code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''leola'': virtualenv)'
# name: python3
# ---
import sys
sys.path.append('../../../')
# +
import warnings
warnings.filterwarnings('ignore')
import fpl_analytics
import plotly.express as px
import pandas as pd
# -
week = 12
fpl_data = fpl_analytics.load_fpl(week+1)
base_feature=fpl_analytics.BaseWeeklyFeature(fpl_data)
avg3_feature=fpl_analytics.Avg3WeeklyFeature(fpl_data)
opp_feature=fpl_analytics.OppWeeklyFeature(fpl_data)
hist_feature=fpl_analytics.HistWeeklyFeature(fpl_data)
# +
pos = ["Goalkeeper", "Defender", "Midfielder", "Forward"]
actual_res = {p: (base_feature.extract(p, [week], x_only=False))[1] for p in pos}
actual_res = pd.concat(list(actual_res.values()), keys=list(actual_res.keys()), names=["position", "player"])
pre_res = pd.read_pickle(f"points_predict_{week}.pkl")
scatter_res = pd.concat([actual_res, pre_res], keys=["actual", "predict"], axis=1)
scatter_res=scatter_res.reset_index().dropna()
# +
pos = ["Goalkeeper", "Defender", "Midfielder", "Forward"]
actual_res = {p: (base_feature.extract(p, [week], x_only=False))[1] for p in pos}
actual_res = pd.concat(list(actual_res.values()), keys=list(actual_res.keys()), names=["position", "player"])
# +
import plotly.express as px
fig = px.scatter(scatter_res, y="actual", x="predict", color="position",
hover_data=['player'],
title=f"FPL week {week} algo prediction vs actual")
fig.show()
# +
actual_res = {p: (base_feature.extract(p, [week], x_only=False))[1] for p in pos}
actual_res = pd.concat(list(actual_res.values()), keys=list(actual_res.keys()), names=["position", "player"])
prev_res = {p: (base_feature.extract(p, [week-1], x_only=False))[1] for p in pos}
prev_res = pd.concat(list(prev_res.values()), keys=list(prev_res.keys()), names=["position", "player"])
import random
prev_res = prev_res.apply(lambda x: x+random.random()/2)
prev_res.index=pd.MultiIndex.from_tuples([(i[0], i[1].replace(f"week {week-1}", f"week {week}")
) for i in prev_res.index],
names=["position", "player"])
scatter_res_pre = pd.concat([actual_res, prev_res], keys=["actual", "previous"], axis=1)
scatter_res_pre=scatter_res_pre.reset_index().dropna()
# +
fig = px.scatter(scatter_res_pre, y="actual", x="previous", color="position",
hover_data=['player'],
title=f"FPL week {week} algo pre-week vs actual")
fig.show()
# -
from sklearn.metrics import ndcg_score
# +
def point_gp(x):
if x>=8:
return 2
elif 5<=x<=7:
return 1
else:
return 0
prev_res = {p: (base_feature.extract(p, [week-1], x_only=False))[1] for p in pos}
prev_res = pd.concat(list(prev_res.values()), keys=list(prev_res.keys()), names=["position", "player"])
prev_res.index=pd.MultiIndex.from_tuples([(i[0], i[1].replace(f"week {week-1}", f"week {week}")
) for i in prev_res.index],
names=["position", "player"])
scatter_res_pre = pd.concat([actual_res, prev_res], keys=["actual", "previous"], axis=1)
scatter_res_pre=scatter_res_pre.reset_index().dropna()
srp = scatter_res_pre.sort_values(by="actual", ascending=False)
srp["actual_rank"] = srp.actual.apply(point_gp)
n_s_b = ndcg_score([srp["actual_rank"].to_numpy()], [srp["previous"].to_numpy()])
print (f"NDCG baseline", n_s_b)
sr = scatter_res.sort_values(by="actual", ascending=False)
sr["actual_rank"] = sr.actual.apply(point_gp)
n_s_a = ndcg_score([sr["actual_rank"].to_numpy()], [sr["predict"].to_numpy()])
print (f"NDCG algo", n_s_a)
res = {f"week {week} Overall NDCG": pd.Series({"baseline": n_s_b, "algo": n_s_a})}
for p in pos:
srp = scatter_res_pre[scatter_res_pre["position"]==p]
srp = srp.sort_values(by="actual", ascending=False)
srp["actual_rank"] = srp.actual.apply(point_gp)
n_s_b = ndcg_score([srp["actual_rank"].to_numpy()],
[srp["previous"].to_numpy()])
print (f"NDCG {p} on baseline", n_s_b)
sr = scatter_res[scatter_res["position"]==p]
sr = sr.sort_values(by="actual", ascending=False)
sr["actual_rank"] = sr.actual.apply(point_gp)
n_s_a = ndcg_score([sr["actual_rank"].to_numpy()],
[sr["predict"].to_numpy()])
print (f"NDCG {p} on algo", n_s_a)
res[f"week {week} {p} NDCG"] = pd.Series({"baseline": n_s_b, "algo": n_s_a})
# -
pd.DataFrame(res)
| report/2021_2022/gw_prediction_gw_12_eval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="copyright"
# Copyright 2020 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="title"
# # Vertex AI client library: AutoML image classification model for online prediction
#
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/GoogleCloudPlatform/ai-platform-samples/blob/master/ai-platform-unified/notebooks/community/gapic/automl/showcase_automl_image_classification_online.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
# </a>
# </td>
# <td>
# <a href="https://github.com/GoogleCloudPlatform/ai-platform-samples/blob/master/ai-platform-unified/notebooks/community/gapic/automl/showcase_automl_image_classification_online.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
# View on GitHub
# </a>
# </td>
# </table>
# <br/><br/><br/>
# + [markdown] id="overview:automl"
# ## Overview
#
#
# This tutorial demonstrates how to use the Vertex AI Python client library to create image classification models and do online prediction using Google Cloud's [AutoML](https://cloud.google.com/ai-platform-unified/docs/start/automl-users).
# + [markdown] id="dataset:flowers,icn"
# ### Dataset
#
# The dataset used for this tutorial is the [Flowers dataset](https://www.tensorflow.org/datasets/catalog/tf_flowers) from [TensorFlow Datasets](https://www.tensorflow.org/datasets/catalog/overview). The version of the dataset you will use in this tutorial is stored in a public Cloud Storage bucket. The trained model predicts the type of flower an image is from a class of five flowers: daisy, dandelion, rose, sunflower, or tulip.
# + [markdown] id="objective:automl,training,online_prediction"
# ### Objective
#
# In this tutorial, you create an AutoML image classification model and deploy for online prediction from a Python script using the Vertex AI client library. You can alternatively create and deploy models using the `gcloud` command-line tool or online using the Google Cloud Console.
#
# The steps performed include:
#
# - Create a Vertex AI `Dataset` resource.
# - Train the model.
# - View the model evaluation.
# - Deploy the `Model` resource to a serving `Endpoint` resource.
# - Make a prediction.
# - Undeploy the `Model`.
# + [markdown] id="costs"
# ### Costs
#
# This tutorial uses billable components of Google Cloud (GCP):
#
# * Vertex AI
# * Cloud Storage
#
# Learn about [Vertex AI
# pricing](https://cloud.google.com/ai-platform-unified/pricing) and [Cloud Storage
# pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
# Calculator](https://cloud.google.com/products/calculator/)
# to generate a cost estimate based on your projected usage.
# + [markdown] id="install_aip"
# ## Installation
#
# Install the latest version of Vertex AI client library.
# + id="install_aip"
import sys
if "google.colab" in sys.modules:
USER_FLAG = ""
else:
USER_FLAG = "--user"
# ! pip3 install -U google-cloud-aiplatform $USER_FLAG
# + [markdown] id="install_storage"
# Install the latest GA version of *google-cloud-storage* library as well.
# + id="install_storage"
# ! pip3 install -U google-cloud-storage $USER_FLAG
# + [markdown] id="restart"
# ### Restart the kernel
#
# Once you've installed the Vertex AI client library and Google *cloud-storage*, you need to restart the notebook kernel so it can find the packages.
# + id="restart"
import os
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
# + [markdown] id="before_you_begin"
# ## Before you begin
#
# ### GPU runtime
#
# *Make sure you're running this notebook in a GPU runtime if you have that option. In Colab, select* **Runtime > Change Runtime Type > GPU**
#
# ### Set up your Google Cloud project
#
# **The following steps are required, regardless of your notebook environment.**
#
# 1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
#
# 2. [Make sure that billing is enabled for your project.](https://cloud.google.com/billing/docs/how-to/modify-project)
#
# 3. [Enable the Vertex AI APIs and Compute Engine APIs.](https://console.cloud.google.com/flows/enableapi?apiid=ml.googleapis.com,compute_component)
#
# 4. [The Google Cloud SDK](https://cloud.google.com/sdk) is already installed in Vertex AI Notebooks.
#
# 5. Enter your project ID in the cell below. Then run the cell to make sure the
# Cloud SDK uses the right project for all the commands in this notebook.
#
# **Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
# + id="set_project_id"
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
# + id="autoset_project_id"
if PROJECT_ID == "" or PROJECT_ID is None or PROJECT_ID == "[your-project-id]":
# Get your GCP project id from gcloud
# shell_output = !gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID:", PROJECT_ID)
# + id="set_gcloud_project_id"
# ! gcloud config set project $PROJECT_ID
# + [markdown] id="region"
# #### Region
#
# You can also change the `REGION` variable, which is used for operations
# throughout the rest of this notebook. Below are regions supported for Vertex AI. We recommend that you choose the region closest to you.
#
# - Americas: `us-central1`
# - Europe: `europe-west4`
# - Asia Pacific: `asia-east1`
#
# You may not use a multi-regional bucket for training with Vertex AI. Not all regions provide support for all Vertex AI services. For the latest support per region, see the [Vertex AI locations documentation](https://cloud.google.com/ai-platform-unified/docs/general/locations)
# + id="region"
REGION = "us-central1" # @param {type: "string"}
# + [markdown] id="timestamp"
# #### Timestamp
#
# If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append onto the name of resources which will be created in this tutorial.
# + id="timestamp"
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
# + [markdown] id="gcp_authenticate"
# ### Authenticate your Google Cloud account
#
# **If you are using Vertex AI Notebooks**, your environment is already authenticated. Skip this step.
#
# **If you are using Colab**, run the cell below and follow the instructions when prompted to authenticate your account via oAuth.
#
# **Otherwise**, follow these steps:
#
# In the Cloud Console, go to the [Create service account key](https://console.cloud.google.com/apis/credentials/serviceaccountkey) page.
#
# **Click Create service account**.
#
# In the **Service account name** field, enter a name, and click **Create**.
#
# In the **Grant this service account access to project** section, click the Role drop-down list. Type "Vertex AI" into the filter box, and select **Vertex AI Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
#
# Click Create. A JSON file that contains your key downloads to your local environment.
#
# Enter the path to your service account key as the GOOGLE_APPLICATION_CREDENTIALS variable in the cell below and run the cell.
# + id="gcp_authenticate"
import os
import sys
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# If on AI Platform, then don't execute this code
if not os.path.exists("/opt/deeplearning/metadata/env_version"):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
# %env GOOGLE_APPLICATION_CREDENTIALS ''
# + [markdown] id="setup_vars"
# ### Set up variables
#
# Next, set up some variables used throughout the tutorial.
# ### Import libraries and define constants
# + [markdown] id="import_aip"
# #### Import Vertex AI client library
#
# Import the Vertex AI client library into our Python environment.
# + id="import_aip"
import os
import sys
import time
import google.cloud.aiplatform_v1 as aip
from google.protobuf import json_format
from google.protobuf.json_format import MessageToJson, ParseDict
from google.protobuf.struct_pb2 import Struct, Value
# + [markdown] id="aip_constants"
# #### Vertex AI constants
#
# Setup up the following constants for Vertex AI:
#
# - `API_ENDPOINT`: The Vertex AI API service endpoint for dataset, model, job, pipeline and endpoint services.
# - `PARENT`: The Vertex AI location root path for dataset, model, job, pipeline and endpoint resources.
# + id="aip_constants"
# API service endpoint
API_ENDPOINT = "{}-aiplatform.googleapis.com".format(REGION)
# Vertex AI location root path for your dataset, model and endpoint resources
PARENT = "projects/" + PROJECT_ID + "/locations/" + REGION
# + [markdown] id="automl_constants"
# #### AutoML constants
#
# Set constants unique to AutoML datasets and training:
#
# - Dataset Schemas: Tells the `Dataset` resource service which type of dataset it is.
# - Data Labeling (Annotations) Schemas: Tells the `Dataset` resource service how the data is labeled (annotated).
# - Dataset Training Schemas: Tells the `Pipeline` resource service the task (e.g., classification) to train the model for.
# + id="automl_constants:icn"
# Image Dataset type
DATA_SCHEMA = "gs://google-cloud-aiplatform/schema/dataset/metadata/image_1.0.0.yaml"
# Image Labeling type
LABEL_SCHEMA = "gs://google-cloud-aiplatform/schema/dataset/ioformat/image_classification_single_label_io_format_1.0.0.yaml"
# Image Training task
TRAINING_SCHEMA = "gs://google-cloud-aiplatform/schema/trainingjob/definition/automl_image_classification_1.0.0.yaml"
# + [markdown] id="tutorial_start:automl"
# # Tutorial
#
# Now you are ready to start creating your own AutoML image classification model.
# + [markdown] id="clients:automl,online_prediction"
# ## Set up clients
#
# The Vertex AI client library works as a client/server model. On your side (the Python script) you will create a client that sends requests and receives responses from the Vertex AI server.
#
# You will use different clients in this tutorial for different steps in the workflow. So set them all up upfront.
#
# - Dataset Service for `Dataset` resources.
# - Model Service for `Model` resources.
# - Pipeline Service for training.
# - Endpoint Service for deployment.
# - Prediction Service for serving.
# + id="clients:automl,online_prediction"
# client options same for all services
client_options = {"api_endpoint": API_ENDPOINT}
def create_dataset_client():
client = aip.DatasetServiceClient(client_options=client_options)
return client
def create_model_client():
client = aip.ModelServiceClient(client_options=client_options)
return client
def create_pipeline_client():
client = aip.PipelineServiceClient(client_options=client_options)
return client
def create_endpoint_client():
client = aip.EndpointServiceClient(client_options=client_options)
return client
def create_prediction_client():
client = aip.PredictionServiceClient(client_options=client_options)
return client
clients = {}
clients["dataset"] = create_dataset_client()
clients["model"] = create_model_client()
clients["pipeline"] = create_pipeline_client()
clients["endpoint"] = create_endpoint_client()
clients["prediction"] = create_prediction_client()
for client in clients.items():
print(client)
# + [markdown] id="create_aip_dataset"
# ## Dataset
#
# Now that your clients are ready, your first step in training a model is to create a managed dataset instance, and then upload your labeled data to it.
#
# ### Create `Dataset` resource instance
#
# Use the helper function `create_dataset` to create the instance of a `Dataset` resource. This function does the following:
#
# 1. Uses the dataset client service.
# 2. Creates an Vertex AI `Dataset` resource (`aip.Dataset`), with the following parameters:
# - `display_name`: The human-readable name you choose to give it.
# - `metadata_schema_uri`: The schema for the dataset type.
# 3. Calls the client dataset service method `create_dataset`, with the following parameters:
# - `parent`: The Vertex AI location root path for your `Database`, `Model` and `Endpoint` resources.
# - `dataset`: The Vertex AI dataset object instance you created.
# 4. The method returns an `operation` object.
#
# An `operation` object is how Vertex AI handles asynchronous calls for long running operations. While this step usually goes fast, when you first use it in your project, there is a longer delay due to provisioning.
#
# You can use the `operation` object to get status on the operation (e.g., create `Dataset` resource) or to cancel the operation, by invoking an operation method:
#
# | Method | Description |
# | ----------- | ----------- |
# | result() | Waits for the operation to complete and returns a result object in JSON format. |
# | running() | Returns True/False on whether the operation is still running. |
# | done() | Returns True/False on whether the operation is completed. |
# | canceled() | Returns True/False on whether the operation was canceled. |
# | cancel() | Cancels the operation (this may take up to 30 seconds). |
# + id="create_aip_dataset"
TIMEOUT = 90
def create_dataset(name, schema, labels=None, timeout=TIMEOUT):
start_time = time.time()
try:
dataset = aip.Dataset(
display_name=name, metadata_schema_uri=schema, labels=labels
)
operation = clients["dataset"].create_dataset(parent=PARENT, dataset=dataset)
print("Long running operation:", operation.operation.name)
result = operation.result(timeout=TIMEOUT)
print("time:", time.time() - start_time)
print("response")
print(" name:", result.name)
print(" display_name:", result.display_name)
print(" metadata_schema_uri:", result.metadata_schema_uri)
print(" metadata:", dict(result.metadata))
print(" create_time:", result.create_time)
print(" update_time:", result.update_time)
print(" etag:", result.etag)
print(" labels:", dict(result.labels))
return result
except Exception as e:
print("exception:", e)
return None
result = create_dataset("flowers-" + TIMESTAMP, DATA_SCHEMA)
# + [markdown] id="dataset_id:result"
# Now save the unique dataset identifier for the `Dataset` resource instance you created.
# + id="dataset_id:result"
# The full unique ID for the dataset
dataset_id = result.name
# The short numeric ID for the dataset
dataset_short_id = dataset_id.split("/")[-1]
print(dataset_id)
# + [markdown] id="data_preparation:image,u_dataset"
# ### Data preparation
#
# The Vertex AI `Dataset` resource for images has some requirements for your data:
#
# - Images must be stored in a Cloud Storage bucket.
# - Each image file must be in an image format (PNG, JPEG, BMP, ...).
# - There must be an index file stored in your Cloud Storage bucket that contains the path and label for each image.
# - The index file must be either CSV or JSONL.
# + [markdown] id="data_import_format:icn,u_dataset,csv"
# #### CSV
#
# For image classification, the CSV index file has the requirements:
#
# - No heading.
# - First column is the Cloud Storage path to the image.
# - Second column is the label.
# + [markdown] id="import_file:u_dataset,csv"
# #### Location of Cloud Storage training data.
#
# Now set the variable `IMPORT_FILE` to the location of the CSV index file in Cloud Storage.
# + id="import_file:flowers,csv,icn"
IMPORT_FILE = (
"gs://cloud-samples-data/vision/automl_classification/flowers/all_data_v2.csv"
)
# + [markdown] id="quick_peek:csv"
# #### Quick peek at your data
#
# You will use a version of the Flowers dataset that is stored in a public Cloud Storage bucket, using a CSV index file.
#
# Start by doing a quick peek at the data. You count the number of examples by counting the number of rows in the CSV index file (`wc -l`) and then peek at the first few rows.
# + id="quick_peek:csv"
if "IMPORT_FILES" in globals():
FILE = IMPORT_FILES[0]
else:
FILE = IMPORT_FILE
count = ! gsutil cat $FILE | wc -l
print("Number of Examples", int(count[0]))
print("First 10 rows")
# ! gsutil cat $FILE | head
# + [markdown] id="import_data"
# ### Import data
#
# Now, import the data into your Vertex AI Dataset resource. Use this helper function `import_data` to import the data. The function does the following:
#
# - Uses the `Dataset` client.
# - Calls the client method `import_data`, with the following parameters:
# - `name`: The human readable name you give to the `Dataset` resource (e.g., flowers).
# - `import_configs`: The import configuration.
#
# - `import_configs`: A Python list containing a dictionary, with the key/value entries:
# - `gcs_sources`: A list of URIs to the paths of the one or more index files.
# - `import_schema_uri`: The schema identifying the labeling type.
#
# The `import_data()` method returns a long running `operation` object. This will take a few minutes to complete. If you are in a live tutorial, this would be a good time to ask questions, or take a personal break.
# + id="import_data"
def import_data(dataset, gcs_sources, schema):
config = [{"gcs_source": {"uris": gcs_sources}, "import_schema_uri": schema}]
print("dataset:", dataset_id)
start_time = time.time()
try:
operation = clients["dataset"].import_data(
name=dataset_id, import_configs=config
)
print("Long running operation:", operation.operation.name)
result = operation.result()
print("result:", result)
print("time:", int(time.time() - start_time), "secs")
print("error:", operation.exception())
print("meta :", operation.metadata)
print(
"after: running:",
operation.running(),
"done:",
operation.done(),
"cancelled:",
operation.cancelled(),
)
return operation
except Exception as e:
print("exception:", e)
return None
import_data(dataset_id, [IMPORT_FILE], LABEL_SCHEMA)
# + [markdown] id="train_automl_model"
# ## Train the model
#
# Now train an AutoML image classification model using your Vertex AI `Dataset` resource. To train the model, do the following steps:
#
# 1. Create an Vertex AI training pipeline for the `Dataset` resource.
# 2. Execute the pipeline to start the training.
# + [markdown] id="create_pipeline:automl"
# ### Create a training pipeline
#
# You may ask, what do we use a pipeline for? You typically use pipelines when the job (such as training) has multiple steps, generally in sequential order: do step A, do step B, etc. By putting the steps into a pipeline, we gain the benefits of:
#
# 1. Being reusable for subsequent training jobs.
# 2. Can be containerized and ran as a batch job.
# 3. Can be distributed.
# 4. All the steps are associated with the same pipeline job for tracking progress.
#
# Use this helper function `create_pipeline`, which takes the following parameters:
#
# - `pipeline_name`: A human readable name for the pipeline job.
# - `model_name`: A human readable name for the model.
# - `dataset`: The Vertex AI fully qualified dataset identifier.
# - `schema`: The dataset labeling (annotation) training schema.
# - `task`: A dictionary describing the requirements for the training job.
#
# The helper function calls the `Pipeline` client service'smethod `create_pipeline`, which takes the following parameters:
#
# - `parent`: The Vertex AI location root path for your `Dataset`, `Model` and `Endpoint` resources.
# - `training_pipeline`: the full specification for the pipeline training job.
#
# Let's look now deeper into the *minimal* requirements for constructing a `training_pipeline` specification:
#
# - `display_name`: A human readable name for the pipeline job.
# - `training_task_definition`: The dataset labeling (annotation) training schema.
# - `training_task_inputs`: A dictionary describing the requirements for the training job.
# - `model_to_upload`: A human readable name for the model.
# - `input_data_config`: The dataset specification.
# - `dataset_id`: The Vertex AI dataset identifier only (non-fully qualified) -- this is the last part of the fully-qualified identifier.
# - `fraction_split`: If specified, the percentages of the dataset to use for training, test and validation. Otherwise, the percentages are automatically selected by AutoML.
# + id="create_pipeline:automl"
def create_pipeline(pipeline_name, model_name, dataset, schema, task):
dataset_id = dataset.split("/")[-1]
input_config = {
"dataset_id": dataset_id,
"fraction_split": {
"training_fraction": 0.8,
"validation_fraction": 0.1,
"test_fraction": 0.1,
},
}
training_pipeline = {
"display_name": pipeline_name,
"training_task_definition": schema,
"training_task_inputs": task,
"input_data_config": input_config,
"model_to_upload": {"display_name": model_name},
}
try:
pipeline = clients["pipeline"].create_training_pipeline(
parent=PARENT, training_pipeline=training_pipeline
)
print(pipeline)
except Exception as e:
print("exception:", e)
return None
return pipeline
# + [markdown] id="task_requirements:automl,icn"
# ### Construct the task requirements
#
# Next, construct the task requirements. Unlike other parameters which take a Python (JSON-like) dictionary, the `task` field takes a Google protobuf Struct, which is very similar to a Python dictionary. Use the `json_format.ParseDict` method for the conversion.
#
# The minimal fields we need to specify are:
#
# - `multi_label`: Whether True/False this is a multi-label (vs single) classification.
# - `budget_milli_node_hours`: The maximum time to budget (billed) for training the model, where 1000 = 1 hour. For image classification, the budget must be a minimum of 8 hours.
# - `model_type`: The type of deployed model:
# - `CLOUD`: For deploying to Google Cloud.
# - `MOBILE_TF_LOW_LATENCY_1`: For deploying to the edge and optimizing for latency (response time).
# - `MOBILE_TF_HIGH_ACCURACY_1`: For deploying to the edge and optimizing for accuracy.
# - `MOBILE_TF_VERSATILE_1`: For deploying to the edge and optimizing for a trade off between latency and accuracy.
# - `disable_early_stopping`: Whether True/False to let AutoML use its judgement to stop training early or train for the entire budget.
#
# Finally, create the pipeline by calling the helper function `create_pipeline`, which returns an instance of a training pipeline object.
# + id="task_requirements:automl,icn"
PIPE_NAME = "flowers_pipe-" + TIMESTAMP
MODEL_NAME = "flowers_model-" + TIMESTAMP
task = json_format.ParseDict(
{
"multi_label": False,
"budget_milli_node_hours": 8000,
"model_type": "CLOUD",
"disable_early_stopping": False,
},
Value(),
)
response = create_pipeline(PIPE_NAME, MODEL_NAME, dataset_id, TRAINING_SCHEMA, task)
# + [markdown] id="pipeline_id:response"
# Now save the unique identifier of the training pipeline you created.
# + id="pipeline_id:response"
# The full unique ID for the pipeline
pipeline_id = response.name
# The short numeric ID for the pipeline
pipeline_short_id = pipeline_id.split("/")[-1]
print(pipeline_id)
# + [markdown] id="get_training_pipeline"
# ### Get information on a training pipeline
#
# Now get pipeline information for just this training pipeline instance. The helper function gets the job information for just this job by calling the the job client service's `get_training_pipeline` method, with the following parameter:
#
# - `name`: The Vertex AI fully qualified pipeline identifier.
#
# When the model is done training, the pipeline state will be `PIPELINE_STATE_SUCCEEDED`.
# + id="get_training_pipeline"
def get_training_pipeline(name, silent=False):
response = clients["pipeline"].get_training_pipeline(name=name)
if silent:
return response
print("pipeline")
print(" name:", response.name)
print(" display_name:", response.display_name)
print(" state:", response.state)
print(" training_task_definition:", response.training_task_definition)
print(" training_task_inputs:", dict(response.training_task_inputs))
print(" create_time:", response.create_time)
print(" start_time:", response.start_time)
print(" end_time:", response.end_time)
print(" update_time:", response.update_time)
print(" labels:", dict(response.labels))
return response
response = get_training_pipeline(pipeline_id)
# + [markdown] id="wait_training_complete"
# # Deployment
#
# Training the above model may take upwards of 20 minutes time.
#
# Once your model is done training, you can calculate the actual time it took to train the model by subtracting `end_time` from `start_time`. For your model, you will need to know the fully qualified Vertex AI Model resource identifier, which the pipeline service assigned to it. You can get this from the returned pipeline instance as the field `model_to_deploy.name`.
# + id="wait_training_complete"
while True:
response = get_training_pipeline(pipeline_id, True)
if response.state != aip.PipelineState.PIPELINE_STATE_SUCCEEDED:
print("Training job has not completed:", response.state)
model_to_deploy_id = None
if response.state == aip.PipelineState.PIPELINE_STATE_FAILED:
raise Exception("Training Job Failed")
else:
model_to_deploy = response.model_to_upload
model_to_deploy_id = model_to_deploy.name
print("Training Time:", response.end_time - response.start_time)
break
time.sleep(60)
print("model to deploy:", model_to_deploy_id)
# + [markdown] id="model_information"
# ## Model information
#
# Now that your model is trained, you can get some information on your model.
# + [markdown] id="evaluate_the_model:automl"
# ## Evaluate the Model resource
#
# Now find out how good the model service believes your model is. As part of training, some portion of the dataset was set aside as the test (holdout) data, which is used by the pipeline service to evaluate the model.
# + [markdown] id="list_model_evaluations:automl,icn"
# ### List evaluations for all slices
#
# Use this helper function `list_model_evaluations`, which takes the following parameter:
#
# - `name`: The Vertex AI fully qualified model identifier for the `Model` resource.
#
# This helper function uses the model client service's `list_model_evaluations` method, which takes the same parameter. The response object from the call is a list, where each element is an evaluation metric.
#
# For each evaluation (you probably only have one) we then print all the key names for each metric in the evaluation, and for a small set (`logLoss` and `auPrc`) you will print the result.
# + id="list_model_evaluations:automl,icn"
def list_model_evaluations(name):
response = clients["model"].list_model_evaluations(parent=name)
for evaluation in response:
print("model_evaluation")
print(" name:", evaluation.name)
print(" metrics_schema_uri:", evaluation.metrics_schema_uri)
metrics = json_format.MessageToDict(evaluation._pb.metrics)
for metric in metrics.keys():
print(metric)
print("logloss", metrics["logLoss"])
print("auPrc", metrics["auPrc"])
return evaluation.name
last_evaluation = list_model_evaluations(model_to_deploy_id)
# + [markdown] id="create_endpoint:automl"
# ## Deploy the `Model` resource
#
# Now deploy the trained Vertex AI `Model` resource you created with AutoML. This requires two steps:
#
# 1. Create an `Endpoint` resource for deploying the `Model` resource to.
#
# 2. Deploy the `Model` resource to the `Endpoint` resource.
# + [markdown] id="create_endpoint"
# ### Create an `Endpoint` resource
#
# Use this helper function `create_endpoint` to create an endpoint to deploy the model to for serving predictions, with the following parameter:
#
# - `display_name`: A human readable name for the `Endpoint` resource.
#
# The helper function uses the endpoint client service's `create_endpoint` method, which takes the following parameter:
#
# - `display_name`: A human readable name for the `Endpoint` resource.
#
# Creating an `Endpoint` resource returns a long running operation, since it may take a few moments to provision the `Endpoint` resource for serving. You call `response.result()`, which is a synchronous call and will return when the Endpoint resource is ready. The helper function returns the Vertex AI fully qualified identifier for the `Endpoint` resource: `response.name`.
# + id="create_endpoint"
ENDPOINT_NAME = "flowers_endpoint-" + TIMESTAMP
def create_endpoint(display_name):
endpoint = {"display_name": display_name}
response = clients["endpoint"].create_endpoint(parent=PARENT, endpoint=endpoint)
print("Long running operation:", response.operation.name)
result = response.result(timeout=300)
print("result")
print(" name:", result.name)
print(" display_name:", result.display_name)
print(" description:", result.description)
print(" labels:", result.labels)
print(" create_time:", result.create_time)
print(" update_time:", result.update_time)
return result
result = create_endpoint(ENDPOINT_NAME)
# + [markdown] id="endpoint_id:result"
# Now get the unique identifier for the `Endpoint` resource you created.
# + id="endpoint_id:result"
# The full unique ID for the endpoint
endpoint_id = result.name
# The short numeric ID for the endpoint
endpoint_short_id = endpoint_id.split("/")[-1]
print(endpoint_id)
# + [markdown] id="instance_scaling"
# ### Compute instance scaling
#
# You have several choices on scaling the compute instances for handling your online prediction requests:
#
# - Single Instance: The online prediction requests are processed on a single compute instance.
# - Set the minimum (`MIN_NODES`) and maximum (`MAX_NODES`) number of compute instances to one.
#
# - Manual Scaling: The online prediction requests are split across a fixed number of compute instances that you manually specified.
# - Set the minimum (`MIN_NODES`) and maximum (`MAX_NODES`) number of compute instances to the same number of nodes. When a model is first deployed to the instance, the fixed number of compute instances are provisioned and online prediction requests are evenly distributed across them.
#
# - Auto Scaling: The online prediction requests are split across a scaleable number of compute instances.
# - Set the minimum (`MIN_NODES`) number of compute instances to provision when a model is first deployed and to de-provision, and set the maximum (`MAX_NODES) number of compute instances to provision, depending on load conditions.
#
# The minimum number of compute instances corresponds to the field `min_replica_count` and the maximum number of compute instances corresponds to the field `max_replica_count`, in your subsequent deployment request.
# + id="instance_scaling"
MIN_NODES = 1
MAX_NODES = 1
# + [markdown] id="deploy_model:automatic"
# ### Deploy `Model` resource to the `Endpoint` resource
#
# Use this helper function `deploy_model` to deploy the `Model` resource to the `Endpoint` resource you created for serving predictions, with the following parameters:
#
# - `model`: The Vertex AI fully qualified model identifier of the model to upload (deploy) from the training pipeline.
# - `deploy_model_display_name`: A human readable name for the deployed model.
# - `endpoint`: The Vertex AI fully qualified endpoint identifier to deploy the model to.
#
# The helper function calls the `Endpoint` client service's method `deploy_model`, which takes the following parameters:
#
# - `endpoint`: The Vertex AI fully qualified `Endpoint` resource identifier to deploy the `Model` resource to.
# - `deployed_model`: The requirements specification for deploying the model.
# - `traffic_split`: Percent of traffic at the endpoint that goes to this model, which is specified as a dictionary of one or more key/value pairs.
# - If only one model, then specify as **{ "0": 100 }**, where "0" refers to this model being uploaded and 100 means 100% of the traffic.
# - If there are existing models on the endpoint, for which the traffic will be split, then use `model_id` to specify as **{ "0": percent, model_id: percent, ... }**, where `model_id` is the model id of an existing model to the deployed endpoint. The percents must add up to 100.
#
# Let's now dive deeper into the `deployed_model` parameter. This parameter is specified as a Python dictionary with the minimum required fields:
#
# - `model`: The Vertex AI fully qualified model identifier of the (upload) model to deploy.
# - `display_name`: A human readable name for the deployed model.
# - `disable_container_logging`: This disables logging of container events, such as execution failures (default is container logging is enabled). Container logging is typically enabled when debugging the deployment and then disabled when deployed for production.
# - `automatic_resources`: This refers to how many redundant compute instances (replicas). For this example, we set it to one (no replication).
#
# #### Traffic Split
#
# Let's now dive deeper into the `traffic_split` parameter. This parameter is specified as a Python dictionary. This might at first be a tad bit confusing. Let me explain, you can deploy more than one instance of your model to an endpoint, and then set how much (percent) goes to each instance.
#
# Why would you do that? Perhaps you already have a previous version deployed in production -- let's call that v1. You got better model evaluation on v2, but you don't know for certain that it is really better until you deploy to production. So in the case of traffic split, you might want to deploy v2 to the same endpoint as v1, but it only get's say 10% of the traffic. That way, you can monitor how well it does without disrupting the majority of users -- until you make a final decision.
#
# #### Response
#
# The method returns a long running operation `response`. We will wait sychronously for the operation to complete by calling the `response.result()`, which will block until the model is deployed. If this is the first time a model is deployed to the endpoint, it may take a few additional minutes to complete provisioning of resources.
# + id="deploy_model:automatic"
DEPLOYED_NAME = "flowers_deployed-" + TIMESTAMP
def deploy_model(
model, deployed_model_display_name, endpoint, traffic_split={"0": 100}
):
deployed_model = {
"model": model,
"display_name": deployed_model_display_name,
"automatic_resources": {
"min_replica_count": MIN_NODES,
"max_replica_count": MAX_NODES,
},
}
response = clients["endpoint"].deploy_model(
endpoint=endpoint, deployed_model=deployed_model, traffic_split=traffic_split
)
print("Long running operation:", response.operation.name)
result = response.result()
print("result")
deployed_model = result.deployed_model
print(" deployed_model")
print(" id:", deployed_model.id)
print(" model:", deployed_model.model)
print(" display_name:", deployed_model.display_name)
print(" create_time:", deployed_model.create_time)
return deployed_model.id
deployed_model_id = deploy_model(model_to_deploy_id, DEPLOYED_NAME, endpoint_id)
# + [markdown] id="make_prediction"
# ## Make a online prediction request
#
# Now do a online prediction to your deployed model.
# + [markdown] id="get_test_item"
# ### Get test item
#
# You will use an arbitrary example out of the dataset as a test item. Don't be concerned that the example was likely used in training the model -- we just want to demonstrate how to make a prediction.
# + id="get_test_item:automl,icn,csv"
# test_item = !gsutil cat $IMPORT_FILE | head -n1
if len(str(test_item[0]).split(",")) == 3:
_, test_item, test_label = str(test_item[0]).split(",")
else:
test_item, test_label = str(test_item[0]).split(",")
print(test_item, test_label)
# + [markdown] id="predict_item:automl,icn"
# ### Make a prediction
#
# Now you have a test item. Use this helper function `predict_item`, which takes the following parameters:
#
# - `filename`: The Cloud Storage path to the test item.
# - `endpoint`: The Vertex AI fully qualified identifier for the `Endpoint` resource where the `Model` resource was deployed.
# - `parameters_dict`: Additional filtering parameters for serving prediction results.
#
# This function calls the prediction client service's `predict` method with the following parameters:
#
# - `endpoint`: The Vertex AI fully qualified identifier for the `Endpoint` resource where the `Model` resource was deployed.
# - `instances`: A list of instances (encoded images) to predict.
# - `parameters`: Additional filtering parameters for serving prediction results.
# - `confidence_threshold`: The threshold for returning predictions. Must be between 0 and 1.
# - `max_predictions`: The maximum number of predictions to return, sorted by confidence.
#
# How does confidence_threshold affect the model accuracy? The threshold won't change the accuracy. What it changes is *recall* and *precision*.
#
# - Precision: The higher the precision the more likely what is predicted is the correct prediction, but return fewer predictions. Increasing the confidence threshold increases precision.
# - Recall: The higher the recall the more likely a correct prediction is returned in the result, but return more prediction with incorrect prediction. Decreasing the confidence threshold increases recall.
#
# In this example, you will predict for precision. You set the confidence threshold to 0.5 and the maximum number of predictions for a classification to two. Since, all the confidence values across the classes must add up to one, there are only two possible outcomes:
#
# 1. There is a tie, both 0.5, and returns two predictions.
# 2. One value is above 0.5 and the rest are below 0.5, and returns one prediction.
#
# #### Request
#
# Since in this example your test item is in a Cloud Storage bucket, you open and read the contents of the image using `tf.io.gfile.Gfile()`. To pass the test data to the prediction service, you encode the bytes into base64 -- which makes the content safe from modification while transmitting binary data over the network.
#
# The format of each instance is:
#
# { 'content': { 'b64': [base64_encoded_bytes] } }
#
# Since the `predict()` method can take multiple items (instances), send your single test item as a list of one test item. As a final step, you package the instances list into Google's protobuf format -- which is what you pass to the `predict()` method.
#
# #### Response
#
# The `response` object returns a list, where each element in the list corresponds to the corresponding image in the request. You will see in the output for each prediction (just one in this case):
#
# - `ids`: The instance ID of each data item.
# - `confidences`: The percent of confidence between 0 and 1 in the prediction for each class.
# - `displayNames`: The corresponding class names.
# + id="predict_item:automl,icn"
import base64
import tensorflow as tf
def predict_item(filename, endpoint, parameters_dict):
parameters = json_format.ParseDict(parameters_dict, Value())
with tf.io.gfile.GFile(filename, "rb") as f:
content = f.read()
# The format of each instance should conform to the deployed model's prediction input schema.
instances_list = [{"content": base64.b64encode(content).decode("utf-8")}]
instances = [json_format.ParseDict(s, Value()) for s in instances_list]
response = clients["prediction"].predict(
endpoint=endpoint, instances=instances, parameters=parameters
)
print("response")
print(" deployed_model_id:", response.deployed_model_id)
predictions = response.predictions
print("predictions")
for prediction in predictions:
print(" prediction:", dict(prediction))
predict_item(test_item, endpoint_id, {"confidenceThreshold": 0.5, "maxPredictions": 2})
# + [markdown] id="undeploy_model"
# ## Undeploy the `Model` resource
#
# Now undeploy your `Model` resource from the serving `Endpoint` resoure. Use this helper function `undeploy_model`, which takes the following parameters:
#
# - `deployed_model_id`: The model deployment identifier returned by the endpoint service when the `Model` resource was deployed to.
# - `endpoint`: The Vertex AI fully qualified identifier for the `Endpoint` resource where the `Model` is deployed to.
#
# This function calls the endpoint client service's method `undeploy_model`, with the following parameters:
#
# - `deployed_model_id`: The model deployment identifier returned by the endpoint service when the `Model` resource was deployed.
# - `endpoint`: The Vertex AI fully qualified identifier for the `Endpoint` resource where the `Model` resource is deployed.
# - `traffic_split`: How to split traffic among the remaining deployed models on the `Endpoint` resource.
#
# Since this is the only deployed model on the `Endpoint` resource, you simply can leave `traffic_split` empty by setting it to {}.
# + id="undeploy_model"
def undeploy_model(deployed_model_id, endpoint):
response = clients["endpoint"].undeploy_model(
endpoint=endpoint, deployed_model_id=deployed_model_id, traffic_split={}
)
print(response)
undeploy_model(deployed_model_id, endpoint_id)
# + [markdown] id="cleanup"
# # Cleaning up
#
# To clean up all GCP resources used in this project, you can [delete the GCP
# project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
#
# Otherwise, you can delete the individual resources you created in this tutorial:
#
# - Dataset
# - Pipeline
# - Model
# - Endpoint
# - Batch Job
# - Custom Job
# - Hyperparameter Tuning Job
# - Cloud Storage Bucket
# + id="cleanup"
delete_dataset = True
delete_pipeline = True
delete_model = True
delete_endpoint = True
delete_batchjob = True
delete_customjob = True
delete_hptjob = True
delete_bucket = True
# Delete the dataset using the Vertex AI fully qualified identifier for the dataset
try:
if delete_dataset and "dataset_id" in globals():
clients["dataset"].delete_dataset(name=dataset_id)
except Exception as e:
print(e)
# Delete the training pipeline using the Vertex AI fully qualified identifier for the pipeline
try:
if delete_pipeline and "pipeline_id" in globals():
clients["pipeline"].delete_training_pipeline(name=pipeline_id)
except Exception as e:
print(e)
# Delete the model using the Vertex AI fully qualified identifier for the model
try:
if delete_model and "model_to_deploy_id" in globals():
clients["model"].delete_model(name=model_to_deploy_id)
except Exception as e:
print(e)
# Delete the endpoint using the Vertex AI fully qualified identifier for the endpoint
try:
if delete_endpoint and "endpoint_id" in globals():
clients["endpoint"].delete_endpoint(name=endpoint_id)
except Exception as e:
print(e)
# Delete the batch job using the Vertex AI fully qualified identifier for the batch job
try:
if delete_batchjob and "batch_job_id" in globals():
clients["job"].delete_batch_prediction_job(name=batch_job_id)
except Exception as e:
print(e)
# Delete the custom job using the Vertex AI fully qualified identifier for the custom job
try:
if delete_customjob and "job_id" in globals():
clients["job"].delete_custom_job(name=job_id)
except Exception as e:
print(e)
# Delete the hyperparameter tuning job using the Vertex AI fully qualified identifier for the hyperparameter tuning job
try:
if delete_hptjob and "hpt_job_id" in globals():
clients["job"].delete_hyperparameter_tuning_job(name=hpt_job_id)
except Exception as e:
print(e)
if delete_bucket and "BUCKET_NAME" in globals():
# ! gsutil rm -r $BUCKET_NAME
| ai-platform-unified/notebooks/unofficial/gapic/automl/showcase_automl_image_classification_online.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Test your code with this notebook; replace your own code blocks in the for loop
# Your code should work if
def RGB_func(X):
'''
This is the 'fake' function to genration RGB from the x_A, x_B, x_C
Please replace this function with the actual experimental data
x is normalized.
'''
X = np.array(X)
RGB = X.dot(np.array([[1,0.1,0.0],[0.0,0.8,0.1],[0.3,0.0,1]]))
return np.around(RGB*255+0.02* np.random.randn(*RGB.shape)*255)
# +
import numpy as np
flag = False
############### TASK 1: Initial Conditions ###################
############### Please Change with your own code #############
X_init = [[10,100,90],[50, 20, 130]]
###############################################################
##******************Check the X_init format****************#
for x in np.array(X_init):
if len(x) != 3:
print('ERROR! each row needs to be 3 elements for color A, B and C')
print('you have', len(x), ' elements in one of the rows')
flag = True
if abs(sum(x)-200)> 0.01*200:
print('ERROR! each row needs to sum up closed to 200 (or 198 - 200) [uL]')
print('you have a sum of', sum(x), ' [uL] in one of the rows')
flag = True
##*********************************************************#
if flag != True:
X_new = np.array(X_init)
X = []
num_rounds = 25
for i in range(num_rounds):
if X ==[]:
X = X_new/200
else:
X = np.concatenate([X, X_new/200])
if len(X)>25:
X = X[0:25]
RGB_ref = [np.array([ 83., 108., 76.])+0.01* np.random.randn(3)*255] # each value is in the range 0-255
RGB_exp = np.array(RGB_func(X))
#######TASK 2: Optimization Metric (e.g. error function)############
############### Please Change with your own code #############
def MSE_func(RGB_exp,RGB_ref):
from sklearn.metrics import mean_squared_error
'''
Caclulate the metric for optimization:
Mean Square Error of experimental RGB measurements and the targeted RGB
'''
Y = []
for i in range(len(RGB_exp)):
Y.append([mean_squared_error(RGB_exp[i],RGB_ref[0])])
return np.array(Y)
Y = MSE_func(RGB_exp, RGB_ref)
####################################################################
##******************Check the Y format****************##
if len(np.shape(Y))==2:
if np.shape(Y)[1] != 1:
print('we need a two dimensional array with 1 element at each row')
print(np.shape(Y)[1])
flag = True
else:
print('we need a two dimensional array with 1 element at each row')
flag = True
if flag == True:
break
##*********************************************************#
#######TASK 3: Optimization Metric (e.g. error function)############
############### Please Change with your own code ##################
def optimizer_func(X, Y, BatchSize):
'''
Bayesian Optimizer Function
BatchSize is the number of suggestions for the next rounds
X should be the input variables
Y should be the metric to be optimized
'''
import GPy
from GPyOpt.methods import BayesianOptimization
bds = [{'name': 'x1', 'type': 'continuous', 'domain': (0, 1)},
{'name': 'x2', 'type': 'continuous', 'domain': (0, 1)},
{'name': 'x3', 'type': 'continuous', 'domain': (0, 1)},
]
constraints = [{'name': 'constr_1',
'constraint': 'x[:,0] + x[:,1] + x[:,2] -(1 + 0.005)'},###<= 0
{'name': 'constr_2',
'constraint': '(1- 0.005) - (x[:,0] + x[:,1] + x[:,2]) '}]###<= 0
kernel = GPy.kern.Matern52(input_dim=len(bds), ARD = True)
optimizer = BayesianOptimization(f=None,
domain=bds,
constraints = constraints,
model_type='GP',
acquisition_type ='EI',
acquisition_jitter = 0.1,
X=X,
Y=Y,
evaluator_type = 'local_penalization',
batch_size = BatchSize,
normalize_Y= True,
#noise_var = 0.02**2,
kernel = kernel
)
return optimizer
batch_size = 2
opt = optimizer_func(np.array(X), np.array(Y), batch_size)
X_new = opt.suggest_next_locations()*200
####################################################################
##******************Check the X_new format****************##
for x in np.array(X_init):
if len(x) != 3:
print('ERROR! each row needs to be 3 elements for color A, B and C')
print('you have', len(x), ' elements in one of the rows')
flag = True
if abs(sum(x)-200)> 0.01*200:
print('ERROR! each row needs to sum up closed to 200 (or 198 - 200) [uL]')
print('you have a sum of', sum(x), ' [uL] in one of the rows')
flag = True
if flag == True:
break
##*********************************************************#
##############TASK 4: Early Stop Condition#########################
############### You can change it or leave it like this #############
if len(RGB_exp)>25:
print('Reach the max number of samples')
break
def color_difference(rgb, rgb_ref):
return abs(rgb_ref[0]-rgb[0]), abs(rgb_ref[1]-rgb[1]), abs(rgb_ref[2]-rgb[2])
diff = []
for rgb in RGB_exp:
rgb_ref = RGB_ref[0]
r_err, g_err, b_err = color_difference(rgb, rgb_ref)
diff.append((r_err<=8) & (g_err<=8) & (b_err<=8))
if np.sum(diff)>=1:
print("Success! Early stop after collecting", len(diff), "samples")
print("Good match at ", np.arange(len(diff))[np.array(diff) == 1])
break
####################################################################
print("Lowest Error:", np.round(np.min(np.sqrt(Y))))
print("Lowest Error RGB", RGB_exp[np.argmin(np.sqrt(Y))])
print("Lowest ref RGB", np.around(RGB_ref[0]))
# -
| Week11/Color_Mixing_Example/test_code.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # d6tstack with Dask
#
# Dask is a great library for out-of-core computing. But if input files are not properly organized it quickly breaks. For example:
#
# 1) if columns are different between files, dask won't even read the data! It doesn't tell you what you need to do to fix it.
#
# 2) if column order is rearranged between files it will read data, but into the wrong columns and you won't notice it
#
# Dask can't handle those scenarios. With d6tstack you can easily fix the situation with just a few lines of code!
#
# For more instructions, examples and documentation see https://github.com/d6t/d6tstack
# ## Base Case: Columns are same between all files
# As a base case, we have input files which have consistent input columns and thus can be easily read in dask.
# +
import dask.dataframe as dd
# consistent format
ddf = dd.read_csv('test-data/input/test-data-input-csv-clean-*.csv')
ddf.compute()
# -
# ## Problem Case 1: Columns are different between files
# That worked well. But what happens if your input files have inconsistent columns across files? Say for example one file has a new column that the other files don't have.
# consistent format
ddf = dd.read_csv('test-data/input/test-data-input-csv-colmismatch-*.csv')
ddf.compute()
# ## Fixing the problem with d6stack
# Urgh! There's no way to use these files in dask. You don't even know what's going on. What file caused the problem? Why did it cause a problem? All you know is one file has more columns than the first file.
#
# You can either manually process those files or use d6tstack to easily check for such a situation and fix it with a few lines of code - no manual processing required. Let's take a look!
# +
import glob
import d6tstack.combine_csv
cfg_fnames = list(glob.glob('test-data/input/test-data-input-csv-colmismatch-*.csv'))
c = d6tstack.combine_csv.CombinerCSV(cfg_fnames)
# check columns
print('all equal',c.is_all_equal())
print('')
c.is_column_present()
# -
# Before using dask you can quickly use d6stack to check if all colums are consistent with `d6tstack.combine_csv.CombinerCSV.is_all_equal()`. If they are not consistent you can easily see which files are causing problems with `d6tstack.combine_csv.CombinerCSV.is_col_present()`, in this case there is a new column "profit2" in "test-data-input-csv-colmismatch-mar.csv".
#
# **Let's use d6stack to fix the situation.** We will use out-of-core processing with `d6tstack.combine_csv.CombinerCSVAdvanced.combine_save()` to save data from all files into one combined file with constistent columns. Any missing data is filled with NaN (to keep only common columns use `cfg_col_sel=c.col_preview['columns_common']`) Just 2 lines of code!
# out-of-core combining
fnames = d6tstack.combine_csv.CombinerCSV(cfg_fnames).to_csv_align(output_dir='test-data/output/')
# NB: Instead of `to_csv_align()` you can also run `to_csv_combine()` which creates a single combined file.
#
# Now you can read this in dask and do whatever you wanted to do in the first place.
# consistent format
ddf = dd.read_csv('test-data/output/d6tstack-test-data-input-csv-colmismatch-*.csv')
ddf.compute()
# ## Problem Case 2: Columns are reordered between files
# This is a sneaky case. The columns are the same but the order is different! Dask will read everything just fine without a warning but your data is totally messed up!
#
# In the example below, the "profit" column contains data from the "cost" column!
# consistent format
ddf = dd.read_csv('test-data/input/test-data-input-csv-reorder-*.csv')
ddf.compute()
# +
cfg_fnames = list(glob.glob('test-data/input/test-data-input-csv-reorder-*.csv'))
c = d6tstack.combine_csv.CombinerCSV(cfg_fnames)
# check columns
col_sniff = c.sniff_columns()
print('all columns equal?' , c.is_all_equal())
print('')
print('in what order do columns appear in the files?')
print('')
col_sniff['df_columns_order'].reset_index(drop=True)
# -
# Again, just a useful check before loading data into dask you can see that the columns don't line up. It's very fast to run because it only reads the headers, there's NO reason for you NOT to do it from a QA perspective.
#
# Same as above, the fix is the same few lines of code with d6stack.
# out-of-core combining
fnames = d6tstack.combine_csv.CombinerCSV(cfg_fnames).to_csv_align(output_dir='test-data/output/')
# consistent format
ddf = dd.read_csv('test-data/output/d6tstack-test-data-input-csv-reorder-*.csv')
ddf.compute()
| examples-dask.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from keras.models import Model
from keras.layers import Input
from keras.layers.convolutional import ZeroPadding2D
from keras import backend as K
def format_decimal(arr, places=6):
return [round(x * 10**places) / 10**places for x in arr]
# ### ZeroPadding2D
# **[convolutional.ZeroPadding2D.0] padding (1,1) on 3x5x2 input, dim_ordering=tf**
# +
data_in_shape = (3, 5, 2)
L = ZeroPadding2D(padding=(1, 1), dim_ordering='tf')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(250)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[convolutional.ZeroPadding2D.1] padding (1,1) on 3x5x2 input, dim_ordering=th**
# +
data_in_shape = (3, 5, 2)
L = ZeroPadding2D(padding=(1, 1), dim_ordering='th')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(250)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[convolutional.ZeroPadding2D.2] padding (3,2) on 2x6x4 input, dim_ordering=tf**
# +
data_in_shape = (2, 6, 4)
L = ZeroPadding2D(padding=(3, 2), dim_ordering='tf')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(252)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
# **[convolutional.ZeroPadding2D.3] padding (3,2) on 2x6x4 input, dim_ordering=th**
# +
data_in_shape = (2, 6, 4)
L = ZeroPadding2D(padding=(3, 2), dim_ordering='th')
layer_0 = Input(shape=data_in_shape)
layer_1 = L(layer_0)
model = Model(input=layer_0, output=layer_1)
# set weights to random (use seed for reproducibility)
np.random.seed(252)
data_in = 2 * np.random.random(data_in_shape) - 1
print('')
print('in shape:', data_in_shape)
print('in:', format_decimal(data_in.ravel().tolist()))
result = model.predict(np.array([data_in]))
print('out shape:', result[0].shape)
print('out:', format_decimal(result[0].ravel().tolist()))
# -
| notebooks/layers/convolutional/ZeroPadding2D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# do oversampling
import numpy as np
import pandas as pd
import competition_helpers
from sklearn import tree
from sklearn.ensemble import BaggingClassifier
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score
from imblearn.over_sampling import SMOTE, ADASYN
from collections import Counter
# +
# I/O configuration here
X_train = competition_helpers.read_csv("train_features.csv")
y_train = competition_helpers.read_csv("train_label.csv", remove_header=True)
X_test = competition_helpers.read_csv("test_features.csv")
submission_col = np.array(pd.read_csv("test_features.csv", header=None).iloc[: , 0]).ravel()
submission_file_name = "results/voting_default_submission1.csv"
print(X_train.shape, y_train.shape, X_test.shape)
print(sorted(Counter(list(y_train.flatten())).items()))
X_resampled, y_resampled = SMOTE().fit_resample(X_train, y_train.ravel())
print(sorted(Counter(list(y_resampled.flatten())).items()))
# -
# 5 fold cross validation
# train_test_split = competition_helpers.kfold_stratified_split(X_train, y_train, 5,False)
# With standardization
standardized_train_test_split = competition_helpers.kfold_stratified_split(X_resampled, y_resampled.reshape((-1, 1)), 5,True)
# +
# # 5 fold train test split results
# results = []
# for estimators_ in [50, 100, 150]:
# for lr in [0.1, 0.5, 1, 5]:
# for [(X_train_cv, y_train_cv), (X_test_cv, y_test_cv)] in train_test_split:
# clf = AdaBoostClassifier(random_state=42,
# base_estimator=tree.DecisionTreeClassifier(
# max_depth=None, min_samples_split=60, min_samples_leaf= 30
# ),
# n_estimators=estimators_,
# learning_rate=lr
# )
# clf.fit(X_train_cv, y_train_cv.ravel())
# prediction = clf.predict(X_test_cv)
# accuracy = accuracy_score(y_test_cv.ravel(), prediction.ravel())
# precision = precision_score(y_test_cv.ravel(), prediction.ravel())
# recall = recall_score(y_test_cv.ravel(), prediction.ravel())
# f1 = f1_score(y_test_cv.ravel(), prediction.ravel())
# results.append([accuracy, precision, recall, f1])
# measures = np.sum(np.array(results), axis=0) / len(results)
# print("n_estimators: {} learning rate: {} measures: {}".format(estimators_, lr, measures))
# +
results = []
for [(X_train_cv, y_train_cv), (X_test_cv, y_test_cv)] in standardized_train_test_split:
clf = BaggingClassifier(base_estimator=tree.DecisionTreeClassifier(), n_estimators=100)
clf.fit(X_train_cv, y_train_cv.ravel())
prediction = clf.predict(X_test_cv)
accuracy = accuracy_score(y_test_cv.ravel(), prediction.ravel())
precision = precision_score(y_test_cv.ravel(), prediction.ravel())
recall = recall_score(y_test_cv.ravel(), prediction.ravel())
f1 = f1_score(y_test_cv.ravel(), prediction.ravel())
results.append([accuracy, precision, recall, f1])
measures = np.sum(np.array(results), axis=0) / len(results)
# -
print(measures)
# +
# fitting the test dataset
clf = BaggingClassifier(base_estimator=tree.DecisionTreeClassifier(), n_estimators=100)
clf.fit(X_resampled, y_resampled.ravel())
prediction = clf.predict(X_test)
# -
pd.DataFrame({"id": submission_col, "label": prediction}).to_csv(submission_file_name, encoding='utf-8', index=False)
| competition/BaggingClassifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="7ivmxKvQCfTG"
# # **Importing necessary library**
# + id="cbeTYHqyCe-M" executionInfo={"status": "ok", "timestamp": 1627283260344, "user_tz": -345, "elapsed": 394, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
from scipy.stats import norm
# + [markdown] id="qMVj8N2f_68K"
# # **Load the dataset**
# + colab={"base_uri": "https://localhost:8080/"} id="zmc3smGsAEuO" executionInfo={"status": "ok", "timestamp": 1627283262202, "user_tz": -345, "elapsed": 1270, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="d6e26c30-14f4-4d53-c347-1e20ce3c513e"
from google.colab import drive
drive.mount('/content/drive')
# + id="b3S_eqh7AGUW" executionInfo={"status": "ok", "timestamp": 1627283262203, "user_tz": -345, "elapsed": 79, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}}
DATA_DIR = "/content/drive/MyDrive/Colab Notebooks/datamining/dataset/dataset.csv"
# + id="po1uljU5AY9H" executionInfo={"status": "ok", "timestamp": 1627283262204, "user_tz": -345, "elapsed": 79, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}}
dataset = pd.read_csv(DATA_DIR)
# + [markdown] id="1tKi9jrPCVhF"
# # **Data Analysis**
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="kxErGL-XAlgi" executionInfo={"status": "ok", "timestamp": 1627283262205, "user_tz": -345, "elapsed": 79, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="891236a1-e7a2-4c4e-e7b8-eec30b461d05"
dataset.head()
# + [markdown] id="dX0VmaoSFid-"
# # understanding the dataset
# ## all feature name:
# 1. brandname
# 2. model
# 1. cpucore
# 2. refreshclock
# 1. gpu
# 2. ram
# 1. ssd
# 2. rom
# 1. display
# 2. displayquality
# 1. price (need to predict)
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="2G8mhZE4AlcN" executionInfo={"status": "ok", "timestamp": 1627283262207, "user_tz": -345, "elapsed": 78, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="004e081c-741e-4e6d-e8dc-b8d6ba0a72b7"
print(dataset.shape)
# + [markdown] id="O4HjtgEAErpU"
#
# ```
# Missing Values
# All The Numerical Variables
# Distribution of the Numerical Variables
# Categorical Variables
# Cardinality of Categorical Variables
# Outliers
# Relationship between independent and dependent feature(SalePrice)
# ```
#
#
#
# + [markdown] id="T7u8CIuSGfW-"
# # Determinig the category of features
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="FE125JgjAlV2" executionInfo={"status": "ok", "timestamp": 1627283262208, "user_tz": -345, "elapsed": 68, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="55ee0732-6f5a-4fc7-c872-ac0182a60aa9"
dataset.head(845)
# + colab={"base_uri": "https://localhost:8080/"} id="JELkHLfdHXax" executionInfo={"status": "ok", "timestamp": 1627283262208, "user_tz": -345, "elapsed": 64, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="a0646748-c97f-4c81-8a37-b69ce8d14569"
dataset.dtypes
# + colab={"base_uri": "https://localhost:8080/"} id="XDklevt8656T" executionInfo={"status": "ok", "timestamp": 1627283262209, "user_tz": -345, "elapsed": 58, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="514ef6c3-94c7-44fc-d3f7-bb4947fad0f0"
dataset.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="rLpCRSmW7Qq2" executionInfo={"status": "ok", "timestamp": 1627283262210, "user_tz": -345, "elapsed": 52, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="3cf34de3-4f79-4464-d726-29a77e5c9c46"
dataset.describe()
# + [markdown] id="VwQ4wLUBGw8A"
# # understanding the dataset
# ## all feature name:
# 1. brandname (categorical)
# 2. model (categorical)
# 1. cpucore (categorical)
# 2. refreshclock (numerical)
# 1. gpu (categorical)
# 2. ram (numerical)
# 1. ssd (numerical)
# 2. rom (numerical)
# 1. display (numerical)
# 2. displayquality (categorical)
# 1. price (numerical)
#
#
# + [markdown] id="lNn_TY5qINUB"
# # **Missing Values**
# + [markdown] id="0FEUHQLSRPfd"
# drop the duplicate rows
# + colab={"base_uri": "https://localhost:8080/"} id="Tyz8m8R1JGWW" executionInfo={"status": "ok", "timestamp": 1627283262212, "user_tz": -345, "elapsed": 52, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="6b24b514-9a2e-4d2d-c048-08b4625ed985"
print(dataset['brandname'].isnull().sum())
print(dataset['cpucore'].isnull().sum())
print(dataset['refreshrate'].isnull().sum())
print(dataset['gpu'].isnull().sum())
print(dataset['ram'].isnull().sum())
print(dataset['ssd'].isnull().sum())
print(dataset['rom'].isnull().sum())
print(dataset['display'].isnull().sum())
print(dataset['displayquality'].isnull().sum())
# + colab={"base_uri": "https://localhost:8080/"} id="rM-Wt3Q82X7P" executionInfo={"status": "ok", "timestamp": 1627283262212, "user_tz": -345, "elapsed": 45, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AO<KEY>FLTjgo=s64", "userId": "17265195811399595189"}} outputId="7ce7e967-1845-46ae-93eb-1a8f68927b6c"
#seeing no.of non-null values and datatype in each column
dataset.info()
# + colab={"base_uri": "https://localhost:8080/"} id="30Ca09zCAlTG" executionInfo={"status": "ok", "timestamp": 1627283262213, "user_tz": -345, "elapsed": 39, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="09f1bc8b-caa3-4be2-c3ce-75476421a027"
## Here we will check the percentage of nan values present in each feature
## 1 -step make the list of features which has missing values
features_with_na=[features for features in dataset.columns if dataset[features].isnull().sum()>1]
## 2- step print the feature name and the percentage of missing values
for feature in features_with_na:
print(feature, np.round(dataset[feature].isnull().mean(), 4), ' % missing values')
# + [markdown] id="n1weoTxlLqyy"
# # **Handling the null values**
# + [markdown] id="2kAy2y79MSw9"
#
# Since they are many missing values, we need to find the relationship between missing values and Price.
#
# 1. Let's plot some diagram for this relationship
#
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="eZatgx2ULp8N" executionInfo={"status": "ok", "timestamp": 1627283263584, "user_tz": -345, "elapsed": 1403, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="8c0caf81-3ed8-408c-f83b-b2ff1d9ce818"
for feature in features_with_na:
data = dataset.copy()
# let's make a variable that indicates 1 if the observation was missing or zero otherwise
data[feature] = np.where(data[feature].isnull(), 1, 0)
# let's calculate the mean SalePrice where the information is missing or present
data.groupby(feature)['price'].median().plot.bar()
plt.title(feature)
plt.show()
# + [markdown] id="F71OILvQPPZf"
#
# Here With the relation between the missing values and the dependent variable is clearly visible.So We need to replace these nan values with something meaningful which we will do in the Feature Engineering section
# + [markdown] id="c6xZZvl15EQR"
#
# [website from](https://analyticsindiamag.com/5-ways-handle-missing-values-machine-learning-datasets/)
#
#
# # **1. Deleting Rows**
#
# This method commonly used to handle the null values. Here, we either delete a particular row if it has a null value for a particular feature and a particular column if it has more than 70-75% of missing values. This method is advised only when there are enough samples in the data set. One has to make sure that after we have deleted the data, there is no addition of bias. Removing the data will lead to loss of information which will not give the expected results while predicting the output.
#
# # Pros:
# Complete removal of data with missing values results in robust and highly accurate model
# Deleting a particular row or a column with no specific information is better, since it does not have a high weightage
# # Cons:
# Loss of information and data
# Works poorly if the percentage of missing values is high (say 30%), compared to the whole dataset
#
#
#
#
# # **2. Replacing With Mean/Median/Mode**
#
# This strategy can be applied on a feature which has numeric data like the age of a person or the ticket fare. We can calculate the mean, median or mode of the feature and replace it with the missing values. This is an approximation which can add variance to the data set. But the loss of the data can be negated by this method which yields better results compared to removal of rows and columns. Replacing with the above three approximations are a statistical approach of handling the missing values. This method is also called as leaking the data while training. Another way is to approximate it with the deviation of neighbouring values. This works better if the data is linear.
#
#
#
# # Pros:
# This is a better approach when the data size is small
# It can prevent data loss which results in removal of the rows and columns
# # Cons:
# Imputing the approximations add variance and bias
# Works poorly compared to other multiple-imputations method
#
#
#
#
# # **3. Assigning An Unique Category**
#
#
#
# A categorical feature will have a definite number of possibilities, such as gender, for example. Since they have a definite number of classes, we can assign another class for the missing values. Here, the features Cabin and Embarked have missing values which can be replaced with a new category, say, U for ‘unknown’. This strategy will add more information into the dataset which will result in the change of variance. Since they are categorical, we need to find one hot encoding to convert it to a numeric form for the algorithm to understand it. Let us look at how it can be done in Python:
#
#
#
# # Pros:
# Less possibilities with one extra category, resulting in low variance after one hot encoding — since it is categorical
# Negates the loss of data by adding an unique category
# # Cons:
# Adds less variance
# Adds another feature to the model while encoding, which may result in poor performance
#
#
#
# # **4. Predicting The Missing Values**
# Using the features which do not have missing values, we can predict the nulls with the help of a machine learning algorithm. This method may result in better accuracy, unless a missing value is expected to have a very high variance. We will be using linear regression to replace the nulls in the feature ‘age’, using other available features. One can experiment with different algorithms and check which gives the best accuracy instead of sticking to a single algorithm.
#
#
#
# # Pros:
# Imputing the missing variable is an improvement as long as the bias from the same is smaller than the omitted variable bias
# Yields unbiased estimates of the model parameters
# # Cons:
# Bias also arises when an incomplete conditioning set is used for a categorical variable
# Considered only as a proxy for the true values
#
#
#
#
# # **5. Using Algorithms Which Support Missing Values**
#
# KNN is a machine learning algorithm which works on the principle of distance measure. This algorithm can be used when there are nulls present in the dataset. While the algorithm is applied, KNN considers the missing values by taking the majority of the K nearest values. In this particular dataset, taking into account the person’s age, sex, class etc, we will assume that people having same data for the above mentioned features will have the same kind of fare.
#
# Unfortunately, the SciKit Learn library for the K – Nearest Neighbour algorithm in Python does not support the presence of the missing values.
#
# Another algorithm which can be used here is RandomForest. This model produces a robust result because it works well on non-linear and the categorical data. It adapts to the data structure taking into consideration of the high variance or the bias, producing better results on large datasets.
#
# # Pros:
# Does not require creation of a predictive model for each attribute with missing data in the dataset
# Correlation of the data is neglected
# ## Cons:
# Is a very time consuming process and it can be critical in data mining where large databases are being extracted
# Choice of distance functions can be Euclidean, Manhattan etc. which is do not yield a robust result
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="gI0TrboW4X1L" executionInfo={"status": "ok", "timestamp": 1627283263586, "user_tz": -345, "elapsed": 120, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="b9279f00-84cd-41e2-cee0-af0f22e6a5f6"
dataset.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="fanXZupq4hs9" executionInfo={"status": "ok", "timestamp": 1627283263587, "user_tz": -345, "elapsed": 115, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="65231708-6dc4-4e1d-9f89-94143c5c524b"
dataset.notnull().sum()
# + id="GIR77-wX2r7u" executionInfo={"status": "ok", "timestamp": 1627283263588, "user_tz": -345, "elapsed": 104, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}}
#setting brandname as index(ie brandname instead of index(symbol) number)
dataset.set_index("brandname",inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="Za7Z10W12w96" executionInfo={"status": "ok", "timestamp": 1627283263590, "user_tz": -345, "elapsed": 101, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="3a418b4b-cfc1-4680-9ae1-19150d4e3d4e"
dataset
# + id="YErU1yU92w5u" executionInfo={"status": "ok", "timestamp": 1627283263591, "user_tz": -345, "elapsed": 96, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}}
#we have total 10 columns now(excluding brandname)
#keeping only those rows which have alteast 5 valid values i.e. deleting rows with 5 or more inavlid(null) values
dataset1=dataset.dropna(thresh=5)
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="pLsl99jp2w22" executionInfo={"status": "ok", "timestamp": 1627283263593, "user_tz": -345, "elapsed": 94, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="d958db64-c00f-4e96-c988-c53a83040c0d"
dataset1
# + id="5NV_YdfR2w0N" executionInfo={"status": "ok", "timestamp": 1627283263595, "user_tz": -345, "elapsed": 91, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}}
# replacing null value for ssd and hdd by 0 as when one has valid value other has 0 in most cases.
dataset2=dataset1.fillna({'ssd':0,
'rom':0
})
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="2pqaxZiR4OAD" executionInfo={"status": "ok", "timestamp": 1627283263595, "user_tz": -345, "elapsed": 90, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="497b270e-769b-4633-f706-c82196b3b72f"
dataset2
# + id="bv4_SBrw4Dv2" executionInfo={"status": "ok", "timestamp": 1627283263596, "user_tz": -345, "elapsed": 67, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}}
#we have total 10 columns now(excluding brandname)
#keeping only those rows which have alteast 5 valid values i.e. deleting rows with 5 or more inavlid(null) values after replacing missing(null) values in rom and ssd with 0.
dataset3=dataset2.dropna(thresh=5)
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="PPKu-fae4Dre" executionInfo={"status": "ok", "timestamp": 1627283263598, "user_tz": -345, "elapsed": 68, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="48e045c7-6922-4753-c238-3cc914d14d5b"
dataset3
# + id="nmanBS-q4DpT" executionInfo={"status": "ok", "timestamp": 1627283263599, "user_tz": -345, "elapsed": 67, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}}
#replacing null values of gpu by intel
dataset4=dataset3.fillna({'gpu':"Intel"
})
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="l0irPVUJ4o0Z" executionInfo={"status": "ok", "timestamp": 1627283263600, "user_tz": -345, "elapsed": 67, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="60319ce2-c742-49be-8bf0-5481f146fe97"
dataset4
# + colab={"base_uri": "https://localhost:8080/"} id="AL01q1Qv4owA" executionInfo={"status": "ok", "timestamp": 1627283263601, "user_tz": -345, "elapsed": 67, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="9d8815b5-8286-4ca4-aaa9-33db174a0b5e"
#checking number of null(NaN) values in each column after filling missing(null) values of gpu,rom,ssd
print("\nTotal missing value in each column in Dataframe:\n",dataset4.isnull().sum())
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="qlUD4tJ74otY" executionInfo={"status": "ok", "timestamp": 1627283263602, "user_tz": -345, "elapsed": 57, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="a87138ff-63ad-4bfd-a65c-1f4e0a0c309c"
#reseting index to default
dataset4.reset_index()
# + colab={"base_uri": "https://localhost:8080/"} id="5192ix-g4Dmf" executionInfo={"status": "ok", "timestamp": 1627283264281, "user_tz": -345, "elapsed": 734, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="47bc1743-a89a-4aab-9b1e-01bbf7c7899a"
#checking number of null(NaN) values in each column
print("\nTotal missing value in each column in Dataframe:\n",dataset4.isnull().sum())
# + id="daVfs4_84Dj5" executionInfo={"status": "ok", "timestamp": 1627283264282, "user_tz": -345, "elapsed": 75, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}}
# Impute display quality and cpucore with mode (Categorical Variable) by "Generalized imputation"
dataset4['displayquality'].fillna(dataset4['displayquality'].mode()[0], inplace = True)
dataset4['cpucore'].fillna(dataset4['cpucore'].mode()[0], inplace = True)
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="SrRBliAP4DhZ" executionInfo={"status": "ok", "timestamp": 1627283264284, "user_tz": -345, "elapsed": 76, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="c8e031db-4963-42b0-90cb-b17d920f68a0"
dataset4
# + id="pWyUSuTg5NW7" executionInfo={"status": "ok", "timestamp": 1627283264285, "user_tz": -345, "elapsed": 76, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}}
# Impute ram with median(Numerical value) by "Generalized imputation"
dataset4['ram'].fillna(dataset4['ram'].median(), inplace = True)
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="cmqwLC4V5NR7" executionInfo={"status": "ok", "timestamp": 1627283264286, "user_tz": -345, "elapsed": 76, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="ae82bac7-946a-449c-c40f-11be7840dbd1"
dataset4
# + id="IzFrL2Vu5NPE" executionInfo={"status": "ok", "timestamp": 1627283264287, "user_tz": -345, "elapsed": 76, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}}
# Impute display with median(Numerical value) correlating it with brandname by "Special case imputation"
dataset4['display'].fillna(dataset4.groupby(["brandname"])['display'].transform('median'),inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="cddRHJjI5NMh" executionInfo={"status": "ok", "timestamp": 1627283264288, "user_tz": -345, "elapsed": 76, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="0bc36bad-220d-418a-96c0-b5d618dec653"
dataset4
# + id="H4Ac_e7Z5NJl" executionInfo={"status": "ok", "timestamp": 1627283264289, "user_tz": -345, "elapsed": 75, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}}
# Impute display with mode(Numerical value) correlating it with both display and displayquality by "Special case imputation"
dataset4['refreshrate'].fillna(dataset4.groupby(["display","displayquality"])['refreshrate'].transform('median'),inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="PEOAhxV05NG8" executionInfo={"status": "ok", "timestamp": 1627283264289, "user_tz": -345, "elapsed": 75, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="deecd8e6-b035-491d-b1fc-f64bb46e0389"
dataset4
# + colab={"base_uri": "https://localhost:8080/"} id="eRaBY5fO6NkD" executionInfo={"status": "ok", "timestamp": 1627283264290, "user_tz": -345, "elapsed": 74, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="48886b6f-5673-4c53-8ef2-38108f2fb276"
#checking number of null(NaN) values in each column
print("\nTotal missing value in each column in Dataframe:\n",dataset4.isnull().sum())
# + id="1G_7P2596Nf2" executionInfo={"status": "ok", "timestamp": 1627283264290, "user_tz": -345, "elapsed": 67, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}}
# Impute display with mode(Numerical value) correlating it with displayquality only by "Special case imputation"
dataset4['refreshrate'].fillna(dataset4.groupby(["displayquality"])['refreshrate'].transform('median'),inplace=True)
# + colab={"base_uri": "https://localhost:8080/"} id="LOUtjCLF6Nc-" executionInfo={"status": "ok", "timestamp": 1627283264291, "user_tz": -345, "elapsed": 67, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="a87b09a0-7e7d-40b9-8715-70f2b3033d47"
#checking number of null(NaN) values in each column
print("\nTotal missing value in each column in Dataframe:\n",dataset4.isnull().sum())
# + id="8pOz85Df6NaU" executionInfo={"status": "ok", "timestamp": 1627283264291, "user_tz": -345, "elapsed": 62, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}}
# Impute display with mode(Numerical value) correlating it with display only by "Special case imputation"
dataset4['refreshrate'].fillna(dataset4.groupby(["display"])['refreshrate'].transform('median'),inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="NAIiSYIm6NXn" executionInfo={"status": "ok", "timestamp": 1627283264292, "user_tz": -345, "elapsed": 63, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="1e76c6bb-1089-484a-e076-0827f3643214"
dataset4
# + colab={"base_uri": "https://localhost:8080/"} id="QL4ANIml7h7k" executionInfo={"status": "ok", "timestamp": 1627283264293, "user_tz": -345, "elapsed": 62, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="8c2a04b0-18a4-4b26-f59f-31910c4fbea1"
#checking number of null(NaN) values in each column
print("\nTotal missing value in each column in Dataframe:\n",dataset4.isnull().sum())
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="ZYgBQMv79fJB" executionInfo={"status": "ok", "timestamp": 1627283264293, "user_tz": -345, "elapsed": 57, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="3a71ee9c-f37d-465b-b30c-13ff4f2a1717"
dataset4
# + id="-lugHngZr6PM" executionInfo={"status": "ok", "timestamp": 1627284077887, "user_tz": -345, "elapsed": 451, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}}
# After null value analysis complition
dataset4.to_csv("/content/drive/MyDrive/Colab Notebooks/datamining/removednullvaluedataset.csv")
# + colab={"base_uri": "https://localhost:8080/", "height": 252} id="zCz7He1LLvsN" executionInfo={"status": "ok", "timestamp": 1627283264294, "user_tz": -345, "elapsed": 56, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="83eaad74-50d7-4390-8834-f476402af53d"
# list of numerical variables
numerical_features = [feature for feature in dataset4.columns if dataset4[feature].dtypes != 'O']
print('Number of numerical variables: ', len(numerical_features))
# visualise the numerical variables
dataset4[numerical_features].head()
# + colab={"base_uri": "https://localhost:8080/"} id="W2VnvFju93xX" executionInfo={"status": "ok", "timestamp": 1627283264295, "user_tz": -345, "elapsed": 51, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="6c61f8ae-adeb-423e-f101-ac70e00bbfae"
dataset4.info()
# + [markdown] id="7uC8zOrQApv4"
# # **outlier detection and removal**
# + id="JxrVQcgoAkt4" colab={"base_uri": "https://localhost:8080/", "height": 450} executionInfo={"status": "ok", "timestamp": 1627283264295, "user_tz": -345, "elapsed": 45, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="2e86468e-d632-4184-e414-b5ac77232700"
dataset4
# + [markdown] id="8OasyeZ5dzTd"
#
#
# ```
# Most common causes of outliers on a data set:
# Data entry errors (human errors)
# Measurement errors (instrument errors)
# Experimental errors (data extraction or experiment planning/executing errors)
# Intentional (dummy outliers made to test detection methods)
# Data processing errors (data manipulation or data set unintended mutations)
# Sampling errors (extracting or mixing data from wrong or various sources)
# Natural (not an error, novelties in data)
# ```
#
#
# + [markdown] id="TO8mcPV1d-tl"
#
#
# ```
# Some of the most popular methods for outlier detection are:
# Z-Score or Extreme Value Analysis (parametric)
# Probabilistic and Statistical Modeling (parametric)
# Linear Regression Models (PCA, LMS)
# Proximity Based Models (non-parametric)
# Information Theory Models
# High Dimensional Outlier Detection Methods (high dimensional sparse data)
# ```
#
#
# + [markdown] id="_ewygenheKMz"
# **Z-Score**
#
# The z-score or standard score of an observation is a metric that indicates how many standard deviations a data point is from the sample’s mean, assuming a gaussian distribution. This makes z-score a parametric method. Very frequently data points are not to described by a gaussian distribution, this problem can be solved by applying transformations to data ie: scaling it.
# Some Python libraries like Scipy and Sci-kit Learn have easy to use functions and classes for a easy implementation along with Pandas and Numpy.
# After making the appropriate transformations to the selected feature space of the dataset, the z-score of any data point can be calculated with the following expression:
#
# z = (x - mean)/standard_deviation
#
#
# When computing the z-score for each sample on the data set a threshold must be specified. Some good ‘thumb-rule’ thresholds can be: 2.5, 3, 3.5 or more standard deviations.
#
#
#
#
# [refrence page](https://towardsdatascience.com/a-brief-overview-of-outlier-detection-techniques-1e0b2c19e561)
#
# + [markdown] id="M-B41y8YfesD"
# # Z-Score or Extreme Value Analysis (parametric)
# + colab={"base_uri": "https://localhost:8080/", "height": 450} id="QdAcOmU-d-S3" executionInfo={"status": "ok", "timestamp": 1627283264296, "user_tz": -345, "elapsed": 45, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="36f03980-d0cc-4892-d89d-af2acdc85511"
dataset4
# + id="BGfPmcbCAkrF" colab={"base_uri": "https://localhost:8080/", "height": 279} executionInfo={"status": "ok", "timestamp": 1627283271894, "user_tz": -345, "elapsed": 1037, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="a18a00ea-d9ad-47a0-d64b-839723673169"
# in refresh rate
plt.hist(dataset4.refreshrate, bins=20, rwidth=0.8)
plt.xlabel('refresh rate (hz)')
plt.ylabel('Count')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="Sw6AULL7ivnH" executionInfo={"status": "ok", "timestamp": 1627283275684, "user_tz": -345, "elapsed": 713, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="88894f5c-e3f6-4796-d71a-386fab920ecf"
plt.hist(dataset4.refreshrate, bins=20, rwidth=0.8, density=True)
plt.xlabel('refresh rate (hz)')
plt.ylabel('Count')
rng = np.arange(dataset4.refreshrate.min(), dataset4.refreshrate.max(), 0.1)
plt.plot(rng, norm.pdf(rng,dataset4.refreshrate.mean(),dataset4.refreshrate.std()))
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="ICJ7u7hLf2hq" executionInfo={"status": "ok", "timestamp": 1627283280604, "user_tz": -345, "elapsed": 922, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="c74c1944-3bc5-4e0f-d603-d161fe6f2c33"
# in ram
plt.hist(dataset4.ram, bins=20, rwidth=0.8)
plt.xlabel('ram (GB)')
plt.ylabel('Count')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="gHBe6vCJjQwq" executionInfo={"status": "ok", "timestamp": 1627283282156, "user_tz": -345, "elapsed": 14, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="e4d5bb90-a791-414c-b1db-536561b79b59"
plt.hist(dataset4.ram, bins=20, rwidth=0.8, density=True)
plt.xlabel('ram (GB)')
plt.ylabel('Count')
rng = np.arange(dataset4.ram.min(), dataset4.ram.max(), 0.1)
plt.plot(rng, norm.pdf(rng,dataset4.ram.mean(),dataset4.ram.std()))
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="UeOEGjT7f2dN" executionInfo={"status": "ok", "timestamp": 1627283287139, "user_tz": -345, "elapsed": 1226, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="24cc71de-1bac-412d-81d1-732c3779f094"
# in rom
plt.hist(dataset4.rom, bins=20, rwidth=0.8)
plt.xlabel('rom (GB)')
plt.ylabel('Count')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="EdV541R3jbGS" executionInfo={"status": "ok", "timestamp": 1627283290097, "user_tz": -345, "elapsed": 909, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="77d83b6b-e40d-4cc6-856a-e6d91a7af601"
plt.hist(dataset4.rom, bins=20, rwidth=0.8, density=True)
plt.xlabel('rom (GB)')
plt.ylabel('Count')
rng = np.arange(dataset4.rom.min(), dataset4.rom.max(), 0.1)
plt.plot(rng, norm.pdf(rng,dataset4.rom.mean(),dataset4.rom.std()))
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="JUGTi4gbh12o" executionInfo={"status": "ok", "timestamp": 1627283292537, "user_tz": -345, "elapsed": 671, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="1d18e018-112c-45de-8c5c-f849aa824ec5"
# in display
plt.hist(dataset4.display, bins=20, rwidth=0.8)
plt.xlabel('display (inch)')
plt.ylabel('Count')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="o73yPUhAjlOf" executionInfo={"status": "ok", "timestamp": 1627283296440, "user_tz": -345, "elapsed": 464, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="bb562bd6-5fb6-4f2a-a511-f25d6a3cae6f"
plt.hist(dataset4.display, bins=20, rwidth=0.8, density=True)
plt.xlabel('display (inch)')
plt.ylabel('Count')
rng = np.arange(dataset4.display.min(), dataset4.display.max(), 0.1)
plt.plot(rng, norm.pdf(rng,dataset4.display.mean(),dataset4.display.std()))
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="mblDPbivh1zF" executionInfo={"status": "ok", "timestamp": 1627283301987, "user_tz": -345, "elapsed": 959, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="e922b07e-f36d-4f32-fbca-d537ca3806fd"
# in price
plt.hist(dataset4.price, bins=20, rwidth=0.8)
plt.xlabel('price (rs)')
plt.ylabel('Count')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 307} id="-4unmZzLh1wi" executionInfo={"status": "ok", "timestamp": 1627283305792, "user_tz": -345, "elapsed": 2066, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="59fc964b-9666-4ffc-b558-d90e01e59292"
plt.hist(dataset4.price, bins=20, rwidth=0.8, density=True)
plt.xlabel('price (rs)')
plt.ylabel('Count')
rng = np.arange(dataset4.price.min(), dataset4.price.max(), 0.1)
plt.plot(rng, norm.pdf(rng,dataset4.price.mean(),dataset4.price.std()))
# + colab={"base_uri": "https://localhost:8080/"} id="GbdwWLbboVcz" executionInfo={"status": "ok", "timestamp": 1627283743185, "user_tz": -345, "elapsed": 415, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}} outputId="8b3fb34a-909b-47ac-f6e7-a5cbea3f396a"
print("mean = {}".format(dataset4.price.mean()))
print("median = {}".format(dataset4.price.median()))
print("mode = {}".format(dataset4.price.mode()))
# Outlier detection and removal using Z Score
dataset4['zscore_price'] = ( dataset4.price - dataset4.price.mean() ) / dataset4.price.std()
dataset4.head(20)
print(dataset4[dataset4['zscore_price']>4])
print(dataset4[dataset4['zscore_price'] < -4])
dataset4_removed_outlier_price = dataset4[(dataset4.zscore_price>-4) & (dataset4.zscore_price<4)]
print(dataset4_removed_outlier_price.head())
# Remove two columns name is "zscore_price"
dataset4_removed_outlier_price = dataset4_removed_outlier_price.drop(['zscore_price'], axis = 1)
print(dataset4_removed_outlier_price.head())
print(dataset4_removed_outlier_price.info())
# + id="hRY9Imxopsye" executionInfo={"status": "ok", "timestamp": 1627283978767, "user_tz": -345, "elapsed": 498, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GjJcF7DevbQJhXjEo6haTw5Ueuuwu-b8cFLTjgo=s64", "userId": "17265195811399595189"}}
dataset4_removed_outlier_price.to_csv("/content/drive/MyDrive/Colab Notebooks/datamining/removed_outlier.csv")
# + id="GZahhHX8rvWp"
| data_mining/lpps/data_preprocessing/outlieranalysis/analysisandregression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # L06: Pruning Decision Trees in Scikit-Learn
# %load_ext watermark
# %watermark -d -u -a '<NAME>' -v -p numpy,scipy,matplotlib,sklearn
# +
from sklearn import datasets
from sklearn.model_selection import train_test_split
import numpy as np
iris = datasets.load_iris()
X = iris.data[:, [2, 3]]
y = iris.target
X_temp, X_test, y_temp, y_test = \
train_test_split(X, y, test_size=0.1,
shuffle=True, random_state=1, stratify=y)
X_train, X_valid, y_train, y_valid = \
train_test_split(X_temp, y_temp, test_size=0.3,
shuffle=True, random_state=1, stratify=y_temp)
print('Train size', X_train.shape, 'class proportions', np.bincount(y_train))
print('Valid size', X_valid.shape, 'class proportions', np.bincount(y_valid))
print('Test size', X_test.shape, 'class proportions', np.bincount(y_test))
# -
# # Baseline tree
# +
from sklearn.tree import DecisionTreeClassifier
tree = DecisionTreeClassifier(random_state=1)
tree.fit(X_train, y_train)
train_acc = tree.score(X_train, y_train)
valid_acc = tree.score(X_valid, y_valid)
test_acc = tree.score(X_test, y_test)
print(f'Training accuracy: {train_acc*100:.2f}%')
print(f'Validation accuracy: {valid_acc*100:.2f}%')
print(f'Depth: {tree.get_depth()}')
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from sklearn.tree import plot_tree
plt.figure(figsize=(5, 7))
plot_tree(tree,
filled=True,
rounded=True,
feature_names=['Sepal length', 'Sepal width',
'Petal length', 'Petal width'],
class_names=['Setosa',
'Versicolor',
'Virginica']
)
#plt.tight_layout()
plt.show()
# -
# # Minimum Samples Per Leaf (Pre-pruning)
# +
all_train_acc = []
all_valid_acc = []
all_hyperparam = []
for i in range(1, 15):
tree = DecisionTreeClassifier(min_samples_leaf=i, random_state=1)
tree.fit(X_train, y_train)
train_acc = tree.score(X_train, y_train)
valid_acc = tree.score(X_valid, y_valid)
all_train_acc.append(train_acc*100)
all_valid_acc.append(valid_acc*100)
all_hyperparam.append(i)
plt.plot(all_hyperparam, all_train_acc, label='Training accuracy')
plt.plot(all_hyperparam, all_valid_acc, ls='--', label='Validation accuracy')
plt.xlabel('min_samples_leaf')
plt.ylabel('Accuracy in %')
plt.legend()
plt.show()
# +
tree = DecisionTreeClassifier(min_samples_leaf=2, random_state=1)
tree.fit(X_train, y_train)
train_acc = tree.score(X_train, y_train)
valid_acc = tree.score(X_valid, y_valid)
test_acc = tree.score(X_test, y_test)
print(f'Training accuracy: {train_acc*100:.2f}%')
print(f'Validation accuracy: {valid_acc*100:.2f}%')
print(f'Depth: {tree.get_depth()}')
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from sklearn.tree import plot_tree
plt.figure(figsize=(5, 7))
plot_tree(tree,
filled=True,
rounded=True,
feature_names=['Sepal length', 'Sepal width',
'Petal length', 'Petal width'],
class_names=['Setosa',
'Versicolor',
'Virginica']
)
#plt.tight_layout()
plt.show()
# -
# # Max Depth (Pre-pruning)
# +
all_train_acc = []
all_valid_acc = []
all_hyperparam = []
for i in range(1, 15):
tree = DecisionTreeClassifier(max_depth=i, random_state=1)
tree.fit(X_train, y_train)
train_acc = tree.score(X_train, y_train)
valid_acc = tree.score(X_valid, y_valid)
all_train_acc.append(train_acc*100)
all_valid_acc.append(valid_acc*100)
all_hyperparam.append(i)
plt.plot(all_hyperparam, all_train_acc, label='Training accuracy')
plt.plot(all_hyperparam, all_valid_acc, ls='--', label='Validation accuracy')
plt.xlabel('max_depth')
plt.ylabel('Accuracy in %')
plt.legend()
plt.show()
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from sklearn.tree import plot_tree
tree = DecisionTreeClassifier(max_depth=2, random_state=1)
tree.fit(X_train, y_train)
train_acc = tree.score(X_train, y_train)
valid_acc = tree.score(X_valid, y_valid)
test_acc = tree.score(X_test, y_test)
print(f'Training accuracy: {train_acc*100:.2f}%')
print(f'Validation accuracy: {valid_acc*100:.2f}%')
print(f'Depth: {tree.get_depth()}')
plt.figure(figsize=(5, 7))
plot_tree(tree,
filled=True,
rounded=True,
feature_names=['Sepal length', 'Sepal width',
'Petal length', 'Petal width'],
class_names=['Setosa',
'Versicolor',
'Virginica']
)
#plt.tight_layout()
plt.show()
# -
# # Cost Complexity Pruning
# +
all_train_acc = []
all_valid_acc = []
all_hyperparam = []
for i in np.arange(0.0, 0.5, 0.005):
tree = DecisionTreeClassifier(ccp_alpha=i, random_state=1)
tree.fit(X_train, y_train)
train_acc = tree.score(X_train, y_train)
valid_acc = tree.score(X_valid, y_valid)
all_train_acc.append(train_acc*100)
all_valid_acc.append(valid_acc*100)
all_hyperparam.append(i)
plt.plot(all_hyperparam, all_train_acc, label='Training accuracy')
plt.plot(all_hyperparam, all_valid_acc, ls='--', label='Validation accuracy')
plt.xlabel('ccp_alpha')
plt.ylabel('Accuracy in %')
plt.legend()
plt.show()
# -
all_hyperparam[np.argmax(all_valid_acc)]
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from sklearn.tree import plot_tree
tree = DecisionTreeClassifier(ccp_alpha=0.015, random_state=1)
tree.fit(X_train, y_train)
train_acc = tree.score(X_train, y_train)
valid_acc = tree.score(X_valid, y_valid)
test_acc = tree.score(X_test, y_test)
print(f'Training accuracy: {train_acc*100:.2f}%')
print(f'Validation accuracy: {valid_acc*100:.2f}%')
print(f'Depth: {tree.get_depth()}')
plt.figure(figsize=(5, 7))
plot_tree(tree,
filled=True,
rounded=True,
feature_names=['Sepal length', 'Sepal width',
'Petal length', 'Petal width'],
class_names=['Setosa',
'Versicolor',
'Virginica']
)
#plt.tight_layout()
plt.show()
# -
# More info about minimal cost complexity pruning at
# - https://scikit-learn.org/stable/auto_examples/tree/plot_cost_complexity_pruning.html
# - https://scikit-learn.org/stable/modules/tree.html#minimal-cost-complexity-pruning
| 06-decision-trees/code/06-trees_pruning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="BZPSH4VkK7J2"
# 欢迎来到HanLP在线交互环境,这是一个Jupyter记事本,可以输入任意Python代码并在线执行。请点击左上角【Run】来运行这篇NLP教程。
#
#
#
#
# + [markdown] id="XxPAiNwSK7J4"
# ## 安装
# 量体裁衣,HanLP提供**RESTful**(云端)和**native**(本地)两种API,分别面向轻量级和海量级两种场景。无论何种API何种语言,HanLP接口在语义上保持一致,你可以**任选一种**API来运行本教程。
#
# ### 轻量级RESTful API
#
# 仅数KB,适合敏捷开发、移动APP等场景。简单易用,无需GPU配环境,**强烈推荐**,秒速安装:
#
# + colab={"base_uri": "https://localhost:8080/"} id="lgMa4kbfK7J5" outputId="5bb662d8-1665-4bcc-c517-70d1c4bc4837"
# !pip install hanlp_restful
# + [markdown] id="N4G6GbNmK7J6"
# 创建客户端,填入服务器地址:
# + id="3XM9-3-oK7J6"
from hanlp_restful import HanLPClient
HanLP = HanLPClient('https://www.hanlp.com/api', auth=None, language='zh') # auth不填则匿名,zh中文,mul多语种
# + [markdown] id="pbeFH9jmK7J7"
# 调用`parse`接口,传入一篇文章,得到HanLP精准的分析结果。
# + colab={"base_uri": "https://localhost:8080/"} id="mNJPvZ_3K7J7" outputId="4048d0d6-2dad-4582-e327-f99338f8f72b"
doc = HanLP.parse("2021年HanLPv2.1为生产环境带来次世代最先进的多语种NLP技术。阿婆主来到北京立方庭参观自然语义科技公司。")
print(doc)
# + [markdown] id="w4E8Kn_nK7J8"
# #### 可视化
# 输出结果是一个可以`json`化的`dict`,键为[NLP任务名](https://hanlp.hankcs.com/docs/data_format.html#naming-convention),值为分析结果。关于标注集含义,请参考[《语言学标注规范》](https://hanlp.hankcs.com/docs/annotations/index.html)及[《格式规范》](https://hanlp.hankcs.com/docs/data_format.html)。我们购买、标注或采用了世界上量级最大、种类最多的语料库用于联合多语种多任务学习,所以HanLP的标注集也是覆盖面最广的。通过`doc.pretty_print`,可以在等宽字体环境中得到可视化,你需要取消换行才能对齐可视化结果。我们已经发布HTML环境的可视化,在Jupyter Notebook中自动对齐中文。
# + colab={"base_uri": "https://localhost:8080/", "height": 575} id="GZ79la4LK7J8" outputId="b9bd5dc0-52f9-4b42-93fd-7c4e49214ace"
doc.pretty_print()
# + [markdown] id="WIKyCLQJK7J9"
# #### 申请秘钥
# 由于服务器算力有限,匿名用户每分钟限2次调用。如果你需要更多调用次数,[建议申请免费公益API秘钥auth](https://bbs.hanlp.com/t/hanlp2-1-restful-api/53)。
# + [markdown] id="PcZAZopQK7J9"
# ### 海量级native API
#
# 依赖PyTorch、TensorFlow等深度学习技术,适合**专业**NLP工程师、研究者以及本地海量数据场景。要求Python 3.6以上,支持Windows,推荐*nix。可以在CPU上运行,推荐GPU/TPU。
#
# 无论是Windows、Linux还是macOS,HanLP的安装只需一句话搞定。
# + colab={"base_uri": "https://localhost:8080/"} id="bjRdHxl1K7J-" outputId="659d7920-c857-4eb8-f45f-dba84366688a"
# !pip install hanlp -U
# + [markdown] id="dHhIRwgqK7J-"
# #### 加载模型
# HanLP的工作流程是先加载模型,模型的标示符存储在`hanlp.pretrained`这个包中,按照NLP任务归类。
# + colab={"base_uri": "https://localhost:8080/"} id="KHY6bsG_K7J-" outputId="208c12b6-2702-4ee7-a03a-f053b7ad3479"
import hanlp
hanlp.pretrained.mtl.ALL # MTL多任务,具体任务见模型名称,语种见名称最后一个字段或相应语料库
# + [markdown] id="WDT3Hks0K7J_"
# 调用`hanlp.load`进行加载,模型会自动下载到本地缓存。自然语言处理分为许多任务,分词只是最初级的一个。与其每个任务单独创建一个模型,不如利用HanLP的联合模型一次性完成多个任务:
# + id="4Cj8a73rK7J_" colab={"base_uri": "https://localhost:8080/"} outputId="a92ac736-6e61-4949-8d35-56c773faf950"
HanLP = hanlp.load(hanlp.pretrained.mtl.CLOSE_TOK_POS_NER_SRL_DEP_SDP_CON_ELECTRA_BASE_ZH)
# + [markdown] id="pBqH_My8K7J_"
# ## 多任务批量分析
# 客户端创建完毕,或者模型加载完毕后,就可以传入一个或多个句子进行分析了:
# + id="B58npfkHK7J_" colab={"base_uri": "https://localhost:8080/"} outputId="69fed02d-39cb-4b4c-d2c8-d0edc25970ea"
doc = HanLP(['2021年HanLPv2.1为生产环境带来次世代最先进的多语种NLP技术。', '阿婆主来到北京立方庭参观自然语义科技公司。'])
print(doc)
# + [markdown] id="tvuxfWPYK7J_"
# ## 可视化
# 输出结果是一个可以`json`化的`dict`,键为[NLP任务名](https://hanlp.hankcs.com/docs/data_format.html#naming-convention),值为分析结果。关于标注集含义,请参考[《语言学标注规范》](https://hanlp.hankcs.com/docs/annotations/index.html)及[《格式规范》](https://hanlp.hankcs.com/docs/data_format.html)。我们购买、标注或采用了世界上量级最大、种类最多的语料库用于联合多语种多任务学习,所以HanLP的标注集也是覆盖面最广的。通过`doc.pretty_print`,可以在等宽字体环境中得到可视化,你需要取消换行才能对齐可视化结果。我们已经发布HTML环境的可视化,在Jupyter Notebook中自动对齐中文。
# + id="M8WxTdlAK7KA" colab={"base_uri": "https://localhost:8080/", "height": 575} outputId="a027a302-74d8-48c9-b30d-45ebf8741c1e"
doc.pretty_print()
# + [markdown] id="_B2HDiZgK7KA"
# ## 指定任务
# 简洁的接口也支持灵活的参数,任务越少,速度越快。如指定仅执行分词:
# + id="9Mnys4t2K7KA" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="88d72a72-c095-4f6d-df0b-d881887087ce"
HanLP('阿婆主来到北京立方庭参观自然语义科技公司。', tasks='tok').pretty_print()
# + [markdown] id="s5RkVkVkK7KA"
# ### 执行粗颗粒度分词
# + id="5R_PwELlK7KA" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="5ce2c037-eb44-481f-9de2-dc0d4122e7c4"
HanLP('阿婆主来到北京立方庭参观自然语义科技公司。', tasks='tok/coarse').pretty_print()
# + [markdown] id="pTrajkHEK7KB"
# ### 执行分词和PKU词性标注
# + id="kkkgVKFqK7KB" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="e9f9879b-47ce-459a-e089-923de1c6436c"
HanLP('阿婆主来到北京立方庭参观自然语义科技公司。', tasks='pos/pku').pretty_print()
# + [markdown] id="YLLTVY0RK7KB"
# ### 执行粗颗粒度分词和PKU词性标注
# + id="5qSlqbcfK7KB" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="66944459-bc22-4bd9-e4af-4d2aba9316f3"
HanLP('阿婆主来到北京立方庭参观自然语义科技公司。', tasks=['tok/coarse', 'pos/pku'], skip_tasks='tok/fine').pretty_print()
# + [markdown] id="3nNojvHiK7KB"
# ### 执行分词和MSRA标准NER
# + id="tTVoEPiAK7KB" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="b8dc8c24-3392-4712-d1b6-e2dc8b7710e8"
HanLP('阿婆主来到北京立方庭参观自然语义科技公司。', tasks='ner/msra').pretty_print()
# + [markdown] id="uG2wYTfmK7KB"
# ### 执行分词、词性标注和依存句法分析
# + id="WXl6f7zyK7KC" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="8671e0e4-d0c3-40f4-a4db-ba9aaec225ab"
doc = HanLP('阿婆主来到北京立方庭参观自然语义科技公司。', tasks=['pos', 'dep'])
doc.pretty_print()
# + [markdown] id="ocxM3LsGK7KC"
# 转换为CoNLL格式:
# + id="NtKmSB_0K7KC" colab={"base_uri": "https://localhost:8080/"} outputId="cc9245b3-32c2-4d35-88a8-a7d91127eca7"
print(doc.to_conll())
# + [markdown] id="PNBo-kETK7KC"
# ### 执行分词、词性标注和短语成分分析
# + id="Ja8dib6XK7KC" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="a972f5bb-ae23-47a9-cd9f-6070a5b39f50"
doc = HanLP('阿婆主来到北京立方庭参观自然语义科技公司。', tasks=['pos', 'con'])
doc.pretty_print()
# + [markdown] id="Mg3DhvjhK7KC"
# #### 将短语结构树以bracketed形式打印
# + id="kE8iBZNUK7KC" colab={"base_uri": "https://localhost:8080/"} outputId="79e2a72d-e473-41ca-c054-9595a4dd5971"
print(doc['con']) # str(doc['con'])会将短语结构列表转换为括号形式
# + [markdown] id="MfleaY_pK7KC"
# 关于标注集含义,请参考[《语言学标注规范》](https://hanlp.hankcs.com/docs/annotations/index.html)及[《格式规范》](https://hanlp.hankcs.com/docs/data_format.html)。我们购买、标注或采用了世界上量级最大、种类最多的语料库用于联合多语种多任务学习,所以HanLP的标注集也是覆盖面最广的。
#
# ## 多语种支持
# 总之,可以通过tasks参数灵活调用各种NLP任务。除了中文联合模型之外,你可以在文档中通过找到许多其他语种的模型,比如日语:
# + id="oJP8dvfvK7KD" colab={"base_uri": "https://localhost:8080/"} outputId="2262ccdb-7cf5-4859-8d6c-18300e54c22e"
ja = hanlp.load(hanlp.pretrained.mtl.NPCMJ_UD_KYOTO_TOK_POS_CON_BERT_BASE_CHAR_JA)
# + id="3WPvCbH2K7KD" colab={"base_uri": "https://localhost:8080/", "height": 991} outputId="46a9435d-ed5b-47ef-99c6-71d7ee0fc6e8"
ja(['2021年、HanLPv2.1は次世代の最先端多言語NLP技術を本番環境に導入します。',
'奈須きのこは1973年11月28日に千葉県円空山で生まれ、ゲーム制作会社「ノーツ」の設立者だ。',]).pretty_print()
# + [markdown] id="NifrOGlNK7KD"
# 以及支持104种语言的多语种联合模型:
# + id="ae-4j5sbK7KD" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="2777cc5d-c1c5-4091-b754-0c220dafea8a"
from hanlp.utils.torch_util import gpus_available
if gpus_available():
mul = hanlp.load(hanlp.pretrained.mtl.UD_ONTONOTES_TOK_POS_LEM_FEA_NER_SRL_DEP_SDP_CON_XLMR_BASE)
mul(['In 2021, HanLPv2.1 delivers state-of-the-art multilingual NLP techniques to production environments.',
'2021年、HanLPv2.1は次世代の最先端多言語NLP技術を本番環境に導入します。',
'2021年 HanLPv2.1为生产环境带来次世代最先进的多语种NLP技术。']).pretty_print()
else:
print(f'建议在GPU环境中运行XLMR_BASE。')
# + [markdown] id="0QV_93CjK7KD"
# 你可以在下面输入你想执行的代码~
| plugins/hanlp_demo/hanlp_demo/zh/tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Notebook to process output from thecontentmine
#
# This notebook illustrates the basic process of processing each resource in each paper and allocates a score.
# +
import pandas
import json
import process_urls
# -
with open('amw_token', 'r') as f:
github_token = f.read()
#Read JSON data into the datastore variable
with open('../sample_data/small_dict_of_papers.json', 'r') as f:
dict_of_papers = json.load(f)
resources_list = process_urls.process_papers_dict(dict_of_papers, verbose=True,
github_token=github_token)
# ## Analysis of the data
#
# We can store the data in a flat SQL database, but here we stuff it into a Pandas dataframe to allow some analysis.
url_df = pandas.DataFrame.from_dict(resources_list)
url_df
| notebooks/resolvre_and_check_resources.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.1
# language: julia
# name: julia-1.6
# ---
# # Performance optimization exercise 1
# Optimize the following code.
#
# (The type and size of the input is fixed/may not be changed.)
# +
function work!(A, N)
D = zeros(N,N)
for i in 1:N
D = b[i]*c*A
b[i] = sum(D)
end
end
N = 100
A = rand(N,N)
b = rand(N)
c = 1.23
work!(A,N)
# -
using BenchmarkTools
@btime work!($A, $N);
# ## Optimizations
# +
# ...
| Day2/3a_exercise_optimization1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (ds4b)
# language: python
# name: pycharm-d7086c1b
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/SoIllEconomist/ds4b/blob/master/python_ds4b/06_sql/02_select_statements.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="mTu1Gb5u99xE" colab_type="code" colab={}
# #!pip install ipython-sql
# #!git clone https://github.com/thomasnield/oreilly_getting_started_with_sql.git
# + id="oU0Rj0Rh99xM" colab_type="code" colab={}
# %load_ext sql
# + id="3sf8ytI-99xP" colab_type="code" colab={} outputId="b9eaf7bd-eb11-4699-d2df-8783a26fb313"
# %sql sqlite:///oreilly_getting_started_with_sql/rexon_metals.db
# + [markdown] id="RirZWFLy99xT" colab_type="text"
# # SELECT
#
# When working with databases and SQL, the most common task is to request data from one or more tables and display it.
#
# The `SELECT` statement accomplishes this.
# + [markdown] id="El7L03FV99xU" colab_type="text"
# ## Retrieving Data with SQL
#
# Let’s write our first SQL statement. The most common SQL operation is a `SELECT` statement, which pulls data from a table and then displays the results.
