code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Q#
# language: qsharp
# name: iqsharp
# ---
# # Superposition Kata
#
# **Superposition** quantum kata is a series of exercises designed
# to get you familiar with the concept of superposition and with programming in Q#.
# It covers the following topics:
# * basic single-qubit and multi-qubit gates,
# * superposition,
# * flow control and recursion in Q#.
#
# It is recommended to complete the [BasicGates kata](./../BasicGates/BasicGates.ipynb) before this one to get familiar with the basic gates used in quantum computing. The list of basic gates available in Q# can be found at [Microsoft.Quantum.Intrinsic](https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic).
#
# Each task is wrapped in one operation preceded by the description of the task.
# Your goal is to fill in the blank (marked with `// ...` comments)
# with some Q# code that solves the task. To verify your answer, run the cell using Ctrl/⌘+Enter.
#
# The tasks are given in approximate order of increasing difficulty; harder ones are marked with asterisks.
# To begin, first prepare this notebook for execution (if you skip this step, you'll get "Syntax does not match any known patterns" error when you try to execute Q# code in the next cells):
%package Microsoft.Quantum.Katas::0.11.2003.3107
# > The package versions in the output of the cell above should always match. If you are running the Notebooks locally and the versions do not match, please install the IQ# version that matches the version of the `Microsoft.Quantum.Katas` package.
# > <details>
# > <summary><u>How to install the right IQ# version</u></summary>
# > For example, if the version of `Microsoft.Quantum.Katas` package above is 0.1.2.3, the installation steps are as follows:
# >
# > 1. Stop the kernel.
# > 2. Uninstall the existing version of IQ#:
# > dotnet tool uninstall microsoft.quantum.iqsharp -g
# > 3. Install the matching version:
# > dotnet tool install microsoft.quantum.iqsharp -g --version 0.1.2.3
# > 4. Reinstall the kernel:
# > dotnet iqsharp install
# > 5. Restart the Notebook.
# > </details>
#
# ### <a name="plus-state"></a> Task 1. Plus state.
#
# **Input:** A qubit in the $|0\rangle$ state.
#
# **Goal:** Change the state of the qubit to $|+\rangle = \frac{1}{\sqrt{2}} \big(|0\rangle + |1\rangle\big)$.
# +
%kata T01_PlusState_Test
operation PlusState (q : Qubit) : Unit {
// Hadamard gate H will convert |0⟩ state to |+⟩ state.
// Type the following: H(q);
// Then run the cell using Ctrl/⌘+Enter.
// ...
}
# -
# *Can't come up with a solution? See the explained solution in the [Superposition Workbook](./Workbook_Superposition.ipynb#plus-state).*
# ### <a name="minus-state"></a> Task 2. Minus state.
#
# **Input**: A qubit in the $|0\rangle$ state.
#
# **Goal**: Change the state of the qubit to $|-\rangle = \frac{1}{\sqrt{2}} \big(|0\rangle - |1\rangle\big)$.
# +
%kata T02_MinusState_Test
operation MinusState (q : Qubit) : Unit {
// ...
}
# -
# *Can't come up with a solution? See the explained solution in the [Superposition Workbook](./Workbook_Superposition.ipynb#minus-state).*
# ### <a name="unequal-superposition"></a> Task 3*. Unequal superposition.
#
# **Inputs:**
#
# 1. A qubit in the $|0\rangle$ state.
# 2. Angle $\alpha$, in radians, represented as `Double`.
#
# **Goal** : Change the state of the qubit to $\cos{α} |0\rangle + \sin{α} |1\rangle$.
#
# <br/>
# <details>
# <summary><b>Need a hint? Click here</b></summary>
# Experiment with rotation gates from Microsoft.Quantum.Intrinsic namespace.
# Note that all rotation operators rotate the state by <i>half</i> of its angle argument.
# </details>
# +
%kata T03_UnequalSuperposition_Test
operation UnequalSuperposition (q : Qubit, alpha : Double) : Unit {
// ...
}
# -
# *Can't come up with a solution? See the explained solution in the [Superposition Workbook](./Workbook_Superposition.ipynb#unequal-superposition).*
# ### <a name="superposition-of-all-basis-vectors-on-two-qubits"></a>Task 4. Superposition of all basis vectors on two qubits.
#
# **Input:** Two qubits in the $|00\rangle$ state (stored in an array of length 2).
#
# **Goal:** Change the state of the qubits to $|+\rangle \otimes |+\rangle = \frac{1}{2} \big(|00\rangle + |01\rangle + |10\rangle + |11\rangle\big)$.
# +
%kata T04_AllBasisVectors_TwoQubits_Test
operation AllBasisVectors_TwoQubits (qs : Qubit[]) : Unit {
// ...
}
# -
# *Can't come up with a solution? See the explained solution in the [Superposition Workbook](./Workbook_Superposition.ipynb#superposition-of-all-basis-vectors-on-two-qubits).*
# ### <a name="superposition-of-basis-vectors-with-phases"></a>Task 5. Superposition of basis vectors with phases.
#
# **Input:** Two qubits in the $|00\rangle$ state (stored in an array of length 2).
#
# **Goal:** Change the state of the qubits to $\frac{1}{2} \big(|00\rangle + i|01\rangle - |10\rangle - i|11\rangle\big)$.
#
# <br/>
# <details>
# <summary><b>Need a hint? Click here</b></summary>
# Is this state separable?
# </details>
# +
%kata T05_AllBasisVectorsWithPhases_TwoQubits_Test
operation AllBasisVectorsWithPhases_TwoQubits (qs : Qubit[]) : Unit {
// ...
}
# -
# *Can't come up with a solution? See the explained solution in the [Superposition Workbook](./Workbook_Superposition.ipynb#superposition-of-basis-vectors-with-phases).*
# ### <a name="bell-state"></a>Task 6. Bell state $|\Phi^{+}\rangle$.
#
# **Input:** Two qubits in the $|00\rangle$ state (stored in an array of length 2).
#
# **Goal:** Change the state of the qubits to $|\Phi^{+}\rangle = \frac{1}{\sqrt{2}} \big (|00\rangle + |11\rangle\big)$.
#
# > You can find detailed coverage of Bell states and their creation [in this blog post](https://blogs.msdn.microsoft.com/uk_faculty_connection/2018/02/06/a-beginners-guide-to-quantum-computing-and-q/).
# +
%kata T06_BellState_Test
operation BellState (qs : Qubit[]) : Unit {
// ...
}
# -
# *Can't come up with a solution? See the explained solution in the [Superposition Workbook](./Workbook_Superposition.ipynb#bell-state).*
# ### <a name="all-bell-states"></a> Task 7. All Bell states.
#
# **Inputs:**
#
# 1. Two qubits in the $|00\rangle$ state (stored in an array of length 2).
# 2. An integer index.
#
# **Goal:** Change the state of the qubits to one of the Bell states, based on the value of index:
#
# <table>
# <col width="50"/>
# <col width="200"/>
# <tr>
# <th style="text-align:center">Index</th>
# <th style="text-align:center">State</th>
# </tr>
# <tr>
# <td style="text-align:center">0</td>
# <td style="text-align:center">$|\Phi^{+}\rangle = \frac{1}{\sqrt{2}} \big (|00\rangle + |11\rangle\big)$</td>
# </tr>
# <tr>
# <td style="text-align:center">1</td>
# <td style="text-align:center">$|\Phi^{-}\rangle = \frac{1}{\sqrt{2}} \big (|00\rangle - |11\rangle\big)$</td>
# </tr>
# <tr>
# <td style="text-align:center">2</td>
# <td style="text-align:center">$|\Psi^{+}\rangle = \frac{1}{\sqrt{2}} \big (|01\rangle + |10\rangle\big)$</td>
# </tr>
# <tr>
# <td style="text-align:center">3</td>
# <td style="text-align:center">$|\Psi^{-}\rangle = \frac{1}{\sqrt{2}} \big (|01\rangle - |10\rangle\big)$</td>
# </tr>
# </table>
# +
%kata T07_AllBellStates_Test
operation AllBellStates (qs : Qubit[], index : Int) : Unit {
// ...
}
# -
# *Can't come up with a solution? See the explained solution in the [Superposition Workbook](./Workbook_Superposition.ipynb#all-bell-states).*
# ### <a name="greenberger-horne-zeilinger"></a> Task 8. Greenberger–Horne–Zeilinger state.
#
# **Input:** $N$ ($N \ge 1$) qubits in the $|0 \dots 0\rangle$ state (stored in an array of length $N$).
#
# **Goal:** Change the state of the qubits to the GHZ state $\frac{1}{\sqrt{2}} \big (|0\dots0\rangle + |1\dots1\rangle\big)$.
#
# > For the syntax of flow control statements in Q#, see [the Q# documentation](https://docs.microsoft.com/quantum/language/statements#control-flow).
# +
%kata T08_GHZ_State_Test
operation GHZ_State (qs : Qubit[]) : Unit {
// You can find N as Length(qs).
// ...
}
# -
# *Can't come up with a solution? See the explained solution in the [Superposition Workbook](./Workbook_Superposition_Part2.ipynb#greenberger-horne-zeilinger).*
# ### <a name="superposition-of-all-basis-vectors"></a> Task 9. Superposition of all basis vectors.
#
# **Input:** $N$ ($N \ge 1$) qubits in the $|0 \dots 0\rangle$ state.
#
# **Goal:** Change the state of the qubits to an equal superposition of all basis vectors $\frac{1}{\sqrt{2^N}} \big (|0 \dots 0\rangle + \dots + |1 \dots 1\rangle\big)$.
#
# > For example, for $N = 2$ the final state should be $\frac{1}{\sqrt{2}} \big (|00\rangle + |01\rangle + |10\rangle + |11\rangle\big)$.
#
# <br/>
# <details>
# <summary><b>Need a hint? Click here</b></summary>
# Is this state separable?
# </details>
# +
%kata T09_AllBasisVectorsSuperposition_Test
operation AllBasisVectorsSuperposition (qs : Qubit[]) : Unit {
// ...
}
# -
# *Can't come up with a solution? See the explained solution in the [Superposition Workbook](./Workbook_Superposition_Part2.ipynb#superposition-of-all-basis-vectors).*
# ### <a name="superposition-of-all-even-or-all-odd-numbers"></a> Task 10. Superposition of all even or all odd numbers.
#
# **Inputs:**
#
# 1. $N$ ($N \ge 1$) qubits in the $|0 \dots 0\rangle$ state (stored in an array of length $N$).
# 2. A boolean `isEven`.
#
# **Goal:** Prepare a superposition of all *even* numbers if `isEven` is `true`, or of all *odd* numbers if `isEven` is `false`.
# A basis state encodes an integer number using [big-endian](https://en.wikipedia.org/wiki/Endianness) binary notation: state $|01\rangle$ corresponds to the integer $1$, and state $|10 \rangle$ - to the integer $2$.
#
# > For example, for $N = 2$ and `isEven = false` you need to prepare superposition $\frac{1}{\sqrt{2}} \big (|01\rangle + |11\rangle\big )$,
# and for $N = 2$ and `isEven = true` - superposition $\frac{1}{\sqrt{2}} \big (|00\rangle + |10\rangle\big )$.
# +
%kata T10_EvenOddNumbersSuperposition_Test
operation EvenOddNumbersSuperposition (qs : Qubit[], isEven : Bool) : Unit {
// ...
}
# -
# *Can't come up with a solution? See the explained solution in the [Superposition Workbook](./Workbook_Superposition_Part2.ipynb#superposition-of-all-even-or-all-odd-numbers).*
# ### <a name="threestates-twoqubits"></a>Task 11*. $\frac{1}{\sqrt{3}} \big(|00\rangle + |01\rangle + |10\rangle\big)$ state.
#
# **Input:** Two qubits in the $|00\rangle$ state.
#
# **Goal:** Change the state of the qubits to $\frac{1}{\sqrt{3}} \big(|00\rangle + |01\rangle + |10\rangle\big)$.
#
# <br/>
# <details>
# <summary><b>Need a hint? Click here</b></summary>
# If you need trigonometric functions, you can find them in Microsoft.Quantum.Math namespace; you'll need to add <pre>open Microsoft.Quantum.Math;</pre> to the code before the operation definition.
# </details>
# +
%kata T11_ThreeStates_TwoQubits_Test
operation ThreeStates_TwoQubits (qs : Qubit[]) : Unit {
// ...
}
# -
# ### Task 12*. Hardy state.
#
# **Input:** Two qubits in the $|00\rangle$ state.
#
# **Goal:** Change the state of the qubits to $\frac{1}{\sqrt{12}} \big(3|00\rangle + |01\rangle + |10\rangle + |11\rangle\big)$.
#
# <br/>
# <details>
# <summary><b>Need a hint? Click here</b></summary>
# If you need trigonometric functions, you can find them in Microsoft.Quantum.Math namespace; you'll need to add <pre>open Microsoft.Quantum.Math;</pre> to the code before the operation definition.
# </details>
# +
%kata T12_Hardy_State_Test
operation Hardy_State (qs : Qubit[]) : Unit {
// ...
}
# -
# ### Task 13. Superposition of $|0 \dots 0\rangle$ and the given bit string.
#
# **Inputs:**
#
# 1. $N$ ($N \ge 1$) qubits in the $|0 \dots 0\rangle$ state.
# 2. A bit string of length $N$ represented as `Bool[]`. Bit values `false` and `true` correspond to $|0\rangle$ and $|1\rangle$ states. You are guaranteed that the first bit of the bit string is `true`.
#
# **Goal:** Change the state of the qubits to an equal superposition of $|0 \dots 0\rangle$ and the basis state given by the bit string.
#
# > For example, for the bit string `[true, false]` the state required is $\frac{1}{\sqrt{2}}\big(|00\rangle + |10\rangle\big)$.
# +
%kata T13_ZeroAndBitstringSuperposition_Test
operation ZeroAndBitstringSuperposition (qs : Qubit[], bits : Bool[]) : Unit {
// ...
}
# -
# ### Task 14. Superposition of two bit strings.
#
# **Inputs:**
#
# 1. $N$ ($N \ge 1$) qubits in the $|0 \dots 0\rangle$ state.
# 2. Two bit strings of length $N$ represented as `Bool[]`s. Bit values `false` and `true` correspond to $|0\rangle$ and $|1\rangle$ states. You are guaranteed that the two bit strings differ in at least one bit.
#
# **Goal:** Change the state of the qubits to an equal superposition of the basis states given by the bit strings.
#
# > For example, for bit strings `[false, true, false]` and `[false, false, true]` the state required is $\frac{1}{\sqrt{2}}\big(|010\rangle + |001\rangle\big)$.
#
# > If you need to define any helper functions, you'll need to create an extra code cell for it and execute it before returning to this cell.
# +
%kata T14_TwoBitstringSuperposition_Test
operation TwoBitstringSuperposition (qs : Qubit[], bits1 : Bool[], bits2 : Bool[]) : Unit {
// ...
}
# -
# ### Task 15*. Superposition of four bit strings.
#
# **Inputs:**
#
# 1. $N$ ($N \ge 1$) qubits in the $|0 \dots 0\rangle$ state.
# 2. Four bit strings of length $N$, represented as `Bool[][]` `bits`. `bits` is an $4 \times N$ which describes the bit strings as follows: `bits[i]` describes the `i`-th bit string and has $N$ elements. You are guaranteed that all four bit strings will be distinct.
#
# **Goal:** Change the state of the qubits to an equal superposition of the four basis states given by the bit strings.
#
# > For example, for $N = 3$ and `bits = [[false, true, false], [true, false, false], [false, false, true], [true, true, false]]` the state required is $\frac{1}{2}\big(|010\rangle + |100\rangle + |001\rangle + |110\rangle\big)$.
#
# <br/>
# <details>
# <summary><b>Need a hint? Click here</b></summary>
# Remember that you can allocate extra qubits. If you do, you'll need to return them to the $|0\rangle$ state before releasing them.
# </details>
# +
%kata T15_FourBitstringSuperposition_Test
operation FourBitstringSuperposition (qs : Qubit[], bits : Bool[][]) : Unit {
// ...
}
# -
# ### Task 16**. W state on $2^k$ qubits.
#
# **Input:** $N = 2^k$ qubits in the $|0 \dots 0\rangle$ state.
#
# **Goal:** Change the state of the qubits to the [W state](https://en.wikipedia.org/wiki/W_state) - an equal superposition of $N$ basis states on $N$ qubits which have Hamming weight of 1.
#
# > For example, for $N = 4$ the required state is $\frac{1}{2}\big(|1000\rangle + |0100\rangle + |0010\rangle + |0001\rangle\big)$.
#
# <br/>
# <details>
# <summary><b>Need a hint? Click here</b></summary>
# You can use Controlled modifier to perform arbitrary controlled gates.
# </details>
# +
%kata T16_WState_PowerOfTwo_Test
operation WState_PowerOfTwo (qs : Qubit[]) : Unit {
// ...
}
# -
# ### Task 17**. W state on an arbitrary number of qubits.
#
# **Input:** $N$ qubits in the $|0 \dots 0\rangle$ state ($N$ is not necessarily a power of 2).
#
# **Goal:** Change the state of the qubits to the [W state](https://en.wikipedia.org/wiki/W_state) - an equal superposition of $N$ basis states on $N$ qubits which have Hamming weight of 1.
#
# > For example, for $N = 3$ the required state is $\frac{1}{\sqrt{3}}\big(|100\rangle + |010\rangle + |001\rangle\big)$.
#
# <br/>
# <details>
# <summary><b>Need a hint? Click here</b></summary>
# You can modify the signature of the given operation to specify its controlled specialization.
# </details>
# +
%kata T17_WState_Arbitrary_Test
operation WState_Arbitrary (qs : Qubit[]) : Unit {
// ...
}
| Superposition/Superposition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import openpnm as op
import openpnm.models.geometry as gm
import openpnm.models.misc as mm
import openpnm.models.physics as pm
import scipy as sp
print(op.__version__)
# %matplotlib inline
# ## Generate Two Networks with Different Spacing
spacing_lg = 0.00006
layer_lg = op.network.Cubic(shape=[10, 10, 1], spacing=spacing_lg)
spacing_sm = 0.00002
layer_sm = op.network.Cubic(shape=[30, 5, 1], spacing=spacing_sm)
# ## Position Networks Appropriately, then Stitch Together
# Start by assigning labels to each network for identification later
layer_sm['pore.small'] = True
layer_sm['throat.small'] = True
layer_lg['pore.large'] = True
layer_lg['throat.large'] = True
# Next manually offset CL one full thickness relative to the GDL
layer_sm['pore.coords'] -= [0, spacing_sm*5, 0]
layer_sm['pore.coords'] += [0, 0, spacing_lg/2 - spacing_sm/2] # And shift up by 1/2 a lattice spacing
# Finally, send both networks to stitch which will stitch CL onto GDL
from openpnm.topotools import stitch
stitch(network=layer_lg, donor=layer_sm,
P_network=layer_lg.pores('bottom'),
P_donor=layer_sm.pores('top'),
len_max=0.00005)
combo_net = layer_lg
combo_net.name = 'combo'
# ## Create Geometry Objects for Each Layer
Ps = combo_net.pores('small')
Ts = combo_net.throats('small')
geom_sm = op.geometry.GenericGeometry(network=combo_net, pores=Ps, throats=Ts)
Ps = combo_net.pores('large')
Ts = combo_net.throats('small', mode='not')
geom_lg = op.geometry.GenericGeometry(network=combo_net, pores=Ps, throats=Ts)
# ### Add Geometrical Properties to the *Small* Domain
# The *small* domain will be treated as a continua, so instead of assigning pore sizes we want the 'pore' to be same size as the lattice cell.
geom_sm['pore.diameter'] = spacing_sm
geom_sm['pore.area'] = spacing_sm**2
geom_sm['throat.diameter'] = spacing_sm
geom_sm['throat.area'] = spacing_sm**2
geom_sm['throat.length'] = 1e-12 # A very small number to represent nearly 0-length
# ### Add Geometrical Properties to the *Large* Domain
geom_lg['pore.diameter'] = spacing_lg*sp.rand(combo_net.num_pores('large'))
geom_lg.add_model(propname='pore.area',
model=gm.pore_area.sphere)
geom_lg.add_model(propname='throat.diameter',
model=mm.misc.from_neighbor_pores,
pore_prop='pore.diameter', mode='min')
geom_lg.add_model(propname='throat.area',
model=gm.throat_area.cylinder)
geom_lg.add_model(propname='throat.length',
model=gm.throat_length.straight)
# ## Create Phase and Physics Objects
air = op.phases.Air(network=combo_net, name='air')
phys_lg = op.physics.GenericPhysics(network=combo_net, geometry=geom_lg, phase=air)
phys_sm = op.physics.GenericPhysics(network=combo_net, geometry=geom_sm, phase=air)
# Add pore-scale models for diffusion to each Physics:
phys_lg.add_model(propname='throat.diffusive_conductance',
model=pm.diffusive_conductance.ordinary_diffusion)
phys_sm.add_model(propname='throat.diffusive_conductance',
model=pm.diffusive_conductance.ordinary_diffusion)
# For the *small* layer we've used a normal diffusive conductance model, which when combined with the diffusion coefficient of air will be equivalent to open-air diffusion. If we want the *small* layer to have some tortuosity we must account for this:
porosity = 0.5
tortuosity = 2
phys_sm['throat.diffusive_conductance'] *= (porosity/tortuosity)
# Note that this extra line is NOT a pore-scale model, so it will be over-written when the `phys_sm` object is regenerated.
# ### Add a Reaction Term to the Small Layer
# A standard n-th order chemical reaction is $ r=k \cdot x^b $, or more generally: $ r = A_1 \cdot x^{A_2} + A_3 $. This model is available in `OpenPNM.Physics.models.generic_source_terms`, and we must specify values for each of the constants.
# Set Source Term
air['pore.A1'] = 1e-10 # Reaction pre-factor
air['pore.A2'] = 2 # Reaction order
air['pore.A3'] = 0 # A generic offset that is not needed so set to 0
phys_sm.add_model(propname='pore.reaction',
model=pm.generic_source_term.power_law,
A1='pore.A1', A2='pore.A2', A3='pore.A3',
X='pore.mole_fraction',
regen_mode='deferred')
# ## Perform a Diffusion Calculation
Deff = op.algorithms.ReactiveTransport(network=combo_net, phase=air)
Ps = combo_net.pores(['large', 'right'], mode='intersection')
Deff.set_value_BC(pores=Ps, values=1)
Ps = combo_net.pores('small')
Deff.set_source(propname='pore.reaction', pores=Ps)
Deff.settings['conductance'] = 'throat.diffusive_conductance'
Deff.settings['quantity'] = 'pore.mole_fraction'
Deff.run()
# ## Visualize the Concentration Distribution
# Save the results to a VTK file for visualization in Paraview:
Deff.results()
op.io.VTK.save(network=combo_net, phases=[air])
# And the result would look something like this:
# 
| Simulations/mixed_continuum_and_pore_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pandana network accessibility simple demo
#
# This notebook uses [pandana](https://udst.github.io/pandana/network.html) (v0.2) to download street network and points-of-interest data from OpenStreetMap and then calculate network accessibility to the points of interest. Note: pandana currently only runs on Python 2.
#
# For a more in-depth demo, check out [pandana-accessibility-demo-full.ipynb](pandana-accessibility-demo-full.ipynb)
import pandana, matplotlib.pyplot as plt
from pandana.loaders import osm
# %matplotlib inline
import matplotlib
print(matplotlib.__version__)
print(pandana.version)
bbox = [48.0616244,11.360777, 48.2481162, 11.7229083]
#[37.76, -122.35, 37.9, -122.17] #lat-long bounding box for berkeley/oakland
amenity = 'pub' #accessibility to this type of amenity
distance = 1500 #max distance in meters
# ## Download points of interest (POIs) and network data from OpenStreetMap
# first download the points of interest corresponding to the specified amenity type
pois = osm.node_query(bbox[0], bbox[1], bbox[2], bbox[3], tags='"amenity"="{}"'.format(amenity))
pois[['amenity', 'name', 'lat', 'lon']].tail()
# query the OSM API for the street network within the specified bounding box
network = osm.network_from_bbox(bbox[0], bbox[1], bbox[2], bbox[3])
pickle.dump(network, open('munich_net.pkl','wb'))
# how many network nodes did we get for this bounding box?
len(network[0].index)
import pickle
#
network = pickle.load(open('munich_net.pkl','rb'))
# +
import pandana as pdna
network=pdna.Network(network[0]["x"], network[0]["y"],
network[1].reset_index()['level_0'],
network[1].reset_index()['level_1'],
network[1].reset_index()[["distance"]])
# -
# ## Process the network data then compute accessibility
# identify nodes that are connected to fewer than some threshold of other nodes within a given distance
# do nothing with this for now, but see full example in other notebook for more
lcn = network.low_connectivity_nodes(impedance=1000, count=10, imp_name='distance')
# precomputes the range queries (the reachable nodes within this maximum distance)
# so, as long as you use a smaller distance, cached results will be used
network.precompute(distance + 1)
# initialize the underlying C++ points-of-interest engine
network.init_pois(num_categories=1, max_dist=distance, max_pois=7)
# initialize a category for this amenity with the locations specified by the lon and lat columns
network.set_pois(category='my_amenity', x_col=pois['lon'], y_col=pois['lat'])
# +
# search for the n nearest amenities to each node in the network
access = network.nearest_pois(distance=distance, category='my_amenity', num_pois=7)
# each df cell represents the network distance from the node to each of the n POIs
access.head()
# -
# ## Plot the accessibility
# +
# keyword arguments to pass for the matplotlib figure
bbox_aspect_ratio = (bbox[2] - bbox[0]) / (bbox[3] - bbox[1])
fig_kwargs = {'facecolor':'w',
'figsize':(10, 10 * bbox_aspect_ratio)}
# keyword arguments to pass for scatter plots
plot_kwargs = {'s':5,
'alpha':0.9,
'cmap':'viridis_r',
'edgecolor':'none'}
# -
# plot the distance to the nth nearest amenity
n = 1
bmap, fig, ax = network.plot(access[n], bbox=bbox, plot_kwargs=plot_kwargs, fig_kwargs=fig_kwargs)
#ax.set_axis_bgcolor('k')
ax.set_title('Walking distance (m) to nearest {} around Munich'.format(amenity), fontsize=15)
fig.savefig('images/accessibility-pub-east-bay.png', dpi=200, bbox_inches='tight')
plt.show()
# For a more in-depth demo, check out [pandana-accessibility-demo-full.ipynb](pandana-accessibility-demo-full.ipynb)
| accessibility/pandana-accessibility-demo-simple.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/GarvRJ/T.Y.MINIPROJECT/blob/master/yolov5.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="Xk5T7drQnZ8z" outputId="19a22226-06e5-49cf-a9a7-d02b35a98341"
# ls
# + colab={"base_uri": "https://localhost:8080/"} id="jdlCyJMOjvkK" outputId="a9fb89c3-2327-42f8-dd71-cbaf79fceb1c"
# cd /content/drive/MyDrive/yolov5_miniproject
# + colab={"base_uri": "https://localhost:8080/"} id="T7qe_L7lnljk" outputId="b95178ef-b8a0-4f7f-bab7-f5193bcd8841"
# !git clone https://github.com/ultralytics/yolov5
# + colab={"base_uri": "https://localhost:8080/"} id="RDYyBTjvlY9i" outputId="df1d3e7e-08ba-4264-e320-0d28badd803f"
# %cd yolov5
# + id="-VDdu1NQlY-t" colab={"base_uri": "https://localhost:8080/"} outputId="3d89c1de-bd4e-4a09-9197-fa6aec12fe1c"
# !git reset --hard 886f1c03d839575afecb059accf74296fad395b6
# + colab={"base_uri": "https://localhost:8080/"} id="DBMLKPHMmtkc" outputId="d9e4e248-7680-48d7-8fc3-291e1735a56f"
# !pip install -qr requirements.txt # install dependencies (ignore errors)
import torch
from IPython.display import Image, clear_output # to display images
from utils.google_utils import gdrive_download # to download models/datasets
# clear_output()
print('Setup complete. Using torch %s %s' % (torch.__version__, torch.cuda.get_device_properties(0) if torch.cuda.is_available() else 'CPU'))
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="1Y1GvdbKoHV_" outputId="e7577539-3e4c-4448-8c8e-20bedc503356"
# !pip install roboflow
from roboflow import Roboflow
rf = Roboflow(api_key="FX0t6K12dztT7aeguf2z")
project = rf.workspace("roboflow-gw7yv").project("vehicles-openimages")
dataset = project.version(1).download("yolov5")
# + colab={"base_uri": "https://localhost:8080/"} id="b-xvGDAupIL_" outputId="1862d298-f572-474d-e235-ea49d660a066"
# %cd yolov5
# %ls
# + colab={"base_uri": "https://localhost:8080/"} id="fYybqZ0Wr091" outputId="df11795b-659a-4ff3-bfdd-badaffc6fee8"
# %cat Vehicles-OpenImages-1/data.yaml
# + id="oR5t_P3otDio"
# define number of classes based on YAML
import yaml
with open("Vehicles-OpenImages-1" + "/data.yaml", 'r') as stream:
num_classes = str(yaml.safe_load(stream)['nc'])
# + colab={"base_uri": "https://localhost:8080/"} id="WUkMLHwZtOii" outputId="0dce510a-eff7-43c2-aa81-2bb4b84a455d"
# %cat /content/yolov5/models/yolov5s.yaml
# + id="aNPaMbbCtVlX"
from IPython.core.magic import register_line_cell_magic
@register_line_cell_magic
def writetemplate(line, cell):
with open(line, 'w') as f:
f.write(cell.format(**globals()))
# + id="M-hjWugAtjLh"
# %%writetemplate /content/yolov5/models/custom_yolov5s.yaml
# parameters
nc: {num_classes} # number of classes
depth_multiple: 0.33 # model depth multiple
width_multiple: 0.50 # layer channel multiple
# anchors
anchors:
- [10,13, 16,30, 33,23] # P3/8
- [30,61, 62,45, 59,119] # P4/16
- [116,90, 156,198, 373,326] # P5/32
# YOLOv5 backbone
backbone:
# [from, number, module, args]
[[-1, 1, Focus, [64, 3]], # 0-P1/2
[-1, 1, Conv, [128, 3, 2]], # 1-P2/4
[-1, 3, BottleneckCSP, [128]],
[-1, 1, Conv, [256, 3, 2]], # 3-P3/8
[-1, 9, BottleneckCSP, [256]],
[-1, 1, Conv, [512, 3, 2]], # 5-P4/16
[-1, 9, BottleneckCSP, [512]],
[-1, 1, Conv, [1024, 3, 2]], # 7-P5/32
[-1, 1, SPP, [1024, [5, 9, 13]]],
[-1, 3, BottleneckCSP, [1024, False]], # 9
]
# YOLOv5 head
head:
[[-1, 1, Conv, [512, 1, 1]],
[-1, 1, nn.Upsample, [None, 2, 'nearest']],
[[-1, 6], 1, Concat, [1]], # cat backbone P4
[-1, 3, BottleneckCSP, [512, False]], # 13
[-1, 1, Conv, [256, 1, 1]],
[-1, 1, nn.Upsample, [None, 2, 'nearest']],
[[-1, 4], 1, Concat, [1]], # cat backbone P3
[-1, 3, BottleneckCSP, [256, False]], # 17 (P3/8-small)
[-1, 1, Conv, [256, 3, 2]],
[[-1, 14], 1, Concat, [1]], # cat head P4
[-1, 3, BottleneckCSP, [512, False]], # 20 (P4/16-medium)
[-1, 1, Conv, [512, 3, 2]],
[[-1, 10], 1, Concat, [1]], # cat head P5
[-1, 3, BottleneckCSP, [1024, False]], # 23 (P5/32-large)
[[17, 20, 23], 1, Detect, [nc, anchors]], # Detect(P3, P4, P5)
]
# + colab={"base_uri": "https://localhost:8080/"} id="6QrlYyFPtnNp" outputId="c59cf490-480a-48b0-8cd0-933f6a34ccd4"
# !pip install wandb
# + colab={"base_uri": "https://localhost:8080/", "height": 133} id="JXNS2SsGtt6M" outputId="ee1f5472-5e8a-4e83-a4df-039e0d749442"
# + colab={"base_uri": "https://localhost:8080/"} id="DUyZeom_uVZb" outputId="252289c4-8f77-4b8f-817a-537b2c5287e8"
# %%time
# %cd /content/yolov5/
# !python train.py --img 416 --batch 16 --epochs 100 --data Vehicles-OpenImages-12/data.yaml --cfg ./models/custom_yolov5s.yaml --weights '' --name yolov5s_results --cache
| yolov5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
import openpathsampling as paths
import numpy as np
import matplotlib.pyplot as plt
import os
import openpathsampling.visualize as ops_vis
from IPython.display import SVG
# ### Advanced analysis techniques
#
# Now we'll move on to a few more advanced analysis techniques. (These are discussed in Paper II.)
#
# With the fixed path length ensemble, we should check for recrossings. To do this, we create an ensemble which represents the recrossing paths: a frame in $\beta$, possible frames in neither $\alpha$ nor $\beta$, and then a frame in $\alpha$.
#
# Then we check whether any subtrajectory of a trial trajectory matches that ensemble, by using the `Ensemble.split()` function. We can then further refine to see which steps that included trials with recrossings were actually accepted.
# %%time
flexible = paths.Storage("tps_nc_files/alanine_dipeptide_tps.nc")
# %%time
fixed = paths.Storage("tps_nc_files/alanine_dipeptide_fixed_tps.nc")
flex_scheme = flexible.schemes[0]
fixed_scheme = fixed.schemes[0]
# +
# TODO: cache trajectories too?
# -
# create the ensemble that identifies recrossings
alpha = fixed.volumes.find('C_7eq')
beta = fixed.volumes.find('alpha_R')
recrossing_ensemble = paths.SequentialEnsemble([
paths.LengthEnsemble(1) & paths.AllInXEnsemble(beta),
paths.OptionalEnsemble(paths.AllOutXEnsemble(alpha | beta)),
paths.LengthEnsemble(1) & paths.AllInXEnsemble(alpha)
])
# %%time
# now we check each step to see if its trial has a recrossing
steps_with_recrossing = []
for step in fixed.steps:
# trials is a list of samples: with shooting, only one in the list
recrossings = [] # default for initial empty move (no trials in step[0].change)
for trial in step.change.trials:
recrossings = recrossing_ensemble.split(trial.trajectory)
# recrossing contains a list with the recrossing trajectories
# (len(recrossing) == 0 if no recrossings)
if len(recrossings) > 0:
steps_with_recrossing += [step] # save for later analysis
accepted_recrossings = [step for step in steps_with_recrossing if step.change.accepted is True]
print "Trials with recrossings:", len(steps_with_recrossing)
print "Accepted trials with recrossings:", len(accepted_recrossings)
# Note that the accepted trials with recrossing does not account for how long the trial remained active. It also doesn't tell us whether the trial represented a new recrossing event, or was correlated with the previous recrossing event.
# Let's take a look at one of the accepted trajectories with a recrossing event. We'll plot the value of $\psi$, since this is what distinguishes the two states. We'll also select the frames that are actually inside each state and color them (red for $\alpha$, blue for $\beta$).
# +
psi = fixed.cvs.find('psi')
trajectory = accepted_recrossings[0].active[0].trajectory
in_alpha_indices = [trajectory.index(s) for s in trajectory if alpha(s)]
in_alpha_psi = [psi(trajectory)[i] for i in in_alpha_indices]
in_beta_indices = [trajectory.index(s) for s in trajectory if beta(s)]
in_beta_psi = [psi(trajectory)[i] for i in in_beta_indices]
plt.plot(psi(trajectory), 'k-')
plt.plot(in_alpha_indices, in_alpha_psi, 'ro') # alpha in red
plt.plot(in_beta_indices, in_beta_psi, 'bo') # beta in blue
# -
# Now let's see how many recrossing events there are in each accepted trial. If there's one recrossing, then the trajectory must go $\alpha\to\beta\to\alpha\to\beta$ to be accepted. Two recrossings would mean $\alpha\to\beta\to\alpha\to\beta\to\alpha\to\beta$.
recrossings_per = []
for step in accepted_recrossings:
for test in step.change.trials:
recrossings_per.append(len(recrossing_ensemble.split(test.trajectory)))
print recrossings_per
# these numbers come from accepted trial steps, not all steps
print sum(recrossings_per)
print len(recrossings_per)
print len([x for x in recrossings_per if x==2])
# # Comparing the fixed and flexible simulations
# %%time
# transition path length distribution
flex_ens = flex_scheme.network.sampling_ensembles[0]
fixed_transition_segments = sum([flex_ens.split(step.active[0].trajectory) for step in fixed.steps],[])
fixed_transition_length = [len(traj) for traj in fixed_transition_segments]
flexible_transition_length = [len(s.active[0].trajectory) for s in flexible.steps]
print len(fixed_transition_length)
bins = np.linspace(0, 400, 80);
plt.hist(flexible_transition_length, bins, alpha=0.5, normed=True, label="flexible");
plt.hist(fixed_transition_length, bins, alpha=0.5, normed=True, label="fixed");
plt.legend(loc='upper right');
# #### Identifying different mechanisms using custom ensembles
#
# We expected the plot above to be very similar for both cases. However, we know that the $\alpha\to\beta$ transition in alanine dipeptide can occur via two mechanisms: since $\psi$ is periodic, the transition can occur due to an overall increase in $\psi$, or due to an overall decrease in $\psi$. We also know that the alanine dipeptide transitions aren't actually all that rare, so they will occur spontaneously in long simulations.
#
#
# This section shows how to create custom ensembles to identify whether the transition occurred with an increasing $\psi$ or a decreasing $\psi$. We also need to account for (unlikely) edge cases where the path starts in one direction but completes the transition from the other.
# First, we'll create a few more `Volume` objects. In this case, we will completely tile the Ramachandran space; while a complete tiling isn't necessary, it is often useful.
# first, we fully subdivide the Ramachandran space
phi = fixed.cvs.find('phi')
deg = 180.0/np.pi
nml_plus = paths.PeriodicCVDefinedVolume(psi, -160/deg, -100/deg, -np.pi, np.pi)
nml_minus = paths.PeriodicCVDefinedVolume(psi, 0/deg, 100/deg, -np.pi, np.pi)
nml_alpha = (paths.PeriodicCVDefinedVolume(phi, 0/deg, 180/deg, -np.pi, np.pi) &
paths.PeriodicCVDefinedVolume(psi, 100/deg, 200/deg, -np.pi, np.pi))
nml_beta = (paths.PeriodicCVDefinedVolume(phi, 0/deg, 180/deg, -np.pi, np.pi) &
paths.PeriodicCVDefinedVolume(psi, -100/deg, 0/deg, -np.pi, np.pi))
# +
#TODO: plot to display where these volumes are
# -
# Next, we'll create ensembles for the "increasing" and "decreasing" transitions. These transitions mark a crossing of either the `nml_plus` or the `nml_minus`. These aren't necessarily $\alpha\to\beta$ transitions. However, any $\alpha\to\beta$ transition must contain at least one subtrajectory which satsifies one of these ensembles.
increasing = paths.SequentialEnsemble([
paths.AllInXEnsemble(alpha | nml_alpha),
paths.AllInXEnsemble(nml_plus),
paths.AllInXEnsemble(beta | nml_beta)
])
decreasing = paths.SequentialEnsemble([
paths.AllInXEnsemble(alpha | nml_alpha),
paths.AllInXEnsemble(nml_minus),
paths.AllInXEnsemble(beta | nml_beta)
])
# Finally, we'll write a little function that characterizes a set of trajectories according to these ensembles. It returns a dictionary mapping the ensemble (`increasing` or `decreasing`) to a list of trajectories that have a subtrajectory that satisfies it (at least one entry in `ensemble.split(trajectory)`). That dictionary also contains keys for `'multiple'` matched ensembles and `None` if no ensemble was matched. Trajectories for either of these keys would need to be investigated further.
def categorize_transitions(ensembles, trajectories):
results = {ens : [] for ens in ensembles + ['multiple', None]}
for traj in trajectories:
matched_ens = None
for ens in ensembles:
if len(ens.split(traj)) > 0:
if matched_ens is not None:
matched_ens = 'multiple'
else:
matched_ens = ens
results[matched_ens].append(traj)
return results
# With that function defined, let's use it!
categorized = categorize_transitions(ensembles=[increasing, decreasing],
trajectories=fixed_transition_segments)
print "increasing:", len(categorized[increasing])
print "decreasing:", len(categorized[decreasing])
print " multiple:", len(categorized['multiple'])
print " None:", len(categorized[None])
# Comparing to the flexible length simulation:
flex_trajs = [step.active[0].trajectory for step in flexible.steps]
flex_categorized = categorize_transitions(ensembles=[increasing, decreasing],
trajectories=flex_trajs[::10])
print "increasing:", len(flex_categorized[increasing])
print "decreasing:", len(flex_categorized[decreasing])
print " multiple:", len(flex_categorized['multiple'])
print " None:", len(flex_categorized[None])
# So the fixed length sampling is somehow capturing both kinds of transitions (probably because they are not really that rare). Let's see what the path length distribution from only the decreasing transitions looks
plt.hist([len(traj) for traj in flex_categorized[decreasing]], bins, alpha=0.5, normed=True);
plt.hist([len(traj) for traj in categorized[decreasing]], bins, alpha=0.5, normed=True);
# Still a little off, although this might be due to bad sampling. Let's see how many of the decorrelated trajectories have this kind of transition.
full_fixed_tree = ops_vis.PathTree(
fixed.steps,
ops_vis.ReplicaEvolution(replica=0)
)
full_history = full_fixed_tree.generator
# start with the decorrelated tragectories
fixed_decorrelated = full_history.decorrelated_trajectories
# find the A->B transitions from the decorrelated trajectories
decorrelated_transitions = sum([flex_ens.split(traj) for traj in fixed_decorrelated], [])
# find the A->B transition from these which are decreasing
decorrelated_decreasing = sum([decreasing.split(traj) for traj in decorrelated_transitions], [])
print len(decorrelated_decreasing)
# So this is based off of 11 decorrelated trajectory transitions. That's not a lot of statistics.
#
# However, we expect to see a *very* different distribution for the "increasing" paths:
plt.hist([len(traj) for traj in categorized[increasing]], bins, normed=True, alpha=0.5, color='g');
# Let's also check whether we go back and forth between the increasing transition and the decreasing transition, or whether there's just a single change from one type to the other.
def find_switches(ensembles, trajectories):
switches = []
last_category = None
traj_num = 0
for traj in trajectories:
category = None
for ens in ensembles:
if len(ens.split(traj)) > 0:
if category is not None:
category = 'multiple'
else:
category = ens
if last_category != category:
switches.append((category, traj_num))
traj_num += 1
last_category = category
return switches
switches = find_switches([increasing, decreasing], fixed_transition_segments)
print [switch[1] for switch in switches], len(fixed_transition_segments)
# So there are a lot of switches early in the simulation, and then it gets stuck in one state for much longer.
# Even though we know the alanine dipeptide transitions are not particularly rare, this does give us reason to re-check the temperature. First we'll check what the intergrator says its temperature is, then we'll calculate the temperature based on the kinetic energy of every 50th trajectory.
#
# Note that the code below is specific to using the OpenMM engine.
every_50th_trajectory = [step.active[0].trajectory for step in fixed.steps[::50]]
# make a set to remove duplicates, if trajs aren't decorrelated
every_50th_traj_snapshots = list(set(sum(every_50th_trajectory, [])))
# sadly, it looks like that trick with set doesn't do any good here
# +
# this is ugly as sin: we need a better way of doing it (ideally as a snapshot feature)
# dof calculation taken from OpenMM's StateDataReporter
import simtk.openmm as mm
import simtk.unit
dof = 0
system = engine.simulation.system
dofs_from_particles = 0
for i in range(system.getNumParticles()):
if system.getParticleMass(i) > 0*simtk.unit.dalton:
dofs_from_particles += 3
dofs_from_constraints = system.getNumConstraints()
dofs_from_motion_removers = 0
if any(type(system.getForce(i)) == mm.CMMotionRemover for i in range(system.getNumForces())):
dofs_from_motion_removers += 3
dof = dofs_from_particles - dofs_from_constraints - dofs_from_motion_removers
#print dof, "=", dofs_from_particles, "-", dofs_from_constraints, "-", dofs_from_motion_removers
kinetic_energies = []
potential_energies = []
temperatures = []
R = simtk.unit.BOLTZMANN_CONSTANT_kB * simtk.unit.AVOGADRO_CONSTANT_NA
for snap in every_50th_traj_snapshots:
engine.current_snapshot = snap
state = engine.simulation.context.getState(getEnergy=True)
ke = state.getKineticEnergy()
temperatures.append(2 * ke / dof / R)
# -
plt.plot([T / T.unit for T in temperatures])
mean_T = np.mean(temperatures)
plt.plot([mean_T / mean_T.unit]*len(temperatures), 'r')
print "Mean temperature:", np.mean(temperatures).format("%.2f")
| examples/alanine_dipeptide_tps/AD_tps_4_advanced.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''c-m_env'': venv)'
# name: python3
# ---
from seqeval.metrics import accuracy_score
from seqeval.metrics import classification_report
from seqeval.metrics import f1_score
y_true = [['O', 'O', 'O', 'B-MISC', 'I-MISC', 'I-MISC', 'O'], ['B-PER', 'I-PER', 'O']]
y_pred = [['O', 'O', 'B-MISC', 'I-MISC', 'I-MISC', 'I-MISC', 'O'], ['B-PER', 'I-PER', 'O']]
f1_score(y_true, y_pred)
accuracy_score(y_true, y_pred)
print(classification_report(y_true, y_pred))
# +
from typing import List, Dict, Sequence
class Matrics:
def __init__(self, sents_true_labels: Sequence[Sequence[Dict]], sents_pred_labels:Sequence[Sequence[Dict]]):
self.sents_true_labels = sents_true_labels
self.sents_pred_labels = sents_pred_labels
self.types = set(entity['type'] for sent in sents_true_labels for entity in sent)
self.confusion_matrices = {type: {'TP': 0, 'TN': 0, 'FP': 0, 'FN': 0} for type in self.types}
self.scores = {type: {'p': 0, 'r': 0, 'f1': 0} for type in self.types}
def cal_confusion_matrices(self) -> Dict[str, Dict]:
"""Calculate confusion matrices for all sentences."""
for true_labels, pred_labels in zip(self.sents_true_labels, self.sents_pred_labels):
for true_label in true_labels:
entity_type = true_label['type']
prediction_hit_count = 0
for pred_label in pred_labels:
if pred_label['type'] != entity_type:
continue
if pred_label['start_idx'] == true_label['start_idx'] and pred_label['end_idx'] == true_label['end_idx'] and pred_label['text'] == true_label['text']: # TP
self.confusion_matrices[entity_type]['TP'] += 1
prediction_hit_count += 1
elif ((pred_label['start_idx'] == true_label['start_idx']) or (pred_label['end_idx'] == true_label['end_idx'])) and pred_label['text'] != true_label['text']: # boundry error, count FN, FP
self.confusion_matrices[entity_type]['FP'] += 1
self.confusion_matrices[entity_type]['FN'] += 1
prediction_hit_count += 1
if prediction_hit_count != 1: # FN, model cannot make a prediction for true_label
self.confusion_matrices[entity_type]['FN'] += 1
prediction_hit_count = 0 # reset to default
def cal_scores(self) -> Dict[str, Dict]:
"""Calculate precision, recall, f1."""
confusion_matrices = self.confusion_matrices
scores = {type: {'p': 0, 'r': 0, 'f1': 0} for type in self.types}
for entity_type, confusion_matrix in confusion_matrices.items():
if confusion_matrix['TP'] == 0 and confusion_matrix['FP'] == 0:
scores[entity_type]['p'] = 0
else:
scores[entity_type]['p'] = confusion_matrix['TP'] / (confusion_matrix['TP'] + confusion_matrix['FP'])
if confusion_matrix['TP'] == 0 and confusion_matrix['FN'] == 0:
scores[entity_type]['r'] = 0
else:
scores[entity_type]['r'] = confusion_matrix['TP'] / (confusion_matrix['TP'] + confusion_matrix['FN'])
if scores[entity_type]['p'] == 0 or scores[entity_type]['r'] == 0:
scores[entity_type]['f1'] = 0
else:
scores[entity_type]['f1'] = 2*scores[entity_type]['p']*scores[entity_type]['r'] / (scores[entity_type]['p']+scores[entity_type]['r'])
self.scores = scores
def print_confusion_matrices(self):
for entity_type, matrix in self.confusion_matrices.items():
print(f"{entity_type}: {matrix}")
def print_scores(self):
for entity_type, score in self.scores.items():
print(f"{entity_type}: f1 {score['f1']:.4f}, precision {score['p']:.4f}, recall {score['r']:.4f}")
if __name__ == "__main__":
sents_true_labels = [[{'start_idx': 0, 'end_idx': 1, 'text': 'Foreign Ministry', 'type': 'ORG'},
{'start_idx': 3, 'end_idx': 4, 'text': '<NAME>', 'type': 'PER'},
{'start_idx': 6, 'end_idx': 6, 'text': 'Reuters', 'type': 'ORG'}]]
sents_pred_labels = [[{'start_idx': 3, 'end_idx': 3, 'text': 'Shen', 'type': 'PER'},
{'start_idx': 6, 'end_idx': 6, 'text': 'Reuters', 'type': 'ORG'}]]
matrics = Matrics(sents_true_labels, sents_pred_labels)
matrics.cal_confusion_matrices()
matrics.print_confusion_matrices()
matrics.cal_scores()
matrics.print_scores()
# PER: {'TP': 0, 'TN': 0, 'FP': 1, 'FN': 1}
# ORG: {'TP': 1, 'TN': 0, 'FP': 0, 'FN': 1}
# PER: f1 0.0000, precision 0.0000, recall 0.0000
# ORG: f1 0.6667, precision 1.0000, recall 0.5000
# -
| notebooks/002_scipyNER.ipynb |
# ---
# title: "Deep Dream"
# output:
# html_notebook:
# theme: cerulean
# highlight: textmate
# jupyter:
# jupytext:
# cell_metadata_filter: name,tags,-all
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# + name="setup" tags=["remove_cell"]
knitr::opts_chunk$set(warning = FALSE, message = FALSE)
# -
# ***
#
# This notebook contains the code samples found in Chapter 8, Section 2 of [Deep Learning with R](https://www.manning.com/books/deep-learning-with-r). Note that the original text features far more content, in particular further explanations and figures: in this notebook, you will only find source code and related comments.
#
# ***
#
# ## Implementing Deep Dream in Keras
#
# We will start from a convnet pre-trained on ImageNet. In Keras, we have many such convnets available: VGG16, VGG19, Xception, ResNet50... albeit the same process is doable with any of these, your convnet of choice will naturally affect your visualizations, since different convnet architectures result in different learned features. The convnet used in the original Deep Dream release was an Inception model, and in practice Inception is known to produce very nice-looking Deep Dreams, so we will use the InceptionV3 model that comes with Keras.
# +
library(keras)
# We will not be training our model,
# so we use this command to disable all training-specific operations
k_set_learning_phase(0)
# Build the InceptionV3 network.
# The model will be loaded with pre-trained ImageNet weights.
model <- application_inception_v3(
weights = "imagenet",
include_top = FALSE,
)
# -
# Next, we compute the "loss", the quantity that we will seek to maximize during the gradient ascent process. In Chapter 5, for filter visualization, we were trying to maximize the value of a specific filter in a specific layer. Here we will simultaneously maximize the activation of all filters in a number of layers. Specifically, we will maximize a weighted sum of the L2 norm of the activations of a set of high-level layers. The exact set of layers we pick (as well as their contribution to the final loss) has a large influence on the visuals that we will be able to produce, so we want to make these parameters easily configurable. Lower layers result in geometric patterns, while higher layers result in visuals in which you can recognize some classes from ImageNet (e.g. birds or dogs). We'll start from a somewhat arbitrary configuration involving four layers -- but you will definitely want to explore many different configurations later on:
# Named mapping layer names to a coefficient
# quantifying how much the layer's activation
# will contribute to the loss we will seek to maximize.
# Note that these are layer names as they appear
# in the built-in InceptionV3 application.
# You can list all layer names using `summary(model)`.
layer_contributions <- list(
mixed2 = 0.2,
mixed3 = 3,
mixed4 = 2,
mixed5 = 1.5
)
# Now let's define a tensor that contains our loss, i.e. the weighted sum of the L2 norm of the activations of the layers listed above.
# +
# Get the symbolic outputs of each "key" layer (we gave them unique names).
layer_dict <- model$layers
names(layer_dict) <- lapply(layer_dict, function(layer) layer$name)
# Define the loss.
loss <- k_variable(0)
for (layer_name in names(layer_contributions)) {
# Add the L2 norm of the features of a layer to the loss.
coeff <- layer_contributions[[layer_name]]
activation <- layer_dict[[layer_name]]$output
scaling <- k_prod(k_cast(k_shape(activation), "float32"))
loss <- loss + (coeff * k_sum(k_square(activation)) / scaling)
}
# -
# Now we can set up the gradient ascent process:
# +
# This holds our generated image
dream <- model$input
# Normalize gradients.
grads <- k_gradients(loss, dream)[[1]]
grads <- grads / k_maximum(k_mean(k_abs(grads)), 1e-7)
# Set up function to retrieve the value
# of the loss and gradients given an input image.
outputs <- list(loss, grads)
fetch_loss_and_grads <- k_function(list(dream), outputs)
eval_loss_and_grads <- function(x) {
outs <- fetch_loss_and_grads(list(x))
loss_value <- outs[[1]]
grad_values <- outs[[2]]
list(loss_value, grad_values)
}
gradient_ascent <- function(x, iterations, step, max_loss = NULL) {
for (i in 1:iterations) {
c(loss_value, grad_values) %<-% eval_loss_and_grads(x)
if (!is.null(max_loss) && loss_value > max_loss)
break
cat("...Loss value at", i, ":", loss_value, "\n")
x <- x + (step * grad_values)
}
x
}
# -
# Finally, here is the actual Deep Dream algorithm.
#
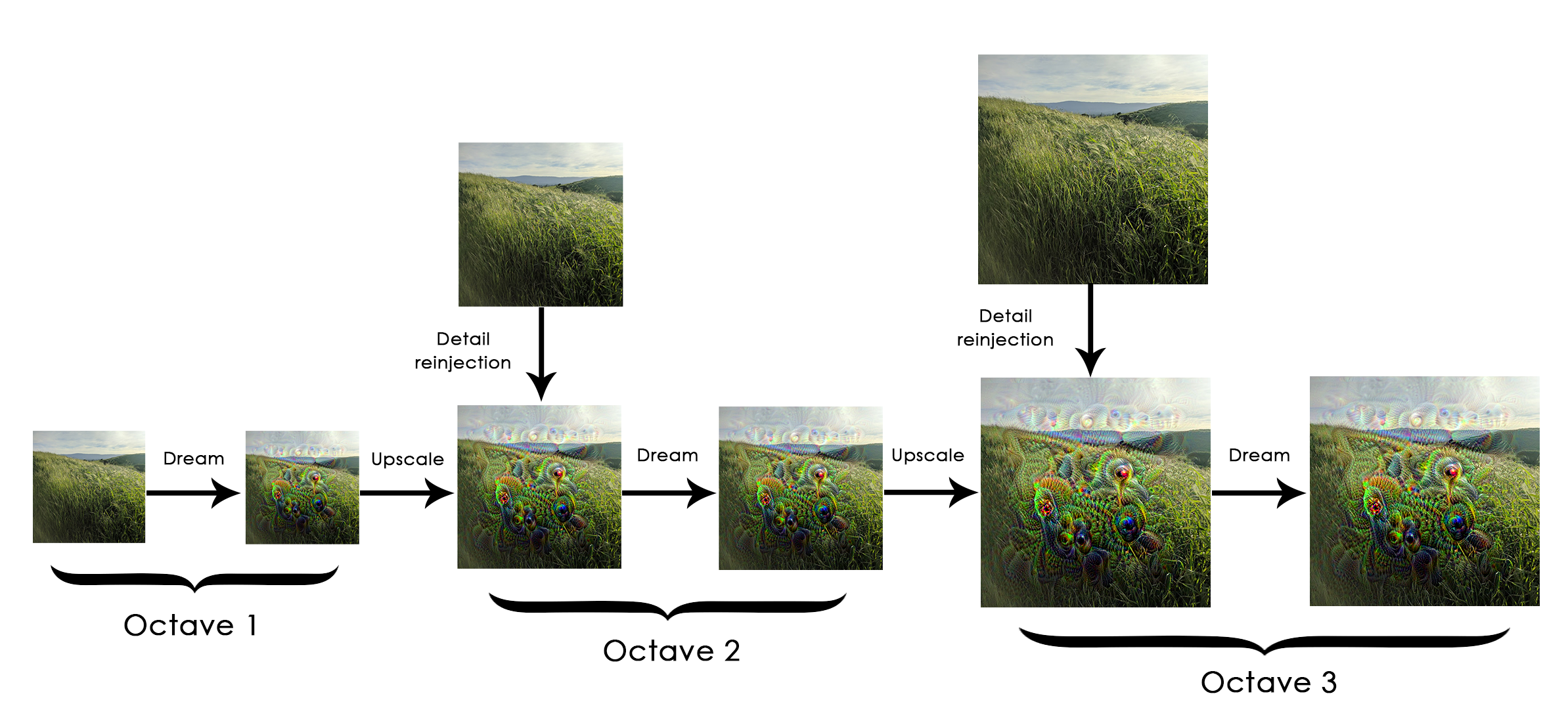
# First, we define a list of "scales" (also called "octaves") at which we will process the images. Each successive scale is larger than previous one by a factor 1.4 (i.e. 40% larger): we start by processing a small image and we increasingly upscale it:
#
# 
#
# Then, for each successive scale, from the smallest to the largest, we run gradient ascent to maximize the loss we have previously defined, at that scale. After each gradient ascent run, we upscale the resulting image by 40%.
#
# To avoid losing a lot of image detail after each successive upscaling (resulting in increasingly blurry or pixelated images), we leverage a simple trick: after each upscaling, we reinject the lost details back into the image, which is possible since we know what the original image should look like at the larger scale. Given a small image S and a larger image size L, we can compute the difference between the original image (assumed larger than L) resized to size L and the original resized to size S -- this difference quantifies the details lost when going from S to L.
# +
resize_img <- function(img, size) {
image_array_resize(img, size[[1]], size[[2]])
}
save_img <- function(img, fname) {
img <- deprocess_image(img)
image_array_save(img, fname)
}
# Util function to open, resize, and format pictures into appropriate tensors
preprocess_image <- function(image_path) {
image_load(image_path) %>%
image_to_array() %>%
array_reshape(dim = c(1, dim(.))) %>%
inception_v3_preprocess_input()
}
# Util function to convert a tensor into a valid image
deprocess_image <- function(img) {
img <- array_reshape(img, dim = c(dim(img)[[2]], dim(img)[[3]], 3))
img <- img / 2
img <- img + 0.5
img <- img * 255
dims <- dim(img)
img <- pmax(0, pmin(img, 255))
dim(img) <- dims
img
}
# +
# Playing with these hyperparameters will also allow you to achieve new effects
step <- 0.01 # Gradient ascent step size
num_octave <- 3 # Number of scales at which to run gradient ascent
octave_scale <- 1.4 # Size ratio between scales
iterations <- 20 # Number of ascent steps per scale
# If our loss gets larger than 10,
# we will interrupt the gradient ascent process, to avoid ugly artifacts
max_loss <- 10
# Fill this to the path to the image you want to use
dir.create("dream")
base_image_path <- "~/Downloads/creative_commons_elephant.jpg"
# Load the image into an array
img <- preprocess_image(base_image_path)
# We prepare a list of shapes
# defining the different scales at which we will run gradient ascent
original_shape <- dim(img)[-1]
successive_shapes <- list(original_shape)
for (i in 1:num_octave) {
shape <- as.integer(original_shape / (octave_scale ^ i))
successive_shapes[[length(successive_shapes) + 1]] <- shape
}
# Reverse list of shapes, so that they are in increasing order
successive_shapes <- rev(successive_shapes)
# Resize the array of the image to our smallest scale
original_img <- img
shrunk_original_img <- resize_img(img, successive_shapes[[1]])
for (shape in successive_shapes) {
cat("Processsing image shape", shape, "\n")
img <- resize_img(img, shape)
img <- gradient_ascent(img,
iterations = iterations,
step = step,
max_loss = max_loss)
upscaled_shrunk_original_img <- resize_img(shrunk_original_img, shape)
same_size_original <- resize_img(original_img, shape)
lost_detail <- same_size_original - upscaled_shrunk_original_img
img <- img + lost_detail
shrunk_original_img <- resize_img(original_img, shape)
save_img(img, fname = sprintf("dream/at_scale_%s.png",
paste(shape, collapse = "x")))
}
save_img(img, fname = "dream/final_dream.png")
# -
plot(as.raster(deprocess_image(img) / 255))
| notebooks/8.2-deep-dream.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cflows
# language: python
# name: cflows
# ---
# ## Config
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '0'
# +
# %load_ext autoreload
# %autoreload 2
from pathlib import Path
from experiment import data_path, device
model_name = 'celeba-cef-joint'
checkpoint_path = data_path / 'cef_models' / model_name
gen_path = data_path / 'generated' / model_name
# -
# ## Load data
# +
import torchvision
from torch.utils.data import DataLoader
from torchvision import transforms
import data
transform = transforms.Compose([
transforms.Resize((64, 64)),
transforms.ToTensor(),
])
train_data = data.CelebA(root=data_path, split='train', transform=transform)
val_data = data.CelebA(root=data_path, split='valid', transform=transform)
test_data = data.CelebA(root=data_path, split='test', transform=transform)
# -
# ## Define model
# +
from nflows import cef_models
flow = cef_models.CelebACEFlow().to(device)
# -
# ## Train
# +
import torch.optim as opt
from experiment import train_injective_flow
optim = opt.Adam(flow.parameters(), lr=0.0001)
scheduler = opt.lr_scheduler.CosineAnnealingLR(optim, 300)
def weight_schedule():
for _ in range(300):
yield 0.001, 10000
scheduler.step()
train_loader = DataLoader(train_data, batch_size=256, shuffle=True, num_workers=30)
val_loader = DataLoader(val_data, batch_size=256, shuffle=True, num_workers=30)
train_injective_flow(flow, optim, scheduler, weight_schedule, train_loader, val_loader,
model_name, checkpoint_path=checkpoint_path, checkpoint_frequency=25)
# -
# ## Generate some samples
# +
from experiment import save_samples
save_samples(flow, num_samples=len(test_data), gen_path=gen_path, checkpoint_epoch=-1, batch_size=512)
| experiments/celeba-cef-joint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf_gpu
# language: python
# name: tf_gpu
# ---
# # 4.2. OpenAI Gym Environments
# ## Gym basics and a simple text environment
# +
import gym # pip install gym, pip install gym[atari]
from IPython import display
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
env = gym.make('Taxi-v3')
env.reset()
# -
env.P # {state: {action: [(probability, next state, reward, done)]}}
for step in range(100):
action = env.action_space.sample()
new_state, reward, done, info = env.step(action)
clear_output(wait=True)
print(env.render(mode='ansi'))
print(f'Timestep: {step + 1}')
print(f'State: {new_state}')
print(f'Action: {action}')
print(f'Reward: {reward}')
sleep(0.2)
| Section 4/Video 4.2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Tutorial 3: Realism and Complexity
# ==================================
#
# Up to now, we've fitted some fairly crude and unrealistic lens models. For example, we've modeled the lens `Galaxy`'s
# mass as a sphere. Given most lens galaxies are `elliptical`s we should probably model their mass as elliptical! We've
# also omitted the lens `Galaxy`'s light, which typically outshines the source galaxy.
#
# In this example, we'll start using a more realistic lens model.
#
# In my experience, the simplest lens model (e.g. that has the fewest parameters) that provides a good fit to real
# strong lenses is as follows:
#
# 1) An _EllipticalSersic `LightProfile` for the lens `Galaxy`'s light.
# 2) A `EllipticalIsothermal` (SIE) `MassProfile` for the lens `Galaxy`'s mass.
# 3) An `EllipticalExponential` `LightProfile`.for the source-`Galaxy`'s light (to be honest, this is too simple,
# but lets worry about that later).
#
# This has a total of 18 non-linear parameters, which is over double the number of parameters we've fitted up to now.
# In future exercises, we'll fit even more complex models, with some 20-30+ non-linear parameters.
# +
# %matplotlib inline
from pyprojroot import here
workspace_path = str(here())
# %cd $workspace_path
print(f"Working Directory has been set to `{workspace_path}`")
from os import path
import autolens as al
import autolens.plot as aplt
import autofit as af
# -
# we'll use new strong lensing data, where:
#
# - The lens `Galaxy`'s `LightProfile` is an `EllipticalSersic`.
# - The lens `Galaxy`'s total mass distribution is an `EllipticalIsothermal`.
# - The source `Galaxy`'s `LightProfile` is an `EllipticalExponential`.
# +
dataset_name = "light_sersic__mass_sie__source_exp"
dataset_path = path.join("dataset", "howtolens", "chapter_2", dataset_name)
imaging = al.Imaging.from_fits(
image_path=path.join(dataset_path, "image.fits"),
noise_map_path=path.join(dataset_path, "noise_map.fits"),
psf_path=path.join(dataset_path, "psf.fits"),
pixel_scales=0.1,
)
# -
# we'll create and use a 2.5" `Mask2D`.
mask = al.Mask2D.circular(
shape_2d=imaging.shape_2d, pixel_scales=imaging.pixel_scales, radius=2.5
)
# When plotted, the lens light`s is clearly visible in the centre of the image.
aplt.Imaging.subplot_imaging(imaging=imaging, mask=mask)
# Like in the previous tutorial, we use a `SettingsPhaseImaging` object to specify our model-fitting procedure uses a
# regular `Grid`.
# +
settings_masked_imaging = al.SettingsMaskedImaging(grid_class=al.Grid, sub_size=2)
settings = al.SettingsPhaseImaging(settings_masked_imaging=settings_masked_imaging)
# -
# Now lets fit the dataset using a phase.
phase = al.PhaseImaging(
search=af.DynestyStatic(
path_prefix="howtolens",
name="phase_t3_realism_and_complexity",
n_live_points=80,
),
settings=settings,
galaxies=af.CollectionPriorModel(
lens_galaxy=al.GalaxyModel(
redshift=0.5, bulge=al.lp.EllipticalSersic, mass=al.mp.EllipticalIsothermal
),
source_galaxy=al.GalaxyModel(redshift=1.0, bulge=al.lp.EllipticalExponential),
),
)
# Lets run the phase.
# +
print(
"Dynesty has begun running - checkout the autolens_workspace/output/3_realism_and_complexity"
" folder for live output of the results, images and lens model."
" This Jupyter notebook cell with progress once Dynesty has completed - this could take some time!"
)
result = phase.run(dataset=imaging, mask=mask)
print("Dynesty has finished run - you may now continue the notebook.")
# -
# And lets look at the fit to the `Imaging` data, which as we are used to fits the data brilliantly!
aplt.FitImaging.subplot_fit_imaging(fit=result.max_log_likelihood_fit)
# Up to now, all of our non-linear searches have been successes. They find a lens model that provides a visibly good fit
# to the data, minimizing the residuals and inferring a high log likelihood value.
#
# These solutions are called `global` maxima, they correspond to the highest likelihood regions of the entirity of
# parameter space. There are no other lens models in parameter space that would give higher likelihoods - this is the
# model we wants to always infer!
#
# However, non-linear searches may not always successfully locate the global maxima lens models. They may instead infer
# a `local maxima`, a solution which has a high log likelihood value relative to the lens models near it in parameter
# space, but whose log likelihood is significantly below the `global` maxima solution somewhere else in parameter space.
#
# Inferring such solutions is essentially a failure of our `NonLinearSearch` and it is something we do not want to
# happen! Lets infer a local maxima, by reducing the number of `live points` Dynesty uses to map out parameter space.
# we're going to use so few that it has no hope of locating the global maxima, ultimating finding and inferring a local
# maxima instead.
# +
phase = al.PhaseImaging(
search=af.DynestyStatic(
path_prefix="howtolens",
name="phase_t3_realism_and_complexity__local_maxima",
n_live_points=5,
),
settings=settings,
galaxies=af.CollectionPriorModel(
lens_galaxy=al.GalaxyModel(
redshift=0.5, bulge=al.lp.EllipticalSersic, mass=al.mp.EllipticalIsothermal
),
source_galaxy=al.GalaxyModel(redshift=1.0, bulge=al.lp.EllipticalExponential),
),
)
print(
"Dynesty has begun running - checkout the autolens_workspace/output/3_realism_and_complexity"
" folder for live output of the results, images and lens model."
" This Jupyter notebook cell with progress once Dynesty has completed - this could take some time!"
)
result_local_maxima = phase.run(dataset=imaging, mask=mask)
print("Dynesty has finished run - you may now continue the notebook.")
# -
# And lets look at the fit to the `Imaging` data, which is clearly worse than our original fit above.
aplt.FitImaging.subplot_fit_imaging(fit=result_local_maxima.max_log_likelihood_fit)
# Finally, just to be sure we hit a local maxima, lets compare the maximum log likelihood values of the two results
#
# The local maxima value is significantly lower, confirming that our `NonLinearSearch` simply failed to locate lens
# models which fit the data better when it searched parameter space.
print("Likelihood of Global Model:")
print(result.max_log_likelihood_fit.log_likelihood)
print("Likelihood of Local Model:")
print(result_local_maxima.max_log_likelihood_fit.log_likelihood)
# In this example, we intentionally made our `NonLinearSearch` fail, by using so few live points it had no hope of
# sampling parameter space thoroughly. For modeling real lenses we wouldn't do this on purpose, but the risk of inferring
# a local maxima is still very real, especially as we make our lens model more complex.
#
# Lets think about *complexity*. As we make our lens model more realistic, we also made it more complex. For this
# tutorial, our non-linear parameter space went from 7 dimensions to 18. This means there was a much larger *volume* of
# parameter space to search. As this volume grows, there becomes a higher chance that our `NonLinearSearch` gets lost
# and infers a local maxima, especially if we don't set it up with enough live points!
#
# At its core, lens modeling is all about learning how to get a `NonLinearSearch` to find the global maxima region of
# parameter space, even when the lens model is extremely complex.
#
# And with that, we're done. In the next exercise, we'll learn how to deal with failure and begin thinking about how we
# can ensure our `NonLinearSearch` finds the global-maximum log likelihood solution. Before that, think about
# the following:
#
# 1) When you look at an image of a strong lens, do you get a sense of roughly what values certain lens model
# parameters are?
#
# 2) The `NonLinearSearch` failed because parameter space was too complex. Could we make it less complex, whilst
# still keeping our lens model fairly realistic?
#
# 3) The source galaxy in this example had only 7 non-linear parameters. Real source galaxies may have multiple
# components (e.g. a bar, disk, bulge, star-forming knot) and there may even be more than 1 source galaxy! Do you
# think there is any hope of us navigating a parameter space if the source contributes 20+ parameters by itself?
| howtolens/chapter_2_lens_modeling/tutorial_3_realism_and_complexity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
p = [2, 4, 10, 6, 8, 4]
max0, min0 = max(p), min(p)
diff = abs(max0 - min0)
# p[0] = (p[0] - min0) / diff
for i in range(len(p)):
p[i] = (p[i] - min0) / diff
print(p)
# -
nmax = 100
for n in range(1, nmax + 1):
message = ''
if not n % 3:
message = 'Fizz'
if not n % 5:
message += 'Buzz'
print(message or n)
| ue7/ue7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import datetime, time
import simpy
import shapely.geometry
import pandas as pd
import openclsim.core as core
import openclsim.model as model
import datetime
import numpy as np
# +
simulation_start = datetime.datetime.now()
my_env = simpy.Environment(initial_time=time.mktime(simulation_start.timetuple()))
my_env.epoch = time.mktime(simulation_start.timetuple())
registry = {}
keep_resources = {}
# +
Site = type(
"Site",
(
core.Identifiable,
core.Log,
core.Locatable,
core.HasContainer,
core.HasResource,
),
{},
)
TransportProcessingResource = type(
"TransportProcessingResource",
(
core.Identifiable,
core.Log,
core.ContainerDependentMovable,
core.Processor,
core.HasResource,
core.LoadingFunction,
core.UnloadingFunction,
),
{},
)
# +
location_from_site = shapely.geometry.Point(5.1, 52)
location_to_site = shapely.geometry.Point(5, 52.1)
location_to_site_2 = shapely.geometry.Point(5, 52.2)
data_from_site = {
"env": my_env,
"name": "Winlocatie",
"geometry": location_from_site,
"capacity": 100,
"level": 100,
}
data_to_site = {
"env": my_env,
"name": "Dumplocatie",
"geometry": location_to_site,
"capacity": 55,
"level": 0,
}
from_site = Site(**data_from_site)
to_site = Site(**data_to_site)
# +
data_cutter = {
"env": my_env,
"name": "Cutter_1",
"geometry": location_from_site,
"capacity": 5,
"compute_v": lambda x: 10,
"loading_rate": 3600/5,
"unloading_rate": 3600/5
}
data_barge_1 = {
"env": my_env,
"name": "Barge_1",
"geometry": location_to_site,
"capacity": 5,
"compute_v": lambda x: 10,
"loading_rate": 3600/5,
"unloading_rate": 3600/5
}
data_barge_2 = {
"env": my_env,
"name": "Barge_2",
"geometry": location_to_site,
"capacity": 5,
"compute_v": lambda x: 10,
"loading_rate": 3600/5,
"unloading_rate": 3600/5
}
cutter = TransportProcessingResource(**data_cutter)
barge_1 = TransportProcessingResource(**data_barge_1)
barge_2 = TransportProcessingResource(**data_barge_2)
# +
requested_resources = {}
single_run = [
model.MoveActivity(**{
"env": my_env,
"name": "sailing empty_1",
"registry": registry,
"mover": barge_1,
"destination": from_site,
"postpone_start": True,
}),
model.ShiftAmountActivity(**{
"env": my_env,
"name": "Transfer MP_1",
"registry": registry,
"processor": cutter,
"origin": from_site,
"destination": barge_1,
"amount": 5,
"duration": 3600,
"postpone_start": True,
# "requested_resources":requested_resources
}),
model.MoveActivity(**{
"env": my_env,
"name": "sailing filled_1",
"registry": registry,
"mover": barge_1,
"destination": to_site,
"postpone_start": True,
}),
model.ShiftAmountActivity(**{
"env": my_env,
"name": "Transfer TP_1",
"registry": registry,
"processor": barge_1,
"origin": barge_1,
"destination": to_site,
"amount": 5,
"duration": 3600,
"postpone_start": True,
# "requested_resources":requested_resources
})
]
sequential_activity_data = {
"env": my_env,
"name": "Single run process_1",
"registry": registry,
"sub_processes": single_run,
"postpone_start": True,
}
activity = model.SequentialActivity(**sequential_activity_data)
while_activity = model.WhileActivity(**{
"env": my_env,
"name": "while_1",
"registry": registry,
"sub_processes": [activity],
"condition_event": [{"type":"container", "concept": to_site, "state":"full"}],
"postpone_start": False,
})
# +
single_run = [
model.MoveActivity(**{
"env": my_env,
"name": "sailing empty_2",
"registry": registry,
"mover": barge_2,
"destination": from_site,
"postpone_start": True,
}),
model.ShiftAmountActivity(**{
"env": my_env,
"name": "Transfer MP_2",
"registry": registry,
"processor": cutter,
"origin": from_site,
"destination": barge_2,
"amount": 5,
"duration": 3600,
"postpone_start": True,
}),
model.MoveActivity(**{
"env": my_env,
"name": "sailing filled_2",
"registry": registry,
"mover": barge_2,
"destination": to_site,
"postpone_start": True,
}),
model.ShiftAmountActivity(**{
"env": my_env,
"name": "Transfer TP_2",
"registry": registry,
"processor": barge_2,
"origin": barge_2,
"destination": to_site,
"amount": 5,
"duration": 3600,
"postpone_start": True,
})
]
activity = model.SequentialActivity(** {
"env": my_env,
"name": "Single run process_2",
"registry": registry,
"sub_processes": single_run,
"postpone_start": True,
# "keep_resources"
})
while_activity = model.WhileActivity(**{
"name": "while_2",
"env": my_env,
"registry": registry,
"sub_processes": [activity],
"condition_event": [{"type":"container", "concept": to_site, "state":"full"}],
"postpone_start": False,
})
# -
my_env.run()
# +
# !! BUGG !!
# why we need reservations:
# !! BUGG !!
print(f"Level of the to_site: {to_site.container.get_level()}, Capacity of the to_site {to_site.container.get_capacity()}")
# -
| notebooks/17_cutters and barges (BUG IN RESERVATIONS).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.2 64-bit (''venv'': venv)'
# name: python3
# ---
import pandas as pd
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
data = pd.read_csv("./voiceDataSet.csv")
data.label = data.label.apply(lambda x: x=="male").astype('int32')
# +
# X = data.iloc[:,:-1]
# Y = data.iloc[:,-1:]
y = data['label']
X = data.drop(['label'], axis = 1)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
# -
scaler = StandardScaler()
scaler.fit(X_train)
X_train = scaler.transform(X_train)
X_test = scaler.transform(X_test)
model = LogisticRegression()
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
accuracy_score(y_test, y_pred)
| unit_4/ml_4.7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from math import *
import numpy as np
from scipy.integrate import odeint
import matplotlib.pyplot as plot
n=3.5 #разница в скорости
s=8.5 #расстояние обнаружения
fi=pi*3/4 #угол движения
# +
def f(tetha, r): #уравнение катера
dr=r/sqrt(n**2 - 1)
return dr
def f2(t): #лодка браконьеров
xt = tan(fi+pi)*t
return xt
# +
r0=s/(n+1) #первый случай
#решение диф уравнения для катера
tetha = np.arange(0, 2*pi, 0.01)
r = odeint(f, r0, tetha)
#вычисление траектории лодки
t=np.arange(0.00000000000001, 20)
r1=np.sqrt(t**2 + f2(t)**2)
tetha1=np.arctan(f2(t)/t)
plot.rcParams["figure.figsize"] = (10, 10)
plot.polar(tetha, r, 'red', label = 'катер')
plot.polar(tetha1, r1, 'green', label = 'лодка')
#вычисление точки пересечения
tmp=0
for i in range(len(tetha)):
if round(tetha[i], 2) == round(fi+pi, 2):
tmp=i
print("Тета:", tetha[tmp], "r:", r[tmp][0])
print("X:", r[tmp][0]/sqrt(2), "Y:", -r[tmp][0]/sqrt(2))
plot.legend()
plot.savefig("01.png",dpi=100)
# +
r0=s/(n-1) #второй случай
#решение диф уравнения для катера
tetha = np.arange(0, 2*pi, 0.01)
r = odeint(f, r0, tetha)
#вычисление траектории лодки
t=np.arange(0.00000000000001, 20)
r1=np.sqrt(t**2 + f2(t)**2)
tetha1=np.arctan(f2(t)/t)
plot.rcParams["figure.figsize"] = (8, 8)
plot.polar(tetha, r, 'red', label = 'катер')
plot.polar(tetha1, r1, 'green', label = 'лодка')
#вычисление точки пересечения
tmp=0
for i in range(len(tetha)):
if round(tetha[i], 2) == round(fi+pi, 2):
tmp=i
print("Тета:", tetha[tmp], "r:", r[tmp][0])
print("X:", r[tmp][0]/sqrt(2), "Y:", -r[tmp][0]/sqrt(2))
plot.legend()
plot.savefig("02.png",dpi=100)
# -
| Lab-02/lab02-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## LAMBDA FUNCTION
#
# anonymous funtion,
#
# a funtion with no name
# +
# faster than normal funtion
# -
def add(a,b):
return a+b
add(4,9)
# +
# we can use lambda function
# we can create lambda funtion only if there is only single expression
# -
add = lambda a,b: a+b
# +
# remember that here add is variable, not function
# -
add(3,4)
def even(num):
if num%2==0:
return True
even(2)
even1 =lambda a: a%2==0
even1(3)
def addition(x,y,z):
return x,y,z
addition(1,2,3)
addition = lambda x,y,z: x+y+z
addition(1,2,3)
| pypractise11.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Imports
import random
import sys
import numpy as np
import time
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import torch.nn as nn
import torch.optim as optim
import torchvision.utils as vutils
import matplotlib.animation as animation
from IPython.display import HTML
import model_v4_small as model
import torch
import keijzer_exogan as ke
# initialize random seeds
manualSeed = 999
random.seed(manualSeed)
torch.manual_seed(manualSeed)
# %matplotlib inline
# %config InlineBackend.print_figure_kwargs={'facecolor' : "w"} # Make sure the axis background of plots is white, this is usefull for the black theme in JupyterLab
# Initialize default seaborn layout
sns.set_palette(sns.hls_palette(8, l=.3, s=.8))
sns.set(style='ticks')
"""
Local variables
"""
workers = 0 # Number of workers for dataloader, 0 when to_vram is enabled
batch_size = 1 # using one image ofcourse
image_size = 32
nz = 100 # size of latent vector
n_iters = 2000 #25*10**3 # number of iterations to do for inpainting
torch.backends.cudnn.benchmark=True # Uses udnn auto-tuner to find the best algorithm to use for your hardware, speeds up training by almost 50%
lr = 1e-1
lamb1 = 1 #1e4
lamb2 = 1e-1 # 1 , total_loss = lamb1*loss_context + lamb2*loss_perceptual
beta1 = 0.5 # Beta1 hyperparam for Adam optimizers
selected_gpus = [4] # Number of GPUs available. Use 0 for CPU mode.
