)
for offset in range(0, num_examples, batch_size):
end = offset + batch_size
batch_x, batch_y = X_train[offset:end], y_train[offset:end]
sess.run(training_operation, feed_dict={x: batch_x, y: batch_y, keep_prob:0.6})
validation_accuracy = evaluate(X_valid, y_valid)
validation_percent = validation_accuracy*100
print("\nEPOCH {} ...".format(i+1))
print("Validation Accuracy = {:.3f}%".format(validation_percent))
end_time = time.time() - start_time
print("Time taken for the last epoch: %.3f seconds" %end_time)
test_accuracy = evaluate(X_test, y_test)
test_percent = test_accuracy*100
print("\n\n\nAccuracy compared to test set = {:.3f}%".format(test_percent))
final_time = time.time() - total_time
print("Total Training: %.3f seconds" %final_time)
saver.save(sess, '.\model')
print('Model successfully Saved to current directory!')
# -
# roload and test the Model
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
test_accuracy = evaluate(X_test, y_test)
test_percent = test_accuracy*100
print("Test Accuracy = {:.3f}%".format(test_percent))
# ### Question 4
#
# _How did you train your model? (Type of optimizer, batch size, epochs, hyperparameters, etc.)_
#
# **Answer:**
#
# The reason why I chose AdamOptimizer is that it is more both quicker and more accurate than a standard stochastic gradient descent optimizer. In addition, I decided to set a relatively large epoch counts of 100 and a small batch size of 64 as I could rely on my powerful nVidia GTX970 GPU to complete the task efficiently. A rather ambitious learning rate of 0.0005 was also chosen. The codes responsible for training is in cell number 19 while preparations for the training took place in cell 14 to 18.
#
# At the end of the training, the training model was saved to three files with prefix 'model'.
#
# The validation accuracy you observed in the result sheet above referred to the accuracy of the training model when it was compared with the data located in a separate file called 'valid.p'. No training-testing data splitting was required.
#
# The test accuracy at the end of the result sheet above referred to the accuracy of the training model when it was compared with the data located in a separate file called 'test.p'
#
# The final test accuracy stood between 95% and 96% after several tests, which was quite satisfactory I thought. However, the validation accuracy kinda plateaued at around epoch 10! Which is good to know!
#
#
#
# ### Question 5
#
#
# _What approach did you take in coming up with a solution to this problem? It may have been a process of trial and error, in which case, outline the steps you took to get to the final solution and why you chose those steps. Perhaps your solution involved an already well known implementation or architecture. In this case, discuss why you think this is suitable for the current problem._
# **Answer:**
#
#
#
# The Convnet LeNet Lab template in the nanodegree course is a solid convolutional neural network upon which my implementation is largely based. I tried various configurations: 1 conv + 1 FC layers, 1 conv + 2 FC layers, 2 conv + 1 FC layers, 2 conv + 2 FC layers, 3 conv + 2 FC (This one didn't work at all due to dimension errors) and 2 conv + 3 FC layers which was the one I chosen for the final showdown.
#
# 1 conv + 1 FC, 1 conv + 2 FC and 2 conv + 1 FC all had significantly worse accuracies than 2 conv + 2 FC and 2 conv + 3 FC.
#
# The 2 conv + 2 FC layers configuration has a very similar performance to that of the 2 conv + 3 FC layers configuration. In the hindsight, maybe I should've chosen the 2 conv + 2 FC one as GPU would have had less work to do but I just wanted to stretch its muscle a bit, lol.
#
#
#
# ---
#
# ## Step 3: Test a Model on New Images
#
# Take several pictures of traffic signs that you find on the web or around you (at least five), and run them through your classifier on your computer to produce example results. The classifier might not recognize some local signs but it could prove interesting nonetheless.
#
# You may find `signnames.csv` useful as it contains mappings from the class id (integer) to the actual sign name.
# ### Implementation
#
# Use the code cell (or multiple code cells, if necessary) to implement the first step of your project. Once you have completed your implementation and are satisfied with the results, be sure to thoroughly answer the questions that follow.
# #### Testing on German Traffic Signs
# +
# load up new test images
df = pd.read_csv('signnames.csv')
import glob
images_resized = []
images = []
for j in glob.glob('./extra_German_sign/*.jpg'):
image = plt.imread(j)
image_resized = cv2.resize(image, (32, 32), interpolation=cv2.INTER_AREA)
images_resized.append(image_resized)
image_preprocessed = preprocess(image_resized)
images.append(image_preprocessed[np.newaxis,...])
images = np.vstack(images)
# -
with tf.Session() as sess:
new_saver = tf.train.import_meta_graph('model.meta')
new_saver.restore(sess, tf.train.latest_checkpoint('./'))
out = sess.run(tf.argmax(logits, 1), feed_dict={x: images, keep_prob: 1})
# +
# Plot Images with prediction
new_label_list = [np.argmax(row) for row in out]
plt.figure(figsize=(12,12))
for i in range(0,images.shape[0]):
with sns.axes_style("white"):
plt.subplot(4, 4, i+1)
plt.imshow(np.squeeze(images_resized[i]), cmap='gray')
plt.tick_params(axis='both', which='both', bottom='on', top='on', labelbottom='off', right='off', left='off', labelleft='off')
plt.xlabel(df.loc[out[i]].SignName)
plt.tight_layout()
# -
# ### Question 6
#
# _Choose five candidate images of traffic signs and provide them in the report. Are there any particular qualities of the image(s) that might make classification difficult? It could be helpful to plot the images in the notebook._
# **Answer:**
#
# I can think of several problems that can make the life extremely difficult for the model.
#
# 1) The orientation of the traffic signs.
#
# 2) The actual clarity of the traffic signs.
#
# 3) The angles and perspective at which the photos were taken.
#
# 4) Multiple signs in one image.
# ### Question 7
#
# _Is your model able to perform equally well on captured pictures when compared to testing on the dataset? The simplest way to do this check the accuracy of the predictions. For example, if the model predicted 1 out of 5 signs correctly, it's 20% accurate._
#
# _**NOTE:** You could check the accuracy manually by using `signnames.csv` (same directory). This file has a mapping from the class id (0-42) to the corresponding sign name. So, you could take the class id the model outputs, lookup the name in `signnames.csv` and see if it matches the sign from the image._
#
# **Answer:**
#
# The model correctly guessed the signs in 1st, 8th,12th, 13th,14th(partially) image, giving an accuracy of 5/15 = 33.3%, which is significantly worse than the test accuracy obtained earlier! Please correct me if I'm wrong as I don't drive due to medical conditions.
k = 5
with tf.Session() as sess:
new_saver = tf.train.import_meta_graph('model.meta')
new_saver.restore(sess, tf.train.latest_checkpoint('./'))
out_prob = sess.run(tf.nn.top_k(tf.nn.softmax(logits), k=k), feed_dict={x: images, keep_prob: 1})
# +
plt.rcParams['figure.figsize'] = (15, 30)
image_indices = (0,1,2,3,4,5,6,7,8,9,10,11,12,13,14)
#image_indices = np.arange(0, len(images_resized))
for i, im in enumerate(image_indices):
with sns.axes_style("white"):
plt.subplot(len(image_indices), 2, (2*i)+1)
plt.imshow(np.squeeze(images_resized[im]), cmap='gray')
plt.axis('on')
plt.xlabel(df.loc[out[i]].SignName)
plt.subplot(len(image_indices) ,2, (2*i)+2)
plt.barh(np.arange(k), out_prob.values[im])
plt.yticks(np.arange(k)+0.3, df.loc[out_prob.indices[im]].SignName)
plt.tight_layout()
# -
# ### Question 8
#
# *Use the model's softmax probabilities to visualize the **certainty** of its predictions, [`tf.nn.top_k`](https://www.tensorflow.org/versions/r0.12/api_docs/python/nn.html#top_k) could prove helpful here. Which predictions is the model certain of? Uncertain? If the model was incorrect in its initial prediction, does the correct prediction appear in the top k? (k should be 5 at most)*
#
# `tf.nn.top_k` will return the values and indices (class ids) of the top k predictions. So if k=3, for each sign, it'll return the 3 largest probabilities (out of a possible 43) and the correspoding class ids.
#
# Take this numpy array as an example:
#
# ```
# # (5, 6) array
# a = np.array([[ 0.24879643, 0.07032244, 0.12641572, 0.34763842, 0.07893497,
# 0.12789202],
# [ 0.28086119, 0.27569815, 0.08594638, 0.0178669 , 0.18063401,
# 0.15899337],
# [ 0.26076848, 0.23664738, 0.08020603, 0.07001922, 0.1134371 ,
# 0.23892179],
# [ 0.11943333, 0.29198961, 0.02605103, 0.26234032, 0.1351348 ,
# 0.16505091],
# [ 0.09561176, 0.34396535, 0.0643941 , 0.16240774, 0.24206137,
# 0.09155967]])
# ```
#
# Running it through `sess.run(tf.nn.top_k(tf.constant(a), k=3))` produces:
#
# ```
# TopKV2(values=array([[ 0.34763842, 0.24879643, 0.12789202],
# [ 0.28086119, 0.27569815, 0.18063401],
# [ 0.26076848, 0.23892179, 0.23664738],
# [ 0.29198961, 0.26234032, 0.16505091],
# [ 0.34396535, 0.24206137, 0.16240774]]), indices=array([[3, 0, 5],
# [0, 1, 4],
# [0, 5, 1],
# [1, 3, 5],
# [1, 4, 3]], dtype=int32))
# ```
#
# Looking just at the first row we get `[ 0.34763842, 0.24879643, 0.12789202]`, you can confirm these are the 3 largest probabilities in `a`. You'll also notice `[3, 0, 5]` are the corresponding indices.
#
#
# **The model is most certain about the following:**
#
# (predicted: bumpy road, actual: bumpy road) correctly classified
#
# (predicted: priority road, actual:slope of gradient of 8%)
# understandably a very tough sign to classify as not many roads share the same slope. The correct label is NOT in the top 5 most likely classifications.
#
# (predicted:Road work, actual:Road work)
# Correctly classified.
#
#
# (predicted: dangerous curve to the right, actual:motor vehicle prohibited)
# no idea why the model is so far off.The correct label is NOT in the top 5 most likely classifications.
#
# (predicted: End of no passing, actual: diversion) The correct label is NOT in the top 5 most likely classifications.
#
#
# (predicted: no passing, actual: overtaking allowed)
# grayscale conversion is definitely answerable for the wrong prediction here. The correct label is NOT in the top 5 most likely classifications.
#
#
#
# (predicted:right-of-way at the next intersection, actual: speed limit of 60km/h) The correct label is NOT in the top 5 most likely classifications.
#
# (predicted: speed limit(50km/h), actual: motor vehicle prohibited)
# This prediction makes absolutely no sense. The correct label is NOT in the top 5 most likely classifications.
#
#
# (predicted: no entry, actual: no entry) correctly classified
#
#
# (predicted: priority road, actual: priority road) correctly classified
#
#
# (predicted: road narrow on the right, actual: road narrow on both side) partially correctly classified, The correct label is NOT in the top 5 most likely classifications.
#
# (predicted: keep right, actual: 30km/h zone) The correct label is NOT in the top 5 most likely classifications.
#
# **The model is most uncertain about:**
#
# (predicted:bicyle crossing, actual:wild animal crossing)
# incorrectly classified, however, the correct label is in the top 5 most likely classifications.
#
# (predicted: roundabout mandatory, actual: bicycle crossing)
# The correct label is NOT in the top 5 most likely classifications.
#
# (predicted: speed limits(50km), actual: roundabout mandatory)
# incorrectly classified, however, the correct label is in the top 5 most likely classifications.
# ## THANK YOU FOR VIEWING.
# > **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to \n",
# "**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
# ## reference:
#
# #### https://www.tensorflow.org/
# #### https://https://github.com/
# #### https://developer.nvidia.com/
#
#
# #### graveyard functions (please ignore)
'''
plot_loss_accuracy(batches, loss_batch, train_acc_batch, valid_acc_batch)
'''
'''
if not offset % 50:
# Calculate Training and Validation accuracy
training_accuracy = sess.run(accuracy_tunning, feed_dict={x: X_train,
y: y_train, keep_prob: 0.8 })
validation_accuracy = sess.run(accuracy_tunning, feed_dict={x: X_valid,
y: y_valid, keep_prob: 1})
# Log batches
previous_batch = batches[-1] if batches else 0
batches.append(50 + previous_batch)
loss_batch.append(c)
train_acc_batch.append(training_accuracy)
valid_acc_batch.append(validation_accuracy)
'''
| Term_1/Project_2_Traffic_Sign_Classification/Traffic_Sign_Classifier_FINAL_submission.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Building Gene Regulatory Networks with BioCRNpyler
# ### _<NAME>_
#
# In this notebook, we will use RegulatedPromoter and CombinatorialPromoter to model Gene Regulation via Transcription Factors.
#
# _Note: For this notebook to function, you need the default_parameter.txt file in the same folder as the notebook._
# + [markdown] slideshow={"slide_type": "slide"}
# # Modeling Gene Regulation by a Single Transcription Factor
# In biology, it is very common for a transcription factor to turn on or off promoter. BioCRNpyler has a number of Promoter Components to do this.
# * RegulatedPromoter: A list of regulators each bind individually to turn a Promoter ON or OFF (No Combinatorial Logic)
# * RepressiblePromoter: A single repressor modelled with a hill function
# * ActivatablePromoter: A single activator modeled with a hill function
#
# In the next example, we will produce and compare the outputs of these kinds of promoters.
# + [markdown] slideshow={"slide_type": "slide"}
# # Example 1: ActivatablePromoter
# A very simple Promoter Component modelled with a hill function. However, this class is not able to accurately capture the binding of Machinery like RNAP and shouldn't be used with Mixtures that include machinery.
# + slideshow={"slide_type": "skip"} tags=[]
from biocrnpyler import *
#ActivatedPromoter Example
activator = Species("activator", material_type = "small_molecule")
S_A = Species("A")
#Create a custom set of parameters
hill_parameters = {"k":1.0, "n":4, "K":20, "kleak":.01}
#By Loading custom parameters into the promoter, we override the default parameters of the Mixture
P_activatable = ActivatablePromoter("P_activtable", activator = activator, leak = True, parameters = hill_parameters)
#Create a DNA assembly "reporter" with P_activatable for its promoter
activatable_assembly = DNAassembly(name="activatable_assembly", promoter=P_activatable, rbs="Strong", protein = S_A)
M = SimpleTxTlExtract(name="SimpleTxTl", parameter_file = "default_parameters.txt", components=[activatable_assembly])
CRN = M.compile_crn();
print(CRN.pretty_print(show_rates = True, show_keys = True))
# + slideshow={"slide_type": "skip"} tags=[]
#Titrate the activator and plot the result
try:
# %matplotlib inline
from biocrnpyler import *
import numpy as np
import pylab as plt
import pandas as pd
for a_c in np.linspace(0, 50, 5):
x0 = {activatable_assembly.dna:1, activator:a_c}
timepoints = np.linspace(0, 100, 100)
R = CRN.simulate_with_bioscrape_via_sbml(timepoints, initial_condition_dict = x0)
plt.plot(R["time"], R[str(S_A)], label = "[activator]="+str(a_c))
plt.ylabel(f"[{S_A}]")
plt.legend()
except ModuleNotFoundError:
print('please install the plotting libraries: pip install biocrnpyler[all]')
# + [markdown] slideshow={"slide_type": "slide"}
# # Example 2: RepressiblePromoter
# A very simple Promoter Component modelled with a hill function. However, this class is not able to accurately capture the binding of Machinery like RNAP and shouldn't be used with Mixtures that include machinery.
# + slideshow={"slide_type": "subslide"} tags=[]
#ActivatedPromoter Example
repressor = S_A #defined in the previous example
reporter = Species("reporter", material_type = "protein")
#Create a custom set of parameters
hill_parameters = {"k":1.0, "n":4, "K":20, "kleak":.01}
#By Loading custom parameters into the promoter, we override the default parameters of the Mixture
P_repressible = RepressiblePromoter("P_repressible", repressor = repressor, leak = True, parameters = hill_parameters)
#Create a DNA assembly "reporter" with P_activatable for its promoter
repressible_assembly = DNAassembly(name="reporter", promoter=P_repressible, rbs="Strong", protein = reporter)
M = SimpleTxTlExtract(name="SimpleTxTl", parameter_file = "default_parameters.txt", components=[repressible_assembly])
CRN = M.compile_crn()
print(CRN.pretty_print(show_rates = True, show_keys = True))
# + slideshow={"slide_type": "skip"}
#Titrate the repressor and plot the result
try:
import biocrnpyler
import numpy as np
import pylab as plt
import pandas as pd
for r_c in np.linspace(0, 50, 5):
x0 = {repressible_assembly.dna:1, repressor:r_c}
timepoints = np.linspace(0, 100, 100)
R = CRN.simulate_with_bioscrape_via_sbml(timepoints, initial_condition_dict = x0)
plt.plot(R["time"], R[str(reporter)], label = f"[{str(S_A)}]={r_c}")
plt.ylabel("[B]")
plt.legend()
except ModuleNotFoundError:
print('please install the plotting libraries: pip install biocrnpyler[all]')
# + [markdown] slideshow={"slide_type": "slide"}
# # Example 3: A Simple Genetic Regulatory Network
# In this example, the activatable_assembly will produce a repressor that represses the repressable_assembly. Notice that activatable_assembly already procues the repressor of the RepressablePromoter...so this is easy!
# + slideshow={"slide_type": "fragment"} tags=[]
M = SimpleTxTlExtract(name="SimpleTxTl", parameter_file = "default_parameters.txt", components=[repressible_assembly, activatable_assembly])
CRN = M.compile_crn()
print(CRN.pretty_print(show_rates = True, show_keys = False))
#Titrate the activator, which in turn will automatically produce the repressor
try:
import biocrnpyler
import numpy as np
import pylab as plt
import pandas as pd
plt.figure(figsize = (10, 6))
ax1, ax2 = plt.subplot(121), plt.subplot(122)#Create two subplots
for a_c in np.linspace(0, 50, 5):
x0 = {activatable_assembly.dna:1, repressible_assembly.dna:1, activator:a_c}
timepoints = np.linspace(0, 100, 100)
R = CRN.simulate_with_bioscrape_via_sbml(timepoints, initial_condition_dict = x0)
plt.sca(ax1)
plt.plot(R["time"], R[str(S_A)], label = "[activator]="+str(a_c))
plt.sca(ax2)
plt.plot(R["time"], R[str(reporter)], label = "[activator]="+str(a_c))
plt.sca(ax1)
plt.ylabel("activatable assembly output: [A]")
plt.legend()
plt.sca(ax2)
plt.ylabel("repressable assembly output: [B]")
plt.legend()
except ModuleNotFoundError:
print('please install the plotting libraries: pip install biocrnpyler[all]')
# + [markdown] slideshow={"slide_type": "slide"}
# # Example 4: RegulatedPromoter
# In the below example, a CRN from RegulatedPromoter is generated. This Component models the detailed binding of regulators to the DNA and has seperate transcription rates for each regulator. It is suitable for complex models that include machinery. Regulators do not act Combinatorically.
# + slideshow={"slide_type": "subslide"} tags=[]
#1 Regulated Promoter Needs lots of parameters!
component_parameters = {
#Promoter Activator Binding Parameters. Note the part_id = [promoter_name]_[regulator_name]
ParameterKey(mechanism = 'binding', part_id = 'regulated_promoter_A', name = 'kb'):100, #Promoter - Activator Binding
ParameterKey(mechanism = 'binding', part_id = "regulated_promoter_A", name = 'ku'):5.0, #Unbinding
ParameterKey(mechanism = 'binding', part_id = "regulated_promoter_A", name = 'cooperativity'):4.0, #Cooperativity
#Activated Promoter Transcription. Note the part_id = [promoter_name]_[regulator_name]
#These regulate RNAP binding to an activated promoter and transcription
ParameterKey(mechanism = 'transcription', part_id = 'regulated_promoter_A', name = 'kb'):100, #Promoter - Activator Binding
ParameterKey(mechanism = 'transcription', part_id = "regulated_promoter_A", name = 'ku'):1.0, #Unbinding
ParameterKey(mechanism = 'transcription', part_id = 'regulated_promoter_A', name = "ktx"): 1., #Transcription Rate
#Promoter Repressor Binding Parameters. Note the part_id = [promoter_name]_[regulator_name]
ParameterKey(mechanism = 'binding', part_id = 'regulated_promoter_R', name = 'kb'):100,
ParameterKey(mechanism = 'binding', part_id = "regulated_promoter_R", name = 'ku'):5.0,
ParameterKey(mechanism = 'binding', part_id = "regulated_promoter_R", name = 'cooperativity'):4.0,
#Repressed Promoter Transcription. Note the part_id = [promoter_name]_[regulator_name]
#These regulate RNAP binding to a repressed promoter and transcription
ParameterKey(mechanism = 'transcription', part_id = 'regulated_promoter_R', name = 'kb'):1,
ParameterKey(mechanism = 'transcription', part_id = "regulated_promoter_R", name = 'ku'):100.0,
ParameterKey(mechanism = 'transcription', part_id = 'regulated_promoter_R', name = "ktx"): 1.0, #Transcription Rate
#Leak Parameters for transcription
#These regulate expression of an unbound promoter
ParameterKey(mechanism = 'transcription', part_id = 'regulated_promoter_leak', name = "kb"): 2.,
ParameterKey(mechanism = 'transcription', part_id = 'regulated_promoter_leak', name = "ku"): 100,
ParameterKey(mechanism = 'transcription', part_id = 'regulated_promoter_leak', name = "ktx"): 1.0, #Transcription Rate
}
repressor = Species("R", material_type = "protein")
activator = Species("A", material_type = "protein")
reporter = Species("reporter", material_type = "protein")
#Create a RegulatedPromoter Object named "P_reg" with regulators "activator" and "repressor"
#By Loading custom parameters into the promoter, we override the default parameters of the Mixture
P_reg = RegulatedPromoter("regulated_promoter", regulators=[activator, repressor], leak=True, parameters = component_parameters)
#Create a DNA assembly "reporter" with P_reg for its promoter
reg_reporter = DNAassembly(name="reporter", promoter=P_reg, rbs="Strong", protein = reporter)
#Use a simple TxTl model with dilution
#M = SimpleTxTlDilutionMixture(name="e coli", parameter_file = "default_parameters.txt", components=[reg_reporter])
M = TxTlExtract(name="e coli extract", parameter_file = "default_parameters.txt", components=[reg_reporter])
CRN = M.compile_crn()
print(CRN.pretty_print(show_rates = True, show_keys = False))
# + slideshow={"slide_type": "skip"} tags=[]
#Lets titrate Repressor and Activator - notice the bahvior is not combinatorial
try:
import biocrnpyler
import numpy as np
import pylab as plt
import pandas as pd
plt.figure(figsize = (20, 10))
ax1, ax2, ax3 = plt.subplot(131), plt.subplot(132), plt.subplot(133)
titration_list = [0, .05, .1, .5, 1.0, 5.]
N = len(titration_list)
HM = np.zeros((N, N))
for a_ind, a_c in enumerate(titration_list):
for r_ind, r_c in enumerate(titration_list):
x0 = {reg_reporter.dna:.1, repressor:r_c, activator:a_c}
timepoints = np.linspace(0, 1000, 1000)
R = CRN.simulate_with_bioscrape_via_sbml(timepoints, initial_condition_dict = x0)
if a_ind == 0:
plt.sca(ax1)
plt.plot(R["time"], R[str(reporter)], label = "[A]="+str(a_c) +" [R]="+str(r_c))
if r_ind == 0:
plt.sca(ax2)
plt.plot(R["time"], R[str(reporter)], label = "[A]="+str(a_c) +" [R]="+str(r_c))
HM[a_ind, r_ind] = R[str(reporter)][len(timepoints)-1]
plt.sca(ax1)
plt.title("Repressor Titration")
plt.legend()
plt.sca(ax2)
plt.title("Activator Titration")
plt.legend()
plt.sca(ax3)
plt.title("Endpoint Heatmap (log)")
cb = plt.pcolor(np.log(HM))
plt.colorbar(cb)
plt.xlabel("Repressor")
plt.ylabel("Activator")
plt.xticks(np.arange(.5, N+.5, 1), [str(i) for i in titration_list])
plt.yticks(np.arange(.5, N+.5, 1), [str(i) for i in titration_list])
except ModuleNotFoundError:
print('please install the plotting libraries: pip install biocrnpyler[all]')
# + [markdown] slideshow={"slide_type": "slide"}
# # Example 5: Induction Model of a Ligand which Activates a Transcription Factor
# In many biological circuits, small molecules (ligands) can bind to a transcription factor modulating its functionality.
#
# In BioCRNpyler, we will model this by creating a ChemicalComplex Component which consists of a Transcription Factor and a ligand. This the ComplexSpecies formed by by binding the transcription will also work as the regulator (activator or repressor) of a regulated promoter. In this example, we will use RepressablePromoter.
#
# In the activating case, the bound form of the ChemicalComplex will induce gene expression.
# + slideshow={"slide_type": "subslide"} tags=[]
inactive_repressor = Species("A", material_type = "protein")
ligand = Species("L", material_type = "ligand")
#Create a ChemicalComplex to model ligand-inactive_repressor bindning
activatable_repressor = ChemicalComplex([inactive_repressor, ligand])
#Other Promoters could also be used
P_repressible = RepressiblePromoter("P_repressible", repressor = activatable_repressor.get_species(), leak = True, parameters = hill_parameters)
#Create a DNA assembly "reporter" with P_activatable for its promoter
repressible_assembly = DNAassembly(name="reporter", promoter=P_repressible, rbs="Strong", protein = "reporter")
M = ExpressionDilutionMixture(name="ExpressionDilutionMixture", parameter_file = "default_parameters.txt", components=[repressible_assembly, activatable_repressor])
CRN = M.compile_crn();print(CRN.pretty_print(show_rates = True, show_keys = False))
# + slideshow={"slide_type": "skip"} tags=[]
#Lets titrate ligand and repressor
try:
import biocrnpyler
import numpy as np
import pylab as plt
import pandas as pd
plt.figure(figsize = (8, 6))
N = 11 #Number of titrations
max_titration = 100
HM = np.zeros((N, N))
for r_ind, R_c in enumerate(np.linspace(0, max_titration, N)):
for l_ind, L_c in enumerate(np.linspace(0, max_titration, N)):
x0 = {repressible_assembly.dna:1, inactive_repressor:R_c, ligand:L_c}
timepoints = np.linspace(0, 1000, 1000)
R = CRN.simulate_with_bioscrape_via_sbml(timepoints, initial_condition_dict = x0)
HM[r_ind, l_ind] = R["protein_reporter"][len(timepoints)-1]
plt.title("Activatable Repressor vs Ligand Endpoint Heatmap\n Exhibits NOT AND Logic")
cb = plt.pcolor(HM)
plt.colorbar(cb)
plt.xlabel("Ligand")
plt.ylabel("Activatbale Repressor")
plt.xticks(np.arange(.5, N+.5, 1), [str(i) for i in np.linspace(0, max_titration, N)])
plt.yticks(np.arange(.5, N+.5, 1), [str(i) for i in np.linspace(0, max_titration, N)])
except ModuleNotFoundError:
print('please install the plotting libraries: pip install biocrnpyler[all]')
# + [markdown] slideshow={"slide_type": "slide"}
# # Example 6: Induction Models of a Ligand which Deactivates a Transcription Factor
# In the inactivating case, the unbound transcription factor will activate the gene and the bound form will not.
# + slideshow={"slide_type": "subslide"} tags=[]
repressor = Species("A", material_type = "protein")
ligand = Species("L", material_type = "ligand")
#Create a ChemicalComplex to model ligand-inactive_repressor bindning
inactive_repressor = ChemicalComplex([repressor, ligand])
#Other Promoters could also be Used
P_repressible = RepressiblePromoter("P_repressible", repressor = repressor, leak = True, parameters = hill_parameters)
#Create a DNA assembly "reporter" with P_activatable for its promoter
repressible_assembly = DNAassembly(name="reporter", promoter=P_repressible, rbs="Strong", protein = "reporter")
M = ExpressionDilutionMixture(name="ExpressionDilutionMixture", parameter_file = "default_parameters.txt", components=[repressible_assembly, activatable_repressor])
CRN = M.compile_crn();print(CRN.pretty_print(show_rates = True, show_keys = False))
# + slideshow={"slide_type": "skip"} tags=[]
#Titration of ligand and repressor
try:
import biocrnpyler
import numpy as np
import pylab as plt
import pandas as pd
plt.figure(figsize = (8, 6))
N = 11 #Number of titrations
max_titration = 100
HM = np.zeros((N, N))
for r_ind, R_c in enumerate(np.linspace(0, max_titration, N)):
for l_ind, L_c in enumerate(np.linspace(0, max_titration, N)):
x0 = {repressible_assembly.dna:1, repressor:R_c, ligand:L_c}
timepoints = np.linspace(0, 1000, 1000)
R = CRN.simulate_with_bioscrape_via_sbml(timepoints, initial_condition_dict = x0)
HM[r_ind, l_ind] = R["protein_reporter"][len(timepoints)-1]
plt.title("Deactivatable Repressor vs Ligand Endpoint Heatmap\nAllows for Tunable Induction")
cb = plt.pcolor(HM)
plt.colorbar(cb)
plt.xlabel("Ligand")
plt.ylabel("Deactivatbale Repressor")
plt.xticks(np.arange(.5, N+.5, 1), [str(i) for i in np.linspace(0, max_titration, N)])
plt.yticks(np.arange(.5, N+.5, 1), [str(i) for i in np.linspace(0, max_titration, N)])
except ModuleNotFoundError:
print('please install the plotting libraries: pip install biocrnpyler[all]')
# + [markdown] slideshow={"slide_type": "slide"}
# # Example 7: Modeling AND, OR, and XOR Promoters with Combinatorial Promoter
# CombinatorialPromoter is a Component designed to model arbitrary combinatorial logic on a promoter. For example, a promoter with 2 transcription factor binding sites can have 4 differents states:
# * Nothing bound
# * just factor 1 bound
# * just factor 2 bound
# * factors 1 and 2 bound
#
# In general, a promoter with $N$ binding sites has up to $2^N$ possible states. Combinatorial promoter enumerates all these states and allows for the modeller to decide which are capable of transcription and which are not. For more details on this class, see the CombinatorialPromoter example ipython notebook in the BioCRNpyler examples folder.