#
# Write the following statement:
# + id="gpyJcCUs99xV" colab_type="code" colab={} outputId="1a2d3046-2128-4b3b-93d8-5b50bce2e078"
# %sql SELECT * FROM CUSTOMER;
# + [markdown] id="O-PNboBg99xZ" colab_type="text"
# Let’s break down exactly what happened. A SELECT statement allows you to choose which columns to pull from a table. So the first part of the SQL shown here should be read as “Select all columns,” where * is a placeholder to specify all columns.
#
# And you are getting these columns from the CUSTOMER table.
#
# You do not have to pull all columns in a SELECT statement. You can also pick and choose only the columns you are interested in. The following query will only pull the CUSTOMER_ID and NAME columns:
# + id="-y-UJgnx99xa" colab_type="code" colab={} outputId="ddb84ce6-3108-4d23-8286-37deb95e9d3f"
# %sql SELECT CUSTOMER_ID, NAME FROM CUSTOMER;
# + [markdown] id="DU6jJb-F99xd" colab_type="text"
# ## Expression in SELECT Statements
#
# The SELECT statement can do far more than simply select columns. You can also do calculations on one or more columns and include them in your query result. Let’s work with another table called PRODUCT.
# + id="SsYpd6F599xe" colab_type="code" colab={} outputId="78bc78cd-516a-41c8-c966-1e2f60a0d5fc"
# %sql SELECT * FROM PRODUCT;
# + [markdown] id="e2v_b6go99xh" colab_type="text"
# Suppose we wanted to generate a calculated column called TAXED_PRICE that is 7% higher than PRICE. We could use a SELECT query to dynamically calculate this for us
# + id="0tKxU23s99xi" colab_type="code" colab={} outputId="a2444ef6-ce7f-44d8-f6ea-fb033a450391" language="sql"
#
# SELECT
# PRODUCT_ID,
# DESCRIPTION,
# PRICE,
# PRICE * 1.07 AS TAXED_PRICE
# FROM PRODUCT;
# + [markdown] id="E6WcvX_G99xk" colab_type="text"
# Notice how the TAXED_PRICE column was dynamically calculated in the SELECT query. This column is not stored in the table, but rather calculated and displayed to us every time we run this query.
#
# Let’s take a look at our TAXED_PRICE column and break down how it was created. We first see the PRICE is multiplied by 1.07 to calculate the taxed amount. We generate this TAXED_PRICE value for every record.
#
# Notice too that we gave this calculated value a name using an AS statement (this is known as an alias):
#
# We can use aliases to give names to expressions. We can also use aliases to apply a new name to an existing column within the query. For example, we can alias the PRICE column to UNTAXED_PRICE. This does not actually change the name of the column in the table, but it gives it a new name within the scope of our SELECT statement:
#
# + id="ncT5NFHO99xm" colab_type="code" colab={} outputId="04aba9ed-59e0-49d4-84d8-9b1a32a1fcc8" language="sql"
# SELECT
# PRODUCT_ID,
# DESCRIPTION,
# PRICE AS UNTAXED_PRICE,
# PRICE * 1.07 AS TAXED_PRICE
# FROM PRODUCT
# + [markdown] id="z5-mhgG099xp" colab_type="text"
# To round the TAXED_PRICE to two decimal places, we can pass the multiplication expression PRICE * 1.07 as the first argument, and a 2 as the second:
# + id="zAq8jyyO99xr" colab_type="code" colab={} outputId="066ac272-44f4-4917-cc62-2f551be51121" language="sql"
# SELECT
# PRODUCT_ID,
# DESCRIPTION,
# PRICE AS UNTAXED_PRICE,
# round(PRICE * 1.07,2) AS TAXED_PRICE
# FROM PRODUCT
# + [markdown] id="snHqKp6i99xt" colab_type="text"
# Here are the mathematical operators you can use in SQL:
#
# | Operator | Description | Example |
# |---------- |----------------------------------------------- |---------------------- |
# | + | Adds two numbers | STOCK + NEW_SHIPMENT |
# | - | Subtracts two numbers | STOCK - DEFECTS |
# | * | Multiplies two numbers | PRICE * 1.07 |
# | / | Divides two numbers | STOCK / PALLET_SIZE |
# | % | Divides two, numbers but return the remainder | STOCK % PALLET_SIZE |
# + [markdown] id="LlUbjVa599xu" colab_type="text"
# ## Text Concatenation
#
# The concatenate operator is specified by a double pipe (||), and you put the data values to concatenate on both sides of it.
# + id="xjJ0q5j699xv" colab_type="code" colab={} outputId="95dfd599-0aa2-4f90-8853-4e1761ee4348" language="sql"
# SELECT NAME,
# CITY || ', ' || STATE AS LOCATION
# FROM CUSTOMER;
# + id="D1_00T8Q99xy" colab_type="code" colab={} outputId="9c302bfc-479f-45d4-f06b-a8d90db54b5a" language="sql"
# SELECT NAME,
# STREET_ADDRESS || ' ' || CITY || ', ' || STATE || ' ' || ZIP AS SHIP_ADDRESS
# FROM CUSTOMER;
# + [markdown] id="vffzCyLk99x0" colab_type="text"
# ## Summary
# In this section, we covered how to use the SELECT statement, the most common SQL operation. It retrieves and transforms data from a table without affecting the table itself. We also learned how to select columns and write expressions.
#
# Within expressions, we can use operators and functions to do tasks such as rounding, math, and concatenation.
| python_ds4b/06_sql/02_select_statements.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# <div class="contentcontainer med left" style="margin-left: -50px;">
# <dl class="dl-horizontal">
# <dt>Title</dt> <dd> ErrorBars Element</dd>
# <dt>Dependencies</dt> <dd>Matplotlib</dd>
# <dt>Backends</dt>
# <dd><a href='./ErrorBars.ipynb'>Matplotlib</a></dd>
# <dd><a href='../bokeh/ErrorBars.ipynb'>Bokeh</a></dd>
# <dd><a href='../plotly/ErrorBars.ipynb'>Plotly</a></dd>
# </dl>
# </div>
import numpy as np
import holoviews as hv
hv.extension('matplotlib')
# ``ErrorBars`` provide a visual indicator for the variability of the plotted data on a graph. They are usually applied on top of other plots such as scatter, curve or bar plots to indicate the variability in each sample.
#
# ``ErrorBars`` may be used to represent symmetric error or asymmetric error. An ``ErrorBars`` Element must have one key dimensions representing the samples along the x-axis and two or three value dimensions representing the value of the sample and positive and negative error values associated with that sample or x-axis. See the [Tabular Datasets](../../../user_guide/08-Tabular_Datasets.ipynb) user guide for supported data formats, which include arrays, pandas dataframes and dictionaries of arrays.
# #### Symmetric error
#
# By default the ``ErrorBars`` Element accepts x- and y-coordinates along with a symmetric error value along y-axis:
np.random.seed(7)
errors = [(0.1*i, np.sin(0.1*i), np.random.rand()/2) for i in np.linspace(0, 100, 11)]
hv.Curve(errors) * hv.ErrorBars(errors)
# #### Assymetric error
# ``ErrorBars`` is a set of x-/y-coordinates with associated error values along y-axis. Error values may be either symmetric or asymmetric, and thus can be supplied as an Nx3 or Nx4 array (or any of the alternative constructors Chart Elements allow).
errors = [(0.1*i, np.sin(0.1*i), np.random.rand()/2, np.random.rand()/4) for i in np.linspace(0, 100, 11)]
hv.Curve(errors) * hv.ErrorBars(errors, vdims=['y', 'yerrneg', 'yerrpos'])
# #### Errors along x-axis
# ``ErrorBars`` can be a set of a set of x-/y-coordinates with associated error values along x-axis. The parameter `horizontal`, when set to `True`, will set supplied errors along x-axis instead of y-axis.
errors = [(0.1*i, np.sin(0.1*i), np.random.rand()/2) for i in np.linspace(0, 100, 11)]
hv.Curve(errors) * hv.ErrorBars(errors, horizontal=True)
# #### Errors along x and y axes
# Two `ErrorBars` with orthogonal errors can be composed together to give x and y errorbars
# +
yerrors = [(0.1*i, np.sin(0.1*i), np.random.rand()/2, np.random.rand()/4) for i in np.linspace(0, 100, 11)]
xerrors = [(0.1*i, np.sin(0.1*i), np.random.rand()/2, np.random.rand()/4) for i in np.linspace(0, 100, 11)]
(hv.Curve(yerrors)
* hv.ErrorBars(yerrors, vdims=['y', 'yerrneg', 'yerrpos'])
* hv.ErrorBars(xerrors, vdims=['y', 'xerrneg', 'xerrpos'], horizontal=True)
)
# -
# For full documentation and the available style and plot options, use ``hv.help(hv.ErrorBars).``
| examples/reference/elements/matplotlib/ErrorBars.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="WdDeRs8nwsse" colab_type="text"
# <b><h1>Data Types and Operators</h1></b>
# <h4>Welcome to this lesson on Data Types and Operators! You'll learn about:
# </h4>
#
# * **Data Types:** Integers, Floats, Booleans,
# Strings, Lists, Tuples, Sets, Dictionaries
#
# * **Operators:** Arithmetic, Assignment, Comparison, Logical, Membership, Identity
#
# * Built-In Functions, Compound Data Structures, Type Conversion
#
# * Whitespace and Style Guidelines
# + [markdown] id="PR-M6YwHxkoW" colab_type="text"
# <br><br>
#
# <h2><b>1. Print() function in python:</b></h2>
#
# You will be seeing this print function very frequently in python programming. It helps us to see what exactly is happening in our code.
#
# Try to run the next lines of code and see what happens next:
# + id="x_R9bFHvvu1f" colab_type="code" colab={}
2+7
2*7
# + [markdown] id="_nKLdWFIyZt-" colab_type="text"
# * You will see that even though you had written 2 lines of code only the last line of code gets seen in the output area.
#
# Now this happens because you haven't told python what to alctually do with it.
#
# This is where *print()* comes in. *print()* in python is a useful builtin function that we can use to display input value as text in the output.
# + id="p2xPUnXMyYA7" colab_type="code" colab={}
print(2+7)
print(2*7)
# + id="ZoMyiceiJTAp" colab_type="code" colab={}
print('Hello World!!!!')
# + [markdown] id="hHx4ul-t1hJu" colab_type="text"
# <br><br>
#
# ## **2. Arithmetic Operators**
#
# An arithmetic operator is a mathematical function that takes 2 operands and performs a calculation on them. Lets take the previous example, we multiplied 2 and 7 here 2 and 7 are the operands and this asterisks is the arithmetic operator for multiplication. Python has several arithmetic operators most of which follow the usual rules of mathematics. Lets look at them now.
#
# * <b>+</b> Addition
# * <b>-</b> Subtraction
# * <b>*</b> Multiplication
# * <b>/</b> Division
# * <b>%</b> Mod (the remainder after dividing)
# * <b>**</b> Exponentiation (note that ^ does not do this operation, as you might have seen in other languages)
# * <b>//</b> Divides and rounds down to the nearest integer
#
# [ Now sometime people might confuse this (^)Caret symbol for Exponentiation as you might have seen it in other languages but in python this symbol is used to do bitwise **XOR operation**. As learning bitwise operation is not necessary for this course we will not be covering it but you can surely check out the link in the resources below to [learn more.](https://wiki.python.org/moin/BitwiseOperators) ]
#
# ###**Task:**
# **Now perform all the arithematic operations on the numbers 7 and 2.**
# + id="2NAGm8p4yjmu" colab_type="code" colab={}
# While showing the output use print()
# add 7 and 2 below
# substract 2 from 7 below
# multiply 7 and 2 below
# divide 7 by 2 below
# remainder after we divide 7 by 2 below
# 7 raise to the power 2 below
# divide 7 by 2 nd round it to the nearest integer
# + [markdown] id="SNltdCZw0oB5" colab_type="text"
# ##### Expected output:<br>
# 9<br>
# 5<br>
# 14<br>
# 3.5<br>
# 1<br>
# 49<br>
# 3
# + [markdown] id="U7a4p_mvxRk4" colab_type="text"
# <br><br>
#
# ## **3. Order of Operations:**
#
# Now Arithmetic operators in python follow the **BEDMAS** order of operations. <br>
# Look at this mathematical equation (2+3*5) what do you think will be the answer 25 or 17. The answer is 17 because in maths we follow a convention called as the order of operations BEDMAS ie.
# * Brackets
# * Exponents
# * Division and Multiplication (left to right)
# * Addition and Substraction (left to right)
#
# Preference given to the operator that comes first in BEDMAS.
#
# + [markdown] id="I28162IMzAj7" colab_type="text"
# **<br><h3>Question:</h3>
# In this quiz you're going to do some calculations for your
# bills. Your net expenditure for the past 5 months has been
# 1200, 2000, 800, 1500 and 1700. What is your average monthly
# expenditure.**
# + id="UJV4fx80ysCU" colab_type="code" colab={}
# write your code here
# + [markdown] id="-jsUpF3MzgeZ" colab_type="text"
# ##### Expected output:<br>1440.0
# + [markdown] id="_AYZ8efT1uq3" colab_type="text"
# <br><h3>Question:</h3>
# In this quiz you're going to do some calculations for a mason. Two rooms in a house needs to be filled. One room is 4 ft wide by 11 ft long, the other is 5 ft wide by 9 ft long. Tiles come in packages of 7 and each tile is 1ftx1ft in size<br><br>
# 1) How many tiles are needed?<br>
# 2) You buy 17 packages each containing 6 tiles each. How many tiles will be left over?
# + id="QqCobpMb0Teg" colab_type="code" colab={}
# write the code that calculates how many tiles are needed down below.
# write the code that calculates how many tiles will be left over.
# + [markdown] id="EgDQ7vYB6sTn" colab_type="text"
# ##### Expected output:<br>89<br>13
# + [markdown] id="uVi1mZvCCuSV" colab_type="text"
# <br><br>
#
# ## **4. Variables:**<br>
# Understanding variables is very important in any programming language they are used all the time in python. Using variables in place of direct numbers have many advantages. Variables are used to store information to be referenced and manipulated in a computer program.
#
# Creating a new variable in python is very simple, lets create one together, here in this example below the variable name is *month*, the equal sign is the assignment operator and the value of the variable is 12.
# + id="kIeQxFP67CNX" colab_type="code" colab={}
month=12
print(month)
# + [markdown] id="hfgtXMqGDI2l" colab_type="text"
# Now its your turn create a variable named *rent* with its value being 1700
# + id="slBwG5njDHik" colab_type="code" colab={}
# create and print the varibale rent down below
# + [markdown] id="dCgFMQt8Dc6a" colab_type="text"
# ##### Expected output:<br>1700
# + [markdown] id="ESJS8yE9J0zB" colab_type="text"
# In any case, whatever term is on the left side, is now a name for whatever value is on the right side. Once a value has been assigned to a variable name, you can access the value from the variable name.
#
# For example if we run this code we will get 3 as the output here as in the first line we assigned 3 to *a* and in the second line we assigned *a* to *b* so when we print *b* we get 3 as the output.
# + id="0Kg1PjaUDaLl" colab_type="code" colab={}
a=3
b=a
print(b)
# + [markdown] id="icTPtDSAKjEr" colab_type="text"
# If we don't declare the variable and try to print the output then we will get the following error
# + id="P_KmqiPBJ5_Z" colab_type="code" colab={}
print(x)
# + [markdown] id="PUE_n529K-Gp" colab_type="text"
# <br><br>
#
# ## **5. Multiple Assignment Operator:**
#
# Suppose you are making a program where in you enter the dimensions of the tank and it will give the volume of the tank as the output. So you can write code as:
# + id="gZHy4uODKsQX" colab_type="code" colab={}
height = 3
length = 6
width = 2
volume = height * length * width
print(volume)
# + [markdown] id="PdSLF0z5N0Bq" colab_type="text"
# Now python has a very useful way to assign multiple variables together in a single line using multiple assignment like this:
# + id="cGr5tmszNwUe" colab_type="code" colab={}
# this will now assign 3 to height, 6 to length and 2 to width just as before.
height , length , width = 3 , 6 , 2
volume = height * length * width
print(volume)
# + [markdown] id="hZdrLa63Ql8Z" colab_type="text"
# ou can use this when you are assigning closely related variables such as height width length or coordinates such as x, y , z of an object.
#
# <br><h3>Question:</h3>
# Find the length of the vector with its start coordinates as the origin and the end coordinates as (4,5,6).
#
# > *length = squareroot(x^2 + y^2 + z^2)*
# + id="jQMAaZT1O0F3" colab_type="code" colab={}
# write your code here
# + [markdown] id="SuOghUR5SbVg" colab_type="text"
# ##### Expected output:<br>8.774964387392123
# + [markdown] id="WbLxkJozSF5g" colab_type="text"
# <br><br>
#
# ## **6. Variable Naming Conventions:**
#
# There are some rules we need to follow while giving a name for a Python variable.
#
# - **Rule-1**: You should start variable name with an alphabet or **underscore(_)** character.
# - **Rule-2:** A variable name can only contain **A-Z,a-z,0-9** and **underscore(_)**.
# - **Rule-3:** You cannot start the variable name with a **number**.
# - **Rule-4:** You cannot use special characters with the variable name such as such as **$,%,#,&,@.-,^** etc.
# - **Rule-5**: Variable names are **case sensitive**. For example str and Str are two different variables.
# - **Rule-6:** Do not use reserve keyword as a variable name for example keywords like **class, for, def, del, is, else,** **try, from,** etc. more examples are given below and as we go through the course we will come across many more. Creating names that are descriptive of the values often will help you avoid using any of these words.
# + id="KtIoAjAKQ0Jh" colab_type="code" colab={}
#Allowed variable names
x=2
y="Hello"
mypython="PythonGuides"
my_python="PythonGuides"
_my_python="PythonGuides"
_mypython="PythonGuides"
MYPYTHON="PythonGuides"
myPython="PythonGuides"
myPython7="PythonGuides"
# + id="HswApcdn-GTJ" colab_type="code" colab={}
#Variable name not Allowed
7mypython="PythonGuides"
-mypython="PythonGuides"
myPy@thon="PythonGuides"
my Python="PythonGuides"
for="PythonGuides"
#It shows invalid syntax.
#It will execute one by one and will show the error.
# + [markdown] id="zoaFd7Jw-gVJ" colab_type="text"
# Also there are some naming convention that needs to be followed like:
#
# * try to keep the name of the variables descriptive short but descriptive. **for example:** when taking inputs for the height of a tree of a box the appropriate variable name will be just *height* not *x* not *h* not *height_of_the_tree*.
#
# * Also the pythonic way to name variables is to use all lowercase letters and underscores to separate words.
# + id="R_5NxrWx-L2E" colab_type="code" colab={}
# pythonic way
my_height = 58
my_lat = 40
my_long = 105
# + id="bufefdio-5N5" colab_type="code" colab={}
# not pythonic way
my height = 58 # wont work
MYLONG = 40 # will work still avoid using it
MyLat = 105 # will work still avoid using it
# + [markdown] id="0vk0r_8m_dWl" colab_type="text"
# Though the last two of these would work in python, they are not pythonic ways to name variables. The way we name variables is called snake case, because we tend to connect the words with underscores.
# + [markdown] id="DAoihPxbFI_d" colab_type="text"
#
#
# What if we want to change or update the value of a variable for example take the example of rent = 1700, suppose the rent has hiked and the new rent is 2000 we can just assign the variable its new value as:
# + id="81F5psRj_kUP" colab_type="code" colab={}
rent = 1700
rent = 2000
print(rent)
# + [markdown] id="MavVAvfNGFOi" colab_type="text"
# This is called overwriting the variable , i.e, When a new value is assigned to a variable, the old one is forgotten.
#
# If we had then caused some damages to the property during our crazy house party and we have to pay for them then we can just apply these changes directly to this variable.
# + id="rkEh72gQFaK2" colab_type="code" colab={}
rent = 1700
rent = 2000
rent =rent + 700
print(rent)
# + [markdown] id="vqwJs-3bHU7S" colab_type="text"
# in the line 3 the variable *rent* is being assigned to itself plus 700 which results to 2700.
#
# Because such increment and assignment operations are very common python has a very special assignment operator for this.
# + id="gUkRhTKnH9mv" colab_type="code" colab={}
rent = 1700
rent = 2000
rent += 700
print(rent)
# + [markdown] id="6N1_qkohIEb2" colab_type="text"
# we can actually use this **+=** operator to tell python that we are incrementing the value on the left by the value on the right. **+=** is a example of assignment operator **-= *= /=** are some more examples of assignment operators. All of these operators just apply arithmetic operation to the variable on the left with the value on the right which makes your code more concise and easier to read and understand.
# + [markdown] id="lAV3yc89IiTE" colab_type="text"
# <br><h3>Question:</h3>
# You are managing your finances using python:
#
# * complete the tasks mentioned in the comments. and then print the amount left in your bank.
#
# > *Note that this code uses scientific notation to define large numbers. 1.2e6 is equal to 1.2 * 10 ** 6 which is equal to 1200000.*
# + id="uodq27HAINVs" colab_type="code" colab={}
# the amount of money in your bank account is 1.2e5
amt = 1.2e6
# money recieved every year in form of salary
salary = 9.3e5
# decrease the salary by 30% as tax paied to the government.
# Add the salary to the bank account amt variable
# increase the amt by 4% that you recieved from the bank as intrest
# substract 1.2e5 from amt as the rent paied
# print the new value of the amt variable
# + [markdown] id="kFgrnORZJIkZ" colab_type="text"
# ##### Expected output:<br>1805040.0
# + [markdown] id="G3q5YcesJ8MV" colab_type="text"
# <br><br>
#
# ## **7. Integers and Floats:**
# So far the numbers that we have dealt with were mostly whole numbers or integers, but as you may have notices that other types of numbers also do exist. For example dividing one integer by another gives us a number that isn't an integer, in python we represent such a number as a float, which is short for floating point number.
# + id="fqai63lgI8v8" colab_type="code" colab={}
print(3/2)
# + [markdown] id="25ijafCzMh4j" colab_type="text"
# Numbers with a decimal point, such as 3.14, are called floating-point numbers (or floats). Note that even though the value 42 is an integer, the value 42.0 would be a floating-point number. And if 2 integers are divided then also we get float as an answer.
#
# You can check the datatype of any value by using the builtin function of type, that returns the type of an object. Here as you can see they type of a number without a decimal and the type of a number with a decimal.
# + id="L6ZPHtSyKogM" colab_type="code" colab={}
a = 3
b = 2.5
print(type(a))
print(type(b))
# + [markdown] id="4zrtRdZtDnj6" colab_type="text"
# An operation involving an int and a float will always give float as its output. We can also covert one datatype to another by constructing new objects of those types with int and float.
#
# When we convert a float to an int the part after the decimal point is dropped and hence there is no rounding. eg 28.9 will be cut to 28.
#
# Similarly converting int to float just adds a decimal at the end of the number and a 0 after that. example 3 will become 3.0
# + id="_17RKbnXMvSZ" colab_type="code" colab={}
a = float(3)
b = int(28.9)
print(a)
print(b)
# + [markdown] id="qlcwxnN0EYgx" colab_type="text"
#
#
# > Another point that you need to keep in mind is float are an approximation to the number they represent. As float can represent very large range of numbers python must use approximation to represent these numbers. For example this floating point number 0.23 is in reality slightly more than 0.23. such that if we add up 0.23 to itself a few times and check its equality to the expected resultant it will be different. Although the difference is very small but it exists never the less and you should know about it.
#
#
# + id="ecx9e1p8DuVJ" colab_type="code" colab={}
print(0.23 + 0.23 + 0.23 + 0.23 + 0.23 + 0.23
+ 0.23 + 0.23 + 0.23 + 0.23 + 0.23 + 0.23
+ 0.23 + 0.23 + 0.23 + 0.23 + 0.23 + 0.23
+ 0.23 + 0.23 + 0.23 + 0.23 + 0.23 + 0.23
+ 0.23 + 0.23 + 0.23 + 0.23+ 0.23 + 0.23 == 6.9)
# + [markdown] id="tqTe9v60FvTh" colab_type="text"
# <br><br>
#
# ##**8. Division by Zero and Errors:**
#
# What happens if you divide by zero in Python? Try it out! Test run this code and see what happens.
# + id="GRd2s9UXF56A" colab_type="code" colab={}
print(4/0)
# + [markdown] id="T7A07CYAGAkB" colab_type="text"
# what you should have seen when you submitted the Divide by Zero code above, Traceback means
# > *"What was the programming doing when it broke"! *
#
# This part is usually less helpful than the very last line of your error. Though you can dig through the rest of the error, looking at just the final line *ZeroDivisionError*, and the message says we divided by zero. Python is enforcing the rules of arithmetic!
# <br><br>
# In general, there are two types of errors to look out for
#
# * **Exceptions**
# * **Syntax**
#
# An Exception is a problem that occurs when the code is running, but a *'Syntax Error'* is a problem detected when Python checks the code before it runs it. For more information, see the Python tutorial page on [Errors and Exceptions](https://docs.python.org/3/tutorial/errors.html).
# + [markdown] id="cs768gAfFM2S" colab_type="text"
# <br><br>
#
# ##**9. Boolean Datatype, Comparison and Logical Operators:**
#
# Bool is another datatype that is commonly used in Python. Bool is short for Boolean which can have a value of either True or False. Boolean algebra is the branch of algebra in which the values of the variables are the truth values true or false. Boolean algebra us the framework on which all electronic devices and built and exists fundamentally in every line of code inside a computer. In python we can easily assign boolean values like this:
#
# + id="JW5nXrxsEnin" colab_type="code" colab={}
python_awsome = True
doumentation_bad = False
# + [markdown] id="a-JN4xZlHmzD" colab_type="text"
# We can use comparison operators to compare 2 values and produce boolean results like:
# + id="mVRNbJecHjgH" colab_type="code" colab={}
a = 3 > 1
print(a)
# + [markdown] id="QK7KL0o3Jqo3" colab_type="text"
# Here 3 is greater than 1 so printing out the output gives us a boolean value to true. There are many comparison operators in python, as you can see here are all of them.
#
# As you will see the function of all these comparison operators are evident from their names itself these are **less than, greater than , less than or equal to, greater than or equal to, not equal to**.
#
# Working with boolean has its own set of operators called as logical operators. These operators very useful when working with boolean, and evaluates if both the sides are true, **OR** evaluates if atleast one side is true and **not** evaluates the inverse of the input boolean.
#
# Lets understand if via an example:
# + id="eN24eeGyHl6k" colab_type="code" colab={}
rent = 1200
is_affordable = rent > 1000 and rent < 2000
print(is_affordable)
# + [markdown] id="XgntpdwgLpzT" colab_type="text"
# Here we check if the *rent* of a house is affordable or not, here in the second line we evaluate both the sides ie *rent > 1000*, yes, so it is **true** while the second condition is *rent < 200*, that too is **true**. as both the condition on the left and right side of and is **true** hence the boolean value of **true** will be assigned to the *is_affordable variable*. In other words if the *rent* is greater than 1000 and less than 2000 then only it is affordable.
#
# And here you can see how not works:
# + id="8c9_zsNvKR0v" colab_type="code" colab={}
rent = 1200
is_affordable = not(rent > 1000 and rent < 2000) #"not" just inverts bool value
print(is_affordable)
# + [markdown] id="NDgIKOVeOPUW" colab_type="text"
# <br><h3>**Question:**</h3>
#
# Is **Mumbai** more **dense** than Delhi?
# > *Print ***True*** if it is and ***False*** if it is not.*
#
# + id="2Fq_HK_mPYkD" colab_type="code" colab={}
mum_population, mum_area = 20411000, 4355
del_population, del_area = 30291000, 1484
mum_francisco_pop_density = mum_population/mum_area
del_de_janeiro_pop_density = del_population/del_area
# Print True if Mumbai is denser than Delhi, and False otherwise
# + [markdown] id="xflyycOOQZUP" colab_type="text"
# ##### Expected output:<br>False
# + [markdown] id="oeq2QDdEMSK8" colab_type="text"
# <br><br>
#
# ## **10. Strings:**
#
# Python has another datatype in its toolkit called as Strings, as the name suggest this datatype deals with characters words and text. String is a immutable order of sequences of characters (eg, Letters, numbers, spaces and symbols. We will be explaining what does immutable order means later on.
#
# You can create a string by using quotes as seen here, you can use either single / double quotes they both work equally well but there are some cases where you might prefer one over the other which we will be discussing below.
# + id="Q84mRQ2_MFGW" colab_type="code" colab={}
# using Double Quotes
print("ShapeAI")
# using Single Quotes
print('ShapeAI')
# + [markdown] id="odFoazduN4rR" colab_type="text"
# In this example we printed the word *ShapeAI* using single and double quotes and got the same output *ShapeAI*.
#
# We can also assign a string to a variable just like float and int.
# + id="Krfhjf9NN0-D" colab_type="code" colab={}
motto = "Learn | Code | Compete | Intern"
print(motto)
# + [markdown] id="YHgDcRm6R0FI" colab_type="text"
# Strings in Python are shown as the variable type str.
# + id="YkyDHgq3RW6K" colab_type="code" colab={}
type(motto)
# + [markdown] id="SjqlkfgzR-gR" colab_type="text"
# > *String can contain any character number symbol space within the quotes.
# However if we want to have quotes inside the string we get an error.*
# + id="788JWDKUR4BD" colab_type="code" colab={}
dialogue = "shiva said, "you learn as you grow""
# + [markdown] id="WjG9dsH5SOFT" colab_type="text"
# Python provides 2 easy ways to handle such problem:
#
# 1. Place the string in single quotes rather than double quotes. This will solve your problem for having double quotes within the string. But sometimes you will want to have both double and single quotes in your string in that case this will prove to be a problem.
# + id="_lPyEIYmSIes" colab_type="code" colab={}
dialogue = 'shiva said, "you learn as you grow"'
print(dialogue)
# + [markdown] id="EMqd6v3xSMno" colab_type="text"
# 2. In that case we can use a backslash to skip quotes as you can see in this example. The backslash helps python to know that the the single quote should be interpreted as part of the string rather than the quote that ends the string.
# + id="otSfYmAoS2TM" colab_type="code" colab={}
dialogue = '"shiva you\'re bag is red"'
print(dialogue)
# + [markdown] id="RA3hh1VhTFUF" colab_type="text"
# There are a few operators that we use on floats and ints that can also be used on strings. For example we can use the **'+'** to combine / concatenate 2 strings together and we can use **'*'** to repeat the string let us look at an example for each.
# + id="bIBYEJ6uS7ko" colab_type="code" colab={}
print("hello" + "world")
print("hello" + " " + "world")
# + [markdown] id="iurj3HxXTkx2" colab_type="text"
# > *here in this example we can see that using the plus arithmetic operator we get *helloworld* written together but this word that is printed out has no meaning, we need to have a space between both the words to have a meaning. We can add another string containing just a space in between the words to do so.*
# + id="jYVDqes0TeCz" colab_type="code" colab={}
word = "hello"
print(word * 5)
# + [markdown] id="78vTvfYSTwap" colab_type="text"
# Now in the second example we can see that using the multiplication operator on a string we get repetition of the same word as many time as the number we multiplied the string by in the output.
# + [markdown] id="fIFnUmZcT3_I" colab_type="text"
# However unlike multiplication and addition operators the other arithmetic operators like division and subtraction cannot be used on strings any attempt to do so would result in an error that string is an unsupported datatype for the division/subtraction operator.
# + id="h2nM7OBxT3he" colab_type="code" colab={}
word_1 = "hello"
word_2 = "world"
print(word_1 / word_2)
# + [markdown] id="9gumg_-mUBfA" colab_type="text"
# A useful builtin function for string datatypes is *len()* which stands for length. As the name suggests (it returns the length of an object ie., it returns the no of characters in a string.
#
# It takes in values in a parenthesis and returns the length of the string. *len()* is a little different from *print()* as the value returned from length can be stored in a variable as seen in the example here. The *len()* function outputs a value 7 that is then stored in a variable called as *word_length* which is then printed out.
# + id="oQNIXXfNTtzl" colab_type="code" colab={}
word_length = len("ShapeAI")
print(word_length)
# + [markdown] id="EEfczWGFzu6Y" colab_type="text"
# ### **Question:**
#
# The line of code in the following code block will cause a *SyntaxError*, thanks to the misuse of quotation marks. First run it with Test Run to view the error message. Then resolve the problem so that the quote (from Mahatama Gandhi) is correctly assigned to the variable *gandhi_quote*.
# + id="MnmykPZy0F-w" colab_type="code" colab={}
# TODO: Fix this string!
gandhi_quote = 'If you don't ask, you don't get it'
# + [markdown] id="GDbOH8th0g7c" colab_type="text"
# ### **Question:**
#
# In this question you have to print the accuracy logs of a model in training.
# + id="tlAI_jTK0ywg" colab_type="code" colab={}
model = "VGG16"
iteration = "150"
accuracy = "67.98"
# TODO: print a log message using the variables above
# This message should have the same format as this one:
# "the accuracy of ResNET50 model in 100th iteration is: 42.16%"
# + [markdown] id="WiPTD1y81uXZ" colab_type="text"
# ### **Question:**
#
# Use string concatination and *len()* function to fing the length of a persons complete name and store it in the variable *name_length*.
#
# As a business card designer find if the name can fit into a business card.
# + id="9Hzh7_vB1tpb" colab_type="code" colab={}
given_name = "Rahul"
middle_names = "Shastri"
family_name = "Mishra"
name_length = #todo: calculate how long this name is
# Now we check to make sure that the name fits within the driving license character limit
# Nothing you need to do here
driving_license_character_limit = 28
print(name_length <= driving_license_character_limit)
# + [markdown] id="Juq_M-ENy6-_" colab_type="text"
# <br><br>
#
# ## **11. Type and Type Conversion Revision:**
#
# Till now we have covered 4 different datatypes int, float, bool and string. As you can recall from the previous classes python has a builtin function called as type that returns the type of an object.
# + id="wsr3y0KgyuWW" colab_type="code" colab={}
print(type(75))
print(type(75.0))
print(type("75"))
print(type(True))
# + [markdown] id="iGgkIXg5zgwN" colab_type="text"
# Look at the code example we can see that even though the first 3 values appear to be same they can be encoded into different datatypes each with their own set of functions operations and uses.
#
# This is to note that here we have called the function print on another function type to output the return value of the function type. In such a case always the function inside the parenthesis is run first ie. here it will be type.
#
# Different types have different properties with their own set of functions operations and uses and hence while choosing a variable you need to choose the correct set of datatype for it depending upon how you care going to use it this is very important.
#
# There might be sometimes when you don't have the control over the type of the data being provided to you like one that has been received from a user as in input. But the good news is that python allows you to create new objects from old and change the datatypes for these new objects. As we had previously seen in the integers and floats video.
# + [markdown] id="bBZGIBLe6kKT" colab_type="text"
# > For example here we created a float ie 3.0 from an int 3 and assigned it to a new variable called decimal
# + id="yUceqN1QzIJ0" colab_type="code" colab={}
decimal = float(3)
print(decimal)
print(type(decimal))
# + [markdown] id="OHQFOeZi6o6c" colab_type="text"
# > In this next example we created a string from the integer variable *marks* and used that to create a larger string.
# + id="xZwhCiLZ6jky" colab_type="code" colab={}
marks = 15
subject = "coding"
semester = "first"
result = "I scored " + str(marks) + " in " + subject + " during my " + semester + " semester."
print(result)
# + [markdown] id="IqgSdyf067yQ" colab_type="text"
# > we can also create an float from string
# + id="Kqsda09k6d4x" colab_type="code" colab={}
marks = "15"
print(type(marks))
marks = float(marks)
print(type(marks))
# + [markdown] id="pzFSq-6W7oZi" colab_type="text"
# ### **Question:**
#
# In this quiz, you’ll need to change the types of the input and output data in order to get the result you want.
#
# Calculate and print the total sales for the week from the data provided. Print out a string of the form "This week's total sales: xxx", where xxx will be the actual total of all the numbers. You’ll need to change the type of the input data in order to calculate that total.
# + id="hu24eSfU7Q3F" colab_type="code" colab={}
mon_sales = "121"
tues_sales = "105"
wed_sales = "110"
thurs_sales = "98"
fri_sales = "95"
#TODO: Print a string with this format: This week's total sales: xxx
# You will probably need to write some lines of code before the print statement.
total_sales = (float(mon_sales) + float(tues_sales) + float(wed_sales)
+ float(thurs_sales) + float(fri_sales))
print("This week\'s total sales: " + str(total_sales))
# + [markdown] id="_tvr0pgf81Ia" colab_type="text"
# <br><br>
#
# ## **12. String Methods:**
#
# Methods are like some of the functions you have already seen:
#
# * len("this")
# * type(12)
# * print("Hello world")
#
# These three above are functions - notice they use parentheses, and accept one or more arguments. Functions will be studied in much more detail in a later lesson!
#
# A method in Python behaves similarly to a function. Methods actually are functions that are called using dot notation. For example, *lower()* is a string method that can be used like this, on a string called **"sample string"**: *sample_string.lower()*.
#
# Methods are specific to the data type for a particular variable. So there are some built-in methods that are available for all strings, different methods that are available for all integers, etc.
#
# Below is an image that shows some methods that are possible with any string.
#
# )
print(my_string.count('a'))
print(my_string.find('a'))
# + [markdown] id="zCSjfToiVx7t" colab_type="text"
# You can see that the count and find methods both take another argument. However, the .islower() method does not accept another argument.
#
# No professional has all the methods memorized, which is why understanding how to use documentation and find answers is so important. Gaining a strong grasp of the foundations of programming will allow you to use those foundations to use documentation to build so much more than someone who tries to memorize all the built-in methods in Python.
# + [markdown] id="4M_Qt7mSWGuD" colab_type="text"
# ### **12.a. One important string method: format()**
#
# We will be using the *format()* string method a good bit in our future work in Python, and you will find it very valuable in your coding, especially with your print statements.
#
# We can best illustrate how to use *format()* by looking at some examples:
# + id="utXx78crUTO0" colab_type="code" colab={}
# Example 1
print("EG:1")
print("Mohammed has {} balloons".format(27))
# Example 2
print("EG:2")
animal = "dog"
action = "bite"
print("Does your {} {}?".format(animal, action))
# Example 3
print("EG:3")
maria_string = "Maria loves {} and {}"
print(maria_string.format("math","statistics"))
# + [markdown] id="G9ybjouxWyQe" colab_type="text"
# Notice how in each example, the number of pairs of curly braces {} you use inside the string is the same as the number of replacements you want to make using the values inside *format()*.
#
# More advanced students can learn more about the formal syntax for using the *format()* string method [here](https://docs.python.org/3.6/library/string.html#format-string-syntax).
# + [markdown] id="PqKTCtYOTMQ8" colab_type="text"
# <br><br>
#
# ## **13. Lists and Membership Operators:**
#
# **Data structures** are containers that organize and group data types together in different ways. A **list** is one of the most common and basic data structures in Python. It is a mutable ordered sequence of elements.
#
# The code below defines a variable *students* which contains a list of strings. Each element in the list is a string that signifies the name of a student.
#
# > The data inside a list can be a mixture of any number and combination of diffrent data types.
# + id="zRS5zyHbWqaP" colab_type="code" colab={}
students = ['sam', 'pam', 'rocky', 'austin', 'steve', 'banner']
# + [markdown] id="fl7PSRXXdhrO" colab_type="text"
# List are ordered, we can look up individual elements by their index, we can look elements from a list just like we have done below.
# + id="uERsCZXNeBbY" colab_type="code" colab={}
print(students[0])
print(students[1])
print(students[2])
# + [markdown] id="cGgwxP3_eWir" colab_type="text"
# Notice that the first element in the list is accessed by the index 0, many programming language follow this convection called as zero based indexing.
#
# We can also access the elements from the end of the list using negative index as seen in the examples below.
# + id="CiGVaf_beRAC" colab_type="code" colab={}
print(students[-1])
print(students[-2])
print(students[-3])
# + [markdown] id="KqJU4qOYgE6A" colab_type="text"
# If you try to access an index in a list that doesn't exist then you will get an Error as seen below.
# + id="C5DSaY0Wf2wT" colab_type="code" colab={}
print(students[20])
# + [markdown] id="nKJIgDrZh0Cn" colab_type="text"
# ### **Question:**
#
# Try to use *len()* to pull the last element from the above list
# + id="NOiBUWAwgV87" colab_type="code" colab={}
# TODO: write your code here
students[len(students)-1]
# + [markdown] id="WZU02jRcFFPN" colab_type="text"
# ## **13.a. Membership Operators:[Lists]**
#
# In addition to accessing individual elements froma a list, we can use pythons sliceing notation to access a subsequence of a list.
#
# Slicing means using indicies to slice off parts of an object like list/string. Look at an example
# + id="dLyKcmXNiLwU" colab_type="code" colab={}
students = ['sam', 'pam', 'rocky', 'austin', 'steve', 'banner', 'tony', 'bruce',
'henry', 'clark', 'diana']
student = "Barry"
# slice a particular range
marvel = students[4:7]
flash = student[1:3]
print(marvel)
print(flash)
# + id="hzDrsBVELAsr" colab_type="code" colab={}
# slice from the end
dc = students[7:]
flash = student[1:]
print(dc)
print(flash)
# + id="NYSNXq7wLtR9" colab_type="code" colab={}
# slice from the begining
normal = students[:4]
flash = student[:3]
print(normal)
print(flash)
# + id="uXeNb-9qNdkL" colab_type="code" colab={}
# length of the list and the string
print(len(students))
print(len(student))
# + [markdown] id="FPqHnXKXMLku" colab_type="text"
# Of the types we have seen lists are most familier to strings, both supports the *len()* function, indexing and slicing.
#
# > Here above you have seen that the length of a string is the no of characters in the string, while the length of a list is the no of elements in the list.
#
# Another thing that they both supports aare membership operators:
#
# * **in:** evaluates if an object on the left side is included in the object on the right side.
# * **not in:** evaluates if object on left side is not included in object on right side.
# + id="3WSj6PZYL3Eg" colab_type="code" colab={}
greeting = "Hello there"
print('her' in greeting, 'her' not in greeting)
# + id="sQIsQ-_PRZQm" colab_type="code" colab={}
print('ShapeAI' in students, 'ShapeAI' not in students)
# + [markdown] id="ykKZQtwXfWRd" colab_type="text"
# ### **13.b. Mutability and Order:**
#
# So how are Lists diffrent from Strings, both supporst slicing, indexing, in and not in operators.
#
# The most obvious diffrence between them is that string is a sequence of characters while list's elements can be any type of bojects string, integers, floats orr bools.
#
# A more important diference is that lists can be modified but string can't. Look at the example below to understand more.
# + id="uEYr9AS0Rlxe" colab_type="code" colab={}
students = ['sam', 'pam', 'rocky', 'austin', 'steve', 'banner', 'tony', 'bruce',
'henry', 'clark', 'diana']
students[2] = 'ben'
print(students)
# + id="FHuFUelijkOp" colab_type="code" colab={}
student = "Barry"
student[1] = "e"
print(student)
# + [markdown] id="xMiN9z9pkAWm" colab_type="text"
# **Mutability** is about whether or not we can change an object once it has been created. If an object (like a list or string) can be changed (like a list can), then it is called **mutable**. However, if an object cannot be changed without creating a completely new object (like strings), then the object is considered immutable.
# + [markdown] id="xm5EHFuykdpP" colab_type="text"
# **Order** is about whether the position of an element in the object can be used to access the element. **Both strings and lists are ordered.** We can use the order to access parts of a list and string.
#
# > However, you will see some data types in the next sections that will be unordered. For each of the upcoming data structures you see, it is useful to understand how you index, are they mutable, and are they ordered. Knowing this about the data structure is really useful!
#
# Additionally, you will see how these each have different methods, so why you would use one data structure vs. another is largely dependent on these properties, and what you can easily do with it!
# + [markdown] id="nQ22-QIIRXsG" colab_type="text"
# Previously when we created a variable that heald an immutable object like string, the value of the immutable object was saved in memory. Like as you can see below
# + id="-JjIXgKuPjlO" colab_type="code" colab={}
student = "pam"
character = student
print(character)
character = "peter"
print(character)
print(student)
# + [markdown] id="T4SpeeBCZAVB" colab_type="text"
# Lists are diffrent from strings as they are mutable as can be seen from the example below
# + id="fDu_EQW3j5yw" colab_type="code" colab={}
students = ['sam', 'pam', 'rocky', 'austin', 'steve', 'banner', 'tony', 'bruce',
'henry', 'clark', 'diana']
characters = students
print(characters)
characters[1]= "peter"
print(characters)
print(students)
# + [markdown] id="VC6ZnlQfZjsF" colab_type="text"
# There are some useful functions for lists that you should get familier with.
#
# 1. ***len():*** returns how many elements does the list has.
# 2. ***max():*** returns the greatest element of a list.
# 3. ***min():*** returns the smallest element of a list.
# 4. ***sorted():*** returns a copy of the list, in order from smallest to the largest. leaving the orignal list unchanged
#
# > max element in a list of integers is the largest integer, while in the case of string is the string that will come last if the list was sorted alphabetically.
# + id="bk55uR0VPzqr" colab_type="code" colab={}
students = ['sam', 'pam', 'rocky', 'austin', 'steve', 'banner', 'tony', 'bruce',
'henry', 'clark', 'diana']
student = "barry"
print(max(students))
print(max(student))
# + [markdown] id="hL-h6v8Nev9j" colab_type="text"
# > a point to note is that even though you can have a list cntaining int and string a max function will be undefined upon such a list.
# + id="qofTAIxvccsm" colab_type="code" colab={}
max([2, 'two'])
# + id="DmQ7YXTse_Le" colab_type="code" colab={}
characters = sorted(students)
print(characters)
# + [markdown] id="SZiwJj5ua7qs" colab_type="text"
# ***Join()*** is an other useful function for lists(string lists), Join is a string method that takes a list of strings as an argument, and returns a string consisting of the list elements joined by a separator string. Look at the example below to understand.
#
#
# + id="tUxIAmUiZgOv" colab_type="code" colab={}
sep_str = "\n".join(["Jack", "O", "Lantern"])
print(sep_str)
# + [markdown] id="mKrXZopBesma" colab_type="text"
# In this example we use the string **"\n"** as the separator so that there is a newline between each element.
# We can also use other strings as separators with .join. Here we use a hyphen.
# + id="GPH5EfbYepKQ" colab_type="code" colab={}
name = "-".join(["Jack", "O", "Lantern"])
print(name)
# + [markdown] id="z4Qml70wfYt5" colab_type="text"
# > It is important to remember to separate each of the items in the list you are joining with a comma (,). Forgetting to do so will not trigger an error, but will also give you unexpected results.
# + [markdown] id="NaEnB22pfci9" colab_type="text"
# ***append()*** is an other useful method that adds an element to the end of a list.
# + id="XcY1vqKLe1FK" colab_type="code" colab={}
letters = ['a', 'b', 'c', 'd']
letters.append('e')
print(letters)
# + [markdown] id="HFY2Ty8_hUdo" colab_type="text"
# <br><br>
#
# ## **14. Tuples:**
#
# A tuple is another useful container. It's a data type for immutable ordered sequences of elements. They are often used to store related pieces of information. Consider this example involving (x, y, z) coordinates:
# + id="_Q0uioLufvgc" colab_type="code" colab={}
vector = (4, 5, 9)
print("x-coordinate:", vector[0])
print("y-coordinate:", vector[1])
print("z-coordinate:", vector[2])
# + [markdown] id="_JtjMmZ2rz1t" colab_type="text"
# Tuples are similar to lists in that they store an ordered collection of objects which can be accessed by their indices. Unlike lists, however, tuples are immutable - you can't add and remove items from tuples, or sort them in place.
#
# > Tuples can also be used to assign multiple variables in a compact way.
#
# > The parentheses are optional when defining tuples, and programmers frequently omit them if parentheses don't clarify the code.
# + id="-a_ISfUIrx65" colab_type="code" colab={}
location = 108.7774, 92.5556
latitude, longtitude = location
print("The coordinates are {} x {}".format(latitude, longtitude))
# + [markdown] id="W5LSu94ItjfD" colab_type="text"
# In the second line, two variables are assigned from the content of the tuple location. This is called tuple unpacking. You can use tuple unpacking to assign the information from a tuple into multiple variables without having to access them one by one and make multiple assignment statements.
#
# If we won't need to use location directly, we could shorten those two lines of code into a single line that assigns three variables in one go!
# + id="0d7u9CZ1tgo-" colab_type="code" colab={}
location = 108.7774, 92.5556
print("The coordinates are {} x {}".format(latitude, longtitude))
# + [markdown] id="_jOE97oTyFTQ" colab_type="text"
# ##**15. Sets:**
# A set is a data type for mutable unordered collections of unique elements. One application of a set is to quickly remove duplicates from a list.
#
# + id="StB2pYuPuaLz" colab_type="code" colab={}
numbers = [1, 2, 6, 3, 1, 1, 6]
unique_nums = set(numbers)
print(unique_nums)
# + [markdown] id="l7hrHkJ0yrxb" colab_type="text"
# Sets support the in operator the same as lists do. You can add elements to sets using the add method, and remove elements using the pop method, similar to lists. Although, when you pop an element from a set, a random element is removed. Remember that sets, unlike lists, are unordered so there is no "last element".
# + id="TQIH3VGXynbz" colab_type="code" colab={}
fruit = {"apple", "banana", "orange", "grapefruit"} # define a set
print("watermelon" in fruit) # check for element
fruit.add("watermelon") # add an element
print(fruit)
print(fruit.pop()) # remove a random element
print(fruit)
# + [markdown] id="DORS4RJ_0W0Z" colab_type="text"
# <br><br>
#
# ## **16. Dictionaries and Identity Operators:**
#
# A dictionary is a mutable data type that stores mappings of unique keys to values. Here's a dictionary that stores elements and their atomic numbers.
# + id="ApfpAVUvzqDe" colab_type="code" colab={}
elements = {"hydrogen": 1, "helium": 2, "carbon": 6}
# + [markdown] id="xTlhGP84yajj" colab_type="text"
# Dictionaries can have keys of any immutable type, like integers or tuples, not just strings. It's not even necessary for every key to have the same type! We can look up values or insert new values in the dictionary using square brackets that enclose the key.
# + id="pAtBqAtPyZzq" colab_type="code" colab={}
print(elements["helium"]) # print the value mapped to "helium"
elements["lithium"] = 3 # insert "lithium" with a value of 3 into the dictionary
# + [markdown] id="8ygwwby4y6Ia" colab_type="text"
# We can check whether a value is in a dictionary the same way we check whether a value is in a list or set with the in keyword. Dicts have a related method that's also useful, *get()*. *get()* looks up values in a dictionary, but unlike square brackets, get returns **None** (or a default value of your choice) if the key isn't found.
# + id="CyBTsiY6yrRA" colab_type="code" colab={}
print("carbon" in elements)
print(elements.get("dilithium"))
# + [markdown] id="d84ZmGfRzy7r" colab_type="text"
# Carbon is in the dictionary, so ***True*** is printed. Dilithium isn’t in our dictionary so ***None*** is returned by get and then printed. If you expect lookups to sometimes fail, get might be a better tool than normal square bracket lookups because errors can crash your program.
# + [markdown] id="vYKbWMN40Hbj" colab_type="text"
# ### **16.a. Keyword Operators:**
#
# * ***is:*** evaluates if both sides have the same identity
# * ***is not*** evaluates if both sides have different identities
#
# You can check if a key returned ***None*** with the ***is*** operator. You can check for the opposite using ***is not.***
# + id="-3Z4D-Przuu2" colab_type="code" colab={}
n = elements.get("dilithium")
print(n is None)
print(n is not None)
# + [markdown] id="WY58GiSVSQAU" colab_type="text"
# ### Task:
# Define a dictionary named population that contains this data:
#
# **Keys** -> **Values**
#
# New York -> 17.8
#
# Spain -> 13.3
#
# Dhaka -> 13.0
#
# Mumbai -> 12.5
# + id="PEGkVp3S0pzm" colab_type="code" colab={}
population = {"New York":17.8, "Spain":13.3, "Dhaka":13.0, "Mumbai":12.5}
print(population)
# + [markdown] id="esVtoMWGTESQ" colab_type="text"
# ## 16.b. ***Get()* with a Default Value:**
#
# Dictionaries have a related method that's also useful, *get()*. *get()* looks up values in a dictionary, but unlike looking up values with square brackets, *get()* returns **None** (or a default value of your choice) if the key isn't found. If you expect lookups to sometimes fail, *get()* might be a better tool than normal square bracket lookups.
# + id="lpgpXnvLS_BO" colab_type="code" colab={}
print(population.get('London'))
# + id="lwUfwma8TvA9" colab_type="code" colab={}
population['London']
# + id="l50bnIi5T1Wl" colab_type="code" colab={}
population.get('London', 'There\'s no such place!')
# + [markdown] id="6mBdBxyQULNO" colab_type="text"
# > *In the last example we specified a default value (the string 'There's no such element!') to be returned instead of None when the key is not found.*
#
#
# + [markdown] id="Yl7JfrMAn3kJ" colab_type="text"
# ## **16.c. Compound Data Structures:**
#
# Previously we have seen a dictonary called *elements* in which the element names are maped to their atomic numbers which are integers. But what if we want to store more information about each element like their atomic weight and symbol. We can do that by adjusting this dictionary so that it maps the element names to an other dictionary, that stores that collection of data.
#
# + id="UUfWjEUIUIKS" colab_type="code" colab={}
elements = {"hydrogen": {"number": 1,
"weight": 1.00794,
"symbol": "H"},
"helium": {"number": 2,
"weight": 4.002602,
"symbol": "He"}}
# + [markdown] id="pHEaNsQMplHp" colab_type="text"
#
# We can look up information about an element using this nested dictionary, using square brackets or the *get()* method.
# + id="N3Icsgotpv_P" colab_type="code" colab={}
helium = elements["helium"] # get the helium dictionary
hydrogen_weight = elements["hydrogen"]["weight"] # get hydrogen's weight
print(helium)
print(hydrogen_weight)
# + [markdown] id="L8id3asCp8Q5" colab_type="text"
# > You can also add a new key to the element dictionary.
# + id="krGwpkyqpyM7" colab_type="code" colab={}
oxygen = {"number":8,"weight":15.999,"symbol":"O"} # create a new oxygen dictionary
elements["oxygen"] = oxygen # assign 'oxygen' as a key to the elements dictionary
print('elements = ', elements)
# + [markdown] id="GBpaX13TrJ0s" colab_type="text"
# ### **Question:**
#
# Try your hand at working with nested dictionaries. Add another entry, *'is_noble_gas,'* to each dictionary in the *elements* dictionary. After inserting the new entries you should be able to perform these lookups:
#
#
#
# ---
#
# > print(elements['hydrogen']['is_noble_gas'])<br>
# False
#
# > print(elements['helium']['is_noble_gas'])<br>
# True
# + id="V8Ynb9OpqGCY" colab_type="code" colab={}
elements = {'hydrogen': {'number': 1, 'weight': 1.00794, 'symbol': 'H'},
'helium': {'number': 2, 'weight': 4.002602, 'symbol': 'He'}}
# todo: Add an 'is_noble_gas' entry to the hydrogen and helium dictionaries
# hint: helium is a noble gas, hydrogen isn't
elements['helium']['is_noble_gas'] = True
elements['hydrogen']['is_noble_gas'] = False
# + id="ZJmnZKwJr3Nw" colab_type="code" colab={}
elements
# + id="42016zIdsXQL" colab_type="code" colab={}
| Day_1_Data_Types_Operators.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [](https://colab.research.google.com/github/giswqs/GEE-Courses/blob/master/docs/gee_intro/ImageCollection/reducing_image_collection.ipynb)
# **Reducing an ImageCollection**
#
# To composite images in an `ImageCollection`, use `imageCollection.reduce()`. This will composite all the images in the collection to a single image representing, for example, the min, max, mean or standard deviation of the images. (See the Reducers section for more information about reducers). For example, to create a median value image from a collection:
# ## Create an interactive map
import ee
import geemap
Map = geemap.Map()
Map
# ## Compute a median image
# +
# Load a Landsat 8 collection for a single path-row.
collection = (
ee.ImageCollection('LANDSAT/LC08/C01/T1_TOA')
.filter(ee.Filter.eq('WRS_PATH', 44))
.filter(ee.Filter.eq('WRS_ROW', 34))
.filterDate('2014-01-01', '2015-01-01')
)
# Compute a median image and display.
median = collection.median()
Map.setCenter(-122.3578, 37.7726, 12)
Map.addLayer(median, {'bands': ['B4', 'B3', 'B2'], 'max': 0.3}, 'median')
# -
collection.size().getInfo()
collection.aggregate_array("system:id").getInfo()
# ## Use median reducer
# At each location in the output image, in each band, the pixel value is the median of all unmasked pixels in the input imagery (the images in the collection). In the previous example, `median()` is a convenience method for the following call:
# +
# Reduce the collection with a median reducer.
median = collection.reduce(ee.Reducer.median())
# Display the median image.
Map.addLayer(
median,
{'bands': ['B4_median', 'B3_median', 'B2_median'], 'max': 0.3},
'also median',
)
Map
# -
# ## Create an image composite
states = ee.FeatureCollection('TIGER/2018/States')
Map.addLayer(states, {}, "US States")
ca = states.filter(ee.Filter.eq("NAME", "California"))
Map.addLayer(ca, {}, "California")
collection = (
ee.ImageCollection('LANDSAT/LC08/C01/T1_TOA')
.filterBounds(ca)
.filterDate('2020-01-01', '2021-01-01')
)
collection.size().getInfo()
image = collection.median().clip(ca)
Map.addLayer(image, {'bands': ['B4', 'B3', 'B2'], 'max': 0.3}, 'Landsat 2020')
| docs/gee_intro/ImageCollection/reducing_image_collection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#%% Libraries
import pandas as pd
import warnings
import pandas as pds
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
from matplotlib import pyplot as plt
from sklearn.utils import resample
from pandas_profiling import ProfileReport
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import cross_val_score, KFold
from pandas.plotting import scatter_matrix
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import ExtraTreesRegressor
from sklearn import metrics
from sklearn.metrics import mean_squared_error
from sklearn.metrics import roc_auc_score, plot_roc_curve
from sklearn.metrics import precision_score, recall_score, f1_score
from sklearn.feature_selection import RFECV as RFECV_SKYLEARN
pd.options.display.max_columns = 100
#pd.set_option('display.max_columns', None)
import warnings
warnings.filterwarnings("ignore")
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
from IPython.display import Audio, display
def allDone():
display(Audio(url='https://sound.peal.io/ps/audios/000/000/537/original/woo_vu_luvub_dub_dub.wav', autoplay=True))
# # Model
#Importing the necessary packages and libaries
from sklearn.metrics import confusion_matrix
from sklearn.model_selection import train_test_split
from sklearn import svm, datasets
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
#Open File
albert = pd.read_csv('C:/Users/user/Documents/1. GitHub/Albert_Einstein/data.csv')
# Order categories
categories = pd.Categorical(albert['target'], categories=['low', 'med', 'high'], ordered=True)
categories
# Label your target with numerical values
labels, unique = pd.factorize(categories, sort=True)
albert['target'] = labels
X = albert[['x1', 'x2', 'x3']]
y = albert['target']
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.8, random_state = 0)
linear = svm.SVC(kernel='linear', C=1, decision_function_shape='ovo').fit(X_train, y_train)
rbf = svm.SVC(kernel='rbf', gamma=1, C=1, decision_function_shape='ovo').fit(X_train, y_train)
poly = svm.SVC(kernel='poly', degree=3, C=1, decision_function_shape='ovo').fit(X_train, y_train)
sig = svm.SVC(kernel='sigmoid', C=1, decision_function_shape='ovo').fit(X_train, y_train)
# retrieve the accuracy and print it for all 4 kernel functions
accuracy_lin = linear.score(X_test, y_test)
accuracy_poly = poly.score(X_test, y_test)
accuracy_rbf = rbf.score(X_test, y_test)
accuracy_sig = sig.score(X_test, y_test)
print('Accuracy Linear Kernel:', accuracy_lin)
print('Accuracy Polynomial Kernel:', accuracy_poly)
print('Accuracy Radial Basis Kernel:', accuracy_rbf)
print('Accuracy Sigmoid Kernel:', accuracy_sig)
# creating a confusion matrix
cm_lin = confusion_matrix(y_test, linear_pred)
cm_poly = confusion_matrix(y_test, poly_pred)
cm_rbf = confusion_matrix(y_test, rbf_pred)
cm_sig = confusion_matrix(y_test, sig_pred)
print(cm_lin)
print(cm_poly)
print(cm_rbf)
print(cm_sig)
#stepsize in the mesh, it alters the accuracy of the plotprint
#to better understand it, just play with the value, change it and print it
h = .01
#create the mesh
x_min, x_max = X[:, 0].min() - 1, X[:, 0].max() + 1
y_min, y_max = X[:, 1].min() - 1, X[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),np.arange(y_min, y_max, h))
# create the title that will be shown on the plot
titles = ['Linear kernel','RBF kernel','Polynomial kernel','Sigmoid kernel']
for i, clf in enumerate((linear, rbf, poly, sig)):
#defines how many plots: 2 rows, 2columns=> leading to 4 plots
plt.subplot(2, 2, i + 1) #i+1 is the index
#space between plots
plt.subplots_adjust(wspace=0.4, hspace=0.4)
Z = clf.predict(np.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=plt.cm.PuBuGn, alpha=0.7)
# Plot also the training points
plt.scatter(X[:, 0], X[:, 1], c=y, cmap=plt.cm.PuBuGn, edgecolors='grey')
plt.xlabel('x1')
plt.ylabel('target')
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.xticks(())
plt.yticks(())
plt.title(titles[i])
plt.show()
linear_pred = linear.predict(X_test)
poly_pred = poly.predict(X_test)
rbf_pred = rbf.predict(X_test)
sig_pred = sig.predict(X_test)
# +
# Hyperarameters
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import classification_report
from sklearn.svm import SVC
# Set the parameters by cross-validation
tuned_parameters = [{'kernel': ['rbf'], 'gamma': [1e-3, 1e-4],
'C': [1, 10, 100, 1000]},
{'kernel': ['linear'], 'C': [1, 10, 100, 1000]}]
scores = ['precision', 'recall']
for score in scores:
print("# Tuning hyper-parameters for %s" % score)
print()
clf = GridSearchCV(
SVC(), tuned_parameters, scoring='%s_macro' % score
)
clf.fit(X_train, y_train)
print("Best parameters set found on development set:")
print()
print(clf.best_params_)
print()
print("Grid scores on development set:")
print()
means = clf.cv_results_['mean_test_score']
stds = clf.cv_results_['std_test_score']
for mean, std, params in zip(means, stds, clf.cv_results_['params']):
print("%0.3f (+/-%0.03f) for %r"
% (mean, std * 2, params))
print()
print("Detailed classification report:")
print()
print("The model is trained on the full development set.")
print("The scores are computed on the full evaluation set.")
print()
y_true, y_pred = y_test, clf.predict(X_test)
print(classification_report(y_true, y_pred))
print()
allDone()
# +
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_squared_log_error
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.metrics import precision_score, recall_score, f1_score
# Set your variables, separating your target from the rest of the features
X = albert[['x1', 'x2', 'x3']]
y = albert['target']
#data_dmatrix = xgb.DMatrix(data=X,label=y)
#Split test and train
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
# Model
sig = svm.SVC(kernel='rbf', C=1, gamma=0.001, decision_function_shape='ovo').fit(X_train, y_train)
# Fit Model
sig.fit(X_train, y_train)
# See your prediction score
preds = sig.predict(X_test)
rmse = np.sqrt(mean_squared_error(y_test, preds))
rmsle = np.sqrt(mean_squared_log_error( y_test, preds))
F1_weight = f1_score(y_test, preds, average='weighted')
F1_macro = f1_score(y_test, preds, average='macro')
F1_micro = f1_score(y_test, preds, average='micro')
print("\n\n\n\nSEE YOUR RMSE SCORE: %f" % (rmse))
print("SEE YOUR RMSEL SCORE: %f" % (rmsle), "\n")
print("SEE YOUR F1_weight SCORE: %f" % (F1_weight))
print("SEE YOUR F1_macro SCORE: %f" % (F1_macro))
print("SEE YOUR F1_micro SCORE: %f" % (F1_micro),"\n" )
# +
#X, y = albert(n_samples=100, noise=0.1, random_state=42)
pipe = make_pipeline(StandardScaler(), SVM(kernel="rbf", C=1, gama=0.001, decision_function_shape='ovo'))
pipe.fit(X, y)
plot_svm(sig, X)
# +
import numpy as np
import matplotlib.pyplot as plt
from sklearn import svm
# figure number
fignum = 1
# plot the line, the points, and the nearest vectors to the plane
plt.figure(fignum, figsize=(4, 3))
plt.clf()
plt.scatter(clf.support_vectors_[:, 0], sig.support_vectors_[:, 1], s=80,
facecolors='none', zorder=10, edgecolors='k')
plt.scatter(X[:, 0], X[:, 1], c=Y, zorder=10, cmap=plt.cm.Paired,
edgecolors='k')
plt.axis('tight')
x_min = -3
x_max = 3
y_min = -3
y_max = 3
XX, YY = np.mgrid[x_min:x_max:200j, y_min:y_max:200j]
Z = sig.decision_function(np.c_[XX.ravel(), YY.ravel()])
# Put the result into a color plot
Z = Z.reshape(XX.shape)
plt.figure(fignum, figsize=(4, 3))
plt.pcolormesh(XX, YY, Z > 0, cmap=plt.cm.Paired)
plt.contour(XX, YY, Z, colors=['k', 'k', 'k'], linestyles=['--', '-', '--'],
levels=[-.5, 0, .5])
plt.xlim(x_min, x_max)
plt.ylim(y_min, y_max)
plt.xticks(())
plt.yticks(())
fignum = fignum + 1
plt.show()
# +
# Oh, wait...our F1-Micro which computes our multiclass accuracy is low...what about class weight trick instead of StandardScaler?
# +
from sklearn.metrics import mean_squared_error
from sklearn.metrics import mean_squared_log_error
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.metrics import precision_score, recall_score, f1_score
# Set your variables, separating your target from the rest of the features
X = albert[['x1', 'x2', 'x3']]
y = albert['target']
#data_dmatrix = xgb.DMatrix(data=X,label=y)
#Split test and train
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
# Model
sig = svm.SVC(kernel='rbf', C=1, gama=0.001, class_weight= 'balanced', decision_function_shape='ovo').fit(X_train, y_train)
# Fit Model
sig.fit(X_train, y_train)
# See your prediction score
preds = sig.predict(X_test)
rmse = np.sqrt(mean_squared_error(y_test, preds))
rmsle = np.sqrt(mean_squared_log_error( y_test, preds))
F1_weight = f1_score(y_test, preds, average='weighted')
F1_macro = f1_score(y_test, preds, average='macro')
F1_micro = f1_score(y_test, preds, average='micro')
print("\n\n\n\nSEE YOUR RMSE SCORE: %f" % (rmse))
print("SEE YOUR RMSEL SCORE: %f" % (rmsle), "\n")
print("SEE YOUR F1_weight SCORE: %f" % (F1_weight))
print("SEE YOUR F1_macro SCORE: %f" % (F1_macro))
print("SEE YOUR F1_micro SCORE: %f" % (F1_micro),"\n" )
# -
# +
import xgboost
from xgboost import XGBClassifier
import shap
import eli5
import numpy
import numpy.core.multiarray as multiarray
#Test new model
xgb = XGBClassifier(n_estimators=100, max_depth=25, importance_type='gain', colsample_bytree=0.3, gamma=0.5, min_child_weight=10, reg_lambda= 5, scale_pos_weight= 1)
xgb.fit(X_train, y_train)
explainer = shap.TreeExplainer(xgb)
explainer
# See your prediction score
preds = xgb.predict(X_test)
rmse = np.sqrt(mean_squared_error(y_test, preds))
rmsle = np.sqrt(mean_squared_log_error( y_test, preds))
F1_weight = f1_score(y_test, preds, average='weighted')
F1_macro = f1_score(y_test, preds, average='macro')
F1_micro = f1_score(y_test, preds, average='micro')
print("\n\n\n\nSEE YOUR RMSE SCORE: %f" % (rmse))
print("SEE YOUR RMSEL SCORE: %f" % (rmsle), "\n")
print("SEE YOUR F1_weight SCORE: %f" % (F1_weight))
print("SEE YOUR F1_macro SCORE: %f" % (F1_macro))
print("SEE YOUR F1_micro SCORE: %f" % (F1_micro),"\n" )
# +
# Not many improvements after one hour of tuning...
# -
# # Predict
# + code_folding=[]
# How many randoms?
#randoms = pd.DataFrame(clf.predict_proba(X_test)[:, 1], columns=['prob']).query('0.2 < prob < 0.6 ')
#randoms
#print(randoms.shape)
# +
# 52 from 10000 is irrelevant
# -
# +
# Get the predictions and put them on the test data.
X_output = X_test.copy()
X_output.loc[:,'predict'] = np.round(clf.predict(X_output),2)
# Randomly pick some observations
#random_picks = np.arange(1,330,50) # Every 50 rows
predict_table = X_output.iloc[:]
predict_table
# +
#%% Predict on your Unseen Data
data = albert.sample(frac=0.95, random_state=786)
data_unseen = albert.drop(data.index)
data.reset_index(inplace=True, drop=True)
data_unseen.reset_index(inplace=True, drop=True)
print('Data for Modeling: ' + str(data.shape))
print('Unseen Data For Predictions ' + str(data_unseen.shape))
data_unseen
# Backup your old column to compare
target_backup = [data_unseen["target"]]
target_backup = pd.DataFrame(target_backup).T
del data_unseen['target']
# Get the predictions and put them with the unseen data.
X_output = data_unseen.copy()
X_output.loc[:,'predict'] = np.round(clf.predict(X_output),2)
# Randomly pick some observations
#random_picks = np.arange(1,330,50) # Every 50 rows
predict_table = X_output.iloc[:]
# Fill your new table with all variables to compare
predict_table['target'] = target_backup
predict_table = predict_table[['x1', 'x2', 'x3', 'target', 'predict']]
# Boolean function for new column
def Check(predict_table):
if predict_table['target']== predict_table['predict']:
return "True"
else:
return "False"
# Result
predict_table['result'] = predict_table.apply(Check, axis=1)
predict_table
predict_table['result'].value_counts()
# +
# It's good to compare but with parcimony: since it's a hierarchy from low/medium/high, True or False doesn't always has the same weight;
# -
# # Visualize
| Backup/Hospital-Albert-Einstein-SVM-Backup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 2: Opening files
# ### Instructions
# Use the open() function to create a File object. The name of the file is "crime_rates.csv" and we want the file to be accessed in read mode ("r"). Assign this File object to the variable f.
# ### Answer
# ! touch test.txt
a = open("test.txt", "r")
print(a)
f = open("crime_rates.csv", "r")
# ## 3: Reading in files
# ### Instructions
# Run the **read()** method on the File object **f** to return the *string* representation of **crime_rates.csv**. Assign the resulting *string* to the variable **data**.
f = open("crime_rates.csv", "r")
data = f.read()
print(data)
# ## 4: Splitting
# ### Instructions
# Split the string object data on the new-line character "\n" and store the result in a variable named rows. Then use the print() function to display the first 5 elements in rows.
# ### Answer
# +
# We can split a string into a list.
sample = "john,plastic,joe"
split_list = sample.split(",")
print(split_list)
# Here's another example.
string_two = "How much wood\ncan a woodchuck chuck\nif a woodchuck\ncan chuck wood?"
split_string_two = string_two.split('\n')
print(split_string_two)
# Code from previous cells
f = open('crime_rates.csv', 'r')
data = f.read()
rows = data.split('\n')
print(rows[0:5])
# -
# ## 5: Loops
# ### Instructions
# ...
# ### Answer
# ## 6: Practice, loops
# ### Instructions
# The variable ten_rows contains the first 10 elements in rows. Write a for loop that iterates over each element in ten_rows and uses the print() function to display each element.
# ### Answer
ten_rows = rows[0:10]
for row in ten_rows:
print(row)
# ## 7: List of lists
# ### Instructions
# For now, explore and run the code we dissected in this step in the code cell below
# ### Answer
three_rows = ["Albuquerque,749", "Anaheim,371", "Anchorage,828"]
final_list = []
for row in three_rows:
split_list = row.split(',')
final_list.append(split_list)
print(final_list)
for elem in final_list:
print(elem)
print(final_list[0])
print(final_list[1])
print(final_list[2])
# ## 8: Practice, splitting elements in a list
# Let's now convert the full dataset, rows, into a list of lists using the same logic from the step before.
# ### Instructions
# Write a for loop that splits each element in rows on the comma delimiter and appends the resulting list to a new list named final_data. Then, display the first 5 elements in final_data using list slicing and the print() function.
# ### Answer
f = open('crime_rates.csv', 'r')
data = f.read()
rows = data.split('\n')
final_data = [row.split(",")
for row in rows]
print(final_data[0:5])
# ## 9: Accessing elements in a list of lists, the manual way
# ### Instructions
# five_elements contains the first 5 elements from final_data. Create a list of strings named cities_list that contains the city names from each list in five_elements.
# ### Answer
five_elements = final_data[:5]
print(five_elements)
cities_list = [city for city,_ in five_elements]
# ## 10: Looping through a list of lists
# ### Instructions
# Create a list of strings named cities_list that contains just the city names from final_data. Recall that the city name is located at index 0 for each list in final_data.
# ### Answer
# +
crime_rates = []
for row in five_elements:
# row is a list variable, not a string.
crime_rate = row[1]
# crime_rate is a string, the city name.
crime_rates.append(crime_rate)
cities_list = [row[0] for row in final_data]
# -
# ## 11: Practice
# ### Instructions
# Create a list of integers named int_crime_rates that contains just the crime rates as integers from the list rows.
#
# First create an empty list and assign it to int_crime_rates. Then, write a for loop that iterates over rows that executes the following:
#
# * uses the split() method to convert each string in rows into a list on the comma delimiter
# * converts the value at index 1 from that list to an integer using the int() function
# * then uses the append() method to add each integer to int_crime_rates
f = open('crime_rates.csv', 'r')
data = f.read()
rows = data.split('\n')
print(rows[0:5])
# +
int_crime_rates = []
for row in rows:
data = row.split(",")
if len(data) < 2:
continue
int_crime_rates.append(int(row.split(",")[1]))
print(int_crime_rates)
# -
| python_introduction/beginner/.ipynb_checkpoints/files and loops-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/baker371/k8-data-visualization/blob/lwasampijja-baker/upwork-devs/lwasampijja-baker/All_Progress_Issues.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="qODUMhkwFqUk"
# #### Table Showing All Open Issues
# + id="Bb8_Q65qoHEs" cellView="form" outputId="de510e4c-64c8-4242-b972-84d694dcd267" colab={"base_uri": "https://localhost:8080/", "height": 614}
#@title
from google.colab import drive
drive.mount("/content/drive", force_remount=True)
import requests
import json
import os
import numpy as np
import pandas as pd
from pandas import json_normalize
import seaborn as sns
import matplotlib.pyplot as plt
import datetime as dt
import datetime
# %reload_ext google.colab.data_table
# !cp "/content/drive/My Drive/models/issues.py" .
from issues import *
repos = Repos.get_repos()
issues = Issues(repos)
dfs = []
for repo in repos:
jtoken = os.getenv('dtollaku', '')
headers = {'Authorization': f'token {jtoken}'}
url = f'https://api.github.com/repositories/{repo[1]}/issues'
res = requests.get(url, headers=headers, params={'state': 'open'}).json()
data = json_normalize(res, max_level=1)
temp_df = pd.DataFrame(data)
temp_df['repo'] = repo[0]
dfs.append(temp_df)
df = pd.concat(dfs, ignore_index=True)
df['created_at'] = pd.to_datetime(df['created_at']).dt.date
df['Aging Days'] = datetime.datetime.now().date() - df['created_at']
df = df[['repo','title','assignee.login', 'created_at', 'Aging Days']]
df = df.rename({'repo':'Project', 'title':'Issue Title', 'assignee.login':'Assignees', 'created_at':'Date Created'}, axis=1)
df
# + id="alYzjuwQkSlS" cellView="form" outputId="2e215fcf-4780-42d1-d8a1-86ac88cc463a" colab={"base_uri": "https://localhost:8080/", "height": 837}
#@title
plt.figure(figsize=(15,10))
df["Project"].value_counts().plot.bar(title="Bar Chart Showing Number of Open Issues By Project")
plt.ylabel('Number of Issues')
plt.xlabel('Project')
plt.show()
# + [markdown] id="7KqN7oWZemE2"
# #### A Table Showing issues assigned to a person across multiple repos and Export to Excel
# + id="g9BNqCQie9h4" cellView="form"
#@title
project_filter = [" "] #@param {type:"raw"}
assignee_filter = ["baker371"] #@param {type:"raw"}
# + id="qkqdPRUHfC1c" cellView="form" outputId="ee99af16-24b4-4906-819a-18c739d4ec82" colab={"base_uri": "https://localhost:8080/", "height": 188}
#@title
filter1 = df["Project"].isin(project_filter)
filter2 = df["Assignees"].isin(assignee_filter)
df[filter1 | filter2]
# + [markdown] id="OQ67tI4ItXKh"
# #### Export to Excel
# + id="1kLJKgsbtJwr" cellView="form"
#@title
df.to_csv('report.csv')
# + [markdown] id="uQ-71Vk7hqDr"
# #### Open Issues per Assignee
# + id="_2FaUm_3hMp9" cellView="form" outputId="95d52462-0d14-40da-a815-88f33cc408e8" colab={"base_uri": "https://localhost:8080/", "height": 624}
#@title
df['Assignees'].value_counts().head(30).plot(kind='barh', colormap='Paired', figsize=(20,10), title="Bar Graph Showing Number of Open Issues Per Assignee")
# + [markdown] id="anvIvtAGO4rv"
# #### Visualising Zenhub Pipelines Per Repos
#
#
# + id="-xR7sXV8OUxJ" cellView="form" outputId="b9dcdacc-0d24-4c7d-dca5-366c31ac2485" colab={"base_uri": "https://localhost:8080/", "height": 598}
#@title
issues.show_tabular_report_by_repo()
# + [markdown] id="F2_xxmpgPaaO"
# #### Visualising Zenhub Pipelines Per User
# + id="-MiNkxPrOqkQ" cellView="form" outputId="e9e34c04-179e-4c8c-aef2-36024f8c8328" colab={"base_uri": "https://localhost:8080/", "height": 598}
#@title
issues.show_tabular_report_by_user()
| upwork-devs/lwasampijja-baker/All_Progress_Issues.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <center> <font size=5> <h1>Define working environment</h1> </font> </center>
# The following cells are used to:
# - Import needed libraries
# - Set the environment variables for Python, Anaconda, GRASS GIS and R statistical computing
# - Define the ["GRASSDATA" folder](https://grass.osgeo.org/grass73/manuals/helptext.html), the name of "location" and "mapset" where you will to work.
# **Import libraries**
## Import libraries needed for setting parameters of operating system
import os
import sys
# <center> <font size=3> <h3>Environment variables when working on Linux Mint</h3> </font> </center>
# **Set 'Python' and 'GRASS GIS' environment variables**
# Here, we set [the environment variables allowing to use of GRASS GIS](https://grass.osgeo.org/grass64/manuals/variables.html) inside this Jupyter notebook. Please change the directory path according to your own system configuration.
# +
### Define GRASS GIS environment variables for LINUX UBUNTU Mint 18.1 (Serena)
# Check is environmental variables exists and create them (empty) if not exists.
if not 'PYTHONPATH' in os.environ:
os.environ['PYTHONPATH']=''
if not 'LD_LIBRARY_PATH' in os.environ:
os.environ['LD_LIBRARY_PATH']=''
# Set environmental variables
os.environ['GISBASE'] = '/home/tais/SRC/GRASS/grass_trunk/dist.x86_64-pc-linux-gnu'
os.environ['PATH'] += os.pathsep + os.path.join(os.environ['GISBASE'],'bin')
os.environ['PATH'] += os.pathsep + os.path.join(os.environ['GISBASE'],'script')
os.environ['PATH'] += os.pathsep + os.path.join(os.environ['GISBASE'],'lib')
#os.environ['PATH'] += os.pathsep + os.path.join(os.environ['GISBASE'],'etc','python')
os.environ['PYTHONPATH'] += os.pathsep + os.path.join(os.environ['GISBASE'],'etc','python')
os.environ['PYTHONPATH'] += os.pathsep + os.path.join(os.environ['GISBASE'],'etc','python','grass')
os.environ['PYTHONPATH'] += os.pathsep + os.path.join(os.environ['GISBASE'],'etc','python','grass','script')
os.environ['PYTHONLIB'] = '/usr/lib/python2.7'
os.environ['LD_LIBRARY_PATH'] += os.pathsep + os.path.join(os.environ['GISBASE'],'lib')
os.environ['GIS_LOCK'] = '$$'
os.environ['GISRC'] = os.path.join(os.environ['HOME'],'.grass7','rc')
os.environ['PATH'] += os.pathsep + os.path.join(os.environ['HOME'],'.grass7','addons')
os.environ['PATH'] += os.pathsep + os.path.join(os.environ['HOME'],'.grass7','addons','bin')
os.environ['PATH'] += os.pathsep + os.path.join(os.environ['HOME'],'.grass7','addons')
os.environ['PATH'] += os.pathsep + os.path.join(os.environ['HOME'],'.grass7','addons','scripts')
## Define GRASS-Python environment
sys.path.append(os.path.join(os.environ['GISBASE'],'etc','python'))
# -
# **Import GRASS Python packages**
# +
## Import libraries needed to launch GRASS GIS in the jupyter notebook
import grass.script.setup as gsetup
## Import libraries needed to call GRASS using Python
import grass.script as gscript
from grass.script import core as grass
# -
# **-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
# **Display current environment variables of your computer**
## Display the current defined environment variables
for key in os.environ.keys():
print "%s = %s \t" % (key,os.environ[key])
# **-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
# <center> <font size=5> <h1>Define functions</h1> </font> </center>
# This section of the notebook is dedicated to defining functions which will then be called later in the script. If you want to create your own functions, define them here.