#n_images = 500
inpaint_n_times = 1
save_array_results = True
load_array_results = False
filename = '0_25_1e-1_wgan_v4_small_complex' # 0:100 lamb1=10, lamb2=1
# debug_0_5000_1_1e-1_* c is exogan data with original brian mask, d is with binary mask
# -
#path = '/datb/16011015/MakeAI/ASPAs//' #notice how you dont put the last folder in here...
path = '/shared/datasets/16011015/MakeAI_SURFsara/ASPAs/' # gpu server
# # Load smaller selection of ASPAs
# +
images = np.load(path+'MakeAI_test_complex_2.npy').astype('float32') # 4.1 is a random selection of 5k images
print('Loaded %s images' % len(images))
print('Batch size: ', batch_size)
# Number of training epochs
# Learning rate for optimizers
ngpu = len(selected_gpus)
print('Number of GPUs used: ', ngpu)
"""
Load data and prepare DataLoader
"""
shuffle = False
if shuffle:
np.random.shuffle(images) # shuffles the images
images = images[0:25] # 200 should take ~ 11 hours
n_images = len(images)
#images = images[:int(len(images)*0.005)]
print('Number of images: ', n_images)
dataset = ke.numpy_dataset(data=images, to_vram=True) # to_vram pins it to all GPU's
#dataset = numpy_dataset(data=images, to_vram=True, transform=transforms.Compose([transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])) # to_vram pins it to all GPU's
# Create the dataloader
dataloader = torch.utils.data.DataLoader(dataset, batch_size=batch_size,
shuffle=True, num_workers=workers, pin_memory=False)
# +
"""
Load and setup models
"""
# Initialize cuda
device = torch.device("cuda:"+str(selected_gpus[0]) if (torch.cuda.is_available() and ngpu > 0) else "cpu")
# Load models, set to evaluation mode since training is not needed (this also allows batchsize 1 to work with batchnorm2d layers)
netG = model.Generator(ngpu).eval().to(device)
netD = model.Discriminator(ngpu).eval().to(device)
# Apply weights
print('Loading weights...')
try:
# Load saved weights
netG.load_state_dict(torch.load('gan_data//weights//netG_complex_22', map_location=device)) #net.module..load_... for parallel model , net.load_... for single gpu model
netD.load_state_dict(torch.load('gan_data//weights//netD_complex_22', map_location=device))
except:
print('Could not load saved weights.')
sys.exit()
"""
Define input training stuff (fancy this up)
"""
G = netG
D = netD
z = torch.randn(1, nz, 1, 1, requires_grad=True, device=device)
# Handle multi-gpu if desired
if (device.type == 'cuda') and (ngpu > 1):
G = nn.DataParallel(G, device_ids=selected_gpus, output_device=device)
D = nn.DataParallel(D, device_ids=selected_gpus, output_device=device)
#z = nn.DataParallel(z, device_ids=selected_gpus, output_device=device)
# Setup Adam optimizers for both G and D
optimizerD = optim.Adam(netD.parameters(), lr=lr, betas=(beta1, 0.999)) # should be sgd
optimizerG = optim.Adam(netG.parameters(), lr=lr, betas=(beta1, 0.999))
print('done')
# -
# # Show generated images
# +
from sklearn.preprocessing import MinMaxScaler
z_tests = [torch.randn(1, nz, 1, 1, device=device) for _ in range(9)]
plt.figure(figsize=(10,10))
for i in range(9):
img = G(z_tests[i]).detach().cpu()[0, 0, :, :]
plt.subplot(3,3,i+1)
#scaler = MinMaxScaler((0, 1.2))
#img = scaler.fit_transform(img)
plt.imshow(img, cmap='gray', vmin=-1.2, vmax=1.2)
#plt.imshow(img, cmap='gray')
plt.tight_layout()
img.min(), img.max(), img.mean(), img.std()
# -
# # Show first 9 selected images
# +
plt.figure(figsize=(10,10))
for i in range(9):
try:
img = images[i]
plt.subplot(3,3,i+1)
plt.imshow(img[0, :, :], cmap='gray', vmin=-1.2, vmax=1.2)
except:
pass
plt.tight_layout()
img.min(), img.max(), img.mean(), img.std()
# -
# # Visualizing the weights
# +
weights = [param.data.cpu().numpy().flatten() for param in netD.parameters()]
plt.figure(figsize=(10,10))
for i,layer_weights in enumerate(weights):
print('Layer: %s \t n_weights: %s \t std: %.4f \t mean: %.4f' % (i, len(layer_weights), layer_weights.std(), layer_weights.mean()))
plt.subplot(3,2,i+1)
plt.title('netD layer %s weights' % i)
plt.hist(layer_weights, bins=100)
plt.grid()
plt.tight_layout()
# +
weights = [param.data.cpu().numpy().flatten() for param in netG.parameters()] # where param.data are the weights of the i-th layer
plt.figure(figsize=(10,10))
for i,layer_weights in enumerate(weights):
print('Layer: %s \t n_weights: %s \t std: %.4f \t mean: %.4f' % (i, len(layer_weights), layer_weights.std(), layer_weights.mean()))
plt.subplot(3,2,i+1)
plt.title('netG layer %s weights' % i)
plt.hist(layer_weights, bins=100)
plt.grid()
plt.tight_layout()
# -
# # Inpainting
# The corrupted image $y$ is mapped to the closest $z$ in the latent representation space, this mapping is denoted as $\hat{z}$.
#
# $\hat{z} = \operatorname{arg\,min}_z \{ \mathcal{L}_c(z |y, M) + \mathcal{L}_p (z) \}$
#
# where
#
# $\mathcal{L}_c(z |y, M) = || M \bigodot G(z) - M \bigodot y||_1 = || M \bigodot (G(z)-y) ||_1 $
#
# with $\mathcal{L}_c$ being contextual loss and $M$ being a binary mask with the same size as $y$,
#
# $\mathcal{L}_p (z) = \lambda \operatorname{log}(1-D(G(z)))$
#
# with $\mathcal{L}_p$ being perceptual loss and $D$ being the discriminator.
#
# Once $G(\hat{z})$ is generated, the final solution $\hat{x}$ is calculated as
#
# $\hat{x} = \operatorname{arg\, min}_x ||\nabla x - \nabla G(\hat{z}) ||^2_2$
#
# (substitute $x_i = y_i$ for $M_i = 1$).
#
# -----
#
# $|| ... ||$ is done by `torch.norm()`.
# $... \bigodot ...$ is done by `torch.mul()`.
# -----
# TODO: Implement $\hat{x} = \operatorname{arg\, min}_x ||\nabla x - \nabla G(\hat{z}) ||^2_2$
# Currently $\hat{x} = G(\hat{z}) \bigodot (1 -M)+y$
# ## Create the mask
def create_mask():
mask = np.full([1,1,32,32], 1) # init array with 0.5's
mask = torch.from_numpy(mask).to(device)
#mask = torch.ones([1, 1, 32, 32]).to(device) # create mask with 1's in the shape of image
#print("mask.shape", mask.shape)
# use a random 'easy' mask
# set all params to 0
mask[:, :, :16, 25:] = 0
# set noise to 0
mask[:, :, 18:, :] = 0
"""Weighted mask"""
# Normalization factors
mask[:, :, 16:18, :] = 6 #6
# Planet mass
mask[:, :, :16, 25:26] = 0
mask = mask.float() # make sure dtype is float, torch was complaining during inpainting that this is a double
return mask
m = create_mask().cpu()[0, 0, :, :]
plt.imshow(m, cmap='gray', vmin=0, vmax=2)
# # Inpaiting functions
def save_inpainting_results():
# save real aspa's
all_reals = []
for selected_aspa in range(len(real_images)):
reals = np.array([real_images[selected_aspa][i].detach().cpu().numpy()[0, 0, :, :] for i in range(inpaint_n_times)])
all_reals.append(reals)
all_reals = np.array(all_reals)
np.save('gan_data//val_errors//'+filename+'_reals.npy', all_reals)
# save inpained aspa's
all_inpainteds = []
for selected_aspa in range(len(real_images)):
inpainteds = np.array([final_inpainted_images[selected_aspa][i].detach().cpu().numpy()[0, 0, :, :] for i in range(inpaint_n_times)])
all_inpainteds.append(inpainteds)
all_inpainteds = np.array(all_inpainteds)
np.save('gan_data//val_errors//'+filename+'_inpainteds.npy', all_inpainteds)
np.save('gan_data//val_errors//'+filename+'_n_iterations.npy', n_iteration)
np.save('gan_data//val_errors//'+filename+'_contextual_losses.npy', contextual_losses)
np.save('gan_data//val_errors//'+filename+'_perceptual_losses.npy', perceptual_losses)
return
# ## Inpainting loop
# 22.33 iters / s
# +
# Lists to keep track of progress
real_images = []
masked_images= []
#inpainted_images = []
final_inpainted_images = [] # last n inpainted images, one index location for each input image [[aspa1, aspa1, aspa1], [aspa2,aspa2,aspa2]] .... where aspa1, aspa1, aspa1 are 3 unique inpaintings
n_iteration = []
perceptual_losses = []
contextual_losses = []
MSELoss = nn.MSELoss()
L1Loss = nn.L1Loss() # MAE
SmoothL1Loss = nn.SmoothL1Loss()
"""
Inpainting
"""
t3 = time.time()
past_s_image = 0
for i, data in enumerate(dataloader, 0): # batches per epoch
real_images_n_times = []
final_inpainted_images_n_times = [] # list of (n) last inpainted image(s), for one aspa
t1 = time.time()
for j in range(inpaint_n_times): # inpaint n times per image
z = torch.randn(1, nz, 1, 1, requires_grad=True, device=device)
opt = optim.Adam([z], lr=lr)
real_cpu = data.to(device)
b_size = real_cpu.size(0) # this is one ofc, it's one image we're trying to inpaint
#print("data.shape: ", data.shape)
image = data.to(device) # select the image (Channel, Height, Width), this is the original unmasked input image
real_images_n_times.append(image)
#print("image.shape: ", image.shape)
"""Mask the image"""
mask = create_mask()
masked_image = torch.mul(image, mask).to(device) #image bigodot mask
masked_images.append(masked_image)
#print('masked image shape', masked_image.shape)
#plt.imshow(masked_image.detach().cpu()[0, 0, :, :], cmap='gray') # plot the masked image
# what's v and m?
v = torch.tensor(0, dtype=torch.float32, device=device)
m = torch.tensor(0, dtype=torch.float32, device=device)
"""Start the inpainting process"""
early_stopping_n_iters = 0
early_stopping_min_loss = 999999999999 # set to random high number to initialize
if j != 0:
n_iteration.append(iteration)
for iteration in range(n_iters):
if z.grad is not None:
z.grad.data.zero_()
G.zero_grad()
D.zero_grad()
image_generated = G(z) # generated image G(z)
image_generated_masked = torch.mul(image_generated, mask) # G(z) bigodot M
image_generated_inpainted = torch.mul(image_generated, (1-mask))+masked_image
#if (iteration % 100 == 0):
# inpainted_images.append(image_generated_inpainted)
#print("image_generated_inpainted.shape : ",image_generated_inpainted.shape)
t = image_generated_inpainted.detach().cpu()[0, 0, :, :]
# TODO: why does this already look real?
#plt.imshow(t, cmap='gray') # plot the masked image
"""Calculate losses"""
loss_context = lamb1*torch.norm(image_generated_masked-masked_image, p=1) #what's p=1?
#loss_context = lamb1*MSELoss(image_generated_masked,masked_image)
#loss_context = L1Loss(image_generated_masked, masked_image)*10
#loss_context = SmoothL1Loss(image_generated_masked, masked_image)*10
discriminator_output = netD(image_generated_inpainted) - 0.005 # -0.005 offset so loss_perceptual doesn't become 1 when D(G(z)) == 1.000000
#print("Discriminator output: ", discriminator_output)
labels = torch.full((b_size,), 1, device=device)
loss_perceptual = lamb2*torch.log(1-discriminator_output)
#if loss_perceptual == -np.inf:
# #print('loss perceptual == -np.inf()')
# loss_perceptual = torch.tensor(-10, dtype=torch.float32, device=device)
#print(loss_perceptual.data.cpu().numpy().flatten()[0])
total_loss = loss_context + loss_perceptual
#total_loss = loss_context + 10*discriminator_output
# grab the values from losses for printing
loss_perceptual = loss_perceptual.data.cpu().numpy().flatten()[0]
#loss_perceptual = 0
loss_context = loss_context.data.cpu().numpy().flatten()[0]
total_loss.sum().backward() # TODO: find out why .sum() is needed (why does the loss tensor have 2 elements?)
opt.step()
total_loss = total_loss.data.cpu().numpy().flatten()[0]
"""Early stopping""" # TODO:
if iteration > 0:
delta_loss = early_stopping_min_loss - total_loss
delta_iters = iteration - iter1
if (delta_loss < 0.1) or (total_loss > early_stopping_min_loss):
early_stopping_n_iters += 1
else:
#print('set to zero')
early_stopping_n_iters = 0
if early_stopping_n_iters > 1000:
#n_iteration.append(iteration)
#break
#z = z_best
#early_stopping_n_iters = 0
#print('z copied')
pass
loss1 = total_loss
iter1 = iteration
if total_loss < early_stopping_min_loss:
early_stopping_min_loss = total_loss
best_inpained_image = image_generated.detach().cpu()
contextual_loss_best = loss_context
perceptual_loss_best = loss_perceptual
early_stopping_n_iters = 0
z_best = z
#print('min loss: ', early_stopping_min_loss)
t2 = time.time()
"""Calculate ETA"""
#t_per_iter = t2 - t1 # time per iteration in seconds
past_time = t2 - t3
#eta = t_per_iter * (n_iters - iteration) + t_per_iter* (len(images)-i+1) * n_iters # time left to finish epoch/image + time left to finish all epochs/images in SECONDS
#eta_h = (eta/ 60) // 60 # divisor integer
#eta_m = eta % 60 # get remainer
past_m = past_time / 120
past_s = past_time % 60
if (iteration % 50 == 0):
print("\r image [{}/{}] inpainting [{}/{}] iteration : {:4} , context_loss: {:.3f}, perceptual_loss: {:3f}, total_loss: {:3f}, min L: {:3f}, {:1f}, D(G(z)): {:3f}, Run time: {:.0f}m {:.0f}s, s per image {:.0f}s".format(i+1,
len(images), j+1, inpaint_n_times, iteration, loss_context,loss_perceptual, total_loss,early_stopping_min_loss, early_stopping_n_iters, discriminator_output.data.cpu().numpy().flatten()[0], past_m, past_s, past_s_image),end="")
"""NaN monitor"""
#if (loss_context or loss_perceptual == np.nan()) and iteration >64:
# print(r'='*10 + ' NaN '+ '='*10)
# print(loss_context, loss_percept ual)
#break+
final_inpainted_images_n_times.append(best_inpained_image.detach().cpu())
past_s_image = (t2-t1) % 60
final_inpainted_images.append(final_inpainted_images_n_times)
real_images.append(real_images_n_times)
contextual_losses.append(contextual_loss_best)
perceptual_losses.append(perceptual_loss_best)
if save_array_results:
save_inpainting_results()
# -
perceptual_losses
# # Error of one ASPA
# +
selected_aspa = 0
reals = [real_images[selected_aspa][i].detach().cpu().numpy()[0, 0, :, :] for i in range(inpaint_n_times)]
inpainteds = [final_inpainted_images[selected_aspa][i].detach().cpu().numpy()[0, 0, :, :] for i in range(inpaint_n_times)]
# +
# :16, :25 is the spectrum location within the ASPA
real = reals[0][:16, :25].flatten()
inpainted = inpainteds[0][:16, :25].flatten()
plt.figure(figsize=(15,5))
plt.plot(real, 'x-', c='r', linewidth=0.5)
plt.plot(inpainted, '.-', linewidth=0.5)
# +
# Pixel difference
# -
plt.plot(inpainted-real, '.-')
# +
i = 0
xhat,yhat = ke.decode_spectrum_from_aspa(reals[i])
x,y = ke.decode_spectrum_from_aspa(inpainteds[i])
plt.plot(xhat, yhat, label='real', c='r')
plt.plot(x,y,label='inpainted')
plt.gca().set_xscale('log')
plt.legend()
# -
reals[0].shape
reals = [ke.decode_params_from_aspa(aspa_real) for aspa_real in reals]
inpainteds = [ke.decode_params_from_aspa(aspa_inpainted) for aspa_inpainted in inpainteds]
# Initialize ExoGAN params with zero's
inpainted_params = {
'planet_mass': [],
'temp_profile': [],
'ch4_mixratio': [],
'planet_radius': [],
'h2o_mixratio': [],
'co2_mixratio': [],
'co_mixratio': []
}
# iterate over all params
for i,param in enumerate(inpainted_params):
# iterate over all inpainted values (of above param)
for j,inpainted in enumerate(inpainteds):
y_hat = reals[j][param] # real value
y = inpainted[param] # inpainted value
percentage_error = ((y - y_hat) / y_hat)*100
inpainted_params[param] += [percentage_error]
df = pd.DataFrame(inpainted_params)
df = df.replace([np.inf, -np.inf], np.nan) # TODO: Fix the occurance of inf later
df.describe()
df
# + active=""
# plt.figure(figsize=((25,10)))
# for i,param in enumerate(inpainted_params):
# #if param == 'temp_profile':
# # pass
# #else:
# plt.subplot(3,3,i+1)
# plt.title(param)
# plt.hist(df[param], bins=25)
#
# # plot mean and median line
# mu = df[param].mean()
# plt.axvline(x=mu, color='black', linestyle='-.', alpha=0.9, label='mean')
# plt.axvline(x=df[param].median(), color='black', linestyle='-', alpha=1, label='median')
#
# # plot std lines
# plt.axvline(x=mu-df[param].std(), color='black', linestyle=':', alpha=1, label=r'$\sigma$')
# plt.axvline(x=mu+df[param].std(), color='black', linestyle=':', alpha=1)
#
# #plt.xlabel(r'Percentage error')
# plt.legend()
# plt.grid()
# plt.tight_layout()
# -
# # Error per ASPA
def calculate_aspa_error(selected_aspa):
"""
Index value of selected aspa -> final_inpainted_images[selected_aspa]
E.g. 0 for first image, 1 for 2nd image etc.
Returns df containing percentage errors per param
"""
#if load_array_results:
# reals = np.load('gan_data//val_errors//'+filename+'_reals.npy')
# inpainteds = np.load('gan_data//val_errors//'+filename+'_inpainteds.npy')
#else:
reals = [real_images[selected_aspa][i].detach().cpu().numpy()[0, 0, :, :] for i in range(inpaint_n_times)]
inpainteds = [final_inpainted_images[selected_aspa][i].detach().cpu().numpy()[0, 0, :, :] for i in range(inpaint_n_times)]
reals = [ke.decode_params_from_aspa(aspa_real) for aspa_real in reals]
inpainteds = [ke.decode_params_from_aspa(aspa_inpainted) for aspa_inpainted in inpainteds]
# Initialize ExoGAN params with zero's
inpainted_params = {
'planet_mass': [],
'temp_profile': [],
'ch4_mixratio': [],
'planet_radius': [],
'h2o_mixratio': [],
'co2_mixratio': [],
'co_mixratio': []
}
# iterate over all params
for i,param in enumerate(inpainted_params):
# iterate over all inpainted values (of above param)
for j,inpainted in enumerate(inpainteds):
y_hat = reals[j][param] # real value
y = inpainted[param] # inpainted value
percentage_error = ((y - y_hat) / y_hat)*100
inpainted_params[param] += [percentage_error]
df = pd.DataFrame(inpainted_params)
return df
dfs = [calculate_aspa_error(selected_aspa) for selected_aspa in range(len(images))]
# # Create df of all mean values
# Dataframe containing the mean value of the n inpaintings, per image.
# +
means = []
for i in range(len(dfs)):
df = dfs[i].describe()
means.append(df[df.index == 'mean'])
means = pd.concat(means)
means = means.replace([np.inf, -np.inf], np.nan) # TODO: Fix the occurance of inf later
means.describe()
# -
# # Create df of all std values
# Dataframe containing the std of the n inpaintings, per image.
# +
stds = []
for i in range(len(dfs)):
df = dfs[i].describe()
stds.append(df[df.index == 'std'])
stds = pd.concat(stds)
stds.describe()
# -
# # Hist of all mean
plt.figure(figsize=((25,10)))
for i,param in enumerate(inpainted_params):
plt.subplot(3,3,i+1)
plt.title(param)
try:
plt.hist(means[param], bins=25)
# plot mean and median line
plt.axvline(x=means[param].mean(), color='black', linestyle='-.', alpha=0.9)
plt.axvline(x=means[param].median(), color='black', linestyle='-', alpha=0.9)
# plot std lines
plt.axvline(x=-means[param].std(), color='black', linestyle=':', alpha=1)
plt.axvline(x=means[param].std(), color='black', linestyle=':', alpha=1)
except:
pass
plt.grid()
plt.tight_layout()
# # n iterations per inpainting
# +
n_iteration = np.array(n_iteration)
if save_array_results:
np.save('gan_data//val_errors//'+filename+'_n_iterations.npy', n_iteration)
_ = plt.hist(n_iteration, bins=50)
n_iteration.mean(), n_iteration.std()
# -
n_iteration
| notebooks/inpainting_v4_wgan_complex.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_tensorflow_p36
# language: python
# name: conda_tensorflow_p36
# ---
# # Compile and Deploy a TensorFlow model on Inf1 instances
#
#
# Amazon SageMaker now supports Inf1 instances for high performance and cost-effective inferences. Inf1 instances are ideal for large scale machine learning inference applications like image recognition, speech recognition, natural language processing, personalization, and fraud detection. In this example, we train a classification model on the MNIST dataset using TensorFlow, compile it using Amazon SageMaker Neo, and deploy the model on Inf1 instances on a SageMaker endpoint and use the Neo Deep Learning Runtime to make inferences in real-time and with low latency.
#
# ### Inf 1 instances
# Inf1 instances are built from the ground up to support machine learning inference applications and feature up to 16 AWS Inferentia chips, high-performance machine learning inference chips designed and built by AWS. The Inferentia chips are coupled with the latest custom 2nd generation Intel® Xeon® Scalable processors and up to 100 Gbps networking to enable high throughput inference. With 1 to 16 AWS Inferentia chips per instance, Inf1 instances can scale in performance to up to 2000 Tera Operations per Second (TOPS) and deliver extremely low latency for real-time inference applications. The large on-chip memory on AWS Inferentia chips used in Inf1 instances allows caching of machine learning models directly on the chip. This eliminates the need to access outside memory resources during inference, enabling low latency without impacting bandwidth.
#
# ### Set up the environment
# +
import os
import sagemaker
from sagemaker import get_execution_role
import boto3
sagemaker_session = sagemaker.Session()
role = get_execution_role()
# -
# ### Download the MNIST dataset
# +
import utils
from tensorflow.contrib.learn.python.learn.datasets import mnist
import tensorflow as tf
data_sets = mnist.read_data_sets("data", dtype=tf.uint8, reshape=False, validation_size=5000)
utils.convert_to(data_sets.train, "train", "data")
utils.convert_to(data_sets.validation, "validation", "data")
utils.convert_to(data_sets.test, "test", "data")
# -
# ### Upload the data
# We use the ```sagemaker.Session.upload_data``` function to upload our datasets to an S3 location. The return value inputs identifies the location -- we will use this later when we start the training job.
inputs = sagemaker_session.upload_data(path="data", key_prefix="data/DEMO-mnist")
# # Construct a script for distributed training
# Here is the full code for the network model:
# !cat 'mnist.py'
# The script here is and adaptation of the [TensorFlow MNIST example](https://github.com/tensorflow/models/tree/master/official/vision/image_classification). It provides a ```model_fn(features, labels, mode)```, which is used for training, evaluation and inference. See [TensorFlow MNIST distributed training notebook](The script here is and adaptation of the [TensorFlow MNIST example](https://github.com/tensorflow/tensorflow/tree/master/tensorflow/examples/tutorials/mnist). It provides a ```model_fn(features, labels, mode)```, which is used for training, evaluation and inference. See [TensorFlow MNIST distributed training notebook](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/sagemaker-python-sdk/tensorflow_script_mode_training_and_serving/tensorflow_script_mode_training_and_serving.ipynb) for more details about the training script.
#
#
# At the end of the training script, there are two additional functions, to be used with Neo Deep Learning Runtime:
# * `neo_preprocess(payload, content_type)`: Function that takes in the payload and Content-Type of each incoming request and returns a NumPy array
# * `neo_postprocess(result)`: Function that takes the prediction results produced by Deep Learining Runtime and returns the response body
#
# <NAME>., <NAME>., & <NAME>. (2010). MNIST handwritten digit databaseATT Labs [Online]. Available: http://yann.lecun.com/exdb/mnist, 2.
# ## Create a training job using the sagemaker.TensorFlow estimator
# +
from sagemaker.tensorflow import TensorFlow
mnist_estimator = TensorFlow(
entry_point="mnist.py",
role=role,
framework_version="1.11.0",
training_steps=1000,
evaluation_steps=100,
train_instance_count=2,
train_instance_type="ml.c5.xlarge",
sagemaker_session=sagemaker_session,
)
mnist_estimator.fit(inputs)
# -
# The **```fit```** method will create a training job in two **ml.c5.xlarge** instances. The logs above will show the instances doing training, evaluation, and incrementing the number of **training steps**.
#
# In the end of the training, the training job will generate a saved model for compilation.
# # Deploy the trained model on Inf1 instance for real-time inferences
#
# Once the training is complete, we compile the model using Amazon SageMaker Neo to optize performance for our desired deployment target. Amazon SageMaker Neo enables you to train machine learning models once and run them anywhere in the cloud and at the edge. To compile our trained model for deploying on Inf1 instances, we are using the ``TensorFlowEstimator.compile_model`` method and select ``'ml_inf1'`` as our deployment target. The compiled model will then be deployed on an endpoint using Inf1 instances in Amazon SageMaker.
#
# ## Compile the model
#
# The ``input_shape`` is the definition for the model's input tensor and ``output_path`` is where the compiled model will be stored in S3. **Important. If the following command result in a permission error, scroll up and locate the value of execution role returned by `get_execution_role()`. The role must have access to the S3 bucket specified in ``output_path``.**
# +
output_path = "/".join(mnist_estimator.output_path.split("/")[:-1])
mnist_estimator.framework_version = "1.15.0"
optimized_estimator = mnist_estimator.compile_model(
target_instance_family="ml_inf1",
input_shape={"data": [1, 784]}, # Batch size 1, 3 channels, 224x224 Images.
output_path=output_path,
framework="tensorflow",
framework_version="1.15.0",
)
# -
# ## Deploy the compiled model on a SageMaker endpoint
#
# Now that we have the compiled model, we will deploy it on an Amazon SageMaker endpoint. Inf1 instances in Amazon SageMaker are available in four sizes: ml.inf1.xlarge, ml.inf1.2xlarge, ml.inf1.6xlarge, and ml.inf1.24xlarge. In this example, we are using ``'ml.inf1.xlarge'`` for deploying our model.
#
optimized_predictor = optimized_estimator.deploy(
initial_instance_count=1, instance_type="ml.inf1.xlarge"
)
# +
import numpy as np
def numpy_bytes_serializer(data):
f = io.BytesIO()
np.save(f, data)
f.seek(0)
return f.read()
optimized_predictor.content_type = "application/vnd+python.numpy+binary"
optimized_predictor.serializer = numpy_bytes_serializer
# -
# ## Invoking the endpoint
#
# Once the endpoint is ready, you can send requests to it and receive inference results in real-time with low latency.
# +
from tensorflow.examples.tutorials.mnist import input_data
from IPython import display
import PIL.Image
import io
mnist = input_data.read_data_sets("/tmp/data/", one_hot=True)
for i in range(10):
data = mnist.test.images[i]
# Display image
im = PIL.Image.fromarray(data.reshape((28, 28)) * 255).convert("L")
display.display(im)
# Invoke endpoint with image
predict_response = optimized_predictor.predict(data)
print("========================================")
label = np.argmax(mnist.test.labels[i])
print("label is {}".format(label))
prediction = predict_response
print("prediction is {}".format(prediction))
# -
# ## Deleting endpoint
#
# Delete the endpoint if you no longer need it.
sagemaker_session.delete_endpoint(optimized_predictor.endpoint)
| sagemaker_neo_compilation_jobs/deploy_tensorflow_model_on_Inf1_instance/tensorflow_distributed_mnist_neo_inf1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Constructing an Optimal Portfolio
# + outputHidden=false inputHidden=false
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.optimize import fmin
import math
import warnings
warnings.filterwarnings("ignore")
# yfinance is used to fetch data
import yfinance as yf
yf.pdr_override()
# + outputHidden=false inputHidden=false
# input
symbols = ['BAC','AAPL', 'JNJ']
start = '2012-01-01'
end = '2019-01-01'
rf = 0.003
# + outputHidden=false inputHidden=false
def annual_returns(symbols, start, end):
df = yf.download(symbols,start,end)['Adj Close']
log_rets = np.log(df) - np.log(df.shift(1))
date = []
d0 = df.index
for i in range(0, len(log_rets)):
date.append(d0[i].strftime("%Y"))
y = pd.DataFrame(log_rets, date, columns = [symbols])
return np.exp(y.groupby(y.index).sum()) - 1
# + outputHidden=false inputHidden=false
def portfolio_var(M, W):
cor = np.corrcoef(M.T)
vol = np.std(M, axis=0)
var = 0.0
for i in range(n):
for j in range(n):
var += W[i] * W[j] * vol[i] * vol[j] * cor[i, j]
return var
# + outputHidden=false inputHidden=false
def sharpe(M, W):
var = portfolio_var(M, W)
mean_return = np.mean(M, axis=0)
ret = np.array(mean_return)
return (np.dot(W, ret) - rf)/ np.sqrt(252)
# + outputHidden=false inputHidden=false
def negative_sharpe_n_minus_1_stock(W):
w2 = np.append(W, 1-sum(W))
return -sharpe(M, w2)
# + outputHidden=false inputHidden=false
n = len(symbols)
x2 = annual_returns(symbols[0], start, end)
for i in range(1,n):
x_ = annual_returns(symbols[i], start, end)
x2 = pd.merge(x2, x_, left_index=True, right_index=True)
M = np.array(x2)
# + outputHidden=false inputHidden=false
print('Efficient Portfolio (Mean-Variance)')
print('Symbols: ', symbols)
print('Sharpe ratio for an equal-weighted portfolio')
equal_weighted = np.ones(n, dtype=float) * 1.0/n
print(equal_weighted)
print(round(sharpe(M, equal_weighted), 4))
# + outputHidden=false inputHidden=false
w0 = np.ones(n-1, dtype=float) * 1.0 / n
w1 = fmin(negative_sharpe_n_minus_1_stock, w0)
final_weight = np.append(w1, 1 - sum(w1))
final_sharpe = sharpe(M, final_weight)
print('Optimal weights:')
print(final_weight)
print('Sharpe ratio:')
print(round(final_sharpe,4))
| Python_Stock/Portfolio_Strategies/Optimal_Portfolio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import nn
import importlib
importlib.reload(nn)
from nn import ANN
import numpy as np
# %matplotlib notebook
import matplotlib.pyplot as plt
import ipywidgets as wgs
def apply_random_weights(model, seed, max_abs_value):
np.random.seed(seed)
for layer in range(1, len(units_per_layer)):
layer_size = units_per_layer[layer]
prev_layer_size = units_per_layer[layer-1]
for i in range(layer_size):
rand_array = (np.random.rand(prev_layer_size) - 0.5) * 2 * max_abs_value # values between [-1,1] * max_abs_value
b = (np.random.rand() - 0.5) * 2 * max_abs_value # values between [-1,1] * max_abs_value
model.set_weights(layer-1, i, rand_array, b)
def get_model():
assert(nn_model != None)
return nn_model
def on_value_change(change):
is_b = change.owner.description == 'b' # description contains either 'w' or 'b'
changed_layer,changed_unit = change.owner.description_tooltip.split(",")
changed_layer = int(changed_layer)
changed_unit = int(changed_unit)
model = get_model()
# change the value of model's weights according to the respective slider's new value
if is_b:
model.Bs[changed_layer][changed_unit] = change['new']
else:
changed_weight = int(change.owner.description[1:])-1 # skip 'w'
model.Ws[changed_layer][changed_unit][changed_weight] = change['new']
# predict and redraw the figure
redraw_figure(model.predict(X), ax)
def create_tab_layout(units_per_layer, model, slider_max_abs_value):
# set layout details
real_time_update = False
# create tab
tab = wgs.Tab()
tab_children = []
fslider_style = {'description_width': 'initial'}
fslider_layout = wgs.Layout(width='200px')
vbox_layout = wgs.Layout(min_width= '210px')
slider_step = slider_max_abs_value * 0.01
min_height,max_height = -slider_max_abs_value,slider_max_abs_value
# set tab children
num_layers = len(units_per_layer) - 1
for layer_index in range(num_layers):
num_units = units_per_layer[layer_index+1]
num_units_prev = units_per_layer[layer_index]
box = wgs.HBox(
[wgs.VBox(
[wgs.Label(value="unit"+str(unit_index+1))] +
[wgs.FloatSlider(min=min_height,max=max_height,step=slider_step,
value=model.Ws[layer_index][unit_index][w_index-1],
style=fslider_style,layout=fslider_layout,
continuous_update=real_time_update,
description_tooltip=str(layer_index)+","+str(unit_index),
description="w" + str(w_index)) for w_index in range(1,num_units_prev+1)] +
[wgs.FloatSlider(min=min_height,max=max_height,step=slider_step,
value=model.Bs[layer_index][unit_index][0],
style=fslider_style,layout=fslider_layout,
continuous_update=real_time_update,
description_tooltip=str(layer_index)+","+str(unit_index),
description="b")],
layout=vbox_layout)
for unit_index in range(num_units)])
tab_children.append(box)
# set tab titles
tab.children = tab_children
for i in range(0,num_layers):
tab.set_title(i, "layer"+str(i+1))
# set sliders events
for hbox_elem in tab.children:
for vbox_elem in hbox_elem.children:
for slidr in vbox_elem.children[1:]: # skip Label
slidr.observe(on_value_change, names='value')
return tab
def redraw_figure(output_data, axis):
axis.cla()
axis.plot(output_data[0])
nn_model = None
# set nn inputs
fig_len_size = 100
X = np.array([range(-int(fig_len_size/2), int(fig_len_size/2))], dtype=float)
# +
### CHANGE HERE ###
units_per_layer = [1, 2, 2, 1] # set the number of units for each layer
# activation types: sigmoid, tanh, relu, relu_limited, mexican_hat, ceil, step
activations = [ANN.Activation_type.mexican_hat, ANN.Activation_type.mexican_hat, ANN.Activation_type.mexican_hat]
###################
slider_max_abs_value = 3
nn_model = ANN(units_per_layer, activations) # initialize nn model
# model prediction with random weights
### CHANGE HERE ###
rand_seed = 9
###################
apply_random_weights(nn_model, rand_seed, slider_max_abs_value)
model_result = nn_model.predict(X)
# create and display layout
tabs = create_tab_layout(units_per_layer, nn_model, slider_max_abs_value)
display(tabs)
# set figure
plt.ion() # enable interactive mode
fig, ax = plt.subplots()
plt.subplots_adjust(left=0.08, bottom=0.08, right=0.99, top=0.99, hspace=0.0, wspace=0.0)
redraw_figure(model_result, ax)
# -
| 1d_nn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SciPy
# The SciPy library is one of the core packages that make up the SciPy stack. It provides many user-friendly and efficient numerical routines such as routines for numerical integration and optimization.
#
# Library documentation: <a>http://www.scipy.org/scipylib/index.html</a>
# needed to display the graphs
# %matplotlib inline
from pylab import *
from numpy import *
from scipy.integrate import quad, dblquad, tplquad
# integration
val, abserr = quad(lambda x: exp(-x ** 2), Inf, Inf)
val, abserr
from scipy.integrate import odeint, ode
# +
# differential equation
def dy(y, t, zeta, w0):
x, p = y[0], y[1]
dx = p
dp = -2 * zeta * w0 * p - w0**2 * x
return [dx, dp]
# initial state
y0 = [1.0, 0.0]
# time coodinate to solve the ODE for
t = linspace(0, 10, 1000)
w0 = 2*pi*1.0
# solve the ODE problem for three different values of the damping ratio
y1 = odeint(dy, y0, t, args=(0.0, w0)) # undamped
y2 = odeint(dy, y0, t, args=(0.2, w0)) # under damped
y3 = odeint(dy, y0, t, args=(1.0, w0)) # critial damping
y4 = odeint(dy, y0, t, args=(5.0, w0)) # over damped
fig, ax = subplots()
ax.plot(t, y1[:,0], 'k', label="undamped", linewidth=0.25)
ax.plot(t, y2[:,0], 'r', label="under damped")
ax.plot(t, y3[:,0], 'b', label=r"critical damping")
ax.plot(t, y4[:,0], 'g', label="over damped")
ax.legend();
# -
from scipy.fftpack import *
# +
# fourier transform
N = len(t)
dt = t[1]-t[0]
# calculate the fast fourier transform
# y2 is the solution to the under-damped oscillator from the previous section
F = fft(y2[:,0])
# calculate the frequencies for the components in F
w = fftfreq(N, dt)
fig, ax = subplots(figsize=(9,3))
ax.plot(w, abs(F));
# -
# ### Linear Algebra
A = array([[1,2,3], [4,5,6], [7,8,9]])
b = array([1,2,3])
# solve a system of linear equations
x = solve(A, b)
x
# +
# eigenvalues and eigenvectors
A = rand(3,3)
B = rand(3,3)
evals, evecs = eig(A)
evals
# -
evecs
svd(A)
# ### Optimization
from scipy import optimize
# +
def f(x):
return 4*x**3 + (x-2)**2 + x**4
fig, ax = subplots()
x = linspace(-5, 3, 100)
ax.plot(x, f(x));
# -
x_min = optimize.fmin_bfgs(f, -0.5)
x_min
# ### Statistics
from scipy import stats
# +
# create a (continous) random variable with normal distribution
Y = stats.norm()
x = linspace(-5,5,100)
fig, axes = subplots(3,1, sharex=True)
# plot the probability distribution function (PDF)
axes[0].plot(x, Y.pdf(x))
# plot the commulative distributin function (CDF)
axes[1].plot(x, Y.cdf(x));
# plot histogram of 1000 random realizations of the stochastic variable Y
axes[2].hist(Y.rvs(size=1000), bins=50);
# -
Y.mean(), Y.std(), Y.var()
# t-test example
t_statistic, p_value = stats.ttest_ind(Y.rvs(size=1000), Y.rvs(size=1000))
t_statistic, p_value
# Testing complete; Gopal
| tests/ipython-notebooks/SciPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Interactive Plotting and Monte Carlo Simulations of CPPI
# #### salimt
# +
import ipywidgets as widgets
from IPython.display import display
import pandas as pd
import edhec_risk_kit_123 as erk
# %load_ext autoreload
# %autoreload 2
# -
# ## GBM Simulations
def show_gbm(n_scenarios, mu, sigma):
"""
Draw the results of a stock price evolution under a Geometric Brownian Motion model
"""
s_0=100
prices = erk.gbm(n_scenarios=n_scenarios, mu=mu, sigma=sigma, s_0=s_0)
ax = prices.plot(legend=False, color="indianred", alpha = 0.5, linewidth=2, figsize=(12,5))
ax.axhline(y=100, ls=":", color="black")
# draw a dot at the origin
ax.plot(0,s_0, marker='o',color='darkred', alpha=0.2)
gbm_controls = widgets.interactive(show_gbm,
n_scenarios=widgets.IntSlider(min=1, max=1000, step=1, value=1),
mu=(0., +.2,.01),
sigma=(0, .3, .01)
)
display(gbm_controls)
# # Interactive CPPI Monte Carlo Testing
# +
def show_cppi(n_scenarios=50, mu=0.07, sigma=0.15, m=3, floor=0., riskfree_rate=0.03, y_max=100):
"""
Plot the results of a Monte Carlo Simulation of CPPI
"""
start = 100
sim_rets = erk.gbm(n_scenarios=n_scenarios, mu=mu, sigma=sigma, prices=False, steps_per_year=12)
risky_r = pd.DataFrame(sim_rets)
# run the "back"-test
btr = erk.run_cppi(risky_r=pd.DataFrame(risky_r),riskfree_rate=riskfree_rate,m=m, start=start, floor=floor)
wealth = btr["Wealth"]
y_max=wealth.values.max()*y_max/100
ax = wealth.plot(legend=False, alpha=0.3, color="indianred", figsize=(12, 6))
ax.axhline(y=start, ls=":", color="black")
ax.axhline(y=start*floor, ls="--", color="red")
ax.set_ylim(top=y_max)
cppi_controls = widgets.interactive(show_cppi,
n_scenarios=widgets.IntSlider(min=1, max=1000, step=5, value=50),
mu=(0., +.2, .01),
sigma=(0, .30, .05),
floor=(0, 2, .1),
m=(1, 5, .5),
riskfree_rate=(0, .05, .01),
y_max=widgets.IntSlider(min=0, max=100, step=1, value=100,
description="Zoom Y Axis")
)
display(cppi_controls)
# -
# # Adding a Histogram and Reporting Floor Violations
# +
import matplotlib.pyplot as plt
def show_cppi(n_scenarios=50, mu=0.07, sigma=0.15, m=3, floor=0., riskfree_rate=0.03, y_max=100):
"""
Plot the results of a Monte Carlo Simulation of CPPI
"""
start = 100
sim_rets = erk.gbm(n_scenarios=n_scenarios, mu=mu, sigma=sigma, prices=False, steps_per_year=12)
risky_r = pd.DataFrame(sim_rets)
# run the "back"-test
btr = erk.run_cppi(risky_r=pd.DataFrame(risky_r),riskfree_rate=riskfree_rate,m=m, start=start, floor=floor)
wealth = btr["Wealth"]
# calculate terminal wealth stats
y_max=wealth.values.max()*y_max/100
terminal_wealth = wealth.iloc[-1]
# Plot!
fig, (wealth_ax, hist_ax) = plt.subplots(nrows=1, ncols=2, sharey=True, gridspec_kw={'width_ratios':[3,2]}, figsize=(24, 9))
plt.subplots_adjust(wspace=0.0)
wealth.plot(ax=wealth_ax, legend=False, alpha=0.3, color="indianred")
wealth_ax.axhline(y=start, ls=":", color="black")
wealth_ax.axhline(y=start*floor, ls="--", color="red")
wealth_ax.set_ylim(top=y_max)
terminal_wealth.plot.hist(ax=hist_ax, bins=50, ec='w', fc='indianred', orientation='horizontal')
hist_ax.axhline(y=start, ls=":", color="black")
cppi_controls = widgets.interactive(show_cppi,
n_scenarios=widgets.IntSlider(min=1, max=1000, step=5, value=50),
mu=(0., +.2, .01),
sigma=(0, .3, .05),
floor=(0, 2, .1),
m=(1, 5, .5),
riskfree_rate=(0, .05, .01),
y_max=widgets.IntSlider(min=0, max=100, step=1, value=100,
description="Zoom Y Axis")
)
display(cppi_controls)
# -
# # Adding Terminal Wealth Statistics
# +
import matplotlib.pyplot as plt
import numpy as np
def show_cppi(n_scenarios=50, mu=0.07, sigma=0.15, m=3, floor=0., riskfree_rate=0.03, steps_per_year=12, y_max=100):
"""
Plot the results of a Monte Carlo Simulation of CPPI
"""
start = 100
sim_rets = erk.gbm(n_scenarios=n_scenarios, mu=mu, sigma=sigma, prices=False, steps_per_year=steps_per_year)
risky_r = pd.DataFrame(sim_rets)
# run the "back"-test
btr = erk.run_cppi(risky_r=pd.DataFrame(risky_r),riskfree_rate=riskfree_rate,m=m, start=start, floor=floor)
wealth = btr["Wealth"]
# calculate terminal wealth stats
y_max=wealth.values.max()*y_max/100
terminal_wealth = wealth.iloc[-1]
tw_mean = terminal_wealth.mean()
tw_median = terminal_wealth.median()
failure_mask = np.less(terminal_wealth, start*floor)
n_failures = failure_mask.sum()
p_fail = n_failures/n_scenarios
e_shortfall = np.dot(terminal_wealth-start*floor, failure_mask)/n_failures if n_failures > 0 else 0.0
# Plot!
fig, (wealth_ax, hist_ax) = plt.subplots(nrows=1, ncols=2, sharey=True, gridspec_kw={'width_ratios':[3,2]}, figsize=(24, 9))
plt.subplots_adjust(wspace=0.0)
wealth.plot(ax=wealth_ax, legend=False, alpha=0.3, color="indianred")
wealth_ax.axhline(y=start, ls=":", color="black")
wealth_ax.axhline(y=start*floor, ls="--", color="red")
wealth_ax.set_ylim(top=y_max)
terminal_wealth.plot.hist(ax=hist_ax, bins=50, ec='w', fc='indianred', orientation='horizontal')
hist_ax.axhline(y=start, ls=":", color="black")
hist_ax.axhline(y=tw_mean, ls=":", color="blue")
hist_ax.axhline(y=tw_median, ls=":", color="purple")
hist_ax.annotate(f"Mean: ${int(tw_mean)}", xy=(.7, .9),xycoords='axes fraction', fontsize=24)
hist_ax.annotate(f"Median: ${int(tw_median)}", xy=(.7, .85),xycoords='axes fraction', fontsize=24)
if (floor > 0.01):
hist_ax.axhline(y=start*floor, ls="--", color="red", linewidth=3)
hist_ax.annotate(f"Violations: {n_failures} ({p_fail*100:2.2f}%)\nE(shortfall)=${e_shortfall:2.2f}", xy=(.7, .7), xycoords='axes fraction', fontsize=24)
cppi_controls = widgets.interactive(show_cppi,
n_scenarios=widgets.IntSlider(min=1, max=1000, step=5, value=50),
mu=(0., +.2, .01),
sigma=(0, .3, .05),
floor=(0, 2, .1),
m=(1, 5, .5),
riskfree_rate=(0, .05, .01),
steps_per_year=widgets.IntSlider(min=1, max=12, step=1, value=12,
description="Rebals/Year"),
y_max=widgets.IntSlider(min=0, max=100, step=1, value=100,
description="Zoom Y Axis")
)
display(cppi_controls)
# -
# ## SOME INSIGHTS
# **Consider the Monte Carlo Simulation we ran for CPPI. Assume there is no floor set (i.e. Floor is set to Zero) As you increase the number of scenarios, which of the following would you expect:**
# - The difference in terminal wealth between the Worst Scenario and the Best Scenario will INCREASE
# **As you increase the FLOOR, the WORST CASE scenario will:**
# - INCREASE
# **Assume a non-zero floor that is less than the starting wealth. As you increase mu and keep other parameters fixed, you would expect that the terminal wealth:**
# - INCREASES
# **All other things being equal, which of these changes will cause an INCREASE in floor violations**
# - Increasing both “m” and “sigma”
# **All other things being equal, which of these changes will cause an INCREASE in floor violations**
# - Increasing “m” but decreasing “rebals per year”
# **All other things being equal, which of these changes will cause in INCREASE in Expected Shortfall**
# - Increasing “m” but decreasing “rebals per year”
# **Parameter changes that increase the probability of floor violations will also tend to increase the Expected Shortfall. This statement is:**
# - TRUE
# **A CPPI Based Principal Protection Strategy aims to return at least the invested principal by setting the floor equal to the initial value of the assets. Which of the following is true:**
# - It is only possible to run a CPPI based Principal Protection Strategy if the risk free rate is greater than or equal to 0
# **A CPPI based Principal Protection Strategy with 12 rebals per year can have a zero expected shortfall only if:**
# - m = 1
# **All other things being equal, A CPPI based Principal Protection Strategy is more likely to have a final negative return if:**
# - sigma increases
# **All other things being equal, A CPPI based Principal Protection Strategy is more likely to have a final negative return if:**
# - sigma increases
| portfolio-construction-and-analysis-with-python/monte-carlo-simulations-of-CPPI-and-diversification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import cv2
import time
import numpy as np
## Preparation for writing the ouput video
#fourcc = cv2.VideoWriter_fourcc(*'XVID')
#out = cv2.VideoWriter('output.avi', fourcc, 20.0, (640, 480))
##reading from the webcam
cap = cv2.VideoCapture(0)
## Allow the system to sleep for 3 seconds before the webcam starts
time.sleep(3)
count = 0
background = 0
## Capture the background in range of 60
for i in range(60):
ret, background = cap.read()
background = np.flip(background, axis=1)
## Read every frame from the webcam, until the camera is open
while (cap.isOpened()):
ret, img = cap.read()
if not ret:
break
count += 1
img = np.flip(img, axis=1)
## Convert the color space from BGR to HSV
hsv = cv2.cvtColor(img, cv2.COLOR_BGR2HSV)
## Generat masks to detect red color
##YOU CAN CHANGE THE COLOR VALUE BELOW ACCORDING TO YOUR CLOTH COLOR
lower_red = np.array([0, 120, 50])
upper_red = np.array([10, 255,255])
mask1 = cv2.inRange(hsv, lower_red, upper_red)
lower_red = np.array([100, 40, 40])
upper_red = np.array([100, 255, 255])
mask1 = cv2.inRange(hsv, lower_red, upper_red)
# setting the lower and upper range for mask2
lower_red = np.array([155, 40, 40])
upper_red = np.array([180, 255, 255])
mask2 = cv2.inRange(hsv, lower_red, upper_red)
#----------------------------------------------------------------------#
# the above block of code could be replaced with
# some other code depending upon the color of your cloth
mask1 = mask1 + mask2
# Refining the mask corresponding to the detected red color
mask1 = cv2.morphologyEx(mask1, cv2.MORPH_OPEN, np.ones((3, 3),
np.uint8), iterations = 2)
mask1 = cv2.dilate(mask1, np.ones((3, 3), np.uint8), iterations = 1)
mask2 = cv2.bitwise_not(mask1)
# Generating the final output
res1 = cv2.bitwise_and(background, background, mask = mask1)
res2 = cv2.bitwise_and(img, img, mask = mask2)
final_output = cv2.addWeighted(res1, 1, res2, 1, 0)
cv2.imshow("INVISIBLE MAN", final_output)
k = cv2.waitKey(1)
if k == 27:
break
cap.release()
cv2.destroyAllWindows()
#colors code
#skin color Values
#lower_red = np.array([0, 0, 70])
#upper_red = np.array([100, 255,255])
# mask1 = cv2.inRange(hsv, lower_red, upper_red)
#-----------------------
# -
| invisible cloak.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial: optimal binning with binary target - LocalSolver
# To get us started, let's load the application_train.csv file from the Kaggle's competition https://www.kaggle.com/c/home-credit-default-risk/data.
import pandas as pd
df = pd.read_csv("data/kaggle/HomeCreditDefaultRisk/application_train.csv", engine='c')
# We choose a variable to discretize and the binary target.
variable = "REGION_POPULATION_RELATIVE"
x = df[variable].values
y = df.TARGET.values
# Import and instantiate an ``OptimalBinning`` object class. We pass the variable name, its data type, and a solver, in this case, we choose the commercial solver LocalSolver. Note that LocalSolver requires a time limit, which is set to 20 seconds (LocalSolver 10.0). Besides, for this example, we require a more granular binning, therefore we allow a large number of prebins with small size.
#
# To use LocalSolver follow the avaiable instructions:
# https://www.localsolver.com/docs/last/quickstart/solvingyourfirstmodelinpython.html
from optbinning import OptimalBinning
optb = OptimalBinning(name=variable, dtype="numerical", solver="ls", max_n_prebins=100,
min_prebin_size=0.001, time_limit=20)
# We fit the optimal binning object with arrays ``x`` and ``y``.
optb.fit(x, y)
# You can check if an optimal or feasible solution has been found via the ``status`` attribute:
optb.status
binning_table = optb.binning_table
binning_table.build()
binning_table.analysis()
binning_table.plot(metric="event_rate")
optb.information(print_level=1)
# Computing the optimal binning starting with a large number of prebins might be challenging in some situations, therefore solvers such as LocalSolver are suitable to find quality feasible solutions in a reasonable amount of time. However, if LocalSolver is not available we can always try solver options "cp" or "mip".
# #### Constraint programming solver
# First, we keep the 5 seconds time limit:
optb = OptimalBinning(name=variable, dtype="numerical", solver="cp", max_n_prebins=100,
min_prebin_size=0.001, time_limit=5)
optb.fit(x, y)
# The status is "UNKNOWN" therefore nor feasible or optimal solutions was found in 5 seconds.
optb.status
optb.information(print_level=1)
# We increase the time limit to 30 seconds.
optb = OptimalBinning(name=variable, dtype="numerical", solver="cp", max_n_prebins=100,
min_prebin_size=0.001, time_limit=30)
optb.fit(x, y)
# In 30 seconds we found a feasible solution
optb.status
optb.information(print_level=1)
binning_table = optb.binning_table
binning_table.build()
binning_table.analysis()
# The current solution is IV = 0.03514331, compared to the LocalSolver solver solution 0.03776231. Let us increase the time limit to 200 seconds.
optb = OptimalBinning(name=variable, dtype="numerical", solver="cp", max_n_prebins=100,
min_prebin_size=0.001, time_limit=200)
optb.fit(x, y)
# The optimal solution is found within the time limit.
optb.status
optb.information(print_level=1)
binning_table = optb.binning_table
binning_table.build()
binning_table.analysis()
# The optimal solution is IV = 0.03776231, matching the LocalSolver solver solution 0.03776231.
binning_table.plot(metric="event_rate")
| doc/source/tutorials/tutorial_binary_localsolver.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# import modules
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
#open csv with pandas
df02 = pd.read_csv("./data/ACS_16_5YR_DP02_with_ann.csv", low_memory=False)
# print columns
df02.columns
# see shape of data
df02.shape
# see column types
df02.dtypes
# print first 5 rows
df02.head()
# delete first row
df02 = df02.drop(df02.index[[0]])
# convert df columns to numeric, ignoring non-numeric values
cols = df02.columns
df02[cols] = df02[cols].apply(pd.to_numeric, errors='ignore')
# see column types
df02.dtypes
# +
# drop columns with "(X)"
# df != "(X)" creates a boolean DataFrame which is True where df is "(X)"
# (df != 0).any(axis=0) returns a boolean Series indicating which columns have "(X)" entries
# The any operation aggregates values along the 0-axis -- i.e. along the rows -- into a single boolean value.
# Hence the result is one boolean value for each column
# df.loc can be used to select those columns
df02 = df02.loc[:, (df02 != '(X)').any(axis=0)]
# -
# get info on data
print(df02.info())
#open zipcode csv with pandas
zipcode = pd.read_csv("./data/zipcode_states.csv")
#merge df02 with zipcodes
df002 = pd.merge(left=df02, right=zipcode, left_on='GEO.id2', right_on='zipcode')
# see first 5 rows
df002.head()
# get info on data
print(df002.info())
# +
# get info stats on df
import pandas_profiling
#save report to file
pfr = pandas_profiling.ProfileReport(df002)
pfr.to_file("./profiling/df002.html")
# -
# DataFrame method to check for missing values in a column
# The .all() method returns True if all values are True
# if using it on a DataFrame, you need to chain another .all()
# method so that you return only one True or False value.
assert pd.notnull(df002['GEO.id2']).all().all()
assert pd.notnull(df002['HC01_VC150']).all().all()
assert (df002.HC01_VC150 >=0).all().all()
df002['HC03_VC166'].unique()
# ls data/
#open metadata csv with pandas
df02m = pd.read_csv("./data/ACS_16_5YR_DP02_metadata.csv")
df02m.head(10)
# +
# create boxplot
df002.boxplot(column='HC01_VC03', by='state', rot=90)
plt.show()
| .ipynb_checkpoints/cleaning data-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ### Imports
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import os
from sklearn.model_selection import train_test_split
from sklearn import preprocessing
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import SGDRegressor
from yellowbrick.regressor import PredictionError
from yellowbrick.features.importances import FeatureImportances
from yellowbrick.model_selection import LearningCurve
from yellowbrick.model_selection import ValidationCurve
from yellowbrick.classifier import ClassificationReport
from sklearn.ensemble import RandomForestClassifier
from yellowbrick.classifier import ConfusionMatrix
from sklearn.model_selection import StratifiedKFold
from sklearn_porter import Porter
# ### Load data
dataset = pd.read_csv('../results/Intel(R) Core(TM) i7-8700 CPU @ 3.20GHz/Intersect_masterset.csv')
dataset.columns = dataset.columns.str.strip()
dataset.head()
# ### Shape datasets
feature_cols = ['average1', 'n1','n2', 'average2']
# +
#feature_cols = ['n1', 'n2']
# -
# All rows, and the feature_cols' columns
X = dataset.loc[:, feature_cols]
#scaler = preprocessing.StandardScaler()
#X = scaler.fit_transform(X)
X.shape
ys = []
algos = ['skewed_1', 'skewed_2', 'non_skewed']
for algo in algos:
y = dataset.loc[:, algo]
ys.append(y)
y.shape
sets = []
for index, y in enumerate(ys):
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
sets.append((X_train, X_test, y_train, y_test, algos[index]))
"train: " + str(y_train.shape) + " test: " + str(y_test.shape)
# # Regression
# ### Train regression model
def regPredictionError():
fig = plt.figure()
fig.subplots_adjust(bottom= 1, top= 3.5, left=1, right=3)
for index, (X_train, X_test, y_train, y_test, filename) in enumerate(sets):
ax = fig.add_subplot(2,2,index+1)
visualizer = PredictionError(LinearRegression(n_jobs=-1), ax=ax)
visualizer.fit(X_train, y_train)
visualizer.score(X_test, y_test)
visualizer.finalize()
ax.set_title(filename)
plt.show()
regPredictionError()
# ### Regression Feature Importances
def regFeatureImportances():
fig = plt.figure()
fig.subplots_adjust(bottom= 1, top= 3.5, left=1, right=3)
for index, y in enumerate(ys):
ax = fig.add_subplot(2,2,index+1)
labels = list(map(lambda s: s.title(), feature_cols))
viz = FeatureImportances(LinearRegression(n_jobs=-1), labels=labels, ax=ax, relative=False)
viz.fit(X, y)
ax.set_title(algos[index])
ax.set_xlabel("Coefficient value")
plt.show()
regFeatureImportances()
# ### Regression Learning Curve
def regLearningCurve():
sizes = np.linspace(0.3, 1.0, 10)
fig = plt.figure()
fig.subplots_adjust(bottom= 1, top= 3.5, left=1, right=3)
for index, y in enumerate(ys):
ax = fig.add_subplot(2,2,index+1)
viz = LearningCurve(LinearRegression(n_jobs=-1), scoring='r2', train_sizes=sizes, n_jobs=4, ax=ax)
viz.fit(X, y)
viz.finalize()
ax.set_title(algos[index])
plt.show()
regLearningCurve()
# ### Regression Validation Curve
def regValidationCurve():
param_range = np.arange(100, 1000, 100)
fig = plt.figure()
fig.subplots_adjust(bottom= 1, top= 3.5, left=1, right=3)
for index, y in enumerate(ys):
ax = fig.add_subplot(2,2,index+1)
viz = ValidationCurve(
SGDRegressor(), param_name="max_iter", param_range=param_range,
scoring="r2", n_jobs=4, ax=ax)
viz.fit(X, y)
viz.finalize()
ax.set_title(algos[index])
plt.show()
regValidationCurve()
# # Classification
# Make images larger
import matplotlib as mpl
plt.rcParams['figure.dpi'] = 125
# ### Build Classification dataset
list(dataset)
feature_cols = ['range',
'n1',
'average1',
'median1',
'std1',
'n2',
'average2',
'median2',
'std2'
]
X = dataset.loc[:, feature_cols]
# scaler = preprocessing.StandardScaler()
# X = scaler.fit_transform(X)
X = X.values
cy = []
cyTimes = []
for i in range(len(ys[0])):
minVal = float("inf")
minName = ""
for j, y in enumerate(ys):
if (minVal > y[i]):
minVal = y[i]
minName = algos[j]
cy.append(minName)
cyTimes.append(minVal)
cy = np.array(cy)
cy.shape
def clfSplit():
return train_test_split(X, cy, test_size=0.2, random_state=0)
# ### Train Classification model
def clfReport():
X_train, X_test, y_train, y_test = clfSplit()
visualizer = ClassificationReport(RandomForestClassifier(n_estimators=5), support='percent')
visualizer.fit(X_train, y_train)
visualizer.score(X_test, y_test)
visualizer.poof()
clfReport()
# ### Confusion Matrix
def confusionMatrix():
X_train, X_test, y_train, y_test = clfSplit()
cm = ConfusionMatrix(RandomForestClassifier(n_estimators=5))
cm.fit(X_train, y_train)
cm.score(X_test, y_test)
cm.poof()
confusionMatrix()
# ### Classification Feature importances
def clfFeatureImportances():
labels = list(map(lambda s: s.title(), feature_cols))
viz = FeatureImportances(RandomForestClassifier(n_estimators=5), labels=labels, relative=True)
viz.fit(X, cy)
viz.poof()
clfFeatureImportances()
# ### Classification learning curve
def clfLearningCurve():
sizes = np.linspace(0.15, 1.0, 10)
viz = LearningCurve(
RandomForestClassifier(n_estimators=10), train_sizes=sizes,
scoring='f1_weighted', n_jobs=-1
)
viz.fit(X, cy)
viz.poof()
clfLearningCurve()
# ### Classification Validation Curve
def clfValidationCurve():
param_range = np.arange(1, 30, 1)
viz = ValidationCurve(
RandomForestClassifier(), param_name="n_estimators", param_range=param_range,
scoring="f1_weighted", n_jobs=-1)
viz.fit(X, cy)
viz.poof()
clfValidationCurve()
# ### Export Classification model to C
def exportClassifierToC():
clf = DecisionTreeClassifier(max_depth=1)
clf.fit(X, cy)
porter = Porter(clf, language='c')
output = porter.export(embed_data=True)
with open('tree.c', 'w') as f:
f.write(output)
exportClassifierToC()
| Machine Learning/Machine Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/prachuryanath/TF-Learning/blob/main/09_SkimLit_nlp_milestone_project_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Yc96wwcKO-El"
# # Milestone Project 2 : SkimLit
#
# In this project, we're going to be putting what we've learned into practice.
#
# More specificially, we're going to be replicating the deep learning model behind the 2017 paper `PubMed 200k RCT: a Dataset for Sequenctial Sentence Classification in Medical Abstracts`.
#
# When it was released, the paper presented a new dataset called `PubMed 200k RCT which consists of ~200,000 labelled Randomized Controlled Trial (RCT) abstracts`.
#
# The goal of the dataset was to explore the ability for NLP models to classify sentences which appear in sequential order.
#
# ### **Problem in a sentence**
# *The number of RCT papers released is continuing to increase, those without structured abstracts can be hard to read and in turn slow down researchers moving through the literature.*
#
# ### **Solution in a sentence**
# *Create an NLP model to classify abstract sentences into the role they play (e.g. objective, methods, results, etc) to enable researchers to skim through the literature (hence SkimLit 🤓🔥) and dive deeper when necessary.*
# + [markdown] id="49iOYPcIPeRI"
# ## What we're going to cover
# Time to take what we've learned in the NLP fundmentals notebook and build our biggest NLP model yet:
#
# * Downloading a text dataset (PubMed RCT200k from GitHub)
# * Writing a preprocessing function to prepare our data for modelling
# * Setting up a series of modelling experiments
# * Making a baseline (TF-IDF classifier)
# * Deep models with different combinations of: token embeddings, character embeddings, pretrained embeddings, positional embeddings
# * Building our first multimodal model (taking multiple types of data inputs)
# * Replicating the model architecture from https://arxiv.org/pdf/1612.05251.pdf
# * Find the most wrong predictions
# * Making predictions on PubMed abstracts from the wild
# + colab={"base_uri": "https://localhost:8080/"} id="vbHryq10QGlA" outputId="2d5c0ebb-2649-4de3-fe20-9b4b22d95fdc"
# Check for GPU
# !nvidia-smi
# + [markdown] id="aXj_Ou67QU0U"
# ## Get data
#
# Before we can start building a model, we've got to download the PubMed 200k RCT dataset.
#
# In a phenomenal act of kindness, the authors of the paper have made the data they used for their research availably publically and for free in the form of .txt files on GitHub.
#
# We can copy them to our local directory using git clone `https://github.com/Franck-Dernoncourt/pubmed-rct.`
# + colab={"base_uri": "https://localhost:8080/"} id="rQ5b5_lMQuJV" outputId="1476113c-f23e-4105-ab86-3d83916e97be"
# !git clone https://github.com/Franck-Dernoncourt/pubmed-rct.git
# !ls pubmed-rct
# + [markdown] id="FSrJNVHRQ0aQ"
# Looking at the README file from the GitHub page, we get the following information:
#
# * `PubMed 20k` is a subset of `PubMed 200k`. I.e., any abstract present in `PubMed 20k` is also present in `PubMed 200k`.
# * `PubMed_200k_RCT` is the same as `PubMed_200k_RCT_numbers_replaced_with_at_sign`, except that in the latter all numbers had been replaced by @. (same for `PubMed_20k_RCT` vs. `PubMed_20k_RCT_numbers_replaced_with_at_sign`).
# * Since Github file size limit is 100 MiB, we had to compress `PubMed_200k_RCT\train.7z` and `PubMed_200k_RCT_numbers_replaced_with_at_sign\train.zip`. To uncompress train.7z, you may use 7-Zip on Windows, Keka on Mac OS X, or p7zip on Linux.
#
# To begin with, the dataset we're going to be focused on is `PubMed_20k_RCT_numbers_replaced_with_at_sign`.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="Z1liAnC0R5Zc" outputId="c890bf4e-15d5-413c-8e68-cb93c2df4045"
# Check what files are in the PubMed_20K dataset
# !ls pubmed-rct/PubMed_20k_RCT_numbers_replaced_with_at_sign
# + [markdown] id="e9v0_aP8SJWc"
# Beautiful, looks like we've got three separate text files:
#
# * **train.txt** - *training samples.*
# * **dev.txt** - *dev is short for development set, which is another name for validation set (in our case, we'll be using and referring to this file as our validation set).*
# * **test.txt** - *test samples.*
# + id="z72jYlzJSATJ"
# Start by using the 20k dataset
data_dir = "pubmed-rct/PubMed_20k_RCT_numbers_replaced_with_at_sign/"
# + colab={"base_uri": "https://localhost:8080/"} id="AXbC3PBRSDHX" outputId="95eb8823-d829-42a6-da4c-abc24733c0c6"
# Check all of the filenames in the target directory
import os
filenames = [data_dir + filename for filename in os.listdir(data_dir)]
filenames
# + [markdown] id="Pc8I6W8nSE75"
# ## Preprocess data
#
# To get familiar and understand how we have to prepare our data for our deep learning models, we've got to visualize it.
#
# Because our data is in the form of text files, let's write some code to read each of the lines in a target file.
# + id="SXXE_zSsShAx"
# Create function to read the lines of a document
def get_lines(filename):
"""
Reads filename (a text file) and returns the lines of text as a list
Args:
filename : a string containing the target filepath to read.
Returns:
A list of strings with one string per line from the target filename.
For example:
["this is the first line of filename",
"this is the second line of filename",
...]
"""
with open(filename, "r") as f:
return f.readlines()
# + colab={"base_uri": "https://localhost:8080/"} id="SLH534ogTwad" outputId="20dae020-fb9b-4cdb-ed19-c1080a7e2792"
train_lines = get_lines(data_dir+"train.txt")
train_lines[:10]
# + [markdown] id="dwXHgushT7pY"
# Let's write a function to perform the following steps:
#
# * Take a target file of abstract samples.
# * Read the lines in the target file.
# * For each line in the target file:
# * If the line begins with ### mark it as an abstract ID and the beginning of a new abstract.
# * Keep count of the number of lines in a sample.
# * If the line begins with \n mark it as the end of an abstract sample.
# * Keep count of the total lines in a sample.
# * Record the text before the \t as the label of the line.
# * Record the text after the \t as the text of the line.
# * Return all of the lines in the target text file as a list of dictionaries containing the key/value pairs:
# * "line_number" - the position of the line in the abstract (e.g. 3).
# * "target" - the role of the line in the abstract (e.g. OBJECTIVE).
# * "text" - the text of the line in the abstract.
# * "total_lines" - the total lines in an abstract sample (e.g. 14).
# * Abstract ID's and newlines should be omitted from the returned preprocessed data.
# + id="S1deHBAEUhMe"
def preprocess_text_with_line_numbers(filename):
"""
Returns a list of dictionaries of abstract line data.
Takes in filename, reads its contents and sorts through each line,
extracting things like the target label, the text of the sentence,
how many sentences are in the current abstract and what sentence
number the target line is.
Args:
filename : a string of the target text file to read and extract line data from.
Returns:
A list of dictionaries each containing a line from an abstract,
the lines label, the lines position in the abstract and the total
number of lines in the abstract where the line is from. For example:
[{"target":'Conclusion',
"text": "The study couldn't have gone better",
"line_number":8,
"total_lines":9}]
"""
input_lines = get_lines(filename) # get all lines from filename
abstract_lines = "" # create an empty abstract
abstract_samples = [] # create an empty list of abstracts
# Loop through each line in target file
for line in input_lines:
if line.startswith('###'): # check to see if line is an ID line
abstract_id = line
abstract_lines = "" # reset abstract string
elif line.isspace(): # check to see if line is a new line
abstract_line_split = abstract_lines.splitlines() # split abstract into separate lines
# Iterate through each line in abstract and count them at the same time
for abstract_line_number, abstract_line in enumerate(abstract_line_split):
line_data = {} # create empty dict to store data from line
target_text_split = abstract_line.split("\t") # split target label from text
line_data["target"] = target_text_split[0] # get target label
line_data["text"] = target_text_split[1].lower() # get target text and lower it
line_data["line_number"] = abstract_line_number # what number line does the line appear in the abstract?
line_data["total_lines"] = len(abstract_line_split) - 1 # how many total lines are in the abstract? (start from 0)
abstract_samples.append(line_data) # add line data to abstract samples list
else: # if the above conditions aren't fulfilled, the line contains a labelled sentence
abstract_lines += line
return abstract_samples
# + colab={"base_uri": "https://localhost:8080/"} id="WOZpSqPEaC0K" outputId="6f4e7274-c864-4463-8210-e3c49b7ffc4c"
# Get data from file and preprocess it
# %%time
train_samples = preprocess_text_with_line_numbers(data_dir + "train.txt")
val_samples = preprocess_text_with_line_numbers(data_dir + "dev.txt") # dev is another name for validation set
test_samples = preprocess_text_with_line_numbers(data_dir + "test.txt")
len(train_samples), len(val_samples), len(test_samples)
# + colab={"base_uri": "https://localhost:8080/"} id="EgZfGCaiaHR0" outputId="2ad7b170-da9a-4776-d16f-2879137e9ed9"
# Check the first abstract of our training data
train_samples[:10]
# + colab={"base_uri": "https://localhost:8080/", "height": 483} id="v0sJK23Wam7F" outputId="204cb14b-e68f-46a1-c3a7-a0a286093d47"
import pandas as pd
train_df = pd.DataFrame(train_samples)
val_df = pd.DataFrame(val_samples)
test_df = pd.DataFrame(test_samples)
train_df.head(14)
# + colab={"base_uri": "https://localhost:8080/"} id="z4A6lpD1a61i" outputId="31b512d1-30e9-4848-8bff-254d4ac44bb8"
# Distribution of labels in training data
train_df.target.value_counts()
# + colab={"base_uri": "https://localhost:8080/", "height": 267} id="5mqVEHKpbH3o" outputId="d436df37-6732-486f-fe60-53e8b94c58b7"
train_df.total_lines.plot.hist();
# + [markdown] id="k-cjhPuebNeF"
# ### Get lists of sentences
# + colab={"base_uri": "https://localhost:8080/"} id="7olPj8Exb1ly" outputId="add884d7-0486-40ba-8fa9-ca590975207b"
# Convert abstract text lines into lists
train_sentences = train_df["text"].tolist()
val_sentences = val_df["text"].tolist()
test_sentences = test_df["text"].tolist()
len(train_sentences), len(val_sentences), len(test_sentences)
# + colab={"base_uri": "https://localhost:8080/"} id="DSU4YdF0b4U_" outputId="f5354887-0459-4320-b61b-ca06d68f3e46"
# View first 10 lines of training sentences
train_sentences[:10]
# + [markdown] id="0WmxmMebb7AV"
# ## Make numeric labels
#
# We're going to create one hot and label encoded labels.
#
# We could get away with just making label encoded labels, however, TensorFlow's CategoricalCrossentropy loss function likes to have one hot encoded labels (this will enable us to use label smoothing later on).
#
# To numerically encode labels we'll use Scikit-Learn's OneHotEncoder and LabelEncoder classes.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="7hqVk3rAcgbJ" outputId="2f6790f7-377c-4d85-b553-4b8573e051fe"
# One hot encode labels
from sklearn.preprocessing import OneHotEncoder
one_hot_encoder = OneHotEncoder(sparse=False)
train_labels_one_hot = one_hot_encoder.fit_transform(train_df["target"].to_numpy().reshape(-1, 1))
val_labels_one_hot = one_hot_encoder.transform(val_df["target"].to_numpy().reshape(-1, 1))
test_labels_one_hot = one_hot_encoder.transform(test_df["target"].to_numpy().reshape(-1, 1))
# Check what training labels look like
train_labels_one_hot
# + [markdown] id="FqZ4a-UCckXH"
# ### Label encode labels
# + colab={"base_uri": "https://localhost:8080/"} id="3_3fhlWVcnVX" outputId="cb50aff6-ff98-4bdf-b705-a7301c6cff33"
# Extract labels ("target" columns) and encode them into integers
from sklearn.preprocessing import LabelEncoder
label_encoder = LabelEncoder()
train_labels_encoded = label_encoder.fit_transform(train_df["target"].to_numpy())
val_labels_encoded = label_encoder.transform(val_df["target"].to_numpy())
test_labels_encoded = label_encoder.transform(test_df["target"].to_numpy())
# Check what training labels look like
train_labels_encoded
# + colab={"base_uri": "https://localhost:8080/"} id="gPHnRKKScpwV" outputId="834674f2-17d8-4cef-8f4c-b016f9a01e81"
# Get class names and number of classes from LabelEncoder instance
num_classes = len(label_encoder.classes_)
class_names = label_encoder.classes_
num_classes, class_names
# + [markdown] id="6XK9k-DHcwpE"
# # Creating a series of model experiments
# + [markdown] id="F4v7aRdvFInv"
# ## Model 0 : Getting a baseline
#
# Our first model we'll be a TF-IDF Multinomial Naive Bayes as recommended by Scikit-Learn's machine learning map.
#
# To build it, we'll create a Scikit-Learn Pipeline which uses the TfidfVectorizer class to convert our abstract sentences to numbers using the TF-IDF (term frequency-inverse document frequecy) algorithm and then learns to classify our sentences using the MultinomialNB aglorithm.
# + id="F2a4ozzxFPKc"
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.naive_bayes import MultinomialNB
from sklearn.pipeline import Pipeline
# Create a pipeline
model_0 = Pipeline([
("tf-idf", TfidfVectorizer()),
("clf", MultinomialNB())
])
# Fit the pipeline to the training data
model_0.fit(X=train_sentences,
y = train_labels_encoded);
# + colab={"base_uri": "https://localhost:8080/"} id="zgkr8gYaF5Pv" outputId="0e95e792-94db-4731-95a5-cb4fe5e3735f"
# Evaluate baseline on validation dataset
model_0.score(X = val_sentences,
y = val_labels_encoded)
# + colab={"base_uri": "https://localhost:8080/"} id="3rHdfS3bGENE" outputId="cb7af861-649d-45c8-bb5c-60c6320ee9ee"
# Make predictions
baseline_preds = model_0.predict(val_sentences)
baseline_preds
# + [markdown] id="FxVpyW4xGNZA"
# ### Download helper functions script
# + colab={"base_uri": "https://localhost:8080/"} id="7dO5dSbPGU-K" outputId="d7eb1354-6bc7-45b4-d449-a2ca0acc0643"
# Download helper functions script
# !wget https://raw.githubusercontent.com/prachuryanath/TF-Learning/main/extras/helper_functions.py
# + colab={"base_uri": "https://localhost:8080/"} id="UeJolLFJGaFG" outputId="05fa39d9-72d5-4fcc-df09-4028d65aa78e"
# Import calculate_results helper function
from helper_functions import calculate_results
# Calculate baseline results
baseline_results = calculate_results(y_true=val_labels_encoded,
y_pred=baseline_preds)
baseline_results
# + [markdown] id="LWE3W4EYGfEy"
# ## Preparing our data for deep sequence models
#
# Excellent! We've got a working baseline to try and improve upon.
#
# But before we start building deeper models, we've got to create vectorization and embedding layers.
#
# The vectorization layer will convert our text to numbers and the embedding layer will capture the relationships between those numbers.
#
# To start creating our vectorization and embedding layers, we'll need to import the appropriate libraries (namely TensorFlow and NumPy).
# + id="0SUIHdX1GrOm"
import numpy as np
import tensorflow as tf
from tensorflow.keras import layers
# + colab={"base_uri": "https://localhost:8080/"} id="Tx7tIeidIbX6" outputId="008dd622-dfff-43d3-b48b-4e686106e6f6"
# How long is each sentence on average ?
sent_lens = [len(sentence.split()) for sentence in train_sentences]
avg_sent_lens = np.mean(sent_lens)
avg_sent_lens
# + colab={"base_uri": "https://localhost:8080/", "height": 0} id="IEbJrjoIIrfy" outputId="35c522cb-650a-4209-a026-dd068e48617c"
# What's the distribution look like ?
import matplotlib.pyplot as plt
plt.hist(sent_lens, bins=15)
# + colab={"base_uri": "https://localhost:8080/"} id="je8OphXJI_4T" outputId="ffab61e2-542b-4b08-b45e-55419163b9af"
# How long of a sentence covers 95% of the lengths?
output_seq_len = int(np.percentile(sent_lens, 95))
output_seq_len
# + [markdown] id="xWAg6f0oJPeA"
# Wonderful! It looks like 95% of the sentences in our training set have a length of 55 tokens or less.
#
# When we create our tokenization layer, we'll use this value to turn all of our sentences into the same length. Meaning sentences with a length below 55 get padded with zeros and sentences with a length above 55 get truncated (words after 55 get cut off).
# + colab={"base_uri": "https://localhost:8080/"} id="FE5cD-M-JYaM" outputId="2179d227-a69c-4b4d-b36f-7084fe473aaa"
# Maximum sentence length in the training set
max(sent_lens)
# + [markdown] id="EnUNv660JaSG"
# ### Create text vectorizer
#
# Now we've got a little more information about our texts, let's create a way to turn it into numbers.
#
# To do so, we'll use the TextVectorization layer from TensorFlow.
#
# We'll keep all the parameters default except for max_tokens (the number of unique words in our dataset) and output_sequence_length (our desired output length for each vectorized sentence).
#
#
# + id="VuMMxPTIJori"
# How many words are in our vocabulary? (taken from 3.2 in https://arxiv.org/pdf/1710.06071.pdf)
max_tokens = 68000
# + id="kX-T69OpJqts"
# Create text vectorizer
from tensorflow.keras.layers.experimental.preprocessing import TextVectorization
text_vectorizer = TextVectorization(max_tokens=max_tokens, # number of words in vocabulary
output_sequence_length=55) # desired output length of vectorized sequences
# + [markdown] id="jS_MU-0KJz9i"
# Great! Looks like our text_vectorizer is ready, let's adapt it to the training data (let it read the training data and figure out what number should represent what word) and then test it out.