#
# Below, we will use a Combinatorial Promoter to Produce OR, AND, and XOR logic with two species, $A$ and $B$ by passing in lists of the transcribable combinations of regulators to the tx_capable_list keyword.
#
# + slideshow={"slide_type": "subslide"} tags=[]
#AND Logic
A = Species("A") ;B = Species("B") #Inducers
#Create the Combinatorial Promoter
Prom_AND = CombinatorialPromoter("combinatorial_promoter",[A,B], tx_capable_list = [[A,B]], leak = True) #the Combination A and B can be transcribed
AND_assembly = DNAassembly("AND",promoter=Prom_AND,rbs="medium",protein="GFP")
#Use an Expression Mixture to focus on Logic, not Transcription & Translation
M = ExpressionExtract(name="expression", parameter_file = "default_parameters.txt", components=[AND_assembly])
CRN = M.compile_crn(); print(CRN.pretty_print(show_rates = True, show_keys = False))
#Lets titrate A and B
try:
import biocrnpyler
import numpy as np
import pylab as plt
import pandas as pd
plt.figure(figsize = (8, 6))
N = 11 #Number of titrations
max_titration = 10
HM = np.zeros((N, N))
for a_ind, A_c in enumerate(np.linspace(0, max_titration, N)):
for b_ind, B_c in enumerate(np.linspace(0, max_titration, N)):
x0 = {AND_assembly.dna:1, A:A_c, B:B_c}
timepoints = np.linspace(0, 1000, 1000)
R = CRN.simulate_with_bioscrape_via_sbml(timepoints, initial_condition_dict = x0)
HM[a_ind, b_ind] = R["protein_GFP"][len(timepoints)-1]
plt.title("AND Endpoint Heatmap")
cb = plt.pcolor(HM)
plt.colorbar(cb)
plt.xlabel("B")
plt.ylabel("A")
plt.xticks(np.arange(.5, N+.5, 1), [str(i) for i in np.linspace(0, max_titration, N)])
plt.yticks(np.arange(.5, N+.5, 1), [str(i) for i in np.linspace(0, max_titration, N)])
except ModuleNotFoundError:
print('please install the plotting libraries: pip install biocrnpyler[all]')
# + slideshow={"slide_type": "subslide"} tags=[]
#Create OR Logic
Prom_OR = CombinatorialPromoter("combinatorial_promoter",[A,B], leak=False,
tx_capable_list = [[A,B], [A], [B]]) #the Combinations A and B or just A or just B be transcribed
ORassembly = DNAassembly("OR",promoter=Prom_OR,rbs="medium",protein="GFP")
print(ORassembly)
#Use an Expression Mixture to focus on Logic, not Transcription & Translation
M = ExpressionExtract(name="expression", parameter_file = "default_parameters.txt", components=[ORassembly])
CRN = M.compile_crn()
print(CRN.pretty_print(show_rates = True, show_keys = False))
#Lets titrate A and B
try:
import biocrnpyler
import numpy as np
import pylab as plt
import pandas as pd
plt.figure(figsize = (6, 6))
N = 11 #Number of titrations
max_titration = 10
HM = np.zeros((N, N))
for a_ind, A_c in enumerate(np.linspace(0, max_titration, N)):
for b_ind, B_c in enumerate(np.linspace(0, max_titration, N)):
x0 = {ORassembly.dna:1, A:A_c, B:B_c}
timepoints = np.linspace(0, 1000, 1000)
R = CRN.simulate_with_bioscrape_via_sbml(timepoints, initial_condition_dict = x0)
HM[a_ind, b_ind] = R["protein_GFP"][len(timepoints)-1]
plt.title("OR Endpoint Heatmap")
cb = plt.pcolor(HM)
plt.colorbar(cb)
plt.xlabel("B")
plt.ylabel("A")
plt.xticks(np.arange(.5, N+.5, 1), [str(i) for i in np.linspace(0, max_titration, N)])
plt.yticks(np.arange(.5, N+.5, 1), [str(i) for i in np.linspace(0, max_titration, N)])
except ModuleNotFoundError:
print('please install the plotting libraries: pip install biocrnpyler[all]')
# + slideshow={"slide_type": "subslide"} tags=[]
#Create XOR Logic
Prom_XOR = CombinatorialPromoter("combinatorial_promoter",[A,B], leak=False,
tx_capable_list = [[A], [B]]) #the Combinations just A or just B can be transcribed
XORassembly = DNAassembly("XOR",promoter=Prom_XOR,rbs="medium",protein="GFP")
#Use an Expression Mixture to focus on Logic, not Transcription & Translation
M = ExpressionExtract(name="expression", parameter_file = "default_parameters.txt", components=[XORassembly])
CRN = M.compile_crn()
print(CRN.pretty_print(show_rates = True, show_keys = False))
#Lets titrate A and B
try:
import biocrnpyler
import numpy as np
import pylab as plt
import pandas as pd
plt.figure(figsize = (6, 6))
N = 11 #Number of titrations
max_titration = 10
HM = np.zeros((N, N))
for a_ind, A_c in enumerate(np.linspace(0, max_titration, N)):
for b_ind, B_c in enumerate(np.linspace(0, max_titration, N)):
x0 = {XORassembly.dna:1, A:A_c, B:B_c}
timepoints = np.linspace(0, 1000, 1000)
R = CRN.simulate_with_bioscrape_via_sbml(timepoints, initial_condition_dict = x0)
HM[a_ind, b_ind] = R["protein_GFP"][len(timepoints)-1]
plt.title("XOR Endpoint Heatmap")
cb = plt.pcolor(HM)
plt.colorbar(cb)
plt.xlabel("B")
plt.ylabel("A")
plt.xticks(np.arange(.5, N+.5, 1), [str(i) for i in np.linspace(0, max_titration, N)])
plt.yticks(np.arange(.5, N+.5, 1), [str(i) for i in np.linspace(0, max_titration, N)])
except ModuleNotFoundError:
print('please install the plotting libraries: pip install biocrnpyler[all]')
# -
| examples/4. Promoters Transcriptional Regulation and Gene Regulatory Networks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] toc="true"
# # Table of Contents
# <p>
# + run_control={"frozen": false, "read_only": false}
# system level
import sys
# arrays
import numpy as np
# keras
from keras.models import Sequential, Model
from keras.layers import Input, Flatten, Dense, Activation, Dropout, merge
from keras.layers.convolutional import Convolution2D, MaxPooling2D
from keras.layers.normalization import BatchNormalization
from keras.optimizers import SGD
from keras.models import model_from_json
from keras.utils import np_utils
from keras import backend as K
from keras.callbacks import Callback, ModelCheckpoint
from keras.preprocessing.image import ImageDataGenerator
# sklearn (for machine learning)
from sklearn import metrics
# plotting
from matplotlib import pyplot as plt
import matplotlib.cm as cm
from mpl_toolkits.axes_grid1 import make_axes_locatable
#import seaborn as sns
# theano
import theano
print "CONFIG:", theano.config.device
# + code_folding=[1, 36] run_control={"frozen": false, "read_only": false}
# utility plots
def remove_tick_labels(axis):
"""Remove tick labels."""
axis.set_xticklabels([])
axis.set_yticklabels([])
for tl in axis.xaxis.get_major_ticks():
tl.tick1On = tl.tick2On = False
for tl in axis.yaxis.get_major_ticks():
tl.tick1On = tl.tick2On = False
def make_mosaic(imgs, nrows, ncols, border=1):
"""Make a nice mosaic.
Given a set of images with all the same shape, makes a
mosaic with nrows and ncols;
intended for use with activation layer
"""
# set up data
nimgs = imgs.shape[0]
imshape = imgs.shape#[1:]
print nimgs, imgs.shape, imshape
mosaic = np.ma.masked_all((nrows * imshape[0] + (nrows - 1) * border,
ncols * imshape[1] + (ncols - 1) * border),
dtype=np.float32)
paddedh = imshape[0] + border
paddedw = imshape[1] + border
for i in xrange(nimgs):
row = int(np.floor(i / ncols))
col = i % ncols
mosaic[row * paddedh:row * paddedh + imshape[0],
col * paddedw:col * paddedw + imshape[1]] = imgs[i]
return mosaic
def nice_imshow(ax, data, vmin=None, vmax=None, cmap=None, norm=None):
"""Image plot."""
if cmap is None:
cmap = cm.jet
if vmin is None:
vmin = data.min()
if vmax is None:
vmax = data.max()
divider = make_axes_locatable(ax)
cax = divider.append_axes("right", size="5%", pad=0.05)
remove_tick_labels(ax)
im = ax.imshow(data, vmin=vmin, vmax=vmax,
#interpolation='nearest',
cmap=cmap,
norm=norm)
plt.colorbar(im, cax=cax)
def get_layers(model, data,
version_data,
ind_layer_in=0,
ind_layer_out=0):
"""Recall activation layer information."""
data_sets = {"train": 0, "test": 1}
layer_input = model.layers[ind_layer_in].input
layer_output = K.function([layer_input, K.learning_phase()],
[model.layers[ind_layer_out].output])
layer_outputs = layer_output([data, data_sets[version_data]])[0]
return layer_outputs
def plot_layer_filter_panel(
layer,
ind_obj_choice,
n_img_y=6,
n_img_x=6):
"""Plot: result of a layer; all filters."""
# style
#plt.style.use(dir_style + 'deeplensing.mplstyle')
#sns.set_palette("coolwarm", n_colors=6)
#n_colors = 300
#cmap = sns.color_palette("coolwarm", n_colors=n_colors)#, color_codes=True)
#cmap = ListedColormap(cmap)
layer_object = layer[ind_obj_choice]
# layer_object = layer_object + np.abs(np.min(layer_object))+1.0
c1 = np.squeeze(layer_object)
# plot
plt.figure(figsize=(12, 12))
#plt.grid(False)
nice_imshow(plt.gca(), make_mosaic(c1, n_img_x, n_img_y), vmin=-120, xmax=40, cmap=cmap) #, norm=LogNorm())
return
# + run_control={"frozen": false, "read_only": false}
# ------------------------------------------------------------------------------
# Input variables
# ------------------------------------------------------------------------------
# training variables
nb_train = 1000
nb_valid = 1000
nb_test = 1000
nb_epoch = 10
nb_classes = 2
batch_size = 32
shuffle = True
# data locations
dir_test = "/Users/nord/Dropbox/deeplensing/Data/Simulation/SimLensPop/Data097/"
file_x_data = dir_test + "xtrain_lenspop.npy" # x data (images)
file_y_data = dir_test + "ytrain_lenspop.npy" # y data (labels)
file_lens_data = dir_test + "lenspars_set.npy" # lens data (physical parameters of lenses/non-lenses)
f_model = dir_test + "model.json" # model data (architecture)
f_weights = dir_test + "weights.h5" # model data (weights that we fit for)
# + run_control={"frozen": false, "read_only": false}
# ------------------------------------------------------------------------------
# Read in Data
# ------------------------------------------------------------------------------
# load data
x_data = np.load(file_x_data)
y_data = np.load(file_y_data)
lens_data = np.load(file_lens_data)
# check data sizes
statement = "#TrainingSamples + #ValidSamples #TestSamples > TotalSamples, exiting!!!"
nb_total = nb_train + nb_test + nb_valid
assert nb_total <= len(x_data), statement
# indices for where to slice the arrays
ind_valid_start = ind_train_end = nb_train
ind_valid_end = ind_test_start = nb_train + nb_valid
ind_test_end = nb_train + nb_valid + nb_test
# slice the image arrays
x_train = x_data[:ind_train_end, :, :, :]
x_valid = x_data[ind_valid_start: ind_valid_end, :, :, :]
x_test = x_data[ind_test_start: ind_test_end, :, :, :]
# slice the label arrays
y_train = y_data[:ind_train_end]
y_valid = y_data[ind_valid_start: ind_valid_end]
y_test = y_data[ind_test_start: ind_test_end]
# cast data types
x_train = x_train.astype('float32')
x_valid = x_valid.astype('float32')
x_test = x_test.astype('float32')
print "Data dimensions: "
print "Input data: ", np.shape(x_data), np.shape(y_data)
print "Training set: ", np.shape(x_train), np.shape(y_train)
print "Validation set: ", np.shape(x_valid), np.shape(y_valid)
print "Test Set: ", np.shape(x_test), np.shape(y_test)
print
# + run_control={"frozen": false, "read_only": false}
# ------------------------------------------------------------------------------
# generate the model architecture
# example: shallow res (enter link to reference)
# ------------------------------------------------------------------------------
# Define architecture for model
data_shape = np.shape(x_data)
input_shape = (3, 64, 64)
x = Input(shape=input_shape)
c0 = Convolution2D(32, 3, 3, activation='softplus', subsample=(4, 4), border_mode='same', dim_ordering='th')(x)
b0 = BatchNormalization()(c0)
d0 = Dropout(0.5)(b0)
c1 = Convolution2D( 8, 3, 3, activation='softplus', subsample=(2, 2), border_mode='same')(d0)
b1 = BatchNormalization()(c1)
d1 = Dropout(0.5)(b1)
f = Flatten()(d1)
z0 = Dense(128, activation='softplus')(f)
z1 = Dense(32, activation='softplus')(z0)
y = Dense(1, activation='sigmoid')(z1)
model = Model(input=x, output=y)
# Compile Model
optimizer = 'adadelta'
metrics = ['accuracy']
loss = 'binary_crossentropy'
model.compile(loss=loss, optimizer=optimizer, metrics=metrics)
# + run_control={"frozen": false, "read_only": false}
# ------------------------------------------------------------------------------
# Train model
# ------------------------------------------------------------------------------
# Train
history = model.fit(
x_train, y_train,
batch_size=batch_size,
nb_epoch=nb_epoch,
validation_data=(x_valid, y_valid),
#shuffle=shuffle,
verbose=True
)
# Save data
model.save_weights(f_weights, overwrite=True)
open(f_model, 'w').write(model.to_json())
# + run_control={"frozen": false, "read_only": false}
# ------------------------------------------------------------------------------
# Evaluate
# ------------------------------------------------------------------------------
# predict
prob = model.predict(x_valid)
pred = (prob > 0.5).astype('int32')
# measure confusion
cm = metrics.confusion_matrix(y_valid, pred, labels=[0, 1])
cm = cm.astype('float')
cm_norm = cm / cm.sum(axis=1)[:, np.newaxis]
print "cm", cm
print "cm_norm", cm_norm
fpr, tpr, thresholds = metrics.roc_curve(y_valid, prob, pos_label=1)
auc = metrics.roc_auc_score(y_valid, prob)
print "AUC:", auc
# + run_control={"frozen": false, "read_only": false}
# ------------------------------------------------------------------------------
# Analyze
# ------------------------------------------------------------------------------
# History
hist = history.history
loss = hist['loss']
val_loss = hist["val_loss"]
epochs = np.arange(nb_epoch)
figsize=(5,3)
fig, axis1 = plt.subplots(figsize=figsize)
plot1_loss = axis1.plot(epochs, loss, 'b', label='loss')
plot1_val_loss = axis1.plot(epochs, val_loss, 'r', label="val loss")
plots = plot1_loss + plot1_val_loss
labs = [l.get_label() for l in plots]
axis1.set_xlabel('Epoch')
axis1.set_ylabel('Loss')
plt.title("Loss History")
plt.tight_layout()
axis1.legend(loc='upper right')
# ROC
figsize=(5,5)
fig, axis1 = plt.subplots(figsize=figsize)
x_onetoone = y_onetoone = [0, 1]
#label_roc = "ROC, AUC = " + auc if auc is not None else "ROC"
plt.plot(fpr, tpr, 'r-')#, label=label_roc)
plt.plot(x_onetoone, y_onetoone, 'k--', label="1-1")
plt.legend(loc=0)
plt.title("Receiver Operator Characteristic (ROC)")
plt.xlabel("False positive (1 - Specificity)")
plt.ylabel("True positive (selectivity)")
plt.tight_layout()
# plot variety of filters for a given image
# print out CM cleanly
# + run_control={"frozen": false, "read_only": false}
layers = get_layers(model, x_valid, "test",
ind_layer_in=1,
ind_layer_out=1)
plot_layer_filter_panel(layers[0], 0)
# + run_control={"frozen": false, "read_only": false}
| notebooks/demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Bases de connaissance : interroger Wikidata à l'aide de requêtes SPARQL
# ## Imports
from datetime import datetime as dt
from SPARQLWrapper import SPARQLWrapper, JSON
# ## Obtenir la liste des politiciens belges
# +
# Retrieve results from SPARQL
endpoint = "https://query.wikidata.org/bigdata/namespace/wdq/sparql"
sparql = SPARQLWrapper(endpoint)
# P27 = country of citizenship
# Q31 = Belgium
# P106 = Opccupation
# Q82955 = Politician
# P569 = date of birth
# P570 = date of death
statement = """
SELECT DISTINCT ?person ?personLabel ?dateBirth ?dateDeath WHERE {
?person wdt:P27 wd:Q31 .
?person wdt:P106 wd:Q82955 .
?person wdt:P569 ?dateBirth .
OPTIONAL {?person wdt:P570 ?dateDeath .}
SERVICE wikibase:label { bd:serviceParam wikibase:language "en" . }
}
ORDER BY ?personLabel
"""
sparql.setQuery(statement)
sparql.setReturnFormat(JSON)
results = sparql.query().convert()
rows = results['results']['bindings']
print(f"\n{len(rows)} Belgian politicians found\n")
print(rows[:10])
# -
# ## Filtrer pour n'afficher que les noms contenant "Bouchez"
# +
name_filter = 'Bouchez'
max_results = 10
date_format = "%Y-%m-%dT%H:%M:%SZ"
filtered_rows = [row for row in rows if name_filter in row['personLabel']['value']]
print(f"Displaying the first {max_results}:\n")
for row in filtered_rows[:max_results]:
try:
birth_date = dt.strptime(row['dateBirth']['value'], date_format)
birth_year = birth_date.year
except ValueError:
birth_year = "????"
try:
death_date = dt.strptime(row['dateDeath']['value'], date_format)
death_year = death_date.year
except ValueError: # unknown death date
death_year = "????"
except KeyError: # still alive
death_year = ""
print(f"{row['personLabel']['value']} ({birth_year}-{death_year})")
# -
# ## Pour en savoir plus
# - Le projet Wikidata : https://www.wikidata.org/wiki/Wikidata:Main_Page
# - Aide à la construction de requêtes : https://query.wikidata.org/
| module1/s2_sparql.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pprint
import numpy
import toyplot.mp4
x = numpy.random.normal(size=100)
y = numpy.random.normal(size=len(x))
# +
canvas = toyplot.Canvas(300, 300, style={"background-color":"white"})
axes = canvas.cartesian()
mark = axes.scatterplot(x, y, size=10)
text_mark = canvas.text(150, 20, "0/5 <small>(0.00)</small>")
def callback(frame):
frame.set_datum_text(text_mark, 0, 0,
text="%s/%s <small>(%.2f)</small>" % (frame.index, frame.count - 1, frame.time),
style={"font-size": "14px", "font-weight":"bold"},
)
if frame.index == 0:
for i in range(len(x)):
frame.set_datum_style(mark, 0, i, style={"opacity":0.2})
else:
frame.set_datum_style(mark, 0, frame.index - 1, style={"opacity":1.0})
canvas.animate(len(x) + 1, callback)
# +
def progress(frame):
print(frame, end=" ")
toyplot.mp4.render(canvas, "test.mp4", progress=progress)
| notebooks/text-animation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Basic Classifier using InferDotNet
# ## Introduction
# This notebook looks at building a `1-feature` probabilistic classifier - a.k.a. a Bayes point machine. Whilst reading the <a href='https://mbmlbook.com/EmailClassifier_A_model_for_classification.html'>Model Based Machine Learning</a> ebook by <a href="https://www.microsoft.com/en-us/research/people/jwinn/?from=http%3A%2F%2Fresearch.microsoft.com%2Fen-us%2Fpeople%2Fjwinn%2F"><NAME></a> et al. from <a href="https://www.microsoft.com/en-us/research/">Microsoft Reseach</a> I found it very confusing diving right into the model based machine learning approach and was looking for a way to tie MBML with traditional ML - this notebook is the result.
# Although the <a href="https://mbmlbook.com/index.html">MBML book</a> is a brilliant book, the first 3 chapters cover very probabilistic approaches to solving problem - as it should, as that is what the book is all about 😅.
#
# However, coming from traditional machine learning I was glad to see that in Chapter 4 they build a binary classifier, albeit in a probabilistic fashion. The MBML book uses <a href="https://en.wikipedia.org/wiki/Factor_graph#:~:text=A%20factor%20graph%20is%20a,the%20factorization%20of%20a%20function.&text=Factor%20graphs%20generalize%20constraint%20graphs,where%20all%20factors%20are%20constraints.">factor graphs</a> across the board - if you're not yet aquited with factor graphs, the TLDR from <a href="https://en.wikipedia.org/wiki/Factor_graph#:~:text=A%20factor%20graph%20is%20a,the%20factorization%20of%20a%20function.&text=Factor%20graphs%20generalize%20constraint%20graphs,where%20all%20factors%20are%20constraints.">wikipedia</a>:
#
# > A factor graph is a bipartite graph representing the factorization of a function. Given a factorization of a function $g$,
# > $$g(X_1,X_2,\dots,X_n) = \prod_{j=1}^m f_j(S_j),$$
# > where ${\displaystyle S_{j}\subseteq \{X_{1},X_{2},\dots ,X_{n}\}}$, the corresponding factor graph ${\displaystyle G=(X,F,E)}$ consists of variable vertices ${\displaystyle X=\{X_{1},X_{2},\dots ,X_{n}\}}$, factor vertices ${\displaystyle F=\{f_{1},f_{2},\dots ,f_{m}\}}$, and edges ${\displaystyle E}$. The edges depend on the factorization as follows: there is an undirected edge between factor vertex ${\displaystyle f_{j}}$ and variable vertex ${\displaystyle X_{k}}$ iff ${\displaystyle X_{k}\in S_{j}}$.
#
# In other words, factor graphs are a way to factorize functions, and factor graphs as used in MBML specifically where $g(X_1,X_2,\dots,X_n)$ is a joint distribution or a joint likelihood function, and the factorization depends on the conditional independencies among the variables. These factor graphs can be solved using various algorithms like Belief propagation, also known as sum-product message passing, and a go-to platform to perform these calculations is C#'s dotnet using the Microsoft funded inferDotNet package.
# ### The Setup
# This notebook duplicates the concepts in chapter 4 of MBML, which is titled: `Uncluttering Your Inbox` and is all about building a classifier that can classify whether a person is going to reply to an email or not, and then using this classifier to, well, uncluttering your inbox. The chapter starts off by building a naive 1 feature classifier with the following factor graph:
#
#
# <p align="center">
# <img src='assets/email-classifier.jpg' width='300px'>
# </p>
#
# The factor graph translates into the following. We've got some `featureValue` that we observe (that's why it is grayed out - it's "observed") which is our feature value, i.e. $X_1$. Then we create a random variable called score, calculated as $score = featureValue \times weight$, i.e. $y=w\times x_1$. After that it gets a bit funky. We are trying to build a classifier, in other words, we want to "learn" some mapping, $f(X)$, that will produce a value (score) when we pass our feature value(s) $x_1$ through. We whould then like to determine a threshold for that score to say whether our feature value(s) comes from one class or the other. The most basic classifier to do this job is Logistic Regression. Here we assume a mapping from $x$ to $y$ as
#
# $${\displaystyle y={\begin{cases}1&\beta _{0}+\beta _{1}x+\varepsilon >0\\0&{\text{else}}\end{cases}}}$$
#
# where $\beta_0$ and $\beta_1$ are parameters that we will estimate in our training process and $\epsilon$ is the standard error term. This is basically what this factor graph is trying to do, build a similiar classifier, although in a very different approach - the Model Based Machine Learning approach. From the <a href="https://mbmlbook.com/EmailClassifier_A_model_for_classification.html">MBML book</a>:
#
#
# > You may be surprised to learn that many classification algorithms can be interpreted as doing approximate inference in some probabilistic model. So rather than running a classification algorithm, we can instead build the corresponding model and use an inference algorithm to do classification. Why would we do this instead of using the classification algorithm? Because a model-based approach to classification gives us several benefits:
# > * The assumptions in the classifier are made explicit. This helps us to understand what the classifier is doing, which can allow us to improve how we use it to achieve better prediction accuracy.
# > * We can modify the model to improve its accuracy or give it new capabilities, beyond those of the original classifier.
# > * We can use standard inference algorithms both to train the model and to make predictions with it. This is particularly useful when modifying the model, since the training and prediction algorithms remain in sync with the modified model.
#
#
# They MBML book does come with code snippets, however most of the time it is very objectified code, which doesn't really help to learn the basics of inferDotNet, you can see <a href='https://github.com/dotnet/mbmlbook/blob/master/src/4.%20Uncluttering%20Your%20Inbox/Models/OneFeatureModel.cs'>here</a>.
#
# So instead of using their code, this repo looks at creating the above factor graph, but instead of classifying whether an email is replied to or not, we dumb it down even further and aim to classify flowers from the Iris dataset as Setosa or Virginica using their sepal length attribute. If you haven't used the Iris dataset before, it consists of 150 flower oberservations for three different Iris species: Setosa, Versicolor and Virginica with their associated attributes: sepal length, sepal width, petal length and petalwidth. Here we are only using the Setosa and Verginica species and their sepal length attribute that is disibtrubed as
#
# <p align="center">
# <img src='assets/sepal-length-dist.jpg' width='70%'>
# </p>
#
# So our factor graph looks like this:
#
# <p align="center">
# <img src='assets/iris-classifier.jpg' width='300px'>
# </p>
#
# So what is happening here? Model based machine learning is all about *assumptions*. The assumptions chapter 4 makes to create this factor graph is:
#
# 1. The feature values can always be calculated, for any email.
# 2. Each email has an associated continuous score which is higher when there is a higher probability of the user replying to the email.
# 3. If an email’s feature value changes by $x$, then its score will change by $weight \times x$ for some fixed, continuous weight.
#
# In our flower case:
#
# 1. The feature values can always be calculated, for any flower.
# 2. Each flower has an associated continuous score which is higher when there is a higher probability of the flower being Setosa.
# 3. If an flower's feature value changes by $x$, then its score will change by $weight \times x$ for some fixed, continuous weight.
#
# Assumption 1 and 2 is just house keeping, but assumption 3 gives rise to:
#
# <p align="center">
# <img src='assets/assumption-3.jpg' width='300px'>
# </p>
#
# From MBML:
# > In drawing the factor graph, we’ve had to assume some prior distribution for weight. In this case, we have assumed that the weight is drawn from a Gaussian distribution with zero mean, so that it is equally likely to be positive or negative.
#
# Assumption 4 becomes: The weight for a feature is equally likely to be positive or negative.
#
# We might be tempted to do the following:
#
# <p align="center">
# <img src='assets/threshold.jpg' width='300px'>
# </p>
#
# Just add a threshold with a lot of variance centered around 0 and let the model go. However, MBML warns against this as any inference algorithm will fail trying to compute this model. The book gives the reason
#
# > Unfortunately, if we attempt to run inference on this model then any inference algorithm we try will fail. This is because some of the observed values have zero probability under the model. In other words, there is no way that the data-generating process encoded by our model could have generated the observed data values. When your data has zero probability under your model, it is a sure sign that the model is wrong!
#
# What this means, is there are some outliers appear to come from one side of the classification, but are really from the other class. Looking at our sepal length distribution below, we highlight which data points will have zero probability under the model.
#
# <p align="center">
# <img src='assets/sepal-length-dist-with-overlap.jpg' width='500px'>
# </p>
#
# As is common in MBML, to combat this, we add some noise to our observation's score calculation. In other words, we take the score that is generated by taking the product of the sepal length ($x$) and the weight ($w$) and we add some Gaussian noise by fixing the mean of a Gaussian random variable to the calculated score ($x \times w$) with variance 10.
#
# So we get the following factor graph with added noise and we threshold the noisyScore using a random Gaussian variable with prior $\mathcal{N}(0,10)$.
#
# <p align="center">
# <img src='assets/iris-classifier.jpg' width='300px'>
# </p>
#
# Coming to grips with factors graphs, their associated assumptions and how to navigate pitfalls like observed data having zero probability under a model is one part of the MBML journey. The next is converting the factor graph into working C# code using the inferDotNet package.