# ### Function for computing processing time
# The "print_processing_time" is used to calculate and display the processing time for various stages of the processing chain. At the beginning of each major step, the current time is stored in a new variable, using [time.time() function](https://docs.python.org/2/library/time.html). At the end of the stage in question, the "print_processing_time" function is called and takes as argument the name of this new variable containing the recorded time at the beginning of the stage, and an output message.
# +
## Import library for managing time in python
import time
## Function "print_processing_time()" compute processing time and printing it.
# The argument "begintime" wait for a variable containing the begintime (result of time.time()) of the process for which to compute processing time.
# The argument "printmessage" wait for a string format with information about the process.
def print_processing_time(begintime, printmessage):
endtime=time.time()
processtime=endtime-begintime
remainingtime=processtime
days=int((remainingtime)/86400)
remainingtime-=(days*86400)
hours=int((remainingtime)/3600)
remainingtime-=(hours*3600)
minutes=int((remainingtime)/60)
remainingtime-=(minutes*60)
seconds=round((remainingtime)%60,1)
if processtime<60:
finalprintmessage=str(printmessage)+str(seconds)+" seconds"
elif processtime<3600:
finalprintmessage=str(printmessage)+str(minutes)+" minutes and "+str(seconds)+" seconds"
elif processtime<86400:
finalprintmessage=str(printmessage)+str(hours)+" hours and "+str(minutes)+" minutes and "+str(seconds)+" seconds"
elif processtime>=86400:
finalprintmessage=str(printmessage)+str(days)+" days, "+str(hours)+" hours and "+str(minutes)+" minutes and "+str(seconds)+" seconds"
return finalprintmessage
# -
# ### Function for creation of configuration file for r.li (landscape units provided as polygons)
def create_rli_configfile(listoflandcoverraster,landscape_polygons,returnlistpath=False):
# Check if 'listoflandcoverraster' is not empty
if len(listoflandcoverraster)==0:
sys.exit("The list of landcover raster is empty and should contain at least one raster name")
# Get the version of GRASS GIS
version=grass.version()['version'].split('.')[0]
# Define the folder to save the r.li configuration files
if sys.platform=="win32":
rli_dir=os.path.join(os.environ['APPDATA'],"GRASS"+version,"r.li")
else:
rli_dir=os.path.join(os.environ['HOME'],".grass"+version,"r.li")
if not os.path.exists(rli_dir):
os.makedirs(rli_dir)
## Create an ordered list with the 'cat' value of landscape units to be processed.
list_cat=[int(x) for x in gscript.parse_command('v.db.select', quiet=True, map=landscape_polygons, column='cat', flags='c')]
list_cat.sort()
# Declare a empty dictionnary which will contains the north, south, east, west values for each landscape unit
landscapeunit_bbox={}
# Declare a empty list which will contain the path of the configation files created
listpath=[]
# Declare a empty string variable which will contains the core part of the r.li configuration file
maskedoverlayarea=""
# Duplicate 'listoflandcoverraster' in a new variable called 'tmp_list'
tmp_list=list(listoflandcoverraster)
# Set the current landcover raster as the first of the list
base_landcover_raster=tmp_list.pop(0) #The pop function return the first item of the list and delete it from the list at the same time
# Loop trough the landscape units
for cat in list_cat:
# Extract the current landscape unit polygon as temporary vector
tmp_vect="tmp_"+base_landcover_raster.split("@")[0]+"_"+landscape_polygons.split("@")[0]+"_"+str(cat)
gscript.run_command('v.extract', overwrite=True, quiet=True, input=landscape_polygons, cats=cat, output=tmp_vect)
# Set region to match the extent of the current landscape polygon, with resolution and alignement matching the landcover raster
gscript.run_command('g.region', vector=tmp_vect, align=base_landcover_raster)
# Rasterize the landscape unit polygon
landscapeunit_rast=tmp_vect[4:]
gscript.run_command('v.to.rast', overwrite=True, quiet=True, input=tmp_vect, output=landscapeunit_rast, use='cat', memory='3000')
# Remove temporary vector
gscript.run_command('g.remove', quiet=True, flags="f", type='vector', name=tmp_vect)
# Set the region to match the raster landscape unit extent and save the region info in a dictionary
region_info=gscript.parse_command('g.region', raster=landscapeunit_rast, flags='g')
n=str(round(float(region_info['n']),5)) #the config file need 5 decimal for north and south
s=str(round(float(region_info['s']),5))
e=str(round(float(region_info['e']),6)) #the config file need 6 decimal for east and west
w=str(round(float(region_info['w']),6))
# Save the coordinates of the bbox in the dictionary (n,s,e,w)
landscapeunit_bbox[cat]=n+"|"+s+"|"+e+"|"+w
# Add the line to the maskedoverlayarea variable
maskedoverlayarea+="MASKEDOVERLAYAREA "+landscapeunit_rast+"|"+landscapeunit_bbox[cat]+"\n"
# Compile the content of the r.li configuration file
config_file_content="SAMPLINGFRAME 0|0|1|1\n"
config_file_content+=maskedoverlayarea
config_file_content+="RASTERMAP "+base_landcover_raster+"\n"
config_file_content+="VECTORMAP "+landscape_polygons+"\n"
# Create a new file and save the content
configfilename=base_landcover_raster.split("@")[0]+"_"+landscape_polygons.split("@")[0]
path=os.path.join(rli_dir,configfilename)
listpath.append(path)
f=open(path, 'w')
f.write(config_file_content)
f.close()
# Continue creation of r.li configuration file and landscape unit raster the rest of the landcover raster provided
while len(tmp_list)>0:
# Reinitialize 'maskedoverlayarea' variable as an empty string
maskedoverlayarea=""
# Set the current landcover raster as the first of the list
current_landcover_raster=tmp_list.pop(0) #The pop function return the first item of the list and delete it from the list at the same time
# Loop trough the landscape units
for cat in list_cat:
# Define the name of the current "current_landscapeunit_rast" layer
current_landscapeunit_rast=current_landcover_raster.split("@")[0]+"_"+landscape_polygons.split("@")[0]+"_"+str(cat)
base_landscapeunit_rast=base_landcover_raster.split("@")[0]+"_"+landscape_polygons.split("@")[0]+"_"+str(cat)
# Copy the the landscape unit created for the first landcover map in order to match the name of the current landcover map
gscript.run_command('g.copy', overwrite=True, quiet=True, raster=(base_landscapeunit_rast,current_landscapeunit_rast))
# Add the line to the maskedoverlayarea variable
maskedoverlayarea+="MASKEDOVERLAYAREA "+current_landscapeunit_rast+"|"+landscapeunit_bbox[cat]+"\n"
# Compile the content of the r.li configuration file
config_file_content="SAMPLINGFRAME 0|0|1|1\n"
config_file_content+=maskedoverlayarea
config_file_content+="RASTERMAP "+current_landcover_raster+"\n"
config_file_content+="VECTORMAP "+landscape_polygons+"\n"
# Create a new file and save the content
configfilename=current_landcover_raster.split("@")[0]+"_"+landscape_polygons.split("@")[0]
path=os.path.join(rli_dir,configfilename)
listpath.append(path)
f=open(path, 'w')
f.write(config_file_content)
f.close()
# Return a list of path of configuration files creates if option actived
if returnlistpath:
return listpath
# ### Function for creation of binary raster from a categorical raster (multiprocessed)
# +
###### Function creating a binary raster for each category of a base raster.
### The function run within the current region. If a category do not exists in the current region, no binary map will be produce
# 'categorical_raster' wait for the name of the base raster to be used. It is the one from which one binary raster will be produced for each category value
# 'prefix' wait for a string corresponding to the prefix of the name of the binary raster which will be produced
# 'setnull' wait for a boolean value (True, False) according to the fact that the output binary should be 1/0 or 1/null
# 'returnlistraster' wait for a boolean value (True, False) regarding to the fact that a list containing the name of binary raster is desired as return of the function
# 'category_list' wait for a list of interger corresponding to specific category of the base raster to be used
# 'ncores' wait for a integer corresponding to the number of desired cores to be used for parallelization
# Import libraries for multiprocessing
import multiprocessing
from multiprocessing import Pool
from functools import partial
def create_binary_raster(categorical_raster,prefix="binary",setnull=False,returnlistraster=True,category_list=None,ncores=2):
# Check if raster exists to avoid error in mutliprocessing
try:
mpset=categorical_raster.split("@")[1]
except:
mpset=""
if categorical_raster not in gscript.list_strings(type='raster',mapset=mpset):
sys.exit('Raster <%s> not found' %categorical_raster)
# Check for number of cores doesnt exceed available
nbcpu=multiprocessing.cpu_count()
if ncores>=nbcpu:
ncores=nbcpu-1
returnlist=[] #Declare empty list for return
#gscript.run_command('g.region', raster=categorical_raster, quiet=True) #Set the region
null='null()' if setnull else '0' #Set the value for r.mapcalc
minclass=1 if setnull else 2 #Set the value to check if the binary raster is empty
if category_list == None: #If no category_list provided
category_list=[cl for cl in gscript.parse_command('r.category',map=categorical_raster,quiet=True)]
for i,x in enumerate(category_list): #Make sure the format is UTF8 and not Unicode
category_list[i]=x.encode('UTF8')
category_list.sort(key=float) #Sort the raster categories in ascending.
p=Pool(ncores) #Create a pool of processes and launch them using 'map' function
func=partial(get_binary,categorical_raster,prefix,null,minclass) # Set the two fixed argument of the function
returnlist=p.map(func,category_list) # Launch the processes for as many items in the 'functions_name' list and get the ordered results using map function
p.close()
p.join()
if returnlistraster:
return returnlist
#### Function that extract binary raster for a specified class (called in 'create_binary_raster' function)
def get_binary(categorical_raster,prefix,null,minclass,cl):
binary_class=prefix+"_"+cl
gscript.run_command('r.mapcalc', expression=binary_class+'=if('+categorical_raster+'=='+str(cl)+',1,'+null+')',overwrite=True, quiet=True)
if len(gscript.parse_command('r.category',map=binary_class,quiet=True))>=minclass: #Check if created binary is not empty
return binary_class
else:
gscript.run_command('g.remove', quiet=True, flags="f", type='raster', name=binary_class)
# -
# ### Function for computation of spatial metrics at landscape level (multiprocessed)
# +
##### Function that compute different landscape metrics (spatial metrics) at landscape level.
### The metric computed are "dominance","pielou","renyi","richness","shannon","simpson".
### It is important to set the computation region before runing this script so that it match the extent of the 'raster' layer.
# 'configfile' wait for the path (string) to the configuration file corresponding to the 'raster' layer.
# 'raster' wait for the name (string) of the landcover map on which landscape metrics will be computed.
# 'returnlistresult' wait for a boolean value (True/False) according to the fact that a list containing the path to the result files is desired.
# 'ncores' wait for a integer corresponding to the number of desired cores to be used for parallelization.
# Import libraries for multiprocessing
import multiprocessing
from multiprocessing import Pool
from functools import partial
def compute_landscapelevel_metrics(configfile, raster, spatial_metric):
filename=raster.split("@")[0]+"_%s" %spatial_metric
outputfile=os.path.join(os.path.split(configfile)[0],"output",filename)
if spatial_metric=='renyi': # The alpha parameter was set to 2 as in https://en.wikipedia.org/wiki/R%C3%A9nyi_entropy
gscript.run_command('r.li.%s' %spatial_metric, overwrite=True,
input=raster,config=configfile,alpha='2', output=filename)
else:
gscript.run_command('r.li.%s' %spatial_metric, overwrite=True,
input=raster,config=configfile, output=filename)
return outputfile
def get_landscapelevel_metrics(configfile, raster, returnlistresult=True, ncores=2):
# Check if raster exists to avoid error in mutliprocessing
try:
mpset=raster.split("@")[1]
except:
mpset=""
if raster not in gscript.list_strings(type='raster',mapset=mpset):
sys.exit('Raster <%s> not found' %raster)
# Check if configfile exists to avoid error in mutliprocessing
if not os.path.exists(configfile):
sys.exit('Configuration file <%s> not found' %configfile)
# Check for number of cores doesnt exceed available
nbcpu=multiprocessing.cpu_count()
if ncores>=nbcpu:
ncores=nbcpu-1
if ncores>6:
ncores=6 #There are only 6 metrics to compute
# List of metrics to be computed
spatial_metric_list=["dominance","pielou","renyi","richness","shannon","simpson"]
#Declare empty list for return
returnlist=[]
# Create a new pool
p=Pool(ncores)
# Set the two fixed argument of the 'compute_landscapelevel_metrics' function
func=partial(compute_landscapelevel_metrics,configfile, raster)
# Launch the processes for as many items in the 'functions_name' list and get the ordered results using map function
returnlist=p.map(func,spatial_metric_list)
p.close()
p.join()
# Return list of paths to result files
if returnlistresult:
return returnlist
# -
# ### Function for computation of spatial metrics at class level (multiprocessed)
# +
##### Function that compute different landscape metrics (spatial metrics) at class level.
### The metric computed are "patch number (patchnum)","patch density (patchdensity)","mean patch size(mps)",
### "coefficient of variation of patch area (padcv)","range of patch area size (padrange)",
### "standard deviation of patch area (padsd)", "shape index (shape)", "edge density (edgedensity)".
### It is important to set the computation region before runing this script so that it match the extent of the 'raster' layer.
# 'configfile' wait for the path (string) to the configuration file corresponding to the 'raster' layer.
# 'raster' wait for the name (string) of the landcover map on which landscape metrics will be computed.
# 'returnlistresult' wait for a boolean value (True/False) according to the fact that a list containing the path to the result files is desired.
# 'ncores' wait for a integer corresponding to the number of desired cores to be used for parallelization.
# Import libraries for multiprocessing
import multiprocessing
from multiprocessing import Pool
from functools import partial
def compute_classlevel_metrics(configfile, raster, spatial_metric):
filename=raster.split("@")[0]+"_%s" %spatial_metric
gscript.run_command('r.li.%s' %spatial_metric, overwrite=True,
input=raster,config=configfile,output=filename)
outputfile=os.path.join(os.path.split(configfile)[0],"output",filename)
return outputfile
def get_classlevel_metrics(configfile, raster, returnlistresult=True, ncores=2):
# Check if raster exists to avoid error in mutliprocessing
try:
mpset=raster.split("@")[1]
except:
mpset=""
if raster not in [x.split("@")[0] for x in gscript.list_strings(type='raster',mapset=mpset)]:
sys.exit('Raster <%s> not found' %raster)
# Check if configfile exists to avoid error in mutliprocessing
if not os.path.exists(configfile):
sys.exit('Configuration file <%s> not found' %configfile)
# Check for number of cores doesnt exceed available
nbcpu=multiprocessing.cpu_count()
if ncores>=nbcpu:
ncores=nbcpu-1
if ncores>8:
ncores=8 #There are only 8 metrics to compute
# List of metrics to be computed
spatial_metric_list=["patchnum","patchdensity","mps","padcv","padrange","padsd","shape","edgedensity"]
# Declare empty list for return
returnlist=[]
# Create a new pool
p=Pool(ncores)
# Set the two fixed argument of the 'compute_classlevel_metrics' function
func=partial(compute_classlevel_metrics,configfile, raster)
# Launch the processes for as many items in the 'functions_name' list and get the ordered results using map function
returnlist=p.map(func,spatial_metric_list)
p.close()
p.join()
# Return list of paths to result files
if returnlistresult:
return returnlist
# -
# **-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
# <center> <font size=5> <h1>User inputs</h1> </font> </center>
## Define a empty dictionnary for saving user inputs
user={}
# +
## Enter the path to GRASSDATA folder
user["gisdb"] = "/home/tais/Documents/GRASSDATA_Spie2017subset_Ouaga"
## Enter the name of the location (existing or for a new one)
user["location"] = "SPIE_subset"
## Enter the EPSG code for this location
user["locationepsg"] = "32630"
## Enter the name of the mapset to use for segmentation
user["mapsetname"] = "test_rli"
# -
# **-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
# # Compute spatial metrics for deriving land use in street blocs
# **Launch GRASS GIS working session**
# +
## Set the name of the mapset in which to work
mapsetname=user["mapsetname"]
## Launch GRASS GIS working session in the mapset
if os.path.exists(os.path.join(user["gisdb"],user["location"],mapsetname)):
gsetup.init(os.environ['GISBASE'], user["gisdb"], user["location"], mapsetname)
print "You are now working in mapset '"+mapsetname+"'"
else:
print "'"+mapsetname+"' mapset doesn't exists in "+user["gisdb"]
# -
# **-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
# ### Create binary rasters from the base landcover map
# +
# Save time for computing processin time
begintime=time.time()
# Set the name of the 'base' landcover map
baselandcoverraster="classif@test_rli"
# Create as many binary raster layer as categorical values existing in the base landcover map
gscript.run_command('g.region', raster=baselandcoverraster, quiet=True) #Set the region
pref=baselandcoverraster.split("@")[0]+"_cl" #Set the prefix
raster_list=[] # Initialize a empty list for results
raster_list=create_binary_raster(baselandcoverraster,
prefix=pref,setnull=True,returnlistraster=True,
category_list=None,ncores=15) #Extract binary raster
# Compute and print processing time
print_processing_time(begintime,"Extraction of binary rasters achieved in ")
# -
# Insert the name of the base landcover map at first position in the list
raster_list.insert(0,baselandcoverraster)
# Display the raster to be used for landscape analysis
raster_list
# + active=""
# raster_list=['classif@test_rli',
# 'classif_cl_11',
# 'classif_cl_13',
# 'classif_cl_14',
# 'classif_cl_20',
# 'classif_cl_30',
# 'classif_cl_31',
# 'classif_cl_41',
# 'classif_cl_51']
# -
# ## Create r.li configuration file for a list of landcover rasters
# Save time for computing processin time
begintime=time.time()
# Set the name of the vector polygon layer containing the
landscape_polygons="streetblocks@PERMANENT"
# Run creation of r.li configuration file and associated raster layers
list_configfile=create_rli_configfile(raster_list,landscape_polygons,returnlistpath=True)
# Compute and print processing time
print_processing_time(begintime,"Extraction of binary rasters achieved in ")
# Display the path to the configuration files created
list_configfile
# + active=""
# list_configfile=[u'/home/tais/.grass7/r.li/landcover_spie17_streetblocks',
# u'/home/tais/.grass7/r.li/landcover_spie17_cl_11_streetblocks',
# u'/home/tais/.grass7/r.li/landcover_spie17_cl_13_streetblocks',
# u'/home/tais/.grass7/r.li/landcover_spie17_cl_14_streetblocks',
# u'/home/tais/.grass7/r.li/landcover_spie17_cl_21_streetblocks',
# u'/home/tais/.grass7/r.li/landcover_spie17_cl_22_streetblocks',
# u'/home/tais/.grass7/r.li/landcover_spie17_cl_31_streetblocks',
# u'/home/tais/.grass7/r.li/landcover_spie17_cl_32_streetblocks',
# u'/home/tais/.grass7/r.li/landcover_spie17_cl_33_streetblocks',
# u'/home/tais/.grass7/r.li/landcover_spie17_cl_34_streetblocks',
# u'/home/tais/.grass7/r.li/landcover_spie17_cl_41_streetblocks',
# u'/home/tais/.grass7/r.li/landcover_spie17_cl_51_streetblocks']
# -
# **-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
# ## Compute spatial metrics at landscape level
# Initialize an empty list which will contains the resultfiles
resultfiles=[]
# Save time for computing processin time
begintime=time.time()
# Get the path to the configuration file for the base landcover raster
currentconfigfile=list_configfile[0]
# Get the name of the base landcover raster
currentraster=raster_list[0]
# Set the region to match the extent of the base raster
gscript.run_command('g.region', raster=currentraster, quiet=True)
# Launch the processes for as many items in the 'functions_name' list and get the ordered results using map function
resultfiles.append(get_landscapelevel_metrics(currentconfigfile, currentraster, returnlistresult=True, ncores=10))
# Compute and print processing time
print_processing_time(begintime,"Computation of spatial metric achieved in ")
resultfiles
# + active=""
# resultfiles=[[u'/home/tais/.grass7/r.li/output/landcover_spie17_dominance',
# u'/home/tais/.grass7/r.li/output/landcover_spie17_pielou',
# u'/home/tais/.grass7/r.li/output/landcover_spie17_renyi',
# u'/home/tais/.grass7/r.li/output/landcover_spie17_richness',
# u'/home/tais/.grass7/r.li/output/landcover_spie17_shannon',
# u'/home/tais/.grass7/r.li/output/landcover_spie17_simpson']]
# -
# ## Compute spatial metrics at class level
# +
# Save time for computing processin time
begintime=time.time()
# Get a list with paths to the configuration file for class level metrics
classlevelconfigfiles=list_configfile[1:]
# Get a list with name of binary landcover raster for class level metrics
classlevelrasters=raster_list[1:]
for x,currentraster in enumerate(classlevelrasters[:]):
# Get the path to the configuration file for the base landcover raster
currentconfigfile=classlevelconfigfiles[x]
# Set the region to match the extent of the base raster
gscript.run_command('g.region', raster=currentraster, quiet=True)
# Launch the processes for as many items in the 'functions_name' list and get the ordered results using map function
resultfiles.append(get_classlevel_metrics(currentconfigfile, currentraster, returnlistresult=True, ncores=10))
# Compute and print processing time
print_processing_time(begintime,"Computation of spatial metric achieved in ")
# -
resultfiles
# Flat the 'resultfiles' list which contains several lists
resultfiles=[item for sublist in resultfiles for item in sublist]
resultfiles
# ## Compute some special metrics
# ### Mean and standard deviation of NDVI
# ### Mean and standard deviation of SAR textures
# ### Mean and standard deviation of building's height
# # Importing the NDVI layer
# +
break
## Saving current time for processing time management
begintime_ndvi=time.time()
## Import nDSM imagery
print ("Importing NDVI raster imagery at " + time.ctime())
gscript.run_command('r.import',
input="/media/tais/data/MAUPP/WorldView3_Ouagadougou/Orthorectified/mosaique_georef/NDVI/ndvi_georef_ordre2.TIF",
output="ndvi", overwrite=True)
# Mask null/nodata values
gscript.run_command('r.null', map="ndvi")
print_processing_time(begintime_ndvi, "imagery has been imported in ")
# -
# # Importing the nDSM layer
# +
break
## Saving current time for processing time management
begintime_ndsm=time.time()
## Import nDSM imagery
print ("Importing nDSM raster imagery at " + time.ctime())
grass.run_command('r.import',
input="/media/tais/data/MAUPP/WorldView3_Ouagadougou/Orthorectified/mosaique_georef/nDSM/nDSM_mosaik_georef_ordre2.tif",
output="ndsm", overwrite=True)
## Define null value for specific value in nDSM raster. Adapt the value to your own data.
# If there is no null value in your data, comment the next line
grass.run_command('r.null', map="ndsm", setnull="-999")
# Make histogram equalisation on grey color.
grass.run_command('r.colors', flags='e', map='ndsm', color='grey')
print_processing_time(begintime_ndsm, "nDSM imagery has been imported in ")
# -
# ### Masking the nDSM artifacts
# +
break
# Import vector with nDSM artifacts zones
grass.run_command('v.in.ogr', overwrite=True,
input="/media/tais/data/MAUPP/WorldView3_Ouagadougou/Masque_artifacts_nDSM/Ouaga_mask_artifacts_nDSM.shp",
output="mask_artifacts_ndsm")
## Set computational region to match the default region
grass.run_command('g.region', overwrite=True, raster="ndsm")
# Rasterize the vector layer, with value "0" on the artifacts zones
grass.run_command('v.to.rast', input='mask_artifacts_ndsm', output='mask_artifacts_ndsm',
use='val', value='0', memory='5000')
## Set computational region to match the default region
grass.run_command('g.region', overwrite=True, raster="ndsm")
## Create a new nDSM with artifacts filled with '0' value
formula='tmp_artifact=nmin(ndsm,mask_artifacts_ndsm)'
grass.mapcalc(formula, overwrite=True)
## Remove the artifact mask
grass.run_command('g.remove', flags='f', type='raster', name="mask_artifacts_ndsm")
## Rename the new nDSM
grass.run_command('g.rename', raster='tmp_artifact,ndsm', overwrite=True)
## Remove the intermediate nDSM layer
grass.run_command('g.remove', flags='f', type='raster', name="tmp_artifact")
# -
# **-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
# # Define input raster for computing statistics of segments
## Display the name of rasters available in PERMANENT and CLASSIFICATION mapset
print grass.read_command('g.list',type="raster", mapset="PERMANENT", flags='rp')
print grass.read_command('g.list',type="raster", mapset=user["classificationA_mapsetname"], flags='rp')
# +
## Define the list of raster layers for which statistics will be computed
inputstats=[]
inputstats.append("opt_blue")
inputstats.append("opt_green")
inputstats.append("opt_red")
inputstats.append("opt_nir")
inputstats.append("ndsm")
inputstats.append("ndvi")
print "Layer to be used to compute raster statistics of segments:\n"+'\n'.join(inputstats)
# +
## Define the list of raster statistics to be computed for each raster layer
rasterstats=[]
rasterstats.append("min")
rasterstats.append("max")
rasterstats.append("range")
rasterstats.append("mean")
rasterstats.append("stddev")
#rasterstats.append("coeff_var") # Seems that this statistic create null values
rasterstats.append("median")
rasterstats.append("first_quart")
rasterstats.append("third_quart")
rasterstats.append("perc_90")
print "Raster statistics to be computed:\n"+'\n'.join(rasterstats)
# +
## Define the list of area measures (segment's shape statistics) to be computed
areameasures=[]
areameasures.append("area")
areameasures.append("perimeter")
areameasures.append("compact_circle")
areameasures.append("compact_square")
areameasures.append("fd")
print "Area measures to be computed:\n"+'\n'.join(areameasures)
# -
# **-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
# <center> <font size=5> <h1>Compute objects' statistics</h1> </font> </center>
## Saving current time for processing time management
begintime_computeobjstat=time.time()
# ## Define the folder where to save the results and create it if necessary
# In the next cell, please adapt the path to the directory where you want to save the .csv output of i.segment.uspo.
# +
## Folder in which save processing time output
outputfolder="/media/tais/My_Book_1/MAUPP/Traitement/Ouagadougou/Segmentation_fullAOI_localapproach/Results/CLASSIF/stats_optical"
## Create the folder if does not exists
if not os.path.exists(outputfolder):
os.makedirs(outputfolder)
print "Folder '"+outputfolder+"' created"
# -
# ### Copy data from other mapset to the current mapset
# Some data need to be copied from other mapsets into the current mapset.
# ### Remove current mask
## Check if there is a raster layer named "MASK"
if not grass.list_strings("rast", pattern="MASK", mapset=mapsetname, flag='r'):
print 'There is currently no MASK'
else:
## Remove the current MASK layer
grass.run_command('r.mask',flags='r')
print 'The current MASK has been removed'
# ***Copy segmentation raster***
## Copy segmentation raster layer from SEGMENTATION mapset to current mapset
grass.run_command('g.copy', overwrite=True,
raster="segmentation_raster@"+user["segmentation_mapsetname"]+",segments")
# ***Copy morphological zone (raster)***
## Copy segmentation raster layer from SEGMENTATION mapset to current mapset
grass.run_command('g.copy', overwrite=True,
raster="zone_morpho@"+user["segmentation_mapsetname"]+",zone_morpho")
# ***Copy morphological zone (vector)***
## Copy segmentation raster layer from SEGMENTATION mapset to current mapset
grass.run_command('g.copy', overwrite=True,
vector="zone_morpho@"+user["segmentation_mapsetname"]+",zone_morpho")
# **-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
# # Compute statistics of segments (Full AOI extend)
# ### Compute statistics of segment using i.segment.stats
# The process is make to compute statistics iteratively for each morphological zones, used here as tiles.
# This section uses the ['i.segment.stats' add-on](https://grass.osgeo.org/grass70/manuals/addons/i.segment.stats.html) to compute statistics for each object.
## Save name of the layer to be used as tiles
tile_layer='zone_morpho'+'@'+mapsetname
## Save name of the segmentation layer to be used by i.segment.stats
segment_layer='segments'+'@'+mapsetname
## Save name of the column containing area_km value
area_column='area_km2'
## Save name of the column containing morphological type value
type_column='type'
## Save the prefix to be used for the outputfiles of i.segment.stats
prefix="Segstat"
## Save the list of polygons to be processed (save the 'cat' value)
listofregion=list(grass.parse_command('v.db.select', map=tile_layer,
columns='cat', flags='c'))[:]
for count, cat in enumerate(listofregion):
print str(count)+" cat:"+str(cat)
# + active=""
# ## Just a print to identify which cat correspond to which area and morpho type
#
# for count, cat in enumerate(listofregion):
# condition="cat="+cat
# typemorpho=(grass.read_command('v.db.select', map=tile_layer,
# columns=type_column, where=condition,flags="c"))
# area=(grass.read_command('v.db.select', map=tile_layer,
# columns=area_column, where=condition,flags="c"))
# print str(count)+" cat:"+str(cat)+" morpho_type:"+str(typemorpho)+" area:"+str(area)
# +
## Initialize a empty string for saving print outputs
txtcontent=""
## Running i.segment.stats
messagetoprint="Start computing statistics for segments to be classified, using i.segment.stats on "+time.ctime()+"\n"
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
begintime_isegmentstats=time.time()
## Compute total area to be processed for process progression information
processed_area=0
nbrtile=len(listofregion)
attributes=grass.parse_command('db.univar', flags='g', table=tile_layer.split("@")[0], column=area_column, driver='sqlite')
total_area=float(attributes['sum'])
messagetoprint=str(nbrtile)+" region(s) will be processed, covering an area of "+str(round(total_area,3))+" Sqkm."+"\n\n"
print (messagetoprint)
txtcontent+=messagetoprint
## Save time before looping
begintime_isegmentstats=time.time()
## Start loop on morphological zones
count=1
for cat in listofregion[:]:
## Save current time at loop' start.
begintime_current_id=time.time()
## Create a computional region for the current polygon
condition="cat="+cat
outputname="tmp_"+cat
grass.run_command('v.extract', overwrite=True, quiet=True,
input=tile_layer, type='area', where=condition, output=outputname)
grass.run_command('g.region', overwrite=True, vector=outputname, align=segment_layer)
grass.run_command('r.mask', overwrite=True, raster=tile_layer, maskcats=cat)
grass.run_command('g.remove', quiet=True, type="vector", name=outputname, flags="f")
## Save size of the current polygon and add it to the already processed area
size=round(float(grass.read_command('v.db.select', map=tile_layer,
columns=area_column, where=condition,flags="c")),2)
## Print
messagetoprint="Computing segments's statistics for tile n°"+str(cat)
messagetoprint+=" ("+str(count)+"/"+str(len(listofregion))+")"
messagetoprint+=" corresponding to "+str(size)+" km2"
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
## Define the csv output file name, according to the optimization function selected
outputcsv=os.path.join(outputfolder,prefix+"_"+str(cat)+".csv")
## Compute statistics of objets using i.segment.stats only with .csv output (no vectormap output).
grass.run_command('i.segment.stats', overwrite=True, map=segment_layer,
rasters=','.join(inputstats), raster_statistics=','.join(rasterstats),
area_measures=','.join(areameasures), csvfile=outputcsv, processes='20')
## Add the size of the zone to the already processed area
processed_area+=size
## Print
messagetoprint=print_processing_time(begintime_current_id,
"i.segment.stats finishes to process th current tile in ")
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
remainingtile=nbrtile-count
if remainingtile>0:
messagetoprint=str(round((processed_area/total_area)*100,2))+" percent of the total area processed. "
messagetoprint+="Still "+str(remainingtile)+" zone(s) to process."+"\n"
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
else:
messagetoprint="\n"
print (messagetoprint)
txtcontent+=messagetoprint
## Adapt the count
count+=1
## Remove current mask
grass.run_command('r.mask', flags='r')
## Compute processing time and print it
messagetoprint=print_processing_time(begintime_isegmentstats, "Statitics computed in ")
print (messagetoprint)
txtcontent+=messagetoprint
#### Write text file with log of processing time
## Create the .txt file for processing time output and begin to write
f = open(os.path.join(outputfolder,mapsetname+"_processingtime_isegmentstats.txt"), 'w')
f.write(mapsetname+" processing time information for i.segment.stats"+"\n\n")
f.write(txtcontent)
f.close()
# -
## print
print_processing_time(begintime_computeobjstat,"Object statistics computed in ")
# ## Concatenate individuals .csv files and replace unwanted values
# BE CAREFUL! Before runing the following cells, please check your data to be sure that it makes sens to replace the 'nan', 'null', or 'inf' values with "0"
## Define the outputfile for .csv containing statistics for all segments
outputfile=os.path.join(outputfolder,"all_segments_stats.csv")
print outputfile
# +
# Create a dictionary with 'key' to be replaced by 'values'
findreplacedict={}
findreplacedict['nan']="0"
findreplacedict['null']="0"
findreplacedict['inf']="0"
# Define pattern of file to concatenate
pat=prefix+"_*.csv"
sep="|"
# +
## Initialize a empty string for saving print outputs
txtcontent=""
## Saving current time for processing time management
begintime_concat=time.time()
## Print
messagetoprint="Start concatenate individual .csv files and replacing unwanted values."
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
# Concatenate and replace unwanted values
messagetoprint=concat_findreplace(outputfolder,pat,sep,findreplacedict,outputfile)
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
## Compute processing time and print it
messagetoprint=print_processing_time(begintime_concat, "Process achieved in ")
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
#### Write text file with log of processing time
## Create the .txt file for processing time output and begin to write
filepath=os.path.join(outputfolder,mapsetname+"_processingtime_concatreplace.txt")
f = open(filepath, 'w')
f.write(mapsetname+" processing time information for concatenation of individual .csv files and replacing of unwanted values."+"\n\n")
f.write(txtcontent)
f.close()
# -
# # Create new database in postgresql
# User for postgresql connexion
dbuser="tais"
# Password of user
dbpassword="<PASSWORD>"
# Host of database
host="localhost"
# Name of the new database
dbname="ouaga_fullaoi_localsegment"
# Set name of schema for objects statistics
stat_schema="statistics"
# Set name of table with statistics of segments - FOR OPTICAL
object_stats_table="object_stats_optical"
# +
break
from psycopg2.extensions import ISOLATION_LEVEL_AUTOCOMMIT
# Connect to postgres database
db=None
db=pg.connect(dbname='postgres', user=dbuser, password=<PASSWORD>, host=host)
# Allow to create a new database
db.set_isolation_level(ISOLATION_LEVEL_AUTOCOMMIT)
# Execute the CREATE DATABASE query
cur=db.cursor()
#cur.execute('DROP DATABASE IF EXISTS ' + dbname) #Comment this to avoid deleting existing DB
cur.execute('CREATE DATABASE ' + dbname)
cur.close()
db.close()
# -
# ### Create PostGIS Extension in the database
break
# Connect to the database
db=pg.connect(database=dbname, user=dbuser, password=<PASSWORD>, host=host)
# Open a cursor to perform database operations
cur=db.cursor()
# Execute the query
cur.execute('CREATE EXTENSION IF NOT EXISTS postgis')
# Make the changes to the database persistent
db.commit()
# Close connection with database
cur.close()
db.close()
# <center> <font size=4> <h2>Import statistics of segments in a Postgresql database</h2> </font> </center>
# ## Create new schema in the postgresql database
schema=stat_schema
# +
from psycopg2.extensions import ISOLATION_LEVEL_AUTOCOMMIT
# Connect to postgres database
db=None
db=pg.connect(dbname=dbname, user='tais', password='<PASSWORD>', host='localhost')
# Allow to create a new database
db.set_isolation_level(ISOLATION_LEVEL_AUTOCOMMIT)
# Execute the CREATE DATABASE query
cur=db.cursor()
#cur.execute('DROP SCHEMA IF EXISTS '+schema+' CASCADE') #Comment this to avoid deleting existing DB
try:
cur.execute('CREATE SCHEMA '+schema)
except Exception as e:
print ("Exception occured : "+str(e))
cur.close()
db.close()
# -
# ## Create a new table
# Connect to an existing database
db=pg.connect(database=dbname, user=dbuser, password=<PASSWORD>, host=host)
# Open a cursor to perform database operations
cur=db.cursor()
# Drop table if exists:
cur.execute("DROP TABLE IF EXISTS "+schema+"."+object_stats_table)
# Make the changes to the database persistent
db.commit()
# +
import csv
# Create a empty list for saving of column name
column_name=[]
# Create a reader for the first csv file in the stack of csv to be imported
pathtofile=os.path.join(outputfolder, outputfile)
readercsvSubset=open(pathtofile)
readercsv=csv.reader(readercsvSubset, delimiter='|')
headerline=readercsv.next()
print "Create a new table '"+schema+"."+object_stats_table+"' with header corresponding to the first row of file '"+pathtofile+"'"
## Build a query for creation of a new table with auto-incremental key-value (thus avoiding potential duplicates of 'cat' value)
# All column data-types are set to 'text' in order to be able to import some 'nan', 'inf' or 'null' values present in statistics files
# This table will allow to import all individual csv files in a single Postgres table, which will be cleaned after
query="CREATE TABLE "+schema+"."+object_stats_table+" ("
query+="key_value serial PRIMARY KEY"
query+=", "+str(headerline[0])+" text"
column_name.append(str(headerline[0]))
for column in headerline[1:]:
if column[0] in ('1','2','3','4','5','6','7','8','9','0'):
query+=","
query+=" "+"W"+str(column)+" double precision"
column_name.append("W"+str(column))
else:
query+=","
query+=" "+str(column)+" double precision"
column_name.append(str(column))
query+=")"
# Execute the CREATE TABLE query
cur.execute(query)
# Make the changes to the database persistent
db.commit()
# Close cursor and communication with the database
cur.close()
db.close()
# -
# ## Copy objects statistics from csv to Postgresql database
# Connect to an existing database
db=pg.connect(database=dbname, user=dbuser, password=<PASSWORD>, host=host)
# Open a cursor to perform database operations
cur=db.cursor()
# +
## Initialize a empty string for saving print outputs
txtcontent=""
## Saving current time for processing time management
begintime_copy=time.time()
## Print
messagetoprint="Start copy of segments' statistics in the postgresql table '"+schema+"."+object_stats_table+"'"
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
# Create query for copy data from csv, avoiding the header, and updating only the column which are in the csv (to allow auto-incremental key value to wokr)
query="COPY "+schema+"."+object_stats_table+"("+', '.join(column_name)+") "
query+=" FROM '"+str(pathtofile)+"' HEADER DELIMITER '|' CSV;"
# Execute the COPY FROM CSV query
cur.execute(query)
# Make the changes to the database persistent
db.commit()
## Compute processing time and print it
messagetoprint=print_processing_time(begintime_copy, "Process achieved in ")
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
#### Write text file with log of processing time
## Create the .txt file for processing time output and begin to write
filepath=os.path.join(outputfolder,mapsetname+"_processingtime_PostGimport.txt")
f = open(filepath, 'w')
f.write(mapsetname+" processing time information for importation of segments' statistics in the PostGreSQL Database."+"\n\n")
f.write(txtcontent)
f.close()
# -
# Close cursor and communication with the database
cur.close()
db.close()
# # Drop duplicate values of CAT
# Here, we will find duplicates. Indeed, as statistics are computed for each tile (morphological area) and computational region aligned to the pixels raster, some objets could appear in two different tile resulting on duplicates on "CAT" column.
#
# We firs select the "CAT" of duplicated objets and then puting them in a list. Then, for each duplicated "CAT", we select the key-value (primary key) of the smallest object (area_min). The row corresponding to those key-values are then remoed using the "DELETE FROM" query.
# Connect to an existing database
db=pg.connect(database=dbname, user=dbuser, password=<PASSWORD>, host=host)
# Open a cursor to perform database operations
cur=db.cursor()
# +
## Initialize a empty string for saving print outputs
txtcontent=""
## Saving current time for processing time management
begintime_removeduplic=time.time()
## Print
messagetoprint="Start removing duplicates in the postgresql table '"+schema+"."+object_stats_table+"'"
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
# Find duplicated 'CAT'
find_duplicated_cat()
# Remove duplicated
count_pass=1
count_removedduplic=0
while len(cattodrop)>0:
messagetoprint="Removing duplicates - Pass "+str(count_pass)
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
find_duplicated_key()
remove_duplicated_key()
messagetoprint=str(len(keytodrop))+" duplicates removed."
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
count_removedduplic+=len(keytodrop)
# Find again duplicated 'CAT'
find_duplicated_cat()
count_pass+=1
messagetoprint="A total of "+str(count_removedduplic)+" duplicates were removed."
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
## Compute processing time and print it
messagetoprint=print_processing_time(begintime_removeduplic, "Process achieved in ")
print (messagetoprint)
txtcontent+=messagetoprint+"\n"
#### Write text file with log of processing time
## Create the .txt file for processing time output and begin to write
filepath=os.path.join(outputfolder,mapsetname+"_processingtime_RemoveDuplic.txt")
f = open(filepath, 'w')
f.write(mapsetname+" processing time information for removing duplicated objects."+"\n\n")
f.write(txtcontent)
f.close()
# -
# Vacuum the current Postgresql database
vacuum(db)
# # Change the primary key from 'key_value' to 'cat'
# Connect to an existing database
db=pg.connect(database=dbname, user=dbuser, password=<PASSWORD>, host=host)
# Open a cursor to perform database operations
cur=db.cursor()
# +
# Build a query to drop the current constraint on primary key
query="ALTER TABLE "+schema+"."+object_stats_table+" \
DROP CONSTRAINT "+object_stats_table+"_pkey"
# Execute the query
cur.execute(query)
# Make the changes to the database persistent
db.commit()
# Build a query to change the datatype of 'cat' to 'integer'
query="ALTER TABLE "+schema+"."+object_stats_table+" \
ALTER COLUMN cat TYPE integer USING cat::integer"
# Execute the query
cur.execute(query)
# Make the changes to the database persistent
db.commit()
# Build a query to add primary key on 'cat'
query="ALTER TABLE "+schema+"."+object_stats_table+" \
ADD PRIMARY KEY (cat)"
# Execute the query
cur.execute(query)
# Make the changes to the database persistent
db.commit()
# Build a query to drop column 'key_value'
query="ALTER TABLE "+schema+"."+object_stats_table+" \
DROP COLUMN key_value"
# Execute the query
cur.execute(query)
# Make the changes to the database persistent
db.commit()
# -
# Vacuum the current Postgresql database
vacuum(db)
# Close cursor and communication with the database
cur.close()
db.close()
# ### Show first rows of statistics
# Connect to an existing database
db=pg.connect(database=dbname, user=dbuser, password=<PASSWORD>, host=host)
# Number of line to show (please limit to 100 for saving computing time)
nbrow=15
# Query
query="SELECT * FROM "+schema+"."+object_stats_table+" \
ORDER BY cat \
ASC LIMIT "+str(nbrow)
# Execute query through panda
df=pd.read_sql(query, db)
# Show dataframe
df.head(15)
# <left> <font size=4> <b> End of classification part </b> </font> </left>
print("The script ends at "+ time.ctime())
print_processing_time(begintime_segmentation_full, "Entire process has been achieved in ")
# **-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-_-**
| .ipynb_checkpoints/Landuse_classification-Copy6-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext Cython
# + language="cython"
# cdef int i,j,k
# cdef float proce,margin
# + language="cython"
# #cdef block
# def integrate(a,b,f):
# cdef:
# int i
# int N=2000
# float dx,s=0.0
# dx=(b-a)/N
# for i in range(N):
# s+=f(a+i*dx)
# return s*dx
# + language="cython"
# def cfib(int n):
# cdef:
# int i
# double a=0.0,b=1.0
# for i in range(n):
# a,b=a+b,a
# return a
# -
cfib(90)
def fib(n):
a,b=0.0,1.0
for i in range(n):
a,b=a+b,a
return a
# %timeit cfib(90)
# %timeit fib(90)
def automatic_inference():
i=1
d=2.0
c=3+4j
r=i*d+c
return r
# +
import cython
@cython.infer_types(True)
def more_inference():
i=1
d=2.0
c=3+4j
r=i*d+c
return r
# -
| cythonTest/syntax/notebook/syntax.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 9.1
# language: sagemath
# metadata:
# cocalc:
# description: Open-source mathematical software system
# priority: 1
# url: https://www.sagemath.org/
# name: sage-9.1
# resource_dir: /ext/jupyter/kernels/sage-9.1
# ---
# ### A geometric sequence is such that its next term is always generated by multiplying a fixed number K to its previous term
[5 , 50, 500, 5000] # We multiply 10 each time
[ 3* 2^k for k in [1..10]] # We multiply 2 each time
# ## Find the sum of $3^1 + 3^2 + 3^3 + \dots + 3^{12} $
sum([3^k for k in [1..12]])
(3^13 - 3) / 2
88572 * 2 + 3
# ## How about $5^1 + 5^2 + 5^3 + \dots + 5^{12} $
sum([5^k for k in [1..12]])
(5^13 - 5) / 4
# ## What if we change the index and base $7^1 + 7^2 + 7^3 + \dots + 7^9$
(7^10 -7)/6
sum([7^k for k in [1..9]])
# ## And, next $9^1 + 9^2 + 9^3 + \dots + 9^{13} $
(9^14 - 9)/8
# ## Compute this immediately without calculator $ 10^0 + 10^1 + 10 ^2 + 10^3 + \dots + 10^{23}$
(10^24 - 1) / 9
# # Sum of geometric sequence
# $$ 1+ x + x^2 + x^3 + \dots + x^n = \frac{x^{n+1} - 1}{x-1}$$ for all numbers $x$
| 20210407/lecture/2 - Geometric sequence.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import ipywidgets as widgets
from IPython.display import display
# #### Note that beyond the basic example most of this notebook is just reference
# ## Basic Example
# Since button clicks are stateless, they are transmitted from the front-end to the back-end using custom messages. By using the `on_click` method, a button that prints a message when it has been clicked is shown below. To capture prints (or any other kind of output) and ensure it is displayed, be sure to send it to an `Output` widget (or put the information you want to display into an HTML widget).
# +
button = widgets.Button(description="Click Me!")
output = widgets.Output()
display(button, output)
def on_button_clicked(b):
with output:
print("Button clicked.")
button.on_click(on_button_clicked)
# -
# ### Create a callback that watches a widget for value changes
# +
int_range = widgets.IntSlider()
output2 = widgets.Output()
display(int_range, output2)
def on_value_change(change):
with output2:
print(change['new'])
int_range.observe(on_value_change, names='value')
# -
# ## Linking two widgets (Reference Only)
# NB 01_Widgets_list shows how to link two widgets using widgets.jslink and the syntax below, it appears there
# are two others, link and dlink that have to do with traitlets but im not sure how they are different
caption = widgets.Label(value='The values of slider1 and slider2 are synchronized')
sliders1, slider2 = widgets.IntSlider(description='Slider 1'),\
widgets.IntSlider(description='Slider 2')
l = widgets.link((sliders1, 'value'), (slider2, 'value'))
display(caption, sliders1, slider2)
caption = widgets.Label(value='Changes in source values are reflected in target1')
source, target1 = widgets.IntSlider(description='Source'),\
widgets.IntSlider(description='Target 1')
dl = widgets.dlink((source, 'value'), (target1, 'value'))
display(caption, source, target1)
# +
caption = widgets.Label(value='The values of range1 and range2 are synchronized')
slider = widgets.IntSlider(min=-5, max=5, value=1, description='Slider')
def handle_slider_change(change):
caption.value = 'The slider value is ' + (
'negative' if change.new < 0 else 'nonnegative'
)
slider.observe(handle_slider_change, names='value')
display(caption, slider)
# -
# ## Linking in kernel vs client
# ### Kernel only works if python is running so nbviewer tutorials may need jslink
# 
# ## Advaned ways to slow down event calls on continuous sliders/textareas
# #### Debouncing, ignores function calls if a certain amount of time hasnt passed
# +
import asyncio
class Timer:
def __init__(self, timeout, callback):
self._timeout = timeout
self._callback = callback
self._task = asyncio.ensure_future(self._job())
async def _job(self):
await asyncio.sleep(self._timeout)
self._callback()
def cancel(self):
self._task.cancel()
def debounce(wait):
""" Decorator that will postpone a function's
execution until after `wait` seconds
have elapsed since the last time it was invoked. """
def decorator(fn):
timer = None
def debounced(*args, **kwargs):
nonlocal timer
def call_it():
fn(*args, **kwargs)
if timer is not None:
timer.cancel()
timer = Timer(wait, call_it)
return debounced
return decorator
slider = widgets.IntSlider()
text = widgets.IntText()
@debounce(0.2)
def value_changed(change):
text.value = change.new
slider.observe(value_changed, 'value')
widgets.VBox([slider, text])
# -
# #### Throttling limits the rate of calls that are made
# +
import asyncio
from time import time
def throttle(wait):
""" Decorator that prevents a function from being called
more than once every wait period. """
def decorator(fn):
time_of_last_call = 0
scheduled = False
new_args, new_kwargs = None, None
def throttled(*args, **kwargs):
nonlocal new_args, new_kwargs, time_of_last_call, scheduled
def call_it():
nonlocal new_args, new_kwargs, time_of_last_call, scheduled
time_of_last_call = time()
fn(*new_args, **new_kwargs)
scheduled = False
time_since_last_call = time() - time_of_last_call
new_args = args
new_kwargs = kwargs
if not scheduled:
new_wait = max(0, wait - time_since_last_call)
Timer(new_wait, call_it)
scheduled = True
return throttled
return decorator
# +
slider = widgets.IntSlider()
text = widgets.IntText()
@throttle(0.2)
def value_changed(change):
text.value = change.new
slider.observe(value_changed, 'value')
widgets.VBox([slider, text])
# -
| notes/03_Widget_Events.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
#Automated reloading of custom modules
# %load_ext autoreload
# %autoreload 2
#Standard modules
import os
import time
import random
import numpy as np
import scipy
import torch
import torch.nn as nn
import torch.nn.functional as F
import matplotlib.pyplot as plt
from datetime import datetime
#Custom modules
import helpers as hlp
import models as mdl
from run import run, benchmark
# -
# # Centralized learning
device = "cuda:6"
seed = 0
pt_MNIST = run(n_clients=1, dataset="MNIST", model="LeNet5", preset="cl", feature_dim=84,reduced=0.02, rounds=100, track_history=10, device=device, seed=seed)
pt_FMNIST = run(n_clients=1, dataset="FMNIST", model="ResNet9", preset="cl", feature_dim=128, reduced=0.1, rounds=20, track_history=5, device=device, seed=seed)
pt_CIFAR10 = run(n_clients=1, dataset="CIFAR10", model="ResNet18", preset="cl", feature_dim=256, reduced=0.2, rounds=20, track_history=5, device=device, seed=seed)
# # MNIST
dataset = "MNIST"
model = "LeNet5"
reduced = 0.02
rounds = 100
n_avg = 10
track_history = False
export_dir = None
device = "cuda:7"
n_clients = 2
lambda_kd = 10
lambda_disc = 1
feature_dim = 84
seed = 0
# Experiment
pt_kd, pt_fl, pt_fd, pt_il = benchmark(n_clients=n_clients, dataset=dataset, model=model, lambda_kd=lambda_kd, lambda_disc=lambda_disc,
reduced=reduced, n_avg=n_avg, rounds=rounds, track_history=track_history,
export_dir=export_dir, device=device, seed=seed)
# +
dataset = "MNIST"
model = "LeNet5"
reduced = 0.02
rounds = 100
n_avg = 10
track_history = 2
export_dir = None
device = "cuda:7"
n_clients = 5
lambda_kd = 10
lambda_disc = 1
feature_dim = 84
seed = 0
# Experiment
pt_kd, pt_fl, pt_fd, pt_il = benchmark(n_clients=n_clients, dataset=dataset, model=model, lambda_kd=lambda_kd, lambda_disc=lambda_disc,
reduced=reduced, n_avg=n_avg, rounds=rounds, track_history=track_history,
export_dir=export_dir, device=device, seed=seed)
hlp.plot_global_training_history(pt_kd, metric="accuracy", which=["Train", "Validation (global)"], savepath="./figures/cfkd.png")
hlp.plot_global_training_history(pt_fl, metric="accuracy", which=["Train", "Validation (global)"], savepath="./figures/fl.png")
hlp.plot_global_training_history(pt_fd, metric="accuracy", which=["Train", "Validation (global)"], savepath="./figures/fd.png")
hlp.plot_global_training_history(pt_il, metric="accuracy", which=["Train", "Validation (global)"], savepath="./figures/il.png")
# -
dataset = "MNIST"
model = "LeNet5"
reduced = 0.02
rounds = 100
n_avg = 10
track_history = False
export_dir = None
device = "cuda:7"
n_clients = 10
lambda_kd = 10
lambda_disc = 1
feature_dim = 84
seed = 0
# Experiment
pt_kd, pt_fl, pt_fd, pt_il = benchmark(n_clients=n_clients, dataset=dataset, model=model, feature_dim=feature_dim, lambda_kd=lambda_kd, lambda_disc=lambda_disc,
reduced=reduced, n_avg=n_avg, rounds=rounds, track_history=track_history,
export_dir=export_dir, device=device, seed=seed)
# # Fashion-MNIST
dataset = "FMNIST"
model = "ResNet9"
reduced = 0.1
rounds = 20
n_avg = 10
track_history = False
export_dir = None
device = "cuda:7"
n_clients = 2
lambda_kd = 10
lambda_disc = 1
feature_dim = 128
seed = 0
# Experiment
pt_kd, pt_fl, pt_fd, pt_il = benchmark(n_clients=n_clients, dataset=dataset, model=model, feature_dim=feature_dim, lambda_kd=lambda_kd, lambda_disc=lambda_disc,
reduced=reduced, n_avg=n_avg, rounds=rounds, track_history=track_history,
export_dir=export_dir, device=device, seed=seed)
dadataset = "FMNIST"
model = "ResNet9"
reduced = 0.1
rounds = 20
n_avg = 10
track_history = False
export_dir = None
device = "cuda:7"
n_clients = 5
lambda_kd = 10
lambda_disc = 1
feature_dim = 128
seed = 0
# Experiment
pt_kd, pt_fl, pt_fd, pt_il = benchmark(n_clients=n_clients, dataset=dataset, model=model, feature_dim=feature_dim, lambda_kd=lambda_kd, lambda_disc=lambda_disc,
reduced=reduced, n_avg=n_avg, rounds=rounds, track_history=track_history,
export_dir=export_dir, device=device, seed=seed)
# Figure MNIST
dataset = "FMNIST"
model = "ResNet9"
reduced = 0.1
rounds = 20
n_avg = 10
track_history = 0
export_dir = None
device = "cuda:7"
n_clients = 10
lambda_kd = 10
lambda_disc = 1
feature_dim = 128
seed = 0
# Experiment
pt_kd, pt_fl, pt_fd, pt_il = benchmark(n_clients=n_clients, dataset=dataset, model=model, feature_dim=feature_dim, lambda_kd=lambda_kd, lambda_disc=lambda_disc,
reduced=reduced, n_avg=n_avg, rounds=rounds, track_history=track_history,
export_dir=export_dir, device=device, seed=seed)
# # CIFAR10
dataset = "CIFAR10"
model = "ResNet18"
reduced = 0.2
rounds = 20
n_avg = 10
track_history = 0
export_dir = None
device = "cuda:2"
n_clients = 2
lambda_kd = 10
lambda_disc = 1
feature_dim = 256
seed = 0
# Experiment
pt_kd, pt_fl, pt_fd, pt_il = benchmark(n_clients=n_clients, dataset=dataset, model=model, feature_dim=feature_dim, lambda_kd=lambda_kd, lambda_disc=lambda_disc,
reduced=reduced, n_avg=n_avg, rounds=rounds, track_history=track_history,
export_dir=export_dir, device=device, seed=seed)
dataset = "CIFAR10"
model = "ResNet18"
reduced = 0.2
rounds = 20
n_avg = 10
track_history = 0
export_dir = None
device = "cuda:7"
n_clients = 5
lambda_kd = 10
lambda_disc = 1
feature_dim = 256
seed = 0
# Experiment
pt_kd, pt_fl, pt_fd, pt_il = benchmark(n_clients=n_clients, dataset=dataset, model=model, feature_dim=feature_dim, lambda_kd=lambda_kd, lambda_disc=lambda_disc,
reduced=reduced, n_avg=n_avg, rounds=rounds, track_history=track_history,
export_dir=export_dir, device=device, seed=seed)
dataset = "CIFAR10"
model = "ResNet18"
reduced = 0.2
rounds = 20
n_avg = 10
track_history = False
export_dir = None
device = "cuda:3"
n_clients = 10
lambda_kd = 10
lambda_disc = 1
feature_dim = 256
seed = 0
# Experiment
pt_kd, pt_fl, pt_fd, pt_il = benchmark(n_clients=n_clients, dataset=dataset, model=model, lambda_kd=lambda_kd, lambda_disc=lambda_disc,
reduced=reduced, n_avg=n_avg, rounds=rounds, track_history=track_history,
export_dir=export_dir, device=device, seed=seed)
# # tSNE
# +
dataset = "MNIST"
model = "LeNet5"
reduced = 0.02
rounds = 100
n_avg = 10
track_history = 1
export_dir = None
device = "cuda:7"
n_clients = 2
lambda_kd = 10
lambda_disc = 1
feature_dim = 84
seed = 0
# Experiment
pt_list, tracker = run(n_clients=n_clients, dataset=dataset, model=model, lambda_kd=lambda_kd, lambda_disc=lambda_disc,
reduced=reduced, n_avg=n_avg, rounds=rounds, track_history=track_history,
export_dir=export_dir, device=device, seed=seed)
pt_il_list, tracker_il = run(n_clients=n_clients, dataset=dataset, model=model, preset="il",
reduced=reduced, rounds=rounds, track_history=track_history,
export_dir=export_dir, device=device, seed=seed)
pt_fl_list, tracker_fl = run(n_clients=n_clients, dataset=dataset, model=model, preset="fl",
reduced=reduced, rounds=rounds, track_history=track_history,
export_dir=export_dir, device=device, seed=seed)
# -
# Seperated figures
r_list = [1, 10, 25, 50, 75, 100]
tracker.plot_tSNE(r_list=r_list, savepath="./figures/tSNE_cfkd.png", title="Ours")
tracker_il.plot_tSNE(r_list=r_list, savepath="./figures/tSNE_il.png", title="IL")
tracker_fl.plot_tSNE(r_list=r_list, savepath="./figures/tSNE_fl.png", title="FL")
tracker_il.plot_tSNE(r_list=r_list, single_client=0, savepath="./figures/tSNE_il0.png", title="IL (Client 0)")
tracker_il.plot_tSNE(r_list=r_list, single_client=1, savepath="./figures/tSNE_il1.png", title="IL (Client 1)")
# Single figure
r_list = [1, 10, 25, 50, 75, 100]
fig, axs = plt.subplots(3, len(r_list), figsize=(3*len(r_list), 3*3))
plt.subplots_adjust(hspace=0, wspace=0)
[axs[0,i].set_title("Round {}".format(r)) for i, r in enumerate(r_list)]
tracker.plot_tSNE(r_list=r_list, title="Ours", fig_axs=(fig, axs[0,:]))
tracker_il.plot_tSNE(r_list=r_list, title="IL", fig_axs=(fig, axs[1,:]))
tracker_fl.plot_tSNE(r_list=r_list, title="FL", savepath="./figures/tSNE.png", fig_axs=(fig, axs[2,:]))
# # Lambda study
# +
dataset = "MNIST"
model = "LeNet5"
reduced = 0.02
rounds = 100
n_avg = 10
track_history = False
export_dir = None
device = "cuda:3"
n_clients = 5
feature_dim = 84
seed = 0
lambda_kd_list = [0, 0.1, 0.5, 1, 2, 4, 8, 10]
lambda_disc_list = [0, 0.1, 0.5, 1, 2, 4, 8, 10]
tr_loss = np.zeros((len(lambda_kd_list), len(lambda_disc_list)))
val_loss = np.zeros((len(lambda_kd_list), len(lambda_disc_list)))
tr_acc = np.zeros((len(lambda_kd_list), len(lambda_disc_list)))
val_acc = np.zeros((len(lambda_kd_list), len(lambda_disc_list)))
for i_kd, lambda_kd in enumerate(lambda_kd_list):
for i_disc, lambda_disc in enumerate(lambda_disc_list):
pt_kd, _ = run(n_clients=n_clients, dataset=dataset, model=model, lambda_kd=lambda_kd, lambda_disc=lambda_disc,
reduced=reduced, n_avg=n_avg, rounds=rounds, track_history=track_history,
export_dir=export_dir, device=device, seed=seed)
tr_loss[i_kd, i_disc] = np.array([pt.perf_histories["Train"]["loss"][-1] for pt in pt_kd]).mean()
val_loss[i_kd, i_disc] = np.array([pt.perf_histories["Validation (global)"]["loss"][-1] for pt in pt_kd]).mean()
tr_acc[i_kd, i_disc] = np.array([pt.perf_histories["Train"]["accuracy"][-1] for pt in pt_kd]).mean()
val_acc[i_kd, i_disc] = np.array([pt.perf_histories["Validation (global)"]["accuracy"][-1] for pt in pt_kd]).mean()
# -
import seaborn as sns
import matplotlib
fig, ax = plt.subplots(1,1, figsize=(5,4))
data = (val_acc-val_acc[0,0]) * 100
sns.heatmap(data, cmap="RdYlGn", annot=False, cbar=True, square=True, ax=ax, xticklabels=lambda_disc_list, yticklabels=lambda_kd_list)
ax.set_xlabel("$\lambda_{disc}$", fontsize=16)
ax.set_ylabel("$\lambda_{KD}$", fontsize=16)
ax.add_patch(matplotlib.patches.Rectangle((3, 7), 1.0, 0.9, edgecolor='red', fill=False, lw=2))
fig.savefig("./figures/lambda_ablation.png", bbox_inches='tight')
# # Number of parameters
# +
print("LeNet5 for MNIST")
m_lenet = mdl.LeNet5(1, 84, 10)
hlp.model_size(m_lenet)
print("ResNet9 for FMNIST")
m_rn9 = mdl.ResNet9(1, 128, 10)
hlp.model_size(m_rn9)
print("ResNet18 for MNIST")
m_rn18 = mdl.ResNet18(3, 256, 10)
hlp.model_size(m_rn18)
# -
| Figures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Clustering
# ## Load data
# Four objects are loaded:
# 1. 'data/prepped.pkl' is a pickled dataframe containing cleaned comments and their subreddit label
# 2. LSA, pLSA, and LDA are numpy arrays containing several features (taking the form of a real number) extracted from each comment. These are the versions with comments agglomerated by subreddit.
# +
import numpy as np
import pandas as pd
from time import time
import pickle
print("Importing data...")
t0 = time()
data_comments = pd.read_pickle('data/prepped.pkl')
data = pd.read_pickle('data/prepped_sub.pkl')
LSA = np.load('data/X_LSA_sub.npy')
pLSA = np.load('data/X_pLSA_sub.npy')
LDA = np.load('data/X_LDA_sub.npy')
doc2vec = np.load('data/doc2vec_sub.npy')
print("done in %0.3fs" % (time()-t0))
# assert(data.shape[0] == LSA.shape[0])
# assert(data.shape[0] == pLSA.shape[0])
# assert(data.shape[0] == LDA.shape[0])
# -
# ## Clulstering method evaluation
# Clustering methods are compared for each feature extraction method.
# 1. A memory-efficient KMeans algorithm
# 2. Birch, a memory-efficient, tree-based method
#
# They are compared using the average of the Silhouette scores.
# +
from sklearn.cluster import MiniBatchKMeans, Birch
from sklearn.metrics import silhouette_score
from sklearn.model_selection import KFold
n_samples = data.shape[0]
splits=[]
for split, _ in KFold(n_splits=5).split(range(n_samples)):
splits.append(split)
extractors = [
("LSA", LSA),
("pLSA", pLSA),
("LDA", LDA),
("doc2vec", doc2vec)
]
rec_range = range(5,105,5)
results = {'extractor' : np.array([], dtype='object'),
'n_recommended' : np.array([], dtype='float64'),
'clusterer' : np.array([], dtype='float64'),
'fit_time' : np.array([], dtype='float64'),
'pred_time' : np.array([], dtype='float64'),
'score_time' : np.array([], dtype='float64'),
'score' : np.array([], dtype='float64'),
'fit_time_std' : np.array([], dtype='float64'),
'pred_time_std' : np.array([], dtype='float64'),
'score_time_std' : np.array([], dtype='float64'),
'score_std' : np.array([], dtype='float64'),
}
counts = len(extractors)*20
count = 0
overall_time = time()
for ex_name, X in extractors:
for n_recommended in rec_range:
n_clusters = round(n_samples / n_recommended)
clusterers = [
('kmeans', MiniBatchKMeans(n_clusters=n_clusters)),
('birch', Birch(threshold=0.01, branching_factor=50, n_clusters=n_clusters))
]
for cl_name, clusterer in clusterers:
var1 = []
var2 = []
var3 = []
var4 = []
for ind in splits:
t0 = time()
clusterer.fit(X[ind])
t1 = time()
preds = clusterer.predict(X[ind])
t2 = time()
score = silhouette_score(X[ind], preds, metric='cosine') #, sample_size=round(0.01*n_samples), random_state=0)
t3 = time()
var1.append((t1-t0))
var2.append((t2-t1))
var3.append((t3-t2))
var4.append(score)
results['extractor']= np.append(results['extractor'], ex_name)
results['n_recommended'] = np.append(results['n_recommended'], n_recommended)
results['clusterer'] = np.append(results['clusterer'], cl_name)
results['fit_time'] = np.append(results['fit_time'], np.mean(var1))
results['pred_time'] = np.append(results['pred_time'], np.mean(var2))
results['score_time'] = np.append(results['score_time'], np.mean(var3))
results['score'] = np.append(results['score'], np.mean(var4))
results['fit_time_std'] = np.append(results['fit_time_std'], np.std(var1))
results['pred_time_std'] = np.append(results['pred_time_std'], np.std(var2))
results['score_time_std'] = np.append(results['score_time_std'], np.std(var3))
results['score_std'] = np.append(results['score_std'], np.std(var4))
count = count + 1
# print("Percent complete: %0d, Elapsed time: %0.1f minutes" % (round(count/counts*100), ((time()-overall_time)/60)))
print()
print("Total Elapsed time: %0.1f" % ((time()-overall_time)/60))
# -
continuous, integral, categorical = (results['n_recommended'], results['extractor'], results['clusterer'])
mask = categorical=='birch'
continuous, integral, categorical, score, score_std, fit_time, fit_time_std = (continuous[mask], integral[mask], categorical[mask], results['score'][mask], results['score_std'][mask], results['fit_time'][mask], results['fit_time_std'][mask])
# +
import matplotlib.pyplot as plt
import matplotlib.cm as mplcm
import matplotlib.colors as colors
# %matplotlib inline
plt.ioff()
# For each subplot, define [var, var_std, xlabel, ylabel, title, legend?]
plot_array = [
[score, score_std, "Average size of cluster", "Silhouette Score", "", False], # Showing legends on leftmost plot shows which integral values are present for each categorical value
[fit_time, fit_time_std, "Average size of cluster", "Fit Time in seconds", "", False],
# [results['pred_time'], results['pred_time_std'], "Average size of cluster", "Prediction Time in seconds", "", False],
# [results['score_time'], results['score_time_std'], "Average size of cluster", "Score Time in seconds", "", False],
]
# Set colormap
scalarMap = mplcm.ScalarMappable(norm=colors.Normalize(vmin=-3,
vmax=len(np.unique(integral))),
cmap=plt.get_cmap('BuPu'))
fs = 18
# Define legend for all values of integral
fig_t = plt.figure();
color_ints = {}
for i, lab in enumerate(['LSA','pLSA','LDA','doc2vec']): #np.unique(integral)):
color_ints[lab] = i
plt.plot([],[],label=lab, color=scalarMap.to_rgba(i))
handles, labels = fig_t.axes[0].get_legend_handles_labels()
plt.close(fig_t);
dim_h = len(plot_array) # horizontal number of subplots
dim_v = len(np.unique(categorical)) # vertical number of subplots
p = 0 # subplot count
fig = plt.figure(figsize=[12,6]);
for i in np.unique(categorical):
vals_used = ['LSA','pLSA','LDA','doc2vec'] #np.unique(integral[categorical == i])
for j in range(dim_h):
p = p+1
ax = fig.add_subplot(dim_v, dim_h, p);
for k in vals_used:
mask = np.logical_and(categorical == i, integral == k)
x = np.array(continuous[mask], dtype='float64') # *SearchCV uses dtype 'object'
ind_sort = np.argsort(x) # order by x values for line plot
x = x[ind_sort]
y = plot_array[j][0][mask][ind_sort]
y_stds = plot_array[j][1][mask][ind_sort]
# Show line plot with error bars
_ = plt.errorbar(x, y, yerr=y_stds, label=k, color=scalarMap.to_rgba(color_ints[k]))
# Show confidence interval as shaded region
_ = plt.fill_between(x, y - y_stds, y + y_stds, color=scalarMap.to_rgba(color_ints[k]), alpha=.25)
_ = plt.xlabel(plot_array[j][2], fontsize=fs);
_ = plt.ylabel(plot_array[j][3], fontsize=fs);
# if j==0:
# subtitle=("%s clustering" % i)
# else:
# subtitle=""
# _ = plt.title(subtitle+plot_array[j][4], fontsize=fs);
# if plot_array[j][5]:
# _ = plt.legend(loc='upper left', fontsize=fs);
# Add legends to right of plots
for i in range(int(p/dim_h)):
fig.axes[(i+1)*dim_h-1].legend(handles, labels, loc='center left', bbox_to_anchor=(1, 0.5), title="Method", fontsize=fs)
plt.tight_layout()
plt.show()
# -
# ## Clustering analysis
# +
from sklearn.cluster import MiniBatchKMeans, Birch
from sklearn.metrics import silhouette_score
n_samples = data.shape[0]
n_recommended = 15
n_clusters = round(n_samples / n_recommended)
extractors = [
("LSA", LSA),
("pLSA", pLSA),
("LDA", LDA),
("doc2vec", doc2vec)
]
clusterers = [
("kmeans", MiniBatchKMeans(n_clusters=n_clusters, # 8
init='k-means++',
max_iter=100,
batch_size=100,
verbose=0,
compute_labels=True,
random_state=0, # None
tol=0.0,
max_no_improvement=10,
init_size=(3*n_clusters), # None
n_init=3,
reassignment_ratio=0.01,
) ),
("birch", Birch(threshold=0.05,
branching_factor=50,
n_clusters=n_clusters,
compute_labels=True,
copy=True,
) ),
]
ex_preds = {}
for ex_name, X in extractors:
print(ex_name)
print()
cl_preds = {}
for cl_name, clusterer in clusterers:
print(cl_name)
t0 = time()
clusterer.fit(X)
t1 = time()
preds = clusterer.predict(X)
t2 = time()
score = silhouette_score(X, preds) #, sample_size=round(0.01*n_samples), random_state=0)
t3 = time()
cl_preds[cl_name] = (clusterer, preds, score)
print("Fit time : %0.3f sec" % (t1-t0))
print("Pred time : %0.3f sec" % (t2-t1))
print("Score time: %0.3f sec" % (t3-t2))
print("Score : %0.3f" % score)
print()
ex_preds[ex_name] = cl_preds
# -
# Choose model to analyze
_, preds, _ = ex_preds["LSA"]["birch"]
# ### List subreddits in each cluster
import pprint
for i in range(max(preds)+1):
pprint.pprint(data.iloc[preds==i,1].tolist())
pprint.pprint(data.iloc[preds==56,1].tolist())
# ### Wordclouds and frequency distributions
# +
from wordcloud import WordCloud
import matplotlib.pyplot as plt
max_words = 20
wc = WordCloud(background_color="white", max_words=max_words)
#for i in range(max(preds)+1):
for i in range(56,58):
print("Cluster number {}:".format(i))
print(data.iloc[preds==i,1].tolist())
wc.generate(data[preds==i].body.str.cat(sep=' '))
plt.imshow(wc, interpolation="bilinear")
plt.axis("off")
plt.show()
words = []
freqs = []
for word in wc.words_:
if wc.words_[word]!=0:
words.append(word)
freqs.append(wc.words_[word])
x = range(max_words,0,-1)
plt.barh(x,freqs[:max_words])
plt.yticks(x, np.asarray(words[:max_words]))
plt.show()
# -
# ### tSNE visualization
# +
does not compile
from sklearn.manifold import TSNE
X_embedded = TSNE(n_components=2).fit_transform(X)
indices = range(max(preds)+1)
cmap = plt.get_cmap('jet')
colors = cmap(np.linspace(0, 1.0, len(indices)))
for i, color in zip(indices, colors):
plt.scatter(X_embedded[preds==i,0],X_embedded[preds==i,1],c=color,label=i,alpha=0.5) # 'C{}'.format(i)
# plt.legend()
plt.show()
# -
# ## Approximate Nearest Neighbors
#
# +
from annoy import AnnoyIndex
ann_input = LSA
indx = AnnoyIndex(ann_input.shape[1], metric='angular')
print("Building Annoy index...")
t0 = time()
for i, row in enumerate(ann_input):
indx.add_item(i,row)
indx.build(50)
print("done in %0.3fs" % (time()-t0))
# +
subname = 'Fitness'
test_ix = data.index[data.subreddit==subname].tolist()[0]
n_recommended = 6
print("Given /r/%s, I recommend..." % data.iloc[test_ix,1])
ixs, dists = indx.get_nns_by_item(test_ix, n_recommended, search_k=-1, include_distances=True)
for ix, dist in zip(ixs[1:], dists[1:]):
print(("/r/%s," % data.iloc[ix,1]).ljust(20) + (" at a distance of %0.2f" % dist))
# -
# ## Example of biased unsupervised metric
# +
from sklearn.cluster import KMeans
from sklearn.cluster import Birch
from sklearn.metrics import silhouette_score
coords_1 = np.random.multivariate_normal([0,0], [[1.5,0],[0,1]], 100)
coords_2 = np.random.multivariate_normal([2,2], [[1,0.5],[0.5,1.5]], 100)
coords_3 = np.random.multivariate_normal([-2,3], [[0.8,0],[0,1]], 100)
coords = np.append(np.append(coords_1,coords_2,axis=0),coords_3,axis=0)
truth = np.append(np.zeros([100,]), np.append(np.ones([100,]), 2*np.ones([100])))
truth_scores = []
km_scores = []
bi_scores = []
for _ in range(50):
coords_1 = np.random.multivariate_normal([0,0], [[1.5,0],[0,1]], 100)
coords_2 = np.random.multivariate_normal([2,2], [[1,0.5],[0.5,1.5]], 100)
coords_3 = np.random.multivariate_normal([-2,3], [[0.8,0],[0,1]], 100)
coords = np.append(np.append(coords_1,coords_2,axis=0),coords_3,axis=0)
truth = np.append(np.zeros([100,]), np.append(np.ones([100,]), 2*np.ones([100])))
truth_scores.append( silhouette_score(coords, truth))
km_labels = KMeans(3).fit_predict(coords)
km_scores.append(silhouette_score(coords, km_labels))
bi_labels = Birch(n_clusters=3, threshold=0.05).fit(coords).predict(coords)
bi_scores.append(silhouette_score(coords, bi_labels))
print("Silhouette Scores")
print("Truth : %0.3f +/- %0.3f" % (np.mean(truth_scores), np.std(truth_scores)) )
print("KMeans : %0.3f +/- %0.3f" % (np.mean(km_scores), np.std(km_scores)) )
print("Birch : %0.3f +/- %0.3f" % (np.mean(bi_scores), np.std(bi_scores)) )
fig = plt.figure(figsize=[5,5])
plt.scatter(coords[truth==0,0],coords[truth==0,1],c='C0')
plt.scatter(coords[truth==1,0],coords[truth==1,1],c='C1')
plt.scatter(coords[truth==2,0],coords[truth==2,1],c='C2')
plt.axis('off')
plt.savefig("presentation/figures/clusters.png", bbox_inches='tight')
plt.show()
fig = plt.figure(figsize=[5,5])
plt.scatter(coords[km_labels==1,0],coords[km_labels==1,1],c='C0')
plt.scatter(coords[km_labels==0,0],coords[km_labels==0,1],c='C1')
plt.scatter(coords[km_labels==2,0],coords[km_labels==2,1],c='C2')
plt.axis('off')
plt.show()
fig = plt.figure(figsize=[5,5])
plt.scatter(coords[bi_labels==0,0],coords[bi_labels==0,1],c='tab:blue')
plt.scatter(coords[bi_labels==1,0],coords[bi_labels==1,1],c='tab:green')
plt.scatter(coords[bi_labels==2,0],coords[bi_labels==2,1],c='tab:orange')
plt.axis('off')
plt.show()
# -
# +
from sklearn.cluster import MiniBatchKMeans, Birch
from sklearn.metrics import silhouette_score
from sklearn.model_selection import KFold
n_samples = data.shape[0]
for split, _ in KFold(n_splits=5).split(range(n_samples)):
splits.append(split)
extractors = [
("LSA", LSA),
# ("pLSA", pLSA),
("LDA", LDA),
("doc2vec", doc2vec)
]
results = {}
for ex_name, X in extractors:
for n_recommended in range(5,105,5):
n_clusters = round(n_samples / n_recommended)
clusterers = [
('kmeans', MiniBatchKMeans(n_clusters=n_clusters)),
('birch', Birch(threshold=0.05, branching_factor=50, n_clusters=n_clusters))
]
for cl_name, clusterer in clusterers:
var1 = []
var2 = []
var3 = []
var4 = []
for ind in splits:
t0 = time()
clusterer.fit(X[ind])
t1 = time()
preds = clusterer.predict(X[ind])
t2 = time()
score = silhouette_score(X[ind], preds) #, sample_size=round(0.01*n_samples), random_state=0)
t3 = time()
var1.append((t1-t0))
var2.append((t2-t1))
var3.append((t3-t2))
var4.append(score)
results[ex_name][cl_name]['fit_time'].append(np.mean(var1))
results[ex_name][cl_name]['pred_time'].append(np.mean(var2))
results[ex_name][cl_name]['score_time'].append(np.mean(var3))
results[ex_name][cl_name]['score'].append(np.mean(var4))
results[ex_name][cl_name]['fit_time_std'].append(np.std(var1))
results[ex_name][cl_name]['pred_time_std'].append(np.std(var2))
results[ex_name][cl_name]['score_time_std'].append(np.std(var3))
results[ex_name][cl_name]['score_std'].append(np.std(var4))
| clustering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Duel of sorcerers
# You are witnessing an epic battle between two powerful sorcerers: Gandalf and Saruman. Each sorcerer has 10 spells of variable power in their mind and they are going to throw them one after the other. The winner of the duel will be the one who wins more of those clashes between spells. Spells are represented as a list of 10 integers whose value equals the power of the spell.
# ```
# gandalf = [10, 11, 13, 30, 22, 11, 10, 33, 22, 22]
# saruman = [23, 66, 12, 43, 12, 10, 44, 23, 12, 17]
# ```
# For example:
# 1. The first clash is won by Saruman: 10 against 23, 23 wins
# 2. The second clash is also won by Saruman: 11 against 66, 66 wins
# 3. etc.
#
#
# You will create two variables, one for each sorcerer, where the sum of clashes won will be stored. Depending on which variable is greater at the end of the duel, you will show one of the following three results on the screen:
# * Gandalf wins
# * Saruman wins
# * Tie
#
# <img src="images/content_lightning_bolt_big.jpg" width="400">
# ## Solution
# +
# Assign spell power lists to variables
gandalf = [10, 11, 13, 30, 22, 11, 10, 33, 22, 22]
saruman = [23, 66, 12, 43, 12, 10, 44, 23, 12, 17]
# -
# Assign 0 to each variable that stores the victories
# Execution of spell clashes
# +
# We check who has won, do not forget the possibility of a draw, in which case, no one wins.
# Print the result based on the winner.
# -
# ## Goals
#
# 1. Treatment of lists
# 2. Use of **for loop**
# 3. Use of conditional **if-elif-else**
# 4. Use of the functions **range(), len()**
# 5. Print
# ## Bonus
#
# 1. Spells now have a name and there is a dictionary that relates that name to a power.
# 2. A sorcerer wins if he succeeds in winning 3 spell clashes in a row.
# 3. Average of each of the spell lists.
# 4. Standard deviation of each of the spell lists.
#
# ```
# POWER = {
# 'Fireball': 50,
# 'Lightning bolt': 40,
# 'Magic arrow': 10,
# 'Black Tentacles': 25,
# 'Contagion': 45
# }
#
# gandalf = ['Fireball', 'Lightning bolt', 'Lightning bolt', 'Magic arrow', 'Fireball',
# 'Magic arrow', 'Lightning bolt', 'Fireball', 'Fireball', 'Fireball']
# saruman = ['Contagion', 'Contagion', 'Black Tentacles', 'Fireball', 'Black Tentacles',
# 'Lightning bolt', 'Magic arrow', 'Contagion', 'Magic arrow', 'Magic arrow']
# ```
#
# Good luck!
# +
# 1. Spells now have a name and there is a dictionary that relates that name to a power.
# variables
POWER = {
'Fireball': 50,
'Lightning bolt': 40,
'Magic arrow': 10,
'Black Tentacles': 25,
'Contagion': 45
}
gandalf = ['Fireball', 'Lightning bolt', 'Lightning bolt', 'Magic arrow', 'Fireball',
'Magic arrow', 'Lightning bolt', 'Fireball', 'Magic arrow', 'Fireball']
saruman = ['Contagion', 'Contagion', 'Black Tentacles', 'Fireball', 'Black Tentacles',
'Lightning bolt', 'Magic arrow', 'Contagion', 'Magic arrow', 'Magic arrow']
# +
# Assign spell power lists to variables
# +
# 2. A sorcerer wins if he succeeds in winning 3 spell clashes in a row.
# Execution of spell clashes
# check for 3 wins in a row
# check the winner
# +
# 3. Average of each of the spell lists.
# +
# 4. Standard deviation of each of the spell lists.
# -
| duel/duel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="1-TOAriPHRIu"
# > Note: KNN is a memory-based model, that means it will memorize the patterns and not generalize. It is simple yet powerful technique and compete with SOTA models like BERT4Rec.
# + id="xWTTsFsu3idp" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1628096233726, "user_tz": -330, "elapsed": 2459, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="31f3b304-3368-405c-9aba-0a9309c20037"
import os
project_name = "reco-tut-itr"; branch = "main"; account = "sparsh-ai"
project_path = os.path.join('/content', project_name)
if not os.path.exists(project_path):
# !cp /content/drive/MyDrive/mykeys.py /content
import mykeys
# !rm /content/mykeys.py
path = "/content/" + project_name;
# !mkdir "{path}"
# %cd "{path}"
import sys; sys.path.append(path)
# !git config --global user.email "<EMAIL>"
# !git config --global user.name "reco-tut"
# !git init
# !git remote add origin https://"{mykeys.git_token}":x-oauth-basic@github.com/"{account}"/"{project_name}".git
# !git pull origin "{branch}"
# !git checkout main
else:
# %cd "{project_path}"
# + id="fZtYfVlgGURe" executionInfo={"status": "ok", "timestamp": 1628096259909, "user_tz": -330, "elapsed": 397, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
import os
import numpy as np
import pandas as pd
import scipy.sparse
from scipy.spatial.distance import correlation
# + colab={"base_uri": "https://localhost:8080/"} id="f6uglOI8Gb-V" executionInfo={"status": "ok", "timestamp": 1628096800010, "user_tz": -330, "elapsed": 454, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="7be89cd3-b007-4049-9c15-a2cd274059ea"
df = pd.read_parquet('./data/silver/rating.parquet.gz')
df.info()
# + colab={"base_uri": "https://localhost:8080/"} id="e0oKCHseIkHG" executionInfo={"status": "ok", "timestamp": 1628096861155, "user_tz": -330, "elapsed": 417, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="b540c2af-c662-4729-b8c7-5209e400b042"
df2 = pd.read_parquet('./data/silver/items.parquet.gz')
df2.info()
# + colab={"base_uri": "https://localhost:8080/"} id="FBkKJvjPIp2q" executionInfo={"status": "ok", "timestamp": 1628096888907, "user_tz": -330, "elapsed": 444, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="8a2c4297-707c-4ac0-e202-edf7a8e630fb"
df = pd.merge(df, df2, on='itemId')
df.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 439} id="e4XoIzO8GWIE" executionInfo={"status": "ok", "timestamp": 1628096367203, "user_tz": -330, "elapsed": 675, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="90ade036-d416-4ba9-d6c9-1ab3dbfb7777"
rating_matrix = pd.pivot_table(df, values='rating',
index=['userId'], columns=['itemId'])
rating_matrix
# + id="QWfR5ZHwGwVI" executionInfo={"status": "ok", "timestamp": 1628096380422, "user_tz": -330, "elapsed": 396, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
def similarity(user1, user2):
try:
user1=np.array(user1)-np.nanmean(user1)
user2=np.array(user2)-np.nanmean(user2)
commonItemIds=[i for i in range(len(user1)) if user1[i]>0 and user2[i]>0]
if len(commonItemIds)==0:
return 0
else:
user1=np.array([user1[i] for i in commonItemIds])
user2=np.array([user2[i] for i in commonItemIds])
return correlation(user1,user2)
except ZeroDivisionError:
print("You can't divide by zero!")