# + id="XmIQcCE1JvO7"
# Adapt text vectorizer to training sentences
text_vectorizer.adapt(train_sentences)
# + colab={"base_uri": "https://localhost:8080/"} id="vbymR2tRJyLZ" outputId="86960fc8-dba4-407f-df98-8a47f8262977"
# Test out text vectorizer
import random
target_sentence = random.choice(train_sentences)
print(f"Text:\n{target_sentence}")
print(f"\nLength of text: {len(target_sentence.split())}")
print(f"\nVectorized text:\n{text_vectorizer([target_sentence])}")
# + colab={"base_uri": "https://localhost:8080/"} id="FAqJhJOJJ32h" outputId="1c6a8940-2766-4f1d-85bb-f05b522d54a9"
# How many words in our training vocabulary?
rct_20k_text_vocab = text_vectorizer.get_vocabulary()
print(f"Number of words in vocabulary: {len(rct_20k_text_vocab)}"),
print(f"Most common words in the vocabulary: {rct_20k_text_vocab[:5]}")
print(f"Least common words in the vocabulary: {rct_20k_text_vocab[-5:]}")
# + colab={"base_uri": "https://localhost:8080/"} id="ESWjEHYMKD5t" outputId="513a1398-87aa-4f24-a150-00cab8aba2fd"
# Get the config of our text vectorizer
text_vectorizer.get_config()
# + [markdown] id="ikIQmf_fKKNI"
# ### Create custom text embedding
#
# Our token_vectorization layer maps the words in our text directly to numbers. However, this doesn't necessarily capture the relationships between those numbers.
#
# To create a richer numerical representation of our text, we can use an **embedding**.
#
# As our model learns (*by going through many different examples of abstract sentences and their labels*), it'll update its embedding to better represent the relationships between tokens in our corpus.
#
# We can create a trainable embedding layer using TensorFlow's `Embedding layer`.
#
# Once again, the main parameters we're concerned with here are the inputs and outputs of our `Embedding layer`.
#
# The input_dim parameter defines the size of our vocabulary. And the output_dim parameter defines the dimension of the embedding output.
#
# Once created, our embedding layer will take the integer outputs of our `text_vectorization` layer as inputs and convert them to feature vectors of size `output_dim`.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="Hq32taDUMUHl" outputId="d3fef36c-c9c7-47da-dc5b-949252e14eb1"
# Create token embedding layer
token_embed = layers.Embedding(input_dim=len(rct_20k_text_vocab), # length of vocabulary
output_dim=128, # Note: different embedding sizes result in drastically different numbers of parameters to train
# Use masking to handle variable sequence lengths (save space)
mask_zero=True,
name="token_embedding")
# Show example embedding
print(f"Sentence before vectorization:\n{target_sentence}\n")
vectorized_sentence = text_vectorizer([target_sentence])
print(f"Sentence after vectorization (before embedding):\n{vectorized_sentence}\n")
embedded_sentence = token_embed(vectorized_sentence)
print(f"Sentence after embedding:\n{embedded_sentence}\n")
print(f"Embedded sentence shape: {embedded_sentence.shape}")
# + [markdown] id="rBbbVDpKMwbH"
# ### Create datasets
# Namely, the tf.data API provides methods which enable faster data loading.
#
# The main steps we'll want to use with our data is to turn it into a `PrefetchDataset` of batches.
#
# Doing so we'll ensure TensorFlow loads our data onto the GPU as fast as possible, in turn leading to faster training time.
#
# To create a batched PrefetchDataset we can use the methods `batch()` and `prefetch()`, the parameter `tf.data.AUTOTUNE` will also allow TensorFlow to determine the optimal amount of compute to use to prepare datasets.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="uZXBLJ62NBZ2" outputId="6a5af520-2fdb-46c8-d0c1-75c10298354b"
# Turn our data into TensorFlow Datasets
train_dataset = tf.data.Dataset.from_tensor_slices((train_sentences, train_labels_one_hot))
valid_dataset = tf.data.Dataset.from_tensor_slices((val_sentences, val_labels_one_hot))
test_dataset = tf.data.Dataset.from_tensor_slices((test_sentences, test_labels_one_hot))
train_dataset
# + colab={"base_uri": "https://localhost:8080/"} id="_rrX-anYNPe7" outputId="c2c89d3a-810a-4f06-9a15-87af734fbe0e"
# Take the TensorSliceDataset's and turn them into prefetched batches
train_dataset = train_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
valid_dataset = valid_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
test_dataset = test_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
train_dataset
# + [markdown] id="_dn9x-rwNRgQ"
# ## Model 1 : Conv1D with token embeddings
#
# All of our deep models will follow a similar structure:
#
# `Input (text) -> Tokenize -> Embedding -> Layers -> Output (label probability)`
#
# The main component we'll be changing throughout is the Layers component. Because any modern deep NLP model requires text to be converted into an embedding before meaningful patterns can be discovered within.
#
# The first model we're going to build is a 1-dimensional Convolutional Neural Network.
#
# + id="StCVFrLwNg4j"
# Create 1D convolutional model to process sequences
inputs = layers.Input(shape=(1,), dtype=tf.string)
text_vectors = text_vectorizer(inputs) # vectorize text inputs
token_embeddings = token_embed(text_vectors) # create embedding
x = layers.Conv1D(64, kernel_size=5, padding="same", activation="relu")(token_embeddings)
x = layers.GlobalAveragePooling1D()(x) # condense the output of our feature vector
outputs = layers.Dense(num_classes, activation="softmax")(x)
model_1 = tf.keras.Model(inputs, outputs)
# Compile
model_1.compile(loss="categorical_crossentropy", # if your labels are integer form (not one hot) use sparse_categorical_crossentropy
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# + colab={"base_uri": "https://localhost:8080/"} id="mPA6RZ65N5dk" outputId="a4c3a260-7394-4a59-fb60-d379d733e3d7"
# Get summary of Conv1D model
model_1.summary()
# + [markdown] id="Bz3ZVL8NN-E8"
# Since our training data contains nearly 200,000 sentences, fitting a deep model may take a while even with a GPU. So to keep our experiments swift, we're going to run them on a subset of the training dataset.
#
# More specifically, we'll only use the first 10% of batches (about 18,000 samples) of the training set to train on and the first 10% of batches from the validation set to validate on.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="K7GdhSyXOQEG" outputId="c1ee618e-0519-48f3-9ad7-61a0864934a4"
# Fit the model
model_1_history = model_1.fit(train_dataset,
steps_per_epoch=int(0.1 * len(train_dataset)), # only fit on 10% of batches for faster training time
epochs=3,
validation_data=valid_dataset,
validation_steps=int(0.1 * len(valid_dataset))) # only validate on 10% of batches
# + colab={"base_uri": "https://localhost:8080/"} id="TwM9iMbNOVkM" outputId="af17d4e0-af2c-443e-b08d-a27b86648cb6"
# Evaluate on whole validation dataset (we only validated on 10% of batches during training)
model_1.evaluate(valid_dataset)
# + colab={"base_uri": "https://localhost:8080/"} id="1m_AXUlQP_Fa" outputId="a5b4755c-4abb-4d78-cbd9-db44a374380d"
# Make predictions (our model outputs prediction probabilities for each class)
model_1_pred_probs = model_1.predict(valid_dataset)
model_1_pred_probs
# + colab={"base_uri": "https://localhost:8080/"} id="on46vMnoQCjy" outputId="a8d65371-d9e1-4b0d-e045-79aee34b1aaf"
# Convert pred probs to classes
model_1_preds = tf.argmax(model_1_pred_probs, axis=1)
model_1_preds
# + colab={"base_uri": "https://localhost:8080/"} id="joubzlk9TfHY" outputId="6d95727d-084f-42d5-e598-16c62db2f84f"
# Calculate model_1 results
model_1_results = calculate_results(y_true=val_labels_encoded,
y_pred=model_1_preds)
model_1_results
# + [markdown] id="ioEtyCWVThue"
# ## Model 2 : Feature extraction with pretrained token embeddings
#
# The model structure will look like:
#
# `Inputs (string) -> Pretrained embeddings from TensorFlow Hub (Universal Sentence Encoder) -> Layers -> Output (prediction probabilities)`
#
# You'll notice the lack of tokenization layer we've used in a previous model. This is because the `Universal Sentence Encoder (USE)` takes care of tokenization for us.
#
# This type of model is called transfer learning, or more specifically, **feature extraction transfer learning**. In other words, taking the patterns a model has learned elsewhere and applying it to our own problem.
# + id="8S5Yh76iT76y"
# Download pretrained TensorFlow Hub USE
import tensorflow_hub as hub
tf_hub_embedding_layer = hub.KerasLayer("https://tfhub.dev/google/universal-sentence-encoder/4",
trainable=False,
name="universal_sentence_encoder")
# + colab={"base_uri": "https://localhost:8080/"} id="rnTm9-ArUPWI" outputId="330360aa-d933-4594-cffc-18f8d4cf5809"
# Test out the embedding on a random sentence
random_training_sentence = random.choice(train_sentences)
print(f"Random training sentence:\n{random_training_sentence}\n")
use_embedded_sentence = tf_hub_embedding_layer([random_training_sentence])
print(f"Sentence after embedding:\n{use_embedded_sentence[0][:30]} (truncated output)...\n")
print(f"Length of sentence embedding:\n{len(use_embedded_sentence[0])}")
# + [markdown] id="0BrYwbV7Ui9W"
# ### Building and fitting an NLP feature extraction model from TensorFlow Hub
# + id="o3PJYUHkUry3"
# Define feature extractor model using TF Hub layer
inputs = layers.Input(shape=[], dtype=tf.string)
pretrained_embedding = tf_hub_embedding_layer(inputs) # tokenize text and create embedding
x = layers.Dense(128, activation="relu")(pretrained_embedding) # add a fully connected layer on top of the embedding
# Note: you could add more layers here if you wanted to
outputs = layers.Dense(5, activation="softmax")(x) # create the output layer
model_2 = tf.keras.Model(inputs=inputs,
outputs=outputs)
# Compile the model
model_2.compile(loss="categorical_crossentropy",
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# + colab={"base_uri": "https://localhost:8080/"} id="JWzn7queUvJn" outputId="3fca8b06-d823-42eb-998e-e4cb7a306d45"
# Get a summary of the model
model_2.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="IFlxT8IjUxGv" outputId="cd1989bd-f14f-40e0-d6e2-558962d9f174"
# Fit feature extractor model for 3 epochs
model_2.fit(train_dataset,
steps_per_epoch=int(0.1 * len(train_dataset)),
epochs=3,
validation_data=valid_dataset,
validation_steps=int(0.1 * len(valid_dataset)))
# + colab={"base_uri": "https://localhost:8080/"} id="oYO8oGtZU1o9" outputId="f6e16259-adb1-4fa4-a5dc-a8bd467bf0b6"
# Evaluate on whole validation dataset
model_2.evaluate(valid_dataset)
# + colab={"base_uri": "https://localhost:8080/"} id="ZUxpOCHyVEhI" outputId="8451e4bf-ccbd-4a3d-9610-31474be6661b"
# Make predictions with feature extraction model
model_2_pred_probs = model_2.predict(valid_dataset)
model_2_pred_probs
# + colab={"base_uri": "https://localhost:8080/"} id="kY2TbJqLVGLd" outputId="5a61e48b-f115-4644-9e4d-91def4f34388"
# Convert the predictions with feature extraction model to classes
model_2_preds = tf.argmax(model_2_pred_probs, axis=1)
model_2_preds
# + id="Env6Ch23VINu" colab={"base_uri": "https://localhost:8080/"} outputId="b1f66511-4f62-47f0-8cb6-c2e60689c8ca"
# Calculate results from TF Hub pretrained embeddings results on validation set
model_2_results = calculate_results(y_true=val_labels_encoded,
y_pred=model_2_preds)
model_2_results
# + [markdown] id="iJiGRfXgVKT0"
# ## Model 3: Conv1D with character embeddings
# + [markdown] id="bC2BUk8p5n_4"
# ### Creating a character-level tokenizer
#
# We've built models with a custom token embedding and a pretrained token embedding, how about we build one using a character embedding?
#
# The difference between a character and token embedding is that the character embedding is created using sequences split into characters *(e.g. hello -> [h, e, l, l, o])* where as a token embedding is created on sequences split into tokens.
#
# We can create a character-level embedding by first vectorizing our sequences (after they've been split into characters) using the `TextVectorization` class and then passing those vectorized sequences through an `Embedding` layer.
#
# Before we can vectorize our sequences on a character-level we'll need to split them into characters. Let's write a function to do so.
# + colab={"base_uri": "https://localhost:8080/", "height": 69} id="lkAYI4dl5yg9" outputId="93e3fd47-d18d-47cc-9b2c-3e84438dc218"
# Make function to split sentences into characters
def split_chars(text):
return " ".join(list(text))
# Test splitting non-character-level sequence into characters
split_chars(random_training_sentence)
# + colab={"base_uri": "https://localhost:8080/"} id="-ltr7Aa_6D18" outputId="bde94515-2822-4cca-a63f-20a2c28df5d0"
# Split sequence-level data splits into character-level data splits
train_chars = [split_chars(sentence) for sentence in train_sentences]
val_chars = [split_chars(sentence) for sentence in val_sentences]
test_chars = [split_chars(sentence) for sentence in test_sentences]
print(train_chars[0])
# + colab={"base_uri": "https://localhost:8080/"} id="lu5fg7Gq6PTL" outputId="37906ae6-7946-4cf7-a72a-0a4067ec8f5e"
# What's the average character length?
char_lens = [len(sentence) for sentence in train_sentences]
mean_char_len = np.mean(char_lens)
mean_char_len
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="8wXXI0Ol6Sk0" outputId="7504df6b-0bcb-47fa-9615-4a93b68a2f92"
# Check the distribution of our sequences at character-level
import matplotlib.pyplot as plt
plt.hist(char_lens, bins=7);
# + colab={"base_uri": "https://localhost:8080/"} id="4ifuLnxi6WWN" outputId="9f0d7f80-1f22-48b6-a908-8c104333ff0c"
# Find what character length covers 95% of seuences
output_seq_char_len = int(np.percentile(char_lens, 95))
output_seq_char_len
# + [markdown] id="6_-KpsgC6cup"
# We'll set max_tokens (the total number of different characters in our sequences) to 28, in other words, 26 letters of the alphabet + space + OOV (out of vocabulary or unknown) tokens.
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="5AH0YKoX6k_D" outputId="6b3bfb91-3c56-4ec2-9cca-e6ac884b0543"
# Get all keyboard characters for char-level embedding
import string
alphabet = string.ascii_lowercase + string.digits + string.punctuation
alphabet
# + id="gkfcUvwY6m8S"
# Create char-level token vectorizer instance
NUM_CHAR_TOKENS = len(alphabet) + 2 # num characters in alphabet + space + OOV token
char_vectorizer = TextVectorization(max_tokens=NUM_CHAR_TOKENS,
output_sequence_length=output_seq_char_len,
standardize="lower_and_strip_punctuation",
name="char_vectorizer")
# Adapt character vectorizer to training characters
char_vectorizer.adapt(train_chars)
# + colab={"base_uri": "https://localhost:8080/"} id="1Rb3poVE6pif" outputId="f33baf86-b769-4847-c298-0079b7912a9d"
# Check character vocabulary characteristics
char_vocab = char_vectorizer.get_vocabulary()
print(f"Number of different characters in character vocab: {len(char_vocab)}")
print(f"5 most common characters: {char_vocab[:5]}")
print(f"5 least common characters: {char_vocab[-5:]}")
# + colab={"base_uri": "https://localhost:8080/"} id="SIK0hHQE6tUX" outputId="a6274524-72e3-4042-a9d3-12a9b335fd6b"
# Test out character vectorizer
random_train_chars = random.choice(train_chars)
print(f"Charified text:\n{random_train_chars}")
print(f"\nLength of chars: {len(random_train_chars.split())}")
vectorized_chars = char_vectorizer([random_train_chars])
print(f"\nVectorized chars:\n{vectorized_chars}")
print(f"\nLength of vectorized chars: {len(vectorized_chars[0])}")
# + [markdown] id="6xCvrv496vtE"
# You'll notice sequences with a length shorter than 290 (`output_seq_char_length`) get padded with zeros on the end, this ensures all sequences passed to our model are the same length.
#
# Also, due to the standardize parameter of `TextVectorization` being "`lower_and_strip_punctuation`" and the split parameter being "`whitespace`" by default, symbols (such as @) and spaces are removed.
# + [markdown] id="wL3O6RWI7yGH"
# ### Creating a character-level embedding
#
# We've got a way to vectorize our character-level sequences, now's time to create a character-level embedding.
#
# Just like our custom token embedding, we can do so using the `tensorflow.keras.layers.Embedding` class.
#
# Our character-level embedding layer requires an input dimension and output dimension.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="S_llxFGT8CWD" outputId="fc3ee612-e10e-417c-8faa-eed8ac85c2ca"
# Create char embedding layer
char_embed = layers.Embedding(input_dim=NUM_CHAR_TOKENS, # number of different characters
output_dim=25, # embedding dimension of each character (same as Figure 1 in https://arxiv.org/pdf/1612.05251.pdf)
mask_zero=True,
name="char_embed")
# Test out character embedding layer
print(f"Charified text (before vectorization and embedding):\n{random_train_chars}\n")
char_embed_example = char_embed(char_vectorizer([random_train_chars]))
print(f"Embedded chars (after vectorization and embedding):\n{char_embed_example}\n")
print(f"Character embedding shape: {char_embed_example.shape}")
# + [markdown] id="SvJnKmS_644R"
# ### Building a Conv1D model to fit on character embeddings
#
# Now we've got a way to turn our character-level sequences into numbers (`char_vectorizer`) as well as numerically represent them as an embedding (`char_embed`) let's test how effective they are at encoding the information in our sequences by creating a character-level sequence model.
#
# The model will have the same structure as our custom token embedding model (`model_1`) except it'll take character-level sequences as input instead of token-level sequences.
#
# `Input (character-level text) -> Tokenize -> Embedding -> Layers (Conv1D, GlobalMaxPool1D) -> Output (label probability)`
#
# + id="hN0SlIG57ORy"
# Make Conv1D on chars only
inputs = layers.Input(shape=(1,), dtype="string")
char_vectors = char_vectorizer(inputs)
char_embeddings = char_embed(char_vectors)
x = layers.Conv1D(64, kernel_size=5, padding="same", activation="relu")(char_embeddings)
x = layers.GlobalMaxPool1D()(x)
outputs = layers.Dense(num_classes, activation="softmax")(x)
model_3 = tf.keras.Model(inputs=inputs,
outputs=outputs,
name="model_3_conv1D_char_embedding")
# Compile model
model_3.compile(loss="categorical_crossentropy",
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# + colab={"base_uri": "https://localhost:8080/"} id="6ofjKXir7cQF" outputId="bf4721b1-c56d-4b39-da31-5f0ddb8c0a3d"
# Check the summary of conv1d_char_model
model_3.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="Z9kZizVf7eHQ" outputId="e6915345-145d-46dd-e487-7d8c4c03c9cd"
# Create char datasets
train_char_dataset = tf.data.Dataset.from_tensor_slices((train_chars, train_labels_one_hot)).batch(32).prefetch(tf.data.AUTOTUNE)
val_char_dataset = tf.data.Dataset.from_tensor_slices((val_chars, val_labels_one_hot)).batch(32).prefetch(tf.data.AUTOTUNE)
train_char_dataset
# + colab={"base_uri": "https://localhost:8080/"} id="iDJ8Jvct7gY1" outputId="d09fbb2a-4cd1-4645-dd96-c41663720ca7"
# Fit the model on chars only
model_3_history = model_3.fit(train_char_dataset,
steps_per_epoch=int(0.1 * len(train_char_dataset)),
epochs=3,
validation_data=val_char_dataset,
validation_steps=int(0.1 * len(val_char_dataset)))
# + colab={"base_uri": "https://localhost:8080/"} id="PXxPl8fwdZ5P" outputId="26a96ffb-3a68-4efd-9597-b86730ef4403"
# Evaluate model_3 on whole validation char dataset
model_3.evaluate(val_char_dataset)
# + colab={"base_uri": "https://localhost:8080/"} id="B6-NoSBmdea8" outputId="eb7f9278-629b-4a1b-b85b-bbc3288047a6"
# Make predictions with character model only
model_3_pred_probs = model_3.predict(val_char_dataset)
model_3_pred_probs
# + colab={"base_uri": "https://localhost:8080/"} id="t2LvtfOvdeJ7" outputId="c6f5963d-29df-429a-c767-8ccb5a296d50"
# Convert predictions to classes
model_3_preds = tf.argmax(model_3_pred_probs, axis=1)
model_3_preds
# + colab={"base_uri": "https://localhost:8080/"} id="QP0_u8w-dh8Q" outputId="9759fc2b-a533-4e4f-c1e3-493501c4ba6b"
# Calculate Conv1D char only model results
model_3_results = calculate_results(y_true=val_labels_encoded,
y_pred=model_3_preds)
model_3_results
# + [markdown] id="Hs03Xb7K7kKt"
# ## Model 4: Combining pretrained token embeddings + character embeddings (hybrid embedding layer)
#
# To start replicating (or getting close to replicating) the model in Figure 1, we're going to go through the following steps:
#
# 1. Create a token-level model (similar to `model_1`)
# 2. Create a character-level model (similar to `model_3` with a slight modification to reflect the paper)
# 3. Combine (using `layers.Concatenate`) the outputs of 1 and 2
# 4. Build a series of output layers on top of 3
# 5. Construct a model which takes token and character-level sequences as input and produces sequence label probabilities as output
# + id="AgTvcwOt8_mZ"
# 1. Setup token inputs/model
token_inputs = layers.Input(shape=[], dtype=tf.string, name="token_input")
token_embeddings = tf_hub_embedding_layer(token_inputs)
token_output = layers.Dense(128, activation="relu")(token_embeddings)
token_model = tf.keras.Model(inputs=token_inputs,
outputs=token_output)
# 2. Setup char inputs/model
char_inputs = layers.Input(shape=(1,), dtype=tf.string, name="char_input")
char_vectors = char_vectorizer(char_inputs)
char_embeddings = char_embed(char_vectors)
char_bi_lstm = layers.Bidirectional(layers.LSTM(25))(char_embeddings) # bi-LSTM shown in Figure 1 of https://arxiv.org/pdf/1612.05251.pdf
char_model = tf.keras.Model(inputs=char_inputs,
outputs=char_bi_lstm)
# 3. Concatenate token and char inputs (create hybrid token embedding)
token_char_concat = layers.Concatenate(name="token_char_hybrid")([token_model.output,
char_model.output])
# 4. Create output layers - addition of dropout discussed in 4.2 of https://arxiv.org/pdf/1612.05251.pdf
combined_dropout = layers.Dropout(0.5)(token_char_concat)
combined_dense = layers.Dense(200, activation="relu")(combined_dropout) # slightly different to Figure 1 due to different shapes of token/char embedding layers
final_dropout = layers.Dropout(0.5)(combined_dense)
output_layer = layers.Dense(num_classes, activation="softmax")(final_dropout)
# 5. Construct model with char and token inputs
model_4 = tf.keras.Model(inputs=[token_model.input, char_model.input],
outputs=output_layer,
name="model_4_token_and_char_embeddings")
# + colab={"base_uri": "https://localhost:8080/"} id="sk9S9v1B9QUz" outputId="9e94b5fc-788a-41b5-94ad-811ac583dc19"
# Get summary of token and character model
model_4.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 856} id="ayc8_xZk9rJU" outputId="3b8c70e5-c261-424d-d079-2d66c18f7395"
# Plot hybrid token and character model
from tensorflow.keras.utils import plot_model
plot_model(model_4)
# + id="bG9W0kEP9ve_"
# Compile token char model
model_4.compile(loss="categorical_crossentropy",
optimizer=tf.keras.optimizers.Adam(), # section 4.2 of https://arxiv.org/pdf/1612.05251.pdf mentions using SGD but we'll stick with Adam
metrics=["accuracy"])
# + [markdown] id="x_Q9rK8I96lF"
# ### Combining token and character data into a tf.data dataset
#
# To keep our experiments fast, we'll fit our token-character-hybrid model on 10% of training and validate on 10% of validation batches. However, the difference with this model is that it requires two inputs, token-level sequences and character-level sequences.
#
# We can do this by create a tf.data.Dataset with a tuple as it's first input, for example:
#
# * `((token_data, char_data), (label))`
# + id="8lmlsJF5_KsT"
# Combine chars and tokens into a dataset
train_char_token_data = tf.data.Dataset.from_tensor_slices((train_sentences, train_chars)) # make data
train_char_token_labels = tf.data.Dataset.from_tensor_slices(train_labels_one_hot) # make labels
train_char_token_dataset = tf.data.Dataset.zip((train_char_token_data, train_char_token_labels)) # combine data and labels
# Prefetch and batch train data
train_char_token_dataset = train_char_token_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
# Repeat same steps validation data
val_char_token_data = tf.data.Dataset.from_tensor_slices((val_sentences, val_chars))
val_char_token_labels = tf.data.Dataset.from_tensor_slices(val_labels_one_hot)
val_char_token_dataset = tf.data.Dataset.zip((val_char_token_data, val_char_token_labels))
val_char_token_dataset = val_char_token_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
# + colab={"base_uri": "https://localhost:8080/"} id="n7u_Zwgf_NON" outputId="a71cee1c-42ff-41ee-8282-d3f4fe46087e"
# Check out training char and token embedding dataset
train_char_token_dataset, val_char_token_dataset
# + [markdown] id="83dkpfH5_PMa"
# ### Fitting a model on token and character-level sequences
# + colab={"base_uri": "https://localhost:8080/"} id="A8omi3OT_eUe" outputId="15426200-7a52-48fb-e112-d22851b3f752"
# Fit the model on tokens and chars
model_4_history = model_4.fit(train_char_token_dataset, # train on dataset of token and characters
steps_per_epoch=int(0.1 * len(train_char_token_dataset)),
epochs=3,
validation_data=val_char_token_dataset,
validation_steps=int(0.1 * len(val_char_token_dataset)))
# + colab={"base_uri": "https://localhost:8080/"} id="CtKW5j6O_gEg" outputId="c18456d0-e38f-461e-e593-6c522be6e0da"
# Evaluate on the whole validation dataset
model_4.evaluate(val_char_token_dataset)
# + colab={"base_uri": "https://localhost:8080/"} id="jWbtU3IA_hpz" outputId="4af93dc2-1c40-4667-aabc-9d1a6c720282"
# Make predictions using the token-character model hybrid
model_4_pred_probs = model_4.predict(val_char_token_dataset)
model_4_pred_probs
# + colab={"base_uri": "https://localhost:8080/"} id="Iskz6lBN_jnU" outputId="cec39ad6-9665-4daf-fee9-601b55ffd175"
# Turn prediction probabilities into prediction classes
model_4_preds = tf.argmax(model_4_pred_probs, axis=1)
model_4_preds
# + colab={"base_uri": "https://localhost:8080/"} id="rPlHumgd_lfU" outputId="3cd71c7e-0f05-421a-ba16-f35c4e7aff54"
# Get results of token-char-hybrid model
model_4_results = calculate_results(y_true=val_labels_encoded,
y_pred=model_4_preds)
model_4_results
# + [markdown] id="yoxYXqHmYRwe"
# ## Model 5: Transfer Learning with pretrained token embeddings + character embeddings + positional embeddings
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="I1wxQygsYfZr" outputId="ffbfab76-f6ca-496a-acdc-9e71606db72e"
# Inspect training dataframe
train_df.head()
# + [markdown] id="Q7XR-gAGYine"
# ### Create positional embeddings
#
# Since our "`line_number`" and "`total_line`" columns are already numerical, we could pass them as they are to our model.
#
# But to avoid our model thinking a line with "`line_number`"=5 is five times greater than a line with "`line_number`"=1, we'll use one-hot-encoding to encode our "`line_number`" and "`total_lines`" features.
#
# To do this, we can use the `tf.one_hot` utility.
#
# `tf.one_hot` returns a one-hot-encoded tensor. It accepts an array (or tensor) as input and the depth parameter determines the dimension of the returned tensor.
#
# To figure out what we should set the depth parameter to, let's investigate the distribution of the "`line_number`" column.
# + colab={"base_uri": "https://localhost:8080/"} id="bp1S771WY2Oe" outputId="7520a10c-75e5-43b3-c80c-4aab838f4d59"
# How many different line numbers are there?
train_df["line_number"].value_counts()
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="wRhZdKemY8Qp" outputId="b2354fdc-1ba4-490f-c7c1-6482bee3613d"
# Check the distribution of "line_number" column
train_df.line_number.plot.hist()
# + id="2LRqNfRfY_SM"
# Use TensorFlow to create one-hot-encoded tensors of our "line_number" column
train_line_numbers_one_hot = tf.one_hot(train_df["line_number"].to_numpy(), depth=15)
val_line_numbers_one_hot = tf.one_hot(val_df["line_number"].to_numpy(), depth=15)
test_line_numbers_one_hot = tf.one_hot(test_df["line_number"].to_numpy(), depth=15)
# + [markdown] id="HpotHQuMY_O0"
# Setting the depth parameter of tf.one_hot to 15 means any sample with a "`line_number`" value of over 15 gets set to a tensor of all 0's, where as any sample with a "`line_number`" of under 15 gets turned into a tensor of all 0's but with a 1 at the index equal to the "`line_number`" value.
#
# + colab={"base_uri": "https://localhost:8080/"} id="87xBjEeHY_MM" outputId="d4b599dc-2c5e-4521-caf6-666100acce3f"
# Check one-hot encoded "line_number" feature samples
train_line_numbers_one_hot.shape, train_line_numbers_one_hot[:20]
# + colab={"base_uri": "https://localhost:8080/"} id="-T4lhMV6Y_Js" outputId="8831e93e-9218-41df-c5b3-48cf32d63690"
# How many different numbers of lines are there?
train_df["total_lines"].value_counts()
# + colab={"base_uri": "https://localhost:8080/", "height": 267} id="EuoXIpdUY_Hn" outputId="4d3b50a8-9e94-4e3a-99d1-ac35c4f4411c"
# Check the distribution of total lines
train_df.total_lines.plot.hist();
# + colab={"base_uri": "https://localhost:8080/"} id="vL87NPSgY-5K" outputId="139440b7-918a-4f80-f26e-dd428c335920"
# Check the coverage of a "total_lines" value of 20
np.percentile(train_df.total_lines, 98) # a value of 20 covers 98% of samples
# + colab={"base_uri": "https://localhost:8080/"} id="gNafwWXDZPce" outputId="4cef5df3-c2df-4bd4-dfdb-acd59167db7d"
# Use TensorFlow to create one-hot-encoded tensors of our "total_lines" column
train_total_lines_one_hot = tf.one_hot(train_df["total_lines"].to_numpy(), depth=20)
val_total_lines_one_hot = tf.one_hot(val_df["total_lines"].to_numpy(), depth=20)
test_total_lines_one_hot = tf.one_hot(test_df["total_lines"].to_numpy(), depth=20)
# Check shape and samples of total lines one-hot tensor
train_total_lines_one_hot.shape, train_total_lines_one_hot[:10]
# + [markdown] id="ejQ1zA2nZXtm"
# ### Building a tribid embedding model
#
# We're going to go through the following steps:
#
# 1. Create a token-level model (similar to `model_1`)
# 2. Create a character-level model (similar to `model_3` with a slight modification to reflect the paper)
# 3. Create a "`line_number`" model (takes in one-hot-encoded "`line_number`" tensor and passes it through a non-linear layer)
# 4. Create a "`total_lines`" model (takes in one-hot-encoded "`total_lines`" tensor and passes it through a non-linear layer)
# 5. Combine (using `layers.Concatenate`) the outputs of 1 and 2 into a token-character-hybrid embedding and pass it series of output to Figure 1 and section 4.2 of Neural Networks for Joint Sentence Classification in Medical Paper Abstracts
# 6. Combine (using `layers.Concatenate`) the outputs of 3, 4 and 5 into a token-character-positional tribrid embedding
# 7. Create an output layer to accept the tribrid embedding and output predicted label probabilities
# 8. Combine the inputs of 1, 2, 3, 4 and outputs of 7 into a tf.keras.Model
# + id="6PPo_dYnZXah"
# 1. Token inputs
token_inputs = layers.Input(shape=[], dtype="string", name="token_inputs")
token_embeddings = tf_hub_embedding_layer(token_inputs)
token_outputs = layers.Dense(128, activation="relu")(token_embeddings)
token_model = tf.keras.Model(inputs=token_inputs,
outputs=token_embeddings)
# 2. Char inputs
char_inputs = layers.Input(shape=(1,), dtype="string", name="char_inputs")
char_vectors = char_vectorizer(char_inputs)
char_embeddings = char_embed(char_vectors)
char_bi_lstm = layers.Bidirectional(layers.LSTM(32))(char_embeddings)
char_model = tf.keras.Model(inputs=char_inputs,
outputs=char_bi_lstm)
# 3. Line numbers inputs
line_number_inputs = layers.Input(shape=(15,), dtype=tf.int32, name="line_number_input")
x = layers.Dense(32, activation="relu")(line_number_inputs)
line_number_model = tf.keras.Model(inputs=line_number_inputs,
outputs=x)
# 4. Total lines inputs
total_lines_inputs = layers.Input(shape=(20,), dtype=tf.int32, name="total_lines_input")
y = layers.Dense(32, activation="relu")(total_lines_inputs)
total_line_model = tf.keras.Model(inputs=total_lines_inputs,
outputs=y)
# 5. Combine token and char embeddings into a hybrid embedding
combined_embeddings = layers.Concatenate(name="token_char_hybrid_embedding")([token_model.output,
char_model.output])
z = layers.Dense(256, activation="relu")(combined_embeddings)
z = layers.Dropout(0.5)(z)
# 6. Combine positional embeddings with combined token and char embeddings into a tribrid embedding
z = layers.Concatenate(name="token_char_positional_embedding")([line_number_model.output,
total_line_model.output,
z])
# 7. Create output layer
output_layer = layers.Dense(5, activation="softmax", name="output_layer")(z)
# 8. Put together model
model_5 = tf.keras.Model(inputs=[line_number_model.input,
total_line_model.input,
token_model.input,
char_model.input],
outputs=output_layer)
# + colab={"base_uri": "https://localhost:8080/"} id="548OtDhVZXXN" outputId="d374c4ae-ccc6-4115-ccaf-c160f089d8c1"
# Get a summary of our token, char and positional embedding model
model_5.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 707} id="0Y4xa9pDZXUQ" outputId="90597d89-2c71-494b-fb52-073ef360cbb6"
# Plot the token, char, positional embedding model
from tensorflow.keras.utils import plot_model
plot_model(model_5)
# + colab={"base_uri": "https://localhost:8080/"} id="jljyZqfnZXQd" outputId="78bc032d-a548-4c78-f016-135793073f9e"
# Check which layers of our model are trainable or not
for layer in model_5.layers:
print(layer, layer.trainable)
# + id="0C4fPi0mZXNA"
# Compile token, char, positional embedding model
model_5.compile(loss=tf.keras.losses.CategoricalCrossentropy(label_smoothing=0.2), # add label smoothing (examples which are really confident get smoothed a little)
optimizer=tf.keras.optimizers.Adam(),
metrics=["accuracy"])
# + [markdown] id="0wzuMn0yaO1N"
# ### Create tribid embedding datasets and fit tribid model
#
# Again, to keep our experiments swift, let's fit on 20,000 examples for 3 epochs.
#
# This time our model requires four feature inputs:
#
# 1. Train line numbers one-hot tensor (`train_line_numbers_one_hot`)
# 2. Train total lines one-hot tensor (`train_total_lines_one_hot`)
# 3. Token-level sequences tensor (`train_sentences`)
# 4. Char-level sequences tensor (`train_chars`)
#
# We can pass these as tuples to our `tf.data.Dataset.from_tensor_slices()` method to create appropriately shaped and batched `PrefetchedDataset`'s.
# + colab={"base_uri": "https://localhost:8080/"} id="lDXhCx8faoMq" outputId="12867070-442c-4d03-cd9e-b33405dfeade"
# Create training and validation datasets (all four kinds of inputs)
train_pos_char_token_data = tf.data.Dataset.from_tensor_slices((train_line_numbers_one_hot, # line numbers
train_total_lines_one_hot, # total lines
train_sentences, # train tokens
train_chars)) # train chars
train_pos_char_token_labels = tf.data.Dataset.from_tensor_slices(train_labels_one_hot) # train labels
train_pos_char_token_dataset = tf.data.Dataset.zip((train_pos_char_token_data, train_pos_char_token_labels)) # combine data and labels
train_pos_char_token_dataset = train_pos_char_token_dataset.batch(32).prefetch(tf.data.AUTOTUNE) # turn into batches and prefetch appropriately
# Validation dataset
val_pos_char_token_data = tf.data.Dataset.from_tensor_slices((val_line_numbers_one_hot,
val_total_lines_one_hot,
val_sentences,
val_chars))
val_pos_char_token_labels = tf.data.Dataset.from_tensor_slices(val_labels_one_hot)
val_pos_char_token_dataset = tf.data.Dataset.zip((val_pos_char_token_data, val_pos_char_token_labels))
val_pos_char_token_dataset = val_pos_char_token_dataset.batch(32).prefetch(tf.data.AUTOTUNE) # turn into batches and prefetch appropriately
# Check input shapes
train_pos_char_token_dataset, val_pos_char_token_dataset
# + colab={"base_uri": "https://localhost:8080/"} id="lZtnZxolaqoy" outputId="b88dd746-9cbb-46fd-dbb2-bd5bf0cc27b4"
# Fit the token, char and positional embedding model
history_model_5 = model_5.fit(train_pos_char_token_dataset,
steps_per_epoch=int(0.1 * len(train_pos_char_token_dataset)),
epochs=3,
validation_data=val_pos_char_token_dataset,
validation_steps=int(0.1 * len(val_pos_char_token_dataset)))
# + colab={"base_uri": "https://localhost:8080/"} id="vnNdrZNOatD5" outputId="439c6743-3d18-4a5b-88b5-a227cc8182c9"
# Make predictions with token-char-positional hybrid model
model_5_pred_probs = model_5.predict(val_pos_char_token_dataset, verbose=1)
model_5_pred_probs
# + colab={"base_uri": "https://localhost:8080/"} id="hIH-bJ5kavQV" outputId="78c2cf7b-c0bc-4f4b-f6b9-2c964f608b31"
# Turn prediction probabilities into prediction classes
model_5_preds = tf.argmax(model_5_pred_probs, axis=1)
model_5_preds
# + colab={"base_uri": "https://localhost:8080/"} id="6xUvADnXaxFk" outputId="6de8fec5-4f71-409a-c73c-8667bfd19852"
# Calculate results of token-char-positional hybrid model
model_5_results = calculate_results(y_true=val_labels_encoded,
y_pred=model_5_preds)
model_5_results
# + [markdown] id="Buf0fTkfazeX"
# ## Compare model results
# + colab={"base_uri": "https://localhost:8080/", "height": 235} id="lv8zu_1DdLtp" outputId="f47ae241-7f30-484a-e241-0de18a6939e0"
# Combine model results into a DataFrame
all_model_results = pd.DataFrame({"baseline": baseline_results,
"custom_token_embed_conv1d": model_1_results,
"pretrained_token_embed": model_2_results,
"custom_char_embed_conv1d": model_3_results,
"hybrid_char_token_embed": model_4_results,
"tribrid_pos_char_token_embed": model_5_results})
all_model_results = all_model_results.transpose()
all_model_results
# + colab={"base_uri": "https://localhost:8080/", "height": 571} id="3Dpy8mGodNmf" outputId="0c668440-f445-4ac5-fc57-634878cc5967"
# Reduce the accuracy to same scale as other metrics
all_model_results["accuracy"] = all_model_results["accuracy"]/100
# Plot and compare all of the model results
all_model_results.plot(kind="bar", figsize=(10, 7)).legend(bbox_to_anchor=(1.0, 1.0));
# + colab={"base_uri": "https://localhost:8080/", "height": 571} id="xgVN3aaVdrkR" outputId="b2e6fca8-8570-4deb-d4f0-9e3866a7ef62"
# Sort model results by f1-score
all_model_results.sort_values("f1", ascending=False)["f1"].plot(kind="bar", figsize=(10, 7));
# + [markdown] id="Z4jCp28Gdto7"
# ### Save and load best performing model
# + id="ZOJrm1UDd0Ry"
# Save best performing model to SavedModel format (default)
model_5.save("skimlit_tribrid_model") # model will be saved to path specified by string
# + colab={"base_uri": "https://localhost:8080/"} id="Ef4xwQ-7d2No" outputId="d0467a3d-161c-46f4-f7ba-85ad4df6a1df"
# Download pretrained model from Google Storage
# !wget https://storage.googleapis.com/ztm_tf_course/skimlit/skimlit_tribrid_model.zip
# !mkdir skimlit_gs_model
# !unzip skimlit_tribrid_model.zip -d skimlit_gs_model
# + colab={"base_uri": "https://localhost:8080/"} id="3cQTdRCJd6Bz" outputId="ac70d2b1-e6cb-4d85-e513-e01f6a7e40a3"
import tensorflow_hub as hub
import tensorflow as tf
from tensorflow.keras.layers.experimental.preprocessing import TextVectorization
model_path = "skimlit_gs_model/skimlit_tribrid_model"
# Load downloaded model from Google Storage
loaded_model = tf.keras.models.load_model(model_path,
custom_objects={"TextVectorization": TextVectorization, # required for char vectorization
"KerasLayer": hub.KerasLayer}) # required for token embedding
# + [markdown] id="fnJE4shbd8OT"
# ### Make predictions and evaluate them against the truth labels
# + colab={"base_uri": "https://localhost:8080/"} id="a0mNuHS-eInK" outputId="640b3b61-2474-4fe2-f87a-d345273a49b4"
# Make predictions with the loaded model on the validation set
loaded_pred_probs = loaded_model.predict(val_pos_char_token_dataset, verbose=1)
loaded_preds = tf.argmax(loaded_pred_probs, axis=1)
loaded_preds[:10]
# + colab={"base_uri": "https://localhost:8080/"} id="xxpFEqHjeKjU" outputId="9fd213cd-be35-4c59-f724-8ca293360388"
# Evaluate loaded model's predictions
loaded_model_results = calculate_results(val_labels_encoded,
loaded_preds)
loaded_model_results
# + colab={"base_uri": "https://localhost:8080/"} id="zzKyDjWYeOzJ" outputId="c15aedb5-0195-4679-c524-577a79b6371f"
# Check loaded model summary (note the number of trainable parameters)
loaded_model.summary()
# + [markdown] id="tOrPrZhueQ6j"
# ## Evaluate model on test dataset
# + colab={"base_uri": "https://localhost:8080/"} id="eqIXA2wWehvj" outputId="84866414-1162-46e0-e6fe-4cbd7442f247"
# Create test dataset batch and prefetched
test_pos_char_token_data = tf.data.Dataset.from_tensor_slices((test_line_numbers_one_hot,
test_total_lines_one_hot,
test_sentences,
test_chars))
test_pos_char_token_labels = tf.data.Dataset.from_tensor_slices(test_labels_one_hot)
test_pos_char_token_dataset = tf.data.Dataset.zip((test_pos_char_token_data, test_pos_char_token_labels))
test_pos_char_token_dataset = test_pos_char_token_dataset.batch(32).prefetch(tf.data.AUTOTUNE)
# Check shapes
test_pos_char_token_dataset
# + colab={"base_uri": "https://localhost:8080/"} id="mvWqEdnhekLl" outputId="22afaefe-2c5d-4537-d48b-9379b7898ecf"
# Make predictions on the test dataset
test_pred_probs = loaded_model.predict(test_pos_char_token_dataset,
verbose=1)
test_preds = tf.argmax(test_pred_probs, axis=1)
test_preds[:10]
# + colab={"base_uri": "https://localhost:8080/"} id="_jUX9PI8el2H" outputId="74135155-6f74-4d91-a887-7067e77c8255"
# Evaluate loaded model test predictions
loaded_model_test_results = calculate_results(y_true=test_labels_encoded,
y_pred=test_preds)
loaded_model_test_results
# + [markdown] id="fC9VMwXWenez"
# ### Find most wrong
#
# One of the best ways to investigate where your model is going wrong (or potentially where your data is wrong) is to visualize the "most wrong" predictions.
#
# The most wrong predictions are samples where the model has made a prediction with a high probability but has gotten it wrong (the model's prediction disagreess with the ground truth label).
#
# Looking at the most wrong predictions can give us valuable information on how to improve further models or fix the labels in our data.
#
# Let's write some code to help us visualize the most wrong predictions from the test dataset.
#
# First we'll convert all of our integer-based test predictions into their string-based class names.
# + colab={"base_uri": "https://localhost:8080/"} id="WZvrKHwIerNZ" outputId="3aaa8440-2bc4-4262-c10b-02d75a86a2e9"
# %%time
# Get list of class names of test predictions
test_pred_classes = [label_encoder.classes_[pred] for pred in test_preds]
test_pred_classes
# + [markdown] id="y_M_9TVyetNf"
# Now we'll enrich our test DataFame with a few values:
#
# * A "`prediction`" (string) column containing our model's prediction for a given sample.
# * A "`pred_prob`" (float) column containing the model's maximum prediction probabiliy for a given sample.
# * A "`correct`" (bool) column to indicate whether or not the model's prediction matches the sample's target label.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="libqHrwpe4AB" outputId="8b6941f2-9228-43c1-bf3e-8fde00316b26"
# Create prediction-enriched test dataframe
test_df["prediction"] = test_pred_classes # create column with test prediction class names
test_df["pred_prob"] = tf.reduce_max(test_pred_probs, axis=1).numpy() # get the maximum prediction probability
test_df["correct"] = test_df["prediction"] == test_df["target"] # create binary column for whether the prediction is right or not
test_df.head(20)
# + colab={"base_uri": "https://localhost:8080/", "height": 589} id="ChgZH2-6e51r" outputId="5f0352b2-c695-46b8-bd43-c5192884ad19"
# Find top 100 most wrong samples (note: 100 is an abitrary number, you could go through all of them if you wanted)
top_100_wrong = test_df[test_df["correct"] == False].sort_values("pred_prob", ascending=False)[:100]
top_100_wrong
# + colab={"base_uri": "https://localhost:8080/"} id="xKUfDdr-e8Cj" outputId="5cf8c04e-9dce-46fb-92da-ddedb43d5543"
# Investigate top wrong preds
for row in top_100_wrong[0:10].itertuples(): # adjust indexes to view different samples
_, target, text, line_number, total_lines, prediction, pred_prob, _ = row
print(f"Target: {target}, Pred: {prediction}, Prob: {pred_prob}, Line number: {line_number}, Total lines: {total_lines}\n")
print(f"Text:\n{text}\n")
print("-----\n")
# + [markdown] id="tHLDP3xde_H6"
# # Make example predictions
#
# Okay, we've made some predictions on the test dataset, now's time to really test our model out.
#
# To do so, we're going to get some data from the wild and see how our model performs.
#
# In other words, were going to find an RCT abstract from PubMed, preprocess the text so it works with our model, then pass each sequence in the wild abstract through our model to see what label it predicts.
#
# For an appropriate sample, we'll need to search PubMed for RCT's (randomized controlled trials) without abstracts which have been split up (on exploring PubMed you'll notice many of the abstracts are already preformatted into separate sections, this helps dramatically with readability).
#
# Looking at the large chunk of text can seem quite intimidating. Now imagine you're a medical researcher trying to skim through the literature to find a study relevant to your work.
#
# Sounds like quite the challenge right?
#
# Enter SkimLit 🤓🔥!
#
# Let's see what our best model so far (`model_5`) makes of the above abstract.
#
# But wait...
#
# As you might've guessed the above abstract hasn't been formatted in the same structure as the data our model has been trained on. Therefore, before we can make a prediction on it, we need to preprocess it just as we have our other sequences.
#
# More specifically, for each abstract, we'll need to:
#
# 1. Split it into sentences (lines).
# 2. Split it into characters.
# 3. Find the number of each line.
# 4. Find the total number of lines.
#
# Starting with number 1, there are a couple of ways to split our abstracts into actual sentences. A simple one would be to use Python's in-built split() string method, splitting the abstract wherever a fullstop appears. However, can you imagine where this might go wrong?
#
# Another more advanced option would be to leverage spaCy's (a very powerful NLP library) sentencizer class. Which is an easy to use sentence splitter based on spaCy's English language model.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="MYmLVGClfFe4" outputId="01b98597-0d22-438a-e2fc-b9e380c9ec59"
import json
# Download and open example abstracts (copy and pasted from PubMed)
# !wget https://raw.githubusercontent.com/mrdbourke/tensorflow-deep-learning/main/extras/skimlit_example_abstracts.json
with open("skimlit_example_abstracts.json", "r") as f:
example_abstracts = json.load(f)
example_abstracts
# + colab={"base_uri": "https://localhost:8080/", "height": 241} id="xiN8yduPfLe2" outputId="536630cf-d89c-4140-b105-84d1da6e7804"
# See what our example abstracts look like
abstracts = pd.DataFrame(example_abstracts)
abstracts
# + colab={"base_uri": "https://localhost:8080/"} id="G277VTmZfiFG" outputId="de1738a4-d856-4068-f384-476240fc27bc"
# Create sentencizer - Source: https://spacy.io/usage/linguistic-features#sbd
from spacy.lang.en import English
nlp = English() # setup English sentence parser
sentencizer = nlp.create_pipe("sentencizer") # create sentence splitting pipeline object
nlp.add_pipe(sentencizer) # add sentence splitting pipeline object to sentence parser
doc = nlp(example_abstracts[0]["abstract"]) # create "doc" of parsed sequences, change index for a different abstract
abstract_lines = [str(sent) for sent in list(doc.sents)] # return detected sentences from doc in string type (not spaCy token type)
abstract_lines
# + [markdown] id="mFwOQJKHfkf8"
# Now our abstract has been split into sentences, how about we write some code to count line numbers as well as total lines.
#
# To do so, we can leverage some of the functionality of our preprocess_text_with_line_numbers() function.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="WSARr-Zhfrqs" outputId="3bc0df73-66dc-4132-9ff6-ea5af57cecbf"
# Get total number of lines
total_lines_in_sample = len(abstract_lines)
# Go through each line in abstract and create a list of dictionaries containing features for each line
sample_lines = []
for i, line in enumerate(abstract_lines):
sample_dict = {}
sample_dict["text"] = str(line)
sample_dict["line_number"] = i
sample_dict["total_lines"] = total_lines_in_sample - 1
sample_lines.append(sample_dict)
sample_lines
# + colab={"base_uri": "https://localhost:8080/"} id="n4lh68-lft89" outputId="01ed5073-0581-47bd-dc26-b403bca6f6e3"
# Get all line_number values from sample abstract
test_abstract_line_numbers = [line["line_number"] for line in sample_lines]
# One-hot encode to same depth as training data, so model accepts right input shape
test_abstract_line_numbers_one_hot = tf.one_hot(test_abstract_line_numbers, depth=15)
test_abstract_line_numbers_one_hot
# + colab={"base_uri": "https://localhost:8080/"} id="58wtRUj8fwCZ" outputId="f88e4f04-2cd4-4fd9-ac71-3b5e0d79222f"
# Get all total_lines values from sample abstract
test_abstract_total_lines = [line["total_lines"] for line in sample_lines]
# One-hot encode to same depth as training data, so model accepts right input shape
test_abstract_total_lines_one_hot = tf.one_hot(test_abstract_total_lines, depth=20)
test_abstract_total_lines_one_hot
# + colab={"base_uri": "https://localhost:8080/"} id="LqzcyoOkfyVK" outputId="4d957d5b-2640-481a-9f70-1235dee63ce4"
# Split abstract lines into characters
abstract_chars = [split_chars(sentence) for sentence in abstract_lines]
abstract_chars
# + colab={"base_uri": "https://localhost:8080/"} id="h0aAr32yf0mn" outputId="6e32b408-d3e0-4565-fad4-5124c4dbe124"
# Make predictions on sample abstract features
# %%time
test_abstract_pred_probs = loaded_model.predict(x=(test_abstract_line_numbers_one_hot,
test_abstract_total_lines_one_hot,
tf.constant(abstract_lines),
tf.constant(abstract_chars)))
test_abstract_pred_probs
# + colab={"base_uri": "https://localhost:8080/"} id="M8oLi2bof26Y" outputId="a1fc6e43-2575-474f-97cb-ae052d576873"
# Turn prediction probabilities into prediction classes
test_abstract_preds = tf.argmax(test_abstract_pred_probs, axis=1)
test_abstract_preds
# + [markdown] id="PGuhUB0kf5GA"
# Now we've got the predicted sequence label for each line in our sample abstract, let's write some code to visualize each sentence with its predicted label.
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="w07KS7bnf7Dr" outputId="eaee2ee0-098e-4488-deaa-b46931ccbf25"
# Turn prediction class integers into string class names
test_abstract_pred_classes = [label_encoder.classes_[i] for i in test_abstract_preds]
test_abstract_pred_classes
# + colab={"base_uri": "https://localhost:8080/"} id="L1GV3Z7yf81u" outputId="92506289-0578-47b2-cb63-0beb92e61e7d"
# Visualize abstract lines and predicted sequence labels
for i, line in enumerate(abstract_lines):
print(f"{test_abstract_pred_classes[i]}: {line}")
# + [markdown] id="UVG96O4Pf-gt"
# Nice! Isn't that much easier to read? I mean, it looks like our model's predictions could be improved, but how cool is that?
#
# Imagine implementing our model to the backend of the PubMed website to format any unstructured RCT abstract on the site.
#
# Or there could even be a browser extension, called "SkimLit" which would add structure (powered by our model) to any unstructured RCT abtract.
#
# And if showed your medical researcher friend, and they thought the predictions weren't up to standard, there could be a button saying "is this label correct?... if not, what should it be?". That way the dataset, along with our model's future predictions, could be improved over time.
#
# Of course, there are many more ways we could go to improve the model, the usuability, the preprocessing functionality (e.g. functionizing our sample abstract preprocessing pipeline) but I'll leave these for the exercises/extensions.
#
#
| 09_SkimLit_nlp_milestone_project_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from scipy.stats import kurtosis as scipy_kurtosis
# # Load Data
# Load Half-Hourly UK System Cashout Prices.
#
# Data Source: https://www.nordpoolgroup.com/historical-market-data/
file_path = 'https://www.nordpoolgroup.com/48cd48/globalassets/marketdata-excel-files/n2ex-day-ahead-auction-prices_2020_hourly_gbp.xls'
#file_path = '../data/nordpool/n2ex-day-ahead-auction-prices_2018_hourly_gbp.xls'
# Load
raw_list = pd.read_html(file_path, decimal=',')
df = raw_list[0]
# ### Format Data
# +
# Name Columns
df.columns = ['date','hour_UK','hour_EU','price']
# Format Date Column
df.date = pd.to_datetime(df.date, format='%d-%m-%Y')
# Format Hour Columns
df.hour_UK = df.hour_UK.str.slice(stop = 2)
df.hour_UK = pd.to_numeric(df.hour_UK)
df.hour_EU = df.hour_EU.str.slice(stop = 2)
df.hour_EU = pd.to_numeric(df.hour_EU)
# Convert Price to GBP
df.price = df.price/100
# +
# Check NA values
nan_value_count = df.isna().any(axis=1).sum()
na_df = df.loc[df.isna().any(axis=1)]
#df.price.isna().sum()
print(f"NaN values count: {nan_value_count}")
print(f"NaN values table:")
display(na_df)
# -
# Drop NA values
df = df.dropna()
# # Analyse
df.head()
# ### Exceedance Probability Plot
# Log(x) vs. Log(Pr(X>x))
from math import log10
# First let's sort the prices from low to high.
#
# Then calculate steps towards an exceedance probability.
sorted_df = df.sort_values(by='price')
sorted_df = sorted_df.reset_index()
sorted_df = sorted_df.rename(columns={'index':'old_index'})
sorted_df['ones'] = 1
sorted_df['cumcount'] = sorted_df.ones.cumsum()
sorted_df['exceedance_count'] = len(sorted_df)-sorted_df.cumcount+1
# Exceedance probability
sorted_df['exceed_prob'] = sorted_df.exceedance_count/len(sorted_df)
# +
#sorted_df['log_price']=sorted_df.price.apply(log10)
# -
sorted_df['log_exceed_prob']=sorted_df.exceed_prob.apply(log10)
sorted_df.head() # Lowest Prices
sorted_df.tail() # Highest Prices
# I'm not sure how it handles tiny values and log(0)
# +
#Maybe Remove tiny values because they blow up when taking log
#sorted_df.price+=-1*sorted_df.price.min()
# -
sorted_df.set_index('price').exceed_prob.plot(loglog=True, marker='*')
#sorted_df.plot.scatter('log_price', 'log_exceed_prob')
# Alternative code, but this runs more slowly:
# ```
# fig, ax = plt.subplots(constrained_layout=True)
# x = np.array([sorted_df.price])
# y = np.array([sorted_df.exceed_prob])
#
# ax.loglog(x, y, '*')
# ax.set_xlabel('x')
# ax.set_ylabel('Probability(price>x)')
# ax.set_title('Title')
# plt.show()
# ```
sorted_df.plot.scatter('price', 'exceed_prob')
# Conclusions:
# * Looks like a power law between ~40 and 70 GBP
# * Tail behaves funny it gets fatter then thinner in this particular dataset.
# # Appendix:
# ### Generic Loglog Plot
# First show one that's not loglog:
fig, ax = plt.subplots(constrained_layout=True)
x = np.array([1,2,3,4,5, 6,7,8,9,10])
y = np.array([1,10,100,1_000,100_000, 1_000_000, 10_000_000, 100_000_000, 1_000_000_000, 10_000_000_000])
ax.plot(x, y, '*')
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_title('Title')
plt.show()
# Now this one IS loglog:
fig, ax = plt.subplots(constrained_layout=True)
x = np.array([1,2,3,4,5, 6,7,8,9,10])
y = np.array([1,10,100,1_000,100_000, 1_000_000, 10_000_000, 100_000_000, 1_000_000_000, 10_000_000_000])
ax.loglog(x, y, '*')
ax.set_xlabel('x')
ax.set_ylabel('y')
ax.set_title('Title')
plt.show()
| notebooks/Notebook-13 - LogLog Exceedance Probability.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Prob2b.py
def gcd(a, b):
""" finds gcd/lcm of two numbers """
if b == 0:
return a
else:
return gcd(b, a%b)
def gcd3(a,b,c):
""" finds gcd of 3 numbers """
x = gcd(a,b)
g = gcd(x, gcd(b,c))
alpha = a % g
beta = b % g
gamma = c % g
return (g, alpha, beta, gamma)
gcd3(6930, 13230, 15760)
| ds/Prob2b.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Observational Write-Up (Results)
# Four mice tumor treatment groups were analyzed in this study (Capomulin, Infubinol, Ketapril, and a placebo). After analyzing the data, Capomulin's treatment group obtained the best results in reducing tumor volume by 19% (45 mm3 to 36.23 mm3), having the least amount of metastatic sites (1.47), and incurring the highest survival rate (84%). Moreover, the other three groups had negative results in regards to tumor volume (increasing between 46% and 57%), metastatic sites (2.1 to 3.3), and survival rate (36% to 66%). From this analysis, Capomulin appears to be the most promising drug candidate for this kind of tumor.
# +
# Dependencies and Setup
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
# Hide warning messages in notebook
import warnings
warnings.filterwarnings('ignore')
# File to Load (Remember to Change These)
mouse_drug_data_to_load = "data/mouse_drug_data.csv"
clinical_trial_data_to_load = "data/clinicaltrial_data.csv"
# Read the Mouse and Drug Data and the Clinical Trial Data
mouse_drug_data = pd.read_csv(mouse_drug_data_to_load)
clinical_trial_data = pd.read_csv(clinical_trial_data_to_load)
# Combine the data into a single dataset
merged_df = pd.merge(mouse_drug_data, clinical_trial_data, on = "Mouse ID")
# Display the data table for preview
merged_df.head()
# -
# ## Tumor Response to Treatment
# +
# Store the Mean Tumor Volume Data Grouped by Drug and Timepoint
mean_tumor_volume = merged_df.groupby(["Drug", "Timepoint"]).mean()["Tumor Volume (mm3)"]
# Convert to DataFrame
mean_tumor_volume_df = pd.DataFrame(mean_tumor_volume).reset_index()
# Preview DataFrame
mean_tumor_volume_df.head()
# +
# Store the Standard Error of Tumor Volumes Grouped by Drug and Timepoint
standard_error_volume = merged_df.groupby(["Drug", "Timepoint"]).sem()["Tumor Volume (mm3)"]
# Convert to DataFrame
standard_error_volume_df = pd.DataFrame(standard_error_volume).reset_index()
# Preview DataFrame
standard_error_volume_df.head()
# +
# Minor Data Munging to Re-Format the Data Frames
reformated_mean_df = mean_tumor_volume_df.pivot_table(index = "Timepoint", columns = "Drug", values = "Tumor Volume (mm3)")
reformated_error_df = standard_error_volume_df.pivot_table(index = "Timepoint", columns = "Drug", values = "Tumor Volume (mm3)")
# Preview that Reformatting worked
reformated_error_df.head()
reformated_mean_df.head(10)
# +
# Generate the Plot (with Error Bars)
plt.errorbar(reformated_mean_df.index, reformated_mean_df["Capomulin"], yerr = reformated_error_df["Capomulin"], color = 'r', marker = "o", label = "Capomulin")
plt.errorbar(reformated_mean_df.index, reformated_mean_df["Infubinol"], yerr = reformated_error_df["Infubinol"], color = 'b', marker = "^", label = "Infubinol")
plt.errorbar(reformated_mean_df.index, reformated_mean_df["Ketapril"], yerr = reformated_error_df["Ketapril"], color = 'g', marker = 's', label = "Ketapril")
plt.errorbar(reformated_mean_df.index, reformated_mean_df["Placebo"], yerr = reformated_error_df["Placebo"], color = 'black', marker = 'd', label= "Placebo")
# Set x-label
plt.xlabel("Time (Days)")
# Set y-label
plt.ylabel("Tumor Volume (mm3)")
# Set title
plt.title("Tumor Response to Treatment")
# Set legend
plt.legend(loc = "upper left")
# Set grid on the y-axis
plt.grid(axis = 'y')
# Save the Figure
plt.savefig("Graphs/Tumor Response to Treatment.png")
# Show Figure
plt.show()
# -
# ## Metastatic Response to Treatment
# +
# Store the Mean Met. Site Data Grouped by Drug and Timepoint
mean_metastatic_data = merged_df.groupby(["Drug", "Timepoint"]).mean()["Metastatic Sites"]
# Convert to DataFrame
mean_metastatic_df = pd.DataFrame(mean_metastatic_data)
# Preview DataFrame
mean_metastatic_df.head()
# +
# Store the Standard Error associated with Met. Sites Grouped by Drug and Timepoint
std_err_metastatic_data = merged_df.groupby(["Drug", "Timepoint"]).sem()["Metastatic Sites"]
# Convert to DataFrame
std_err_metastatic_df = pd.DataFrame(std_err_metastatic_data)
# Preview DataFrame
std_err_metastatic_df.head()
# +
# Minor Data Munging to Re-Format the Data Frames
reformated_mean_metastatic_df = mean_metastatic_df.pivot_table(index = "Timepoint", columns = "Drug", values = "Metastatic Sites")
reformated_err_metastatic_df = std_err_metastatic_df.pivot_table(index = "Timepoint", columns = "Drug", values = "Metastatic Sites")
# Preview that Reformatting worked
reformated_err_metastatic_df.head()
reformated_mean_metastatic_df.head(10)
# +
# Generate the Plot (with Error Bars)
plt.errorbar(reformated_mean_metastatic_df.index, reformated_mean_metastatic_df["Capomulin"], yerr = reformated_err_metastatic_df["Capomulin"], label = "Capomulin", color = 'r', marker = 'o')
plt.errorbar(reformated_mean_metastatic_df.index, reformated_mean_metastatic_df["Infubinol"], yerr = reformated_err_metastatic_df["Infubinol"], label = "Infubinol", color = 'b', marker = '^')
plt.errorbar(reformated_mean_metastatic_df.index, reformated_mean_metastatic_df["Ketapril"], yerr = reformated_err_metastatic_df["Ketapril"], label = "Ketapril", color = 'g', marker = 's')
plt.errorbar(reformated_mean_metastatic_df.index, reformated_mean_metastatic_df["Placebo"], yerr = reformated_err_metastatic_df["Placebo"], label = "Placebo", color = 'black', marker = 'd')
# Set x-label
plt.xlabel("Treatment Duration (Days)")
# Set y-label
plt.ylabel("Met. Sites")
# Set title
plt.title("Metastatic Spread During Treatment")
# Set legend
plt.legend(loc = "upper left")
# Set grid on y-axis
plt.grid(axis = 'y')
# Save the Figure
plt.savefig("Graphs/Metastatic Spread During Treatment.png")
# Show the Figure
plt.show()
# -
# ## Survival Rates
# +
# Store the Count of Mice Grouped by Drug and Timepoint (We can pass any metric)
mouse_count_data = merged_df.groupby(["Drug", "Timepoint"]).count()["Mouse ID"]
# Convert to DataFrame
mouse_count_df = pd.DataFrame(mouse_count_data).reset_index()
mouse_count_df = mouse_count_df.rename(columns = {"Mouse ID":"Mouse Count"})
# Preview DataFrame
mouse_count_df.head()
# +
# Minor Data Munging to Re-Format the Data Frames
reformated_mouse_count_df = mouse_count_df.pivot_table(index="Timepoint", columns = "Drug", values = "Mouse Count")
# Preview the Data Frame
reformated_mouse_count_df.head(10)
# +
# Generate the Plot (Accounting for percentages)
plt.errorbar(reformated_mouse_count_df.index, round(reformated_mouse_count_df['Capomulin']/reformated_mouse_count_df['Capomulin'][0] * 100, 2), marker = 'o', color = 'r', label = 'Capomunil')
plt.errorbar(reformated_mouse_count_df.index, round(reformated_mouse_count_df['Infubinol']/reformated_mouse_count_df['Infubinol'][0] * 100, 2), marker = '^', color = 'b', label = 'Infubinol')
plt.errorbar(reformated_mouse_count_df.index, round(reformated_mouse_count_df['Ketapril']/reformated_mouse_count_df['Ketapril'][0] * 100, 2), marker = 's', color = 'g', label = 'Ketapril')
plt.errorbar(reformated_mouse_count_df.index, round(reformated_mouse_count_df['Placebo']/reformated_mouse_count_df['Placebo'][0] * 100, 2), marker = 'd', color = 'k', label = 'Placebo')
# Set x-label
plt.xlabel('Time (Days)')
# Set y-label
plt.ylabel('Survival Rate (%)')
# Set title
plt.title('Survival During Treatment')
# Set legend
plt.legend(loc = 'lower left')
# Set grid
plt.grid()
# Save the Figure
plt.savefig('Graphs/Survival During Treatment.png')
# Show the Figure
plt.show()
# -
# ## Summary Bar Graph
# +
# Calculate the percent changes for each drug
drug_effectiveness = (reformated_mean_df.iloc[-1] - reformated_mean_df.iloc[0])/reformated_mean_df.iloc[0] * 100
# Display the data to confirm
drug_effectiveness
# +
# Store all Relevant Percent Changes into a Tuple
relevant_drugs = drug_effectiveness.loc[['Capomulin', 'Infubinol', 'Ketapril', 'Placebo']]
relevant_drugs
# Splice the data between passing and failing drugs
bar_color = []
for value in relevant_drugs:
if value > 0:
bar_color.append('r')
else:
bar_color.append('g')
# Orient widths. Add labels, tick marks, etc.
x_axis = np.arange(0, len(relevant_drugs.index))
tick_locations = []
for x in x_axis:
tick_locations.append(x)
plt.xticks(tick_locations, relevant_drugs.index)
graph = plt.bar(x_axis, relevant_drugs.values, color = bar_color, width = 1)
plt.xlabel('Drugs')
plt.ylabel('% Tumor Volume Change')
plt.title('Tumor Change Over 45 Day Treatment')
plt.grid()
plt.ylim(-30, max(relevant_drugs.values) + 20)
plt.xlim(-0.75, len(relevant_drugs.index) - 0.25)
# Use functions to label the percentages of changes
for i in graph:
yvalue = i.get_height()
plt.text(i.get_x() + .35, yvalue - (.5 * yvalue), "{:.0%}".format(yvalue/100))
# Save the Figure
plt.savefig('Graphs/Tumor Change Results')
# Show the Figure
plt.show()
# -
| Pymaceuticals/pymaceuticals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Load pickled data
import pickle
import numpy as np
# TODO: Fill this in based on where you saved the training and testing data
training_file = "traffic-signs-data/train.p"
validation_file= "traffic-signs-data/valid.p"
testing_file = "traffic-signs-data/test.p"
with open(training_file, mode='rb') as f:
train = pickle.load(f)
with open(validation_file, mode='rb') as f:
valid = pickle.load(f)
with open(testing_file, mode='rb') as f:
test = pickle.load(f)
X_train, y_train = train['features'], train['labels']
X_valid, y_valid = valid['features'], valid['labels']
X_test, y_test = test['features'], test['labels']
# -
# ### Step 1: Dataset Summary & Exploration¶
#
# The pickled data is a dictionary with 4 key/value pairs:
#
# > - 'features' is a 4D array containing raw pixel data of the traffic sign images, (num examples, width, height, channels).
# > - 'labels' is a 1D array containing the label/class id of the traffic sign. The file signnames.csv contains id -> name mappings for each id.
# > - 'sizes' is a list containing tuples, (width, height) representing the original width and height the image.
# > - 'coords' is a list containing tuples, (x1, y1, x2, y2) representing coordinates of a bounding box around the sign in the image. THESE COORDINATES ASSUME THE ORIGINAL IMAGE. THE PICKLED DATA CONTAINS RESIZED VERSIONS (32 by 32) OF THESE IMAGES
#
# ### Provide a Basic Summary of the Data Set Using Python, Numpy and/or Pandas
# +
### Use python, pandas or numpy methods rather than hard coding the results
# Number of training examples
n_train = len(X_train)
# Number of validation examples
n_validation = len(X_valid)
# Number of testing examples.
n_test = len(X_test)
# What's the shape of an traffic sign image?
image_shape = X_train[0].shape
# How many unique classes/labels there are in the dataset.
n_classes = len(set(y_train))
print("Number of training images =", n_train)
print("Number of validation images =", n_validation)
print("Number of testing images =", n_test)
print("Image data shape =", image_shape)
print("Number of classes =", n_classes)
# -
# ### Include an exploratory visualization of the dataset
#
# Visualize the German Traffic Signs Dataset using the pickled file(s). This is open ended, suggestions include: plotting traffic sign images, plotting the count of each sign, etc.
# +
### Data exploration visualization code
import matplotlib.pyplot as plt
# %matplotlib inline
def show_images(X, end, total, images_per_row = 30, images_per_col = 15,
H = 20, W = 1, its_gray = False):
number_of_images = images_per_row * images_per_col
figure, axis = plt.subplots(images_per_col, images_per_row, figsize=(H, W))
figure.subplots_adjust(hspace = .2, wspace=.001)
axis = axis.ravel()
for i in range(number_of_images):
index = np.random.randint(end - total, end - 1)
image = X[index]
axis[i].axis('off')
if its_gray:
axis[i].imshow(image.reshape(32, 32), cmap='gray')
else:
axis[i].imshow(image)
show_images(X_train, len(X_train), len(X_train),
images_per_row = 20, images_per_col = 5,
H = 32, W = 10)
# -
# ### Data Set Summary
#
# +
# Count frequency of each label
labels, counts = np.unique(y_train, return_counts=True)
# Plot the histogram
plt.rcParams["figure.figsize"] = [15, 5]
axes = plt.gca()
axes.set_xlim([-1,43])
plt.bar(labels, counts, tick_label=labels, width=0.8, align='center')
plt.title('PLOT: Class Distribution across Training set Data')
plt.show()
# -
#
# ### Step 2: Design and Test a Model Architecture
#
# Design and implement a deep learning model that learns to recognize traffic signs. Train and test your model on the [German Traffic Sign Dataset](http://benchmark.ini.rub.de/?section=gtsrb&subsection=dataset).
#
# The LeNet-5 implementation shown in the classroom at the end of the CNN lesson is a solid starting point. You'll have to change the number of classes and possibly the preprocessing, but aside from that it's plug and play!
#
# With the LeNet-5 solution from the lecture, you should expect a validation set accuracy of about 0.89. To meet specifications, the validation set accuracy will need to be at least 0.93. It is possible to get an even higher accuracy, but 0.93 is the minimum for a successful project submission.
#
# There are various aspects to consider when thinking about this problem:
#
# * Neural network architecture (is the network over or underfitting?)
# * Play around preprocessing techniques (normalization, rgb to grayscale, etc)
# * Number of examples per label (some have more than others).
# * Generate fake data.
#
#
#
# ### Pre-process the Data Set (normalization, grayscale, etc.)
#
# Minimally, the image data should be normalized so that the data has mean zero and equal variance. For image data, (pixel - 128)/ 128 is a quick way to approximately normalize the data and can be used in this project.
#
# Other pre-processing steps are optional. You can try different techniques to see if it improves performance.
#
# Use the code cell (or multiple code cells, if necessary) to implement the first step of your project.
#
#
#
# After plotting some images from the dataset it is observed that the certain amount of images are either too dark or too bright. Because of this an open cv based auto contrast correction technique is applied as preprocessing. As required by the network, all images are resized to 32x32 size. This step is also considered as part of preprocessing.
#
# My preprocessing pipeline is as follows:
#
# 1. Resize the image to 32x32
# 2. Apply auto contrast adjustment
#
#
# +
### Preprocess the data here. It is required to normalize the data. Other preprocessing steps could include
### converting to grayscale, etc.
### Feel free to use as many code cells as needed.
import cv2
#print("Using OpenCV version: %s" % (cv2.__version__))
#auto correction technique
def auto_contrast(img):
# CLAHE (Contrast Limited Adaptive Histogram Equalization)
clahe = cv2.createCLAHE(clipLimit=3., tileGridSize=(8, 8))
lab = cv2.cvtColor(img, cv2.COLOR_RGB2LAB) # convert from BGR to LAB color space
l, a, b = cv2.split(lab) # split on 3 different channels
l2 = clahe.apply(l) # apply CLAHE to the L-channel
lab = cv2.merge((l2, a, b)) # merge channels
img2 = cv2.cvtColor(lab, cv2.COLOR_LAB2RGB) # convert from LAB to RGB
return img2
def resize(image):
return cv2.resize(image, (32, 32))
def image_preprocess(image):
image = resize(image)
image = auto_contrast(image)
return image
# -
# Pre-processing on (training, test, validation sets)
X_train = np.array([image_preprocess(x) for x in X_train])
X_valid = np.array([image_preprocess(x) for x in X_valid])
X_test = np.array([image_preprocess(x) for x in X_test])
show_images(X_train, len(X_train), len(X_train),
images_per_row = 20, images_per_col = 5,
H = 32, W = 10, its_gray=False)
# ### Model Architecture
#
def preprocess_op(x):
p0 = tf.image.convert_image_dtype(x, tf.float32)
p0 = tf.divide(tf.subtract(p0, 127.5), 127.5)
return p0
# ### final model consist of the following layers:
# Layer Input Output
# Input 32x32x3
# Convolution 32x32x3 28x28x6
# Relu
# Dropout
# Max Pooling 28x28x6 14x14x6
# Convolution 14x14x6 10x10x16
# Relu
# Dropout
# Max Pooling 10x10x16 5x5x16
# Convolution 5x5x16 120(flatten)
# Relu
# Dropout
# Fully Connected 120 84
# Relu
# Dropout
# Fully Connected 84 43
#
#
#
# +
### Define your architecture here.
### Feel free to use as many code cells as needed.
import tensorflow as tf
#print("Using TensorFlow version: %s" % (tf.__version__))
def LeNet(x, keep_prob=1.0):
mu = 0
sigma = 0.1
print("LeNet!")
p0 = preprocess_op(x)
# C1: Input 32x32x3, Output: 28x28x6
weight1 = tf.Variable(tf.truncated_normal(shape=(5, 5, 3, 6), mean=mu, stddev=sigma))
bias1 = tf.Variable(tf.zeros(shape=(6)))
conv1 = tf.nn.conv2d(p0, weight1, strides=(1, 1, 1, 1), padding='VALID')
conv1 = tf.add(conv1, bias1)
conv1 = tf.nn.relu(conv1)
conv1 = tf.nn.dropout(conv1, keep_prob=keep_prob)
print("C1: Input %s Output %s" % (x.get_shape(), conv1.get_shape()))
# P2 Input 28x28x6, Output: 14x14x6
pool1 = tf.nn.max_pool(conv1, ksize=(1, 2, 2, 1), strides=(1, 2, 2, 1), padding='VALID')
print("P2: Input %s Output %s" % (conv1.get_shape(), pool1.get_shape()))
# C3 Input 14x14x6, Output: 10x10x16
weight2 = tf.Variable(tf.truncated_normal(shape=(5, 5, 6, 16), mean=mu, stddev=sigma))
bias2 = tf.Variable(tf.zeros(shape=(16)))
conv2 = tf.nn.conv2d(pool1, weight2, strides=(1, 1, 1, 1), padding='VALID')
conv2 = tf.add(conv2, bias2)
conv2 = tf.nn.relu(conv2)
conv2 = tf.nn.dropout(conv2, keep_prob=keep_prob)
print("C3: Input %s Output %s" % (pool1.get_shape(), conv2.get_shape()))
# P4 Input 10x10x16, Output 5x5x16
pool2 = tf.nn.max_pool(conv2, ksize=(1, 2, 2, 1), strides=(1, 2, 2, 1), padding='VALID')
print("P4: Input %s Output %s" % (conv2.get_shape(), pool2.get_shape()))
# C5: Input 5x5x16, Output 1x120
fc1 = tf.contrib.layers.flatten(pool2)
weight3 = tf.Variable(tf.truncated_normal(shape=(400, 120), mean=mu, stddev=sigma))
bias3 = tf.Variable(tf.zeros(shape=(120)))
fc1 = tf.matmul(fc1, weight3)
fc1 = tf.add(fc1, bias3)
fc1 = tf.nn.relu(fc1)
fc1 = tf.nn.dropout(fc1, keep_prob=keep_prob)
print("C5: Input %s Output %s" % (pool2.get_shape(), fc1.get_shape()))
# F6 Input 1x120, Output 1x84
weights4 = tf.Variable(tf.truncated_normal(shape=(120, 84), mean=mu, stddev=sigma))
bias4 = tf.Variable(tf.zeros(shape=(84)))
fc2 = tf.matmul(fc1, weights4)
fc2 = tf.add(fc2, bias4)
fc2 = tf.nn.relu(fc2)
fc2 = tf.nn.dropout(fc2, keep_prob=keep_prob)
print("F6: Input %s Output %s" % (fc1.get_shape(), fc2.get_shape()))
# F7 Input 1x84, Output 1x10
weight5 = tf.Variable(tf.truncated_normal(shape=(84, 43), mean=mu, stddev=sigma))
bias5 = tf.Variable(tf.zeros(shape=(43)))
logits = tf.matmul(fc2, weight5)
logits = tf.add(logits, bias5)
print("F7: Input %s Output %s" % (fc2.get_shape(), logits.get_shape()))
return logits
# -
#
# ### Train, Validate and Test the Model
#
# A validation set can be used to assess how well the model is performing. A low accuracy on the training and validation sets imply underfitting. A high accuracy on the training set but low accuracy on the validation set implies overfitting.