#
# ### The C# Model Code
# The first piece of C# code in Program.cs is pure C# (no inferDotNet) and just reads in the CSV generated by this notebook:
#
# ```C#
# string dataDir = args[0];
# string datasetFilename = dataDir+args[1];
# string[] lines = File.ReadAllLines(datasetFilename);
# bool[] isSetosaLabel = new bool[lines.Length];
# double[] featureVal = new double[lines.Length];
#
# for (int i = 0; i < lines.Length; i++)
# {
# string[] strArray = lines[i].Split('|');
# isSetosaLabel[i] = strArray[1] == "1";
# featureVal[i] = float.Parse(strArray[0].Replace(".", ","));
# }
# ```
#
# Next we create the model.
#
# We start by creating a `Range` variable from the `Microsoft.ML.Probabilistic.Models.Range` namespace that we alias as `Range`. We will use this range to iterate over all of our observations/flowers.
#
# ```C#
# int numberOfFlowers = lines.Length;
# Range flower = new Range(numberOfFlowers).Named("flower");
# ```
#
# The next bit of inferDotNet code is to enure our range gets handled sequentially. This is a bit of an artifact from bringing across code from the email example as for email temporal consistency is preferred. For flowers' sepal lengths, maybe less so, but it can't hurt.
#
# ```C#
# flower.AddAttribute(new Sequential());
# ```
#
# Next we declare the variables in our factor graph within InferDotNet, i.e. all the rounded squares. We can use our `flower Range` along with `Variable.Array<T>` to create the variables for our feature $x$ and our label $y$.
#
# ```C#
# // The feature - x
# VariableArray<double> featureValues = Variable.Array<double>(flower).Named("featureValue").Attrib(new DoNotInfer());
# // The label - y
# VariableArray<bool> isSetosa = Variable.Array<bool>(flower).Named("isSetosa");
# ```
#
# For our random variables, weight and threshold, we initialise them using `Variable.GaussianFromMeanAndVariance`.
#
# ```C#
# // The weight - w
# Variable<double> weight = Variable.GaussianFromMeanAndVariance(0,1).Named("weight");
# // The threshold
# Variable<double> threshold = Variable.GaussianFromMeanAndVariance(-5,10).Named("threshold");
# ```
#
# Next we can loop over our `flower Range` using `Variable.ForEach` and calculate the score, and consequently our noisyScore. We can then make our "prediction" by checking whether the noisy score is above or below our threshold. Something to note here is that the `Variable.ForEach` method takes a range as argument, and within the `using` block, that same variable, i.e. `flower`, is the iterating variable - this was a bit confusing to me and took some getting use to. So no counter variable `i` for a loop like `for(i=0, ...)`, instead just `using (Variable.ForEach(flower)){}` and you can use the `flower` variable inside the block.
#
# ```C#
# using (Variable.ForEach(flower))
# {
# var score = (featureValues[flower] * weight).Named("score");
#
# var noisyScore = Variable.GaussianFromMeanAndVariance(score, 10).Named("noisyScore");
# isSetosa[flower] = noisyScore > threshold;
# }
# ```
#
# We've got two more steps to go before we've converted our factor graph into some code, setting our observed values and running inference. To observe values within InferDotNet we do the following:
#
# ```C#
# isSetosa.ObservedValue = isSetosaLabel;
# featureValues.ObservedValue = featureVal;
# ```
#
# Something to note here is that `isSetosa` and `featureValues` are our inferDotNet variables, whereas `isSetosaLabel` and `featureVal` are our vanilla C# arrays.
#
# The final bit in the puzzle is to run inference, i.e. let the energy flow our model and let the priors get updated with evidence - if there is any evidence. For this example we'll be using the `ExpectationPropagation` (EP) algorithm to do our message passing. Another option is to use `VariationalMessagePassing` (VMP), however this doesn't work for our current setup and my knowledge on these algorithms is still a bit vague at this stage to know why.
#
# ```C#
# var InferenceEngine = new InferenceEngine(new ExpectationPropagation());
# InferenceEngine.NumberOfIterations = 50;
# ```
#
# We can then infer our posterior weight and threshold Gaussians by running:
#
# ```C#
# Gaussian postWeight = InferenceEngine.Infer<Gaussian>(weight);
# Gaussian postThreshold = InferenceEngine.Infer<Gaussian>(threshold);
# ```
#
# To be able to get the results back into this notebook I use the following to create a CSV with the means and variances of our posterior Gaussians.
#
# ```C#
# var results = new StringBuilder();
#
# results.AppendLine("variable;mean;variance");
# var line = string.Format("postWeight;{0};{1}", postWeight.GetMean(), postWeight.GetVariance());
# results.AppendLine(line.Replace(',', '.'));
# line = string.Format("postThreshold;{0};{1}", postThreshold.GetMean(), postThreshold.GetVariance());
# results.AppendLine(line.Replace(',', '.'));
#
# File.WriteAllText(dataDir+"results.csv", results.ToString());
# ```
# ## Imports
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from pathlib import Path
from sklearn.preprocessing import MinMaxScaler
from scipy import special
import numpy as np
# Gaussian CDF
def cdf(x,mu,sig):
return 0.5*(1+special.erf((x-mu)/(sig*np.sqrt(2))))
# ### Config
data_dir = Path('../data')
input_dataset = data_dir/'iris.csv'
output_dataset = data_dir/'iris-one-feature.csv'
model_output = data_dir/'model-output.csv'
# ### Data Acquisition
# +
df = pd.read_csv(input_dataset)
# We subset the data to only have Setosa and Virginica
df_2_class = df[df['class'].\
isin(['Iris-virginica', 'Iris-setosa'])].copy()
# -
df_2_class.sample(2)
f = sns.displot(data=df_2_class,
x='sepallength',
aspect=2,
hue='class')
plt.title('Sepal Length Distribution between Setosa and Virginica');
df_2_class.loc[df_2_class['class'] == 'Iris-setosa', 'is_setosa'] = 1
df_2_class.loc[df_2_class['class'] == 'Iris-virginica', 'is_setosa'] = 0
# When scaling this go a bit weird - still need to figure out why
from sklearn.preprocessing import StandardScaler
# scaler = MinMaxScaler(feature_range=(-1,1))
scaler = StandardScaler()
df_2_class.sepallength = scaler.fit_transform(df_2_class.sepallength.values.reshape(-1,1)).flatten()
df_2_class[['sepallength', 'is_setosa']].\
to_csv(output_dataset,
header=False,
sep='|',
index=False)
f = sns.displot(data=df_2_class,
x='sepallength',
aspect=2,
hue='class')
plt.title('Sepal Length Distribution between Setosa and Virginica');
df_2_class.sepallength.describe()
# ## Run Model
# ls ../data
cmd = f'dotnet run --project ../models/bayes-point-classifier/ ../data/ iris-one-feature.csv'
# !{cmd}
# ## Analyze Results
df_results = pd.read_csv(data_dir/'results.csv', sep=';')
df_results
postWeightMean = df_results[df_results.variable == "postWeight"]['mean'].values[0]
postWeightVariance = df_results[df_results.variable == "postWeight"]['variance'].values[0]
postThresholdMean = df_results[df_results.variable == "postThreshold"]['mean'].values[0]
postThresholdVariance = df_results[df_results.variable == "postThreshold"]['variance'].values[0]
postThresholdStandardDev = np.sqrt(postThresholdVariance)
postWeightMean, postWeightVariance, postThresholdMean, postThresholdVariance, postThresholdStandardDev
sampledWeight = np.random.normal(postWeightMean, np.sqrt(postWeightVariance))
sampledWeight, postWeightMean
df_2_class['score'] = df_2_class.sepallength * postWeightMean
# +
x_min_max = 20
x = np.linspace(-1*x_min_max, x_min_max, 1000)
fig = plt.figure(figsize=(15,5))
for sigma in [postThresholdStandardDev* v for v in np.linspace(0.0001, 1, 4)]:
sig = str(round(sigma, 2))
mean = str(round(postThresholdMean, 2))
label = "${\\mathcal {N}_{thresh}}("+mean+","+sig+")$"
y = cdf(x, postThresholdMean, sigma)
plt.plot(x, y, label=label)
sig = str(round(np.sqrt(postWeightVariance), 2))
mean = str(round(postWeightMean, 2))
plt.scatter(x=df_2_class.score,
y=df_2_class.is_setosa,
c=df_2_class.is_setosa.map({0:'red',
1:'green'}))
plt.legend()
plt.title("Noisy Scores with sampled weight $w="+str(round(sampledWeight,2))+"$"+""" from ${\\mathcal {N}_{weight}}("""+mean+","+sig+")$"+"""
Along with the threshold CDFs for various sigma's""");
# -
| notebooks/iris-bayes-point-classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Dependencies:
# pip: scikit-learn, anndata, scanpy
#
# Modified from the Python starter kit for the NeurIPS 2021 Single-Cell Competition.
# Parts with `TODO` are supposed to be changed by you.
#
# More documentation:
#
# https://viash.io/docs/creating_components/python/
# +
import logging
import anndata as ad
import sys
from scipy.sparse import csc_matrix
from sklearn.decomposition import TruncatedSVD
from sklearn.linear_model import LinearRegression
import numpy as np
logging.basicConfig(level=logging.INFO)
# +
## VIASH START
# Anything within this block will be removed by `viash` and will be
# replaced with the parameters as specified in your config.vsh.yaml.
meta = { 'resources_dir': '.' }
par = {
'input_train_mod1': 'sample_data/openproblems_bmmc_cite_starter/openproblems_bmmc_cite_starter.train_mod1.h5ad',
'input_train_mod2': 'sample_data/openproblems_bmmc_cite_starter/openproblems_bmmc_cite_starter.train_mod2.h5ad',
'input_test_mod1': 'sample_data/openproblems_bmmc_cite_starter/openproblems_bmmc_cite_starter.test_mod1.h5ad',
'distance_method': 'minkowski',
'output': 'output.h5ad',
'n_pcs': 50,
}
## VIASH END
test_mod2_file = 'sample_data/openproblems_bmmc_cite_starter/openproblems_bmmc_cite_starter.test_mod2.h5ad '
# +
## VIASH START
# Anything within this block will be removed by `viash` and will be
# replaced with the parameters as specified in your config.vsh.yaml.
meta = { 'resources_dir': '.' }
par = {
'input_train_mod1': 'sample_data/openproblems_bmmc_multiome_starter/openproblems_bmmc_multiome_starter.train_mod1.h5ad',
'input_train_mod2': 'sample_data/openproblems_bmmc_multiome_starter/openproblems_bmmc_multiome_starter.train_mod2.h5ad',
'input_test_mod1': 'sample_data/openproblems_bmmc_multiome_starter/openproblems_bmmc_multiome_starter.test_mod1.h5ad',
'distance_method': 'minkowski',
'output': 'output.h5ad',
'n_pcs': 50,
}
## VIASH END
test_mod2_file = 'sample_data/openproblems_bmmc_multiome_starter/openproblems_bmmc_multiome_starter.test_mod2.h5ad'
# -
method_id = 'basic_beans'
sys.path.append(meta['resources_dir'])
# +
logging.info('Reading `h5ad` files...')
input_train_mod1 = ad.read_h5ad(par['input_train_mod1'])
input_train_mod2 = ad.read_h5ad(par['input_train_mod2'])
input_test_mod1 = ad.read_h5ad(par['input_test_mod1'])
# +
# TODO: implement own method
from beans import method
adata = method(input_train_mod1, input_train_mod2, input_test_mod1)
adata.uns["method_id"] = method_id
# +
from scipy.sparse import issparse
issparse(adata.X)
# -
# +
logging.info('Storing annotated data...')
adata.write_h5ad(par['output'], compression = "gzip")
# -
from pygam import LinearGAM
# +
from sklearn.decomposition import TruncatedSVD
logging.info('Performing dimensionality reduction on modality 1 values...')
input_mod1 = ad.concat(
{"train": input_train_mod1, "test": input_test_mod1},
axis=0,
join="outer",
label="group",
fill_value=0,
index_unique="-"
)
embedder_mod1 = TruncatedSVD(n_components=50)
mod1_pca = embedder_mod1.fit_transform(input_mod1.X)
logging.info('Performing dimensionality reduction on modality 2 values...')
embedder_mod2 = TruncatedSVD(n_components=50)
mod2_pca = embedder_mod2.fit_transform(input_train_mod2.X)
# split dimred back up
X_train = mod1_pca[input_mod1.obs['group'] == 'train']
X_test = mod1_pca[input_mod1.obs['group'] == 'test']
y_train = mod2_pca
logging.info('Running Linear regression...')
# -
from matplotlib import pyplot as plt
import pandas as pd
print(input_train_mod1.uns['dataset_id'])
print(input_train_mod2.uns['dataset_id'])
print(input_test_mod1.uns['dataset_id'])
m1_tr = pd.DataFrame(X_train)
m1_te = pd.DataFrame(X_test)
m2_tr = pd.DataFrame(X_train)
input_train_mod2.shape
# +
import colorsys
import numpy as np
N = m1_tr.shape[0]
HSV = [(float(x)/N, 1, 1) for x in range(1,N+1)]
RGB = map(lambda x: colorsys.hsv_to_rgb(*x), HSV)
# -
RGB
# +
plt.scatter(m1_tr[0], m1_tr[1], c=m1_tr.index, cmap='Set2')
plt.title("RNA")
plt.show()
plt.scatter(m1_te[0], m1_te[1], c=m1_te.index, cmap='Set2')
plt.title("RNA")
plt.show()
plt.scatter(m2_tr[0], m2_tr[1], c=m2_tr.index, cmap='Set2')
plt.title("ATAC")
plt.show()
# -
m1_tr.iloc[[0]]
for i in [0,5,10,150]:
plt.scatter(m1_tr.iloc[[i]], m1_te.iloc[[i]])
plt.title("GEX vs ATAC components in one cell")
axes = plt.gca()
axes.set_xlim([-4,5])
axes.set_ylim([-4,5])
plt.show()
input_train_mod2.X.A
for i in [0,5,10,150]:
plt.scatter(input_train_mod1.X[[i]].A, input_train_mod2.X[[i]].A)
plt.title("GEX vs ATAC in one cell")
axes = plt.gca()
axes.set_xlim([-4,5])
axes.set_ylim([-1,1.25])
plt.show()
y_train.shape
input_train_mod2.X.shape
input_train_mod2
from statistics import sqrt
from math import ceil, floor
N = X_train.shape[0]
k = ceil(sqrt(N))
kf = floor(sqrt(N))
kr = round(sqrt(N))
from sklearn.neighbors import KNeighborsRegressor
neigh = KNeighborsRegressor(n_neighbors=k)
neigh.fit(X_train, y_train)
y_pred = neigh.predict(X_test)
# +
# Project the predictions back to the modality 2 feature space
y_pred = y_pred @ embedder_mod2.components_
# Store as sparse matrix to be efficient. Note that this might require
# different classifiers/embedders before-hand. Not every class is able
# to support such data structures.
y_pred = csc_matrix(y_pred)
adata2 = ad.AnnData(
X=y_pred,
obs=input_test_mod1.obs,
var=input_train_mod2.var,
uns={
'dataset_id': input_train_mod1.uns['dataset_id'],
'method_id': 'starter_kit'
},
)
# -
true_test_mod2 = ad.read_h5ad(test_mod2_file)
# +
from sklearn.metrics import mean_squared_error
def calculate_rmse(true_test_mod2, pred_test_mod2):
return mean_squared_error(true_test_mod2.X.toarray(), pred_test_mod2.X.toarray(), squared=False)
# -
calculate_rmse(true_test_mod2, adata)
calculate_rmse(true_test_mod2, adata2)
#forward
tests_f={}
for i in range (1, 200,10):
pred_data = method(input_train_mod1, input_train_mod2, input_test_mod1, k=i, d=50)
tests_f[i]=calculate_rmse(true_test_mod2, pred_data)
tests_f
# #go backwards
# tests={}
# for i in range (2, 200, 10):
# pred_data = method(input_train_mod2, input_train_mod1, true_test_mod2, k=i)
# tests[i]=calculate_rmse(input_test_mod1, pred_data)
# tests
plt.scatter(tests_f.keys(), tests_f.values())
tests_f.keys()
| script.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.9 64-bit (''torch'': conda)'
# name: python3
# ---
# +
# #!/usr/bin/env python3
# -*- coding: utf-8 -*-
# @Date : Jun-01-21 15:11
# @Author : <NAME> (<EMAIL>)
# @RefLink : https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html
# @RefLink : https://discuss.pytorch.org/t/how-to-split-dataset-into-test-and-validation-sets/33987/5
import numpy as np
import torch
from torchvision.datasets import ImageFolder
from torchvision.transforms import Compose, ToTensor, Resize
from torch.utils.data import DataLoader
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
# -
# ## train_test_split
#
# - numpy arrays
X, y = np.arange(10).reshape((5, 2)), range(5)
print(X) # Column first
print(y)
# ### 用于生成数据索引
num_samples = 50000 # Same as MNIST
val_split = 0.2
# train_test_split
# Input: a list of int
# Outputs: train indices, validation indices
train_idx, val_idx = train_test_split(list(range(num_samples)), test_size=val_split, shuffle=False)
# #### 结合 PyTorch Datasets 使用
def train_val_dataset(dataset, val_split=0.25):
train_idx, val_idx = train_test_split(list(range(len(dataset))), test_size=val_split)
datasets = {}
datasets['train'] = Subset(dataset, train_idx)
datasets['val'] = Subset(dataset, val_idx)
return datasets
# +
len_dataset = 1000
val_split = 0.25
# 随机的,因此固定random_state
random_state = 42
train_idx, val_idx = train_test_split(
list(range(len_dataset)), test_size=val_split, random_state=random_state)
print(train_idx[:10]) # 只要random_state一致结果就一直
# -
#### 可视化索引序列
plt.scatter(list(range(len(train_idx))), sorted(train_idx))
# - 稀疏性
# - 随机性
# +
from torchvision import datasets, transforms
kwargs = {'batch_size': 32}
# prepare transform
transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,)),
transforms.Lambda(lambda x: torch.flatten(x)
) # Add for 1D inputs
])
data_root = os.path.expanduser("~/.datasets")
mnist_train = datasets.MNIST(
data_root, train=True, download=True, transform=transform)
mnist_test = datasets.MNIST(
data_root, train=False, transform=transform)
train_loader = torch.utils.data.DataLoader(mnist_train, **kwargs)
test_loader = torch.utils.data.DataLoader(mnist_test, **kwargs)
# -
# mnist_train.__dir__()
mnist_train.targets
from torch.utils.data import Subset
from sklearn.model_selection import train_test_split
def train_val_dataset(dataset, val_split=0.25, random_state=None):
train_idx, val_idx = train_test_split(list(range(len(dataset))), test_size=val_split, random_state=random_state)
datasets = {}
datasets['train'] = Subset(dataset, train_idx)
datasets['val'] = Subset(dataset, val_idx)
return datasets
mnist_train_val = train_val_dataset(mnist_train, random_state=42)
train_loader = torch.utils.data.DataLoader(mnist_train_val["train"], **kwargs)
valid_loader = torch.utils.data.DataLoader(mnist_train_val["val"], **kwargs)
| sklearn_basics/sklearn.model_selection.train_test_split_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Non-massaction propensities in BioCRNpyler
# The propensity $\rho(s)$ (or rate function) of a chemical reaction determines how quickly that reaction occurs. It is assumed to be a function of the chemical species $s$ as well as internal parameter specific to the propensity type.
#
# \begin{equation}
# \{\textrm{Input Species}\} \xrightarrow{\rho(s)} \{\textrm{Output Species}\}
# \end{equation}
#
# By default, BioCRNpyler assumes that propensities are massaction with only one parameter, the rate constant $k$. However, non-massaction propensities are also supported. When creating a reaction with non-massaction propensities, a propensity_params dictionary must be passed into the chemical_reaction_network.reaction constructor with the appropriate parameters for that particular propensity type. The supported propensity types and their relevant parameters are:
#
# 1. "massaction: $\rho(S) = k \Pi_{s} s^{I_s}$. Required parameters: "k" the rate constant. Note: for stochastic models mass action propensities are $\rho(S) = k \Pi_{s} s!/(s - I_s)!$.
# 2. "positivehill": $\rho(s) = k \frac{s^n}{(K^n+s^n)}$. Requried parameters: rate constant "k", offset "K", hill coefficient "n", hill species "s1".
# 3. "negativehill": $\rho(s) = k \frac{1}{(K^n+s^n)}$. Requried parameters: rate constant "k", offset "K", hill coefficient "n", hill species "s1".
# 4. "proportionalpositivehill": $\rho(s) = k d \frac{s^n}{(K^n+s^n)}$. Requried parameters: rate constant "k", offset "K", hill coefficient "n", hill species "s1", propritional species "d".
# 5. "proportionalnegativehill": $\rho(s) = k d \frac{1}{(K^n+s^n)}$. Requried parameters: rate constant "k", offset "K", hill coefficient "n", hill species "s1", propritional species "d".
# 6. "general": $\rho(s) = f(s)$ where $f$ can be any algebraic function typed as a string. Required parameters: "rate" an algebraic expression including species and model parameters written as a string.
#
# The following notebook contains examples of the propensity types support by BioCRNpyler.
# +
from biocrnpyler.chemical_reaction_network import Species, Reaction, ComplexSpecies, ChemicalReactionNetwork
#Names of different supported propensities
propensity_types = ['hillpositive', 'proportionalhillpositive', 'hillnegative', 'proportionalhillnegative', 'massaction', 'general']
#Parameter Values
kb = 100
ku = 10
kex = 1.
kd = .1
#Species
G = Species(name = "G", material_type = "dna") #DNA
A = Species(name = "A", material_type = "protein") #Activator
GA = ComplexSpecies([G, A, A]) #Activated Gene
X = Species(name = "X", material_type = "protein")
rxnd = Reaction([X], [], kd)
#Massaction Activation
species1 = [G, A, GA, X]
rxn0_1 = Reaction([G, A, A], [GA], k=kb, k_rev = ku)
rxn0_2 = Reaction([GA], [GA, X], k=kex)
CRN0 = ChemicalReactionNetwork(species1, [rxn0_1, rxn0_2, rxnd])
print("\nMassaction Activation", repr(CRN0))
#Massaction Repressed
rxn1_1 = Reaction([G, A, A], [GA], k=kb, k_rev = ku)
rxn1_2 = Reaction([G], [G, X], k=kex)
CRN1 = ChemicalReactionNetwork(species1, [rxn1_1, rxn1_2, rxnd])
print("\nMassaction Repression", repr(CRN1))
#hill positive
species2 = [G, A, X]
rxn2_1 = Reaction([G], [G, X], propensity_type = "hillpositive", propensity_params = {"k":kex, "n":2, "K":float(kb/ku), "s1":A})
CRN2 = ChemicalReactionNetwork(species2, [rxn2_1, rxnd])
print("\nHill Positive CRN", repr(CRN2))
#proportional hill positive
rxn3_1 = Reaction([G], [G, X], propensity_type = "proportionalhillpositive", propensity_params = {"k":kex, "n":2, "K":float(kb/ku), "s1":A, "d":G})
CRN3 = ChemicalReactionNetwork(species2, [rxn3_1, rxnd])
print("\nProportional Hill Positive CRN", repr(CRN3))
#hill Negative
rxn4_1 = Reaction([G], [G, X], propensity_type = "hillnegative", propensity_params = {"k":kex, "n":2, "K":float(kb/ku), "s1":A})
CRN4 = ChemicalReactionNetwork(species2, [rxn4_1, rxnd])
print("\nHill Negative CRN", repr(CRN4))
#proportional hill negative
rxn5_1 = Reaction([G], [G, X], propensity_type = "proportionalhillnegative", propensity_params = {"k":kex, "n":2, "K":float(kb/ku), "s1":A, "d":G})
CRN5 = ChemicalReactionNetwork(species2, [rxn5_1, rxnd])
print("\nProportional Hill Negative CRN", repr(CRN5))
# -
| examples/Non-Massaction Propensities.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: dowhy
# language: python
# name: dowhy
# ---
# # Do-sampler Introduction
# by <NAME>
#
# The "do-sampler" is a new feature in do-why. While most potential-outcomes oriented estimators focus on estimating the specific contrast $E[Y_0 - Y_1]$, Pearlian inference focuses on more fundamental quantities like the joint distribution of a set of outcomes Y, $P(Y)$, which can be used to derive other statistics of interest.
#
# Generally, it's hard to represent a probability distribution non-parametrically. Even if you could, you wouldn't want to gloss over finite-sample problems with you data you used to generate it. With these issues in mind, we decided to represent interventional distributions by sampling from them with an object called to "do-sampler". With these samples, we can hope to compute finite-sample statistics of our interventional data. If we bootstrap many such samples, we can even hope for good sampling distributions for these statistics.
#
# The user should note that this is still an area of active research, so you should be careful about being too confident in bootstrapped error bars from do-samplers.
#
# Note that do samplers sample from the outcome distribution, and so will vary significantly from sample to sample. To use them to compute outcomes, it's recommended to generate several such samples to get an idea of the posterior variance of your statistic of interest.
#
# ## Pearlian Interventions
#
# Following the notion of an intervention in a Pearlian causal model, our do-samplers implement a sequence of steps:
#
# 1. Disrupt causes
# 2. Make Effective
# 3. Propagate and sample
#
# In the first stage, we imagine cutting the in-edges to all of the variables we're intervening on. In the second stage, we set the value of those variables to their interventional quantities. In the third stage, we propagate that value forward through our model to compute interventional outcomes with a sampling procedure.
#
# In practice, there are many ways we can implement these steps. They're most explicit when we build the model as a linear bayesian network in PyMC3, which is what underlies the MCMC do sampler. In that case, we fit one bayesian network to the data, then construct a new network representing the interventional network. The structural equations are set with the parameters fit in the initial network, and we sample from that new network to get our do sample.
#
# In the weighting do sampler, we abstractly think of "disrupting the causes" by accounting for selection into the causal state through propensity score estimation. These scores contain the information used to block back-door paths, and so have the same statistics effect as cutting edges into the causal state. We make the treatment effective by selecting the subset of our data set with the correct value of the causal state. Finally, we generated a weighted random sample using inverse propensity weighting to get our do sample.
#
# There are other ways you could implement these three steps, but the formula is the same. We've abstracted them out as abstract class methods which you should override if you'd like to create your own do sampler!
#
# ## Statefulness
#
# The do sampler when accessed through the high-level pandas API is stateless by default.This makes it intuitive to work with, and you can generate different samples with repeated calls to the `pandas.DataFrame.causal.do`. It can be made stateful, which is sometimes useful.
#
# The 3-stage process we mentioned before is implemented by passing an internal `pandas.DataFrame` through each of the three stages, but regarding it as temporary. The internal dataframe is reset by default before returning the result.
#
# It can be much more efficient to maintain state in the do sampler between generating samples. This is especially true when step 1 requires fitting an expensive model, as is the case with the MCMC do sampler, the kernel density sampler, and the weighting sampler.
#
# Instead of re-fitting the model for each sample, you'd like to fit it once, and then generate many samples from the do sampler. You can do this by setting the kwarg `stateful=True` when you call the `pandas.DataFrame.causal.do` method. To reset the state of the dataframe (deleting the model as well as the internal dataframe), you can call the `pandas.DataFrame.causal.reset` method.
#
# Through the lower-level API, the sampler is stateful by default. The assumption is that a "power user" who is using the low-level API will want more control over the sampling process. In this case, state is carried by internal dataframe `self._df`, which is a copy of the dataframe passed on instantiation. The original dataframe is kept in `self._data`, and is used when the user resets state.
#
# ## Integration
#
# The do-sampler is built on top of the identification abstraction used throughout do-why. It uses a `dowhy.CausalModel` to perform identification, and builds any models it needs automatically using this identification.
#
# ## Specifying Interventions
#
# There is a kwarg on the `dowhy.do_sampler.DoSampler` object called `keep_original_treatment`. While an intervention might be to set all units treatment values to some specific value, it's often natural to keep them set as they were, and instead remove confounding bias during effect estimation. If you'd prefer not to specify an intervention, you can set the kwarg like `keep_original_treatment=True`, and the second stage of the 3-stage process will be skipped. In that case, any intervention specified on sampling will be ignored.
#
# If the `keep_original_treatment` flag is set to false (it is by default), then you must specify an intervention when you sample from the do sampler. For details, see the demo below!
#
#
# ## Demo
#
# First, let's generate some data and a causal model. Here, Z confounds our causal state, D, with the outcome, Y.
import os, sys
sys.path.append(os.path.abspath("../../"))
import numpy as np
import pandas as pd
import dowhy.api
# +
N = 5000
z = np.random.uniform(size=N)
d = np.random.binomial(1., p=1./(1. + np.exp(-5. * z)))
y = 2. * z + d + 0.1 * np.random.normal(size=N)
df = pd.DataFrame({'Z': z, 'D': d, 'Y': y})
# -
(df[df.D == 1].mean() - df[df.D == 0].mean())['Y']
# So the naive effect is around 60% high. Now, let's build a causal model for this data.
# +
from dowhy import CausalModel
causes = ['D']
outcomes = ['Y']
common_causes = ['Z']
model = CausalModel(df,
causes,
outcomes,
common_causes=common_causes)
# -
# Now that we have a model, we can try to identify the causal effect.
identification = model.identify_effect()
# Identification works! We didn't actually need to do this yet, since it will happen internally with the do sampler, but it can't hurt to check that identification works before proceeding. Now, let's build the sampler.