# + id="24XPWqs7G0I_" executionInfo={"status": "ok", "timestamp": 1628097264660, "user_tz": -330, "elapsed": 470, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
def nearestNeighbourRatings(activeUser, K):
try:
similarityMatrix=pd.DataFrame(index=rating_matrix.index,columns=['Similarity'])
for i in rating_matrix.index:
similarityMatrix.loc[i]=similarity(rating_matrix.loc[activeUser],rating_matrix.loc[i])
similarityMatrix=pd.DataFrame.sort_values(similarityMatrix,['Similarity'],ascending=[0])
nearestNeighbours=similarityMatrix[:K]
neighbourItemRatings=rating_matrix.loc[nearestNeighbours.index]
predictItemRating=pd.DataFrame(index=rating_matrix.columns, columns=['Rating'])
for i in rating_matrix.columns:
predictedRating=np.nanmean(rating_matrix.loc[activeUser])
for j in neighbourItemRatings.index:
if rating_matrix.loc[j,i]>0:
predictedRating += (rating_matrix.loc[j,i]-np.nanmean(rating_matrix.loc[j]))*nearestNeighbours.loc[j,'Similarity']
predictItemRating.loc[i,'Rating']=predictedRating
except ZeroDivisionError:
print("You can't divide by zero!")
return predictItemRating
# + id="HA09XRRfHOZ5" executionInfo={"status": "ok", "timestamp": 1628097419122, "user_tz": -330, "elapsed": 384, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
def topNRecommendations(activeUser, N):
try:
predictItemRating = nearestNeighbourRatings(activeUser,N)
placeAlreadyWatched = list(rating_matrix.loc[activeUser].loc[rating_matrix.loc[activeUser]>0].index)
predictItemRating = predictItemRating.drop(placeAlreadyWatched)
topRecommendations = pd.DataFrame.sort_values(predictItemRating,['Rating'],ascending = [0])[:N]
topRecommendationTitles = (df.loc[df.itemId.isin(topRecommendations.index)])
except ZeroDivisionError:
print("You can't divide by zero!")
return list([topRecommendationTitles.location,
topRecommendationTitles.place,
topRecommendationTitles.state,
topRecommendationTitles.location_rating])
# + id="xp-VaE0pIHIl" executionInfo={"status": "ok", "timestamp": 1628097633307, "user_tz": -330, "elapsed": 1651, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
def favoritePlace(activeUser,N):
topPlace=pd.DataFrame.sort_values(df[df.userId==activeUser],['rating'],ascending=[0])[:N]
return list([topPlace.location,
topPlace.place,
topPlace.state,
topPlace.location_rating])
# + id="w_klxgDlH7o4" executionInfo={"status": "ok", "timestamp": 1628097422831, "user_tz": -330, "elapsed": 4, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}}
activeUser = 4
# + colab={"base_uri": "https://localhost:8080/", "height": 190} id="KMj2yGvtLk7n" executionInfo={"status": "ok", "timestamp": 1628097662575, "user_tz": -330, "elapsed": 458, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="5195a7f3-4b8d-4fd0-e59a-b48a788dd649"
print("Your favorite places are: ")
fav_place=pd.DataFrame(favoritePlace(str(activeUser),4))
fav_place=fav_place.T
fav_place=fav_place.sort_values(by='location_rating', ascending=False)
fav_place
# + colab={"base_uri": "https://localhost:8080/", "height": 190} id="ZyMlJdYIH9dB" executionInfo={"status": "ok", "timestamp": 1628097678985, "user_tz": -330, "elapsed": 488, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "13037694610922482904"}} outputId="98c5b7ff-b122-4402-dc5f-3ca3c4ecd19d"
print("The recommended places for you are: ")
topN = pd.DataFrame(topNRecommendations(str(activeUser), 4))
topN = topN.T
topN = topN.sort_values(by = 'location_rating', ascending=False).drop_duplicates().reset_index(drop=True)
topN
| notebooks/reco-tut-itr-03-modeling-collaborative-knn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Purpose: To implement the code for Stable Count Thresholding Algorithm
# +
import matplotlib.pyplot as plt
import matplotlib.cm as cm
import numpy as np
from skimage import io
from skimage import filters
# -
# *Defining the File Location*
file_location = '../../qdbev_lectin.tif'
# *Reading in the image as a numpy array*
im = io.imread(file_location)
im.shape
# *Splitting the color channels of the image*
cell_ims = im[0::2]
qd_ims = im[1::2]
# ## Testing for the cell_ims
# *Creating a numpy array for Threshold values & Defining Delta*
# +
#Defining delta according to the number provided by the paper
i = 0
delta = 1
T_array = np.arange(2*delta, 255, delta)
WT_array = np.zeros(256)
#initializing variables to decrease the processing for the rest of the for loop
im_binary_low = (cell_ims >= 0).astype(np.int_)
im_binary = (cell_ims >= delta).astype(np.int_)
im_binary_high = (cell_ims >= (2*delta)).astype(np.int_)
# +
#Calculating the first WT:
NT_minus_delta = np.count_nonzero(im_binary_low)
NT = np.count_nonzero(im_binary)
NT_plus_delta = np.count_nonzero(im_binary_high)
WT = ((NT_minus_delta - 2 * NT + NT_plus_delta)/(delta**2))
WT_array[i] = WT
# -
# *Current code for quickly binarizing the 3D image and getting the count of non-zero values*
im_binary_low = im_binary
im_binary = im_binary_high
i = 1
for T in T_array:
#getting a binary image
im_binary_high = (cell_ims >= (T + delta)).astype(np.int_)
NT_minus_delta = np.count_nonzero(im_binary_low)
NT = np.count_nonzero(im_binary)
NT_plus_delta = np.count_nonzero(im_binary_high)
WT_plus = ((NT_minus_delta - 2 * NT + NT_plus_delta)/(delta**2))
WT_array[i] = WT_plus
#increased the alpha because I think that we have bigger pictures than those used in the paper
alpha = NT/20000
if WT_plus < alpha:
if abs(WT - WT_plus) in range(0,10):
print(T - delta, abs(WT-WT_plus))
else:
pass
else:
pass
#Resetting variable names for next loop
im_binary_low = im_binary
im_binary = im_binary_high
WT = WT_plus
i += 1
cell_hist = plt.hist(cell_ims[0].ravel(), bins=256, range = (0, 256), fc='k', ec='k')
plt.imshow(cell_ims[3])
# +
plt.imshow(cell_ims[5], cmap = 'gray', clim=(185,256))
plt.title('First Cell Image')
plt.xticks([])
plt.yticks([])
plt.show()
# -
IM_MAX= np.max(cell_ims, axis=0)
plt.imshow(IM_MAX)
plt.imshow(IM_MAX, clim=(185,256))
# ## Testing for the qd ims
# +
#Defining delta according to the number provided by the paper
i = 0
delta = 1
T_array = np.arange(2*delta, 255, delta)
WT_array = np.zeros(256)
#initializing variables to decrease the processing for the rest of the for loop
im_binary_low = (qd_ims >= 0).astype(np.int_)
im_binary = (qd_ims >= delta).astype(np.int_)
im_binary_high = (qd_ims >= (2*delta)).astype(np.int_)
# +
#Calculating the first WT:
NT_minus_delta = np.count_nonzero(im_binary_low)
NT = np.count_nonzero(im_binary)
NT_plus_delta = np.count_nonzero(im_binary_high)
WT = ((NT_minus_delta - 2 * NT + NT_plus_delta)/(delta**2))
WT_array[i] = WT
# -
# *Current code for quickly binarizing the 3D image and getting the count of non-zero values*
im_binary_low = im_binary
im_binary = im_binary_high
i = 1
for T in T_array:
#getting a binary image
im_binary_high = (qd_ims >= (T + delta)).astype(np.int_)
NT_minus_delta = np.count_nonzero(im_binary_low)
NT = np.count_nonzero(im_binary)
NT_plus_delta = np.count_nonzero(im_binary_high)
WT_plus = ((NT_minus_delta - 2 * NT + NT_plus_delta)/(delta**2))
WT_array[i] = WT_plus
#increased the alpha because I think that we have bigger pictures than those used in the paper
alpha = NT/20000
if WT_plus < alpha:
if abs(WT - WT_plus) in range(0,10):
print(T - delta, abs(WT-WT_plus))
else:
pass
else:
pass
#Resetting variable names for next loop
im_binary_low = im_binary
im_binary = im_binary_high
WT = WT_plus
i += 1
# +
fig = plt.figure()
ax1 = fig.add_subplot(1,1,1)
ax1.imshow(qd_ims[5], clim=(106, 256))
ax1.set_xticks([])
ax1.set_yticks([])
ax1.set_title('QD ims')
fig.set_size_inches(15, 15, forward=True)
# -
IM_MAX_QD = np.max(qd_ims, axis=0)
# +
fig = plt.figure()
ax1 = fig.add_subplot(1,1,1)
ax1.imshow(IM_MAX_QD, clim=(106, 256))
ax1.set_xticks([])
ax1.set_yticks([])
ax1.set_title('QD ims')
fig.set_size_inches(15, 15, forward=True)
# +
fig = plt.figure()
ax1 = fig.add_subplot(1,1,1)
ax1.imshow(IM_MAX_QD)
ax1.set_xticks([])
ax1.set_yticks([])
ax1.set_title('QD ims')
fig.set_size_inches(10, 12, forward=True)
# -
| scripts/SCT_thresholding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Find if a string is made of unique characters
# ## hashmap
#
string='holaa'
# +
#def isUnique(string):
mapa={}
for word in string:
if word not in mapa:
mapa[word]=1
else:
mapa[word]+=1
# -
mapa
for i in mapa.values():
if i>1:
print('False')
print('True')
for keys in mapa:
print(keys)
string='mundo'
def isUnique(string):
mapa={}
for word in string:
if word not in mapa:
mapa[word]=1
else:
mapa[word]+=1
for i in mapa.values():
if i>1:
return False
break
return True
isUnique(string)
def isUnique2(string):
mapa={}
for i in string:
if i not in mapa:
mapa[i]=1
else:
break
return False
return True
isUnique2(string)
# ## O(n)
# ### bueno es n+m (tamaño del string+ numero de caracteres)
# # check permutation
string1='abc'
string2='cba'
string3='a cb'
def crearMapa(texto):
mapa_palabras={}
for word in texto:
if word not in mapa_palabras:
mapa_palabras[word]=1
else:
mapa_palabras[word]+=1
return mapa_palabras
# +
def check_permutation(texto1,texto2):
if len(texto1)!=len(texto2):
return False
mapaTexto1=crearMapa(texto1)
mapaTexto2=crearMapa(texto2)
for key in mapaTexto1:
if key not in mapaTexto2 or mapaTexto1[key]!=mapaTexto2[key]:
return False
return True
# -
check_permutation(string1,string3)
# ### run time es O(n+n) osea O(n) peor escenario
# ## URLfy
input1='<NAME> '
# +
resultado=''
for word in input1:
if word==' ':
word='%20'
resultado+=word
print(resultado)
# -
input1.replace(' ','%20')[0:17]
input1='<NAME> '
length=13
def urlify(input1,length):
resultado=''
count=0
for word in input1:
if word==' ':
word='%20'
resultado+=word
count+=1
if count==length:
break
return resultado
return resultado
urlify(input1,length)
# ### resultado en O(n)
#
# # Palindrome Permutation
#
# se busca que todos los elementos del string sean pares, solo se permite un elemento impar.
def PalindromePermutation(string):
string=string.replace(' ','')
string=string.lower()
mapa={}
for i in string:
if i not in mapa:
mapa[i]=1
else:
mapa[i]+=1
odd_count=0
for key, value in mapa.items():
if value%2!=0 and odd_count==0:
odd_count+=1
elif value%2!=0 and odd_count!=0:
return False
return True
inputPalindrome="Tact Coa"
PalindromePermutation(inputPalindrome)
# ### O(n) time and memory
# ## One Way
input1='pale'
input2='ale'
input3='pales'
input4='bake'
# cuando son del mismo largo
def oneWaySame(input1,input2):
flag=0
for i in range(len(input2)):
if input1[i]!=input2[i]:
flag+=1
if flag>1:
return False
else:
return True
oneWaySame(input1,input4)
# cuando son de diferente tamaño
def oneWayDiff(input1,input2):
flag=0
for i in range(len(input2)):
if input1[i+flag]!=input2[i]:
flag+=1
if flag>1:
return False
else:
return True
oneWayDiff(input1,input2)
def oneWay(string1,string2):
if len(string1) <=(len(string2)-2) or len(string1)>=(len(string2)+2):
return False
elif len(string1)==len(string2):
return oneWaySame(string1,string2)
elif len(string1)>len(string2):
return oneWayDiff(string1,string2)
else:
return oneWayDiff(string2,string1)
oneWay(input1,input4)
oneWay(input1,input3)
# ### runtime O(n)
# # String Compression
string1='aabcccccaaa'
def compress(string1):
S = len(string1)
result = []
if S < 2:
return string1
c = 1
for i in range(S):
# First pass
if not result:
result.append(string1[i])
continue
if result[-1] == string1[i]:
c += 1
else:
result.append(str(c))
result.append(string1[i])
c = 1
# Last pass
if S-1 == i:
result.append(str(c))
# Short-circuit if compression ends up bigger
if S < len(result):
return string1
return ''.join(result)
compress(string1)
# runtime O(n) peor escenario
# # rotate a matrix in place
import numpy as np
A=[[1,2,3],[4,5,6],[7,8,9]]
len(A)
def rotationInPlace(A):
#encontrar el largo de la matriz
n=len(A)
# recorrer por bloques la matriz
# dividir 4 partes a partir del centro
# encontrar el centro n/2 redondeo al entero proximo ceil
for layer in range(int(np.ceil(n/2))):
# ancho del bloque a modificar
first=layer
last=n-1-layer
for i in range(first,last):
# para iniciar desde [0,0] quitamos un offset
offset=i-first
#variable temporal guardar el primer elemento del bloque
top=A[first][i]
# manda la izquierda del bloque al primer elemento
#left->top
A[first][i]=A[last-offset][first]
#bottom->left
# manda el elemento a la izquierda al fondo
A[last-offset][first]=A[last][last-offset]
# right-> bottom
# manda elemento de la derecha a fondo
A[last][last-offset]=A[i][last]
# top->right
# elemento top a la derecha
A[i][last]=top
def rotatatev2(matrix):
matrix=matrix[::-1]
n=len(matrix)
for i in range(n):
for j in range(i):
matrix[i][j],matrix[j][i]=matrix[j][i],matrix[i][j]
return matrix
rotationInPlace(A)
A
A=[[1,2,3],[4,5,6],[7,8,9]]
rotatatev2(A)
A[::-1]
A
# runtime O(n^2)
# # zero Matrix
B=[[1,2,3],[4,5,0],[7,8,9]]
C=[[1,2,3],[4,5,0],[7,8,9],[7,8,9]]
B
C
len(C[0])
def MatrixZero2(matrix):
M=len(matrix[:])
N=len(matrix[0])
for i in range(M):
for j in range(N):
if matrix[i][j]==0:
row=i
col=j
break
for l in range(N):
matrix[row][l]=0
for k in range(M):
matrix[k][col]=0
return matrix
MatrixZero2(C)
# +
def MatrixZero(B):
M,N=np.shape(B)
for i in range(M):
for j in range(N):
if B[i][j]==0:
c1=i
c2=j
break
for l in range(N):
B[c1][l]=0
for k in range(M):
B[k][c2]=0
# -
MatrixZero(B)
B
B=[[0,2,3],[4,5,0],[7,8,9]]
# +
c1=[]
c2=[]
M,N=np.shape(B)
for i in range(M):
for j in range(N):
if B[i][j]==0:
c1.append(i)
c2.append(j)
#for l in range(N):
for row in c1:
for l in range(N):
B[row][l]=0
for col in c2:
for k in range(M):
B[k][col]=0
# -
B
# no estoy seguro creo que es O(n) pero podria ser O(n^2)
# # String Rotation
#
original='waterbottle'
rotate='erbottlewat'
false_rotate='esbottlewat'
x=original[:3]
y=original[3:]
def isSubstring(original,substring):
x=original[:3]
y=original[3:]
for i in range(len(y)):
if y[i]!=substring[i]:
return False
return True
isSubstring(original,rotate)
s1 = "barfoo"
s2 = "foobar"
isSubstring(s2,s1)
# run time O(n)
def is_rotation(str1, str2):
return(len(str1) == len(str2) and str1 in str2*2)
is_rotation(original,rotate)
is_rotation(original,false_rotate)
def is_rotation_v2(string1,string2):
if len(string1)!=len(string2):
return False
for index in range(len(string1)):
if string2.startswith(string1[index:]) and string2.endswith(string1[:index]):
return True
return False
is_rotation_v2(original,rotate)
is_rotation_v2(original,false_rotate)
# O(n) en el caso de is_rotation_v2
# ## version estilo python
def isSubstring2(s2,s1):
s1=s1+s1
return s2 in s1
isSubstring2(original,rotate)
isSubstring2(s2,s1)
| CTCI/c1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.3 64-bit
# language: python
# name: python37364bit4a0e5dd9ae1a4de69f3c061dd783db34
# ---
# Modify this line to briefly discribe the functionality of new_script.ipynb<br/><br/>Copyright (C) 2017 <NAME> Lab<br/>This program is free software: you can redistribute it and/or modify<br/>it under the terms of the GNU General Public License as published by<br/>the Free Software Foundation, either version 3 of the License, or<br/>(at your option) any later version.<br/>This program is distributed in the hope that it will be useful,<br/>but WITHOUT ANY WARRANTY; without even the implied warranty of<br/>MERCHANTABILITY or FITNESS FOR A PARTICULAR PURPOSE. See the<br/>GNU General Public License for more details.<br/>You should have received a copy of the GNU General Public License<br/>along with this program. If not, see <http://www.gnu.org/licenses/>.
# +
import os
from dotenv import load_dotenv, find_dotenv
from os.path import join, dirname, basename, exists, isdir
### Load environmental variables from the project root directory ###
# find .env automagically by walking up directories until it's found
dotenv_path = find_dotenv()
# load up the entries as environment variables
load_dotenv(dotenv_path)
# now you can get the variables using their names
# Check whether a network drive has been specified
DATABASE = os.environ.get("NETWORK_URL")
if DATABASE == 'None':
pass
else:
pass
#mount network drive here
# set up directory paths
CURRENT_DIR = os.getcwd()
PROJ = dirname(dotenv_path) # project root directory
DATA = join(PROJ, 'data') #data directory
RAW_EXTERNAL = join(DATA, 'raw_external') # external data raw directory
RAW_INTERNAL = join(DATA, 'raw_internal') # internal data raw directory
INTERMEDIATE = join(DATA, 'intermediate') # intermediate data directory
FINAL = join(DATA, 'final') # final data directory
RESULTS = join(PROJ, 'results') # output directory
FIGURES = join(RESULTS, 'figures') # figure output directory
PICTURES = join(RESULTS, 'pictures') # picture output directory
# make folders specific for certain data
folder_name = ''
if folder_name != '':
#make folders if they don't exist
if not exists(join(RAW_EXTERNAL, folder_name)):
os.makedirs(join(RAW_EXTERNAL, folder_name))
if not exists(join(INTERMEDIATE, folder_name)):
os.makedirs(join(INTERMEDIATE, folder_name))
if not exists(join(FINAL, folder_name)):
os.makedirs(join(FINAL, folder_name))
print('Standard variables loaded, you are good to go!')
# -
# # Read in data
# here, data is read in as well as translations from uniprot ID to NCBI Entrez (mapper obtained from: https://www.uniprot.org/uploadlists/)
# +
## Your code here ##
import pandas as pd
# de results loading
#de_results = pd.read_csv("~/Documents/masters/thesis/thesis/data/final/de_edger_results_stress.csv")
# protein to gene translation file read in
uniprot_to_entrez = pd.read_csv("~/Documents/masters/thesis/thesis/data/intermediate/uniprot_to_entrez.tsv", sep="\t")
uniprot_to_entrez = uniprot_to_entrez.drop_duplicates(subset ="From", keep = False)\
.set_index("From").to_dict('index')
# -
uniprot_to_entrez
# de results translated to gene IDs
de_results_entrez = de_results
de_results_entrez["Unnamed: 0"] = [uniprot_to_entrez[i]['To'] if i in uniprot_to_entrez else "nan" for i in de_results_entrez["Unnamed: 0"]]
de_results_entrez = de_results_entrez[de_results_entrez["Unnamed: 0"] != "nan"]
de_results_ids = [list(de_results[de_results[i] != 0]["Unnamed: 0"]) for i in de_results.columns]
# +
# Get ontologies (from http://geneontology.org/ontology/go-basic.obo)
from goatools.base import download_go_basic_obo
from goatools.obo_parser import GODag
import os
obo_fname = download_go_basic_obo() # Download data
obodag = GODag(obo_fname) # Load data
# Get associations genes <-> ontologies (from ftp://ftp.ncbi.nlm.nih.gov/gene/DATA/gene2go.gz)
from goatools.base import download_ncbi_associations
from goatools.anno.genetogo_reader import Gene2GoReader
gene2go_fname = download_ncbi_associations() # Download data
objanno = Gene2GoReader(gene2go_fname, taxids=[511145]) # Read data (only for e. coli)
ns2assoc = objanno.get_ns2assc()
#Remove downloaded data:
os.remove(obo_fname)
os.remove(gene2go_fname)
# -
# # Read in translation from gene to go
#
# chosen taxid E. coli corresponds to Escherichia coli str. K-12 substr. MG1655, as found here: https://www.ncbi.nlm.nih.gov/Taxonomy/Browser/wwwtax.cgi?mode=Info&id=511145&lvl=3&lin=f&keep=1&srchmode=1&unlock
#
# +
from __future__ import print_function
from goatools.anno.genetogo_reader import Gene2GoReader
for nspc, id2gos in ns2assoc.items():
print("{NS} {N:,} annotated e. coli genes".format(NS=nspc, N=len(id2gos)))
# +
# Over-representation object:
from goatools.goea.go_enrichment_ns import GOEnrichmentStudyNS
population = de_results_entrez["Unnamed: 0"]
goeaobj = GOEnrichmentStudyNS(
population, # Population (all mapped yeast genes in Entrez format)
ns2assoc, # geneid/GO associations
obodag, # Ontologies
propagate_counts = False, # ???
alpha = 0.05, # default significance cut-off
methods = ['fdr_bh']) # default multipletest correction method
# -
# # Run Gene ontology
# run gene ontology enrichment on the top genes contained within the
# +
# %%capture
# 'p_' means "pvalue". 'fdr_bh' is the multipletest method we are currently using.
# genes to be analysed
geneids_study = list(de_results_entrez["Unnamed: 0"][0:400])
goea_results_sig = []
for i in de_results_ids:
goea_results_all = goeaobj.run_study(i)
goea_results_sig.append([r for r in goea_results_all if r.p_fdr_bh < 0.05])
# +
from goatools.godag_plot import plot_gos, plot_results, plot_goid2goobj
# goeaobj.wr_tsv(join(FINAL, "p_values_" + group_name + ".tsv"), goea_results_sig)
# save results of goe analysis
j = 0
for i in de_results.columns[1:]:
goeaobj.wr_tsv("/Users/jonas/Documents/masters/thesis/thesis/data/final/go_analysis/p_values_go_{}.tsv".format(i), goea_results_sig[j])
j += 1
# +
# plot_results("/Users/jonas/Documents/masters/thesis/thesis/results/figures/nbt3102_{NS}.png", goea_results_sig)
| data_science/code/go_term_analysis/go_term_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from matplotlib.ticker import PercentFormatter
def reformat_google(file):
# Get file
filepath = '../files/'
fileloc = filepath + file
# Read csv and parse dates
df = pd.read_csv(fileloc, header=1, parse_dates=[0])
df.columns=['date', 'search']
# Get ratio to pre-2020 mean
pre_2020_mean = df.loc[df.date.dt.year != 2020].search.mean()
df.search /= pre_2020_mean
# Get month by month for seaborn's 95% confidence interval
df.date = df.date.apply(lambda x: x.strftime('%Y-%m'))
return df
cond = reformat_google('condolences_5_15.csv')
short = reformat_google('shortness_5_15.csv')
push = reformat_google('pushups_5_15.csv')
mask = reformat_google('mask_5_15.csv')
assert cond.shape == short.shape == push.shape == mask.shape
# +
plt.style.use('default')
sns.lineplot(x='date', y='search', data=cond, label='condolences')
sns.lineplot(x='date', y='search', data=mask, label='mask')
sns.lineplot(x='date', y='search', data=short, label='shortness of breath')
sns.lineplot(x='date', y='search', data=push, label='pushups')
# Start from 12th month and show every 2 months
plt.xticks(list(range(12, cond.date.nunique(), 2)))
plt.xlim(12, cond.date.nunique() - 1)
plt.xlabel('Date')
plt.ylabel('Ratio to pre-2020 average')
plt.legend()
plt.title('Frequency of COVID-19 related Google searches in NYC')
plt.grid(0.4)
# plt.tight_layout()
plt.figtext(0.05, 0.01, 'Error bars give 95% confidence intervals for each month.',
horizontalalignment='left', fontsize=7)
plt.savefig('plots/covid_searches.png', dpi=300)
plt.show()
# -
| condolences.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import turicreate as tc
import os
# Change if applicable
ig02_path = os.getcwd()
# Load all images in random order
raw_sf = tc.image_analysis.load_images(ig02_path, recursive=True,
random_order=True)
# Split file names so that we can determine what kind of image each row is
# E.g. bike_005.mask.0.png -> ['bike_005', 'mask']
info = raw_sf['path'].apply(lambda path: os.path.basename(path).split('.')[:2])
# Rename columns to 'name' and 'type'
info = info.unpack().rename({'X.0': 'name', 'X.1': 'type'})
# Add to our main SFrame
raw_sf = raw_sf.add_columns(info)
# Extract label (e.g. 'bike') from name (e.g. 'bike_003')
raw_sf['label'] = raw_sf['name'].apply(lambda name: name.split('_')[0])
# Original path no longer needed
del raw_sf['path']
# Split into images and masks
sf_images = raw_sf[raw_sf['type'] == 'image']
sf_masks = raw_sf[raw_sf['type'] == 'mask']
def mask_to_bbox_coordinates(img):
"""
Takes a tc.Image of a mask and returns a dictionary representing bounding
box coordinates: e.g. {'x': 100, 'y': 120, 'width': 80, 'height': 120}
"""
import numpy as np
mask = img.pixel_data
if mask.max() == 0:
return None
# Take max along both x and y axis, and find first and last non-zero value
x0, x1 = np.where(mask.max(0))[0][[0, -1]]
y0, y1 = np.where(mask.max(1))[0][[0, -1]]
return {'x': (x0 + x1) / 2, 'width': (x1 - x0),
'y': (y0 + y1) / 2, 'height': (y1 - y0)}
# Convert masks to bounding boxes (drop masks that did not contain bounding box)
sf_masks['coordinates'] = sf_masks['image'].apply(mask_to_bbox_coordinates)
# +
# There can be empty masks (which returns None), so let's get rid of those
sf_masks = sf_masks.dropna('coordinates')
# Combine label and coordinates into a bounding box dictionary
sf_masks = sf_masks.pack_columns(['label', 'coordinates'],
new_column_name='bbox', dtype=dict)
# Combine bounding boxes of the same 'name' into lists
sf_annotations = sf_masks.groupby('name',
{'annotations': tc.aggregate.CONCAT('bbox')})
# Join annotations with the images. Note, some images do not have annotations,
# but we still want to keep them in the dataset. This is why it is important to
# a LEFT join.
sf = sf_images.join(sf_annotations, on='name', how='left')
# The LEFT join fills missing matches with None, so we replace these with empty
# lists instead using fillna.
sf['annotations'] = sf['annotations'].fillna([])
# Remove unnecessary columns
del sf['type']
# Save SFrame
sf.save('ig02.sframe')
# -
import turicreate as tc
# Load the data
data = tc.SFrame('ig02.sframe')
# Make a train-test split
train_data, test_data = data.random_split(0.8)
import sys
#sys.stdout = open('test.txt', 'w')
# Create a model
model = tc.object_detector.create(train_data, batch_size= 12, max_iterations= 10, verbose=True, all_iterations=True)
#sys.stdout.close()
import sys
#sys.stdout = open('test.txt', 'w')
# Create a model
model1 = tc.object_detector.create(train_data, batch_size= 12, max_iterations= 10, verbose=True, all_iterations=False)
#sys.stdout.close()
#all iterations turned off prints occasionally
import sys
#sys.stdout = open('test.txt', 'w')
# Create a model
model = tc.object_detector.create(train_data, batch_size= 12, max_iterations= 20, verbose=True, all_iterations=True)
#sys.stdout.close()
import sys
#sys.stdout = open('test.txt', 'w')
# Create a model
model1 = tc.object_detector.create(train_data, batch_size= 12, max_iterations= 20, verbose=True, all_iterations=False)
#sys.stdout.close()
import sys
#sys.stdout = open('test.txt', 'w')
# Create a model
model = tc.object_detector.create(train_data, batch_size= 12, max_iterations= 20, verbose=False, all_iterations=True)
#sys.stdout.close()
# Save predictions to an SArray
predictions = model.predict(test_data)
# Evaluate the model and save the results into a dictionary
metrics = model.evaluate(test_data)
# Save the model for later use in Turi Create
model.save('mymodel.model')
model.summary()
model.summary(output='dict')
metrics.summary()
metrics
scores = model.evaluate(data)
# +
sf = tc.SFrame('test.sframe')
# -
docs = docs.dict_trim_by_keys(turicreate.text_analytics.stopwords(), exclude=True)
sa = tc.SArray('test.txt')
sa
# +
import re
import pandas as pd
with open('test.txt') as f, open('outfile.csv', 'w') as outfile:
for line in f:
line = re.sub('[|]', '', line)
line = re.sub('[+]', '', line)
line = re.sub('[-]','', line)
for word in line.split():
outfile.write(word + ',')
if not line.isspace():
outfile.write('\n')
df = pd.read_csv("outfile.csv", usecols = ['Iteration', 'Loss', 'Elapsed'])
# +
import matplotlib.pyplot as plt
x = df['Iteration']
y1 = df['Loss']
x1 = df['Elapsed']
plt.xlabel('Iteration')
plt.ylabel('Loss')
plt.plot(x,y1)
#plt.plot(x1,y1)
# -
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <td>
# <a target="_blank" href="https://labelbox.com" ><img src="https://labelbox.com/blog/content/images/2021/02/logo-v4.svg" width=256/></a>
# </td>
# <td>
# <a href="https://colab.research.google.com/github/Labelbox/labelbox-python/blob/develop/examples/basics/labels.ipynb" target="_blank"><img
# src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"></a>
# </td>
#
# <td>
# <a href="https://github.com/Labelbox/labelbox-python/tree/develop/examples/basics/labels.ipynb" target="_blank"><img
# src="https://img.shields.io/badge/GitHub-100000?logo=github&logoColor=white" alt="GitHub"></a>
# </td>
# # Labels
# #### *** This section explains how to use the Label object but It is reccomended that you use bulk export for exporting labels ***
# * [Bulk export examples](https://github.com/Labelbox/labelbox-python/tree/develop/examples/label_export)
# * [Label format documentation](https://docs.labelbox.com/data-model/en/index-en#label)
# !pip install labelbox
from labelbox import Client, Label
import json
import os
# * Set the following cell with your data to run this notebook
# Pick a project that has labels
PROJECT_ID = "ckmdx02yhzsrt0757owj3aprx"
# # API Key and Client
# Provide a valid api key below in order to properly connect to the Labelbox Client.
# Add your api key
API_KEY = None
client = Client(api_key=API_KEY)
project = client.get_project(PROJECT_ID)
# ### Read
# Use relationship to get labels
labels = project.labels()
# Get the first label in the dataset
label = next(labels)
label
# Json result
print(label.seconds_to_label)
# Print first annotation
json.loads(label.label)['objects'][0]
# ... any other field supported by a Label
# See all available fields [here](https://github.com/Labelbox/labelbox-python/blob/develop/labelbox/schema/label.py)
# ### Create
# * Create is not supported from the api.
# * See model assisted upload tutorials for attaching labels to a project
# ### Update
# * We can create reviews and benchmarks for a label
#### Review
label.create_review(score=1)
next(label.reviews()).score
# We also can set this label to be a benchmark
# Labelers will be compared against this label to determine if they are properly trained
label.create_benchmark()
# ### Delete
# * Grab a list of labels
# * Either a whole project worth
# ```labels = list(project.labels())```
# * Or a custom set of labels and bulk delete
# * ```Label.bulk_delete([labels])```
# * Delete a single label with the following
# * `label.delete()`
# * This is not reccomended unless you are only deleting a handful. Otherwise use bulk_delete
| examples/basics/labels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 텐서(Tensors)
# > 2.2.1 장에 해당하는 코드
#
# 해당 챕터에서는 파이썬의 기본 문법과 numpy 패키지의 사용법은 알고 있다고 가정하고 진행합니다. 파이썬의 문법을 배우시고 싶은 분은 아래 링크를 참고
#
# * [모두를 위한 파이썬](https://www.edwith.org/pythonforeverybody)
# 패키지를 불러온다.
import matplotlib.pyplot as plt
import numpy as np
import torch
print(" torch: {} \n numpy: {}".format(torch.__version__, np.__version__))
print(" Use GPU? {}".format(torch.cuda.is_available()))
# ## 텐서를 만드는 방법
# +
# list 로부터 2x3 텐서 생성
x_list = [[1, 2, 3], [4, 5, 6]]
x = torch.Tensor(x_list)
print(x)
# numpy array 로부터 2x3 텐서 생성
x_numpy = np.array([[1, 2, 3], [4, 5, 6]])
x = torch.Tensor(x_numpy)
print(x)
# -
# 다시 원래 자료형태로 복귀
# +
# .tolist()
x_back2list = x.tolist()
print(type(x_back2list))
# .numpy()
x_back2numpy = x.numpy()
print(type(x_back2numpy))
# -
# gpu 사용하기
# +
# 기본 device 정보
print(x.device)
device = 'cuda'
# GPU 사용
x = x.to(device)
print(x.device)
device = 'cpu'
# CPU 사용
x = x.to(device)
print(x.device)
# -
# 랜덤 텐서 생성
# +
# 랜덤 숫자로 구성된 크기가 2x3 인 텐서 생성
# 0과 1사이의 랜덤한 숫자
print(torch.rand(2, 3))
# 0과 8 사이의 정수형 랜덤한 숫자
print(torch.randint(low=0, high=8, size=(2, 3)))
# -
# `torch.zeros_like`, `torch.ones_like`
# GPU를 사용하고 크기가 x 와 같은 0으로 채워진 텐서 생성
x_zeros = torch.zeros_like(x.cuda())
print(x_zeros.device)
print(x_zeros)
# ## 텐서의 타입(type)
# +
# 실수형 텐서
a = torch.FloatTensor(np.array([[1, 2, 3], [4, 5, 6]]))
print(a.type())
print(a)
# 정수형 텐서
b = torch.LongTensor(np.array([[1, 2, 3], [4, 5, 6]]))
print(b.type())
print(b)
# 8 bit 정수형
c = torch.ByteTensor([True, False, True, True])
print(c.type())
print(c)
# -
# ## Tensor Manipulation
#
# ### Slicing
torch.manual_seed(777)
x = torch.randint(0, 10, size=(2, 3, 4))
print(x)
print(x[0, 2, 3])
def draw_tensor(temp):
"""tensor drawing in ch3"""
fig, axes = plt.subplots(len(temp), 1)
for i in range(len(temp)):
axes[i].matshow(temp.numpy()[i], vmin=0, vmax=1, cmap='gray')
plt.show()
# 선택한 텐서를 시각화 한다.
temp = torch.ones_like(x)
temp[0, 2, 3] = 0
draw_tensor(temp)
print(x[0, :2, 3])
# 선택한 텐서를 시각화 한다.
temp = torch.ones_like(x)
temp[0, :2, 3] = 0
draw_tensor(temp)
x[0, 2, 1:]
temp = torch.ones_like(x)
temp[0, 2, 1:] = 0
fig = draw_tensor(temp)
print(x[0, 2, 1:2])
temp = torch.ones_like(x)
temp[0, 2, 1:2] = 0
fig = draw_tensor(temp)
print(x[0, 2, :])
temp = torch.ones_like(x)
temp[0, 2, :] = 0
fig = draw_tensor(temp)
# ### view
# +
# 크기가 (2, 3, 4) 3차원 텐서를 (2, 2, 6) 으로 변경
print(x.view(2, 2, 6))
# 크기가 (2, 3, 4) 3차원 텐서를 (1, 1, 12) 으로 변경
print(x.view(-1, 1, 12))
# -
# ### transpose
# (2, 3, 4) 크기의 텐서의 첫번째 차원과 두번째 차원이 바뀐다.
x_transposed = x.transpose(0, 1)
print(x_transposed.size())
print(x_transposed)
# (2, 3) 크기의 행렬 생성
A = torch.Tensor([[1, 2, 3],
[4, 5, 6]])
# 크기가 (3, 2) 이 되었다.
A.t()
# ## squeeze & unsqueeze
# +
# 크기가 (2, 1, 3, 4, 1) 인 5차원 텐서를 생성한다
x = torch.rand((2, 1, 3, 4, 1))
# 모든 차원에서 크기가 1인 차원을 squeeze 한다.
print(x.squeeze().size()) # 크기확인
# 두번째 차원(크기 = 1)을 squeeze 한다.
print(x.squeeze(1).size()) # 크기확인
# 4번째 차원에 크기를 1 추가, 6차원 텐서가 된다.
print(x.unsqueeze(3).size()) # 크기확인
# -
# # ### cat & stack
def draw_tensor2(x):
"""tensor drawing in ch3: cat&stack"""
subplot_size = 1 if x.ndimension() <= 2 else x.size(0)
fig, axes = plt.subplots(subplot_size, 1)
if subplot_size == 1:
axes.matshow(x.numpy(), vmin=0, vmax=1, cmap='gray')
else:
for i in range(subplot_size):
axes[i].matshow(x.numpy()[i], vmin=0, vmax=1, cmap='gray')
plt.show()
# +
torch.manual_seed(777)
# 크기가 (2, 3) 인 A, B 텐서를 만든다
A = torch.rand((2, 3))
B = torch.rand((2, 3))
# 첫번째 차원을 기준으로 텐서를 concatenate 한다.
AB_cated = torch.cat([A, B], dim=0)
print(AB_cated)
# 시각화
draw_tensor2(AB_cated)
# +
# 첫번째 차원을 기준으로 텐서를 stack 한다.
AB_stacked = torch.stack([A, B], dim=0)
print(AB_stacked)
# 각 텐서를 첫번째 차원 기준으로 unsqueeze 후, cat 한것과 같은 결과
AB_unsqueeze_cat = torch.cat([A.unsqueeze(0), B.unsqueeze(0)], dim=0)
print(AB_unsqueeze_cat)
# 시각화
draw_tensor2(AB_stacked)
# -
# ## 텐서의 사칙연산: add / multiplication
#
# ### 텐서의 덧셈과 뺄셈
# +
# x, y 크기가 3인 1차원 벡터 생성
x = torch.Tensor([1, 2, 3])
y = torch.Tensor([4, 5, 6])
# 같은 크기의 벡터간 덧셈
print(torch.add(x, y))
# 같은 크기의 벡터간 뺄셈
print(torch.sub(x, y))
# 벡터 + 스칼라
print(x+1)
# 벡터 - 스칼라
print(x-1)
# -
# ### 텐서의 곱셈
# +
# 곱셈
print(x * y)
print(torch.mul(x, y))
# 나눗셈
print(x / y)
print(torch.div(x, y))
# -
print("inner product: ")
print(torch.dot(x, y))
print(x.dot(y))
# +
# (3, 2) 크기의 X 텐서와 (2, 2) 크기의 Y 텐서를 생성한다.
X = torch.Tensor([[1, 4], [2, 5], [3, 6]])
Y = torch.Tensor([[7, 9], [8, 10]])
# 행렬의 곱셈을 한다.
print(torch.mm(X, Y))
print(X.mm(Y))
# -
# ## 기초연산:
#
# ### sum / mean
# +
# (2, 2, 2) 크기의 3차원 텐서 Z 를 생성한다.
Z = torch.Tensor([[[1, 2], [3, 4]], [[5, 6], [7, 8]]])
# 텐서의 모든 원소의 합 혹은 평균을 구한다.
print(torch.sum(Z))
print(torch.mean(Z))
# 지정한 차원의 합을 축소시키면서 합 혹은 평균을 구한다.
print(torch.sum(Z, dim=0))
print(torch.mean(Z, dim=0))
# -
# ### max & argmax / min & argmin
# +
# 텐서의 모든 원소중 최대값 및 최대값의 위치 구하기
print(torch.max(Z))
print(torch.argmax(Z))
# 텐서의 모든 원소중 최소값 및 최소값의 위치 구하기
print(torch.min(Z))
print(torch.argmin(Z))
# 차원 지정시 지정된 차원을 기준으로 차원이 축소 되면서
# 최대값 및 위치 혹은 최소값 및 위치를 가진 튜플(tuple)을 반환한다.
print(torch.max(Z, dim=0))
print(torch.min(Z, dim=0))
# -
# ### 논리연산
# +
# 크기가 3인 벡터를 생성
z = torch.Tensor([-3, 2, 0])
# 0과 같다
print(z.eq(0))
print(z == 0)
# 0 보다 크거나 같다
print(z.ge(0))
print(z >= 0)
# 0보다 크다
print(z.gt(0))
print(z > 0)
# 0보다 작거나 같다
print(z.le(0))
print(z <= 0)
# 0보다 작다
print(z.lt(0))
print(z < 0)
# -
| 01-tensor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
import datetime
def rain_warning(precip_chance, precip_type):
warning=''
if precip_chance >= 0.9:
warning="Whoa, it's almost certainly "+precip_type+"ing today. Bring an umbrella or something."
elif precip_chance >= 0.5:
warning="Better than even chance of "+precip_type+"ing today. Might want to look out for that"
elif precip_chance >= 0.1:
warning="Non-zero chance of "+precip_type+"ing today. Just sayin'."
else:
warning="Pretty much nothing going on today. Might see some "+ precip_type+", but don't count on it."
return(warning)
def send_mail(message_text, subject):
return requests.post(
"https://api.mailgun.net/v3/sandbox5375a212cf1648e59f6321df8fdb88da.mailgun.org/messages",
auth=("api", "key-<KEY>"),
data={"from": "weather bot <<EMAIL>>",
"to": "<EMAIL>",
"subject": subject,
"text": message_text})
url = 'https://api.forecast.io/forecast/'
apikey = '53343f8442d30d598f50f5910124610a'
latlong = '40.8117150,-73.9578630'
response = requests.get(url+apikey+'/'+latlong)
forecast_complete = response.json()
current_conditions=forecast_complete['currently']
forecast_today=forecast_complete['daily']['data'][0]
forecast_email=["Right now it is ", current_conditions['summary'].lower(), " out and feels like ", str(current_conditions['temperature']),
" degrees. Today it will be ", forecast_today['summary'][:-1].lower()+", with a high of ",
str(forecast_today['temperatureMax']), " and a low of ", str(forecast_today['temperatureMin'])," degrees. "]
if 'precipType' in forecast_today:
forecast_email.append(rain_warning(forecast_today['precipProbability'], forecast_today['precipType']))
else:
forecast_email.append("Literally no chance of precipitation today. ")
today_raw = datetime.datetime.today()
today = today_raw.strftime("%A %B %d %Y").split(' ')
msg_subject=('8AM Weather forecast for ', today[0]+', ', today[1], ' ', today[2]+', ', today[3])
send_mail(''.join(forecast_email), ''.join(msg_subject))
| 10/homework_10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 0.0.IMPORTS
# +
import math
import datetime
import inflection
import numpy as np
import pandas as pd
import seaborn as sns
from scipy import stats as ss
from boruta import BorutaPy
from tabulate import tabulate
from matplotlib import pyplot as plt
from IPython.display import Image
from sklearn.ensemble import RandomForestRegressor
from IPython.core.display import HTML
from sklearn.preprocessing import RobustScaler, MinMaxScaler, LabelEncoder
# -
# ## 0.1.HELPER FUNCTIONS
# +
def jupyter_settings():
# %matplotlib inline
# %pylab inline
plt.stlye.use ('bmh')
plt.rcParams['figure.figsize'] = [40,20]
plt.rcParams['font.size'] = 24
display(HTML( '<style>. container {width: 100% !importante; }</style>' ) )
pd.options.display.max_columns = None
pd.options.display.max_rows = None
pd.set_option( 'display.expand_frame_repr', False )
sns.set()
def cramer_v (x,y):
cm = pd.crosstab( x, y ).values
n = cm.sum()
r,k = cm.shape
chi2 = ss.chi2_contingency (cm) [0]
chi2corr = max(0,chi2 - (k-1)*(r-1)/(n-1))
kcorr = k - (k-1) **2/(n-1)
rcorr = r - (r-1) **2/(n-1)
return np.sqrt( (chi2corr/n) / (min(kcorr-1, rcorr-1) ) )
# -
# ## 0.2.LOADING DATA
#
# +
df_sales_raw = pd.read_csv ( 'data/train.csv', low_memory=False)
df_stores_raw = pd.read_csv ( 'data/store.csv', low_memory=False)
#MERGE DE DADOS
df_raw = pd.merge ( df_sales_raw, df_stores_raw, how= 'left', on= 'Store' )
# -
df_raw.head()
# # 1.0 DESCRICAO DOS DADOS
# ## 1.1 RENAME COLUMNS
df1 = df_raw.copy()
df1.columns
# + code_folding=[]
cols_old = ['Store', 'DayOfWeek', 'Date', 'Sales', 'Customers', 'Open', 'Promo', 'StateHoliday', 'SchoolHoliday',
'StoreType', 'Assortment', 'CompetitionDistance', 'CompetitionOpenSinceMonth', 'CompetitionOpenSinceYear',
'Promo2', 'Promo2SinceWeek','Promo2SinceYear', 'PromoInterval']
snakecase = lambda x: inflection.underscore (x)
cols_new = list( map( snakecase, cols_old) )
#rename
df1.columns = cols_new
# -
df1.columns
# ## 1.2 DIMENSIONS
# +
##shape 0 = linhas 1 = colunas
print('Number of Rows: {}'.format ( df1.shape[0] ) )
print('Number of Columns: {}'.format ( df1.shape[1] ) )
# -
# ## 1.3 DATA TYPES
df1['date'] = pd.to_datetime( df1['date'] )
df1.dtypes
# ## 1.4 CHECK NA
df1.isna().sum()
# ## 1.5 FILLOUT NA
df1 ['competition_distance'].max()
# + code_folding=[]
#competition_distance
df1['competition_distance'] = df1 ['competition_distance'].apply( lambda x: 20000.0 if math.isnan (x) else x)
#competition_open_since_month
df1['competition_open_since_month'] = df1.apply( lambda x: x ['date'].month if math.isnan( x['competition_open_since_month'] ) else x['competition_open_since_month'], axis = 1 )
#competition_open_since_year
df1['competition_open_since_year'] = df1.apply( lambda x: x ['date'].year
if math.isnan( x['competition_open_since_year'] )
else x['competition_open_since_year'], axis = 1 )
#promo2_since_week
df1['promo2_since_week'] = df1.apply( lambda x: x ['date'].week
if math.isnan( x['promo2_since_week'] )
else x['promo2_since_week'], axis = 1 )
#promo2_since_year
df1['promo2_since_year'] = df1.apply( lambda x: x ['date'].year
if math.isnan( x['promo2_since_year'] )
else x['promo2_since_year'], axis = 1 )
#promo_interval
month_map = {1:'Jan', 2:'Feb', 3:'Mar', 4:'Apr', 5:'May', 6:'Jun', 7:'Jul', 8:'Aug', 9:'Sep', 10:'Oct', 11:'Nov', 12:'Dec'}
df1['promo_interval'].fillna(0,inplace = True)
df1['month_map'] = df1['date'].dt.month.map( month_map )
#promo_interval - Se a promo está dentro do intervalo indicado
df1['is_promo'] = df1[['month_map', 'promo_interval']].apply( lambda x: 0
if x['promo_interval'] == 0
else 1 if x['month_map'] in x['promo_interval'].split (',')
else 0, axis = 1)
# -
# ## 1.6. CHANGE TYPES
df1.dtypes
# +
#Mudar os tipos dos dados das colunas
df1['competition_open_since_month'] = df1['competition_open_since_month'].astype (int)
df1['competition_open_since_year'] = df1['competition_open_since_year'].astype (int)
df1['promo2_since_week'] = df1['promo2_since_week'].astype (int)
df1['promo2_since_year'] = df1['promo2_since_year'].astype (int)
# -
# ## 1.7. DESCRIPTIVE STATISTICAL
#Separar variaveis numéricas e categórigas
num_attributes = df1.select_dtypes( include = ['int32', 'int64' , 'float64'] )
cat_attributes = df1.select_dtypes( exclude = ['int32', 'int64' , 'float64', 'datetime64[ns]'] )
# +
#Central Tendency - mean, median
ct1 = pd.DataFrame( num_attributes.apply ( np.mean ) ).T
ct2 = pd.DataFrame( num_attributes.apply ( np.median) ).T
#Dispersion - std, min, max, range, skew, kurtosis
d1 = pd.DataFrame( num_attributes.apply( np.std ) ).T
d2 = pd.DataFrame( num_attributes.apply( min ) ).T
d3 = pd.DataFrame( num_attributes.apply( max ) ).T
d4 = pd.DataFrame( num_attributes.apply( lambda x: x.max() - x.min() ) ).T
d5 = pd.DataFrame( num_attributes.apply( lambda x: x.skew() ) ).T
d6 = pd.DataFrame( num_attributes.apply( lambda x: x.kurtosis() ) ).T
#Concatenate
m = pd.concat( [d2,d3,d4,ct1,ct2,d1,d5,d6] ).T.reset_index()
m.columns = ['attributes', 'min', 'max', 'range','mean', 'median', 'std', 'skew', 'kurtosis']
# -
m.head(9)
sns.displot(df1['competition_distance'])
cat_attributes.head()
#variáveis categóricas sendo analisadas por boxplot.
cat_attributes.apply ( lambda x: x.unique().shape[0] )
# +
aux1 = df1 [ ( df1['state_holiday'] != '0' ) & ( df1['sales'] > 0 ) ]
plt.subplot(1,3,1)
sns.boxplot( x= 'store_type', y= 'sales' , data= aux1 )
plt.subplot(1,3,2)
sns.boxplot( x= 'state_holiday', y= 'sales' , data= aux1 )
plt.subplot(1,3,3)
sns.boxplot( x= 'assortment', y= 'sales' , data= aux1 )
# -
df1.sample(5).T
df2 = df1.copy()
# # 2.0 Feature Engeneering
# + [markdown] heading_collapsed=true
# ## 2.1. Mapa Mental de Hipóteses
# + hidden=true
Image('image/Minmap_hipotese.png')
# -
# ## 2.2. Criação das Hipóteses
# + [markdown] heading_collapsed=true
# ### 2.2.1. Hipótese de Loja
# + [markdown] hidden=true
# **1.** Lojas com maior quadro de funcionários deveriam vernder mais.
# + [markdown] hidden=true
# **2.** Lojas com maior estoque deveriam vender mais.
# + [markdown] hidden=true
# **3.** Lojas com maior porte deveriam vender mais.
# + [markdown] hidden=true
# **4.** Lojas com menor porte deveriam vender mais.
# + [markdown] hidden=true
# **5.** Lojas com maior sortimento deveriam vender mais.
# + [markdown] hidden=true
# **6.** Lojas com competidores à mais tempo deveriam vender mais
# + [markdown] hidden=true
# **7.** Lojas com competidores mais próximos deveriam vender menos
# + [markdown] heading_collapsed=true
# ### 2.2.2. Hipótese de Produtos
# + [markdown] hidden=true
# **1.** Lojas que investem mais em marketing, deveriam vender mais
# + [markdown] hidden=true
# **2.** Lojas que expõem mais os produtos na vitrine, deveriam vender mais
# + [markdown] hidden=true
# **3.** Lojas que tem preços menores, deveriam vender mais
# + [markdown] hidden=true
# **4.** Lojas que tem preços menores deveriam vender mais
# + [markdown] hidden=true
# **5.** Lojas com promoçao mais agressiva (descontos maiores), deveriam vender mais
# + [markdown] hidden=true
# **6.** Lojas com promoções ativas por mais tempo, deveriam vender mais
# + [markdown] hidden=true
# **7.** Lojas com mais dias de promoção deveriam vender mais
# + [markdown] hidden=true
# **8.** Lojas com mais promoções consecutivas deveriam vender mais
# + [markdown] heading_collapsed=true
# ### 2.2.3. Hipótese de Tempo
# + [markdown] hidden=true
# **1.** Lojas abertas durante o feriado de Natal deveriam vender mais
# + [markdown] hidden=true
# **2.** Lojas deveriam vender mais ao longo dos anos
# + [markdown] hidden=true
# **3.** Lojas deveriam vender mais no segundo semestre do ano
# + [markdown] hidden=true
# **4.** Lojas deveriam vender mais depois do dia 10 de cada mês
# + [markdown] hidden=true
# **5.** Lojas deveriam vender menos aos finais de semana
# + [markdown] hidden=true
# **6.** Lojas deveriam vender menos durante os feriados escolares
# + [markdown] heading_collapsed=true
# ## 2.3. Lista Final de Hipóteses
# + [markdown] hidden=true
# **1.** Lojas com maior sortimento deveriam vender mais.
# + [markdown] hidden=true
# **2.** Lojas com competidores à mais tempo deveriam vender mais
# + [markdown] hidden=true
# **3.** Lojas com competidores mais próximos deveriam vender menos
# + [markdown] hidden=true
# **4.** Lojas com competidores a mais tempo deveriam vender mais
# + [markdown] hidden=true
# **5.** Lojas com promoções ativas por mais tempo, deveriam vender mais
# + [markdown] hidden=true
# **6.** Lojas com mais dias de promoção deveriam vender mais
# + [markdown] hidden=true
# **7.** Lojas com mais promoções consecutivas deveriam vender mais
# + [markdown] hidden=true
# **8.** Lojas abertas durante o feriado de Natal deveriam vender mais
# + [markdown] hidden=true
# **9.** Lojas deveriam vender mais ao longo dos anos
# + [markdown] hidden=true
# **10.** Lojas deveriam vender mais no segundo semestre do ano
# + [markdown] hidden=true
# **11.** Lojas deveriam vender mais depois do dia 10 de cada mês
# + [markdown] hidden=true
# **12.** Lojas deveriam vender menos aos finais de semana
# + [markdown] hidden=true
# **13.** Lojas deveriam vender menos durante os feriados escolares
# -
# ## 2.4. Feature Engineering
# +
# year
df2['year'] = df2 ['date'].dt.year
# month
df2['month'] = df2 ['date'].dt.month
# day
df2['day'] = df2 ['date'].dt.day
# week of the year
df2['week_of_year'] = df2 ['date'].dt.isocalendar().week
# year week
df2['year_week'] = df2 ['date'].dt.strftime('%Y-%W')
# competittion since
df2['competition_since'] = df2.apply (lambda x: datetime.datetime ( year= x['competition_open_since_year'], month= x['competition_open_since_month'], day=1 ), axis = 1 )
df2['competition_time_month'] = ((df2['date'] - df2['competition_since'])/30 ).apply( lambda x: x.days).astype (int)
# promo since
df2['promo_since'] = df2['promo2_since_year'].astype(str) + '-' + df2['promo2_since_week'].astype(str)
df2['promo_since'] = df2['promo_since'].apply(lambda x: datetime.datetime.strptime(x + '-1', '%Y-%W-%w') - datetime.timedelta ( days=7))
df2['promo_time_week']= ((df2['date'] - df2['promo_since'])/7).apply (lambda x: x.days ).astype(int)
# assortment
df2['assortment'] = df2['assortment'].apply(lambda x: 'basic' if x == 'a' else 'extra' if x == 'b' else 'extended')
# state holiday
df2['state_holiday'] = df2['state_holiday'].apply(lambda x: 'public_holiday' if x == 'a' else 'easter_holiday' if x == 'b' else 'christmas' if x == 'c' else 'regular_day')
# -
df2.head().T
# # 3.0. PASSO 03 - FILTRAGEM DE VARIÁVEIS
# ## 3.1. SELECAO DE LINHAS
df3 = df2.copy()
df3 = df3[(df3['open'] != 0) & (df3['sales'] > 0)]
# ## 3.1. SELECAO DE COLUNAS
cols_drop = ['customers', 'open', 'promo_interval', 'month_map']
df3 = df3.drop(cols_drop, axis = 1)
df3.columns
# # 4.0. PASSO 04 - ANALISE EXPLORATORIA
df4 = df3.copy()
# ## 4.1. Analise Univariada
# ### 4.1.1. Response Variable
# + hide_input=false
sns.displot(df4['sales'])
# -
# ### 4.1.2. Numerical Variable
# + hide_input=false
num_attributes.hist(bins = 25)
# -
# ### 4.1.3. Categorical Variable
cat_attributes.head()
df4['state_holiday'].drop_duplicates()
# +
##state_holiday
plt.subplot ( 3,2,1 )
a = df4[df4['state_holiday'] != 'regular_day']
sns.countplot( a['state_holiday'])
plt.subplot ( 3,2,2 )
sns.kdeplot( df4[ df4['state_holiday'] == 'public_holiday']['sales'], label = 'public_holiday', shade = True)
sns.kdeplot( df4[ df4['state_holiday'] == 'easter_holiday']['sales'], label = 'easter_holiday', shade = True)
sns.kdeplot( df4[ df4['state_holiday'] == 'christmas']['sales'], label = 'christmas', shade = True)
#store_type
plt.subplot ( 3,2,3 )
sns.countplot( df4['store_type'])
plt.subplot ( 3,2,4 )
sns.kdeplot( df4[ df4['store_type'] == 'a']['sales'], label = 'a', shade = True)
sns.kdeplot( df4[ df4['store_type'] == 'b']['sales'], label = 'b', shade = True)
sns.kdeplot( df4[ df4['store_type'] == 'c']['sales'], label = 'c', shade = True)
sns.kdeplot( df4[ df4['store_type'] == 'd']['sales'], label = 'd', shade = True)
##assortment
plt.subplot ( 3,2,5 )
sns.countplot( df4['assortment'])
plt.subplot ( 3,2,6 )
sns.kdeplot( df4[ df4['assortment'] == 'extended']['sales'], label = 'extended', shade = True)
sns.kdeplot( df4[ df4['assortment'] == 'basic']['sales'], label = 'basic', shade = True)
sns.kdeplot( df4[ df4['assortment'] == 'extra']['sales'], label = 'extra', shade = True)
# -
# ## 4.2. Analise Bivariada
# + [markdown] heading_collapsed=true
# ### H1. Lojas com maior sortimento deveriam vender mais.
# **FALSA** Lojas com MAIOR SORTIMENTO vendem MENOS.
# + hidden=true hide_input=true
aux1 = df4[['assortment', 'sales']].groupby( 'assortment').sum().reset_index()
sns.barplot (x = 'assortment', y = 'sales', data = aux1)
aux2 = df4[['year_week', 'assortment' ,'sales']].groupby( ['year_week' , 'assortment'] ).sum().reset_index()
aux2.pivot ( index = 'year_week', columns = 'assortment' , values = 'sales').plot()
## plotado separadamente devido a escala - observar melhor como as vendas do tipo extra ocorre.
aux3 = aux2[aux2 ['assortment'] == 'extra']
aux3.pivot ( index = 'year_week', columns = 'assortment' , values = 'sales').plot()
# + [markdown] heading_collapsed=true
# ### H2. Lojas com competidores à mais tempo deveriam vender mais
# **FALSA** Lojas com COMPETIDORES MAIS PRÓXIMOS vendem MAIS
# + hidden=true hide_input=true
aux1 = df4[['competition_distance', 'sales']].groupby('competition_distance').sum().reset_index()
sns.barplot(x= 'competition_distance', y='sales', data=aux1);
# + hidden=true hide_input=false
aux1 = df4[['competition_distance', 'sales']].groupby('competition_distance').sum().reset_index()
plt.subplot (1,3,1)
sns.scatterplot( x= 'competition_distance', y= 'sales', data = aux1);
plt.subplot (1,3,2)
bins = list(np.arange (0,20000, 1500) )
aux1['competition_distance_binned'] = pd.cut(aux1['competition_distance'], bins = bins)
aux2 = aux1[['competition_distance_binned', 'sales']].groupby('competition_distance_binned').sum().reset_index()
sns.barplot(x= 'competition_distance_binned', y='sales', data=aux2);
plt.xticks (rotation = 90);
plt.subplot (1,3,3)
sns.heatmap(aux1.corr(method = 'pearson'), annot = True);
# + hidden=true
aux1.sample(4)
# + [markdown] heading_collapsed=true
# ### H3. Lojas com competidores há mais tempo deveriam vender mais
# **FALSA** Lojas com COMPETIDORES HÁ MAIS TEMPO vendem MENOS
# + hidden=true
plt.subplot(1,3,1)
aux1 = df4 [['competition_time_month', 'sales']].groupby ('competition_time_month').sum().reset_index()
aux2 = aux1[( aux1 ['competition_time_month'] < 120 ) & ( aux1 ['competition_time_month'] != 0 )]
sns.barplot(x='competition_time_month' , y='sales', data = aux2);
plt.xticks (rotation = 90);
plt.subplot(1,3,2)
sns.regplot(x= 'competition_time_month' , y= 'sales', data= aux2);
plt.subplot(1,3,3)
sns.heatmap(aux1.corr(method = 'pearson'), annot = True);
# + [markdown] heading_collapsed=true
# ### H4. Lojas com promoções ativas por mais tempo deveriam vender mais.
# **FALSA** Lojas compromocoes ativas por mais tempo vendem menos, depois de um certo periodo de promocao
# + hidden=true
aux1 = df4[['promo_time_week', 'sales']].groupby('promo_time_week').sum().reset_index()
grid = plt.GridSpec(2,3)
plt.subplot(grid[0,0])
aux2 = aux1 [aux1['promo_time_week'] > 0] #promo extendido
sns.barplot (x = 'promo_time_week', y = 'sales', data = aux2);
plt.xticks (rotation = 90);
plt.subplot(grid[0,1])
sns.regplot (x = 'promo_time_week', y = 'sales', data = aux2);
plt.subplot(grid[1,0])
aux3 = aux1 [aux1['promo_time_week'] < 0] #promo regular
sns.barplot (x = 'promo_time_week', y = 'sales', data = aux3);
plt.xticks (rotation = 90);
plt.subplot(grid[1,1])
sns.regplot (x = 'promo_time_week', y = 'sales', data = aux3);
plt.subplot(grid[:,2])
sns.heatmap(aux1.corr(method = 'pearson'), annot = True);
# + [markdown] heading_collapsed=true
# ### <s> H5. Lojas com mais dias de promoção deveriam vender mais <s>
# + [markdown] heading_collapsed=true hide_input=true
# ### H7. Lojas com mais promoções consecutivas deveriam vender mais
# **FALSA** Lojas com promocoes consecutivas vendem menos
# + hidden=true hide_input=false
df4[['promo', 'promo2', 'sales']].groupby( [ 'promo', 'promo2'] ).sum().reset_index()
# + hidden=true
aux1 = df4[( df4 ['promo'] == 1 ) & ( df4['promo2'] == 1 )][['year_week', 'sales']].groupby('year_week').sum().reset_index()
ax = aux1.plot()
aux2 = df4[( df4 ['promo'] == 1 ) & ( df4['promo2'] == 0 )][['year_week', 'sales']].groupby('year_week').sum().reset_index()
aux2.plot(ax=ax)
ax.legend(labels = ['Tradicional & Extendida', 'Extendida']);
# + [markdown] heading_collapsed=true
# ### H8. Lojas abertas durante o feriado de Natal deveriam vender mais
# **FALSA** Lojas vendem menos no feriado de natal
# + hidden=true
aux = df4[df4['state_holiday'] != 'regular_day']
plt.subplot(1,2,1)
aux1 = aux[['state_holiday', 'sales']].groupby('state_holiday').sum().reset_index()
sns.barplot( x= 'state_holiday', y ='sales', data = aux1);
plt.subplot(1,2,2)
aux2 = aux[['year', 'state_holiday', 'sales']].groupby( [ 'year', 'state_holiday'] ).sum().reset_index()
sns.barplot( x= 'year', y= 'sales', hue = 'state_holiday', data= aux2 );
# + hidden=true
aux = df4[(df4['state_holiday'] != 'regular_day') & (df4['state_holiday'] != 'public_holiday')]
plt.subplot(1,2,1)
aux1 = aux[['state_holiday', 'sales']].groupby('state_holiday').mean().reset_index()
sns.barplot( x= 'state_holiday', y ='sales', data = aux1);
plt.subplot(1,2,2)
aux2 = aux[[ 'state_holiday', 'sales']].groupby( [ 'state_holiday'] ).mean().reset_index()
sns.barplot( x= 'state_holiday', y= 'sales', hue = 'state_holiday', data= aux2 );
# + [markdown] heading_collapsed=true
# ### H9. Lojas deveriam vender mais ao longo dos anos
# **FALSA** Lojas vendem menos ao longo dos anos
# + hidden=true
aux1 = df4[['year', 'sales']].groupby('year').sum().reset_index()
plt.subplot(1,3,1)
sns.barplot(x = 'year', y='sales', data = aux1);
plt.subplot(1,3,2)
sns.regplot(x = 'year', y='sales', data = aux1);
plt.subplot(1,3,3)
sns.heatmap(aux1.corr(method = 'pearson'), annot = True);
# + hidden=true
aux1 = df4[['year', 'sales']].groupby('year').sum().reset_index()
plt.subplot(1,3,1)
sns.barplot(x = 'year', y='sales', data = aux1);
plt.subplot(1,3,2)
sns.regplot(x = 'year', y='sales', data = aux1);
plt.subplot(1,3,3)c
sns.heatmap(aux1.corr(method = 'pearson'), annot = True);
# -
# ### H10.** Lojas deveriam vender mais no segundo semestre do ano
# **FALSA** Lojas vendem menos no segundo semestre do ano
# + hide_input=true
aux1 = df4[['month', 'sales']].groupby('month').sum().reset_index()
plt.subplot(1,3,1)
sns.barplot(x = 'month', y='sales', data = aux1);
plt.subplot(1,3,2)
sns.regplot(x = 'month', y='sales', data = aux1);
plt.subplot(1,3,3)
sns.heatmap(aux1.corr(method = 'pearson'), annot = True);
# -
# ### H11.** Lojas deveriam vender mais depois do dia 10 de cada mês
# **Verdadeiro** Lojas vendem mais após o dia 10
# + hide_input=true
aux1 = df4[['day', 'sales']].groupby('day').sum().reset_index()
plt.subplot(2,2,1)
sns.barplot(x = 'day', y='sales', data = aux1);
plt.subplot(2,2,2)
sns.regplot(x = 'day', y='sales', data = aux1);
plt.subplot(2,2,3)
sns.heatmap(aux1.corr(method = 'pearson'), annot = True);
plt.subplot(2,2,4)
aux1['before_after'] = aux1['day'].apply(lambda x: 'before_10_days'if x<= 10 else 'after_10_days')
aux2 = aux1[['before_after', 'sales']].groupby('before_after').sum().reset_index()
sns.barplot(x='before_after', y='sales', data = aux2);
# -
# ### H12.** Lojas deveriam vender menos aos finais de semana
# **Verdadeiro** Lojas vendem menos nos finais de semana
# + hide_input=true
aux1 = df4[['day_of_week', 'sales']].groupby('day_of_week').sum().reset_index()
plt.subplot(1,3,1)
sns.barplot(x = 'day_of_week', y='sales', data = aux1);
plt.subplot(1,3,2)
sns.regplot(x = 'day_of_week', y='sales', data = aux1);
plt.subplot(1,3,3)
sns.heatmap(aux1.corr(method = 'pearson'), annot = True);
# + [markdown] heading_collapsed=true
# ### H13.** Lojas deveriam vender menos durante os feriados escolares
# **Verdadeiro** Lojas vendem vendem menos durante os feriados escolares, exceto os meses de Julho e Agosto.
# + hidden=true
aux1 = df4[['school_holiday', 'sales']].groupby('school_holiday').sum().reset_index()
plt.subplot(2,1,1)
sns.barplot(x = 'school_holiday', y='sales', data = aux1);
aux2 = df4[['month','school_holiday', 'sales']].groupby(['month','school_holiday']).sum().reset_index()
plt.subplot(2,1,2)
sns.barplot(x = 'month', y='sales', hue='school_holiday' ,data = aux2);
# -
# ### 4.2.1. Resumo das Hipóteses
tab = [['Hipotese', 'Conclusao', 'Relevancia'],
['H1', 'Falsa', 'Baixa'],
['H2', 'Falsa', 'Media'],
['H3', 'Falsa', 'Media'],
['H4', 'Falsa', 'Baixa'],
['H5', ' -', '-'],
['H7', 'Falsa', 'Baixa'],
['H8', 'Falsa', 'Media'],
['H9', 'Falsa', 'Alta'],
['H10', 'Verdadeira', 'Alta'],
['H11', 'Verdadeira', 'Alta'],
['H12', 'Verdadeira', 'Alta'],
['H13', 'Verdadeira', 'Baixa'],
]
print(tabulate (tab, headers = 'firstrow'))
# ## 4.3. Analise Multivariada
# ### 4.3.1. Numerical Attributes
##person - categorias numericas
correlation = num_attributes.corr(method = 'pearson')
sns.heatmap (correlation, annot = True);
# ### 4.3.1. Categorical Attributes
# +
#Only Categorical Data
a = df4.select_dtypes ( include = 'object')
#Calculate Cramer_v
a1 = cramer_v(a['state_holiday'], a['state_holiday'])
a2 = cramer_v(a['state_holiday'], a['store_type'])
a3 = cramer_v(a['state_holiday'], a['assortment'])
a4 = cramer_v(a['store_type'], a['state_holiday'])
a5 = cramer_v(a['store_type'], a['store_type'])
a6 = cramer_v(a['store_type'], a['assortment'])
a7 = cramer_v(a['assortment'], a['state_holiday'])
a8 = cramer_v(a['assortment'], a['store_type'])
a9 = cramer_v(a['assortment'], a['assortment'])
# Final Dataset
d = pd.DataFrame ({'state_holiday': [a1,a2,a3],
'store_type': [a4,a5,a6],
'assortment': [a7,a8,a9]})
d=d.set_index(d.columns)
# -
sns.heatmap(d, annot = True)
# # 5.0. PASSO 05 - DATA PREPARATION
df5 = df4.copy()
# ## 5.1. Normalização
# ## 5.2. Rescaling
print(df5.dtypes)
df5['week_of_year'] = df5['week_of_year'].astype('Int64')
df5[ 'competition_time_month'] = df5['competition_time_month'].astype('Int64')
df5[ 'promo_time_week'] = df5['promo_time_week'].astype('Int64')
a = df5.select_dtypes( include = ['int64', 'float64'] )
#Check for outliers
sns.boxplot(a['promo_time_week'])
# +
rs = RobustScaler()
mms = MinMaxScaler()
#Natureza não cíclica - ROBUST SCALER
#competition_distance
df5['competition_distance'] = rs.fit_transform( df5[['competition_distance']].values)
#competition_time_month
df5['competition_time_month'] = rs.fit_transform( df5[['competition_time_month']].values)
#promo_time_week
df5['promo_time_week'] = mms.fit_transform( df5[['promo_time_week']].values)
#year
df5['year'] = mms.fit_transform( df5[['year']].values)
# -
# ## 5.3. Transformacao
# ### 5.3.1. Encoding
# +
#Natureza Linear
#state_holiday - One Hot Encodiing
df5 = pd.get_dummies(df5, prefix = ['state_holiday'], columns = ['state_holiday'])
#store_type - Label Encoding
le = LabelEncoder()
df5['store_type'] = le.fit_transform( df5[ 'store_type' ] )
#assortment - Ordinal Encoding
assortment_dict = {'basic':1 , 'extra' : 2 , 'extended' : 3}
df5['assortment'] = df5 ['assortment'].map( assortment_dict )
# +
##<NAME>
#day_of_week
df5['day_of_week_sin'] = df5['day_of_week'].apply( lambda x: np.sin( x* ( 2. * np.pi/7) ) )
df5['day_of_week_cos'] = df5['day_of_week'].apply( lambda x: np.cos( x* ( 2. * np.pi/7) ) )
#month
df5['month_sin'] = df5['month'].apply( lambda x: np.sin( x* ( 2. * np.pi/12) ) )
df5['month_cos'] = df5['month'].apply( lambda x: np.cos( x* ( 2. * np.pi/12) ) )
#day
df5['day_sin'] = df5['day'].apply( lambda x: np.sin( x* ( 2. * np.pi/30) ) )
df5['day_cos'] = df5['day'].apply( lambda x: np.cos( x* ( 2. * np.pi/30) ) )
#week_of_year
df5['week_of_year_sin'] = df5['week_of_year'].apply( lambda x: np.sin( x * ( 2. * np.pi/52) ) )
df5['week_of_year_cos'] = df5['week_of_year'].apply( lambda x: np.cos( x * ( 2. * np.pi/52) ) )
# -
# # 6.0. PASSO 06 - DATA PREPARATION
df6 = df5.copy()
# ## 6.1. Split dataframe into training and test dataset
cols_drop = ['week_of_year', 'day', 'month', 'day_of_week', 'promo_since', 'competition_since', 'year_week']
df6 = df6.drop( cols_drop, axis=1)
df6[['store', 'date']].groupby ( 'store' ).max().reset_index() ['date'][0] - datetime.timedelta( days = 6*7)
# + code_folding=[]
#training dataset
X_train = df6[df6['date'] < '2015-06-19']
y_train = X_train['sales']
#test dataset
X_test = df6[df6['date'] > '2015-06-19']
y_test = X_test['sales']
print( 'Training Min Date: {}'.format( X_train['date'].min() ) )
print( 'Training Max Date: {}'.format( X_train['date'].max() ) )
print( '\nTest Min Date: {}'.format( X_test['date'].min() ) )
print( 'Test Max Date: {}'.format( X_test['date'].max() ) )
# -
# ## 6.2. Boruta as Feature Selector
# +
#training and test dataset for boruta
X_train_n = X_train.drop( ['date','sales'], axis=1).values
y_train_n = y_train.values.ravel()
#define RandomFOrestRegressor
rf = RandomForestRegressor( n_jobs = -1 )
#define Boruta
boruta = BorutaPy( rf, n_estimators = 'auto', verbose=2, random_state = 42).fit( X_train_n, y_train_n)
# +
cols_selected = boruta.support_.tolist()
#best features
X_train_fs = X_train.drop( ['date', 'sales'], axis = 1 )
cols_selected_boruta = X_train_fs.iloc[:,cols_selected].columns.to_list()
#not selected boruta
cols_not_selected_boruta = list(np.setdiff1d( X_train_fs.columns, cols_selected_boruta ))
# -
# ## 6.3. Manual Feature Selection
cols_not_selected_boruta = [
'is_promo',
'month_sin',
'school_holiday',
'state_holiday_christmas',
'state_holiday_easter_holiday',
'state_holiday_public_holiday',
'state_holiday_regular_day',
'year']
# +
cols_selected_boruta = [
'store',
'promo',
'store_type',
'assortment',
'competition_distance',
'competition_open_since_month',
'competition_open_since_year',
'promo2',
'promo2_since_week',
'promo2_since_year',
'competition_time_month',
'promo_time_week',
'day_of_week_sin',
'day_of_week_cos',
'week_of_year_cos',
'week_of_year_sin',
'month_cos',
'day_sin',
'day_cos']
# Columns to add
feat_to_add = ['date', 'sales']
# final features
#cols_selected_boruta.extend( feat_to_add)
# -
cols_selected_boruta
# # 7.0. PASSO 07 - MACHINE LEARNING MODELLING
x_train = X_train[ cols_selected_boruta ]
x_test = X_test[ cols_selected_boruta ]
x_test.dtypes
x_test.isna().sum()
# + active=""
# ## 7.1. Average Model
# -
from sklearn.metrics import mean_absolute_error, mean_squared_error
# +
def mean_absolute_percentage_error( y, yhat ):
return np.mean( np.abs( ( y - yhat ) / y))
def ml_error( model_name, y, yhat ):
mae = mean_absolute_error (y,yhat)
mape = mean_absolute_percentage_erro(y, yhat)
rmse = np.sqrt(mean_squared_error( y, yhat ) )
return pd.DataFrame( {
'Model Name': model_name,
'MAE' : mae,
'MAPE': mape,
'RMSE': rmse}, index = [0] )
# +
aux1= x_test.copy()
aux1['sales'] = y_test.copy()
#predictions
aux2 = aux1[['store', 'sales']].groupby('store').mean().reset_index().rename( columns ={'sales': 'predictions'} )
aux1 = pd.merge(aux1, aux2, how = 'left', on='store')
yhat_baseline = aux1['predictions']
#performance
baseline_result = ml_error( 'Average Model', np.expm1( y_test ), np.expm1( yhat_baseline ) )
baseline_result
# -
# ## 7.1. Linear Regression Model
# ## 7.1. Linear Regression Regularized Model
| n07_v01_store_sales_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/KorStats/classification-project/blob/main/result/Fq_prediciont.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="U3wrPXQ_ORLb"
# ## 구글드라이브에서 파일 읽어오기
# + [markdown] id="myLUQojpTcBX"
# **F inc2 예측**
# + id="gWN17JM7BHC7" colab={"base_uri": "https://localhost:8080/"} outputId="b1b88102-835d-4ff8-e28e-1c6536c30e20"
# !pip install mxnet
# !pip install gluonnlp pandas tqdm
# !pip install sentencepiece
# !pip install transformers==3.0.2
# !pip install torch
#깃허브에서 KoBERT 파일 로드
# !pip install git+https://git@github.com/SKTBrain/KoBERT.git@master
import torch
from torch import nn
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import Dataset, DataLoader
import gluonnlp as nlp
import numpy as np
from tqdm import tqdm, tqdm_notebook
#kobert
from kobert.utils import get_tokenizer
from kobert.pytorch_kobert import get_pytorch_kobert_model
#transformers
from transformers import AdamW
from transformers.optimization import get_cosine_schedule_with_warmup
#GPU 사용
device = torch.device("cuda:0")
#BERT 모델, Vocabulary 불러오기
bertmodel, vocab = get_pytorch_kobert_model()
# + id="dqJbzoq_OPxK" colab={"base_uri": "https://localhost:8080/"} outputId="baa90fd1-a064-4cf5-9cd4-fd1f95e0ca06"
from google.colab import drive
drive.mount('/content/drive')
# + id="wmtIZqMGOdVX" colab={"base_uri": "https://localhost:8080/"} outputId="36a4c0d0-7457-4957-caf5-3fdf44838009"
from google.colab import drive
drive.mount('/content/gdrive')
# !ln -s /content/gdrive/My\ Drive/ /mydrive
# !ls /mydrive
# + id="xV_iSLl7QFgx" colab={"base_uri": "https://localhost:8080/"} outputId="75073b96-c628-4288-cb19-35be6241223e"
# !unzip /content/gdrive/'My Drive'/산업분류자동화/'모델 개발용 자료'.zip -d /content/dataset
# !unzip /content/gdrive/'My Drive'/산업분류자동화/'실습용 자료'.zip -d /content/dataset
# + id="-lGZy4fGpXlo"
import pandas as pd
ts=pd.read_csv('/content/gdrive/MyDrive/답안 작성용 파일.csv' ,encoding = "euc-kr")
# + id="YxbNybjyIrb0"
test=ts[ts['digit_1']=='F']
test=test.reset_index(drop=True)
# + [markdown] id="l9sDYuTPPhQE"
# # 제출용 파일 만들기
# + id="PJTDOJ7PIOwS"
class BERTClassifier(nn.Module):
def __init__(self,
bert,
hidden_size = 768,
num_classes=8, ##클래스 수 조정##
dr_rate=None,
params=None):
super(BERTClassifier, self).__init__()
self.bert = bert
self.dr_rate = dr_rate
self.classifier = nn.Linear(hidden_size , num_classes)
if dr_rate:
self.dropout = nn.Dropout(p=dr_rate)
def gen_attention_mask(self, token_ids, valid_length):
attention_mask = torch.zeros_like(token_ids)
for i, v in enumerate(valid_length):
attention_mask[i][:v] = 1
return attention_mask.float()
def forward(self, token_ids, valid_length, segment_ids):
attention_mask = self.gen_attention_mask(token_ids, valid_length)
_, pooler = self.bert(input_ids = token_ids, token_type_ids = segment_ids.long(), attention_mask = attention_mask.float().to(token_ids.device))
if self.dr_rate:
out = self.dropout(pooler)
return self.classifier(out)
class BERTDataset(Dataset):
def __init__(self, dataset, sent_idx, label_idx, bert_tokenizer, max_len,
pad, pair):
transform = nlp.data.BERTSentenceTransform(
bert_tokenizer, max_seq_length=max_len, pad=pad, pair=pair)
self.sentences = [transform([i[sent_idx]]) for i in dataset]
self.labels = [np.int32(i[label_idx]) for i in dataset]
def __getitem__(self, i):
return (self.sentences[i] + (self.labels[i], ))
def __len__(self):
return (len(self.labels))
# Setting parameters
max_len = 64
batch_size = 64
warmup_ratio = 0.1
num_epochs = 5 #추후 num_epochs 증가할 필요 있음
max_grad_norm = 1
log_interval = 200
learning_rate = 5e-5
# + id="puEIYgU4I-OH" colab={"base_uri": "https://localhost:8080/"} outputId="83e33222-875c-4434-8935-d15fde534bc6"
model = torch.load('/content/gdrive/MyDrive/F_inc_model_2.pt')
model.eval()
# + id="iFIdmbZ3Ql1w" colab={"base_uri": "https://localhost:8080/"} outputId="e3ebcb45-6aec-4113-cb2a-95fcae8391e9"
#토큰화
tokenizer = get_tokenizer()
tok = nlp.data.BERTSPTokenizer(tokenizer, vocab, lower=False)
def predict(predict_sentence):
data = [predict_sentence, '0']
dataset_another = [data]
another_test = BERTDataset(dataset_another, 0, 1, tok, max_len, True, False)
test_dataloader = torch.utils.data.DataLoader(another_test, batch_size=batch_size, num_workers=5)
model.eval()
for batch_id, (token_ids, valid_length, segment_ids, label) in enumerate(test_dataloader):
token_ids = token_ids.long().to(device)
segment_ids = segment_ids.long().to(device)
valid_length= valid_length
label = label.long().to(device)
out = model(token_ids, valid_length, segment_ids)
test_eval=[]
for i in out:
logits=i
logits = logits.detach().cpu().numpy()
return np.argmax(logits)
# + [markdown] id="p67LGw4M6n2d"
# # 제출하기
# + colab={"base_uri": "https://localhost:8080/", "height": 143} id="kheYXl3WqBeT" outputId="1b1a3a56-6ff5-4854-8b89-cb9ee959591b"
test['text']=test['text_obj'].map(str)+" "+test['text_mthd'].map(str)+" "+test['text_deal'].map(str)
i=[]
for sentence in test['text']:
a=sentence.replace('nan', '')
i.append(a)
test['text']=pd.DataFrame(i)
test=test.drop(['text_obj','text_mthd', 'text_deal'], axis=1) # 기존에 있던 세개의 text변수 삭제
test.head(3)
# + id="eq6JjCO7CcZN" colab={"base_uri": "https://localhost:8080/"} outputId="6096a256-c499-4aa3-8b84-cfeb01ecd54a"
result=[]
for sentence in test['text']:
temp=predict(sentence)
result.append(temp)
# + id="S-XuWlx16jS4"
a=pd.DataFrame(result, columns=['digit_3'])
a.loc[(a['digit_3'] == 0), 'digit_3'] = 411
a.loc[(a['digit_3'] == 1), 'digit_3'] = 412
a.loc[(a['digit_3'] == 2), 'digit_3'] = 421
a.loc[(a['digit_3'] == 3), 'digit_3'] = 422
a.loc[(a['digit_3'] == 4), 'digit_3'] = 423
a.loc[(a['digit_3'] == 5), 'digit_3'] = 424
a.loc[(a['digit_3'] == 6), 'digit_3'] = 425
a.loc[(a['digit_3'] == 7), 'digit_3'] = 426
# + id="Ts01zGPNLUB0"
test['digit_3']=a
# + id="DvplvVddk7kT"
test.to_csv('result_F_inc2.csv', index=False, encoding='utf-8-sig')
# + id="7e3fPjp2Spjd"
# + id="M_Dxs9j-Spl6"
# + [markdown] id="4-3lej8MTVOR"
# **Q 예측**
# + id="qCYV6zi3Spor"
import pandas as pd
ts=pd.read_csv('/content/gdrive/MyDrive/답안 작성용 파일.csv' ,encoding = "euc-kr")
test=ts[ts['digit_1']=='Q']
test=test.reset_index(drop=True)
# + id="3vFzH1nXSprY"
class BERTClassifier(nn.Module):
def __init__(self,
bert,
hidden_size = 768,
num_classes=6, ##클래스 수 조정##
dr_rate=None,
params=None):
super(BERTClassifier, self).__init__()
self.bert = bert
self.dr_rate = dr_rate
self.classifier = nn.Linear(hidden_size , num_classes)
if dr_rate:
self.dropout = nn.Dropout(p=dr_rate)
def gen_attention_mask(self, token_ids, valid_length):
attention_mask = torch.zeros_like(token_ids)
for i, v in enumerate(valid_length):
attention_mask[i][:v] = 1
return attention_mask.float()
def forward(self, token_ids, valid_length, segment_ids):
attention_mask = self.gen_attention_mask(token_ids, valid_length)
_, pooler = self.bert(input_ids = token_ids, token_type_ids = segment_ids.long(), attention_mask = attention_mask.float().to(token_ids.device))
if self.dr_rate:
out = self.dropout(pooler)
return self.classifier(out)
class BERTDataset(Dataset):
def __init__(self, dataset, sent_idx, label_idx, bert_tokenizer, max_len,
pad, pair):
transform = nlp.data.BERTSentenceTransform(
bert_tokenizer, max_seq_length=max_len, pad=pad, pair=pair)
self.sentences = [transform([i[sent_idx]]) for i in dataset]
self.labels = [np.int32(i[label_idx]) for i in dataset]
def __getitem__(self, i):
return (self.sentences[i] + (self.labels[i], ))
def __len__(self):
return (len(self.labels))
# Setting parameters
max_len = 64
batch_size = 64
warmup_ratio = 0.1
num_epochs = 5 #추후 num_epochs 증가할 필요 있음
max_grad_norm = 1
log_interval = 200
learning_rate = 5e-5
# + colab={"base_uri": "https://localhost:8080/"} id="6-xCTTWjSpuR" outputId="a760d89e-03c6-46dd-9a96-fb827cad9b21"
model = torch.load('/content/gdrive/MyDrive/Q_inc_model.pt')
model.eval()
# + colab={"base_uri": "https://localhost:8080/"} id="du0waWajSpxA" outputId="50a3845c-df99-4ab6-bfbf-5443d0703e6c"
#토큰화
tokenizer = get_tokenizer()
tok = nlp.data.BERTSPTokenizer(tokenizer, vocab, lower=False)
def predict(predict_sentence):
data = [predict_sentence, '0']
dataset_another = [data]
another_test = BERTDataset(dataset_another, 0, 1, tok, max_len, True, False)
test_dataloader = torch.utils.data.DataLoader(another_test, batch_size=batch_size, num_workers=5)
model.eval()
for batch_id, (token_ids, valid_length, segment_ids, label) in enumerate(test_dataloader):
token_ids = token_ids.long().to(device)
segment_ids = segment_ids.long().to(device)
valid_length= valid_length
label = label.long().to(device)
out = model(token_ids, valid_length, segment_ids)
test_eval=[]
for i in out:
logits=i
logits = logits.detach().cpu().numpy()
return np.argmax(logits)
# + colab={"base_uri": "https://localhost:8080/", "height": 143} id="RhSm1rIHSpzs" outputId="725c9715-8f4c-43e9-c953-ef2a75f7b43e"
test['text']=test['text_obj'].map(str)+" "+test['text_mthd'].map(str)+" "+test['text_deal'].map(str)
i=[]
for sentence in test['text']:
a=sentence.replace('nan', '')
i.append(a)
test['text']=pd.DataFrame(i)
test=test.drop(['text_obj','text_mthd', 'text_deal'], axis=1) # 기존에 있던 세개의 text변수 삭제
test.head(3)
# + id="uEMvMMTLSp2e" colab={"base_uri": "https://localhost:8080/"} outputId="82f8e3bc-b527-443b-f153-5b812948204b"
result=[]
for sentence in test['text']:
temp=predict(sentence)
result.append(temp)
# + id="062EktyJTLFn"
a=pd.DataFrame(result, columns=['digit_3'])
a.loc[(a['digit_3'] == 0), 'digit_3'] = 861
a.loc[(a['digit_3'] == 1), 'digit_3'] = 862
a.loc[(a['digit_3'] == 2), 'digit_3'] = 863
a.loc[(a['digit_3'] == 3), 'digit_3'] = 869
a.loc[(a['digit_3'] == 4), 'digit_3'] = 871
a.loc[(a['digit_3'] == 5), 'digit_3'] = 872
# + id="tbVAjHGHTLIH"
test['digit_3']=a
# + id="vXRdipYuTLKi"
test.to_csv('result_Q.csv', index=False, encoding='utf-8-sig')
# + id="hxWoCAwtTLQV"
# + id="vSrnnHXNUWRp"
# + id="VoP_cISvWDI5"
# + id="ib0fzH2FWDMa"
# + id="aURhTonaaNvk"
# + id="ppeJY9XQaNye"
| result/Fq_prediciont.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# +
mat = np.random.normal(0, 1, size=(100, 2))
mat2 = np.random.normal(0, 1, size=(150, 2))
mat = np.dot(mat, mat2.T)
plt.matshow(mat)
plt.axis('off')
plt.savefig("mat.pdf", transparency=True)
mat = np.random.normal(0, 1, size=(100, 2))
mat2 = np.random.normal(0, 1, size=(150, 2))
mat = np.dot(mat, mat2.T)
plt.matshow(mat)
plt.axis('off')
plt.savefig("mat2.pdf", transparency=True)
# -
vec = np.random.normal(0, 1, size=(100, 1))
vec = np.hstack((vec, vec, vec))
plt.matshow(vec, cmap=plt.cm.get_cmap("tab10"))
plt.axis('off')
plt.savefig("vec.pdf", transparency=True)
plt.show()
from sklearn.datasets import make_blobs
# Not corrected
X, y = make_blobs(n_samples = 500, centers = [[0, 0], [10, 10], [ 0, 10], [10, 0]], cluster_std=1)
print("batch_no")
plt.figure(figsize=(5, 5))
plt.scatter(X[:, 0][y<=1], X[:, 1][y<= 1])
plt.scatter(X[:, 0][y> 1], X[:, 1][y> 1])
# plt.scatter(X[:, 0][y== 2], X[:, 1][y== 2])
# plt.scatter(X[:, 0][y== 3], X[:, 1][y== 3])
plt.axis("off")
plt.tight_layout()
plt.savefig("batch_no.pdf", transparency=True)
plt.show()
print("biology_no")
plt.figure(figsize=(5, 5))
plt.scatter(X[:, 0][y%2 == 1], X[:, 1][y%2 == 1], c="r")
plt.scatter(X[:, 0][y%2 == 0], X[:, 1][y%2 == 0], c="g")
# plt.scatter(X[:, 0][y== 2], X[:, 1][y== 2])
# plt.scatter(X[:, 0][y== 3], X[:, 1][y== 3])
plt.axis("off")
plt.tight_layout()
plt.savefig("bio_no.pdf", transparency=True)
# well corrected
X, y = make_blobs(n_samples = 500, centers = [[0, 0], [10, 10], [ 0, 2], [8, 10]], cluster_std=1)
print("batch_ok")
plt.figure(figsize=(5, 5))
plt.scatter(X[:, 0][y<=1], X[:, 1][y<= 1])
plt.scatter(X[:, 0][y> 1], X[:, 1][y> 1])
# plt.scatter(X[:, 0][y== 2], X[:, 1][y== 2])
# plt.scatter(X[:, 0][y== 3], X[:, 1][y== 3])
plt.axis("off")
plt.tight_layout()
plt.savefig("batch_ok.pdf", transparency=True)
plt.show()
print("biology_ok")
plt.figure(figsize=(5, 5))
plt.scatter(X[:, 0][y%2 == 1], X[:, 1][y%2 == 1], c="r")
plt.scatter(X[:, 0][y%2 == 0], X[:, 1][y%2 == 0], c="g")
# plt.scatter(X[:, 0][y== 2], X[:, 1][y== 2])
# plt.scatter(X[:, 0][y== 3], X[:, 1][y== 3])
plt.axis("off")
plt.tight_layout()
plt.savefig("bio_ok.pdf", transparency=True)
# too corrected
plt.figure(figsize=(5, 5))
X, y = make_blobs(n_samples = 500, centers = [[0, 0], [1, 1], [ 0, 2], [1, 2]], cluster_std=1)
print("batch_over")
plt.scatter(X[:, 0][y<=1], X[:, 1][y<= 1])
plt.scatter(X[:, 0][y> 1], X[:, 1][y> 1])
# plt.scatter(X[:, 0][y== 2], X[:, 1][y== 2])
# plt.scatter(X[:, 0][y== 3], X[:, 1][y== 3])
plt.axis("off")
plt.tight_layout()
plt.savefig("batch_over.pdf", transparency=True)
plt.show()
print("biology_over")
plt.figure(figsize=(5, 5))
plt.scatter(X[:, 0][y%2 == 1], X[:, 1][y%2 == 1], c="r")
plt.scatter(X[:, 0][y%2 == 0], X[:, 1][y%2 == 0], c="g")
# plt.scatter(X[:, 0][y== 2], X[:, 1][y== 2])
# plt.scatter(X[:, 0][y== 3], X[:, 1][y== 3])
plt.axis("off")
plt.tight_layout()
plt.savefig("bio_over.pdf", transparency=True)
| notebooks/.ipynb_checkpoints/Marketing-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.0 32-bit
# name: python3
# ---
# +
import numpy as np
import matplotlib as mtp
from Layer import Dense
from Activation import ReLU, SoftMax
from NeuralNetwork import NeuralNetwork
from Loss import CategoricalCrossEntropy
X = np.random.randn(100,10)
nn = NeuralNetwork()
nn.addLayer(Dense(10,16), ReLU())
nn.addLayer(Dense(16,4), SoftMax())
nn.forward(X)
nn.calculateLoss(CategoricalCrossEntropy())
print(nn.loss)
print(nn.output)
# -
| categorization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + run_control={"frozen": false, "read_only": false}
import sys
sys.path[0:0] = ['../..','../../3rdparty'] # Put these at the head of the search path
from jove.DotBashers import *
from jove.Def_md2mc import *
from jove.Def_DFA import addtosigma_dfa, totalize_dfa
# + [markdown] run_control={"frozen": false, "read_only": false}
# # This is now a universal markdown!
# + [markdown] run_control={"frozen": false, "read_only": false}
# This is now a markdown that supports all machines!
# + [markdown] run_control={"frozen": false, "read_only": false}
# # A Frequently Committee Mistake
# + [markdown] run_control={"frozen": false, "read_only": false}
# I am expecting a frequent mistake to be committed in PDA specifications. So I list this mistake and how to avoid it, first.
# + [markdown] run_control={"frozen": false, "read_only": false}
# """
# THE FOLLOWING PDA syntax is in error. The stack part should be a single string
# WITHOUT any blanks
#
# brpda = md2mc('''PDA
# I : '', '' ; S -> M
#
# M : '', S ; (S) -> M !! <--- THIS IS RIGHT
#
# M : '', S ; ( S ) -> M !! <--- THIS IS WRONG
#
# M : '', S ; SS -> M !! <--- THIS IS RIGHT
#
# M : '', S ; S S -> M !! <--- THIS IS WRONG
#
# M : '', S ; e -> M
# M : (, ( ; '' -> M
# M : ), ) ; '' -> M
# M : e, e ; '' -> M
# M : '', # ; '' -> F''')
# dotObj_pda(brpda, FuseEdges=True)
# """
#
# + [markdown] run_control={"frozen": false, "read_only": false}
# """
#
# HERE IS ANOTHER MISTAKE!
#
# THE FOLLOWING PDA syntax is in error. The stack part should be a single string
# WITHOUT any blanks
#
# brpda = md2mc('''PDA
# I : '', '' ; S -> M
#
# M : '', S ; (S) -> M
#
# M : '', S ; SS -> M
#
# M : '', S ; 'e' -> M # <--- THIS IS WRONG: no need to quote e
#
# M : (, '(' ; '' -> M # <--- THIS IS WRONG: no quotes around (
#
# M : ')', ) ; '' -> M # <--- THIS IS WRONG: no quotes around )
#
# M : e, e ; '' -> M
#
# M : '', '#' ; '' -> F''') # <--- THIS IS WRONG: no quotes around #
#
# dotObj_pda(brpda, FuseEdges=True)
#
#
# """
# + run_control={"frozen": false, "read_only": false}
brpda = md2mc('''PDA
I : '', '' ; S -> M
M : '', S ; (S) -> M
M : '', S ; SS -> M
M : '', S ; e -> M
M : (, ( ; '' -> M
M : ), ) ; '' -> M
M : e, e ; '' -> M
M : '', # ; '' -> F''')
dotObj_pda(brpda) # <-- Ugh, without FuseEdges as a parameter, the drawing is a mess...
# + run_control={"frozen": false, "read_only": false}
brpda = md2mc('''PDA
I : '', '' ; S -> M
M : '', S ; (S) -> M
M : '', S ; SS -> M
M : '', S ; e -> M
M : (, ( ; '' -> M
M : ), ) ; '' -> M
M : e, e ; '' -> M
M : '', # ; '' -> F''')
dotObj_pda(brpda, FuseEdges=True) # <-- what a difference FuseEdges makes !!
# + [markdown] run_control={"frozen": false, "read_only": false}
# # We begin with several examples
# + run_control={"frozen": false, "read_only": false}
ev0end1 = md2mc('''
DFA
I : 0 -> A
A : 0 | 1 -> I
I : 1 -> F
F : 0 | 1 -> I
''')
dotObj_dfa(ev0end1)
# + run_control={"frozen": false, "read_only": false}
ev0end1plus = addtosigma_dfa(ev0end1, {'2'})
ev0end1plus
# + run_control={"frozen": false, "read_only": false}
dotObj_dfa_w_bh(totalize_dfa(ev0end1plus))
# + run_control={"frozen": false, "read_only": false}
dotObj_dfa_w_bh(totalize_dfa(ev0end1plus), FuseEdges=True)
# + run_control={"frozen": false, "read_only": false}
third1dfa=md2mc(src="File", fname="machines/dfafiles/thirdlastis1.dfa")
dotObj_dfa(third1dfa)
# + run_control={"frozen": false, "read_only": false}
# An NFA with multiple initial states
nfaMultiQ0 = md2mc('''
NFA
I0 : a | b | c -> A, B
I0 : c -> F
I1 : a | b -> A, B
A : c -> F
B : d -> F
''')
dotObj_nfa(nfaMultiQ0)
# + run_control={"frozen": false, "read_only": false}
nfa_ends0101 = md2mc(src="File", fname="machines/nfafiles/endsin0101.nfa")
dotObj_nfa(nfa_ends0101)
# + run_control={"frozen": false, "read_only": false}
# generates syntax error correctly: md2mc(src="File", fname="machines/pdafiles/erroneous3.pda")
# + [markdown] run_control={"frozen": false, "read_only": false}
# # Code for DFA Markdown
# + run_control={"frozen": false, "read_only": false}
nfa_ends0101 = md2mc(src="File", fname="machines/nfafiles/endsin0101.nfa")
# + run_control={"frozen": false, "read_only": false}
nfa_ends0101
# + run_control={"frozen": false, "read_only": false}
dotObj_nfa(nfa_ends0101)
# + [markdown] run_control={"frozen": false, "read_only": false}
# __What's needed now : More testing, more use!__
# + run_control={"frozen": false, "read_only": false}
last1dfa=md2mc('''
NFA
I : 0 -> I
I : 1 -> F
F : 0 -> I
F : 1 -> F
''')
# + run_control={"frozen": false, "read_only": false}
second1dfa=md2mc(src="File", fname="machines/dfafiles/secondlastis1.dfa")
# + run_control={"frozen": false, "read_only": false}
second1dfa_do = dotObj_dfa(second1dfa)
second1dfa_do
# + run_control={"frozen": false, "read_only": false}
second1dfa_do.source
# + run_control={"frozen": false, "read_only": false}
third1dfa=md2mc(src="File", fname="machines/dfafiles/thirdlastis1.dfa")
# + run_control={"frozen": false, "read_only": false}
third1dfa
# + run_control={"frozen": false, "read_only": false}
third1dfa_do = dotObj_dfa(third1dfa)
# + run_control={"frozen": false, "read_only": false}
third1dfa_do.source
# + run_control={"frozen": false, "read_only": false}
pdasip=md2mc(src="File", fname="machines/pdafiles/f27sip.pda")
# + run_control={"frozen": false, "read_only": false}
pdasip
# + run_control={"frozen": false, "read_only": false}
dotObj_pda(pdasip)
# + run_control={"frozen": false, "read_only": false}
dotObj_pda(pdasip, True)
# + run_control={"frozen": false, "read_only": false}
wwndtm=md2mc(src="File", fname="machines/tmfiles/wwndtm.tm")
# + run_control={"frozen": false, "read_only": false}
wwndtm
# + run_control={"frozen": false, "read_only": false}
dotObj_tm(wwndtm)
# + run_control={"frozen": false, "read_only": false}
wpwtm = '''
TM
!!---------------------------------------------------------------------------
!! This is a DTM for recognizing strings of the form w#w where w is in {0,1}*
!! The presence of the "#" serves as the midpoint-marker, thus allowing the
!! TM to deterministically match around it.
!!