#
### Train your model here.
### Calculate and report the accuracy on the training and validation set.
### Once a final model architecture is selected,
### the accuracy on the test set should be calculated and reported as well.
height, width, channels = X_train[0].shape
x = tf.placeholder(tf.float32, shape=(None, height, width, channels))
y = tf.placeholder(tf.int32, shape=(None))
keep_prob = tf.placeholder(tf.float32)
one_hot_y = tf.one_hot(y, 43)
# +
rate = 0.001
logits = LeNet(x, keep_prob)
cross_entropy = tf.nn.softmax_cross_entropy_with_logits_v2(logits=logits, labels=one_hot_y)
loss_operations = tf.reduce_mean(cross_entropy)
optimizer = tf.train.AdamOptimizer(learning_rate = rate)
training_operation = optimizer.minimize(loss_operations)
# +
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1))
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
saver = tf.train.Saver()
def evaluate(X_data, y_data):
num_examples = len(X_data)
total_accuracy = 0
total_loss = 0
sess = tf.get_default_session()
for offset in range(0, num_examples, BATCH_SIZE):
batch_x, batch_y = X_data[offset:offset+BATCH_SIZE], y_data[offset:offset+BATCH_SIZE]
accuracy, loss = sess.run([accuracy_operation, loss_operations], feed_dict={x: batch_x, y: batch_y, keep_prob: 1.0})
total_accuracy += (accuracy * len(batch_x))
total_loss += (loss * len(batch_x))
return total_accuracy / num_examples, total_loss / num_examples
# -
# Following are the parameters are used to train my model:
#
# 1. Learning Rate = 0.001
# 2. Number of Epochs = 30
# 3. Batch Size = 128
# 4. Optimizer = Adam Optimizer
#
# +
### Calculate and report the accuracy on the training and validation set.
from sklearn.utils import shuffle
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
num_examples = len(X_train)
print("Training...")
print()
for i in range(EPOCHS):
X_train, y_train = shuffle(X_train, y_train)
for offset in range(0, num_examples, BATCH_SIZE):
end = offset + BATCH_SIZE
batch_x, batch_y = X_train[offset:end], y_train[offset:end]
sess.run(training_operation, feed_dict={x: batch_x, y: batch_y, keep_prob: 0.5})
training_accuracy, training_loss = evaluate(X_train, y_train)
validation_accuracy, validation_loss = evaluate(X_valid, y_valid)
print("EPOCH {} ...".format(i+1))
print("Training Accuracy = {:.3f} Loss = {:.3f}".format(training_accuracy, training_loss))
print("Validation Accuracy = {:.3f} Loss = {:.3f}".format(validation_accuracy, validation_loss))
print()
saver.save(sess, './lenet')
print("Model saved")
# -
# Training ...
#
# EPOCH 1 ...
# Training Accuracy = 0.557 Loss = 2.024
# Validation Accuracy = 0.514 Loss = 2.086
#
# EPOCH 2 ...
# Training Accuracy = 0.738 Loss = 1.407
# Validation Accuracy = 0.707 Loss = 1.474
#
# EPOCH 3 ...
# Training Accuracy = 0.816 Loss = 1.072
# Validation Accuracy = 0.796 Loss = 1.140
#
# EPOCH 4 ...
# Training Accuracy = 0.869 Loss = 0.866
# Validation Accuracy = 0.829 Loss = 0.943
#
# EPOCH 5 ...
# Training Accuracy = 0.905 Loss = 0.711
# Validation Accuracy = 0.864 Loss = 0.806
#
# EPOCH 6 ...
# Training Accuracy = 0.921 Loss = 0.612
# Validation Accuracy = 0.893 Loss = 0.698
#
# EPOCH 7 ...
# Training Accuracy = 0.925 Loss = 0.531
# Validation Accuracy = 0.904 Loss = 0.608
#
# EPOCH 8 ...
# Training Accuracy = 0.937 Loss = 0.476
# Validation Accuracy = 0.917 Loss = 0.552
#
# EPOCH 9 ...
# Training Accuracy = 0.943 Loss = 0.437
# Validation Accuracy = 0.910 Loss = 0.530
#
# EPOCH 10 ...
# Training Accuracy = 0.947 Loss = 0.407
# Validation Accuracy = 0.919 Loss = 0.498
#
# EPOCH 11 ...
# Training Accuracy = 0.951 Loss = 0.378
# Validation Accuracy = 0.919 Loss = 0.475
#
# EPOCH 12 ...
# Training Accuracy = 0.958 Loss = 0.341
# Validation Accuracy = 0.915 Loss = 0.457
#
# EPOCH 13 ...
# Training Accuracy = 0.963 Loss = 0.311
# Validation Accuracy = 0.925 Loss = 0.411
#
# EPOCH 14 ...
# Training Accuracy = 0.966 Loss = 0.312
# Validation Accuracy = 0.925 Loss = 0.411
#
# EPOCH 15 ...
# Training Accuracy = 0.968 Loss = 0.294
# Validation Accuracy = 0.922 Loss = 0.408
#
# EPOCH 16 ...
# Training Accuracy = 0.967 Loss = 0.259
# Validation Accuracy = 0.934 Loss = 0.355
#
# EPOCH 17 ...
# Training Accuracy = 0.968 Loss = 0.257
# Validation Accuracy = 0.927 Loss = 0.374
#
# EPOCH 18 ...
# Training Accuracy = 0.968 Loss = 0.235
# Validation Accuracy = 0.927 Loss = 0.339
#
# EPOCH 19 ...
# Training Accuracy = 0.972 Loss = 0.237
# Validation Accuracy = 0.935 Loss = 0.343
#
# EPOCH 20 ...
# Training Accuracy = 0.976 Loss = 0.212
# Validation Accuracy = 0.936 Loss = 0.329
#
# EPOCH 21 ...
# Training Accuracy = 0.977 Loss = 0.206
# Validation Accuracy = 0.939 Loss = 0.311
#
# EPOCH 22 ...
# Training Accuracy = 0.977 Loss = 0.207
# Validation Accuracy = 0.936 Loss = 0.323
#
# EPOCH 23 ...
# Training Accuracy = 0.979 Loss = 0.185
# Validation Accuracy = 0.941 Loss = 0.287
#
# EPOCH 24 ...
# Training Accuracy = 0.979 Loss = 0.192
# Validation Accuracy = 0.937 Loss = 0.311
#
# EPOCH 25 ...
# Training Accuracy = 0.978 Loss = 0.171
# Validation Accuracy = 0.941 Loss = 0.277
#
# EPOCH 26 ...
# Training Accuracy = 0.980 Loss = 0.179
# Validation Accuracy = 0.946 Loss = 0.288
#
# EPOCH 27 ...
# Training Accuracy = 0.980 Loss = 0.161
# Validation Accuracy = 0.946 Loss = 0.274
#
# EPOCH 28 ...
# Training Accuracy = 0.981 Loss = 0.159
# Validation Accuracy = 0.945 Loss = 0.262
#
# EPOCH 29 ...
# Training Accuracy = 0.983 Loss = 0.155
# Validation Accuracy = 0.948 Loss = 0.271
#
# EPOCH 30 ...
# Training Accuracy = 0.984 Loss = 0.151
# Validation Accuracy = 0.944 Loss = 0.265
#
# Model saved
### the accuracy on the test set should be calculated and reported as well.
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
test_accuracy, _ = evaluate(X_test, y_test)
print("Test Accuracy = {:.3f}".format(test_accuracy))
# INFO:tensorflow:Restoring parameters from ./lenet
# Test Accuracy = 0.93
print("Training set accuracy = {:.3f}".format(training_accuracy))
print("Validation set accuracy = {:.3f}".format(validation_accuracy))
print("Test set accuracy = {:.3f}".format(test_accuracy))
# Training set accuracy = 0.984
# Validation set accuracy = 0.944
# Test set accuracy = 0.938
#
# ### Step 3: Test a Model on New Images
#
# To give yourself more insight into how your model is working, download at least five pictures of German traffic signs from the web and use your model to predict the traffic sign type.
#
# You may find signnames.csv useful as it contains mappings from the class id (integer) to the actual sign name.
# ### Load and Output the Images
# +
### Load the images and plot them here.
### Feel free to use as many code cells as needed.
import glob
import os
filelist = glob.glob("*.jpg")
test_images = []
test_labels = []
for filename in filelist:
print(filename)
image = cv2.imread(filename)
if image is None:
continue
image = cv2.cvtColor(image, cv2.COLOR_BGR2RGB)
test_images.append(image)
sign_name = os.path.splitext(filename)[0]
test_labels.append(int(sign_name))
print("Number of test images =", len(test_images))
#show_images(test_images, len(test_images), len(test_images),
# images_per_row = len(filelist), images_per_col = 1,
# H = 10, W = 10, its_gray=False)
print("Test Labels: ", test_labels)
#test_images[0].shape
# -
#
# 4.jpg
# 17.jpg
# 14.jpg
# 12.jpg
# 2.jpg
# 1.jpg
# Number of test images = 6
# Test Labels: [4, 17, 14, 12, 2, 1]
# I have selected 6 random German Traffic signs. They are of classes -
#
# 1. speed limit 30,
# 2. speed limit 50,
# 4. speed limit 70,
# 17. No Entry
# 14. Stop
# 12. Priority.
#
#
# Out of this 6 signs, 5 got classified correctly. Misclassified sign is speed limit 30
# ### Predict the Sign Type for Each Image
#
test_images_pre = [image_preprocess(i) for i in test_images]
# +
### Run the predictions here and use the model to output the prediction for each image.
### Make sure to pre-process the images with the same pre-processing pipeline used earlier.
### Feel free to use as many code cells as needed.
import csv
signs_class=[]
with open('signnames.csv', 'rt') as csvfile:
reader = csv.DictReader(csvfile, delimiter=',')
for row in reader:
signs_class.append((row['SignName']))
# -
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
pred = sess.run(logits, feed_dict={x: test_images_pre, keep_prob: 1.0})
output = np.argmax(pred, axis=1)
print("Predicted :", output)
print("Ground Truth:", np.array(test_labels))
# INFO:tensorflow:Restoring parameters from ./lenet
# Predicted : [ 4 17 14 12 2 2]
# Ground Truth: [ 4 17 14 12 2 1]
# ### Analyze Performance
#
# +
### Calculate the accuracy for these 5 new images.
### For example, if the model predicted 1 out of 5 signs correctly, it's 20% accurate on these new images.
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
test_accuracy, _ = evaluate(test_images_pre, test_labels)
print("Test Accuracy = {:.3f}".format(test_accuracy))
# -
# INFO:tensorflow:Restoring parameters from ./lenet
# Test Accuracy = 0.833
#
# ### Output Top 5 Softmax Probabilities For Each Image Found on the Web
#
#
#
# For each of the new images, print out the model's softmax probabilities to show the certainty of the model's predictions (limit the output to the top 5 probabilities for each image). tf.nn.top_k could prove helpful here.
#
# The example below demonstrates how tf.nn.top_k can be used to find the top k predictions for each image.
#
# tf.nn.top_k will return the values and indices (class ids) of the top k predictions. So if k=3, for each sign, it'll return the 3 largest probabilities (out of a possible 43) and the correspoding class ids.
#
# Take this numpy array as an example. The values in the array represent predictions. The array contains softmax probabilities for five candidate images with six possible classes. tf.nn.top_k is used to choose the three classes with the highest probability:
#
# (5, 6) array
# a = np.array([[ 0.24879643, 0.07032244, 0.12641572, 0.34763842, 0.07893497,
# 0.12789202],
# [ 0.28086119, 0.27569815, 0.08594638, 0.0178669 , 0.18063401,
# 0.15899337],
# [ 0.26076848, 0.23664738, 0.08020603, 0.07001922, 0.1134371 ,
# 0.23892179],
# [ 0.11943333, 0.29198961, 0.02605103, 0.26234032, 0.1351348 ,
# 0.16505091],
# [ 0.09561176, 0.34396535, 0.0643941 , 0.16240774, 0.24206137,
# 0.09155967]])
#
# Running it through sess.run(tf.nn.top_k(tf.constant(a), k=3)) produces:
#
# TopKV2(values=array([[ 0.34763842, 0.24879643, 0.12789202],
# [ 0.28086119, 0.27569815, 0.18063401],
# [ 0.26076848, 0.23892179, 0.23664738],
# [ 0.29198961, 0.26234032, 0.16505091],
# [ 0.34396535, 0.24206137, 0.16240774]]), indices=array([[3, 0, 5],
# [0, 1, 4],
# [0, 5, 1],
# [1, 3, 5],
# [1, 4, 3]], dtype=int32))
#
# Looking just at the first row we get [ 0.34763842, 0.24879643, 0.12789202], you can confirm these are the 3 largest probabilities in a. You'll also notice [3, 0, 5] are the corresponding indices.
#
### Print out the top five softmax probabilities for the predictions on the German traffic sign images found on the web.
### Feel free to use as many code cells as needed.
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
signs_top_5 = sess.run(tf.nn.top_k(tf.nn.softmax(logits), k = 5), feed_dict={x: test_images_pre, keep_prob: 1.0})
print("Predicted :", signs_top_5)
# INFO:tensorflow:Restoring parameters from ./lenet
#
#
# Predicted : TopKV2(values=array([[9.11246359e-01, 7.43291378e-02, 9.22355428e-03, 1.22498453e-03,
# 1.21638970e-03],
# [9.99999166e-01, 8.85979489e-07, 8.92348140e-10, 3.81197712e-10,
# 3.67172237e-10],
# [9.67177510e-01, 1.20190699e-02, 6.96608843e-03, 2.49543856e-03,
# 1.96893746e-03],
# [1.00000000e+00, 2.24168470e-12, 2.62942093e-13, 7.94377110e-14,
# 2.43091561e-14],
# [9.10831332e-01, 4.48558368e-02, 2.58830953e-02, 1.74687281e-02,
# 4.76850866e-04],
# [2.02121958e-01, 1.20504126e-01, 8.39806125e-02, 8.37136507e-02,
# 4.91460077e-02]], dtype=float32), indices=array([[ 4, 1, 0, 5, 2],
# [17, 14, 12, 1, 29],
# [14, 15, 2, 1, 13],
# [12, 13, 26, 17, 42],
# [ 2, 3, 5, 1, 7],
# [ 2, 1, 3, 5, 15]], dtype=int32))
# ### Step 4 (Optional): Visualize the Neural Network's State with Test Images
# This Section is not required to complete but acts as an additional excersise for understaning the output of a neural network's weights. While neural networks can be a great learning device they are often referred to as a black box. We can understand what the weights of a neural network look like better by plotting their feature maps. After successfully training your neural network you can see what it's feature maps look like by plotting the output of the network's weight layers in response to a test stimuli image. From these plotted feature maps, it's possible to see what characteristics of an image the network finds interesting. For a sign, maybe the inner network feature maps react with high activation to the sign's boundary outline or to the contrast in the sign's painted symbol.
#
# Provided for you below is the function code that allows you to get the visualization output of any tensorflow weight layer you want. The inputs to the function should be a stimuli image, one used during training or a new one you provided, and then the tensorflow variable name that represents the layer's state during the training process, for instance if you wanted to see what the LeNet lab's feature maps looked like for it's second convolutional layer you could enter conv2 as the tf_activation variable.
#
# For an example of what feature map outputs look like, check out NVIDIA's results in their paper End-to-End Deep Learning for Self-Driving Cars in the section Visualization of internal CNN State. NVIDIA was able to show that their network's inner weights had high activations to road boundary lines by comparing feature maps from an image with a clear path to one without. Try experimenting with a similar test to show that your trained network's weights are looking for interesting features, whether it's looking at differences in feature maps from images with or without a sign, or even what feature maps look like in a trained network vs a completely untrained one on the same sign image.
# +
### Visualize your network's feature maps here.
### Feel free to use as many code cells as needed.
# image_input: the test image being fed into the network to produce the feature maps
# tf_activation: should be a tf variable name used during your training procedure that represents the calculated state of a specific weight layer
# activation_min/max: can be used to view the activation contrast in more detail, by default matplot sets min and max to the actual min and max values of the output
# plt_num: used to plot out multiple different weight feature map sets on the same block, just extend the plt number for each new feature map entry
def outputFeatureMap(image_input, tf_activation, activation_min=-1, activation_max=-1 ,plt_num=1):
# Here make sure to preprocess your image_input in a way your network expects
# with size, normalization, ect if needed
# image_input =
# Note: x should be the same name as your network's tensorflow data placeholder variable
# If you get an error tf_activation is not defined it may be having trouble accessing the variable from inside a function
activation = tf_activation.eval(session=sess,feed_dict={x : image_input})
featuremaps = activation.shape[3]
plt.figure(plt_num, figsize=(15,15))
for featuremap in range(featuremaps):
plt.subplot(6,8, featuremap+1) # sets the number of feature maps to show on each row and column
plt.title('FeatureMap ' + str(featuremap)) # displays the feature map number
if activation_min != -1 & activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin =activation_min, vmax=activation_max, cmap="gray")
elif activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmax=activation_max, cmap="gray")
elif activation_min !=-1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin=activation_min, cmap="gray")
else:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", cmap="gray")
# -
| Traffic_Sign_Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/intel-analytics/BigDL/blob/branch-2.0/python/orca/colab-notebook/quickstart/keras_lenet_mnist.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="v6GU7UKOxwIm"
# 
# ---
# + [markdown] id="4kFGdO9QY04B"
# ##### Copyright 2016 The BigDL Authors.
# + id="mGdz5eaqY52t"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# + [markdown] id="hC_eOavHPXwQ"
# ## **Environment Preparation**
# + [markdown] id="oDiGYRkSx_Mj"
# **Install Java 8**
#
# Run the cell on the **Google Colab** to install jdk 1.8.
#
# **Note:** if you run this notebook on your computer, root permission is required when running the cell to install Java 8. (You may ignore this cell if Java 8 has already been set up in your computer).
# + id="ATZFtoV7l51r"
# Install jdk8
# !apt-get install openjdk-8-jdk-headless -qq > /dev/null
import os
# Set environment variable JAVA_HOME.
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
# !update-alternatives --set java /usr/lib/jvm/java-8-openjdk-amd64/jre/bin/java
# !java -version
# + [markdown] id="0Si5BdI7yJ2A"
# **Install BigDL Orca**
#
# You can install the latest pre-release version using `!pip install --pre --upgrade bigdl-orca`.
# + id="YlvUQ-BIPd54"
# Install latest pre-release version of BigDL Orca
# Installing BigDL Orca from pip will automatically install pyspark, bigdl, and their dependencies.
# !pip install --pre --upgrade bigdl-orca
# + id="qAWBOUouPj8S"
# Install python dependencies
# The tutorial below only supports TensorFlow 1.15
# !pip install tensorflow==1.15.0 tensorflow-datasets==2.1.0
# + [markdown] id="oMXB8SmUPtvr"
# ## **Distributed Keras (v2.3) using Orca APIs**
#
# In this guide we will describe how to scale out Keras (v2.3) programs using Orca in 4 simple steps.
# + id="BLUja9IKzOlK"
# import necesary libraries and modules
import argparse
from bigdl.orca import init_orca_context, stop_orca_context
from bigdl.orca.learn.tf.estimator import Estimator
from bigdl.orca import OrcaContext
# + [markdown] id="QtQjv6ypzvOg"
# ### **Step 1: Init Orca Context**
# + id="c4SvqWvKz4pI"
OrcaContext.log_output = True # recommended to set it to True when running BigDL in Jupyter notebook (this will display terminal's stdout and stderr in the Jupyter notebook).
cluster_mode = "local"
if cluster_mode == "local":
init_orca_context(cluster_mode="local", cores=4) # run in local mode
elif cluster_mode == "k8s":
init_orca_context(cluster_mode="k8s", num_nodes=2, cores=2) # run on K8s cluster
elif cluster_mode == "yarn":
init_orca_context(cluster_mode="yarn-client", num_nodes=2, cores=2) # run on Hadoop YARN cluster
# + [markdown] id="1PfwVyJsFJ6c"
# This is the only place where you need to specify local or distributed mode. View [Orca Context](https://bigdl.readthedocs.io/en/latest/doc/Orca/Overview/orca-context.html) for more details.
#
# **Note**: You should export HADOOP_CONF_DIR=/path/to/hadoop/conf/dir when you run on Hadoop YARN cluster.
# + [markdown] id="YsTyFBA10OVJ"
# ### **Step 2: Define the Model**
# You may define your model, loss and metrics in the same way as in any standard (single node) Keras program.
# + id="QWSDG7Ln0co7"
from tensorflow import keras
model = keras.Sequential(
[keras.layers.Conv2D(20, kernel_size=(5, 5), strides=(1, 1), activation='tanh',
input_shape=(28, 28, 1), padding='valid'),
keras.layers.MaxPooling2D(pool_size=(2, 2), strides=(2, 2), padding='valid'),
keras.layers.Conv2D(50, kernel_size=(5, 5), strides=(1, 1), activation='tanh',
padding='valid'),
keras.layers.MaxPooling2D(pool_size=(2, 2), strides=(2, 2), padding='valid'),
keras.layers.Flatten(),
keras.layers.Dense(500, activation='tanh'),
keras.layers.Dense(10, activation='softmax'),
]
)
model.compile(optimizer=keras.optimizers.RMSprop(),
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# + [markdown] id="HaKNUnXW3lMG"
# ### **Step 3: Define Train Dataset**
# You can define the dataset using standard tf.data.Dataset.
# + id="7Mc4V3-Y31YI"
import tensorflow as tf
import tensorflow_datasets as tfds
def preprocess(data):
data['image'] = tf.cast(data["image"], tf.float32) / 255.
return data['image'], data['label']
# get DataSet
dataset_dir = "~/tensorflow_datasets"
mnist_train = tfds.load(name="mnist", split="train", data_dir=dataset_dir)
mnist_test = tfds.load(name="mnist", split="test", data_dir=dataset_dir)
mnist_train = mnist_train.map(preprocess)
mnist_test = mnist_test.map(preprocess)
# + [markdown] id="AOGlwwku4jjL"
# ### **Step 4: Fit with Orca Estimator**
#
# First, create an Estimator
# + id="1j8MtW5V4qNm"
from bigdl.orca.learn.tf.estimator import Estimator
est = Estimator.from_keras(keras_model=model)
# + [markdown] id="T3_FzzUu5xIq"
# Next, fit and evaluate using the Estimator
# + id="mqhyRK4h5y5g"
max_epoch = 1
est.fit(data=mnist_train,
batch_size=320,
epochs=max_epoch,
validation_data=mnist_test)
# + [markdown] id="JH7sFDg4l7Mx"
# Finally, evaluate using the Estimator.
# + id="jRuELE-B6Vju"
# evaluate and print result
result = est.evaluate(mnist_test)
print(result)
est.save_keras_model("/tmp/mnist_keras.h5")
# + [markdown] id="mFTZiZSFZxI5"
# Now, the accuracy of this model has reached 97%.
# + id="2yFtIG526g1E"
# stop orca context when program finishes
stop_orca_context()
| python/orca/colab-notebook/quickstart/keras_lenet_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # OUTLINE
# * Introduce DALBAR study, criticisms
# * Methodology for using ETFs instead of MFs. Reasoning - precise, daily, ascendant
# * Choose a basket of ETFs: SPY, sectors, vol & other hot fads, small, trader funds (2x and 3x) etc...
# * Pick one major fund in depth, track across time,
# * calculate a composite of many funds, find consistent level. This is headline takeaway
# * try comparing Vanguard to Powershares, etc... Generalize
# * try comparing boring funds to trendy funds. Generalize
#
#
#
#
# # Timing of ETF Inflows and Outflows
# # Title = Our Own Worst Enemies
#
#
# ## Introduction
#
# I recently heard an investment advisor remark "We don't worry so much about an _investment's_ returns, we are most concerned with the _investor's_ returns." The advi
#
#
#
#
# +
from __future__ import print_function
from __future__ import division
from importlib import reload
import pandas as pd
import numpy as np
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
# %matplotlib inline
from IPython import display
import seaborn as sns
import datetime
import os
os.getcwd()
## NOTE: Add paths to dependencies on any machine being used here. Invalid paths will be silently ignored
import sys
sys.path.append('/anaconda2/') # either place your config.py in this path, or add another sys.path.append() pointing to your path for config.py
import config ## TODO: place file called config.py in the /anaconda2/ folder
sys.path.append(config.REPO_ROOT+'data/') #Chad: Home
from prices.eod import iqfeed
from prices.eod.read import get_adjustment_factors
# -
# +
reload(iqfeed)
#symbols = ['IBB','TLT','EEM','XLE','GLD','AGG','QQQ','FXI']
symbols = ['XLK','XLY','XLB','XLE','XLV','XLU','XLI','XLP']
#price = iqfeed.read_etf_daily(symbol)
price = iqfeed.read_etf_daily(symbols)
so = iqfeed.read_etf_so(symbols)
df = price[['close']].join(so)
df = df[df.so>0]
adj_factors = get_adjustment_factors(symbols).stack()
adj_factors.name = 'adj_factors'
#df = df.join(adj_factors)
df['close_adj'] = df.close * adj_factors
df['daily_ret'] = df.close.pct_change()
df['daily_ret_adj'] = df.close_adj.pct_change()
df['flow'] = np.log(df.so).diff()
df['mkt_cap'] = df.so*df.close
df['mkt_cap_adj'] = df.so*df.close_adj
df = df[df.daily_ret.abs()<0.2] # filter likely outliers
df = df[df.flow.abs()<0.50] # catch and exclude any rows with massive (erroneous) flows
df = df.xs(slice('2010-01-01',None),level='date',drop_level=False)
#df.dropna().tail()
#df.groupby(level='date').sum().resample('BM').sum().flow.abs().hist(bins=50)
# -
# +
## fix problem with SO counts
# off by 1000 at some date
# -
df.flow.unstack().resample('BM').sum()['2010':].sum()
# +
def calc_twr(df,div_adj=True):
if div_adj == True:
cum_ret = (df.daily_ret_adj+1).product() - 1
else:
cum_ret = (df.daily_ret+1).product() - 1
return cum_ret
def calc_approx_mwr(df,div_adj=True):
''' Calculate the return for the full time period normalized for amount of capital deployed.
'''
if div_adj == True:
# First, calculate the invested "basis" in terms of day 0 dollars.
adj_flows = df.flow*(df.mkt_cap_adj/df.mkt_cap_adj.iloc[0])
adj_flows.iloc[0] = 1 # initial principal
basis = (adj_flows).cumsum() #cumulative contributions in beginning of period dollars
# Next, convert daily returns to log basis and weight by amount of capital deployed
# to calculate an average compound return rate (log)
log_rets = np.log(df.daily_ret_adj+1)
else:
# First, calculate the invested "basis" in terms of day 0 dollars.
adj_flows = df.flow*(df.mkt_cap/df.mkt_cap.iloc[0])
adj_flows.iloc[0] = 1 # initial principal
basis = (adj_flows).cumsum() #cumulative contributions in beginning of period dollars
# Next, convert daily returns to log basis and weight by amount of capital deployed
# to calculate an average compound return rate (log)
log_rets = np.log(df.daily_ret+1)
avg_daily_log_ret = (log_rets*basis / basis.mean()).mean()
# finally, convert into a simple return for the full period
cum_ret = (avg_daily_log_ret+1)**(len(log_rets))-1
cr2 = np.exp(avg_daily_log_ret*len(basis))-1
#print(cr2)
return cum_ret
df.sort_index(inplace=True)
for symbol in symbols:
#tmp = df.xs(slice('2014-01-01','2016-12-31'),level='date',drop_level=False).xs(symbol,level='symbol',drop_level=True)
tmp = df.xs(slice('2009-01-01','2017-12-31'),level='date',drop_level=False).xs(symbol,level='symbol',drop_level=True)
print("For: {}".format(symbol))
print(" MWR (adj): {}".format(calc_approx_mwr(tmp,True)))
print(" TWR (adj): {}".format(calc_twr(tmp,True)))
#print(" MWR: {}".format(calc_approx_mwr(tmp,False)))
#print(" TWR: {}".format(calc_twr(tmp,False)))
print()
# +
def compare_annual(df):
symbols = df.index.get_level_values('symbol').unique().tolist()
out = pd.DataFrame()
for symbol in symbols:
twr = df.xs(symbol,level='symbol').resample('A').apply(calc_twr,div_adj=True)
twr.name = 'twr'
mwr = df.xs(symbol,level='symbol').resample('A').apply(calc_approx_mwr,div_adj=True)
mwr.name = 'mwr'
both = pd.concat([twr,mwr],axis=1).reset_index()
both['symbol'] = symbol
both.set_index(['date','symbol'],inplace=True)
both['timing_impact'] = both.mwr - both.twr
out = pd.concat([out,both],axis=0)
return out
by_year = compare_annual(df).timing_impact.unstack().round(3)*100
by_year.index = by_year.index.year
title = ''
sns.heatmap(by_year,center =0.00, cmap = sns.diverging_palette(10, 220, sep=2, n=21),annot=True)
#.xs(slice('2009-01-01','2009-12-31'),level='date',drop_level=False)
# -
df.flow.unstack().resample('A').sum()
df.xs('MUB',level='symbol').flow.plot()
df.loc['2009'].xs('MUB',level='symbol')['2009-12']
# +
#cash_flow_series = np.array([begin_val]),net_flows.values,end_val])
#np.irr(cash_flow_series)
# -
df = ibb.xs(slice('2012-01-01',None),level='date',drop_level=False)#ibb.loc[slice('2014-01-04',None)]
adj_flows = df.flow*(df.mkt_cap/df.mkt_cap.iloc[0]) #contributions in beginning of period dollars
adj_flows.iloc[0] = 1 # initial principal
basis = (adj_flows).cumsum() #cumulative contributions in beginning of period dollars
adj_ret = df.daily_ret*basis # daily returns weighted by amount of capital
adj_growth_factor = adj_ret + 1
print("MWR: {}".format(((adj_ret+1).product()-1)/basis.mean()))
print("TWR: {}".format(calc_twr(df)))
# +
def calc_approx_mwr(df):
''' Calculate the return for the full time period normalized for amount of capital deployed.
'''
# First, calculate the invested "basis" in terms of day 0 dollars.
adj_flows = df.flow*(df.mkt_cap/df.mkt_cap.iloc[0])
adj_flows.iloc[0] = 1 # initial principal
basis = (adj_flows).cumsum() #cumulative contributions in beginning of period dollars
# Next, convert daily returns to log basis and weight by amount of capital deployed
# to calculate an average compound return rate (log)
log_rets = np.log(df.daily_ret+1)
avg_daily_log_ret = (log_rets*basis / basis.mean()).mean()
# finally, convert into a simple return for the full period
cum_ret = (avg_daily_log_ret+1)**(len(log_rets))-1
return cum_ret
df = spy.loc['2012']
print("MWR: {}".format(calc_approx_mwr(df)))
print("TWR: {}".format(calc_twr(df)))
# -
# +
twr = ibb.xs('IBB',level='symbol').resample('A').apply(calc_twr)
twr.name = 'twr'
mwr = ibb.xs('IBB',level='symbol').resample('A').apply(calc_dollar_wtd_return)
mwr.name = 'mwr'
a = pd.concat([twr,mwr],axis=1)
a
# -
mwr
def OLD_calc_mwr(df):
''' irr in period'''
begin_val = 10000.
end_val = df.mkt_cap[-1]/df.mkt_cap[0]*10000.
cum_flow = (df.flow+1).cumprod()-1
balance = pd.Series(index=net_flows.index)
balance.iloc[0] = begin_val #+ net_flows.iloc[0]
net_flows.iloc[-1] = end_val + net_flows.iloc[-1]
irr = np.irr(net_flows)
print(irr)
print(net_flows)
cum_ret = (1+irr)**len(net_flows)-1
return cum_ret
(np.log((df.daily_ret+1)).mean()+1)**(len(df.daily_ret))-1
# +
df = ibb.loc['2012']#.xs(slice('2012-01-01',None),level='date',drop_level=False)
# TWR (weight all equally)
log_rets = np.log((df.daily_ret+1))
avg_daily_log_ret = log_rets.mean()
compound_growth = (avg_daily_log_ret+1)**(len(df.daily_ret))-1
print("TWR: {}".format(compound_growth))
# MWR (weight by dollar values)
log_rets = np.log(df.daily_ret+1)
avg_daily_log_ret = (log_rets*basis / basis.mean()).mean()
compound_growth = (avg_daily_log_ret+1)**(len(df.daily_ret))-1
print("MWR: {}".format(compound_growth))
# -
df['log_ret'] = np.log(1+df.daily_ret)
df['adj_flow'] = df.flow*(df.mkt_cap/df.mkt_cap.iloc[0])
df['future_avg_log_ret'] = df.sort_index(ascending=False).log_ret.expanding().mean()
df.future_avg_log_ret*df.flow
np.e**(log_rets.cumsum())-1
# ## Summary
# This post presented the concept of organizing data into a `features` dataframe and `outcome` dataframe, and then showed how simple it is to join these two dataframes together to train a model.
#
# True, the convention may take a few examples to get used to. However, after trial and error, I've found this to be the most error-resistant, flexible, and high-performance way to go.
#
# In the [next post], I will share some methods of feature engineering and feature selection.
#
#
| content-draft/XX_ETF_timing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Face Detection using a Pytorch CNN - Using LFW and AID Datasets
#
# LFW Dataset - https://www.kaggle.com/jessicali9530/lfw-dataset
#
# AID Dataset - https://www.kaggle.com/ashishsaxena2209/animal-image-datasetdog-cat-and-panda
import pandas as pd
import os
import numpy as np
from PIL import Image
from matplotlib.pyplot import imshow
import math
######
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
# +
class NeetoCNN(nn.Module):
def __init__(self, inchannels = 3):
super(NeetoCNN, self).__init__()
# 3 input image channel then 6 ch then 9 and finnally 12, 3x3 square convolution
# kernel
self.conv1 = nn.Conv2d(inchannels, inchannels * 2, 7)
self.conv2 = nn.Conv2d(inchannels * 2, inchannels * 3, 7)
self.conv3 = nn.Conv2d(inchannels * 3, inchannels * 4, 7)
self.conv4 = nn.Conv2d(inchannels * 4, inchannels * 5, 7)
# an affine operation: y = Wx + b
# Remember after a conv operation nout = (nin+2*npadding-nfilter)+1
# in this example
# conv1 = 250-7+1 = 244 => Maxpool => 122
# conv2 = 122-7+1 = 122 => maxpool => 58
# conv3 = 58-7+1 = 52 => maxpool => 26
# conv4 = 26-7+1 = 20 => maxpool => 10
self.fc1 = nn.Linear(inchannels * 5 * 10 * 10, 256) # 10*10 from image dimension
self.fc2 = nn.Linear(256, 100)
self.fc3 = nn.Linear(100, 25)
self.fc4 = nn.Linear(25, 4)
def forward(self, x):
x = F.max_pool2d(F.relu(self.conv1(x.float())), 2)
x = F.max_pool2d(F.relu(self.conv2(x)), 2)
x = F.max_pool2d(F.relu(self.conv3(x)), 2)
x = F.max_pool2d(F.relu(self.conv4(x)), 2)
x = x.view(-1, self.num_flat_features(x))
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = F.relu(self.fc3(x))
x = torch.sigmoid(self.fc4(x))
return x
def num_flat_features(self, x):
size = x.size()[1:] # all dimensions except the batch dimension
num_features = 1
for s in size:
num_features *= s
return num_features
myCNN = NeetoCNN()
print(myCNN)
# -
def str_converter(num, sz):
num = str(num)
while len(num) < sz:
num = "0" + num
return num
data_folder = "./Data/Data/"
face_folder = "lfw-deepfunneled/lfw-deepfunneled/"
name_ds = pd.read_csv(data_folder + "lfw/lfw_allnames.csv")
name_list = name_ds.values.tolist()
name = "Ricardo_Lazo_fd"
name_list.append([name, 7])
acum = 0
for i in name_list:
acum += i[1]
print("imagenes totales de personas", acum)
im = Image.open(data_folder + "lfw/" + face_folder + name + "/" + name + "_" + str_converter(1, 4) + ".jpg")
imshow(np.asarray(im))
aid_aug_folder = "aid_aug/animals/"
animals = ["cats", "dogs", "panda"]
print("imagenes por animales: 1000 c/u")
im = Image.open(data_folder + aid_aug_folder + animals[0] + "/" + animals[0] + "_" + str_converter(1,5) + ".jpg")
imshow(np.asarray(im))
#im = Image.open(data_folder + aid_aug_folder + animals[1] + "/" + animals[1] + "_" + str_converter(1,5) + ".jpg")
#imshow(np.asarray(im))
#im = Image.open(data_folder + aid_aug_folder + animals[2] + "/" + animals[2] + "_" + str_converter(1,5) + ".jpg")
#imshow(np.asarray(im))
Y_t = np.array([[1.,0.,0.,0.], [0.,1.,0.,0.], [0.,0.,1.,0.], [0.,0.,0.,1.]], dtype=np.float64)
# loss
loss_fn = nn.MSELoss()
# learning rate
learning_rate = 1e-4
# optimizer
optimizer = torch.optim.Adam(myCNN.parameters(), lr=learning_rate)
for t in range(200):
for i in animals:
if i == "cats":
Y_case = np.array([Y_t[1]])
elif i == "dogs":
Y_case = np.array([Y_t[2]])
else:
Y_case = np.array([Y_t[3]])
Y_case = torch.from_numpy(Y_case).to(torch.float32)
for j in range(1, 101):
im = Image.open(data_folder + aid_aug_folder + i + "/" + i + "_" + str_converter(j, 5) + ".jpg")
temp_np = np.reshape(np.array(im.convert("RGB")), (1,3,250,250))
x = Variable(torch.from_numpy(temp_np)).to(torch.float32)
# Forward pass: compute predicted y by passing x to the model.
optimizer.zero_grad()
y_pred = myCNN(x) #4*1
#print(y_pred)
# Compute and print loss.
#print(Y_case, y_pred)
loss = loss_fn(y_pred, Y_case)
loss.backward()
optimizer.step()
Y_case = np.array([Y_t[0]])
Y_case = torch.from_numpy(Y_case).double().to(torch.float32)
for i in name_list:
for j in range(1, math.ceil(i[1] * 0.1)):
im = Image.open(data_folder + "lfw/" + face_folder + i[0] + "/" + i[0] + "_" + str_converter(j, 4) + ".jpg")
temp_np = np.reshape(np.array(im), (1,3,250,250))
x = Variable(torch.from_numpy(temp_np)).to(torch.float32)
# Forward pass: compute predicted y by passing x to the model.
optimizer.zero_grad()
y_pred = myCNN(x) #4*1
# Compute and print loss.
loss = loss_fn(y_pred, Y_case)
loss.backward()
optimizer.step()
if t % 10 == 9:
print ("it ==", t + 1)
conf_mat = np.zeros((4,4))
for i in animals:
for j in range(201, 1001):
im = Image.open(data_folder + aid_aug_folder + i + "/" + i + "_" + str_converter(j, 5) + ".jpg")
temp_np = np.reshape(np.array(im.convert("RGB")), (1,3,250,250))
x = Variable(torch.from_numpy(temp_np)).to(torch.float32)
# Forward pass: compute predicted y by passing x to the model.
y_pred = myCNN(x) #4*1
# Compute and print loss.
Y = y_pred.tolist()[0]
Y = [1 if i > 0.5 else 0 for i in Y]
if i == "cats":
conf_mat[1] += Y
elif i == "dogs":
conf_mat[2] += Y
else:
conf_mat[3] += Y
for i in name_list:
for j in range(math.ceil(i[1] * 0.1), i[1] + 1):
im = Image.open(data_folder + "lfw/" + face_folder + i[0] + "/" + i[0] + "_" + str_converter(j, 4) + ".jpg")
temp_np = np.reshape(np.array(im), (1,3,250,250))
x = Variable(torch.from_numpy(temp_np)).to(torch.float32)
# Forward pass: compute predicted y by passing x to the model.
y_pred = myCNN(x) #4*1
Y = y_pred.tolist()[0]
Y = [1 if i > 0.5 else 0 for i in Y]
conf_mat[0] += Y
print(conf_mat)
| Face Detection using a Pytorch CNN - Using LFW and AID Datasets At. 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/alfmorais/estrutura_de_dados_em_python/blob/main/secao_6/aula_68.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="4XPZmW_y5Hlm"
# #Pilhas - Aula 67
#
# Empilhar um elemento em cima do outro.
#
# A - você estpa ocupado com um projeto de longo prazo
#
# B - é interrompido por um colega solicitando ajuda em um outro projeto.
#
# C - enquanto estiver trabalhando em B, alguém da contabilidade aparece para uma reunião sobre despesas de viagem
#
# D - durante a reunidão, recebe um telefonema de emergência de alguém de vendas e passa alguns minutos resolvendo um problema relacionado a um novo produto
#
# RESOLUÇÃO:
#
# D
#
# C
#
# B
#
# A
#
# Quando tiver terminado o telefonema D, voltará para a reunião C, quando tiver acabado com C, voltará para o projeto B e quando tiver terminado com B, poderá finalmente voltar para o projeto A.
#
#
#
# * Permite acesso a um item de dados: o último item inserido
# * Se o último item for removido, o item anterior ao último inserido poderá ser acessado.
# * Aplicações:
# > * Correção da expressões aritméticas, tais como 3 * (4 + 5)
# > * Percorrimento de uma árvore binária
# > * Pesquisa do vértice de um grafo
# > * Microprocessadores com arquitetura baseada em pilhas. Quando um método é chamado, seu endereço de retorno e seus parâmetros são empilhados em uma pilha e quando ele retorna, são desempilhados.
#
# # Operações
#
# - Empilhar: colocar um item de dados no topo da pilha
# - Desempilhar: Remover um item do topo da pilha
# - Ver o topo: Mostra o elemento que está no topo da pilha
# - Último a entrar Primeiro a a Sair (LIFO - Last In First Out)
#
# + id="7KF0fq1z4vzt"
# Aula 68
import numpy as np
# + id="1mcB61-G9u-e"
class Pilha:
def __init__(self, capacidade):
self.__capacidade = capacidade
self.__topo = -1
self.__valores = np.empty(self.__capacidade, dtype=int)
def __pilha_cheia(self):
if self.__topo == self.__capacidade - 1:
return True
else:
return False
def __pilha_vazia(self):
if self.__topo == -1:
return True
else:
return False
def empilhar(self, valor):
if self.__pilha_cheia():
print('A pilha está cheia')
else:
self.__topo += 1
self.__valores[self.__topo] = valor
def desempilhar(self):
if self.__pilha_vazia():
print('A pilha está vazia')
else:
self.__topo -= 1
def ver_topo(self):
if self.__topo != -1:
return self.__valores[self.__topo]
else:
return -1
# + id="78wJ6iVS-Lft"
pilha = Pilha(5)
# + colab={"base_uri": "https://localhost:8080/"} id="sLm86R1P-OsT" outputId="0d9013a3-0919-4c88-dba6-1a26e0f68f0b"
pilha.ver_topo()
# + colab={"base_uri": "https://localhost:8080/"} id="Ly9c2GBNBVVW" outputId="c4cb21e4-8edb-4a77-bd5c-6c963c1c423e"
pilha.empilhar(1)
pilha.ver_topo()
# + id="Qnn0fbe-DP7D"
pilha.empilhar(1)
pilha.empilhar(2)
pilha.empilhar(3)
pilha.empilhar(4)
# + colab={"base_uri": "https://localhost:8080/"} id="xA4_87wyDSev" outputId="009b78b1-3598-4c71-9ebe-fea10b27a1e9"
pilha.empilhar(6)
# + colab={"base_uri": "https://localhost:8080/"} id="HsAbSpOyDT2c" outputId="8712be80-a07f-44ef-d239-68334ba7c39f"
pilha.ver_topo()
# + id="rEGtWeFoDVQO"
pilha.desempilhar()
# + id="adgCGj6nDWzO"
pilha.desempilhar()
pilha.desempilhar()
pilha.desempilhar()
# + colab={"base_uri": "https://localhost:8080/"} id="CQzphTYoDYDq" outputId="f283b211-f21f-43ad-c5a7-8c8a8a543004"
pilha.ver_topo()
# + id="7Srcc3dKDaWt"
pilha.desempilhar()
# + colab={"base_uri": "https://localhost:8080/"} id="-FEI2bARDwI1" outputId="24465fbe-6a59-4998-c6f0-ac05103a1d13"
pilha.ver_topo()
| secao_6/aula_68.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Cvxpylayers tutorial
# +
import cvxpy as cp
import matplotlib.pyplot as plt
import numpy as np
import torch
from cvxpylayers.torch import CvxpyLayer
torch.set_default_dtype(torch.double)
np.set_printoptions(precision=3, suppress=True)
# -
# ## Parametrized convex optimization problem
#
# $$
# \begin{array}{ll} \mbox{minimize} & f_0(x;\theta)\\
# \mbox{subject to} & f_i(x;\theta) \leq 0, \quad i=1, \ldots, m\\
# & A(\theta)x=b(\theta),
# \end{array}
# $$
# with variable $x \in \mathbf{R}^n$ and parameters $\theta\in\Theta\subseteq\mathbf{R}^p$
#
# * objective and inequality constraints $f_0, \ldots, f_m$ are *convex* in $x$ for each $\theta$, *i.e.*, their graphs curve upward
# * equality constraints are linear
# * for a given value of $\theta$, find a value for $x$ that minimizes objective, while satisfying constraints
# * we can efficiently solve these globally with near total reliability
# ## Solution map
# * Solution $x^\star$ is an implicit function of $\theta$
# * When unique, define solution map as function
# $x^\star = \mathcal S(\theta)$
# * Need to call numerical solver to evaluate
# * This function is often differentiable
# * In a series of papers we showed how to analytically differentiate this function, using the implicit function theorem
# * Benefits of analytical differentiation: works with nonsmooth objective/constraints, low memory usage, don't compound errors
# ## CVXPY
# * High level domain-specific language (DSL) for convex optimization
# * Define variables, parameters, objective and constraints
# * Synthesize into problem object, then call solve method
# * We've added derivatives to CVXPY (forward and backward)
# ## CVXPYlayers
# 
# * Convert CVXPY problems into callable, differentiable Pytorch and Tensorflow modules in one line
# ## Applications
# * learning convex optimization models (structured prediction): https://stanford.edu/~boyd/papers/learning_copt_models.html
# * learning decision-making policies (reinforcement learning): https://stanford.edu/~boyd/papers/learning_cocps.html
# * machine learning hyper-parameter tuning and feature engineering: https://stanford.edu/~boyd/papers/lsat.html
# * repairing infeasible or unbounded optimization problems: https://stanford.edu/~boyd/papers/auto_repair_cvx.html
# * as protection layers in neural networks: http://physbam.stanford.edu/~fedkiw/papers/stanford2019-10.pdf
# * custom neural network layers (sparsemax, csoftmax, csparsemax, LML): https://locuslab.github.io/2019-10-28-cvxpylayers/
# * and many more...
# ## Average example
# Find the average of a vector:
# \begin{equation}
# \begin{array}{ll}
# \mbox{minimize} & \sum_{i=1}^n (y_i - x)^2
# \end{array}
# \end{equation}
# Variable $x$, parameters $y\in\mathbf{R}^n$
#
# The solution map is clearly:
# $$x=\sum_{i=1}^n y_i / n$$
# +
n = 7
# Define variables & parameters
x = cp.Variable()
y = cp.Parameter(n)
# Define objective and constraints
objective = cp.sum_squares(y - x)
constraints = []
# Synthesize problem
prob = cp.Problem(cp.Minimize(objective), constraints)
# Set parameter values
y.value = np.random.randn(n)
# Solve problem in one line
prob.solve(requires_grad=True)
print("solution:", "%.3f" % x.value)
print("analytical solution:", "%.3f" % np.mean(y.value))
# -
# The gradient is simply:
# $$\nabla_y x = (1/n)\mathbf{1}$$
# +
# Set gradient wrt x
x.gradient = np.array([1.])
# Differentiate in one line
prob.backward()
print("gradient:", y.gradient)
print("analytical gradient:", np.ones(y.size) / n)
# -
# ## Median example
# Finding the median of a vector:
# \begin{equation}
# \begin{array}{ll}
# \mbox{minimize} & \sum_{i=1}^n |y_i - x|,
# \end{array}
# \end{equation}
# Variable $x$, parameters $y\in\mathbf{R}^n$
#
# Solution:
# $$x=\mathbf{median}(y)$$
#
# Gradient (no duplicates):
# $$(\nabla_y x)_i = \begin{cases}
# 1 & y_i = \mathbf{median}(y) \\
# 0 & \text{otherwise}.
# \end{cases}$$
# +
n = 7
# Define variables & parameters
x = cp.Variable()
y = cp.Parameter(n)
# Define objective and constraints
objective = cp.norm1(y - x)
constraints = []
# Synthesize problem
prob = cp.Problem(cp.Minimize(objective), constraints)
# Set parameter values
y.value = np.random.randn(n)
# Solve problem in one line
prob.solve(requires_grad=True)
print("solution:", "%.3f" % x.value)
print("analytical solution:", "%.3f" % np.median(y.value))
# +
# Set gradient wrt x
x.gradient = np.array([1.])
# Differentiate in one line
prob.backward()
print("gradient:", y.gradient)
g = np.zeros(y.size)
g[y.value == np.median(y.value)] = 1.
print("analytical gradient:", g)
# -
# ## Elastic-net regression example
# We are given training data $(x_i, y_i)_{i=1}^{N}$,
# where $x_i\in\mathbf{R}$ are inputs and $y_i\in\mathbf{R}$ are outputs.
# Suppose we fit a model for this regression problem by solving the elastic-net problem
# \begin{equation}
# \begin{array}{ll}
# \mbox{minimize} & \frac{1}{N}\sum_{i=1}^N (ax_i + b - y_i)^2 + \lambda |a| + \alpha a^2,
# \end{array}
# \label{eq:trainlinear}
# \end{equation}
# where $\lambda,\alpha>0$ are hyper-parameters.
#
# We hope that the test loss $\mathcal{L}^{\mathrm{test}}(a,b) =
# \frac{1}{M}\sum_{i=1}^M (a\tilde x_i + b - \tilde y_i)^2$ is small, where
# $(\tilde x_i, \tilde y_i)_{i=1}^{M}$ is our test set.
#
# First, we set up our problem, where $\{x_i, y_i\}_{i=1}^N$, $\lambda$, and $\alpha$ are our parameters.
# +
from sklearn.datasets import make_blobs
from sklearn.model_selection import train_test_split
torch.manual_seed(0)
np.random.seed(0)
n = 2
N = 60
X, y = make_blobs(N, n, centers=np.array([[2, 2], [-2, -2]]), cluster_std=3)
Xtrain, Xtest, ytrain, ytest = train_test_split(X, y, test_size=.5)
Xtrain, Xtest, ytrain, ytest = map(
torch.from_numpy, [Xtrain, Xtest, ytrain, ytest])
Xtrain.requires_grad_(True)
m = Xtrain.shape[0]
a = cp.Variable((n, 1))
b = cp.Variable((1, 1))
X = cp.Parameter((m, n))
Y = ytrain.numpy()[:, np.newaxis]
log_likelihood = (1. / m) * cp.sum(
cp.multiply(Y, X @ a + b) - cp.logistic(X @ a + b)
)
regularization = - 0.1 * cp.norm(a, 1) - 0.1 * cp.sum_squares(a)
prob = cp.Problem(cp.Maximize(log_likelihood + regularization))
fit_logreg = CvxpyLayer(prob, [X], [a, b])
torch.manual_seed(0)
np.random.seed(0)
n = 1
N = 60
X = np.random.randn(N, n)
theta = np.random.randn(n)
y = X @ theta + .5 * np.random.randn(N)
Xtrain, Xtest, ytrain, ytest = train_test_split(X, y, test_size=.5)
Xtrain, Xtest, ytrain, ytest = map(
torch.from_numpy, [Xtrain, Xtest, ytrain, ytest])
Xtrain.requires_grad_(True)
m = Xtrain.shape[0]
# set up variables and parameters
a = cp.Variable(n)
b = cp.Variable()
X = cp.Parameter((m, n))
Y = cp.Parameter(m)
lam = cp.Parameter(nonneg=True)
alpha = cp.Parameter(nonneg=True)
# set up objective
loss = (1/m)*cp.sum(cp.square(X @ a + b - Y))
reg = lam * cp.norm1(a) + alpha * cp.sum_squares(a)
objective = loss + reg
# set up constraints
constraints = []
prob = cp.Problem(cp.Minimize(objective), constraints)
# -
# convert into pytorch layer in one line
fit_lr = CvxpyLayer(prob, [X, Y, lam, alpha], [a, b])
# this object is now callable with pytorch tensors
fit_lr(Xtrain, ytrain, torch.zeros(1), torch.zeros(1))
# sweep over values of alpha, holding lambda=0, evaluating the gradient along the way
alphas = np.logspace(-3, 2, 200)
test_losses = []
grads = []
for alpha_vals in alphas:
alpha_tch = torch.tensor([alpha_vals], requires_grad=True)
alpha_tch.grad = None
a_tch, b_tch = fit_lr(Xtrain, ytrain, torch.zeros(1), alpha_tch)
test_loss = (Xtest @ a_tch.flatten() + b_tch - ytest).pow(2).mean()
test_loss.backward()
test_losses.append(test_loss.item())
grads.append(alpha_tch.grad.item())
plt.semilogx()
plt.plot(alphas, test_losses, label='test loss')
plt.plot(alphas, grads, label='analytical gradient')
plt.plot(alphas[:-1], np.diff(test_losses) / np.diff(alphas), label='numerical gradient', linestyle='--')
plt.legend()
plt.xlabel("$\\alpha$")
plt.show()
# sweep over values of lambda, holding alpha=0, evaluating the gradient along the way
lams = np.logspace(-3, 2, 200)
test_losses = []
grads = []
for lam_vals in lams:
lam_tch = torch.tensor([lam_vals], requires_grad=True)
lam_tch.grad = None
a_tch, b_tch = fit_lr(Xtrain, ytrain, lam_tch, torch.zeros(1))
test_loss = (Xtest @ a_tch.flatten() + b_tch - ytest).pow(2).mean()
test_loss.backward()
test_losses.append(test_loss.item())
grads.append(lam_tch.grad.item())
plt.semilogx()
plt.plot(lams, test_losses, label='test loss')
plt.plot(lams, grads, label='analytical gradient')
plt.plot(lams[:-1], np.diff(test_losses) / np.diff(lams), label='numerical gradient', linestyle='--')
plt.legend()
plt.xlabel("$\\lambda$")
plt.show()
# compute the gradient of the test loss wrt all the training data points, and plot
plt.figure(figsize=(10, 6))
a_tch, b_tch = fit_lr(Xtrain, ytrain, torch.tensor([.05]), torch.tensor([.05]), solver_args={"eps": 1e-8})
test_loss = (Xtest @ a_tch.flatten() + b_tch - ytest).pow(2).mean()
test_loss.backward()
a_tch_test, b_tch_test = fit_lr(Xtest, ytest, torch.tensor([0.]), torch.tensor([0.]), solver_args={"eps": 1e-8})
plt.scatter(Xtrain.detach().numpy(), ytrain.numpy(), s=20)
plt.plot([-5, 5], [-3*a_tch.item() + b_tch.item(),3*a_tch.item() + b_tch.item()], label='train')
plt.plot([-5, 5], [-3*a_tch_test.item() + b_tch_test.item(), 3*a_tch_test.item() + b_tch_test.item()], label='test')
Xtrain_np = Xtrain.detach().numpy()
Xtrain_grad_np = Xtrain.grad.detach().numpy()
ytrain_np = ytrain.numpy()
for i in range(Xtrain_np.shape[0]):
plt.arrow(Xtrain_np[i], ytrain_np[i],
-.1 * Xtrain_grad_np[i][0], 0.)
plt.legend()
plt.show()
# move the training data points in the direction of their gradients, and see the train line get closer to the test line
plt.figure(figsize=(10, 6))
Xtrain_new = torch.from_numpy(Xtrain_np - .15 * Xtrain_grad_np)
a_tch, b_tch = fit_lr(Xtrain_new, ytrain, torch.tensor([.05]), torch.tensor([.05]), solver_args={"eps": 1e-8})
plt.scatter(Xtrain_new.detach().numpy(), ytrain.numpy(), s=20)
plt.plot([-5, 5], [-3*a_tch.item() + b_tch.item(),3*a_tch.item() + b_tch.item()], label='train')
plt.plot([-5, 5], [-3*a_tch_test.item() + b_tch_test.item(), 3*a_tch_test.item() + b_tch_test.item()], label='test')
plt.legend()
plt.show()
| examples/torch/tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# %matplotlib inline
from sklearn import tree
from sklearn import metrics,model_selection,preprocessing
from IPython.display import Image,display
import matplotlib.pyplot as plt,pydotplus
data=pd.read_csv("prity12.csv")
data.head()
data.info()
# # DataPreprocessing
df=data
df=df.drop(['Model'],axis=1)
# +
df.Engine=pd.Categorical(df.Engine)
df['EngineCode']=df.Engine.cat.codes
df['SC/Turbo']=pd.Categorical(df['SC/Turbo'])
df['SC/Turbo_code']=df['SC/Turbo'].cat.codes
df.Weight=pd.Categorical(df.Weight)
df['weight_Code']=df.Weight.cat.codes
df['Fuel Economy']=pd.Categorical(df['Fuel Economy'])
df['Fuel_Economy_code']=df['Fuel Economy'].cat.codes
df.Fast=pd.Categorical(df.Fast)
df['Fast_code']=df.Fast.cat.codes
# -
df.info()
df.columns
df_new=df.drop(['Engine', 'SC/Turbo', 'Weight', 'Fuel Economy', 'Fast'],axis=1)
df_new.head()
# # Train and Test data for the model
X=df_new.drop(['Fast_code'],axis=1)
Y=df['Fast_code']
X.shape,Y.shape
df_test=df_new.sample(n=5)
x_test=df_test.drop(['Fast_code'],axis=1)
y_test=df_test['Fast_code']
model_tree=tree.DecisionTreeClassifier()
model_tree.fit(X,Y)
y_pred=model_tree.predict(x_test)
print(y_pred)
y_test
# # Metrics Evaluation
# +
wrong_pred=(y_test != y_pred).sum()
print("Total Wrongly predicted = {}".format(wrong_pred))
accuracy=metrics.accuracy_score(y_test,y_pred)
print("Accuracy of this model = {:.3f}".format(accuracy))
# -
# # Graphical view of the tree
ddata=tree.export_graphviz(model_tree,out_file=None,filled=True,rounded=True,
feature_names=['Engine','SC/Turbo','weight','Fuel Economy'],
class_names=['YES','NO'])
graph=pydotplus.graph_from_dot_data(ddata)
display(Image(graph.create_png()))
# +
import os
os.environ['PATH'] = os.environ['PATH']+';'+os.environ['CONDA_PREFIX']+r"\Library\bin\graphviz"
# -
ddata=tree.export_graphviz(model_tree,out_file=None,filled=True,rounded=True,
feature_names=['Engine','SC/Turbo','weight','Fuel Economy'],
class_names=['YES','NO'])
graph=pydotplus.graph_from_dot_data(ddata)
display(Image(graph.create_png()))
# # Save Model
# +
import pickle
# Save the trained model as a pickle string.
model_file="model.bkp"
pickle.dump(model_tree, open(model_file, 'wb'))
#loading the model
loaded_model=pickle.load(open(model_file, 'rb'))
loaded_model.predict(x_test)
# -
| final lab code and report/decision tree/decision tree 0001 (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tensorflow_p36)
# language: python
# name: conda_tensorflow_p36
# ---
import tensorflow as tf
from keras.backend.tensorflow_backend import set_session
config = tf.ConfigProto()
config.gpu_options.per_process_gpu_memory_fraction = 0.1
set_session(tf.Session(config=config))
from tensorflow.python.client import device_lib
print(device_lib.list_local_devices())
from keras import backend as K
print(K.tensorflow_backend._get_available_gpus())
# +
# #!/usr/bin/env python3
import pandas as pd
import lz4.frame
import gzip
import io
import pyarrow.parquet as pq
import pyarrow as pa
import numpy as np
'''
filepath = 'cboe/lz4_test/BTCUSD_order_book_20170627.csv.lz4'
#filepath = 'cboe/lz4_test/BTCUSD_order_book_20170627.csv.gz'
df = pandas.read_csv(io.TextIOWrapper(lz4.frame.open(filepath)))
#df = pandas.read_csv(filepath)
#df = paratext.load_csv_to_pandas(gzip.open(filepath).read())
print((df))
'''
from glob import glob
from plumbum.cmd import rm
# -
| check_if_gpu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
import matplotlib
import seaborn as sns
from annotations.CONSTANTS import *
from main_feature_generation import do_generate_feature_vector_and_labels_for_brushing
from main_LOSOCV_brushing import evaluate_LOSOCV
from main_train_ML_model_and_export import train_and_save_AB_model
from main_mORAL_testing import process_mORAL_and_validate, process_mORAL
font = {'family': 'calibri',
'weight': 'bold',
'size': 22}
matplotlib.rc('font', **font)
matplotlib.rcParams['pdf.fonttype'] = 42
pids = [d for d in os.listdir(sensor_data_dir) if os.path.isdir(os.path.join(sensor_data_dir, d)) and len(d) == 4]
skipped_pids = ['8337', 'a764', 'aebb', 'b0e8']
print(pids)
# -
# # Directory structure for data:
#
# ## Base directory for sensor data, annotation, and output files
# CONSTANTS.data_dir = 'your_dir/data/'
#
# <br>data_dir</br> contains three folders:
# <ul>
# <li>1. annotation</li>
# <li>2. sensor_data</li>
# <li>3. features_and_MLresults</li>
# </ul>
#
# ### 1. annotation directory
# This directory contains one annotation file for each video and one metadata file for all the video
# ##### 1a. Configurations.csv
# contains all the metadata information for each video annotation
# ##### 1b. Annotation files
# For any video file one annotation file is generated in the form of 'uuuu_YYMMDD_HHmmSS.csv'
# Here 'uuuu' is the four character user id and YYMMDD_HHmmSS is the video start time.
# Each CSV file has five columns:
# <ul>
# <li> start_offset(sec) </li>
# <li>end_offset(sec) </li>
# <li>start_timestamp</li>
# <li>end_timestamp</li>
# <li>label</li>
# </ul>
#
# ### 2. sensor_data
#
# ##### 2a. one folder for each participants
# ###### 2a-i. Inside each participant's folder one folder for each day (After daywise splitting)
# Each of these directory contains 12 files (for each stream of the inertial sensor data):
# <ul>
# <li> ax_left_filename = 'left-wrist-accelx.csv' </li>
# <li> ay_left_filename = 'left-wrist-accely.csv' </li>
# <li> az_left_filename = 'left-wrist-accelz.csv' </li>
# <li> gx_left_filename = 'left-wrist-gyrox.csv' </li>
# <li> gy_left_filename = 'left-wrist-gyroy.csv' </li>
# <li> gz_left_filename = 'left-wrist-gyroz.csv' </li>
# <li> ax_right_filename = 'right-wrist-accelx.csv' </li>
# <li> ay_right_filename = 'right-wrist-accely.csv' </li>
# <li> az_right_filename = 'right-wrist-accelz.csv' </li>
# <li> gx_right_filename = 'right-wrist-gyrox.csv' </li>
# <li> gy_right_filename = 'right-wrist-gyroy.csv' </li>
# <li> gz_right_filename = 'right-wrist-gyroz.csv' </li>
# </ul>
#
#
# ### 3. Output files:
# CONSTANTS.feature_dir = data_dir + 'features_and_MLresults/'
# 
# ## Step 1: Feature Generation
#
# #### Generate features and brushing labels for all the participants, i.e., pids
# #### Export the features and labels as CSV files participantwise
do_generate_feature_vector_and_labels_for_brushing(pids)
# ## Step 2: Evaluate different models
#
# #### Evaluate different models by Leave-One-Subject_Out_Cross_validation
#
# evaluate_LOSOCV(pids, skipped_pids, do_feature_selection=True)
res_modelwise, AB_res = evaluate_LOSOCV(pids, skipped_pids, do_feature_selection=False)
# ##### LOSOCV Results
# +
fig = matplotlib.pyplot.gcf()
fig.set_size_inches(15, 10)
sns.boxplot(x='Models', y='value', data=res_modelwise, hue='Metrics', width=0.8)
plt.show()
# -
# ##### Participantwise Ada-Boost model's output
print('participantwise results....')
for p, v in AB_res.items():
print(p, v)
# ## Step 3: Training
# #### Train the best model (from previous step) and export as pickle file
#
model_filename = 'trained_model_files/brushingAB.model'
train_and_save_AB_model(pids, skipped_pids, model_filename=model_filename)
# ## Step 4: Testing
# #### Use the trained brushing model and get brushing events
# +
# Run the model and evaluate with Groundtruth
process_mORAL_and_validate(pids)
# Just testing
process_mORAL(pids)
| main_train_and_test_brushing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import
# ## Packages
# +
# General
import itertools
import os
from joblib import dump, load
# -
# ## Models
# ## In Package
# +
dct_models_package = {}
# Causality
dir_path = "./../../extdata/models/"
file_tag = "causality"
file_name = file_tag + ".joblib"
file_path = os.path.join(dir_path, file_name)
dct_models_package[file_tag] = load(file_path)
# Direction
file_tag = "direction"
file_name = file_tag + ".joblib"
file_path = os.path.join(dir_path, file_name)
dct_models_package[file_tag] = load(file_path)
# Output
dct_models_package
# -
# ## In Analysis / Development
# +
dct_models_development = {}
# Causality
dir_path = "./../data/output/"
folder_tag = "causality_direction"
file_tag= "causality"
file_name = file_tag + ".joblib"
file_path = os.path.join(dir_path, folder_tag, file_name)
dct_models_development[file_tag] = load(file_path)
# Direction
file_tag = "direction"
file_name = file_tag + ".joblib"
file_path = os.path.join(dir_path, folder_tag, file_name)
dct_models_development[file_tag] = load(file_path)
# Output
dct_models_development
# -
| inst/model_development/notebooks/direction_classification_model_verification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-1:081325390199:image/datascience-1.0
# ---
# # Hosting multiple models with multi-model endpoints
# <img align="left" width="130" src="https://raw.githubusercontent.com/PacktPublishing/Amazon-SageMaker-Cookbook/master/Extra/cover-small-padded.png"/>
#
# This notebook contains the code to help readers work through one of the recipes of the book [Machine Learning with Amazon SageMaker Cookbook: 80 proven recipes for data scientists and developers to perform ML experiments and deployments](https://www.amazon.com/Machine-Learning-Amazon-SageMaker-Cookbook/dp/1800567030)
# ### How to do it...
# %store -r model_a_s3_path
# %store -r model_b_s3_path
# %store -r s3_bucket
# %store -r prefix
# +
import sagemaker
from sagemaker import get_execution_role
session = sagemaker.Session()
role = get_execution_role()
# +
from sagemaker.image_uris import retrieve
image_uri = retrieve(
"xgboost",
region="us-east-1",
version="0.90-2"
)
image_uri
# -
models_path = f"s3://{s3_bucket}/model-artifacts/"
# +
from sagemaker.multidatamodel import MultiDataModel
multi_model = MultiDataModel(
name="chapter09-multi",
model_data_prefix=models_path,
image_uri=image_uri,
role=role
)
# -
multi_model.add_model(model_a_s3_path)
multi_model.add_model(model_b_s3_path)
list(multi_model.list_models())
# +
model_a, model_b = list(
multi_model.list_models()
)
print(model_a)
print(model_b)
# +
# %%time
endpoint_name = "chapter09-mma"
multi_model.deploy(
initial_instance_count=1,
instance_type='ml.t2.medium',
endpoint_name=endpoint_name
)
# +
from sagemaker.predictor import Predictor
from sagemaker.serializers import CSVSerializer
from sagemaker.deserializers import JSONDeserializer
predictor = Predictor(
endpoint_name=endpoint_name
)
predictor.serializer = CSVSerializer()
predictor.deserializer = JSONDeserializer()
# -
a, b = list(multi_model.list_models())
a
b
predictor.predict(data="10,-5", target_model=a)
predictor.predict(data="10,-5", target_model=b)
predictor.delete_endpoint()
| Chapter09/03 - Hosting multiple models with multi-model endpoints.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.6 64-bit
# language: python
# name: python37664bitf0c61f1e05474c16bf06d0152d368821
# ---
import networkx as nx
import pandas as pd
import numpy as np
from collections import defaultdict, Counter
import matplotlib.pyplot as plt
import networkx as nx
# %matplotlib inline
plt.style.use('ggplot')
import torch
import torch.nn.functional as F
from torch_geometric.nn import GCNConv, graclus
from torch_cluster import graclus_cluster
df2011raw = pd.read_csv('E:\Github\MLN\FinalData/2011raw.csv')
dfgraph = pd.read_csv('E:\Github\MLN\FinalData/GraphProp.csv')
dfcensus = pd.read_csv('E:\Github\MLN\FinalData/census.csv')
df2001raw = pd.read_csv('E:\Github\MLN\FinalData/2001.csv')
df2011raw.head()
df2011raw.shape
dfgraph.head()
dfcensus.head()
for i in range(34):
print('{:28} {}'.format(dfcensus['State'].tolist()[i], dfgraph.Id.tolist()[i]))
dfgraph['Pop Size'] = dfcensus['Population']
dfgraph = dfgraph.drop(columns=['Label', 'timeset'])
dfgraph.columns
dffeatures = dfgraph[['weighted indegree', 'weighted outdegree','indegree', 'outdegree', 'Degree', 'pageranks','clustering', 'Pop Size']]
dffeatures[['Area','Density','DecadalGrowth']] = dfcensus[['Area','Density','DecadalGrowth']]
dffeatures.head()
# +
decadalgrowth = []
for i in dffeatures['DecadalGrowth'].str[:-1].tolist():
try:
decadalgrowth.append(np.float(i))
except:
decadalgrowth.append(-1 * float(i[1:]))
# -
dffeatures['DecadalGrowth'] = decadalgrowth
dffeatures.head()
dffeatures.dtypes
dffeatures['Area'] = pd.to_numeric(dffeatures['Area'].str.replace(',',''))
dffeatures['Density'] = pd.to_numeric(dffeatures['Density'].str.replace(',',''))
# ## 2011
df2011raw.columns[:29]
top5 = df2011raw.groupby('Area Name')['Business Females'].apply(lambda grp: grp.nlargest(10)).index
top5data = []
for i in top5:
top5data.append([df2011raw['Last residence'].loc[i[1]], i[0], df2011raw['Business Females'].loc[i[1]]])
t = pd.DataFrame(top5data, columns=['Source', 'Target','Weight'])
t.to_csv('E:/Github/MLN/FinalData/top10'+'BusinessFemale'+'2011.csv', index=False)
# ## 2001
df2001raw.columns[:29]
top5 = df2001raw.groupby('Area Name')['Business Females'].apply(lambda grp: grp.nlargest(10)).index
top5data = []
for i in top5:
top5data.append([df2001raw['Last residence'].loc[i[1]], i[0], df2001raw['Business Females'].loc[i[1]]])
t = pd.DataFrame(top5data, columns=['Source', 'Target','Weight'])
t.to_csv('E:/Github/MLN/FinalData/top10'+'BusinessFemale'+'2001.csv', index=False)
| code/Baseline Scratch Files And Analysis Source File/GraphGeneration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 349} id="9OsVp1dWJNRr" outputId="83dd2431-1809-42a2-bbf9-af9e9a339d4f"
import numpy as np
from matplotlib import pyplot as plt
x = np.array([3.2, 3.2, 3.7, 3.9, 4, 4, 4.1, 4.5, 4.9, 5.1, 5.3, 5.9, 6])
y = np.array([54445, 64445, 57189, 63218, 55794, 56957, 57081, 61111, 67938, 66029, 83088, 81363, 93940])
b0 = np.array([1235, 1500, 1850, 2000])
b1 = np.array([1900, 1950, 2000, 2100])
plt.plot(x,y)
plt.xlabel(' X1 ')
plt.ylabel(' Y ')
plt.show()
beta_values = [(b0[i],b1[i]) for i in range(len(b0))]
for i in range(len(b0)):
values = [((y[j] - (b0[i] + (b1[i] * x[j])))**2) for j in range(len(x))]
print(f"Mean square error for 'Beta' values {beta_values[i]} : {np.mean(values)}")
| j_components_assignment2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The operating system
# 
import os
import sys
# ## name of operating system
print('os.name',',',os.name,'\n')
# ## get current dir
print('os.getcwd()',',',os.getcwd(),'\n')
# ## list files in dir
print('os.listdir()',',',os.listdir(),'\n')
# ## make a directory
# +
this_dir=os.getcwd()
directory='test'
#this adds either / or \ depending on os
path = os.path.join(this_dir, directory)
try:
os.mkdir(path)
except:
pass
#or
try:
os.mkdir('test2')
except:
pass
# -
# ## list files in dir
print('os.listdir()',',',os.listdir(),'\n')
# ## Check file/dir exists
print(os.path.exists("test2"),os.path.exists("test"))
# ## change directory
os.chdir(directory)
print('os.getcwd()',',',os.getcwd(),'\n')
# ## Create and Rename a file
# +
# create a file 'a' is append
open('Old.txt','a').close()
# check file exists
print('old=',os.path.exists("Old.txt"),'. new=',os.path.exists("New.txt"))
# rename a file
fd = "Old.txt"
os.rename(fd,'New.txt')
# check file exists
print('old=',os.path.exists("Old.txt"),'. new=',os.path.exists("New.txt"))
# -
# ## Copy a file-
# ### can be done in os but easier in shutil
#
# https://stackabuse.com/how-to-copy-a-file-in-python/
# +
import shutil
shutil.copyfile('New.txt', 'Old.txt')
# check file exists
print('old=',os.path.exists("Old.txt"),'. new=',os.path.exists("New.txt"))
# -
# ## Remove a file
# +
# remove a file
os.remove("New.txt")
# os.remove("Old.txt")
# check file exists
print('old=',os.path.exists("Old.txt"),'. new=',os.path.exists("New.txt"))
# -
# ## Go back up in directory
# +
os.path.dirname(os.path.dirname( ))
# -
#remove directory
os.rmdir(path)
#list files in dir
print('os.listdir()',',',os.listdir(),'\n')
# ## Create new files
#create new file and edit
nano file.txt
#create file
touch file.txt
# !cd
# # Subprocess module
# +
import subprocess
# subprocess.run("date",shell=True)
print(subprocess.run(['cmd', '/c', 'date']))
# -
# # Pointers for Getting Your Environment Setup
#
# ### Learning more about operating systems
#
# We’ve talked briefly about what an operating system is and what we'll need to know about operating systems for this course. If you want to learn some additional operating system concepts, check out the videos on this subject in the Technical Support Fundamentals course (https://www.coursera.org/lecture/technical-support-fundamentals/module-introduction-I3n9l). If you want to dive deeper onto how to manage Windows and Linux, check out the Operating Systems and You: Becoming a Power User course (https://www.coursera.org/learn/os-power-user).
#
# If you want to discover more about the history of Unix, you can read all the details on the Unix Wikipedia page https://en.wikipedia.org/wiki/History_of_Unix.
#
#
# ### Installing Python and additional modules
#
# If you don't have Python installed yet, we recommend that you visit the official Python website (http://www.python.org/) and download the installer that corresponds to your operating system.
#
# There’s a bunch of guides out there for installing Python and they all follow a similar process to the one we described in the videos. This guide from Real Python (https://realpython.com/installing-python/) includes instructions on how to install python on a range of different operating systems and distributions.
#
# Once you have Python installed on your operating system, it's a good idea to familiarize yourself with pip and the associated tools. You can find more info about these here https://packaging.python.org/guides/installing-using-pip-and-virtual-environments/.
#
# ### Using package management systems
#
# Package management systems help you better manage the software installed on your machine. These management systems vary a lot from operating system to operating system. So, you need to pick the one that works for the OS you’re using. Check out these guides for help with this:
#
# * Installing Python 3 on Windows 10 with Chocolatey
# https://www.digitalocean.com/community/tutorials/how-to-install-python-3-and-set-up-a-local-programming-environment-on-windows-10
#
#
# * Installing Python 3 on MacOS with Homebrew
# http://www.pyladies.com/blog/Get-Your-Mac-Ready-for-Python-Programming/
#
#
# * Package management basics on Linux
# https://www.digitalocean.com/community/tutorials/package-management-basics-apt-yum-dnf-pkg
#
#
# Other information
#
# * Python in the Microsoft Store for Windows 10
# https://devblogs.microsoft.com/python/python-in-the-windows-10-may-2019-update/
# # Setting up Your Environment
#
# After you’ve installed Python and checked that it works, the next step to set up your developer environment is to choose your main code editor.
#
# These are some of the common editors for Python, available for all platforms:
#
# * Atom https://atom.io/
#
# * Eclipse https://www.eclipse.org/
#
# * PyCharm https://www.jetbrains.com/pycharm/
#
# * Sublime Text http://www.sublimetext.com/
#
# * Visual Studio Code https://code.visualstudio.com/
#
# You can read more about these editors, and others, in these overview comparatives:
#
# * Python IDEs and Code Editors (Guide) https://realpython.com/python-ides-code-editors-guide/#pycharm
#
# * Best Python IDEs and Code Editors https://www.softwaretestinghelp.com/python-ide-code-editors/
#
# * Top 5 Python IDEs for Data Science https://www.datacamp.com/community/tutorials/data-science-python-ide
#
# We encourage you to try out these editors and pick your favorite. Then, install it on your computer and experiment with writing and executing Python scripts locally.