# +
from dowhy.do_samplers.weighting_sampler import WeightingSampler
sampler = WeightingSampler(df,
causal_model=model,
keep_original_treatment=True,
variable_types={'D': 'b', 'Z': 'c', 'Y': 'c'})
# -
# Now, we can just sample from the interventional distribution! Since we set the `keep_original_treatment` flag to `False`, any treatment we pass here will be ignored. Here, we'll just pass `None` to acknowledge that we know we don't want to pass anything.
#
# If you'd prefer to specify an intervention, you can just put the interventional value here instead as a list or numpy array.
#
interventional_df = sampler.do_sample(None)
(interventional_df[interventional_df.D == 1].mean() - interventional_df[interventional_df.D == 0].mean())['Y']
# Now we're much closer to the true effect, which is around 1.0!
| docs/source/example_notebooks/do_sampler_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
import numpy as np
import pygfunction as gt
from math import exp, log
# -
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('seaborn-bright')
plt.rcParams['figure.figsize'] = [15, 9]
plt.rcParams['font.size'] = 12
# 
B = 5
D = 1
H = 100
r_b = 0.05
bf_A = gt.boreholes.rectangle_field(N_1=1, N_2=2, B_1=B, B_2=B, H=H, D=D, r_b=r_b)
bf_A
alpha = 1e-6
ts = H**2/(9.*alpha)
lntts = np.arange(-8.5, 3.5, 0.5)
lntts_py = np.arange(-8.25, 3.75, 0.5)
time = np.exp(lntts_py) * ts
pyg_A = gt.gfunction.uniform_heat_extraction(bf_A, time, alpha, disp=True)
cpp_data_A = np.genfromtxt('../examples/1x2-self_1x2_cross_out.csv',
delimiter=',',
skip_header=1,
unpack=True)
cpp_g_self_A = cpp_data_A[1][:]
cpp_g_cross_A = cpp_data_A[2][:]
# +
fig = plt.figure(figsize=(7, 5), dpi=200)
ax = fig.add_subplot(1, 1, 1)
ax.plot(lntts_py, pyg_A, linestyle='-', marker='X', label=r'$g_{A\to A}$ (Reference)')
ax.plot(lntts, cpp_g_self_A, linestyle='--', marker='s', label=r'$g_{A\to A}$')
ax.plot(lntts, cpp_g_cross_A, linestyle=':', marker='v', label=r'$g_{B\to A}$')
plt.grid()
plt.legend()
plt.ylabel('g')
plt.xlabel('ln(t/ts)')
plt.title(r'Field A-B; $X_B$=5m, D=1m; H=100m; $r_b$=0.05m')
plt.savefig('Field_A-B_g_functions.PNG', bbox_inches='tight')
plt.show()
# -
bf_C = gt.boreholes.rectangle_field(N_1=2, N_2=2, B_1=B, B_2=B, H=H, D=D, r_b=r_b)
bf_C
pyg_C = gt.gfunction.uniform_heat_extraction(bf_C, time, alpha, disp=True)
# +
fig = plt.figure(figsize=(7, 5), dpi=200)
ax = fig.add_subplot(1, 1, 1)
ax.plot(lntts_py, pyg_C, linestyle='-', marker='X', label=r'$g_{C\to C}$ (Reference)')
ax.plot(lntts, cpp_g_self_A, linestyle='--', marker='s', label=r'$g_{A\to A}$')
ax.plot(lntts, cpp_g_cross_A, linestyle=':', marker='v', label=r'$g_{B\to A}$')
ax.plot(lntts, cpp_g_self_A + cpp_g_cross_A, linestyle='-.', marker='p', label=r'$g_{A\to A} + g_{B\to A}$')
plt.grid()
plt.legend()
plt.ylabel('g')
plt.xlabel('ln(t/ts)')
plt.title(r'Fields C, A-B; $X_B$=5m, D=1m; H=100m; $r_b$=0.05m')
plt.savefig('Field_A-B-C_g_functions.PNG', bbox_inches='tight')
plt.show()
# -
# 
cpp_data_E = np.genfromtxt('../examples/2x2_L-self_single_cross_out.csv',
delimiter=',',
skip_header=1,
unpack=True)
cpp_g_self_E = cpp_data_E[1][:]
cpp_g_cross_E = cpp_data_E[2][:]
bf_E = gt.boreholes.L_shaped_field(N_1=2, N_2=2, B_1=B, B_2=B, H=H, D=D, r_b=r_b)
bf_E
pyg_E = gt.gfunction.uniform_heat_extraction(bf_E, time, alpha, disp=True)
# +
fig = plt.figure(figsize=(7, 5), dpi=200)
ax = fig.add_subplot(1, 1, 1)
ax.plot(lntts_py, pyg_C, linestyle='-', marker='X', label=r'$g_{C\to C}$ (Reference)')
ax.plot(lntts, cpp_g_self_E, linestyle='--', marker='s', label=r'$g_{E\to E}$')
ax.plot(lntts, cpp_g_cross_E, linestyle=':', marker='v', label=r'$g_{D\to E}$')
ax.plot(lntts, cpp_g_self_E + cpp_g_cross_E, linestyle='-.', marker='p', label=r'$g_{E\to E} + g_{D\to E}$')
plt.grid()
plt.legend()
plt.ylabel('g')
plt.xlabel('ln(t/ts)')
plt.title(r'Fields C, E-D; $X_B$=5m D=1m; H=100m; $r_b$=0.05m')
plt.savefig('Field_C-E-D_g_functions.PNG', bbox_inches='tight')
plt.show()
# -
cpp_data_D = np.genfromtxt('../examples/single_self_2x2_L-cross_out.csv',
delimiter=',',
skip_header=1,
unpack=True)
cpp_g_self_D = cpp_data_D[1][:]
cpp_g_cross_D = cpp_data_D[2][:]
bf_D = gt.boreholes.rectangle_field(N_1=1, N_2=1, B_1=B, B_2=B, H=H, D=D, r_b=r_b)
bf_D
pyg_D = gt.gfunction.uniform_heat_extraction(bf_D, time, alpha, disp=True)
# +
fig = plt.figure(figsize=(7, 5), dpi=200)
ax = fig.add_subplot(1, 1, 1)
ax.plot(lntts_py, pyg_C, linestyle='-', marker='X', label=r'$g_{C\to C}$ (Reference)')
ax.plot(lntts, cpp_g_self_D, linestyle='--', marker='s', label=r'$g_{D\to D}$')
ax.plot(lntts, cpp_g_cross_D, linestyle=':', marker='v', label=r'$g_{E\to D}$')
ax.plot(lntts, cpp_g_self_D + cpp_g_cross_D, linestyle='-.', marker='p', label=r'$g_{D\to D} + g_{E\to D}$')
plt.grid()
plt.legend()
plt.ylabel('g')
plt.xlabel('ln(t/ts)')
plt.title(r'Fields C, D-E; $X_B$=5m, D=1m; H=100m; $r_b$=0.05m')
plt.savefig('Field_C-D-E_g_functions.PNG', bbox_inches='tight')
plt.show()
# -
| notebooks/Validate_pygfunction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Gridded Datasets
import xarray as xr
import numpy as np
import holoviews as hv
hv.extension('matplotlib')
# %opts Scatter3D [size_index=None color_index=3] (cmap='fire')
# In the [Tabular Data](./07-Tabular_Datasets.ipynb) guide we covered how to work with columnar data in HoloViews. Apart from tabular or column based data there is another data format that is particularly common in the science and engineering contexts, namely multi-dimensional arrays. The gridded data interfaces allow working with grid-based datasets directly.
#
# Grid-based datasets have two types of dimensions:
#
# * they have coordinate or key dimensions, which describe the sampling of each dimension in the value arrays
# * they have value dimensions which describe the quantity of the multi-dimensional value arrays
#
# There are many different types of gridded datasets, which each approximate or measure a surface or space at discretely specified coordinates. In HoloViews, gridded datasets are typically one of three possible types: Regular rectilinear grids, irregular rectilinear grids, and curvilinear grids. Regular rectilinear grids can be defined by 1D coordinate arrays specifying the spacing along each dimension, while the other types require grid coordinates with the same dimensionality as the underlying value arrays, specifying the full n-dimensional coordinates of the corresponding array value. HoloViews provides many different elements supporting regularly spaced rectilinear grids, but currently only QuadMesh supports irregularly spaced rectilinear and curvilinear grids.
#
# The difference between uniform, rectilinear and curvilinear grids is best illustrated by the figure below:
#
# <figure>
# <img src="http://www.earthsystemmodeling.org/esmf_releases/public/ESMF_6_3_0r/ESMC_crefdoc/img9.png" alt="grid-types">
# <figcaption>Types of logically rectangular grid tiles. Red circles show the values needed to specify grid coordinates for each type. Reproduced from <a href="http://www.earthsystemmodeling.org/esmf_releases/public/ESMF_6_3_0r/ESMC_crefdoc/node5.html">ESMF documentation</a></figcaption>
# </figure>
#
#
# In this section we will first discuss how to work with the simpler rectilinear grids and then describe how to define a curvilinear grid with 2D coordinate arrays.
#
# ## Declaring gridded data
#
# All Elements that support a ColumnInterface also support the GridInterface. The simplest example of a multi-dimensional (or more precisely 2D) gridded dataset is an image, which has implicit or explicit x-coordinates, y-coordinates and an array representing the values for each combination of these coordinates. Let us start by declaring an Image with explicit x- and y-coordinates:
img = hv.Image((range(10), range(5), np.random.rand(5, 10)), datatype=['grid'])
img
# In the above example we defined that there would be 10 samples along the x-axis, 5 samples along the y-axis and then defined a random ``5x10`` array, matching those dimensions. This follows the NumPy (row, column) indexing convention. When passing a tuple HoloViews will use the first gridded data interface, which stores the coordinates and value arrays as a dictionary mapping the dimension name to a NumPy array representing the data:
img.data
# However HoloViews also ships with an interface for ``xarray`` and the [GeoViews](https://geoviews.org) library ships with an interface for ``iris`` objects, which are two common libraries for working with multi-dimensional datasets:
# +
arr_img = img.clone(datatype=['image'])
print(type(arr_img.data))
try:
xr_img = img.clone(datatype=['xarray'])
print(type(xr_img.data))
except:
print('xarray interface could not be imported.')
# -
# In the case of an Image HoloViews also has a simple image representation which stores the data as a single array and converts the x- and y-coordinates to a set of bounds:
print("Array type: %s with bounds %s" % (type(arr_img.data), arr_img.bounds))
# To summarize the constructor accepts a number of formats where the value arrays should always match the shape of the coordinate arrays:
#
# 1. A simple np.ndarray along with (l, b, r, t) bounds
# 2. A tuple of the coordinate and value arrays
# 3. A dictionary of the coordinate and value arrays indexed by their dimension names
# 3. XArray DataArray or XArray Dataset
# 4. An Iris cube
# # Working with a multi-dimensional dataset
# A gridded Dataset may have as many dimensions as desired, however individual Element types only support data of a certain dimensionality. Therefore we usually declare a ``Dataset`` to hold our multi-dimensional data and take it from there.
dataset3d = hv.Dataset((range(3), range(5), range(7), np.random.randn(7, 5, 3)),
['x', 'y', 'z'], 'Value')
dataset3d
# This is because even a 3D multi-dimensional array represents volumetric data which we can display easily only if it contains few samples. In this simple case we can get an overview of what this data looks like by casting it to a ``Scatter3D`` Element (which will help us visualize the operations we are applying to the data:
hv.Scatter3D(dataset3d)
# ### Indexing
# In order to explore the dataset we therefore often want to define a lower dimensional slice into the array and then convert the dataset:
dataset3d.select(x=1).to(hv.Image, ['y', 'z']) + hv.Scatter3D(dataset3d.select(x=1))
# ### Groupby
#
# Another common method to apply to our data is to facet or animate the data using ``groupby`` operations. HoloViews provides a convient interface to apply ``groupby`` operations and select which dimensions to visualize.
(dataset3d.to(hv.Image, ['y', 'z'], 'Value', ['x']) +
hv.HoloMap({x: hv.Scatter3D(dataset3d.select(x=x)) for x in range(3)}, kdims='x'))
# ### Aggregating
# Another common operation is to aggregate the data with a function thereby reducing a dimension. You can either ``aggregate`` the data by passing the dimensions to aggregate or ``reduce`` a specific dimension. Both have the same function:
hv.Image(dataset3d.aggregate(['x', 'y'], np.mean)) + hv.Image(dataset3d.reduce(z=np.mean))
# By aggregating the data we can reduce it to any number of dimensions we want. We can for example compute the spread of values for each z-coordinate and plot it using a ``Spread`` and ``Curve`` Element. We simply aggregate by that dimension and pass the aggregation functions we want to apply:
hv.Spread(dataset3d.aggregate('z', np.mean, np.std)) * hv.Curve(dataset3d.aggregate('z', np.mean))
# It is also possible to generate lower-dimensional views into the dataset which can be useful to summarize the statistics of the data along a particular dimension. A simple example is a box-whisker of the ``Value`` for each x-coordinate. Using the ``.to`` conversion interface we declare that we want a ``BoxWhisker`` Element indexed by the ``x`` dimension showing the ``Value`` dimension. Additionally we have to ensure to set ``groupby`` to an empty list because by default the interface will group over any remaining dimension.
dataset3d.to(hv.BoxWhisker, 'x', 'Value', groupby=[])
# Similarly we can generate a ``Distribution`` Element showing the ``Value`` dimension, group by the 'x' dimension and then overlay the distributions, giving us another statistical summary of the data:
dataset3d.to(hv.Distribution, 'Value', [], groupby='x').overlay()
# ## Categorical dimensions
# The key dimensions of the multi-dimensional arrays do not have to represent continuous values, we can display datasets with categorical variables as a ``HeatMap`` Element:
heatmap = hv.HeatMap((['A', 'B', 'C'], ['a', 'b', 'c', 'd', 'e'], np.random.rand(5, 3)))
heatmap + heatmap.table()
# ## Non-uniform rectilinear grids
#
# As discussed above, there are two main types of grids handled by HoloViews. So far, we have mainly dealt with uniform, rectilinear grids, but we can use the ``QuadMesh`` element to work with non-uniform rectilinear grids and curvilinear grids.
#
# In order to define a non-uniform, rectilinear grid we can declare explicit irregularly spaced x- and y-coordinates. In the example below we specify the x/y-coordinate bin edges of the grid as arrays of shape ``M+1`` and ``N+1`` and a value array (``zs``) of shape ``NxM``:
n = 8 # Number of bins in each direction
xs = np.logspace(1, 3, n)
ys = np.linspace(1, 10, n)
zs = np.arange((n-1)**2).reshape(n-1, n-1)
print('Shape of x-coordinates:', xs.shape)
print('Shape of y-coordinates:', ys.shape)
print('Shape of value array:', zs.shape)
hv.QuadMesh((xs, ys, zs))
# ## Curvilinear grids
#
# To define a curvilinear grid the x/y-coordinates of the grid should be defined as 2D arrays of shape ``NxM`` or ``N+1xM+1``, i.e. either as the bin centers or the bin edges of each 2D bin.
# +
n=20
coords = np.linspace(-1.5,1.5,n)
X,Y = np.meshgrid(coords, coords);
Qx = np.cos(Y) - np.cos(X)
Qy = np.sin(Y) + np.sin(X)
Z = np.sqrt(X**2 + Y**2)
print('Shape of x-coordinates:', Qx.shape)
print('Shape of y-coordinates:', Qy.shape)
print('Shape of value array:', Z.shape)
qmesh = hv.QuadMesh((Qx, Qy, Z))
qmesh
# -
# ## Working with xarray data types
# As demonstrated previously, `Dataset` comes with support for the `xarray` library, which offers a powerful way to work with multi-dimensional, regularly spaced data. In this example, we'll load an example dataset, turn it into a HoloViews `Dataset` and visualize it. First, let's have a look at the xarray dataset's contents:
xr_ds = xr.tutorial.load_dataset("air_temperature")
xr_ds
# It is trivial to turn this xarray Dataset into a Holoviews `Dataset` (the same also works for DataArray):
hv_ds = hv.Dataset(xr_ds)[:, :, "2013-01-01"]
print(hv_ds)
# We have used the usual slice notation in order to select one single day in the rather large dataset. Finally, let's visualize the dataset by converting it to a `HoloMap` of `Images` using the `to()` method. We need to specify which of the dataset's key dimensions will be consumed by the images (in this case "lat" and "lon"), where the remaing key dimensions will be associated with the HoloMap (here: "time"). We'll use the slice notation again to clip the longitude.
# %%opts Image [colorbar=True]
# %%output size=200
hv_ds.to(hv.Image, kdims=["lon", "lat"], dynamic=False)[:, 220:320, :]
# Here, we have explicitly specified the default behaviour `dynamic=False`, which returns a HoloMap. Note, that this approach immediately converts all available data to images, which will take up a lot of RAM for large datasets. For these situations, use `dynamic=True` to generate a [DynamicMap](./06-Live_Data.ipynb) instead. Additionally, [xarray features dask support](http://xarray.pydata.org/en/stable/dask.html), which is helpful when dealing with large amounts of data.
#
# It is also possible to render curvilinear grids with xarray, and here we will load one such example. The dataset below defines a curvilinear grid of air temperatures varying over time. The curvilinear grid can be identified by the fact that the ``xc`` and ``yc`` coordinates are defined as two-dimensional arrays:
rasm = xr.tutorial.load_dataset("rasm")
rasm.coords
# To simplify the example we will select a single timepoint and add explicit coordinates for the x and y dimensions:
rasm = rasm.isel(time=0, x=slice(0, 200)).assign_coords(x=np.arange(200), y=np.arange(205))
rasm.coords
# Now that we have defined both rectilinear and curvilinear coordinates we can visualize the difference between the two by explicitly defining which set of coordinates to use:
hv.QuadMesh(rasm, ['x', 'y']) + hv.QuadMesh(rasm, ['xc', 'yc'])
#
#
# Additional examples of visualizing xarrays in the context of geographical data can be found in the GeoViews documentation: [Gridded Datasets I](http://geo.holoviews.org/Gridded_Datasets_I.html) and
# [Gridded Datasets II](http://geo.holoviews.org/Gridded_Datasets_II.html). These guides also contain useful information on the interaction between xarray data structures and HoloViews Datasets in general.
# # API
# ## Accessing the data
# In order to be able to work with data in different formats Holoviews defines a general interface to access the data. The dimension_values method allows returning underlying arrays.
#
# #### Key dimensions (coordinates)
#
# By default ``dimension_values`` will return the expanded columnar format of the data:
heatmap.dimension_values('x')
# To access just the unique coordinates along a dimension simply supply the ``expanded=False`` keyword:
heatmap.dimension_values('x', expanded=False)
# Finally we can also get a non-flattened, expanded coordinate array returning a coordinate array of the same shape as the value arrays
heatmap.dimension_values('x', flat=False)
# #### Value dimensions
# When accessing a value dimension the method will similarly return a flat view of the data:
heatmap.dimension_values('z')
# We can pass the ``flat=False`` argument to access the multi-dimensional array:
heatmap.dimension_values('z', flat=False)
| examples/user_guide/08-Gridded_Datasets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LBA: Probability and Simulation
#
# ## Analysis
#
# Solve the following problems, one of which you will write yourself! Full solutions require clear steps with written justifications as well as an interpretation of the result. There are multiple approaches to solving these problems; you are encouraged to try solving the problems using Python. For example, the scipy.stats library includes useful tools such as a z-score calculator.
#
# ### 1) Volleyball
#
# “*There’s only a 35% chance that the team will win the point when they are serving.*” Contrast how Frequentists and Bayesians would interpret this statement (<200 words). [#probability]
# #### What would a Frequentist say?
#
# "Although I don't know the exact number of times the team will win the point when serving, I know that its value is fixed (not a random one). As a frequentist, I assert that the probable is that which happens often. Therefore, if I collect data from a sample of all volleyball games and estimate its expected value, 35% must be the relative frequency of winning while serving which is most consistent with the data."
#
# #### What about the Bayesian?
#
# "I agree that the mean is a fixed and unknown value, but I see no problem in representing the uncertainty probabilistically. I will do so by defining a probability distribution over the possible values of the expected value and use sample data to update the distribution that the team will win the point. Therefore, 35% is the degree of belief I have that the team will win the point when they are serving"
# Frequentists argue that probability is fundamentally related to the frequencies of repeated events where the parameters of interest are fixed and unchanging, whereas Bayesians concede that probability is fundamentally related to our knowledge about an event i.e. the prior and likelihood of observed data and the state of the world can always be updated.
# +
from IPython.display import Image
Image("Table.png")
# -
# ### 2) Basketball
#
# Tversky and his colleagues studied the records of 48 of the Philadelphia 76ers basketball games in the 1980–81 season to see whether a player had times when he was hot and every shot went in, and other times when he was cold and barely able to hit the backboard. The players estimated that they were about 25 percent more likely to make a shot after a hit than after a miss. In fact, the opposite was true—the 76ers were 6 percent more likely to score after a miss than after a hit. The authors report that the number of hot and cold streaks was about what one would expect by purely random effects. Assuming that a player has a fifty-fifty chance of making a shot and makes 20 shots a game, estimate by simulation (in Python) the proportion of the games in which the player will have a streak of 5 or more hits. Explain the result using a suitable interpretation of probability. [#probability, #simulation]
# +
import random
def shoot(p = 0.5):
return 1 if random.random() < p else 0
def one_game():
score = []
shot = 0
while shot < 20:
score.append(shoot())
shot += 1
return score
def n_game(n):
game = 0
desired = [1,1,1,1,1]
streak = []
while game < n:
outcome = one_game()
if str(desired)[1:-1] in str(outcome):
streak.append(1)
else:
streak.append(0)
game += 1
return streak
record = n_game(10000000)
probability = record.count(1)/len(record)
print(probability)
# -
# Since the free throw is a repeatable event, I propose that this probability is best viewed as a long run frequency. Therefore, the long-run average (i.e. in the limit as the number of shots approaches infinity) of a player having a streak of 5 or more hits is 25%. We assume as Frequentists that the repetitions occur independently and under essentially identical conditions. Note that in terms of relative frequency, there is no real difference between taking a shot 10 times in succession and taking shots all at once by 10 idential 76ers. The limitations of a frequentist approach is that we do now know what the limiting frequency will be, or that even one exists since it is not defined behorehand and we can't prove convergence. Furthermore, we cannot assign probabilities to single-case events.
#
# Through the above Monte Carlo simulation, we were able to test various outcome possibilities. This is useful because analysis of possibilities should be done before the fact, especially since human beings do not have a very good intuition for random processes. (Tversky et al., 1985) Studies across a number of sports show that 'streaks' don't exist. Independent attempts may yield occasional long strings of success or failures such as in the case of making a streak of 5 or more hits, but these are not sufficient evidence to conclude a relationship between successive shot attempts.
#
# On the other hand, Bayesian approaches are useful when it is difficult to assume that the event is repeatable and the collection of studies is a one-time phenomenon. In essence, a Bayesian would consider the 25% as the degree of belief that in 25% of the games, the player will have a streak of 5 or more hits.
# #### Source:
# <NAME>., <NAME>., & <NAME>. (1985). The hot hand in basketball: On the misperception of random sequences. Cognitive Psychology, 17(3), 295–314. https://doi.org/10.1016/0010-0285(85)90010-6
#
# ### 3) Baseball
#
# A rookie is brought to a baseball club on the assumption that he will have a 0.300 batting average based on his past performance. (Batting average is the ratio of the number of hits to the number of times at bat.) In the first year, he comes to bat 400 times during the season and his batting average is .348. Assume that his at bats can be considered Bernoulli trials with probability 0.3 for success. Give a both qualitative and quantitative argument about what is likely to happen to the player’s batting performance next season. Be sure to discuss the merits of any assumptions that are made. [#distributions]
# #### Quantitative
# We are using a binomial distribution to describe the mean and standard deviation of batting average. The assumptions relevant here include that each trial is independent. Although the belief of a streak or more accurately, 'hot-hand' persists, we have reasonable evidence to assume that the outcome of the player's fifth at-bat is unaffected by his performance in the first four at-bats. There are a fixed number of trials i.e. 400 and each at-bat can be considered a trial of the experiment. At each at-bat, there are two outcomes of interest – either the player gets a hit (success) or he doesn’t get a hit (failure). Since the player’s batting average is .300, the probability that he will get a hit in a single at-bat is p = .300 and this is the same for each trial.
# +
import math
from scipy import stats
np = 0.3*400
sd = math.sqrt(400*0.3*0.7)
nq = 0.7 * 400
print(np, nq)
# -
# When the sample size is large enough, the binomial distribution with parameters n and p can be approximated by the normal model. From the above cell, we can see that: $$np ≥ 10 $$ $$n(1-p)≥ 10$$
#
# Therefore, there are at least 10 expected successes and failures and a normal approximation is appropriate.
Image("Area.png")
prob = ((stats.norm(np, sd).cdf(139.2)) - 0.5)*2
print(prob)
# According to the normal approximation, the probability that the player will deviate from his batting average more than the same amount next season is 0.036. Therefore, there is a 0.963 likelihood that his batting average next season is between 100.8 and 139.2.
# #### Qualitative
# Because a player’s batting average in a given year of his career is an average of a very large number of statistically independent random variables, it might be expected to be normally distributed around its expected value. From this, the conclusion follows that the likelihood the player's batting performance for the next season is closer to the career batting average (population mean) is 0.963.
#
# This makes sense because a batting average of 0.348 is exceptional, but very unlikely. Therefore, following an extreme random event, the next random event is likely to be less extreme. This phenomena is called regression to the mean.
#
# Although baseball's best hitters can exceed 0.300, the league-wide batting average is typically around .260 ('What is a Batting Average (AVG)?', n.d.). Intuitively, most players who do much better than their peers are also performing better than their own career averages. These atheletes tend to be above average in both skill and in luck and only the skill portion is relevant to future performance. The regression-toward-the-mean explanation is that their skills did not deteriorate, but rather that their unusually good performance as a result of chance during the first season exaggerated their skills.
#
# Interestingly, since batting average regress towards the mean, baseball performances measures such as the batting average are considered imperfect assessments of underlying skill. Schall et al. suggests that by using 'correlation coefficients estimated from earlier seasons', predictions of batting averages can be improved to show reduced regression to the mean.
#
# Furthermore, the greater the size of trials, the more probable it will be that his battling average is closer to the population mean. This is because according to the central limit theorem, as the sample size exceeds 30, we closely approximate a normal distribution. Conversely, smaller sample sizes lead to a greater standard error and a wider distribution, such that the sample means are less likely to be clustered around the mean. This is because the Gaussian distribution is constrained by an area of 1. Therefore, for larger samples (400 at-bats), it is more probable that the mean is closer to the expected value.
#
# The degree to which the batting average is expected to regress towards the mean depends on the relative contribution of chance to the outcome: the greather the role of chance, the more the regression towards the mean. However, regression toward the mean does not occur for every individual i.e. we cannot entirely predict the athlete's performance in the next season.
# #### Source:
#
# What is a Batting Average (AVG)? | Glossary. (n.d.). Retrieved February 4, 2018, from http://m.mlb.com/glossary/standard-stats/batting-average
#
# <NAME>., & <NAME>. (2000). Do baseball players regress toward the mean? The American Statistician; Alexandria, 54(4), 231–235.
#
# ### 4) Ski jumping
#
# The distribution of all observed competitive ski jumps in Zakopane, Poland during 2010-2016 is nearly normal with an average jump distance of 120.5 m and a standard deviation of 10 m. [#distributions]
#
# 1. What is the probability that the next random jumper will jumper further than 135 m?
# 2. What is the probability that the mean distance of the next 15 random jumpers is further than 135 m?
# 3. Could you still estimate the probabilities above if the distribution was actually negatively skewed by the presence of some shorter distance jumps?
#
# +
m = 120.5
sd = 10
prob = 1 - (stats.norm(m, sd).cdf(135))
print('The probability that the next random jumper will jump further than 135 m is', prob)
se = 10/math.sqrt(15)
prob1 = 1 - (stats.norm(m, se).cdf(135))
#We are assuming that if we take a samples of size 15 from the population many many times
print('The probability that the mean distance of the next 15 random jumpers is further than 135 m is', prob1)
# -
# No, because we can't assume that the distribution of sample means is well approximated by a normal model since the data are strongly skewed and the sample size is very small (less than 30). The average of our sample means is no longer the population mean as the central limit theorem doesn't hold.
#
# The central limit theorem (CLT) indicates that if the sample size is sufficiently large (at least 30), the means of samples obtained using a random sampling with replacement are distributed normally regardless of the shape of the population distribution. Another condition for the CLT is independence onf observation. Generally, since the probability of ski jumping a certain distance stays the same regardless of the outcomes of previous ski jumps, we can assume that this conditions holds true.
#
# According to the CLT, even if the population distribution is skewed, the sampling distribution will be approximately normally distributed if the sample size is large enough. The larger each sample, the less spread out around the true population mean this distribution will be. The more closely the sample means gather symmetrically around the population mean, they have a corresponding reduction in the standard error. However, if our sample is strongly skewed, the CLT no longer applies and we cannot estimate the probabilities accurately.
#
# In principle, the researcher could decide to exclude extreme outliers to reduce skewness though such reasons need to be explicitly stated and its implications thoroughly considered. In such instances, the potential effects of any psychological biases e.g. seeking confirmatory information and giving dimished weight to evidence that contradicts one's positions (#confirmationbias) need to be carefully checked.