!!---------------------------------------------------------------------------
!!---------------------------------------------------------------------------
!! State : rd ; wr , mv -> tostates !! comment
!!---------------------------------------------------------------------------
Iq0 : 0 ; X , R -> q1 !! All 0s are converted to X, and matching
!! 0s are then sought to the right of the #
Iq0 : 0 ; Y , R -> q7 !! All 1s are converted to Y, and matching
!! 1s are then sought to the right of the #
Iq0 : # ; # , R -> q5 !! If we see # rightaway, we are in the
!! situation of having to match eps # eps
q1 : 0 ; 0,R | 1 ; 1,R -> q1 !! In q1, skip over the remaining 0s and 1s
q1 : # ; # , R -> q2 !! But upon seeing a #, look for a matching
!! 0 (since we are in q2, we know this).
q2 : X ; X,R | Y ; Y,R -> q2 !! All X and Y are "past stuff" to skip over
q2 : 0 ; X , L -> q3 !! When we find a matching 0, turn that to
!! an X, and sweep left to do the next pass
q3 : X ; X,L | Y ; Y,L -> q3 !! In q3, we move over all past X, Y
q3 : # ; # , L -> q4 !! but when we reach the middle marker, we
!! know that the next action is to seek the
!! next unprocessed 0 or 1
q4 : 0 ; 0,L | 1 ; 1,L -> q4 !! In q4, wait till we hit the leftmost 0/1
q4 : X ; X,R | Y ; Y,R -> Iq0 !! When we hit an X or Y, we know that we've
!! found the leftmost 0/1. Another pass begins.
q5 : X ; X,R | Y ; Y,R -> q5 !! In q5, we skip over X and Y (an equal number
!! of X and Y lie to the left of the #)
q5 : . ; . , R -> Fq6 !! .. and we accept when we see a blank (.)
q7 : 0 ; 0,R | 1 ; 1,R -> q7 !! q7 is similar to q1
q7 : # ; # , R -> q8 !! and q8 is similar to q2
q8 : X ; X,R | Y ; Y,R -> q8
q8 : 0 ; X , L -> q9 !! and q9 is similar to q3
q9 : X ; X,L | Y ; Y,L -> q9 !! In q9, we move over all past X, Y
q9 : # ; # , L -> q10 !! and q10 is similar to q4
q10 : 0 ; 0,L | 1 ; 1,L -> q10 !! In q10, wait till we hit the leftmost 0/1
q10 : X ; X,R | Y ; Y,R -> Iq0 !! When we hit an X or Y, we know that we've
!! found the leftmost 0/1. Another pass begins.
!!---------------------------------------------------------------------------
!! You may use the line below as an empty shell to populate for your purposes
!! Also serves as a syntax reminder for entering DFAs.
!!
!! State : r1 ; w1 , m1 | r2 ; w2 , m2 -> s1 , s2 !! comment
!!
!! .. : .. ; .. , .. | .. ; .. , .. -> .. , .. !! ..
!!---------------------------------------------------------------------------
!!
!! Good commenting and software-engineering methods, good clean indentation,
!! grouping of similar states, columnar alignment, etc etc. are HUGELY
!! important in any programming endeavor -- especially while programming
!! automata. Otherwise, you can easily make a mistake in your automaton
!! code. Besides, you cannot rely upon others to find your mistakes, as
!! they will find your automaton code impossible to read!
!!
!!---------------------------------------------------------------------------
'''
# + run_control={"frozen": false, "read_only": false}
wpwmd = md2mc(wpwtm)
# + run_control={"frozen": false, "read_only": false}
do_wpwmd = dotObj_tm(wpwmd)
# + run_control={"frozen": false, "read_only": false}
#do_wpwmd.source
# + run_control={"frozen": false, "read_only": false}
wwndtm_do=dotObj_tm(wwndtm)
# + run_control={"frozen": false, "read_only": false}
#wwndtm_do.source
# + run_control={"frozen": false, "read_only": false}
do_wpwmd
# + run_control={"frozen": false, "read_only": false}
ev0end1 = md2mc('''
DFA
I : 0 -> A
I : 1 -> F
A : 0 | 1 -> I
F : 0 | 1 -> I
''')
# + run_control={"frozen": false, "read_only": false}
dotObj_dfa(ev0end1)
# + run_control={"frozen": false, "read_only": false}
pedpda = md2mc(src="File",fname="machines/pdafiles/f27sip.pda")
dotObj_pda(pedpda)
# + run_control={"frozen": false, "read_only": false}
dotObj_pda(pedpda)
# + run_control={"frozen": false, "read_only": false}
pedpda
# + run_control={"frozen": false, "read_only": false}
pedpda = md2mc(src="File", fname="machines/pdafiles/pedagogical2.pda")
dotObj_pda(pedpda)
# + run_control={"frozen": false, "read_only": false}
dotObj_tm(md2mc(src="File", fname="machines/tmfiles/shift_left_tm.tm"))
# + run_control={"frozen": false, "read_only": false}
dotObj_tm(md2mc(src="File", fname="machines/tmfiles/shift_right_tm.tm"))
# + run_control={"frozen": false, "read_only": false}
dotObj_tm(md2mc(src="File", fname="machines/tmfiles/decimal_double_tm.tm"))
# + run_control={"frozen": false, "read_only": false}
dotObj_tm(md2mc(src="File", fname="machines/tmfiles/decimal_double_tm.tm"), FuseEdges=True)
# + run_control={"frozen": false, "read_only": false}
dotObj_tm(md2mc(src="File", fname="machines/tmfiles/collatz_tm.tm"))
# + run_control={"frozen": false, "read_only": false}
dotObj_tm(md2mc(src="File", fname="machines/tmfiles/collatz_tm.tm"), FuseEdges = True)
# + run_control={"frozen": false, "read_only": false}
brpda = md2mc('''PDA
I : '', '' ; S -> M
M : '', S ; (S) -> M
M : '', S ; SS -> M
M : '', S ; e -> M
M : (, ( ; '' -> M
M : ), ) ; '' -> M
M : e, e ; '' -> M
M : '', # ; '' -> F''')
dotObj_pda(brpda, FuseEdges=True)
# + run_control={"frozen": false, "read_only": false}
| notebooks/driver/Drive_md2mc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Análisis de cambio climático
# ## Introducción
#
# En este documento se analizará el cambio climatico a traves de las variaciones de las temperaturas mundiales desde 1970, ademas se incluiran distintas variables que puedan ser de utilidad para generar un analisis más profundo, de este modo dar un mejor entendiento de los factores que se afectan al calentamiento global.
#
# Los datos son obtenidos a traves de:
# - https://www.kaggle.com/berkeleyearth/climate-change-earth-surface-temperature-data?select=GlobalLandTemperaturesByCountry.csv
# - https://data.worldbank.org/
#
# Lo siguientes análisis contendrán comentarios de los que se esta realizando para que cualquiera pueda entenderlo.
# ## Cargando datos
# Importamos las librerias a usar en este análisis, las cuales son:
# - Pandas: Ayuda a transformar y tratar los archivos separados por coma (csv) en paneles de datos como tablas de excel
# - Numpy: Libreria que nos permite hacer operaciones matematicas y estaditicas de manera más sencilla
# - Seaborn: Permite generar graficas llamativas con los datos obtenidos
import pandas as pd
import numpy as np
import seaborn as sns
import os
# Usamo la libreia **OS** para cambiar el directorio, de este modo accedemos a la carpeta **data** para cargar los datos
os.chdir('/Users/angel/Desktop/proyectos/c_global/data')
os.getcwd()
print(os.listdir('.'))
# Leemos los datos de la temperatura registradas a nivel global
df = pd.read_csv('GlobalLandTemperaturesByCountry.csv')
df
# ### Descripción del Dataset principal
#
# Aquí podemos ver las dimensiones del dataset:
# - Más de 50.000 filas
# - 4 columnas
# - Contiene 2 datos tipo "objeto"
# - Contiene 2 datos tipos "float" (número)
# - La media de las temperaturas es de 17.19
# - La mediana de las temperaturas es de 20.09
# - Hay datos de 243 paises
# - Los datos son tomados desde el año 1743 al 2013. Solamente se usaran los datos desde 1970 en adelante
#
df.shape
df.info()
df.describe()
len(df['Country'].unique())
df['Country'].unique()
# Cambiamos el tipo de dato de las fechas con el comando *to_datetime()*, con esto nos aseguramos que se encuentre en el formato correcto. Esto loa añadimos a otra variable que se llamara "year".
df['year']= pd.to_datetime(df['dt'])
df['year']
idx = df['year'] > '1970-01-01'
df = df[idx]
df
# #### Agrupación y Agregación
# Los datos se agrupan por país con el metodo *groupby*, estos se sub agrupan a traves de las fechas con el metodo *Grouper* que es especial para agrupar por fechas, se le da una frecuencia de agrupación de 1 año con *freq=1*
#
# El metodo *agg* nos sirve para poder agregar nuevas funciones, ya sea a traves de funciones lambdas o a traves de numpy. En este aso se agregro un diccionario con el nombre, calculando con numpy la media y la mediana
df_temp_avg = df.groupby(['Country',
pd.Grouper(key='year', freq='1Y')
]).agg({'AverageTemperature':[np.mean,np.median]})
df_temp_avg
# Aqui podemos observar un análisis de las temperaturas medias (promedio y mediana) de Chile
df_temp_avg.xs('Chile')['AverageTemperature'].plot(figsize = (10,10))
# ### Generando gráficas (Parte 1)
#
# A continuación se presentaran distintas gráficas, alguna de ellas solamente para mostrar el poder del las librerias y otras para comenzar a obtener insight significativos sobre los datos.
#
# Cabe destacar que en esta sección solo se vera el primer dataset y luego se incorporarán los siguientes.
#
# El dataframe se manipulará agregando la media de las temperaturas por año, se resetea el índice y de este modo obtener un dataframe que pueda ser útil
df_temp_med = df_temp_avg['AverageTemperature'][['median']].reset_index()
df_temp_med
df_temp_med['date'] = df_temp_med['year'].dt.year
df_temp_med['date']
df_temp_med.rename(columns={'median':'temperature'},inplace=True)
df_temp_med
# #### Tablas Dinámicas con Pandas
#
# - Se usan con el metodo *pivot_table*
# - Se le ingresan los valores con la variable *values*
# - Las filas se ingresan con la variable *index*
# - Las columnas se ingresan con la variable *columns*}
# - Estas funcionan de la misma forma que las tablas dinámicas de Excel
# - Nos permiten generar graficas mejores
df_temp_pivot = df_temp_med.pivot_table(values='temperature',index='date',columns='Country')
df_temp_pivot.head()
# Esta grafica de caja solo nos muestra las posibilidades que tiene Pandas para graficar las tablas dinámicas.
df_temp_pivot.boxplot(figsize=(10,10), rot = 45)
df_temp_pivot.T
df_temp_pivot.T.sample(5)
df_temp_pivot.T.sample(5).T.boxplot(figsize=(12,15), rot=45)
df_agri = pd.read_csv('API_AG.LND.AGRI.K2_DS2_en_csv_v2_716226.csv', header = 2)
df_fore = pd.read_csv('API_AG.LND.FRST.K2_DS2_en_csv_v2_716262.csv', header = 2)
df_elec = pd.read_csv('API_EG.USE.ELEC.KH.PC_DS2_en_csv_v2_715482.csv', header = 2)
df_co2e = pd.read_csv('API_EN.ATM.CO2E.KT_DS2_en_csv_v2_713263.csv', header = 2)
df_popu = pd.read_csv('API_SP.POP.TOTL_DS2_en_csv_v2_713131.csv', header = 2)
df_popu.describe()
range(1971,2015)
map(str,range(1971,2015))
list(map(str,range(1971,2015)))
cols = ['Country Name','Country Code'] + list(map(str,range(1971,2015)))
cols[:5]
df_agri.loc[:,cols].melt(id_vars=['Country Name','Country Code']).rename(
columns={'variable':'date',
'Country Name':'Country',
'Country Code':'name',
'value':'agriculture'})
def fun_format(df,col='agriculture'):
return df.loc[:,cols].melt(id_vars=['Country Name','Country Code']).rename(
columns={'variable':'date',
'Country Name':'Country',
'Country Code':'name',
'value':col})
df_agri = fun_format(df_agri,col='agriculture')
df_fore = fun_format(df_fore,col='forestal')
df_elec = fun_format(df_elec,col='electricprod')
df_co2e = fun_format(df_co2e,col='co2')
df_popu = fun_format(df_popu,col='population')
df_popu
df_temp_med
df_popu.info()
df_temp_med.info()
df_popu['date'] = df_popu['date'].astype(float)
df_fore['date'] = df_fore['date'].astype(float)
df_elec['date'] = df_elec['date'].astype(float)
df_co2e['date'] = df_co2e['date'].astype(float)
df_agri['date'] = df_agri['date'].astype(float)
df_merge = pd.merge(df_temp_med[['Country','temperature','date']],
df_popu, on = ['Country', 'date'],how = 'inner')
# +
df_merge = pd.merge(df_merge,
df_fore,
on = ['Country','name', 'date'],
how = 'inner')
df_merge = pd.merge(df_merge,
df_elec,
on = ['Country','name', 'date'],
how = 'inner')
df_merge = pd.merge(df_merge,
df_co2e,
on = ['Country','name', 'date'],
how = 'inner')
df_merge = pd.merge(df_merge,
df_agri,
on = ['Country','name', 'date'],
how = 'inner')
df_merge
# -
df_climate = df_merge.dropna()
pd.set_option('display.float_format','{:,.1f}'.format)
df_climate.groupby('Country')['co2'].median().sort_values(ascending=False)
first_10_total = df_climate.groupby('Country')['co2'].median().sort_values(ascending=False).head(10)
first_10_total
first_10_index = df_climate.groupby('Country')['co2'].median().sort_values(ascending=False).head(10).index
first_10_index
df_max_co2=df_climate[df_climate['Country'].isin(first_10_index)]
df_max_co2
df_topten = df_max_co2.groupby('Country').mean()
df_max_co2.set_index('Country').plot.scatter(
x='population',
y='co2',
c='date',
colormap = 'viridis',
figsize=(15,10)
)
sns.heatmap(df_topten.reset_index().corr(),annot=True)
# +
scatter_matrix(df_topten.reset_index(),diagonal='kde', figsize=(20,10))
plt.suptitle('scatter-matrix')
plt.show()
# -
sns.heatmap(df_climate.corr(),annot = True)
df_med = df_climate.groupby('date').median()
df_med
sns.heatmap(df_med.reset_index().corr(),annot=True)
from pandas.plotting import scatter_matrix
import matplotlib.pyplot as plt
# +
scatter_matrix(df_med.reset_index(),diagonal='kde', figsize=(20,10))
plt.suptitle('scatter-matrix')
plt.show()
# -
from pandas.plotting import parallel_coordinates
df_samerica = df_climate[
df_climate['Country'].isin(['Colombia',
'Argentina',
'Bolivia',
'Mexico',
'Peru',
'Chile'])
][['Country',
'temperature',
'co2',
'agriculture',
'forestal']]
parallel_coordinates(df_samerica,'Country', colormap='jet')
df_climate.info()
df_climate.groupby(['Country']).median()
sns.set(color_codes=True)
sns.regplot(x="electricprod", y="co2", data=df_climate.groupby(['Country']).median());
sns.lmplot(x="electricprod", y="co2", data=df_climate.groupby(['Country']).median());
| .ipynb_checkpoints/Data Analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Azure-Sentinel-API-Calls" data-toc-modified-id="Azure-Sentinel-API-Calls-1">Azure Sentinel API Calls</a></span><ul class="toc-item"><li><ul class="toc-item"><li><span><a href="#Description" data-toc-modified-id="Description-1.0.1">Description</a></span></li><li><span><a href="#Installation-and-imports" data-toc-modified-id="Installation-and-imports-1.0.2">Installation and imports</a></span></li><li><span><a href="#Authentication" data-toc-modified-id="Authentication-1.0.3">Authentication</a></span></li></ul></li></ul></li></ul></div>
# -
# # Azure Sentinel API Calls
#
# MSTICpy versions > 0.8.5
#
# ### Description
#
# This Notebook provides an example of using the Azure Sentinel API features of MSTICpy in order retrieve specific data from Azure Sentinel
#
# ### Installation and imports
# %pip install --upgrade msticpy[azsentinel]
from msticpy.data.azure_sentinel import AzureSentinel
import msticpy.nbtools.nbwidgets as widgets
from msticpy.data import data_obfus as mask
# ### Authentication
# The first step to be able to use the features is to call the AzureSentinel class and connect to it. Authentication uses the standardized Azure authentication options of using environment variables, Azure CLI credentials, Managed Identities, and interactive logons.
azs = AzureSentinel()
azs.connect()
# Once connected we need to select an Azure Sentinel workspace to get details from. The easies way to do this is with the get_subscriptions() and get_sentinel_workspaces() functions to select the subscription and workspace you with to connect to. If you already know which workspace you wish to connect to you can skip straight to the other functions and enter these details.
# +
# Query for our subscriptions
subs = azs.get_subscriptions()
subs = subs.mp_obf.obfuscate(column_map={"Display Name": "str"})
# Display subscriptions (masked names) in a pick list
print("Select a subscription:")
sub = widgets.SelectItem(
item_list=subs['Display Name'].to_list(),
auto_display=True
)
# -
# Get the subscription ID
sub_id = subs[subs['Display Name'] == sub.value].iloc[0]['Subscription ID']
# Query for workspaces in that subscription
workspaces = azs.get_sentinel_workspaces(sub_id = sub_id)
# Display workspaces in a list
print("Select an Azure Sentinel Workspace:")
ws = widgets.SelectItem(
item_dict=workspaces,
auto_display=True
)
# Now that we have selected our workspace we can call various functions to get details about content in the workspace. These are typically returned as DataFrames. Below we get a list of hunting queries configured in our workspace.
queries = azs.get_hunting_queries(ws.value)
queries.head().drop(columns=["id", "etag", "name"])
# Hunting queries return the raw queries associated with them, this allows us to pass the query directly to a QueryProvider in order to get the results of the hunting query within the notebook.
from msticpy.data.data_providers import QueryProvider
from msticpy.common.wsconfig import WorkspaceConfig
qry_prov = QueryProvider('LogAnalytics')
wkspace = WorkspaceConfig()
qry_prov.connect(wkspace.code_connect_str)
qry_prov.exec_query(queries['properties.Query'].iloc[2])
# We can also get a list of configured alert rules:
alert_rules = azs.get_alert_rules(ws.value)
alert_rules.head().drop(columns=["id", "etag", "name"])
# We can also get a list of saved bookmarks. To see the events these bookmarks relate to you can pass the query value to a QueryProvider.
bkmarks = azs.get_bookmarks(ws.value)
bkmarks.head().drop(columns=["id", "etag", "name"])
# We can also interact with Incidents via the API to get a set of all incidents, or a single incident:
incidents = azs.get_incidents(res_id=ws.value)
display(incidents.head())
incident = azs.get_incident(incident_id = incidents.iloc[0]['name'] , res_id=ws.value)
display(incident)
# You can also interact with an incident - adding comments or changing properties such as severity or status:
azs.post_comment(incident_id = incident.iloc[0]['name'], comment="This is a test comment", res_id=ws.value)
azs.update_incident(incident_id = incident.iloc[0]['name'], update_items={"severity":"High"}, res_id=ws.value)
| docs/notebooks/AzureSentinelAPIs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Now You Code 4: Email Harvest Training
#
# Let's teach you how to extract emails from text. This has a variety of applications. The most common being buiding a list of emails for spamming... er, I meant "mass marketing."
#
# The best way to find the emails in the mailbox file is to search for lines in the file that begin with "From: (similar to what we did in the lab). When you find an email write just the email address (not "From:" and the address) to "NYC4-emails.txt", and don't worry about duplicates.
#
# The program should print the number of emails it wrote to the file.
#
# Example Run:
#
# ```
# Wrote 27 emails to NYC4-emails.txt
# ```
# ## Step 1: Problem Analysis
#
# Input: (No user input, but we do read from the file `NYC4-mbox-short.txt`
#
# Output (The number of emails collected from the file, and a new file `NYC4-emails.txt` with one email per line.
#
# Algorithm:
#
# ```
# (write here)
# ```
#
# +
## Step 2: Write the code
# +
## Step 3: Write another program to read the emails from NYC4-emails.txt and print them to the console
# -
# ## Step 4: Questions
#
# 1. Does this code actually "detect" emails? How does it find emails in the text?
#
# Answer:
#
#
# 2. Explain how this program can be improved to prompt for an email file at runtime?
#
# Answer:
#
#
# 3. Devise an approach to remove duplicate emails from the output file. You don't have to write as code, just explain it.
#
# Answer:
# ## Step 5: Reflection
#
# Reflect upon your experience completing this assignment. This should be a personal narrative, in your own voice, and cite specifics relevant to the activity as to help the grader understand how you arrived at the code you submitted. Things to consider touching upon: Elaborate on the process itself. Did your original problem analysis work as designed? How many iterations did you go through before you arrived at the solution? Where did you struggle along the way and how did you overcome it? What did you learn from completing the assignment? What do you need to work on to get better? What was most valuable and least valuable about this exercise? Do you have any suggestions for improvements?
#
# To make a good reflection, you should journal your thoughts, questions and comments while you complete the exercise.
#
# Keep your response to between 100 and 250 words.
#
# `--== Write Your Reflection Below Here ==--`
#
#
| content/lessons/07/Now-You-Code/NYC4-Email-Harvest-Training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # **Chapter 03 시계열 모델링**
# 정상정 **(Stationary)** 의 개념을 분석한다
# +
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
import warnings
# plt.style.use('seaborn')
# plt.rcParams['figure.dpi'] = 300
warnings.simplefilter(action='ignore', category=FutureWarning)
# -
# ## **1 시계열 데이터 분해**
# ### **01 Loading the DataSet**
# - 'GC=F' : Gold Price Code
# +
DATA_FILENAME = 'data/stock-amazon.pkl'
# Loading the DataSet
import pandas as pd
import yfinance as yf
try:
data = pd.read_pickle(DATA_FILENAME)
except FileNotFoundError:
data = yf.download('AMZN', start='2010-01-01', end='2022-01-01',
progress=False, auto_adjust=True)
data.to_pickle(DATA_FILENAME)
data.head(3)
# -
# ### **02 Data Pre Processing**
# +
# Data Pre Processing
data.rename(columns={'Close':'price'}, inplace=True)
df = data[['price']]
df = df.resample('M').last() # 월간 데이터로 변환하기
# 이동평균과 표준편차를 추가 한다
WINDOW_SIZE = 12
df['rolling_mean'] = df.price.rolling(window=WINDOW_SIZE).mean()
df['rolling_std'] = df.price.rolling(window=WINDOW_SIZE).std()
# -
# ### **03 Visualization**
# ```python
# import cufflinks as cf
# from plotly.offline import iplot, init_notebook_mode
# cf.set_config_file(world_readable=True, theme='pearl', offline=True) # set up settings (run it once)
# init_notebook_mode(connected=True) # initialize notebook display
# df.iplot(title="Gold Price") # Plotly from DataFrame
# ```
# Visualization
plt.rcParams['figure.figsize'] = [20, 5]
df.plot(title='Gold Price')
plt.show()
# ### **04 승산적 모델을 활용하여 계절성 특징을 찾는다**
# +
from statsmodels.tsa.seasonal import seasonal_decompose
plt.rcParams['figure.figsize'] = [20, 8]
decomposition_results = seasonal_decompose(df.price, model='multiplicative')
decomposition_results.plot().suptitle('Multiplicative Decomposition', fontsize=18)
plt.show()
# -
# ## **2 ProPhet 을 활용한 시계열 데이터 분해**
#
# [SettingWithCopyWarning: DataFrame 에 대해서](https://emilkwak.github.io/pandas-dataframe-settingwithcopywarning)
#
# https://github.com/facebook/prophet
# ```r
# # ! pip install pystan
# # ! pip install fbprophet
# ```
# ### **01 Loading the DataSet**
# 위 아마존 데이터를 시계열 분석한 뒤, 1년 변화를 예측합니다
df = data[['price']]
df.head(3)
df.reset_index(drop=False, inplace=True)
df.head(3)
df.rename(columns={'Date':'ds', 'price':'y'}, inplace=True)
df.head(3)
| stock-books/03-Timeseries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:Python 2]
# language: python
# name: conda-env-Python 2-py
# ---
# +
from numpy.linalg import inv
import numpy as np
from scipy.linalg import eig
from sklearn.datasets import make_blobs
from sklearn.metrics import pairwise_distances
from diffmaps_util import k, diag
# -
# Diffusion Distance <br />
# A distance function between any two points based on the random walk on the graph [1].
#
#
# Diffusion map <br />
# Low dimensional description of the data by the first few eigenvectors [1].
#
n = 3
X, y = make_blobs(n_samples=n, cluster_std=.1, centers=[[1,1]])
X
# Define a pairwise similarity matrix between points...
L = k(X, .9)
# and a diagonal normalization matrix $D_{i,i} = \sum_j L_{i,j}$
D = diag(L)
# Matrix M <br />
# $M = D^{-1}L$
M = inv(D).dot(L)
# The matrix M is adjoint to a symmetric matrix <br />
# $M_s = D^{1/2}MD^{-1/2}$
#
# M and M<sub>s</sub> share the same eigenvalues. <br />
# Since M<sub>s</sub> is symmetric, it is diagonalizable and has a set of _n_ real eigenvalues {$\lambda_{j=0}^{n-1}$} whose corresponding eigenvectors form an orthonormal basis of $\mathbf{R}^n$. <br />
# The left and right eigenvectors of M, denoted $\phi_j$ and $\psi_j$ are related to those of M<sub>s</sub>.
#
# $$ \phi_j = \mathbf{v}_j D^{1/2}, \psi_j = \mathbf{v}_j D^{-1/2} $$
Ms = diag(D, .5).dot(M).dot(diag(D,-.5))
# ---
# Now we utilize the fact that by constrution M is a stochastic matrix
p0 = np.eye(n)
# *The stationary probability distribution $\Phi_0$ *
e = p0
for i in range(1000):
e = e.dot(M)
print e
p1 = p0.dot(M)
p1
w, v = eig(M)
w = w.real
print w
print v
# sorting the eigenvalues and vectors
temp = {_:(w[_], v[:,_]) for _ in range(len(w))}
w = []
v = []
for _ in sorted(temp.items(), key=lambda x:x[1], reverse=True):
w.append(_[1][0])
v.append(_[1][1])
w = np.array(w)
v = np.array(v).T
print w
print v
psi = v / v[:,0]
print psi
# ## Diffusion Map
# $$ \Psi_t(x) = (\lambda_1^t\psi(x), \lambda_2^t\psi(x), ..., \lambda_k^t\psi(x)) $$
diffmap = (w.reshape(-1,1) * psi.T).T[:,1:]
diffmap
# ## Diffusion Distance
# Defined by Euclidean distance in the diffusion map
# $$ D_t^2(x_0, x_1) = ||\Psi_t(x_0) - \Psi_t(x_1)||^2 $$
dt0 = pairwise_distances(diffmap)**2
dt0
# ## Diffusion Distance [2]
# Defined by probability distribution on time _t_.
# $$
# D_t^2(x_0, x_1) = ||p(t, y|x_0) - p(t, y|x_1)||_w^2 \\
# = \sum_y (p(t, y|x_0) - p(t, y|x_1))^2 w(y)
# $$
dt = []
for i in range(n):
_ = []
for j in range(n):
_.append(sum((p1[i]-p1[j])**2 / v[:,0]**2))
dt.append(_)
dt = np.array(dt)
dt
(dt0 - dt)
print M
M.sum(axis=1)
w, v = eig(M)
w = w.real
print w
print v
p0*w[0]*v[:,0]**2 + p0*w[1]*v[:,1]**2 + p0*w[2]*v[:,2]**2
| subjects/diffusion maps/Diffusion Maps 02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="GG3_2HyRVY4Y" tags=["header", "comment"]
# # Introduction
#
# This guide will show you how to:
#
# * Initialize Neptune and create a `run`,
# * Log `altair` charts to Neptune.
# + [markdown] id="1tBfsYCm05rl"
# # Before you start
#
# Install `neptune-client` and `altair`
# + id="xm0WX77L1Nii"
# ! pip install altair==4.1.0 neptune-client
# + [markdown] id="0NQaVXuoWA-E" tags=["header", "installation"]
# # Import and initialize `neptune`
# + id="quJ_tJDk7R3e"
import neptune.new as neptune
# + id="asktsEotnTlR"
run = neptune.init(api_token="<PASSWORD>", project="common/altair-support")
# + [markdown] id="VnSWXRbXHUz3" tags=["comment", "exclude"]
# Click on the link above to open this run in Neptune. For now it is empty but keep the tab with run open to see what happens next.
# + [markdown] id="QYakxenbPkrR"
# **Note:**<br>
# To [create your own private projects](https://docs.neptune.ai/administration/workspace-project-and-user-management/projects#create-project), you will need a [Neptune account](https://neptune.ai/register) and your [personal Neptune API token](https://docs.neptune.ai/getting-started/installation#get-api-token).
# + [markdown] id="pk4ZgmW9Q7Mg" tags=["header"]
# # Log `altair` charts to Neptune
# + [markdown] id="E3XjDiMB-XDK"
# ## Create a sample chart
# + id="0mEZs-60mhxc"
# ! pip install vega_datasets==0.9.0
# + id="V6i1BJfL96ww"
import altair as alt
from vega_datasets import data
source = data.cars()
brush = alt.selection(type="interval")
points = (
alt.Chart(source)
.mark_point()
.encode(
x="Horsepower:Q",
y="Miles_per_Gallon:Q",
color=alt.condition(brush, "Origin:N", alt.value("lightgray")),
)
.add_selection(brush)
)
bars = (
alt.Chart(source)
.mark_bar()
.encode(y="Origin:N", color="Origin:N", x="count(Origin):Q")
.transform_filter(brush)
)
chart = points & bars
chart
# + [markdown] id="tqqF7gsP_Sh9"
# ## Log interactive image
# + id="8WWCJy73_Sh_"
run["interactive_img"].upload(neptune.types.File.as_html(chart))
# + [markdown] id="XSHLD6AkI7LW"
# # Stop logging
#
# <font color=red>**Warning:**</font><br>
# Once you are done logging, you should stop tracking the run using the `stop()` method.
# This is needed only while logging from a notebook environment. While logging through a script, Neptune automatically stops tracking once the script has completed execution.
# + id="Zgh3NwDoJAuG"
run.stop()
# + [markdown] id="swXKaHZmASyd"
# # Explore the charts in Neptune
#
# The images can be found in the **All metadata** section
# 
# + [markdown] id="had9MNRtQ7Mr" tags=["comment"]
# # Conclusion
#
# You’ve learned how to:
#
# * Initialize Neptune and create a run,
# * Log `altair` charts to Neptune.
# + [markdown] id="_3ibcYhPQ7Mr" tags=["comment"]
# # What's next
#
# Now that you know how to log charts to Neptune, you can learn:
#
# * [How to log other types of metadata to Neptune](https://docs.neptune.ai/you-should-know/logging-and-managing-runs-results/logging-runs-data#what-objects-can-you-log-to-neptune)
# * [How to download runs data from Neptune](https://docs.neptune.ai/user-guides/logging-and-managing-runs-results/downloading-runs-data)
# * [How to connect Neptune to the ML framework you are using](https://docs.neptune.ai/essentials/integrations)
| integrations-and-supported-tools/altair/notebooks/Neptune_Altair_Support.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 编译序列重要采样
#
# **Compiled Sequential Importance Sampling:**
#
# 编译序列重要性采样[1]或推理编译是一种通过学习重要性抽样的建议分布来减少推断的计算成本的技术。
# Compiled sequential importance sampling [1], or inference compilation, is a technique to amortize the computational cost of inference by learning a proposal distribution for importance sampling.
# The proposal distribution is learned to minimise the KL divergence between the model and the guide, $\rm{KL}\!\left( p({\bf z} | {\bf x}) \lVert q_{\phi, x}({\bf z}) \right)$. This differs from variational inference, which would minimise $\rm{KL}\!\left( q_{\phi, x}({\bf z}) \lVert p({\bf z} | {\bf x}) \right)$. Using this loss encourages the approximate proposal distribution to be broader than the true posterior (mass covering), whereas variational inference typically learns a narrower approximation (mode seeking). Guides for importance sampling are usually desired to have heavier tails than the model (see this [stackexchange question](https://stats.stackexchange.com/questions/76798/in-importance-sampling-why-should-the-importance-density-have-heavier-tails)). Therefore, the inference compilation loss is usually more suited to compiling a guide for importance sampling.
#
# Another benefit of CSIS is that, unlike many types of variational inference, it has no requirement that the model is differentiable. This allows it to be used for inference on arbitrarily complex programs (e.g. a Captcha renderer [1]).
#
# This example shows CSIS being used to speed up inference on a simple problem with a known analytic solution.
# +
import torch
import torch.nn as nn
import torch.functional as F
import pyro
import pyro.distributions as dist
import pyro.infer
import pyro.optim
import os
smoke_test = ('CI' in os.environ)
n_steps = 2 if smoke_test else 2000
# -
# ## Specify the model:
#
# The model is specified in the same way as any Pyro model, except that a keyword argument, `observations`, must be used to input a dictionary with each observation as a key. Since inference compilation involves learning to perform inference for any observed values, it is not important what the values in the dictionary are. `0` is used here.
def model(prior_mean, observations={"x1": 0, "x2": 0}):
x = pyro.sample("z", dist.Normal(prior_mean, torch.tensor(5**0.5)))
y1 = pyro.sample("x1", dist.Normal(x, torch.tensor(2**0.5)), obs=observations["x1"])
y2 = pyro.sample("x2", dist.Normal(x, torch.tensor(2**0.5)), obs=observations["x2"])
return x
# ## And the guide:
#
# The guide will be trained (a.k.a. compiled) to use the observed values to make proposal distributions for each unconditioned `sample` statement. In the paper [1], a neural network architecture is automatically generated for any model. However, for the implementation in Pyro the user must specify a task-specific guide program structure. As with any Pyro guide function, this should have the same call signature as the model. It must also encounter the same unobserved `sample` statements as the model. So that the guide program can be trained to make good proposal distributions, the distributions at `sample` statements should depend on the values in `observations`. In this example, a feed-forward neural network is used to map the observations to a proposal distribution for the latent variable.
#
# `pyro.module` is called when the guide function is run so that the guide parameters can be found by the optimiser during training.
# +
class Guide(nn.Module):
def __init__(self):
super(Guide, self).__init__()
self.neural_net = nn.Sequential(
nn.Linear(2, 10),
nn.ReLU(),
nn.Linear(10, 20),
nn.ReLU(),
nn.Linear(20, 10),
nn.ReLU(),
nn.Linear(10, 5),
nn.ReLU(),
nn.Linear(5, 2))
def forward(self, prior_mean, observations={"x1": 0, "x2": 0}):
pyro.module("guide", self)
x1 = observations["x1"]
x2 = observations["x2"]
v = torch.cat((x1.view(1, 1), x2.view(1, 1)), 1)
v = self.neural_net(v)
mean = v[0, 0]
std = v[0, 1].exp()
pyro.sample("z", dist.Normal(mean, std))
guide = Guide()
# -
# ## Now create a `CSIS` instance:
# The object is initialised with the model; the guide; a PyTorch optimiser for training the guide; and the number of importance-weighted samples to draw when performing inference. The guide will be optimised for a particular value of the model/guide argument, `prior_mean`, so we use the value set here throughout training and inference.
optimiser = pyro.optim.Adam({'lr': 1e-3})
csis = pyro.infer.CSIS(model, guide, optimiser, num_inference_samples=50)
prior_mean = torch.tensor(1.)
# ## Now we 'compile' the instance to perform inference on this model:
# The arguments given to `csis.step` are passed to the model and guide when they are run to evaluate the loss.
for step in range(n_steps):
csis.step(prior_mean)
# ## And now perform inference by importance sampling:
#
# The compiled guide program should now be able to propose a distribution for `z` that approximates the posterior, $p(z | x_1, x_2)$, for any $x_1, x_2$. The same `prior_mean` is entered again, as well as the observed values inside `observations`.
posterior = csis.run(prior_mean,
observations={"x1": torch.tensor(8.),
"x2": torch.tensor(9.)})
marginal = pyro.infer.EmpiricalMarginal(posterior, "z")
# ## We now plot the results and compare with importance sampling:
#
# We observe $x_1 = 8$ and $x_2 = 9$. Inference is performed by taking 50 samples using CSIS, and 50 using importance sampling from the prior. We then plot the resulting approximations to the posterior distributions, along with the analytic posterior.
# +
import numpy as np
import scipy.stats
import matplotlib.pyplot as plt
# Draw samples from empirical marginal for plotting
csis_samples = [marginal().detach() for _ in range(1000)]
# Calculate empirical marginal with importance sampling
is_posterior = pyro.infer.Importance(model, num_samples=50).run(prior_mean,
observations={"x1": torch.tensor(8.),
"x2": torch.tensor(9.)})
is_marginal = pyro.infer.EmpiricalMarginal(is_posterior, "z")
is_samples = [is_marginal().detach() for _ in range(1000)]
# Calculate true prior and posterior over z
true_posterior_z = np.arange(-10, 10, 0.05)
true_posterior_p = np.array([np.exp(scipy.stats.norm.logpdf(p, loc=7.25, scale=(5/6)**0.5)) for p in true_posterior_z])
prior_z = true_posterior_z
prior_p = np.array([np.exp(scipy.stats.norm.logpdf(z, loc=1, scale=5**0.5)) for z in true_posterior_z])
plt.rcParams['figure.figsize'] = [30, 15]
plt.rcParams.update({'font.size': 30})
fig, ax = plt.subplots()
plt.plot(prior_z, prior_p, 'k--', label='Prior')
plt.plot(true_posterior_z, true_posterior_p, color='k', label='Analytic Posterior')
plt.hist(csis_samples, range=(-10, 10), bins=100, color='r', normed=1, label="Inference Compilation")
plt.hist(is_samples, range=(-10, 10), bins=100, color='b', normed=1, label="Importance Sampling")
plt.xlim(-8, 10)
plt.ylim(0, 5)
plt.xlabel("z")
plt.ylabel("Estimated Posterior Probability Density")
plt.legend()
plt.show()
# -
# Using $x_1 = 8$ and $x_2 = 9$ gives a posterior far from the prior, and so using the prior as a guide for importance sampling is inefficient, giving a very small effective sample size. By first learning a suitable guide function, CSIS has a proposal distribution much more closely matched to the true posterior. This allows samples to be drawn with far better coverage of the true posterior, and greater effective sample size, as shown in the graph above.
#
# For other examples of inference compilation, see [1] or <https://github.com/probprog/anglican-infcomp-examples>.
#
# ## References
#
# [1] `Inference compilation and universal probabilistic programming`,<br />
# <NAME>, <NAME>, and <NAME>
| tutorial/source/D07-csis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Network Intrusion Detection
# %matplotlib inline
# to remove
# %load_ext autoreload
# %autoreload 2
# +
import pandas as pd
import experiments
import numpy as np
from modAL.models import ActiveLearner
from modAL.uncertainty import entropy_sampling, uncertainty_sampling
import matplotlib.pyplot as plt
# -
pd.set_option('display.max_columns', 200)
# %%time
ni = experiments.NetworkIntrusionDetection()
ni.df.shape
# ## Dataset stats
# %%time
ni.report_labels()
# ## Baseline and Oracle
# %%time
df_baseline_oracle = ni.report_baseline_oracle()
df_baseline_oracle[['baseline_unsupervised_f1', 'oracle_f1']].round(2)
df_baseline_oracle.mean().round(2)
df_baseline_oracle.std().round(2)
# ## Active Learning
df_active_learning = ni.report_active_learning()
ni.report_active_learning_across_labels()
ni.report_active_learning_query_time()
ni.plot_active_learning_time_series_overlapping(
attribute='f1', label='smurf',
learner1='RandomForestClassifier', sampling1='entropy',
learner2='RandomForestClassifier', sampling2='random',
title='Random forest for smurf attack',
ylim=[0.9980, 1.0001], ylabel='F1',
legend=['Entropy sampling', 'Random sampling']
)
ni.plot_active_learning_time_series_overlapping(
attribute='f1', label='smurf',
learner1='LogisticRegression', sampling1='entropy',
learner2='LogisticRegression', sampling2='random',
title='Logistic regression for smurf attack',
ylim=[0.9980, 1.0001], ylabel='F1',
legend=['Entropy sampling', 'Random sampling']
)
ni.plot_active_learning_time_series_overlapping(
attribute='f1', label='nmap',
learner1='RandomForestClassifier', sampling1='entropy',
learner2='RandomForestClassifier', sampling2='random',
title='Random forest for nmap attack',
ylim=[0, 1], ylabel='F1',
legend=['Entropy sampling', 'Random sampling']
)
ni.plot_active_learning_time_series_overlapping(
attribute='f1', label='nmap',
learner1='LogisticRegression', sampling1='entropy',
learner2='LogisticRegression', sampling2='random',
title='Logistic regression for nmap attack',
ylim=[0, 1], ylabel='F1',
legend=['Entropy sampling', 'Random sampling']
)
# ## Ensemble
for label in ni.features:
try:
ni.plot_active_learning_time_series_overlapping(
attribute='f1', label=label.replace('.', ''),
learner1='RandomForestClassifier', sampling1='entropy',
learner2='VotingClassifier', sampling2='entropy',
title=f'RF vs. ensemble for {label} attack',
ylim=[0, 1], ylabel='F1',
legend=['RF', 'Ensemble of RF, LR, and IF']
)
plt.show()
except:
pass
# ## Error analysis
# ### Uncertainty sampling drawbacks
label = 'nmap.'
# +
def get_predictions(estimator):
predictions = estimator.predict(active_learning_data.x_dev)
print(ni._get_metrics(active_learning_data.y_dev,
predictions, None))
return predictions
def get_feature_importance(estimator, n=10):
df = pd.DataFrame({'feature': ni.splits[label]['x_train'].columns,
'importance': estimator.feature_importances_}).sort_values(by='importance', ascending=False)[:n][::-1]
df.set_index('feature').plot(kind='barh', legend=False)
plt.title(f'Top {n} features by importance')
plt.xlabel('Gini importance')
def get_gets_wrong(predictions):
return pd.DataFrame(active_learning_data.x_dev[predictions != active_learning_data.y_dev],
columns=ni.splits['nmap.']['x_train'].columns)
def get_right_wrong_comp(predictions, n=10):
# doesn't get right
gets_wrong = get_gets_wrong(predictions)
gets_right = pd.DataFrame(active_learning_data.x_dev[(predictions == active_learning_data.y_dev) & (active_learning_data.y_dev)],
columns=ni.splits['nmap.']['x_train'].columns)
z = (gets_wrong.mean() - gets_right.mean()) / gets_right.std()
z = z.abs()
return z[pd.notnull(z) & np.isfinite(z)].sort_values(ascending=False)[:n]
# -
active_learning_data = ni._active_learning_data_split(label)
active_learner = ActiveLearner(ni.active_learning_rf, uncertainty_sampling,
active_learning_data.x_train_start.values,
active_learning_data.y_train_start.values)
predictions = get_predictions(active_learner)
indices = np.array(range(len(active_learning_data.y_dev)))[predictions != active_learning_data.y_dev]
active_learner.predict_proba(active_learning_data.x_dev[indices])
get_feature_importance(active_learner.estimator, n=10)
get_right_wrong_comp(predictions, n=10)
index = active_learner.query(active_learning_data.x_train_pool.values)[0][0]
index
pd.DataFrame(
active_learning_data.x_train_pool.values[index].reshape(1, -1),
columns=ni.splits['nmap.']['x_dev'].columns
)
active_learning_data.y_train_pool[index]
x_instance, y_instance = ni._get_active_learning_instance(active_learning_data.x_train_pool,
active_learning_data.y_train_pool,
[index])
active_learner.teach(x_instance, y_instance)
predictions = get_predictions(active_learner)
get_feature_importance(active_learner.estimator, n=10)
index = active_learner.query(active_learning_data.x_train_pool.values)[0][0]
index
active_learning_data.y_train_pool[index]
x_instance, y_instance = ni._get_active_learning_instance(active_learning_data.x_train_pool,
active_learning_data.y_train_pool,
[index])
active_learner.teach(x_instance, y_instance)
predictions = get_predictions(active_learner)
pd.DataFrame(
active_learning_data.x_train_pool.values[index].reshape(1, -1),
columns=ni.splits['nmap.']['x_dev'].columns
).src_bytes
get_feature_importance(active_learner.estimator)
get_right_wrong_comp(predictions, n=10)
clf = ni.active_learning_rf
clf.fit(ni.splits[label]['x_train'], ni.splits[label]['y_train'])
predictions = get_predictions(clf)
get_feature_importance(clf, n=10)
get_right_wrong_comp(predictions)
# ### Unsupervised sampling strategy
df_learn_anomalies = ni.report_learn_anomalies()
cols = ['label', 'initial_f1', 'sample_10_f1', 'sample_50_f1', 'sample_100_f1']
q = df_learn_anomalies[cols]
q.mean().round(2)
q.std().round(2)
qq = df_active_learning[(df_active_learning.learner == 'RandomForestClassifier')
& (df_active_learning['sampling strategy'] == 'entropy_sampling')][cols]
qq.mean().round(2)
qq.std().round(2)
for label in ni.features:
ni.plot_active_learning_time_series_overlapping(
attribute='f1', label=label.replace('.', ''),
learner1='RandomForestClassifier', sampling1='entropy',
learner2='RandomForestClassifier', sampling2='iforest',
title=f'Entropy vs. unsupervised sampling for {label} attack',
ylim=[0, 1], ylabel='F1',
legend=['RF', 'IF']
)
plt.show()
| final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "4a463c67-7543-42d2-a116-e70e8451b09b", "showTitle": false, "title": ""}
# ## Interpretability - Tabular SHAP explainer
#
# In this example, we use Kernel SHAP to explain a tabular classification model built from the Adults Census dataset.
#
# First we import the packages and define some UDFs we will need later.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "bf0fdfc2-97b2-48e4-b3d9-794b0cb3da67", "showTitle": false, "title": ""}
import pyspark
from synapse.ml.explainers import *
from pyspark.ml import Pipeline
from pyspark.ml.classification import LogisticRegression
from pyspark.ml.feature import StringIndexer, OneHotEncoder, VectorAssembler
from pyspark.sql.types import *
from pyspark.sql.functions import *
import pandas as pd
vec_access = udf(lambda v, i: float(v[i]), FloatType())
vec2array = udf(lambda vec: vec.toArray().tolist(), ArrayType(FloatType()))
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "ae47e1f9-0672-47ed-94de-10970e1b14b5", "showTitle": false, "title": ""}
# Now let's read the data and train a simple binary classification model.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "58807448-d8e0-4818-adc8-27536d561fb3", "showTitle": false, "title": ""}
df = spark.read.parquet("wasbs://publicwasb@mmlspark.blob.core.windows.net/AdultCensusIncome.parquet")
labelIndexer = StringIndexer(inputCol="income", outputCol="label", stringOrderType="alphabetAsc").fit(df)
print("Label index assigment: " + str(set(zip(labelIndexer.labels, [0, 1]))))
training = labelIndexer.transform(df)
display(training)
categorical_features = [
"workclass",
"education",
"marital-status",
"occupation",
"relationship",
"race",
"sex",
"native-country",
]
categorical_features_idx = [col + "_idx" for col in categorical_features]
categorical_features_enc = [col + "_enc" for col in categorical_features]
numeric_features = ["age", "education-num", "capital-gain", "capital-loss", "hours-per-week"]
strIndexer = StringIndexer(inputCols=categorical_features, outputCols=categorical_features_idx)
onehotEnc = OneHotEncoder(inputCols=categorical_features_idx, outputCols=categorical_features_enc)
vectAssem = VectorAssembler(inputCols=categorical_features_enc + numeric_features, outputCol="features")
lr = LogisticRegression(featuresCol="features", labelCol="label", weightCol="fnlwgt")
pipeline = Pipeline(stages=[strIndexer, onehotEnc, vectAssem, lr])
model = pipeline.fit(training)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f617f9a4-7e67-43f8-8fa9-92680b635b3d", "showTitle": false, "title": ""}
# After the model is trained, we randomly select some observations to be explained.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f55757a6-6204-4f64-a91e-65bfbacf62bc", "showTitle": false, "title": ""}
explain_instances = model.transform(training).orderBy(rand()).limit(5).repartition(200).cache()
display(explain_instances)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "48a0c8ee-8e36-4bd3-9a04-eded6d2c8894", "showTitle": false, "title": ""}
# We create a TabularSHAP explainer, set the input columns to all the features the model takes, specify the model and the target output column we are trying to explain. In this case, we are trying to explain the "probability" output which is a vector of length 2, and we are only looking at class 1 probability. Specify targetClasses to `[0, 1]` if you want to explain class 0 and 1 probability at the same time. Finally we sample 100 rows from the training data for background data, which is used for integrating out features in Kernel SHAP.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "7e097552-e617-4e1c-a085-b66eca5bcb69", "showTitle": false, "title": ""}
shap = TabularSHAP(
inputCols=categorical_features + numeric_features,
outputCol="shapValues",
numSamples=5000,
model=model,
targetCol="probability",
targetClasses=[1],
backgroundData=broadcast(training.orderBy(rand()).limit(100).cache()),
)
shap_df = shap.transform(explain_instances)
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "6933b52b-7d46-4210-810a-f984b76dd4a2", "showTitle": false, "title": ""}
# Once we have the resulting dataframe, we extract the class 1 probability of the model output, the SHAP values for the target class, the original features and the true label. Then we convert it to a pandas dataframe for visisualization.
# For each observation, the first element in the SHAP values vector is the base value (the mean output of the background dataset), and each of the following element is the SHAP values for each feature.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "05e01f98-e44c-46c9-a8ae-26ba892f85b3", "showTitle": false, "title": ""}
shaps = (
shap_df.withColumn("probability", vec_access(col("probability"), lit(1)))
.withColumn("shapValues", vec2array(col("shapValues").getItem(0)))
.select(["shapValues", "probability", "label"] + categorical_features + numeric_features)
)
shaps_local = shaps.toPandas()
shaps_local.sort_values("probability", ascending=False, inplace=True, ignore_index=True)
pd.set_option("display.max_colwidth", None)
shaps_local
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "f9317a27-900a-4d1d-9e9f-9fe906eae75c", "showTitle": false, "title": ""}
# We use plotly subplot to visualize the SHAP values.
# + application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "c9b4c03e-eac8-4314-a6c2-0a451525e6a4", "showTitle": false, "title": ""}
from plotly.subplots import make_subplots
import plotly.graph_objects as go
import pandas as pd
features = categorical_features + numeric_features
features_with_base = ["Base"] + features
rows = shaps_local.shape[0]
fig = make_subplots(
rows=rows,
cols=1,
subplot_titles="Probability: " + shaps_local["probability"].apply("{:.2%}".format) + "; Label: " + shaps_local["label"].astype(str),
)
for index, row in shaps_local.iterrows():
feature_values = [0] + [row[feature] for feature in features]
shap_values = row["shapValues"]
list_of_tuples = list(zip(features_with_base, feature_values, shap_values))
shap_pdf = pd.DataFrame(list_of_tuples, columns=["name", "value", "shap"])
fig.add_trace(
go.Bar(x=shap_pdf["name"], y=shap_pdf["shap"], hovertext="value: " + shap_pdf["value"].astype(str)),
row=index + 1,
col=1,
)
fig.update_yaxes(range=[-1, 1], fixedrange=True, zerolinecolor="black")
fig.update_xaxes(type="category", tickangle=45, fixedrange=True)
fig.update_layout(height=400 * rows, title_text="SHAP explanations")
fig.show()
# + [markdown] application/vnd.databricks.v1+cell={"inputWidgets": {}, "nuid": "8f22fceb-0fc0-4a86-a0ca-2a7b47b4795a", "showTitle": false, "title": ""}
# Your results will look like:
#
# <img src="https://mmlspark.blob.core.windows.net/graphics/explainers/tabular-shap.png" style="float: right;"/>
| notebooks/features/responsible_ai/Interpretability - Tabular SHAP explainer.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.2.0
# language: julia
# name: julia-1.2
# ---
# In this notebook, we compute invariant sets for the planar system with univariate input of Example ?.? of [???].
# We need to select an semidefinite programming solver to find the invariant set. A complete list of available solvers is [available here](https://www.juliaopt.org/JuMP.jl/stable/installation/#Getting-Solvers-1).
using MosekTools
using JuMP
solver = with_optimizer(Mosek.Optimizer, QUIET=true)
# We first define the safe sets and input sets for our system using [Polyhedra](https://github.com/JuliaPolyhedra/Polyhedra.jl).
using Polyhedra
G = [-0.7238 -0.5117
0.2558 -0.8091
-0.0528 0.7725
-0.1060 -0.7190
-0.1252 0.1868
0.7232 -0.9371
0.4235 0.6708]
F = [0.2990;
0.0983;
0.0276;
0.1202;
0.0348;
0.0921;
0.0240]
safe_set = polyhedron(hrep(G, F), DefaultLibrary{Float64}(solver))
# The difference between example `Planar_System_bounded_univariate_control.ipynb` is that here
# the `safe_set` have the `chebyshevcenter` at the origin and the system is linear.
# For the other example, the system becomes affine if we recenter it.
cheby_center, cheby_radius = chebyshevcenter(safe_set, solver)
safe_set = Polyhedra.translate(safe_set, -cheby_center)
input_set = polyhedron(convexhull([-2], [2]))
# We new define the dynamic of our system and create it using [MathematicalSystems](https://github.com/JuliaReach/MathematicalSystems.jl/).
using MathematicalSystems
A = [1 0.1
0 1]
B = reshape([0, 0.05], 2, 1)
system = ConstrainedLinearControlDiscreteSystem(A, B, safe_set, input_set)
# We now use SwitchOnSafety to compute the invariant set. We start with symmetric ellipsoid centered at the origin.
using SwitchOnSafety
sym_variable = Ellipsoid(symmetric = true)
max_vol_sym_ell = invariant_set(system, solver, sym_variable)
# This computes the maximal *volume* ellipsoid.
# We can instead maximize the integral of the quadratic form defining the ellipsoid over the hypercube.
# This corresponds to maximizing the sum of squares of its semi-axes instead or equivalently the trace of the positive definite matrix defining the quadratic form.
max_tr_sym_ell = invariant_set(system, solver, sym_variable, volume_heuristic = ell -> L1_heuristic(ell, ones(3)))
# We can see the result below.
using Plots
plot(safe_set)
plot!(max_vol_sym_ell, color=:orange)
plot!(max_tr_sym_ell, color=:green)
# We can see that forcing the center of the ellipsoid to coincide with the Chebyshev center of the safe set is quite conservative.
# We can ask instead to search for any ellipsoid with the origin in its interior.
# As the system is reformulated into an algebraic system with safe set `safe_set * input_set`, the Chebyshev center is `(0, 0, 0)` (not that `Polyhedra.chebyshevcenter(safe * input)` is currently incorrect because of https://github.com/JuliaPolyhedra/Polyhedra.jl/issues/125).
# To avoid having to solve Bilinear Matrix Inequalities, we set the S-procedure scaling to `1.0`.
using SwitchOnSafety
variable = Ellipsoid(point = SetProg.InteriorPoint(zeros(3)))
max_vol_ell = invariant_set(system, solver, variable, λ = 1.0)
# Instead of maximizing the volume, we can maximize the L1 norm as above.
using SwitchOnSafety
max_tr_ell = invariant_set(system, solver, variable, λ = 1.0, volume_heuristic = ell -> L1_heuristic(ell, ones(3)))
# We can see that we get a larger ellipsoids for the volume maximization but not for the L1 norm as maximizing the L1 integral over the hypercube centered at the origin is not a very good heuristic. We should instaed maximize the L1 integral over the safe set but this is not implemented yet in [SetProg](https://github.com/blegat/SetProg.jl).
using Plots
plot(safe_set)
plot!(max_vol_ell, color=:orange)
plot!(max_tr_ell, color=:green)
| examples/Controlled_Invariant_Sets/Planar_System_bounded_univariate_control_centered.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''fhl_py38'': conda)'
# name: python3810jvsc74a57bd0eadf4def0511d16cff3b403ebad030b79626359e624f736446cdd8402c7991c1
# ---
# # Create BoundingBox Labeled Datasets from CSV ground truth
#
#
# This script will create two labeled datasets: <dataset_name>_training and <dataset_name>_test
#
# 1. split training and test labeled datapoints
# 2. create jason-line files for training and test
# 3. upload jason-line files to default workspace blobstore, under fhl/datasets/<dataset_name>/label folder
# 4. register labeled datasets
import json
import pandas as pd
import sklearn
from sklearn.model_selection import train_test_split
datastore_name = 'workspaceblobstore'
dataset_name = 'NoWindowOD'
# +
# read ground truth from csv file
raw_df = pd.read_csv('./labels/stage_2_train_labels.csv')
# adding label
def get_label(row):
if row['Target'] == 0:
val = 'Not Lung Opacity'
elif row['Target'] == 1:
val = 'Lung Opacity'
return val
raw_df['Label'] = raw_df.apply(get_label, axis=1)
# Filling NaN with average values
raw_df['x'].fillna(value=raw_df['x'].mean(), inplace=True)
raw_df['y'].fillna(value=raw_df['y'].mean(), inplace=True)
raw_df['width'].fillna(value=raw_df['width'].mean(), inplace=True)
raw_df['height'].fillna(value=raw_df['height'].mean(), inplace=True)
# -
# merge by
IMG_WIDTH = 1024
IMG_HEIGHT = 1024
jsonline_obj = []
tags = []
target_count = 0
for patient_id in raw_df['patientId'].unique():
obj = {}
target = False
obj['image_url'] = 'AmlDatastore://' + datastore_name + '/fhl/datasets/' + dataset_name + '/image/' + patient_id + '.png'
obj['label'] = []
obj['label_confidence'] = []
sub_df = raw_df[raw_df['patientId'] == patient_id]
for _, row in sub_df.iterrows():
target = row['Label'] == 'Lung Opacity'
label = {'label': row['Label'], 'topX': row['x']/IMG_WIDTH, 'topY': row['y']/IMG_HEIGHT, 'bottomX': (row['x'] + row['width'])/IMG_WIDTH, 'bottomY': (row['y'] + row['height'])/IMG_HEIGHT}
obj['label'].append(label)
obj['label_confidence'].append(1.0)
jsonline_obj.append(obj)
if target:
tags.append('Lung Opacity')
target_count += 1
else:
tags.append('Not Lung Opacity')
image_df = pd.DataFrame({'obj':jsonline_obj,'tag':tags})
image_df['tag'].value_counts()
# Down-sample Not Lung Opacit
from sklearn.utils import resample
major_df = image_df[image_df['tag'] == 'Not Lung Opacity']
minor_df = image_df[image_df['tag'] == 'Lung Opacity']
down_sample_df = resample(major_df, replace=False, n_samples=minor_df.shape[0])
source_df = pd.concat([down_sample_df, minor_df])
train, test = train_test_split(source_df, test_size = 0.2, random_state = 2021)
train['tag'].value_counts()
train_obj = train['obj']
test_obj = test['obj']
import os
import json
def save_jasonline_file(objects: list, file_name: str):
base_path = os.path.dirname(file_name)
if not os.path.exists(base_path):
os.makedirs(base_path)
with open(file_name, 'w') as jf:
for obj in objects:
jf.write('{}\n'.format(json.dumps(obj)))
# Generate jsonline files, which can be registered as labeled dataset
label_folder = os.path.abspath(os.path.join(os.path.curdir, 'label'))
save_jasonline_file(train_obj, os.path.join(label_folder, 'labeleddatapoints_training.jsonl'))
save_jasonline_file(test_obj, os.path.join(label_folder, 'labeleddatapoints_test.jsonl'))
import azureml.core
from azureml.core import Workspace
# Load workspace
ws = Workspace.from_config()
ds = ws.datastores['workspaceblobstore']
ds.upload(src_dir = label_folder, target_path= '/fhl/datasets/' + dataset_name + '/label/', overwrite= True)
# +
# Create/register labeled dataset for training and test(inference)
from azureml.contrib.dataset.labeled_dataset import _LabeledDatasetFactory, LabeledDatasetTask
tags = {}
tags['labelingCreatedBy'] = "FHL Notebook"
tags['labelingProjectType'] = 'Object Identification (Bounding Box)'
tags['SourceDatastoreName'] = 'workspaceblobstore'
tags['SourceRelativePath'] = 'fhl/datasets/' + dataset_name + '/image/'
tags['labelingLabelName'] = '["Lung Opacity","No Lung Opacity"]'
training_dataset = _LabeledDatasetFactory.from_json_lines(task=LabeledDatasetTask.IMAGE_CLASSIFICATION, path=ds.path('fhl/datasets/' + dataset_name + '/label/labeleddatapoints_training.jsonl'))
training_dataset.register(ws, dataset_name + '_training', tags= tags)
test_dataset = _LabeledDatasetFactory.from_json_lines(task=LabeledDatasetTask.IMAGE_CLASSIFICATION, path=ds.path('fhl/datasets/' + dataset_name + '/label/labeleddatapoints_test.jsonl'))
test_dataset.register(ws, dataset_name + '_test', tags= tags)
# -
| notebooks/fhl_dicom_od_create_labeled_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (oat-use)
# language: python
# name: oat-use
# ---
# %run "Common setup.ipynb"
from SALib.analyze.rbd_fast import analyze as rbd_analyze
# +
### Analyze extreme results
extreme_numeric_samples = pd.read_csv(f'{DATA_DIR}extreme_numeric_samples.csv', index_col=0)
extreme_numeric_vals = extreme_numeric_samples[perturbed_cols].values
# Results from extremity testing with coupling disabled
extreme_results = pd.read_csv(f'{DATA_DIR}no_irrigation_extreme_results.csv', index_col=0)
extreme_results['Avg. $/ML'].fillna(extreme_results["Avg. Annual Profit ($M)"], inplace=True)
extreme_disabled_res = extreme_results.values
target_result_idx = extreme_results.columns.tolist().index(tgt_metric)
extremity_disabled_results = rbd_analyze(CIM_SPEC, extreme_numeric_vals,
extreme_disabled_res[:, target_result_idx],
M=4, seed=101)
# +
# Extreme values with interactions
extreme_results = pd.read_csv(f'{DATA_DIR}with_irrigation_extreme_results.csv', index_col=0)
extreme_results['Avg. $/ML'].fillna(extreme_results["Avg. Annual Profit ($M)"], inplace=True)
extreme_enabled_res = extreme_results.values
extremity_enabled_results = rbd_analyze(CIM_SPEC, extreme_numeric_vals,
extreme_enabled_res[:, target_result_idx],
M=4, seed=101)
# +
fig, axes = plt.subplots(1,2, figsize=(12,4), sharey=True, sharex=True)
disabled = extremity_disabled_results.to_df().loc[tgt_param, ['S1']]
disabled.name = 'Irrigation Efficiency'
enabled = extremity_enabled_results.to_df().loc[tgt_param, ['S1']]
enabled.name = 'Irrigation Efficiency'
disabled.plot(kind='bar',
legend=None,
title='Disabled interactions',
ax=axes[0],
rot=0)
enabled.plot(kind='bar',
legend=None,
title='Enabled interactions',
ax=axes[1],
rot=0).legend(
bbox_to_anchor=(1.05, 0.65)
)
fig.suptitle("EASI Extremity Testing", x=0.51, y=1.05, fontsize=20);
# -
fig.savefig(f'{FIG_DIR}single_param_ext_EASI_results.png', dpi=300, bbox_inches='tight')
# ---
def incremental_easi_analysis(lower, upper, step=1):
res, idx = [], []
for reps in range(lower, upper, step):
try:
results = rbd_analyze(CIM_SPEC, numeric_vals[:reps, :],
np_res[:reps],
M=4, seed=101)
except ZeroDivisionError:
res.append(np.nan)
idx.append(reps)
continue
# End try
enabled = results.to_df()
tmp = enabled.loc[tgt_param, 'S1']
res.append(tmp)
idx.append(reps)
# End for
return res, idx
numeric_samples = pd.read_csv(f'{DATA_DIR}oat_mc_10_numeric_samples.csv', index_col=0)
numeric_samples = numeric_samples[perturbed_cols]
numeric_vals = numeric_samples.values
# +
# Coupling disabled
oat_10_no_irrigation_results = pd.read_csv(f'{DATA_DIR}oat_no_irrigation_10_results.csv', index_col=0)
oat_10_no_irrigation_results['Avg. $/ML'].fillna(oat_10_no_irrigation_results["Avg. Annual Profit ($M)"], inplace=True)
np_res = oat_10_no_irrigation_results.loc[:, tgt_metric].values
res, idx = incremental_easi_analysis(2, 541)
disabled = pd.DataFrame({'S1': res}, index=idx)
# +
# Coupling enabled
oat_10_with_irrigation_results = pd.read_csv(f'{DATA_DIR}oat_with_irrigation_10_results.csv', index_col=0)
oat_10_with_irrigation_results['Avg. $/ML'].fillna(oat_10_with_irrigation_results["Avg. Annual Profit ($M)"], inplace=True)
np_res = oat_10_with_irrigation_results.loc[:, tgt_metric].values
# np_res = np_res.astype('float64')
res, idx = incremental_easi_analysis(2, 541)
enabled = pd.DataFrame({'S1': res}, index=idx)
# +
fig, axes = plt.subplots(1,2, figsize=(12,4), sharey=True, sharex=True)
labels = [str(i) if i % 54 == 0 else '' for i in idx]
disabled.loc[:, 'S1'].plot(kind='bar',
legend=None,
title='Disabled',
ax=axes[0],
use_index=False,
rot=45,
width=1.1,
edgecolor='C0')
enabled.loc[:, 'S1'].plot(kind='bar',
legend=None,
title='Enabled',
ax=axes[1],
use_index=False,
rot=45,
width=1.1,
edgecolor='C0').legend(
bbox_to_anchor=(1.35, 0.65)
)
fig.suptitle("EASI Analysis", x=0.5, y=1.05, fontsize=22)
# fig.tight_layout()
plt.xlabel("$N$", x=-0.05, labelpad=15);
axes[0].set_xticklabels(labels)
axes[1].set_xticklabels(labels);
# -
fig.savefig(FIG_DIR+'EASI_larger_sample.png', dpi=300, bbox_inches='tight')
# ---
#
# Using full inputs/outputs
# +
target_result_idx = all_outputs.columns.tolist().index("SW Allocation Index")
numeric_samples = to_numeric_samples(all_inputs)
numeric_vals = numeric_samples.values
np_res = all_outputs.values[:, target_result_idx]
np_res = np_res.astype('float64')
# -
res, idx = incremental_easi_analysis(2, 5625)
# +
full_run_plot = pd.DataFrame({'S1': res}, index=idx).plot(
kind='bar',
# legend=None,
title='Full model runs',
use_index=False,
rot=45,
width=1.1,
edgecolor='C0')
full_run_plot.set_xlabel("$N$")
labels = [str(i) if i % 500 == 0 else '' for i in idx]
full_run_plot.set_xticklabels(labels)
full_run_plot.set_xticklabels(labels);
# -
| notebooks/5a EASI Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.2 64-bit
# language: python
# name: python3
# ---
planeta = 'Marte '
gravedad = 0.00143
nombre = 'Ganímedes'
title = f'datos de gravedad sobre {nombre}'
hechos = f"""{'-'*80}
Nombre del planeta: {planeta}
Gravedad en {nombre}: {gravedad * 1000} m/s2
"""
template = f"""{title.title()}
{hechos}
"""
print(hechos)
planeta = 'Marte '
gravedad = 0.00143
nombre = 'Ganímedes'
print(hechos)
new_template = """
Datos de Gravedad sobre: {nombre}
-------------------------------------------------------------------------------
Nombre del planeta: {planeta}
Gravedad en {nombre}: {gravedad} m/s2
"""
print(new_template.format(nombre=nombre, planeta=planeta, gravedad=gravedad))
print(new_template.format(nombre=nombre, planeta=planeta, gravedad=gravedad*1000))
| Kata_4_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import glob
from stempy import io, image
import matplotlib.pyplot as plt
files = glob.glob('/data/4dstem/smallScanningDiffraction/data*.dat')
inner_radii = [0, 40]
outer_radii = [288, 288]
reader = io.reader(files)
imgs = image.create_stem_images(reader, inner_radii, outer_radii, scan_dimensions=(160, 160))
for img in imgs:
fig,ax=plt.subplots(figsize=(6,6))
ax.matshow(img, cmap=plt.cm.gray)
plt.show()
| examples/stem_image.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: RAPIDS Stable
# language: python
# name: rapids-stable
# ---
# # SQL Reference
# ## Functions
# ### Aggregators
# Aggregators perform calculations of a set of elements. They can be used with and without a GROUP BY clause
# #### COUNT
# Counts non null-elements
from blazingsql import BlazingContext
bc = BlazingContext()
bc.create_table('nation', '../../../../data/nation.parquet')
query = '''
SELECT
COUNT(n_nationkey)
FROM
nation
'''
bc.sql(query)
query = '''
SELECT
n_regionkey, COUNT(n_nationkey)
FROM
nation
GROUP BY n_regionkey
'''
bc.sql(query)
# #### SUM
# Sums non null-elements
from blazingsql import BlazingContext
bc = BlazingContext()
bc.create_table('nation', '../../../../data/nation.parquet')
query = '''
SELECT
SUM(n_nationkey)
FROM
nation
'''
bc.sql(query)
query = '''
SELECT
n_regionkey, SUM(n_nationkey)
FROM
nation
GROUP BY n_regionkey
'''
bc.sql(query)
# #### MIN and MAX
# Takes the minimum and maximum of non-null elements respectively
from blazingsql import BlazingContext
bc = BlazingContext()
bc.create_table('nation', '../../../../data/nation.parquet')
query = '''
SELECT
MIN(n_nationkey), MAX(n_nationkey)
FROM
nation
'''
bc.sql(query)
query = '''
SELECT
n_regionkey, MIN(n_nationkey), MAX(n_nationkey)
FROM
nation
GROUP BY n_regionkey
'''
bc.sql(query)
# #### AVG
# Takes the average of non-null elements
from blazingsql import BlazingContext
bc = BlazingContext()
bc.create_table('nation', '../../../../data/nation.parquet')
query = '''
SELECT
AVG(n_nationkey)
FROM
nation
'''
bc.sql(query)
query = '''
SELECT
n_regionkey, AVG(n_nationkey)
FROM
nation
GROUP BY n_regionkey
'''
bc.sql(query)
# #### STDDEV, STDDEV_SAMP, STDDEV_POP
# STDDEV_SAMP takes the sample standard deviation of the elements.
#
# STDDEV_POP takes the population standard deviation of the elements.
#
# STDDEV is an alias of STDDEV_SAMP.
#
# NOTE: The difference between STDDEV_SAMP and STDDEV_POP is that STDDEV_SAMP is scaled by 1/(N-1) while STDDEV_POP is scaled by 1/N.
from blazingsql import BlazingContext
bc = BlazingContext()
bc.create_table('nation', '../../../../data/nation.parquet')
query = '''
SELECT
STDDEV(n_nationkey) AS std_dev,
STDDEV_SAMP(n_nationkey) AS std_dev_samp,
STDDEV_POP(n_nationkey) as std_dev_pop
FROM
nation
'''
bc.sql(query)
query = '''
SELECT
n_regionkey,
STDDEV(n_nationkey) AS std_dev,
STDDEV_SAMP(n_nationkey) AS std_dev_samp,
STDDEV_POP(n_nationkey) as std_dev_pop
FROM
nation
GROUP BY n_regionkey
'''
bc.sql(query)
# #### VARIANCE, VAR_SAMP, VAR_POP
# VAR_SAMP takes the sample variance of the elements.
#
# VAR_POP takes the population variance of the elements.
#
# VARIANCE is an alias of VAR_SAMP.
#
# NOTE: The difference between VAR_SAMP and VAR_POP is that VAR_SAMP is scaled by 1/(N*(N-1)) while VAR_POP is scaled by 1/(N^2).
from blazingsql import BlazingContext
bc = BlazingContext()
bc.create_table('nation', '../../../../data/nation.parquet')
query = '''
SELECT
VARIANCE(n_nationkey) AS var,
VAR_SAMP(n_nationkey) AS varsamp,
VAR_POP(n_nationkey) as varpop
FROM
nation
'''
bc.sql(query)
query = '''
SELECT
n_regionkey,
VARIANCE(n_nationkey) AS var,
VAR_SAMP(n_nationkey) AS varsamp,
VAR_POP(n_nationkey) as varpop
FROM
nation
GROUP BY n_regionkey
'''
bc.sql(query)
| docs/blazingsql/sql_reference/functions/aggregators.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jaynarayan94/All-ML-Code-And-Mini-Projects/blob/master/Multi_Class_Classification_using_Brainwave_Dataset.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="9_7V2ieBsk3t" colab_type="text"
# ### From the Blog : [Multi-Class classification with Sci-kit learn & XGBoost: A case study using Brainwave data](https://medium.com/free-code-camp/multi-class-classification-with-sci-kit-learn-xgboost-a-case-study-using-brainwave-data-363d7fca5f69)
# + id="sllA6qhMpzBZ" colab_type="code" colab={}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from collections import Counter
import warnings
warnings.filterwarnings('ignore')
np.random.seed(1234)
# + id="Ay2rkIf7soin" colab_type="code" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": "OK"}}, "base_uri": "https://localhost:8080/", "height": 91} outputId="430ed528-b781-490c-a529-86a71ebaa59d"
from google.colab import files
uploaded = files.upload()
# + id="ICVTIBrEt2oD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="71088183-cfdf-4691-8e47-227dccd0f711"
for fn in uploaded.keys():
print('User uploaded file "{name}" with length {length} bytes'.format(
name=fn, length=len(uploaded[fn])))
# + id="6hgjFq20s8gn" colab_type="code" colab={}
import io
brainwave_df = pd.read_csv(io.StringIO(uploaded['emotions.csv'].decode('utf-8')))
# + id="DFw2rmKR2Yvx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 261} outputId="e569beba-8889-406d-cdcf-bd6de3f6842e"
brainwave_df.head()
# + id="zTjoVkHR2bII" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="617fa63d-78df-448b-eea7-fc0ffd3c0d88"
brainwave_df.shape
# + id="TF2rfzih2fjc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 101} outputId="aeacd6ac-5de0-45bc-be6d-e942c0390622"
brainwave_df.info()
# + id="5jnrX4_R2gx3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d3e0a6e4-ef15-4dd6-b98c-0879f0083aee"
Counter(brainwave_df['label'])
# + [markdown] id="TZZYlVGS2oF-" colab_type="text"
# There are 2549 columns in the dataset and ‘label’ is the target column for our classification problem. All other columns like ‘mean_d_1_a’, ‘mean_d2_a’ etc are describing features of brainwave signal readings. Columns starting with the ‘fft’ prefix are most probably ‘Fast Fourier transforms’ of original signals. Our target column ‘label’ describes the degree of emotional sentiment.
#
# As per Kaggle, here is the challenge: “Can we predict emotional sentiment from brainwave readings?”
#
# Let’s first understand class distributions from column ‘label’:
# + id="KvAlrX3G2i8E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 316} outputId="ab79ad3f-1d3c-45d3-ea5b-9b3a3ed8a0dd"
sns.countplot(brainwave_df['label']);
plt.xticks(rotation='45')
plt.xlabel('EEG Brainwave Label')
plt.show()
# + [markdown] id="_AydT3JE3ev9" colab_type="text"
# So, there are three classes, ‘POSITIVE’, ‘NEGATIVE’ & ‘NEUTRAL’, for emotional sentiment. From the bar chart, it is clear that class distribution is not skewed and it is a ‘multi-class classification’ problem with target variable ‘label’. We will try with different classifiers and see the accuracy levels.
# + id="c__wNMVS2pIj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 261} outputId="5bc78054-289c-4710-c7ee-dfed2a82b938"
# let's split depedent & Indepent Variables
x = brainwave_df.drop(['label'],axis=1)
y = brainwave_df['label']
x.head()
# + id="Ft-0ZIns3fXQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 113} outputId="c611e1e6-2036-421b-f5e1-5ab81e9d9408"
#Check for missing values.
x[x.isnull().any(axis=1)]
# + [markdown] id="GL3ZCBqK3n6H" colab_type="text"
# As it is a ‘classification’ problem, we will follow the below conventions for each ‘classifier’ to be tried:
# * We will use a ‘cross validation’ (in our case will use 10 fold cross validation) approach over the dataset and take average accuracy. This will give us a holistic view of the classifier’s accuracy.
# * We will use a ‘Pipeline’ based approach to combine all pre-processing and main classifier computation. A ML ‘Pipeline’ wraps all processing stages in a single unit and act as a ‘classifier’ itself. By this, all stages become re-usable and can be put in forming other ‘pipelines’ also.
# * We will track total time in building & testing for each approach. We will call this ‘time taken’.
#
# ## RandomForest Classifier
# ‘RandomForest’ is a tree & bagging approach-based ensemble classifier. It will automatically reduce the number of features by its probabilistic entropy calculation approach. Let’s see that:
# + id="uVlLCxIp3joj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="5265eba6-2837-4196-bdc6-1e402b9afefc"
# %%time
from sklearn.pipeline import Pipeline
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score,train_test_split
pl_random_forest = Pipeline(steps=[('random_forest',RandomForestClassifier())])
scores = cross_val_score(pl_random_forest,x,y,cv=10,scoring='accuracy')
print('Accuracy for Random forest Classifier : ',scores.mean())
# + [markdown] id="thSmRDQj3wIj" colab_type="text"
# Accuracy is very good at 97.09% and ‘total time taken’ is quite short (5.13 s only).
#
# For this classifier, no pre-processing stages like scaling or noise removal are required, as it is completely probability-based and not at all affected by scale factors.
#
# ## Logistic Regression Classifier
# ‘Logistic Regression’ is a linear classifier and works in same way as linear regression.
# + id="2UB996bR3oel" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="b51a065f-49e8-4cb1-e458-29ccabe4574a"
# %%time
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
pl_log_reg = Pipeline(steps=[('scalre',StandardScaler()),\
('log_reg',LogisticRegression(multi_class='multinomial', solver='sag', max_iter=200))])
scores = cross_val_score(pl_log_reg,x,y,cv=10,scoring='accuracy')
print('Accuracy for Logisitic Regression : ',scores.mean())
# + [markdown] id="lfZ8jalW34HE" colab_type="text"
# We can see accuracy (94.51%) is lower than ‘RandomForest’ and ‘time taken’ is higher (3min 49s).
#
# ‘Logistic Regression’ is heavily affected by different value ranges across dependent variables, thus forces ‘feature scaling’. That’s why ‘StandardScaler’ from scikit-learn has been added as a preprocessing stage. It automatically scales features according to a Gaussian Distribution with zero mean & unit variance, and thus values for all variables range from -1 to +1.
#
# The reason for high time taken is high-dimensionality and scaling time required. There are 2549 variables in the dataset and the coefficient of each one should be optimised as per the Logistic Regression process. Also, there is a question of multi-co-linearity. This means linearly co-related variables should be grouped together instead of considering them separately.
#
# The presence of multi-col-linearity affects accuracy. So now the question becomes, “Can we reduce the number of variables, reduce multi-co-linearity, & improve ‘time taken?”
#
# ## Principal Component Analysis (PCA)
# PCA can transform original low level variables to a higher dimensional space and thus reduce the number of required variables. All co-linear variables get clubbed together. Let’s do a PCA of the data and see what are the main PC’s:
# + id="RKNeM8oM3xPX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 353} outputId="5ca3e881-34b9-44bd-bde0-ee6fc9b7a3f5"
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaled_df = scaler.fit_transform(x)
pca = PCA(n_components = 20)
pca_vectors = pca.fit_transform(scaled_df)
for index,var in enumerate(pca.explained_variance_ratio_):
print('Explained Variance ratio by Principal Component ',(index+1)," : ",var)
# + id="qQPjGRfO35es" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="764d98dd-759c-4221-9c6a-bf3276ed4b33"
pca.explained_variance_ratio_.sum()
# + [markdown] id="QdBf6vsH4D2n" colab_type="text"
# We mapped 2549 variables to 20 Principal Components. From the above result, it is clear that first 10 PCs are a matter of importance. The total percentage of the explained variance ratio by the first 10 PCs is around 0.737 (0.36 + 0.095 + ..+ 0.012). Or it can be said that the first 10 PCs explain 82% variance of the entire dataset.
#
# So, with this we are able to reduce 2549 variables to 10 variables. That’s a dramatic change, isn’t it? In theory, Principal Components are virtual variables generated from mathematical mapping. From a business angle, it is not possible to tell which physical aspect of the data is covered by them. That means, physically, that Principal Components don’t exist. But, we can easily use these PCs as quantitative input variables to any ML algorithm and get very good results.
#
# For visualisation, let’s take the first two PCs and see how can we distinguish different classes of the data using a ‘scatterplot’.
# + id="MM4mRfb84B4u" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 325} outputId="224110ea-6866-4d27-a0d7-3a1346ac6d1d"
import seaborn as sns; sns.set()
sns.scatterplot(x=pca_vectors[:, 0], y=pca_vectors[:, 1], hue=brainwave_df['label'])
plt.title('Principal Components vs Class distribution', fontsize=16)
plt.ylabel('Principal Component 2', fontsize=16)
plt.xlabel('Principal Component 1', fontsize=16)
plt.xticks(rotation='vertical');
# + id="gLf4t5Q34FRh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="7b0c6f41-3bf2-45b4-e7f8-ee99a59becf2"
# %%time
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
pl_log_reg_pca = Pipeline(steps=[('scaler',StandardScaler()),
('pca',PCA(n_components=2)),
('log_reg',LogisticRegression(multi_class='multinomial',solver='sag',max_iter=200))])
scores = cross_val_score(pl_log_reg_pca,x,y,cv=10,scoring='accuracy')
print('Accuracy for Logistic Regression with 2 Principal Components: ',scores.mean())
# + [markdown] id="ZH9Ce2vu4MHb" colab_type="text"
# Time taken ( 6.69 s) was reduced but accuracy (81.51%) decreased but its better with components.
#
# Now, let’s take all 10 PCs and run:
# + id="5z7HkmEP4KaI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="24ab22cc-15dd-4faa-91fd-04f16576f55e"
# %%time
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
pl_log_reg_pca = Pipeline(steps=[('scaler',StandardScaler()),
('pca',PCA(n_components=10)),
('log_reg',LogisticRegression(multi_class='multinomial',solver='sag',max_iter=200))])
scores = cross_val_score(pl_log_reg_pca,x,y,cv=10,scoring='accuracy')
print('Accuracy for Logistic Regression with 10 Principal Components: ',scores.mean())
# + [markdown] id="MAma5XsA4TY0" colab_type="text"
# We see an improvement in accuracy (87.80%) compared to 2 PC cases with a marginal increase in ‘time taken’.
#
# So, in both cases we saw low accuracy compared to normal logistic regression, but a significant improvement in ‘time taken’.
#
# Accuracy can be further tested with a different ‘solver’ & ‘max_iter’ parameter. We used ‘sag’ as ‘solver’ with L2 penalty and 200 as ‘max_iter’. These values can be changed to get a variable effect on accuracy.
#
# Though ‘Logistic Regression’ is giving low accuracy, there are situations where it may be needed specially with PCA. In datasets with a very large dimensional space, PCA becomes the obvious choice for ‘linear classifiers’.
# + [markdown] id="QpEuLhaN4k82" colab_type="text"
# ## Artificial Neural Network Classifier (ANN)
# An ANN classifier is non-linear with automatic feature engineering and dimensional reduction techniques. ‘MLPClassifier’ in scikit-learn works as an ANN. But here also, basic scaling is required for the data. Let’s see how it works:
#
#
# + id="O3gVNfHD8wEL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="c33e0e76-0b6c-429a-f966-ae2723e52bdf"
# %%time
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.neural_network import MLPClassifier
pl_mlp = Pipeline(steps=[('scaler',StandardScaler()),
('mlp',MLPClassifier(hidden_layer_sizes=(1275,637)))])
scores = cross_val_score(pl_mlp,x,y,cv=10,scoring='accuracy')
print('Accuracy for ANN: ',scores.mean())
# + [markdown] id="2de6Ub004nuG" colab_type="text"
# Accuracy (97.18%) is very good, though running time is high ( 7min 27s).
#
# The reason for high ‘time taken’ is the rigorous training time required for neural networks, and that too with a high number of dimensions.
#
# It is a general convention to start with a hidden layer size of 50% of the total data size and subsequent layers will be 50% of the previous one. In our case these are (1275 = 2549 / 2, 637 = 1275 / 2). The number of hidden layers can be taken as hyper-parameter and can be tuned for better accuracy. In our case it is 2.
#
# ## Linear Support Vector Machines Classifier (SVM)
# We will now apply ‘Linear SVM’ on the data and see how accuracy is coming along. Here also scaling is required as a preprocessing stage.
# + id="WuVYDXBB8zds" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="4de8e8ff-cb7c-4988-8b31-444638b79f6f"
# %%time
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.svm import LinearSVC
pl_svm = Pipeline(steps=[('scaler',StandardScaler()),
('svc',LinearSVC())])
scores = cross_val_score(pl_svm,x,y,cv=10,scoring='accuracy')
print('Accuracy for Linear SVM : ',scores.mean())
# + [markdown] id="8Bp21NJF4udC" colab_type="text"
# Accuracy is coming in at 96.4% which is little less than ‘RandomForest’ or ‘ANN’. ‘time taken’ is 1min 19s which is in far better than ‘ANN’.
#
# ## Extreme Gradient Boosting Classifier (XGBoost)
# XGBoost is a boosted tree based ensemble classifier. Like ‘RandomForest’, it will also automatically reduce the feature set. For this we have to use a separate ‘xgboost’ library which does not come with scikit-learn.
#
# Let’s see how it works:
# + id="q8NaH9tR4v-z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="c320c0d6-f1f2-437e-de44-4354cfc3bef4"
# %%time
import xgboost as xgb
pl_xgb = Pipeline(steps=[('xgboost',xgb.XGBClassifier(objective='multi::softmax'))])
scores = cross_val_score(pl_xgb,x,y,cv=10)
print('Accuracy for XGBoost Classifier : ',scores.mean())
# + [markdown] id="ZWP8pjUE4zC3" colab_type="text"
# Accuracy (99.4%) is exceptionally good, but ‘time taken’(15 min) is quite high. Nowadays, for complicated problems, XGBoost is becoming a default choice for Data Scientists for its accurate results. It has high running time due to its internal ensemble model structure. However, XGBoost performs well in GPU machines.
# + [markdown] id="KXgWnDtK42Nb" colab_type="text"
# # Conclusion
# ### From all of the classifiers, it is clear that for accuracy ‘XGBoost’ is the winner. But if we take ‘time taken’ along with ‘accuracy’, then ‘RandomForest’ is a perfect choice. But we also saw how to use a simple linear classifier like ‘logistic regression’ with proper feature engineering to give better accuracy. Other classifiers don’t need that much feature engineering effort.
# ### It depends on the requirements, use case, and data engineering environment available to choose a perfect ‘classifier’.
| Multi_Class_Classification_using_Brainwave_Dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problem Set
#
# "For the things we have to learn before we can do them, we learn by doing them." - Aristotle
#
# There's nothing quite like working with a new tool to really understand it, so we have put together some exercises through this book to give you a chance to put into practice what you learned in the previous lesson(s).
# ## Problems using NDarray [(Official Documentation)](https://mxnet.incubator.apache.org/api/python/ndarray/ndarray.html)
#
#
# Problem 1: Initialize an ndarray of dimension 1x256 on the GPU without overwriting its memory. Then, find the index corresponding to the maximum value in the array (argmax)
# +
# Problem 1 Work Area
# -
# ## Problems from Linear Algebra
#
# Problem 2: Create a 4x4 matrix of random values (where values are uniformly random on the iterval [0,1]. Then create an 4x4 identity matrix (an identity of size n is the n × n square matrix with ones on the main diagonal and zeros elsewhere). Multiply the two together and verify that you get the original matrix back.
# +
# Problem 2 Work Area
# -
# Problem 3: Create a 3x3x20 tensor such that at every x,y coordinate, moving through the z coordinate lists the [Fibonacci sequence](https://en.wikipedia.org/wiki/Fibonacci_number). So, at a z position of 0, the 3x3 matrix will be all 1s. At z-position 1, the 3x3 matrix will be all 1s. At z-position 2, the 3x3 matrix will be all 2s, at z-position 3, the 3x3 matrix will be all 3s and so forth.
#
# Hint: Create the first 2 matrices by hand and then use element-wise operations in a loop to construct the rest of the tensor.
# +
# Problem 3 Work Area
# -
# Problem 4: What is the sum of the vector you created? What is the mean?
# +
# Problem 4 Work Area
# -
# Problem 5: Create a vector [0,1], and another vector [1,0], and use mxnet to calculate the angle between them. Remember that the dot product of two vectors is equal to the cossine of the angle between the vectors, and that the arccos function is the inverse of cosine.
# +
# Problem 5 Work Area
# -
# ## Problems from Probability
#
# Problem 6: In the classic game of Risk, the attacker can roll a maximum of three dice, while the defender can roll a maximum of two dice. Simulate the attacking and defending dice using `sample_multinomial` to try to estimate the odds that an attacker will win against a defender when both are rolling the maximum number of dice.
# +
# Problem 6 Work Area
# -
# ## Problems from Automatic differentiation with ``autograd``
#
# Problem 7: The formula for a parabola is y=ax^2+bx+c. If a=5 and b = 13, what is the slope of y when x=0. How about when x=7?
# +
# Problem 7 Work Area
# -
# Problem 8: Graph the parabola described in Problem 6 and inspect the slope of y when x = 0 and x = 7. Does it match up with your answer from Problem 6?
#
# +
# Problem 8 Work Area
# -
# ## Next
# [Chapter 2: Linear regression from scratch](../chapter02_supervised-learning/linear-regression-scratch.ipynb)
# For whinges or inquiries, [open an issue on GitHub.](https://github.com/zackchase/mxnet-the-straight-dope)
| chapter01_crashcourse/chapter-one-problem-set.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd ###hist_600###
import matplotlib.pyplot as plt
data = pd.read_csv("sensor_data_600.txt", delimiter=" ",
header=None, names = ("date","time","ir","lidar"))
data["lidar"].hist(bins = max(data["lidar"]) - min(data["lidar"]),align='left')
plt.show()
# -
data.lidar.plot() ###plot_all_data##
plt.show()
data.ir.plot()
plt.show()
data["hour"] = [e//10000 for e in data.time] ###hourly_mean###
d = data.groupby("hour")
d.lidar.mean().plot()
plt.show()
d.lidar.get_group(6).hist() ###two_mode_hist###
d.lidar.get_group(14).hist()
plt.show()
# +
each_hour = { i : d.lidar.get_group(i).value_counts().sort_index() for i in range(24)} #時間ごとにデータフレームを作成 ###calc_joint_probs
freqs = pd.concat(each_hour, axis=1) #concatで連結
freqs = freqs.fillna(0) #欠損値(NaN)を0で埋める
probs = freqs/len(data) #頻度を確率で
probs #表示
# +
import seaborn as sns ###2d_hist (下のセルも)
sns.heatmap(probs)
plt.show()
# -
sns.jointplot(data["hour"], data["lidar"], data, kind="kde")
plt.show()
p_t = pd.DataFrame(probs.sum()) #各列を合計
p_t.plot()
p_t.transpose() #紙面の関係で表を横並びに
p_t.sum() # 1になる
p_z = pd.DataFrame(probs.transpose().sum()) #行と列を転置して各列を合計
p_z.plot()
p_z.transpose()
p_z.sum()
cond_z_t = probs/p_t[0] #列(時間)ごとにP(t)で割るとP(x|t)となる ###lidar600cond
cond_z_t
(cond_z_t[6]).plot.bar(color="blue", alpha=0.5) ###lidar600pxt###
(cond_z_t[14]).plot.bar(color="orange", alpha=0.5)
plt.show()
# +
cond_t_z = probs.transpose()/probs.transpose().sum() #行と列を入れ替えて同様に計算するとP(t|z)となる ###lidar600bayes1
print("P(z=630) = ", p_z[0][630]) #センサ値が630になる確率(何時かの情報はない)
print("P(t=13) = ", p_t[0][13]) #時間が13時である確率
print("P(t=13 | z = 630) = ", cond_t_z[630][13])
print("Bayes P(z=630 | t = 13) = ", cond_t_z[630][13]*p_z[0][630]/p_t[0][13])
print("answer P(z=630 | t = 13) = ", cond_z_t[13][630]) #13時にセンサ値が630
# -
def bayes_estimation(sensor_value, current_estimation): ###lidar600bayes2
new_estimation = []
for i in range(24):
new_estimation.append(cond_z_t[i][sensor_value]*current_estimation[i])
print(new_estimation)
return new_estimation/sum(new_estimation) #正規化
estimation = bayes_estimation(630, p_t[0]) ###lidar600bayesonestep
plt.plot(estimation)
# +
values_5 = [630,632,636] #sensor_data_600.txtから拾ってきた5時台のセンサ値 ###lidar600bayesestm1
estimation = p_t[0]
for v in values_5:
estimation = bayes_estimation(v, estimation)
plt.plot(estimation)
# +
values_11 = [617,624,619] #sensor_data_600.txtから拾ってきた11時台のセンサ値 ###lidar600bayesestm2
estimation = p_t[0]
for v in values_11:
estimation = bayes_estimation(v, estimation)
plt.plot(estimation)
# -
estimation
| section_sensor/lidar_600.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
students_performans = pd.read_csv('https://stepik.org/media/attachments/course/4852/StudentsPerformance.csv')
students_performans.head()
students_performans.describe()
students_performans.dtypes
students_performans.shape
students_performans.groupby('gender').aggregate({'writing score': 'mean'})
students_performans.head()
students_performans.iloc[0:5, 1:3]
students_performans.iloc[[1,3,5], [0, 5, -1]]
students_performans.select_dtypes(include=[int], exclude=[object]).head(5)
students_performans.dtypes
students_performans.index
students_performans.get_dtype_counts()
titanic = pd.read_csv('https://stepik.org/media/attachments/course/4852/titanic.csv')
titanic.shape
titanic.get_dtype_counts()
students_performans_with_names = students_performans.iloc[[1,3,5]]
students_performans_with_names.index = ["Liza", "Pavel", "Oleg"]
students_performans_with_names
students_performans_with_names.loc[['Liza'], ['gender']]
type(students_performans.iloc[:, 0])
serie1 = pd.Series([1, 2, 3], index=['INDEX1', 'INDEX2', 'INDEX3'])
serie2 = pd.Series([4, 5, 6], index=['INDEX1', 'INDEX2', 'INDEX3'])
df = pd.DataFrame({'col_name_1': serie1, 'col_name_2': serie2})
df
students_performans_with_names['gender']
students_performans_with_names[['gender']]
| notebooks/1.4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 1: Simple Inference
#
# In this exercise, you will be running inference on a few images.
# The primary goal of the first exercise is to familiarize you with the workflow for inference.
#
# You will be creating a vehicle detection application where the model counts how many vehicles are found in an image. The image you will use is:
#
# <img src="resources/cars_1900_first_frame.jpg">
#
# There appear to be 9 vehicles in the image. Let's see how the computer vision models do.
#
# ### Important! The quiz will ask you how many vehicles were detected in the last step.
#
# ## Step 1: Running on the DevCloud
#
# In the exercises of this tutorial including this one, we will be taking a look at running inferences on a variety of devices like VPU and FPGA.