# # Reading and Writing Files Cheat-Sheet
#
# Check out the following link for more information:
#
# * https://docs.python.org/3/library/functions.html#open
# # Files and Directories Cheat-Sheet
#
# Check out the following links for more information:
#
# * https://docs.python.org/3/library/os.html
#
# * https://docs.python.org/3/library/os.path.html
#
# * https://en.wikipedia.org/wiki/Unix_time
# # CSV Files Cheat Sheet
#
# Check out the following links for more information:
#
# * https://docs.python.org/3/library/csv.html
#
# * https://realpython.com/python-csv/
| _notebooks/2021-10-04-OS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # COVID-19 OPEN RESEARCH DATASET
import os
import json
import pandas as pd
# %cd '/home/myilmaz/devel/covid551982_1475446_bundle_archive/'
# ## Papers researching chronic kidney disease as a comorbidity risk
import pandas as pd
#Removed hypertension
kags=pd.DataFrame(None)
for i in os.listdir('/home/myilmaz/devel/covid551982_1475446_bundle_archive/Kaggle/target_tables/8_risk_factors/'):
kag=pd.read_csv('Kaggle/target_tables/8_risk_factors/'+i)
kags=kags.append(kag)
kags.head()
# %ls document_parses
keep=['Epidemiology, clinical course, and outcomes of critically ill adults with COVID-19 in New York City: a prospective cohort study','Psychiatric Predictors of COVID-19 Outcomes in a Skilled Nursing Facility Cohort','COVID-19 in Iran, a comprehensive investigation from exposure to treatment outcomes']
arts=set(kags['Study'])
os.path.getsize('document_parses/pdf_json')/1000000
# creat=[]
# alltext=[]
# studies=[]
# for i in os.listdir('document_parses/pdf_json'):
#
# with open('document_parses/pdf_json/'+i) as json_file:
# data = json.load(json_file)
#
# study=data['metadata']['title']
#
# if study not in list(arts):
#
# if study not in studies:
# studies.append(study)
#
#
# doc=[]
# text=''
# savee=0
# for c,j in enumerate(data['body_text']):
#
# if data['metadata']['title']=='':
# print('no title!')
# text=''
# doc=[]
# words=''
# row=[]
#
# break
#
#
# words=data['body_text'][c]['text']
#
#
#
#
# if words.lower().find('covid')>-1:
# savee+=1
# if words.lower().find('sars-cov-2')>-1:
# savee+=1
#
# if (words.lower().find('sanitation')>=0 or words.lower().find('chronic')>=0 or words.lower().find('heart disease')>=0 or words.lower().find('diabetic')>=0 or words.lower().find('ckd')>=0 or words.lower().find('cardiovascular disease')>=0 or words.lower().find('diabetes')>=0):
# text=''
# doc=[]
# words=''
# row=[]
# break
# row=[i,data['metadata']['title'],data['body_text'][c]['section'],c,data['body_text'][c]['text']]
# doc.append(row)
# text+=words
#
# if savee>0:
# creat.append(doc)
# alltext.append(text)
# else:
# pass
#
# else:
# pass
# else:
# pass
jsons=[j[0] for i in creat for j in i]
titles=[j[1] for i in creat for j in i]
sections=[j[2] for i in creat for j in i]
sectionNo=[j[3] for i in creat for j in i]
text=[j[4] for i in creat for j in i]
doc=pd.DataFrame(None,columns=['jsons','titles','sections','sectionNo','text'])
doc=pd.DataFrame(None,columns=['jsons','titles','sections','sectionNo','text'])
doc.jsons=jsons
doc.titles=titles
doc.sections=sections
doc.sectionNo=sectionNo
doc.text=text
docs=doc.copy(deep=True)
docs.drop_duplicates(keep='first',inplace=True)
# NUMBER OF UNIQUE DOCUMENTS IN THE DATA SET
docs['jsons'].nunique()
#docs.to_csv('covids.csv',index=False)
docs=pd.read_csv('covids.csv')
docs.head()
# +
import numpy as np
np.min([len(i) for i in alltext])
np.max([len(i) for i in alltext])
print('Average document length is {} words'.format(np.mean([len(i) for i in alltext])))
# -
docs['case']=docs['sections'].apply(lambda x: (str(x).lower().find('case study')>=0)|(str(x).lower().find('case report')>=0)|(str(x).lower().find('case studies')>=0))
case=pd.DataFrame(docs[docs['case']==1])
cases=set(case['jsons'])
docs['kidney']=docs['text'].apply(lambda x: str(x).lower().find('creatinine')>=0)
docs['heart']=docs['text'].apply(lambda x: (str(x).lower().find('cardiac')>=0)|(str(x).lower().find('heart')>=0))
kidney=pd.DataFrame(docs[docs['kidney']==1])
kidneys=set(kidney['jsons'])
heart=pd.DataFrame(docs[docs['heart']==1])
hearts=set(heart['jsons'])
select=set(list(hearts)[0:1000]+list(kidneys)+list(cases))
len(select)
selection=pd.DataFrame(docs[docs['jsons'].isin(select)]).reset_index(drop=True)
selection.tail(40)
selection['lenSection']=selection['text'].apply(lambda x: len(x))
# ## Use pretrained NER model to find Problems, Tests, and Treatments
# +
import os
import pyspark.sql.functions as F
from pyspark.sql.functions import monotonically_increasing_id
import json
import os
from pyspark.ml import Pipeline
from pyspark.sql import SparkSession
from sparknlp.annotator import *
from sparknlp.base import *
# Install pyspark
# ! pip install --ignore-installed -q pyspark==2.4.4
# Install Spark NLP
# ! pip install --ignore-installed -q spark-nlp==2.5
import sparknlp
print (sparknlp.version())
import json
import os
from pyspark.ml import Pipeline
from pyspark.sql import SparkSession
from sparknlp.annotator import *
from sparknlp.base import *
from pyspark.sql.functions import monotonically_increasing_id
import pyspark.sql.functions as F
import pyspark.sql.types as t
# -
spark=sparknlp.start()
selection.fillna('',inplace=True)
selection.head(1)
selection.drop(columns=['case','kidney','heart'],inplace=True)
sparkdocs=spark.createDataFrame(selection).toDF('docid','title','section','sectionNo','text','lenSection')
# +
document_assembler = DocumentAssembler() \
.setInputCol("text")\
.setOutputCol('document')
sentence_detector = SentenceDetector() \
.setInputCols(["document"]) \
.setOutputCol("sentence")
tokenizer = Tokenizer() \
.setInputCols(["sentence"]) \
.setOutputCol("token")
word_embeddings = WordEmbeddingsModel.load("/home/myilmaz/cache_pretrained/embeddings_clinical_en_2.4.0_2.4_1580237286004")\
.setInputCols(["sentence", "token"])\
.setOutputCol("embeddings")
embeddingsSentence = SentenceEmbeddings() \
.setInputCols(["sentence", "embeddings"]) \
.setOutputCol("sent_embeddings") \
.setPoolingStrategy("AVERAGE")
clinical_ner = NerDLModel.load('/home/myilmaz/cache_pretrained/ner_clinical_en_2.4.0_2.4_1580237286004') \
.setInputCols(["sentence", "token", "embeddings"]) \
.setOutputCol("ner")
ner_converter = NerConverter() \
.setInputCols(["sentence", "token", "ner"]) \
.setOutputCol("ner_chunk")
nlpPipeline = Pipeline(stages=[document_assembler,sentence_detector,tokenizer,
word_embeddings,embeddingsSentence,
clinical_ner,ner_converter
])
empty_data = spark.createDataFrame([[""]]).toDF("text")
model = nlpPipeline.fit(empty_data)
# -
results=model.transform(sparkdocs)
embeds = results.select('docid','section','sectionNo','lenSection',F.explode(F.arrays_zip('sent_embeddings.metadata','sent_embeddings.embeddings')).alias("cols")) \
.select('docid','section','sectionNo','lenSection',F.expr("cols['0'].sentence").alias("sentid"),
F.col('cols.1').alias("sent_embeddings"))
exploded = results.select('docid','title','section','sectionNo','lenSection',F.explode(F.arrays_zip('token.metadata','token.result','token.begin','ner.result')).alias("cols")) \
.select('docid','title','section','sectionNo','lenSection',F.expr("cols['0'].sentence").alias("sentid"),
F.col('cols.1').alias("token"),F.col('cols.2').alias('tokenStart'),F.col('cols.3').alias("label"))
# +
from pyspark.sql.functions import udf, col
join1=udf(lambda x:','.join(x))
stringify = udf(lambda x: [str(j) for j in x])
embeds=embeds.withColumn("sent_embeddings", stringify(col("sent_embeddings")))
embeds=embeds.withColumn("sent_embeddings", join1(col("sent_embeddings")))
# -
exploded.show()
# ## Save annotated documents for further analysis
# exploded.write.option("header", "true").csv("covids3.csv")
# embeds.write.option("header", "true").csv("embeddingsCov.csv")
# please=exploded.join(embeds,['docid','section','sectionNo','sentid'],'left')
exploded.columns
import pyspark.sql.types as t
myschema = t.StructType(
[
t.StructField('docid', t.StringType(), True),
t.StructField('title', t.StringType(), True),
t.StructField('section', t.StringType(), True),
t.StructField('sectionNo', t.StringType(), True),
t.StructField('lenSection', t.StringType(), True),
t.StructField('sentid', t.IntegerType(), True),
t.StructField('token', t.StringType(), True),
t.StructField('tokenStart', t.StringType(), True),
t.StructField('label', t.StringType(), True)
]
)
# import pyspark.sql.types as t
# myschema2 = t.StructType(
# [
# t.StructField('docid', t.StringType(), True),
# t.StructField('section', t.StringType(), True),
# t.StructField('sectionNo', t.StringType(), True),
# t.StructField('sentid', t.StringType(), True),
# t.StructField('sent_embeddings', t.StringType(), True)
#
# ]
# )
# import os
# csvs=os.listdir('covids2.csv')
# big=spark.read.csv('covids2.csv/'+csvs[0],sep=',',schema=myschema,header=True)
#
#
# for i in csvs[1:]:
#
# dfs=spark.read.csv('covids2.csv/'+i,sep=',',schema=myschema,header=True)
#
# big=big.union(dfs)
#
#
# df1.join(df2, usingColumns=Seq(“col1”, …), joinType=”left”)
# import os
# csvs=os.listdir('covids3.csv')
# covs=pd.DataFrame(None)
# for i in csvs:
#
# dfs=spark.read.csv('covids3.csv/'+i,sep=',',schema=myschema,header=True)
# one=dfs.toPandas()
# covs=covs.append(one)
#
# covs.to_csv('covs.csv',index=False)
# import os
# csvs=os.listdir('embeddingsCov.csv')
# embed=pd.DataFrame(None)
# for i in csvs:
#
# dfs=spark.read.csv('embeddingsCov.csv/'+i,sep=',',schema=myschema2,header=True)
# one=dfs.toPandas()
# embed=embed.append(one)
# embed.to_csv('embed2.csv',index=False)
covs=pd.read_csv('covs.csv')
covs=covs.dropna().reset_index(drop=True)
# +
import numpy as np
tokens=[]
savei=''
save=0
for i,j in zip(covs.token,covs.label):
if j.split('-')[0]!='I':
if save<0:
tokens[save]=savei
tokens.append(None)
savei=i
save=0
continue
else:
tokens.append(savei)
savei=i
save=0
continue
elif j.split('-')[0]=='I':
savei+=' '+i
save-=1
tokens.append(None)
else:
tokens.append(None)
if save<0:
tokens[save]=savei
tokens.append(None)
else:
tokens.append(savei)
tokens=tokens[1:]
# -
covs['chunks']=tokens
risks=pd.read_csv('allrisks.csv')
testdf=covs[covs['chunks'].notnull()]
testdf=testdf[testdf['label']!='O']
#testdf['chunks'].value_counts()
testdf=testdf.drop_duplicates()
testdf2=risks[risks['chunks'].notnull()]
testdf2=testdf2[testdf2['label']!='O']
#testdf2['chunks'].value_counts()
testdf2=testdf2.drop_duplicates()
covdf=testdf.copy(deep=True)
riskdf=testdf2.copy(deep=True)
import pandas as pd
pd.set_option('display.max_colwidth', None)
pd.set_option('display.max_rows', None)
covdf['study']=covdf['chunks'].apply(lambda x: str(x).lower().find('study')>=0)
covdf['studies']=covdf['chunks'].apply(lambda x: str(x).lower().find('studies')>=0)
covdf['meta']=covdf['chunks'].apply(lambda x: str(x).lower().find('meta-analysis')>=0)
covdf['data']=covdf['chunks'].apply(lambda x: str(x).lower().find('analysis')>=0 )
covdf['stats']=covdf['chunks'].apply(lambda x: str(x).lower().find('analyses')>=0 )
covdf['review']=covdf['chunks'].apply(lambda x: str(x).lower().find('a systematic review')>=0 )
covdf['review2']=covdf['chunks'].apply(lambda x: str(x).lower().find('a systemic review')>=0 )
covdf['predict']=covdf['chunks'].apply(lambda x: str(x).lower().find('prediction')>=0 )
covdf['data2']=covdf['chunks'].apply(lambda x: str(x).lower().find('data')>=0)
covdf['vars']=covdf['chunks'].apply(lambda x: str(x).lower().find('continuous variables')>=0)
import pandas as pd
pd.set_option('display.max_colwidth', None)
pd.set_option('display.max_rows', None)
riskdf['study']=riskdf['chunks'].apply(lambda x: str(x).lower().find('study')>=0)
riskdf['studies']=riskdf['chunks'].apply(lambda x: str(x).lower().find('studies')>=0)
riskdf['meta']=riskdf['chunks'].apply(lambda x: str(x).lower().find('meta-analysis')>=0)
riskdf['data']=riskdf['chunks'].apply(lambda x: str(x).lower().find('analysis')>=0 )
riskdf['stats']=riskdf['chunks'].apply(lambda x: str(x).lower().find('analyses')>=0 )
riskdf['review']=riskdf['chunks'].apply(lambda x: str(x).lower().find('a systematic review')>=0 )
riskdf['review2']=riskdf['chunks'].apply(lambda x: str(x).lower().find('a systemic review')>=0 )
riskdf['predict']=riskdf['chunks'].apply(lambda x: str(x).lower().find('prediction')>=0 )
riskdf['data2']=riskdf['chunks'].apply(lambda x: str(x).lower().find('data')>=0)
riskdf['vars']=riskdf['chunks'].apply(lambda x: str(x).lower().find('continuous variables')>=0)
covdf['remove']=covdf['study']+covdf['studies']+covdf['meta']+covdf['stats']+covdf['vars']+covdf['data']+covdf['review']+covdf['predict']+covdf['review2']+covdf['data2']
riskdf['remove']=riskdf['study']+riskdf['studies']+riskdf['meta']+riskdf['stats']+riskdf['vars']+riskdf['data']+riskdf['review']+riskdf['predict']+riskdf['review2']+riskdf['data2']
# +
newlabel=[]
for i,j in zip(covdf['remove'],covdf['label']):
if i==1:
newlabel.append('O')
else:
newlabel.append(j)
# -
newlabel2=[]
for i,j in zip(riskdf['remove'],riskdf['label']):
if i==1:
newlabel2.append('O')
else:
newlabel2.append(j)
bigdf2=covdf.drop(columns=['study','studies','meta','stats','data','vars','review','review2','predict','remove','data2'])
riskdf2=riskdf.drop(columns=['study','studies','meta','stats','data','vars','review','review2','predict','remove','data2'])
chunks=[]
labels=[]
chunkStart=[]
for i,j,k in zip(bigdf2['chunks'],bigdf2['label'],bigdf2['tokenStart']):
if (j=='O' or j.split('-')[0]=='I'):
chunks.append('O')
labels.append('O')
chunkStart.append('O')
else:
if i!='O':
chunks.append(i)
else:
chunks.append('O')
labels.append(j)
chunkStart.append(k)
bigdf2['label']=labels
bigdf2['chunks']=chunks
bigdf2['chunkStart']=chunkStart
chunks=[]
labels=[]
chunkStart=[]
for i,j,k in zip(riskdf2['chunks'],riskdf2['label'],riskdf2['tokenStart']):
if (j=='O' or j.split('-')[0]=='I'):
chunks.append('O')
labels.append('O')
chunkStart.append('O')
else:
if i!='O':
chunks.append(i)
else:
chunks.append('O')
labels.append(j)
chunkStart.append(k)
riskdf2['label']=labels
riskdf2['chunks']=chunks
riskdf2['chunkStart']=chunkStart
bigdf2['label']=newlabel
bigdf2=pd.DataFrame(bigdf2[bigdf2['label']!='O']).reset_index()
riskdf2['label']=newlabel2
riskdf2=pd.DataFrame(riskdf2[riskdf2['label']!='O']).reset_index()
# +
riskdf3=riskdf2.dropna().reset_index(drop=True)
bigdf3=bigdf2.dropna().reset_index(drop=True)
riskdf3=riskdf3.drop_duplicates().reset_index()
bigdf3=bigdf3.drop_duplicates().reset_index()
# -
riskdf.head()
# +
bigdf3['label']=bigdf3['label'].apply(lambda x: [x])
bigdf3['chunks']=bigdf3['chunks'].apply(lambda x: [x])
bigdf4=bigdf3.groupby(['docid','title','section','sectionNo','lenSection'],as_index=False)['token','chunks','label'].sum()
# +
#riskdf3['token']=riskdf2['token'].apply(lambda x: [x])
riskdf3['label']=riskdf3['label'].apply(lambda x: [x])
riskdf3['chunks']=riskdf3['chunks'].apply(lambda x: [x])
riskdf4=riskdf3.groupby(['docid','title','section','sectionNo','lenSection'],as_index=False)['token','chunks','label'].sum()
# -
bigdf4.head()
# +
riskdf4['preexist']=1
bigdf4['preexist']=0
alldf=riskdf4.append(bigdf4)
# -
alldf=alldf.reset_index()
alldf['preexist'].value_counts()
alldf.drop(columns=['token'],inplace=True)
labeldic={'B-PROBLEM':[],'B-TEST':[],'B-TREATMENT':[]}
for row in alldf['label']:
try:
labeldic['B-PROBLEM'].append(len([i for i in row if i=='B-PROBLEM']))
except:
labeldic['B-PROBLEM'].append([])
try:
labeldic['B-TEST'].append(len([i for i in row if i=='B-TEST']))
except:
labeldic['B-TEST'].append([])
try:
labeldic['B-TREATMENT'].append(len([i for i in row if i=='B-TREATMENT']))
except:
labeldic['B-TREATMENT'].append([])
labelcols=pd.DataFrame(labeldic).reset_index()
labelcols['enttotals']=labelcols['B-PROBLEM']+labelcols['B-TEST']+labelcols['B-TREATMENT']
vocab=[item for sublist in alldf['chunks'] if type(sublist)!=float for item in sublist]
# import json
# with open('vocab.json','r') as file:
# js=file.read()
# #
# file.close()
#
# vocab=json.loads(js)
vocabset=list(set(vocab))
import collections, numpy
freq=collections.Counter(vocab)
freq=dict(freq)
a=len(alldf)
vocabdict={}
vocabdict={key:([0]*a) for (key,val) in freq.items() if val>40}
len(vocabdict)
for c,row in enumerate(alldf['chunks']):
for chunk in row:
try:
vocabdict[chunk][c]+=1
except:
pass
chunkdf=pd.DataFrame(vocabdict)
# +
from sklearn.feature_extraction.text import TfidfTransformer
tfidf = TfidfTransformer()
result=tfidf.fit_transform(chunkdf).toarray()
# +
from sklearn.decomposition import TruncatedSVD
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import Normalizer
svd = TruncatedSVD(n_components=12)
normalizer = Normalizer(copy=False)
lsa = make_pipeline(svd, normalizer)
lsas = lsa.fit_transform(result)
# -
lsas.shape
final=pd.DataFrame(lsas).reset_index().merge(labelcols.reset_index(),on='index',how='right')
final['preexist']=alldf['preexist']
final['lenSection']=alldf['lenSection']
alldf.columns
# +
slimdict2=pd.DataFrame(pd.DataFrame(result,columns=chunkdf.columns).reset_index().merge(labelcols.reset_index(),on='index',how='right'))
slimdict2['preexist']=alldf['preexist']
slimdict2['lenSection']=alldf['lenSection']
# -
slimdict2['B-PROBLEM']=slimdict2['B-PROBLEM']/alldf['lenSection']
slimdict2['B-TEST']=slimdict2['B-TEST']/alldf['lenSection']
slimdict2['B-TREATMENT']=slimdict2['B-TREATMENT']/alldf['lenSection']
slimdict2['enttotals']=slimdict2['enttotals']/alldf['lenSection']
final=final.drop(columns=['level_0','index'])
slimdict2=slimdict2.drop(columns=['level_0','index','lenSection'])
# +
###START HERE FOR
# -
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
# +
#final2=final.dropna().reset_index(drop=True)
slimdict3=slimdict2.dropna().reset_index(drop=True)
#X=final2.drop(columns=['preexist'])
X=slimdict3.drop(columns=['preexist'])
#X, y = X, final2['preexist']
X, y = X, slimdict3['preexist']
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
clf = RandomForestClassifier(n_estimators=200, random_state=42,n_jobs=-1)
clf.fit(X_train, y_train)
print("Accuracy on test data: {:.2f}".format(clf.score(X_test, y_test)))
print("Accuracy on train data: {:.2f}".format(clf.score(X_train, y_train)))
# -
list(chunkdf.columns)[44]
list(np.argsort(feature_importance)[-34:-24])
# +
from matplotlib import pyplot as plt
feature_importance = clf.feature_importances_
# Make importances relative to max importance.
feature_importance = 100.0 * (feature_importance / feature_importance.max())
sorted_idx = np.argsort(feature_importance)[-50:]
pos = np.arange(sorted_idx.shape[0]) + .5
fig, ax = plt.subplots(figsize =(10,10))
plt.barh(pos, feature_importance[sorted_idx], align='center')
plt.yticks(pos, X.columns[sorted_idx])
plt.xlim(5,100)
plt.xlabel('Relative Importance')
plt.title('Variable Importance')
plt.show()
# +
from matplotlib import pyplot as plt
feature_importance = clf.feature_importances_
# Make importances relative to max importance.
feature_importance = 100.0 * (feature_importance / feature_importance.max())
sorted_idx = np.argsort(feature_importance)[-50:]
pos = np.arange(sorted_idx.shape[0]) + .5
fig, ax = plt.subplots(figsize =(10,10))
plt.barh(pos, feature_importance[sorted_idx], align='center')
plt.yticks(pos, X.columns[sorted_idx])
plt.xlabel('Relative Importance')
plt.title('Variable Importance')
plt.show()
# +
## Removed data analysis related features
## Added length of section feature since number of entities in section was such an important feature. This increased score from 83 to...
# -
# ## Prep for visualizations
problems=bigdf2[bigdf2['label']=='B-PROBLEM']
tests=bigdf2[bigdf2['label']=='B-TEST']
# +
#len(problems[problems['chunks']=='proteinuria'])
# -
probhist=pd.DataFrame(problems['chunks'].value_counts())
probhist=probhist.rename(columns={'chunks':'counts'})
probhist2=probhist.iloc[0:100]
testhist=pd.DataFrame(tests['chunks'].value_counts())
testhist=testhist.rename(columns={'chunks':'counts'})
testhist2=testhist.iloc[0:100]
# ## Look at most frequent "Test" entities
testhist2.head(40)
import matplotlib.pyplot as plt
import seaborn as sns
fig, ax = plt.subplots(figsize =(24,12))
chart=sns.barplot(testhist2.index,testhist2['counts'])
chart.set_xticklabels(testhist2.index,rotation=90)
plt.title('Test Entities',fontsize=18)
plt.show()
# The pretrained model is returning a lot of false positive for Test entities, but you can still see that kidney related tests such as "creatinine" are well represented in the dataset.
# ## Look at most frequent "Problem" entities
probhist2.iloc[39:79]
import seaborn as sns
import seaborn as sns
fig, ax = plt.subplots(figsize =(24,12))
chart=sns.barplot(probhist2.index,probhist2['counts'])
chart.set_xticklabels(probhist2.index,rotation=90)
plt.title('Problem Entities',fontsize=18)
plt.show()
# You can see that kidney related problems such as "AKI" are well represented in the dataset.
# ## Find 'Test' entities near the most frequent kidney related 'Problem' entity
problems=pd.DataFrame(problems).reset_index(drop=True)
problems['sectionid']=problems.docid+'-'+problems.section
tests=pd.DataFrame(tests).reset_index(drop=True)
tests['sectionid']=tests.docid+'-'+tests.section
akis=pd.DataFrame(problems[problems['chunks']=='AKI']).reset_index(drop=True)
a=list(set(akis['sectionid']))
akitest=tests[tests['sectionid'].isin(a)]
# +
akicount=pd.DataFrame(akitest.groupby(['chunks'])['label'].count()).reset_index()
akicount=akicount.sort_values(by='label',ascending=False).reset_index(drop=True)
akicount.columns=['chunk','counts']
# -
akicount
# +
import seaborn as sns
import seaborn as sns
fig, ax = plt.subplots(figsize =(24,12))
chart= sns.barplot(akicount['chunk'][0:50],akicount['counts'][0:50])
chart.set_xticklabels(akicount.chunk,rotation=90)
plt.title("Clinical Tests Near 'AKI'",fontsize=20)
plt.show()
# -
# Our clinical tests NER is returning a lot of false positives but we still see that creatinine, CRP, and PCR tests are well represented in the dataset, appearing in the same section as "AKI". This tells me the information is probably not historical and I will have measurements that I can use for predictions as well as terms to use for topic modelling and text classification.
# ## Find 'Problem' entities near the most frequent kidney related 'Test' entity
# +
creatins=pd.DataFrame(tests[tests['chunks']=='creatinine']).reset_index(drop=True)
b=list(set(creatins['sectionid']))
creatprob=problems[problems['sectionid'].isin(b)]
creatcount=pd.DataFrame(creatprob.groupby(['chunks'])['label'].count()).reset_index()
creatcount=creatcount.sort_values(by='label',ascending=False).reset_index(drop=True)
creatcount.columns=['chunk','counts']
# -
creatcount
creatcounts=creatcount.iloc[0:50]
# +
import seaborn as sns
import seaborn as sns
fig, ax = plt.subplots(figsize =(24,12))
chart= sns.barplot(creatcounts.chunk,creatcounts['counts'])
chart.set_xticklabels(creatcounts.chunk,rotation=90)
plt.title("Patient Problems Near 'Creatinine' Test",fontsize=20)
plt.show()
# -
# AKI, hypertension, diabetes, and acute kidney injury are all well represented in the dataset, appearing in the same section as "creatinine" tests. This tells me the information is probably not historical and I will have measurements that I can use for predictions as well as terms to use for topic modelling and text classification.
# ## Frequency of 'patient' mentions in documents
patient=pd.DataFrame(big[(big['token'].str.lower()=='patient')|(big['token'].str.lower()=='patients')]).reset_index(drop=True)
patients=patient.groupby(['docid'])['token'].count()
patients=patients.reset_index()
patients=patients.rename(columns={'token':'counts'})
len(patients)
sns.boxplot(patients['counts'])
plt.title('Frequency of Patient Mentions in 1568 Documents',fontsize=14)
# ## Frequency of 'case report' mentions in documents
case=pd.DataFrame(big[(big['section'].str.lower()=='case report')|(big['section']=='case study')|(big['chunks'].str.lower()=='case report')|(big['chunks'].str.lower()=='case study')|(big['section'].str.lower()=='case reports')|(big['section']=='case studies')|(big['chunks'].str.lower()=='case reports')|(big['chunks'].str.lower()=='case studies')]).reset_index(drop=True)
cases=case.groupby(['docid'])['section'].count()
cases=cases.reset_index()
cases=cases.rename(columns={'section':'counts'})
len(cases)
sns.boxplot(cases['counts'])
plt.title('Frequency of Case Report/Study Mentions in 78 Documents',fontsize=14)
# 78 documents refer to case reports a median of about 550 times (The average document length is about 30,000 words.) I think I will have enough patient data to attempt some predictions.
# +
artlist=kag['Study']
pres=[]
doc=[]
for i in os.listdir('document_parses/pdf_json'):
with open('document_parses/pdf_json/'+i) as json_file:
data = json.load(json_file)
if data['metadata']['title'] in list(artlist):
for c,j in enumerate(data['body_text']):
row=[i,data['metadata']['title'],data['body_text'][c]['section'],data['body_text'][c]['text']]
doc.append(row)
pres.append(doc)
# -
jsons=[j[0] for i in pres for j in i]
titles=[j[1] for i in pres for j in i]
sections=[j[2].lower() for i in pres for j in i]
text=[j[1].lower()+'. '+j[2].lower()+'. '+j[3].lower() for i in pres for j in i]
pres2=pd.DataFrame(None,columns=['jsons','titles','sections','text'])
pres2['jsons']=jsons
pres2['titles']=titles
pres2['section']=sections
pres2['text']=text
pres2.head(1)
case=pd.DataFrame(pres2[(pd.Series(pres2['section']).str.contains('case report'))|(pd.Series(pres2['section']).str.contains('case study'))|(pd.Series(pres2['text']).str.contains('case report'))|(pd.Series(pres2['text']).str.contains('case study'))|(pd.Series(pres2['section']).str.contains('case reports'))|(pd.Series(pres2['section']).str.contains('case studies'))|(pd.Series(pres2['text']).str.contains('case reports'))|(pd.Series(pres2['text']).str.contains('case studies'))]).reset_index(drop=True)
case.head()
len(case)
case['jsons'].nunique()
case['titles'].value_counts()
| covidModel-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Imports here
import torch
# from torch import nn
# from torchvision import datasets, transforms, models
import matplotlib.pyplot as plt
from time import time
from PIL import Image
import numpy as np
# import copy
import seaborn as sns
import train_utils
import json
# %matplotlib inline
# -
def load_checkpoint(checkpoint):
'''
This function loads saved parameters and state from previously trained Neural Network.
Input: file name of checkpoint
Output: model, optimizer, device (GPU or CPU), loss criterion, and checkpoint dictionary with secondary information
'''
# Read from checkpoint.pth saved file
filename = checkpoint #f'checkpoint2_{model_name}.pth'
# Deserialize 'pickled' file (reading saved checkpoint)
### Tip: use map location to enable run on CPU model trained in GPU
checkpoint = torch.load(filename, map_location=lambda storage, loc: storage)
# Initialize model, applying custom setup for each one
model, optimizer, criterion = train_utils.set_nn(checkpoint['class_to_idx'], checkpoint['model_name'])
# load model and optimizer saved state data
model.load_state_dict(checkpoint['model_state_dict'])
optimizer.load_state_dict(checkpoint['optimizer_state_dict'])
return model, optimizer, criterion, checkpoint
def load_cat_to_name(cat_to_name_file='cat_to_name.json'):
with open(cat_to_name_file, 'r') as f:
cat_to_name = json.load(f)
return cat_to_name
def process_image(image_path):
''' Scales, crops, and normalizes a PIL image for a PyTorch model,
returns an Numpy array, as expected by Torch.Tensor.
'''
# resize to 256 pixel
image = Image.open(image_path).resize((256,256))
# Center crop to 224 pixel
width, height = image.size # Get dimensions
final_size = 224
left = (width - final_size)/2
top = (height - final_size)/2
right = (width + final_size)/2
bottom = (height + final_size)/2
image = image.crop((left, top, right, bottom))
# Transform image into np.array
im = np.array(image)
# Normalize pixels from [0 - 255] to [0 - 1.0] float range
im = (im - im.min()) / (im.max() - im.min())
# Normalize as expected by pre-trained model
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
im = (im - mean ) / std
# Transpose moving color channel from third (matplotlib) to first position (pytorch)
im = im.transpose((2, 0, 1)) # color, x, y
return im
def imshow(image, ax=None):
"""Transforms back from Tensor to Image format and display."""
if ax is None:
fig, ax = plt.subplots()
# PyTorch tensors assume the color channel is the first dimension
# but matplotlib assumes is the third dimension
image = image.transpose((1, 2, 0))
# Undo preprocessing
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
image = (std * image) + mean
# Image needs to be clipped between 0 and 1 or it looks like noise when displayed
image = np.clip(image, 0, 1)
# Remove spines/axis
ax.axis('off')
ax.imshow(image)
return ax
# +
def predict(image_path, model, device, cat_to_name, k=5):
''' Predict the class (or classes) of an image using a trained deep learning model.
'''
# Bring model to device (CPU or GPU, if available)
model.to(device)
# Make sure model is ready to be used for classifying/evaluation
model.eval()
# Transform original image into numpy array, as expected by Torch.Tensor
processed_image = process_image(image_path)
# Load processed image as Tensor. Fix: cast input image as float
input_tensor = torch.tensor(processed_image).float()
# Use GPU if available
input_tensor = input_tensor.to(device)
# # As recommended, convert input image to FloatTensor
# input_tensor = input_tensor.float()
# Add expected batch information for a single image
input_tensor = input_tensor.unsqueeze_(0)
output = model.forward(input_tensor)
probabilities = torch.exp(output)
top_p, top_class = probabilities.topk(k, dim=1)
# unpack from Tensor back to simple list
top_class = top_class.squeeze().tolist()
top_p = top_p.squeeze().tolist()
# Convert indices to actual classes
idx_to_class = {val: key for key,val in model.class_to_idx.items()}
top_label = [idx_to_class[class_] for class_ in top_class]
top_name = [cat_to_name[label] for label in top_label]
return top_p, top_label, top_name
# -
# Display an image along with the top 5 classes
import seaborn as sns
def display_result(image_path, model):
fig, axes = plt.subplots(2,1, figsize=(5,8))
# Set up title
flower_num = image_path.split('/')[2]
title = checkpoint['class_to_idx'].get(str(flower_num))
# Plot flower
img = process_image(image_path)
axes[0].set_title(title)
imshow(img, ax=axes[0]);
# Make prediction
probs, classes, names = predict(image_path, model)
# Plot bar chart
sns.barplot(x=probs, y=names, ax=axes[1],color=sns.color_palette()[0])
plt.show();
| TransferLearning/2_FlowerClassifier/image_utils.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n"}
import time
import copy
import numpy as np
# + pycharm={"name": "#%%\n"}
def draw_fill(puzzle, patternLength, patternWidth, start, count, solList):
count += 1
puzzleLength, puzzleWidth = puzzle.shape
patternNum = (puzzleWidth*puzzleLength)/(patternWidth*patternLength)
horizonal = False
if start[0] + patternLength <= puzzleLength and start[1] + patternWidth <= puzzleWidth:
horizonal = True
#if (puzzle[start[0]:start[0]+patternLength, start[1]:start[1]+patternWidth] != 0).any():
for i in range(start[0], start[0]+patternLength):
for j in range(start[1], start[1]+patternWidth):
if puzzle[i][j] != 0:
horizonal = False
if horizonal:
newPuzzle = copy.deepcopy(puzzle)
for i in range(start[0], start[0]+patternLength):
for j in range(start[1], start[1]+patternWidth):
newPuzzle[i][j] = count
if count == patternNum:
solList.append(newPuzzle)
return
for i in range(start[0], puzzleLength):
for j in range(0, puzzleWidth):
if newPuzzle[i][j] == 0:
newStart = (i, j)
break
else:
continue
break
draw_fill(newPuzzle, patternLength, patternWidth, newStart, count, solList)
vertical = False
if patternLength != patternWidth and start[0]+patternWidth <= puzzleLength and start[1]+patternLength <= puzzleWidth:
vertical = True
for i in range(start[0], start[0]+patternWidth):
for j in range(start[1], start[1]+patternLength):
if puzzle[i][j] != 0:
vertical = False
if vertical:
newPuzzle = copy.deepcopy(puzzle)
for i in range(start[0], start[0]+patternWidth):
for j in range(start[1], start[1]+patternLength):
newPuzzle[i][j] = count
if count == patternNum:
solList.append(newPuzzle)
return
for i in range(start[0], puzzleLength):
for j in range(0, puzzleWidth):
if newPuzzle[i][j] == 0:
newStart = (i, j)
break
else:
continue
break
draw_fill(newPuzzle, patternLength, patternWidth, newStart, count, solList)
def backtrack(puzzleLength, puzzleWidth, patternLength, patternWidth):
patternNum = (puzzleWidth*puzzleLength)/(patternWidth*patternLength)
solList = []
if patternNum%1 == 0:
inputPuzzle = np.zeros((puzzleLength, puzzleWidth))
draw_fill(inputPuzzle, patternLength, patternWidth, (0, 0), 0, solList)
#solList = np.asarray(solList).reshape((puzzleLength, puzzleWidth))
return solList
# +
def get_cost_c(conf, L, cluster_info=None):
# homogeneous setting; in real setting, we access cluster to get cost_c
num_stages = int(np.max(conf))
stage_cost = []
for i in range(1, num_stages):
b = np.where(conf == i)
c = np.where(conf == i+1)
# All pairs of GPU in the same node
if (b[1] == c[1]).all():
stage_cost.append(0)
else:
stage_cost.append(1)
stage_cost = np.asarray(stage_cost).reshape((1,-1))
ret = copy.deepcopy(stage_cost)
for i in range(L-1):
ret = np.concatenate((ret, stage_cost), axis=0)
return ret
def get_cost_e(conf, L, cluster_info=None):
# homogeneous setting; in real setting, we access cluster to get cost_e
# return amp_simulator()
#print(conf.shape[0] * conf.shape[1])
num_gpus_per_pipeline = conf.shape[0] * conf.shape[1] / np.max(conf)
return np.ones(L) / num_gpus_per_pipeline
def generate_initial(M, N, threads=2):
h_w_list = []
h_w_list.append((M, 1))
h_w_list.append((1, N))
known = {}
configs = []
for (h, w) in h_w_list:
solution = backtrack(M, N, h, w)
assert len(solution) > 0
config_idx = np.random.choice(len(solution), size=1)[0]
config = solution[config_idx]
configs.append(config)
solution.pop(config_idx)
known[(h, w)] = solution
#print(np.asarray(configs[0]))
return h_w_list, configs, known
def cool_down(iter, max_iter, init_temp):
return init_temp * (1 - iter / max_iter)
def neighbor(cur, known, M, N, maximum_try = 10):
h, w = cur
time_s = time.time()
while time.time() - time_s < 10:
index = np.random.choice([0,1], size=1)[0]
if index == 0:
valid = []
upper = min(M, N)
upper = min((M*N) // w, upper) + 1
for i in range(1, upper):
if (i, w) in known.keys():
solution = known[(i, w)]
else:
solution = backtrack(M, N, i, w)
known[(i, w)] = solution
if len(solution) > 0:
valid.append(i)
if len(valid) == 0:
continue
#return
new_h = np.random.choice(valid, size=1)[0]
# TODO
new_config_idx = np.random.choice(len(known[(new_h, w)]), size=1)[0]
ret = known[(new_h, w)].pop(new_config_idx)
return new_h, w, ret
else:
valid = []
upper = min(M, N)
upper = min((M*N) // h, upper) + 1
for i in range(1, upper):
if (h, i) in known.keys():
solution = known[(h, i)]
else:
solution = backtrack(M, N, h, i)
known[(h, i)] = solution
if len(solution) > 0:
valid.append(i)
if len(valid) == 0:
continue
new_w = np.random.choice(valid, size=1)[0]
new_config_idx = np.random.choice(len(known[(h, new_w)]), size=1)[0]
ret = known[(h, new_w)].pop(new_config_idx)
return h, new_w, ret
return None
def predict(configs, L, B):
costs = []
for i in range(len(configs)):
config = configs[i]
config = np.asarray(config)
#config = config.reshape((config.shape[0], config.shape[2]))
cost_e = get_cost_e(config, L)
cost_c = get_cost_c(config, L)
k = int(np.max(config))
# refer to pipeling slicing
cost = pipe_dp(L, cost_e, cost_c, k, B)[1]
costs.append(cost)
return np.asarray(costs)
# number of GPU per node
M = 8
#
N = 4
num_iter = 500
init_t = 1
# 16 layers network, 3 microbatches
L = 16
B = 3
h_w, configs, known = generate_initial(M, N)
costs = predict(configs, L, B)
for i in range(num_iter):
cur_t = cool_down(i, num_iter, init_t)
new_configs = []
new_h_w = []
for (h, w) in h_w:
step = neighbor((h, w), known, M, N)
if step is None:
new_h, new_w, new_config = (h, w, configs[h_w.index((h,w))])
else:
new_h, new_w, new_config = step
if step is None:
assert False
else:
pass
#print(step)
new_h_w.append((new_h, new_w))
new_configs.append(new_config)
new_costs = predict(new_configs, L, B)
acc_prob = np.exp(np.minimum((costs - new_costs)/ (cur_t+1e-5) , 0))
acc_index = (np.random.random(len(acc_prob)) < acc_prob)
for j in range(len(configs)):
if acc_index[j]:
configs[j] = new_configs[j]
costs[j] = new_costs[j]
configs, costs
# -
# +
# Scratch code below
# -
# +
def placement_reachable(M, N, m, n, s_joint):
#horizontal_tile = np.asarray(list(range(m * n))).reshape((m, n))
#vertical_tile = np.transpose(horizontal_tile)
horizontal_tile = np.ones((m,n))
vertical_tile = np.ones((n,m))
vertical_tile[0] = 0
t = True
i = 0
while i < N:
match = False
# Check whether horizontal
if i <= N - n:
for j in range(n-m, n):
#print(s_joint[j:, i:i+n])
match_height = n-j
# print(match_height)
if (s_joint[j:, i:i+n] == horizontal_tile[:match_height,:]).all():
# print(i, j, "h", s_joint[j:, i:i+n], horizontal_tile[:match_height,:], match_height)
i += n
if j != n-m:
t = False
match = True
break
if i <= N - m:
for j in range(n):
#print(s_joint,j,i,m, s_joint[j:, i:i+m])
match_height = n-j
if (s_joint[j:, i:i+m] == vertical_tile[:match_height,:]).all():
# print(i, j, "v", s_joint[j:, i:i+n], vertical_tile[:match_height,:], match_height)
i += m
if j != 0:
t = False
match = True
break
if not match:
return False, _
return True, t
# # ! Always assume m < n
def init(M, N, m, n, s_array):
h, w = s_array.shape
checked = np.zeros((h, w))
i = 0
j = 0
# horizontal_tile = np.asarray(list(range(m * n))).reshape((m, n))
# vertical_tile = np.transpose(horizontal_tile)
horizontal_tile = np.ones((m,n))
vertical_tile = np.ones((n,m))
vertical_tile[0] = 0
#print(s_array)
terminate = True
for i in range(h):
for j in range(w):
if checked[i][j] == 1:
continue
# Check horizontal
if i <= M - m and j <= N - n:
match_height = min(h-i, m)
if (s_array[i:i+match_height, j:j+n] == horizontal_tile[:match_height,:]).all() and (checked[i:i+m, j:j+n] != 1).all():
checked[i:i + m, j: j + n] = 1
if match_height != m:
terminate = False
continue
# Check vertical
if i <= M - n and j <= N - m:
match_height = min(h-i, n)
if (s_array[i:i+match_height, j:j+m] == vertical_tile[:match_height,:]).all() and (checked[i:i+n, j:j+m] != 1).all():
checked[i:i + n, j: j + m] = 1
if match_height != n:
terminate = False
continue
#print(i, j, s_array, checked)
return False, _
return True, terminate
# returns all possible pipe group configurations
def generate_placement(grid, len_1, len_2):
tot_len = len_1 * len_2
# possible configuration number for a row
from itertools import product
#possible_s = list(product(range(tot_len),repeat = grid.shape[1]*(len_2-1)))
#single_possible_s = list(product(list(range(tot_len)),repeat = grid.shape[1]))
possible_s = list(product(range(2),repeat = grid.shape[1]*(len_2-1)))
single_possible_s = list(product(list(range(2)),repeat = grid.shape[1]))
#print(possible_s, single_possible_s)
for i in range(len(possible_s)):
possible_s[i] = np.asarray(list(possible_s[i])).reshape(1,-1)
for i in range(len(single_possible_s)):
single_possible_s[i] = np.asarray(list(single_possible_s[i])).reshape(1,-1)
# the solution will be the union of all possible configurations in the last row
dp = [[None for j in range(len(possible_s))] for i in range(grid.shape[0])]
# initialize the first (len_1 -1) row
for s_index in range(len(possible_s)):
valid, terminate = init(grid.shape[0], grid.shape[1], len_1, len_2, possible_s[s_index].reshape(-1, grid.shape[1]))
if valid:
dp[0][s_index] = [(possible_s[s_index].reshape(-1, grid.shape[1]), terminate)]
#print(possible_s[s_index])
print(dp[0])
# dp by row index
for i in range(len_2-1, grid.shape[0]):
print(" ")
print(dp[i-1], i)
print(" ")
# iterate through all possibly reachable row?
#j = i - 1
for s_index_1 in range(len(possible_s)):
# print("haha", s_index_1, len(possible_s))
for s_index_2 in range(len(single_possible_s)):
s_1 = possible_s[s_index_1]
s_2 = single_possible_s[s_index_2]
# print(s_1, s_2)
s_joint = np.concatenate((s_1, s_2), axis=0)
# early return if the last rows themselves are not possible
#print(s_joint, valid)
if dp[i-1][s_index_1] is None:
print(i-1, s_index_1)
continue
#valid, terminate = placement_reachable(grid.shape[0], grid.shape[1], len_1, len_2, s_joint)
#valid, terminate = init(grid.shape[0], grid.shape[1], len_1, len_2, s_joint)
valid, terminate = placement_reachable(grid.shape[0], grid.shape[1], len_1, len_2, s_joint)
# print(s_joint, valid)
if valid:
if dp[i][s_index_2] is None:
dp[i][s_index_2] = []
for solution in dp[i-1][s_index_1]:
#print(i-1,solution)
sol, _ = solution
s_joint_sol = np.concatenate((copy.deepcopy(sol), s_2), axis=0)
dp[i][s_index_2].append((s_joint_sol, terminate))
# print(dp[0])
# print(dp[1])
# print(dp[2])
ret_sol = []
for i in range(len(single_possible_s)):
s = possible_s[i]
if dp[grid.shape[0]-1][i] is None:
continue
for (sol, t) in dp[grid.shape[0]-1][i]:
if t:
ret_sol.append(sol)
return ret_sol
# for len_1 in factors:
# # Genarate all possible configuratinos
# remain = num_gpu / len_1
# factors_2 = []
# for i in range(1, min(cluster_shape) + 1):
# if remain % i == 0:
# factors_2.append(i)
# for len_2 in factors_2:
# num_cut = num_gpu / (len_1*len_2)
# confs = generate_placement(grid, len_1. len_2)
# for conf in confs:
# cost_c = get_cost_c(conf)
# cost_e = get_cost_e(conf)
# opt_pipe = pipe_dp(L, cost_e, cost_c, num_cut, B)
# cost = amp_simulator(conf, opt_pipe)
| playground/pipeline/mesh_slicing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Universal Approximation Theorem:
#
# Let $\phi(\cdot)$ be a non-constant, bounded and monotone-increasing continuous function. Let $I_{m_0}$ denote the $m_0$-dimensional unit hypercube $[0, 1]^{m_0}$. The space of continuous functions on $I_{m_0}$ is denoted by $C(I_{m_0})$. Then, given any function $f \in C(I_{m_0})$ and $\epsilon > 0$, there exists an integer $m_1$ and sets of real constants $\alpha_i, \beta_i,$ and $w_{ij}$, where $i = 1, \ldots, m_1$ and $j = 1, \ldots, m_0$ such that we may define
# \begin{equation}
# F(x_1, \ldots, x_{m_0}) = \sum_{i=1}^{m_1} \alpha_i \phi\left( \sum_{j=1}^{m_0} w_{ij}x_j + b_i\right)
# \end{equation}
# as an approximation realization of the function $f(\cdot)$: that is,
# \begin{equation}
# \left| F(x_1, \ldots, x_{m_0}) - f(x_1, \ldots, x_{m_0}) \right| < \epsilon
# \end{equation}
# for all $x_1, x_2, \ldots, x_{m_0}$ in the input space.
#
# * Essentially, the Universal Approximation Theorem states that a single hidden layer is sufficient for a multilayer perceptron to compute a uniform $\epsilon$ approximation to a given training set - provided you have the *right* number of neurons and the *right* activation function. (However, this does not say that a single hidden layer is optimal with regards to learning time, generalization, etc.)
#
#
# ## Background for Error Back-Propagation
#
# * Error Back-Propagation is based on *gradient descent*.
# * Let's review/learn gradient descent:
#
# *Method of Gradient/Steepest Descent:*
#
# *move in direction opposite to the gradient vector, $g = \bigtriangledown E(\mathbf{w})$
# \begin{eqnarray}
# w(n+1) &=& w(n) - \eta g(n)\\
# \Delta w(n) &=& w(n+1) - w(n)\\
# \Delta w(n) &=& - \eta g(n) \quad \text{ Error correction rule }
# \end{eqnarray}
# * Show that using steepest descent, $E(\mathbf{w}(n+1)) < E(\mathbf{w}(n)) $
# * Recall: Taylor Series Expansion: $f(x) = f(a) + f'(a)(x-a) + \frac{f''(a)}{2!}(x-a)^2 + ....$
# * Approximate $E(\mathbf{w}(n+1))$ with Taylor Series around $w(n)$
# \begin{eqnarray}
# E(w(n+1)) &\approx& E(w(n)) + \Delta E(w(n))(w(n+1) - w(n))\\
# &\approx& E(w(n)) + g^T(n)(\Delta w(n))\\
# &\approx& E(w(n)) - \eta g^T(n)g(n)\\
# &\approx& E(w(n)) - \eta \left\| g(n) \right\|^2
# \end{eqnarray}
# * For positive, small $\eta$, the cost function is decreased
#
#
# ## Error Back-propagation
#
# * There are many approaches to train a neural network.
# * One of the most commonly used is the *Error Back-Propagation Algorithm*.
#
#
# * Two kinds of signals:
# 1. Function Signals: presumed to perform useful function at the output of the network, also called input signal
# 2. Error Signals: propagates backwards, involves an error-dependent function
#
# * Each hidden or output neuron performs two computations:
# 1. Computation of function signal going out of this neuron
# 2. Computation of an estimate of the gradient vector
#
#
# * First let's consider the output layer...
#
# * Given a training set, $\left\{ \mathbf{x}_n, d_n\right\}_{n = 1}^N$, we want to find the parameters of our network that minimizes the squared error:
# \begin{equation}
# E(w) = \frac{1}{2} \sum_{n=1}^N (d_n - y_n)^2
# \end{equation}
#
# * What is a common optimization approach to estimate the parameters that minimize an objective/error function? *gradient descent*
# * To use gradient descent, what do we need? The analytic form of the gradient.
#
# \begin{eqnarray}
# \frac{ \partial E}{\partial w_i} &=& \frac{\partial}{\partial w_i} \left[ \frac{1}{2} \sum_{n=1}^N (d_n - y_n)^2 \right]\\
# &=& \frac{1}{2} \sum_{n=1}^N \frac{\partial}{\partial w_i} (d_n - y_n)^2 \\
# &=& \frac{1}{2} \sum_{n=1}^N 2(d_n - y_n) \frac{\partial}{\partial w_i} (d_n - y_n) \\
# &=& \sum_{n=1}^N (d_n - y_n) \left( \frac{\partial}{\partial w_i} d_n - \frac{\partial}{\partial w_i} y_n \right) \\
# &=& \sum_{n=1}^N (d_n - y_n) \left( - \frac{\partial }{\partial w_i} y_n \right)
# \end{eqnarray}
#
# * What is $y_n$ in terms of $w_i$? (At first let's assume we have no hidden layers, only the output layer to deal with)
#
# \begin{equation}
# y_n = \phi(v_n) = \phi(\mathbf{w}^T \mathbf{x}_n)
# \end{equation}
#
# * Going back to computing our gradient...
# \begin{eqnarray}
# &=& \sum_{n=1}^N (d_n - y_n) \left( - \frac{\partial }{\partial w_i} y_n \right) \\
# &=& \sum_{n=1}^N - (d_n - y_n) \frac{\partial y_n}{\partial v_n} \frac{\partial v_n}{\partial w_i}
# \end{eqnarray}
#
# * So, $\frac{\partial y_n}{\partial v_n}$ will depend on the form of the activation function we use. If we use the sigmoid: $y_n = \frac{1}{1 + \exp(-\alpha v_n)}$, then *what is* $\frac{\partial y_n}{\partial v_n}$ ?
#
# \begin{eqnarray}
# \frac{\partial y_n}{\partial v_n} &=& \frac{\partial \phi(v_n)}{\partial v_n}\\
# &=& \frac{ \partial }{\partial v_n} \frac{1}{1 + \exp(-\alpha v_n)} \\
# &=& \frac{ \left(1 + \exp(- \alpha v_n)\right)\left(\frac{ \partial }{\partial v_n} 1\right) - \left(1\right)\left( \frac{ \partial }{\partial v_n} 1 + \exp(- \alpha v_n) \right)}{(1 + \exp(-\alpha v_n))^2}\\
# &=& \frac{ - \frac{ \partial }{\partial v_n} (1 + \exp(- \alpha v_n) ) }{(1 + \exp(-\alpha v_n))^2}\\
# &=& \frac{ -1 }{(1 + \exp(-\alpha v_n))^2} \exp(-\alpha v_n)(-\alpha)\\
# &=&\frac{ 1 }{1 + \exp(-\alpha v_n)} \frac{ 1 }{1 + \exp(- \alpha v_n)} \exp(-\alpha v_n)\\
# &=&\frac{ 1 }{1 + \exp(-\alpha v_n)} \frac{ \exp(-\alpha v_n) }{1 + \exp(-\alpha v_n)}\\
# &=& y_n (1-y_n)
# \end{eqnarray}
#
# * Going back to computing our gradient...
# \begin{eqnarray}
# &=& \sum_{n=1}^N - (d_n - y_n) \frac{\partial y_n}{\partial v_n} \frac{\partial v_n}{\partial w_i} \\
# &=& \sum_{n=1}^N - (d_n - y_n) y_n (1-y_n) \frac{\partial v_n}{\partial w_i}\\
# &=& \sum_{n=1}^N - (d_n - y_n) y_n (1-y_n) \frac{\partial }{\partial w_i} \mathbf{w}^T \mathbf{x}_n\\
# &=& \sum_{n=1}^N - (d_n - y_n) y_n (1-y_n) x_{ni}
# \end{eqnarray}
#
# * *Now that we have the gradient, how do we use this to update the output layer weights in our MLP?*
# * *How will this update equation (for the output layer) change if the network is a multilayer perceptron with hidden units?*
# * *Can you write this in vector form to update all weights simultaneously?*
#
#
#
# ## Backpropagation through hidden layers
#
# * Now to address hidden layers... We have to deal with the credit assignment problem.
#
# <img src="HiddenLayerImage.png" width="500">
#
# * Suppose we want to update $w_{ji}$ where $j$ is a hidden layer.
#
# * The error objective function over all $N$ data points is
#
# \begin{equation}
# E(n) = \frac{1}{2}\sum_{n=1}^N e_n^2 = \sum_{n=1}^N \left(d_n - y_n \right)^2 = \sum_{n=1}^N \left(d_n - \phi_n(v_n(n)) \right)^2
# \end{equation}
#
# \begin{eqnarray}
# \frac{\partial E(n)}{\partial w_{lj}} &=& \frac{\partial E(n)}{\partial e_l(n)} \frac{\partial e_l(n)}{\partial y_l(n)}\frac{\partial y_l(n)}{\partial v_{l}(n)}\frac{\partial v_l(n)}{\partial w_{lj}}\\
# &=& [e_l][-1][\phi^{\prime}(v_l(n))][y_{jl}(n)] \\
# \end{eqnarray}
#
# * Let's define a *local gradient* $\delta_l(n)$:
#
# \begin{eqnarray}
# \delta_l(n) &=& - \frac{\partial E(n)}{\partial v_l(n)}\\
# &=& e_l(n) \phi^{\prime}(v_l(n))
# \end{eqnarray}
#
# * Similarly,
#
# \begin{eqnarray}
# \delta_j(n) &=& - \frac{\partial E(n)}{\partial v_j(n)}\\
# &=& - \frac{\partial E(n)}{\partial y_j(n)}\frac{\partial y_j(n)}{\partial v_j(n)} \\
# &=& - \frac{\partial E(n)}{\partial y_j(n)} \phi^{\prime}(v_j(n))
# \end{eqnarray}
#
# * Note that:
# \begin{eqnarray}
# \frac{\partial E(n)}{\partial y_j(n)} &=& \sum_{l}e_l(n)\frac{\partial e_l(n)}{\partial y_j(n)} \\
# &=& \sum_{l}e_l(n)\frac{\partial e_l(n)}{\partial v_l(n)} \frac{\partial v_l(n)}{\partial y_j(n)} \\
# &=& \sum_{l}e_l(n)[-\phi^{\prime}(v_l(n))][w_{lj}(n)]
# \end{eqnarray}
#
# * So,
#
# \begin{eqnarray}
# \delta_j(n) &=& - \frac{\partial E(n)}{\partial y_j(n)} \phi^{\prime}(v_j(n)) \\
# &=& - \left[ \sum_{l}e_l(n)[-\phi^{\prime}(v_l(n))][w_{lj}(n)] \right] \phi^{\prime}(v_j(n))\\
# &=& \phi^{\prime}(v_j(n)) \sum_l \delta_l(n)w_{lj}(n)
# \end{eqnarray}
#
# * So, you can write the gradient at a hidden neuron in terms of the local gradient and the connected neurons in the next layer
#
# \begin{equation}
# \delta w_{ij}(n) = \eta \delta_j(n) y_i(n)
# \end{equation}
#
| Lecture17_Backpropagation/Lecture 17 - Backpropagation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: RAPIDS Stable
# language: python
# name: rapids-stable
# ---
# # Data Visualization
#
# The RAPIDS AI ecosystem and `cudf.DataFrame` are built on a series of standards that simplify interoperability with established and emerging data science tools.
#
# With a growing number of libraries adding GPU support, and a `cudf.DataFrame`’s ability to convert `.to_pandas()`, a large portion of the Python Visualization ([PyViz](pyviz.org/tools.html)) stack is immediately available to display your data.
#
# In this Notebook, we’ll walk through some of the data visualization possibilities with BlazingSQL.
#
# Blog post: [Data Visualization with BlazingSQL](https://blog.blazingdb.com/data-visualization-with-blazingsql-12095862eb73?source=friends_link&sk=94fc5ee25f2a3356b4a9b9a49fd0f3a1)
#
# #### Overview
# - [Matplotlib](#Matplotlib)
# - [Datashader](#Datashader)
# - [HoloViews](#HoloViews)
# - [cuxfilter](#cuxfilter)
from blazingsql import BlazingContext
bc = BlazingContext()
# ### Dataset
#
# The data we’ll be using for this demo comes from the [NYC Taxi dataset](https://www1.nyc.gov/site/tlc/about/tlc-trip-record-data.page) and is stored in a public AWS S3 bucket.
# +
bc.s3('blazingsql-colab', bucket_name='blazingsql-colab')
bc.create_table('taxi', 's3://blazingsql-colab/yellow_taxi/taxi_data.parquet')
# -
# Let's give the data a quick look to get a clue what we're looking at.
bc.sql('select * from taxi').tail()
# ## Matplotlib
#
# [GitHub](https://github.com/matplotlib/matplotlib)
#
# > _Matplotlib is a comprehensive library for creating static, animated, and interactive visualizations in Python._
#
# By calling the `.to_pandas()` method, we can convert a `cudf.DataFrame` into a `pandas.DataFrame` and instantly access Matplotlib with `.plot()`.
#
# For example, **does the `passenger_count` influence the `tip_amount`?**
bc.sql('SELECT * FROM taxi').to_pandas().plot(kind='scatter', x='passenger_count', y='tip_amount')
# Other than the jump from 0 to 1 or outliers at 5 and 6, having more passengers might not be a good deal for the driver's `tip_amount`.
# Let's see what demand is like. Based on dropoff time, **how many riders were transported by hour?** i.e. column `7` will be the total number of passengers dropped off from 7:00 AM through 7:59 AM for all days in this time period.
riders_by_hour = '''
select
sum(passenger_count) as sum_riders,
hour(cast(tpep_dropoff_datetime || '.0' as TIMESTAMP)) as hour_of_the_day
from
taxi
group by
hour(cast(tpep_dropoff_datetime || '.0' as TIMESTAMP))
order by
hour(cast(tpep_dropoff_datetime || '.0' as TIMESTAMP))
'''
bc.sql(riders_by_hour).to_pandas().plot(kind='bar', x='hour_of_the_day', y='sum_riders', title='Sum Riders by Hour', figsize=(12, 6))
# Looks like the morning gets started around 6:00 AM, and builds up to a sustained lunchtime double peak from 12:00 PM - 3:00 PM. After a quick 3:00 PM - 5:00 PM siesta, we're right back for prime time from 6:00 PM to 8:00 PM. It's downhill from there, but tomorrow is a new day!
# +
solo_rate = len(bc.sql('select * from taxi where passenger_count = 1')) / len(bc.sql('select * from taxi')) * 100
print(f'{solo_rate}% of rides have only 1 passenger.')
# -
# The overwhelming majority of rides have just 1 passenger. How consistent is this solo rider rate? **What's the average `passenger_count` per trip by hour?**
#
# And maybe time of day plays a role in `tip_amount` as well, **what's the average `tip_amount` per trip by hour?**
#
# We can run both queries in the same cell and the results will display inline.
# +
xticks = [n for n in range(24)]
avg_riders_by_hour = '''
select
avg(passenger_count) as avg_passenger_count,
hour(dropoff_ts) as hour_of_the_day
from (
select
passenger_count,
cast(tpep_dropoff_datetime || '.0' as TIMESTAMP) dropoff_ts
from
taxi
)
group by
hour(dropoff_ts)
order by
hour(dropoff_ts)
'''
bc.sql(avg_riders_by_hour).to_pandas().plot(kind='line', x='hour_of_the_day', y='avg_passenger_count', title='Avg. # Riders per Trip by Hour', xticks=xticks, figsize=(12, 6))
avg_tip_by_hour = '''
select
avg(tip_amount) as avg_tip_amount,
hour(dropoff_ts) as hour_of_the_day
from (
select
tip_amount,
cast(tpep_dropoff_datetime || '.0' as TIMESTAMP) dropoff_ts
from
taxi
)
group by
hour(dropoff_ts)
order by
hour(dropoff_ts)
'''
bc.sql(avg_tip_by_hour).to_pandas().plot(kind='line', x='hour_of_the_day', y='avg_tip_amount', title='Avg. Tip ($) per Trip by Hour', xticks=xticks, figsize=(12, 6))
# -
# Interestingly, they almost resemble each other from 8:00 PM to 9:00 AM, but where average `passenger_count` continues to rise until 3:00 PM, average `tip_amount` takes a dip until 3:00 PM.
#
# From 3:00 PM - 8:00 PM average `tip_amount` starts rising and average `passenger_count` waits patiently for it to catch up.
#
# Average `tip_amount` peaks at midnight, and bottoms out at 5:00 AM. Average `passenger_count` is highest around 3:00 AM, and lowest at 6:00 AM.
# ## Datashader
#
# [GitHub](https://github.com/holoviz/datashader)
#
# > Datashader is a data rasterization pipeline for automating the process of creating meaningful representations of large amounts of data.
#
# As of [holoviz/datashader#793](https://github.com/holoviz/datashader/pull/793), the following Datashader features accept `cudf.DataFrame` and `dask_cudf.DataFrame` input:
#
# - `Canvas.points`, `Canvas.line` and `Canvas.area` rasterization
# - All reduction operations except `var` and `std`.
# - `transfer_functions.shade` (both 2D and 3D) inputs
#
# #### Colorcet
#
# [GitHub](https://github.com/holoviz/colorcet)
#
# > Colorcet is a collection of perceptually uniform colormaps for use with Python plotting programs like bokeh, matplotlib, holoviews, and datashader based on the set of perceptually uniform colormaps created by <NAME> at the Center for Exploration Targeting.
from datashader import Canvas, transfer_functions as tf
from colorcet import fire
# **Do dropoff locations change based on the time of day?** Let's say 6AM-4PM vs 6PM-4AM.
# Dropoffs from 6:00 AM to 4:00 PM
query = '''
select
dropoff_x, dropoff_y
from
taxi
where
hour(cast(tpep_pickup_datetime || '.0' as TIMESTAMP)) BETWEEN 6 AND 15
'''
nyc = Canvas().points(bc.sql(query), 'dropoff_x', 'dropoff_y')
tf.set_background(tf.shade(nyc, cmap=fire), "black")
# Dropoffs from 6:00 PM to 4:00 AM
query = '''
select
dropoff_x, dropoff_y
from
taxi
where
hour(cast(tpep_pickup_datetime || '.0' as TIMESTAMP)) BETWEEN 18 AND 23
OR hour(cast(tpep_pickup_datetime || '.0' as TIMESTAMP)) BETWEEN 0 AND 3
'''
nyc = Canvas().points(bc.sql(query), 'dropoff_x', 'dropoff_y')
tf.set_background(tf.shade(nyc, cmap=fire), "black")
# While Manhattan makes up the majority of the dropoff geography from 6:00 AM to 4:00 PM, Midtown's spark grows and spreads deeper into Brooklyn and Queens in the 6:00 PM to 4:00 AM window.
#
# Consistent with the more decentralized look across the map, dropoffs near LaGuardia Airport (upper-middle right side) also die down relative to surrounding areas as the night rolls in.
# ## HoloViews
#
# [GitHub](https://github.com/holoviz/holoviews)
#
# > HoloViews is an open-source Python library designed to make data analysis and visualization seamless and simple. With HoloViews, you can usually express what you want to do in very few lines of code, letting you focus on what you are trying to explore and convey, not on the process of plotting.
#
# By calling the `.to_pandas()` method, we can convert a `cudf.DataFrame` into a `pandas.DataFrame` and hand off to HoloViews or other CPU visualization packages.
# +
from holoviews import extension, opts
from holoviews import Scatter, Dimension
import holoviews.operation.datashader as hd
extension('bokeh')
opts.defaults(opts.Scatter(height=425, width=425), opts.RGB(height=425, width=425))
cmap = [(49,130,189), (107,174,214), (123,142,216), (226,103,152), (255,0,104), (50,50,50)]
# -
# With HoloViews, we can easily explore the relationship of multiple scatter plots by saving them as variables and displaying them side-by-side with the same code cell.
#
# For example, let's reexamine `passenger_count` vs `tip_amount` next to a new `holoviews.Scatter` of `fare_amount` vs `tip_amount`.
#
# **Does `passenger_count` affect `tip_amount`?**
# +
s = Scatter(bc.sql('select passenger_count, tip_amount from taxi').to_pandas(), 'passenger_count', 'tip_amount')
# 0-6 passengers, $0-$100 tip
ranged = s.redim.range(passenger_count=(-0.5, 6.5), tip_amount=(0, 100))
shaded = hd.spread(hd.datashade(ranged, x_sampling=0.25, cmap=cmap))
riders_v_tip = shaded.redim.label(passenger_count="Passenger Count", tip_amount="Tip ($)")
# -
# **How do `fare_amount` and `tip_amount` relate?**
# +
s = Scatter(bc.sql('select fare_amount, tip_amount from taxi').to_pandas(), 'fare_amount', 'tip_amount')
# 0-30 miles, $0-$60 tip
ranged = s.redim.range(fare_amount=(0, 100), tip_amount=(0, 100))
shaded = hd.spread(hd.datashade(ranged, cmap=cmap))
fare_v_tip = shaded.redim.label(fare_amount="Fare Amount ($)", tip_amount="Tip ($)")
# -
# Display the answers to both side by side.
riders_v_tip + fare_v_tip
# ## cuxfilter
#
# [GitHub](https://github.com/rapidsai/cuxfilter)
#
# > cuxfilter (ku-cross-filter) is a RAPIDS framework to connect web visualizations to GPU accelerated crossfiltering. Inspired by the javascript version of the original, it enables interactive and super fast multi-dimensional filtering of 100 million+ row tabular datasets via cuDF.
#
# cuxfilter allows us to culminate these charts into a dashboard.
import cuxfilter
# Create `cuxfilter.DataFrame` from a `cudf.DataFrame`.
cux_df = cuxfilter.DataFrame.from_dataframe(bc.sql('SELECT passenger_count, tip_amount, dropoff_x, dropoff_y FROM taxi'))
# Create some charts & define a dashboard object.
# +
chart_0 = cuxfilter.charts.datashader.scatter_geo(x='dropoff_x', y='dropoff_y')
chart_1 = cuxfilter.charts.bokeh.bar('passenger_count', x_range=[-0.5, 6.5], add_interaction=False)
chart_2 = cuxfilter.charts.datashader.heatmap(x='passenger_count', y='tip_amount', x_range=[-0.5, 6.5], y_range=[0, 100],
color_palette=cmap, title='Passenger Count vs Tip Amount ($)')
# -
dashboard = cux_df.dashboard([chart_0, chart_1, chart_2], title='NYC Yellow Cab')
# Display charts in Notebook with `.view()`.
chart_0.view()
chart_2.view()
# ## Multi-GPU Data Visualization
#
# Packages like Datashader and cuxfilter support dask_cudf distributed objects (Series, DataFrame).
# +
from dask_cuda import LocalCUDACluster
from dask.distributed import Client
cluster = LocalCUDACluster()
client = Client(cluster)
bc = BlazingContext(dask_client=client, network_interface='lo')
# +
bc.s3('blazingsql-colab', bucket_name='blazingsql-colab')
bc.create_table('distributed_taxi', 's3://blazingsql-colab/yellow_taxi/taxi_data.parquet')
# -
# Dropoffs from 6:00 PM to 4:00 AM
# +
query = '''
select
dropoff_x, dropoff_y
from
distributed_taxi
where
hour(cast(tpep_pickup_datetime || '.0' as TIMESTAMP)) BETWEEN 18 AND 23
OR hour(cast(tpep_pickup_datetime || '.0' as TIMESTAMP)) BETWEEN 0 AND 3
'''
nyc = Canvas().points(bc.sql(query), 'dropoff_x', 'dropoff_y')
tf.set_background(tf.shade(nyc, cmap=fire), "black")
# -
# ## That's the Data Vizualization Tour!
#
# You've seen the basics of Data Visualization in BlazingSQL Notebooks and how to utilize it. Now is a good time to experiment with your own data and see how to parse, clean, and extract meaningful insights from it.
#
# We'll now get into how to run Machine Learning with popular Python and GPU-accelerated Python packages.
#
# Continue to the [Machine Learning introductory Notebook](machine_learning.ipynb)
| intro_notebooks/data_visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from scipy.linalg import hankel, eigh
from matplotlib import pyplot as plt
from matplotlib import style
# ## Data fetching and cleaning for analysis.
xmeas = np.asarray(pd.read_csv('xmv10_359_data_1.csv', usecols=[14], header=None))
X_train_data = xmeas[:500] # For training
X_theta = xmeas[250:4000] # For threshold calculation
X_test = xmeas[3750:] # For detection
# ## Embedding (Creating Lag Covariance Matrix)
N = len(X_train_data)
L = N // 2
X_train = hankel(X_train_data[:L],X_train_data[L-1:]) # Creating trajectory matrix
# ## Singular Value Decomposition of Log Covariance matrix (Trajectory Matrix above)
eigenValues, eigenVectors = eigh(np.matmul(X_train, X_train.T))
idx = eigenValues.argsort()[::-1]
eigenValues = eigenValues[idx]
eigenVectors = eigenVectors[:,idx]
# Sree plot
fig, ax = plt.subplots(1, 1, figsize=(10,8))
ax.plot(eigenValues)
ax.set_title("Screeplot")
ax.set_ylabel("Eigen value ->")
ax.set_xlabel("Cardinal Number of Eigen value ->")
# ## Projection onto signal subspace
# +
# From the above scree plot, it is seen that the first eigen value is more
# significant and the other eigen values are close to zero. Thus we choose r as 1
# and then we compute the U for projection and centroid in the signal subspace
r = 1
# Extracted Training signals
U, Sigma, V = np.linalg.svd(X_train)
V = V.T
# d = np.linalg.matrix_rank(X_train)
X_elem = np.array( [Sigma[i] * np.outer(U[:,i], V[:,i]) for i in range(0,r)] )
X_train_extracted = X_elem.sum(axis=0)
X_train_extracted_data = np.asarray(list(X_train_extracted[:,0]) + list(X_train_extracted[:,-1]))
U = eigenVectors[:,:r] # r as statistical dimension
UT = U.T
pX = np.matmul(UT,X_train_extracted)
centroid = np.mean(pX, axis=1)
centroid = centroid[:,np.newaxis]
# -
# ## Distance Tracking and Threshold Calculation
# +
# Calculating the departure threshold in signal subspace using centroid and UT
#For training phase
Xtrg = hankel(X_train_data[:L], X_train_data[L-1:])
pXtrg = np.matmul(UT,Xtrg)
dtrg_matrix = centroid - pXtrg
dtrg_scores = np.linalg.norm(dtrg_matrix, axis=0, ord=2)
# For Validation phase and threshold calculation
Xt = hankel(X_theta[:L],X_theta[L-1:])
pXt = np.matmul(UT,Xt)
dt_matrix = centroid - pXt
dt_scores = np.linalg.norm(dt_matrix, axis=0, ord=2)
# d_scores = np.asarray([np.matmul(d_matrix[:,i].T, d_matrix[:,i]) for i in range(d_matrix.shape[1])])
dt_theta = np.max(dt_scores)
dt_theta
# -
# ## Testing Phase
Xj = hankel(X_test[:L],X_test[L-1:])
pXj = np.matmul(UT, Xj)
dj_matrix = centroid - pXj
dj_scores = np.linalg.norm(dj_matrix, axis=0, ord=2)
# dj_scores = [np.matmul(dj_matrix[:,i].T, dj_matrix[:,i]) for i in range(dj_matrix.shape[1])]
dj_scores = np.asarray(dj_scores)
np.max(dj_scores)
# ## Plotting and Visualizing
# +
#Testing Plots and Subplots
style.use('default')
box = dict(facecolor='yellow', pad=3, alpha=0.2)
fig = plt.figure(figsize=(10,7))
ax1 = fig.add_subplot(211)
ax2 = fig.add_subplot(212)
ax1.set_xlim(0,5000)
ax2.set_xlim(0,5000)
# ax2.set_ylim(0,10)
plt.subplots_adjust(hspace =0.3)
xlables = list(range(0,5000,10)) # for both plots
# Plotting signal reading
xmeasx_1 = list(range(501))
xmeasx_2 = list(range(501, 4001))
xmeasx_3 = list(range(4001,len(xmeas)))
ax1.plot(xmeasx_1, xmeas[:501] ,'b', label='Training') # Plot of Training Data
ax1.plot(xmeasx_2, xmeas[501:4001] ,'k', label='Threshold calculation') # Plot of Threshold Determination Data
ax1.plot(xmeasx_3, xmeas[4001:] ,'r', label='Detection') # Plot of Detection Phase
ax1.plot(X_train_extracted_data, 'g', linewidth=1, label='Extracted Signal' )
ax1.set_xticklabels(xlables)
ax1.title.set_text('Direct Attack 1 Scenario')
ax1.set_ylabel('Sensor Reading', bbox=box)
ylim = list(ax1.get_ylim())
ax1.vlines(4000,ylim[0],ylim[1],linestyles='dashed', colors='r')
X = np.array([[4000,5000],[4000,5000]])
Y = np.array([[ylim[0],ylim[0]],[ylim[1],ylim[1]]])
C = np.array([[4000,4500]])
ax1.pcolormesh(X, Y, C, cmap='cool_r', alpha=0.2)
ax1.legend(loc='best', ncol=4)
# Plotting departure score
dy = dtrg_scores
dx = list(range(L,len(dy)+L))
ax2.plot(dx, dy, 'c', label='Training phase')
dy = dt_scores
dx = list(range(500,len(dy)+500))
ax2.plot(dx, dy, 'b', label='Threshold calculation')
dy = dj_scores
dx = list(range(4000,len(dy)+4000))
ax2.plot(dx, dy, 'r', label='Detection Phase')
ylim = list(ax2.get_ylim())
ax2.vlines(4000,ylim[0],ylim[1],linestyles='dashed', colors='r')
ax2.set_xticklabels(xlables)
ax2.hlines(dt_theta,0,5000,linestyles='dashed', label='Alarm Threshold')
ax2.set_xlabel('Time in hours', bbox=box)
ax2.set_ylabel('Departure Score', bbox=box)
X = np.array([[4000,5000],[4000,5000]])
Y = np.array([[ylim[0],ylim[0]],[ylim[1],ylim[1]]])
C = np.array([[4000,4500]])
ax2.pcolormesh(X, Y, C, cmap='cool_r', alpha=0.2)
ax2.legend(loc='upper left')
fig.align_ylabels([ax1,ax2])
# -
| data + Solutions/1 - Scenario DA1/.ipynb_checkpoints/da1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Taken from https://www.kaggle.com/rikdifos/nba-players-salary-prediction/
# -
# <font size=5 >NBA Players Salary Prediction</font>
# <font size=4 >Python Application Using AdaBoost Algorithm</font>
# Author:[<NAME>](https://xsong.ltd/en) a.k.a. Malcolm
#
# > [Another Version](https://xsong.ltd/en/nba/) is available on my personal website.
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Data-Wrangling" data-toc-modified-id="Data-Wrangling-1"><span class="toc-item-num">1 </span>Data Wrangling</a></span></li><li><span><a href="#Descriptive-Statistics-&--Data-Visualization" data-toc-modified-id="Descriptive-Statistics-&--Data-Visualization-2"><span class="toc-item-num">2 </span>Descriptive Statistics & Data Visualization</a></span><ul class="toc-item"><li><span><a href="#Average-Points-Per-Game-Ranking" data-toc-modified-id="Average-Points-Per-Game-Ranking-2.1"><span class="toc-item-num">2.1 </span>Average Points Per Game Ranking</a></span></li><li><span><a href="#Player-Efficiency-Rating-Ranking" data-toc-modified-id="Player-Efficiency-Rating-Ranking-2.2"><span class="toc-item-num">2.2 </span>Player Efficiency Rating Ranking</a></span></li><li><span><a href="#High-Age-Players-Ranking" data-toc-modified-id="High-Age-Players-Ranking-2.3"><span class="toc-item-num">2.3 </span>High Age Players Ranking</a></span></li><li><span><a href="#Turnover-Ranking" data-toc-modified-id="Turnover-Ranking-2.4"><span class="toc-item-num">2.4 </span>Turnover Ranking</a></span></li><li><span><a href="#Correlation-Plot" data-toc-modified-id="Correlation-Plot-2.5"><span class="toc-item-num">2.5 </span>Correlation Plot</a></span></li><li><span><a href="#Correlation-between-Points-per-game-and-age" data-toc-modified-id="Correlation-between-Points-per-game-and-age-2.6"><span class="toc-item-num">2.6 </span>Correlation between Points per game and age</a></span></li><li><span><a href="#Which-position-is-the-most-prone-to-turnovers?" data-toc-modified-id="Which-position-is-the-most-prone-to-turnovers?-2.7"><span class="toc-item-num">2.7 </span>Which position is the most prone to turnovers?</a></span></li><li><span><a href="#Who-has-the-most-blocks?" data-toc-modified-id="Who-has-the-most-blocks?-2.8"><span class="toc-item-num">2.8 </span>Who has the most blocks?</a></span></li></ul></li><li><span><a href="#Preprocessing" data-toc-modified-id="Preprocessing-3"><span class="toc-item-num">3 </span>Preprocessing</a></span><ul class="toc-item"><li><span><a href="#Dependent-Variable" data-toc-modified-id="Dependent-Variable-3.1"><span class="toc-item-num">3.1 </span>Dependent Variable</a></span></li></ul></li><li><span><a href="#Algorithms" data-toc-modified-id="Algorithms-4"><span class="toc-item-num">4 </span>Algorithms</a></span><ul class="toc-item"><li><span><a href="#CART" data-toc-modified-id="CART-4.1"><span class="toc-item-num">4.1 </span>CART</a></span></li><li><span><a href="#AdaBoost" data-toc-modified-id="AdaBoost-4.2"><span class="toc-item-num">4.2 </span>AdaBoost</a></span></li></ul></li></ul></div>
# -
# # Data Wrangling
# + [Data from Kaggle](https://www.kaggle.com/koki25ando/nba-salary-prediction-using-multiple-regression/comments)
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from pylab import *
# %matplotlib inline
# %config InlineBackend.figure_format = 'svg'
salary_table = pd.read_csv("NBA_season1718_salary.csv",encoding = 'utf-8')
seasons = pd.read_csv("Seasons_Stats.csv",encoding = 'utf-8')
# + Variable Explaining
#
#
# Variable | meaning
# ------------- | -------------
# Pos | Position
# MPG | Average Minutes played Per Game
# PPG | Average Points Per Game
# APG | Average Assists Per Game
# RPG | Average Rebounds Per Game
# TOPG | Average Turnovers Per Game
# BPG | Average Blocks Per Game
# SPG | Average Steals Per Game
# PER | Player Efficiency Rating
# G | Games Attended
# AGE | Age
# salary17_18 | 2017-2018 Salary(million)
# +
salary_table = salary_table[['Player','season17_18']]
salary_table.rename(columns={'season17_18':'salary17_18'},inplace = True) #variable rename
salary_table['salary17_18'] = salary_table['salary17_18']/1000000 #transform salary to 'million'
seasons = seasons[seasons['Year']>=2017]
stats17 = seasons[['Year','Player','Pos','Age','G','PER',
'MP','PTS','AST','TRB','TOV','BLK','STL']]
stats17.drop_duplicates(subset=['Player'], keep='first',inplace=True) #drop duplicate data
c = ['MPG','PPG','APG','RPG','TOPG','BPG','SPG']
w = ['MP','PTS','AST','TRB','TOV','BLK','STL']
for i,s in zip(c,w):
stats17[i] = stats17[s] / stats17['G']
stats17.drop(w,axis=1,inplace=True)
#stats17.drop(['G'],axis=1,inplace=True)
stats17.loc[stats17['Pos'] == 'PF-C','Pos'] = 'PF'
stats_salary = pd.merge(stats17, salary_table)
# -
stats_salary.columns
# # Descriptive Statistics & Data Visualization
# ## Average Points Per Game Ranking
stats_salary.drop_duplicates(subset=['Player'],keep='first',inplace=True)
stats_salary.sort_values(by='PPG',ascending=False,inplace = True)
stats_salary[['Player','PPG']].head(10)
# ## Player Efficiency Rating Ranking
stats_salary.sort_values(by='PER',ascending = False,inplace = True)
stats_salary[['Player','PER']].head(10)
# ## High Age Players Ranking
stats_salary.sort_values(by='Age',ascending = False,inplace = True)
stats_salary[['Player','Age']].head(10)
# 
# ## Turnover Ranking
# + In the turnover list per game, <NAME> ranked first, this should be related to the ball, and will be verified later.
stats_salary.sort_values(by='TOPG',ascending=False,inplace = True)
stats_salary[['Player','TOPG']].head(10)
# ## Correlation Plot
sns.set_style("white")
heat_salary= stats_salary[['salary17_18','Pos','MPG','PPG','APG','RPG','TOPG',
'BPG','SPG','Age','PER']]
dfData = heat_salary.corr()
sns.heatmap(dfData)
# + Age and multiple data indicators are weakly correlated.
# + Average turnovers per game and playing time have strong correlation, and it might explain why <NAME> has some many turnovers averagely.