#
# ### 5) Construct your own problem.
#
# Write (and solve) a probability problem based on the data collected at the sporting event you attended. The analysis should contain multiple parts, ***enough to demonstrate deep knowledge of all three HC’s: #probability, #distributions, and #simulation.*** It must also involve computations done ***both analytically and via simulation***. Note that the same problem can be approached using multiple techniques. First discuss the context: explain the game you attended and what you measured. Also discuss all assumptions made and any potential flaws in your analysis. The (mostly complete) model problem linked in the assignment instructions is an example of what you could do. Do not copy this example exactly. If you’d like more ideas on what you could calculate, use textbook problems and class activities for inspiration, and brainstorm with your classmates or your professor.
#
# **Notes**:
#
# - The calculation may require some information not obtained in your data collection. You can make additional assumptions as long as the problem still uses your observation in some way.
# - Your problem and solution can have multiple parts but should not be much longer than the model problem provided.
# - Choose an appropriate level of difficulty. The problem should be solvable by your classmates, and WILL be solved by your classmates in lesson 3.2! So, if your analysis involves a distribution or concept not covered in class, include an APA citation as well as a description of the distribution and its relevance for the problem.
# - Don’t forget to include a photo of you at the game!
# I attended the men's basketball match between Arizona State Sun Devils and California Golden Bears College in Haas Pavilion, Berkeley.
#
# The objective of a basketball game is for one team to score more than the opposing team by the end of play time. Teams are made of 15 players with 5 players on the court at any one time. Shots made outside the arc score 3 points, while shots made inside the arc score 2 points. Any free throws score 1 point. We will assume the outcome of the game is the result of a series of random trials.
from IPython.display import Image
filename = "CalBears.jpg" #add the file name for the photo of you at the game
Image(filename)
# #### Assumptions and Limitations
#
# Here, each shot is a Bernoulli trial and trials are assumed to be independent:
# the outcome of a second attempt is not affected by the first attempt. There is a repeated, fixed number, n, of trials. In this case, n is the number of shots taken. Each trial has two outcomes: basket (success) or no basket (failure). The probability of success are assumed the same (identical) for each trial since free throws are always shot from the same distance and there is no defensive pressure. Each time the player lines up at the free throw line, he has the same probability of success. These probabilities are different for free throws and three-pointers, as determined by career statistics.
#
# However, the assumption of identical probabilities may be problematic for field goals, which are shot from varying distances and this can affect outcomes. Nonetheless, the law of large numbers states that for sufficiently large number of independent trials, the effect of such extraneous factors cancels each other out. As a result, our analysis holds.
#
# Here, we are going to use career statistics for each athlete instead of my observations at the game since the former average figures are taken over a longer period of time i.e. their entire professional career. According to the law of large numbers, they are likely to be closer to the expected values for each outcome. For instance, if a player has an intrinsic ability of 80% to make a free throw as the number of free throw attempts gets very large, the percent of successes will likely converge to 80% over their career.
# 1) If <NAME> shoots 3 free throws in a game, what is the probability that he will make at most 2 of them?
Image("NickHamilton2.png")
prob = stats.binom.cdf(2, 3, 0.667, loc = 0)
print(prob)
# 2) Does <NAME> have better odds of making 3 free throws in a row or one three pointer?
prob2 = (stats.binom.pmf(3, 3, 0.667, loc = 0))
print(prob2)
# If we imagine a billion <NAME>'s all taking a first free throw, we would expect on average that 66.7% would make that first free throw and the remaining would miss it. Of the ones that made the first one, we would expect 66.7% to make that first free throw and the second free throw and similarly of the 66.7% of the Nick Hamilton's that made the first and second free throw would also make the third free throw.
#
# Therefore, <NAME> has a probability of 0.25 to score one three pointer while a probability of 0.297 of making 3 free throws in a row. Therefore, he has better odds of making 3 free throws in a row.
# 3) <NAME> and <NAME> attempt 3 free throws. Their probability of making a shot is 66.7% and 76.7% respectively. Which player has a higher probability of making at least two shots?
Image("RemyMartin.png")
# +
nick = stats.binom.cdf(2,3,0.667, loc = 0)
remy = stats.binom.cdf(2,3,0.767, loc = 0)
print(nick)
print(remy)
# -
# <NAME> has a higher probability of making at least two shots.
# 4) How much spread is there in 10000 samples of size 100 <NAME> free throws? In other words, what is the standard error of the proportion?
Image("MarcusLee.png")
# +
import matplotlib.pyplot as plt
import numpy as np
import matplotlib.style
import matplotlib as mpl
mpl.style.use('classic')
def score(p = 0.495):
return 0 if random.random() < p else 1
def hundred ():
hits = []
shots = 0
while shots <100:
hits.append(score())
shots += 1
return hits.count(1)/shots #sample percentage from 100 free throws
def runsim(n):
sim = 0
samplemeans = []
while sim <n:
samplemeans.append(hundred())
sim += 1
return samplemeans
data = runsim(10000)
plt.hist(data)
plt.ylabel('Frequency')
plt.xlabel('Sample Means')
plt.show()
# -
print('The standard deviation of sample means is', np.std(data))
# +
#Method 2
sesp = math.sqrt((0.495*(1-0.495))/100) #standard error of sample proportion
print(sesp)
# -
# #### Source:
#
# The Standard Error of a Proportion. (n.d.). Retrieved February 5, 2018, from http://www.jerrydallal.com/lhsp/psd.htm
#
# We wanted to know how much spread there was in the sample proportion. We used both Monte Carlo simulation strategy and analytical methods. In the former, we repeatedly resampled and thereby obtained an approximation to the standard error of the sample proportion as 0.05. Analytical methods yielded an answer of approximately 0.05 as well.
# ### 6) Reflection
#
# Write a short reflection paragraph (< 200 words) about the insights you gained pertaining to the HCs used in this assignment. Which 1-2 learning principle(s) were the most beneficial for deepening your understanding of the connection between sports, probability, distributions, and simulation? [#scienceoflearning]
# Throughout this assignment, learning through storytelling was a beneficial strategy to understand the distinction between Bayesian and Frequentist interpretations of statistics. In practice, I created two active agents in my mind, referring to one as "Frequentist Frank" and the other as "Bayesian Bella" as characters I would learn more about while preparing for the assignment. Representing chunked information through active agents facilitates perception and recall (Kahneman, 2011). In particular, each piece of information can serve as a retrieval cue to the next related piece of information for the agent, creating an 'interlocking sequence of associations' (Kosslyn, 2017) and I was able to access these mental representations of abstract concepts easily after storytelling.
#
# Additionally, interleaved practice was very useful. By spacing out my studying over time instead of a single crammed session, I felt more confident engaging in the material with a clear understanding of the nuances of #probability and #distribution. For example, on one occasion I was struggling with the law of large numbers and decided to analyze a speech about racial inequality for my Multimodal Communications assignment. During the process, I had a revelation: getting the actual probability that an individual of color will suffer abuse by the police in the US is difficult because it requires gathering data on every single encounter of that subpopulation with law enforcement officers. Therefore, if the sample size is large enough, it will approach the expected value, and statistical analysis *is* a reliable method to characterize institutional racism. Geared with this new appreciation, I was far more motivated to work on my assignment, and it was a valuable learning experience.
# #### Source:
#
# <NAME>. (2011). Thinking, fast and slow (First edition.). New York: Farrar, Straus and Giroux.
#
#
# <NAME>. (2017). The science of learning. In <NAME> & <NAME> (Eds.), Working universities: Minerva and the future of higher education. Cambridge, MA: MIT Press. Retrieved from https://course-resources.minerva.kgi.edu/uploaded_files/mke/YRpz1r/chapter11-science-learning.pdf
#
| CS51_LBA (2).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: ml
# language: python
# name: ml
# ---
import pandas as pd
data = pd.read_csv('Data/notasCurso.csv')
data.info()
data.describe()
data[data['Nota primera oportunidad']>=60].count()
data[data['Nota segunda oportunidad']>=60].count()
data.loc[(data['Nota primera oportunidad']!=0) | (data['Nota segunda oportunidad']!=0),['Nota primera oportunidad','Nota segunda oportunidad']].count()
data.loc[(data['Nota primera oportunidad']==0) & (data['Nota segunda oportunidad']==0),['Nota primera oportunidad','Nota segunda oportunidad']].count()
data.loc[(data['Nota primera oportunidad']!=0) | (data['Nota segunda oportunidad']!=0),['Nota primera oportunidad','Nota segunda oportunidad']]
data.loc[(data['Nota primera oportunidad']>=60) & (data['Nota segunda oportunidad']>=60),['Nota primera oportunidad','Nota segunda oportunidad']]
# ### Se evalua los siguientes caso
# > El estudiante aprobo en el primer intento
# > El estudiante aprobo en el segundo intento
# > El estudiante aprobo en el primer intento y repitio para mejorar el puntaje
pasa = lambda x,y:int(x>=60 or y>=60)
def contar(nota1,nota2):
if nota2!=0:
return 2
if nota1!=0:
return 1
return 0
# ### Se considera una nota final mayor de 60 para aprobar el curso
data['aprobar'] = data.apply(lambda x: pasa(x['Nota primera oportunidad'],x['Nota segunda oportunidad']), axis=1)
data['notaMax']=data.apply(lambda x:max(x['Nota primera oportunidad'],x['Nota segunda oportunidad']), axis=1)
data['intento']=data.apply(lambda x:contar(x['Nota primera oportunidad'],x['Nota segunda oportunidad']), axis=1)
data
data.to_csv('Data/aprobados_reprobados.csv', index=False)
data['Nota primera oportunidad']
| tratamiento_de_datos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# # PyRadiomics toolbox in Python
# + [markdown] deletable=true editable=true
# In this notebook, we will load 2 brain datasets with segmentations of brain tumors. The `radiomics` package will be used to extract a set of features, and the "signatures" will be compared.
#
# For more detailed examples and links to source code, visit http://radiomics.io
# + deletable=true editable=true
# Radiomics package
from radiomics import featureextractor
import six, numpy as np
# + [markdown] deletable=true editable=true
# ## Setting up data
#
# Here we use `SimpleITK` (referenced as `sitk`, see http://www.simpleitk.org/ for details) to load two brain images and the corresponding segmentations as label maps.
# + deletable=true editable=true
import os
import SimpleITK as sitk
from radiomics import getTestCase
# repositoryRoot points to the root of the repository. The following line gets that location if this Notebook is run
# from it's default location in \pyradiomics\examples\Notebooks
repositoryRoot = os.path.abspath(os.path.join(os.getcwd(), ".."))
imagepath_1, labelpath_1 = getTestCase('brain1', repositoryRoot)
imagepath_2, labelpath_2 = getTestCase('brain2', repositoryRoot)
image_1 = sitk.ReadImage(imagepath_1)
label_1 = sitk.ReadImage(labelpath_1)
image_2 = sitk.ReadImage(imagepath_2)
label_2 = sitk.ReadImage(labelpath_2)
# + [markdown] deletable=true editable=true
# ## Show the images
#
# Using `matplotlib.pyplot` (referenced as `plt`), display the images in grayscale and labels in color.
# + deletable=true editable=true
# Display the images
# %matplotlib inline
import matplotlib.pyplot as plt
plt.figure(figsize=(20,20))
# First image
plt.subplot(2,2,1)
plt.imshow(sitk.GetArrayFromImage(image_1)[12,:,:], cmap="gray")
plt.title("Brain #1")
plt.subplot(2,2,2)
plt.imshow(sitk.GetArrayFromImage(label_1)[12,:,:])
plt.title("Segmentation #1")
# Second image
plt.subplot(2,2,3)
plt.imshow(sitk.GetArrayFromImage(image_2)[8,:,:], cmap="gray")
plt.title("Brain #2")
plt.subplot(2,2,4)
plt.imshow(sitk.GetArrayFromImage(label_2)[8,:,:])
plt.title("Segmentation #2")
plt.show()
# + [markdown] deletable=true editable=true
# ## Extract the features
#
# Using the `radiomics` package, first construct an `extractor` object from the parameters set in `Params.yaml`. Then generate features for the two images.
# + deletable=true editable=true
import os
# Instantiate the extractor
params = os.path.join(os.getcwd(), '..', 'examples', 'exampleSettings', 'Params.yaml')
extractor = featureextractor.RadiomicsFeaturesExtractor(params)
result_1 = extractor.execute(image_1, label_1)
result_2 = extractor.execute(image_2, label_2)
# + [markdown] deletable=true editable=true
# ## Prepare for plotting
#
# Because we'd like to plot the feature vectors, create `numpy` arrays for features starting with `original_` (excluding meta-features).
# + deletable=true editable=true
# Make an array of the values
feature_1 = np.array([])
feature_2 = np.array([])
for key, value in six.iteritems(result_1):
if key.startswith("original_"):
feature_1 = np.append ( feature_1, result_1[key])
feature_2 = np.append ( feature_2, result_2[key])
# + [markdown] deletable=true editable=true
# ## Plot
#
# Plot the two feature vectors and the difference. Feature values have a wide range of magnitudes and are plotted on a log scale.
# + deletable=true editable=true
plt.figure(figsize=(20,20))
plt.subplot(3,1,1)
plt.plot(feature_1)
plt.yscale('log')
plt.title ( "Features from brain #1")
plt.subplot(3,1,2)
plt.plot(feature_2)
plt.yscale('log')
plt.title ( "Features from brain #1")
plt.subplot(3,1,3)
plt.plot(feature_1 - feature_2)
plt.yscale('log')
plt.title ( "Difference")
plt.show()
| notebooks/RadiomicsExample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 作業 : (Kaggle)鐵達尼生存預測
# # [作業目標]
# - 試著完成三種不同特徵類型的三種資料操作, 觀察結果
# - 思考一下, 這三種特徵類型, 哪一種應該最複雜/最難處理
# # [作業重點]
# - 完成剩餘的八種 類型 x 操作組合 (In[6]~In[13], Out[6]~Out[13])
# - 思考何種特徵類型, 應該最複雜
# +
# 載入基本套件
import pandas as pd
import numpy as np
# 讀取訓練與測試資料
data_path = 'data/'
df_train = pd.read_csv(data_path + 'titanic_train.csv')
df_test = pd.read_csv(data_path + 'titanic_test.csv')
df_train.shape
# -
# 重組資料成為訓練 / 預測用格式
train_Y = df_train['Survived']
ids = df_test['PassengerId']
df_train = df_train.drop(['PassengerId', 'Survived'] , axis=1)
df_test = df_test.drop(['PassengerId'] , axis=1)
df = pd.concat([df_train,df_test])
df.head()
# 秀出資料欄位的類型與數量
dtype_df = df.dtypes.reset_index()
dtype_df.columns = ["Count", "Column Type"]
dtype_df = dtype_df.groupby("Column Type").aggregate('count').reset_index()
dtype_df
#確定只有 int64, float64, object 三種類型後, 分別將欄位名稱存於三個 list 中
int_features = []
float_features = []
object_features = []
for dtype, feature in zip(df.dtypes, df.columns):
if dtype == 'float64':
float_features.append(feature)
elif dtype == 'int64':
int_features.append(feature)
else:
object_features.append(feature)
print(f'{len(int_features)} Integer Features : {int_features}\n')
print(f'{len(float_features)} Float Features : {float_features}\n')
print(f'{len(object_features)} Object Features : {object_features}')
# # 作業1
# * 試著執行作業程式,觀察三種類型 (int / float / object) 的欄位分別進行( 平均 mean / 最大值 Max / 相異值 nunique )
# 中的九次操作會有那些問題? 並試著解釋那些發生Error的程式區塊的原因?
#
# Ans: object type 無平均值可言
#
# # 作業2
# * 思考一下,試著舉出今天五種類型以外的一種或多種資料類型,你舉出的新類型是否可以歸在三大類中的某些大類?
# 所以三大類特徵中,哪一大類處理起來應該最複雜?
#
# Ans: 有向圖或樹狀結構類型,可以歸類再object中,object中涵蓋非整數及浮點數類別,所以處理上必須費心思探究其實際的意義。
# 例 : 整數 (int) 特徵取平均 (mean)
df[int_features].mean()
# 請依序列出 三種特徵類型 (int / float / object) x 三種方法 (平均 mean / 最大值 Max / 相異值 nunique) 的其餘操作
"""
Your Code Here
"""
print('-' * 80)
print('[INT]')
print('mean:\n', df[int_features].mean(), '\n')
print('max:\n', df[int_features].max(), '\n')
print('nunique:\n', df[int_features].nunique(), '\n')
print('-' * 80)
print('[float]')
print('mean:\n', df[float_features].mean(), '\n')
print('max:\n', df[float_features].max(), '\n')
print('nunique:\n', df[int_features].nunique(), '\n')
print('-' * 80)
print('[object]')
print('mean:\n', df[object_features].mean(), '\n')
print('max:\n', df[object_features].max(), '\n')
print('nunique:\n', df[object_features].nunique(), '\n')
print('-' * 80)
| homeworks/D018/Day_018_HW.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="bTwRw3xNX5cj" colab_type="code" colab={}
import tensorflow as tf
# + id="X2m2QySzYw0W" colab_type="code" outputId="61b7b210-e5de-4c4b-bf6e-87be6f6bc0b1" colab={"base_uri": "https://localhost:8080/", "height": 68}
import nltk
nltk.download('stopwords')
# + id="8BbO35yiYA01" colab_type="code" colab={}
import csv
import tensorflow as tf
import numpy as np
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
from nltk.corpus import stopwords
from string import punctuation
from sklearn.model_selection import train_test_split
#STOPWORDS = set(stopwords.words('portuguese'))
STOPWORDS = set(stopwords.words('portuguese') + list(punctuation))
# + id="aCM22r2KZExP" colab_type="code" colab={}
# Parâmetros Principais
# Tamanho do vocabulário a ser criado com o tokenizer. Este considera as N palavras mais comuns (ou frequentes nos textos)
vocab_size = 5000
# Tamanho do vetor de características que representa cada palavra. Cada palavra é transformada para um vetor com 64 entradas.
embedding_dim = 64
# Tamanho máximo da sequência de códigos que representa um texto (tamanho do feature vector)
max_length = 35
# Tipo de truncagem a ser utilizado, 'post' significa remover valores do final da sequência.
trunc_type = 'post'
# Tipo de padding a ser utilizado, 'post' significa completar no final da sequência.
padding_type = 'post'
# Token a ser utilizado quando uma palavra não presente no vocabulário é encontrada no texto
Not_known = '<NKN>'
# Porcentagem de instâncias a ser utilizada no treinamento
training_portion = .7
# + id="loZOoQUMZJZE" colab_type="code" outputId="86796e5f-7d49-4423-e534-dcff9ed89cdb" colab={"base_uri": "https://localhost:8080/", "height": 139}
# Carrega os textos. Em X o texto de entrada e em y os rótulos.
# Cada linha do texto carrega é filtrada para retirada de stopwords.
def ler(file):
with open(file, 'r', encoding='latin-1') as csvfile:
texto = csv.reader(csvfile, delimiter=';') # lê um texto do arquivo
print(texto)
next(texto)
for linha in texto: # processa cada linha do texto
y.append(linha[2])
aux = linha[0]
for word in STOPWORDS: # retirada de stopwords
token = ' ' + word + ' '
aux = aux.replace(token, ' ')
aux = aux.replace(' ', ' ')
X.append(aux)
X = []
y = []
ler('g1_v2_ws.csv')
ler('g1_v1_ws.csv')
print(len(y)) # quantidade de rótulos
print(len(X)) # quantidade de textos
train_X, validation_X, train_y, validation_y = train_test_split(X,y,test_size=.3, random_state=42, stratify=y)
print(train_y)
print(validation_y)
# + id="eCZn9vA_Z2Zn" colab_type="code" colab={}
# Cria o vocabulário a partir da base de treinamento considerando o tamanho definido em vocab_size.
# Utiliza como coringa o símbolo Not_known
tokenizer = Tokenizer(num_words = vocab_size, oov_token=Not_known)
tokenizer.fit_on_texts(train_X)
word_index = tokenizer.word_index
# + id="IJ5RBL6NZ81j" colab_type="code" outputId="0ccdf8c0-75a4-4be3-b7d6-dc32e64f3b94" colab={"base_uri": "https://localhost:8080/", "height": 187}
# Lista os N primeiros vocábulos do dicionários (os N mais frequentes)
N=10
dict(list(word_index.items())[0:N])
# + id="BDSnnl_MaC5e" colab_type="code" colab={}
# Converte uma linha de texto em uma sequência de valores
train_sequences = tokenizer.texts_to_sequences(train_X)
# + id="Zids558vaHzB" colab_type="code" outputId="7b9d719c-9d22-499f-f012-a5b17a106d8d" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Mostra uma linha de texto convertida para sequência de valores
# Cada valor representa uma palavra do vocabulário
print(train_sequences[5])
# + id="s2jxWfWVaNPR" colab_type="code" colab={}
# Transforma todas as sequências para um tamanho fixo. Sequências pequenas são completadas e sequências maiores que o limite são truncadas
train_padded = pad_sequences(train_sequences, maxlen=max_length, padding=padding_type, truncating=trunc_type)
# + id="eXpamqjxaTfL" colab_type="code" outputId="ec1bacd4-d980-438d-845d-f9e5fb49c329" colab={"base_uri": "https://localhost:8080/", "height": 119}
print(len(train_sequences[0]))
print(len(train_padded[0]))
print(len(train_sequences[1]))
print(len(train_padded[1]))
print(len(train_sequences[10]))
print(len(train_padded[10]))
# + id="o4uSyQj2aZrY" colab_type="code" outputId="67fe20db-73d7-4364-c297-7f6dda8eb5a8" colab={"base_uri": "https://localhost:8080/", "height": 68}
# Imprime uma sequência
print(train_padded[6])
# + id="Job1hIYQafUq" colab_type="code" outputId="674986d1-bd7c-4d4f-b319-8f75b1ea9371" colab={"base_uri": "https://localhost:8080/", "height": 51}
# Tokeniza a base de validação.
validation_sequences = tokenizer.texts_to_sequences(validation_X)
validation_padded = pad_sequences(validation_sequences, maxlen=max_length, padding=padding_type, truncating=trunc_type)
print(len(validation_sequences))
print(validation_padded.shape)
# + id="PJmSci_Ram7C" colab_type="code" outputId="12985b5f-25c2-4745-d825-69b6598a3eb0" colab={"base_uri": "https://localhost:8080/", "height": 34}
# Mostra o conjunto de rótulos
print(set(y))
# + id="El9KrDsdasAU" colab_type="code" colab={}
# Tokeniza os rótulos
label_tokenizer = Tokenizer()
label_tokenizer.fit_on_texts(y)
# Observe que subtraímos 1 dos códigos para estes comecem em 0
training_label_seq = np.array(label_tokenizer.texts_to_sequences(train_y))-1
validation_label_seq = np.array(label_tokenizer.texts_to_sequences(validation_y))-1
# + id="KtA4DaEpawPj" colab_type="code" outputId="ba4b4602-9604-41c5-bef8-f2402ffa6d05" colab={"base_uri": "https://localhost:8080/", "height": 153}
print(training_label_seq[0])
print(training_label_seq[1])
print(training_label_seq[400])
print(training_label_seq.shape)
print(validation_label_seq[0])
print(validation_label_seq[1])
print(validation_label_seq[50])
print(validation_label_seq.shape)
# + id="KN6zfnEVa2FN" colab_type="code" outputId="e50e6790-13be-4f0e-9c7a-afd614cfa51e" colab={"base_uri": "https://localhost:8080/", "height": 88}
# Confere como ficaram as nossas frases depois de transformdas
# Apenas para conferência.
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
def decode_article(text):
return ' '.join([reverse_word_index.get(i, '?') for i in text])
print(decode_article(train_padded[10]))
print('---')
print(train_X[10])
# + id="7G3sPURXa-V7" colab_type="code" outputId="141dba50-6e72-449f-cd24-c71b84097bd7" colab={"base_uri": "https://localhost:8080/", "height": 629}
# Criando a rede LSTM (Long Short Term Memory)
hidden_size=64
model = tf.keras.Sequential()
model.add(tf.keras.layers.Embedding(vocab_size, embedding_dim))
#model.add(tf.keras.layers.LSTM(embedding_dim, dropout = 0.25 , recurrent_dropout=0.25))
model.add(tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(embedding_dim)))
model.add(tf.keras.layers.Dense(64, activation='relu'))
model.add(tf.keras.layers.Dense(7, activation='softmax'))
model.summary()
# Compilando a LSTM
model.compile(loss='sparse_categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# Treinamento da LSTM
num_epochs = 10
history = model.fit(train_padded, training_label_seq, epochs=num_epochs, validation_data=(validation_padded, validation_label_seq), verbose=2)
# + id="tPxVfIYSzD2j" colab_type="code" outputId="f15c6b4f-8645-4d5f-8aeb-d695e59f7a0e" colab={"base_uri": "https://localhost:8080/", "height": 629}
# Criando uma RNN (Recurrent Neural Network)
model = tf.keras.Sequential()
model.add(tf.keras.layers.Embedding(input_dim=vocab_size, output_dim=64, input_length=max_length))
# The output of SimpleRNN will be a 2D tensor of shape (batch_size, 128)
model.add(tf.keras.layers.SimpleRNN(units=64))
model.add(tf.keras.layers.Dense(64, activation='relu'))
model.add(tf.keras.layers.Dense(7, activation='softmax'))
model.summary()
# Compilando a RNN
#model.compile(optimizer='rmsprop', loss='binary_crossentropy', metrics=['acc'])
model.compile(loss='sparse_categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# Treinamento da RNN
history = model.fit(train_padded, training_label_seq, batch_size=32, epochs=10, validation_data=(validation_padded, validation_label_seq), verbose=2)
# + id="lA5tWjiAbOmn" colab_type="code" outputId="ccfff88d-854c-495a-e781-3e4c13e7878d" colab={"base_uri": "https://localhost:8080/", "height": 542}
# Imprime gráfico histórico do treinamento
import matplotlib.pyplot as plt
def plot_graphs(history, string):
plt.plot(history.history[string])
plt.plot(history.history['val_'+string])
plt.xlabel("Epochs")
plt.ylabel(string)
plt.legend([string, 'val_'+string])
plt.show()
plot_graphs(history, "accuracy")
plot_graphs(history, "loss")
# + id="ODeyBE44bVnT" colab_type="code" outputId="db83ddc9-c296-4ee4-dca9-33921b1628e1" colab={"base_uri": "https://localhost:8080/", "height": 102}
# Testando uma frase
txt = ["Clientes de shopping se assustam com suposta bomba deixada por trio durante assalto. Os bandidos deixaram a suposta bomba em cima do balcão da loja, deixando os funcionários e clientes assustados."]
seq = tokenizer.texts_to_sequences(txt)
padded = pad_sequences(seq, maxlen=max_length, padding=padding_type, truncating=trunc_type)
pred = model.predict(padded)
labels = ['desgosto', 'medo', 'surpresa', 'raiva', 'tristeza', 'alegria', 'neutro']
print(pred, labels[np.argmax(pred)])
print("Resultado na validacao:")
score=model.evaluate(validation_padded, validation_label_seq, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
# + id="-Lyitqlc2eIr" colab_type="code" colab={}
# Plotar a matrix de confusão
def plot_confusion_matrix(cm, classes,
normalize=False,
title=None,
cmap=plt.cm.Blues):
plt.imshow(cm, interpolation = 'nearest',cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation = 45)
plt.yticks(tick_marks, classes)
if normalize:
cm = cm.astype('float')/cm.sum(axis=1)[:,np.newaxis]
print("Normalized confusion matrix")
else:
print("Confusion matrix, without normalization")
thresh = cm.max()/2
for i, j in itertools.product(range(cm.shape[0]),range(cm.shape[1])):
plt.text(j, i, cm[i,j], horizontalalignment="center", color="white" if cm[i,j]>thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# + id="JxYCvgOi16r2" colab_type="code" outputId="d8cd1332-77a7-454d-e216-9e8de16bf6b6" colab={"base_uri": "https://localhost:8080/", "height": 328}
# Preparar Matriz de Confusão
import itertools
from sklearn.metrics import confusion_matrix
y_predict=model.predict_classes(validation_padded)
val=validation_label_seq.reshape(len(validation_label_seq))
cm=confusion_matrix(val, y_predict)
cm_plot_labels=['desgosto', 'medo', 'surpresa', 'raiva', 'tristeza', 'alegria', 'neutro']
plot_confusion_matrix(cm, cm_plot_labels, title='Confusion matrix')
| 04-deep-learning/02-rnn-lstm/Text_Classification_LSTM_emotions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.0 64-bit
# language: python
# name: python38064bit1060d4750c904259afeb7847dfa8ded2
# ---
# + id="yJ-CkY5ghQW-"
import numpy as np
import pandas as pd
import scipy.signal
# + colab={"base_uri": "https://localhost:8080/"} id="X3AlPtoNhQXA" outputId="605b70ba-4ead-4e75-fc45-32cf1d44b33e"
pip install vmdpy
# + id="CAwClapVhQXB"
from vmdpy import VMD
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.preprocessing import LabelEncoder
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import classification_report,accuracy_score,f1_score,precision_score,recall_score
# + id="y4eaDOYxhQXB"
train = pd.read_csv('train.tsv',sep='\t')
# + id="XqVEFjXuhQXC"
def format_text(df,col):
#Remove @ tags
comp_df = df.copy()
# remove all the punctuation
comp_df[col] = comp_df[col].str.replace(r'(@\w*)','')
#Remove URL
comp_df[col] = comp_df[col].str.replace(r"http\S+", "")
#Remove # tag and the following words
comp_df[col] = comp_df[col].str.replace(r'#\w+',"")
#Remove all non-character
comp_df[col] = comp_df[col].str.replace(r"[^a-zA-Z ]","")
# Remove extra space
comp_df[col] = comp_df[col].str.replace(r'( +)'," ")
comp_df[col] = comp_df[col].str.strip()
# Change to lowercase
comp_df[col] = comp_df[col].str.lower()
comp_df[col] = comp_df[col].str.replace('httpurl', '')
return comp_df
# + id="wjpgPX7QhQXC"
train = format_text(train,'Text')
# + id="MU9EkEbVhQXE"
test = pd.read_csv('test.tsv',sep='\t',header=None)
test = format_text(test,1)
# + id="jAXYAb5JhQXE"
X_X = train['Text'].tolist()
Y_train = train['Label']
# + id="0S7q2aLzhQXF"
le = LabelEncoder()
le.fit(Y_train)
Y_train = le.transform(Y_train)
# + id="wAaaeOALhQXF"
tfidf = TfidfVectorizer(sublinear_tf=True, min_df=5, norm='l2', encoding='latin-1', ngram_range=(1, 2), stop_words='english')
features = tfidf.fit_transform(X_X).toarray()
# + id="ebi1wD6xhQXF"
def energy(u):
# Estimate PSD `S_xx_welch` at discrete frequencies `f_welch`
f_welch, S_xx_welch = scipy.signal.welch(u)
# Integrate PSD over spectral bandwidth
# to obtain signal power `P_welch`
df_welch = f_welch[1] - f_welch[0]
return np.sum(S_xx_welch) * df_welch
# + id="HS5Ln_BHhQXG"
def maxvdm(f,alpha):
tau = 0.