# In order to access these devices, we will be taking advatnage of the Intel DevCloud for Edge.
# This is a free cloud service from Intel Corporation where users get to try some of the newest edge devices from Intel.
# For more information as well as instructions to sign up for an account, visit [devcloud.intel.com](https://devcloud.intel.com).
# For the exercises in this tutorial you do not need to sign up for an account on the DevCloud.
# We will just be sending your submission to the cloud to run inference.
#
# To run a workload on the DevCloud, we must submit a **job** to a job queue.
# A job consists of a bash script to run the workload, along with any auxillary file used in the workload.
# When submitting a job to a cloud, you can request a certail hardware resource such as VPU or FPGA.
# Then these files are sent over to the DevCloud, the bash script is run, and finally the result is made available to us.
#
# For this first step, we'll start by running a simple workload on the DevCloud.
# Using the `model_downloader.py`, let's get the list of available models for inference.
# The `model_downloader.py` is already available on the DevCloud, so for this we just need to create the bash script.
# In jupyter, cells beginning with `%%writefile myfile.txt` will dump the contents of the cell into a file, named `myfile.txt` in this case.
# So the next code cell creates a file `show_all_models.sh`.
#
# `show_all_models.sh` is currently set up to print the help menu of the `model_downloader.py`.
# Modify the following cell so that it displays the list of all the available models instead.
# %%writefile show_all_models.sh
/opt/intel/openvino/deployment_tools/tools/model_downloader/downloader.py --help
# To submit this to the DevCloud we need to make a REST API call.
# However this part is beyond the scope of this tutorial.
# So we have provided the utility function `submitToDevCloud()` for submitting your job.
# This function will submit your job, and (by default) print the stdout and stderr.
#
# This function has two required arguments, and one optional argument.
# - script: String. Relative path to the bash script to run. Required.
# - node_type: String. Selects the node to request. Must be one of CPU, GPU, VPU, FPGA. Required.
# - script_args: List of strings. Arguments to pass to the bash script. Optional. Default is empty array.
# - files: List of strings. Specifies the auxiliary files. Optional. Default is empty array.
#
# Run the following cell to submit `show_all_models.sh` to the job queue.
from devcloud_utils import submitToDevCloud
submitToDevCloud("show_all_models.sh", "CPU")
# That's it! You have just run a workload on the DevCloud.
#
# ## Step 2: Running Inference
#
# Next step is running the inference itself.
# This section will make use of the Inference Engine that we have covered.
#
# To run the workload on the DevCloud, we need to create two files.
# We need a python script that will run the inference using Inference Engine.
# Additionally, we need a shell script that will start this python file.
# Let's start with the python file, which we call `main.py`.
#
# The instructions for completing the file are broken into steps. In the cell, the parts that need to be modified are signified by `##! ... !##` The number in parentheses shows the step in the instruction that this corresponds to.
#
# **You can click on the steps to get detailed instructions.**
# Follow the instructions to complete `main.py`.
# If you get stuck on any of the stps, refer to the slide deck from course 1 video 6.
# <details>
# <summary><b>(2.1)</b> Complete the <code>prepImage()</code> function by finding the NCHW values from the network.</summary>
#
# The input image will be loaded using OpenCV, so several image processing steps needs to be done.
# First, it will have the wrong size. so the image must be reshaped using `cv2.resize()` function.
# Second, OpenCV loads an image in HWC format whereas the network expects an NCHW format.
# So the image must first be transposed using `transpose()` method of numpy arrays.
# Then the N dimention must be added using the `reshape()` method.
#
# As the preprocesisng is outside of the toolkit, they are already implemented for you.
# However it is missing the sizes for dimensions NCHW of the network input.
#
# Complete the `prepImage()` function by getting the values for `n`, `c`, `h` and `w` from the function input `net`.
#
# </details><br/>
#
# <details>
# <summary><b>(2.2)</b> Create IECore and use it to create IENetwork object for <code>vehicle-detection-adas-0002</code>. Use the provided paths for the models.</summary>
#
# Remember that you need to create IECore object first.
# The model `vehicle-detection-adas-0002` has already been downloaded for you.
# The `IENetwork` is created with `read_network()` method of `IECore` which takes the path to the two files as input.
#
# </details><br/>
#
# <details>
# <summary><b>(2.3)</b> Preprocess the image with <code>prepImage()</code>.</summary>
#
# Prepare the image for inference using the prepImage() function from earlier. Remember that this function outputs a modiified image, instead of doing the modification in place.
#
# </details><br/>
#
# <details>
# <summary><b>(2.4)</b> Create ExecutableNetwork object.</summary>
#
# ExecutableNetwork is created from the load_network() method of IECore object. The IENetwork should be the one created earlier, and the device should be the one in `device` variable. This variable is set by the commandline input to the main.py script.
#
# </details><br/>
#
# <details>
# <summary><b>(2.5)</b> Run synchronous inference.</summary>
#
# Synchronous, or blocking, run is started with `infer()` method. Remember that for the `inputs` argument, you will need the name of the input layer so that you can input the image.
#
# </details><br/>
#
# <details>
# <summary><b>(2.6)</b> Get the output array and run <code>printCount()</code> to see how many vehicles were detected.</summary>
#
# Remmber that the output is a dictionary, with output layer name as key and the result array as value. You will need the name of the output layer to get the output array.
#
# `printCount()` takes the output array and counts the number of vehicle locations with confidence greater than the `prob_threshold` argument (default 50%).
#
# A quick explanation of what `prinCount` does. `vehicle-detection-adas-0002` returns 100 potential regions where an object might be.
# For every potential object, the model assigns a probability that it is an object.
# So to find the vehicles in the image, you need to look for entries over a certain threshold probability.
# The function simply loops over the potential regions and counts the number of regions with confidence values greater than the `prob_threshold`.
#
# The model also provides bounding boxes for where the potential object is, and it returns an index to the detected object.
# Though not shown in this example, this information can be processed and placed on the original image.
#
# </details><br/>
#
# +
# %%writefile main.py
import os
import sys
from openvino.inference_engine import IECore, IENetwork
import cv2
# Prepares image for imference
# inputs:
# orig_image - numpy array containing the original, unprocessed image
# net - IENetwork object
# output:
# preprocessed image.
def prepImage(orig_image, net):
##! (2.1) Find n, c, h, w from net !##
input_blob = next(iter(net.inputs))
n, c, h, w = net.inputs[input_blob].shape
input_image = cv2.resize(orig_image, (w, h))
input_image = input_image.transpose((2, 0, 1))
input_image.reshape((n, c, h, w))
return input_image
# Processes the result. Prints the number of detected vehices.
# inputs:
# detected_obects - numpy array containing the ooutput of the model
# prob_threashold - Required probability for "detection"
# output:
# numpy array of image wtth rectangles drawn around the vehicles.
def printCount(detected_objects, prob_threshold=0.5):
detected_count = 0
for obj in detected_objects[0][0]:
# Draw only objects when probability more than specified threshold
if obj[2] > prob_threshold:
detected_count+=1
print("{} vehicles detected.".format(detected_count))
# Getting the device as commandline argument
device = sys.argv[1]
##! (2.2) create IECore and IENetwork objects for vehicle-detection-adas-0002 !##
xml_path="/data/intel/vehicle-detection-adas-0002/FP16-INT8/vehicle-detection-adas-0002.xml"
bin_path="/data/intel/vehicle-detection-adas-0002/FP16-INT8/vehicle-detection-adas-0002.bin"
image_path = "cars_1900_first_frame.jpg"
original_image = cv2.imread(image_path)
##! (2.3) Preprocess the image. !##
ie = IECore()
net = ie.read_network(model = xml_path, weights = bin_path)
preprocessed_img = prepImage(original_image, net)
##! (2.4) Create ExecutableNetwork object. Use the device variable for targetted device !##
exec_net = ie.load_network(network = net, device_name = device, num_requests = 1)
input_blob = next(iter(net.inputs))
output_blob = next(iter(net.outputs))
##! (2.5) Run synchronous inference. !##
exec_net.infer({input_blob: preprocessed_img})
##! (2.6) Run printCount. Make sure you extracted the array result form the dictionary returned by infer(). !##
result = exec_net.requests[0].outputs[output_blob]
printCount(result, prob_threshold = 0.01)
# -
# Now that you have the python script, we need the bash script.
# This script is provided for you.
# Run the following cell to create `run.sh`.
# %%writefile run.sh
DEVICE=$1
source /opt/intel/openvino/bin/setupvars.sh
python3 main.py $DEVICE
# One important note here is that the bash script takes in one argument (`$1` is the first positional argument). This input is then passed on to the Python script so that you can set the device argument. Inputs to the script are passed in using the `script_args` argument.
#
# Run the following cell to run the job on the CPU.
from devcloud_utils import submitToDevCloud
submitToDevCloud("run.sh", "CPU", script_args=["CPU"], files=["cars_1900_first_frame.jpg","main.py"])
# Now run the following cell to run the job on the GPU. Note that this one will take noticeably longer. There is some on-the-fly compiling of OpenCL for GPU, which takes some time.
from devcloud_utils import submitToDevCloud
submitToDevCloud("run.sh", "GPU", script_args=["GPU"], files=["cars_1900_first_frame.jpg","main.py"])
#
# Congratulations! You have successfully run the inference workload on a GPU and a CPU.
#
# ## Step 4: Quiz question
#
# For the final step, let's try lowering the required confidence to 0.01 (e.g. 1%) and see how many vehicles are detected by the model.
# You will have to go back to the main.py cell, and add a `prob_threshold` argument to the `printCount()` function.
# **The quiz will ask you how many vehicles were detected by the `vehicle-detection-adas-0002` on the CPU device with this setting.**
| Lab_Exercise1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # mmdcornea
# Cornea cells marking.
# ## Description
# This procedure creates a maker for each cell in a very poor quality microscopic image of a cornea. The composition of an opening with the regional maximum is used to create the markers.
import numpy as np
from PIL import Image
import ia870 as ia
# # Reading and topographic view
# The gray-scale image of the cornea is read and displayed. A topographic model is also displayed. We can notice that the cells are formed by small hills in the topographic model. We can also notice that the image is very noisy.
a_pil = Image.open('../data/corneacells.tif').convert('L')
# b = mmsurf(a);
a_pil
a = np.array (a_pil)
# # Filtering and cell detection
# The image is filtered by an alternating sequential filtering with size 2. This filter is composed by openings and closings, removing small peaks and valleys. Next, the regional maxima are detected. For illustrative purpose, they are displayed overlayed on the topographic image view. These regional maxima are the markers for each cell. If anything goes wrong in this step, the error will be propagated throughout the process.
c = ia.iaasf(a,'OC',ia.iasecross(),2)
d = ia.iaregmax(c)
Image.fromarray(ia.iagsurf(c))
Image.fromarray(ia.iagshow(ia.iagsurf(c), d).transpose(1,2,0))
# # Find the background marker
# Following the paradigm of segmentation by watershed, the background marker is detected by applying the constrained watershed on the negation of the cells image using the markers detected in the last step. These watershed lines partition the image in regions of influence of each cell. For illustrative display, the negative of the cell image is displayed overlayed by the markers on the left, and also overlayed by the watershed lines on the right.
e = ia.ianeg(a);
f = ia.iacwatershed(e, d, ia.iasebox());
Image.fromarray(ia.iagshow(e, d).transpose(1, 2, 0))
Image.fromarray(ia.iagshow(e, f, d).transpose(1, 2, 0))
# # Labeling the markers and gradient
# As the internal and external markers can be touching, we combine the external marker with value 1 with the labeling of the internal markers added by 1. The labeled marker image is shown on the left. The final watershed will be applied on the gradient of the original image, which is shown on the right.
g = ia.iagray(f, 'uint16', 1);
h1 = ia.iaaddm(ia.ialabel(d), np.uint16(1));
h = ia.iaintersec(ia.iagray(d,'uint16'), h1);
i = ia.iaunion( g, h);
Image.fromarray(ia.iaglblshow(i).transpose(1, 2, 0))
#mmshow(j);
j = ia.iagradm( a)
Image.fromarray((255.*j/j.max()).astype(np.uint8))
# ## Constrained watershed of the gradient from markers
# Apply the constrained watershed on the gradient from the labeled internal and external markers. Show the watershed lines on the left and the results overlayed on the original image, on the right.
k = ia.iacwatershed(j, i)
Image.fromarray(k.astype(np.uint8)*255)
Image.fromarray(ia.iagshow(a, k, k).transpose(1, 2, 0))
| notebooks/dcorneacells.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import ipytest
import ipytest.magics
import pytest
from tests.src.data.test_dbpedia_utils import expected_harvard_university_data
from tests.src.data.test_dbpedia_utils import expected_institute_for_quantum_computing_data
from tests.src.data.test_dbpedia_utils import expected_multiple_country_url_data
from tests.src.data.test_dbpedia_utils import expected_multiple_country_no_url_data
from tests.src.data.test_dbpedia_utils import expected_two_lat_long_data
from tests.src.data.test_dbpedia_utils import expected_no_data
ipytest.config.rewrite_asserts = True
__file__ = '../../tests/notebooks/test_2.3-process-places-raw-data.ipynb'
# -
@pytest.fixture
def expected_imputed_lat_long():
expected = places.copy()
expected.loc[expected.fullName == 'Banaras Hindu University',
['lat', 'long']] = [25.28000068664551, 82.95999908447266]
expected.loc[expected.fullName == 'Bryn Mawr College', ['lat', 'long']] = (
expected.loc[expected.fullName == 'Bryn Mawr, Pennsylvania', ['lat', 'long']].values)
expected.loc[expected.fullName == 'City University of New York',
['lat', 'long']] = [40.71269989013672, -74.00589752197266]
expected.loc[expected.fullName == 'École Normale Supérieure', ['lat', 'long']] = (
expected.loc[expected.fullName == 'Paris', ['lat', 'long']].values)
expected.loc[expected.fullName == 'Forman Christian College', ['lat', 'long']] = (
expected.loc[expected.fullName == 'Lahore', ['lat', 'long']].values)
expected.loc[expected.fullName == 'Ghent University', ['lat', 'long']] = (
expected.loc[expected.fullName == 'Ghent', ['lat', 'long']].values)
expected.loc[
expected.fullName == 'Gh<NAME>han Institute of Engineering Sciences and Technology',
['lat', 'long']] = [34.11666488647461, 72.46666717529297]
expected.loc[expected.fullName == 'Government College University (Lahore)',
['lat', 'long']] = (
expected.loc[expected.fullName == 'Lahore', ['lat', 'long']].values)
expected.loc[expected.fullName == 'Indian Association for the Cultivation of Science',
['lat', 'long']] = (
expected.loc[expected.fullName == 'Kolkata', ['lat', 'long']].values)
expected.loc[expected.fullName == 'International Centre for Theoretical Physics',
['lat', 'long']] = [45.63333511352539, 13.80000019073486]
expected.loc[expected.fullName == 'Kanagawa University',
['lat', 'long']] = [35.47694396972656, 139.6294403076172]
expected.loc[expected.fullName == 'Leiden University', ['lat', 'long']] = (
expected.loc[expected.fullName == 'Leiden', ['lat', 'long']].values)
expected.loc[expected.fullName == 'Lower Canada College', ['lat', 'long']] = (
expected.loc[expected.fullName == 'Montreal', ['lat', 'long']].values)
expected.loc[expected.fullName == 'Luther High School North', ['lat', 'long']] = (
expected.loc[expected.fullName == 'Chicago', ['lat', 'long']].values)
expected.loc[expected.fullName == 'National and Kapodistrian University of Athens',
['lat', 'long']] = [37.983810, 23.727539]
expected.loc[expected.fullName == 'National University of Kharkiv',
['lat', 'long']] = (
expected.loc[expected.fullName == 'Kharkiv', ['lat', 'long']].values)
expected.loc[expected.fullName == 'Radboud University Nijmegen', ['lat', 'long']] = (
expected.loc[expected.fullName == 'Nijmegen', ['lat', 'long']].values)
expected.loc[expected.fullName == 'Saha Institute of Nuclear Physics',
['lat', 'long']] = (
expected.loc[expected.fullName == 'Kolkata', ['lat', 'long']].values)
expected.loc[expected.fullName == 'The Albany Academy', ['lat', 'long']] = (
expected.loc[expected.fullName == 'Albany, New York', ['lat', 'long']].values)
expected.loc[expected.fullName == 'The Bronx High School of Science',
['lat', 'long']] = (
expected.loc[expected.fullName == 'The Bronx', ['lat', 'long']].values)
expected.loc[expected.fullName == 'University of Azad Jammu and Kashmir',
['lat', 'long']] = [34.36100006103516, 73.46199798583984]
expected.loc[expected.fullName == 'University of California, Santa Barbara', ['lat', 'long']] = (
expected.loc[expected.fullName == 'Santa Barbara, California', ['lat', 'long']].values)
expected.loc[expected.fullName == 'University of Hawaii',
['lat', 'long']] = [21.29999923706055, -157.8166656494141]
expected.loc[expected.fullName == 'University of Lisbon',
['lat', 'long']] = [38.71381759643555, -9.139386177062988]
expected.loc[expected.fullName == 'University of Manchester Institute of Science and Technology',
['lat', 'long']] = (
expected.loc[expected.fullName == 'Manchester', ['lat', 'long']].values)
expected.loc[expected.fullName == 'University of Science and Technology of China',
['lat', 'long']] = (
expected.loc[expected.fullName == 'Hefei', ['lat', 'long']].values)
expected.loc[expected.fullName == 'Victoria University of Manchester', ['lat', 'long']] = (
expected.loc[expected.fullName == 'Manchester', ['lat', 'long']].values)
expected.loc[expected.fullName == 'Virtual University of Pakistan', ['lat', 'long']] = (
expected.loc[expected.fullName == 'Lahore', ['lat', 'long']].values)
return expected
# +
# %%run_pytest[clean] -v --color=yes
def test_create_place_data_harvard_university(
expected_harvard_university_data):
assert(create_place_data(json_lines[612]) ==
expected_harvard_university_data)
def test_create_place_data_institute_for_quantum_computing(
expected_institute_for_quantum_computing_data):
assert(create_place_data(json_lines[684]) ==
expected_institute_for_quantum_computing_data)
def test_create_place_data_multiple_country_url(
expected_multiple_country_url_data):
assert(create_place_data(json_lines[132]) ==
expected_multiple_country_url_data)
def test_create_place_data_multiple_country_no_url(
expected_multiple_country_no_url_data):
assert(create_place_data(json_lines[789]) ==
expected_multiple_country_no_url_data)
def test_create_place_data_two_lat_long(
expected_two_lat_long_data):
assert(create_place_data(json_lines[574]) ==
expected_two_lat_long_data)
def test_create_place_data_no_data(
expected_no_data):
assert(create_place_data(json_lines[801]) ==
expected_no_data)
def test_impute_lat_long(expected_imputed_lat_long):
imputed_places = impute_lat_long(places)
assert(imputed_places.equals(expected_imputed_lat_long))
| tests/notebooks/test_2.3-process-places-raw-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (py2)
# language: python
# name: py2
# ---
# +
import igraph
import glob
import copy
import networkx as nx
import numpy as np
import pandas as pd
from matplotlib import pylab as plt
import matplotlib.patches as mpatches
import matplotlib.lines as mlines
# %matplotlib inline
# +
""" get all files """
files = glob.glob('./matrix_csvs/*')
g_MW_files = sorted([f for f in files if ((f.split('/')[-1].split('_')[1]=='green')and (f.split('/')[-1].split('_')[2]=='projection.csv') ) ])
b_MW_files =sorted([f for f in files if ((f.split('/')[-1].split('_')[1]=='blue') and (f.split('/')[-1].split('_')[2]=='projection.csv') )])
all_MW_files = sorted([f for f in files if ((f.split('/')[-1].split('_')[1]=='all') and (f.split('/')[-1].split('_')[2]=='projection.csv'))])
print all_MW_files
# +
""" get a dict going with files and data """
data = {}
years = range(2007,2018)
years_m = [2007,2008,2009,2010,2012,2013,2014,2015,2016,2017]
for y in years_m:
fname_all = [f for f in all_MW_files if ((str(y)+'.75') in f)][0]
fname_g = [f for f in g_MW_files if ((str(y)+'.75') in f)][0]
fname_b = [f for f in b_MW_files if ((str(y)+'.75') in f)][0]
data[y]={'year':y,
'fname_all':fname_all,
'data_all':0.0,
'fname_g':fname_g,
'data_g':0.0,
'fname_b':fname_b,
'data_b':0.0}
#2011 notes
data[2011]={'data_b': 0.0,
'data_g': 0.0,
'year': 2011,
'fname_g': './matrix_csvs/2011.5_green_projection.csv',
'fname_b': './matrix_csvs/2011.5_blue_projection.csv',
'data_all': 0.0,
'fname_all': './matrix_csvs/2011.5_all_projection.csv'}
for y in years:
print data[y]
# +
""" read in all the data """
for y in years:
print y
data[y]['data_all']=pd.read_csv(data[y]['fname_all'], encoding='utf-8').set_index('Unnamed: 0')
data[y]['data_g']=pd.read_csv(data[y]['fname_g'], encoding='utf-8').set_index('Unnamed: 0')
data[y]['data_b']=pd.read_csv(data[y]['fname_b'], encoding='utf-8').set_index('Unnamed: 0')
#data[y]['data_ff'] = data[y]['data_all'].subtract(data[y]['data_g'], fill_value=0.0).subtract(data[y]['data_b'], fill_value=0.0)
print data[2017]['data_all'][data[2017]['data_all'].index=='UA'].sum().sum()
# -
def softmax(x):
return np.exp(x) / np.sum(np.exp(x), axis=0)
def degree(x):
#print x
#print 'softmax: ', softmax(x)
#raw_input('-->')
return np.sum(softmax(x)/np.max(x))/x.shape[0]
# +
""" get dregree for all data"""
for y in years:
print y
for data_str in ['data_all','data_g','data_b']:
df = data[y][data_str]
#degree just with connections
df['degree_simple'] = df[df>0].count(axis=1)
#degree with algo
df['degree_norm'] = df.apply(lambda row: degree(row/np.sum(row)), axis=1)
cols = [c for c in list(df) if len(c)<3]
df['sum'] = df[cols].sum(axis=1)
df['cumsum'] = df['sum'].cumsum()
data[y][data_str] = df.sort_values('sum', ascending=False)
# +
"""join with region and portions"""
country_info = pd.read_csv('all_portions.csv', encoding='utf-8').set_index('iso2')
print country_info
for y in years:
print y
for data_str in ['data_all','data_g','data_b']:
data[y][data_str] = data[y][data_str].join(country_info[['SIPS_REGION',str(y)+'_all',str(y)+'_g',str(y)+'_b',str(y)+'_portion']])
# -
print list(data[2017]['data_all'])
print data[2017]['data_all'].SIPS_REGION.unique()
print data[2017]['data_all'].degree_norm
print data[2017]['data_all']['2017_portion']
# +
""" portion green vs. degree vs. region vs. size """
cols_dict = {
'OTHERX':'#7700a3',
'US':'#00ffcb',
'LAM':'#ffb200',
'OECD_EUR':'#0021ff',
'OECD_PAC':'#00cbff',
'OECD_AMX':'#0f0066',
'AFRICA':'#00ff08',
'ME':'#3ea535',
'CN':'#ff0043',
'IN':'#ff00d4',
'TE':'#6b5900'
}
f, axarr = plt.subplots(2,sharex=True, figsize=(16,9))
for r in axarr:
#for c in r:
#c.set_yscale('log')
r.set_xscale('log')
ii=0
for y in [2007,2017]:
#print data[y]['data_g'].shape
#print data[y]['data_all'].SIPS_REGION
#print data[y]['data_all'].loc['UA']
#print data[y]['data_all'].shape
data[y]['data_all']['color'] = data[y]['data_all'].apply(lambda row: cols_dict[row['SIPS_REGION']], axis=1)
#print data[y]['data_all']['color']
df = data[y]['data_all']
print df.color
print df.degree_simple
axarr[ii].scatter(df['degree_simple'],df[str(y)+'_portion'],s=df[str(y)+'_all']/1000, c=df['color'])
ii+=1
plt.show()
#print data[2017]['data_all']['por_green']
# +
""" degree vs time """
cols_dict = {
'OTHERX':'#7700a3',
'US':'#00ffcb',
'LAM':'#ffb200',
'OECD_EUR':'#0021ff',
'OECD_PAC':'#00cbff',
'OECD_AMX':'#0f0066',
'AFRICA':'#00ff08',
'ME':'#3ea535',
'CN':'#ff0043',
'IN':'#ff00d4',
'TE':'#6b5900'
}
f, axarr = plt.subplots(3,sharex=True, figsize=(16,9))
axarr[0].set_xticks(years)
#for r in axarr:
#for c in r:
#c.set_yscale('log')
# r.set_xscale('log')
print data[2017]['data_g'].loc['IN'].degree_simple
ii=0
for data_str in ['data_all','data_b','data_g']:
degree_means = {}
degree_std = {}
for y in years:
print data[y]['data_g'].loc['IN'].degree_simple
#print data[y]['data_g'].shape
#print data[y]['data_all'].SIPS_REGION
#print data[y]['data_all'].loc['UA']
#print data[y]['data_all'].shape
#data[y]['data_all']['color'] = data[y]['data_all'].apply(lambda row: cols_dict[row['SIPS_REGION']], axis=1)
#print data[y]['data_all']['color']
df = data[y][data_str]
#print list(df)
#print df.color
degree_means[y] = df.groupby('SIPS_REGION').mean().degree_simple.to_dict()
degree_std[y] = df.groupby('SIPS_REGION').std().degree_simple.to_dict()
#print len(df.groupby('SIPS_REGION').mean().degree_simple.to_dict())
for r in df.SIPS_REGION.unique():
print r,[degree_means[y][r] for y in years]
axarr[ii].plot(years, np.array([degree_means[y][r] for y in years]), c=cols_dict[r])
ii+=1
axarr[0].set_ylabel('%$D_{All Generation}$')
axarr[1].set_ylabel('%$D_{Nuclear & Hydro}$')
axarr[2].set_ylabel('%$D_{Renewables}$')
lines = []
labels=[]
for k,v in cols_dict.items():
labels.append(k)
lines.append(mlines.Line2D([0],[0],color=v))
f.legend(lines, labels, loc='center', bbox_to_anchor=(0.5,0.92), ncol=len(cols_dict))
f.suptitle('Degree Distributions for all Countries', fontsize=20)
f.savefig('country_degree.png')
plt.show()
# +
import matplotlib.markers as mmarkers
f, axarr = plt.subplots(nrows=3,ncols=2,sharex=True, figsize=(16,9))
for r in axarr:
for c in r:
#c.set_yscale('log')
c.set_xscale('log')
c.set_ylim(0,60)
#axarr[0].set_yscale('log')
#axarr[1].set_yscale('log')
#axarr[2].set_yscale('log')
#axarr[0].set_xscale('log')
#axarr[1].set_xscale('log')
#axarr[2].set_xscale('log')
#axarr[0].set_xtickabels(years)
#cols_dict={0:'black',1:'blue',2:'green'}
MW_dict = {0:'_all',1:'_b',2:'_g'}
ii=0
jj=-1
for data_str in ['data_all','data_b','data_g']:
jj+=1
ii=0
for y in [2007,2017]:
print data[y]['data_g'].loc['IN'].degree_simple
#print data[y]['data_g'].shape
#print data[y]['data_all'].SIPS_REGION
#print data[y]['data_all'].loc['UA']
#print data[y]['data_all'].shape
#data[y]['data_all']['color'] = data[y]['data_all'].apply(lambda row: cols_dict[row['SIPS_REGION']], axis=1)
#print data[y]['data_all']['color']
df = data[y][data_str]
#print list(df)
#print df.color
#degree_means[y] = df.groupby('SIPS_REGION').mean().degree_simple.to_dict()
#degree_std[y] = df.groupby('SIPS_REGION').std().degree_simple.to_dict()
#print len(df.groupby('SIPS_REGION').mean().degree_simple.to_dict())
for r in df.SIPS_REGION.unique():
axarr[jj,ii].scatter(df[df.SIPS_REGION==r][str(y)+MW_dict[jj]], df[df.SIPS_REGION==r]['degree_simple'], color=cols_dict[r])
ii+=1
lines = []
labels=[]
for k,v in cols_dict.items():
labels.append(k)
lines.append(mlines.Line2D([0],[0],marker=".", linestyle='none', color=v))
f.legend(lines, labels, loc='center', bbox_to_anchor=(0.5,0.93), ncol=len(cols_dict))
axarr[0,0].set_ylabel('$Degree_{Fossil Fuel}$')
axarr[1,0].set_ylabel('$Degree_{Nuclear & Hydro}$')
axarr[2,0].set_ylabel('$Degree_{Renewables}$')
axarr[0,0].set_title('2007')
axarr[0,1].set_title('2017')
axarr[2,0].set_xlabel('Generating Capacity [MW]')
axarr[2,1].set_xlabel('Generating Capacity [MW]')
f.suptitle('Degree Distributions for all Countries', fontsize=20)
#f.legend((black_line,gray_patch), ('$Degree_{Mean}$','$Degree_{Std}}$'), loc='center', bbox_to_anchor=(0.5,0.92), ncol=2)
f.savefig('country_dd_simple.png')
plt.show()
#print np.mean(degree_means)
# -
for y in years:
pos = np.diagonal(data[y]['data_all'])
print pos
plt.imshow(data[y]['data_all'])
plt.show()
# +
#PageRank centrality with igraph
for y in years:
df_all_pairs = [ {k:v for k,v in m.items() if v>0.0} for m in data[y]['data_all'].to_dict(orient='rows')]
df_all_pairs = dict(zip(data[y]['data_all'].index.values,df_all_pairs))
G = igraph.Graph(directed=True)
countries = list(data[y]['data_all'].index)
for c in countries:
G.add_vertex(c)
for k,v in df_all_pairs.items():
#print k
#print v
v.pop(k,None)
for k2,v2 in v.items():
G.add_edge(k, k2, weight=v2)
print len(G.es['weight']), len(pr_all)
pr_all = G.personalized_pagerank(vertices=G.vs, weights=G.es["weight"], directed=True)
#for e in G.vs:
# print e
#print pr_all
#print df_all_pairs
names = list(data[y]['data_all'])
#names_copy = copy.copy(names)
#inds = np.array(pr_all).argsort()[::-1][:]
#sort_names_pr = np.array(names_copy)[inds]
#sort_centrality_pr = np.array(pr_all)[inds]
#sort_names_pr, sort_centrality_pr
plt.plot(pr_all, "o")
plt.show()
# -
# ### Loading matrices and making projections
# +
#Extracting bipartite adj mat
ADJ_bip_green = np.loadtxt(open(path_to_matrices + current_year + "_green_MW.csv", "rb"), delimiter=",", skiprows=1,
usecols=range(1,n_countries+1)) #getting the adjacency matrix
ADJ_bip_blue = np.loadtxt(open(path_to_matrices + current_year + "_blue_MW.csv", "rb"), delimiter=",", skiprows=1,
usecols=range(1,n_countries+1)) #getting the adjacency matrix
#Does the all category include the green and blue or is it just the dirty ones? In principle we would like to have
#only the dirty ones, right?
ADJ_bip_all = np.loadtxt(open(path_to_matrices + current_year + "_all_MW.csv", "rb"), delimiter=",", skiprows=1,
usecols=range(1,n_countries+1))
#To get the high emission plants we use the all matrix and remove the "clean" part
ADJ_bip_dirty = ADJ_bip_all - (ADJ_bip_green + ADJ_bip_blue)
#Making the company projection
ADJ_comp_green = (ADJ_bip_green/ADJ_bip_green.sum()).dot(ADJ_bip_green.T/(ADJ_bip_green.T.sum()))
ADJ_comp_blue = (ADJ_bip_blue/ADJ_bip_blue.sum()).dot(ADJ_bip_blue.T/(ADJ_bip_blue.T.sum()))
ADJ_comp_all = (ADJ_bip_all/ADJ_bip_all.sum()).dot(ADJ_bip_all.T/(ADJ_bip_all.T.sum()))
ADJ_comp_dirty = (ADJ_bip_dirty /ADJ_bip_dirty .sum()).dot(ADJ_bip_dirty.T/(ADJ_bip_dirty .T.sum()))
#Making the country projection
ADJ_country_green = (ADJ_bip_green.T/ADJ_bip_green.T.sum()).dot(ADJ_bip_green/(ADJ_bip_green.sum()))
ADJ_country_blue = (ADJ_bip_blue.T/ADJ_bip_blue.T.sum()).dot(ADJ_bip_blue/(ADJ_bip_blue.sum()))
ADJ_country_all = (ADJ_bip_all.T/ADJ_bip_all.T.sum()).dot(ADJ_bip_all/(ADJ_bip_all.sum()))
ADJ_country_dirty = (ADJ_bip_dirty.T /ADJ_bip_dirty.T .sum()).dot(ADJ_bip_dirty/(ADJ_bip_dirty.sum()))
#Getting the percentage of emissions of each type for each country
green_part = [sum(ADJ_bip_green[:, i])/sum(ADJ_bip_all[:, i]) for i in range(n_countries)]
blue_part = [sum(ADJ_bip_blue[:, i])/sum(ADJ_bip_all[:, i]) for i in range(n_countries)]
dirty_part = [sum(ADJ_bip_dirty[:, i])/sum(ADJ_bip_all[:, i]) for i in range(n_countries)]
# -
# # Multilayer analysis
# +
def make_supra_adj(ADJ_list, COUP_list):
"""Function that takes list of adj matrix and coupling and generates the supra adjacency matrix of the
multiplex network
Args:
ADJ_list: list of numpy 2D array's. Each of them is an adjacency matrix
COUP_list: list of numpy 1D array's. Each of them is the vector to be used for coupling the matrices
Return:
Supra Adjacency matrix. Numpy 2D array.
"""
n_layers = len(ADJ_list)
n_nodes = ADJ_list[0].shape[0]
#stacking up adjacency matrix and coupling accordingly
sup_list = []
for i in range(n_layers):
row = []
for j in range(n_layers):
if i == j: #if in diagonal add adjacency matrix
row.append(ADJ_list[i])
#print(ADJ_list[i].shape)
else: #otherwise add the coupling (diagonal since multiplex network)
row.append(np.diag(COUP_list[i])) #if i or j here defined direction of coupling
#print(np.diag(COUP_list[i]).shape)
sup_list.append(row)
return np.bmat(sup_list)
n_nodes = n_countries
n_layers = 3
#The following function is taylored for igraph centrality
def flatten_centrality(centrality_vector, n=n_nodes, l=n_layers, names=names, hubauth=False):
'''Centrality networks in a Multiplex are commonly expressed as a vector of
nlx1 dimension. To have a centrality for each node it is important to flatten
the vector into nx1 dimension. Also, it is important return a list of ranked
nodes and another of their score.
Args
centrality_vector(numpy array): the centrality measure for each node-layer
n(int): number of nodes
l(int): number of layers
names(list of strings): name of nodes (countries)
hubauth(boolean): indicating if the centrality is hub or authority
Return:
sort_names_multi(list of strings): names of countries ordered by centrality
sort_centrality_multi(list of flots): sorted score of nodes
'''
multi_centrality = []
for i in range(n):
cent = 0
for k in range(l):
cent += centrality_vector[i + n*k]
if hubauth:
multi_centrality.append(cent[0])
else:
multi_centrality.append(cent)
node_names = np.array(copy.deepcopy(names))
inds = np.array(multi_centrality).argsort()[::-1][:]
sort_names_multi = node_names[inds]
sort_centrality_multi = np.array(multi_centrality)[inds]
return sort_names_multi, sort_centrality_multi
# -
#Making the supra adjacency matrix
SUPADJ_country = make_supra_adj([ADJ_country_green, ADJ_country_blue, ADJ_country_dirty], [green_part, blue_part, dirty_part])
#making the graph
G_supra = igraph.Graph.Weighted_Adjacency( SUPADJ_country.tolist() ,mode="directed")
#computing pagerank
pr_muliplex = G_supra.personalized_pagerank(weights=G_supra.es["weight"])
#sorting it appropriately
pr_multiplex_sorted = flatten_centrality(pr_muliplex)
pr_multiplex_sorted
# Comparing it with the aggregate pagerank centrality
G_all = igraph.Graph.Weighted_Adjacency( ADJ_all.tolist() ,mode="directed")
pr_all = G_all.personalized_pagerank(weights=G_all.es["weight"])
names_copy = copy.copy(names)
inds = np.array(pr_all).argsort()[:]
#If small countries are first, try including the [::-1 as below]
#inds = np.array(pr_all).argsort()[::-1][:]
sort_names_pr = np.array(names_copy)[inds]
sort_centrality_pr = np.array(pr_all)[inds]
sort_names_pr, sort_centrality_pr
# # Inconsistency between networkx and igraph?
# I tend to trust igraph more, however the discrepancy might mean that values in general are too close. Perhaps networkx power method is not converging?
# +
#PageRank centrality with igraph
G_all = igraph.Graph.Weighted_Adjacency(ADJ_all.tolist() ,mode="directed")
pr_all = G_all.personalized_pagerank(weights=G_all.es["weight"], directed=True)
names_copy = copy.copy(names)
#inds = np.array(pr_all).argsort()[::-1][:]
inds = np.array(pr_all).argsort()[:]
sort_names_pr = np.array(names_copy)[inds]
sort_centrality_pr = np.array(pr_all)[inds]
sort_names_pr, sort_centrality_pr
#PageRank centrality with networkx
G_all_nx = nx.from_numpy_matrix(ADJ_all)
pr_all_nx = nx.pagerank(G_all_nx, weight="weight")
plt.plot(pr_all_nx.values(), "o")
plt.plot(pr_all, ".")
# -
plt.plot(pr_all, pr_all_nx.values(), "o")
for i in range(n_countries):
print i, "nx = ", names[sorted(pr_all_nx.items(), key=lambda x:x[1])[::-1][i][0]], "ig = ", sort_names_pr[i]
#If we want to do Hubs and Auth the following code will be usefull
"""def G_list_hub(G_adj_list):
'''
Args(list): list of graphs adjacency matrices
Return(list): list of adjacency matrices to be used in the block diagonal
of the supra adjacency matrix for hub score
'''
G_hub = []
for g in G_adj_list:
new = np.dot(g, g.transpose())
G_hub.append(new)
return G_hub
def G_list_auth(G_adj_list):
'''
Args(list): list of graphs adjacency matrices
Return(list): list of adjacency matrices to be used in the block diagonal
of the supra adjacency matrix for auth score
'''
G_auth = []
for g in G_adj_list:
new = np.dot( g.transpose(), g)
G_auth.append(new)
return G_auth
def hub_auth_diag(Coup_list):
coup = []
for c in Coup_list:
coup.append(c*c)
return coup"""
pd.__version__
| Degree_Country.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Nexa And Time Series
# So this is a brief show on the work with time series so far. The first thing that we have to do is to import the classical libraries. We also do a little trick to work with the librarires in the directory above.
# ## Main Libraries
# Scientific Python libraries
import numpy as np
import matplotlib.pyplot as plt
import mpld3
import seaborn as sn
mpld3.enable_notebook()
import sys
sys.path.append("../")
# Nexa in-house libraries
from signals.time_series_class import MixAr
from signals.aux_functions import sidekick
from input.sensors import PerceptualSpace, Sensor
from nexa.nexa import Nexa
# Now we have a couple of imports here. I will explain what al libraries does:
#
# ### Signals
# This is the module to put time series. In this case I am importing a class that allows us to build an autoregressive process (*MixAr*) that can be mix in space with a simpler series (*sidekick*) as Pawell suggested.
#
# ### Input
# This module takes care of the input sides to Nexa, that is a group of sensors. Here I created some classes that allow easy organization and implementation of data with a time dimension. In particular the class *Sensor* represents a single receptive unit that reads a time series from the exterior whereas *Perceptual Space* allows us to deal with a group of them and their interactions.
#
# ### Nexa
# Finally, Nexa. Building in Benjaminsson's previous work I implemented (in far more simpler terms, there is still of testing and optimization to be done) a Nexa framework. The Nexa object contains here a perceptual space which represents a -as stated before- a group of sensors with information on time. The Object contains all the operations that allow the creation of vector codes from the ground up:
#
# * Formation of a *Spatio Temporal Distance Matrix* (STDM) that captures the cross-correlations of a percpetual space.
# * Clustering / Vector quantization in the vector space
# * Index creation. That is, utilities to transform the data from the whole preceptual space to the particular set of indexes of a cluster and the other way around.
# * Clustering / Vector quantization in the data / time space.
# * Code creation.
#
# ## Program Execution and Workflow
# So first we declare and discuss the parameters and setup requiered for a run of Nexa. We declare the time resoultion of the system and the total amount of time that our system will be simulated. In a real data analysis task this will be determined from the domain of the problem but given that we are in the development, toy example phase we determine those quantites by ourselves.
dt = 0.1
Tmax = 100
# ### Time series to analyze
# Now we input the necessary setup for our time series. We present the code here and explain it bellow together with a visualization of both of them.
# +
# Let's get the sideckick function
amplitude = 1
w1 = 1
w2 = 5
beta = sidekick(w1, w2, dt, Tmax, amplitude)
# Now we will get the AR proccess mixed with the sidekick
# First we need the phi's vector
phi0 = 0.0
phi1 = -0.8
phi2 = 0.3
phi = np.array((phi0, phi1, phi2))
# Now we need the initial conditions
x0 = 1
x1 = 1
x2 = 0
initial_conditions = np.array((x0, x1, x2))
# Second we construct the series with the mix
A = MixAr(phi, dt=dt, Tmax=Tmax, beta=beta)
A.initial_conditions(initial_conditions)
mix_series = A.construct_series()
# mix_series = beta
time = A.time
# -
# First we describe the sideckick function, it is specified by two frquencies and the amplitude. Under the hood it is simple a the mix of two sine waves with the given frequency. We visualize it bellow.
# %matplotlib inline
plt.plot(time, beta)
plt.show()
# Now we will visualiza the Auto Regresive process which is a little bit more complicated. In order to specify an autoregresive process we need as many initial conditions as the order of the process. In concrete our AR is:
#
# $$x(t) = \phi_0 + x(t - 1) * \phi_1 + x(t - 2) * \phi_2 $$
#
# It is easy to imagine how to generalize this to any order. Now, the particularity that we introduce is to add also an **spatial** term to this equation.
#
# $$x(t) = \phi_0 + x(t - 1) * \phi_1 + x(t - 2) * \phi_2 + \beta(t)$$
#
# Where beta is our sidekick function.
#
# Our **AR** class therefore takes in its constructor three initial conditions and the corresponding values of phi. We show the plot below and we see the characteristic plot of an AR process.
#
plt.plot(time, mix_series)
# ## Nexa worflow
# Now we present here the nexa worflow but first we need to initialize a couple of parameters and the setup
# Here we will calculate correlations
Nlags = 100
Nspatial_clusters = 2 # Number of spatial clusters
Ntime_clusters = 2 # Number of time clusters
Nembedding = 3 # Dimension of the embedding space
# +
# We create the here perceptual space
aux_sensors = [Sensor(mix_series, dt), Sensor(beta, dt)]
perceptual_space = PerceptualSpace(aux_sensors, Nlags)
# Now the Nexa object
nexa_object = Nexa(perceptual_space, Nlags, Nspatial_clusters,
Ntime_clusters, Nembedding)
# -
# We execute the whole nexa workflow with a single routine
# Calculate all the quantities
nexa_object.calculate_all()
# I decided to implement the routine to calculate the code vectors separte however (discuss this!)
# Build the code vectors
code_vectors = nexa_object.build_code_vectors()
# ## Visualization
# Now in order to discuss this with more detail and show how the whole process looks in at the graph level I present the plots.
#
# First we import all the required libraries.
#
from visualization.sensor_clustering import visualize_cluster_matrix
from visualization.sensors import visualize_SLM
from visualization.sensors import visualize_STDM_seaborn
from visualization.time_cluster import visualize_time_cluster_matrix
from visualization.code_vectors import visualize_code_vectors
# ### Visualize SLM
# First we present the plot of the Sensor Lagged Matrix, which just represents the sensors in our system and all the possible lags until the klags quantity in order to show the overall structure of the time series
# %matplotlib inline
fig = visualize_SLM(nexa_object)
plt.show(fig)
# ### Visualize STDM (Spatio Temporal Distance Matrix)
#
# Now we get the usual correlation matrix between the data with the novelty that we also calculate the correlation between all the possible pairs of laggins and sensors.
# %matplotlib qt
# fig = visualize_STDM(nexa_object)
fig = visualize_STDM_seaborn(nexa_object)
plt.show(fig)
# ### Visualize of Sensor Clusterings
# Now we show how the lagged sensors cluster
# %matplotlib inline
fig = visualize_cluster_matrix(nexa_object)
# ### Visualize the time cluster
# This one is a little bit more tricky. Here we take on of the centers (in this case the second center of the first cluster) and show how the center (code vector) of that and show how it looks. So here we have a center of the first cluster in other words.
# +
# %matplotlib inline
cluster = 0
time_center = 1
fig = visualize_time_cluster_matrix(nexa_object, cluster, time_center,
cmap='coolwarm', inter='none',
origin='upper', fontsize=16)
# -
#
# ### Visualize the Code Vectors
# Here we visualize the code vectors. We show as different cells the different cluster in the sensor space and as different colors the particular code vector that encode the signal at each particular moment in time.
# %matplotlib inline
fig = visualize_code_vectors(code_vectors)
# ### Statistics of the Code Vectors
# Now we calculate the correlation between the two clusters. We expect them to have a very low correlation coefficient.
#
np.corrcoef(code_vectors, rowvar=0)
# ## Discussion and Works To Do
# * New functions instead of an AR or autoregresive process (Anders suggested a square function with variable period)
# * What to do, predict? identify structure? recover the generating patterns?
# * Check more statistics of the code vectors (non all-or-none competition), check distribution of the data in the clusters.
# * Different clusternig algorithms using (Japanese guy paper's)
# # Non-Academic Points to Discuss
# * Got funding for the BrainDisc conference at Freiburg (http://www.bcf.uni-freiburg.de/events/conferences-workshops/20151001-phd-conference). Good point to know other students and interested in some of the talks.
# * NIPS - Funding.
# * Jeannette or Alex, insurance.
# ## Things to do
# * Play with Phis
# * Play with other periodical functions
# * Play with spatial correlaiton that days exponentially -or with another time function-
# * Create lags with arbitrary distribution and distance instead of with linear one.
#
| presentations/.ipynb_checkpoints/2015-august-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <NAME>
# ## Convex Optimization - Prof. Morand
# ### Assignment: Maximize and plot Cobb-Douglas Production Function
import cvxpy as cvx
import numpy as np
import matplotlib.pyplot as plt
# +
# Output(s)
y = cvx.Variable() # Product
# Inputs to production
k = cvx.Variable() # Capital
l = cvx.Variable() # Labor
# Costs of variables
w = cvx.Parameter() # wage
r = cvx.Parameter() # rent
# Budget
M = cvx.Parameter()
# Parameters
A = cvx.Parameter(nonneg=True, value=1) # Factor productivity
# alpha = cvx.Parameter(nonneg=True, value=0.5) # Elasticity
alpha = 0.5
beta = 1 - alpha
# Constraints
constraints = [0 <= k, 0 <= l, w*l + r*k <= M]
# -
# Cobb-Douglas Production
obj = cvx.Maximize(y - A * (k**(alpha) * l**(1 - alpha)))
# Form and solve problem
prob = cvx.Problem(obj, constraints)
# Results
print("status:", prob.status)
print("optimal value: ", prob.value)
print("optimal var: ", k.value, l.value)
| Homework/DaigleMaxCD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pyvisa as visa
rm = visa.ResourceManager()
rm.list_resources()
from ekpmeasure.experiments.ferroelectric._relaxation import Relaxation
# open Scope
scope = rm.open_resource('GPIB0::14::INSTR')
scope.query("*idn?")
# open pulse gen
pg = rm.open_resource('TCPIP0::PULSE-RIDER::inst0::INSTR')
pg.query("*idn?")
# +
exp = Relaxation(pg, scope)
# configure save path
exp.config_path('./computer_control/data/af686free/111121/dataset2/')
# +
# specify parameters
# -
voltages = ['3000mv', '4000mv']
delays = ['200ns', '500ns']
# +
scan_params = {
'delay':short_delays,
'high_voltage':voltages,
}
run_function_args = {
'pg':pg,
'scope':scope,
'pulsewidth':'1000ns',
'identifier':'identifier',
'scope_channel': 'Ch3',
'polarity':'up',
}
order = ['delay', 'high_voltage', ]
# -
# run an n_param_scan
exp.n_param_scan(scan_params, run_function_args, order, ntrials = 1, plot=True)
| src/ekpmeasure/experiments/ferroelectric/_relaxation/_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn.ensemble import RandomForestClassifier
from impreciseshap import ImpreciseShap
from sklearn.metrics import accuracy_score
import pandas as pd
df = pd.read_csv("./data/clust4_1000_250.csv")
df.head()
df_train = df[df['is_test'] == False]
df_test = df[df['is_test'] == True]
X_train, y_train = df_train[['x', 'y']], df_train[['class']]
X_test, y_test = df_test[['x', 'y']], df_test[['class']]
import matplotlib.pyplot as plt
color_dict = {0: "r", 1: "b", 2: "g", 3: "y"}
colors = [color_dict[i] for i in df_train['class']]
plt.scatter(x=df_train['x'], y=df_train['y'], c=colors)
model = RandomForestClassifier(max_depth=8, max_features=2, random_state=40).fit(X_train, y_train)
y_pred = model.predict(X_test)
print(f"Accuracy = {accuracy_score(y_test, y_pred)*100}%")
# # Simple example with intervals
points_to_explain = pd.DataFrame({'x':[1.0, 2.0], 'y':[1.0, 2.0]})
explainer = ImpreciseShap(model=model.predict_proba, masker=X_train, eps=0.15)
result_dataframe = explainer.calculate_shapley_values(points_to_explain)
result_dataframe
# # Example with different epsilon values
from impreciseshap.visualization import get_df_for_eps
eps_arr = [1e-3, 1e-2, 5e-2, 0.1, 0.15]
example_with_eps = get_df_for_eps(model, X_train, points_to_explain, eps_arr)
display(example_with_eps)
| example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import os
pd.set_option('display.max_rows', None)
pd.set_option('display.max_columns', None)
test = pd.read_csv('/Users/omarramos/Documents/Data_Mural_Project/Data/UCUES-Camp-Cli-2/Camp_Cli_(2)_data_Arts.csv',
encoding='utf-16le', sep = '\t'
)
# +
responses1 = {-1 : 'Not applicable',
1 : 'Strongly disagree',
2 : 'Disagree',
3 : 'Somewhat disagree',
4 : 'Somewhat agree',
5 : 'Agree',
6 : 'Strongly'
}
def relabel(dataframe, responses, column_label):
'''
Relabels Pivot Field Values for a given dataframe, column label, and labeled response strings.
'''
print(dataframe[column_label])
for i in dataframe[column_label]:
try:
responses[i]
except KeyError:
print('KeyError occured for responses[i], data unchanged')
return dataframe
else:
dataframe[column_label] = dataframe[column_label].replace(i, responses[i])
return dataframe
relabel(test, responses1 ,'Pivot Field Values')
# -
test
| Data/relabel/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Example 1 (簡單線性回歸)
# 先從簡單的線性回歸舉例, , 稱為斜率, 稱為截距。
# +
# imports
import numpy as np
import matplotlib.pyplot as plt
# 亂數產生資料
np.random.seed(0)
noise = np.random.rand(100, 1)
x = np.random.rand(100, 1)
y = 3 * x + 15 + noise
# y=ax+b Target function a=3, b=15
# plot
plt.scatter(x,y,s=10)
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# -
# ## KNN Regression
# KNN 不僅能夠作為分類器,也可以做回歸連續性的數值預測。其預測值為k個最近鄰居的值的平均值。
#
# Parameters:
# - n_neighbors: 設定鄰居的數量(k),選取最近的k個點,預設為5。
# - algorithm: 搜尋數演算法{'auto','ball_tree','kd_tree','brute'},可選。
# - metric: 計算距離的方式,預設為歐幾里得距離。
#
# Attributes:
# - classes_: 取得類別陣列。
# - effective_metric_: 取得計算距離的公式。
#
# Methods:
# - fit: 放入X、y進行模型擬合。
# - predict: 預測並回傳預測類別。
# - score: 預測成功的比例。
#
# +
from sklearn.neighbors import KNeighborsRegressor
# 建立 KNN 模型
knnModel = KNeighborsRegressor(n_neighbors=3)
# 使用訓練資料訓練模型
knnModel.fit(x,y)
# 使用訓練資料預測
predicted= knnModel.predict(x)
# -
# ### 模型評估
# scikit-learn KNN迴歸模型的score函式是R2 score,可作為模型評估依據,其數值越接近於1代表模型越佳。
# 除了R2 score還有其他許多回歸模型的評估方法,例如: MSE、MAE、RMSE。
from sklearn import metrics
print('R2 score: ', knnModel.score(x, y))
mse = metrics.mean_squared_error(y, predicted)
print('MSE score: ', mse)
# plot
plt.scatter(x, y, s=10, label='True')
plt.scatter(x, predicted, color="r",s=10, label='Predicted')
plt.xlabel('x')
plt.ylabel('y')
plt.legend()
plt.show()
# ## Example 2 (非線性回歸)
# 上面的例子資料都很均勻分布在一條直線上,但現實的資料可能並只會發生在一條線上。下面示範一個多項次的回歸模型例子。
x = np.array([[1.40280301e-01],[9.03857692e-01],[5.35815131e-01],[3.58391981e-01],[2.43418162e-02],[2.43342904e-02],[3.37370600e-03],[7.50261116e-01],[3.61339257e-01],[5.01366775e-01],[4.23721405e-04],[9.40725121e-01],[6.92960750e-01],[4.50878979e-02],[3.30603187e-02],[3.36372142e-02],[9.25633424e-02],[2.75369313e-01],[1.86576499e-01],[8.48144121e-02],[3.74363965e-01],[1.94585372e-02],[8.53484957e-02],[1.34221000e-01],[2.07999831e-01],[6.16501290e-01],[3.98696193e-02],[2.64437058e-01],[3.50955021e-01],[2.15764084e-03],[3.69110747e-01],[2.90784768e-02],[4.23170975e-03],[9.00383763e-01],[9.32445223e-01],[6.53506272e-01],[9.27895484e-02],[9.53984185e-03],[4.68174835e-01],[1.93734218e-01]])
y = np.array([ 5.82469676e+00, 7.94613194e+00, 9.24976070e+00, 6.59761731e+00,
2.16651685e+00, -2.50365745e-03, -1.00182588e+00, 9.02075194e+00,
8.57086436e+00, 8.50848958e+00, -7.34549241e-02, 8.73802779e+00,
7.26038154e+00, 2.38778217e+00, 2.02397265e+00, 3.57417666e+00,
5.15052189e+00, 5.57291682e+00, 6.83461431e+00, 4.20408429e+00,
7.21499207e+00, 2.24057093e+00, 5.63575746e+00, 6.66180813e+00,
5.91402744e+00, 8.29511673e+00, 3.18174801e+00, 8.23158707e+00,
7.30330971e+00, 2.55480191e-02, 6.76197223e+00, 1.05656839e+00,
1.21851645e+00, 1.03566236e+01, 8.95941549e+00, 9.67640393e+00,
5.17463285e+00, 2.25781800e-01, 8.60137397e+00, 8.13359834e+00])
#測試資料集
x_test = np.linspace(-0.1,1.1,500)[:,None]
plt.scatter(x.ravel(),y,color='black')
#測試1,3,7的degree
for k in [1,3,9]:
y_test=knnModel = KNeighborsRegressor(n_neighbors=k).fit(x,y).predict(x_test)
plt.plot(x_test.ravel(),y_test,label='n_neighbors={}'.format(k))
plt.xlim(-0.1,1.0)
plt.ylim(-2,12)
plt.legend(loc='lower right')
| _posts/ithome/2021/10.KNN/10.2.KNN(Regression).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="copyright"
# Copyright 2020 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR asdfCONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="title"
# # Custom training and online prediction
#
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/GoogleCloudPlatform/ai-platform-samples/blob/master/ai-platform-unified/notebooks/official/custom-training-online-prediction.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
# </a>
# </td>
# <td>
# <a href="https://github.com/GoogleCloudPlatform/ai-platform-samples/blob/master/ai-platform-unified/notebooks/official/custom-training-online-prediction.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
# View on GitHub
# </a>
# </td>
# </table>
# <br/><br/><br/>
# + [markdown] id="overview:custom"
# ## Overview
#
#
# This tutorial demonstrates how to use the Vertex SDK for Python to train and deploy a custom image classification model for online prediction.
# + [markdown] id="dataset:custom,cifar10,icn"
# ### Dataset
#
# The dataset used for this tutorial is the [cifar10 dataset](https://www.tensorflow.org/datasets/catalog/cifar10) from [TensorFlow Datasets](https://www.tensorflow.org/datasets/catalog/overview). The version of the dataset you will use is built into TensorFlow. The trained model predicts which type of class an image is from ten classes: airplane, automobile, bird, cat, deer, dog, frog, horse, ship, truck.
# + [markdown] id="objective:custom,training,online_prediction"
# ### Objective
#
# In this notebook, you create a custom-trained model from a Python script in a Docker container using the Vertex SDK for Python, and then do a prediction on the deployed model by sending data. Alternatively, you can create custom-trained models using `gcloud` command-line tool, or online using the Cloud Console.
#
# The steps performed include:
#
# - Create a Vertex AI custom job for training a model.
# - Train a TensorFlow model.
# - Deploy the `Model` resource to a serving `Endpoint` resource.
# - Make a prediction.
# - Undeploy the `Model` resource.
# + [markdown] id="costs"
# ### Costs
#
# This tutorial uses billable components of Google Cloud (GCP):
#
# * Vertex AI
# * Cloud Storage
#
# Learn about [Vertex AI
# pricing](https://cloud.google.com/vertex-ai/pricing) and [Cloud Storage
# pricing](https://cloud.google.com/storage/pricing), and use the [Pricing
# Calculator](https://cloud.google.com/products/calculator/)
# to generate a cost estimate based on your projected usage.
# + [markdown] id="install_aip"
# ## Installation
#
# Install the latest (preview) version of Vertex SDK for Python.
# + id="YsxCgt1zlugo"
# ! pip3 install -U git+https://github.com/googleapis/python-aiplatform.git@mb-release
# + [markdown] id="install_storage"
# Install the latest GA version of *google-cloud-storage* library as well.
# + id="qssss-KSlugo"
# ! pip3 install -U google-cloud-storage
# + [markdown] id="install_pillow"
# Install the *pillow* library for loading images.
# + id="vhP4dtWUlugp"
# ! pip3 install -U pillow
# + [markdown] id="install_numpy"
# Install the *numpy* library for manipulation of image data.
# + id="80-_pO4olugp"
# ! pip3 install -U numpy
# + [markdown] id="restart"
# ### Restart the kernel
#
# Once you've installed everything, you need to restart the notebook kernel so it can find the packages.
# + id="bzPxhxS5lugp"
import os
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
# + [markdown] id="before_you_begin"
# ## Before you begin
#
# ### Select a GPU runtime
#
# **Make sure you're running this notebook in a GPU runtime if you have that option. In Colab, select "Runtime --> Change runtime type > GPU"**
#
# ### Set up your Google Cloud project
#
# **The following steps are required, regardless of your notebook environment.**
#
# 1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
#
# 2. [Make sure that billing is enabled for your project](https://cloud.google.com/billing/docs/how-to/modify-project).
#
# 3. [Enable the Vertex AI API and Compute Engine API](https://console.cloud.google.com/flows/enableapi?apiid=aiplatform.googleapis.com,compute_component).
#
# 4. If you are running this notebook locally, you will need to install the [Cloud SDK](https://cloud.google.com/sdk).
#
# 5. Enter your project ID in the cell below. Then run the cell to make sure the
# Cloud SDK uses the right project for all the commands in this notebook.
#
# **Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
# + [markdown] id="project_id"
# #### Set your project ID
#
# **If you don't know your project ID**, you may be able to get your project ID using `gcloud`.
# + id="autoset_project_id"
import os
PROJECT_ID = ""
if not os.getenv("IS_TESTING"):
# Get your Google Cloud project ID from gcloud
# shell_output=!gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID: ", PROJECT_ID)
# + [markdown] id="set_project_id"
# Otherwise, set your project ID here.
# + id="USd_pUT0lugr"
if PROJECT_ID == "" or PROJECT_ID is None:
PROJECT_ID = "[your-project-id]" # @param {type:"string"}
# + [markdown] id="timestamp"
# #### Timestamp
#
# If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append it onto the name of resources you create in this tutorial.
# + id="c-pX32xalugs"
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
# + [markdown] id="gcp_authenticate"
# ### Authenticate your Google Cloud account
#
# **If you are using Google Cloud Notebooks**, your environment is already
# authenticated. Skip this step.
#
# **If you are using Colab**, run the cell below and follow the instructions
# when prompted to authenticate your account via oAuth.
#
# **Otherwise**, follow these steps:
#
# 1. In the Cloud Console, go to the [**Create service account key**
# page](https://console.cloud.google.com/apis/credentials/serviceaccountkey).
#
# 2. Click **Create service account**.
#
# 3. In the **Service account name** field, enter a name, and
# click **Create**.
#
# 4. In the **Grant this service account access to project** section, click the **Role** drop-down list. Type "Vertex AI"
# into the filter box, and select
# **Vertex AI Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
#
# 5. Click *Create*. A JSON file that contains your key downloads to your
# local environment.
#
# 6. Enter the path to your service account key as the
# `GOOGLE_APPLICATION_CREDENTIALS` variable in the cell below and run the cell.
# + id="vF60K5v1lugs"
import os
import sys
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# If on Google Cloud Notebooks, then don't execute this code
if not os.path.exists("/opt/deeplearning/metadata/env_version"):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
# %env GOOGLE_APPLICATION_CREDENTIALS ''
# + [markdown] id="bucket:custom"
# ### Create a Cloud Storage bucket
#
# **The following steps are required, regardless of your notebook environment.**
#
# When you submit a training job using the Cloud SDK, you upload a Python package
# containing your training code to a Cloud Storage bucket. Vertex AI runs
# the code from this package. In this tutorial, Vertex AI also saves the
# trained model that results from your job in the same bucket. Using this model artifact, you can then
# create Vertex AI model and endpoint resources in order to serve
# online predictions.
#
# Set the name of your Cloud Storage bucket below. It must be unique across all
# Cloud Storage buckets.
#
# You may also change the `REGION` variable, which is used for operations
# throughout the rest of this notebook. Make sure to [choose a region where Vertex AI services are
# available](https://cloud.google.com/vertex-ai/docs/general/locations#available_regions). You may
# not use a Multi-Regional Storage bucket for training with Vertex AI.
# + id="bucket"
BUCKET_NAME = "gs://[your-bucket-name]" # @param {type:"string"}
REGION = "[your-region]" # @param {type:"string"}
# + id="autoset_bucket"
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP
# + [markdown] id="create_bucket"
# **Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
# + id="Oz8J0vmSlugt"
# ! gsutil mb -l $REGION $BUCKET_NAME
# + [markdown] id="validate_bucket"
# Finally, validate access to your Cloud Storage bucket by examining its contents:
# + id="oadE10x2lugu"
# ! gsutil ls -al $BUCKET_NAME
# + [markdown] id="setup_vars"
# ### Set up variables
#
# Next, set up some variables used throughout the tutorial.
# + [markdown] id="import_aip"
# #### Import Vertex SDK for Python
#
# Import the Vertex SDK for Python into your Python environment and initialize it.
# + id="cNEiwLd0lugu"
import os
import sys
from google.cloud import aiplatform
from google.cloud.aiplatform import gapic as aip
aiplatform.init(project=PROJECT_ID, location=REGION, staging_bucket=BUCKET_NAME)
# + [markdown] id="accelerators:training,prediction"
# #### Set hardware accelerators
#
# You can set hardware accelerators for both training and prediction.
#
# Set the variables `TRAIN_GPU/TRAIN_NGPU` and `DEPLOY_GPU/DEPLOY_NGPU` to use a container image supporting a GPU and the number of GPUs allocated to the virtual machine (VM) instance. For example, to use a GPU container image with 4 Nvidia Tesla K80 GPUs allocated to each VM, you would specify:
#
# (aip.AcceleratorType.NVIDIA_TESLA_K80, 4)
#
# See the [locations where accelerators are available](https://cloud.google.com/vertex-ai/docs/general/locations#accelerators).
#
# Otherwise specify `(None, None)` to use a container image to run on a CPU.
#
# *Note*: TensorFlow releases earlier than 2.3 for GPU support fail to load the custom model in this tutorial. This issue is caused by static graph operations that are generated in the serving function. This is a known issue, which is fixed in TensorFlow 2.3. If you encounter this issue with your own custom models, use a container image for TensorFlow 2.3 or later with GPU support.
# + id="xd5PLXDTlugv"
TRAIN_GPU, TRAIN_NGPU = (aip.AcceleratorType.NVIDIA_TESLA_K80, 1)
DEPLOY_GPU, DEPLOY_NGPU = (aip.AcceleratorType.NVIDIA_TESLA_K80, 1)
# + [markdown] id="container:training,prediction"
# #### Set pre-built containers
#
# Vertex AI provides pre-built containers to run training and prediction.
#
# For the latest list, see [Pre-built containers for training](https://cloud.google.com/vertex-ai/docs/training/pre-built-containers) and [Pre-built containers for prediction](https://cloud.google.com/vertex-ai/docs/predictions/pre-built-containers)
# + id="1u1mr18jlugv"
TRAIN_VERSION = "tf-gpu.2-1"
DEPLOY_VERSION = "tf2-gpu.2-1"
TRAIN_IMAGE = "gcr.io/cloud-aiplatform/training/{}:latest".format(TRAIN_VERSION)
DEPLOY_IMAGE = "gcr.io/cloud-aiplatform/prediction/{}:latest".format(DEPLOY_VERSION)
print("Training:", TRAIN_IMAGE, TRAIN_GPU, TRAIN_NGPU)
print("Deployment:", DEPLOY_IMAGE, DEPLOY_GPU, DEPLOY_NGPU)
# + [markdown] id="machine:training,prediction"
# #### Set machine types
#
# Next, set the machine types to use for training and prediction.
#
# - Set the variables `TRAIN_COMPUTE` and `DEPLOY_COMPUTE` to configure your compute resources for training and prediction.
# - `machine type`
# - `n1-standard`: 3.75GB of memory per vCPU
# - `n1-highmem`: 6.5GB of memory per vCPU
# - `n1-highcpu`: 0.9 GB of memory per vCPU
# - `vCPUs`: number of \[2, 4, 8, 16, 32, 64, 96 \]
#
# *Note: The following is not supported for training:*
#
# - `standard`: 2 vCPUs
# - `highcpu`: 2, 4 and 8 vCPUs
#
# *Note: You may also use n2 and e2 machine types for training and deployment, but they do not support GPUs*.
# + id="YAXwbqKKlugv"
MACHINE_TYPE = "n1-standard"
VCPU = "4"
TRAIN_COMPUTE = MACHINE_TYPE + "-" + VCPU
print("Train machine type", TRAIN_COMPUTE)
MACHINE_TYPE = "n1-standard"
VCPU = "4"
DEPLOY_COMPUTE = MACHINE_TYPE + "-" + VCPU
print("Deploy machine type", DEPLOY_COMPUTE)
# + [markdown] id="tutorial_start:custom"
# # Tutorial
#
# Now you are ready to start creating your own custom-trained model with CIFAR10.
# + [markdown] id="train_custom_model"
# ## Train a model
#
# There are two ways you can train a custom model using a container image:
#
# - **Use a Google Cloud prebuilt container**. If you use a prebuilt container, you will additionally specify a Python package to install into the container image. This Python package contains your code for training a custom model.
#
# - **Use your own custom container image**. If you use your own container, the container needs to contain your code for training a custom model.
# + [markdown] id="train_custom_job_args"
# ### Define the command args for the training script
#
# Prepare the command-line arguments to pass to your training script.
# - `args`: The command line arguments to pass to the corresponding Python module. In this example, they will be:
# - `"--epochs=" + EPOCHS`: The number of epochs for training.
# - `"--steps=" + STEPS`: The number of steps (batches) per epoch.
# - `"--distribute=" + TRAIN_STRATEGY"` : The training distribution strategy to use for single or distributed training.
# - `"single"`: single device.
# - `"mirror"`: all GPU devices on a single compute instance.
# - `"multi"`: all GPU devices on all compute instances.
# + id="1npiDcUtlugw"
JOB_NAME = "custom_job_" + TIMESTAMP
MODEL_DIR = "{}/{}".format(BUCKET_NAME, JOB_NAME)
if not TRAIN_NGPU or TRAIN_NGPU < 2:
TRAIN_STRATEGY = "single"
else:
TRAIN_STRATEGY = "mirror"
EPOCHS = 20
STEPS = 100
CMDARGS = [
"--epochs=" + str(EPOCHS),
"--steps=" + str(STEPS),
"--distribute=" + TRAIN_STRATEGY,
]
# + [markdown] id="taskpy_contents"
# #### Training script
#
# In the next cell, you will write the contents of the training script, `task.py`. In summary:
#
# - Get the directory where to save the model artifacts from the environment variable `AIP_MODEL_DIR`. This variable is set by the training service.
# - Loads CIFAR10 dataset from TF Datasets (tfds).
# - Builds a model using TF.Keras model API.
# - Compiles the model (`compile()`).
# - Sets a training distribution strategy according to the argument `args.distribute`.
# - Trains the model (`fit()`) with epochs and steps according to the arguments `args.epochs` and `args.steps`
# - Saves the trained model (`save(MODEL_DIR)`) to the specified model directory.
# + id="72rUqXNFlugx"
# %%writefile task.py
# Single, Mirror and Multi-Machine Distributed Training for CIFAR-10
import tensorflow_datasets as tfds
import tensorflow as tf
from tensorflow.python.client import device_lib
import argparse
import os
import sys
tfds.disable_progress_bar()
parser = argparse.ArgumentParser()
parser.add_argument('--lr', dest='lr',
default=0.01, type=float,
help='Learning rate.')
parser.add_argument('--epochs', dest='epochs',
default=10, type=int,
help='Number of epochs.')
parser.add_argument('--steps', dest='steps',
default=200, type=int,
help='Number of steps per epoch.')
parser.add_argument('--distribute', dest='distribute', type=str, default='single',
help='distributed training strategy')
args = parser.parse_args()
print('Python Version = {}'.format(sys.version))
print('TensorFlow Version = {}'.format(tf.__version__))
print('TF_CONFIG = {}'.format(os.environ.get('TF_CONFIG', 'Not found')))
print('DEVICES', device_lib.list_local_devices())
# Single Machine, single compute device
if args.distribute == 'single':
if tf.test.is_gpu_available():
strategy = tf.distribute.OneDeviceStrategy(device="/gpu:0")
else:
strategy = tf.distribute.OneDeviceStrategy(device="/cpu:0")
# Single Machine, multiple compute device
elif args.distribute == 'mirror':
strategy = tf.distribute.MirroredStrategy()
# Multiple Machine, multiple compute device
elif args.distribute == 'multi':
strategy = tf.distribute.experimental.MultiWorkerMirroredStrategy()
# Multi-worker configuration
print('num_replicas_in_sync = {}'.format(strategy.num_replicas_in_sync))
# Preparing dataset
BUFFER_SIZE = 10000
BATCH_SIZE = 64
def make_datasets_unbatched():
# Scaling CIFAR10 data from (0, 255] to (0., 1.]
def scale(image, label):
image = tf.cast(image, tf.float32)
image /= 255.0
return image, label
datasets, info = tfds.load(name='cifar10',
with_info=True,
as_supervised=True)
return datasets['train'].map(scale).cache().shuffle(BUFFER_SIZE).repeat()
# Build the Keras model
def build_and_compile_cnn_model():
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(32, 3, activation='relu', input_shape=(32, 32, 3)),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Conv2D(32, 3, activation='relu'),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(
loss=tf.keras.losses.sparse_categorical_crossentropy,
optimizer=tf.keras.optimizers.SGD(learning_rate=args.lr),
metrics=['accuracy'])
return model
# Train the model
NUM_WORKERS = strategy.num_replicas_in_sync
# Here the batch size scales up by number of workers since
# `tf.data.Dataset.batch` expects the global batch size.
GLOBAL_BATCH_SIZE = BATCH_SIZE * NUM_WORKERS
MODEL_DIR = os.getenv("AIP_MODEL_DIR")
train_dataset = make_datasets_unbatched().batch(GLOBAL_BATCH_SIZE)
with strategy.scope():
# Creation of dataset, and model building/compiling need to be within
# `strategy.scope()`.
model = build_and_compile_cnn_model()
model.fit(x=train_dataset, epochs=args.epochs, steps_per_epoch=args.steps)
model.save(MODEL_DIR)
# + [markdown] id="train_custom_job"
# ### Train the model
#
# Define your custom training job on Vertex AI.
#
# Use the `CustomTrainingJob` class to define the job, which takes the following parameters:
#
# - `display_name`: The user-defined name of this training pipeline.
# - `script_path`: The local path to the training script.
# - `container_uri`: The URI of the training container image.
# - `requirements`: The list of Python package dependencies of the script.
# - `model_serving_container_image_uri`: The URI of a container that can serve predictions for your model — either a prebuilt container or a custom container.
#
# Use the `run` function to start training, which takes the following parameters:
#
# - `args`: The command line arguments to be passed to the Python script.
# - `replica_count`: The number of worker replicas.
# - `model_display_name`: The display name of the `Model` if the script produces a managed `Model`.
# - `machine_type`: The type of machine to use for training.
# - `accelerator_type`: The hardware accelerator type.
# - `accelerator_count`: The number of accelerators to attach to a worker replica.
#
# The `run` function creates a training pipeline that trains and creates a `Model` object. After the training pipeline completes, the `run` function returns the `Model` object.
# + id="mxIxvDdglugx"
job = aiplatform.CustomTrainingJob(
display_name=JOB_NAME,
script_path="task.py",
container_uri=TRAIN_IMAGE,
requirements=["tensorflow_datasets==1.3.0"],
model_serving_container_image_uri=DEPLOY_IMAGE,
)
MODEL_DISPLAY_NAME = "cifar10-" + TIMESTAMP
# Start the training
if TRAIN_GPU:
model = job.run(
model_display_name=MODEL_DISPLAY_NAME,
args=CMDARGS,
replica_count=1,
machine_type=TRAIN_COMPUTE,
accelerator_type=TRAIN_GPU.name,
accelerator_count=TRAIN_NGPU,
)
else:
model = job.run(
model_display_name=MODEL_DISPLAY_NAME,
args=CMDARGS,
replica_count=1,
machine_type=TRAIN_COMPUTE,
accelerator_count=0,
)
# + [markdown] id="deploy_model:dedicated"
# ### Deploy the model
#
# Before you use your model to make predictions, you need to deploy it to an `Endpoint`. You can do this by calling the `deploy` function on the `Model` resource. This will do two things:
#
# 1. Create an `Endpoint` resource for deploying the `Model` resource to.
# 2. Deploy the `Model` resource to the `Endpoint` resource.
#
#
# The function takes the following parameters:
#
# - `deployed_model_display_name`: A human readable name for the deployed model.
# - `traffic_split`: Percent of traffic at the endpoint that goes to this model, which is specified as a dictionary of one or more key/value pairs.
# - If only one model, then specify as **{ "0": 100 }**, where "0" refers to this model being uploaded and 100 means 100% of the traffic.
# - If there are existing models on the endpoint, for which the traffic will be split, then use `model_id` to specify as **{ "0": percent, model_id: percent, ... }**, where `model_id` is the model id of an existing model to the deployed endpoint. The percents must add up to 100.
# - `machine_type`: The type of machine to use for training.
# - `accelerator_type`: The hardware accelerator type.
# - `accelerator_count`: The number of accelerators to attach to a worker replica.
# - `starting_replica_count`: The number of compute instances to initially provision.
# - `max_replica_count`: The maximum number of compute instances to scale to. In this tutorial, only one instance is provisioned.
#
# ### Traffic split
#
# The `traffic_split` parameter is specified as a Python dictionary. You can deploy more than one instance of your model to an endpoint, and then set the percentage of traffic that goes to each instance.
#
# You can use a traffic split to introduce a new model gradually into production. For example, if you had one existing model in production with 100% of the traffic, you could deploy a new model to the same endpoint, direct 10% of traffic to it, and reduce the original model's traffic to 90%. This allows you to monitor the new model's performance while minimizing the distruption to the majority of users.
#
# ### Compute instance scaling
#
# You can specify a single instance (or node) to serve your online prediction requests. This tutorial uses a single node, so the variables `MIN_NODES` and `MAX_NODES` are both set to `1`.
#
# If you want to use multiple nodes to serve your online prediction requests, set `MAX_NODES` to the maximum number of nodes you want to use. Vertex AI autoscales the number of nodes used to serve your predictions, up to the maximum number you set. Refer to the [pricing page](https://cloud.google.com/vertex-ai/pricing#prediction-prices) to understand the costs of autoscaling with multiple nodes.
#
# ### Endpoint
#
# The method will block until the model is deployed and eventually return an `Endpoint` object. If this is the first time a model is deployed to the endpoint, it may take a few additional minutes to complete provisioning of resources.
# + id="WMH7GrYMlugy"
DEPLOYED_NAME = "cifar10_deployed-" + TIMESTAMP
TRAFFIC_SPLIT = {"0": 100}
MIN_NODES = 1
MAX_NODES = 1
if DEPLOY_GPU:
endpoint = model.deploy(
deployed_model_display_name=DEPLOYED_NAME,
traffic_split=TRAFFIC_SPLIT,
machine_type=DEPLOY_COMPUTE,
accelerator_type=DEPLOY_GPU.name,
accelerator_count=DEPLOY_NGPU,
min_replica_count=MIN_NODES,
max_replica_count=MAX_NODES,
)
else:
endpoint = model.deploy(
deployed_model_display_name=DEPLOYED_NAME,
traffic_split=TRAFFIC_SPLIT,
machine_type=DEPLOY_COMPUTE,
accelerator_type=DEPLOY_COMPUTE.name,
accelerator_count=0,
min_replica_count=MIN_NODES,
max_replica_count=MAX_NODES,
)
# + [markdown] id="make_prediction"
# ## Make an online prediction request
#
# Send an online prediction request to your deployed model.
# + [markdown] id="get_test_item:test"
# ### Get test data
#
# Download images from the CIFAR dataset and preprocess them.
#
# #### Download the test images
#
# Download the provided set of images from the CIFAR dataset:
# + id="E1EQBPGnlugz"
# Download the images
# ! gsutil -m cp -r gs://cloud-samples-data/ai-platform-unified/cifar_test_images .
# + [markdown] id="prepare_test_item:test,image"
# #### Preprocess the images
# Before you can run the data through the endpoint, you need to preprocess it to match the format that your custom model defined in `task.py` expects.
#
# `x_test`:
# Normalize (rescale) the pixel data by dividing each pixel by 255. This replaces each single byte integer pixel with a 32-bit floating point number between 0 and 1.
#
# `y_test`:
# You can extract the labels from the image filenames. Each image's filename format is "image_{LABEL}_{IMAGE_NUMBER}.jpg"
# + id="cl59KGnXlugz"
import numpy as np
from PIL import Image
# Load image data
IMAGE_DIRECTORY = "cifar_test_images"
image_files = [file for file in os.listdir(IMAGE_DIRECTORY) if file.endswith(".jpg")]
# Decode JPEG images into numpy arrays
image_data = [
np.asarray(Image.open(os.path.join(IMAGE_DIRECTORY, file))) for file in image_files
]
# Scale and convert to expected format
x_test = [(image / 255.0).astype(np.float32).tolist() for image in image_data]
# Extract labels from image name
y_test = [int(file.split("_")[1]) for file in image_files]
# + [markdown] id="send_prediction_request:image"
# ### Send the prediction request
#
# Now that you have test images, you can use them to send a prediction request. Use the `Endpoint` object's `predict` function, which takes the following parameters:
#
# - `instances`: A list of image instances. According to your custom model, each image instance should be a 3-dimensional matrix of floats. This was prepared in the previous step.
#
# The `predict` function returns a list, where each element in the list corresponds to the corresponding image in the request. You will see in the output for each prediction:
#
# - Confidence level for the prediction (`predictions`), between 0 and 1, for each of the ten classes.
#
# You can then run a quick evaluation on the prediction results:
# 1. `np.argmax`: Convert each list of confidence levels to a label
# 2. Compare the predicted labels to the actual labels
# 3. Calculate `accuracy` as `correct/total`
# + id="UywuX7fRlugz"
predictions = endpoint.predict(instances=x_test)
y_predicted = np.argmax(predictions.predictions, axis=1)
correct = sum(y_predicted == np.array(y_test))
accuracy = len(y_predicted)
print(
f"Correct predictions = {correct}, Total predictions = {accuracy}, Accuracy = {correct/accuracy}"
)
# + [markdown] id="undeploy_model"
# ## Undeploy the model
#
# To undeploy your `Model` resource from the serving `Endpoint` resource, use the endpoint's `undeploy` method with the following parameter:
#
# - `deployed_model_id`: The model deployment identifier returned by the endpoint service when the `Model` resource was deployed. You can retrieve the deployed models using the endpoint's `deployed_models` property.
#
# Since this is the only deployed model on the `Endpoint` resource, you can omit `traffic_split`.
# + id="khPSAO1tlug0"
deployed_model_id = endpoint.list_models()[0].id
endpoint.undeploy(deployed_model_id=deployed_model_id)
# + [markdown] id="cleanup:custom"
# # Cleaning up
#
# To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
#
# Otherwise, you can delete the individual resources you created in this tutorial:
#
# - Training Job
# - Model
# - Endpoint
# - Cloud Storage Bucket
# + id="NNmebHf7lug0"
delete_training_job = True
delete_model = True
delete_endpoint = True
# Warning: Setting this to true will delete everything in your bucket
delete_bucket = False
# Delete the training job
job.delete()
# Delete the model
model.delete()
# Delete the endpoint
endpoint.delete()
if delete_bucket and "BUCKET_NAME" in globals():
# ! gsutil -m rm -r $BUCKET_NAME
| ai-platform-unified/notebooks/official/custom/sdk-custom-image-classification-online.ipynb |
% ---
% jupyter:
% jupytext:
% text_representation:
% extension: .m
% format_name: light
% format_version: '1.5'
% jupytext_version: 1.14.4
% kernelspec:
% display_name: Matlab
% language: matlab
% name: matlab
% ---
addpath('../');
cat = rgb2gray(imread('../pictures/cat.jpg'));
dog = rgb2gray(imread('../pictures/dog.jpg'));
figure, imshow(cat);
figure, imshow(dog);
dog_low_freq = imboxfilt(dog, 23);
imshow(dog_low_freq);
cat_high_freq = histeq(cat - imboxfilt(cat, 23));
imshow(cat_high_freq);
imshow(cat_high_freq ./ 10 + dog_low_freq ./ 1.1);
| Linear Filters and Convolution/demo/hybrid_images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/MohamadTaghizadeh/11th-international-conference-on-computer-and-knowledge-engineering-workshop/blob/main/Transfer_Learning_using_TensorFlow_with_the_cats_and_dogs_dataset.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="PxRXRCXkj69W"
# #Transfer Learning using TensorFlow with the cats and dogs dataset
# + [markdown] id="Zaofm5BRkAjU"
# import packages and download data
# + id="svB8BOwZj-MN"
import os
from skimage import color
import zipfile
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
import seaborn as sns
from tensorflow.keras.preprocessing.image import ImageDataGenerator
sns.set()
# + colab={"base_uri": "https://localhost:8080/"} id="uqXw8tall2sP" outputId="e5a00fcc-4edf-482f-c54a-428e4d432ded"
# !pip list
# + colab={"base_uri": "https://localhost:8080/"} id="A7QiW9S0mAVo" outputId="398d7af8-d190-412d-ecfd-6b41fb29f30b"
# !nvidia-smi
# + colab={"base_uri": "https://localhost:8080/"} id="iGh2Qrthj-Os" outputId="f829cc0b-6a18-4e63-c9fa-030a61aa2341"
# !wget --no-check-certificate \
# https://storage.googleapis.com/mledu-datasets/cats_and_dogs_filtered.zip \
# -O ./cats_and_dogs_filtered.zip
# + id="rTuhC5L-kdjt"
dataset_path = "./cats_and_dogs_filtered.zip"
# + id="KuBjEx6QkfQL"
zip_object = zipfile.ZipFile(file=dataset_path, mode="r")
# + id="Sa1sypg4kfUt"
zip_object.extractall()
# + id="Om1uGWCQkkbt"
zip_object.close()
# + id="qG6t5CS2kkd6"
dataset_path_new = "./cats_and_dogs_filtered/"
# + colab={"base_uri": "https://localhost:8080/", "height": 665} id="dK8JpOrCkkgq" outputId="6a5e51e8-eb83-442c-f453-a438bb7e0db4"
img_dir = dataset_path_new + '/train/'
cat = 'cats/cat.0.jpg'
dog = 'dogs/dog.0.jpg'
images = [cat, dog]
plt.figure(figsize=(20,10))
for i in range(2):
plt.subplot(1, 2, i + 1)
img = plt.imread(os.path.join(img_dir, images[i]))
plt.imshow(img)
plt.axis('off')
plt.colorbar()
print(img.shape)
print(f"The dimensions of the image are {img.shape[0]} pixels width and {img.shape[1]} pixels height, three single color channel")
print(f"The maximum pixel value is {img.max()} and the minimum is {img.min():}")
print(f"The mean value of the pixels is {img.mean():.4f} and the standard deviation is {img.std():.4f}")
print()
# + colab={"base_uri": "https://localhost:8080/", "height": 355} id="NOm1aTMektGt" outputId="a03e089d-b386-40c7-ea72-b09da3a15867"
for i in range(2):
rgb_img = plt.imread(os.path.join(img_dir, images[i]))
grayscale_img = color.rgb2gray(rgb_img)
sns.distplot(grayscale_img.ravel(), kde=False)
plt.title('Distribution of Pixel Intensities in the Image')
plt.xlabel('Pixel Intensity')
plt.ylabel('Number Pixels in Image')
# + [markdown] id="nQbCOHltkxuU"
# Data preparation
# + id="25ECeT5TkvgD"
train_dir = os.path.join(dataset_path_new, "train")
validation_dir = os.path.join(dataset_path_new, "validation")
# + id="PwJsE2Jak2cx"
data_gen_train = ImageDataGenerator(rescale=1/255.)
data_gen_valid = ImageDataGenerator(rescale=1/255.)
# + colab={"base_uri": "https://localhost:8080/"} id="Tz8-d2C3k5JU" outputId="71b383f4-c684-4cc3-a58f-9c1c9c73ec97"
train_generator = data_gen_train.flow_from_directory(train_dir, target_size=(128,128),
batch_size=128, class_mode="binary")
# + colab={"base_uri": "https://localhost:8080/"} id="GOrTmXXdk7NG" outputId="b5cd18dc-bffb-4e6e-bd2d-f5359879562a"
valid_generator = data_gen_valid.flow_from_directory(validation_dir, target_size=(128,128),
batch_size=128, class_mode="binary")
# + [markdown] id="oHVidL9_k8_z"
# Model
# + id="ZzzBvd_7lAaw"
IMG_SHAPE = (128, 128, 3)
# + colab={"base_uri": "https://localhost:8080/"} id="k6kXfAE6lAfN" outputId="a28eaf91-41b5-495e-d596-a164c991c383"
base_model = tf.keras.applications.MobileNetV2(input_shape=IMG_SHAPE, include_top=False,
weights="imagenet")
# + colab={"base_uri": "https://localhost:8080/"} id="cZ-XhqV6lAlf" outputId="6b3dff8c-fde5-4270-c428-e1287cff753a"
base_model.summary()
# + id="tGOfiSCrlH8l"
base_model.trainable = False
# + colab={"base_uri": "https://localhost:8080/"} id="GYBUD_ualH_R" outputId="45aaed91-c77f-43b9-a7b7-1627427ef82b"
base_model.output
# + id="eWjld5vKlLKO"
global_average_layer = tf.keras.layers.GlobalAveragePooling2D()(base_model.output)
# + id="6PvRdvFElLMf"
prediction_layer = tf.keras.layers.Dense(units=1, activation='sigmoid')(global_average_layer)
# + id="DQy6XgUxlLO0"
model = tf.keras.models.Model(inputs=base_model.input, outputs=prediction_layer)
# + colab={"base_uri": "https://localhost:8080/"} id="Snq1sIkglQZl" outputId="7e83b09b-5d34-4ba5-de8e-3c47de5108ec"
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="xSgZ7VmZlUJn" outputId="32c4f3a2-dea1-4a9e-b054-dfacd22f149a"
model.compile(optimizer=tf.keras.optimizers.RMSprop(lr=0.0001), loss="binary_crossentropy",
metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="Gt-EMcqMlUwO" outputId="6d16917d-99a6-4fc7-f2d5-e231a35e471b"
history = model.fit_generator(train_generator, epochs=100, validation_data=valid_generator)
# + colab={"base_uri": "https://localhost:8080/", "height": 301} id="NbBPj1GxlWJV" outputId="b5cfbb95-c0a5-4275-cc91-cfe19c00d69c"
plt.plot(history.history['loss'], label='train loss')
plt.plot(history.history['val_loss'], label='val loss')
plt.legend()
plt.xlabel("Epoch")
plt.ylabel("Loss")
plt.title('Loss Curve')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 301} id="XD9fitemlYRO" outputId="912d1077-01b5-46cb-b10f-6e3f8fd55bf4"
plt.plot(history.history['accuracy'], label='train acc')
plt.plot(history.history['val_accuracy'], label='val acc')
plt.legend()
plt.xlabel("Epoch")
plt.ylabel("Accuracy")
plt.title('Accuracy Curve')
plt.show()
# + [markdown] id="qJXVESfElcbW"
# #Model Fine-tuning
# + id="rS5kQqrQlZ7V"
base_model.trainable = True
# + colab={"base_uri": "https://localhost:8080/"} id="fv4vJI51lk2u" outputId="16944cfe-487a-414c-bcf8-da923ff00699"
len(base_model.layers)
# + id="08eCdcIKlk5P"
fine_tune_at = 120
# + id="SWyAaO9rlk7K"
for layer in base_model.layers[:fine_tune_at]:
layer.trainable = False
# + colab={"base_uri": "https://localhost:8080/"} id="BRSakGDxlk9I" outputId="d0ea8a70-00f8-4840-dd7b-8b8dc5882b48"
model.compile(optimizer=tf.keras.optimizers.RMSprop(lr=0.0001), loss="binary_crossentropy", metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/"} id="9Ix1CYkplk_Y" outputId="c3879e72-c2ae-4943-894f-8557ec0493cc"
history = model.fit_generator(train_generator, epochs=3, validation_data=valid_generator)
# + colab={"base_uri": "https://localhost:8080/", "height": 301} id="BQUBm0W0llBe" outputId="bfbcf474-93c7-4ef3-dd1c-37832695527e"
plt.plot(history.history['loss'], label='train loss')
plt.plot(history.history['val_loss'], label='val loss')
plt.legend()
plt.xlabel("Epoch")
plt.ylabel("Loss")
plt.title('Loss Curve')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 301} id="G99fGLlOlvh0" outputId="f925325a-1d39-419d-f525-7427f1aac5b3"
plt.plot(history.history['accuracy'], label='train acc')
plt.plot(history.history['val_accuracy'], label='val acc')
plt.legend()
plt.xlabel("Epoch")
plt.ylabel("Accuracy")
plt.title('Accuracy Curve')
plt.show()
| Transfer_Learning_using_TensorFlow_with_the_cats_and_dogs_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 8.3
# language: ''
# name: sagemath
# ---
# # CÁLCULO DIFERENCIAL E INTEGRAL
# # FUNCIONES Y SUS LÍMITES
# ## CÓMPUTO DE LÍMITES
#
# El cálculo preciso de límites requiere reglas precisas, que a continuación enunciamos e ilustramos:
#
# * El límite de una suma es la suma de los límites
# * El límite de una resta es la resta de los límites
# * El límite de un constante por una función es la constante por el límite de la función
# * El límite del producto es el producto de los límites
# * El límite de un cociente es el cociente de los límite **siempre y cuando el límite del denominador sea diferente de cero**
# Cómo se observa, calcular límites con operaciones básicas es sencillo, **excepto cuando tenemos divisiones entre cero**. Aquí tenemos que distinguir dos casos fundamentales:
# ### Singularidades removibles
# En el primero, podemos remover la división entre cero con algún procedimiento álgebraico o geométrico. Consideremos un par de ejemplos.
# #### Ejemplo
# Sea $$f(x)=\dfrac{x^2-1}{x-1}$$. Cálcule el límite cuando $x\to 1$.
#definimos f
f(x) = (x^2-1)/(x-1)
show(f)
#trazamos su gráfica
grafica = plot(f, (x,0,2), gridlines=True)
show(grafica)
# Al observar la gráfica, no observamos irregularidad alguna. De hecho, parecería que la función no es más que una línea recta y, en efecto, lo es...
show(f.full_simplify())
# ... excepto en un punto:
try:
f(0)
print("Cálculo exitoso")
except ValueError:
print("División entre cero")
# Aún así, es posible calcular el límite que esperaríamos de la simplificación, a saber
#
# $$ \lim_{x\to 1} f(x) = \lim_{x\to 1} x+1 = 2$$
L = limit(f(x), x=1)
print(L)
# En el ejemplo anterior, bastaba hacer una simplificación algebraica. Aunque esto siempre es posible, se pueden obtener límites de manera geométrica, para lo cuál siempre es bueno explorar la gráfica.