# + Because of position, there is almost no correlation between averaging blocks and assists per game.
# ## Correlation between Points per game and age
sns.lmplot(x="Age", y="PPG",hue="Pos",col="Pos",col_wrap=3,
data=stats_salary,lowess=True).set(
xlabel='Position',
ylabel='Average Points Per Game')
# For point guards, age and average scores are inverse U-shaped.
# ## Which position is the most prone to turnovers?
sns.boxplot(x="Pos", y="TOPG", data=stats_salary).set(
xlabel='Position',
ylabel='Average Turnovers Per Game')
# Answer:Point Guards
# ## Who has the most blocks?
# no doubt
sns.violinplot(x="Pos", y="BPG", data=stats_salary).set(
xlabel='Position',
ylabel='Average Blocks Per Game')
# # Preprocessing
from sklearn.preprocessing import Normalizer
# +
#from mpl_toolkits.mplot3d import Axes3D
#sns.pairplot(heat_salary)
# -
# Most variables are skew, thus we should rescale them.
# %config InlineBackend.figure_format = 'png'
from pandas.plotting import scatter_matrix
#scatter_matrix(heat_salary)
scatter_matrix(heat_salary, alpha=0.2, figsize=(10,10), diagonal='kde')
# %config InlineBackend.figure_format = 'svg'
# ## Dependent Variable
salary_table['salary17_18'].describe()
plt.hist(stats_salary['salary17_18'],density=True,bins=50)
plt.xlabel('2017-2018 Salary(million)')
plt.ylabel('Density')
plt.show()
# to convert categorical feature to dummy. make new dummies into name_value.
# it will automatic delete a value group as reference, which default to be biggest group.
def convert_dummy(df, feature,rank=0):
pos = pd.get_dummies(df[feature], prefix=feature)
mode = df[feature].value_counts().index[rank]
biggest = feature + '_' + str(mode)
pos.drop([biggest],axis=1,inplace=True)
df.drop([feature],axis=1,inplace=True)
df=df.join(pos)
return df
stats_salary = convert_dummy(stats_salary,'Pos')
# # Algorithms
# ## CART
# +
#from sklearn.externals.six import StringIO
#import pydotplus
#import graphviz
#from IPython.display import Image
#import os
stats_salary = stats_salary.dropna()
Y = stats_salary['salary17_18']
X = stats_salary.drop(['salary17_18','Year', 'Player'],axis=1)
# -
X.columns
# Now let us rescale our data
from sklearn.preprocessing import Normalizer
from sklearn.preprocessing import RobustScaler
from sklearn.preprocessing import MaxAbsScaler
from sklearn.preprocessing import MinMaxScaler
transformer = MaxAbsScaler().fit(X) # Scale each feature by its maximum absolute value.
newX = transformer.transform(X)
newX = pd.DataFrame(newX,columns = X.columns)
X.head()
newX.head() # transformed data
newX.describe()
# The new transformed features has been standardize, for example, age.
# +
#transformer = MinMaxScaler().fit(heat)
#newX = transformer.transform(heat)
#newX = pd.DataFrame(newX)
#scatter_matrix(newX, alpha=0.2, figsize=(10,10), diagonal='kde')
#newX.head()
#X.head()
# +
#transformer = RobustScaler().fit(heat)
#newX = transformer.transform(heat)
#newX = pd.DataFrame(newX)
#scatter_matrix(newX, alpha=0.2, figsize=(10,10), diagonal='kde')
#sns.pairplot(pd.DataFrame(newX))
#newX
# +
#from sklearn.preprocessing import StandardScaler
#transformer = StandardScaler().fit(heat)
#newX2 = transformer.transform(heat)
#newX2 = pd.DataFrame(newX2)
#scatter_matrix(newX2, alpha=0.2, figsize=(10,10), diagonal='kde')
#newX2
# -
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(X, Y, test_size = 0.3)
x_train_NEW, x_test_NEW, y_train_NEW, y_test_NEW = train_test_split(newX, Y, test_size = 0.3)
from sklearn.metrics import mean_squared_error
from sklearn.metrics import r2_score
import numpy as np
# Define a function to evaluation regression algorithms, model is fitted algorithms
# predict is for if display comparison of prediction and true value of test data.
def RegEvaluation(model, ytest, xtest, nameindex, yname,totaldt, predict=True):
ypred = model.predict(xtest)
xtest['Pred_Y'] = model.predict(xtest)
dt = pd.merge(totaldt,xtest,how = 'right')
xtest.drop(['Pred_Y'],axis=1,inplace=True)
dt = dt[[nameindex, yname,'Pred_Y']]
dt.sort_values(by = yname, ascending = False,inplace=True)
rmse = np.sqrt(mean_squared_error(ytest, ypred))
r2 = r2_score(ytest, ypred)
print('RMSE is', rmse)
print('R sequared is', r2)
if predict:
return dt.head(20)
# + Comparison of real and predicted values (Test data)
# +
from sklearn import tree
clf = tree.DecisionTreeRegressor(max_depth=4, criterion="mse")
dtree = clf.fit(x_train, y_train)
RegEvaluation(dtree, y_test, x_test, 'Player', 'salary17_18',stats_salary)
# -
dtree = clf.fit(x_train_NEW, y_train_NEW)
RegEvaluation(dtree, y_test_NEW, x_test_NEW,
'Player', 'salary17_18',stats_salary,predict=False)
# The most important feature deciding players' salary is minutes per game. When play time longer than 31 minuates,which is standard for starting players, the expectation of salary is 18 millions dollars. When the time is less than 31.4 minuates, the average salary is just 5.18 millions dollars. For those who have more playing time, next important features are PER and Age. Players are more efficient and more aged tend to have higher salary. Using Standardized features, $RMSE$ decreases and $R^2$ imporves. Data preprocessing is really important for predicting.
sns.set_style("whitegrid")
from tabulate import tabulate
values = sorted(zip(x_train.columns, clf.feature_importances_), key=lambda x: x[1] * -1)
imp = pd.DataFrame(values,columns = ["Name", "Score"])
imp.sort_values(by = 'Score',inplace = True)
sns.scatterplot(x='Score',y='Name',linewidth=0,
data=imp,s = 30, color='red').set(
xlabel='Importance',
ylabel='Variables')
# + Unsurprisingly, the average score has the greatest impact on salary levels. The second factor is age. Average turnover and steals have no effect on salary.
# ## AdaBoost
# > [How does AdaBoost Classifer predict?](https://machinelearningmastery.com/boosting-and-adaboost-for-machine-learning/): Adaboost iteratively generates K weak classifiers. The final model is the addition of K weak classifiers. Of course, each base classifier is multiplied by its own weight. For the two-class problem (-1,1), each weak classifier will calculate a prediction result of -1 or 1. These prediction results are multiplied by the weighting coefficients of the respective weak classifiers and added. If the final result is greater than 0, it is predicted to be positive (1); otherwise, it is predicted to be negative (-1).
# + display some of test data prediction results
# +
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import AdaBoostRegressor
stats_salary = stats_salary.dropna()
reg = AdaBoostRegressor(DecisionTreeRegressor(max_depth=5),
n_estimators=500)
adaboost = reg.fit(x_train, y_train)
RegEvaluation(adaboost, y_test, x_test, 'Player', 'salary17_18',stats_salary)
# -
ada = reg.fit(x_train_NEW, y_train_NEW)
RegEvaluation(ada, y_test_NEW, x_test_NEW,
'Player', 'salary17_18',stats_salary,predict=False)
# Apperantly,AdaBoost has a higher prediction accuracy. Using Standardized features, $R^2$ has imporved.
# + Variable Importance
values = sorted(zip(x_train.columns, reg.feature_importances_), key = lambda x: x[1] * -1)
imp = pd.DataFrame(values,columns = ["Name", "Score"])
imp.sort_values(by = 'Score',inplace = True)
sns.scatterplot(x='Score',y='Name',linewidth=0,
data=imp,s = 30, color='red').set(
xlabel='Importance',
ylabel='Variables')
# + The results of the AdaBoost algorithm show that the average playing time and efficiency rate have the most important impact on salary, while the average score is only ranked third. This is quite different from the decision tree results.
# +
# #?pd.DataFrame.sort_values
# +
import joblib
joblib.dump(reg, "model.pkl")
joblib.dump(x_train_NEW, "x_train.pkl")
joblib.dump(x_test_NEW, "x_test.pkl")
joblib.dump(y_train_NEW, "y_train.pkl")
joblib.dump(y_test_NEW, "y_test.pkl")
# -
| Final Test Models/nba/nba-players-salary-prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Change directory to VSCode workspace root so that relative path loads work correctly. Turn this addition off with the DataScience.changeDirOnImportExport setting
# ms-python.python added
import os
try:
os.chdir(os.path.join(os.getcwd(), '..'))
print(os.getcwd())
except:
pass
# # X To A Summary
import os
import numpy as np
import pandas as pd
from larval_gonad.io import pickle_load, shelve_load, feather_to_cluster_matrix
# +
pd.options.display.max_rows = 200
try:
os.chdir(os.path.join(os.getcwd(), 'docs'))
print(os.getcwd())
except:
pass
# -
fbgn2chrom = (
pd.read_feather("../references/gene_annotation_dmel_r6-26.feather")
.set_index("FBgn")
.FB_chrom
.rename("chrom")
)
# +
expressed = pickle_load("../output/cellselection-wf/expressed_genes.pkl")
print(f"Number of expressed genes: {len(expressed):,}")
# Expressed by chrom
fbgn2chrom.reindex(expressed).value_counts()
# +
commonly_expressed = pickle_load("../output/cellselection-wf/commonly_expressed_genes.pkl")
print(f"Number of commonly expressed genes: {len(commonly_expressed):,}")
# Commonly expressed by chrom
fbgn2chrom.reindex(commonly_expressed).value_counts()
# -
# Y genes in other clusters
y_genes = fbgn2chrom[fbgn2chrom == "Y"].index.tolist()
tpm = feather_to_cluster_matrix("../output/seurat3-cluster-wf/tpm_by_cluster.feather").reindex(y_genes)
_num = tpm[(tpm.iloc[:, 4:] > 0).any(axis=1)].shape[0]
print(f"Number of Y genes in somatic clusters: {_num:,}")
cb = shelve_load("../output/x-to-a-wf/db/commonly_expressed.bak")
# Median Ratios
cb['data'].groupby(['cluster', 'ratio_type']).ratio.median().unstack().dropna(axis=1)
# Log2FC
np.log2(cb['data'].groupby(['cluster', 'ratio_type']).ratio.median().unstack().dropna(axis=1))
| docs/x_to_a_summary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Create Training Job (Hyperparamter Injection)
#
# In this notebook, we discuss more complicated set ups for `CreateTrainingJob` API. It assumes you are confortable with the set ups discussed in [the notebook on basics of `CreateTrainingJob`](https://github.com/hsl89/amazon-sagemaker-examples/blob/sagemaker-fundamentals/sagemaker-fundamentals/create-training-job/create_training_job.ipynb)
#
#
# ## What is Hyperparameter Injection?
#
# With hyperparamter injection, you don't need to hard code hyperparameters of your ML training in the training image, instead you can pass your hyperparamters through `CreateTrainingJob` API and SageMaker will makes them available to your training container. This way you can experiment a list of hyperparameters for your training job without rebuilding the image for each experiment. More importantly, this is the mechanism used by `CreateHyperParameterTuningJob` API to (you guessed right) create many training jobs to search for the best hyperparameters. We will discuss `CreateHyperParameterTuningJob` in a different notebook.
#
# If you remember from [the notebook on basics of `CreateTrainingJob`](https://github.com/hsl89/amazon-sagemaker-examples/blob/sagemaker-fundamentals/sagemaker-fundamentals/create-training-job/create_training_job.ipynb), SageMaker reserves `/opt/ml` directory "to talk to your container", i.e. provide training information to your training job and retrieve output from it.
#
# You will pass hyperparamters of your training job as a dictionary to the `create_training_job` of boto3 SageMaker client, and it will become availble in `/opt/ml/input/config/hyperparameters.json`. See [reference in the official docs](https://docs.aws.amazon.com/sagemaker/latest/dg/your-algorithms-training-algo-running-container.html).
# ### Set ups
#
# You will build a training image and push it to ECR like in [the notebook on basics of `CreateTrainingJob`](https://github.com/hsl89/amazon-sagemaker-examples/blob/sagemaker-fundamentals/sagemaker-fundamentals/create-training-job/create_training_job.ipynb). The only difference is, the python script for runing the training will print out the hyperparamters in `/opt/ml/input/config/hyperparameters.json` to confirm that container does have access to the hyperparamters you passed to `CreateTrainingJob` API.
#
# This training job does not require any data. Therefore, you don't need to confgure `InputDataConfig` parameter for `CreateTrainingJob`. However, SageMaker always needs an S3 URI to save your model artifact, i.e. you still need to configure `OutputDataConfig` parameter.
# +
import boto3 # your gateway to AWS APIs
import datetime
import pprint
import os
import time
import re
pp = pprint.PrettyPrinter(indent=1)
iam = boto3.client('iam')
# +
# some helper functions
def current_time():
ct = datetime.datetime.now()
return str(ct.now()).replace(":", "-").replace(" ", "-")[:19]
def account_id():
return boto3.client('sts').get_caller_identity()['Account']
# -
# ### Set up a service role for SageMaker
#
# Review [notebook on execution role](https://github.com/hsl89/amazon-sagemaker-examples/blob/execution-role/sagemaker-fundamentals/execution-role/execution-role.ipynb) for step-by-step instructions on how to create an IAM Role.
#
# The service role is intended to be assumed by the SageMaker service to procure resources in your AWS account on your behalf.
#
# 1. If you are running this this notebook on SageMaker infrastructure like Notebook Instances or Studio, then we will use the role you used to spin up those resources
#
# 2. If you are running this notebook on an EC2 instance, then we will create a service role attach `AmazonSageMakerFullAccess` to it. If you already have a SageMaker service role, you can paste its `role_arn` here.
#
# First, let's get some helper functions for creating execution role. We discussed those functions in the [notebook on execution role](https://github.com/hsl89/amazon-sagemaker-examples/blob/execution-role/sagemaker-fundamentals/execution-role/execution-role.ipynb).
# + language="bash"
# cp ../execution-role/iam_helpers.py .
# +
# set up service role for SageMaker
from iam_helpers import create_execution_role
sts = boto3.client('sts')
caller = sts.get_caller_identity()
if ':user/' in caller['Arn']: # as IAM user
# either paste in a role_arn with or create a new one and attach
# AmazonSageMakerFullAccess
role_name = 'sm'
role_arn = create_execution_role(role_name=role_name)['Role']['Arn']
# attach the permission to the role
# skip it if you want to use a SageMaker service that
# already has AmazonFullSageMakerFullAccess
iam.attach_role_policy(
RoleName=role_name,
PolicyArn='arn:aws:iam::aws:policy/AmazonSageMakerFullAccess'
)
elif 'assumed-role' in caller['Arn']: # on SageMaker infra
assumed_role = caller['Arn']
role_arn = re.sub(r"^(.+)sts::(\d+):assumed-role/(.+?)/.*$", r"\1iam::\2:role/\3", assumed_role)
else:
print("I assume you are on an EC2 instance launched with an IAM role")
role_arn = caller['Arn']
# -
# ## Build a training image and push to ECR
#
# You will build a training image here like in [the notebook on basics of `CreateTrainingJob`](https://github.com/hsl89/amazon-sagemaker-examples/blob/sagemaker-fundamentals/sagemaker-fundamentals/create-training-job/create_training_job.ipynb)
# View the Dockerfile
# !cat container_hyperparameter_injection/Dockerfile
# View the "training alogrithm"
# !pygmentize container_hyperparameter_injection/train.py
# The algorithm simply print out hyperparameters in the json file `/opt/ml/input/config/hyperparameters.json` as a verification that it can indeed access those hyperparamters
# + language="sh"
# # build the image
# cd container_hyperparameter_injection/
#
# # tag it as example-image:latest
# docker build -t example-image:latest .
# -
# ## Test the container locally
# Before pushing the image to ECR, it is always a good practice to test it locally. You need to create a `hyperparameters.json` file and make it available to the container at `/opt/ml/input/config/hyperparameters.json`. To do so, you can mount a local directory to `/opt/ml` as a docker volume like in [the notebook on basics of `CreateTrainingJob`](https://github.com/hsl89/amazon-sagemaker-examples/blob/sagemaker-fundamentals/sagemaker-fundamentals/create-training-job/create_training_job.ipynb).
# Checkout the test we provide:
# !pygmentize container_hyperparameter_injection/local_test/test_container.py
# We made some realistic looking hyperparameters in `container_hyperparameter_injection/local_test/ml/input/config/hyperparameters.json` and mounted `container_hyperparamter_injection/local_test/ml` to `/opt/ml` as a docker volume to the container, so that the file container can access the hyperparamters at `/opt/ml/input/config/hyperparameters.json`.
#
# Note that the json file `container_hyperparameter_injection/local_test/ml/input/config/hyperparameters.json` is not nested and the values are all strings, even they meant to be other data types. This is because when calling `CreateTrainingJob` with hyperparameter injection, the hyperparameters can only be a dictionary of key-value pairs, and both key and value need to be a string. See [API reference](https://docs.aws.amazon.com/sagemaker/latest/APIReference/API_CreateTrainingJob.html).
# run the test
# !python container_hyperparameter_injection/local_test/test_container.py
# Now you tested your container, you can push it to ECR and be confident that it will work for a SageMaker training job.
# +
# create a repo in ECR called example-image
ecr = boto3.client('ecr')
try:
# The repository might already exist
# in your ECR
cr_res = ecr.create_repository(
repositoryName='example-image')
pp.pprint(cr_res)
except Exception as e:
print(e)
# + language="bash"
# account=$(aws sts get-caller-identity --query Account | sed -e 's/^"//' -e 's/"$//')
# region=$(aws configure get region)
# ecr_account=${account}.dkr.ecr.${region}.amazonaws.com
#
# # Give docker your ECR login password
# aws ecr get-login-password --region $region | docker login --username AWS --password-stdin $ecr_account
#
# # Fullname of the repo
# fullname=$ecr_account/example-image:latest
#
# #echo $fullname
# # Tag the image with the fullname
# docker tag example-image:latest $fullname
#
# # Push to ECR
# docker push $fullname
# + jupyter={"outputs_hidden": true}
# Inspect the ECR repository
repo_res = ecr.describe_images(
repositoryName='example-image')
pp.pprint(repo_res)
# -
# ## Prepare an S3 bucket for model artifact
# Even you are not training a real model, SageMaker still requires you to give it an S3 URI to upload model artifact in `/opt/ml/model`. So let's create a temporary bucket for this.
# +
# create a bucket
def create_tmp_bucket():
"""Create an S3 bucket that is intended to be used for short term"""
bucket = f"sagemaker-{current_time()}" # accessible by SageMaker
region = boto3.Session().region_name
boto3.client('s3').create_bucket(
Bucket=bucket,
CreateBucketConfiguration={
'LocationConstraint': region
})
return bucket
bucket = create_tmp_bucket()
# -
# ## Put everything together
#
# Now you have everything you need to create a training job that can ingest hyperparamters from the boto3 call. Let's review what you have done. You have
# * created an execution role for SageMaker service
# * built and tested a docker image that includes the runtime and logic of your model training
# * made the image accessible to SageMaker by hosting it on ECR
# * created an S3 bucket for saving model artifact
# +
# set up
import json
sm_boto3 = boto3.client('sagemaker')
# name training job
training_job_name = 'example-training-job-{}'.format(current_time())
# location that SageMaker saves the model artifacts
output_prefix = 'example/output/'
output_path = "s3://" + bucket + '/' + output_prefix
# ECR URI of your image
region = boto3.Session().region_name
account = account_id()
image_uri = "{}.dkr.ecr.{}.amazonaws.com/example-image:latest".format(account, region)
algorithm_specification = {
'TrainingImage': image_uri,
'TrainingInputMode': 'File',
}
# inject the following hyperparamters to your container
# you can define `hyperparameters` in whatever way
# you want as long as it can be parsed to a json file (not nested)
# and both key and value are strings
hyperparamters = {
"num_trees" : "15",
"max_depth" : "4",
"n_iter": "30",
"your_parameter_1": "1",
"your_parameter_2" : "0.01"
}
output_data_config = {
'S3OutputPath': output_path
}
resource_config = {
'InstanceType': 'ml.m5.large',
'InstanceCount':1,
'VolumeSizeInGB':10
}
stopping_condition={
'MaxRuntimeInSeconds':120,
}
enable_network_isolation=False
# -
ct_res = sm_boto3.create_training_job(
TrainingJobName=training_job_name,
AlgorithmSpecification=algorithm_specification,
HyperParameters=hyperparameters, # look here
RoleArn=role_arn,
OutputDataConfig=output_data_config,
ResourceConfig=resource_config,
StoppingCondition=stopping_condition,
EnableNetworkIsolation=enable_network_isolation,
EnableManagedSpotTraining=False,
)
# +
# check training job status every 30 seconds
stopped = False
while not stopped:
tj_state = sm_boto3.describe_training_job(
TrainingJobName=training_job_name)
if tj_state['TrainingJobStatus'] in ['Completed', 'Stopped', 'Failed']:
stopped=True
else:
print("Training in progress")
time.sleep(30)
if tj_state['TrainingJobStatus'] == 'Failed':
print("Training job failed ")
print("Failed Reason: {}".tj_state['FailedReason'])
else:
print("Training job completed")
# -
# ## Inspect the trained model artifact
# +
print("== Output config:")
print(tj_state['OutputDataConfig'])
print()
s3 = boto3.client('s3')
print("== Model artifact:")
pp.pprint(s3.list_objects_v2(Bucket=bucket, Prefix=output_prefix))
# +
# print out logs from Cloud Watch
logs = boto3.client('logs')
log_res= logs.describe_log_streams(
logGroupName='/aws/sagemaker/TrainingJobs',
logStreamNamePrefix=training_job_name)
for log_stream in log_res['logStreams']:
# get one log event
log_event = logs.get_log_events(
logGroupName='/aws/sagemaker/TrainingJobs',
logStreamName=log_stream['logStreamName'])
# print out messages from the log event
for ev in log_event['events']:
for k, v in ev.items():
if k == 'message':
print(v)
# -
# ## Conclusion
#
# Congratulations! You now understand how to avoid hard-code hyperparamters in your training image. To recap,
#
# - Hyperparamter injection allows you to quickly experiment your ML algorithm with different hyperparameters
# - When calling `CreateTrainingJob` with hyperparamter injection, the hyperparameters you passed to `HyperParameter` needs to be a dictionary of string : string
# - To avoid hating yourself, always test your container before pushing it to ECR
# ## Clean up resources
# delete the ECR repo
del_repo_res = ecr.delete_repository(
repositoryName='example-image',
force=True)
pp.pprint(del_repo_res)
# +
# delete the S3 bucket
def delete_bucket_force(bucket_name):
objs = s3.list_objects_v2(Bucket=bucket_name)['Contents']
for obj in objs:
s3.delete_object(
Bucket=bucket_name,
Key=obj['Key'])
return s3.delete_bucket(Bucket=bucket_name)
del_buc_res = delete_bucket_force(bucket)
pp.pprint(del_buc_res)
| sagemaker-fundamentals/create-training-job/create_training_job_hyperparameter_injection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Advanced Lane Finding Project
#
# The goals / steps of this project are the following:
#
# * Compute the camera calibration matrix and distortion coefficients given a set of chessboard images.
# * Apply a distortion correction to raw images.
# * Use color transforms, gradients, etc., to create a thresholded binary image.
# * Apply a perspective transform to rectify binary image ("birds-eye view").
# * Detect lane pixels and fit to find the lane boundary.
# * Determine the curvature of the lane and vehicle position with respect to center.
# * Warp the detected lane boundaries back onto the original image.
# * Output visual display of the lane boundaries and numerical estimation of lane curvature and vehicle position.
#
# ---
# ## First, I'll compute the camera calibration using chessboard images
# +
import numpy as np
import cv2
import glob
import matplotlib.pyplot as plt
def compute_points_for_calibration(images):
# termination criteria
criteria = (cv2.TERM_CRITERIA_EPS + cv2.TERM_CRITERIA_MAX_ITER, 30, 0.001)
# prepare object points, like (0,0,0), (1,0,0), (2,0,0) ....,(6,5,0)
objp = np.zeros((6*9,3), np.float32)
objp[:,:2] = np.mgrid[0:9,0:6].T.reshape(-1,2)
# Arrays to store object points and image points from all the images.
objpoints = [] # 3d points in real world space
imgpoints = [] # 2d points in image plane.
# Step through the list and search for chessboard corners
for fname in images:
img = cv2.imread(fname)
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
# Find the chessboard corners
ret, corners = cv2.findChessboardCorners(gray, (9,6), None)
# If found, add object points, image points
if ret == True:
objpoints.append(objp)
corners2 = cv2.cornerSubPix(gray, corners, (11,11), (-1,-1), criteria)
imgpoints.append(corners2)
return objpoints, imgpoints
# -
# ## Apply a distortion correction to raw images.
# Define a function that takes an image, number of x and y points,
# camera matrix and distortion coefficients
def corners_unwarp(img, nx, ny, mtx, dist):
# Use the OpenCV undistort() function to remove distortion
undist = cv2.undistort(img, mtx, dist, None, mtx)
# Convert undistorted image to grayscale
gray = cv2.cvtColor(undist, cv2.COLOR_BGR2GRAY)
# Search for corners in the grayscaled image
ret, corners = cv2.findChessboardCorners(gray, (nx, ny), None)
warped = M = None
if ret == True:
# If we found corners, draw them! (just for fun)
# cv2.drawChessboardCorners(undist, (nx, ny), corners, ret)
# Choose offset from image corners to plot detected corners
# This should be chosen to present the result at the proper aspect ratio
# My choice of 100 pixels is not exact, but close enough for our purpose here
offset = 100 # offset for dst points
# Grab the image shape
img_size = (gray.shape[1], gray.shape[0])
# For source points I'm grabbing the outer four detected corners
src = np.float32([corners[0], corners[nx-1], corners[-1], corners[-nx]])
# For destination points, I'm arbitrarily choosing some points to be
# a nice fit for displaying our warped result
# again, not exact, but close enough for our purposes
dst = np.float32([[offset, offset], [img_size[0]-offset, offset],
[img_size[0]-offset, img_size[1]-offset],
[offset, img_size[1]-offset]])
# Given src and dst points, calculate the perspective transform matrix
M = cv2.getPerspectiveTransform(src, dst)
# Warp the image using OpenCV warpPerspective()
warped = cv2.warpPerspective(undist, M, img_size)
# Return the resulting image and matrix
return warped, M
# +
# Compute the camera calibration matrix and distortion coefficients
# Make a list of calibration images
images = glob.glob('./camera_cal/calibration*.jpg')
image_shape = cv2.imread(images[0]).shape[1::-1]
objpoints, imgpoints = compute_points_for_calibration(images)
ret, mtx, dist, rvecs, tvecs = cv2.calibrateCamera(objpoints, imgpoints, image_shape, None, None)
if not ret:
print('Error')
# -
# ### Calibration Test
# +
import matplotlib.image as mpimg
# %matplotlib inline
nx, ny = 9, 6
cal_test_img = cv2.imread('./camera_cal/calibration17.jpg')
cal_warped, cal_M = corners_unwarp(cal_test_img, nx, ny, mtx, dist)
output_image_path = './output_images/cal_test.png'
cv2.imwrite(output_image_path, cal_warped)
plt.figure(figsize=(12,12))
plt.subplot(1,2,1)
plt.imshow(cal_test_img)
plt.subplot(1,2,2)
plt.imshow(mpimg.imread(output_image_path))
# +
# straight
birds_eye_test_image = mpimg.imread('./test_images/straight_lines1.jpg')
undist_test = cv2.undistort(birds_eye_test_image, mtx, dist, None, mtx)
output_image_path = './output_images/undist_test.jpg'
cv2.imwrite(output_image_path, cv2.cvtColor(undist_test, cv2.COLOR_BGR2RGB))
plt.imshow(mpimg.imread(output_image_path))
# -
# ## Use color transforms, gradients, etc., to create a thresholded binary image.
def color_pipeline(img, s_thresh=(170, 255), sx_thresh=(20, 100)):
img = np.copy(img)
# Convert to HLS color space and separate the V channel
hls = cv2.cvtColor(img, cv2.COLOR_RGB2HLS)
#l_channel = hls[:,:,1]
s_channel = hls[:,:,2]
# Sobel x
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# Appy CLAHE algorithm to equalize contrast
# clahe = cv2.createCLAHE(clipLimit=4.0, tileGridSize=(8,8))
# gray = clahe.apply(gray)
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0) # Take the derivative in x
abs_sobelx = np.absolute(sobelx) # Absolute x derivative to accentuate lines away from horizontal
scaled_sobel = np.uint8(255*abs_sobelx/np.max(abs_sobelx))
# Threshold x gradient
sxbinary = np.zeros_like(scaled_sobel)
sxbinary[(scaled_sobel >= sx_thresh[0]) & (scaled_sobel <= sx_thresh[1])] = 1
# Threshold color channel
s_binary = np.zeros_like(s_channel)
s_binary[(s_channel >= s_thresh[0]) & (s_channel <= s_thresh[1])] = 1
# Stack each channel
color_binary = np.dstack(( np.zeros_like(sxbinary), sxbinary, s_binary)) * 255
# Combine the two binary thresholds
combined_binary = np.zeros_like(sxbinary)
combined_binary[(s_binary == 1) | (sxbinary == 1)] = 1
return combined_binary
# ### Color Pipeline test
color_test_image = mpimg.imread('./test_images/test4.jpg')
undist_test = cv2.undistort(color_test_image, mtx, dist, None, mtx)
color_test_result = color_pipeline(undist_test)
plt.imshow(color_test_result, cmap='gray')
# ## Apply a perspective transform to rectify binary image ("birds-eye view").
# +
def get_image_src_dst(img):
h, w = img.shape[0], img.shape[1]
src = np.array([[w*0.445, h*0.627], [0, h], [w, h], [w*0.555, h*0.629]], dtype=np.float32)
dst = np.array([[0, 0], [0, h], [w, h], [w, 0]], dtype=np.float32)
return src, dst
def get_birds_eye_matrix(img):
src, dst = get_image_src_dst(img)
return cv2.getPerspectiveTransform(src, dst)
def get_birds_eye_inverse_matrix(img):
src, dst = get_image_src_dst(img)
return cv2.getPerspectiveTransform(dst, src)
def warper(img, src, dst):
# Given src and dst points, calculate the perspective transform matrix
M = get_birds_eye_matrix(img)
# Warp the image using OpenCV warpPerspective()
h, w = img.shape[0], img.shape[1]
warped = cv2.warpPerspective(img, M, (w, h))
return warped
def birds_eye_view(img):
src, dst = get_image_src_dst(img)
return warper(img, src, dst)
# +
# straight
birds_eye_test_image = mpimg.imread('./test_images/straight_lines1.jpg')
undist_test = cv2.undistort(birds_eye_test_image, mtx, dist, None, mtx)
birds_eye_test_result = color_pipeline(undist_test)
birds_eye_trans_img = birds_eye_view(birds_eye_test_result)
output_image_path = './output_images/birds_eye_test.jpg'
cv2.imwrite(output_image_path, birds_eye_trans_img*255)
plt.figure(figsize=(12,12))
plt.subplot(1,2,1)
plt.imshow(undist_test)
plt.subplot(1,2,2)
plt.imshow(mpimg.imread(output_image_path))
plt.imshow(mpimg.imread(output_image_path))
# +
# curve
birds_eye_test_curve = mpimg.imread('./test_images/test5.jpg')
undist_test = cv2.undistort(birds_eye_test_curve, mtx, dist, None, mtx)
birds_eye_test_curve_result = color_pipeline(undist_test)
birds_eye_trans_curve = birds_eye_view(birds_eye_test_curve_result)
output_image_path = './output_images/birds_eye_curve.jpg'
cv2.imwrite(output_image_path, birds_eye_trans_curve*255)
plt.imshow(mpimg.imread(output_image_path))
# -
# ## Detect lane pixels and fit to find the lane boundary.
# +
def fit_poly(img_shape, leftx, lefty, rightx, righty):
left_fit = np.polyfit(lefty, leftx, 2)
right_fit = np.polyfit(righty, rightx, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, img_shape[0]-1, img_shape[0] )
try:
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
except TypeError:
# Avoids an error if `left` and `right_fit` are still none or incorrect
print('The function failed to fit a line!')
left_fitx = 1*ploty**2 + 1*ploty
right_fitx = 1*ploty**2 + 1*ploty
return left_fitx, right_fitx, ploty, left_fit, right_fit
def search_around_poly(binary_warped, left_fit, right_fit):
# HYPERPARAMETER
# Choose the width of the margin around the previous polynomial to search
# The quiz grader expects 100 here, but feel free to tune on your own!
margin = 50
# Grab activated pixels
nonzero = binary_warped.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
left_fit_x = left_fit[0]*(nonzeroy**2) + left_fit[1]*nonzeroy + left_fit[2]
left_lane_inds = ((nonzerox > left_fit_x - margin)
& (nonzerox < left_fit_x + margin))
right_fix_x = right_fit[0]*(nonzeroy**2) + right_fit[1]*nonzeroy + right_fit[2]
right_lane_inds = ((nonzerox > right_fix_x - margin)
& (nonzerox < right_fix_x + margin))
# Again, extract left and right line pixel positions
leftx = nonzerox[left_lane_inds]
lefty = nonzeroy[left_lane_inds]
rightx = nonzerox[right_lane_inds]
righty = nonzeroy[right_lane_inds]
return leftx, lefty, rightx, righty,
# +
def find_lane_pixels(binary_warped, visualize=False):
# Take a histogram of the bottom half of the image
histogram = np.sum(binary_warped[binary_warped.shape[0]//2:,:], axis=0)
# Create an output image to draw on and visualize the result
out_img = np.dstack((binary_warped, binary_warped, binary_warped))
# Find the peak of the left and right halves of the histogram
# These will be the starting point for the left and right lines
midpoint = np.int(histogram.shape[0]//2)
leftx_base = np.argmax(histogram[:midpoint])
rightx_base = np.argmax(histogram[midpoint:]) + midpoint
# HYPERPARAMETERS
# Choose the number of sliding windows
nwindows = 9
# Set the width of the windows +/- margin
margin = 100
# Set minimum number of pixels found to recenter window
minpix = 50
# Set height of windows - based on nwindows above and image shape
window_height = np.int(binary_warped.shape[0]//nwindows)
# Identify the x and y positions of all nonzero pixels in the image
nonzero = binary_warped.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
# Current positions to be updated later for each window in nwindows
leftx_current = leftx_base
rightx_current = rightx_base
# Create empty lists to receive left and right lane pixel indices
left_lane_inds = []
right_lane_inds = []
# Step through the windows one by one
for window in range(nwindows):
# Identify window boundaries in x and y (and right and left)
win_y_low = binary_warped.shape[0] - (window+1)*window_height
win_y_high = binary_warped.shape[0] - window*window_height
win_xleft_low = leftx_current - margin
win_xleft_high = leftx_current + margin
win_xright_low = rightx_current - margin
win_xright_high = rightx_current + margin
# Draw the windows on the visualization image
if (visualize):
cv2.rectangle(out_img,(win_xleft_low,win_y_low), (win_xleft_high,win_y_high),(0,255,0), 2)
cv2.rectangle(out_img,(win_xright_low,win_y_low), (win_xright_high,win_y_high),(0,255,0), 2)
# Identify the nonzero pixels in x and y within the window
good_left_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_xleft_low) & (nonzerox < win_xleft_high)).nonzero()[0]
good_right_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_xright_low) & (nonzerox < win_xright_high)).nonzero()[0]
# Append these indices to the lists
left_lane_inds.append(good_left_inds)
right_lane_inds.append(good_right_inds)
# If you found > minpix pixels, recenter next window on their mean position
if len(good_left_inds) > minpix:
leftx_current = np.int(np.mean(nonzerox[good_left_inds]))
if len(good_right_inds) > minpix:
rightx_current = np.int(np.mean(nonzerox[good_right_inds]))
# Concatenate the arrays of indices (previously was a list of lists of pixels)
try:
left_lane_inds = np.concatenate(left_lane_inds)
right_lane_inds = np.concatenate(right_lane_inds)
except ValueError:
# Avoids an error if the above is not implemented fully
pass
# Extract left and right line pixel positions
leftx = nonzerox[left_lane_inds]
lefty = nonzeroy[left_lane_inds]
rightx = nonzerox[right_lane_inds]
righty = nonzeroy[right_lane_inds]
return leftx, lefty, rightx, righty, out_img
def fit_polynomial(binary_warped, visualize=False):
# Find our lane pixels first
leftx, lefty, rightx, righty, out_img = find_lane_pixels(binary_warped, visualize)
left_fitx, right_fitx, ploty, _, _ = fit_poly(binary_warped.shape, leftx, lefty, rightx, righty)
if (not visualize):
return out_img
## Visualization ##
# Colors in the left and right lane regions
out_img[lefty, leftx] = [255, 0, 0]
out_img[righty, rightx] = [0, 0, 255]
# Plots the left and right polynomials on the lane lines
plt.plot(left_fitx, ploty, color='yellow')
plt.plot(right_fitx, ploty, color='yellow')
return out_img
# -
# ### Find lanes test
# +
# straight
# Load our image - this should be a new frame since last time!
binary_warped = mpimg.imread('./output_images/birds_eye_test.jpg')
# Polynomial fit values from the previous frame
result_ = fit_polynomial(binary_warped, True)
# View your output
cv2.imwrite('./output_images/find_lane_test_straight.jpg', cv2.cvtColor(result_, cv2.COLOR_BGR2RGB))
plt.imshow(result_)
# +
# curve
# Load our image - this should be a new frame since last time!
binary_warped = mpimg.imread('./output_images/birds_eye_curve.jpg')
# Polynomial fit values from the previous frame
# Make sure to grab the actual values from the previous step in your project!
#left_fit = np.array([ 2.13935315e-04, -3.77507980e-01, 4.76902175e+02])
#right_fit = np.array([4.17622148e-04, -4.93848953e-01, 1.11806170e+03])
result_ = fit_polynomial(binary_warped, True)
#result_ = search_around_poly(binary_warped//255)
# View your output
cv2.imwrite('./output_images/find_lane_test_curve.jpg', cv2.cvtColor(result_, cv2.COLOR_BGR2RGB))
plt.imshow(result_)
# -
# ## Determine the curvature of the lane and vehicle position with respect to center.
# +
# Define conversions in x and y from pixels space to meters
ym_per_pix = 30/720 # meters per pixel in y dimension
xm_per_pix = 3.7/700 # meters per pixel in x dimension
def measure_curvature_real(left_fit_cr, right_fit_cr, ploty):
'''
Calculates the curvature of polynomial functions in meters.
'''
# Define y-value where we want radius of curvature
# We'll choose the maximum y-value, corresponding to the bottom of the image
y_eval = np.max(ploty)
# Calculation of R_curve (radius of curvature)
left_curverad = ((1 + (2*left_fit_cr[0]*y_eval*ym_per_pix + left_fit_cr[1])**2)**1.5) / np.absolute(2*left_fit_cr[0])
right_curverad = ((1 + (2*right_fit_cr[0]*y_eval*ym_per_pix + right_fit_cr[1])**2)**1.5) / np.absolute(2*right_fit_cr[0])
return left_curverad, right_curverad
def vehicle_position_real(img, left_fit_cr, right_fit_cr, ploty):
y_eval = np.max(ploty) * ym_per_pix
left_base = left_fit_cr[0] * y_eval**2 + left_fit_cr[1] * y_eval + left_fit_cr[2]
right_base = right_fit_cr[0] * y_eval**2 + right_fit_cr[1] * y_eval + right_fit_cr[2]
center_cr = xm_per_pix * img.shape[1] / 2
return center_cr - (left_base + right_base) / 2
# -
# ### Curvature Test
binary_warped = mpimg.imread('./output_images/birds_eye_curve.jpg')
leftx, lefty, rightx, righty, out_img = find_lane_pixels(binary_warped, True)
plt.imshow(out_img)
# Define conversions in x and y from pixels space to meters
left_fit_cr = np.polyfit(lefty * ym_per_pix, leftx * xm_per_pix, 2)
right_fit_cr = np.polyfit(righty * ym_per_pix, rightx * xm_per_pix, 2)
ploty = np.linspace(0, binary_warped.shape[0]-1, binary_warped.shape[0])
left_curverad, right_curverad = measure_curvature_real(left_fit_cr, right_fit_cr, ploty)
print(left_curverad, 'm', right_curverad, 'm')
# ## Warp the detected lane boundaries back onto the original image.
def back_projection(undist, warped, left_fitx, right_fitx, ploty, Minv):
# Create an image to draw the lines on
warp_zero = np.zeros_like(warped).astype(np.uint8)
color_warp = np.dstack((warp_zero, warp_zero, warp_zero))
# Recast the x and y points into usable format for cv2.fillPoly()
pts_left = np.array([np.transpose(np.vstack([left_fitx, ploty]))])
pts_right = np.array([np.flipud(np.transpose(np.vstack([right_fitx, ploty])))])
pts = np.hstack((pts_left, pts_right))
# Draw the lane onto the warped blank image
cv2.fillPoly(color_warp, np.int_([pts]), (0,255, 0))
# Warp the blank back to original image space using inverse perspective matrix (Minv)
newwarp = cv2.warpPerspective(color_warp, Minv, (undist.shape[1], undist.shape[0]))
# Combine the result with the original image
return cv2.addWeighted(undist, 1, newwarp, 0.3, 0)
# ### Test
# +
# birds_eye_test_curve = mpimg.imread('./test_images/test5.jpg')
birds_eye_test_curve = mpimg.imread('./test_images/test5.jpg')
undist_test = cv2.undistort(birds_eye_test_curve, mtx, dist, None, mtx)
binary_warped = mpimg.imread('./output_images/birds_eye_curve.jpg')
leftx, lefty, rightx, righty, out_img = find_lane_pixels(binary_warped)
left_fitx, right_fitx, ploty, _, _ = fit_poly(binary_warped.shape, leftx, lefty, rightx, righty)
Minv = get_birds_eye_inverse_matrix(binary_warped)
back_image = back_projection(undist_test, binary_warped, left_fitx, right_fitx, ploty, Minv)
plt.imshow(back_image)
# vehicle position
left_fit_cr = np.polyfit(lefty * ym_per_pix, leftx * xm_per_pix, 2)
right_fit_cr = np.polyfit(righty * ym_per_pix, rightx * xm_per_pix, 2)
ploty = np.linspace(0, binary_warped.shape[0]-1, binary_warped.shape[0])
cv2.imwrite('./output_images/warp_test.jpg', cv2.cvtColor(back_image, cv2.COLOR_BGR2RGB))
print(vehicle_position_real(back_image, left_fit_cr, right_fit_cr, ploty))
# -
# ## Output visual display of the lane boundaries and numerical estimation of lane curvature and vehicle position.
# Import everything needed to edit/save/watch video clips
from moviepy.editor import VideoFileClip
from IPython.display import HTML
# +
from collections import deque
recent_left_fitx = deque([], 20)
recent_right_fitx = deque([], 20)
recent_radius = deque([], 20)
def init_pipeline():
global recent_left_fitx
global recent_right_fitx
global recent_radius
recent_left_fitx = deque([], 20)
recent_right_fitx = deque([], 20)
recent_radius = deque([], 20)
def pipeline(img):
# Apply a distortion correction to raw images.
# Make sure mtx and dist are calculated
undist = cv2.undistort(img, mtx, dist, None, mtx)
# Use color transforms, gradients, etc., to create a thresholded binary image.
color_result = color_pipeline(undist)
# Apply a perspective transform to rectify binary image ("birds-eye view").
birds_eye = birds_eye_view(color_result)
# Detect lane pixels and fit to find the lane boundary.
leftx, lefty, rightx, righty, _ = find_lane_pixels(birds_eye, False)
left_fitx, right_fitx, ploty, left_fit, right_fit = fit_poly(birds_eye.shape, leftx, lefty, rightx, righty)
left_fit_cr = np.polyfit(lefty * ym_per_pix, leftx * xm_per_pix, 2)
right_fit_cr = np.polyfit(righty * ym_per_pix, rightx * xm_per_pix, 2)
# Determine the curvature of the lane and vehicle position with respect to center.
left_curverad, right_curverad = measure_curvature_real(left_fit_cr, right_fit_cr, ploty)
# Smoothing
recent_left_fitx.append(left_fitx)
recent_right_fitx.append(right_fitx)
recent_radius.append((left_curverad + right_curverad)//2)
avg_left_fitx = np.mean(recent_left_fitx, axis=0)
avg_right_fitx = np.mean(recent_right_fitx, axis=0)
avg_radius = np.mean(recent_radius, axis=0)
# Warp the detected lane boundaries back onto the original image.
Minv = get_birds_eye_inverse_matrix(undist)
back_image = back_projection(undist, birds_eye, avg_left_fitx, avg_right_fitx, ploty, Minv)
# Output visual display of the lane boundaries and numerical estimation of lane curvature and vehicle position.
curv_text = 'Radius of Curvature = {}(m)'.format(int(avg_radius))
vehicle_pos = round(vehicle_position_real(back_image, left_fit_cr, right_fit_cr, ploty), 2)
if vehicle_pos >= 0:
vehicle_text = 'Vehicle is {}m left of center'.format(vehicle_pos)
else:
vehicle_text = 'Vehicle is {}m right of center'.format(-vehicle_pos)
cv2.putText(back_image, curv_text, (0, 50), cv2.FONT_HERSHEY_PLAIN, 4, (255, 255, 255), 5, cv2.LINE_AA)
cv2.putText(back_image, vehicle_text, (0, 100), cv2.FONT_HERSHEY_PLAIN, 4, (255, 255, 255), 5, cv2.LINE_AA)
return back_image
# -
# ### Pipeline Test
pipeline_test = mpimg.imread('./test_images/test5.jpg')
init_pipeline()
result_ = pipeline(pipeline_test)
cv2.imwrite('./output_images/pipeline_test.jpg', cv2.cvtColor(result_, cv2.COLOR_BGR2RGB))
plt.imshow(result_)
# ### project_video
init_pipeline()
project_video = VideoFileClip("./project_video.mp4")
project_clip = project_video.fl_image(pipeline)
project_output = './output_images/project_video_output.mp4'
# %time project_clip.write_videofile(project_output, audio=False, logger=None)
# ### challenge_video
init_pipeline()
challenge_video = VideoFileClip("./challenge_video.mp4")
challenge_clip = challenge_video.fl_image(pipeline)
challenge_output = './output_images/challenge_video_output.mp4'
# %time challenge_clip.write_videofile(challenge_output, audio=False, logger=None)
# ### harder_challenge_video
init_pipeline()
harder_video = VideoFileClip("./harder_challenge_video.mp4")
harder_clip = harder_video.fl_image(pipeline)
harder_output = './output_images/harder_video_output.mp4'
# %time harder_clip.write_videofile(harder_output, audio=False, logger=None)
| Lane_line.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# First set of Problems
1. True and True # True
2. False and True # False
3. 1 == 1 and 2 == 1 # True and False making it a False
4. "test" == "test" # True
5. 1 == 1 or 2 != 1 # True and True making it a True
6. True and 1 == 1 # True and True making it a True
7. False and 0 != 0 # False and False making it a False
8. True or 1 == 1 # True and True making it a True
9. "test" == "testing" # False
10. 1 != 0 and 2 == 1 # True and False making it a False
11. "test" != "testing" # True
12. "test" == 1 # False
13. not (True and False) # not False so True
14. not (1 == 1 and 0 != 1) # not (True and True) so False
15. not (10 == 1 or 1000 == 1000) # not (False and True) so True
16. not (1 != 10 or 3 == 4) # not (True and False) so True
17. not ("testing" == "testing" and "Zed" == "Cool Guy") # not (True and False) so True
18. 1 == 1 and (not ("testing" == 1 or 1 == 0))
# True and (not(False or False)) is True and True so True
19. "chunky" == "bacon" and (not (3 == 4 or 3 == 3))
# False and (not(False or True)) is False and False so False
20. 3 == 3 and (not ("testing" == "testing" or "Python" == "Fun"))
# True and (not(True or False)) is True and False so False
# -
| E28.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Function |
#1 it will print 5
def a():
return 5
print(a())
#2 it will print 10
def a():
return 5
print(a()+a())
#3 it will print 5
def a():
return 5
return 10
print(a())
#4 it will print 5
def a():
return 5
print(10)
print(a())
#5 it will print 5
def a():
print(5)
x = a()
print(x)
#6 it will print 3,5
def a(b,c):
print(b+c)
print(a(1,2) + a(2,3))
#7 it will print 25
def a(b,c):
return str(b)+str(c)
print(a(2,5))
#8 will print 10 and retrn 10
def a():
b = 100
print(b)
if b < 10:
return 5
else:
return 10
return 7
print(a())
#9 it will return 7,14,21
def a(b,c):
if b<c:
return 7
else:
return 14
return 3
print(a(2,3))
print(a(5,3))
print(a(2,3) + a(5,3))
#10 it will return 8
def a(b,c):
return b+c
return 10
print(a(3,5))
#11 it will print 500 ,500,300,500
b = 500
print(b)
def a():
b = 300
print(b)
print(b)
a()
print(b)
#12 it will print 500 ,500,300,500
b = 500
print(b)
def a():
b = 300
print(b)
return b
print(b)
a()
print(b)
#13 it will print 500,500,300,300
b = 500
print(b)
def a():
b = 300
print(b)
return b
print(b)
b=a()
print(b)
#14 it will print 1,2,3
def a():
print(1)
b()
print(2)
def b():
print(3)
a()
#15 it will return 1,3,5,10
def a():
print(1)
x = b()
print(x)
return 10
def b():
print(3)
return 5
y = a()
print(y)
| function_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from qiskit.optimization import QuadraticProgram
from docplex.mp.advmodel import AdvModel
from docplex.mp.model_reader import ModelReader
from qiskit.optimization.algorithms import MinimumEigenOptimizer
from dwave.plugins.qiskit import DWaveMinimumEigensolver
from dwave.system import DWaveCliqueSampler, LeapHybridSampler
import numpy as np
# -
# # Test binary instances from Burkardt
# ## taken from https://people.math.sc.edu/Burkardt/datasets/datasets.html
# # Test 10 bit binary with leap
model = ModelReader.read(filename='BURKARDT_DATA/BINARY/reid.lp',model_name = "Test_Bin_reid", model_class=AdvModel)
qp = QuadraticProgram()
qp.from_docplex(model)
dwave_solver_hybrid = DWaveMinimumEigensolver(sampler = LeapHybridSampler())
optimizer = MinimumEigenOptimizer(dwave_solver_hybrid)
result = optimizer.solve(qp) # Problem ID : 8f677f6a-7c23-481c-ad1e-e2887cc8758f
result
print(qp.export_as_lp_string())
# # Solve with DWaveCliqueSampler
# Use D-Wave QPU as a minimum eigen solver
# See https://github.com/dwavesystems/dwave-qiskit-plugin
num_reads = 1000
cliqueSampler = DWaveCliqueSampler()
dwave_solver_clique = DWaveMinimumEigensolver(sampler = cliqueSampler, num_reads = num_reads)
optimizer = MinimumEigenOptimizer(dwave_solver_clique)
result = optimizer.solve(qp) # Problem ID : 74108957-e013-40d5-a7ff-e88c9c552ab2
result
result.min_eigen_solver_result.sampleset.to_pandas_dataframe()
# # Test 20 bit binary
# ## Leap
model = ModelReader.read(filename='BURKARDT_DATA/BINARY/two_by_four.lp',model_name = "Test_Bin_16", model_class=AdvModel)
qp = QuadraticProgram()
qp.from_docplex(model)
optimizer_hybrid = MinimumEigenOptimizer(dwave_solver_hybrid)
result = optimizer_hybrid.solve(qp) # Problem ID : 1270f318-2655-4417-a2de-5daf6c4e017a
result
# # Solve with DWaveCliqueSampler
result = optimizer.solve(qp) # Problem ID : 93df81b0-9ca4-450e-a921-edf22bd6b5ee
result
result.min_eigen_solver_result.sampleset.to_pandas_dataframe()
model.solve()
| comparison/Ocean/LinearProgramming/DWave_Qiskit_Plugin_Test/External/Binary_LP_BURKARDT.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 1: Arrays and Strings
### Two Sum ###
def twoSum(nums, target):
"""
:type nums: List[int]
:type target: int
:rtype: List[int]
"""
d = {}
for i in range(len(nums)):
if target - nums[i] in d:
return (d[target - nums[i]], i)
d[nums[i]] = i
return None
nums = [2, 7, 11, 15]
target = 9
print(twoSum(nums, target))
### Is Unique ###
# Description: Implement an algorithm to determine if a string has all unique characters.
# What if you cannot use additional datastructures
def is_unique(s):
return len(s.upper()) == len(list(set(s.upper())))
tests = ["AAA", "ABCD", "dog", "BaAt", " ", ""]
for test in tests:
print(is_unique(test))
### Check Permuatations ###
# Description: Given two strings check if one sting is a permutation of the other.
# O(n log n)
def check_perm(s1, s2):
#if s1 or s2 == None:
#return False
return ''.join(sorted(s1.upper())) == ''.join(sorted(s2.upper()))
check_perm('abac', 'abca')
### URLify ###
# Description: Replace all spaces in a string with '%20'.
## Solution is valid but can be more space effecient and doesnt use all the provided information and not in place
# Missing input info
import queue
def urlify(s, count):
q = queue.Queue()
ret = ''
for i in s:
if i == ' ':
q.put('%20')
else:
q.put(i)
while not q.empty():
popped = q.get()
ret += popped
return ret
urlify('Mr <NAME>', 13)
## Palendrome Permutation ##
## Characteristics of palendrom:
# Max contains one letter of odd count
def palendrome_perm(s):
d = {}
flag = 0
for i in s.replace(' ',''):
if i not in d:
d[i] = 1
else:
d[i] += 1
for k in d:
if flag > 1:
return False
if d[k] % 2 != 0:
flag += 1
return True
palendrome_perm('carerac')
# One Away #
def one_away(s1,s2):
# Come back
# String Compression #
def string_compress(s):
d = {}
string = []
counter = 0
for i in range(len(s)):
if i > 0 and s[i] != s[i - 1]:
string.append(s[i - 1]+str(counter))
counter = 0
counter += 1
string.append(s[-1]+str(counter))
return min(s, ''.join(string), key=len)
string_compress("aabcccccaaa")
# Rotate Matrix #
def rotate_matrix(W):
ret = []
for j in range(len(W)):
r = []
for i in range(len(W), 0, -1):
r.append(W[i - 1][j])
ret.append(r)
return ret
rotate_matrix([
[1,2],
[3,4]
])
rotate_matrix([
[1,2,3],
[4,5,6],
[7,8,9]
])
| test_prep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/1995subhankar1995/AE-VAE-GAN-implementation/blob/master/VAE_Torch.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="-RbBcMcNXOJv" colab_type="code" colab={}
import torch
import torchvision
from torch import nn
from torch.utils.data import DataLoader
from torchvision import transforms
from torchvision.datasets import MNIST
from matplotlib import pyplot as plt
# + id="nMl-5KxfXSwe" colab_type="code" colab={}
# Displaying routine
def display_images(in_, out, n=1, label=None, count=False):
for N in range(n):
if in_ is not None:
in_pic = in_.data.cpu().view(-1, 28, 28)
plt.figure(figsize=(18, 4))
plt.suptitle(label + ' – real test data / reconstructions', color='w', fontsize=16)
for i in range(4):
plt.subplot(1,4,i+1)
plt.imshow(in_pic[i+4*N])
plt.axis('off')
out_pic = out.data.cpu().view(-1, 28, 28)
plt.figure(figsize=(18, 6))
for i in range(4):
plt.subplot(1,4,i+1)
plt.imshow(out_pic[i+4*N])
plt.axis('off')
if count: plt.title(str(4 * N + i), color='w')
# + id="w2STCgnwXZS1" colab_type="code" colab={}
# Set random seeds
torch.manual_seed(1)
torch.cuda.manual_seed(1)
# + id="51QqxA_MXctn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 411, "referenced_widgets": ["<KEY>", "<KEY>", "44fc0d498ee943628a8b674920d0e6f2", "<KEY>", "374e1dc3c0ca442ab6e0656caa02b125", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "05d8618aad534c5585e810affb075a3a", "2640ce861f3a45cbb11b53e6c3d19081", "4206d2b035a94fe597559906c210c06c", "1be56ffc8392407593b189565393a854", "ffec8956c55b4f9f861964f983edee6a", "<KEY>", "<KEY>", "df1aa2e087cf4966aba7de4ff8dbbb02", "eca1494e5dad4524b2a0bb9236e239db", "<KEY>", "331d0e096ac34e609d9a1162e30c1b24", "<KEY>", "fa79f4cc5d844e398ff73740a21d0fd4", "<KEY>", "4d2f6613023040df817690fe1196d9af", "<KEY>", "<KEY>", "90d800527549416f8a1c903a606eaff6", "<KEY>", "3340850d9731477ea6b1c6c998ad7095", "af7d855c9abd4e9b9796a161873716fd", "7948603a3a364012bf710d081a6390e6", "4726cbff5d1241fe8e43f685f6be0f15"]} outputId="48b00962-f7d4-457f-e306-e0072769d7bb"
# Define data loading step
batch_size = 256
kwargs = {'num_workers': 1, 'pin_memory': True}
train_loader = torch.utils.data.DataLoader(
MNIST('./data', train=True, download=True,
transform=transforms.ToTensor()),
batch_size=batch_size, shuffle=True, **kwargs)
test_loader = torch.utils.data.DataLoader(
MNIST('./data', train=False, transform=transforms.ToTensor()),
batch_size=batch_size, shuffle=True, **kwargs)
# + id="VmEmjP5cXh2F" colab_type="code" colab={}
# Defining the device
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# + id="EJ0BVV-wXld9" colab_type="code" colab={}
# Defining the model
d = 20
class VAE(nn.Module):
def __init__(self):
super().__init__()
self.encoder = nn.Sequential(
nn.Linear(784, d ** 2),
nn.ReLU(),
nn.Linear(d ** 2, d * 2)
)
self.decoder = nn.Sequential(
nn.Linear(d, d ** 2),
nn.ReLU(),
nn.Linear(d ** 2, 784),
nn.Sigmoid(),
)
def reparameterise(self, mu, logvar):
if self.training:
std = logvar.mul(0.5).exp_()
eps = std.data.new(std.size()).normal_()
return eps.mul(std).add_(mu)
else:
return mu
def forward(self, x):
mu_logvar = self.encoder(x.view(-1, 784)).view(-1, 2, d)
mu = mu_logvar[:, 0, :]
logvar = mu_logvar[:, 1, :]
z = self.reparameterise(mu, logvar)
return self.decoder(z), mu, logvar
model = VAE().to(device)
# + id="HKKisaUnNWWN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 246} outputId="04876e28-b6f6-4ab2-e07c-0e7af046dbe0"
print(model)
# + id="7xrgA-0cXuwt" colab_type="code" colab={}
# Setting the optimiser
learning_rate = 1e-3
optimizer = torch.optim.Adam(
model.parameters(),
lr=learning_rate,
)
# + id="pAM4VJhiX1I2" colab_type="code" colab={}
# Reconstruction + KL divergence losses summed over all elements and batch
def loss_function(x_hat, x, mu, logvar):
BCE = nn.functional.binary_cross_entropy(
x_hat, x.view(-1, 784), reduction='sum'
)
KLD = 0.5 * torch.sum(logvar.exp() - logvar - 1 + mu.pow(2))
return BCE + KLD
# + id="18dAWvTKX2ML" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="b5ddaeda-523f-4e19-b1b4-cef53fa3fd83"
# Training and testing the VAE
epochs = 20
codes = dict(μ=list(), logσ2=list(), y=list())
for epoch in range(0, epochs + 1):
# Training
if epoch > 0: # test untrained net first
model.train()
train_loss = 0
for x, _ in train_loader:
x = x.to(device)
# ===================forward=====================
x_hat, mu, logvar = model(x)
loss = loss_function(x_hat, x, mu, logvar)
train_loss += loss.item()
# ===================backward====================
optimizer.zero_grad()
loss.backward()
optimizer.step()
# ===================log========================
print(f'====> Epoch: {epoch} Average loss: {train_loss / len(train_loader.dataset):.4f}')
# Testing
means, logvars, labels = list(), list(), list()
with torch.no_grad():
model.eval()
test_loss = 0
for x, y in test_loader:
x = x.to(device)
# ===================forward=====================
x_hat, mu, logvar = model(x)
test_loss += loss_function(x_hat, x, mu, logvar).item()
# =====================log=======================
means.append(mu.detach())
logvars.append(logvar.detach())
labels.append(y.detach())
# ===================log========================
codes['μ'].append(torch.cat(means))
codes['logσ2'].append(torch.cat(logvars))
codes['y'].append(torch.cat(labels))
test_loss /= len(test_loader.dataset)
print(f'====> Test set loss: {test_loss:.4f}')
display_images(x, x_hat, 1, f'Epoch {epoch}')
# + id="_UZ7qaeuX-Pm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 896} outputId="950313f5-2d39-471d-bb49-be416324ffe3"
# Generating a few samples
N = 16
z = torch.randn((N, d)).to(device)
sample = model.decoder(z)
display_images(None, sample, N // 4, count=True)
# + id="NoKu7QQHYGHB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 896} outputId="1c251873-6cdc-469d-bd1b-be4253684aa0"
# Display last test batch
display_images(None, x, 4, count=True)
# + id="GURO2oKMYh7H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 223} outputId="420c4653-979d-4cc0-af66-266bb6db79b0"
# Choose starting and ending point for the interpolation -> shows original and reconstructed
A, B = 1, 14
sample = model.decoder(torch.stack((mu[A].data, mu[B].data), 0))
display_images(None, torch.stack(((
x[A].data.view(-1),
x[B].data.view(-1),
sample.data[0],
sample.data[1]
)), 0))
# + id="gfR6s4L8YqKw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 896} outputId="a3476e89-aa87-4190-cb25-f5a3cfbad81f"
# Perform an interpolation between input A and B, in N steps
N = 16
code = torch.Tensor(N, 20).to(device)
sample = torch.Tensor(N, 28, 28).to(device)
for i in range(N):
code[i] = i / (N - 1) * mu[B].data + (1 - i / (N - 1) ) * mu[A].data
# sample[i] = i / (N - 1) * x[B].data + (1 - i / (N - 1) ) * x[A].data
sample = model.decoder(code)
display_images(None, sample, N // 4, count=True)
| VAE_Torch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:skit]
# language: python
# name: python3
# ---
# +
import numpy as np
import os
import pandas as pd
df = pd.read_csv("tts_data.csv") #differece = Train - Gen
df
# -
df["Difference"].describe()
| Notebooks/pre-processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # Command Line
#
# This is a reference for all the `--help` message from all of OpenPifPaf's command line interfaces (CLIs).
# (cli-help-predict)=
# ## predict
# !python3 -m openpifpaf.predict --help
# (cli-help-video)=
# ## video
# !python3 -m openpifpaf.video --help
# ## train
# !python3 -m openpifpaf.train --help
# ## eval
# !python3 -m openpifpaf.eval --help
# ## export_onnx
# !python3 -m openpifpaf.export_onnx --help
# ## export_coreml
# !python3 -m openpifpaf.export_coreml --help
# ## benchmark
# !python3 -m openpifpaf.benchmark --help
# ## logs
# !python3 -m openpifpaf.logs --help
| guide/cli_help.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
all_aufgaben = []
solved_aufgaben = []
max_aufgaben = [4,7,7,6,7,3,5,5,2,3,3,6]
for i in range(1,11+1):
for n in range(1,max_aufgaben[i]+1):
all_aufgaben.append(float(f"{i}.{n}"))
print(all_aufgaben)
all_aufgaben_backup = [1.1, 1.2, 1.3, 1.4, 1.5, 1.6, 1.7, 2.1, 2.2, 2.3, 2.4, 2.5, 2.6, 2.7, 3.1, 3.2, 3.3, 3.4, 3.5, 3.6, 4.1, 4.2, 4.3, 4.4, 4.5, 4.6, 4.7, 5.1, 5.2, 5.3, 6.1, 6.2, 6.3, 6.4, 6.5, 7.1, 7.2, 7.3, 7.4, 7.5, 8.1, 8.2, 9.1, 9.2, 9.3, 10.1, 10.2, 10.3, 11.1, 11.2, 11.3, 11.4, 11.5, 11.6]
to_solve = np.random.choice(all_aufgaben, size = len(all_aufgaben), replace=False)
print(np.array(to_solve))
11.2,
| Lernphase/Aufgaben_zu_loesen.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
DATA_CONTAINER = "data/"
# -
# loading data
def load_csv_data(file_name, data_container=DATA_CONTAINER):
csv_path = os.path.join(data_container, file_name)
return pd.read_csv(csv_path)
def plot_boxplox(data_time, data_temp, labels, title):
f, (ax1, ax2) = plt.subplots(1, 2, sharey=True)
bp = ax1.boxplot(data_time, labels=labels, notch=True, bootstrap=10000)
ax1.set_ylabel('time to find all solutions (s)', color='b')
ax1.tick_params('y', colors='b')
for element in ['boxes', 'whiskers', 'fliers', 'means', 'medians', 'caps']:
plt.setp(bp[element], color='b')
ax2 = ax2.twinx()
bp = ax2.boxplot(data_temp, labels=labels, notch=True, bootstrap=10000)
ax2.set_ylabel('temperature (°C)', color='r', )
ax2.tick_params('y', colors='r')
for element in ['boxes', 'whiskers', 'fliers', 'means', 'medians', 'caps']:
plt.setp(bp[element], color='r')
plt.show()
def plot_data(timestamp, time, temp, title):
fig, ax1 = plt.subplots()
maxtime = timestamp[-1:]
ax1.plot(timestamp, time, 'b-')
ax1.set_xlabel('experiment time (min) - (duration: %.2f min)' %(maxtime))
ax1.set_ylabel('time to find all solutions (s)', color='b')
ax1.tick_params('y', colors='b')
ax2 = ax1.twinx()
ax2.plot(timestamp, temp, 'r')
ax2.set_ylabel('temperature (°C)', color='r')
ax2.tick_params('y', colors='r')
ax1.set_title(title)
plt.show()
def plot_barh(y_pos, performance, error, labels, title, color, xlabel):
fig, ax = plt.subplots()
plt.grid()
# Example data
people = labels
ax.barh(y_pos, performance, xerr=error, align='center', color=color, ecolor='black', height=0.1)
ax.set_yticks(y_pos)
ax.set_yticklabels(labels)
ax.invert_yaxis()
ax.set_xlabel(xlabel)
ax.set_title(title)
plt.show()
def plot_barh(y_pos, performance, error, labels, title, color, xlabel):
fig, ax = plt.subplots()
plt.grid()
# Example data
people = labels
ax.barh(y_pos, performance, xerr=error, align='center', color=color, ecolor='black', height=0.1)
ax.set_yticks(y_pos)
ax.set_yticklabels(labels)
ax.invert_yaxis()
ax.set_xlabel(xlabel)
ax.set_title(title)
plt.show()
path_3b = "data/performance/raspberry_pi/stretch/3B"
path_3b_p = "data/performance/raspberry_pi/stretch/3B+"
# **Standard Raspbian Kernel**
# +
# loading data standard raspbian kernel multi-thread solution
kernel_std_bplus_mt = load_csv_data("std_kernel_bplus_multithread.csv", path_3b_p)
kernel_std_bplus_mt['time'] -= kernel_std_bplus_mt['time'][0]
kernel_std_b_mt = load_csv_data("std_kernel_b_multithread.csv", path_3b)
kernel_std_b_mt['time'] -= kernel_std_b_mt['time'][0]
# +
# loading data standard raspbian kernel single-thread solution
kernel_std_bplus_st = load_csv_data("std_kernel_bplus_singlethread.csv", path_3b_p)
kernel_std_bplus_st['time'] -= kernel_std_bplus_st['time'][0]
kernel_std_b_st = load_csv_data("std_kernel_b_singlethread.csv", path_3b)
kernel_std_b_st['time'] -= kernel_std_b_st['time'][0]
# +
# loading data standard raspbian kernel multi-thread solution with networking+usb load
kernel_std_bplus_mt_network = load_csv_data("std_kernel_bplus_networking.csv", path_3b_p)
kernel_std_bplus_mt_network['time'] -= kernel_std_bplus_mt_network['time'][0]
kernel_std_bplus_mt_network_client = load_csv_data("send_statistics_std_kernel_bplus.csv", path_3b_p)
kernel_std_bplus_mt_network_client['time'] -= kernel_std_bplus_mt_network_client['time'][0]
# -
# **Preempt-RT Raspbian Kernel**
# +
# loading data preempt-rt raspbian kernel multi-thread solution
kernel_rt_bplus_mt = load_csv_data("rt_kernel_bplus_multithread.csv", path_3b_p)
kernel_rt_bplus_mt['time'] -= kernel_rt_bplus_mt['time'][0]
kernel_rt_b_mt = load_csv_data("rt_kernel_b_multithread.csv", path_3b)
kernel_rt_b_mt['time'] -= kernel_rt_b_mt['time'][0]
# +
# loading data preempt-rt raspbian kernel single-thread solution
kernel_rt_bplus_st = load_csv_data("rt_kernel_bplus_singlethread.csv", path_3b_p)
kernel_rt_bplus_st['time'] -= kernel_rt_bplus_st['time'][0]
kernel_rt_b_st = load_csv_data("rt_kernel_b_singlethread.csv", path_3b)
kernel_rt_b_st['time'] -= kernel_rt_b_st['time'][0]
# +
# loading data preempt-rt raspbian kernel multi-thread solution with networking+usb load
kernel_rt_bplus_mt_network = load_csv_data("rt_kernel_bplus_networking.csv", path_3b_p)
kernel_rt_bplus_mt_network['time'] -= kernel_rt_bplus_mt_network['time'][0]
kernel_rt_bplus_mt_network_client = load_csv_data("send_statistics_rt_kernel_bplus.csv", path_3b_p)
kernel_rt_bplus_mt_network_client['time'] -= kernel_rt_bplus_mt_network_client['time'][0]
# -
# **Standard Raspbian Kernel 4.14.27-v7+**
# +
# plotting standard raspbian kernel with model b+ multi-thread solution
timestamp_std_bplus_mt = kernel_std_bplus_mt.time/60
time_std_bplus_mt = kernel_std_bplus_mt.seconds + kernel_std_bplus_mt.microseconds/1000000
temp_std_bplus_mt = (kernel_std_bplus_mt.cpu_temp + kernel_std_bplus_mt.gpu_temp)/2
title = "Raspberry Pi 3 Model B+\nusing Stardard Raspbian Kernel 4.14.27-v7+\n(Queens: 12, Threads: 4, Iterations: 100)"
plot_data(timestamp_std_bplus_mt[:99], time_std_bplus_mt[:99], temp_std_bplus_mt[:99], title)
# -
print("Time to 45 iterations: %.2f" %timestamp_std_bplus_mt[44])
# plotting standard raspbian kernel with model b multi-thread solution
timestamp_std_b_mt = (kernel_std_b_mt.time)/60
time_std_b_mt = kernel_std_b_mt.seconds + kernel_std_b_mt.microseconds/1000000
temp_std_b_mt = (kernel_std_b_mt.cpu_temp + kernel_std_b_mt.gpu_temp)/2
title = "Raspberry Pi 3 Model B\nusing Stardard Raspbian Kernel 4.14.27-v7+\n(Queens: 12, Threads: 4, Iterations: 100)"
plot_data(timestamp_std_b_mt[:99], time_std_b_mt[:99], temp_std_b_mt[:99], title)
print("Time to 45 iterations: %.2f" %timestamp_std_b_mt[44])
# +
# plotting standard raspbian kernel with model b+ single-thread solution
timestamp_std_bplus_st = kernel_std_bplus_st.time/60
time_std_bplus_st = kernel_std_bplus_st.seconds + kernel_std_bplus_st.microseconds/1000000
temp_std_bplus_st = (kernel_std_bplus_st.cpu_temp + kernel_std_bplus_st.gpu_temp)/2
title = "Raspberry Pi 3 Model B+\nusing Stardard Raspbian Kernel 4.14.27-v7+\n(Queens: 12, Threads: 1, Iterations: 45)"
plot_data(timestamp_std_bplus_st[:44], time_std_bplus_st[:44], temp_std_bplus_st[:44], title)
# -
print("Time to 45 iterations: %.2f" %timestamp_std_bplus_st[44])
# +
# plotting standard raspbian kernel with model b single-thread solution
timestamp_std_b_st = kernel_std_b_st.time/60
time_std_b_st = kernel_std_b_st.seconds + kernel_std_b_st.microseconds/1000000
temp_std_b_st = (kernel_std_b_st.cpu_temp + kernel_std_b_st.gpu_temp)/2
title = "Raspberry Pi 3 Model B\nusing Stardard Raspbian Kernel 4.14.27-v7+\n(Queens: 12, Threads: 1, Iterations: 45)"
plot_data(timestamp_std_b_st[:44], time_std_b_st[:44], temp_std_b_st[:44], title)
# -
print("Time to 50 iterations: %.2f" %timestamp_std_b_st[44])
# +
# plotting standard raspbian kernel with model b+ multi-thread solution
timestamp_std_bplus_mt_net = kernel_std_bplus_mt_network.time/60
time_std_bplus_mt_net = kernel_std_bplus_mt_network.seconds + kernel_std_bplus_mt_network.microseconds/1000000
temp_std_bplus_mt_net = (kernel_std_bplus_mt_network.cpu_temp + kernel_std_bplus_mt_network.gpu_temp)/2
title = "Raspberry Pi 3 Model B+\nusing Stardard Raspbian Kernel 4.14.27-v7+\nwith Network and USB loads\n(Queens: 12, Threads: 4, Iterations: 100)"
plot_data(timestamp_std_bplus_mt_net, time_std_bplus_mt_net, temp_std_bplus_mt_net, title)
# -
print("Time to 50 iterations: %.2f" %timestamp_std_bplus_mt_net[44])
# +
# multi-thread vs single-thread
mt_timing_std_bplus = (time_std_bplus_st[:44,None]/time_std_bplus_mt[:44,None])
mt_timing_std_b = time_std_b_st[:44]/time_std_b_mt[:44]
print("Multi-thread B+: %.2f | Multi-thread B: %.2f" % (np.mean(mt_timing_std_bplus), np.mean(mt_timing_std_b)))
print("Multi-thread B+: %.2f | Multi-thread B: %.2f" % (np.mean(time_std_bplus_mt), np.mean(time_std_b_mt)))
# +
# model B vs B+ single- multi-thread
mt_timing_std_bplus_vs_b = time_std_b_mt[:99]/time_std_bplus_mt[:99]
st_timing_std_bplus_vs_b = time_std_b_st[:44]/time_std_bplus_st[:44]
print("Multi-thread: %.2f | Single-thread B: %.2f" % (np.mean(mt_timing_std_bplus_vs_b), np.mean(st_timing_std_bplus_vs_b)))
print("Multi-thread B+: %.2f | Multi-thread B: %.2f" % (np.mean(time_std_bplus_st), np.mean(time_std_b_st)))
# +
max_std_mt_bplus_temp = np.max(temp_std_bplus_mt)
max_std_mt_b_temp = np.max(temp_std_b_mt)
max_std_st_bplus_temp = np.max(temp_std_bplus_st)
max_std_st_b_temp = np.max(temp_std_b_st)
print("Max. temp. %.2f | Max. temp. %.2f" % (max_std_mt_bplus_temp, max_std_mt_b_temp))
print("Max. temp. %.2f | Max. temp. %.2f" % (max_std_st_bplus_temp, max_std_st_b_temp))
# +
max_std_mt_bplus_net_temp = np.max(temp_std_bplus_mt_net)
print("Max. temp. %.2f" % (max_std_mt_bplus_net_temp))
# +
# multi thread standard
mean_time_std_mt_bplus = np.mean(time_std_bplus_mt[:99])
var_time_std_mt_bplus = np.std(time_std_bplus_mt[:99])
mean_time_std_mt_b = np.mean(time_std_b_mt[:99])
var_time_std_mt_b = np.std(time_std_b_mt[:99])
# multi thread standard with load
mean_time_std_mt_bplus_net = np.mean(time_std_bplus_mt_net)
var_time_std_mt_bplus_net = np.std(time_std_bplus_mt_net)
#single thread standard
mean_time_std_st_bplus = np.mean(time_std_bplus_st[:44])
var_time_std_st_bplus = np.std(time_std_bplus_st[:44])
mean_time_std_st_b = np.mean(time_std_b_st[:44])
var_time_std_st_b = np.std(time_std_b_st[:44])
ypos = [0, 0.2, 0.4, 0.6, 0.8]
performance = [mean_time_std_mt_bplus, mean_time_std_mt_bplus_net, mean_time_std_st_bplus, mean_time_std_mt_b, mean_time_std_st_b ]
error = [var_time_std_mt_bplus, var_time_std_mt_bplus_net, var_time_std_st_bplus, var_time_std_mt_b, var_time_std_st_b]
labels = ['rPi v3 B+\n(Thread: 4, Repeat: 100)',
'rPi v3 B+ with Network & USB load\n(Thread: 4, Repeat: 100)',
'rPi v3 B+\n(Thread: 1, Repeat: 45)',
'rPi v3 B\n(Thread: 4, Repeat: 100)',
'rPi v3 B\n(Thread: 1, Repeat: 45)']
title = 'Mean time to find all solutions (12 Queens)\nusing Standard Raspbian Kernel 4.14.27-v7+'
color = 'blue'
xlabel = 'time (s)'
plot_barh(ypos, performance, error, labels, title, color, xlabel)
# +
# multi thread standard
mean_temp_std_mt_bplus = np.mean(temp_std_bplus_mt[:99])
var_temp_std_mt_bplus = np.std(temp_std_bplus_mt[:99])
mean_temp_std_mt_b = np.mean(temp_std_b_mt[:99])
var_temp_std_mt_b = np.std(temp_std_b_mt[:99])
# multi thread standard with load
mean_temp_std_mt_bplus_net = np.mean(temp_std_bplus_mt_net)
var_temp_std_mt_bplus_net = np.std(temp_std_bplus_mt_net)
#single thread
mean_temp_std_st_bplus = np.mean(temp_std_bplus_st[:44])
var_temp_std_st_bplus = np.std(temp_std_bplus_st[:44])
mean_temp_std_st_b = np.mean(temp_std_b_st[:44])
var_temp_std_st_b = np.std(temp_std_b_st[:44])
ypos = [0, 0.2, 0.4, 0.6, 0.8]
performance = [mean_temp_std_mt_bplus, mean_temp_std_mt_bplus_net, mean_temp_std_st_bplus, mean_temp_std_mt_b, mean_temp_std_st_b ]
error = [var_temp_std_mt_bplus, var_temp_std_mt_bplus_net, var_temp_std_st_bplus, var_temp_std_mt_b, var_temp_std_st_b]
labels = ['rPi v3 B+\n(Thread: 4, Repeat: 150)',
'rPi v3 B+ with Network & USB load\n(Thread: 4, Repeat: 100)',
'rPi v3 B+\n(Thread: 1, Repeat: 50)',
'rPi v3 B\n(Thread: 4, Repeat: 150)',
'rPi v3 B\n(Thread: 1, Repeat: 45)']
title = 'Mean temperature (12 Queens)\nusing Standard Kernel 4.14.27-v7+'
color = 'red'
xlabel = 'temperature (°C)'
plot_barh(ypos, performance, error, labels, title, color, xlabel)
# -
# **Preempt-RT Raspbian Kernel 4.14.27-rt21-v7**
# +
# plotting preempt-rt raspbian kernel with model b+ multi-thread solution
timestamp_rt_bplus_mt = kernel_rt_bplus_mt.time/60
time_rt_bplus_mt = kernel_rt_bplus_mt.seconds + kernel_rt_bplus_mt.microseconds/1000000
temp_rt_bplus_mt = (kernel_rt_bplus_mt.cpu_temp + kernel_rt_bplus_mt.gpu_temp)/2
title = "Raspberry Pi 3 Model B+\nusing Preempt-RT Raspbian Kernel 4.14.27-rt21-v7+\n(Queens: 12, Threads: 4, Iterations: 100)"
plot_data(timestamp_rt_bplus_mt[:99], time_rt_bplus_mt[:99], temp_rt_bplus_mt[:99], title)
# -
print("Time to 45 iterations: %.2f" %timestamp_rt_bplus_mt[44])
# +
# plotting preempt-rt raspbian kernel with model b multi-thread solution
timestamp_rt_b_mt = kernel_rt_b_mt.time/60
time_rt_b_mt = kernel_rt_b_mt.seconds + kernel_rt_b_mt.microseconds/1000000
temp_rt_b_mt = (kernel_rt_b_mt.cpu_temp + kernel_rt_b_mt.gpu_temp)/2
title = "Raspberry Pi 3 Model B\nusing Preempt-RT Raspbian Kernel 4.14.27-rt21-v7+\n(Queens: 12, Threads: 4, Iterations: 100)"
plot_data(timestamp_rt_b_mt[:99], time_rt_b_mt[:99], temp_rt_b_mt[:99], title)
# -
print("Time to 45 iterations: %.2f" %timestamp_rt_b_mt[44])
# +
# plotting preempt-rt raspbian kernel with model b+ single-thread solution
timestamp_rt_bplus_st = kernel_rt_bplus_st.time/60
time_rt_bplus_st = kernel_rt_bplus_st.seconds + kernel_rt_bplus_st.microseconds/1000000
temp_rt_bplus_st = (kernel_rt_bplus_st.cpu_temp + kernel_rt_bplus_st.gpu_temp)/2
title = "Raspberry Pi 3 Model B+\nusing Preempt-RT Raspbian Kernel 4.14.27-rt21-v7+\n(Queens: 12, Threads: 1, Iterations: 45)"
plot_data(timestamp_rt_bplus_st[:44], time_rt_bplus_st[:44], temp_rt_bplus_st[:44], title)
# -
print("Time to 45 iterations: %.2f" %timestamp_rt_bplus_st[44])
# +
# plotting preempt-rt raspbian kernel with model b single-thread solution
timestamp_rt_b_st = kernel_rt_b_st.time/60
time_rt_b_st = kernel_rt_b_st.seconds + kernel_rt_b_st.microseconds/1000000
temp_rt_b_st = (kernel_rt_b_st.cpu_temp + kernel_rt_b_st.gpu_temp)/2
title = "Raspberry Pi 3 Model B\nusing Preempt-RT Raspbian Kernel 4.14.27-rt21-v7+\n(Queens: 12, Threads: 1, Iterations: 45)"
plot_data(timestamp_rt_b_st[:44], time_rt_b_st[:44], temp_rt_b_st[:44], title)
# -
print("Time to 45 iterations: %.2f" %timestamp_rt_b_st[44])
# +
# plotting standard raspbian kernel with model b+ multi-thread solution
timestamp_rt_bplus_mt_net = kernel_rt_bplus_mt_network.time/60
time_rt_bplus_mt_net = kernel_rt_bplus_mt_network.seconds + kernel_rt_bplus_mt_network.microseconds/1000000
temp_rt_bplus_mt_net = (kernel_rt_bplus_mt_network.cpu_temp + kernel_rt_bplus_mt_network.gpu_temp)/2
title = "Raspberry Pi 3 Model B+\nusing Preempt-RT Raspbian Kernel 4.14.27-rt21-v7+\nwith Network and USB loads\n(Queens: 12, Threads: 4, Iterations: 100)"
plot_data(timestamp_rt_bplus_mt_net, time_rt_bplus_mt_net, temp_rt_bplus_mt_net, title)
# +
# multi thread standard
mean_time_rt_mt_bplus = np.mean(time_rt_bplus_mt)
var_time_rt_mt_bplus = np.std(time_rt_bplus_mt)
mean_time_rt_mt_b = np.mean(time_rt_b_mt)
var_time_rt_mt_b = np.std(time_rt_b_mt)
# multi thread standard with load
mean_time_rt_mt_bplus_net = np.mean(time_rt_bplus_mt_net)
var_time_rt_mt_bplus_net = np.std(time_rt_bplus_mt_net)
#single thread standard
mean_time_rt_st_bplus = np.mean(time_rt_bplus_st)
var_time_rt_st_bplus = np.std(time_rt_bplus_st)
mean_time_rt_st_b = np.mean(time_rt_b_st)
var_time_rt_st_b = np.std(time_rt_b_st)
ypos = [0, 0.2, 0.4, 0.6, 0.8]
performance = [mean_time_rt_mt_bplus, mean_time_rt_mt_bplus_net, mean_time_rt_st_bplus, mean_time_rt_mt_b, mean_time_rt_st_b ]
error = [var_time_rt_mt_bplus, var_time_rt_mt_bplus_net, var_time_rt_st_bplus, var_time_rt_mt_b, var_time_rt_st_b]
labels = ['rPi v3 B+\n(Thread: 4, Repeat: 150)',
'rPi v3 B+ with Network & USB load\n(Thread: 4, Repeat: 100)',
'rPi v3 B+\n(Thread: 1, Repeat: 45)',
'rPi v3 B\n(Thread: 4, Repeat: 150)',
'rPi v3 B\n(Thread: 1, Repeat: 45)']
title = 'Mean Time to find all solutions (12 Queens)\nusing Preempt-RT Raspbian Kernel 4.14.27-rt21-v7+'
color = 'blue'
xlabel = 'time (s)'
plot_barh(ypos, performance, error, labels, title, color, xlabel)
# +
# multi-thread vs single-thread
mt_timing_rt_bplus = (time_rt_bplus_st[:44,None]/time_rt_bplus_mt[:44,None])
mt_timing_rt_b = time_rt_b_st[:44]/time_rt_b_mt[:44]
print("Multi-thread B+: %.2f | Multi-thread B: %.2f" % (np.mean(mt_timing_rt_bplus), np.mean(mt_timing_rt_b)))
print("Multi-thread B+: %.2f | Multi-thread B: %.2f" % (np.mean(time_rt_bplus_mt), np.mean(time_rt_b_mt)))
# model B vs B+ single- multi-thread
mt_timing_rt_bplus_vs_b = time_rt_b_mt[:99]/time_rt_bplus_mt[:99]
st_timing_rt_bplus_vs_b = time_rt_b_st[:44]/time_rt_bplus_st[:44]
print("Multi-thread: %.2f | Single-thread B: %.2f" % (np.mean(mt_timing_rt_bplus_vs_b), np.mean(st_timing_rt_bplus_vs_b)))
print("Multi-thread B+: %.2f | Multi-thread B: %.2f" % (np.mean(time_rt_bplus_st), np.mean(time_rt_b_st)))
max_rt_mt_bplus_temp = np.max(temp_rt_bplus_mt)
max_rt_mt_b_temp = np.max(temp_rt_b_mt)
max_rt_st_bplus_temp = np.max(temp_rt_bplus_st)
max_rt_st_b_temp = np.max(temp_rt_b_st)
print("Max. temp. %.2f | Max. temp. %.2f" % (max_rt_mt_bplus_temp, max_rt_mt_b_temp))
print("Max. temp. %.2f | Max. temp. %.2f" % (max_rt_st_bplus_temp, max_rt_st_b_temp))
max_rt_mt_bplus_net_temp = np.max(temp_rt_bplus_mt_net)
print("Max. temp. %.2f" % (max_rt_mt_bplus_net_temp))
# +
# multi thread
mean_temp_rt_mt_bplus = np.mean(temp_rt_bplus_mt)
var_temp_rt_mt_bplus = np.std(temp_rt_bplus_mt)
mean_temp_rt_mt_b = np.mean(temp_rt_b_mt)
var_temp_rt_mt_b = np.std(temp_rt_b_mt)
# multi thread standard with load
mean_temp_rt_mt_bplus_net = np.mean(temp_rt_bplus_mt_net)
var_temp_rt_mt_bplus_net = np.std(temp_rt_bplus_mt_net)
#single thread
mean_temp_rt_st_bplus = np.mean(temp_rt_bplus_st)
var_temp_rt_st_bplus = np.std(temp_rt_bplus_st)
mean_temp_rt_st_b = np.mean(temp_rt_b_st)
var_temp_rt_st_b = np.std(temp_rt_b_st)
ypos = [0, 0.2, 0.4, 0.6, 0.8]
performance = [mean_temp_rt_mt_bplus, mean_temp_rt_mt_bplus_net, mean_temp_rt_st_bplus, mean_temp_rt_mt_b, mean_temp_rt_st_b ]
error = [var_temp_rt_mt_bplus, var_temp_rt_mt_bplus_net, var_temp_rt_st_bplus, var_temp_rt_mt_b, var_temp_rt_st_b]
labels = ['rPi v3 B+\n(Thread: 4, Repeat: 150)',
'rPi v3 B+ with Network & USB load\n(Thread: 4, Repeat: 100)',
'rPi v3 B+\n(Thread: 1, Repeat: 45)',
'rPi v3 B\n(Thread: 4, Repeat: 150)',
'rPi v3 B\n(Thread: 1, Repeat: 45)']
title = 'Mean Temperature (12 Queens)\nusing Preempt-RT Kernel 4.14.27-rt21-v7+'
color = 'red'
xlabel = 'temperature (°C)'
plot_barh(ypos, performance, error, labels, title, color, xlabel)
# -
# **Comparison**
# +
mt_timing_rt_std_bplus = time_rt_bplus_mt[:99,None]/time_std_bplus_mt[:99,None]
mt_timing_rt_std_b = time_rt_b_mt[:44]/time_std_b_mt[:44]
st_timing_rt_std_bplus = time_rt_bplus_st[:99,None]/time_std_bplus_st[:99,None]
st_timing_rt_std_b = time_rt_b_st[:44]/time_std_b_st[:44]
print("RT B+ MT: %.2f | RT B MT: %.2f" % (np.mean(mt_timing_rt_std_bplus), np.mean(mt_timing_rt_std_b)))
print("RT B+ ST: %.2f | RT B ST: %.2f" % (np.mean(st_timing_rt_std_bplus), np.mean(st_timing_rt_std_b)))
# -
# **Networking**
# +
# multi thread rt
mean_time_rt_mt_bplus_net = np.mean(time_rt_bplus_mt_net)
var_time_rt_mt_bplus_net = np.std(time_rt_bplus_mt_net)
# multi thread standard
mean_time_std_mt_bplus_net = np.mean(time_std_bplus_mt_net)
var_time_std_mt_bplus_net = np.std(time_std_bplus_mt_net)
ypos = [0, 0.4]
performance = [mean_time_std_mt_bplus_net, mean_time_rt_mt_bplus_net]
error = [var_time_std_mt_bplus_net, var_time_rt_mt_bplus_net]
labels = ['rPi v3 B+\nStandard Raspbian Kernel\n(Thread: 4, Repeat: 100)',
'rPi v3 B+\nPreempt-RT Raspbian Kernel\n(Thread: 4, Repeat: 100)']
title = 'Mean Time to find all solutions (12 Queens)\nusing Standard & Standard Raspbian Kernel 4.14.27-(rt21)-v7+'
color = 'blue'
xlabel = 'time (s)'
plot_barh(ypos, performance, error, labels, title, color, xlabel)
# +
# multi thread rt
mean_temp_rt_mt_bplus_net = np.mean(temp_rt_bplus_mt_net)
var_temp_rt_mt_bplus_net = np.std(temp_rt_bplus_mt_net)
# multi thread standard
mean_temp_std_mt_bplus_net = np.mean(temp_std_bplus_mt_net)
var_temp_std_mt_bplus_net = np.std(temp_std_bplus_mt_net)
ypos = [0, 0.4]
performance = [mean_temp_std_mt_bplus_net, mean_temp_rt_mt_bplus_net]
error = [var_temp_std_mt_bplus_net, var_temp_rt_mt_bplus_net]
labels = ['rPi v3 B+\nStandard Raspbian Kernel\n(Thread: 4, Repeat: 100)',
'rPi v3 B+\nPreempt-RT Raspbian Kernel\n(Thread: 4, Repeat: 100)']
title = 'Mean Temperature (12 Queens)\nusing Standard & Preempt-RT Kernel 4.14.27-(rt21)-v7+'
color = 'red'
xlabel = 'temperature (°C)'
plot_barh(ypos, performance, error, labels, title, color, xlabel)
# -
print("Max. temp: %.2f | Max. temp: %.2f" %(np.max(temp_std_bplus_mt_net),np.max(temp_rt_bplus_mt_net) ))
# +
mt_timing_rt_std_bplus_net = time_rt_bplus_mt_net[:99,None]/time_std_bplus_mt_net[:99,None]
mt_temp_rt_std_bplus_net = temp_rt_bplus_mt_net[:99,None]/temp_std_bplus_mt_net[:99,None]
print("RT B+ MT: %.2f" % (np.mean(mt_timing_rt_std_bplus_net)))
print("RT B+ MT: %.2f" % (np.mean(mt_temp_rt_std_bplus_net)))
# -
kernel_rt_bplus_mt_network_client.head()
experiment_time = kernel_rt_bplus_mt_network_client.time/60
transfer_time_rt = (kernel_rt_bplus_mt_network_client.seconds+kernel_rt_bplus_mt_network_client.microseconds/1000000)
transfer_rate_rt = kernel_rt_bplus_mt_network_client.size/transfer_time_rt/1024
fig, ax1 = plt.subplots()
ax1.plot(experiment_time, transfer_rate_rt)
ax1.set_xlabel('experiment time (min)')
ax1.set_ylabel('transfer rate (kb/s)')
ax1.set_title('Ethernet transfer data rate\nusing Preempt-RT Raspbian Kernel 4.14.27-rt21-v7+\nMin.: %.2f kb/s, Max.: %.2f kb/s, Mean: %.2f kb/s' %(np.min(transfer_rate_rt), np.max(transfer_rate_rt), np.mean(transfer_rate_rt)))
experiment_time = kernel_rt_bplus_mt_network_client.time/60
transfer_time_std = (kernel_std_bplus_mt_network_client.seconds+kernel_std_bplus_mt_network_client.microseconds/1000000)
transfer_rate_std = kernel_std_bplus_mt_network_client.size/transfer_time_std/1024
fig, ax1 = plt.subplots()
ax1.plot(experiment_time[:2223], transfer_rate_std[:2223]) # 2223 -> I forgot to stop the clock! ;)
ax1.set_xlabel('experiment time (min)')
ax1.set_ylabel('transfer rate (kb/s)')
ax1.set_title('Ethernet transfer data rate\nusing Standard Raspbian Kernel 4.14.27-v7+\n Min.: %.2f kb/s, Max.: %.2f kb/s, Mean: %.2f kb/s' %(np.min(transfer_rate_std[:2223]), np.max(transfer_rate_std[:2223]), np.mean(transfer_rate_std[:2223])))
| performance_test_std_preempt_rt_rpi_3B_3Bp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score,classification_report,confusion_matrix
from sklearn.naive_bayes import GaussianNB
# %matplotlib inline
# -
# ### 1.Load the kinematics dataset as measured on mobile sensors from the file “run_or_walk.csv”. List out the columns in the dataset.