K = 3
DC = 0
init = 1
tol = 1e-7
u, u_hat, omega = VMD(f, alpha, tau, K, DC, init, tol)
energy_array=[]
for i in u:
energy_array.append(energy(i))
ind = np.argmax(energy_array)
return u[ind]
# + id="abePeSUehQXG"
X_X_1 = test[1].tolist()
# + id="p6-XC6d7hQXG"
features_1 = tfidf.transform(X_X_1).toarray()
# + id="_mJ4qixuhQXG"
Y_test = le.transform(test[2])
# + id="cdrB1sgFhQXH"
models = [
RandomForestClassifier(n_estimators=200, max_depth=3, random_state=0),
LogisticRegression(random_state=0),
KNeighborsClassifier(n_neighbors=3)
]
# + id="anWlxO10hQXH"
models_acc = {"RandomForestClassifier":[],"LogisticRegression":[],"KNeighborsClassifier":[]}
models_f1 = {"RandomForestClassifier":[],"LogisticRegression":[],"KNeighborsClassifier":[]}
models_pre = {"RandomForestClassifier":[],"LogisticRegression":[],"KNeighborsClassifier":[]}
models_re = {"RandomForestClassifier":[],"LogisticRegression":[],"KNeighborsClassifier":[]}
# + colab={"base_uri": "https://localhost:8080/"} id="EfFAZLLBhQXI" outputId="0efb93b4-ee64-4be9-b0d5-c6bb3cb390ac"
for k in range(1,20):
print("K-Value Tuning :",k,"- out of 20 ")
X_train = []
for i in features:
X_train.append(maxvdm(i,k))
print("\tTraining Data Done..")
X_test = []
for i in features_1:
X_test.append(maxvdm(i,k))
print("\tTesting Data Done..")
for i in models:
i.fit(X_train, Y_train)
y_pred = i.predict(X_test)
acc = accuracy_score(Y_test, y_pred)
f1 = f1_score(Y_test, y_pred)
pre = precision_score(Y_test, y_pred)
re = recall_score(Y_test, y_pred)
name = i.__class__.__name__
models_acc[name].append(acc)
models_f1[name].append(f1)
models_pre[name].append(pre)
models_re[name].append(re)
# + id="cc8I9BT7hQXI"
A = pd.DataFrame([models_acc,models_f1,models_pre,models_re])
# + id="iuyFIAoEhQXI"
A.to_csv("Hyper_alpha-1to20.csv")
| notebooks/ScratchPad/TFIDF_VMD_Hyptertune-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ***Introduction to Radar Using Python and MATLAB***
# ## <NAME> - Copyright (C) 2019 Artech House
# <br/>
#
# # LFM Train Ambiguity Function
# ***
# Referring to Section 8.6.2, the ambiguity function for a LFM train is found using the generic waveform method outlined in Section 8.6.3.
# ***
# Begin by getting the library path
import lib_path
# Set the pulsewidth (s), the bandwidth (Hz), the pulse repetition interval (s) and the number of pulses
# +
pulsewidth = 0.4
bandwidth = 10.0
pri = 1.0
number_of_pulses = 6
# -
# Generate the time delay (s) using the `linspace` routine from `scipy`
# +
from numpy import linspace
# Set the time delay
time_delay = linspace(-number_of_pulses * pri, number_of_pulses * pri, 5000)
# -
# Calculate the ambiguity function for the LFM train
# +
from Libs.ambiguity.ambiguity_function import lfm_train
from numpy import finfo
ambiguity = lfm_train(time_delay, finfo(float).eps, pulsewidth, bandwidth, pri, number_of_pulses)
# -
# Plot the zero-Doppler cut using the `matplotlib` routines
# +
from matplotlib import pyplot as plt
# Set the figure size
plt.rcParams["figure.figsize"] = (15, 10)
# Plot the ambiguity function
plt.plot(time_delay, ambiguity, '')
# Set the x and y axis labels
plt.xlabel("Time (s)", size=12)
plt.ylabel("Relative Amplitude", size=12)
# Turn on the grid
plt.grid(linestyle=':', linewidth=0.5)
# Set the plot title and labels
plt.title('LFM Train Ambiguity Function', size=14)
# Set the tick label size
plt.tick_params(labelsize=12)
# -
# Set the Doppler mismatch frequencies using the `linspace` routine
doppler_frequency = linspace(-bandwidth, bandwidth, 5000)
# Calculate the ambiguity function for the LFM train
ambiguity = lfm_train(finfo(float).eps, doppler_frequency, pulsewidth, bandwidth, pri, number_of_pulses)
# Display the zero-range cut for the LFM train
# +
plt.plot(doppler_frequency, ambiguity, '')
# Set the x and y axis labels
plt.xlabel("Doppler (Hz)", size=12)
plt.ylabel("Relative Amplitude", size=12)
# Turn on the grid
plt.grid(linestyle=':', linewidth=0.5)
# Set the plot title and labels
plt.title('LFM Train Ambiguity Function', size=14)
# Set the tick label size
plt.tick_params(labelsize=12)
# -
# Set the time delay and Doppler mismatch frequency and create the two-dimensional grid using the `meshgrid` routine from `scipy`
# +
from numpy import meshgrid
# Set the time delay
time_delay = linspace(-number_of_pulses * pri, number_of_pulses * pri, 1000)
# Set the Doppler mismatch
doppler_frequency = linspace(-bandwidth, bandwidth, 1000)
# Create the grid
t, f = meshgrid(time_delay, doppler_frequency)
# -
# Calculate the ambiguity function for the LFM train
ambiguity = lfm_train(t, f, pulsewidth, bandwidth, pri, number_of_pulses)
# Display the two-dimensional contour plot for the LFM train ambiguity function
# +
# Plot the ambiguity function
from numpy import finfo
plt.contour(t, f, ambiguity + finfo('float').eps, 30, cmap='jet', vmin=-0.2, vmax=1.0)
# Set the x and y axis labels
plt.xlabel("Time (s)", size=12)
plt.ylabel("Doppler (Hz)", size=12)
# Turn on the grid
plt.grid(linestyle=':', linewidth=0.5)
# Set the plot title and labels
plt.title('LFM Pulse Ambiguity Function', size=14)
# Set the tick label size
plt.tick_params(labelsize=12)
# -
| jupyter/Chapter08/lfm_train_ambiguity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from sklearn.externals.joblib import Memory
from sklearn.datasets import load_svmlight_file
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import random
#数据读取和分割
data = load_svmlight_file("a9a.txt")
x = data[0]
y = data[1]
x = x.toarray()
y = y.T
#补全X
add_num = np.zeros(int(x.size/122))
x = np.column_stack([x,add_num])
x_train,x_test,y_train,y_test = train_test_split(x,y,test_size=0.33, random_state=1)
#学习率、C、权重的设置
rating = 0.003
C = 0.9
weight = np.ones(124)
weight_NAG = weight
weight_RMSProp = weight
weight_AdaDelta = weight
weight_Adam = weight
#训练集和测试集的长度
train_length = int(x_train.size/123)
test_length = int(x_test.size/123)
#测试集和训练集的截距设置
weight0 = np.ones(train_length)
weight1 = np.ones(test_length)
x_train = np.column_stack([x_train,weight0])
x_test = np.column_stack([x_test,weight1])
#验证集的loss数组初始化
loss_test = np.array([])
loss_NAG = np.array([])
loss_RMSProp = np.array([])
loss_AdaDelta = np.array([])
loss_Adam = np.array([])
#累计Adam迭代次数
iter = 1
#定义分类器
def svm_classification(x,y,length,weight,rating,C,k,loss,ways,iter):
#准确率计数
temp = 0
#四种优化方法的参数初始化
jw = 0
Vt_NAG = 0
Gt_RMSProp = 0
Gt_AdaDelta = 0
theta_AdaDelta = 0
delta_AdaDelta = 0
Gt_Adam = 0
Mt_Adam = 0
alpha_Adam = 0
epsilon = 0.00001
#500批梯度下降
for j in range(500):
upsilon = 1 - np.dot(y[k + j],np.dot(weight.T,x[k + j].T))
if(upsilon > 0):
gradient = 1/C * weight + np.dot(x[k + j],y[k + j])
#损失函数累加
jw = jw + upsilon
if(upsilon <= 0):
gradient = 1/C * weight
#NAG优化方法
if(ways == 1):
gradient = (gradient - 1/C * 0.9 * Vt_NAG)/500
Vt_NAG = 0.9*Vt_NAG + rating * gradient
weight = weight - Vt_NAG
#RMSProp优化方法
if(ways == 2):
gradient = gradient/500
Gt_RMSProp = 0.9*Gt_RMSProp + 0.1*np.dot(gradient,gradient)
weight = weight - (rating/np.sqrt(Gt_RMSProp + epsilon)) * gradient
#AdaDelta
if(ways == 3):
gradient = gradient/500
Gt_AdaDelta = 0.95*Gt_AdaDelta + 0.05*np.dot(gradient,gradient)
theta_AdaDelta = -np.dot((np.sqrt(delta_AdaDelta + epsilon)/np.sqrt(Gt_AdaDelta + epsilon)),gradient)
weight = weight + theta_AdaDelta
delta_AdaDelta = 0.95*delta_AdaDelta + 0.05*np.dot(theta_AdaDelta,theta_AdaDelta)
#Adam优化方法
if(ways == 4):
gradient = gradient/500
Mt_Adam = 0.9*Mt_Adam + 0.1*gradient
Gt_Adam = 0.999*Gt_Adam + 0.001*np.dot(gradient,gradient)
alpha_Adam = rating*(np.sqrt(1-0.999/(np.sqrt(iter)))/(1-(0.9/np.sqrt(iter))))
weight = weight - alpha_Adam * (Mt_Adam/(np.sqrt(Gt_Adam + epsilon)))
iter = iter + 1
#预测值
y_p = np.dot(weight.T,x[k + j].T)
#验证预测结果(阈值设定,计算准确率)
if(y_p > 6):
y_predict = 1.0
if(y_p <= 6):
y_predict = -1.0
if(y_predict == y[k + j]):
temp = temp + 1
#损失函数
jw = np.dot(weight,weight)/2 + C * jw
loss = np.append(loss,jw/500)
return loss,weight
#训练和测试四种优化方法,并迭代200次
for i in range(200):
loss_NAG,weight_NAG = svm_classification(x_test,y_test,test_length,weight_NAG,rating,C,np.random.randint(0,test_length - 502),loss_NAG,1,iter)
loss_RMSProp,weight_RMSProp = svm_classification(x_test,y_test,test_length,weight_RMSProp,rating,C,np.random.randint(0,test_length - 502),loss_RMSProp,2,iter)
loss_AdaDelta,weight_AdaDelta = svm_classification(x_test,y_test,test_length,weight_AdaDelta,rating,C,np.random.randint(0,test_length - 502),loss_AdaDelta,3,iter)
loss_Adam,weight_Adam = svm_classification(x_test,y_test,test_length,weight_Adam,rating,C,np.random.randint(0,test_length - 502),loss_Adam,4,iter)
#图显示
plt.figure()
plt.yscale('log')
plt.gca().set_xlabel('iter times')
plt.gca().set_ylabel('loss scale')
#蓝色的为NAG,黑色的为RMSProp,绿色的为AdaDelta,红色的为Adam
plt.plot(loss_NAG,'blue',label='loss_NAG')
plt.plot(loss_RMSProp,'black',label='loss_RMSProp')
plt.plot(loss_AdaDelta,'green',label='loss_AdaDelta')
plt.plot(loss_Adam,'red',label='loss_Adam')
plt.legend()
plt.show()
# -
| ClassificationExperiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from unidecode import unidecode
import nltk
from nltk.corpus import stopwords
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.feature_extraction.text import CountVectorizer
nltk.download('stopwords')
df = pd.read_csv('../base/review.csv',encoding='latin-1')
df.head()
# +
import string
from nltk.stem.snowball import SnowballStemmer
import swifter
import nltk
stemmer = SnowballStemmer("english")
stop = set(stopwords.words('english'))
def lower(texto):
return texto.lower()
def normalize(texto):
return unidecode(texto)
def remove_ponctuation(texto):
for punc in string.punctuation:
texto = texto.replace(punc," ")
return texto
def remove_stopwords(texto):
ret = []
for palavra in texto.split():
if palavra not in stop:
ret.append(palavra)
return ' '.join(ret)
def stem(texto):
ret = []
for palavra in texto.split():
ret.append(stemmer.stem(palavra))
return ' '.join(ret)
def remove_number(texto):
result = ''.join([i for i in texto if not i.isdigit()])
return result
def pipeline(texto):
texto = normalize(texto)
texto = lower(texto)
texto = remove_ponctuation(texto)
texto = remove_stopwords(texto)
texto = remove_number(texto)
texto = stem(texto)
return texto
# -
df['SentimentText'].apply(lower).head()
remove_ponctuation("é, ué!")
len(df)
df['preproc'] = df['SentimentText'].swifter.apply(pipeline)
# +
# vectorizer = CountVectorizer()
# X = vectorizer.fit_transform(df['preproc'])
# len(vectorizer.get_feature_names())
# -
vectorizer_tfidf = TfidfVectorizer()
X = vectorizer_tfidf.fit_transform(df['preproc'])
len(vectorizer_tfidf.get_feature_names())
y = df['Sentiment']
# +
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import cross_val_score
import numpy as np
clf = LogisticRegression(solver='liblinear')
np.mean(cross_val_score(clf,X, y, cv=10,scoring='balanced_accuracy'))
# -
from sklearn.naive_bayes import MultinomialNB
clf = MultinomialNB()
np.mean(cross_val_score(clf,X, y, cv=10,scoring='balanced_accuracy'))
clf.fit(X,y)
import pickle
filename = 'clf.pickle'
outfile = open(filename,'wb')
pickle.dump(clf,outfile)
outfile.close()
filename = 'vectorizer.pickle'
outfile = open(filename,'wb')
pickle.dump(vectorizer_tfidf,outfile)
outfile.close()
# +
#I just love this movie. Specially the climax, seriously one of the best climax I have ever seen.
# +
#I just want to say how amazing this film is from start to finish. This will take you on a emotional ride.You will not he disappointed
# +
#LITERALLY , one of the best movies i have seen in my entire life , filled with a tone of action and emotions . you will love avenger endgame . ' i love you 3000 '
| 4.1_review_texts/review_filmes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python3
# name: python3
# ---
# # Notebook without solutions
#
# The idea is with the use of classes, we can decide whether to show or not the solutions
# of a particular lecture, creating two different types of jupyter notebooks. For now it only
# works with *code blocks*, you have to include **:class: solution**, and set in the conf.py file
# *jupyter_drop_solutions=True*.
#
# Here is a small example
# ## Question 1
#
# Plot the area under the curve
#
# $$
# f(x)=\sin(4\pi x) exp(-5x)
# $$
#
# when $ x \in [0,1] $
| tests/base/ipynb/solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf
# language: python
# name: tf
# ---
# Importing the Keras libraries and packages
from keras.models import Sequential
from keras.layers import Convolution2D
from keras.layers import MaxPooling2D
from keras.layers import Flatten
from keras.layers import Dense
import matplotlib.pyplot as plt
from PIL import Image
import numpy as np
from sklearn.metrics import classification_report, confusion_matrix
import pandas as pd
import os
# Initialize CNN
model = Sequential()
# +
# building model
model.add(Convolution2D(64, 3, 3, input_shape = (64, 64, 3), activation = 'relu'))
# pooling
model.add(MaxPooling2D(pool_size = (2,2)))
# adding a second convol layer
model.add(Convolution2D(64, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2,2)))
# Flattening
model.add(Flatten())
# Full connection
model.add(Dense(units=128, activation='relu'))
# output layer
model.add(Dense(units=9, activation='softmax'))
# -
# compile
model.compile(optimizer = 'adam', loss='categorical_crossentropy', metrics=['accuracy'])
# Fitting the CNN to the images(Image Augmentation, Image Preprocessing)
from keras.preprocessing.image import ImageDataGenerator
train_datagen = ImageDataGenerator(
rescale=1./255,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True)
# normalizing
test_datagen = ImageDataGenerator(rescale=1./255)
# path to your experimental data
os.chdir('/home/sachin_sharma/Desktop/jpg_data')
# This section will create Training set( Training_Set:TestSet ratio 80:20)
training_set = train_datagen.flow_from_directory(
'TrainingSet',
target_size=(64, 64),
batch_size=32,
class_mode='categorical')
# This section will create the Test set
test_set = test_datagen.flow_from_directory(
'TestSet',
target_size=(64, 64),
batch_size=32,
class_mode='categorical',
shuffle=False)
history = model.fit_generator(training_set,
steps_per_epoch=(training_set.samples/32),
epochs=25,
validation_data=test_set,
validation_steps=(test_set.samples/32))
# saving the model
model.save('exp1_b_1.h5')
# Visualizing the mapping between labels
training_set.class_indices
# +
# Confusion Matrix
Y_pred = model.predict_generator(test_set, test_set.samples//32 +1 )
y_pred = np.argmax(Y_pred, axis=1)
print('Confusion Matrix')
#cm = confusion_matrix(test_set.classes, y_pred)
#visulaizing
def cm2df(cm, labels):
df = pd.DataFrame()
# rows
for i, row_label in enumerate(labels):
rowdata={}
# columns
for j, col_label in enumerate(labels):
rowdata[col_label]=cm[i,j]
df = df.append(pd.DataFrame.from_dict({row_label:rowdata}, orient='index'))
return df[labels]
#calling
df = cm2df(cm, ["AnnualCrop", "Buildup", "Forest", "HerbaceousVegetation", "Highway", "Pasture", "PermanentCrop", "River", "SeaLake"])
print(df)
# -
# Classification report
print('Classification Report')
target_names = ['AnnualCrop','Buildup','Forest', 'HerbaceousVegetation', 'Highway', 'Pasture', 'PermanentCrop', 'River', 'SeaLake']
classificn_report = classification_report(test_set.classes, y_pred, target_names=target_names)
print(classificn_report)
# +
# Plotting the Loss and Classification Accuracy
model.metrics_names
print(history.history.keys())
# "Accuracy"
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('Model Accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# "Loss"
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# -
| Results/exp1_b_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
import os
import numpy as np
import pandas as pd
import torch
import torch.nn as nn
import torch.optim as optim
import torch.utils.data
from torch.autograd import Variable
from sklearn.utils import shuffle
from torchsample.initializers import Uniform
from torchsample.modules import ModuleTrainer
from torchsample.metrics import CategoricalAccuracy
# %aimport torchsample.modules
# %matplotlib inline
# -
use_cuda = False
batch_size = 64
# # Setup data
# We're going to look at the IMDB dataset, which contains movie reviews from IMDB, along with their sentiment. Keras comes with some helpers for this dataset.
from keras.datasets import imdb
idx = imdb.get_word_index()
# This is the word list:
idx_arr = sorted(idx, key=idx.get)
idx_arr[:10]
# ...and this is the mapping from id to word
idx2word = {v: k for k, v in idx.items()}
# We download the reviews using code copied from keras.datasets:
# +
from keras.utils.data_utils import get_file
import pickle
path = get_file('imdb_full.pkl',
origin='https://s3.amazonaws.com/text-datasets/imdb_full.pkl',
md5_hash='d091312047c43cf9e4e38fef92437263')
f = open(path, 'rb')
(x_train, labels_train), (x_test, labels_test) = pickle.load(f)
# -
len(x_train)
# Here's the 1st review. As you see, the words have been replaced by ids. The ids can be looked up in idx2word.
', '.join(map(str, x_train[0]))
# The first word of the first review is 23022. Let's see what that is.
idx2word[23022]
# Here's the whole review, mapped from ids to words.
' '.join([idx2word[o] for o in x_train[0]])
# The labels are 1 for positive, 0 for negative.
labels_train_tensor = torch.from_numpy(np.array(labels_train))
labels_test_tensor = torch.from_numpy(np.array(labels_test))
labels_train[:10]
# Reduce vocab size by setting rare words to max index.
# +
vocab_size = 5000
trn = [np.array([i if i < vocab_size - 1 else vocab_size - 1 for i in s]) for s in x_train]
test = [np.array([i if i < vocab_size - 1 else vocab_size - 1 for i in s]) for s in x_test]
# -
# Look at distribution of lengths of sentences.
lens = np.array(list(map(len, trn)))
(lens.max(), lens.min(), lens.mean())
# Pad (with zero) or truncate each sentence to make consistent length.
# +
seq_len = 500
from keras.preprocessing import sequence
trn = sequence.pad_sequences(trn, maxlen=seq_len, value=0)
test = sequence.pad_sequences(test, maxlen=seq_len, value=0)
trn_tensor = torch.from_numpy(trn).long()
test_tensor = torch.from_numpy(test).long()
# -
# This results in nice rectangular matrices that can be passed to ML algorithms. Reviews shorter than 500 words are pre-padded with zeros, those greater are truncated.
trn_tensor.size()
# ## Create simple models
# ### Single hidden layer NN
# The simplest model that tends to give reasonable results is a single hidden layer net. So let's try that. Note that we can't expect to get any useful results by feeding word ids directly into a neural net - so instead we use an embedding to replace them with a vector of 32 (initially random) floats for each word in the vocab.
# +
import torch.nn as nn
import torch.nn.functional as F
class SingleHiddenLayerModule(nn.Module):
def __init__(self):
super().__init__()
num_dimensions = 32
self.embedding = nn.Embedding(vocab_size, num_dimensions)
self.fc1 = nn.Linear(seq_len * num_dimensions, 100)
self.dropout = nn.Dropout(0.7)
self.fc2 = nn.Linear(100, 2)
self.init()
def forward(self, words_ids):
x = self.embedding(words_ids) # x => torch.Size([64, 500, 32])
x = x.view(x.size(0), -1) # x => torch.Size([64, 16000])
x = self.fc1(x)
x = F.relu(x, True)
x = self.dropout(x)
x = self.fc2(x)
# result = F.sigmoid(x)
result = x
return result
def init(self):
torch.nn.init.constant(self.fc1.bias, val=0.0)
torch.nn.init.constant(self.fc2.bias, val=0.0)
# +
# %autoreload 2
# criterion = nn.BCELoss()
criterion = nn.CrossEntropyLoss()
model = SingleHiddenLayerModule()
if(use_cuda):
model.cuda()
criterion.cuda()
trainer = ModuleTrainer(model)
trainer.set_optimizer(optim.Adam, lr=1e-3)
trainer.set_loss(criterion)
trainer.set_initializers([Uniform(module_filter="embedding*", a=-0.05, b=0.05), XavierUniform(module_filter="fc*")])
trainer.set_metrics([CategoricalAccuracy()])
# trainer.summary((trn_tensor.size(0), labels_train_tensor.size(0)))
model
# -
trainer.fit(trn_tensor, labels_train_tensor, validation_data=(test_tensor, labels_test_tensor),
nb_epoch=2, batch_size=batch_size, shuffle=True)
# The [stanford paper](http://ai.stanford.edu/~amaas/papers/wvSent_acl2011.pdf) that this dataset is from cites a state of the art accuracy (without unlabelled data) of 0.883. ~~So we're short of that, but on the right track.~~ We've already beaten the state of the art in 2011 with a simple Neural Net.
# ### Single conv layer with max pooling
# A CNN is likely to work better, since it's designed to take advantage of ordered data. We'll need to use a 1D CNN, since a sequence of words is 1D.
# +
import torch.nn as nn
import torch.nn.functional as F
class CnnMaxPoolingModule(nn.Module):
def __init__(self):
super().__init__()
num_dimensions = 32
self.embedding = nn.Embedding(vocab_size, num_dimensions)
self.drop1 = nn.Dropout(0.2)
self.conv1 = nn.Conv1d(in_channels=32, out_channels=64, kernel_size=5, padding=2, groups=1)
self.fc1 = nn.Linear(seq_len * num_dimensions, 100)
self.dropout = nn.Dropout(0.7)
self.fc2 = nn.Linear(100, 2)
self.init()
def forward(self, words_ids):
x = self.embedding(words_ids) # x => torch.Size([B, 500, 32])
x = x.permute(0, 2, 1)
# print('emb', x.size())
x = self.drop1(x) # x => torch.Size([B, 500, 32])
x = self.conv1(x) # x => torch.Size([B, 500, 64])
x = F.relu(x, True)
# print('conv1', x.size())
x = self.drop1(x) # x => torch.Size([B, 500, 64])
x = F.max_pool1d(x, kernel_size=2)
# print('max', x.size())
x = x.view(x.size(0), -1)
# print(x.size())
x = self.fc1(x)
x = F.relu(x, True)
x = self.dropout(x)
x = self.fc2(x)
# result = F.sigmoid(x)
result = x
#raise 'Error'
return result
def init(self):
torch.nn.init.constant(self.conv1.bias, val=0.0)
torch.nn.init.constant(self.fc1.bias, val=0.0)
torch.nn.init.constant(self.fc2.bias, val=0.0)
# +
# %autoreload 2
# criterion = nn.BCELoss()
criterion = nn.CrossEntropyLoss()
model = CnnMaxPoolingModule()
if(use_cuda):
model.cuda()
criterion.cuda()
trainer = ModuleTrainer(model)
trainer.set_optimizer(optim.Adam, lr=1e-3)
trainer.set_loss(criterion)
trainer.set_initializers([Uniform(module_filter="embedding*", a=-0.05, b=0.05), XavierUniform(module_filter="fc*"), XavierUniform(module_filter="conv*")])
trainer.set_metrics([CategoricalAccuracy()])
# trainer.summary((trn_tensor.size(0), labels_train_tensor.size(0)))
model
# -
trainer.fit(trn_tensor, labels_train_tensor, validation_data=(test_tensor, labels_test_tensor),
nb_epoch=2, batch_size=batch_size, shuffle=True)
trainer.fit(trn_tensor, labels_train_tensor, validation_data=(test_tensor, labels_test_tensor),
nb_epoch=4, batch_size=batch_size, shuffle=True)
# ### Pre-trained vectors
# You may want to look at wordvectors.ipynb before moving on.
#
# In this section, we replicate the previous CNN, but using pre-trained embeddings.
# +
import torch
import re
from torchtext.vocab import load_word_vectors
wv_dict, wv_arr, wv_size = load_word_vectors('.', 'glove.6B', 50)
print('Loaded', len(wv_arr), 'words')
# -
#
# The glove word ids and imdb word ids use different indexes. So we create a simple function that creates an embedding matrix using the indexes from imdb, and the embeddings from glove (where they exist).