# #### Ejercicio
# Definamos la función
#
# $$g(x) = \dfrac{\sin(x)}{x}$$
#
# * Trace su gráfica alrededor de $x=0$
# * Intente cálcular el valor en $x=0$
# * Intente simplificar la expresión
# * De la gráfica, intuya cual es su límite cuando $x \to 0$
# * Cálcule dicho límite
# ### Singularidades irremovibles
#
# Existen otro tipo de singularidades al intentar dividir entre cero, que no es posible evitar.
#
# El ejemplo típico de esto sería
# $$ f(x) = \dfrac{1}{x} $$ cuando $x=0$.
### definimos f y graficamos
f(x) = 1/x
grafica = plot(f, (x,-1,1), thickness = 2, color = "red")
show(grafica, ymin=-1000, ymax=1000)
# Incluso podemos tratar de calcular el límite, pero no obtendremos algún número real
print(limit(f, x=0))
# **Observación**
#
# En el ejemplo anterior, $f(x)\to \infty$ si $x\to 0$ por la derecha. ¿Cuál es el límite por la izquierda?
# #### Ejercicio
#
# Determine las sigularidade de
# $$ g(x)= \dfrac{x^{2} + x - 6}{x^{2} + 6 \, x + 9} $$
# y su tipo.
#
# *Sugerencia: Utilice el método `factor` para simplificar la expresión.*
p(x) = x^2+x-6
factor(p)
q(x)= x^2+6*x+9
factor(q)
g(x) = p(x)/q(x)
factor(g(x))
grafica = plot(g, -4,-2)
show(grafica, ymin=-100, ymax=100)
# ### Límites de polinomios
#
# De las reglas anteriores, es fácil cálcular el límite de cualquier combinación lineal de límites
#
# $$
# \lim_{x\to a} c_1 f(x) + c_2 g(x) =
# c_1 \lim_{x\to a} f(x) + c_2 \lim_{x\to a} g(x)
# $$
#
# donde $c_1,c_2$ son constantes y el límite de potencias *naturales*
#
# $$
# \lim_{x\to a} \left(f(x)\right)^n =
# \left( \lim_{x\to a} f(x) \right)^n
# $$
#
# donde $n=0,1,2,...$.
# Otro par de reglas intuitivas que nos ayudarán a aterrizar nuestras ideas son las siguientes:
#
# $$ \lim_{x \to a} c = c$$
#
# $$ \lim_{x \to a} x = a$$
# Con esto, podemos concluir que si $p(x)$ es un polinomio, entonces
# $$ \lim_{x \to a} p(x) = p(a)$$.
# +
### ejemplo
p(x) = 2*x^2-3*x+1
print(p(0))
L = limit(p, x=0)
print(L)
# -
# Más aun, si $R(x)$ es una función racional de la forma
# $$ R(x) = \dfrac{p(x)}{q(x)} $$
# con, $p,q$ polinomios, entonces:
# $$ \lim_{x \to a} R(x) = R(a)$$
# **siempre y cuando $$q(a)\neq 0$$** (¿porqué?)
# +
### ejemplo
p(x) = x-2
q(x) = x-3
x0 = 1
print(q(x0))
print(p(x0))
# -
R(x) = p(x)/q(x)
print(R(x0))
L = limit(R, x=x0)
print(L)
# ¿Qué sucede cuando cambiamos $x_0=3$?
# ### Potencias reales
#
# Finalmente, tenemos que considerar funciones de la forma
# $$ f(x) = x^{\alpha} $$
# donde $\alpha$ es cualquier número real.
#
# Como casos particulares tenemos
# * $\alpha$ es una número racional $$x^{n/m} = \sqrt[m]{x^n}$$
# * $\alpha$ es un número negativo $$x^{-n} = \dfrac{1}{x^n} $$
#
# En cualquier caso, debemos tomar en cuenta sus dominio y que no es posible calcular límites en el interior de su complemento.
# ### Ejemplo
# ¿Es posible calcular
# $\lim_{x \to 0} (x-1)^{5/2}$?
f(x) = (x-1)^(5/2)
plot(f)
plot(f, (x,1,5))
# Para determinar su dominio, tendríamos que averiguar para que valores de $x$, la expresión $x-1$ es positiva:
solve(x-1>0, x)
# #### Ejercicio
#
# Para los siguientes límites:
# * Trace la gráfica de la función correspondiente
# * Determine si existe una singularidad y de que tipo
# * Cálcule el límite
# 1. $$ \lim_{h \to 0} \dfrac{(3+h)^2-9}{h}$$
# 2. $$ \lim_{t \to 0} \dfrac{\sqrt{t^2+9}-3}{t^2} $$
# 3. $$ \lim_{x \to 0} |x| $$
# 4. $$ \lim_{x \to 0} \dfrac{|x|}{x} $$
# #### Ejercicio
#
# * Investigue las [funciones de parte entera](https://es.wikipedia.org/wiki/Funciones_de_parte_entera) y sus implementaciones respectivas en `Sagemath` o `Python`.
# * Trace sus gráficas correspondientes y discuta los límites laterales para cada número entero.
# ### Teorema del sandwich
#
# Tratemos de averiguar gráficamente el valor de
#
# $$ \lim_{x\to 0} x^2\sin\dfrac{1}{x} $$
#
# el cual es claro no podemos obtener sólo por sustitución.
f(x) = x^2*sin(1/x)
intervalo = (x,-0.1,0.1)
grafica = plot(f, intervalo)
show(grafica)
# Podríamos adivinar que se acerca a cero. Pero podemos hacer esto más claro, comparando la gráfica de $f(x)$ con la de las funciones $x\mapsto \pm x^2$
grafica += plot(x^2, intervalo, color="green")
grafica += plot(-x^2, intervalo, color="red")
show(grafica)
# De hecho, se puede demostrar formalmente que
# $$ -x^2 \leq x^2\sin(1/x) \leq x^2 $$ para todo $$x\neq 0$$ y como
# $$ \lim_{x\to 0}-x^2 = \lim_{x\to 0}x^2 = 0 $$ entonces
# deducimos que
# $$ \lim_{x\to 0} x^2\sin\dfrac{1}{x} = 0 $$
#
# Este resultado se conoce como **teorema del sandwich** que se enuncia de la siguiente manera
# #### Teorema
#
# Si $$m(x) \leq f(x) \leq M(x)$$ cuando $x \neq c$ pero es cercano a $c$ y
# $$ \lim_{x\to c} m(x) = \lim_{x\to c} M(x) = L$$
# entonces
# $$ \lim_{x\to c} f(x) = L $$
# #### Ejercicio
# 1. Trace la gráfica de $m(x)=\cos(t)$ cuando $t\approx 0$
# 2. Trace la gráfica de $M(x)=1$ cuando $t\approx 0$
# 3. Trace la gráfica de $f(x)=\dfrac{\sin(t)}{t}$ cuando $t\approx 0$
# 4. Valore el límite de $f(x)$ cuando $t\to 0$ utilizando el teorema del sandwich.
# #### Ejercicio
# Evalue
# 1. $$ \lim_{t \to 0}\dfrac{\sin(7t)}{4t} $$
# 2. $$ \lim_{t \to 0}\dfrac{\cos(t)-1}{t} $$
| CALCULO 101 LECCION 4 COMPUTO DE LIMITES.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use(['seaborn-darkgrid'])
import pymc3 as pm
import numpy as np
import pandas as pd
# # Model averaging
#
# When confronted with more than one model we have several options. One of them is to perform model selection, using for example a given Information Criterion as exemplified [in this notebook](model_comparison.ipynb) and this other [example](GLM-model-selection.ipynb). Model selection is appealing for its simplicity, but we are discarding information about the uncertainty in our models. This is somehow similar to computing the full posterior and then just keep a point-estimate like the posterior mean; we may become overconfident of what we really know.
#
# One alternative is to perform model selection but discuss all the different models together with the computed values of a given Information Criterion. It is important to put all these numbers and tests in the context of our problem so that we and our audience can have a better feeling of the possible limitations and shortcomings of our methods. If you are in the academic world you can use this approach to add elements to the discussion section of a paper, presentation, thesis, and so on.
#
# Yet another approach is to perform model averaging. The idea now is to generate a meta-model (and meta-predictions) using a weighted average of the models. There are several ways to do this and PyMC3 includes 3 of them that we are going to briefly discuss, you will find a more thorough explanation in the work by [<NAME> et. al.](https://arxiv.org/abs/1704.02030)
#
# ## Pseudo Bayesian model averaging
#
# Bayesian models can be weighted by their marginal likelihood, this is known as Bayesian Model Averaging. While this is theoretically appealing, is problematic in practice: on the one hand the marginal likelihood is highly sensible to the specification of the prior, in a way that parameter estimation is not, and on the other computing the marginal likelihood is usually a challenging task. An alternative route is to use the values of WAIC (Widely Applicable Information Criterion) or LOO (pareto-smoothed importance sampling Leave-One-Out cross-validation), which we will call generically IC, to estimate weights. We can do this by using the following formula:
#
# $$w_i = \frac {e^{ - \frac{1}{2} dIC_i }} {\sum_j^M e^{ - \frac{1}{2} dIC_j }}$$
#
# Where $dIC_i$ is the difference between the i-esim information criterion value and the lowest one. Remember that the lowest the value of the IC, the better. We can use any information criterion we want to compute a set of weights, but, of course, we cannot mix them.
#
# This approach is called pseudo Bayesian model averaging, or Akaike-like weighting and is an heuristic way to compute the relative probability of each model (given a fixed set of models) from the information criteria values. Look how the denominator is just a normalization term to ensure that the weights sum up to one.
#
# ## Pseudo Bayesian model averaging with Bayesian Bootstrapping
#
# The above formula for computing weights is a very nice and simple approach, but with one major caveat it does not take into account the uncertainty in the computation of the IC. We could compute the standard error of the IC (assuming a Gaussian approximation) and modify the above formula accordingly. Or we can do something more robust, like using a [Bayesian Bootstrapping](http://www.sumsar.net/blog/2015/04/the-non-parametric-bootstrap-as-a-bayesian-model/) to estimate, and incorporate this uncertainty.
#
# ## Stacking
#
# The third approach implemented in PyMC3 is know as _stacking of predictive distributions_ and it has been recently [proposed](https://arxiv.org/abs/1704.02030). We want to combine several models in a metamodel in order to minimize the diverge between the meta-model and the _true_ generating model, when using a logarithmic scoring rule this is equivalently to:
#
# $$\max_{n} \frac{1}{n} \sum_{i=1}^{n}log\sum_{k=1}^{K} w_k p(y_i|y_{-i}, M_k)$$
#
# Where $n$ is the number of data points and $K$ the number of models. To enforce a solution we constrain $w$ to be $w_k \ge 0$ and $\sum_{k=1}^{K} w_k = 1$.
#
# The quantity $p(y_i|y_{-i}, M_k)$ is the leave-one-out predictive distribution for the $M_k$ model. Computing it requires fitting each model $n$ times, each time leaving out one data point. Fortunately we can approximate the exact leave-one-out predictive distribution using LOO (or even WAIC), and that is what we do in practice.
#
# ## Weighted posterior predictive samples
#
# Once we have computed the weights, using any of the above 3 methods, we can use them to get a weighted posterior predictive samples. PyMC3 offers functions to perform these steps in a simple way, so let see them in action using an example.
#
# The following example is taken from the superb book [Statistical Rethinking](http://xcelab.net/rm/statistical-rethinking/) by <NAME>. You will find more PyMC3 examples from this book in this [repository](https://github.com/aloctavodia/Statistical-Rethinking-with-Python-and-PyMC3). We are going to explore a simplified version of it. Check the book for the whole example and a more thorough discussion of both, the biological motivation for this problem and a theoretical/practical discussion of using Information Criteria to compare, select and average models.
#
# Briefly, our problem is as follows: We want to explore the composition of milk across several primate species, it is hypothesized that females from species of primates with larger brains produce more _nutritious_ milk (loosely speaking this is done _in order to_ support the development of such big brains). This is an important question for evolutionary biologists and try to give and answer we will use 3 variables, two predictor variables: the proportion of neocortex compare to the total mass of the brain and the logarithm of the body mass of the mothers. And for predicted variable, the kilocalories per gram of milk. With these variables we are going to build 3 different linear models:
#
# 1. A model using only the neocortex variable
# 2. A model using only the logarithm of the mass variable
# 3. A model using both variables
#
# Let start by uploading the data and centering the `neocortex` and `log mass` variables, for better sampling.
d = pd.read_csv('../data/milk.csv')
d.iloc[:,1:] = d.iloc[:,1:] - d.iloc[:,1:].mean()
d.head()
# Now that we have the data we are going to build our first model using only the `neocortex`.
with pm.Model() as model_0:
alpha = pm.Normal('alpha', mu=0, sigma=10)
beta = pm.Normal('beta', mu=0, sigma=10)
sigma = pm.HalfNormal('sigma', 10)
mu = alpha + beta * d['neocortex']
kcal = pm.Normal('kcal', mu=mu, sigma=sigma, observed=d['kcal.per.g'])
trace_0 = pm.sample(2000)
# The second model is exactly the same as the first one, except we now use the logarithm of the mass
with pm.Model() as model_1:
alpha = pm.Normal('alpha', mu=0, sigma=10)
beta = pm.Normal('beta', mu=0, sigma=1)
sigma = pm.HalfNormal('sigma', 10)
mu = alpha + beta * d['log_mass']
kcal = pm.Normal('kcal', mu=mu, sigma=sigma, observed=d['kcal.per.g'])
trace_1 = pm.sample(2000)
# And finally the third model using the `neocortex` and `log_mass` variables
with pm.Model() as model_2:
alpha = pm.Normal('alpha', mu=0, sigma=10)
beta = pm.Normal('beta', mu=0, sigma=1, shape=2)
sigma = pm.HalfNormal('sigma', 10)
mu = alpha + pm.math.dot(beta, d[['neocortex','log_mass']].T)
kcal = pm.Normal('kcal', mu=mu, sigma=sigma, observed=d['kcal.per.g'])
trace_2 = pm.sample(2000)
# Now that we have sampled the posterior for the 3 models, we are going to compare them visually. One option is to use the `forestplot` function that supports plotting more than one trace.
traces = [trace_0, trace_1, trace_2]
pm.forestplot(traces, figsize=(10, 5));
# Another option is to plot several traces in a same plot is to use `densityplot`. This plot is somehow similar to a forestplot, but we get truncated KDE plots (by default 95% credible intervals) grouped by variable names together with a point estimate (by default the mean).
pm.densityplot(traces, var_names=['alpha', 'sigma']);
# Now that we have sampled the posterior for the 3 models, we are going to use WAIC (Widely applicable information criterion) to compare the 3 models. We can do this using the `compare` function included with PyMC3.
model_dict = dict(zip([model_0, model_1, model_2], traces))
comp = pm.compare(model_dict, method='BB-pseudo-BMA')
comp
# We can see that the best model is `model_2`, the one with both predictor variables. Notice the DataFrame is ordered from lowest to highest WAIC (_i.e_ from _better_ to _worst_ model). Check [this notebook](model_comparison.ipynb) for a more detailed discussing on model comparison.
#
# We can also see that we get a column with the relative `weight` for each model (according to the first equation at the beginning of this notebook). This weights can be _vaguely_ interpreted as the probability that each model will make the correct predictions on future data. Of course this interpretation is conditional on the models used to compute the weights, if we add or remove models the weights will change. And also is dependent on the assumptions behind WAIC (or any other Information Criterion used). So try to do not overinterpret these `weights`.
#
# Now we are going to use copmuted `weights` to generate predictions based not on a single model but on the weighted set of models. This is one way to perform model averaging. Using PyMC3 we can call the `sample_posterior_predictive_w` function as follows:
ppc_w = pm.sample_posterior_predictive_w(traces, 1000, [model_0, model_1, model_2],
weights=comp.weight.sort_index(ascending=True),
progressbar=False)
# Notice that we are passing the weights ordered by their index. We are doing this because we pass `traces` and `models` ordered from model 0 to 2, but the computed weights are ordered from lowest to highest WAIC (or equivalently from larger to lowest weight). In summary, we must be sure that we are correctly pairing the weights and models.
#
# We are also going to compute PPCs for the lowest-WAIC model
ppc_2 = pm.sample_posterior_predictive(trace_2, 1000, model_2,
progressbar=False)
# A simple way to compare both kind of predictions is to plot their mean and hpd interval
# +
mean_w = ppc_w['kcal'].mean()
hpd_w = pm.hpd(ppc_w['kcal']).mean(0)
mean = ppc_2['kcal'].mean()
hpd = pm.hpd(ppc_2['kcal']).mean(0)
plt.errorbar(mean, 1, xerr=[[mean - hpd]], fmt='o', label='model 2')
plt.errorbar(mean_w, 0, xerr=[[mean_w - hpd_w]], fmt='o', label='weighted models')
plt.yticks([])
plt.ylim(-1, 2)
plt.xlabel('kcal per g')
plt.legend();
# -
# As we can see the mean value is almost the same for both predictions but the uncertainty in the weighted model is larger. We have effectively propagated the uncertainty about which model we should select to the posterior predictive samples. You can now try with the other two methods for computing weights `stacking` (the default and recommended method) and `pseudo-BMA`.
#
# **Final notes:**
#
# There are other ways to average models such as, for example, explicitly building a meta-model that includes all the models we have. We then perform parameter inference while jumping between the models. One problem with this approach is that jumping between models could hamper the proper sampling of the posterior.
#
# Besides averaging discrete models we can sometimes think of continuous versions of them. A toy example is to imagine that we have a coin and we want to estimated it's degree of bias, a number between 0 and 1 being 0.5 equal chance of head and tails. We could think of two separated models one with a prior biased towards heads and one towards tails. We could fit both separate models and then average them using, for example, IC-derived weights. An alternative, is to build a hierarchical model to estimate the prior distribution, instead of contemplating two discrete models we will be computing a continuous model that includes these the discrete ones as particular cases. Which approach is better? That depends on our concrete problem. Do we have good reasons to think about two discrete models, or is our problem better represented with a continuous bigger model?
# %load_ext watermark
# %watermark -n -u -v -iv -w
| docs/source/notebooks/model_averaging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Language Identifier Using Word Bigrams
# Based on [Language Identifier by asif31iqbal](https://github.com/asif31iqbal/language-identifier)
# ## 0. Importing libraries and creating helper tokenize method
import pickle
import string
import os
from nltk import ngrams, FreqDist, word_tokenize
from numpy import arange
import matplotlib.pyplot as plt
# %matplotlib inline
def ultimate_tokenize(sentence):
# Remove punctuation and digits
sentence = sentence.translate(str.maketrans('', '', string.punctuation + string.digits))
return word_tokenize(sentence.lower())
# ## 1. Understanding the process
# +
simple_example_text = 'Oh, then, I see Queen Mab hath been with you.'
simple_example_tokens_words = ultimate_tokenize(simple_example_text)
simple_example_tokens_words
# -
simple_example_tokens_chars = list(simple_example_tokens_words[0])
simple_example_tokens_chars
simple_example_tokens_words_unigrams = list(ngrams(simple_example_tokens_words, 1))
simple_example_tokens_words_unigrams
simple_example_tokens_words_bigrams = list(ngrams(simple_example_tokens_words, 2, pad_left=True, pad_right=True, left_pad_symbol='_', right_pad_symbol='_'))
simple_example_tokens_words_bigrams
fdist = FreqDist(simple_example_tokens_words_unigrams)
fdist
unigram_dict = dict()
for k, v in fdist.items():
unigram_dict[' '.join(k)] = v
unigram_dict
file = 'ngram_langid_files/LangId.train.English.txt'
with open(file, encoding='utf8') as f:
content = f.read().lower()
content.replace('\n', '')[:100]
with open('ngram_langid_files/English.unigram.pickle', 'rb') as handle:
unigram_english_dict = pickle.load(handle)
unigram_english_dict
with open('ngram_langid_files/English.bigram.pickle', 'rb') as handle:
bigram_english_dict = pickle.load(handle)
bigram_english_dict
bigram_english_dict.get('of the')
import operator
english_unigram_freqs = sorted(unigram_english_dict.items(), key=operator.itemgetter(1), reverse=True)
english_unigram_freqs[:10]
# +
labels, values = zip(*english_unigram_freqs[:10])
indexes = arange(len(labels))
width = 0.8 # width = 1 would give bars that overlap because they are too close.
fig = plt.figure(figsize=(10,7))
ax = fig.gca() # Get current axis
rects = ax.bar(indexes, values, width)
# Add title and axis labels
fig.suptitle('Top 10 English word unigrams', fontsize=20)
plt.xlabel('Word unigram', fontsize=14)
plt.ylabel('Frequency', fontsize=14)
# Display value of each bar on bar
for rect in rects:
height = rect.get_height()
ax.text(rect.get_x() + rect.get_width() / 2., 50 + height, '%d' % int(height), ha='center', va='bottom') # Can also add color and fontweight arguments.
# Remove the default x-axis tick numbers and use tick numbers of your own choosing:
ax.set_xticks(indexes)
# Replace the tick numbers with strings:
ax.set_xticklabels(labels)
plt.show()
# plt.savefig('top10EnglishWordUnigrams.png')
# -
# ## 1. Generating unigram and bigram frequencies for English, French and Italian from training files
# +
def get_ngram_count_dict(tokens, n):
if n == 1:
n_grams = ngrams(tokens, n)
else:
n_grams = ngrams(tokens, n, pad_left=True, pad_right=True, left_pad_symbol='_', right_pad_symbol='_') # Fun fact: If I remove padding here and later when testing, and also remove the '_' from the unigram dicts, the accuracy rises slightly. However, it's not statistically significant due to the small size of the data.
fdist = FreqDist(n_grams)
ngram_dict = dict()
for k,v in fdist.items():
ngram_dict[' '.join(k)] = v
return ngram_dict
# Calls get_ngram_count_dict to get a unigram and bigram dict from file.
def get_unigram_bigram_dicts(file):
with open(file, encoding='utf8') as f:
content = f.read()
tokens = ultimate_tokenize(content)
unigram_dict = get_ngram_count_dict(tokens, 1)
bigram_dict = get_ngram_count_dict(tokens, 2)
return (unigram_dict, bigram_dict)
# Dumps unigram and bigram dictionary of training data of given language to .pickle files.
def dump_pickle(language):
file = 'ngram_langid_files/LangId.train.' + language + '.txt'
unigram_dict, bigram_dict = get_unigram_bigram_dicts(file)
with open('ngram_langid_files/' + language + '.unigram.pickle', 'wb') as handle:
pickle.dump(unigram_dict, handle, protocol=pickle.HIGHEST_PROTOCOL) # HIGHEST_PROTOCOL instructs pickle to use the highest protocol version available.
with open('ngram_langid_files/' + language + '.bigram.pickle', 'wb') as handle:
pickle.dump(bigram_dict, handle, protocol=pickle.HIGHEST_PROTOCOL)
dump_pickle('English')
dump_pickle('French')
dump_pickle('Italian')
# -
# Later, it will also be required to know how many sentences there are in the training data for each language. This is because of the method used to calculate probabilities (incorporating the probability of the bigram among other bigrams starting with the same word) and the fact we use padding for our bigrams.
#
# In our training data each line is a sentence, which is very convenient for calculating the number of sentences.
#
# We go ahead and get the number of sentences (for more efficiency, the following code could be added to `get_unigram_bigram_dicts`):
# +
with open('ngram_langid_files/LangId.train.English.txt', encoding='utf8') as f:
for i, l in enumerate(f):
pass
number_of_sents_en = i + 1
with open('ngram_langid_files/LangId.train.French.txt', encoding='utf8') as f:
for i, l in enumerate(f):
pass
number_of_sents_fr = i + 1
with open('ngram_langid_files/LangId.train.Italian.txt', encoding='utf8') as f:
for i, l in enumerate(f):
pass
number_of_sents_it = i + 1
print('NUMBER OF SENTENCES IN TRAINING DATA')
print('English:', number_of_sents_en)
print('French:', number_of_sents_fr)
print('Italian:', number_of_sents_it)
# -
# ## 2. Identifying language for each line of the test file using bigram probabilities
# +
with open('ngram_langid_files/English.unigram.pickle', 'rb') as handle:
unigram_english_dict = pickle.load(handle)
with open('ngram_langid_files/English.bigram.pickle', 'rb') as handle:
bigram_english_dict = pickle.load(handle)
with open('ngram_langid_files/French.unigram.pickle', 'rb') as handle:
unigram_french_dict = pickle.load(handle)
with open('ngram_langid_files/French.bigram.pickle', 'rb') as handle:
bigram_french_dict = pickle.load(handle)
with open('ngram_langid_files/Italian.unigram.pickle', 'rb') as handle:
unigram_italian_dict = pickle.load(handle)
with open('ngram_langid_files/Italian.bigram.pickle', 'rb') as handle:
bigram_italian_dict = pickle.load(handle)
vocabulary_size = len(unigram_english_dict) + len(unigram_french_dict) + len(unigram_italian_dict)
vocabulary_size
# +
# Get probability of given bigram belonging to the language which bigram_dict is in
def get_bigram_probability(bigram, first_word, bigram_dict, first_word_dict): # first_word is the first word of the word bigram.
bigram_count = bigram_dict.get(bigram)
if bigram_count is None:
bigram_count = 0
first_word_count = first_word_dict.get(first_word)
if first_word_count is None:
first_word_count = 0
return (bigram_count + 1) / (first_word_count + vocabulary_size) # To get the logic of this formula, note how the proability is used in the function below. Without the + 1 in the Nr, if you find a bigram which is not in our known bigrams for a language, the probability of it being in that language would become 0. So we would like to assign a small probability of 1 / vocabulary_size in that case. Also note the arbitrariness of this 'probability'. We're saying "Given a bigram and a language, what is the probability that the bigram is of that language?" This is arbitrary because to get a meaningful probability we need to know which are the other languages considered and what their bigram frequencies are. That would be another way to do it, but arguable a worse one because it wouldn't be able to give a confidence score for a particular language. The formula just uses common sense to get to a number which works for the purposes. In the denominator, we have both first_word_count and vocabulary_size. Why? We have vocabulary_size for all langs in the denom because the larger this is, the less significant it is that for this particular language the bigram appears so many times. Could we have used a vocab_size of bigrams instead of unigrams? Sure, and the 'probabilities' would end up being much smaller numbers. What about first_word_count? This gives us a way to compare this bigram against other bigrams in this language starting with the same word. In general though, for a given bigram, it's more important to consider how many times it exists than to consider whether it is the usual bigram given a certain first word. The formula achieves that. Take the bigram 'le monseiur' and the English language. Let's say the bigram appears once and 'le' also appears once, while in French 'le monseiur' appears 100 times and le appears 100,000 times. Probability for English = (1 + 1) / (1 + 20,000) = 0.000099995. Probability for French = (100 + 1) / (100,000 + 20,000) = 0.00084166666. Note how the probability for French is still low because 100/100,000 is quite low and maybe it's not French after all if in French le is usually followed by other words. However, it's still significantly higher than the probability for English where both 'le' and 'le monseiur' only appear once.
# Get probability that a given bigram list is of a language (specified by its bigram_dict)
def get_language_probability(bigram_list, first_words, bigram_dict, first_word_dict):
result = 1.0
index = 0
for bigram in bigram_list:
result *= get_bigram_probability(bigram, first_words[index], bigram_dict, first_word_dict)
index += 1
return result
# Load correct solutions
solution_dict = dict()
with open('ngram_langid_files/LangId.sol.txt') as f:
for line in f:
(key, val) = line.split()
solution_dict[int(key)] = val
line_no = 1
result_dict = dict()
correct = 0
incorrect_line_numbers = []
# This needs to be done because I'm using padding for bigrams so the unigram dicts in their raw forms can't be used in get_bigram_probability():
unigram_english_dict['_'] = number_of_sents_en
unigram_french_dict['_'] = number_of_sents_fr
unigram_italian_dict['_'] = number_of_sents_it
with open('ngram_langid_files/LangId.test.txt', encoding='utf8') as f:
for line in f:
tokens = ultimate_tokenize(line)
bigrams = ngrams(tokens, 2, pad_left=True, pad_right=True, left_pad_symbol='_', right_pad_symbol='_')
bigram_list = [] # bigram_list will be exactly like bigrams but instead of [('_', 'this'), ...] it will be ['_ this', ...]. It is required because this is how bigrams are represented in the dictionary.
first_words = [] # The first words of each bigram. This is the similar to making a unigram_list. We use it because we don't want something in the form [(this,), ...]. Also because we want this to include '_'. We want it to include '_' because we're not using the unigrams for classification but as part of a formula to judge bigram frequency based on the starting word.
for b in bigrams:
bigram_list.append(' '.join(b))
first_words.append(b[0])
english_prob = get_language_probability(bigram_list, first_words, bigram_english_dict, unigram_english_dict)
french_prob = get_language_probability(bigram_list, first_words, bigram_french_dict, unigram_french_dict)
italian_prob = get_language_probability(bigram_list, first_words, bigram_italian_dict, unigram_italian_dict)
max_prob = max(english_prob, french_prob, italian_prob)
if max_prob == english_prob:
result_dict[line_no] = 'English'
elif max_prob == french_prob:
result_dict[line_no] = 'French'
else:
result_dict[line_no] = 'Italian'
if solution_dict[line_no] == result_dict[line_no]:
correct += 1
else:
incorrect_line_numbers.append(line_no)
line_no += 1
# Storing results from result_dict to file:
with open('ngram_langid_files/LangId.result.txt', 'w') as f:
for (key, val) in result_dict.items():
f.write(' '.join([str(key), val]) + '\n')
print('Accuracy: {:2.2f}%'.format(correct * 100 / len(solution_dict)))
# -
print('Line numbers for incorrectly classified languages: {}'.format(str(incorrect_line_numbers)))
# ## 3. Testing with our own sentence
sent = "This is a sentence."
sent_tokens = ultimate_tokenize(sent)
sent_bigrams_pre = ngrams(sent_tokens, 2, pad_left=True, pad_right=True, left_pad_symbol='_', right_pad_symbol='_')
sent_bigrams = []
sent_bigrams_first_words = []
for b in sent_bigrams_pre:
sent_bigrams.append(' '.join(b))
sent_bigrams_first_words.append(b[0])
print('Sentence bigrams:', sent_bigrams)
print('Sentence bigrams first words:', sent_bigrams_first_words)
sent_english_prob = get_language_probability(sent_bigrams, sent_bigrams_first_words, bigram_english_dict, unigram_english_dict)
sent_french_prob = get_language_probability(sent_bigrams, sent_bigrams_first_words, bigram_french_dict, unigram_french_dict)
sent_italian_prob = get_language_probability(sent_bigrams, sent_bigrams_first_words, bigram_italian_dict, unigram_italian_dict)
print("RAW 'PROBABILITIES'")
print('English:', sent_english_prob)
print('French:', sent_french_prob)
print('Italian:', sent_italian_prob)
# As we can see, these 'probabilities' are arbitrary. We can try to convert them to percentages since we are classifying only among these 3 languages:
# +
def get_normalized_probabilities(list_of_probabilities):
sum_of_probabilities = sum(list_of_probabilities)
result = []
for probability in list_of_probabilities:
result.append(probability / sum_of_probabilities)
return result
probabilities = [sent_english_prob, sent_french_prob, sent_italian_prob]
normalized_probabilities = get_normalized_probabilities(probabilities)
print('RELATIVE PROBABILITIES')
print('English: ', round(normalized_probabilities[0] * 100, 2), '%', sep='') # I use sep because I don't want a space before the % sign.
print('French: ', round(normalized_probabilities[1] * 100, 2), '%', sep='')
print('Italian: ', round(normalized_probabilities[2] * 100, 2), '%', sep='')
# -
# **PS:** For a state-of-the-art Greek dialect classifier using n-grams, take a look at [Greek Dialect Classifier](https://github.com/hb20007/greek-dialect-classifier).
| 2-3-Language-Identifier-Using-Word-Bigrams.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#default_exp medical.imaging
# -
# # Medical Imaging
#
# > Helpers for working with DICOM files
# +
#export
from fastai2.basics import *
from fastai2.vision.all import *
import pydicom,kornia,skimage
from pydicom.dataset import Dataset as DcmDataset
from pydicom.tag import BaseTag as DcmTag
from pydicom.multival import MultiValue as DcmMultiValue
from PIL import Image
try:
import cv2
cv2.setNumThreads(0)
except: pass
# -
from nbdev.showdoc import *
matplotlib.rcParams['image.cmap'] = 'bone'
#export
_all_ = ['DcmDataset', 'DcmTag', 'DcmMultiValue', 'dcmread']
#export
@patch
def dcmread(self:Path): return pydicom.dcmread(str(self))
# +
# #export
# @patch
# def png16read(self:Path): return array(Image.open(self), dtype=np.uint16)
# -
TEST_DCM = Path('images/sample.dcm')
dcm = TEST_DCM.dcmread()
#export
@patch_property
def pixels(self:DcmDataset):
"`pixel_array` as a tensor"
return tensor(self.pixel_array.astype(np.float32))
#export
@patch_property
def scaled_px(self:DcmDataset):
"`pixels` scaled by `RescaleSlope` and `RescaleIntercept"
img = self.pixels
return img*self.RescaleSlope + self.RescaleIntercept
#export
def array_freqhist_bins(self, n_bins=100):
"A numpy based function to split the range of pixel values into groups, such that each group has around the same number of pixels"
imsd = np.sort(self.flatten())
t = np.array([0.001])
t = np.append(t, np.arange(n_bins)/n_bins+(1/2/n_bins))
t = np.append(t, 0.999)
t = (len(imsd)*t+0.5).astype(np.int)
return np.unique(imsd[t])
#export
@patch
def freqhist_bins(self:Tensor, n_bins=100):
"A function to split the range of pixel values into groups, such that each group has around the same number of pixels"
imsd = self.view(-1).sort()[0]
t = torch.cat([tensor([0.001]),
torch.arange(n_bins).float()/n_bins+(1/2/n_bins),
tensor([0.999])])
t = (len(imsd)*t).long()
return imsd[t].unique()
#export
@patch
def hist_scaled_pt(self:Tensor, brks=None):
# Pytorch-only version - switch to this if/when interp_1d can be optimized
if brks is None: brks = self.freqhist_bins()
brks = brks.to(self.device)
ys = torch.linspace(0., 1., len(brks)).to(self.device)
return self.flatten().interp_1d(brks, ys).reshape(self.shape).clamp(0.,1.)
#export
@patch
def hist_scaled(self:Tensor, brks=None):
if self.device.type=='cuda': return self.hist_scaled_pt(brks)
if brks is None: brks = self.freqhist_bins()
ys = np.linspace(0., 1., len(brks))
x = self.numpy().flatten()
x = np.interp(x, brks.numpy(), ys)
return tensor(x).reshape(self.shape).clamp(0.,1.)
#export
@patch
def hist_scaled(self:DcmDataset, brks=None, min_px=None, max_px=None):
px = self.scaled_px
if min_px is not None: px[px<min_px] = min_px
if max_px is not None: px[px>max_px] = max_px
return px.hist_scaled(brks=brks)
#export
@patch
def windowed(self:Tensor, w, l):
px = self.clone()
px_min = l - w//2
px_max = l + w//2
px[px<px_min] = px_min
px[px>px_max] = px_max
return (px-px_min) / (px_max-px_min)
#export
@patch
def windowed(self:DcmDataset, w, l):
return self.scaled_px.windowed(w,l)
#export
# From https://radiopaedia.org/articles/windowing-ct
dicom_windows = types.SimpleNamespace(
brain=(80,40),
subdural=(254,100),
stroke=(8,32),
brain_bone=(2800,600),
brain_soft=(375,40),
lungs=(1500,-600),
mediastinum=(350,50),
abdomen_soft=(400,50),
liver=(150,30),
spine_soft=(250,50),
spine_bone=(1800,400)
)
#export
class TensorCTScan(TensorImageBW): _show_args = {'cmap':'bone'}
#export
class PILCTScan(PILBase): _open_args,_tensor_cls,_show_args = {},TensorCTScan,TensorCTScan._show_args
#export
@patch
@delegates(show_image)
def show(self:DcmDataset, scale=True, cmap=plt.cm.bone, min_px=-1100, max_px=None, **kwargs):
px = (self.windowed(*scale) if isinstance(scale,tuple)
else self.hist_scaled(min_px=min_px,max_px=max_px,brks=scale) if isinstance(scale,(ndarray,Tensor))
else self.hist_scaled(min_px=min_px,max_px=max_px) if scale
else self.scaled_px)
show_image(px, cmap=cmap, **kwargs)
scales = False, True, dicom_windows.brain, dicom_windows.subdural
titles = 'raw','normalized','brain windowed','subdural windowed'
for s,a,t in zip(scales, subplots(2,2,imsize=4)[1].flat, titles):
dcm.show(scale=s, ax=a, title=t)
dcm.show(cmap=plt.cm.gist_ncar, figsize=(6,6))
#export
@patch
def pct_in_window(dcm:DcmDataset, w, l):
"% of pixels in the window `(w,l)`"
px = dcm.scaled_px
return ((px > l-w//2) & (px < l+w//2)).float().mean().item()
dcm.pct_in_window(*dicom_windows.brain)
#export
def uniform_blur2d(x,s):
w = x.new_ones(1,1,1,s)/s
# Factor 2d conv into 2 1d convs
x = unsqueeze(x, dim=0, n=4-x.dim())
r = (F.conv2d(x, w, padding=s//2))
r = (F.conv2d(r, w.transpose(-1,-2), padding=s//2)).cpu()[:,0]
return r.squeeze()
ims = dcm.hist_scaled(), uniform_blur2d(dcm.hist_scaled(),50)
show_images(ims, titles=('orig', 'blurred'))
#export
def gauss_blur2d(x,s):
s2 = int(s/4)*2+1
x2 = unsqueeze(x, dim=0, n=4-x.dim())
res = kornia.filters.gaussian_blur2d(x2, (s2,s2), (s,s), 'replicate')
return res.squeeze()
#export
@patch
def mask_from_blur(x:Tensor, window, sigma=0.3, thresh=0.05, remove_max=True):
p = x.windowed(*window)
if remove_max: p[p==1] = 0
return gauss_blur2d(p, s=sigma*x.shape[-1])>thresh
#export
@patch
def mask_from_blur(x:DcmDataset, window, sigma=0.3, thresh=0.05, remove_max=True):
return to_device(x.scaled_px).mask_from_blur(window, sigma, thresh, remove_max=remove_max)
# +
mask = dcm.mask_from_blur(dicom_windows.brain)
wind = dcm.windowed(*dicom_windows.brain)
_,ax = subplots(1,1)
show_image(wind, ax=ax[0])
show_image(mask, alpha=0.5, cmap=plt.cm.Reds, ax=ax[0]);
# -
#export
def _px_bounds(x, dim):
c = x.sum(dim).nonzero().cpu()
idxs,vals = torch.unique(c[:,0],return_counts=True)
vs = torch.split_with_sizes(c[:,1],tuple(vals))
d = {k.item():v for k,v in zip(idxs,vs)}
default_u = tensor([0,x.shape[-1]-1])
b = [d.get(o,default_u) for o in range(x.shape[0])]
b = [tensor([o.min(),o.max()]) for o in b]
return torch.stack(b)
#export
def mask2bbox(mask):
no_batch = mask.dim()==2
if no_batch: mask = mask[None]
bb1 = _px_bounds(mask,-1).t()
bb2 = _px_bounds(mask,-2).t()
res = torch.stack([bb1,bb2],dim=1).to(mask.device)
return res[...,0] if no_batch else res
bbs = mask2bbox(mask)
lo,hi = bbs
show_image(wind[lo[0]:hi[0],lo[1]:hi[1]]);
#export
def _bbs2sizes(crops, init_sz, use_square=True):
bb = crops.flip(1)
szs = (bb[1]-bb[0])
if use_square: szs = szs.max(0)[0][None].repeat((2,1))
overs = (szs+bb[0])>init_sz
bb[0][overs] = init_sz-szs[overs]
lows = (bb[0]/float(init_sz))
return lows,szs/float(init_sz)
#export
def crop_resize(x, crops, new_sz):
# NB assumes square inputs. Not tested for non-square anythings!
bs = x.shape[0]
lows,szs = _bbs2sizes(crops, x.shape[-1])
if not isinstance(new_sz,(list,tuple)): new_sz = (new_sz,new_sz)
id_mat = tensor([[1.,0,0],[0,1,0]])[None].repeat((bs,1,1)).to(x.device)
with warnings.catch_warnings():
warnings.filterwarnings('ignore', category=UserWarning)
sp = F.affine_grid(id_mat, (bs,1,*new_sz))+1.
grid = sp*unsqueeze(szs.t(),1,n=2)+unsqueeze(lows.t()*2.,1,n=2)
return F.grid_sample(x.unsqueeze(1), grid-1)
px256 = crop_resize(to_device(wind[None]), bbs[...,None], 128)[0]
show_image(px256)
px256.shape
#export
@patch
def to_nchan(x:Tensor, wins, bins=None):
res = [x.windowed(*win) for win in wins]
if not isinstance(bins,int) or bins!=0: res.append(x.hist_scaled(bins).clamp(0,1))
dim = [0,1][x.dim()==3]
return TensorCTScan(torch.stack(res, dim=dim))
#export
@patch
def to_nchan(x:DcmDataset, wins, bins=None):
return x.scaled_px.to_nchan(wins, bins)
#export
@patch
def to_3chan(x:Tensor, win1, win2, bins=None):
return x.to_nchan([win1,win2],bins=bins)
#export
@patch
def to_3chan(x:DcmDataset, win1, win2, bins=None):
return x.scaled_px.to_3chan(win1, win2, bins)
show_images(dcm.to_nchan([dicom_windows.brain,dicom_windows.subdural,dicom_windows.abdomen_soft]))
#export
@patch
def save_jpg(x:(Tensor,DcmDataset), path, wins, bins=None, quality=90):
fn = Path(path).with_suffix('.jpg')
x = (x.to_nchan(wins, bins)*255).byte()
im = Image.fromarray(x.permute(1,2,0).numpy(), mode=['RGB','CMYK'][x.shape[0]==4])
im.save(fn, quality=quality)
#export
@patch
def to_uint16(x:(Tensor,DcmDataset), bins=None):
d = x.hist_scaled(bins).clamp(0,1) * 2**16
return d.numpy().astype(np.uint16)
#export
@patch
def save_tif16(x:(Tensor,DcmDataset), path, bins=None, compress=True):
fn = Path(path).with_suffix('.tif')
Image.fromarray(x.to_uint16(bins)).save(str(fn), compression='tiff_deflate' if compress else None)
_,axs=subplots(1,2)
with tempfile.TemporaryDirectory() as f:
f = Path(f)
dcm.save_jpg(f/'test.jpg', [dicom_windows.brain,dicom_windows.subdural])
show_image(Image.open(f/'test.jpg'), ax=axs[0])
dcm.save_tif16(f/'test.tif')
show_image(Image.open(str(f/'test.tif')), ax=axs[1]);
#export
@patch
def set_pixels(self:DcmDataset, px):
self.PixelData = px.tobytes()
self.Rows,self.Columns = px.shape
DcmDataset.pixel_array = property(DcmDataset.pixel_array.fget, set_pixels)
#export
@patch
def zoom(self:DcmDataset, ratio):
with warnings.catch_warnings():
warnings.simplefilter("ignore", UserWarning)
self.pixel_array = ndimage.zoom(self.pixel_array, ratio)
#export
@patch
def zoom_to(self:DcmDataset, sz):
if not isinstance(sz,(list,tuple)): sz=(sz,sz)
rows,cols = sz
self.zoom((rows/self.Rows,cols/self.Columns))
#export
@patch_property
def shape(self:DcmDataset): return self.Rows,self.Columns
dcm2 = TEST_DCM.dcmread()
dcm2.zoom_to(90)
test_eq(dcm2.shape, (90,90))
dcm2 = TEST_DCM.dcmread()
dcm2.zoom(0.25)
dcm2.show()
# +
#export
def _cast_dicom_special(x):
cls = type(x)
if not cls.__module__.startswith('pydicom'): return x
if cls.__base__ == object: return x
return cls.__base__(x)
def _split_elem(res,k,v):
if not isinstance(v,DcmMultiValue): return
res[f'Multi{k}'] = 1
for i,o in enumerate(v): res[f'{k}{"" if i==0 else i}']=o
# -
#export
@patch
def as_dict(self:DcmDataset, px_summ=True, window=dicom_windows.brain):
pxdata = (0x7fe0,0x0010)
vals = [self[o] for o in self.keys() if o != pxdata]
its = [(v.keyword,v.value) for v in vals]
res = dict(its)
res['fname'] = self.filename
for k,v in its: _split_elem(res,k,v)
if not px_summ: return res
stats = 'min','max','mean','std'
try:
pxs = self.pixel_array
for f in stats: res['img_'+f] = getattr(pxs,f)()
res['img_pct_window'] = self.pct_in_window(*window)
except Exception as e:
for f in stats: res['img_'+f] = 0
print(res,e)
for k in res: res[k] = _cast_dicom_special(res[k])
return res
#export
def _dcm2dict(fn, **kwargs): return fn.dcmread().as_dict(**kwargs)
#export
@delegates(parallel)
def _from_dicoms(cls, fns, n_workers=0, **kwargs):
return pd.DataFrame(parallel(_dcm2dict, fns, n_workers=n_workers, **kwargs))
pd.DataFrame.from_dicoms = classmethod(_from_dicoms)
# ## Export -
#hide
from nbdev.export import notebook2script
notebook2script()
| nbs/60_medical.imaging.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PostgreSQL in Python using SQLAlchemy
#
# ## The Old Way of Writing Database Code in Python
# We're going to use the library `sqlite3` to create a simple database with two tables **Person** and **Address** in the following design:
#
# 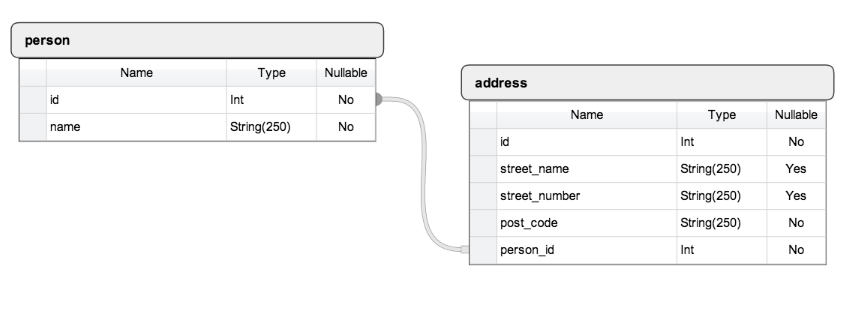
#
# To create tables and insert data into tables, type:
# +
import sqlite3
conn = sqlite3.connect('example.db')
c = conn.cursor()
c.execute('''
CREATE TABLE person
(id INTEGER PRIMARY KEY ASC, name varchar(250) NOT NULL)
''')
c.execute('''
CREATE TABLE address
(id INTEGER PRIMARY KEY ASC, street_name varchar(250), street_number varchar(250),
post_code varchar(250) NOT NULL, person_id INTEGER NOT NULL,
FOREIGN KEY(person_id) REFERENCES person(id))
''')
c.execute('''
INSERT INTO person VALUES(1, '<NAME>')
''')
c.execute('''
INSERT INTO address VALUES(1, 'Newington Road', '15', '12121', 1)
''')
conn.commit()
conn.close()
# -
# To retrieve data from databases, type:
conn = sqlite3.connect('example.db')
c = conn.cursor()
c.execute('SELECT * FROM person')
print(c.fetchall())
c.execute('SELECT * FROM address')
print(c.fetchall())
c.execute('DROP TABLE person')
c.execute('DROP TABLE address')
conn.close()
# ----
#
# ## Python's SQLAlchemy and Declarative
# There are three most important components in writing SQLAlchemy code:
#
# * A Table that represents a table in a database.
# * A mapper that maps a Python class to a table in a database.
# * A class object that defines how a database record maps to a normal Python object.
#
# Instead of having to write code for Table, mapper and the class object at different places, SQLAlchemy's declarative allows a Table, a mapper and a class object to be defined at once in one class definition.
#
# The following declarative definitions specify the same tables:
# !pip install sqlalchemy
# +
from sqlalchemy import Column, ForeignKey, Integer, String
from sqlalchemy.ext.declarative import declarative_base
from sqlalchemy.orm import relationship
from sqlalchemy import create_engine
Base = declarative_base()
class Person(Base):
__tablename__ = 'person'
# Here we define columns for the table person
# Notice that each column is also a normal Python instance attribute.
id = Column(Integer, primary_key=True)
name = Column(String(250), nullable=False)
class Address(Base):
__tablename__ = 'address'
# Here we define columns for the table address.
# Notice that each column is also a normal Python instance attribute.
id = Column(Integer, primary_key=True)
street_name = Column(String(250))
street_number = Column(String(250))
post_code = Column(String(250), nullable=False)
person_id = Column(Integer, ForeignKey('person.id'))
person = relationship(Person)
# Create an engine that stores data in the local directory's
# sqlalchemy_example.db file.
engine = create_engine('sqlite:///sqlalchemy_example.db')
# Create all tables in the engine. This is equivalent to "Create Table"
# statements in raw SQL.
Base.metadata.create_all(engine)
# -
# Add records into the database.
# +
from sqlalchemy import create_engine
from sqlalchemy.orm import sessionmaker
engine = create_engine('sqlite:///sqlalchemy_example.db')
# Bind the engine to the metadata of the Base class so that the
# declaratives can be accessed through a DBSession instance
Base.metadata.bind = engine
DBSession = sessionmaker(bind=engine)
# A DBSession() instance establishes all conversations with the database
# and represents a "staging zone" for all the objects loaded into the
# database session object. Any change made against the objects in the
# session won't be persisted into the database until you call
# session.commit(). If you're not happy about the changes, you can
# revert all of them back to the last commit by calling
# session.rollback()
session = DBSession()
# Insert a Person in the person table
new_person = Person(name='<NAME>')
session.add(new_person)
session.commit()
# Insert an Address in the address table
new_address = Address(street_name='North Street', street_number='21',
post_code='46202', person=new_person)
session.add(new_address)
session.commit()
# -
# To query, type:
# +
engine = create_engine('sqlite:///sqlalchemy_example.db')
Base.metadata.bind = engine
DBSession = sessionmaker()
DBSession.bind = engine
session = DBSession()
# Make a query to find all Persons in the database
print(session.query(Person).all())
# Return the first Person from all Persons in the database
person = session.query(Person).first()
print(person.name)
# Find all Address whose person field is pointing to the person object
print(session.query(Address).filter(Address.person == person).all())
# Retrieve one Address whose person field is point to the person object
print(session.query(Address).filter(Address.person == person).one())
address = session.query(Address).filter(Address.person == person).one()
print(address.id, address.street_name, address.street_number,
address.post_code, address.person_id)
| 01day02_database/postgresql-example.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Apache Toree - Scala
// language: scala
// name: apache_toree_scala
// ---
// + [markdown] deletable=true editable=true
// # Spark Lab Assignment
//
// ## General instructions
// The purpose of this assignment is to develop some basic Spark skills. The assignment is composed by two mandatory tasks and one challenge. Only PhD students are required to go through the challenge part. However, completing the challenges will improve the final grade for Master students too.
//
// **Warning:** all of the tasks **must** be solved using the Spark RDD API, in order to distribute the computations in the Spark workers.
//
// ## How to get help for this assignment
// [GitHub](https://github.com/) is a web-based software development collaboration tool. It is very important for your career that you get familiar with it. This is why Q&A for this assignment will be based on GitHub issues. If you encounter any problem setting up the environment, or you need some additional pointers to solve the tasks please open an issue here: https://github.com/SNICScienceCloud/LDSA-Spark/issues.
//
// ## Getting started
//
// ### Setup the environment
// Before you begin with the assignment you need to get the Docker-based Spark deployment that we have seen in the lecture. You can either install the environment on the SNIC cloud or on your local computer. For Linux and Mac users, installing the environment locally should be straightforward, while on Windows machines the installation could be slightly more challenging. If you are unsure on how to proceed, the best way is to deploy the envirnment on SNIC.
//
// **Setup the environment on the SNIC cloud (recommended)**
//
// 1. Log into https://hpc2n.cloud.snic.se
// 2. Start an instance from *TheSparkBox* image:
//
// * Type an instance name
// * Use the `scc.large` flavor
// * Select the internal IPv4 network
// * Add the `Spark` security group
// * Select your Key Pair
// * Launch the instance
//
// 3. Associate a floating IP to the newly create instance, and SSH into it:
//
// ```
// ssh core@<your-floating-ip>
// ```
//
// 4. Deploy Spark and Jupyter running:
//
// ```
// export SPARK_PUBLIC_DNS="<your-floating-ip>"
// export TSB_JUPYTER_TOKEN="<<PASSWORD>>"
// tsb up -d
// ```
//
// If everithing went well, within a couple of minutes you should be able to access the web UIs.
//
// * **Spark UI**: `http://<your-floating-ip>:8080`
// * **Jupyter**: `http://<your-floating-ip>:8888`
//
//
// **Setup the environment locally**
//
// If you like to work on you local computer, and you can install [Docker](https://www.docker.com/), here you can find detailed instruction to get the environment on your machine: https://github.com/mcapuccini/TheSparkBox. The procedure should be straightforward for Linux and Mac, however on Windows this could a little bit more challenging.
//
// ### Import the course material
// The material for the Spark module in this course is stored in this GitHub repository: https://github.com/SNICScienceCloud/LDSA-Spark. You can import it in your deployment following this steps:
//
// 1. Log into *Jupyter*
// 2. Start a new Jupyter terminal: `New > Terminal`
// 3. Clone the course material running:
//
// ```
// git clone https://github.com/SNICScienceCloud/LDSA-Spark.git
// ```
//
// If everithing went well, you should be able to open this assignment as a Jupyter notebook. Please go back to the Jupyter home page and navigate to `LDSA-Spark > Lab Assignment.ipynb`. The `LDSA-Spark` folder includes also the `Lecture Examples.ipynb` notebook that we have seen in the lecture, so you can play with it. However, keep in mind that only one notebook at the time will manage to send tasks to the Spark Master, so make sure to shutdown notebooks when you are not using them.
//
// ### How to submit the assignment
// Please complete the tasks in this assignment editing this notebook, both for the code implementations and the theory questions. If you are not familiar with Jupyter, you may want to give a look to the [Notebook Basics tutorial](http://nbviewer.jupyter.org/github/jupyter/notebook/blob/master/docs/source/examples/Notebook/Notebook%20Basics.ipynb). When you are done with the tasks, please save this notebook with your solutions, download it as `.ipynb` (`File > Download as > Notebook`), and upload it to the student portal.
//
// ## Task 1: DNA G+C precentage
// DNA is a molecule that carries genetic information used in the growth, development, functioning and reproduction of all known living organisms (including some viruses). The DNA information is coded in a language of 4 bases: cytosine (C), guanine (G), adenine (A), thymine (T). The percentage of G+C bases in a DNA sequence has valuable biological mening (wikipedia: https://goo.gl/kCLvDp), hence it is important to be able to compute it for long DNA sequences.
//
// ### Task
// Given an input DNA sequence, represented as a text file: `data/dna.txt`, compute the percentage of `g` + `c` occurrences into it. An example follows:
//
// **Input file:**
// ```
// atcg
// ccgg
// ttat
// ```
// **result:**
// $$\frac{C_{count} + G_{count}}{C_{count} + G_{count} + A_{count} + T_{count}} = \frac{6}{12} = 0.5$$
//
// **Tip 1:** when you load an input file as an RDD, each line will be loaded into a distinct string RDD record. In Scala you can count the occurrences of a certain character in a string as it follows:
// + deletable=true editable=true
"atttccgg".count(c => c == 'g')
// + [markdown] deletable=true editable=true
// **Question 1:** Is the previous operation parallel, or is computed locally? Why?
//
// *Answer goes here*
// + [markdown] deletable=true editable=true
// **Tip 2:** sums form different RDD records can be aggregated using the RDD *reduce* method. An example follows:
// + deletable=true editable=true
val sumsRDD = sc.parallelize(Array(3,5,2))
sumsRDD.reduce(_+_)
// + [markdown] deletable=true editable=true
// ### Your solution
// + deletable=true editable=true
{
// Implementation goes here
}
// + [markdown] deletable=true editable=true
// **Question 2:** What is an RDD in Spark?
//
// *Answer goes here*
//
// **Question 3:** Is *reduce* a trasformation or an action? What are RDD transformations and RDD actions? How do they differ from each other?
//
// *Answer goes here*
//
// **Question 4:** What are some of the advanteges of Spark over Hadoop (and MapReduce)?
//
// *Answer goes here*
// + [markdown] deletable=true editable=true
// ## Task 2: Monte Carlo integration
// Large dataset analysis is the main use case of Spark. However, Spark can be used to perform compute intensive tasks as well. Numerical integration is a good example problem that falls in this group of use cases.
//
// <img src="img/montecarlo_int.png" width="550"/>
// + [markdown] deletable=true editable=true
// The **Monte Carlo integration** method, is a way to get an approximation of the definite integral of a function $f(x)$, over an interval $[A,B]$. Given a value $Max_{f(x)},$ which $f(x)$ never exceeds, we first randomly draw $N$ uniformly distributed points $(x_1,y_2) … (x_N,y_N)$ s.t. $x_1 … x_N \in [A,B], y_1 … y_N \in [0,Max_{f(x)}]$. Then, assuming that $f(x)$ is positive over $[A,B]$, the fraction of points that fell under $f(x)$ will be roughly equal to the area under the curve, divided by the total area of the rectangle in which we randomly drew points. Hence, the definite integral of $f(x)$ over $[A,B]$ is roughly equal to:
//
// $$(B-A) Max_{f(x)}\frac{n_{P}}{tot_{P}},$$
// where $n_{P}$ is the number of points that fell under $f(x)$, and $tot_{P}$ is the total number of randomly drawn points.
//
// ### Task
// Write a program in Spark to approximate the definite integral of $f(x) = (1 + sin(x))$ / $cos(x)$ over $[0,1].$ Such function is positive and it is lower than $4$ over $[0,1]$.
//
// 
//
// For the purpose of this assignment drawing 1000 points is good enough.
//
// **Question** What does the Spark's parallelize function do? What is it good for?
//
// *Answer goes here*
//
// ### Your solution
// + deletable=true editable=true
{
// Implementation goes here
}
// + [markdown] deletable=true editable=true
// ## Challenge: Iris flower classification
//
// The well-known **Iris flower dataset** (wiki: https://goo.gl/OQjope) contains measurements for 150 *Iris* flowers form 3 different species: *Iris setosa*, *Iris virginica* and *Iris versicolor*. For each example in the dataset, 4 measurements were performed: *sepal length*, *sepal width*, *petal length* and *petal width*.
//
// 
//
// You are given a copy of this dataset in Comma Separated Values (CSV) format: `data/iris_csv.txt`. In the CSV text file each line contains *sepal length*, *sepal width*, *petal length*, *petal width* and *species name* separated by a comma. The file looks something like the following example:
//
// ```
// 5.1,3.5,1.4,0.2,Iris-setosa
// 4.9,3.0,1.4,0.2,Iris-setosa
// 4.7,3.2,1.3,0.2,Iris-setosa
// 7.0,3.2,4.7,1.4,Iris-versicolor
// 6.4,3.2,4.5,1.5,Iris-versicolor
// 6.9,3.1,4.9,1.5,Iris-versicolor
// 6.7,3.3,5.7,2.5,Iris-virginica
// 6.7,3.0,5.2,2.3,Iris-virginica
// 6.3,2.5,5.0,1.9,Iris-virginica
// ```
//
// **Task 1:** Build and evaluate a 3NN classifier for the **Iris flower dataset** in Spark. In order to evaluate your classifier, you can save 20% of the data for testing like we did in the lecture examples. For simplicity, you are allowed to collect the test data.
//
// **N.B.** Part of the challenge is to figure out how to parse the input dataset into an RDD. Google is your friend!
//
// ### Your solution
// + deletable=true editable=true
{
// Implementation goes here
}
| Lab Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Standard imports
# +
# Some fairly standard modules
import os, csv, lzma
import numpy as np
import matplotlib.pyplot as plt
import scipy
import datetime
# The geopandas module does not come standard with anaconda,
# so you'll need to run the anaconda prompt as an administrator
# and install it via "conda install -c conda-forge geopandas".
# That installation will include pyproj and shapely automatically.
# These are useful modules for plotting geospatial data.
import geopandas as gpd
import pyproj
import shapely.geometry
# These modules are useful for tracking where modules are
# imported from, e.g., to check we're using our local edited
# versions of open_cp scripts.
import sys
import inspect
import importlib
# In order to use our local edited versions of open_cp
# scripts, we insert the parent directory of the current
# file ("..") at the start of our sys.path here.
sys.path.insert(0, os.path.abspath(".."))
# -
# Imports from open_cp
import open_cp
import open_cp.naive as naive
# Set seed for randomization
np.random.seed(1)
# Generate random data
import datetime
size = 30
times = [datetime.datetime(2017,3,10) + datetime.timedelta(days=np.random.randint(0,10)) for _ in range(size)]
times.sort()
xc = np.random.random(size=size) * 500
yc = np.random.random(size=size) * 500
points = open_cp.TimedPoints.from_coords(times, xc, yc)
# Create naive predictor array that only uses counts of events in a region to estimate its risk. So, not doing anything interesting here.
#
# type(prediction) == GridPredictionArray
region = open_cp.RectangularRegion(0,500, 0,500)
print("a")
predictor = naive.CountingGridKernel(50, region=region)
print("b")
predictor.data = points
print("c")
prediction = predictor.predict()
print("d")
print("all done")
# Plot visualization of the basic predictor
# +
fig, ax = plt.subplots(figsize=(10,10))
m = ax.pcolor(*prediction.mesh_data(), prediction.intensity_matrix)
ax.scatter(points.xcoords, points.ycoords, marker="+", color="white")
ax.set(xlim=[0, 500], ylim=[0, 500])
cb = plt.colorbar(m, ax=ax)
cb.set_label("Relative risk")
None
# -
# Now we instead use `scipy` Gaussian KDE, so that the predictor is smoothed to use a continuous function still aiming to have the highest values centered at the cells containing the most events.
predictor = naive.ScipyKDE()
predictor.data = points
prediction = predictor.predict()
gridpred = open_cp.predictors.GridPredictionArray.from_continuous_prediction_region(prediction, region, 50)
# Plot the visualization
# +
fig, ax = plt.subplots(figsize=(10,10))
m = ax.pcolor(*gridpred.mesh_data(), gridpred.intensity_matrix)
ax.scatter(points.xcoords, points.ycoords, marker="+", color="white")
ax.set(xlim=[0, 500], ylim=[0, 500])
cb = plt.colorbar(m, ax=ax)
cb.set_label("Relative risk")
None
# -
# Challenge to myself: Can I figure out a way to repeat the above experiment, but with a 2D normal distribution for the data I generate instead?
"""
def measure(n):
m1 = np.random.normal(size=n)
m2 = np.random.normal(scale=0.5, size=n)
return m1 + m2, m1 - m2
def actual_kernel(point):
x, y = point[0], point[1]
# m2 = 0.5 * np.random.normal
# Transform matrix is: A = 1 1/2
# 1 -1/2
# So covariance matrix is AA^* = 5/4 3/4
# 3/4 5/4
a = x * (5 * x - 3 * y) / 4 + y * (-3 * x + 5 * y) / 4
return np.exp(-a/2) / 2*np.pi
m1, m2 = measure(2000)
xmin, xmax = min(m1), max(m1)
ymin, ymax = min(m2), max(m2)
data_2d = np.vstack([m1, m2])
kernel_2d = scipy.stats.gaussian_kde(data_2d)
X, Y = np.mgrid[xmin:xmax:100j, ymin:ymax:100j]
positions = np.vstack([X.ravel(), Y.ravel()])
Z = np.reshape(kernel_2d(positions).T, X.shape)
Z_actual = np.reshape(actual_kernel(positions).T, X.shape)
fig, ax = plt.subplots(ncols=3, figsize=(16,10))
for i, z in enumerate([Z, Z, Z_actual]):
ax[i].imshow(np.rot90(z), cmap=plt.cm.gist_earth_r, extent=[xmin, xmax, ymin, ymax])
ax[i].set_aspect(1)
ax[0].plot(m1, m2, 'k.', markersize=2, alpha=0.3)
ax[0].set_title("Estimated kernel and data")
ax[1].set_title("Estimated kernel")
ax[2].set_title("Actual kernel")
None
"""
# Let's try to do that same naive algorithm from the top, but using UK data instead of random data.
# +
# Obtain UK data
import open_cp.sources.ukpolice as ukpolice
print(inspect.getfile(ukpolice))
points = ukpolice.default_burglary_data()
len(points.timestamps)
# -
# Use pyproj to make a more properly projected visualization of the data
projected_points = open_cp.data.points_from_lon_lat(points, epsg=7405)
points = projected_points
bbox = points.bounding_box
fig, ax = plt.subplots(figsize=(10, 10 * bbox.aspect_ratio))
ax.scatter(points.xcoords, points.ycoords, s=10, alpha=0.2)
print(bbox)
print(type(bbox))
# +
region = open_cp.RectangularRegion(np.floor(bbox.xmin), np.ceil(bbox.xmax), np.floor(bbox.ymin), np.ceil(bbox.ymax))
predictor = naive.CountingGridKernel(2500, region=region)
predictor.data = points
prediction = predictor.predict()
# Plot the figure, 15in x 10in image
fig, ax = plt.subplots(figsize=(15,10))
m = ax.pcolor(*prediction.mesh_data(), prediction.intensity_matrix)
ax.scatter(points.xcoords, points.ycoords, marker="+", color="white")
ax.set(xlim=[bbox.xmin, bbox.xmax], ylim=[bbox.ymin, bbox.ymax])
cb = plt.colorbar(m, ax=ax)
cb.set_label("Relative risk")
None
# +
importlib.reload(open_cp.predictors)
predictor = naive.ScipyKDE()
predictor.data = points
prediction = predictor.predict()
print(type(prediction))
print(prediction)
print(region.height)
print(region.width)
print(region.grid_size)
print(prediction.samples)
gridpred = open_cp.predictors.GridPredictionArray.from_continuous_prediction_region(prediction, region, 2500)
print("Finished making gridpred!!!")
#gridpred
#newpred = prediction.rebase(cell_width=2500, cell_height=2500, xoffset=bbox.xmin, yoffset=bbox.ymin, samples=25)
# +
fig, ax = plt.subplots(figsize=(15,10))
m = ax.pcolor(*gridpred.mesh_data(), gridpred.intensity_matrix)
ax.scatter(points.xcoords, points.ycoords, marker="+", color="white")
ax.set(xlim=[region.xmin, region.xmax], ylim=[region.ymin, region.ymax])
cb = plt.colorbar(m, ax=ax)
cb.set_label("Relative risk")
None
# -
| sandbox/TryingNaive.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from rdkit import Chem
from syba.syba import SybaClassifier
import pandas as pd
import matplotlib.pyplot as plt
syba = SybaClassifier()
syba.fitDefaultScore()
data = pd.read_excel('All_generated_SMILES_SYBA_filtration.xlsx')
data.head()
whole_data_after_QED_Lipinski = pd.read_excel('All_generated_SMILES_QED_Lipinski.xlsx')
mols = [Chem.MolFromSmiles(smi) for smi in whole_data_after_QED_Lipinski['SMILES']]
SYBA_scores = [syba.predict(mol=mol) for mol in mols]
whole_data_after_QED_Lipinski['SYBA_score'] = SYBA_scores
# +
plt.hist(whole_data_after_QED_Lipinski['SYBA_score'],bins=100)
plt.xlabel('SYBA score')
plt.ylabel('Number of compounds')
#plt.title('Histogram of SYBA score distribution')
plt.axvline(x=0.0, color='r')
plt.grid(True)
plt.savefig("tmp/SYBA score after QED LIPINSKI all generated.svg")
plt.show()
print(whole_data_after_QED_Lipinski['SYBA_score'].describe())
print('Kurtosis:')
print(round(whole_data_after_QED_Lipinski['SYBA_score'].kurtosis(),4))
print('Skewness:')
print(round(whole_data_after_QED_Lipinski['SYBA_score'].skew(),4))
# +
plt.hist(whole_data_after_QED_Lipinski['QED'],bins=100)
plt.xlabel('Quantitative Estimation of drug-likeness')
plt.ylabel('Number of compounds')
plt.title('Histogram of QED distribution')
plt.axvline(x=0.5, color='r')
plt.grid(True)
plt.savefig("tmp/Quantitative Estimation of Drug-likeness after QED LIPINSKI all generated.svg")
plt.show()
print(whole_data_after_QED_Lipinski['QED'].describe())
print('Kurtosis:')
print(round(whole_data_after_QED_Lipinski['QED'].kurtosis(),4))
print('Skewness:')
print(round(whole_data_after_QED_Lipinski['QED'].skew(),4))
# +
plt.figure(figsize=(10,5))
plt.hist(whole_data_after_QED_Lipinski['SYBA_score'],bins=100)
plt.xlabel('SYBA score', fontsize=14)
plt.ylabel('Number of compounds', fontsize=14)
plt.axvline(x=0.0, color='r')
plt.grid(True)
plt.savefig("tmp/SYBA score after QED LIPINSKI.svg")
plt.show()
print(whole_data_after_QED_Lipinski['SYBA_score'].describe())
print('Kurtosis:')
print(round(whole_data_after_QED_Lipinski['SYBA_score'].kurtosis(),4))
print('Skewness:')
print(round(whole_data_after_QED_Lipinski['SYBA_score'].skew(),4))
# -
whole_data_after_QED_Lipinski['QED normalized'] =(whole_data_after_QED_Lipinski['QED']-whole_data_after_QED_Lipinski['QED'].min())/(whole_data_after_QED_Lipinski['QED'].max()-whole_data_after_QED_Lipinski['QED'].min())
whole_data_after_QED_Lipinski['SYBA_score normalized'] =(whole_data_after_QED_Lipinski['SYBA_score']-whole_data_after_QED_Lipinski['SYBA_score'].min())/(whole_data_after_QED_Lipinski['SYBA_score'].max()-whole_data_after_QED_Lipinski['SYBA_score'].min())
whole_data_after_QED_Lipinski.head()
whole_data_after_QED_Lipinski['My_score'] = (round(((whole_data_after_QED_Lipinski['SYBA_score normalized']+whole_data_after_QED_Lipinski['QED normalized']))*100/2,2))
# +
plt.figure(figsize=(10,5))
plt.hist(whole_data_after_QED_Lipinski['My_score'],bins=100)
plt.xlabel('My score', fontsize=14)
plt.ylabel('Number of compounds', fontsize=14)
#plt.title('Histogram of My score distribution')
plt.grid(True)
plt.savefig("tmp/My score distribution.svg")
plt.show()
print(whole_data_after_QED_Lipinski['My_score'].describe())
print('Kurtosis:')
print(round(whole_data_after_QED_Lipinski['My_score'].kurtosis(),4))
print('Skewness:')
print(round(whole_data_after_QED_Lipinski['My_score'].skew(),4))
# -
whole_data_after_QED_Lipinski[(whole_data_after_QED_Lipinski['My_score'] >0) & (whole_data_after_QED_Lipinski['SYBA_score'] >0)]
whole_data_after_QED_Lipinski.head()
new__ = ()
data_ = list(data["SMILES"])
for element in range(len(data_)):
if data_[element] in list(whole_data_after_QED_Lipinski['SMILES']):
print(whole_data_after_QED_Lipinski[whole_data_after_QED_Lipinski['SMILES'] == str(data_[element])][['SMILES', "QED", 'SYBA_score', 'QED normalized', 'SYBA_score normalized', 'My_score']])
else:
pass
| prediction_and_selection/My_score_to_final_structures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Deep Reinforcement Learning
# Book: "Introducción al aprendizaje por refuerzo profundo", <NAME>, 2021, Ed. Watch This Space
# ## Part 1, Chapter 2: Formalización del aprendizaje por refuerzo
# ### Frozen Lake: agent following the good plan in a environment with uncertainty (slippery lake)
# ### <font color='darkblue'>PRELIMINAR ELEMENTS</font>
# Import Gym library: https://gym.openai.com
import gym
# Creation of a class Agent that follows the good plan
class Agent:
def __init__(self):
self.actions = {'left':0, 'down':1, 'right':2, 'up':3}
self.good_plan = 2 * ['down'] + ['right'] + ['down'] + 2 * ['right']
self.step = 0
def select_action(self):
action = self.good_plan[self.step]
self.step = (self.step + 1) % 6 # Trick for running a loop of 'n' episodes without reseting the agent
return self.actions[action]
def reset(self):
self.step = 0
# ### <font color='darkblue'>RUNNING THE AGENT FOLLOWING GOOD WAY IN A NON-SLIPPERY ENVIRONMENT</font>
# +
# Initialize variable 'agent' with the 'Agent' class
agent = Agent()
# Creation of environment FrozenLake, from Gym library, with no splippery
env = gym.make('FrozenLake-v0', is_slippery = False)
# Running and rendering one single episode
env.reset() # reset environment before running episode
env.render() # render the episode
is_done = False # episode completion
t = 0 # time step
while not is_done: # loop of experiences until episode finishes
action = agent.select_action() # passing the decided action
state, reward, is_done, _ = env.step(action) # interaction with environment acc. to decided action
env.render() # render environment state
t += 1
print('\nTotal time steps: ', t)
if state == 15:
print('SOLVED!!! :)')
else:
print('NOT SOLVED :(')
# -
# ### <font color='darkblue'>RUNNING THE AGENT IN A SLIPPERY ENVIRONMENT FOR A NUMBER OF EPISODES AND COUNTING SUCCESS RATE</font>
# Function for running one whole episode
def run_episode(agent, env):
env.reset() # reset environment before running episode
agent.reset() # reset the agent before running
is_done = False # episode completion
t = 0 # time step
while not is_done: # loop of experiences until episode finishes
action = agent.select_action() # passing the decided action
state, reward, is_done, _ = env.step(action) # interaction with environment acc. to decided action
t += 1
return (state, reward, is_done)
# +
# Initialize variable 'agent' with the 'Agent' class
agent = Agent()
# Creation of environment FrozenLake, from Gym library, with splippery condition
env = gym.make('FrozenLake-v0', is_slippery = True)
n_episodes = 10000 # Definition of number of episode to run
solved = 0
for episode in range(n_episodes):
state, reward, is_done = run_episode(agent, env)
if state == 15:
solved += 1
print(f'\n Solved {solved} times in {n_episodes} episodes. Success rate: {solved / n_episodes * 100 :.2f}%')
| P1-C2-FrozenLake_splippery-agent_follows_good_plan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analyse datasets in Timbuctoo-instances
# %%capture
# %run func.ipynb
# ## List properties per database
# +
import pandas as pd
def get_ds_props(url):
ds_names = []
cap_locs = []
desc_hrefs = []
desc_types = []
desc_lens = []
rl_num = []
cl_num = []
rl_locs = []
cl_locs = []
rl_ats = []
rl_completeds = []
resources = []
#n_stats = []
# source description
source_desc = get_sitemap(url)
for c_url in source_desc.rs_urls:
desc_url = None
desc_type = None
ds_names.append(c_url.loc.split('/')[6])
cap_locs.append(c_url.loc) # loc of capability list
desc_url = c_url.describedby_href
desc_type = c_url.describedby_type
# capability list
cap_list = get_sitemap(c_url.loc)
# urls
rl_count = 0
cl_count = 0
rl_url = None
cl_url = None
for rs_url in cap_list.rs_urls:
if rs_url.capability == 'resourcelist':
rl_count += 1
rl_url = rs_url.loc
if rs_url.capability == 'changelist':
cl_count += 1
cl_url = rs_url.loc
rl_num.append(rl_count)
cl_num.append(cl_count)
rl_locs.append(rl_url)
cl_locs.append(cl_url)
# dataset description is in RDF
if desc_url is None:
desc_url = cap_list.describedby_href
desc_type = cap_list.describedby_type
desc_len = None
if desc_url:
desc_graph = get_graph(desc_url, desc_type)
desc_len = len(desc_graph)
desc_hrefs.append(desc_url)
desc_types.append(desc_type)
desc_lens.append(desc_len)
# resourcelist
res_list = get_sitemap(rl_url)
rl_ats.append(res_list.at)
rl_completeds.append(res_list.completed)
# up till now we only encounter 1 resource-url in a resource list.
if len(res_list.rs_urls) != 1:
print('warning, %s has not expected number of 1 resource, but %d' % (rl_url, en(res_list.rs_urls)))
resource = res_list.rs_urls[0].loc
resources.append(resource)
# resource
# n_statements = None
# try:
# graph = get_quads(resource)
# n_statements = len(graph)
# except:
# print('could not load', resource)
# n_stats.append(n_statements)
return pd.DataFrame({'ds_name': ds_names, 'cap_loc': cap_locs,
'desc_href': desc_hrefs, 'desc_type': desc_types, 'desc_len': desc_lens,
'n_rl': rl_num, 'n_cl': cl_num, 'rl_loc': rl_locs, 'cl_loc': cl_locs,
'rl_at': rl_ats, 'rl_completed': rl_completeds, 'resource': resources}) #, 'n_statements': n_stats})
# -
pd.options.display.max_colwidth = 0
df = get_ds_props(ANANSI_URL)
display(df.shape, df.head())
df.to_csv('data/anansi_resources.csv')
df = get_ds_props(HUYDAT_URL)
df.info()
df.to_csv('data/huydat_resources.csv')
# ## Inspecting datasets
# +
from rdflib import Literal
import requests
from IPython.core.display import display, HTML
import urllib
from urllib.parse import urlparse
def fr(x):
return '{:12,}'.format(x).replace(',', '.')
def isresolvable(url):
try:
response = requests.get(url)
return response.ok
except:
return False
class GraphProps(object):
def __init__(self, g):
self.statements = len(g)
self.contexts = list(g.contexts())
self.netlocs_s = dict()
self.netlocs_p = dict()
self.netlocs_o = dict()
self.literals = 0
self.predicates = dict()
for s,p,o in g.triples((None,None,None)):
self.predicates[str(p)] = self.predicates.get(str(p), 0) + 1
nl = urlparse(str(s)).netloc
self.netlocs_s[nl] = self.netlocs_s.get(nl, 0) + 1
nl = urlparse(str(p)).netloc
self.netlocs_p[nl] = self.netlocs_p.get(nl, 0) + 1
if isinstance(o, Literal):
self.literals += 1
else:
nl = urlparse(str(o)).netloc
self.netlocs_o[nl] = self.netlocs_o.get(nl, 0) + 1
def print_props(self):
print('contexts :', fr(len(self.contexts)))
print('statements:', fr(self.statements))
print('literals :', fr(self.literals))
nar = self.literals/self.statements
print('literal ratio:', '{:7.2f}'.format(nar).replace('.', ','))
print()
print('subject netlocs:', len(self.netlocs_s), self.netlocs_s)
print('object netlocs:', len(self.netlocs_o), self.netlocs_o)
print()
edo = 'example.org'
edos = self.netlocs_s.get(edo, 0) + self.netlocs_p.get(edo, 0) + self.netlocs_o.get(edo, 0)
print('example.org', '| s', self.netlocs_s.get(edo, 0), '| p', self.netlocs_p.get(edo, 0), '| o', self.netlocs_o.get(edo, 0), '| total', edos)
print()
print('predicate netlocs:', len(self.netlocs_p), '|', self.netlocs_p)
print('predicates:', len(self.predicates))
for key in sorted(self.predicates):
if urlparse(key).netloc != 'example.org' and isresolvable(key):
display(HTML("""<a target="_blank" href="{}">{}</a>""".format(key, key) + ': ' + str(self.predicates[key])))
else:
print('%s: %s' % (key, self.predicates[key]))
# -
def fingerprint(resource_url):
display(HTML("""<h3>fingerprint</h3>"""))
display(HTML("""<a target="_blank" href="{}">{}</a>""".format(resource_url, resource_url)))
g = get_quads(resource_url)
gp = GraphProps(g)
gp.print_props()
# +
# fingerprint('https://repository.huygens.knaw.nl/v5/resourcesync/u519bd710306620fa7c56d541ae7b9f5b7f57a706/test_wwdocument/dataset.nq')
# +
# rurl ='https://data.anansi.clariah.nl/v5/resourcesync/u74ccc032adf8422d7ea92df96cd4783f0543db3b/gemeentegeschiedenisnl/dataset.nq'
# g = get_quads(rurl)
# +
# from rdflib import URIRef
# max = 5
# tel = 0
# for p, o in g.predicate_objects(URIRef('https://gemeentegeschiedenis.nl/def/geometry_type')):
# tel += 1
# print(p, o)
# if tel >= max:
# break
# -
dfhuydat = pd.read_csv('data/huydat_resources.csv', index_col=0)
for index, row in dfhuydat.iterrows():
resource_url = row['resource']
try:
fingerprint(resource_url)
except Exception as ex:
print(type(ex))
| resourcesync/tim02.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # BIO101 Epidemiology Assignment
# ### By <NAME> and <NAME>
#installing the covirsphy package from github
pip install "git+https://github.com/lisphilar/covid19-sir#egg=covsirphy"
# +
#importing libraries
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn import preprocessing
from scipy import integrate, optimize
import warnings
warnings.filterwarnings('ignore') #to ignore the cases where division by 0 occurs
# ML Supervised learning libraries
import lightgbm as lgb
import xgboost as xgb
from xgboost import plot_importance, plot_tree
from sklearn.model_selection import RandomizedSearchCV, GridSearchCV
from sklearn import linear_model
from sklearn.metrics import mean_squared_error
# +
# importing data
# The Day 1 is starting from 26/2/2020 (26th February 2020)
d=pd.read_csv("C:/Users/uttam/anaconda3/BIOProj/GithubData.csv")
d.tail(10)
# -
d.info()
# There is missing data(45)in the 'Active' column
d.corr()
# The correlation between Confirmed and Days Passed,Death and Days Passed,Recovered and Days Passed,Confirmed and Deaths, Confirmed and Recovered,Recovered and Deaths is VERY HIGH. This implies that they are heavily correlated.
# ## Visualising Data with respect to Days Passed
# confirmed cases
plt.figure(figsize=(20,10))
plt.title("Time vs Confirmed cases",fontsize=20)
sns.barplot(data=d, y="Confirmed",x='Days Passed',palette='gnuplot')
plt.show()
# deceased cases
plt.figure(figsize=(20,10))
plt.title("Time vs Deceased cases",fontsize=20)
sns.barplot(data=d, y="Deaths",x='Days Passed',palette='gnuplot')
plt.show()
#recovered cases
plt.figure(figsize=(20,10))
plt.title("Time vs Recovered cases",fontsize=20)
sns.barplot(data=d, y="Recovered",x='Days Passed',palette='gnuplot')
plt.show()
# ## Visualising Together
#Plotting all three columns together
d[0:114].plot(x='Days Passed', y=["Confirmed","Recovered","Deaths"] ,figsize=(12,8), grid=False,title="Confirmed vs Recovered vs Deaths")
plt.show()
# Clearly, Suuth Carolina's number of infected people's curve has not peaked.. and as the recovered curve has not crossed the confirmed curve, the situation is still an outbreak
# +
# Plotting the rates of fatality and recovery
d[0:].plot(x='Days Passed', y=["Fatal per Confirmed","Recovered per Confirmed","Fatal per(Fatal or recovered)"] ,figsize=(12,8), grid=True,title="Rates")
plt.show()
# -
# ## Growth factor
# Where C is the number of confirmed cases,<br>
# Growth Factor =ΔC(n)/ΔC(n−1)
plt.figure(figsize=(15,10))
plt.title("Growth Factor with respect to Time")
sns.lineplot(data=d,y='Growth Rate',x='Days Passed')
plt.show()
# We see that eventually,the growth rate is approaching 1, ie, earlier there was an outbreak of the coronavirus in South Carolina, but it stabilised with time.
# last 7 days
plt.figure(figsize=(10,8))
plt.title("Growth Rate with respect to Time")
sns.lineplot(data=d[107:],y='Growth Rate',x='Days Passed')
plt.show()
# ## ==============================EDA ENDS================================
# # SIR COVID Model
# There's a lot of information to be extracted from this data; for example, we haven't analyzed the effects of long/lat of countries. However, since our main purpose is to develop a predective model in order to understand the key factors that impact the COVID-19 transmission, we will use the SIR model.<br><br>
# SIR is a simple model that considers a population that belongs to one of the following states:<br>
#
# 1. Susceptible (S). The individual hasn't contracted the disease, but she can be infected due to transmisison from infected people<br>
# 2. Infected (I). This person has contracted the disease<br>
# 3. Recovered/Deceased (R). The disease may lead to one of two destinies: either the person survives, hence developing inmunity to the disease, or the person is deceased.<br>
# ## Defining parameters
# +
# Susceptible equation
def fs(N, a, b, beta):
dsdt = -beta*a*b
return dsdt
# Infected equation
def fi(N, a, b, beta, gamma):
didt = beta*a*b - gamma*b
return didt
# Recovered/deceased equation
def fr(N, b, gamma):
drdt = gamma*b
return drdt
# -
# I preferred to use the <b>Runge Kutta method</b> to solve the SIR model as it is widely used and I found the below online resource particularly helpful<br>
# https://blog.tonytsai.name/blog/2014-11-24-rk4-method-for-solving-sir-model/
# ## Runge Kutta Function Definition
# The Runge-Kutta method finds approximate value of y for a given x. Only first order ordinary differential equations can be solved by using the Runge Kutta 4th order method.
# +
# Defining the Runge Kutta function (4th order to solve for 3 dimensions (S(t),I(t),R(t)))
def rungekutta(N, a, b, c, fs, fi, fr, beta, gamma, hs):
a1 = fs(N, a, b, beta)*hs
b1 = fi(N, a, b, beta, gamma)*hs
c1 = fr(N, b, gamma)*hs
ak = a + a1*0.5
bk = b + b1*0.5
ck = c + c1*0.5
a2 = fs(N, ak, bk, beta)*hs
b2 = fi(N, ak, bk, beta, gamma)*hs
c2 = fr(N, bk, gamma)*hs
ak = a + a2*0.5
bk = b + b2*0.5
ck = c + c2*0.5
a3 = fs(N, ak, bk, beta)*hs
b3 = fi(N, ak, bk, beta, gamma)*hs
c3 = fr(N, bk, gamma)*hs
ak = a + a3
bk = b + b3
ck = c + c3
a4 = fs(N, ak, bk, beta)*hs
b4 = fi(N, ak, bk, beta, gamma)*hs
c4 = fr(N, bk, gamma)*hs
a = a + (a1 + 2*(a2 + a3) + a4)/6
b = b + (b1 + 2*(b2 + b3) + b4)/6
c = c + (c1 + 2*(c2 + c3) + c4)/6
return a, b, c
# -
# ## Parameters and their definitions
#
# N = Total population<br>
# Beta = Rate of transition from Susceptible to Infected (S->I)<br>
# Gamma = Rate of transition from Infected to Recovered/Deceased (I->R)<br>
# K = denotes the constant degree distribution of the network (average value for networks in which the probability of finding a node with a different connectivity decays exponentially fast<br>
# hs = jump step of the numerical integration<br>
#
# ### Definining SIR Model function
#
# +
def sirmodel (N, b0, beta, gamma, hs):
# Initial condition
a = float(N-1)/N - b0 #Susceptible fraction of population considering only 1 person is initiallly infected
b = float(1)/N + b0
c = 0.0
sus, inf, rec= [],[],[]
for i in range(10000): # Run for a certain number of time-steps
sus.append(a)
inf.append(b)
rec.append(c)
a,b,c = rungekutta(N, a, b, c, fs, fi, fr, beta, gamma, hs)
return sus, inf, rec
# -
# ### Sample Example of SIR Model
# +
# Values (Just to showcase an example)
N = 7800*(10**4)
b0 = 0 # Initial fraction of population infected (In factors of N)
beta = 0.5
gamma = 0.2
hs = 0.1
sus, inf, rec = sirmodel(N, b0, beta, gamma, hs)
plt.figure(figsize=(8,6))
plt.title("SIR model with beta "+str(beta)+" and gamma "+str(gamma))
plt.plot(sus, 'b.', label='Susceptible (S)');
plt.plot(inf, 'r.', label='Infected (I)');
plt.plot(rec, 'c.', label='Recovered/deceased (R)');
plt.xlabel("Time(Days Passed)", fontsize=10);
plt.ylabel("Fraction of Population", fontsize=10);
plt.xlim(0,1500)
plt.show()
# -
# ## ========================================================================
# ## Fitting SIR Paramaters to South Carolina Dataset
d.Active.fillna(0) # Filling missing values in Active column to 0
# ## CURVE FITTING
# +
population = float(5150000) #Population of south carolina
country_df = pd.DataFrame()
country_df['ConfirmedCases'] = d['Infected']
country_df = country_df[27:]
country_df['day_count'] = list(range(1,len(country_df)+1))
ydata = [i for i in country_df.ConfirmedCases]
xdata = country_df.day_count
ydata = np.array(ydata, dtype=float)
xdata = np.array(xdata, dtype=float)
N = population
inf0 = ydata[0]
sus0 = N - inf0
rec0 = 0.0
def sir_model(y, x, beta, gamma):
sus = -beta * y[0] * y[1] / N
rec = gamma * y[1]
inf = -(sus + rec)
return sus, inf, rec
def fit_odeint(x, beta, gamma):
return integrate.odeint(sir_model, (sus0, inf0, rec0), x, args=(beta, gamma))[:,1]
popt, pcov = optimize.curve_fit(fit_odeint, xdata, ydata)
fitted = fit_odeint(xdata, *popt)
plt.figure(figsize=(10,8))
plt.plot(xdata, ydata, 'o')
plt.plot(xdata, fitted)
plt.title("Curve Fit of SIR model ")
plt.ylabel("Population infected")
plt.xlabel("Days")
plt.show()
print("Optimal parameters: beta =", popt[0], " and gamma = ", popt[1]/3)
# +
N = 5150000
b0 = 0 # Initial fraction of population infected (In factors of N)
beta = 0.3495254812527788
gamma = 0.10387727709107177
hs = 0.1
sus, inf, rec = sirmodel(N, b0, beta, gamma, hs)
plt.figure(figsize=(8,6))
plt.title("SIR model with beta "+str(beta)+" and gamma "+str(gamma))
plt.plot(sus, 'b.', label='Susceptible (S)');
plt.plot(inf, 'r.', label='Infected (I)');
plt.plot(rec, 'c.', label='Recovered/deceased (R)')
plt.xlabel("Time(Days Passed)", fontsize=10)
plt.ylabel("Fraction of Population", fontsize=10)
plt.xlim(0,1500)
plt.show()
# -
# ## LOGISTIC Regression (Predicted what the classicial SIR model predicted)
# +
from scipy.optimize import curve_fit
x_data = range(len(d.index))
y_data = d['Infected']
def log_curve(x, k, x_0, ymax):
return ymax / (1 + np.exp(-k*(x-x_0)))
# Fit the curve
popt, pcov = curve_fit(log_curve, x_data, y_data, bounds=([0,0,0],np.inf), maxfev=50000)
estimated_k, estimated_x_0, ymax= popt
# Plot the fitted curve
k = estimated_k
x_0 = estimated_x_0
y_fitted = log_curve(range(0,365), k, x_0, ymax)
#print("Optimal K obtained is : "+str(k))
print("Days after which infected curve hits inflection point is : "+str(round(x_0,1)))
print("Maximum number of infected people are : "+str(int(ymax)))
fig = plt.figure(figsize=(10,8))
plt.title("Logisitc Regression Curve for Total Infected Cases",fontsize=15)
ax.legend()
ax = fig.add_subplot(111)
ax.plot(range(0,365), y_fitted, '--', label='Fitted Curve')
ax.plot(x_data, y_data, 'o', label='Confirmed Data')
ax.plot()
# +
population = float(5150000) #Population of south carolina
country_df = pd.DataFrame()
country_df['ConfirmedCases'] = d['Infected']
country_df = country_df[45:]
country_df['day_count'] = list(range(1,len(country_df)+1))
ydata = [i for i in country_df.ConfirmedCases]
xdata = country_df.day_count
ydata = np.array(ydata, dtype=float)
xdata = np.array(xdata, dtype=float)
N = population
inf0 = ydata[0]
sus0 = N - inf0
rec0 = 0.0
def sir_model(y, x, beta, gamma):
sus = -beta * y[0] * y[1] / N
rec = gamma * y[1]
inf = -(sus + rec)
return sus, inf, rec
beta=0
gamma=0.1
def fit_odeint(x, beta, gamma):
return integrate.odeint(sir_model, (sus0, inf0, rec0), x, args=(beta, gamma))[:,1]
popt, pcov = optimize.curve_fit(fit_odeint, xdata, ydata)
fitted = fit_odeint(xdata, *popt)
plt.figure(figsize=(10,8))
plt.plot(xdata, ydata, 'o')
plt.plot(xdata, fitted)
plt.title("Curve Fit of SIR model")
plt.ylabel("Population infected")
plt.xlabel("Days")
plt.show()
print("Optimal parameters: beta =", popt[0], " and gamma = ", popt[1])
# +
rho= 1577169.2522
N = 5150000
b0 = 0 # Initial fraction of population infected (In factors of N)
gamma = 0.031098479389070215
beta = 0.06367276709852013
hs = 0.1
sus, inf, rec = sirmodel(N, b0, beta, gamma, hs)
plt.figure(figsize=(8,6))
plt.title("SIR model with beta "+str(beta)+" and gamma "+str(gamma))
plt.plot(sus, 'b.', label='Susceptible (S)');
plt.plot(inf, 'r.', label='Infected (I)');
plt.plot(rec, 'c.', label='Recovered/deceased (R)');
plt.xlabel("Time(Days Passed)", fontsize=10);
plt.ylabel("Fraction of Population", fontsize=10);
plt.xlim(0,1500)
plt.show()
# -
# ## Clearly, the two advanced algorithms did not work. Hence, I had to proceed with the conventional maths as suggested in the Google Classroom video lectures
# ### -------------------------------------------------------------END OF NOTEBOOK--------------------------------------------------------------------
| DataScience/COVID_SIR/COVID_Logistic_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from PIL import Image
import pytesseract
import argparse
import cv2
import os
pytesseract.pytesseract.tesseract_cmd = 'C:/Program Files (x86)/Tesseract-OCR/tesseract'
file = 'Malawi'
text = ''
for i in range(1,24):
if i<10:
image = cv2.imread(file+'-0'+str(i)+'.jpg')
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
gray = cv2.threshold(gray, 0, 255,cv2.THRESH_BINARY | cv2.THRESH_OTSU)[1]
#gray = cv2.medianBlur(gray, 3)
filename = "{}.png".format(os.getpid())
cv2.imwrite(filename, gray)
text = text + pytesseract.image_to_string(Image.open(filename))
text = text + '\x0c'+'\n'
else:
image = cv2.imread(file+'-'+str(i)+'.jpg')
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
gray = cv2.threshold(gray, 0, 255,cv2.THRESH_BINARY | cv2.THRESH_OTSU)[1]
#gray = cv2.medianBlur(gray, 3)
filename = "{}.png".format(os.getpid())
cv2.imwrite(filename, gray)
text = text + pytesseract.image_to_string(Image.open(filename))
text = text + '\x0c'+'\n'
for i in range(1,29):
print(i)
text_file = open("Malawi.txt", "w",encoding = 'utf-8')
text_file.write(text)
text_file.close()
# +
# print(text)
# -
| 2 Code/image transformation/Malawi/Malawi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os, warnings
import numpy as np
import datetime as dt
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.base import clone
from sklearn.pipeline import Pipeline
from dask_ml.preprocessing import StandardScaler
from dask_ml.decomposition import PCA
from joblib import dump, load
import keras
from keras.layers.core import Dropout
from keras.models import load_model
import geopandas
from rasterio import features
from affine import Affine
import dask
dask.config.set(scheduler='multiprocessing')
import xarray as xr
from dask.diagnostics import ProgressBar
# -
import sys
print(sys.executable)
def sfloat(f):
return str(float(f))
def sint(i):
return str(int(i))
def read_glofas_danube():
glofas = xr.open_dataset('../data/danube/glofas_reanalysis_danube_1981-2002.nc')
glofas = glofas.rename({'lat': 'latitude', 'lon': 'longitude'}) # to have the same name like in era5
glofas = shift_time(glofas, -dt.timedelta(days=1)) # the discharge is the mean of the previous 24h of the timestamp
return glofas
def shift_time(ds, value):
ds.coords['time'].values = pd.to_datetime(ds.coords['time'].values) + value
return ds
def select_riverpoints(dis):
return (dis > 10)
def get_mask_of_basin(da, kw_basins='Danube'):
"""
Parameters:
-----------
da : xr.DataArray
contains the coordinates
kw_basins : str
identifier of the basin in the basins dataset
"""
def transform_from_latlon(lat, lon):
lat = np.asarray(lat)
lon = np.asarray(lon)
trans = Affine.translation(lon[0], lat[0])
scale = Affine.scale(lon[1] - lon[0], lat[1] - lat[0])
return trans * scale
def rasterize(shapes, coords, fill=np.nan, **kwargs):
"""Rasterize a list of (geometry, fill_value) tuples onto the given
xray coordinates. This only works for 1d latitude and longitude
arrays.