data = pd.read_csv('run_or_walk.csv')
columns = data.columns
columns
data.head()
data.shape
data.info()
data.describe().transpose()
# Feature engineering: drop the columns- 'date', 'time', 'username'
data = data.drop(['date', 'time', 'username'],axis=1)
# Target variable:
data['activity'].value_counts()
# ### 2. Let the target variable ‘y’ be the activity and assign all the columns after it to ‘x’.
# +
x = data.drop(['activity'],axis=1)
y = data['activity']
x_train, x_test , y_train, y_test = train_test_split(x,y,train_size=0.8)
print(x_train.shape)
print(x_test.shape)
print(y_train.shape)
print(y_test.shape)
# -
# ### 3.Using Scikit-learn fit a Gaussian Naive Bayes model and observe the accuracy. Generate a classification report using scikit learn.
#
# +
gnb = GaussianNB()
gnb = gnb.fit(x_train,y_train)
y_predict = gnb.predict(x_test)
print('Accuracy:',accuracy_score(y_test,y_predict))
print('classification report:', classification_report(y_test,y_predict))
print('confusion_matrix:', confusion_matrix(y_test,y_predict))
# -
# ### 4.Repeat the model once using only the acceleration values as predictors and then using only the gyro values as predictors. Comment on the difference in accuracy between both the models.
# +
x = data[['acceleration_x','acceleration_y','acceleration_z']]
y = data['activity']
x_train, x_test , y_train, y_test = train_test_split(x,y,train_size=0.8)
print(x_train.shape)
print(x_test.shape)
print(y_train.shape)
print(y_test.shape)
gnb = GaussianNB()
gnb = gnb.fit(x_train,y_train)
y_predict = gnb.predict(x_test)
print('Accuracy:',accuracy_score(y_test,y_predict))
print('classification report:', classification_report(y_test,y_predict))
print('confusion_matrix:', confusion_matrix(y_test,y_predict))
# +
x = data[['gyro_x','gyro_y','gyro_z']]
y = data['activity']
x_train, x_test , y_train, y_test = train_test_split(x,y,train_size=0.8)
print(x_train.shape)
print(x_test.shape)
print(y_train.shape)
print(y_test.shape)
gnb = GaussianNB()
gnb = gnb.fit(x_train,y_train)
y_predict = gnb.predict(x_test)
print('Accuracy:',accuracy_score(y_test,y_predict))
print('classification report:', classification_report(y_test,y_predict))
print('confusion_matrix:', confusion_matrix(y_test,y_predict))
# +
# From the comparision of accuracy score between both the models it is evident that the acceleration variables have significant
# impact in the prediction of target variable activity
# -
| Module 9/Module_9-Case+study+2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Docker-compose
#
# 
#
# ## Intoduction
# Docker-compose is a tool that allows you to easily manage multiple containers at once.
#
# ## Context
# Let's say that you have multiple container for your app which is stored in a single Github repo:
# * A container that extracts data from a website
# * A container that receives data and cleans it
# * A container for your Database
# * A container that manages an API
#
# If you want to run your entire system, you will need to start each container manually, if you change the code of all your containers, you need to re-build all of them one by one.
# If you want to share volumes between them, it's complicated.
# And so on...
#
# As you can imagine, it becomes really messy and it's a pain to maintain...
#
# ## Solution
# To tackle that, you can use `docker-compose`. It allows you to manage all those containers at once.
#
# You can create a single `docker-compose` file where you define all your containers and where the corresponding Dockerfile are.
#
# You can also create volumes in it and bind them to every container you want. Then you can simply run one command to build all your images, run all your containers at the same time.
#
# ## Conclusion
# I will not go too deep about it now. Let's master the basics first!
#
# But if one day you come to a point where you need to use multiple containers/images and you feel that's a lot to manage individually, remember docker-compose!
#
# It could also be a super interesting watch/workshop!
#
# ## Some documentation
# * [Official documentation](https://docs.docker.com/compose/)
# * [Awesome article about it](https://towardsdatascience.com/docker-compose-44a8112c850a)
#
# 
| Bonus_resources/deployment/2.Docker/5.docker-compose.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework 0
# In this homework, we will go through basic linear algebra and image manipulation using python to get everyone on the same page for the prerequisite skills for this class.
#
# One of the aims of this homework assignment is to get you to start getting comfortable searching for useful library functions online. So in many of the functions you will implement, you will have to look up helper functions.
# +
#Imports the print function from newer versions of python
from __future__ import print_function
#Setup
# The Random module for implements pseudo-random number generators
import random
# Numpy is the main package for scientific computing with Python.
# This will be one of our most used libraries in this class
import numpy as np
#Imports all the methods in each of the files: linalg.py and imageManip.py
from linalg import *
from imageManip import *
#Matplotlib is a useful plotting library for python
import matplotlib.pyplot as plt
# This code is to make matplotlib figures appear inline in the
# notebook rather than in a new window.
# %matplotlib inline
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# Some more magic so that the notebook will reload external python modules;
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
# %load_ext autoreload
# %autoreload 2
# %reload_ext autoreload
# -
# ## Question 1.1 (5 points)
# First, let's test whether you can define the following matrices and vectors using numpy. Look up `np.array()` for help. In the next code block, define $M$ as a $(4, 3)$ matrix, $a$ as a $(1, 3)$ row vector and $b$ as a $(3, 1)$ column vector:
#
# $$M = \begin{bmatrix}
# 1 & 2 & 3 \\
# 4 & 5 & 6 \\
# 7 & 8 & 9 \\
# 10 & 11 & 12 \end{bmatrix}
# $$
#
# $$a = \begin{bmatrix}
# 1 & 1 & 0
# \end{bmatrix}
# $$
#
# $$b = \begin{bmatrix}
# -1 \\ 2 \\ 5
# \end{bmatrix}
# $$
### YOUR CODE HERE
M = np.array([[1,2,3],[4,5,6],[7,8,9],[10,11,12]])
a = np.array([[1,1,0]])
b = np.array([[-1],[2],[5]])
### END CODE HERE
print("M = \n", M)
print("The size of M is: ", M.shape)
print()
print("a = ", a)
print("The size of a is: ", a.shape)
print()
print("b = ", b)
print("The size of b is: ", b.shape)
# ## Question 1.2 (5 points)
# Implement the `dot_product()` method in `linalg.py` and check that it returns the correct answer for $a^Tb$.
# +
# Let's remove the spurious dimensions along vectors a and b to convert them both to row vectors:
squeezed_a = np.squeeze(a)
squeezed_b = np.squeeze(b)
# Now, let's test out this dot product. Your answer should be 1.
aDotB = dot_product(squeezed_a, squeezed_b)
print(aDotB)
# -
# # Question 1: Linear Algebra Review
# In this section, we will review linear algebra and learn how to use vectors and matrices in python using numpy. By the end of this section, you will have implemented all the required methods in `linalg.py`.
# ## Question 1.3 (5 points)
# Implement the `complicated_matrix_function()` method in `linalg.py` and use it to compute $(a^T b)Ma^T$
# Your answer should be $[[3], [9], [15], [21]]$ of shape(4, 1).
ans = complicated_matrix_function(M, a, b)
print(ans)
print()
print("The size is: ", ans.shape)
# ## Question 1.4 (10 points)
# Implement `svd()` and `get_singular_values()` methods. In this method, perform singular value decomposition on the input matrix and return the largest k singular values (k is specified in the method calls below).
# +
# Let's first only get the first singular value and print it out. It should be ~ 25.46.
only_first_singular_value = get_singular_values(M, 1)
print(only_first_singular_value)
# Now, let's get the first two singular values.
# Notice the first singular value is a lot larger than the second one.
first_two_singular_values = get_singular_values(M, 2)
print(first_two_singular_values)
# Let's make sure that the first singular value in both is the same.
assert only_first_singular_value[0] == first_two_singular_values[0]
# -
# ## Question 1.5 (10 points)
# Implement `eigen_decomp()` and `get_eigen_values_and_vectors()` methods. In this method, perform eigenvalue decomposition on the following matrix and return the largest k eigen values and corresponding eigen vectors (k is specified in the method calls below).
#
# $$M = \begin{bmatrix}
# 1 & 2 & 3 \\
# 4 & 5 & 6 \\
# 7 & 8 & 9 \end{bmatrix}
# $$
#
# +
# Let's define M.
M = np.array([[1,2,3],[4,5,6],[7,8,9]])
# Now let's grab the first eigenvalue and first eigenvector.
# You should get back a single eigenvalue and a single eigenvector.
val, vec = get_eigen_values_and_vectors(M[:,:3], 1)
print("First eigenvalue =", val[0])
print()
print("First eigenvector =", vec[0])
print()
# Now, let's get the first two eigenvalues and eigenvectors.
# You should get back a list of two eigenvalues and a list of two eigenvector arrays.
val, vec = get_eigen_values_and_vectors(M[:,:3], 2)
print("Eigenvalues =", val)
print()
print("Eigenvectors =", vec)
# -
# # Part 2: Image Manipulation
#
# Now that you are familiar with using matrices and vectors. Let's load some images and treat them as matrices and do some operations on them. By the end of this section, you will have implemented all the methods in `imageManip.py`
# +
# Run this code to set the locations of the images we will be using.
# You can change these paths to point to your own images if you want to try them out for fun.
image1_path = './image1.jpg'
image2_path = './image2.jpg'
def display(img):
# Show image
plt.figure(figsize = (5,5))
plt.imshow(img)
plt.axis('off')
plt.show()
# -
# ## Question 2.1 (5 points)
# Implement the load method in imageManip.py and read the display method below. We will use these two methods through the rest of the notebook to visualize our work.
# +
image1 = load(image1_path)
image2 = load(image2_path)
display(image1)
display(image2)
# -
# ## Question 2.2 (10 points)
# Implement the `dim_image()` method by converting images according to $x_n = 0.5*x_p^2$ for every pixel, where $x_n$ is the new value and $x_p$ is the original value.
#
# Note: Since all the pixel values of the image are in the range $[0, 1]$, the above formula will result in reducing these pixels values and therefore make the image dimmer.
new_image = dim_image(image1)
display(new_image)
# ## Question 2.3 (10 points)
# Implement the convert_to_grey_scale method and convert the image into grey scale.
grey_image = convert_to_grey_scale(image1)
display(grey_image)
# ## Question 2.4 (10 points)
# Implement the `rgb_exclusion()`, in which the input image is decomposed into the three channels: R, G and B and return the image excluding the specified channel.
# +
without_red = rgb_exclusion(image1, 'R')
without_blue = rgb_exclusion(image1, 'B')
without_green = rgb_exclusion(image1, 'G')
print("Below is the image without the red channel.")
display(without_red)
print("Below is the image without the green channel.")
display(without_green)
print("Below is the image without the blue channel.")
display(without_blue)
# -
# ## Question 2.5 (10 points)
# Implement the lab_decomposition, in which the input image is decomposed into the three channels: L, A and B and return the values for the specified channel.
# +
image_l = lab_decomposition(image1, 'L')
image_a = lab_decomposition(image1, 'A')
image_b = lab_decomposition(image1, 'B')
print("Below is the image with only the L channel.")
display(image_l)
print("Below is the image with only the A channel.")
display(image_a)
print("Below is the image with only the B channel.")
display(image_b)
# -
# ### Question:
# Explain in 2-3 sentences what the L, A and B channels are and what happens when you take away the L and A channels.
# ### Answer:
# Write your answer here.
# ## Question 2.6 (10 points)
# Implement the `hsv_decomposition()`, in which the input image is decomposed into the three channels: H, S and V and return the values for the specified channel.
# +
image_h = hsv_decomposition(image1, 'H')
image_s = hsv_decomposition(image1, 'S')
image_v = hsv_decomposition(image1, 'V')
print("Below is the image with only the H channel.")
display(image_h)
print("Below is the image with only the S channel.")
display(image_s)
print("Below is the image with only the V channel.")
display(image_v)
# -
# ### Question:
# Explain in 2-3 sentences what the H, S and V channels are and what happens when you take away the H and S channels.
# ### Answer:
# Write your answer here.
# ## Question 2.7 (10 points)
# In mix_images method, create a new image such that the left half of the image is the left half of image1 and the
# right half of the image is the right half of image2. Exclude the specified channel for the given image.
#
# You should see the left half of the monkey without the red channel and the right half of the house image with no green channel.
image_mixed = mix_images(image1, image2, channel1='R', channel2='G')
display(image_mixed)
# ## Extra credit (10 points)
#
# The following questions are optional and will go towards you extra credit grade.
#
# Implement `mix_quadrants` function in `imageManip.py`.
mixed_quadrants = mix_quadrants(image1)
display(mixed_quadrants)
| hw0_release/.ipynb_checkpoints/hw0-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bot Check Check the followers count using bot score from BOT checker API
# You need a rapid api key for [botometer](https://botometer.osome.iu.edu/) from https://botometer.osome.iu.edu/api
# You need also twitter API keys from https://developer.twitter.com with upgraded plan.
#
# Create a file with your secrets called 'settings.json'
#
# ```json
# {
# "rapidapi_key" : "xxxx",
# "consumer_key": "xxxx",
# "consumer_secret": "xxx",
# "access_token": "xxx",
# "access_token_secret": "xxx"
# }
# ```
# +
import botometer
import json
with open("settings.json") as file:
twitter_app_auth = json.load(file)
rapidapikey = twitter_app_auth.pop("rapidapi_key")
# -
# ## The reach and the influence of the account is huge, let's list the top verified followers of the account
import pandas as pd
df = pd.read_csv("./data/narkosedoc_followers.csv")
## Do some data cleansing, drop the NANs in followers
df.dropna(subset = ["followers_count"], inplace=True)
df = df.sort_values(by=["verified", "followers_count"])
df.tail(10)
# +
# remove verified accounts from the data set
df = df[df["verified"] != True]
samples = df.sample(n=100)
samples
# -
# +
bom = botometer.Botometer(wait_on_ratelimit=True,
rapidapi_key=rapidapikey,
**twitter_app_auth)
f = open("botscores.txt", 'w')
for screen_name, result in bom.check_accounts_in(samples["screen_name"]):
output["id_str"] = result["user_data"]["id_str"];
output["id_str"] = result["user_data"]["id_str"];
universalScore = flatten_json(result["raw_scores"])
print (universalScore)
json_object = json.dumps(result)
if "error" not in result:
f.write(json_object + "\n")
print (result)
f.close()
# -
data = pd.read_json ("botscores_backup.txt", lines=True)
data = data[["display_scores", "user"]]
def flatten_nested_json_df(df):
df = df.reset_index()
print(f"original shape: {df.shape}")
print(f"original columns: {df.columns}")
# search for columns to explode/flatten
s = (df.applymap(type) == list).all()
list_columns = s[s].index.tolist()
s = (df.applymap(type) == dict).all()
dict_columns = s[s].index.tolist()
print(f"lists: {list_columns}, dicts: {dict_columns}")
while len(list_columns) > 0 or len(dict_columns) > 0:
new_columns = []
for col in dict_columns:
print(f"flattening: {col}")
# explode dictionaries horizontally, adding new columns
horiz_exploded = pd.json_normalize(df[col]).add_prefix(f'{col}.')
horiz_exploded.index = df.index
df = pd.concat([df, horiz_exploded], axis=1).drop(columns=[col])
new_columns.extend(horiz_exploded.columns) # inplace
for col in list_columns:
print(f"exploding: {col}")
# explode lists vertically, adding new columns
df = df.drop(columns=[col]).join(df[col].explode().to_frame())
new_columns.append(col)
# check if there are still dict o list fields to flatten
s = (df[new_columns].applymap(type) == list).all()
list_columns = s[s].index.tolist()
s = (df[new_columns].applymap(type) == dict).all()
dict_columns = s[s].index.tolist()
print(f"lists: {list_columns}, dicts: {dict_columns}")
print(f"final shape: {df.shape}")
print(f"final columns: {df.columns}")
return df
data=flatten_nested_json_df(data)[['display_scores.universal.fake_follower',
'display_scores.universal.financial', 'display_scores.universal.other',
'display_scores.universal.overall',
'display_scores.universal.self_declared',
'display_scores.universal.spammer', 'user.majority_lang',
'user.user_data.id_str', 'user.user_data.screen_name']];
data["url"] = "https://twitter.com/" + data["user.user_data.screen_name"]
data[["display_scores.universal.overall", "url"]]
data
| BotScoresDoc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## ewf-wfp-02-01-04 - Land Surface Temperature Anomalies Time Series
# Land Surface Temperature Anomalies Time Series
# ---
# ### <a name="service">Service definition
service = dict([('title', 'Land Surface Temperature Anomalies Time Series'),
('abstract', 'Land Surface Temperature Anomalies Time Series'),
('id', 'ewf-wfp-02-01-04')])
# ### <a name="parameter">Parameter Definition
N_1 = dict([('id', 'N_1'),
('value', 'False'),
('title', 'No Aggregation'),
('abstract', 'No aggregation')])
N_3 = dict([('id', 'N_3'),
('value', 'True'),
('title', '30 Day Aggregation'),
('abstract', 'Get a 30 day aggregation')])
N_6 = dict([('id', 'N_6'),
('value', 'False'),
('title', '60 Day Aggregation'),
('abstract', 'Get a 30 day aggregation')])
N_9 = dict([('id', 'N_9'),
('value', 'False'),
('title', '90 Day Aggregation'),
('abstract', 'Get a 90 day aggregation')])
N_12 = dict([('id', 'N_12'),
('value', 'False'),
('title', '120 Day Aggregation'),
('abstract', 'Get a 120 day aggregation')])
N_15 = dict([('id', 'N_15'),
('value', 'False'),
('title', '150 Day Aggregation'),
('abstract', 'Get a 150 day aggregation')])
N_18 = dict([('id', 'N_18'),
('value', 'False'),
('title', '180 Day Aggregation'),
('abstract', 'Get a 180 day aggregation')])
N_27 = dict([('id', 'N_27'),
('value', 'False'),
('title', '270 Day Aggregation'),
('abstract', 'Get a 270 day aggregation')])
N_36 = dict([('id', 'N_36'),
('value', 'False'),
('title', '360 Day Aggregation'),
('abstract', 'Get a 360 day aggregation')])
regionOfInterest = dict([('id', 'regionOfInterest'),
('value', 'POLYGON((-179.999 89.999, 179.999 89.999, 179.999 -89.999, -179.999 -89.999, -179.999 89.999))'),
('title', 'WKT Polygon for the Region of Interest'),
('abstract', 'Set the value of WKT Polygon')])
nameOfRegion = dict([('id', 'nameOfRegion'),
('value', 'Global'),
('title', 'Name of Region'),
('abstract', 'Name of the region of interest'),
('minOccurs', '1')])
startdate = dict([('id', 'startdate'),
('value', '2016-01-01T00:00Z'),
('title', 'Start date'),
('abstract', 'Start date')])
enddate = dict([('id', 'enddate'),
('value', '2016-01-31T23:59Z'),
('title', 'End date'),
('abstract', 'End date')])
indexAgg = dict([('id', 'indexAgg'),
('value', 'better-wfp-02-01-02'),
('title', 'Aggregation user'),
('abstract', 'user to access aggregations catalog'),
('minOccurs', '1')])
apikeyAgg = dict([('id', 'apikeyAgg'),
('value', ''),
('title', 'Aggregation apikey'),
('abstract', 'apikey to access aggregations catalog'),
('minOccurs', '1')])
indexLTA = dict([('id', 'indexLTA'),
('value', 'better-wfp-02-01-03'),
('title', 'LTA user'),
('abstract', 'user to access LTAs catalog'),
('minOccurs', '1')])
apikeyLTA = dict([('id', 'apikeyLTA'),
('value', ''),
('title', 'LTA apikey'),
('abstract', 'apikey to access LTAs catalog'),
('minOccurs', '1')])
# ### <a name="runtime">Runtime parameter definition
# **Input identifiers**
#
# This is the MDOIS stack of products' identifiers
input_identifiers = ('dummy')
# **Input references**
#
# This is the MODIS stack catalogue references
# + slideshow={"slide_type": "subslide"}
input_references = ['dummy']
# + [markdown] slideshow={"slide_type": "slide"}
# **Data path**
#
# This path defines where the data is staged-in.
# -
data_path = "/workspace/modis"
# #### Aux folders
output_folder = ''
temp_folder = 'temp'
# #### Import Modules
# +
import os
import shutil
import cioppy
import sys
import string
import numpy as np
from osgeo import gdal, ogr, osr
from shapely.wkt import loads
import pandas as pd
import geopandas as gpd
import datetime
ciop = cioppy.Cioppy()
# -
# #### Auxiliary vars
check_results = False
# #### Auxiliary methods
# +
def rm_cfolder(folder):
for the_file in os.listdir(folder):
file_path = os.path.join(folder, the_file)
try:
if os.path.isfile(file_path):
os.unlink(file_path)
elif os.path.isdir(file_path): shutil.rmtree(file_path)
except Exception as e:
print(e)
# get metadata from catalog
def get_input_metadata (input_refs, apikeys):
# for each product get metadata
Result_Prod = []
for index,product_ref in enumerate(input_refs):
for index in apikeys:
if index in product_ref:
cat_index = index
cat_apikey = apikeys[index]
# since the search is by identifier
Result_Prod.append(ciop.search(end_point = product_ref,params =[],output_fields='self,identifier,startdate,enclosure,startdate,enddate,wkt,title',creds='{}:{}'.format(cat_index,cat_apikey))[0] )
input_metadata = gpd.GeoDataFrame.from_dict(Result_Prod)
input_metadata['startdate'] = pd.to_datetime(input_metadata['startdate'])
input_metadata['enddate'] = pd.to_datetime(input_metadata['enddate'])
return input_metadata
def get_metadata(filepath):
ds = gdal.Open(filepath)
projection = ds.GetProjection()
geotransform = ds.GetGeoTransform()
no_data_value = ds.GetRasterBand(1).GetNoDataValue()
data_type = ds.GetRasterBand(1).DataType
return projection, geotransform, no_data_value, data_type
def get_matrix_list(image_list):
mat_list = []
for img in image_list:
dataset = gdal.Open(img)
product_array = dataset.GetRasterBand(1).ReadAsArray()
mat_list.append(product_array)
dataset = None
print(type(product_array))
return mat_list
def calc_anomaly(agg_file, LTA_file):
if agg_file and LTA_file:
agg_and_LTA = get_matrix_list([agg_file, LTA_file])
print('Aggregation and LTA converted to matrices')
print(agg_and_LTA[0].dtype)
print(agg_and_LTA[1].dtype)
anomaly_values = (agg_and_LTA[0] * 1.0) - (agg_and_LTA[1] * 1.0)
print(anomaly_values.dtype)
projection, geotransform, no_data_value, data_type = get_metadata(agg_file)
return anomaly_values, projection, geotransform, no_data_value, data_type
else:
return None, None, None
def write_output_image(filepath, output_matrix, image_format, data_format, mask=None, output_projection=None, output_geotransform=None, no_data_value=None):
driver = gdal.GetDriverByName(image_format)
out_rows = np.size(output_matrix, 0)
out_columns = np.size(output_matrix, 1)
if mask is not None and mask is not 0:
output = driver.Create(filepath, out_columns, out_rows, 2, data_format)
mask_band = output.GetRasterBand(2)
mask_band.WriteArray(mask)
if no_data_value is not None:
output_matrix[mask > 0] = no_data_value
else:
output = driver.Create(filepath, out_columns, out_rows, 1, data_format)
if output_projection is not None:
output.SetProjection(output_projection)
if output_geotransform is not None:
output.SetGeoTransform(output_geotransform)
raster_band = output.GetRasterBand(1)
if no_data_value is not None:
raster_band.SetNoDataValue(no_data_value)
raster_band.WriteArray(output_matrix)
gdal.Warp(filepath, output, format="GTiff", outputBoundsSRS='EPSG:4326', xRes=output_geotransform[1], yRes=-output_geotransform[5], targetAlignedPixels=True)
def write_anomaly_output(anomaly, output_folder, product_name, first_date, last_date, lta_start_year, lta_end_year, aggregation, mask_no_value, N_value, regionOfInterest, roi_name, projection, geo_transform, no_data_value):
filename = os.path.join(output_folder, product_name + '_Anomaly_' + roi_name + '_N' + str(N_value) + '_' + aggregation + '_' + first_date + '_' + last_date + '_LTA' + str(lta_start_year) + '_' + str(lta_end_year) + '.tif')
write_output_image(filename, anomaly, 'GTiff', gdal.GDT_Float32, mask_no_value, projection, geo_transform, no_data_value)
return filename
def get_formatted_date(date_str):
date = datetime.datetime.strftime(date_str, '%Y-%m-%dT00:00:00Z')
return date
def write_properties_file(output_name, first_date, last_date, region_of_interest):
title = 'Output %s' % output_name
first_date = get_formatted_date(first_date)
last_date = get_formatted_date(last_date)
with open(output_name + '.properties', 'wb') as file:
file.write('title=%s\n' % title)
file.write('date=%s/%s\n' % (first_date, last_date))
file.write('geometry=%s' % (region_of_interest))
# -
# #### Auxiliary folders
# +
if len(output_folder) > 0:
if not os.path.isdir(output_folder):
os.mkdir(output_folder)
if not os.path.isdir(temp_folder):
os.mkdir(temp_folder)
# -
# #### Workflow
# ##### Get metadata
# +
message = 'Getting metadata from catalog'
ciop.log('INFO', message)
# organize indexes and apikeys in a python dictionary
apikeys = {indexAgg['value']: apikeyAgg['value'], indexLTA['value']: apikeyLTA['value']}
# get input data from catalog
input_metadata = get_input_metadata (input_references, apikeys)
input_metadata_LTA = input_metadata[input_metadata['title'].str.find('LTA') != -1]
input_metadata_Agg = input_metadata[input_metadata['title'].str.find('LTA') == -1]
# -
# ##### Define pairs
len(input_metadata_LTA)
len(input_metadata_Agg)
# +
agg_lat_idx = {}
# N time steps
nlist = [N_1['value'], N_3['value'], N_6['value'], N_9['value'], N_12['value'], N_15['value'], N_18['value'], N_27['value'], N_36['value']]
nlist = [n == 'True' for n in nlist]
nvalues = [1, 3, 6, 9, 12, 15, 18, 27, 36]
for bl,nv in zip(nlist, nvalues):
# only works for selected N time steps
if bl:
subN_input_metadata_Agg = input_metadata_Agg[input_metadata_Agg['title'].str.contains('_N(?:{})_'.format(nv))]
subN_input_metadata_LTA = input_metadata_LTA[input_metadata_LTA['title'].str.contains('_N(?:{})_'.format(nv))]
for index_agg, row_agg in subN_input_metadata_Agg.iterrows():
#print(row['c1'], row['c2'])
agg_lat_idx[index_agg] = []
sm = pd.to_datetime(row_agg['startdate']).month
sd = pd.to_datetime(row_agg['startdate']).day
em = pd.to_datetime(row_agg['enddate']).month
ed = pd.to_datetime(row_agg['enddate']).day
for index_lta, row_lta in subN_input_metadata_LTA.iterrows():
if sm == pd.to_datetime(row_lta['startdate']).month and sd == pd.to_datetime(row_lta['startdate']).day and em == pd.to_datetime(row_lta['enddate']).month and ed == pd.to_datetime(row_lta['enddate']).day:
if row_agg['title'].split('_')[3] in row_lta['title']:
agg_lat_idx[index_agg].append(index_lta)
# -
for agg_idx in agg_lat_idx:
print((input_metadata_Agg.loc[agg_idx]['title']) , (input_metadata_LTA.loc[agg_lat_idx[agg_idx][0]]['title']))
region_of_interest = regionOfInterest['value']
name_of_region = nameOfRegion['value']
# TODO: add something to choose between LTAs list
for agg_idx in agg_lat_idx:
#print((input_metadata_Agg.loc[agg_idx]['title']) , (input_metadata_LTA.loc[agg_lat_idx[agg_idx][0]]['title']))
if len(agg_lat_idx[agg_idx]) < 1:
continue
sub_input_metadata_Agg = input_metadata_Agg.loc[agg_idx]
sub_input_metadata_LTA = input_metadata_LTA.loc[agg_lat_idx[agg_idx][0]]
# get data paths from catalog metadata
filepath_agg = os.path.join(data_path, sub_input_metadata_Agg['enclosure'].split('/')[-1])
filepath_LTA = os.path.join(data_path, sub_input_metadata_LTA['enclosure'].split('/')[-1])
print(filepath_agg)
print(filepath_LTA)
# get metadata from catalog metadata (Agg and LTA)
# Agg
file_name_elements = os.path.basename(filepath_agg).split('.')[0].split('_')
agg_type = file_name_elements[-3]
Nn = file_name_elements[-4]
nv = int(Nn.split('N')[-1])
first_date = sub_input_metadata_Agg['startdate'].strftime('%Y-%m-%d')
last_date = sub_input_metadata_Agg['enddate'].strftime('%Y-%m-%d')
print(nv)
print(first_date)
print(last_date)
# LTA
file_name_elements = os.path.basename(filepath_LTA).split('.')[0].split('_')
agg_type_LTA = file_name_elements[-5]
Nn_LTA = file_name_elements[-6]
start_year = str(sub_input_metadata_LTA['startdate'].year)
end_year = str(sub_input_metadata_LTA['enddate'].year)
print(start_year)
print(end_year)
message = 'Computing Anomaly'
ciop.log('INFO', message)
anomaly_values, projection, geotransform, no_data_value, data_type = calc_anomaly(filepath_agg, filepath_LTA)
message = 'Writing anomaly image'
ciop.log('INFO', message)
filename = write_anomaly_output(anomaly_values, output_folder, 'LST', first_date, last_date, start_year, end_year, agg_type, None, nv, region_of_interest, name_of_region, projection, geotransform, no_data_value)
write_properties_file(filename, datetime.datetime.strptime(first_date, "%Y-%m-%d").date(), datetime.datetime.strptime(last_date, "%Y-%m-%d").date(), region_of_interest)
if check_results:
import matplotlib
import matplotlib.pyplot as plt
fig = plt.figure()
plt.imshow(anomaly_values)
plt.show()
# #### Remove temporay files and folders
# +
rm_cfolder(temp_folder)
os.rmdir(temp_folder)
| src/main/app-resources/notebook/libexec/input.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Classifier drift detector on CIFAR-10
#
# ### Method
#
# The classifier-based drift detector simply tries to correctly classify instances from the reference data vs. the test set. If the classifier does not manage to significantly distinguish the reference data from the test set according to a chosen metric (defaults to the classifier accuracy), then no drift occurs. If it can, the test set is different from the reference data and drift is flagged. To leverage all the available reference and test data, stratified cross-validation can be applied and the out-of-fold predictions are used to compute the drift metric. Note that a new classifier is trained for each test set or even each fold within the test set.
#
# ### Dataset
#
# [CIFAR10](https://www.cs.toronto.edu/~kriz/cifar.html) consists of 60,000 32 by 32 RGB images equally distributed over 10 classes. We evaluate the drift detector on the CIFAR-10-C dataset ([Hendrycks & Dietterich, 2019](https://arxiv.org/abs/1903.12261)). The instances in
# CIFAR-10-C have been corrupted and perturbed by various types of noise, blur, brightness etc. at different levels of severity, leading to a gradual decline in the classification model performance. We also check for drift against the original test set with class imbalances.
# +
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
from tensorflow.keras.layers import Conv2D, Dense, Flatten, Input
from alibi_detect.cd import ClassifierDrift
from alibi_detect.utils.saving import save_detector, load_detector
from alibi_detect.datasets import fetch_cifar10c, corruption_types_cifar10c
# -
# ### Load data
#
# Original CIFAR-10 data:
(X_train, y_train), (X_test, y_test) = tf.keras.datasets.cifar10.load_data()
X_train = X_train.astype('float32') / 255
X_test = X_test.astype('float32') / 255
y_train = y_train.astype('int64').reshape(-1,)
y_test = y_test.astype('int64').reshape(-1,)
# For CIFAR-10-C, we can select from the following corruption types at 5 severity levels:
corruptions = corruption_types_cifar10c()
print(corruptions)
# Let's pick a subset of the corruptions at corruption level 5. Each corruption type consists of perturbations on all of the original test set images.
corruption = ['gaussian_noise', 'motion_blur', 'brightness', 'pixelate']
X_corr, y_corr = fetch_cifar10c(corruption=corruption, severity=5, return_X_y=True)
X_corr = X_corr.astype('float32') / 255
# We split the original test set in a reference dataset and a dataset which should not be flagged as drift. We also split the corrupted data by corruption type:
np.random.seed(0)
n_test = X_test.shape[0]
idx = np.random.choice(n_test, size=n_test // 2, replace=False)
idx_h0 = np.delete(np.arange(n_test), idx, axis=0)
X_ref,y_ref = X_test[idx], y_test[idx]
X_h0, y_h0 = X_test[idx_h0], y_test[idx_h0]
print(X_ref.shape, X_h0.shape)
X_c = []
n_corr = len(corruption)
for i in range(n_corr):
X_c.append(X_corr[i * n_test:(i + 1) * n_test])
# We can visualise the same instance for each corruption type:
# +
i = 6
n_test = X_test.shape[0]
plt.title('Original')
plt.axis('off')
plt.imshow(X_test[i])
plt.show()
for _ in range(len(corruption)):
plt.title(corruption[_])
plt.axis('off')
plt.imshow(X_corr[n_test * _+ i])
plt.show()
# -
# ### Detect drift
#
# We use a simple classification model and try to distinguish between the reference data and the corrupted test sets. Initially we'll use an accuracy threshold set at $0.55$, use $75$% of the shuffled reference and test data for training and evaluate the detector on the remaining $25$%. We only train for 1 epoch.
# +
tf.random.set_seed(0)
model = tf.keras.Sequential(
[
Input(shape=(32, 32, 3)),
Conv2D(8, 4, strides=2, padding='same', activation=tf.nn.relu),
Conv2D(16, 4, strides=2, padding='same', activation=tf.nn.relu),
Conv2D(32, 4, strides=2, padding='same', activation=tf.nn.relu),
Flatten(),
Dense(2, activation='softmax')
]
)
cd = ClassifierDrift(threshold=.55, model=model, X_ref=X_ref, train_size=.75, epochs=1)
# we can also save/load an initialised detector
filepath = 'my_path' # change to directory where detector is saved
save_detector(cd, filepath)
cd = load_detector(filepath)
# +
preds_h0 = cd.predict(X_h0)
labels = ['No!', 'Yes!']
print('Drift? {}'.format(labels[preds_h0['data']['is_drift']]))
# -
# As expected, no drift occurred. The accuracy of the classifier is close to random and below the threshold:
print(f"accuracy: {preds_h0['data']['accuracy']} -- threshold: {preds_h0['data']['threshold']}")
# Let's now check the predictions on the corrupted data:
for x, c in zip(X_c, corruption):
preds = cd.predict(x)
print(f'Corruption type: {c}')
print(f"Drift? {labels[preds['data']['is_drift']]}")
print(f"Accuracy: {preds['data']['accuracy']}")
print('')
# The classifier could easily distinguish the corrupted from the reference data.
#
# ### Use all the available data
#
# So far we've only used $25$% of the data to detect the drift since $75$% is used for training purposes. At the cost of additional training time we can however leverage all the data via stratified cross-validation. We just need to set the number of folds and keep everything else the same. So for each test set `n_folds` models are trained, and the out-of-fold predictions combined for the final drift metric (in this case the accuracy):
cd = ClassifierDrift(threshold=.55, model=model, X_ref=X_ref, n_folds=5, epochs=1)
for x, c in zip(X_c, corruption):
preds = cd.predict(x)
print(f'Corruption type: {c}')
print(f"Drift? {labels[preds['data']['is_drift']]}")
print(f"Accuracy: {preds['data']['accuracy']}")
print('')
preds_h0 = cd.predict(X_h0)
print(f"Drift? {labels[preds_h0['data']['is_drift']]}")
print(f"Accuracy: {preds_h0['data']['accuracy']}")
# ### Customize the drift metric
#
# The drift metric can be adjusted which is very helpful in cases when there is for instance class imbalance as the test dataset sizes can vary by batch. Any function taking `y_true` and `y_pred` als input can be used as drift metric. In the following we'll use the $F1$-score as an illustration:
# +
from sklearn.metrics import f1_score
def f1_adj(y_true: np.ndarray, y_pred: np.ndarray) -> float:
return f1_score(y_true, np.round(y_pred)) # model returns soft predictions, not class labels
# -
cd = ClassifierDrift(threshold=.55, model=model, X_ref=X_ref, n_folds=5, epochs=1, metric_fn=f1_adj)
for x, c in zip(X_c, corruption):
preds = cd.predict(x)
print(f'Corruption type: {c}')
print(f"Drift? {labels[preds['data']['is_drift']]}")
print(f"F1 score: {preds['data']['f1_adj']}")
print('')
# +
preds_h0 = cd.predict(X_h0)
print('Drift? {}'.format(labels[preds_h0['data']['is_drift']]))
print(f"F1 score: {preds_h0['data']['f1_adj']}")
| examples/cd_clf_cifar10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import re
# ## Population preprocessing
# +
file = "../data/population_suisse_clean.xlsx"
df_pop = pd.read_excel(file)
df_pop = df_pop.reset_index()
# -
# Remove district information
df_pop.drop(df_pop['index'].str.startswith(">>"), inplace=True)
# Remove NPZ information and parenthesis information
# +
clean_name = lambda s: s.partition(' ')[2]
remove_sup = lambda s: re.sub(r'\(*\ [^)]*\)', '', s)
df_pop['index'] = df_pop['index'].apply(clean_name)
df_pop['index'] = df_pop['index'].apply(remove_sup)
# -
name2pop = {}
def create_dict(row):
name2pop.update({row['index']: row["Etat de la population"]})
a = df_pop.apply(create_dict, axis=1)
# --------
# ## Capacity preprocessing
file = '../data/2017-01-30_out.csv'
df_train = pd.read_csv(file)
# Clean stop id for aggregation
# +
clean_id = lambda s : s.partition(':')[0]
df_train['stop_id'] = df_train['stop_id'].apply(clean_id)
# -
# Aggregate
most_present = lambda x: x.value_counts().index[0]
df = df_train.groupby(['stop_id']).agg({'name': most_present,'id': 'count'})
df["amount_of_train"] = df.id
df = df[["amount_of_train", "name"]]
df
def get_pop(x):
if x in name2pop:
return name2pop[x]
x1 = x.split(" ")[0]
if x1 in name2pop:
return name2pop[x1]
x2 = x.split("-")[0]
if x2 in name2pop:
return name2pop[x2]
df['pop'] = df.name.apply(get_pop)
df_okay = df[~df['pop'].isnull()]
sum(df['pop'].isnull())
df_okay.plot.scatter(x='amount_of_train', y='pop', logy=True);
name2pop["Prilly"]
df_okay[df_okay.amount_of_train < 3]
df_okay
| preprocessing/population_processing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# +
# Plots Pareto distribution
import numpy as np
import matplotlib.pyplot as plt
try:
import probml_utils as pml
except ModuleNotFoundError:
# %pip install -qq git+https://github.com/probml/probml-utils.git
import probml_utils as pml
from scipy.stats import pareto
params = [(0.1, 1), (0.1, 2), (0.2, 1), (0.2, 2)]
styles = ["b-", "r:", "k-.", "g--"]
labels = ["m={:.2f}, k={:.2f}".format(m, k) for m, k in params]
x = np.linspace(0, 1, 1000)
for i, param in enumerate(params):
m, k = param
probabilities = pareto.pdf(x, k, scale=m)
plt.plot(x, probabilities, styles[i], label=labels[i])
plt.title("Pareto Distribution")
plt.legend()
plt.axis((0.0, 0.5, 0, 20))
pml.savefig("paretoPdf.pdf")
plt.show()
for i, param in enumerate(params):
m, k = param
probabilities = pareto.pdf(x, k, scale=m)
plt.loglog(x, probabilities, styles[i], label=labels[i])
plt.xlim(0.05, 1)
plt.title("Log Pareto Distribution")
plt.legend()
pml.savefig("paretoLogPdf.pdf")
plt.show()
| notebooks/book2/02/pareto_dist_plot.ipynb |
# +
"""
Play RPS w/Input
"""
p1 = input('player 1 choice: ') # from user input
p2 = input('player 2 choice: ') # from user input
if p1 == p2:
print(0)
elif p1 == 'r' and p2 == 's':
print(1)
elif p1 == 'r' and p2 == 'p':
print(2)
elif p1 == 'p' and p2 == 's':
print(2)
elif p1 == 'p' and p2 == 'r':
print(1)
elif p1 == 's' and p2 == 'r':
print(2)
elif p1 == 's' and p2 == 'p':
print(1)
# Given a p1 and p2
# print 1 if p1 has won
# print 2 if p2 has won
# print 0 if tie
# print -1 if invalid input
# expects both p1 and p2 inputs to be either
# "r", "p", or "s"
| pset_conditionals/rps/solution/nb/p3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
import pandas as pd
import numpy as np
# File to Load
file_to_load = "Resources/purchase_data.csv"
# Read Purchasing File and store into Pandas data frame- display the CSV with Pandas
purchase_data = pd.read_csv(file_to_load)
purchase_data.head(40)
# -
# ## Player Count
# * Display the total number of players
#
purchase_data["Price"]
# +
#
players_data = purchase_data.loc[:,['SN','Age','Gender']]
players_data = players_data.drop_duplicates()
players_data
#Count the number of players
player_counts = players_data.count()[0]
player_counts
#Create a data frame
player_counts_df = pd.DataFrame({"Player Counts": [player_counts]})
player_counts_df
# -
# ## Purchasing Analysis (Total)
# * Run basic calculations to obtain number of unique items, average price, etc.
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
#
# +
#average price
item_price = purchase_data["Price"].mean()
item_price
#Total revenue
total_revenue = purchase_data["Price"].sum()
total_revenue
#number of purchases
total_purchases = purchase_data["Price"].count()
total_purchases
#number of unique items
unique_items = purchase_data["Item ID"].unique()
unique_items = len(unique_items)
#create a data frame
purchasing_analysis_df = pd.DataFrame({"Item Price": [item_price],"Total Revenue": [total_revenue],
"Number of Purchases": [total_purchases], "Unique Items": unique_items})
purchasing_analysis_df
# -
# ## Gender Demographics
# * Percentage and Count of Male Players
#
#
# * Percentage and Count of Female Players
#
#
# * Percentage and Count of Other / Non-Disclosed
#
#
#
# +
#Count of male players
male_players = players_data["Gender"].value_counts()
male_players
percentage_counts = male_players/player_counts*100
percentage_counts
gender_demographics_df = pd.DataFrame({"Player Gender Count": male_players,
"Percent Distribution": percentage_counts})
gender_demographics_df
#Count of female players
#female_players - players_data["Female"].count()
#female_players
# -
#
# ## Purchasing Analysis (Gender)
# * Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. by gender
#
#
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
# +
basic_calculations = purchase_data.groupby("Gender").count()["Price"]
basic_calculations
basic_sum = purchase_data.groupby("Gender").sum()["Price"]
basic_sum
purchase_gender = basic_sum/male_players
purchase_gender
mean_counts = purchase_data.groupby("Gender").mean()["Price"]
mean_counts
summary_df = pd.DataFrame({"Purchase Total": basic_sum, "Purchase Count": basic_calculations, "Avg. Purchase Price by Person": purchase_gender,
"Avg. Purchase By Gender": mean_counts})
summary_df
# -
# ## Age Demographics
# * Establish bins for ages
#
#
# * Categorize the existing players using the age bins. Hint: use pd.cut()
#
#
# * Calculate the numbers and percentages by age group
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: round the percentage column to two decimal points
#
#
# * Display Age Demographics Table
#
# +
#establish bins for ages
bins = [0,9.9,14.9,19.9,24.9,29.9,34.9,39.9,150]
labels = ["<10","10-14","15-19","20-24","25-29","30-34","35-39","40+"]
purchase_data["Age Ranges"] = pd.cut(purchase_data["Age"],bins,labels=labels)
purchase_data
bin_counts = purchase_data.groupby(["Age Ranges"])
bin_counts
#count players by ages
age_count = bin_counts["SN"].nunique()
bin_percents = (age_count/int(player_counts)) * 100
bin_percents
#Create a summary data frame for results
age_summary = pd.DataFrame({"Percent of Players": bin_percents, "Total": age_count})
#optional- format with two decimal places
age_summary.style.format({"Percent of Players":"{:,.2f}"})
# -
# ## Purchasing Analysis (Age)
# * Bin the purchase_data data frame by age
#
#
# * Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. in the table below
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
# +
#Count purchases by age group
purchase_countby_age = bin_counts["Purchase ID"].count()
#Avg purchase price by bin
avg_priceby_bin = bin_counts["Price"].mean()
#Purchase sum by bin
sum_purchase_age = bin_counts["Price"].sum()
#Avg purchase price per person, per bin
avg_purch_by_bin = (sum_purchase_age/age_count)
#create df
purchasing_analysis = pd.DataFrame({"Purchase Count": purchase_countby_age,
"Average Purchase Price": avg_priceby_bin,
"Total Purchase": sum_purchase_age,
"Average Purchases per Person": avg_purch_by_bin})
purchasing_analysis
# -
# ## Top Spenders
# * Run basic calculations to obtain the results in the table below
#
#
# * Create a summary data frame to hold the results
#
#
# * Sort the total purchase value column in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the summary data frame
#
#
# +
#find the top spenders
top_spenders = purchase_data.groupby(["SN"])
#Count total purchases by player name
purchase_count_player = top_spenders["Purchase ID"].count()
#Calculate total purchases
total_purchases_player = top_spenders["Price"].sum()
#calculate average purchase price
avg_purchby_player = (total_purchases_player/player_counts)
#avg_purchby_player.style.format{(('${0:,.0f}'))}
#Create data frame
top_spenders_summary = pd.DataFrame({"Purchase Count": purchase_count_player,
"Total Purchases in Dollars": total_purchases_player,
"Average Purchase per Player": avg_purchby_player})
#top_spenders_summary_df.style.format(('${0:,.0f}'))
#top_spenders_summary_df=(by="Total Purchases in Dollars", ascending=False)
spenders_df = top_spenders_summary.sort_values("Total Purchases in Dollars", ascending=False)
spenders_df.head()
# -
# ## Most Popular Items
# * Retrieve the Item ID, Item Name, and Item Price columns
#
#
# * Group by Item ID and Item Name. Perform calculations to obtain purchase count, item price, and total purchase value
#
#
# * Create a summary data frame to hold the results
#
#
# * Sort the purchase count column in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the summary data frame
#
#
# +
#Group by item id and item name
popular_group_df = purchase_data.groupby(["Item ID", "Item Name"])
popular_group_df.count()
#add more information in
popular_analysis_df = pd.DataFrame(popular_group_df["Purchase ID"].count())
popular_analysis_df
#total purchases by item
total_purchase_value = popular_group_df["Price"].sum()
total_purchase_value
total_purchase_value_dollars = total_purchase_value.map("${:,.2f}".format)
total_purchase_value_dollars
#purchase price of most popular items
popular_purchase_price = popular_group_df["Price"].mean()
popular_purchase_price
popular_purchase_price_dollars = popular_purchase_price.map("${:,.2f}".format)
popular_purchase_price_dollars
#popular items summary data- combine data frames
popular_analysis_df["Item Price"] = popular_purchase_price_dollars
popular_analysis_df["Total Purchase Value"] = total_purchase_value_dollars
popular_analysis_df.head()
#sort in ascending order
#popular_sorted = popular_analysis_df.sort_values("Purchase Count", ascending=True)
#popular_sorted.head()
# -
# ## Most Profitable Items
# * Sort the above table by total purchase value in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the data frame
#
#
# +
#sort table in descending order
popular_analysis_df = popular_analysis_df.sort_values(by="Total Purchase Value", ascending=False)
popular_analysis_df["Item Price"] = popular_analysis_df["Item Price"]
popular_analysis_df["Total Purchase Value"] = popular_analysis_df["Total Purchase Value"]
popular_analysis_df.head()
# -
| HeroesOfPymoli/Pandas_Challenge_Final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # What’s new in Python 3.8?
#
# In Python 3.8, the syntax is simplified and support for C libraries is also improved. Below is a brief overview of some of the new features. You can get a complete overview in [What’s New In Python 3.8](https://docs.python.org/3/whatsnew/3.8.html).
# ## Installation
#
# ### Check
# !python3 -V
# or
import sys
assert sys.version_info[:2] >= (3, 8)
# ## Assignment Expressions: Walrus operator `:=`
#
# So far, e.g. `env_base` can be determined by pip as follows:
import os
def _getuserbase():
env_base = os.environ.get("PYTHONUSERBASE", None)
if env_base:
return env_base
# This can now be simplified with:
def _getuserbase():
if env_base := os.environ.get("PYTHONUSERBASE", None):
return env_base
# Multiple nested `if`, such as in [cpython/Lib/copy.py](https://github.com/python/cpython/blob/master/Lib/copy.py), can also be avoided. This
from copyreg import dispatch_table
def copy(x):
cls = type(x)
reductor = dispatch_table.get(cls)
if reductor:
rv = reductor(x)
else:
reductor = getattr(x, "__reduce_ex__", None)
if reductor:
rv = reductor(4)
else:
reductor = getattr(x, "__reduce__", None)
if reductor:
rv = reductor()
else:
raise Error(
"un(deep)copyable object of type %s" % cls)
# becomes that:
def copy(x):
cls = type(x)
reductor = dispatch_table.get(cls)
if reductor := dispatch_table.get(cls):
rv = reductor(x)
elif reductor := getattr(x, "__reduce_ex__", None):
rv = reductor(4)
elif reductor := getattr(x, "__reduce__", None):
rv = reductor()
else:
raise Error("un(deep)copyable object of type %s" % cls)
# ## *Positional-only* parameters
#
# In Python 3.8 a function parameter can be specified position-related with `/`. Several Python functions implemented in C do not allow keyword arguments. This behavior can now be emulated in Python itself, e.g. for the [pow()](https://docs.python.org/3/library/functions.html#pow) function:
def pow(x, y, z=None, /):
"Emulate the built in pow() function"
r = x ** y
return r if z is None else r%z
# ## `f-strings` support `=` for self-documenting expressions and debugging
user = 'veit'
member_since = date(2012, 1, 30)
f'{user=} {member_since=}'
# ## Debug and release build use the same ABI
#
# So far, a consistent application binary interface (ABI) should be guaranteed by [Spack](../../reproduce/spack/). However, this did not include using Python in the debug build. Python 3.8 now also supports ABI compatibility for debug builds. The `Py_TRACE_REFS` macro can now be set with the `./configure --with-trace-refs` option.
# ## New C API
#
# [PEP 587](https://www.python.org/dev/peps/pep-0587/) adds a new C API for configuring the Python initialisation, which offers more precise control of the entire configuration and better error reports.
# ## Vectorcall - a fast protocol for CPython
#
# The protocol is not yet fully implemented; this will probably come with Python 3.9. However, you can already get a full description in [PEP 590](https://www.python.org/dev/peps/pep-0590).
# ## Update – or not?
#
# The following is a brief overview of the problems you may encounter when switching to Python 3.8:
#
# ### Missing packages
#
# * [opencv-python](https://pypi.org/project/opencv-python/#files)
#
# ### Bugs
#
# * Python 3.7.1 was released 4 months after the first major release with a [long list of bug fixes](https://docs.python.org/3.7/whatsnew/changelog.html#python-3-7-1-final) . Something similar is to be expected with Python 3.8.
#
# ### Syntax
#
# * Very few code analysis tools and autoformatters can already handle the syntax changes of Python 3.8
#
# ### Why update anyway?
#
# Since the upgrade will take some time, it can be tempting to postpone the move indefinitely. Why should you concern yourself with incompatibilities in new versions when your current version works reliably?
#
# The problem is that your Python is not supported indefinitely, nor will the libraries you use will support all older Python versions indefinitely. And the longer you delay an update, the bigger and riskier it will be. Therefore, the update to the new major version of Python is usually recommended a few months after the first release.
# ## Porting
#
# > See [Porting to Python 3.8](https://docs.python.org/3.8/whatsnew/3.8.html#porting-to-python-3-8)
| docs/workspace/jupyter/kernels/python38.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
from random import choices
import pickle
import scipy
# -
df_tl = pd.read_excel('D:/UW/project/6ppd-experiments/tl/20201021_Tire_leaching_data.xlsx')
df_oz = pd.read_excel('D:/UW/project/6ppd-experiments/ozonation/20201021-ozonation-6ppd.xlsx')
sns.distplot(df_tl.RT,bins=100);
sns.distplot(df_tl.MZ,bins=128);
sns.distplot(df_oz.RT,bins=100);
sns.distplot(df_oz.MZ,bins=128);
df_na = pd.read_excel('D:/UW/massmotif/mzml/20201106_raw_DRO_DIE.xlsx')
sns.distplot(df_na.mz,bins=128);
sns.distplot(df_na.RT,bins=100);
df_0815 = pd.read_csv('D:/UW/directproject/example_data/clustering/sample0815.csv')
df_1114 = pd.read_csv('D:/UW/directproject/example_data/clustering/sample1114.csv')
df_kathy = pd.read_csv('D:/UW/project/New projects 20200714/exported data in wide format/MSpos-AFFF4_wide_features_all-data_for-Ximin.csv')
sns.distplot(df_0815['Average Rt(min)'],bins=100);
sns.distplot(df_0815['Average Mz'],bins=128);
sns.distplot(df_1114['Average Rt(min)'],bins=100);
sns.distplot(df_1114['Average Mz'],bins=128);
sns.distplot(df_kathy.rt,bins=100);
sns.distplot(df_kathy.am,bins=128);
totrt = list(df_tl.RT) + list(df_oz.RT) + list(df_na.RT) + list(df_kathy.rt) + list(df_0815['Average Rt(min)']) + list(df_1114['Average Rt(min)'])
totmz = list(df_tl.MZ) + list(df_oz.MZ) + list(df_na.mz) + list(df_kathy.am) + list(df_0815['Average Mz']) + list(df_1114['Average Mz'])
counts_mz, bins_mz, bars = plt.hist(df_oz.MZ, bins=128)
counts_rt, bins_rt, bars = plt.hist(df_oz.RT, bins=100)
sns.distplot(totrt,bins=100);
print('tl data',df_tl.shape)
print('oz data',df_oz.shape)
print('nina data',df_na.shape)
print('0815 data',df_0815.shape)
print('1114 data',df_1114.shape)
print('kathy data',df_kathy.shape)
#Generating random mz & rt -- source
source_pair = []
for i in np.arange(0,20,1):
s = 200
rdm_mz = np.random.choice(bins_mz[:-1], size=s, p=[i/counts_mz.sum() for i in counts_mz])
rdm_mz_adj = [i+np.random.uniform(0, bins_mz[1] - bins_mz[0]) for i in rdm_mz]
rdm_rt = np.random.choice(bins_rt[:-1], size=s, p=[i/counts_rt.sum() for i in counts_rt])
rdm_rt_adj = [i+np.random.uniform(0, bins_rt[1] - bins_rt[0]) for i in rdm_rt]
source_pair.append([rdm_mz_adj, rdm_rt_adj])
#Generating random mz & rt -- background tot
background_pair = []
for i in np.arange(0,200,1):
s = 10000
rdm_mz = np.random.choice(bins_mz[:-1], size=s, p=[i/counts_mz.sum() for i in counts_mz])
rdm_mz_adj = [i+np.random.uniform(0, bins_mz[1] - bins_mz[0]) for i in rdm_mz]
rdm_rt = np.random.choice(bins_rt[:-1], size=s, p=[i/counts_rt.sum() for i in counts_rt])
rdm_rt_adj = [i+np.random.uniform(0, bins_rt[1] - bins_rt[0]) for i in rdm_rt]
background_pair.append([rdm_mz_adj, rdm_rt_adj])
with open('./dummydata/randsource_missing_pattern.data', 'wb') as filehandle:
# store the data as binary data stream
pickle.dump(randsource, filehandle)
with open('./dummydata/sourcepair.data', 'rb') as filehandle:
# read the data as binary data stream
test = pickle.load(filehandle)
random.uniform(-200*10*1e-6,200*10*1e-6)
# +
#Generate missing source
import random
randsource = []
for s in test:
for j in np.arange(1,6):
randindex = sorted(random.sample(list(np.arange(0,200)),random.randrange(40, 200)))
r_l = [rt + random.uniform(-0.2,0.2) for rt in s[1]]
m_l = [mz + random.uniform(-200*10*1e-6,200*10*1e-6)for mz in s[0]]
randlist = [[m_l[i] for i in randindex],[r_l[i] for i in randindex]]
randsource.append(randlist)
# -
#Generate missing source
import random
shiftsource = []
for s in test:
for j in np.arange(1,6):
shiftlist = [i + random.uniform(-5, 5) for i in s[1]]
for rt in shiftlist:
rt += random.uniform(-0.2,0.2)
retainindex = [i for i,j in enumerate(shiftlist) if j > 0 and j < 22]
mzlist = s[0]
for mz in mzlist:
mz += random.uniform(-200*5*1e-6,200*5*1e-6)
randlist = [[mzlist[i] for i in retainindex],[shiftlist[j] for j in retainindex]]
shiftsource.append(randlist)
with open('./dummydata/randsource_missing_pattern.data', 'wb') as filehandle:
# store the data as binary data stream
pickle.dump(randsource, filehandle)
scipy.stats.ks_2samp(df_tl.MZ,df_na.mz)
#Steps: 1 alignment 2 cos similarity/msdial algorithm 3 distribution check? Supplement: check coverage and do shift check?
#Method 1
from sklearn.metrics.pairwise import cosine_similarity
A=np.array([7,3]).reshape(1,-1)
B=np.array([7,3]).reshape(1,-1)
cosine_similarity(A,B)
#Method 2 msdial algorithm
plt.scatter(randsource[0][1], randsource[0][0],s=3)
plt.scatter(bg[0][1], bg[0][0],s=3)
plt.scatter(test[0][1],test[0][0],s=3)
plt.scatter(test[0][1],test[0][0],s=3)
plt.scatter(bg[0][1], bg[0][0],s=3,alpha=0.1)
# +
#generate shifting source
#Generate intensity?
| dev/Fingerprint-dummy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.11 64-bit (''pythae_dev'': conda)'
# name: python3
# ---
# +
# If you run on colab uncomment the following line
# #!pip install git+https://github.com/clementchadebec/benchmark_VAE.git
# +
import torch
import torchvision.datasets as datasets
# %load_ext autoreload
# %autoreload 2
# +
mnist_trainset = datasets.MNIST(root='../../data', train=True, download=True, transform=None)
train_dataset = mnist_trainset.data[:-10000].reshape(-1, 1, 28, 28) / 255.
eval_dataset = mnist_trainset.data[-10000:].reshape(-1, 1, 28, 28) / 255.
# -
from pythae.models import VAMP, VAMPConfig
from pythae.trainers import BaseTrainerConfig
from pythae.pipelines.training import TrainingPipeline
from pythae.models.nn.benchmarks.mnist import Encoder_VAE_MNIST, Decoder_AE_MNIST
# +
config = BaseTrainerConfig(
output_dir='my_model',
learning_rate=1e-3,
batch_size=100,
num_epochs=100,
)
model_config = VAMPConfig(
input_dim=(1, 28, 28),
latent_dim=16,
number_components=50
)
model = VAMP(
model_config=model_config,
encoder=Encoder_VAE_MNIST(model_config),
decoder=Decoder_AE_MNIST(model_config)
)
# -
pipeline = TrainingPipeline(
training_config=config,
model=model
)
pipeline(
train_data=train_dataset,
eval_data=eval_dataset
)
import os
last_training = sorted(os.listdir('my_model'))[-1]
trained_model = VAMP.load_from_folder(os.path.join('my_model', last_training, 'final_model'))
from pythae.samplers import VAMPSampler
# create vamp sampler
vamp_sampler = VAMPSampler(
model=trained_model
)
gen_data = vamp_sampler.sample(
num_samples=25
)
import matplotlib.pyplot as plt
# +
# show results with vamp sampler
fig, axes = plt.subplots(nrows=5, ncols=5, figsize=(10, 10))
for i in range(5):
for j in range(5):
axes[i][j].imshow(gen_data[i*5 +j].cpu().squeeze(0), cmap='gray')
axes[i][j].axis('off')
plt.tight_layout(pad=0.)
# -
from pythae.samplers import GaussianMixtureSampler, GaussianMixtureSamplerConfig
# +
# set up gmm sampler config
gmm_sampler_config = GaussianMixtureSamplerConfig(
n_components=10
)
# create gmm sampler
gmm_sampler = GaussianMixtureSampler(
sampler_config=gmm_sampler_config,
model=trained_model
)
# fit the sampler
gmm_sampler.fit(train_dataset)
# -
# sample
gen_data = gmm_sampler.sample(
num_samples=25
)
# +
# show results with gmm sampler
fig, axes = plt.subplots(nrows=5, ncols=5, figsize=(10, 10))
for i in range(5):
for j in range(5):
axes[i][j].imshow(gen_data[i*5 +j].cpu().squeeze(0), cmap='gray')
axes[i][j].axis('off')
plt.tight_layout(pad=0.)
# -
# ## ... the other samplers work the same
| examples/notebooks/models_training/vamp_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import skimage.io as io
img = io.imread('D:\\Dataset\\4.2.05.tiff')
print(type(img))
io.imshow(img)
io.show()
# +
import skimage.data as data
img = data.astronaut()
io.imshow(img)
io.show()
# -
img = data.binary_blobs(length=512, blob_size_fraction=0.1, seed=5)
io.imshow(img)
io.show()
# +
import matplotlib.pyplot as plt
img = data.coffee()
plt.imshow(img)
plt.title('Coffee')
plt.axis('off')
plt.show()
# +
img1 = data.coffee()
img2 = data.rocket()
img3 = data.horse()
img4 = data.astronaut()
titles = ['Coffee', 'Rocket', 'Horse', 'Astronaut']
images = [img1, img2, img3, img4]
for i in range(4):
plt.subplot(2, 2, i+1)
if i == 2:
plt.imshow(images[i], cmap='gray')
else:
plt.imshow(images[i])
plt.title(titles[i])
plt.axis('off')
plt.show()
# -
| Chapter 10/Section16_01_Getting_Started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## JanataHack Cross Sell Prediction
# Cross-selling identifies products or services that satisfy additional, complementary needs that are unfulfilled by the original product that a customer possesses. As an example, a mouse could be cross-sold to a customer purchasing a keyboard. Oftentimes, cross-selling points users to products they would have purchased anyways; by showing them at the right time, a store ensures they make the sale.
#
# Cross-selling is prevalent in various domains and industries including banks. For example, credit cards are cross-sold to people registering a savings account. In ecommerce, cross-selling is often utilized on product pages, during the checkout process, and in lifecycle campaigns. It is a highly-effective tactic for generating repeat purchases, demonstrating the breadth of a catalog to customers. Cross-selling can alert users to products they didn't previously know you offered, further earning their confidence as the best retailer to satisfy a particular need.
# ### Problem Statement
# Your client is an Insurance company that has provided Health Insurance to its customers now they need your help in building a model to predict whether the policyholders (customers) from past year will also be interested in Vehicle Insurance provided by the company.
#
# An insurance policy is an arrangement by which a company undertakes to provide a guarantee of compensation for specified loss, damage, illness, or death in return for the payment of a specified premium. A premium is a sum of money that the customer needs to pay regularly to an insurance company for this guarantee.
#
# For example, you may pay a premium of Rs. 5000 each year for a health insurance cover of Rs. 200,000/- so that if, God forbid, you fall ill and need to be hospitalised in that year, the insurance provider company will bear the cost of hospitalisation etc. for upto Rs. 200,000. Now if you are wondering how can company bear such high hospitalisation cost when it charges a premium of only Rs. 5000/-, that is where the concept of probabilities comes in picture. For example, like you, there may be 100 customers who would be paying a premium of Rs. 5000 every year, but only a few of them (say 2-3) would get hospitalised that year and not everyone. This way everyone shares the risk of everyone else.
#
# Just like medical insurance, there is vehicle insurance where every year customer needs to pay a premium of certain amount to insurance provider company so that in case of unfortunate accident by the vehicle, the insurance provider company will provide a compensation (called ‘sum assured’) to the customer.
#
# Building a model to predict whether a customer would be interested in Vehicle Insurance is extremely helpful for the company because it can then accordingly plan its communication strategy to reach out to those customers and optimise its business model and revenue.
#
# Now, in order to predict, whether the customer would be interested in Vehicle insurance, you have information about demographics (gender, age, region code type), Vehicles (Vehicle Age, Damage), Policy (Premium, sourcing channel) etc.
# ### Hypothesis Generation for Cross-Sell Prediction
# After understanding the problem statement and gathering the required domain knowledge, The next step comes, the hypothesis generation. This will directly spring from the problem statement.
#
# After structured thinking, below are some hypotheses stated from our problem statement-
#
# 1. Male customers are more tend to buy vehicle insurance than females.
# 2. The middle-aged customers would be more interested in the insurance offer.
# 3. Customers having a driving license are more prone to convert.
# 4. Those with new vehicles would be more interested in getting insurance.
# 5. The customers who already have vehicle insurance won’t be interested in getting another.
# 6. If the Customer got his/her vehicle damaged in the past, they would be more interested in buying insurance.
#
# The hypotheses at ready at out end, it is time to look into the data and validate the statements.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics import accuracy_score, f1_score, auc
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.ensemble import RandomForestClassifier
df = pd.read_csv('C:/Users/Princy/Downloads/car_train.csv')
df.shape
df.head()
df.info()
df.isna().sum()
# ### Exploratory Data Analysis
fix, axes = plt.subplots(2, 2, figsize = (10,10))
sns.countplot(ax = axes[0,0], x = 'Gender', hue = 'Response', data = df, palette = 'mako')
sns.countplot(ax = axes[0,1], x = 'Driving_License', hue = 'Response', data = df, palette = 'mako')
sns.countplot(ax = axes[1,0], x = 'Previously_Insured', hue = 'Response', data = df, palette = 'mako')
sns.countplot(ax = axes[1,1], x = 'Vehicle_Age', hue = 'Response', data = df, palette = 'mako')
# From the above visualizations, we can make the following inferences -
#
# 1. The male customers own slightly more vehicles than thet tend to buy insurance in comparison to their female counterparts.
#
# 2. Similarly, the customers who have driving licences will opt for insurance instead of those who don’t have it.
#
# 3. The third visualization depicts that the customers want to have only an insurance policy. It means those who already have insurance won’t convert.
#
# 4. In the last chart, the customers with vehicle age lesser than the 2 years are more likely to buy insurance.
sns.countplot(x = 'Vehicle_Damage', hue = 'Response', data = df, palette = 'mako')
# From the above plot, we can infer that if the vehicle has been damaged previously then the customer will be more interested in buying the insurance as they know the cost.
#
# It is also important to look at the target column, as it will tell us whether the problem is a balanced or an imbalanced.