# +
def get_word(word):
return wv_arr[wv_dict[word]]
def create_emb():
num_dimensions_glove = wv_arr.size()[1]
embedding = nn.Embedding(vocab_size, num_dimensions_glove)
# If we can't find the word in glove, randomly initialize
torch.nn.init.uniform(embedding.weight, a=-0.05, b=0.05)
num_found, num_not_found = 0, 0
for i in range(1,len(embedding.weight)):
word = idx2word[i]
if word and re.match(r"^[a-zA-Z0-9\-]*$", word):
embedding.weight.data[i] = get_word(word)
num_found += 1
else:
num_not_found +=1
# This is our "rare word" id - we want to randomly initialize
torch.nn.init.uniform(embedding.weight.data[-1], a=-0.05, b=0.05)
embedding.weight.requires_grad = False
# This speeds up training. Can it be replaced by BatchNorm1d?
embedding.weight.data /= 3
print("Words found: {}, not found: {}".format(num_found, num_not_found))
return embedding
# -
#
# We pass our embedding matrix to the Embedding constructor, and set it to non-trainable.
# +
import torch.nn as nn
import torch.nn.functional as F
class CnnMaxPoolingModuleWithEmbedding(nn.Module):
def __init__(self, embedding):
super().__init__()
num_dimensions = 32
self.embedding = embedding
self.drop1 = nn.Dropout(0.25)
self.batchnorm = nn.BatchNorm1d(500)
self.conv1 = nn.Conv1d(in_channels=embedding.weight.size()[1], out_channels=64, kernel_size=5, padding=2, groups=1)
self.fc1 = nn.Linear(seq_len * num_dimensions, 100)
self.dropout = nn.Dropout(0.7)
self.fc2 = nn.Linear(100, 2)
self.init()
def forward(self, words_ids):
x = self.embedding(words_ids)
# x = self.batchnorm(x)
x = x.permute(0, 2, 1)
x = self.drop1(x)
x = self.conv1(x)
x = F.relu(x, True)
x = self.drop1(x)
x = F.max_pool1d(x, kernel_size=2)
x = x.view(x.size(0), -1)
x = self.fc1(x)
x = F.relu(x, True)
x = self.dropout(x)
x = self.fc2(x)
result = x
return result
def init(self):
torch.nn.init.constant(self.conv1.bias, val=0.0)
torch.nn.init.constant(self.fc1.bias, val=0.0)
torch.nn.init.constant(self.fc2.bias, val=0.0)
def parameters(self):
p = filter(lambda p: p.requires_grad, nn.Module.parameters(self))
return p
# +
# %autoreload 2
emb = create_emb()
# criterion = nn.BCELoss()
criterion = nn.CrossEntropyLoss()
model = CnnMaxPoolingModuleWithEmbedding(emb)
if(use_cuda):
model.cuda()
criterion.cuda()
trainer = ModuleTrainer(model)
trainer.set_optimizer(optim.Adam, lr=1e-3)
trainer.set_loss(criterion)
trainer.set_initializers([XavierUniform(module_filter="fc*"), XavierUniform(module_filter="conv*")])
trainer.set_metrics([CategoricalAccuracy()])
# trainer.summary((trn_tensor.size(0), labels_train_tensor.size(0)))
# -
trainer.fit(trn_tensor, labels_train_tensor, validation_data=(test_tensor, labels_test_tensor),
nb_epoch=10, batch_size=batch_size, shuffle=True)
# We already have beaten our previous model! But let's fine-tune the embedding weights - especially since the words we couldn't find in glove just have random embeddings.
model.embedding.weight.requires_grad = True
trainer = ModuleTrainer(model)
trainer.set_optimizer(optim.Adam, lr=1e-4)
trainer.set_loss(criterion)
trainer.set_metrics([CategoricalAccuracy()])
trainer.fit(trn_tensor, labels_train_tensor, validation_data=(test_tensor, labels_test_tensor),
nb_epoch=1, batch_size=batch_size, shuffle=True)
# ### Multi-size CNN
# This is an implementation of a multi-size CNN as shown in <NAME>' excellent blog post.
# We create multiple conv layers of different sizes, and then concatenate them.
# +
import torch.nn as nn
import torch.nn.functional as F
class CnnMaxPoolingModuleMultiSizeWithEmbedding(nn.Module):
def __init__(self, embedding):
super().__init__()
num_dimensions = 32
self.embedding = embedding
self.drop1 = nn.Dropout(0.25)
self.batchnorm = nn.BatchNorm1d(500)
self.convs = [self.create_conv(embedding, fsz) for fsz in range (3, 6)]
self.fc1 = nn.Linear(25000, 100)
self.dropout = nn.Dropout(0.7)
self.fc2 = nn.Linear(100, 2)
self.init()
def create_conv(self, embedding, fsz):
return nn.Conv1d(in_channels=embedding.weight.size()[1], out_channels=64, kernel_size=5, padding=2, groups=1)
def conv(self, c, x):
x = c(x)
x = F.relu(x, True)
x = self.drop1(x)
x = F.max_pool1d(x, kernel_size=2)
return x
def forward(self, words_ids):
x = self.embedding(words_ids)
x = x.permute(0, 2, 1)
x = self.drop1(x)
convs = [self.conv(conv, x) for conv in self.convs]
torch.cat(convs, dim=1)
x = x.view(x.size(0), -1)
x = self.fc1(x)
x = F.relu(x, True)
x = self.dropout(x)
x = self.fc2(x)
result = x
return result
def init(self):
torch.nn.init.constant(self.fc1.bias, val=0.0)
torch.nn.init.constant(self.fc2.bias, val=0.0)
for conv in self.convs:
torch.nn.init.xavier_uniform(conv.weight.data, gain=1.0)
torch.nn.init.constant(conv.bias, val=0.0)
def parameters(self):
p = filter(lambda p: p.requires_grad, nn.Module.parameters(self))
return p
# +
# %autoreload 2
emb = create_emb()
criterion = nn.CrossEntropyLoss()
model = CnnMaxPoolingModuleMultiSizeWithEmbedding(emb)
model.embedding.weight.requires_grad = True
if(use_cuda):
model.cuda()
criterion.cuda()
trainer = ModuleTrainer(model)
trainer.set_optimizer(optim.Adam, lr=1e-3)
trainer.set_loss(criterion)
trainer.set_initializers([XavierUniform(module_filter="fc*")])
trainer.set_metrics([CategoricalAccuracy()])
# -
trainer.fit(trn_tensor, labels_train_tensor, validation_data=(test_tensor, labels_test_tensor),
nb_epoch=10, batch_size=batch_size, shuffle=True)
# This is clearly over-fitting. But it does get the highest accuracy on validation set.
# ### LSTM
# We haven't covered this bit yet!
# +
import torch.nn as nn
import torch.nn.functional as F
class LstmEmbeddingModule(nn.Module):
def __init__(self):
super().__init__()
num_dimensions = 32
self.num_hidden = 100
self.embedding = nn.Embedding(vocab_size, num_dimensions)
self.drop1 = nn.Dropout(0.2)
self.lstm1 = nn.LSTM(input_size=32, hidden_size=self.num_hidden, num_layers=1, batch_first=True)
self.fc1 = nn.Linear(50000, 2)
self.hidden = self.init_hidden(batch_size)
self.init()
def forward(self, words_ids):
# We detach the hidden state from how it was previously produced.
# If we didn't, the model would try backpropagating all the way to start of the dataset.
# self.hidden = self.repackage_hidden(self.hidden)
x = self.embedding(words_ids)
x = self.drop1(x)
#print('embd', x.size())
self.hidden = self.init_hidden(x.size(0))
#lenghts = [vocab_size for _ in range(x.size(0))]
#x = torch.nn.utils.rnn.pack_padded_sequence(x, lenghts, batch_first=True)
#print('pack', x.data.size())
x, self.hidden = self.lstm1(x, self.hidden)
#print('lstm', x.data.size())
#x, _ = torch.nn.utils.rnn.pad_packed_sequence(x, batch_first=True)
#print('unpk', x.size())
# print(self.hidden)
# TODO can we get rid of contiguous?
x = x.contiguous().view(x.size(0), -1)
#print('view', x.size())
x = self.fc1(x)
x = F.relu(x, True)
return x
def init(self):
torch.nn.init.constant(self.fc1.bias, val=0.0)
def init_hidden(self, batch_size):
num_layers = 1
weight = next(self.parameters()).data
return (Variable(weight.new(num_layers, batch_size, self.num_hidden).zero_()),
Variable(weight.new(num_layers, batch_size, self.num_hidden).zero_()))
def repackage_hidden(self, h):
"""Wraps hidden states in new Variables, to detach them from their history."""
if type(h) == Variable:
return Variable(h.data)
else:
return tuple(self.repackage_hidden(v) for v in h)
# +
# %autoreload 2
criterion = nn.CrossEntropyLoss()
model = LstmEmbeddingModule()
if(use_cuda):
model.cuda()
criterion.cuda()
trainer = ModuleTrainer(model)
trainer.set_optimizer(optim.Adam, lr=1e-3)
trainer.set_loss(criterion)
# TODO init LSTM
trainer.set_initializers([Uniform(module_filter="embedding*", a=-0.05, b=0.05), XavierUniform(module_filter="fc*"), XavierUniform(module_filter="conv*")])
trainer.set_metrics([CategoricalAccuracy()])
# -
# TODO figure out how to do this in PyTorch
trainer.fit(trn_tensor, labels_train_tensor, validation_data=(test_tensor, labels_test_tensor),
nb_epoch=5, batch_size=batch_size, shuffle=True)
| lesson5-pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: masif_env
# language: python
# name: masif_env
# ---
# +
# #!/usr/bin/python
import sys
import os
import subprocess as sp
import io
import networkx as nx
from importlib import reload
root_path = sp.run(['git', 'rev-parse', '--show-toplevel'], stdout=sp.PIPE).stdout.decode('utf-8')[:-1]
sys.path.append(os.path.join(root_path, 'source'))
sys.path.append(os.path.join(root_path, 'source', 'matlab_libs'))
import my_utils as my
from geometry.vertices_graph import vertices_graph
from data_preparation.extract_and_triangulate_lib import *
class IpyExit(SystemExit):
"""Exit Exception for IPython.
Exception temporarily redirects stderr to buffer.
"""
def __init__(self):
# print("exiting") # optionally print some message to stdout, too
# ... or do other stuff before exit
sys.stderr = io.StringIO()
def __del__(self):
sys.stderr.close()
sys.stderr = sys.__stderr__ # restore from backup
# +
args = ['1Z0K_uR_1000.pdb', 'A', '1Z0K_C_36457.pdb', 'A', '2.0']
ground_truth_cut_dist = float(args[4]) if len(args) > 4 else 2.0
to_reload = (args[5] == '1') if (len(args) > 5) else True
u_pdb_filename, u_chain_name, u_pdb_filepath, u_chain_filepath_base, u_chain_filepath = \
parse_names(args[0:2], tmp_dir=masif_opts["tmp_dir"])
if(to_reload):
# Extract chains of interest.
extractPDB(u_pdb_filepath, u_chain_filepath, u_chain_name)
# construct the mesh.
u_regular_mesh, u_vertex_normals, u_vertices, u_names = \
msms_wrap(u_chain_filepath)
u_vertex_hbond, u_vertex_hphobicity, u_vertex_charges = \
compute_features(u_chain_filepath_base, u_vertices, u_names, u_regular_mesh)
ply_filepath = u_chain_filepath_base + '_d' + str(ground_truth_cut_dist) + '.ply'
if 'compute_iface' in masif_opts and masif_opts['compute_iface']:
C_pdb_filename, C_chain_name, C_pdb_filepath, C_chain_filepath_base, C_chain_filepath = \
parse_names(args[2:4])
C_regular_mesh, C_vertex_normals, C_vertices, C_names = \
msms_wrap(C_pdb_filepath)
#iface = find_iface(C_regular_mesh, u_regular_mesh, ground_truth_cut_dist)
kdt = KDTree(C_regular_mesh.vertices)
d, r = kdt.query(u_regular_mesh.vertices)
d = np.square(d) # Square d, because this is how it was in the pyflann version.
assert(len(d) == len(u_regular_mesh.vertices))
iface_v = np.where(d >= ground_truth_cut_dist)[0]
iface = np.zeros(len(u_regular_mesh.vertices))
iface[iface_v] = 1.0
G, edges = vertices_graph(u_regular_mesh, weighted=False)
save_ply(ply_filepath, u_regular_mesh.vertices,\
u_regular_mesh.faces, normals=u_vertex_normals, charges=u_vertex_charges,\
normalize_charges=True, hbond=u_vertex_hbond, hphob=u_vertex_hphobicity,\
iface=iface)
else:
save_ply(ply_filepath, u_regular_mesh.vertices,\
u_regular_mesh.faces, normals=u_vertex_normals, charges=u_vertex_charges,\
normalize_charges=True, hbond=u_vertex_hbond, hphob=u_vertex_hphobicity)
copy_tmp2dst(ply_filepath, masif_opts['ply_chain_dir'])
copy_tmp2dst(u_chain_filepath, masif_opts['pdb_chain_dir'])
# +
import geometry.vertices_graph
geometry.vertices_graph = reload(geometry.vertices_graph)
G = geometry.vertices_graph.vertices_graph(u_regular_mesh, weighted=False)
N_verts = len(u_regular_mesh.vertices)
print(nx.is_connected(G))
not_iface_v = []
for v in range(N_verts):
if(not v in iface_v):
not_iface_v.append(v)
not_iface_v = np.array(not_iface_v)
G.remove_nodes_from(not_iface_v)
print(nx.is_connected(G))
G_comps = [np.array(list(c)) for c in nx.connected_components(G)]
t_iface = []
for Gc in G_comps:
G_comps_len = len(Gc)
if(G_comps_len > 20):
t_iface.append(Gc)
t_iface = [max(G_comps, key=len)]
t_iface = np.concatenate(t_iface)
print(t_iface)
#print(nx.is_connected(G0))
# +
#list(G.nodes(data=True))
# -
G0 = G
iface_v
# +
# #vertices_graph??
# -
| source/data_preparation/02-ecrt&trg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Python Language Basics, IPython, and Jupyter Notebooks
import numpy as np
np.random.seed(12345)
np.set_printoptions(precision=4, suppress=True)
# ## The Python Interpreter
# ```python
# $ python
# Python 3.6.0 | packaged by conda-forge | (default, Jan 13 2017, 23:17:12)
# [GCC 4.8.2 20140120 (Red Hat 4.8.2-15)] on linux
# Type "help", "copyright", "credits" or "license" for more information.
# >>> a = 5
# >>> print(a)
# 5
# ```
# ```python
# print('Hello world')
# ```
# ```python
# $ python hello_world.py
# Hello world
# ```
# ```shell
# $ ipython
# Python 3.6.0 | packaged by conda-forge | (default, Jan 13 2017, 23:17:12)
# Type "copyright", "credits" or "license" for more information.
#
# IPython 5.1.0 -- An enhanced Interactive Python.
# ? -> Introduction and overview of IPython's features.
# # %quickref -> Quick reference.
# help -> Python's own help system.
# object? -> Details about 'object', use 'object??' for extra details.
#
# In [1]: %run hello_world.py
# Hello world
#
# In [2]:
# ```
# ## IPython Basics
# ### Running the IPython Shell
# $
import numpy as np
data = {i : np.random.randn() for i in range(7)}
data
# >>> from numpy.random import randn
# >>> data = {i : randn() for i in range(7)}
# >>> print(data)
# {0: -1.5948255432744511, 1: 0.10569006472787983, 2: 1.972367135977295,
# 3: 0.15455217573074576, 4: -0.24058577449429575, 5: -1.2904897053651216,
# 6: 0.3308507317325902}
# ### Running the Jupyter Notebook
# ```shell
# $ jupyter notebook
# [I 15:20:52.739 NotebookApp] Serving notebooks from local directory:
# /home/wesm/code/pydata-book
# [I 15:20:52.739 NotebookApp] 0 active kernels
# [I 15:20:52.739 NotebookApp] The Jupyter Notebook is running at:
# http://localhost:8888/
# [I 15:20:52.740 NotebookApp] Use Control-C to stop this server and shut down
# all kernels (twice to skip confirmation).
# Created new window in existing browser session.
# ```
# ### Tab Completion
# ```
# In [1]: an_apple = 27
#
# In [2]: an_example = 42
#
# In [3]: an
# ```
# ```
# In [3]: b = [1, 2, 3]
#
# In [4]: b.
# ```
# ```
# In [1]: import datetime
#
# In [2]: datetime.
# ```
# ```
# In [7]: datasets/movielens/
# ```
# ### Introspection
# ```
# In [8]: b = [1, 2, 3]
#
# In [9]: b?
# Type: list
# String Form:[1, 2, 3]
# Length: 3
# Docstring:
# list() -> new empty list
# list(iterable) -> new list initialized from iterable's items
#
# In [10]: print?
# Docstring:
# print(value, ..., sep=' ', end='\n', file=sys.stdout, flush=False)
#
# Prints the values to a stream, or to sys.stdout by default.
# Optional keyword arguments:
# file: a file-like object (stream); defaults to the current sys.stdout.
# sep: string inserted between values, default a space.
# end: string appended after the last value, default a newline.
# flush: whether to forcibly flush the stream.
# Type: builtin_function_or_method
# ```
# ```python
# def add_numbers(a, b):
# """
# Add two numbers together
#
# Returns
# -------
# the_sum : type of arguments
# """
# return a + b
# ```
# ```python
# In [11]: add_numbers?
# Signature: add_numbers(a, b)
# Docstring:
# Add two numbers together
#
# Returns
# -------
# the_sum : type of arguments
# File: <ipython-input-9-6a548a216e27>
# Type: function
# ```
# ```python
# In [12]: add_numbers??
# Signature: add_numbers(a, b)
# Source:
# def add_numbers(a, b):
# """
# Add two numbers together
#
# Returns
# -------
# the_sum : type of arguments
# """
# return a + b
# File: <ipython-input-9-6a548a216e27>
# Type: function
# ```
# ```python
# In [13]: np.*load*?
# np.__loader__
# np.load
# np.loads
# np.loadtxt
# np.pkgload
# ```
# ### The %run Command
# ```python
# def f(x, y, z):
# return (x + y) / z
#
# a = 5
# b = 6
# c = 7.5
#
# result = f(a, b, c)
# ```
# ```python
# In [14]: %run ipython_script_test.py
# ```
# ```python
# In [15]: c
# Out [15]: 7.5
#
# In [16]: result
# Out[16]: 1.4666666666666666
# ```
# ```python
# >>> %load ipython_script_test.py
#
# def f(x, y, z):
# return (x + y) / z
#
# a = 5
# b = 6
# c = 7.5
#
# result = f(a, b, c)
# ```
# #### Interrupting running code
# ### Executing Code from the Clipboard
# ```python
# x = 5
# y = 7
# if x > 5:
# x += 1
#
# y = 8
# ```
# ```python
# In [17]: %paste
# x = 5
# y = 7
# if x > 5:
# x += 1
#
# y = 8
# ## -- End pasted text --
# ```
# ```python
# In [18]: %cpaste
# Pasting code; enter '--' alone on the line to stop or use Ctrl-D.
# :x = 5
# :y = 7
# :if x > 5:
# : x += 1
# :
# : y = 8
# :--
# ```
# ### Terminal Keyboard Shortcuts
# ### About Magic Commands
# ```python
# In [20]: a = np.random.randn(100, 100)
#
# In [20]: %timeit np.dot(a, a)
# 10000 loops, best of 3: 20.9 µs per loop
# ```
# ```python
# In [21]: %debug?
# Docstring:
# ::
#
# # %debug [--breakpoint FILE:LINE] [statement [statement ...]]
#
# Activate the interactive debugger.
#
# This magic command support two ways of activating debugger.
# One is to activate debugger before executing code. This way, you
# can set a break point, to step through the code from the point.
# You can use this mode by giving statements to execute and optionally
# a breakpoint.
#
# The other one is to activate debugger in post-mortem mode. You can
# activate this mode simply running %debug without any argument.
# If an exception has just occurred, this lets you inspect its stack
# frames interactively. Note that this will always work only on the last
# traceback that occurred, so you must call this quickly after an
# exception that you wish to inspect has fired, because if another one
# occurs, it clobbers the previous one.
#
# If you want IPython to automatically do this on every exception, see
# the %pdb magic for more details.
#
# positional arguments:
# statement Code to run in debugger. You can omit this in cell
# magic mode.
#
# optional arguments:
# --breakpoint <FILE:LINE>, -b <FILE:LINE>
# Set break point at LINE in FILE.
#
# ```
# ```python
# In [22]: %pwd
# Out[22]: '/home/wesm/code/pydata-book
#
# In [23]: foo = %pwd
#
# In [24]: foo
# Out[24]: '/home/wesm/code/pydata-book'
# ```
# ### Matplotlib Integration
# ```python
# In [26]: %matplotlib
# Using matplotlib backend: Qt4Agg
# ```
# ```python
# In [26]: %matplotlib inline
# ```
# ## Python Language Basics
# ### Language Semantics
# #### Indentation, not braces
# ```python
# for x in array:
# if x < pivot:
# less.append(x)
# else:
# greater.append(x)
# ```
# ```python
# a = 5; b = 6; c = 7
# ```
# #### Everything is an object
# #### Comments
# ```python
# results = []
# for line in file_handle:
# # keep the empty lines for now
# # if len(line) == 0:
# # continue
# results.append(line.replace('foo', 'bar'))
# ```
# ```python
# print("Reached this line") # Simple status report
# ```
# #### Function and object method calls
# ```
# result = f(x, y, z)
# g()
# ```
# ```
# obj.some_method(x, y, z)
# ```
# ```python
# result = f(a, b, c, d=5, e='foo')
# ```
# #### Variables and argument passing
a = [1, 2, 3]
b = a
a.append(4)
b
# ```python
# def append_element(some_list, element):
# some_list.append(element)
# ```
# ```python
# In [27]: data = [1, 2, 3]
#
# In [28]: append_element(data, 4)
#
# In [29]: data
# Out[29]: [1, 2, 3, 4]
# ```
# #### Dynamic references, strong types
a = 5
type(a)
a = 'foo'
type(a)
'5' + 5
a = 4.5
b = 2
# String formatting, to be visited later
print('a is {0}, b is {1}'.format(type(a), type(b)))
a / b
a = 5
isinstance(a, int)
a = 5; b = 4.5
isinstance(a, (int, float))
isinstance(b, (int, float))
# #### Attributes and methods
# ```python
# In [1]: a = 'foo'
#
# In [2]: a.<Press Tab>
# a.capitalize a.format a.isupper a.rindex a.strip
# a.center a.index a.join a.rjust a.swapcase
# a.count a.isalnum a.ljust a.rpartition a.title
# a.decode a.isalpha a.lower a.rsplit a.translate
# a.encode a.isdigit a.lstrip a.rstrip a.upper
# a.endswith a.islower a.partition a.split a.zfill
# a.expandtabs a.isspace a.replace a.splitlines
# a.find a.istitle a.rfind a.startswith
# ```
a = 'foo'
getattr(a, 'split')
# #### Duck typing
def isiterable(obj):
try:
iter(obj)
return True
except TypeError: # not iterable
return False
isiterable('a string')
isiterable([1, 2, 3])
isiterable(5)
# if not isinstance(x, list) and isiterable(x):
# x = list(x)
# #### Imports
# ```python
# # some_module.py
# PI = 3.14159
#
# def f(x):
# return x + 2
#
# def g(a, b):
# return a + b
# ```
# import some_module
# result = some_module.f(5)
# pi = some_module.PI
# from some_module import f, g, PI
# result = g(5, PI)
# import some_module as sm
# from some_module import PI as pi, g as gf
#
# r1 = sm.f(pi)
# r2 = gf(6, pi)
# #### Binary operators and comparisons
5 - 7
12 + 21.5
5 <= 2
a = [1, 2, 3]
b = a
c = list(a)
a is b
a is not c
a == c
a = None
a is None
# #### Mutable and immutable objects
a_list = ['foo', 2, [4, 5]]
a_list[2] = (3, 4)
a_list
a_tuple = (3, 5, (4, 5))
a_tuple[1] = 'four'
# ### Scalar Types
# #### Numeric types
ival = 17239871
ival ** 6
fval = 7.243
fval2 = 6.78e-5
3 / 2
3 // 2
# #### Strings
# a = 'one way of writing a string'
# b = "another way"
c = """
This is a longer string that
spans multiple lines
"""
c.count('\n')
a = 'this is a string'
a[10] = 'f'
b = a.replace('string', 'longer string')
b
a
a = 5.6
s = str(a)
print(s)
s = 'python'
list(s)
s[:3]
s = '12\\34'
print(s)
s = r'this\has\no\special\characters'
s
a = 'this is the first half '
b = 'and this is the second half'
a + b
template = '{0:.2f} {1:s} are worth US${2:d}'
template.format(4.5560, 'Argentine Pesos', 1)
# #### Bytes and Unicode
val = "español"
val
val_utf8 = val.encode('utf-8')
val_utf8
type(val_utf8)
val_utf8.decode('utf-8')
val.encode('latin1')
val.encode('utf-16')
val.encode('utf-16le')
bytes_val = b'this is bytes'
bytes_val
decoded = bytes_val.decode('utf8')
decoded # this is str (Unicode) now
# #### Booleans
True and True
False or True
# #### Type casting
s = '3.14159'
fval = float(s)
type(fval)
int(fval)
bool(fval)
bool(0)
# #### None
a = None
a is None
b = 5
b is not None
# def add_and_maybe_multiply(a, b, c=None):
# result = a + b
#
# if c is not None:
# result = result * c
#
# return result
type(None)
# #### Dates and times
from datetime import datetime, date, time
dt = datetime(2011, 10, 29, 20, 30, 21)
dt.day
dt.minute
dt.date()
dt.time()
dt.strftime('%m/%d/%Y %H:%M')
datetime.strptime('20091031', '%Y%m%d')
dt.replace(minute=0, second=0)
dt2 = datetime(2011, 11, 15, 22, 30)
delta = dt2 - dt
delta
type(delta)
dt
dt + delta
# ### Control Flow
# #### if, elif, and else
# if x < 0:
# print('It's negative')
# if x < 0:
# print('It's negative')
# elif x == 0:
# print('Equal to zero')
# elif 0 < x < 5:
# print('Positive but smaller than 5')
# else:
# print('Positive and larger than or equal to 5')
a = 5; b = 7
c = 8; d = 4
if a < b or c > d:
print('Made it')
4 > 3 > 2 > 1
# #### for loops
# for value in collection:
# # do something with value
# sequence = [1, 2, None, 4, None, 5]
# total = 0
# for value in sequence:
# if value is None:
# continue
# total += value
# sequence = [1, 2, 0, 4, 6, 5, 2, 1]
# total_until_5 = 0
# for value in sequence:
# if value == 5:
# break
# total_until_5 += value
for i in range(4):
for j in range(4):
if j > i:
break
print((i, j))
# for a, b, c in iterator:
# # do something
# #### while loops
# x = 256
# total = 0
# while x > 0:
# if total > 500:
# break
# total += x
# x = x // 2
# #### pass
# if x < 0:
# print('negative!')
# elif x == 0:
# # TODO: put something smart here
# pass
# else:
# print('positive!')
# #### range
range(10)
list(range(10))
list(range(0, 20, 2))
list(range(5, 0, -1))
# seq = [1, 2, 3, 4]
# for i in range(len(seq)):
# val = seq[i]
# sum = 0
# for i in range(100000):
# # % is the modulo operator
# if i % 3 == 0 or i % 5 == 0:
# sum += i
# #### Ternary expressions
# value =
# if
x = 5
'Non-negative' if x >= 0 else 'Negative'
| ch02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# # Pandas series
ser=pd.Series([22,34,76],["Deepak","Amit","Rashmi"])
ser
ser.index
ser.loc[['Deepak','Amit']]
# ## Series is 1D labelled array
ser.loc['Deepak']
ser[[2]]
ser.iloc[0]
for i in (0:3):
ser[[i]]
"Deepak" in ser
22 in ser
ser*2
ser**2
ser[['Amit','Deepak']]**2
# # pandas DataFrame
# Creating DataFrame from dictionary of Python series
d={'Age':pd.Series([22,23,45],index=['D','A','C']),
'Salary':pd.Series([1000,2000,3000],index=['D','A','C'])}
df=pd.DataFrame(d)
df
df.index
df.columns
pd.DataFrame(d,index=['D','A'])
pd.DataFrame(d,index=['D','A','C'],columns=['Age'])
data=[{'Deepak':1,'Ashish':2},{'Kavita':5,'Anmole':10,"Sanyam":20,"Deepak":20}]
pd.DataFrame(data)
pd.DataFrame(data,index=["Section 1","Section 2"])
pd.DataFrame(data,columns=["Anmole","Sanyam"])
# # Basic DataFrame Operations
df
df["Age"]
df["Higher Salary"]=df["Salary"]>1000
df
Age=df.pop("Age")
Age
df
del df["Higher Salary"]
df
df.insert(0,'Copy of Salary',df['Salary'])
df
df.insert(1,'Copy of Salary 3',df['Salary'])
# Can duplicate values but col name should be unique
df
# # Case Study: Movie Data Analysis
movies=pd.read_csv("movies.csv",sep=',')
print(type(movies))
movies.head(15)
movies.tail(10)
# # Day 3 Coding
# +
tags=pd.read_csv("tags.csv",sep=",")
tags.head()
# -
ratings=pd.read_csv("ratings.csv",sep=",",parse_dates=['timestamp'])
ratings.head()
del ratings["timestamp"]
del tags["timestamp"]
tags.head()
ratings.head()
# +
#Extract 0th row: notice that it is infact a Series
row_0 = tags.iloc[0]
type(row_0)
# -
row_0
row_0.index
row_0['userId']
row_0['movieId']
'rating' in row_0
for x in row_0:
print (x)
row_0.name
row_0 = row_0.rename('first_row')
row_0.name
range(0,tags.size)
s=tags.size-1
s
for i in range(0,10,1):
row=tags.iloc[i]
row
# # DataFrames
tags.head()
tags.index
tags.columns
# +
# Extract row 0, 11, 2000 from DataFrame
tags.iloc[ [0,11,2000] ]
# -
rows=tags.iloc[range(0,10,1)]
rows
# # Descriptive statistics
ratings['rating'].describe() #Only Rating wil be reflected
ratings.describe()
ratings['rating'].mean()
ratings.mean()
ratings['rating'].min()
ratings['rating'].max()
ratings['rating'].std()
ratings['rating'].mode()
ratings.corr()
filter_1 = ratings['rating'] > 5
print(filter_1)
filter_1.any()
filter_2 = ratings['rating'] > 0
print(filter_2)
filter_2.all()
type(filter_1)
print (all([False, True, False, False])) # if any 1 is false, all() returns false
print (any([False, True, False, False])) # if any 1 is true, any() returns true
# # Data Cleaning
movies.shape
# +
#is any row NULL ?
movies.isnull().any()
# -
tags.isnull().any()
tags.shape
#
#
tags=tags.dropna() # never drop rows directly
tags.shape #no of rows and column
ratings.isnull().any()
tags.shape[0]
rows=tags.iloc[range(0,tags.shape[0],1)]
rows.head()
# No null values
# # Data visualisation
# +
# %matplotlib inline
ratings.hist(column='rating', figsize=(15,10))
# -
ratings.boxplot(column='rating', figsize=(15,10))
# # Slicing out columns
tags['tag'].head()
movies[['title','genres']].head()
ratings[-10:]
tag_counts = tags['tag'].value_counts()
tag_counts
# %matplotlib inline
tag_counts[:10].plot(kind='bar', figsize=(15,10))
# <h1 style="font-size:2em;color:#2467C0">Filters for Selecting Rows</h1>
# +
is_highly_rated = ratings['rating'] >= 4.0
ratings[is_highly_rated][30:50]
# +
is_animation = movies['genres'].str.contains('Animation','IMAX')
movies[is_animation][5:15]
# +
is_animation = movies['genres'].str.contains('Animation')
movies[is_animation][5:15]
# -
movies[is_animation].head(15)
# <h1 style="font-size:2em;color:#2467C0">Group By and Aggregate </h1>
ratings_count = ratings[['movieId','rating']].groupby('rating').count()
ratings_count
ratings.shape
ratings_count.rename(columns = {"movieId": "No. of movies"})
ratings_count
average_rating = ratings[['movieId','rating']].groupby('movieId').mean()
average_rating.head()
# <h1 style="font-size:2em;color:#2467C0">Merge Dataframes</h1>
tags.head()
movies.head()
t = movies.merge(tags, on='movieId', how='inner')
t.head()
# <p style="font-family: Arial; font-size:1.35em;color:#2462C0; font-style:bold"><br>
#
# Split 'genres' into multiple columns
#
# <br> </p>
movie_genres = movies['genres'].str.split('|', expand=True)
movie_genres[:10]
movie_genres = movies['genres'].str.split('|', expand=False)
movie_genres[:10]
# <p style="font-family: Arial; font-size:1.35em;color:#2462C0; font-style:bold"><br>
#
# Add a new column for comedy genre flag
#
# <br> </p>
movie_genres['isComedy'] = movies['genres'].str.contains('Comedy')
movie_genres[:10]
movies['year'] = movies['title'].str.extract('.*\((.*)\).*', expand=True)
movies.tail()
| Python/Henry Harvin/Pandas Library.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="K2s1A9eLRPEj"
# ##### Copyright 2018 The TensorFlow Authors.
#
#
# + cellView="form" colab={} colab_type="code" id="VRLVEKiTEn04"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="Cffg2i257iMS"
# # Image Captioning with Attention
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/beta/tutorials/text/image_captioning">
# <img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
# View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r2/tutorials/text/image_captioning.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
# Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r2/tutorials/text/image_captioning.ipynb">
# <img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
# View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/r2/tutorials/text/image_captioning.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="QASbY_HGo4Lq"
# Given an image like the example below, our goal is to generate a caption such as "a surfer riding on a wave".
#
# 
#
# *[Image Source](https://commons.wikimedia.org/wiki/Surfing#/media/File:Surfing_in_Hawaii.jpg); License: Public Domain*
#
# To accomplish this, you'll use an attention-based model, which enables us to see what parts of the image the model focuses on as it generates a caption.