"""
transform = transform_from_latlon(coords['latitude'], coords['longitude'])
out_shape = (len(coords['latitude']), len(coords['longitude']))
raster = features.rasterize(shapes, out_shape=out_shape,
fill=fill, transform=transform,
dtype=float, **kwargs)
return xr.DataArray(raster, coords=coords, dims=('latitude', 'longitude'))
# this shapefile is from natural earth data
# http://www.naturalearthdata.com/downloads/10m-cultural-vectors/10m-admin-1-states-provinces/
shp2 = '/raid/home/srvx7/lehre/users/a1303583/ipython/ml_flood/data/drainage_basins/Major_Basins_of_the_World.shp'
basins = geopandas.read_file(shp2)
# print(basins)
single_basin = basins.query("NAME == '"+kw_basins+"'").reset_index(drop=True)
# print(single_basin)
shapes = [(shape, n) for n, shape in enumerate(single_basin.geometry)]
da['basins'] = rasterize(shapes, da.coords)
da = da.basins == 0
return da.drop('basins')
def select_upstream(is_river, lat, lon, basin='Danube'):
# longitude condition
is_west = (~np.isnan(is_river.where(is_river.longitude <= lon))).astype(bool)
mask_basin = get_mask_of_basin(is_river, kw_basins=basin)
nearby_mask = is_river*0.
nearby_mask.loc[dict(latitude=slice(lat+1.5, lat-1.5),
longitude=slice(lon-1.5, lon+1.5))] = 1.
nearby_mask = nearby_mask.astype(bool)
mask = mask_basin & nearby_mask & is_west #mask_box_mean_greater &
if 'basins' in mask.coords:
mask = mask.drop('basins')
if 'time' in mask.coords:
mask = mask.drop('time') # time and basins dimension make no sense here
return mask
def add_shifted_predictors(ds, shifts, variables='all'):
"""Adds additional variables to an array which are shifted in time.
Parameters
----------
ds : xr.Dataset
shifts : list of integers
variables : str or list
"""
if variables == 'all':
variables = ds.data_vars
for var in variables:
for i in shifts:
if i == 0: continue # makes no sense to shift by zero
newvar = var+'-'+str(i)
ds[newvar] = ds[var].shift(time=i)
return ds
def preprocess_reshape_flowmodel(X_dis, y_dis):
"""Reshape, merge predictor/predictand in time, drop nans."""
X_dis = X_dis.to_array(dim='time_feature')
#print('X before feature-stacking', X_dis)
X_dis = X_dis.stack(features=['latitude', 'longitude', 'time_feature'])
#print('X before featuredrop', X_dis)
Xar = X_dis.dropna('features', how='all')
yar = y_dis
yar = yar.drop(['latitude', 'longitude'])
yar.coords['features'] = 'dis'
#print('X, y before concat for time nan dropping', Xar, yar)
Xy = xr.concat([Xar, yar], dim='features')
Xyt = Xy.dropna('time', how='any') # drop them as we cannot train on nan values
time = Xyt.time
Xda = Xyt[:,:-1]
yda = Xyt[:,-1]
return Xda, yda, time
from aux.floodmodels import FlowModel
static = xr.open_dataset('../data/danube/era5_slt_z_slor_lsm_stationary_field.nc')
# +
# era5 = xr.open_dataset('../data/usa/era5_lsp_cp_1981-2017_daysum.nc')
# era5 = shift_time(era5, -dt.timedelta(hours=23))
era5 = xr.open_dataset('../data/danube/era5_danube_pressure_and_single_levels.nc')
# -
glofas = read_glofas_danube()
glofas = glofas.isel(time=slice(0, 365*15)) # just to reduce the amount of data
if 'tp' in era5:
tp = era5['tp']*1000
else:
tp = (era5['cp']+era5['lsp'])*1000
tp.name = 'total precip [mm]'
tp = tp.interp(latitude=glofas.latitude,
longitude=glofas.longitude)
# +
def mkdir(d):
if not os.path.isdir(d):
os.makedirs(d)
def replace(string: str, old_new: dict):
for o, n in old_new.items():
string = string.replace(o, str(n))
return string
# -
# ## prepare training data for the FlowModel
# ### features = discharge upstream (t-1, ... t-3)
# ### target = discharge (t)
#
# +
shifts = range(1,4)
X = add_shifted_predictors(glofas, shifts, variables='all')
X = X.drop('dis') # current dis is to be predicted, is not a feature
y = glofas['dis'] # just this variable as dataarray
# -
# ### for rgp in riverpoints:
# +
N_train = dict(time=slice(None, '1990'))
N_valid = dict(time=slice('1990', '1995'))
# kind, lat, lon will be replaced!
ff_mod = '../models/flowmodel/danube/kind/point_lat_lon_flowmodel.pkl'
ff_hist = '../models/flowmodel/danube/kind/point_lat_lon_history.png'
ff_valid = '../models/flowmodel/danube/kind/point_lat_lon_validation.png'
ff_upstream = '../models/flowmodel/danube/kind/point_lat_lon_upstream.png'
#model = FlowModel('Ridge', dict(alphas=np.logspace(-3, 2, 6)))
model = FlowModel('neural_net', dict(epochs=1000,
#filepath=filepath,
))
pipe = Pipeline([#('scaler', StandardScaler()),
#('pca', PCA(n_components=6)),
('model', model),])
# -
syncr = dask.config.get('scheduler') == 'synchronous'
# +
from joblib import Parallel, delayed
@delayed
def train_flowmodel(lat, lon):
global ff_mod, ff_hist, ff_valid, ff_upstream, is_river
f_mod = replace(ff_mod, dict(lat=lat, lon=lon, kind=model.kind))
if os.path.isfile(f_mod):
return # dont go any further
f_hist = replace(ff_hist, dict(lat=lat, lon=lon, kind=model.kind))
f_valid = replace(ff_valid, dict(lat=lat, lon=lon, kind=model.kind))
f_upstream = replace(ff_upstream, dict(lat=lat, lon=lon, kind=model.kind))
upstream = select_upstream(is_river, lat, lon, basin='Danube')
N_upstream = int(upstream.sum())
if syncr: print(N_upstream)
if N_upstream <= 5:
if syncr:
print(lats, lons, 'is spring.')
#mask_springs.loc[dict(latitude=lat, longitude=lon)] = 1.
#plt.imshow(mask_springs.astype(int))
#plt.title('springs')
#plt.show()
else:
if os.path.isfile(f_mod):
if syncr:
print('already trained.')
else:
if syncr:
print(lats, lons, 'is danube river -> train flowmodel')
try:
fig, ax = plt.subplots()
ax.imshow(upstream.astype(int))
plt.title(str(N_upstream)+' upstream points for '+str(lat)+' '+str(lon))
fig.savefig(f_upstream); plt.close('all')
except:
pass
tp_box = tp.sel(latitude=slice(lat+1.5, lat-1.5),
longitude=slice(lon-1.5, lon+1.5))
noprecip = tp_box.mean(['longitude', 'latitude']) < 0.1
Xt = X.copy()
yt = y.copy()
Xt = Xt.where(noprecip, drop=True)
Xt = Xt.where(upstream, drop=True)
yt = yt.sel(latitude=float(lat), longitude=float(lon))
Xda, yda, time = preprocess_reshape_flowmodel(Xt, yt)
X_train = Xda.loc[N_train]
y_train = yda.loc[N_train]
X_valid = Xda.loc[N_valid]
y_valid = yda.loc[N_valid]
if syncr:
print(X_train.shape, y_train.shape)
print(X_valid.shape, y_valid.shape)
ppipe = clone(pipe)
history = ppipe.fit(X_train.values, y_train.values,
model__validation_data=(X_valid.values,
y_valid.values))
mkdir(os.path.dirname(f_mod))
dump(ppipe, f_mod)
try:
h = history.named_steps['model'].m.model.history
# Plot training & validation loss value
fig, ax = plt.subplots()
ax.plot(h.history['loss'], label='loss')
ax.plot(h.history['val_loss'], label='val_loss')
plt.title('Model loss')
ax.set_ylabel('Loss')
ax.set_xlabel('Epoch')
plt.legend() #['Train', 'Test'], loc='upper left')
ax.set_yscale('log')
fig.savefig(f_hist); plt.close('all')
except Exception as e:
warnings.warn(str(e))
ppipe = load(f_mod)
y_m = ppipe.predict(X_valid)
try:
fig, ax = plt.subplots(figsize=(10,4))
y_m.to_pandas().plot(ax=ax)
y_valid.name = 'reanalysis'
y_valid.to_pandas().plot(ax=ax)
plt.legend()
fig.savefig(f_valid); plt.close('all')
except Exception as e:
warnings.warn(str(e))
# -
# +
danube_gridpoints = get_mask_of_basin(glofas['dis'].isel(time=0), 'Danube')
plt.imshow(danube_gridpoints.astype(int))
plt.show()
#mask_springs = glofas['dis'].isel(time=0)
#mask_springs.values[:] = 0.
dis_map_mean = glofas['dis'].mean('time')
is_river = select_riverpoints(dis_map_mean)
plt.imshow(is_river.astype(int))
plt.title('is_river')
plt.show()
# +
task_list = []
for lon in danube_gridpoints.longitude:
for lat in danube_gridpoints.latitude:
#print(danube_gridpoints.sel(latitude=lat, longitude=lon))
if danube_gridpoints.sel(latitude=lat, longitude=lon) == 1:
lat, lon = float(lat), float(lon)
task_list.append(train_flowmodel(lat, lon))
# -
len(task_list)
# + active=""
# with ProgressBar():
# dask.compute(task_list)
# -
Parallel(n_jobs=20, verbose=10)(task_list)
| python/ml-combimodel#2+flowmodel-final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#IMPORT SEMUA LIBARARY
# -
#IMPORT LIBRARY PANDAS
import pandas as pd
#IMPORT LIBRARY UNTUK POSTGRE
from sqlalchemy import create_engine
import psycopg2
#IMPORT LIBRARY CHART
from matplotlib import pyplot as plt
from matplotlib import style
#IMPORT LIBRARY BASE PATH
import os
import io
#IMPORT LIBARARY PDF
from fpdf import FPDF
#IMPORT LIBARARY CHART KE BASE64
import base64
#IMPORT LIBARARY EXCEL
import xlsxwriter
# +
#FUNGSI UNTUK MENGUPLOAD DATA DARI CSV KE POSTGRESQL
# -
def uploadToPSQL(columns, table, filePath, engine):
#FUNGSI UNTUK MEMBACA CSV
df = pd.read_csv(
os.path.abspath(filePath),
names=columns,
keep_default_na=False
)
#APABILA ADA FIELD KOSONG DISINI DIFILTER
df.fillna('')
#MENGHAPUS COLUMN YANG TIDAK DIGUNAKAN
del df['kategori']
del df['jenis']
del df['pengiriman']
del df['satuan']
#MEMINDAHKAN DATA DARI CSV KE POSTGRESQL
df.to_sql(
table,
engine,
if_exists='replace'
)
#DIHITUNG APABILA DATA YANG DIUPLOAD BERHASIL, MAKA AKAN MENGEMBALIKAN KELUARAN TRUE(BENAR) DAN SEBALIKNYA
if len(df) == 0:
return False
else:
return True
# +
#FUNGSI UNTUK MEMBUAT CHART, DATA YANG DIAMBIL DARI DATABASE DENGAN MENGGUNAKAN ORDER DARI TANGGAL DAN JUGA LIMIT
#DISINI JUGA MEMANGGIL FUNGSI MAKEEXCEL DAN MAKEPDF
# -
def makeChart(host, username, password, db, port, table, judul, columns, filePath, name, subjudul, limit, negara, basePath):
#TEST KONEKSI DATABASE
try:
#KONEKSI KE DATABASE
connection = psycopg2.connect(user=username,password=password,host=host,port=port,database=db)
cursor = connection.cursor()
#MENGAMBL DATA DARI TABLE YANG DIDEFINISIKAN DIBAWAH, DAN DIORDER DARI TANGGAL TERAKHIR
#BISA DITAMBAHKAN LIMIT SUPAYA DATA YANG DIAMBIL TIDAK TERLALU BANYAK DAN BERAT
postgreSQL_select_Query = "SELECT * FROM "+table+" ORDER BY tanggal ASC LIMIT " + str(limit)
cursor.execute(postgreSQL_select_Query)
mobile_records = cursor.fetchall()
uid = []
lengthx = []
lengthy = []
#MELAKUKAN LOOPING ATAU PERULANGAN DARI DATA YANG SUDAH DIAMBIL
#KEMUDIAN DATA TERSEBUT DITEMPELKAN KE VARIABLE DIATAS INI
for row in mobile_records:
uid.append(row[0])
lengthx.append(row[1])
if row[2] == "":
lengthy.append(float(0))
else:
lengthy.append(float(row[2]))
#FUNGSI UNTUK MEMBUAT CHART
#bar
style.use('ggplot')
fig, ax = plt.subplots()
#MASUKAN DATA ID DARI DATABASE, DAN JUGA DATA TANGGAL
ax.bar(uid, lengthy, align='center')
#UNTUK JUDUL CHARTNYA
ax.set_title(judul)
ax.set_ylabel('Total')
ax.set_xlabel('Tanggal')
ax.set_xticks(uid)
#TOTAL DATA YANG DIAMBIL DARI DATABASE, DIMASUKAN DISINI
ax.set_xticklabels((lengthx))
b = io.BytesIO()
#CHART DISIMPAN KE FORMAT PNG
plt.savefig(b, format='png', bbox_inches="tight")
#CHART YANG SUDAH DIJADIKAN PNG, DISINI DICONVERT KE BASE64
barChart = base64.b64encode(b.getvalue()).decode("utf-8").replace("\n", "")
#CHART DITAMPILKAN
plt.show()
#line
#MASUKAN DATA DARI DATABASE
plt.plot(lengthx, lengthy)
plt.xlabel('Tanggal')
plt.ylabel('Total')
#UNTUK JUDUL CHARTNYA
plt.title(judul)
plt.grid(True)
l = io.BytesIO()
#CHART DISIMPAN KE FORMAT PNG
plt.savefig(l, format='png', bbox_inches="tight")
#CHART YANG SUDAH DIJADIKAN PNG, DISINI DICONVERT KE BASE64
lineChart = base64.b64encode(l.getvalue()).decode("utf-8").replace("\n", "")
#CHART DITAMPILKAN
plt.show()
#pie
#UNTUK JUDUL CHARTNYA
plt.title(judul)
#MASUKAN DATA DARI DATABASE
plt.pie(lengthy, labels=lengthx, autopct='%1.1f%%',
shadow=True, startangle=180)
plt.axis('equal')
p = io.BytesIO()
#CHART DISIMPAN KE FORMAT PNG
plt.savefig(p, format='png', bbox_inches="tight")
#CHART YANG SUDAH DIJADIKAN PNG, DISINI DICONVERT KE BASE64
pieChart = base64.b64encode(p.getvalue()).decode("utf-8").replace("\n", "")
#CHART DITAMPILKAN
plt.show()
#MENGAMBIL DATA DARI CSV YANG DIGUNAKAN SEBAGAI HEADER DARI TABLE UNTUK EXCEL DAN JUGA PDF
header = pd.read_csv(
os.path.abspath(filePath),
names=columns,
keep_default_na=False
)
#MENGHAPUS COLUMN YANG TIDAK DIGUNAKAN
header.fillna('')
del header['tanggal']
del header['total']
#MEMANGGIL FUNGSI EXCEL
makeExcel(mobile_records, header, name, limit, basePath)
#MEMANGGIL FUNGSI PDF
makePDF(mobile_records, header, judul, barChart, lineChart, pieChart, name, subjudul, limit, basePath)
#JIKA GAGAL KONEKSI KE DATABASE, MASUK KESINI UNTUK MENAMPILKAN ERRORNYA
except (Exception, psycopg2.Error) as error :
print (error)
#KONEKSI DITUTUP
finally:
if(connection):
cursor.close()
connection.close()
# +
#FUNGSI MAKEEXCEL GUNANYA UNTUK MEMBUAT DATA YANG BERASAL DARI DATABASE DIJADIKAN FORMAT EXCEL TABLE F2
#PLUGIN YANG DIGUNAKAN ADALAH XLSXWRITER
# -
def makeExcel(datarow, dataheader, name, limit, basePath):
#MEMBUAT FILE EXCEL
workbook = xlsxwriter.Workbook(basePath+'jupyter/BLOOMBERG/SektorKeuangandanPerbankan/excel/'+name+'.xlsx')
#MENAMBAHKAN WORKSHEET PADA FILE EXCEL TERSEBUT
worksheet = workbook.add_worksheet('sheet1')
#SETINGAN AGAR DIBERIKAN BORDER DAN FONT MENJADI BOLD
row1 = workbook.add_format({'border': 2, 'bold': 1})
row2 = workbook.add_format({'border': 2})
#MENJADIKAN DATA MENJADI ARRAY
data=list(datarow)
isihead=list(dataheader.values)
header = []
body = []
#LOOPING ATAU PERULANGAN, KEMUDIAN DATA DITAMPUNG PADA VARIABLE DIATAS
for rowhead in dataheader:
header.append(str(rowhead))
for rowhead2 in datarow:
header.append(str(rowhead2[1]))
for rowbody in isihead[1]:
body.append(str(rowbody))
for rowbody2 in data:
body.append(str(rowbody2[2]))
#MEMASUKAN DATA DARI VARIABLE DIATAS KE DALAM COLUMN DAN ROW EXCEL
for col_num, data in enumerate(header):
worksheet.write(0, col_num, data, row1)
for col_num, data in enumerate(body):
worksheet.write(1, col_num, data, row2)
#FILE EXCEL DITUTUP
workbook.close()
# +
#FUNGSI UNTUK MEMBUAT PDF YANG DATANYA BERASAL DARI DATABASE DIJADIKAN FORMAT EXCEL TABLE F2
#PLUGIN YANG DIGUNAKAN ADALAH FPDF
# -
def makePDF(datarow, dataheader, judul, bar, line, pie, name, subjudul, lengthPDF, basePath):
#FUNGSI UNTUK MENGATUR UKURAN KERTAS, DISINI MENGGUNAKAN UKURAN A4 DENGAN POSISI LANDSCAPE
pdf = FPDF('L', 'mm', [210,297])
#MENAMBAHKAN HALAMAN PADA PDF
pdf.add_page()
#PENGATURAN UNTUK JARAK PADDING DAN JUGA UKURAN FONT
pdf.set_font('helvetica', 'B', 20.0)
pdf.set_xy(145.0, 15.0)
#MEMASUKAN JUDUL KE DALAM PDF
pdf.cell(ln=0, h=2.0, align='C', w=10.0, txt=judul, border=0)
#PENGATURAN UNTUK UKURAN FONT DAN JUGA JARAK PADDING
pdf.set_font('arial', '', 14.0)
pdf.set_xy(145.0, 25.0)
#MEMASUKAN SUB JUDUL KE PDF
pdf.cell(ln=0, h=2.0, align='C', w=10.0, txt=subjudul, border=0)
#MEMBUAT GARIS DI BAWAH SUB JUDUL
pdf.line(10.0, 30.0, 287.0, 30.0)
pdf.set_font('times', '', 10.0)
pdf.set_xy(17.0, 37.0)
#PENGATURAN UNTUK UKURAN FONT DAN JUGA JARAK PADDING
pdf.set_font('Times','',10.0)
#MENGAMBIL DATA HEADER PDF YANG SEBELUMNYA SUDAH DIDEFINISIKAN DIATAS
datahead=list(dataheader.values)
pdf.set_font('Times','B',12.0)
pdf.ln(0.5)
th1 = pdf.font_size
#MEMBUAT TABLE PADA PDF, DAN MENAMPILKAN DATA DARI VARIABLE YANG SUDAH DIKIRIM
pdf.cell(100, 2*th1, "Kategori", border=1, align='C')
pdf.cell(177, 2*th1, datahead[0][0], border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Jenis", border=1, align='C')
pdf.cell(177, 2*th1, datahead[0][1], border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Pengiriman", border=1, align='C')
pdf.cell(177, 2*th1, datahead[0][2], border=1, align='C')
pdf.ln(2*th1)
pdf.cell(100, 2*th1, "Satuan", border=1, align='C')
pdf.cell(177, 2*th1, datahead[0][3], border=1, align='C')
pdf.ln(2*th1)
#PENGATURAN PADDING
pdf.set_xy(17.0, 75.0)
#PENGATURAN UNTUK UKURAN FONT DAN JUGA JARAK PADDING
pdf.set_font('Times','B',11.0)
data=list(datarow)
epw = pdf.w - 2*pdf.l_margin
col_width = epw/(lengthPDF+1)
#PENGATURAN UNTUK JARAK PADDING
pdf.ln(0.5)
th = pdf.font_size
#MEMASUKAN DATA HEADER YANG DIKIRIM DARI VARIABLE DIATAS KE DALAM PDF
pdf.cell(50, 2*th, str("Negara"), border=1, align='C')
for row in data:
pdf.cell(40, 2*th, str(row[1]), border=1, align='C')
pdf.ln(2*th)
#MEMASUKAN DATA ISI YANG DIKIRIM DARI VARIABLE DIATAS KE DALAM PDF
pdf.set_font('Times','B',10.0)
pdf.set_font('Arial','',9)
pdf.cell(50, 2*th, negara, border=1, align='C')
for row in data:
pdf.cell(40, 2*th, str(row[2]), border=1, align='C')
pdf.ln(2*th)
#MENGAMBIL DATA CHART, KEMUDIAN CHART TERSEBUT DIJADIKAN PNG DAN DISIMPAN PADA DIRECTORY DIBAWAH INI
#BAR CHART
bardata = base64.b64decode(bar)
barname = basePath+'jupyter/BLOOMBERG/SektorKeuangandanPerbankan/img/'+name+'-bar.png'
with open(barname, 'wb') as f:
f.write(bardata)
#LINE CHART
linedata = base64.b64decode(line)
linename = basePath+'jupyter/BLOOMBERG/SektorKeuangandanPerbankan/img/'+name+'-line.png'
with open(linename, 'wb') as f:
f.write(linedata)
#PIE CHART
piedata = base64.b64decode(pie)
piename = basePath+'jupyter/BLOOMBERG/SektorKeuangandanPerbankan/img/'+name+'-pie.png'
with open(piename, 'wb') as f:
f.write(piedata)
#PENGATURAN UNTUK UKURAN FONT DAN JUGA JARAK PADDING
pdf.set_xy(17.0, 75.0)
col = pdf.w - 2*pdf.l_margin
widthcol = col/3
#MEMANGGIL DATA GAMBAR DARI DIREKTORY DIATAS
pdf.image(barname, link='', type='',x=8, y=100, w=widthcol)
pdf.set_xy(17.0, 75.0)
col = pdf.w - 2*pdf.l_margin
pdf.image(linename, link='', type='',x=103, y=100, w=widthcol)
pdf.set_xy(17.0, 75.0)
col = pdf.w - 2*pdf.l_margin
pdf.image(piename, link='', type='',x=195, y=100, w=widthcol)
pdf.ln(2*th)
#MEMBUAT FILE PDF
pdf.output(basePath+'jupyter/BLOOMBERG/SektorKeuangandanPerbankan/pdf/'+name+'.pdf', 'F')
# +
#DISINI TEMPAT AWAL UNTUK MENDEFINISIKAN VARIABEL VARIABEL SEBELUM NANTINYA DIKIRIM KE FUNGSI
#PERTAMA MANGGIL FUNGSI UPLOADTOPSQL DULU, KALAU SUKSES BARU MANGGIL FUNGSI MAKECHART
#DAN DI MAKECHART MANGGIL FUNGSI MAKEEXCEL DAN MAKEPDF
# +
#DEFINISIKAN COLUMN BERDASARKAN FIELD CSV
columns = [
"kategori",
"jenis",
"tanggal",
"total",
"pengiriman",
"satuan",
]
#UNTUK NAMA FILE
name = "SektorKeuangandanPerbankan1_1"
#VARIABLE UNTUK KONEKSI KE DATABASE
host = "localhost"
username = "postgres"
password = "<PASSWORD>"
port = "5432"
database = "bloomberg_SektorKeuangandanPerbankan"
table = name.lower()
#JUDUL PADA PDF DAN EXCEL
judul = "Data Sektor Keuangan Dan Perbankan"
subjudul = "Badan Perencanaan Pembangunan Nasional"
#LIMIT DATA UNTUK SELECT DI DATABASE
limitdata = int(8)
#NAMA NEGARA UNTUK DITAMPILKAN DI EXCEL DAN PDF
negara = "Indonesia"
#BASE PATH DIRECTORY
basePath = 'C:/Users/ASUS/Documents/bappenas/'
#FILE CSV
filePath = basePath+ 'data mentah/BLOOMBERG/SektorKeuangandanPerbankan/' +name+'.csv';
#KONEKSI KE DATABASE
engine = create_engine('postgresql://'+username+':'+password+'@'+host+':'+port+'/'+database)
#MEMANGGIL FUNGSI UPLOAD TO PSQL
checkUpload = uploadToPSQL(columns, table, filePath, engine)
#MENGECEK FUNGSI DARI UPLOAD PSQL, JIKA BERHASIL LANJUT MEMBUAT FUNGSI CHART, JIKA GAGAL AKAN MENAMPILKAN PESAN ERROR
if checkUpload == True:
makeChart(host, username, password, database, port, table, judul, columns, filePath, name, subjudul, limitdata, negara, basePath)
else:
print("Error When Upload CSV")
# -
| jupyter/BLOOMBERG/SektorKeuangandanPerbankan/script/SektorKeuangandanPerbankan1_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
import numpy as np
x = np.linspace(-3, 3)
y = np.linspace(-3, 3)
xv, yv = np.meshgrid(x, y)
def rosenbrock(x, y):
return 100 * (y - x ** 2) ** 2 + (1 - x) ** 2
def sphere(x, y):
return (x - np.array([1] * len(x))) ** 2 + (y - np.array([2] * len(y))) ** 2
# z = rosenbrock(xv, yv)
z = sphere(xv, yv)
import matplotlib.pyplot as plt
# %pylab inline
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.plot_surface(xv, yv, z)
data = []
for x_, y_, z_ in zip(xv.flatten().tolist(), yv.flatten().tolist(), z.flatten().tolist()):
data.append([x_, y_, z_])
data
var = [1.0, 2.1, 3.2]
var_str = [str(val) for val in var]
" ".join(var_str)
# write data to file
file = open("dataset.txt", "w")
for entry in data:
entry_str = "%.4f %.4f %.4f\n" % (entry[0], entry[1], entry[2])
# entry_str.append('\n')
file.write(entry_str)
| DeepModelTraining/dataset/create_dataset.py.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Machine learning models – Exercises
#
# In these exercises, we'll load a cleaned and featurized dataframe then use scikit-learn to predict materials properties.
#
# Before starting, we need to use matminer's `load_dataframe_from_json()` function to load a cleaned and featurized version of the `dielectric_constant` dataset. We will use this dataset for all the exercises.
# +
import os
from matminer.utils.io import load_dataframe_from_json
df = load_dataframe_from_json(os.path.join("resources", "dielectric_constant_featurized.json"))
df.head()
# -
# ## Exercise 1: Split dataset in target property and features
#
# You first need to partition the data into the target property and features used for learning. For this dataset, the target property is contained in the `total_dielectric` column. The features are all other columns, except `structure`, and `composition`.
#
# The target property data should be stored in the `y` variable. The set of features used for learning should be stored in the `X` variable.
#
# *Hint remember to exclude the target property from the feature set.*
# +
# Fill in the blanks
y = df[_____].values
X = df.drop(_____, axis=1)
# -
# ## Exercise 2: Train a random forest model on the dataset
#
# Train a random forest model with 150 estimators on the dataset. Next, use the model to get predictions for all samples and store them to the `y_pred` variable.
# +
from sklearn.ensemble import RandomForestRegressor
# Fill in the blanks below
rf = RandomForestRegressor(____)
rf.fit(__, __)
y_pred = rf.predict(___)
# -
# To see how well your model is performing, run the next cell.
# +
import numpy as np
from sklearn.metrics import mean_squared_error
mse = mean_squared_error(y, y_pred)
print('training RMSE = {:.3f}'.format(np.sqrt(mse)))
# -
# ## Exercise 3: Evaluate the model using cross validation
# Evaluate your random forest model using cross validation with 5 splits. This will give a more realistic idea of how well your model will perform in practice.
# +
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
# Fill in the blanks below
kfold = KFold(___)
scores = cross_val_score(__, __, __, scoring='neg_mean_squared_error', cv=___)
# -
# The final cross validation score can be printed by running the cell below.
rmse_scores = [np.sqrt(abs(s)) for s in scores]
print('Mean RMSE: {:.3f}'.format(np.mean(rmse_scores)))
| workshop/lessons/08_ml_matminer/unit-3-exercies.ipynb |
# ---
# title: "What Is The Probability An Economy Class Seat Is An Aisle Seat?"
# author: "<NAME>"
# date: 2017-12-20T11:53:49-07:00
# description: "What Is The Probability An Economy Class Seat Is An Aisle Seat?"
# type: technical_note
# draft: false
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# There are two types of people in the world, aisle seaters and window seaters. I am an aisle seater, nothing is worse than limited bathroom access on a long flight. The first thing I do when I get my ticket is check to see if I have a window seat. If not, I immediately head over to the airline counter and try to get one.
#
# Last flight, on Turkish Airlines, I ran into a curious situation. I recieved my boarding pass with my seat number, 18C, but the ticket did not specify if C was an aisle seat or not. Making matters worse, the airline counter was swamped with a few dozen people. So I asked myself: **given only the seat letter, C, what is the probability that it is an aisle seat?**
#
# Later, on the flight, I decided to find out.
# ## Preliminaries
# +
# Import required modules
import pandas as pd
import numpy as np
# Set plots to display in the iPython notebook
# %matplotlib inline
# -
# ## Setup possible seat configurations
# I am a pretty frequently flyer on a variety of airlines and aircraft. There are a variety of seating configurations out there, but typically they follow some basic rules:
#
# - No window cluster of seats has more than three seats.
# - On small flights with three seats, the single seat is on the left side.
# - No flight has more than nine rows.
#
# Based on these rules, here are the "typical" seating configurations from aircraft with between two and nine seats per row. A '1' codifies that a seat is an aisle seat, a '0' codifies that it is a non-aisle seat (i.e. window or middle), and 'np.nan' denotes that the aircraft has less than nine seats (this is so all the list lengths are the same).
# +
# An aircraft with two seats per row
rows2 = [1,1, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan]
# An aircraft with three seats per row
rows3 = [1,1,0, np.nan, np.nan, np.nan, np.nan, np.nan, np.nan,]
# An aircraft with four seats per row
rows4 = [0,1,1,0, np.nan, np.nan, np.nan, np.nan, np.nan]
# An aircraft with five seats per row
rows5 = [0,1,1,0,0, np.nan, np.nan,np.nan, np.nan]
# An aircraft with six seats per row
rows6 = [0,1,1,1,1,0, np.nan, np.nan, np.nan]
# An aircraft with seven seats per row
rows7 = [0,1,1,0,1,1,0, np.nan, np.nan]
# An aircraft with eight seats per row
rows8 = [0,0,1,1,1,1,0,0, np.nan]
# An aircraft with nine seats per row
rows9 = [0,0,1,1,0,1,1,0,0]
# -
# For example, in an aircraft with five seats per row, `rows5`, the seating arrangement would be:
#
# 1. window
# 2. aisle
# 3. aisle
# 4. middle
# 5. window
# 6. no seat
# 7. no seat
# 8. no seat
# 9. no seat
# Next, I'm take advantage of pandas row summation options, but to do this I need to wrangle the data into a pandas dataframe. Essentially I am using the pandas dataframe as a matrix.
# Create a list variable of all possible aircraft configurations
seating_map = [rows2, rows3, rows4, rows5, rows6, rows7, rows8, rows9]
# Create a dataframe from the seating_map variable
df = pd.DataFrame(seating_map,
columns=['A', 'B', 'C', 'D', 'E', 'F', 'G', 'H', 'I'],
index=['rows2', 'rows3', 'rows4', 'rows5', 'rows6', 'rows7', 'rows8', 'rows9'])
# Here is all the data we need to construct our probabilities. The columns represent individual seat letters (A, B, etc.) while the rows represent the number of seats-per-row in the aircraft.
# View the dataframe
df
# ## Calculate aisle probability
#
# Because each aircraft seats-per-row configuration (i.e. row) is binary (1 if aisle, 0 if non-aisle), the probability that a seat is an aisle is simply the mean value of each seat letter (i.e. column).
# Create a list wherein each element is the mean value of a column
aisle_probability = [df['A'].mean(),
df['B'].mean(),
df['C'].mean(),
df['D'].mean(),
df['E'].mean(),
df['F'].mean(),
df['G'].mean(),
df['H'].mean(),
df['I'].mean()]
# Display the variable
aisle_probability
# So there you have it, the probability that each seat letter is an aisle. However, we can make the presentation a little more intituative.
# ## Visualize seat letter probabilities
# The most obvious visualization to convey the probabilities would be seat letters on the x-axis and probabilities on the y-axis. Panda's plot function makes that easy.
# Create a list of strings to use as the x-axis labels
seats = ['Seat A', 'Seat B', 'Seat C', 'Seat D',
'Seat E', 'Seat F', 'Seat G', 'Seat H', 'Seat I']
# Plot the probabilities, using 'seats' as the index as a bar chart
pd.Series(aisle_probability, index=seats).plot(kind='bar', # set y to range between 0 and 1
ylim=[0,1],
# set the figure size
figsize=[10,6],
# set the figure title
title='Probabilty of being an Aisle Seat in Economy Class')
# So there we have it! If given a boarding pass with seat C you have a 86% probability of being in an aisle seat!
#
# I hope this was helpful!
| docs/python/other/aisle_seat_probabilities.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Case Study: Analysis of a Double Pipe Heat Exchanger
#
# A stalwart of undergraduate chemical engineering laboratories is study of a double-pipe heat exchanger in counter-current flow. In this case, a student group collected multiple measurements of flow and temperature data from a heat exchanger with sensors configured as shown in the following diagram. (Note: The diagram shows co-current flow. The data was collected with the valves configured for counter-current flow of the hot stream.)
#
# 
# Source: [Gunt WL315C Product Description](https://www.gunt.de/en/products/comparison-of-various-heat-exchangers/060.315C0/wl315c/glct-1:pa-148:pr-1495)
# ## Reading Data
#
# The raw data was copied to a new sheet in the same Google Sheets file, edited to conform with Tidy Data, and a link created using the procedures outlined above for reading data from Google Sheets. The data is read in the following cell.
hx = pd.read_csv("https://docs.google.com/spreadsheets/d/e/2PACX-1vSNUCEFMaGZ-y18p-AnDoImEeenMLbRxXBABwFNeP8I3xiUejolPJx-kr4aUywD0szRel81Kftr8J0R/pub?gid=865146464&single=true&output=csv")
hx.head()
# ## Energy Balances
#
# The first step in this analysis is to verify the energy balance.
#
# $$
# \begin{align*}
# Q_h & = \dot{q}_h \rho C_p (T_{h,in} - T_{h,out}) \\
# Q_c & = \dot{q}_c \rho C_p (T_{c,out} - T_{c, in})
# \end{align*}
# $$
#
# The next cell creates two new calculated variables in the dataframe for $Q_h$ and $Q_c$, and uses the pandas plotting facility to visualize the results. This calculation takes advantage of the "one variable per column" rule of Tidy Data which enables calculations for all observations to be done in a single line of code.
# +
# heat capacity of water
rho = 1.00 # kg / liter
Cp = 4.18 # kJ/ kg / deg C
# heat balances
hx["Qh"] = rho * Cp * hx["Hot Flow (L/hr)"] * (hx["H Inlet"] - hx["H Outlet"]) / 3600
hx["Qc"] = rho * Cp * hx["Cold Flow (L/hr)"] * (hx["C Outlet"] - hx["C Inlet"]) / 3600
hx["Loss (%)"] = 100 * (1 - hx["Qc"]/hx["Qh"])
# plot
display(hx[["Qh", "Qc", "Loss (%)"]].style.format(precision=2))
hx.plot(y = ["Qh", "Qc"], ylim = (0, 15), grid=True, xlabel="Observation", ylabel="kW")
# -
# ## Overall Heat Transfer Coefficient $UA$
#
# The performance of a counter-current heat exchanger is given the relationship
#
# $$Q = U A \Delta T_{lm} $$
#
# where $\Delta T_{lm}$ is the log-mean temperature given by
#
# $$
# \begin{align*}
# \Delta T_0 & = T_{h, out} - T_{c, in} \\
# \Delta T_1 & = T_{h, in} - T_{c, out} \\
# \\
# \Delta T_{lm} & = \frac{\Delta T_1 - \Delta T_0}{\ln\frac{\Delta T_1}{\Delta T_0}}
# \end{align*}
# $$
# +
dT0 = hx["H Outlet"] - hx["C Inlet"]
dT1 = hx["H Inlet"] - hx["C Outlet"]
hx["LMTD"] = (dT1 - dT0) / np.log(dT1/dT0)
Q = (hx.Qh + hx.Qc)/2
hx["UA"] = Q/hx.LMTD
hx.plot(y="UA", xlabel="Observation", ylabel="kW/deg C", grid=True)
# -
# ## How does $UA$ depend on flowrates?
#
# The data clearly demonstrate that the heat transfer coefficient in the double pipe heat exchanger depends on flowrates of both the cold and hot liquid streams. We can see this by inspecting the data.
hx[["Flow Rate H", "Flow Rate C", "Hot Flow (L/hr)", "Cold Flow (L/hr)", "UA"]]
# The replicated measurements provide an opportunity to compute averages. Here we use the pandas `.groupby()` function to group observations and compute means. The data will be used to plot results, so we'll save the results of these calculations as a new dataframe for reuse.
sx = hx.groupby(["Flow Rate H", "Flow Rate C"]).mean()[["Hot Flow (L/hr)", "Cold Flow (L/hr)", "UA"]]
sx
# +
import matplotlib.pyplot as plt
fig, ax = plt.subplots(1, 1)
sx.sort_values("Cold Flow (L/hr)").groupby("Flow Rate H").plot(x = "Cold Flow (L/hr)", y = "UA",
style={"UA": 'ro-'}, ax=ax)
# -
fig, ax = plt.subplots(1, 1)
sx.sort_values("Hot Flow (L/hr)").groupby("Flow Rate C").plot(x = "Hot Flow (L/hr)", y = "UA",
style={"UA": 'ro-'}, ax=ax)
# ## Fitting a Model for $UA$
#
# For a series of transport mechanisms, the overall heat transfer coefficient
#
# $$\frac{1}{UA} = \frac{1}{U_hA} + \frac{1}{U_{tubeA}} + \frac{1}{U_cA}$$
#
# $U_{tube}A$ is a constant for this experiment. $U_h$A and $U_c$A varying with flowrate and proporitonal to dimensionless Nusselt number. The hot and cold liquid flows in the double pipe heat exchanger are well within the range for fully developed turbulent flow. Under these conditions for flows inside closed tubes, the Dittus-Boelter equation provides an explicit expression for Nusselt number
#
# $$Nu = C \cdot Re ^{4/5} Pr ^ n$$
#
# where $C$ is a constant, $Re$ is the Reynold's number that is proportional to flowrate, and $Pr$ is the Prandtl number determined by fluid properties.
#
# Experimentally, consider a set of values for $UA$ determined by varying $\dot{m}_h$ and $\dot{m}_c$ over range of values. Because Reynold's number is proportional to flowrate, we can propose a model
#
# $$\frac{1}{UA} = R = R_{t} + r_h \dot{q}_h^{-0.8} + r_c \dot{q}_h^{-0.8}$$
#
# This suggests a linear regression for $R = \frac{1}{UA}$ in terms of $X_h = \dot{q}_h^{-0.8}$ and $X_c = \dot{q}_c^{-0.8}$.
#
# :::{admonition} Enhancement: Convert to Skikit-learn
#
# The following model fit should be converted to Scikit-learn to enable use of ML tools to the model building process.
#
# :::
hx["R"] = 1.0/hx["UA"]
hx["Xh"] = hx["Hot Flow (L/hr)"]**(-0.8)
hx["Xc"] = hx["Cold Flow (L/hr)"]**(-0.8)
# +
import statsmodels.formula.api as sm
result = sm.ols(formula="R ~ Xh + Xc", data = hx).fit()
print(result.params)
print(result.summary())
# +
hx["Rh"] = 115.3 * hx["Xh"]
hx["Rc"] = 186.3 * hx["Xc"]
hx["Rt"] = 0.142
hx["R_pred"] = hx["Rt"] + hx["Rh"] + hx["Rc"]
hx[["R", "R_pred", "Rt", "Rh", "Rc"]]
# -
# ## Comparison of Model to Experimental Data
hx["UA_pred"] = 1/hx["R_pred"]
hx.plot(y = ["UA", "UA_pred"], grid=True, title="Heat Transfer Coefficient")
| _build/jupyter_execute/notebooks/01/04_case-study.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import math
import matplotlib.pyplot as plt
# x values
x = range(1, 20)
# O(1)
y1 = [1 for n in x]
# O(log n)
y2 = [math.log(n) for n in x]
# O(n)
y3 = x
# O(n log n)
y4 = [n * math.log(n) for n in x]
# O(n^2)
y5 = [n**2 for n in x]
# O(2^n)
y6 = [2**n for n in x]
# O(n!)
y7 = [math.factorial(n) for n in x]
# O(n^n)
y8 = [n**n for n in x]
plt.figure(figsize = (18, 8))
plt.title('Big O Notation', fontsize = 15)
plt.xlabel('Elements', fontsize=12)
plt.ylabel('Operations', fontsize=12)
# O(1)
plt.plot(x, y1, label='$\mathcal{O}(1)$', color='navy')
# lightcoral, orange, mediumseagreen, cadetblue, dodgerblue, slateblue, violet, crimson
# O(log n)
plt.plot(x, y2, label='$\mathcal{O}(logn)$', color='crimson')
# O(n)
plt.plot(x, y3, label='$O(n)$', color='darkgreen')
# O(n log n)
plt.plot(x, y4, label='$\mathcal{O}(nlogn)$', color='cadetblue')
# O(n^{2})
plt.plot(x, y5, label='$\mathcal{O}(n^{2})$', color='slateblue')
# O(2^{n})
plt.plot(x, y6, label='$\mathcal{O}(2^{n})$', color='slategray')
# O(n!)
plt.plot(x, y7, label='$\mathcal{O}(n!)$', color='goldenrod')
# O(n^{n})
plt.plot(x, y8, label='$\mathcal{O}(n^{n})$', color='rebeccapurple')
plt.xlim(1, 11)
plt.ylim(0, 50)
plt.legend(loc='upper right', fontsize=14)
plt.show()
# -
| big-o-notation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Kort introduksjon til jupyter notebook
#
# Dokumentet du nå leser er en [Jupyter notebook](https://jupyter.org/), satt opp via [binder](https://mybinder.org) og som kjører på [Google Cloud](https://cloud.google.com). Dokumentet er ikke en statisk nettside, men et interaktivt miljø som lar deg skrive og kjøre kode i Python.
#
# For eksempel, her er en kort Python-kode som regner ut en verdi, lagrer den i en variabel og skriver ut verdien:
sekunder_i_en_dag = 24 * 60 * 60
print(sekunder_i_en_dag)
# En jupyter notebook er delt inn i flere celler som inneholder kode eller tekst. Denne cellen inneholder bare tekst, mens cellen over inneholder Python-kode.
#
# For å kjøre koden i cellen over, velg cellen (ved å klikke på den) og så trykk på **Run**-knappen eller bruk hurtigtaster: "Command/Ctrl+Enter". Du kan også bruke hurtigtastene "Shift+Enter" for å kjøre cellen og automatisk flytte til neste celle.
#
# Alle celler modifiserer den globale tilstanden, så variabler du definerer ved å kjøre en celle kan brukes i alle andre celler:
sekunder_i_en_uke = 7 * sekunder_i_en_dag
print(sekunder_i_en_uke)
# ## Lagre arbeidet ditt
#
# Siden denne siden kjører via Binder, så vil endringene du gjør her **ikke** lagres dersom du lukker ned nettleseren eller trykker *Logout* over.
#
# Så dersomdu ønsker å ta vare på arbeidet ditt, så må du laste det ned. Det kan gjøre via *File* -> *Download as* -> *Velg ønsket filformat*.
# # Lage plott
#
# For å lage figurer i Python bruker vi en pakke som heter [matplotlib](https://matplotlib.org/). Videre bruker vi ofte en egen pakke for matematiske operasjoner som heter [numpy](https://numpy.org/).
#
# Som et eksempel skal vi nå plotte funksjonen $1 - x^2$ mellom -1 og 1.
#
# Vi importerer først pakkene vi skal bruke:
import numpy as np # Importerer numpy og gir den lokalt navn "np".
import matplotlib.pyplot as plt # Importerer matplotlib.pyplot og gir den lokalt navn "plt".
# Her gir vi pakkene lokale navn når vi importerer de, dette er bare for å spare litt skriving!
# La oss lage en liste x-verdier fra -1 til 1, i steg av 0.01:
x = np.arange(-1, 1 + 0.01, 0.01)
print(x[:20]) # Skriv ut de 20 første x-verdiene.
# **Tips:**
# Dersom du vil vite mer om en funksjon vi bruker, f.eks. `numpy.arange()`, så kan du klikke i koden der du skriver funksjonen og taste "Shift+Tab" for å få opp help for funksjonen (trykker du mer enn en gang får du mer fyldig informasjon).
# Vi skulle plotte $1-x^2$:
y = 1.0 - x**2
fig, ax = plt.subplots()
ax.plot(x, y, label='Min første funksjon: y = 1 - x²')
ax.legend() # Vis figurtekst.
ax.set_xlabel('x') # Legg til tekst på x-aksen.
ax.set_ylabel('y'); # Legg til tekst på y-aksen.
# ## Lagre figuren
# For å lagre plottet ditt, kan du kjøre koden ``fig.savefig('min_figur.png')``. Dette lagrer figuren din med navnet ``min_figur.png``.
#
# For å få lastet ned den lagrede figuren, velg `File -> Open`. Der du vil få en oversikt over hvilke filer du har tilgang til og du kan her velge figuren og laste den ned ved å trykke på *Download*.
fig.savefig('min_figur.png')
# Dette dokumentet er inspirert av introduksjonen til [Google Colaboratory](https://colab.research.google.com/notebooks/intro.ipynb)
# En grei måte å lære seg Python på er å eksperimentere litt, og kanskje bruke det til å løse oppgaver fra øvinger eller liknende.
| jupyter/introduksjon/Introduksjon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from bqplot import Pie, Figure
from IPython.display import display
import numpy as np
# ## Basic Pie Chart
# Generating the simplest possible pie chart and plotting it in a `Figure`.
#
# The area and angle of each slice will be proportional to the data passed to `sizes`
data = range(1, 6)
pie = Pie(sizes=data)
fig = Figure(marks=[pie], animation_duration=1000)
# Add `animation_duration` (in milliseconds) to have smooth transitions
display(fig)
# As with all bqplot Marks, pie data can be dynamically modified
Nslices = 5
pie.sizes = np.random.rand(Nslices)
# Sort the pie slices by ascending size
pie.sort = True
# Setting different styles for selected slices
pie.selected_style = {"opacity": "1", "stroke": "white", "stroke-width": "2"}
pie.unselected_style = {"opacity": "0.2"}
pie.selected = [3]
pie.selected = None
# For more on piechart interactions, see the [Mark Interactions notebook](Mark Interactions.ipynb)
# ### Adding labels
pie.labels = ['{:.2f}'.format(d) for d in pie.sizes]
fig
# Modify label styling
pie.label_color = 'white'
pie.font_size = '20px'
pie.font_weight = 'normal'
# ## Updating pie shape and style
pie1 = Pie(sizes=np.random.rand(6), inner_radius=0.05)
fig1 = Figure(marks=[pie1], animation_duration=1000)
display(fig1)
# ### Change pie dimensions
# As of now, the radius sizes are absolute, in pixels
pie1.radius = 250
pie1.inner_radius = 80
# Angles are in radians, 0 being the top vertical
pie1.start_angle = -90
pie1.end_angle = 90
# ### Moving the pie around
# `x` and `y` attributes control the position of the pie in the figure.
# If no scales are passed for `x` and `y`, they are taken in absolute
# figure coordinates, between 0 and 1.
pie1.y = 0.9
pie1.x = 0.4
pie1.radius = 320
# ### Changing slice styles
# Pie slice colors cycle through the `colors` and `opacities` attribute, as the `Lines` Mark.
pie1.stroke = 'brown'
pie1.colors = ['orange', 'darkviolet']
pie1.opacities = [.1, 1]
display(fig1)
# ## Representing an additional dimension using Color
# The `Pie` allows for its colors to be determined by data, that is passed to the `color` attribute.
# A `ColorScale` with the desired color scheme must also be passed.
# +
from bqplot import ColorScale, ColorAxis
Nslices = 7
size_data = np.random.rand(Nslices)
color_data = np.random.randn(Nslices)
sc = ColorScale(scheme='Reds')
# The ColorAxis gives a visual representation of its ColorScale
ax = ColorAxis(scale=sc)
pie2 = Pie(sizes=size_data, scales={'color': sc}, color=color_data)
Figure(marks=[pie2], axes=[ax])
# -
# ## Positioning the Pie using custom scales
# Pies can be positioned, via the `x` and `y` attributes,
# using either absolute figure scales or custom 'x' or 'y' scales
# +
from datetime import datetime
from bqplot.traits import convert_to_date
from bqplot import DateScale, LinearScale, Axis
avg_precipitation_days = [(d/30., 1-d/30.) for d in [2, 3, 4, 6, 12, 17, 23, 22, 15, 4, 1, 1]]
temperatures = [9, 12, 16, 20, 22, 23, 22, 22, 22, 20, 15, 11]
dates = [datetime(2010, k, 1) for k in range(1, 13)]
sc_x = DateScale()
sc_y = LinearScale()
ax_x = Axis(scale=sc_x, label='month', tick_format='%B')
ax_y = Axis(scale=sc_y, orientation='vertical', label='average temperature')
pies = [Pie(sizes=precipit, x=date, y=temp,
scales={"x": sc_x, "y": sc_y}, radius=30., stroke='navy',
colors=['navy', 'navy'], opacities=[1, .1])
for precipit, date, temp in zip(avg_precipitation_days, dates, temperatures)]
Figure(title='Kathmandu precipitation', marks=pies, axes=[ax_x, ax_y],
padding_x=.05, padding_y=.1)
| examples/Pie.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SARS strains and proteins co-occurence
# ## Protocol
#
# Goal of this notebook is analysis co-occurence of different types, stains of SARS and SARS-related proteins and genes
#
# We are using fantastic labeled dataset by SiBiteLabs:
# https://github.com/SciBiteLabs/CORD19
#
# ### To do:
# 1. Buld dataframe from json-files. Output: data frame with columns ['paper_id','block_id','entity_type_1','..','entity_type_n']. Each row matchs a block. Block is a part of paper's abstract or full_text
# 2. Group SARS strains. Covid-19 is a main group. Output: lists of strains fo each grooup
# 3. Filter papers related to any of SARS group and build dataframe. Output: dataframe with columns ['paper_id','title','authors','year','SARS strain','CVPROT']. Each row matchs paper.
# 4. Plot piechart for strains grous: percent of papers (not blocks) mentioned each strains group. Histogram for strains: top 5 most mentioned strains across the groups.
# 5. Calculate and plot co-ocurence matrix for strains group and proteins (CVPROT). Calcultae andand plot co-ocurence matrix for top 5 most mentioned strains across and proteins (CVPROT). Use data from the block level(not paper)
# 6. Plot histogram: number of CVPROT mentioned at block level for SARS in general
# +
import pandas as pd
import matplotlib.pyplot as plt
import re
from tqdm import tqdm
import os
import json
import numpy as np
import collections
# -
# ## Helper functions
# +
def load_files(dirname):
filenames = os.listdir(dirname)
raw_files = []
for filename in tqdm(filenames):
filename = dirname + filename
file = json.load(open(filename, 'rb'))
raw_files.append(file)
return raw_files
def get_all_files(dirname):
all_files = []
filenames = os.listdir(dirname)
for filename in tqdm(filenames):
filename = dirname + filename
file = json.load(open(filename, 'rb'))
all_files.append(file)
return all_files
def get_cat_vocab(cat):
df_cat = df[cat]
items = df_cat.dropna().tolist()
vocab_list = []
for element in items:
item = element.split(",")
for e in item:
vocab_list.append(e)
c = collections.Counter()
for word in vocab_list:
c[word] += 1
result_dic = dict(c)
return result_dic
# -
# ## 1. Buld dataframe from json-files
# +
#load all files
dirs = [
'data/scibitelabs/biorxiv_medrxiv/biorxiv_medrxiv/',
'data/scibitelabs/comm_use_subset/comm_use_subset/',
'data/scibitelabs/custom_license/custom_license/',
'data/scibitelabs/noncomm_use_subset/noncomm_use_subset/'
]
files_stack = []
for dir_ in dirs:
files = get_all_files(dir_)
files_stack.append(files)
# +
#build list of entities types
c = collections.Counter()
cat_vocab = []
for files in tqdm(files_stack):
for file in files:
for block in file['body_text']:
dict_file = block['termite_hits'].keys()
for key in dict_file:
cat_vocab.append(key)
for word in cat_vocab:
c[word] += 1
vocab_list = (set(list(c.elements())))
# -
vocab_list = sorted(list(vocab_list))
print(vocab_list)
# +
#build dataframe: entity mentions by blocks ignoring hint count
features = []
for files in tqdm(files_stack):
for file in files:
paper_id = file['paper_id']
i = 0
sections = ['abstract', 'body_text']
for section in sections:
for block in file[section]:
block_id = section + '_' + str(i)
block_features = []
block_features.append(paper_id)
block_features.append(block_id)
termite_hits = block['termite_hits']
block_categories = termite_hits.keys()
block_categories = list(block_categories)
for cat in vocab_list:
if cat in block_categories:
cat_entities = []
for hit in termite_hits[cat]:
entity = hit.get('name')
if entity not in cat_entities:
cat_entities.append(entity)
cat_entities = ",".join(cat_entities)
else:
cat_entities = None
block_features.append(cat_entities)
features.append(block_features)
i += 1
col_names = ['paper_id', 'block_id']
for cat in vocab_list:
col_names.append(cat)
df = pd.DataFrame(features, columns=col_names)
df.head()
# -
#save data
df.to_csv('data/data_ner.csv')
# ## 2. Group SARS strains
def get_cat_vocab(cat):
df_cat = df[cat]
items = df_cat.dropna().tolist()
vocab_list = []
for element in items:
item = element.split(",")
for e in item:
vocab_list.append(e)
c = collections.Counter()
for word in vocab_list:
c[word] += 1
result_dic = dict(c)
return result_dic
vocab_sars = get_cat_vocab('SARSCOV')
vocab_sars
# +
sars = ['SARS coronavirus']
covid_19 = ['Severe acute respiratory syndrome coronavirus 2']
sars_sin_strains = [
'SARS coronavirus Sin2748',
'SARS coronavirus Sin2774',
'SARS coronavirus Sin3725V',
'SARS coronavirus Sin0409',
'SARS coronavirus Sin_WNV',
'SARS coronavirus Sin2500',
'SARS coronavirus Sin2677',
'SARS coronavirus Sin2679',
'SARS coronavirus Sin846',
'SARS coronavirus Sin847',
'SARS coronavirus Sin842',
'SARS coronavirus Sin845',
'SARS coronavirus Sin852',
'SARS coronavirus Sin848',
'SARS coronavirus Sin850',
'SARS coronavirus Sin849',
'SARS coronavirus Sin3408',
'SARS coronavirus SinP2',
'SARS coronavirus Sin3408L',
'SARS coronavirus SinP5',
'SARS coronavirus SinP3',
'SARS coronavirus SinP4',
]
sars_betacov_strains = [
'BtRf-BetaCoV/JL2012',
'BtRf-BetaCoV/SX2013',
'BtRf-BetaCoV/HeB2013',
]
sars_tw_strains = [
'SARS coronavirus TW1',
'SARS coronavirus TW2',
'SARS coronavirus TW4',
'SARS coronavirus TW5',
'SARS coronavirus TW10',
'SARS coronavirus TWC2',
'SARS coronavirus TWC3',
'SARS coronavirus TW9',
'SARS coronavirus TW8',
'SARS coronavirus TW7',
'SARS coronavirus TW6',
'SARS coronavirus TW3',
'SARS coronavirus TW4',
]
sars_shanghai_strains = [
'SARS coronavirus ShanghaiQXC1',
'SARS coronavirus ShanghaiQXC2',
]
sars_gz_strains = [
'SARS coronavirus GZ02',
'SARS coronavirus GZ-C',
'SARS coronavirus GZ-B',
'SARS coronavirus GZ50',
'SARS coronavirus GZ0402',
]
sars_bj_stains = [
'SARS coronavirus BJ04',
'SARS coronavirus BJ302',
'SARS coronavirus BJ01',
'SARS coronavirus BJ182-12',
'SARS coronavirus BJ02',
'SARS coronavirus BJ03',
'SARS coronavirus BJ202',
]
sars_lc_stains = [
'SARS coronavirus LC3',
'SARS coronavirus LC2',
'SARS coronavirus LC5',
'SARS coronavirus LC1',
]
sars_other_strains = [
'SARS coronavirus NS-1',
'SARS coronavirus Tor2',
'Bat SARS-like coronavirus',
'SARS coronavirus Urbani',
'SARS coronavirus CUHK-W1',
'SARS coronavirus MA15',
'SARS coronavirus ZS-C',
'SARS coronavirus Sino1-11',
'SARS coronavirus HSR 1',
'Bat SARS-like coronavirus WIV1',
'SARS coronavirus ZJ01',
'SARS coronavirus Frankfurt 1',
'SARS coronavirus HC/SZ/61/03',
'SARS coronavirus AS',
'SARS coronavirus GD03T0013',
'SARS coronavirus GD01': 71,
'SARS coronavirus HKU-39849',
'SARS coronavirus CUHK-AG01',
'SARS coronavirus HZS2-Fb',
'SARS coronavirus PUMC01',
'SARS coronavirus B012',
'SARS coronavirus ExoN1',
'SARS coronavirus C025',
'SARS coronavirus PUMC03',
'SARS coronavirus wtic-MB',
'SARS coronavirus HSZ-Cb',
'SARS coronavirus A022',
'SARS coronavirus SZ1',
'SARS coronavirus WH20',
'SARS coronavirus SoD',
'SARS coronavirus BJ01': 174,
'SARS coronavirus LC4': 5,
'SARS coronavirus JMD': 1,
'SARS coronavirus ES191': 1,
'SARS coronavirus PUMC02': 7,
'SARS coronavirus Taiwan TC2': 1,
'SARS coronavirus Taiwan TC3': 1,
'SARS coronavirus Taiwan TC1': 3,
'SARS coronavirus ZMY 1': 15,
'SARS coronavirus GZ43': 17,
'SARS coronavirus SZ13': 7,
'SARS coronavirus CUHK-L2': 1,
'SARS coronavirus HSZ-A': 1,
'SARS coronavirus HKU-65806': 1,
'SARS coronavirus ZS-B': 4,
'SARS coronavirus GD69': 6,
'SARS coronavirus TW11': 5,
'SARS coronavirus HGZ8L1-A': 4,
'SARS coronavirus Sino3-11': 4,
'SARS coronavirus CUHK-AG02': 5,
'SARS coronavirus CUHK-AG03': 5,
'SARS coronavirus LLJ-2004': 2,
'SARS coronavirus GZ60': 11,
'SARS coronavirus Rs_672/2006': 1,
'SARS coronavirus GZ0401': 2,
'SARS coronavirus HZS2-C': 3,
'SARS coronavirus HZS2-Fc': 3,
'SARS coronavirus GZ-A': 1,
'SARS Coronavirus CDC#200301157': 2,
'SARS coronavirus A030': 2,
'SARS coronavirus A013': 1,
'SARS coronavirus B039': 1,
'SARS coronavirus PC4-227': 3,
'SARS coronavirus PC4-136': 2,
'SARS coronavirus civet020': 2,
'SARS coronavirus civet010': 1,
'SARS coronavirus PC4-13': 2,
'SARS coronavirus HSZ-Bc': 3,
'SARS coronavirus HSZ-Bb': 3,
'SARS coronavirus HSZ-Cc': 3,
'SARS coronavirus C028': 2,
'SARS coronavirus A001': 1,
'SARS coronavirus B024': 1,
'SARS coronavirus civet014': 1,
'SARS coronavirus ZS-A': 4,
'SARS coronavirus TWC': 1,
'SARS coronavirus PC4-115': 1,
'SARS coronavirus HSZ2-A': 2,
'SARS coronavirus HGZ8L2': 2,
'SARS coronavirus HZS2-E': 2,
'SARS coronavirus HZS2-D': 2
]
# -
vocab_cvprot = get_cat_vocab('CVPROT')
vocab_cvprot
vocab_genes = get_cat_vocab('GENE')
vocab_genes
vocab_hpo = get_cat_vocab('HPO')
vocab_hpo
| antigen_scoring/notebooks/SARS_proteins_co-occurence.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: privacy-evaluator-venv
# language: python
# name: privacy-evaluator-venv
# ---
# +
import numpy as np
import privacy_evaluator.models.torch.dcti.dcti as torch_dcti
from privacy_evaluator.datasets.cifar10 import CIFAR10
from privacy_evaluator.classifiers.classifier import Classifier
from privacy_evaluator.attacks.membership_inference.black_box import MembershipInferenceBlackBoxAttack
from privacy_evaluator.attacks.membership_inference.black_box_rule_based import MembershipInferenceBlackBoxRuleBasedAttack
from privacy_evaluator.attacks.membership_inference.label_only_decision_boundary import MembershipInferenceLabelOnlyDecisionBoundaryAttack
# -
# # Membership Inference Attack Examples
# ### Prepare datasets and Pytorch target model
# +
# CIFAR datasets
x_train, y_train, x_test, y_test = CIFAR10.numpy(model_type='torch')
# Wrapped Pytorch target model
target_model = Classifier(torch_dcti.load_dcti(), nb_classes=CIFAR10.N_CLASSES, input_shape=CIFAR10.INPUT_SHAPE)
# -
# ## Membership Inference Black Box Attack
# ### Prepare, fit and attack
attack = MembershipInferenceBlackBoxAttack(target_model, x_train[:100], y_train[:100], x_test[:100], y_test[:100])
attack.fit()
attack.attack(x_train[:100], y_train[:100])
# ### Create output
attack.attack_output(x_train[:100], y_train[:100], np.ones((100,)))
# ## Membership Inference Black Box Rule Based Attack
# ### Prepare and attack
attack = MembershipInferenceBlackBoxRuleBasedAttack(target_model, x_train, y_train, x_test, y_test)
attack.attack(x_train, y_train)
# ### Create output
attack.attack_output(x_train, y_train, np.ones((len(y_train),)))
# ## Membership Inference Label Only Decision Boundary Attack
# ### Prepare, fit and attack
attack = MembershipInferenceLabelOnlyDecisionBoundaryAttack(target_model, x_train[:1], y_train[:1], x_test[:1], y_test[:1])
attack.fit(max_iter=1, max_eval=1, init_eval=1)
attack.attack(x_train[:1], y_train[:1])
# ### Create output
attack.attack_output(x_train[:1], y_train[:1], np.ones((1,)))
| notebooks/membership_inference_attack.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.7 64-bit
# name: python3
# ---
from femda import FEMDA
# # 1. Using FEMDA in a sklearn.pipeline
# +
from sklearn.datasets import load_iris, load_digits
from sklearn.pipeline import make_pipeline
from sklearn.model_selection import train_test_split
from sklearn.decomposition import PCA
X, y = load_digits(return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
# -
pipe = make_pipeline(PCA(n_components=5), FEMDA())
pipe.fit(X_train, y_train)
# +
print(pipe.score(X_test, y_test))
y_pred = pipe.predict(X_test)
#from sklearn.metrics import classification_report
#print(classification_report(y_test, y_pred))
# -
# # 2. Testing FEMDA on other datasets
import femda.experiments.preprocessing as pre
from femda._models_lda import LDA, QDA
from femda._models_t_lda import t_QDA
X_train, y_train, X_test, y_test = pre.ionosphere(r"data\Paper\\")
#Choose between ionosphere, statlog, ecoli, breast_cancer, spam_base
print("FEMDA", FEMDA().fit(X_train, y_train).score(X_test, y_test))
print("t_QDA", t_QDA().fit(X_train, y_train).score(X_test, y_test))
print("LDA", LDA() .fit(X_train, y_train).score(X_test, y_test))
print("QDA", QDA() .fit(X_train, y_train).score(X_test, y_test))
# # 3. Running experiments presented in the paper
from femda.experiments import run_experiments
run_experiments()
# # Check estimator
from sklearn.utils.estimator_checks import check_estimator
f = FEMDA()
print(check_estimator(f) == None)
| demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: conda-env-py37_pytorch-py
# kernelspec:
# display_name: py37_pytorch
# language: python
# name: conda-env-py37_pytorch-py
# ---
# %matplotlib inline
# # Datasets and Dataloaders
#
# Code for processing data samples can get messy and hard to maintain; we ideally want our dataset code
# to be decoupled from our model training code for better readability and modularity.
# PyTorch provides two data primitives: ``torch.utils.data.DataLoader`` and ``torch.utils.data.Dataset``
# that allow you to use pre-loaded datasets as well as your own data.
# ``Dataset`` stores the samples and their corresponding labels, and ``DataLoader`` wraps an iterable around
# the ``Dataset`` to enable easy access to the samples.
#
# PyTorch domain libraries provide a number of pre-loaded datasets (such as FashionMNIST) that
# subclass ``torch.utils.data.Dataset`` and implement functions specific to the particular data.
# They can be used to prototype and benchmark your model. You can find them
# here: [Image Datasets](https://pytorch.org/vision/stable/datasets.html),
# [Text Datasets](https://pytorch.org/text/stable/datasets.html), and
# [Audio Datasets](https://pytorch.org/audio/stable/datasets.html)
#
# ## Loading a dataset
#
# Here is an example of how to load the [Fashion-MNIST](https://research.zalando.com/welcome/mission/research-projects/fashion-mnist/) dataset from TorchVision.
# Fashion-MNIST is a dataset of Zalando’s article images consisting of of 60,000 training examples and 10,000 test examples.
# Each example comprises a 28×28 grayscale image and an associated label from one of 10 classes.
#
# We load the [FashionMNIST Dataset](https://pytorch.org/vision/stable/datasets.html#fashion-mnist) with the following parameters:
# - `root` is the path where the train/test data is stored,
# - `train` specifies training or test dataset,
# - `download=True` downloads the data from the Internet if it's not available at `root`.
# - `transform` and `target_transform` specify the feature and label transformations
#
#
# +
import torch
from torch.utils.data import Dataset
from torchvision import datasets
from torchvision.transforms import ToTensor, Lambda
import matplotlib.pyplot as plt
training_data = datasets.FashionMNIST(
root="data",
train=True,
download=True,
transform=ToTensor()
)
test_data = datasets.FashionMNIST(
root="data",
train=False,
download=True,
transform=ToTensor()
)
# -
# Iterating and Visualizing the Dataset
# -----------------
#
# We can index ``Datasets`` manually like a list: ``training_data[index]``.
# We use ``matplotlib`` to visualize some samples in our training data.
#
#
labels_map = {
0: "T-Shirt",
1: "Trouser",
2: "Pullover",
3: "Dress",
4: "Coat",
5: "Sandal",
6: "Shirt",
7: "Sneaker",
8: "Bag",
9: "Ankle Boot",
}
figure = plt.figure(figsize=(8, 8))
cols, rows = 3, 3
for i in range(1, cols * rows + 1):
sample_idx = torch.randint(len(training_data), size=(1,)).item()
img, label = training_data[sample_idx]
figure.add_subplot(rows, cols, i)
plt.title(labels_map[label])
plt.axis("off")
plt.imshow(img.squeeze(), cmap="gray")
plt.show()
# Creating a Custom Dataset for your files
# ---------------------------------------------------
#
# A custom Dataset class must implement three functions: `__init__`, `__len__`, and `__getitem__`.
# Take a look at this implementation; the FashionMNIST images are stored
# in a directory ``img_dir``, and their labels are stored separately in a CSV file ``annotations_file``.
#
# In the next sections, we'll break down what's happening in each of these functions.
#
#
# +
import os
import pandas as pd
import torchvision.io as tvio
class CustomImageDataset(Dataset):
def __init__(self, annotations_file, img_dir, transform=None, target_transform=None):
self.img_labels = pd.read_csv(annotations_file)
self.img_dir = img_dir
self.transform = transform
self.target_transform = target_transform
def __len__(self):
return len(self.img_labels)
def __getitem__(self, idx):
img_path = os.path.join(self.img_dir, self.img_labels.iloc[idx, 0])
image = tvio.read_image(img_path)
label = self.img_labels.iloc[idx, 1]
if self.transform:
image = self.transform(image)
if self.target_transform:
label = self.target_transform(label)
sample = {"image": image, "label": label}
return sample
# -
# ## init
#
# The `__init__` function is run once when instantiating the Dataset object. We initialize
# the directory containing the images, the annotations file, and both transforms (covered
# in more detail in the next section).
#
# The labels.csv file looks like:
# ```
#
# tshirt1.jpg, 0
# tshirt2.jpg, 0
# ......
# ankleboot999.jpg, 9
# ```
# Example:
# ```
# def __init__(self, annotations_file, img_dir, transform=None, target_transform=None):
# self.img_labels = pd.read_csv(annotations_file)
# self.img_dir = img_dir
# self.transform = transform
# self.target_transform = target_transform
# ```
# ## len
#
# The `__len__` function returns the number of samples in our dataset.
#
# Example:
# ```
# def __len__(self):
# return len(self.img_labels)
# ```
#
# ## getitem
#
# The `__getitem__` function loads and returns a sample from the dataset at the given index `idx`.
# Based on the index, it identifies the image's location on disk, converts that to a tensor using `read_image`, retrieves the
# corresponding label from the csv data in `self.img_labels`, calls the transform functions on them (if applicable), and returns the
# tensor image and corresponding label in a Python `dict`.
#
# Example:
# ```
# def __getitem__(self, idx):
# img_path = os.path.join(self.img_dir, self.img_labels.iloc[idx, 0])
# image = read_image(img_path)
# label = self.img_labels.iloc[idx, 1]
# if self.transform:
# image = self.transform(image)
# if self.target_transform:
# label = self.target_transform(label)
# sample = {"image": image, "label": label}
# return sample
# ```
# Preparing your data for training with DataLoaders
# -------------------------------------------------
# The ``Dataset`` retrieves our dataset's features and labels one sample at a time. While training a model, we typically want to
# pass samples in "minibatches", reshuffle the data at every epoch to reduce model overfitting, and use Python's ``multiprocessing`` to
# speed up data retrieval.
#
# ``DataLoader`` is an iterable that abstracts this complexity for us in an easy API.
#
#
# +
from torch.utils.data import DataLoader
train_dataloader = DataLoader(training_data, batch_size=64, shuffle=True)
test_dataloader = DataLoader(test_data, batch_size=64, shuffle=True)
# -
# ## Iterate through the DataLoader
#
# We have loaded that dataset into the `Dataloader` and can iterate through the dataset as needed.
# Each iteration below returns a batch of `train_features` and `train_labels`(containing `batch_size=64` features and labels respectively). Because we specified `shuffle=True`, after we iterate over all batches the data is shuffled (for finer-grained control over the data loading order, take a look at [Samplers](https://pytorch.org/docs/stable/data.html#data-loading-order-and-sampler>).
#
#
# Display image and label.
train_features, train_labels = next(iter(train_dataloader))
print(f"Feature batch shape: {train_features.size()}")
print(f"Labels batch shape: {train_labels.size()}")
img = train_features[0].squeeze()
label = train_labels[0]
plt.imshow(img, cmap="gray")
plt.show()
print(f"Label: {label}")
| intro-to-pytorch/3-data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 (tensorflow)
# language: python
# name: tensorflow
# ---
# + [markdown] colab_type="text" id="V8-yl-s-WKMG"
# # Object Detection Demo
# Welcome to the object detection inference walkthrough! This notebook will walk you step by step through the process of using a pre-trained model to detect objects in an image. Make sure to follow the [installation instructions](https://github.com/tensorflow/models/blob/master/research/object_detection/g3doc/installation.md) before you start.
# + [markdown] colab_type="text" id="kFSqkTCdWKMI"
# # Imports
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="hV4P5gyTWKMI"
import numpy as np
import os
import six.moves.urllib as urllib
import sys
import tarfile
import tensorflow as tf
import zipfile
from distutils.version import StrictVersion
from collections import defaultdict
from io import StringIO
from matplotlib import pyplot as plt
from PIL import Image
from imageai.Detection import VideoObjectDetection
import cv2
cap = cv2.VideoCapture(0)
execution_path = os.getcwd()
# This is needed since the notebook is stored in the object_detection folder.
sys.path.append("..")
from object_detection.utils import ops as utils_ops
if StrictVersion(tf.__version__) < StrictVersion('1.12.0'):
raise ImportError('Please upgrade your TensorFlow installation to v1.12.*.')
# + [markdown] colab_type="text" id="r5FNuiRPWKMN"
# ## Object detection imports
# Here are the imports from the object detection module.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="bm0_uNRnWKMN"
from utils import label_map_util
from utils import visualization_utils as vis_util
# + [markdown] colab_type="text" id="cfn_tRFOWKMO"
# # Model preparation
# + [markdown] colab_type="text" id="X_sEBLpVWKMQ"
# ## Variables
#
# Any model exported using the `export_inference_graph.py` tool can be loaded here simply by changing `PATH_TO_FROZEN_GRAPH` to point to a new .pb file.
#
# By default we use an "SSD with Mobilenet" model here. See the [detection model zoo](https://github.com/tensorflow/models/blob/master/research/object_detection/g3doc/detection_model_zoo.md) for a list of other models that can be run out-of-the-box with varying speeds and accuracies.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="VyPz_t8WWKMQ"
# What model to download.
MODEL_NAME = 'ssd_mobilenet_v1_coco_2017_11_17'
MODEL_FILE = MODEL_NAME + '.tar.gz'
DOWNLOAD_BASE = 'http://download.tensorflow.org/models/object_detection/'
# Path to frozen detection graph. This is the actual model that is used for the object detection.
PATH_TO_FROZEN_GRAPH = MODEL_NAME + '/frozen_inference_graph.pb'
# List of the strings that is used to add correct label for each box.
PATH_TO_LABELS = os.path.join('data', 'mscoco_label_map.pbtxt')
# + [markdown] colab_type="text" id="7ai8pLZZWKMS"
# ## Download Model
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="KILYnwR5WKMS"
opener = urllib.request.URLopener()
opener.retrieve(DOWNLOAD_BASE + MODEL_FILE, MODEL_FILE)
tar_file = tarfile.open(MODEL_FILE)
for file in tar_file.getmembers():
file_name = os.path.basename(file.name)
if 'frozen_inference_graph.pb' in file_name:
tar_file.extract(file, os.getcwd())
# + [markdown] colab_type="text" id="YBcB9QHLWKMU"
# ## Load a (frozen) Tensorflow model into memory.
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="KezjCRVvWKMV"
detection_graph = tf.Graph()
with detection_graph.as_default():
od_graph_def = tf.GraphDef()
with tf.gfile.GFile(PATH_TO_FROZEN_GRAPH, 'rb') as fid:
serialized_graph = fid.read()
od_graph_def.ParseFromString(serialized_graph)
tf.import_graph_def(od_graph_def, name='')
# + [markdown] colab_type="text" id="_1MVVTcLWKMW"
# ## Loading label map
# Label maps map indices to category names, so that when our convolution network predicts `5`, we know that this corresponds to `airplane`. Here we use internal utility functions, but anything that returns a dictionary mapping integers to appropriate string labels would be fine
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="hDbpHkiWWKMX"
category_index = label_map_util.create_category_index_from_labelmap(PATH_TO_LABELS, use_display_name=True)
# + [markdown] colab_type="text" id="EFsoUHvbWKMZ"
# ## Helper code
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="aSlYc3JkWKMa"
# + [markdown] colab_type="text" id="H0_1AGhrWKMc"
# # Detection
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="jG-zn5ykWKMd"
# Detection
with detection_graph.as_default():
with tf.Session(graph=detection_graph) as sess:
while True:
# Read frame from camera
ret, image_np = cap.read()
# Expand dimensions since the model expects images to have shape: [1, None, None, 3]
image_np_expanded = np.expand_dims(image_np, axis=0)
# Extract image tensor
image_tensor = detection_graph.get_tensor_by_name('image_tensor:0')
# Extract detection boxes
boxes = detection_graph.get_tensor_by_name('detection_boxes:0')
# Extract detection scores
scores = detection_graph.get_tensor_by_name('detection_scores:0')
# Extract detection classes
classes = detection_graph.get_tensor_by_name('detection_classes:0')
# Extract number of detectionsd
num_detections = detection_graph.get_tensor_by_name(
'num_detections:0')
# Actual detection.
(boxes, scores, classes, num_detections) = sess.run(
[boxes, scores, classes, num_detections],
feed_dict={image_tensor: image_np_expanded})
# Visualization of the results of a detection.
vis_util.visualize_boxes_and_labels_on_image_array(
image_np,
np.squeeze(boxes),
np.squeeze(classes).astype(np.int32),
np.squeeze(scores),
category_index,
use_normalized_coordinates=True,
min_score_thresh=.7,
line_thickness=8)
# Display output
cv2.imshow('object detection', cv2.resize(image_np, (800, 600)))
#print(num_detections)
if cv2.waitKey(25) & 0xFF == ord('q'):
cap.release()
cv2.destroyAllWindows()
break
min_score_thresh = 0.5
print(i in classes[0])
# -
| object_detection_tutorial - Copy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Qcodes example with Ithaco
# +
# # %matplotlib nbagg
# %gui qt
import matplotlib.pyplot as plt
import time
import numpy as np
import qcodes as qc
from qcodes.utils.validators import Enum, Strings
import qcodes.instrument_drivers.tektronix.Keithley_2600 as keith
import qcodes.instrument_drivers.agilent.Agilent_34400A as agi
import qcodes.instrument_drivers.ithaco.Ithaco_1211 as ithaco
from qcodes.instrument.parameter import Parameter, StandardParameter
# -
import time
class Timer(object):
def __init__(self, name=None):
self.name = name
def __enter__(self):
self.tstart = time.time()
def __exit__(self, type, value, traceback):
if self.name:
print('[%s]' % self.name,)
print('Elapsed: %s' % (time.time() - self.tstart))
# +
# create Instruments
k1 = keith.Keithley_2600('Keithley1', 'GPIB0::15::INSTR',channel='a')
k2 = keith.Keithley_2600('Keithley2', 'GPIB0::15::INSTR',channel='b')
a1 = agi.Agilent_34400A('Agilent1', 'GPIB0::11::INSTR')
a2 = agi.Agilent_34400A('Agilent2', 'GPIB0::6::INSTR')
camp = ithaco.Ithaco_1211('camp1')
camp.sens.set(1e-4)
curr = ithaco.CurrentParameter(a2.volt, camp)
# set integration time (number of line cycles)
a1.NPLC.set(1)
a2.NPLC.set(1)
# -
camp.snapshot()
a2.volt.units
curr.units
data = qc.Loop(k1.volt[-5:5:1], 0).each(curr).run(location='testsweep', overwrite=True)
plotQ = qc.QtPlot(data.current,windowTitle='YEAH')
plotQ
data.sync()
data.arrays
station1 = qc.Station(a1,a2)
station1.set_measurement(a1.volt)
station2 = qc.Station(a1,a2)
station2.set_measurement(a1.volt, a2.volt)
# Time single readings
with Timer('Time s1'):
station1.measure()
with Timer('Time s2'):
station2.measure()
# Time single readings
with Timer('Time a1'):
a1.volt.get()
with Timer('Time a2'):
a2.volt.get()
# +
with Timer('Time Loop 1'):
data = qc.Loop(k1.volt[-5:5:1], 0).each(a1.volt).run(location='testsweep', overwrite=True,background=False)
with Timer('Time Loop 2'):
data = qc.Loop(k1.volt[-5:5:1], 0).each(a1.volt, a2.volt).run(location='testsweep', overwrite=True,background=False)
# -
with Timer('Time Loop 1'):
data = qc.Loop(k1.volt[-5:5:1], 0).each(a1.volt).run(location='testsweep', overwrite=True)
while data.sync():
time.sleep(0.1)
with Timer('Time Loop 2'):
data = qc.Loop(k1.volt[-5:5:1], 0).each(a1.volt, a2.volt).run(location='testsweep', overwrite=True)
while data.sync():
time.sleep(0.1)
| docs/examples/driver_examples/Qcodes example with Ithaco.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Q1
ft= input()
ft=int(ft)
if ft<=1000:
print("Safe to land")
elif ft<=5000:
print("Bring down to 1000ft")
else:
print("Turn around")
# Q2
for i in range(1,201):
count=0
for j in range(1,i+1):
if(i%j==0):
count=count+1
if (count==2):
print(i)
| Assignment of Day 3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # utils
# +
import vectorbt as vbt
from vectorbt.utils import checks, config, decorators
# + Collapsed="false"
import numpy as np
import pandas as pd
from numba import njit
# +
v1 = 0
a1 = np.array([1])
a2 = np.array([1, 2, 3])
a3 = np.array([[1, 2, 3]])
a4 = np.array([[1], [2], [3]])
a5 = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
sr1 = pd.Series([1], index=pd.Index(['x1'], name='i1'))
print(sr1)
sr2 = pd.Series([1, 2, 3], index=pd.Index(['x2', 'y2', 'z2'], name='i2'))
print(sr2)
df1 = pd.DataFrame(
[[1]],
index=pd.Index(['x3'], name='i3'),
columns=pd.Index(['a3'], name='c3'))
print(df1)
df2 = pd.DataFrame(
[[1], [2], [3]],
index=pd.Index(['x4', 'y4', 'z4'], name='i4'),
columns=pd.Index(['a4'], name='c4'))
print(df2)
df3 = pd.DataFrame(
[[1, 2, 3]],
index=pd.Index(['x5'], name='i5'),
columns=pd.Index(['a5', 'b5', 'c5'], name='c5'))
print(df3)
df4 = pd.DataFrame(
[[1, 2, 3], [4, 5, 6], [7, 8, 9]],
index=pd.Index(['x6', 'y6', 'z6'], name='i6'),
columns=pd.Index(['a6', 'b6', 'c6'], name='c6'))
print(df4)
multi_i = pd.MultiIndex.from_arrays([['x7', 'y7', 'z7'], ['x8', 'y8', 'z8']], names=['i7', 'i8'])
multi_c = pd.MultiIndex.from_arrays([['a7', 'b7', 'c7'], ['a8', 'b8', 'c8']], names=['c7', 'c8'])
df5 = pd.DataFrame([[1, 2, 3], [4, 5, 6], [7, 8, 9]], index=multi_i, columns=multi_c)
print(df5)
# -
# ## config
# +
conf = config.Config({'a': 0, 'b': {'c': 1}}, frozen=False)
conf['b']['d'] = 2
conf = config.Config({'a': 0, 'b': {'c': 1}}, frozen=True)
conf['a'] = 2
try:
conf['d'] = 2
except Exception as e:
print(e)
try:
conf.update(d=2)
except Exception as e:
print(e)
conf.update(d=2, force_update=True)
# +
conf = config.Config({'a': 0, 'b': {'c': 1}}, read_only=True)
try:
conf['a'] = 2
except Exception as e:
print(e)
try:
del conf['a']
except Exception as e:
print(e)
try:
conf.pop('a')
except Exception as e:
print(e)
try:
conf.popitem()
except Exception as e:
print(e)
try:
conf.clear()
except Exception as e:
print(e)
try:
conf.update(a=2)
except Exception as e:
print(e)
print(conf.merge_with(dict(b=dict(d=2))))
# -
print(config.merge_dicts({'a': 1}, {'b': 2}))
print(config.merge_dicts({'a': 1}, {'a': 2}))
print(config.merge_dicts({'a': {'b': 2}}, {'a': {'c': 3}}))
print(config.merge_dicts({'a': {'b': 2}}, {'a': {'b': 3}}))
# +
class H(config.Configured):
def __init__(self, a, b=2, **kwargs):
super().__init__(a=a, b=b, **kwargs)
print(H(1).config)
print(H(1).copy(b=3).config)
print(H(1).copy(c=4).config)
# -
# ## decorators
# +
class G():
@decorators.class_or_instancemethod
def g(self_or_cls):
if isinstance(self_or_cls, type):
print("class")
else:
print("instance")
G.g()
G().g()
# +
class G():
@decorators.cached_property(hello="world", hello2="world2")
def cache_me(self): return np.random.uniform(size=(10000, 10000))
G.cache_me.kwargs
# -
g = G()
# %time _ = g.cache_me
# %time _ = g.cache_me
dir(g)
G.cache_me.clear_cache(g)
# %time _ = g.cache_me
# %time _ = g.cache_me
G.cache_me.clear_cache(g)
G.cache_me.disabled = True
# %time _ = g.cache_me
# %time _ = g.cache_me
G.cache_me.disabled = False
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append((g, 'cache_me'))
# %time _ = g.cache_me
# %time _ = g.cache_me
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append('cache_me')
# %time _ = g.cache_me
# %time _ = g.cache_me
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append(g)
# %time _ = g.cache_me
# %time _ = g.cache_me
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append(G)
# %time _ = g.cache_me
# %time _ = g.cache_me
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append('G')
# %time _ = g.cache_me
# %time _ = g.cache_me
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append('g')
# %time _ = g.cache_me
# %time _ = g.cache_me
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append({'hello': 'world'})
# %time _ = g.cache_me
# %time _ = g.cache_me
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append({'hello': 'world', 'hello2': 'world2'})
# %time _ = g.cache_me
# %time _ = g.cache_me
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append({'hello': 'world', 'hello2': 'world2', 'hello3': 'world3'})
# %time _ = g.cache_me
# %time _ = g.cache_me
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['enabled'] = False
# %time _ = g.cache_me
# %time _ = g.cache_me
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['enabled'] = False
vbt.settings.caching['whitelist'].append((g, 'cache_me'))
# %time _ = g.cache_me
# %time _ = g.cache_me
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['enabled'] = False
vbt.settings.caching['whitelist'].append('cache_me')
# %time _ = g.cache_me
# %time _ = g.cache_me
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['enabled'] = False
vbt.settings.caching['whitelist'].append(g)
# %time _ = g.cache_me
# %time _ = g.cache_me
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['enabled'] = False
vbt.settings.caching['whitelist'].append(G)
# %time _ = g.cache_me
# %time _ = g.cache_me
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['enabled'] = False
vbt.settings.caching['whitelist'].append('G')
# %time _ = g.cache_me
# %time _ = g.cache_me
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['enabled'] = False
vbt.settings.caching['whitelist'].append({'hello': 'world'})
# %time _ = g.cache_me
# %time _ = g.cache_me
vbt.settings.caching.reset()
# +
class G():
@decorators.cached_method(hello="world", hello2="world2")
def cache_me(self, a): return np.random.uniform(size=(10000, 10000)) * a
G.cache_me.kwargs
# -
g = G()
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
dir(g)
G.cache_me.clear_cache(g)
# %time _ = g.cache_me(2)
G.cache_me.clear_cache(g)
G.cache_me.disabled = True
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
G.cache_me.disabled = False
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append(g.cache_me)
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append((g, 'cache_me'))
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append('cache_me')
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append(g)
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append(G)
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append('G')
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append('g')
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append({'hello': 'world'})
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append({'hello': 'world', 'hello2': 'world2'})
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['blacklist'].append({'hello': 'world', 'hello2': 'world2', 'hello3': 'world3'})
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['enabled'] = False
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['enabled'] = False
vbt.settings.caching['whitelist'].append((g, 'cache_me'))
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['enabled'] = False
vbt.settings.caching['whitelist'].append('cache_me')
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['enabled'] = False
vbt.settings.caching['whitelist'].append(g)
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['enabled'] = False
vbt.settings.caching['whitelist'].append(G)
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['enabled'] = False
vbt.settings.caching['whitelist'].append('G')
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
G.cache_me.clear_cache(g)
vbt.settings.caching['enabled'] = False
vbt.settings.caching['whitelist'].append({'hello': 'world'})
# %time _ = g.cache_me(2)
# %time _ = g.cache_me(2)
vbt.settings.caching.reset()
# Non-hashable arguments won't cache
# %time _ = g.cache_me(np.asarray(2))
# %time _ = g.cache_me(np.asarray(2))
# +
class A:
@decorators.custom_property(some_key=0)
def a(self): pass
class B:
@decorators.cached_property(some_key=0, child_cls=A)
def a(self): pass
@decorators.custom_method(some_key=1)
def b(self): pass
class C:
@decorators.cached_method(some_key=0, child_cls=B)
def b(self): pass
@decorators.custom_property(some_key=1)
def c(self): pass
# -
str(decorators.traverse_attr_kwargs(C))
decorators.traverse_attr_kwargs(C, key='some_key')
decorators.traverse_attr_kwargs(C, key='some_key', value=1)
decorators.traverse_attr_kwargs(C, key='some_key', value=(0, 1))
# ## checks
print(checks.is_series(v1))
print(checks.is_series(a1))
print(checks.is_series(sr1))
print(checks.is_series(df1))
print(checks.is_frame(v1))
print(checks.is_frame(a1))
print(checks.is_frame(sr1))
print(checks.is_frame(df1))
print(checks.is_pandas(v1))
print(checks.is_pandas(a1))
print(checks.is_pandas(sr1))
print(checks.is_pandas(df1))
print(checks.is_array(v1))
print(checks.is_array(a1))
print(checks.is_array(sr1))
print(checks.is_array(df1))
print(checks.is_numba_func(lambda x: x))
print(checks.is_numba_func(njit(lambda x: x)))
print(checks.is_hashable(2))
print(checks.is_hashable(np.asarray(2)))
checks.assert_in(0, (0, 1))
checks.assert_numba_func(njit(lambda x: x))
checks.assert_not_none(v1)
checks.assert_type(v1, int)
checks.assert_type(a1, np.ndarray)
checks.assert_type(sr1, (np.ndarray, pd.Series))
checks.assert_type_equal(v1, v1)
checks.assert_type_equal(a1, a2)
checks.assert_type_equal(sr1, sr1)
checks.assert_type_equal(df1, df2)
checks.assert_dtype(a1, np.int)
checks.assert_dtype_equal(v1, a1)
checks.assert_dtype_equal(a1, df1)
checks.assert_dtype_equal(df1, df2)
checks.assert_dtype_equal(df2, df3)
checks.assert_ndim(v1, 0)
checks.assert_ndim(a1, 1)
checks.assert_ndim(df1, 2)
checks.assert_len_equal([[1]], [[2]])
checks.assert_shape_equal(a1, sr1)
checks.assert_shape_equal(df2, df4, axis=0)
checks.assert_shape_equal(df3, df4, axis=1)
checks.assert_shape_equal(df2, df3, axis=(0, 1))
checks.assert_index_equal(df3.index, df3.index)
checks.assert_meta_equal(df3, df3)
checks.assert_array_equal(df3, df3)
checks.assert_level_not_exists(df3.columns, 'a')
| tests/notebooks/utils.ipynb |