#
# 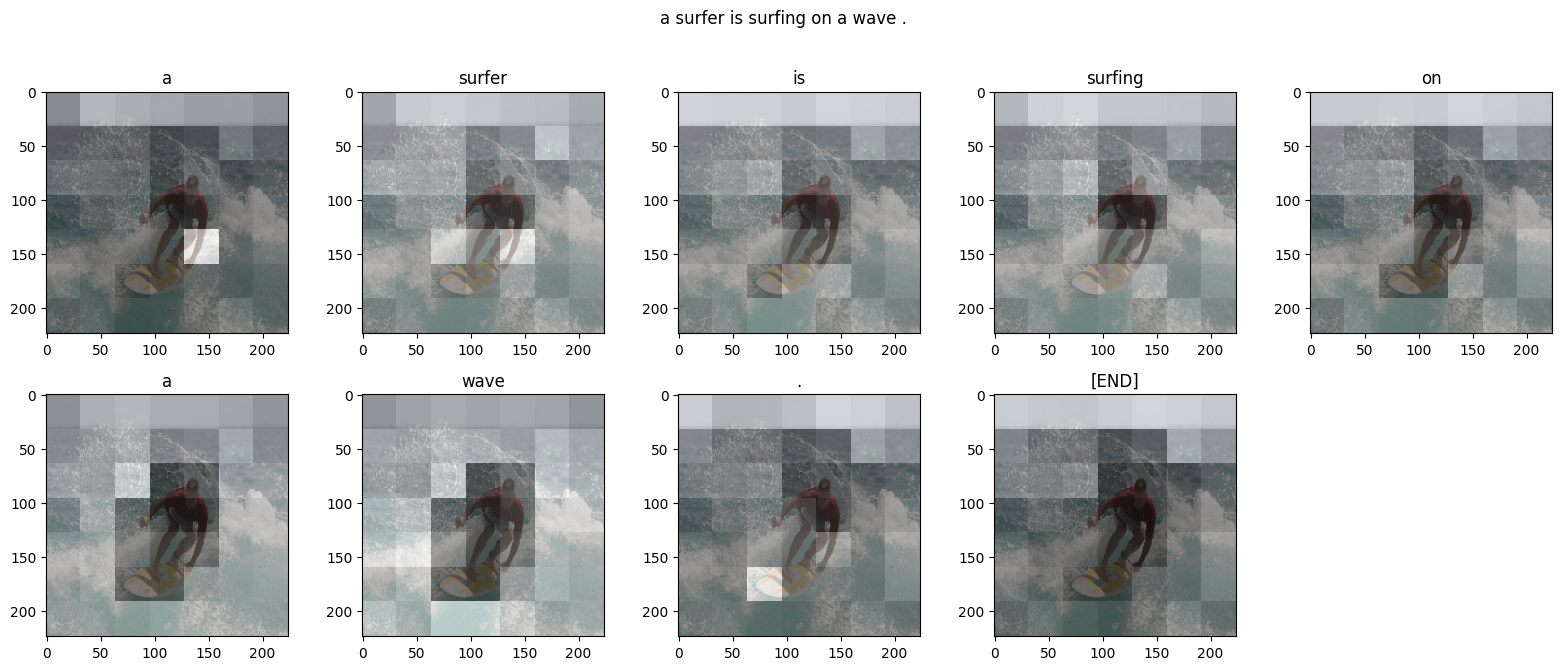
#
# The model architecture is similar to [Show, Attend and Tell: Neural Image Caption Generation with Visual Attention](https://arxiv.org/abs/1502.03044).
#
# This notebook is an end-to-end example. When you run the notebook, it downloads the [MS-COCO](http://cocodataset.org/#home) dataset, preprocesses and caches a subset of images using Inception V3, trains an encoder-decoder model, and generates captions on new images using the trained model.
#
# In this example, you will train a model on a relatively small amount of data—the first 30,000 captions for about 20,000 images (because there are multiple captions per image in the dataset).
# + colab={} colab_type="code" id="6svGEEek67ds"
from __future__ import absolute_import, division, print_function, unicode_literals
# + colab={} colab_type="code" id="U8l4RJ0XRPEm"
# !pip install tensorflow-gpu==2.0.0-beta1
import tensorflow as tf
# You'll generate plots of attention in order to see which parts of an image
# our model focuses on during captioning
import matplotlib.pyplot as plt
# Scikit-learn includes many helpful utilities
from sklearn.model_selection import train_test_split
from sklearn.utils import shuffle
import re
import numpy as np
import os
import time
import json
from glob import glob
from PIL import Image
import pickle
# + [markdown] colab_type="text" id="b6qbGw8MRPE5"
# ## Download and prepare the MS-COCO dataset
#
# You will use the [MS-COCO dataset](http://cocodataset.org/#home) to train our model. The dataset contains over 82,000 images, each of which has at least 5 different caption annotations. The code below downloads and extracts the dataset automatically.
#
# **Caution: large download ahead**. You'll use the training set, which is a 13GB file.
# + colab={} colab_type="code" id="krQuPYTtRPE7"
annotation_zip = tf.keras.utils.get_file('captions.zip',
cache_subdir=os.path.abspath('.'),
origin = 'http://images.cocodataset.org/annotations/annotations_trainval2014.zip',
extract = True)
annotation_file = os.path.dirname(annotation_zip)+'/annotations/captions_train2014.json'
name_of_zip = 'train2014.zip'
if not os.path.exists(os.path.abspath('.') + '/' + name_of_zip):
image_zip = tf.keras.utils.get_file(name_of_zip,
cache_subdir=os.path.abspath('.'),
origin = 'http://images.cocodataset.org/zips/train2014.zip',
extract = True)
PATH = os.path.dirname(image_zip)+'/train2014/'
else:
PATH = os.path.abspath('.')+'/train2014/'
# + [markdown] colab_type="text" id="aANEzb5WwSzg"
# ## Optional: limit the size of the training set
# To speed up training for this tutorial, you'll use a subset of 30,000 captions and their corresponding images to train our model. Choosing to use more data would result in improved captioning quality.
# + colab={} colab_type="code" id="4G3b8x8_RPFD"
# Read the json file
with open(annotation_file, 'r') as f:
annotations = json.load(f)
# Store captions and image names in vectors
all_captions = []
all_img_name_vector = []
for annot in annotations['annotations']:
caption = '<start> ' + annot['caption'] + ' <end>'
image_id = annot['image_id']
full_coco_image_path = PATH + 'COCO_train2014_' + '%012d.jpg' % (image_id)
all_img_name_vector.append(full_coco_image_path)
all_captions.append(caption)
# Shuffle captions and image_names together
# Set a random state
train_captions, img_name_vector = shuffle(all_captions,
all_img_name_vector,
random_state=1)
# Select the first 30000 captions from the shuffled set
num_examples = 30000
train_captions = train_captions[:num_examples]
img_name_vector = img_name_vector[:num_examples]
# + colab={} colab_type="code" id="mPBMgK34RPFL"
len(train_captions), len(all_captions)
# + [markdown] colab_type="text" id="8cSW4u-ORPFQ"
# ## Preprocess the images using InceptionV3
# Next, you will use InceptionV3 (which is pretrained on Imagenet) to classify each image. You will extract features from the last convolutional layer.
#
# First, you will convert the images into InceptionV3's expected format by:
# * Resizing the image to 299px by 299px
# * [Preprocess the images](https://cloud.google.com/tpu/docs/inception-v3-advanced#preprocessing_stage) using the [preprocess_input](https://www.tensorflow.org/api_docs/python/tf/keras/applications/inception_v3/preprocess_input) method to normalize the image so that it contains pixels in the range of -1 to 1, which matches the format of the images used to train InceptionV3.
# + colab={} colab_type="code" id="zXR0217aRPFR"
def load_image(image_path):
img = tf.io.read_file(image_path)
img = tf.image.decode_jpeg(img, channels=3)
img = tf.image.resize(img, (299, 299))
img = tf.keras.applications.inception_v3.preprocess_input(img)
return img, image_path
# + [markdown] colab_type="text" id="MDvIu4sXRPFV"
# ## Initialize InceptionV3 and load the pretrained Imagenet weights
#
# Now you'll create a tf.keras model where the output layer is the last convolutional layer in the InceptionV3 architecture. The shape of the output of this layer is ```8x8x2048```. You use the last convolutional layer because you are using attention in this example. You don't perform this initialization during training because it could become a bottleneck.
#
# * You forward each image through the network and store the resulting vector in a dictionary (image_name --> feature_vector).
# * After all the images are passed through the network, you pickle the dictionary and save it to disk.
#
#
#
# + colab={} colab_type="code" id="RD3vW4SsRPFW"
image_model = tf.keras.applications.InceptionV3(include_top=False,
weights='imagenet')
new_input = image_model.input
hidden_layer = image_model.layers[-1].output
image_features_extract_model = tf.keras.Model(new_input, hidden_layer)
# + [markdown] colab_type="text" id="rERqlR3WRPGO"
# ## Caching the features extracted from InceptionV3
#
# You will pre-process each image with InceptionV3 and cache the output to disk. Caching the output in RAM would be faster but also memory intensive, requiring 8 \* 8 \* 2048 floats per image. At the time of writing, this exceeds the memory limitations of Colab (currently 12GB of memory).
#
# Performance could be improved with a more sophisticated caching strategy (for example, by sharding the images to reduce random access disk I/O), but that would require more code.
#
# The caching will take about 10 minutes to run in Colab with a GPU. If you'd like to see a progress bar, you can:
#
# 1. install [tqdm](https://github.com/tqdm/tqdm):
#
# ```!pip install tqdm```
#
# 2. Import tqdm:
#
# ```from tqdm import tqdm```
#
# 3. Change the following line:
#
# ```for img, path in image_dataset:```
#
# to:
#
# ```for img, path in tqdm(image_dataset):```.
# + colab={} colab_type="code" id="Dx_fvbVgRPGQ"
# Get unique images
encode_train = sorted(set(img_name_vector))
# Feel free to change batch_size according to your system configuration
image_dataset = tf.data.Dataset.from_tensor_slices(encode_train)
image_dataset = image_dataset.map(
load_image, num_parallel_calls=tf.data.experimental.AUTOTUNE).batch(16)
for img, path in image_dataset:
batch_features = image_features_extract_model(img)
batch_features = tf.reshape(batch_features,
(batch_features.shape[0], -1, batch_features.shape[3]))
for bf, p in zip(batch_features, path):
path_of_feature = p.numpy().decode("utf-8")
np.save(path_of_feature, bf.numpy())
# + [markdown] colab_type="text" id="nyqH3zFwRPFi"
# ## Preprocess and tokenize the captions
#
# * First, you'll tokenize the captions (for example, by splitting on spaces). This gives us a vocabulary of all of the unique words in the data (for example, "surfing", "football", and so on).
# * Next, you'll limit the vocabulary size to the top 5,000 words (to save memory). You'll replace all other words with the token "UNK" (unknown).
# * You then create word-to-index and index-to-word mappings.
# * Finally, you pad all sequences to the be same length as the longest one.
# + colab={} colab_type="code" id="HZfK8RhQRPFj"
# Find the maximum length of any caption in our dataset
def calc_max_length(tensor):
return max(len(t) for t in tensor)
# + colab={} colab_type="code" id="oJGE34aiRPFo"
# Choose the top 5000 words from the vocabulary
top_k = 5000
tokenizer = tf.keras.preprocessing.text.Tokenizer(num_words=top_k,
oov_token="<unk>",
filters='!"#$%&()*+.,-/:;=?@[\]^_`{|}~ ')
tokenizer.fit_on_texts(train_captions)
train_seqs = tokenizer.texts_to_sequences(train_captions)
# + colab={} colab_type="code" id="8Q44tNQVRPFt"
tokenizer.word_index['<pad>'] = 0
tokenizer.index_word[0] = '<pad>'
# + colab={} colab_type="code" id="0fpJb5ojRPFv"
# Create the tokenized vectors
train_seqs = tokenizer.texts_to_sequences(train_captions)
# + colab={} colab_type="code" id="AidglIZVRPF4"
# Pad each vector to the max_length of the captions
# If you do not provide a max_length value, pad_sequences calculates it automatically
cap_vector = tf.keras.preprocessing.sequence.pad_sequences(train_seqs, padding='post')
# + colab={} colab_type="code" id="gL0wkttkRPGA"
# Calculates the max_length, which is used to store the attention weights
max_length = calc_max_length(train_seqs)
# + [markdown] colab_type="text" id="M3CD75nDpvTI"
# ## Split the data into training and testing
# + colab={} colab_type="code" id="iS7DDMszRPGF"
# Create training and validation sets using an 80-20 split
img_name_train, img_name_val, cap_train, cap_val = train_test_split(img_name_vector,
cap_vector,
test_size=0.2,
random_state=0)
# + colab={} colab_type="code" id="XmViPkRFRPGH"
len(img_name_train), len(cap_train), len(img_name_val), len(cap_val)
# + [markdown] colab_type="text" id="uEWM9xrYcg45"
# ## Create a tf.data dataset for training
#
#
# + [markdown] colab_type="text" id="horagNvhhZiy"
# Our images and captions are ready! Next, let's create a tf.data dataset to use for training our model.
# + colab={} colab_type="code" id="Q3TnZ1ToRPGV"
# Feel free to change these parameters according to your system's configuration
BATCH_SIZE = 64
BUFFER_SIZE = 1000
embedding_dim = 256
units = 512
vocab_size = len(tokenizer.word_index) + 1
num_steps = len(img_name_train) // BATCH_SIZE
# Shape of the vector extracted from InceptionV3 is (64, 2048)
# These two variables represent that vector shape
features_shape = 2048
attention_features_shape = 64
# + colab={} colab_type="code" id="SmZS2N0bXG3T"
# Load the numpy files
def map_func(img_name, cap):
img_tensor = np.load(img_name.decode('utf-8')+'.npy')
return img_tensor, cap
# + colab={} colab_type="code" id="FDF_Nm3tRPGZ"
dataset = tf.data.Dataset.from_tensor_slices((img_name_train, cap_train))
# Use map to load the numpy files in parallel
dataset = dataset.map(lambda item1, item2: tf.numpy_function(
map_func, [item1, item2], [tf.float32, tf.int32]),
num_parallel_calls=tf.data.experimental.AUTOTUNE)
# Shuffle and batch
dataset = dataset.shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
dataset = dataset.prefetch(buffer_size=tf.data.experimental.AUTOTUNE)
# + [markdown] colab_type="text" id="nrvoDphgRPGd"
# ## Model
#
# Fun fact: the decoder below is identical to the one in the example for [Neural Machine Translation with Attention](../sequences/nmt_with_attention.ipynb).
#
# The model architecture is inspired by the [Show, Attend and Tell](https://arxiv.org/pdf/1502.03044.pdf) paper.
#
# * In this example, you extract the features from the lower convolutional layer of InceptionV3 giving us a vector of shape (8, 8, 2048).
# * You squash that to a shape of (64, 2048).
# * This vector is then passed through the CNN Encoder (which consists of a single Fully connected layer).
# * The RNN (here GRU) attends over the image to predict the next word.
# + colab={} colab_type="code" id="ja2LFTMSdeV3"
class BahdanauAttention(tf.keras.Model):
def __init__(self, units):
super(BahdanauAttention, self).__init__()
self.W1 = tf.keras.layers.Dense(units)
self.W2 = tf.keras.layers.Dense(units)
self.V = tf.keras.layers.Dense(1)
def call(self, features, hidden):
# features(CNN_encoder output) shape == (batch_size, 64, embedding_dim)
# hidden shape == (batch_size, hidden_size)
# hidden_with_time_axis shape == (batch_size, 1, hidden_size)
hidden_with_time_axis = tf.expand_dims(hidden, 1)
# score shape == (batch_size, 64, hidden_size)
score = tf.nn.tanh(self.W1(features) + self.W2(hidden_with_time_axis))
# attention_weights shape == (batch_size, 64, 1)
# you get 1 at the last axis because you are applying score to self.V
attention_weights = tf.nn.softmax(self.V(score), axis=1)
# context_vector shape after sum == (batch_size, hidden_size)
context_vector = attention_weights * features
context_vector = tf.reduce_sum(context_vector, axis=1)
return context_vector, attention_weights
# + colab={} colab_type="code" id="AZ7R1RxHRPGf"
class CNN_Encoder(tf.keras.Model):
# Since you have already extracted the features and dumped it using pickle
# This encoder passes those features through a Fully connected layer
def __init__(self, embedding_dim):
super(CNN_Encoder, self).__init__()
# shape after fc == (batch_size, 64, embedding_dim)
self.fc = tf.keras.layers.Dense(embedding_dim)
def call(self, x):
x = self.fc(x)
x = tf.nn.relu(x)
return x
# + colab={} colab_type="code" id="V9UbGQmERPGi"
class RNN_Decoder(tf.keras.Model):
def __init__(self, embedding_dim, units, vocab_size):
super(RNN_Decoder, self).__init__()
self.units = units
self.embedding = tf.keras.layers.Embedding(vocab_size, embedding_dim)
self.gru = tf.keras.layers.GRU(self.units,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
self.fc1 = tf.keras.layers.Dense(self.units)
self.fc2 = tf.keras.layers.Dense(vocab_size)
self.attention = BahdanauAttention(self.units)
def call(self, x, features, hidden):
# defining attention as a separate model
context_vector, attention_weights = self.attention(features, hidden)
# x shape after passing through embedding == (batch_size, 1, embedding_dim)
x = self.embedding(x)
# x shape after concatenation == (batch_size, 1, embedding_dim + hidden_size)
x = tf.concat([tf.expand_dims(context_vector, 1), x], axis=-1)
# passing the concatenated vector to the GRU
output, state = self.gru(x)
# shape == (batch_size, max_length, hidden_size)
x = self.fc1(output)
# x shape == (batch_size * max_length, hidden_size)
x = tf.reshape(x, (-1, x.shape[2]))
# output shape == (batch_size * max_length, vocab)
x = self.fc2(x)
return x, state, attention_weights
def reset_state(self, batch_size):
return tf.zeros((batch_size, self.units))
# + colab={} colab_type="code" id="Qs_Sr03wRPGk"
encoder = CNN_Encoder(embedding_dim)
decoder = RNN_Decoder(embedding_dim, units, vocab_size)
# + colab={} colab_type="code" id="-bYN7xA0RPGl"
optimizer = tf.keras.optimizers.Adam()
loss_object = tf.keras.losses.SparseCategoricalCrossentropy(
from_logits=True, reduction='none')
def loss_function(real, pred):
mask = tf.math.logical_not(tf.math.equal(real, 0))
loss_ = loss_object(real, pred)
mask = tf.cast(mask, dtype=loss_.dtype)
loss_ *= mask
return tf.reduce_mean(loss_)
# + [markdown] colab_type="text" id="6A3Ni64joyab"
# ## Checkpoint
# + colab={} colab_type="code" id="PpJAqPMWo0uE"
checkpoint_path = "./checkpoints/train"
ckpt = tf.train.Checkpoint(encoder=encoder,
decoder=decoder,
optimizer = optimizer)
ckpt_manager = tf.train.CheckpointManager(ckpt, checkpoint_path, max_to_keep=5)
# + colab={} colab_type="code" id="fUkbqhc_uObw"
start_epoch = 0
if ckpt_manager.latest_checkpoint:
start_epoch = int(ckpt_manager.latest_checkpoint.split('-')[-1])
# + [markdown] colab_type="text" id="PHod7t72RPGn"
# ## Training
#
# * You extract the features stored in the respective `.npy` files and then pass those features through the encoder.
# * The encoder output, hidden state(initialized to 0) and the decoder input (which is the start token) is passed to the decoder.
# * The decoder returns the predictions and the decoder hidden state.
# * The decoder hidden state is then passed back into the model and the predictions are used to calculate the loss.
# * Use teacher forcing to decide the next input to the decoder.
# * Teacher forcing is the technique where the target word is passed as the next input to the decoder.
# * The final step is to calculate the gradients and apply it to the optimizer and backpropagate.
#
# + colab={} colab_type="code" id="Vt4WZ5mhJE-E"
# adding this in a separate cell because if you run the training cell
# many times, the loss_plot array will be reset
loss_plot = []
# + colab={} colab_type="code" id="sqgyz2ANKlpU"
@tf.function
def train_step(img_tensor, target):
loss = 0
# initializing the hidden state for each batch
# because the captions are not related from image to image
hidden = decoder.reset_state(batch_size=target.shape[0])
dec_input = tf.expand_dims([tokenizer.word_index['<start>']] * BATCH_SIZE, 1)
with tf.GradientTape() as tape:
features = encoder(img_tensor)
for i in range(1, target.shape[1]):
# passing the features through the decoder
predictions, hidden, _ = decoder(dec_input, features, hidden)
loss += loss_function(target[:, i], predictions)
# using teacher forcing
dec_input = tf.expand_dims(target[:, i], 1)
total_loss = (loss / int(target.shape[1]))
trainable_variables = encoder.trainable_variables + decoder.trainable_variables
gradients = tape.gradient(loss, trainable_variables)
optimizer.apply_gradients(zip(gradients, trainable_variables))
return loss, total_loss
# + colab={} colab_type="code" id="UlA4VIQpRPGo"
EPOCHS = 20
for epoch in range(start_epoch, EPOCHS):
start = time.time()
total_loss = 0
for (batch, (img_tensor, target)) in enumerate(dataset):
batch_loss, t_loss = train_step(img_tensor, target)
total_loss += t_loss
if batch % 100 == 0:
print ('Epoch {} Batch {} Loss {:.4f}'.format(
epoch + 1, batch, batch_loss.numpy() / int(target.shape[1])))
# storing the epoch end loss value to plot later
loss_plot.append(total_loss / num_steps)
if epoch % 5 == 0:
ckpt_manager.save()
print ('Epoch {} Loss {:.6f}'.format(epoch + 1,
total_loss/num_steps))
print ('Time taken for 1 epoch {} sec\n'.format(time.time() - start))
# + colab={} colab_type="code" id="1Wm83G-ZBPcC"
plt.plot(loss_plot)
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.title('Loss Plot')
plt.show()
# + [markdown] colab_type="text" id="xGvOcLQKghXN"
# ## Caption!
#
# * The evaluate function is similar to the training loop, except you don't use teacher forcing here. The input to the decoder at each time step is its previous predictions along with the hidden state and the encoder output.
# * Stop predicting when the model predicts the end token.
# * And store the attention weights for every time step.
# + colab={} colab_type="code" id="RCWpDtyNRPGs"
def evaluate(image):
attention_plot = np.zeros((max_length, attention_features_shape))
hidden = decoder.reset_state(batch_size=1)
temp_input = tf.expand_dims(load_image(image)[0], 0)
img_tensor_val = image_features_extract_model(temp_input)
img_tensor_val = tf.reshape(img_tensor_val, (img_tensor_val.shape[0], -1, img_tensor_val.shape[3]))
features = encoder(img_tensor_val)
dec_input = tf.expand_dims([tokenizer.word_index['<start>']], 0)
result = []
for i in range(max_length):
predictions, hidden, attention_weights = decoder(dec_input, features, hidden)
attention_plot[i] = tf.reshape(attention_weights, (-1, )).numpy()
predicted_id = tf.argmax(predictions[0]).numpy()
result.append(tokenizer.index_word[predicted_id])
if tokenizer.index_word[predicted_id] == '<end>':
return result, attention_plot
dec_input = tf.expand_dims([predicted_id], 0)
attention_plot = attention_plot[:len(result), :]
return result, attention_plot
# + colab={} colab_type="code" id="fD_y7PD6RPGt"
def plot_attention(image, result, attention_plot):
temp_image = np.array(Image.open(image))
fig = plt.figure(figsize=(10, 10))
len_result = len(result)
for l in range(len_result):
temp_att = np.resize(attention_plot[l], (8, 8))
ax = fig.add_subplot(len_result//2, len_result//2, l+1)
ax.set_title(result[l])
img = ax.imshow(temp_image)
ax.imshow(temp_att, cmap='gray', alpha=0.6, extent=img.get_extent())
plt.tight_layout()
plt.show()
# + colab={} colab_type="code" id="7x8RiPHe_4qI"
# captions on the validation set
rid = np.random.randint(0, len(img_name_val))
image = img_name_val[rid]
real_caption = ' '.join([tokenizer.index_word[i] for i in cap_val[rid] if i not in [0]])
result, attention_plot = evaluate(image)
print ('Real Caption:', real_caption)
print ('Prediction Caption:', ' '.join(result))
plot_attention(image, result, attention_plot)
# opening the image
Image.open(img_name_val[rid])
# + [markdown] colab_type="text" id="Rprk3HEvZuxb"
# ## Try it on your own images
# For fun, below we've provided a method you can use to caption your own images with the model we've just trained. Keep in mind, it was trained on a relatively small amount of data, and your images may be different from the training data (so be prepared for weird results!)
#
# + colab={} colab_type="code" id="9Psd1quzaAWg"
image_url = 'https://tensorflow.org/images/surf.jpg'
image_extension = image_url[-4:]
image_path = tf.keras.utils.get_file('image'+image_extension,
origin=image_url)
result, attention_plot = evaluate(image_path)
print ('Prediction Caption:', ' '.join(result))
plot_attention(image_path, result, attention_plot)
# opening the image
Image.open(image_path)
# + [markdown] colab_type="text" id="VJZXyJco6uLO"
# # Next steps
#
# Congrats! You've just trained an image captioning model with attention. Next, take a look at this example [Neural Machine Translation with Attention](../sequences/nmt_with_attention.ipynb). It uses a similar architecture to translate between Spanish and English sentences. You can also experiment with training the code in this notebook on a different dataset.
| site/en/r2/tutorials/text/image_captioning.ipynb |