Response = df.loc[:,'Response'].value_counts().rename('Count')
plt.xlabel('Response')
plt.ylabel('Count')
sns.barplot(Response.index, Response.values, palette = 'mako')
sns.distplot(df['Age'])
sns.distplot(df['Annual_Premium'])
# ### Data preprocessing
# 1. Converting the categorical features into dummies or doing categorical encoding.
# 2. Binning the numerical features.
# 3. Dropping the unnecessary columns like ids.
# +
def data_prep(df):
df = df.drop(columns = ['id', 'Policy_Sales_Channel', 'Vintage'])
df = pd.get_dummies(df, columns = ['Gender'], prefix = 'Gender')
df = pd.get_dummies(df, columns = ['Vehicle_Damage'], prefix = 'Damage')
df = pd.get_dummies(df, columns = ['Driving_License'], prefix = 'License')
df = pd.get_dummies(df, columns = ['Previously_Insured'], prefix = 'prev_insured')
df['Age'] = pd.cut(df['Age'], bins = [0,29, 35, 50, 100])
df['Age']= df['Age'].cat.codes
df['Annual_Premium'] = pd.cut(df['Annual_Premium'], bins = [0, 30000, 35000, 40000, 45000, 50000, np.inf])
df['Annual_Premium'] = df['Annual_Premium'].cat.codes
df['Vehicle_Age'] = df['Vehicle_Age'].map({'< 1 Year': 0, '1-2 Year': 1, '> 2 Years': 2})
df.drop(columns =['Region_Code'], inplace = True)
return df
df1 = data_prep(df)
# -
df1.head()
# ### Feature Selection
features = ['Age', 'Vehicle_Age', 'Annual_Premium', 'Gender_Female', 'Gender_Male', 'Damage_No', 'Damage_Yes',
'License_0', 'License_1', 'prev_insured_0', 'prev_insured_1']
# ### Train-Test Split
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(df1[features], df1['Response'],
test_size = 0.3, random_state = 101)
X_train.shape, X_test.shape
# ### Handling Class Imbalance using Undersampling data
from imblearn.under_sampling import RandomUnderSampler
RUS = RandomUnderSampler(sampling_strategy = .5, random_state = 3,)
X_train, Y_train = RUS.fit_resample(df1[features], df1['Response'])
# ### Model Training and Prediction
#using accuracy and F-1 score as performace metrics
#f1 score more significant for imbalanced classification problems
#defining function for models' peroformance measurement
def performance_met(model, X_train, Y_train, X_test, Y_test):
acc_train= accuracy_score(Y_train, model.predict(X_train))
f1_train = f1_score(Y_train, model.predict(X_train))
acc_test = accuracy_score(Y_test, model.predict(X_test))
f1_test = f1_score(Y_test, model.predict(X_test))
print('train score: accuracy: {} f1: {}'.format(acc_train, f1_train))
print('test_score: accuracy: {} f1: {}'.format(acc_test, f1_test))
# #### Training 3 models -
# 1. Logistic Regression
# 2. Decision Tree
# 3. Random Forest
#Logistic Regression
model = LogisticRegression()
model.fit(X_train, Y_train)
performance_met(model, X_train, Y_train, X_test, Y_test)
#Decision Tree
model_DT = DecisionTreeClassifier(random_state = 1)
model_DT.fit(X_train, Y_train)
performance_met(model_DT, X_train, Y_train, X_test, Y_test)
#Random Forest
Forest = RandomForestClassifier(random_state = 1)
Forest.fit(X_train, Y_train)
performance_met(Forest, X_train, Y_train, X_test, Y_test)
# We observe low performance from Logistic Regression and significantly higher and simiar performance from Decision Tree and Random Forest models.
# ### Hyperparameter Tuning
#Using GridSearch for finding best parameters for random forest classifier
rf = RandomForestClassifier(random_state = 1)
parameters = {
'bootstrap': [True],
'max_depth': [20,25],
'min_samples_leaf': [3,4],
'min_samples_split': [100,300],
}
grid_search_1 = GridSearchCV(rf, parameters, cv = 3, verbose = 2, n_jobs = -1)
grid_search_1.fit(X_train, Y_train)
performance_met(grid_search_1, X_train, Y_train, X_test, Y_test)
| JanataHack-Cross-Sell-Prediction (1).ipynb |
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# %matplotlib inline
import seaborn as sns
import matplotlib.pyplot as plt
sns.set_context("talk")
sns.set_style("white")
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
import numpy as np
import torch
import torch.nn as nn
import torch.distributions as dist
from torch.nn import functional as F
import pandas as pd
import math
class CorrelatedPosteriorLogProb(nn.Module):
"""Log probability of a 2D posterior that is a mixture of three Poisson distributions."""
def __init__(self):
super().__init__()
big1 = 9
big2 = 13
self.d1 = dist.Poisson(torch.Tensor([0.01, 0.01]))
self.d2 = dist.Poisson(torch.Tensor([0.01, big1]))
self.d3 = dist.Poisson(torch.Tensor([big2, 0.01]))
self.d4 = dist.Poisson(torch.Tensor([big2, big1]))
def forward(self, z):
exp_terms = [self.d1.log_prob(z) + math.log(0.25),
self.d2.log_prob(z) + math.log(0.25),
self.d3.log_prob(z) + math.log(0.25),
self.d4.log_prob(z) + math.log(0.25)]
res = torch.stack(exp_terms)
return torch.logsumexp(res, dim=0)
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
log_prob = CorrelatedPosteriorLogProb()
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
log_prob(torch.Tensor([9, 13]))
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
L = 20
grid = np.zeros((L, L))
for i in range(L):
for j in range(L):
grid[i, j] = log_prob(torch.Tensor([i, j])).sum(-1).numpy()
data = pd.DataFrame({j: grid[:, j] for j in range(grid.shape[1])})
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
cmap = sns.cubehelix_palette(8, start=2, rot=0, dark=0, light=.95, reverse=False)
fig, ax = plt.subplots(figsize=(10, 10))
sns.heatmap(data, ax=ax, cmap=cmap, linewidths=0.1)
ax.set(title='Correlated posterior', xlabel='z1', ylabel='z2')
plt.savefig('log_p_z.png', bbox_inches='tight')
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
| notebook/plot-toy-correlated-posterior.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hamiltonian Monte Carlo
#
# As seen in previous notebooks, Metropolis sampler with proposals generated by random-walk has few (related) problems:
# - it fails in many dimensional problems
# - it often proposes states with low probability, resulting in high rejection rates
#
# To solve these problems, sampling algorithm called Hamiltonian Monte Carlo was developed ([wiki](https://en.wikipedia.org/wiki/Hamiltonian_Monte_Carlo)). There are two possible intuitive motivations for how it should work. One is purely abstract:
#
# Random walk metropolis proposes unprobable states because the exploration of the state-space is *blind*- the proposal process is completely independent on the density distribution (only rejection-acceptance depends on it). We may solve this problem by taking the shape of the distribution into account when proposing states, for example by letting the gradient of the distribution guide us towards more probable states (more accurately, towards the states with the highest contribution to any integration, i.e.: typical set).
# This motivation is explained in detail in [[1](https://arxiv.org/pdf/1701.02434.pdf)] (great paper, give it a try).
#
# Another motivation is based on analogy with physics- ordinary physical systems are great at sampling from random distributions. For example a movement of an atom in a gas- its movement consists of two processes:
# - movement according to Newton's laws of motion under the influence of external potential field (completely deterministic).
# - random collisions with other particles in a gas
#
# With this process, probability distribution of the particle's position and velocity will be such, that:
#
# $$ \begin{aligned} P(x) \propto e^{-\frac{U(x)}{T}} \\
# P(v) \propto e^{-\frac{K(v)}{T}}
# \end{aligned}$$
#
# where $U(x)$ is potential energy of the particle, $K(v)$ is it's kinetic energy (in general, it may also depend on $x$, but for an atom in a gas, it depends only on $v$) and $T$ temperature of the system. In physics this formula is called Boltzman (also canonical) distribution and is valid for all systems in thermal equilibrium, not just particle in a gas [[4](http://assets.cambridge.org/97805218/11194/excerpt/9780521811194_excerpt.pdf)]. So the idea is this:
#
# ** Let's devise such potential $U(x)$ that $P(x)\propto e^{-\frac{U(x)}{T}}$ will be our target distribution ( log probability unexpectedly appears here), simulate the motion of the physical system, add some random collisions (so that $v$ is from $P(v)$) and we should get a valid sampler for $x$.**
#
# Let us repeat this thinking. If we have a high-dimensional problem where we need to sample from some $P(x)$, which is hard, we propose to double state size by adding in a momentum term $p$ with the same dimensionality and work in the joint probability space $\pi(x,v)$. This notebook is a very high-level and introductory exposition into why this works.
#
# Both motivations would lead us to the same sampling algorithm. In this notebook we will show one possible implementation of such sampler, introduce some theoretical concepts usefull for reasoning about it and try it on some problem where random walk metropolis fails. You can find links to further reading at the end of the notebook.
# ## Hamiltonian Monte Carlo algorithm
#
# HMC sampler algorithm works conceptually like this:
#
# - devise suitable (physical) system with potential energy $U(x)/T=-log(P(x))$ and embed it in a joint probability space $\pi(x,v)$ together wih a velocity term such that $\pi(x,v) \propto P(x)P(v)$.
# - at each iteration:
# 1. give the system random kick by sampling the new velocity $v \sim P(v)$
# 2. simulate its evolution according to mechanical laws of motion for some predetermined time
# 3. correct numerical errors
# 4. return its position
#
# We can view the above algorithm as a two-step process [3,7].
#
# The item 1 is a Gibbs step. This is true although we sample the velocity independently. As noted above, the HMC algorithm works in the joint probability space $\pi(x,v)$, which we then decompose as
# $$\pi(x,v) = \pi(v|x)P(x) = P(v)P(x).$$
# In other words sampling from $P(v)$ is the same as sampling from $\pi(v|x)$ which we recognize as the *full conditional* required for Gibbs sampling of $v$ that leaves the distribution $\pi(x,v)$ invariant. Gibbs sampling does not entail an accept/reject state, all samples are accepted.
#
# The items 2 and 3 together constitute a Metropolis step which proposes a new value for both $x$ and $v$ at the same time. If we satisfy some technical conditions, then the HMC integration is a valid reversible (symmetric) proposal that satisfies detailed balance. Thus the proposal satisfies the requirements of being a Metropolis proposal distribution. The proposal is subject to an accept-reject step as required for Metropolis proposals. However there is a twist: exact integration would leave the value $\pi(x,v)$ unchanged and thus the acceptance ration would always be 1. In practice, however, roundoff error can introduce slight changes in the value of $\pi(x,v)$ for the proposal $\pi(x', v')$ *but* we can use the Metropolis accept/reject step to correct for this error.
#
# The original name for HMC was Hybrid Monte Carlo [8], because the method is a hybrid of the two steps: a Gibbs sampling step for velocity/momentum and then a Metropolis step for both variables at the same time.
# We will now go through these steps (in slightly different order).
# ### Simulation of the time evolution of the system
# The HMC is named by the fact that time evolution of the system is usually described using Hamiltonian formulation of classical mechanics. If a system has potential energy $U(x)$ and kinetic energy $K(x,p)$, it's total energy (also called hamiltonian) is $H=U(x)+K(x,p)$, where $x$ and $p$ are generalised position and momentum vectors (can be many dimensional). Then the evolution of the system in time will be a solution of these equations:
# $$\begin{aligned}\dot{x} &= \frac{\partial H}{\partial p} \\
# \dot{p} &= - \frac{\partial H}{\partial x}
# \end{aligned}$$
#
# For mechanical systems, these equations are just a reformulation of ordinary laws of motion- the first one is definition of momentum in terms of velocity and second equation is the second Newton's law (with $- \frac{d U(x)}{d x}$ being the force acting on the system). Hamiltonian formulation is used because:
# - equations of motion have elegant symmetry that makes theoretical proofs easier
# - energy is natural concept in Hamiltonian formulation, which is good for us because potential energy corresponds directly to target probability distribution
#
# Good way to solve (integrate) these equations numerically is *leapfrog algorithm* (also called Velocity-Verlet and many other different names). It has one property quite important for this application: it conserves the energy of the system over long integration times, thus avoiding the slow drift in total energy due to finite time step. One can find claims that it's due to time reversibility of the leapfrog algorithm. This [link](http://physics.ucsc.edu/~peter/242/leapfrog.pdf) suggests it's not the only necessary condition.
#
# Single step of leapfrog algorithm for the equations above (with time step $\delta$) looks like this:
#
# $$\begin{aligned} p(t+\frac{\delta}{2}) &= p(t)-\frac{\delta}{2} \frac{\partial H}{\partial x}\Bigr|_{t} \\
# x(t+\delta) &= x(t) + \delta \frac{\partial H}{\partial p}\Bigr|_{t+\frac{\delta}{2}}\\
# p(t+\delta) &= p(t+\frac{\delta}{2})-\frac{\delta}{2} \frac{\partial H}{\partial x}\Bigr|_{t+\delta}\end{aligned}$$
#
# Keep in mind that this algorithm doesn't prevent numerical errors due to rounding, just drift in energy due to finite time step, so the rounding error has to be corrected in another step of the HMC.
#
# ### Choice of correct physical system ( = correct Hamiltonian)
#
# How to choose hamiltonian which will lead to sampling from my target distribution? Simplest hamiltonian is that of a single particle in a potential field (=particle in an ideal gas):
# $$H(x,v)=U(x)+\frac{1}{2}mv^2$$
# where we started to use velocity $v=\dot{x}=\frac{p}{m}$ instead of momentum. **Remember that $x$ and $v$ can be many dimensional vectors (both with the same number of dimensions).** Hamiltonians with different kinetic energy term can lead to the same distribution $P(x)$ (see the Appendix), however this choice is usually the most natural and easiest.
# HMC with this hamiltonian will lead to probability distribution $ P(x)= \frac{1}{Z} e^{-\frac{U(x)}{T}}$, so if we set $T=1$, we just have to choose U(x)=-log(P(x)) (normalization constant $Z$ can be safely left out, since it has no effect on the result).
#
# Equations of motion for this hamiltonian will be simply:
#
# $$\begin{aligned}\dot{x} &= v \\
# \dot{v} &= a= -\frac{1}{m} \frac{d U(x)}{d x}
# \end{aligned}$$
#
# (See? Just definition of velocity and Newton's second law, nothing scary.) These equations lead to the following numerical integration scheme (using leapfrog algorithm):
#
# $$\begin{aligned} v(t+\frac{\delta}{2}) &= v(t)+\frac{\delta}{2}a(t) \\
# x(t+\delta) &= x(t) + \delta v(t+\frac{\delta}{2})\\
# v(t+\delta) &= v(t+\frac{\delta}{2})+\frac{\delta}{2} a(t+\delta)\end{aligned}$$
#
# So in order to use HMC, we will have to supply not only potential function, but also its gradient (or compute it numerically).
#
# ### Random perturbations
#
# Without some random perturbation of the system its evolution would be completely deterministic and thus useless for sampling. However, random perturbations must be such that they will lead to the correct sampling of $P(x)$. This will be satisfied if during each "random kick" step, we resample momentum from (Boltzman) probability distribution $P(p)=\frac{1}{Z} exp(-K(x,p)/T)$, where $T$ is temperature (see the Appendix or trust the physics). Since we allready set $T=1$, for hamiltonian above we will sample velocity $v$ from distribution $P(v)=\frac{1}{Z}exp(-\frac{m v^2}{2})$, which is a normal distribution with variance $\sigma^2=\frac{1}{m}$.
#
#
# ### Correction of rounding errors
#
# On an ideal machine, this step would not be necessary, but in reality numerical integration will introduce rounding errors which will in time bias the sampling. Turns out that this can be exactly corrected by adding a Metropolis accept/reject stage after the leapfrog integration [[2](http://www.cs.toronto.edu/~radford/ftp/review.pdf)]. The proposed new state $(x_{prop},p_{prop})$ will be accepted with probability $$\frac{exp(-H(x_{prop},p_{prop}))}{exp(-H(x_{old},p_{old}))}$$. If it's not accepted, the system stays in its old state from before leapfrog integration.
#
# The fact that this is only correction step means that rejection should be rare and the **HMC should accept almost all proposed states**. This can be seen by the fact that if the integration would be exact, the total energy $H(x(t),p(t))$ would be still the same (since hamiltonian dynamics preserves energy), and the probability of acceptance would be exactly one.
#
#
# ## Implementation
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
def target_distribution_to_potential(f, gradient_f):
#doing it by hand is numerically more efficient,
#but if you are lazy, here is a helper method
potential=lambda x: -np.log(f(x))
gradient= lambda x: -gradient_f(x)/f(x)
return potential,gradient
class HMC_sampler:
def __init__(self,
potential,
gradient,
num_leapfrog_steps=100,
leapfrog_stepsize=0.1,
mass=1.): #inverse of variance of velocity distribution
self.potential=potential
self.gradient=gradient
self.acceleration=lambda x: -gradient(x)/self.mass
self.num_leapfrog_steps=num_leapfrog_steps
self.leapfrog_stepsize=leapfrog_stepsize
self.mass=mass
def total_energy(self,x,v):
return self.potential(x)+0.5*self.mass*np.dot(v,v)
def do_leapfrog_step(self,x,v,a): # a is for acceleration
new_v= v + 0.5 * self.leapfrog_stepsize * a
new_x= x + new_v * self.leapfrog_stepsize
new_a=self.acceleration(new_x)
new_v= new_v + 0.5 * self.leapfrog_stepsize * new_a
return new_x,new_v,new_a
def should_accept(self, energy, proposed_energy):
#since probability=exp(-energy), this is just metropolis criteria
u = np.random.uniform()
return u < np.exp( energy - proposed_energy )
def sample_velocity(self,shape):
return np.random.normal(0,(1./self.mass)**0.5,size=shape)
def run(self,x_init,num_steps=1000):
x_trace=np.zeros(shape=(num_steps,x_init.shape[0]))
x_trace[0,:]=x_init
logs={'x':[],'v':[],'E':[],'rejected':[]} #just for diagnostics purposes
x=x_init
for i in range(num_steps):
#resample velocity (= random kick)
v=self.sample_velocity(x_init.shape)
#simulate hamiltonian dynamics with leapfrog algorithm
#to get proposal values of x and v
x_prop = x
v_prop = v
a = self.acceleration(x)
for n in range(self.num_leapfrog_steps):
x_prop, v_prop, a = self.do_leapfrog_step(x_prop, v_prop,a)
logs['x'].append(x_prop)
logs['v'].append(v_prop)
#accept or reject
E = self.total_energy(x,v)
E_prop = self.total_energy(x_prop,v_prop)
logs['E'].append(E)
if self.should_accept(E,E_prop):
x= x_prop
else:
logs['rejected'].append((x_prop,v_prop))
x_trace[i,:]= x
return x_trace,logs
# -
# # Simplest example
# Let's try to sample univariate normal distribution $P(x) = \frac{1}{\sqrt{\pi}} e^-x^2$ . Potential energy and it's gradient are:
# +
def potential(x):
return x**2 #So P(x)~exp(-potential(x))
def gradient(x):
return 2*x
#number of leapfrog steps = 50, timestep = 0.1 and m=1
sampler=HMC_sampler(potential,gradient,num_leapfrog_steps=50,leapfrog_stepsize=0.1,mass=1)
# -
# sample 1000 samples, with initial position x=0
x_trace,logs=sampler.run(np.array([0.]),1000)
# +
import seaborn as sns
plt.figure()
plt.subplot(2,1,1)
sns.kdeplot(x_trace[:,0],label='distribution of samples')
plt.plot(np.linspace(-5,5,500),np.exp(-np.linspace(-5,5,500)**2)/(np.pi)**0.5,label='target_distribution')
plt.title('distribution of samples of x')
plt.ylabel('$x_1$', fontsize=14)
plt.legend()
plt.subplot(2,1,2)
plt.plot(x_trace, 'b-')
plt.title('trace of x')
print("number of rejected samples: ",len(logs['rejected']))
# -
# Have a look at the number of rejected samples. Play around with the three parameters that can be tweaked
# - mass (the bigger the mass, the smaller the kicks => slower exploration of the state space)
# - number of leapfrog steps
# - size of leapfrog step
#
# and see what happens.
#
#
# Also have a look at the values of $x$ through which the system passed during leapfrog integration.
# Can you explain why it looks like it does? What about energy log (```logs["E"]```)? What is the distribution of energy?
# +
#number of rejected samples:
print('rejected samples:', len(logs['rejected']))
plt.figure()
plt.plot(logs['x'][:500])
_=plt.title('x during leapfrog integrations')
# -
# # Complicated multidimensional distribution
# Let's try some more complicated distribution, which could be difficult to sample by random walk Metropolis.
# The distribution is:
#
# $$P(x)=e^{-a(r-1)^2 - b \cos(2 \phi) r}$$
#
# Potential for it consists of two equaly deep curved valeys ( in the shape of "bratislavský rožok") at the bottom of a bowl.
# +
a=30.
b=1.5 #play around and try also the value b=6 (or higher). What will happen? How to solve that?
def fancy_potential(x):
r=np.dot(x,x)**0.5
if r<0.1: #potential numerical instability?
return a*(r-1)**2
return a*(r-1)**2 + b*(x[0]**2-x[1]**2)/ r #=a(r-1)**2+b*cos(2*phi)*r
def fancy_gradient(x):
r=np.dot(x,x)**0.5
if r<0.1:
return np.array([0.,0.])
dUdx=2*a*(r-1)*x[0]/r + b*(2*x[0]/r -(x[0]**2-x[1]**2)*x[0]/r**3)
dUdy=2*a*(r-1)*x[1]/r + b*(-2*x[1]/r -(x[0]**2-x[1]**2)*x[1]/r**3)
return np.array([dUdx,dUdy])
xg, yg = np.meshgrid(np.linspace(-2,2,100), np.linspace(-2,2,100))
zg = np.zeros_like(xg)
for i,j in np.ndindex(xg.shape):
zg[i,j] = fancy_potential(np.array([xg[i,j], yg[i,j]]))
plt.figure(figsize=(10,5))
plt.subplot(1,2,1)
plot1=plt.contourf(xg, yg, np.exp(-zg), 20, cmap=plt.cm.viridis)
plt.colorbar(plot1)
plt.title('probability distribution')
plt.subplot(1,2,2)
plot2=plt.contourf(xg, yg, zg, 20,cmap=plt.cm.viridis)
plt.colorbar(plot2)
_=plt.title('potential')
# +
#number of leapfrog steps = 20, timestep = 0.1 and m=10 , number of samples=1000
sampler=HMC_sampler(fancy_potential,fancy_gradient,
num_leapfrog_steps=20,
leapfrog_stepsize=0.1,
mass=10)
x_init=np.array([0.,0.1])
x_trace,logs=sampler.run(x_init,1000)
print('rejected: ', len(logs['rejected']))
# -
g = sns.JointGrid(x=x_trace[:,0], y=x_trace[:,1], size=10)
g.ax_joint.contourf(xg, yg, np.exp(-zg), 20, alpha=0.7, cmap=plt.cm.viridis) #plotting probability distribution
#g.ax_joint.contourf(xg, yg, zg, 20, alpha= 0.7, cmap=plt.cm.viridis) #plotting potential
g = g.plot_joint(plt.scatter, color="r",marker='.')
_ = g.ax_marg_x.hist(x_trace[:,0], color="r", alpha=.4, bins=20)
_ = g.ax_marg_y.hist(x_trace[:,1], color="r", alpha=.4, bins=20, orientation="horizontal")
x,y=zip(*logs['x'][:20*30]) #path of the first 30 leapfrog simulations
_=plt.plot(x,y,linewidth=0.5,color='w')
# HMC sampler clearly follows the regions of high probability quite efficiently without almost any rejections. The white trajectory is trajectory traversed during leapfrogging and it can be clearly seen that it returns to the areas with low potential. Play around with it:
# - try to change parameters a and b of the potential (set b to above 6)
# - change mass, leapfrog_stepsize and num_leapfrog_steps
#
# What exactly does changing the mass do? When does the sampler fail?
#
# # Appendix
#
# ## Why is HMC better than random walk Metropolis in many dimensions?
#
# TODO: read [3](https://arxiv.org/pdf/1206.1901.pdf)
#
# In short: allmost no rejections, so the time to simulate hamiltonian dynamics is worth it.
#
# ## How to finetune it's parameters?
#
# TODO: read [1](https://arxiv.org/pdf/1701.02434.pdf )
#
# ## When not to use HMC?
#
# - unless you understand [5](https://arxiv.org/pdf/1705.08510.pdf), the HMC is usable only with *continuous* state space
# - unless you understand [6](https://papers.nips.cc/paper/5801-reflection-refraction-and-hamiltonian-monte-carlo.pdf), the HMC is not suitable for non-smooth probability densities
# - sharp changes in potential, even if smooth in theory, can lead to numerical instability during leapfrog integration
# - If computation of gradient is difficult / numerically expensive
# - Be carefull if the target distribution have multiple islands with very high probability, since the HMC may become trapped in one of them. Carefull finetuning may take care of that, or you can use HMC with time dependent temperature (tempering).
# - If the probability is exactly zero anywhere. But I believe there exist constrained variations of HMC. Also, reflections at the borders of the area with zero probability as in [6] or change of parametrisation could solve that.
# - if the number of dimensions is low and the shape of target distribution simple.
#
# ## Choice of kinetic energy and momentum sampling distribution
#
# When choosing a suitable hamiltonian, you don't have to constraint yourself to a particle in a potential field ( $H = U(x)+ \frac{1}{2}mv^2$) as we did in this notebook. Only the potential energy $U(x)$ is constrained by it's relationship with the target probability density $U(x)=-log(P(x))$, so it seems you can pick as your kinetic energy any function $K(x,p)$. That is true, but you would have to change the resampling distribution of momentum accordingly. For in depth explanation, see [1](https://arxiv.org/pdf/1701.02434.pdf). In short:
#
# You want to sample from $P(x)$, which is difficult.
# You add new variable $p$.
# Instead of sampling $x$ you sample tuples $(x,p)$ from distribution $P(x,p)$ such, that $P(x)=\int P(x,p) dp$
# In other words, $P(x,p)=P(p|x)P(x)$
# We introduce new function conveniently named $H(x,p)$ such that $P(x,p)=e^{-H(x,p)}$
# From the equations above, we get $H(x,p)=-log(P(p | x))-log(P(x))=K(x,p)+V(x)$
#
# Keep in mind that all introduced functions are just arbitrally functions devised to reproduce our target probability distribution and any analogy with physical energies is purely coincidental ([link](https://vignette.wikia.nocookie.net/zootopia/images/7/74/Coincidence-i-think-not.jpg/revision/latest/scale-to-width-down/640?cb=20160507025359)).
#
# TODO: hmm, it feels like some step is missing here, see [1]
#
# However we already see that kinetic energy must be equal to $K(x,p)=-log(P(p|x))$, so by choosing some $K(x,p)$, we allready define the probability distribution of momentum. Our sampler must respect this constraint, so the momentum must be sampled from the distribution $P(p|x)=e^{-K(x,p)}$.
#
# By choosing $K(x,p)=\frac{1}{2}mv^2$, we get gaussian sampling distribution (which is nice). We could theoretically prefer different sampling distribution, in which case we would have to pick kinetic energy term accordingly.
#
# However, kinetic energy dependent on both $x$ and $p$ should probably be avoided in practice, since the leapfrog algorithm is not time reversible in such case [citation or proof needed].
#
# If you want experimental proof that the kinetic energy function and velocity sampling distribution are interdependent, check out the HMC_sampler implementation and try to rescale one of them (but not the other) and see what happens.
#
# # Further reading
#
# 1. <NAME>: [Conceptual introduction to Hamiltonian Monte Carlo](https://arxiv.org/pdf/1701.02434.pdf), 2017.
# 2. <NAME>: [Probabilistic inference using Markov Chain Monte Carlo methods](http://www.cs.toronto.edu/~radford/ftp/review.pdf), tech report, 1993.
# 3. <NAME>: [MCMC using Hamiltonian dynamics](https://arxiv.org/pdf/1206.1901.pdf), 2012.
# 4. <NAME>: [The Boltzmann distribution law and
# statistical thermodynamics](http://assets.cambridge.org/97805218/11194/excerpt/9780521811194_excerpt.pdf).
# 5. <NAME> et al: [Discontinuous Hamiltonian Monte Carlo for models with
# discrete parameters and discontinuous likelihoods](https://arxiv.org/pdf/1705.08510.pdf), 2018.
# 6. <NAME> and <NAME>: [Reflection, Refraction, and Hamiltonian Monte Carlo](https://papers.nips.cc/paper/5801-reflection-refraction-and-hamiltonian-monte-carlo.pdf), NIPS 2015.
# 7. <NAME> and <NAME>: [The No-U-Turn Sampler: Adaptively Setting Path Lengths in Hamiltonian Monte Carlo](http://jmlr.org/papers/volume15/hoffman14a/hoffman14a.pdf), JMLR 15, 2014.
# 8. <NAME>, et al: [Hybrid Monte Carlo](https://www.sciencedirect.com/science/article/pii/037026938791197X), Physics Letters B 195(2), 1987. This article is behind a paywall.
#
# Some gentler reading and nice pictures:
# https://am207.github.io/2017/wiki/hmcexplore.html
# http://arogozhnikov.github.io/2016/12/19/markov_chain_monte_carlo.html
| notebooks/3j.2 Hamiltonian Monte-Carlo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JSRist0028/animalmigration/blob/main/code/Migration_Data_Prediction_V2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="ew8IxpzYFguF" outputId="952aed24-7271-4fe8-dfc3-155aa87e321c"
# !pip install geopandas
# !pip install shapely
# !pip install meteostat
# + id="-dA5jdXkWLxO"
import pandas as pd
import sklearn.preprocessing
import sklearn.manifold
import matplotlib.pyplot as plt
import numpy as np
import math
import datetime
from datetime import datetime
import meteostat
from meteostat import Stations, Daily
from dateutil.relativedelta import relativedelta
import datetime as dt
from datetime import datetime
# + [markdown] id="IEoacm-CyUH_"
# **Change the csv file and whether or not to include temperature here:**
# + id="paYMy5ZddIYn" colab={"base_uri": "https://localhost:8080/", "height": 608} outputId="749eea6d-6222-4883-e678-200f34338681"
# import data with combined temperature and tracking information
data_raw = pd.read_csv('https://github.com/JSRist0028/animalmigration/blob/0175410b123d51c6463a7d5e1130bdf68373d68c/data/barnacle_geese_data_matrix?raw=true')
# geese data: ('https://github.com/JSRist0028/animalmigration/blob/main/data/barnacle_geese_data_matrix?raw=true')
# whale data: ('https://raw.githubusercontent.com/JSRist0028/animalmigration/main/data/Azores%20Great%20Whales%20Satellite%20Telemetry%20Program%20.csv')
includeTemp = True;
data_raw
# + id="H1MhL2YDI2HT"
# creates dataframe with only unique dates
def date_to_nth_day(date):#, format='%Y%m%d'):
date = pd.to_datetime(date)#, format=format)
new_year_day = pd.Timestamp(year=date.year, month=1, day=1)
return (date - new_year_day).days + 1
# source: https://codereview.stackexchange.com/questions/154140/interpret-yyyymmdd-as-the-nth-day-of-the-year
# create a new DataFrame with only location data from unique days
# this is now redundant
data = pd.DataFrame(columns=data_raw.columns) # create header
for i, row in data_raw.iterrows():
if i>0: # skip first row
if (date_to_nth_day(row['timestamp']) != date_to_nth_day(data_raw['timestamp'][i-1])):
data = data.append(row, ignore_index=True)
#print(i)
# + id="Ruzq8gYuIK6X"
# Splits data into features (today) and targets (tomorrow)
def get_todaytomorrow(data, includeTemp):
num_rows = len(data)
today_timestamp = np.empty((0,1))
tomorrow_timestamp = np.empty((0,1))
if includeTemp:
today = np.empty((0,7), dtype='float')#, 'int') #[]
else:
today = np.empty((0,4), 'int') #[]
tomorrow = np.empty((0,3), dtype='float') #, 'int') #[]
for i, row in data.iterrows():
if (i<(num_rows-3)):
if (pd.notnull(row['tavg']) & pd.notnull(data['tavg'][i+1])): # only add pair if both contain basic weather data (avg temp)
if (data['birdID'][i]==data['birdID'][i+1]): # only add if birdIDs match
today_day = date_to_nth_day(row['timestamp'])
tomorrow_day = date_to_nth_day(data['timestamp'][i+1])
# if (tomorrow_day-today_day == 1): # makes sure that the days are consecutive -> already done ^
birdID_today = row['birdID'].replace('a','1')
birdID_today = birdID_today.replace('b', '2')
birdID_tomorrow = data['birdID'][i+1].replace('a','1')
birdID_tomorrow = birdID_tomorrow.replace('b','2')
today_timestamp = np.append(today_timestamp, row['timestamp'])
tomorrow_timestamp = np.append(tomorrow_timestamp, data['timestamp'][i+1])
if includeTemp:
today_entry = np.asarray([int(birdID_today), today_day, row['latitude'], row['longitude'], row['tavg'], row['tmin'], row['tmax'] ])
tomorrow_entry = np.asarray([int(birdID_tomorrow), data['latitude'][i+1], data['longitude'][i+1]]) #no change
else:
today_entry = np.asarray([int(birdID_today), today_day, row['latitude'], row['longitude']])
tomorrow_entry = np.asarray([int(birdID_tomorrow), data['latitude'][i+1], data['longitude'][i+1]]) # no change
today_entry[np.isnan(today_entry)]=0
tomorrow_entry[np.isnan(tomorrow_entry)]=0
today = np.vstack((today,today_entry))
tomorrow = np.vstack((tomorrow, tomorrow_entry))
#print(i)
return today, tomorrow, today_timestamp, tomorrow_timestamp
today, tomorrow, today_timestamp, tomorrow_timestamp = get_todaytomorrow(data, includeTemp=includeTemp)
# + id="2a8pE1w8WEFo"
# Split the data into training and testing sets
features = today[:,1:] #data.values[:,:-1]
labels = tomorrow[:,1:] #[:,[1,2]] #data.values[:, 60]
#features = features.reshape(features.shape[0], features.shape[1], 1) if using CNN, may need to use this
import math
# Split the data into training and testing sets cronologically
todaydf = pd.DataFrame(today)
tomorrowdf = pd.DataFrame(tomorrow)
def train_test_split(datafile, tomorrow, train_ratio):
# Define dataframes to return
xheaders = ['AnimalID', 'TS', 'Lat - 1', 'Long - 1', 'TAvg', 'TMin', 'TMax']
yheaders = ['AnimalID', 'Lat', 'Long']
trainx = pd.DataFrame(columns = xheaders)
trainy = pd.DataFrame(columns = yheaders)
testx = pd.DataFrame(columns = xheaders)
testy = pd.DataFrame(columns = yheaders)
# Split each unique animal tracking info into training and testing sets
for animal in datafile[0].unique():
traincount = math.ceil(float(datafile[datafile[0] == animal].shape[0]) * train_ratio)
obscount = datafile[datafile[0] == animal].shape[0]
obslist = datafile.index[datafile[0] == animal].tolist()
for obs in range(0, traincount, 1):
dfx = [animal, datafile[1][obslist[obs]], datafile[2][obslist[obs]], datafile[3][obslist[obs]],
datafile[4][obslist[obs]], datafile[5][obslist[obs]], datafile[6][obslist[obs]]]
dfy = [animal, tomorrow[1][obslist[obs]], tomorrow[2][obslist[obs]]]
trainx.loc[len(trainx.index)] = dfx
trainy.loc[len(trainy.index)] = dfy
for obs in range(traincount, obscount, 1):
dfx = [animal, datafile[1][obslist[obs]], datafile[2][obslist[obs]], datafile[3][obslist[obs]],
datafile[4][obslist[obs]], datafile[5][obslist[obs]], datafile[6][obslist[obs]]]
dfy = [animal, tomorrow[1][obslist[obs]], tomorrow[2][obslist[obs]]]
testx.loc[len(testx.index)] = dfx
testy.loc[len(testy.index)] = dfy
return(trainx, trainy, testx, testy)
trainin_x, trainin_y, testin_x, testin_y = train_test_split(todaydf, tomorrowdf, 0.8)
# + id="ZBYtzSx84g8L"
# converts data into correct type for keras model
training_x = np.transpose(np.asarray([trainin_x['TS'], trainin_x['Lat - 1'], trainin_x['Long - 1']]).astype('float32'))
training_y = np.transpose(np.asarray([trainin_y['Lat'], trainin_y['Long']]).astype('float32'))
testing_x = np.transpose(np.asarray([testin_x['TS'], testin_x['Lat - 1'], testin_x['Long - 1']]).astype('float32'))
testing_y = np.transpose(np.asarray([testin_y['Lat'], testin_y['Long']]).astype('float32'))
# from https://stackoverflow.com/questions/48851558/tensorflow-estimator-valueerror-logits-and-labels-must-have-the-same-shape
# + [markdown] id="5cPP4XbxbrSb"
# # Regression Model
# + id="nLm0DyC9mcwT"
# Regression packages
import keras
from keras.models import Sequential
from keras.layers import Dense, Conv1D, Flatten
from keras.wrappers.scikit_learn import KerasRegressor
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import KFold
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from tensorflow.keras.layers.experimental import preprocessing
# + id="DsbPqC3581Em"
# Normalization
normalizer = preprocessing.Normalization()
normalizer.adapt(np.array(training_x))
# Build model, add layers
input_size = training_x.shape[1]
my_model = Sequential([normalizer, Dense(units=1)])
my_model.add(Dense(32, input_dim=input_size, kernel_initializer='normal', activation='relu'))
#my_model.add(Conv1D(32, 3, activation="relu"))
my_model.add(Dense(16, kernel_initializer='normal', activation='relu'))
#my_model.add(Dense(16, kernel_initializer='normal', activation='relu'))
my_model.add(Dense(8, kernel_initializer='normal', activation='relu'))
my_model.add(Dense(2, kernel_initializer='normal'))
# + id="-Hijc_DiCOlP" colab={"base_uri": "https://localhost:8080/", "height": 198} outputId="5395a0b2-6531-42cb-cb64-436f03913a74"
# Train model
from tensorflow import optimizers
my_model.compile(optimizer=optimizers.Adam(learning_rate=0.1),loss='mean_absolute_error')
history = my_model.fit(training_x, training_y,epochs=50,verbose=0,validation_split = 0.2)
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
hist.tail()
# + id="VmlA3hLsCL6L" colab={"base_uri": "https://localhost:8080/", "height": 279} outputId="ff4addb4-106e-488e-df2e-734e5b3214d9"
# Plot loss by Epoch to evaluate model improvement
def plot_loss(history):
plt.plot(history.history['loss'], label='loss')
plt.plot(history.history['val_loss'], label='val_loss')
#plt.ylim([0, 10])
plt.xlabel('Epoch')
plt.ylabel('Error')
plt.legend()
plt.grid(True)
plot_loss(history)
# + id="wTPpms_MDcIj" colab={"base_uri": "https://localhost:8080/"} outputId="d5737c15-08c8-440c-e8a4-02ecae6dd12c"
my_model.evaluate(testing_x,testing_y, verbose=0)
# + id="oNi758gMEg7Q" colab={"base_uri": "https://localhost:8080/", "height": 283} outputId="0d2f8380-6adc-4de7-8480-31774ac3a847"
# Make predictions
train_predictions = my_model.predict(training_x)#.flatten()
test_predictions = my_model.predict(testing_x)#.flatten()
test_predictions
fig,ax = plt.subplots()
ax.plot(testing_y[:,0], testing_y[:,1], 'o', label='Actual')
ax.plot(test_predictions[:,0], test_predictions[:,1], 'o', label='Predicted')
ax.legend()
# + id="_IwLMsR4VKZY" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="d444bea0-d826-42fc-e02d-27c49dbd095d"
testing_error = np.sqrt((test_predictions[:,0] - testing_y[:,0])**2 + (test_predictions[:,1] - testing_y[:,1])**2)
training_error = np.sqrt((train_predictions[:,0] - training_y[:,0])**2 + (train_predictions[:,1] - training_y[:,1])**2)
# need to change this to account for lat/long to get real distance between points
print(f'Mean testing error: {np.mean(testing_error)}')
print(f'Mean training error: {np.mean(training_error)}')
#fig = plt.figure()
#ax = plt.axes(projection='3d')
#ax.scatter3D(training_y[:,0],training_y[:,1], training_error, label='Training Error', marker='x', depthshade=False)
#ax.scatter3D(testing_y[:,0],testing_y[:,1], testing_error,label='Testing Error', marker='x',depthshade=False)
#ax.legend()
from matplotlib.ticker import PercentFormatter
bins_list = [0,1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20]
fig = plt.figure()
plt.hist([training_error, testing_error], bins=bins_list, label=['Training Error', 'Testing Error'], weights=[np.ones(len(training_error))/len(training_error), np.ones(len(testing_error))/len(testing_error)])
#plt.hist(training_error, weights=np.ones(len(training_error))/len(training_error), label='Training Error', bins=bins_list )
#plt.hist(testing_error, weights=np.ones(len(testing_error))/len(testing_error), label='Testing Error',bins=bins_list)
plt.xlabel('Error')
plt.ylabel('Percentage of Prediction')
plt.gca().yaxis.set_major_formatter(PercentFormatter(1))
plt.xlim(0,15)
plt.legend()
# + id="uoRCNQdCFmF0" colab={"base_uri": "https://localhost:8080/", "height": 454} outputId="b6aaacfb-470a-4aa8-97de-d156655435f0"
# Plot prediction on map
from shapely.geometry import Point, LineString, MultiPoint
import geopandas as gpd
from geopandas import GeoDataFrame
actual_plot=gpd.GeoDataFrame([[MultiPoint(np.flip(testing_y, axis=1))]],columns=['geometry'])
predicted_plot=gpd.GeoDataFrame([[MultiPoint(np.flip(test_predictions, axis=1))]],columns=['geometry'])
training_plot = gpd.GeoDataFrame([[MultiPoint(np.flip(training_y, axis=1))]], columns=['geometry'])
minx, miny, maxx, maxy = actual_plot.geometry.total_bounds
fig, ax = plt.subplots(figsize=(20,12))
world = gpd.read_file(gpd.datasets.get_path('naturalearth_lowres'))
world.plot(ax=ax)
training_plot.plot(ax=ax,marker='o',label='Training', color='cyan', markersize=15)
actual_plot.plot(ax=ax, marker='o', label='Actual', color='orange')
predicted_plot.plot(ax=ax,marker='o',label='Predicted', color='red')
ax.set_xlim(minx - 5, maxx + 5) # added/substracted value is to give some margin around total bounds
ax.set_ylim(miny - 5, maxy + 5)
ax.legend()
#https://gis.stackexchange.com/questions/332624/geopandas-plot-two-layers-but-only-to-the-extent-of-the-smaller-one
# + [markdown] id="Ir1uL6vtSFE4"
# # Predicting consecutive days of migration
# + id="hQ11rG2Syz3m"
def getWeather(stations, lat, long, mig_datetime):
#stations = Stations()
stations = stations.nearby(lat = 33.52068, lon = -86.81176,radius=40000) #note: radius is in meters (40k meters ~ 25 miles)
station = stations.fetch(1)
weather_data = Daily(station, start=mig_datetime,end=(mig_datetime+relativedelta(days=0)))
weather_data = weather_data.fetch()
weather_data_temp = weather_data[['tavg', 'tmin', 'tmax', 'prcp', 'snow', 'wdir', 'wspd', 'wpgt', 'pres', 'tsun']].values.tolist()
if (not weather_data_temp): # sometimes it just creates an empty list. This checks/fixes it
weather_data_temp = np.empty((1,10), dtype='float')
weather_data_temp[:] = np.NaN
for i in range(2, stations.count()-1):
if (np.isnan(weather_data_temp).any):
station = stations.fetch(i)
weather_data = Daily(station, start=mig_datetime,end=(mig_datetime + relativedelta(days=1)))
weather_data = weather_data.fetch()
#a = weather_data['wdir'].values[0].tolist()
#print(a)
try:
if np.isnan(weather_data_temp[0][0]): #tavg
[weather_data_temp[0][0]] = weather_data['tavg'].values.tolist()
if np.isnan(weather_data_temp[0][1]): #tmin tmax
[weather_data_temp[0][1]] = weather_data['tmin'].values.tolist()
[weather_data_temp[0][2]] = weather_data['tmax'].values.tolist()
if np.isnan(weather_data_temp[0][3]): #prcp
[weather_data_temp[0][3]] = weather_data['prcp'].values.tolist()
if np.isnan(weather_data_temp[0][4]): #snow
[weather_data_temp[0][4]] = weather_data['snow'].values.tolist()
if np.isnan(weather_data_temp[0][5]): #wdir
[weather_data_temp[0][5]] = weather_data['wdir'].values.tolist()
if np.isnan(weather_data_temp[0][6]): #wspd
[weather_data_temp[0][6]] = weather_data['wspd'].values.tolist()
if np.isnan(weather_data_temp[0][7]): #wpgt
[weather_data_temp[0][7]] = weather_data['wpgt'].values.tolist()
if np.isnan(weather_data_temp[0][8]): #pres
[weather_data_temp[0][8]] = weather_data['pres'].values.tolist()
if np.isnan(weather_data_temp[0][9]): #tsun
[weather_data_temp[0][9]] = weather_data['tsun'].values.tolist()
except:
print("exception: ", weather_data)
station_data = station.values[0]
return weather_data_temp
# + id="ujqyBbt8ydOR"
#getWeather(stations, longtest_x[rows-1][0], longtest_x[rows-1][1], dt.datetime.strptime(tomorrow_timestamp[:][rows-1], "%Y-%m-%d %H:%M:%S"))
#getWeather(stations, longtest_x[rows][0], longtest_x[rows][1], dt.datetime.strptime(tomorrow_timestamp[:][rows], "%Y-%m-%d %H:%M:%S"))
# + id="yFmePQQrST6Z"
from dateutil.relativedelta import relativedelta
longtest_x = testing_x
longtest_y = my_model.predict([longtest_x[:][0].tolist()])
for rows in range(1, len(testin_x), 1):
if includeTemp:
stations = Stations()
weather_data = getWeather(stations, longtest_x[rows][0], longtest_x[rows][1], dt.datetime.strptime(tomorrow_timestamp[:][rows], "%Y-%m-%d %H:%M:%S"))
longtest_x_row = [longtest_x[:][rows].tolist()]
else:
longtest_x_row = [longtest_x[:][rows].tolist()]
predict = my_model.predict(longtest_x_row)
longtest_y = np.vstack((longtest_y, predict))
if len(testin_x) > rows + 1:
if testin_x['AnimalID'][rows] == testin_x['AnimalID'][rows + 1]:
longtest_x[rows + 1, 1] = predict[0][0]
longtest_x[rows + 1, 2] = predict[0][1]
# + id="tfX4ol1YbpHK" colab={"base_uri": "https://localhost:8080/"} outputId="1c8b830b-b321-4199-9622-1733b846ac24"
my_model.evaluate(testing_x, testing_y, verbose=0)
# + id="IfHgLC9KrikH" colab={"base_uri": "https://localhost:8080/", "height": 500} outputId="adc6e827-df51-4b30-c8cd-9bf302c4b124"
# Plot updated prediction on map
actual_plot=gpd.GeoDataFrame([[MultiPoint(np.flip(testing_y, axis=1))]],columns=['geometry'])
predicted_plot=gpd.GeoDataFrame([[MultiPoint(np.flip(longtest_y, axis=1))]],columns=['geometry'])
training_plot = gpd.GeoDataFrame([[MultiPoint(np.flip(training_y, axis=1))]], columns=['geometry'])
minx, miny, maxx, maxy = actual_plot.geometry.total_bounds
fig, ax = plt.subplots(figsize=(20,12))
world = gpd.read_file(gpd.datasets.get_path('naturalearth_lowres'))
world.plot(ax=ax)
training_plot.plot(ax=ax,marker='o',label='Training', color='cyan', markersize=15)
actual_plot.plot(ax=ax, marker='o', label='Actual', color='orange')
predicted_plot.plot(ax=ax,marker='o',label='Predicted', color='red')
ax.set_xlim(minx - 5, maxx + 5) # added/substracted value is to give some margin around total bounds
ax.set_ylim(miny - 5, maxy + 5)
ax.legend()
#https://gis.stackexchange.com/questions/332624/geopandas-plot-two-layers-but-only-to-the-extent-of-the-smaller-one
# + id="FImkX-xrxkkG"
np.savetxt("longtest_y.csv", longtest_y, delimiter=",")
np.savetxt("testing_y.csv", testing_y, delimiter=",")
| code/Migration_Data_Prediction_V2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import argparse
import pickle
import os
import scipy.stats
import numpy as np
import pandas as pd
from tabulate import tabulate
from scikit_posthocs import posthoc_conover
import sys
import matplotlib.pyplot as plt
from matplotlib import cm
from pathlib import Path
from joblib import load
import multiprocessing
import logging
logging.basicConfig(level=logging.INFO)
# +
def dev(approx, true):
return abs(approx - true)
def compute_stats(dataset, alpha, log=False):
# load results
logging.info(f'Loading results from {dataset}...')
with open(dataset, 'rb') as fh:
results = pickle.load(fh)
# analyze results
for i, (name, r) in enumerate(results.items()):
# compute errors
errors = pd.DataFrame({
h: dev(r[h], r['true_ce'])
for h in r.head()
})
# perform ANOVA
data = [errors[h] for h in errors.head()]
_, anova_pvalue = scipy.stats.friedmanchisquare(*data)
ret = (name,)
if anova_pvalue < alpha:
# perform post-hoc analysis
if log:
print("\n")
print(f'{name}: ANOVA p-value {anova_pvalue} is significant @ {alpha}\n')
table = []
for h in errors.head():
for v in errors[h]:
table.append(pd.DataFrame({'Metric': [h], 'Value': [v]}))
table = pd.concat(table)
p_values = posthoc_conover(table, p_adjust='holm', group_col='Metric', val_col='Value')
medians = errors.median(axis=0)
if log:
print(tabulate([
[h, np.median(errors[h]), np.std(errors[h])] + ['diff' if abs(p) < alpha else 'same' for p in p_values[h]]
for h in p_values.columns
], headers=['Metric', 'Bias', 'Std'] + list(p_values.columns)))
ret = name, p_values, medians
else:
if log:
print(f'{name}: ANOVA p-value {anova_pvalue} is not significant @ {alpha}')
if log:
print("\n")
yield ret
# -
def process_single(dataset, alpha):
with open(os.path.splitext(dataset)[0]+".log", "w") as log:
print(f"Writing output to {log.name}")
gen = compute_stats(dataset, alpha, log)
for stat in gen:
if len(stat) > 1:
print(stat[0])
else:
print(f"No result for {stat[0]}")
# +
def process_run(input_tuple):
run, dataset, alpha = input_tuple
f = os.path.join(dataset, run["filename"])
data = []
for ret in compute_stats(f, alpha):
row = run./home/maximl/projects/reliability/notebookscopy()
# if anova was significant, check median bias of ece_v{2,3} compared to ece
if len(ret) > 1:
name, p_values, medians = ret
row["clf"] = name
for m, p in p_values["ece"].drop(['true_ce', 'ece']).iteritems():
# if ece_v{2,3} - ece is significant, check which one is more biased
if p < alpha:
if medians[m] > medians["ece"]:
# ece is less biased
row[m] = -1
else:
# ece is more biased
row[m] = 1
else:
# similar bias
row[m] = 0
data.append(row)
return pd.DataFrame(data)
def process_multi(dataset, alpha):
assert Path(dataset).is_dir(), "Dataset needs to be directory containing grid results."
# load all meta data from runs into one dataframe
runs = pd.concat([
load(p) for p in
Path(dataset).glob("*runs*")
]).reset_index(drop=True)
# compute statistics for every run
with multiprocessing.Pool(processes=multiprocessing.cpu_count()) as pool:
data = pool.map(process_run, map(lambda row: (row[1], dataset, alpha), runs.iterrows()))
data = pd.concat(data)
with open(os.path.join(dataset, "analysis.dat"), "wb") as pkl:
pickle.dump(data, pkl)
return data
# -
data = process_multi("/home/maximl/Data/Experiment_data/results/riverrel/artificial_data/899c1651a0155fcfd0bfb86b16607dde3ac19e49/", 0.05)
| experiments/2_artificial_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# # Week 01: Introduction & the Cool Programming Language
#
# ## How to Run the Program
#
# - Compiler (Offline)
# - $Program \rightarrow Compiler \rightarrow Execute$
# - $Data \rightarrow Execute \rightarrow Output$
# - Interpreter (Online)
# - $Program + Data \rightarrow Interpreter \rightarrow Output$
#
# ## Compiler Concept
#
# - (Syntactic) **Lexical Analysis**
# - Concept: Divide program text into words or tokens.
# - Input: text
# - Output: words or tokens
# - Sample Input: `if x == y then z = 1; else z = 2;`
# - Sample Output: `#IF #ID(x) #EQAUL #ID(y) #THEN ...`
# - (Syntactic) **Parsing**
# - Concept: Diagramming Sentences.
# - Input: Tokens
# - Output: Abstruct Semantic Tree
# - Sample Input: #INT(5) #PLUS #INT(3) #MULTIPLY #INT(5)
# - Sample Output: `(#PLUS (#INT(5)) (#MULTIPLY (#INT(3)) (#INT(5))))`
# - (Types scope) **Semantic Analysis**
# - Concept: Catch inconsistencies.
# - Sample Input: `{ int Jack=3; { int Jack=4; cout << Jack; } }`
# - Question: What is the value?
# - **Optimization**
# - Concept: Run faster/Use less memory/Use low power/network.
# - (Translation) **Code Generation**
# - Concept: Produce assembly code.
#
# ## Related Questions
#
# - Why are there so many programming languages?
# - Application domains have distinct/conflicting needs.
# - Why are there new programming languages?
# - Programming training is the dominant cost for a programming language.
# - Wild-used languages are **slow to change**.
# - Easy to start a new language: when **productivity** > **training cost**
# - Languages adopted to fill a void. (**Void** means new techniques.)
# - What is a good programming languages?
# - There is **no** universally accepted metric for language design.
#
#
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
| OCW/[Stanford]CS143_Compilers/Notes/Week01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Deep Learning Bootcamp November 2017, GPU Computing for Data Scientists
#
# <img src="images/bcamp.png" align="center">
#
# ## Using CUDA, Jupyter, PyCUDA and PyTorch
#
# ### 03 PyCUDA Sigmoid()
#
# Web: https://www.meetup.com/Tel-Aviv-Deep-Learning-Bootcamp/events/241762893/
#
# Notebooks: <a href="https://github.com/QuantScientist/Data-Science-PyCUDA-GPU"> On GitHub</a>
#
# *<NAME>*
#
# <img src="images/gtx.png" width="35%" align="center">
# + [markdown] slideshow={"slide_type": "slide"}
# # PyCUDA Imports
# + slideshow={"slide_type": "skip"}
# # !pip install pycuda
# %reset -f
import pycuda
from pycuda import compiler
import pycuda.driver as cuda
import numpy
import numpy as np
from pycuda.compiler import SourceModule
cuda.init()
print("%d device(s) found." % cuda.Device.count())
for ordinal in range(cuda.Device.count()):
dev = cuda.Device(ordinal)
print "Device #%d: %s" % (ordinal, dev.name())
print cuda
# -
# ! watch --color -n1.0 gpustat
# + [markdown] slideshow={"slide_type": "slide"}
# # Simple addition on the GPU: CUDA Kernel definition
# +
import pycuda.autoinit
# a = np.random.uniform(low=1, high=20, size=(10,))
a = numpy.arange(-100000, 100000, 1)
a = a.astype(numpy.float32)
ARR_SIZE = numpy.int32(a.shape[-1])
print ARR_SIZE
a_gpu = cuda.mem_alloc(a.nbytes)
xout_gpu = cuda.mem_alloc(a.nbytes)
cuda.memcpy_htod(a_gpu, a)
xout_gpu=cuda.mem_alloc_like(a)
# size_gpu=cuda.mem_alloc_like(size)
mod = SourceModule("""
__global__ void sigmoid(float* a, float* b, int size)
{
int index = blockDim.x * blockIdx.x + threadIdx.x;
if (index < size)
b[index] = 1.0f / (1.0f + exp(-1.0f * a[index]));
}
""")
func = mod.get_function("sigmoid")
def sigmoidGPU():
func(a_gpu, xout_gpu,ARR_SIZE, block=(ARR_SIZE/1024,1,1))
a_sigmoid = numpy.empty_like(a)
cuda.memcpy_dtoh(a_sigmoid, xout_gpu)
return a_sigmoid
# print sigmoidGPU()
from scipy.special import expit
y = expit(a)
# print ("__________________________________")
# print y
# + [markdown] slideshow={"slide_type": "slide"}
# # Plot the Sigmoid function
# +
import matplotlib.pyplot as plt
plt.plot(a,y)
plt.text(4,0.8,r'$\sigma(x)=\frac{1}{1+e^{-x}}$',fontsize=15)
plt.legend(loc='lower right')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# # Timing Numpy vs. PyCUDA ...
# +
import timeit
n_iter = ARR_SIZE
rounds = 1000 # for timeit
print 'numpy', timeit.timeit(lambda:
expit(a),
number=rounds)
print 'pycuda', timeit.timeit(lambda:
sigmoidGPU(),
number=rounds)
# -
| day 02 PyTORCH and PyCUDA/PyCUDA/03 PyCUDA sigmoid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import requests
os.chdir("C:/Users/abewo/Documents/GitHub/gpt-2")
# +
# #%run -i download_model.py 345M
# -
# !pip3 install -r requirements.txt
os.chdir('src')
import json
import os
import numpy as np
import tensorflow as tf
import model, sample, encoder
def interact_model(
model_name,
seed,
nsamples,
batch_size,
length,
temperature,
top_k,
models_dir
):
models_dir = os.path.expanduser(os.path.expandvars(models_dir))
if batch_size is None:
batch_size = 1
assert nsamples % batch_size == 0
enc = encoder.get_encoder(model_name, models_dir)
hparams = model.default_hparams()
with open(os.path.join(models_dir, model_name, 'hparams.json')) as f:
hparams.override_from_dict(json.load(f))
if length is None:
length = hparams.n_ctx // 2
elif length > hparams.n_ctx:
raise ValueError("Can't get samples longer than window size: %s" % hparams.n_ctx)
with tf.Session(graph=tf.Graph()) as sess:
context = tf.placeholder(tf.int32, [batch_size, None])
np.random.seed(seed)
tf.set_random_seed(seed)
output = sample.sample_sequence(
hparams=hparams, length=length,
context=context,
batch_size=batch_size,
temperature=temperature, top_k=top_k
)
saver = tf.train.Saver()
ckpt = tf.train.latest_checkpoint(os.path.join(models_dir, model_name))
saver.restore(sess, ckpt)
while True:
raw_text = input("Model prompt >>> ")
while not raw_text:
print('Prompt should not be empty!')
raw_text = input("Model prompt >>> ")
context_tokens = enc.encode(raw_text)
generated = 0
for _ in range(nsamples // batch_size):
out = sess.run(output, feed_dict={
context: [context_tokens for _ in range(batch_size)]
})[:, len(context_tokens):]
for i in range(batch_size):
generated += 1
text = enc.decode(out[i])
print("=" * 40 + " SAMPLE " + str(generated) + " " + "=" * 40)
print(text)
print("=" * 80)
# __model_name:__ This indicates which model we are using. In our case, we are using the GPT-2 model with 345 million parameters or weights
#
# __seed:__ Integer seed for random number generators, fix seed to reproduce results
#
# __nsamples:__ This represents the number of sample texts generated in our output
#
# __batch_size:__ This only affects speed/memory. This must also divide nsamples
# Note: To generate more than one sample, you need to change the values of both nsamples and batch_size and also have to keep them equal.
#
# __length:__ It represents the number of tokens in the generated text. If the length is None, then the number of tokens is decided by model hyperparameters
#
# __temperature:__ This controls randomness in Boltzmann distribution. Lower temperature results in less random completions. As the temperature approaches zero, the model will become deterministic and repetitive. Higher temperature results in more random completions
#
# __top_k:__ This parameter controls diversity. If the value of top_k is set to 1, this means that only 1 word is considered for each step (token). If top_k is set to 40, that means 40 words are considered at each step. 0 (default) is a special setting meaning no restrictions. top_k = 40 generally is a good value
#
# __models_dir:__ It represents the path to parent folder containing model subfolders (contains the <model_name> folder)
os.chdir("C:/Users/abewo/Documents/GitHub/gpt-2")
interact_model(
model_name='345M',
seed=None,
nsamples=1,
batch_size=1,
length=300,
temperature=1,
top_k=0,
models_dir='models'
)
| .ipynb_checkpoints/Exploratory Notebook-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **PROGETTO**
# # FEATURES NUMERICHE, CATEGORIALI, DATA
# In questo notebook tratto ed introduco nel modello le features numeriche, categoriali e di tipo data. Le varie features verranno aggiunte in modo incrementale. Nel successivo notebook verranno introdotte ulteriore features: di tipo insiemistico e di tipo testuale.
#
# Spesso è necessario valutare se vale la pena aggiungere certe features o è necessario capire che alternativa è migliore nella lavorazione di certe features : uso lo score sul validation set per capire cosa è meglio fare e per orientarmi tra le varie possibili scelte.
#
# Verranno presi in considerazione 4 tipi di algoritmi di learning : kNN, regressione lineare, albero di decisione, random forest. Dunque per ogni possibile alternativa ho 4 modelli diversi e dunque ho 4 score sul validation set diversi. Scelgo l'alternativa e il modello che hanno score sul validation minore : questo è il modello migliore fino a quel momento ottenuto. Dunque la mia guida è sempre lo score sul validation e scelgo ciò che minimizza ciò.
# ### FUNZIONI VALUTAZIONE E SELEZIONE MODELLI
#
# Importo le **funzioni per la valutazione e selezione dei modelli**.
#
# Le funzioni compute_train_val_test e model_selection_TrainValTest effettuano la valutazione tramite gli score su training/validation/test, rispettivamente su un modello solo o su una lista di modelli. Le funzioni compute_bias_variance_erroe e model_selection_BiasVarianceError effettuano la valutazione tramite il calcolo di bias/variance/error, rispettivamente su un modello solo o su una lista di modelli.
#
# Nel progetto uso lo **score sul validation** come misura principale per selezionare un modello. Uso il calcolo di bias/variance/error come misura ulteriore di bontà, in particolare per capire come poter migliorare il modello stesso.
from valutazione_modelli import compute_train_val_test, model_selection_TrainValTest, compute_bias_variance_error, \
model_selection_BiasVarianceError
# # PRIMA LETTURA E FEATURES NUMERICHE
# Per prima cosa effettuiamo la prima lettura del dataset e aggiungiamo nel modello le features numeriche: "budget", "popularity", "runtime".
#
# La funzione **cleaning_data_numeric** effettua la prima lettura e lavora ed estrae le features numeriche. Ritorna X (matrice delle features numeriche selezionate), y (vettore con "revenue") e df (dataframe pandas completo di tutte le features). Il dataframe df ha tutte le features, ma con già lavorate solo le features numeriche.
#
# La matrice X ha 4 features: "budget", "budget_dummy", "popularity", "runtime".
#
# Il vettore y è scalato con MinMaxScaler.
#
# In tutto il progetto usa sia dataframe pandas (come df) che array numpy (come X e y). I datframe li uso per gestire, lavorare e visualizzare meglio il dataset. Gli array numpy li uso per valutare i modelli.
# +
from lavorazione_dataset import cleaning_data_numeric
X, y, df = cleaning_data_numeric()
# -
df.info()
df.describe()
print("X shape: ",X.shape)
print("y shape: ",y.shape)
# X per ora ha 4 features : "budget", "budget_dummy", "popularity", "runtime". (Sono in questo preciso ordine).
print(X[0:6,])
# # PREDITTORE BANALE : revenue come funzione lineare del solo budget.
#
# Il primo modello che prendiamo in considerazione è un modello che considera solo "budget" come feature per spiegare "revenue". Predittore banale. In particolare consideriamo la regressione lineare tra "revenue" e "budget".
# +
from sklearn.linear_model import LinearRegression
model = LinearRegression(fit_intercept=True) # Modello regressione lineare
# Calcolo gli score su training/validation/test della regressione lineare con solo "budget".
train, val, test = compute_train_val_test(model, X[:,0:1], y)
print("MSE : ",val)
# -
# Questo è il nostro primo score ottenuto.
# # SOLO FEATURES NUMERICHE
# Consideriamo ora tutte e 4 le features numeriche messe in X. Valutiamo i 4 algoritmi di learning su tale dataset.
# ## **1) KNN (con preprocessing)**
# +
from sklearn.neighbors import KNeighborsRegressor
models = [KNeighborsRegressor(n_neighbors=k) for k in range(1,50)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X ,y ,scale=True, plotta=True, plottaTrain=True,
plottaTest=True, xvalues=range(1,50), xlabel="Numero vicini",
title="Valutazione modelli kNN con score sul training/validation/test" )
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(1,50))[best_model])
# -
# Meglio del predittore banale.
# ## 2) DECISION TREE REGRESSOR
# +
from sklearn.tree import DecisionTreeRegressor
models = [DecisionTreeRegressor(max_leaf_nodes=k) for k in range(2,52)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X, y, plotta=True, plottaTrain=True, plottaTest=True,
xvalues=range(2,52), xlabel="Numero massimo foglie",
title="Valutazione modelli decision tree con score sul training/validation/test" )
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(2,52))[best_model])
# -
# ## 3) LINEAR REGRESSION
# +
from sklearn.linear_model import LinearRegression
model = LinearRegression(fit_intercept=True) # Modello regressione lineare
train, val, test = compute_train_val_test(model, X, y)
print("MSE : ",val)
# -
# ## 4) RANDOM FOREST
# +
from sklearn.ensemble import RandomForestRegressor
models = [RandomForestRegressor(n_estimators=k) for k in range(1,51)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X, y, plotta=True, plottaTrain=True, plottaTest=True,
xvalues=range(1,51), xlabel="Numero alberi",
title="Valutazione modelli decision tree con score sul training/validation/test" )
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(1,51))[best_model])
# -
# Tutti e 4 modelli sono migliori del predittore banale. Il modello migliore è random forest, con un MSE sul validation di 0.00319.
# # AGGIUNTA FEATURES CATEGORIALI
# Aggiungiamo ora le features categoriali: "belongs_to_collection", "homepage", "original_language".
#
# Le features "belongs_to_collection" e "homepage" ho già descritto come le tratto. Invece abbiamo due alternative su come trattare "original_language". Per prima cosa allora aggiungiamo "belongs_to_collection" e "homepage".
#
# La funzione **add_categorial** prende il dataframe df e ritorna newdf, che è il datframe con le sole features categoriali selezionate e da aggiungere (appunto "belongs_to_collection" e "homepage"). Ritorna dunque solo le feature oggetto di studio, lavorate e processate.
#
# Da newdf passiamo alla versione numpy: newX. Concateniamo X e newX in X_tmp.
#
# Sovrascriveremo X con X_tmp solo se vale la pena aggiungere tali features. In questo caso aggiungeremo anche newdf a df.
# +
from lavorazione_dataset import add_categorial
import numpy as np
newdf = add_categorial(df) # newdf è il datframe delle sole feature categoriali in questione, processate e lavorate.
newX = newdf.values # versione numpy
X_tmp = np.append(X,newX,axis=1) # Matrice numpy da valutare
# -
print(newdf.info())
newdf.describe()
# Ora abbiamo 6 features in tutto.
print("X shape: ",X_tmp.shape)
X_tmp[:6,:]
# Valutiamo i 4 algoritmi di learning su tale dataset.
# ## **1) KNN (con preprocessing)**
# +
from sklearn.neighbors import KNeighborsRegressor
models = [KNeighborsRegressor(n_neighbors=k) for k in range(1,50)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X_tmp ,y ,scale=True, plotta=True, plottaTrain=True,
plottaTest=True, xvalues=range(1,50), xlabel="Numero vicini",
title="Valutazione modelli kNN con score sul training/validation/test" )
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(1,50))[best_model])
# -
# ## 2) DECISION TREE REGRESSOR
# +
from sklearn.tree import DecisionTreeRegressor
models = [DecisionTreeRegressor(max_leaf_nodes=k) for k in range(2,52)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X_tmp, y, plotta=True, plottaTrain=True, plottaTest=True,
xvalues=range(2,52), xlabel="Numero massimo foglie",
title="Valutazione modelli decision tree con score sul training/validation/test" )
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(2,52))[best_model])
# -
# ## 3) LINEAR REGRESSION
# +
from sklearn.linear_model import LinearRegression
model = LinearRegression(fit_intercept=True) # Modello regressione lineare
train, val, test = compute_train_val_test(model, X_tmp, y)
print("MSE : ",val)
# -
# ## 4) RANDOM FOREST
# +
from sklearn.ensemble import RandomForestRegressor
models = [RandomForestRegressor(n_estimators=k) for k in range(1,51)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X_tmp, y, plotta=True, plottaTrain=True, plottaTest=True,
xvalues=range(1,51), xlabel="Numero alberi",
title="Valutazione modelli decision tree con score sul training/validation/test" )
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(1,51))[best_model])
# -
# kNN, albero di decisione e regressione lineare rimangono piuttosto stabili rispetto a prima. La random forest migliora il suo MSE sul validation. Dunque ora il modello migliore ora risulta la random forest con anche le features categoriali.
#
# Dunque **aggiungiamo tali features**: riportiamo ciò su X e df.
# +
X = X_tmp
df["belongs_to_collection"] = newdf["belongs_to_collection"]
df["homepage"] = newdf["homepage"]
# -
# **ORIGINAL_LANGUAGE**
#
# Aggiungiamo ora la feature "original_language". Abbiamo due alternative su come trattare "original_language". La funzione **add_language_1** esegue la prima alternativa, mentre la funzione **add_language_2** esegue la seconda alternativa.
#
# Entrambe le funzioni prendono in input il dataframe df e ritornano newdf, ovvero il datframe delle features selezionate e lavorate. Come prima, da newdf estraiamo newX. add_language_1 --> newdf1 --> newX1 ; add_language_2 --> newdf2 --> newX2.
#
# Concateniamo X con newX1 in X_tmp1 e X con con newX2 in X_tmp2. Valutiamo quale alternativa è migliore e sovrascriviamo sulla base di ciò X e df.
# **Alternativa 1**
# "original_language" diventa semplicemente una feature dummy : vale 1 se il film è in lingua inglese, 0 altrimenti.
# Aggiungiamo dunque un ulteriore feature.
# +
from lavorazione_dataset import add_language_1
newdf1 = add_language_1(df)
newX = newdf1.values
X_tmp1 = np.append(X,newX,axis=1)
# -
print(newdf1.info())
newdf1.describe()
# Ora abbiamo 7 features selezionate in tutto.
print("X shape: ",X_tmp1.shape)
X_tmp1[:6,:]
# ## **1) KNN (con preprocessing)**
# +
from sklearn.neighbors import KNeighborsRegressor
models = [KNeighborsRegressor(n_neighbors=k) for k in range(1,50)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X_tmp1 ,y ,scale=True, plotta=True, plottaTrain=True,
plottaTest=True, xvalues=range(1,50), xlabel="Numero vicini",
title="Valutazione modelli kNN con score sul training/validation/test" )
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(1,50))[best_model])
# -
# ## 2) DECISION TREE REGRESSOR
# +
from sklearn.tree import DecisionTreeRegressor
models = [DecisionTreeRegressor(max_leaf_nodes=k) for k in range(2,52)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X_tmp1, y, plotta=True, plottaTrain=True, plottaTest=True,
xvalues=range(2,52), xlabel="Numero massimo foglie",
title="Valutazione modelli decision tree con score sul training/validation/test" )
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(2,52))[best_model])
# -
# ## 3) LINEAR REGRESSION
# +
from sklearn.linear_model import LinearRegression
model = LinearRegression(fit_intercept=True) # Modello regressione lineare
train, val, test = compute_train_val_test(model, X_tmp1, y)
print("MSE : ",val)
# -
# ## 4) RANDOM FOREST
# +
from sklearn.ensemble import RandomForestRegressor
models = [RandomForestRegressor(n_estimators=k) for k in range(1,51)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X_tmp1, y, plotta=True, plottaTrain=True, plottaTest=True,
xvalues=range(1,51), xlabel="Numero alberi",
title="Valutazione modelli decision tree con score sul training/validation/test" )
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(1,51))[best_model])
# -
# C'è un miglioramento complessivo dei modelli. Ed in particolare la random forest migliora molto il suo MSE sul validation. Dunque ora il modello migliore risulta la random forest con anche "original_language" trattata nella prima alternativa.
# **Alternativa 2**
# Le prime 6 lingue rispetto al revenue medio le tengo come valori categoriali distinti. Tutte le altre lingue le accorpo nella categoria "other_language". Ho dunque una variabile categoriale con 7 valori distinti: tale feature la trasformo in 7 variabili binarie(dummy).
#
# Aggiungo in totale 7 feature in più.
# +
from lavorazione_dataset import add_language_2
newdf2 = add_language_2(df)
newX = newdf2.values
X_tmp2 = np.append(X,newX,axis=1)
# -
print(newdf2.info())
newdf2.describe()
# Ora abbiamo 14 features selezionate in tutto (6 di prima + 8 di ora).
print("X shape: ",X_tmp2.shape)
X_tmp2[:6,:]
# ## **1) KNN (con preprocessing)**
# +
from sklearn.neighbors import KNeighborsRegressor
models = [KNeighborsRegressor(n_neighbors=k) for k in range(1,50)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X_tmp2 ,y ,scale=True, plotta=True, plottaTrain=True,
plottaTest=True, xvalues=range(1,50), xlabel="Numero vicini",
title="Valutazione modelli kNN con score sul training/validation/test" )
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(1,50))[best_model])
# -
# ## 2) DECISION TREE REGRESSOR
# +
from sklearn.tree import DecisionTreeRegressor
models = [DecisionTreeRegressor(max_leaf_nodes=k) for k in range(2,52)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X_tmp2, y, plotta=True, plottaTrain=True, plottaTest=True,
xvalues=range(2,52), xlabel="Numero massimo foglie",
title="Valutazione modelli decision tree con score sul training/validation/test" )
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(2,52))[best_model])
# -
# ## 3) LINEAR REGRESSION
# +
from sklearn.linear_model import LinearRegression
model = LinearRegression(fit_intercept=True) # Modello regressione lineare
train, val, test = compute_train_val_test(model, X_tmp2, y)
print("MSE : ",val)
# -
# ## 4) RANDOM FOREST
# +
from sklearn.ensemble import RandomForestRegressor
models = [RandomForestRegressor(n_estimators=k) for k in range(1,51)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X_tmp2, y, plotta=True, plottaTrain=True, plottaTest=True,
xvalues=range(1,51), xlabel="Numero alberi",
title="Valutazione modelli decision tree con score sul training/validation/test" )
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(1,51))[best_model])
# -
# Gli score sono molto simili a quelli dell'alternativa 1. In ogni caso, lo score sul validation della random forest è peggiorato: dunque il MSE migliore c'era con l'alternativa 1. **Scegliamo l'alternativa 1.**
X = X_tmp1
df["original_language"] = newdf1["original_language"]
# # AGGIUNTA FEATURES DATA
# Aggiungiamo l'unica feature di tipo data: "release_date". Come visto, estraiamo da tale feature sia l'anno che il mese. Per l'anno lo trattiamo banalmente come feature numerica, per il mese invece abbiamo 6 alternative diverse da valutare.
# ### ALTERNATIVA 1
def int_to_month(n):
s = ""
if(n==0):
s = "gen"
elif(n==1):
s = "feb"
elif(n==2):
s = "mar"
elif(n==3):
s = "apr"
elif(n==4):
s = "may"
elif(n==5):
s = "jun"
elif(n==6):
s = "jul"
elif(n==7):
s = "aug"
elif(n==8):
s = "sep"
elif(n==9):
s = "oct"
elif(n==10):
s = "nov"
elif(n==11):
s = "dec"
else:
raise RuntimeError("Month error: ",n)
return s
X , y, df = cleaning_data()
scaler=MinMaxScaler()
scaler.fit(y.reshape(df.shape[0],1))
y = scaler.transform(y.reshape(df.shape[0],1)).reshape(df.shape[0],)
newX = add_categorial(df)
X = np.append(X,newX,axis=1)
# +
# year , month_group_1 , month_group_2 , month_group_3
def add_data(df):
feat_list = []
# PULIZIA FEATURE DATA
# year
df["year"] = df["release_date"].map(lambda s : int(s.split("/")[2]))
df["year"] = df["year"].map(lambda y : 1900+y if (y>=20 and y<=99) else 2000+y)
# month
df["month"] = df["release_date"].map(lambda s : int(s.split("/")[0])-1) # da 0 a 11, per mapping migliore con indici
best_months = np.argsort([df["revenue"][df["month"]==m].mean() for m in range(0,12)])[::-1]
def transform(m): # ALT 4
if m in best_months[0:4]:
return "month_group_1"
elif m in list(best_months[4:8]):
return "month_group_2"
elif m in list(best_months[8:12]):
return "month_group_3"
else:
raise RuntimeError("Mese invalido: ",m)
df["month"].map(transform)
newdf = pd.get_dummies(df["month"].map(transform))
for col in newdf.columns:
df[col] = newdf[col]
feat_list.extend(["year","month_group_1","month_group_2","month_group_3"])
return df[feat_list].values
newX = add_data(df)
X_tmp = np.append(X,newX,axis=1)
X_tmp.shape
# -
print(df.info())
df.describe()
X_tmp[:6,:]
# ## **1) KNN (con preprocessing)**
# +
from sklearn.neighbors import KNeighborsRegressor
models = [KNeighborsRegressor(n_neighbors=k) for k in range(1,50)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X_tmp ,y ,scale=True,regr=True, plotta=True, plottaTrain=True,
plottaTest=True, xvalues=range(1,50), xlabel="k")
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(1,50))[best_model])
# -
# Provare standard scaler
# ## 2) DECISION TREE REGRESSOR
# +
from sklearn.tree import DecisionTreeRegressor
models = [DecisionTreeRegressor(max_leaf_nodes=k) for k in range(2,52)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X_tmp, y, scale=False,
regr=True, plotta=True, plottaTrain=True, plottaTest=True,
xvalues=range(2,52), xlabel="Numero massimo foglie")
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(2,52))[best_model])
# -
# ## 3) LINEAR REGRESSION
# +
from sklearn.linear_model import LinearRegression
models = [LinearRegression(fit_intercept=True)]
lista_trainValTest , best_k = model_selection_TrainValTest(models, X_tmp, y, regr=True, plotta=False, plottaTrain=True)
print("MSE : ",lista_trainValTest[best_k][1])
# -
# ## 4) RANDOM FOREST
# +
from sklearn.ensemble import RandomForestRegressor
models = [RandomForestRegressor(n_estimators=k) for k in range(1,51)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X_tmp, y, scale=False,
regr=True, plotta=True, plottaTrain=True, plottaTest=True,
xvalues=range(1,51), xlabel="Numero alberi")
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(1,51))[best_model])
# -
# ### ALTERNATIVA 2
X , y, df = cleaning_data()
scaler=MinMaxScaler()
scaler.fit(y.reshape(df.shape[0],1))
y = scaler.transform(y.reshape(df.shape[0],1)).reshape(df.shape[0],)
newX = add_categorial(df)
X = np.append(X,newX,axis=1)
# +
# 'year', 'sep', 'oct', 'dec', 'aug', 'apr', 'mar', 'jun', 'may', 'feb', 'other_month'
def add_data(df):
feat_list = []
# PULIZIA FEATURE DATA
# year
df["year"] = df["release_date"].map(lambda s : int(s.split("/")[2]))
df["year"] = df["year"].map(lambda y : 1900+y if (y>=20 and y<=99) else 2000+y)
# month
df["month"] = df["release_date"].map(lambda s : int(s.split("/")[0])-1) # da 0 a 11, per mapping migliore con indici
# ALT 5
numb_per_month = [ df[df["month"]==m].shape[0] for m in range(0,12)]
months_sorted = np.argsort(numb_per_month)[::-1]
for m in months_sorted[:9]: #5 #10
df[int_to_month(m)] = (df["month"]==m).astype(int)
df["other_month"] = df["month"].map(lambda m : 1 if m in months_sorted[9:] else 0)
feat_list.append("year")
feat_list.extend([ int_to_month(m) for m in months_sorted[:9]])
feat_list.append("other_month")
return df[feat_list].values
newX = add_data(df)
X_tmp = np.append(X,newX,axis=1)
X.shape
# -
print(df.info())
df.describe()
X_tmp[:6,:]
# ## **1) KNN (con preprocessing)**
# +
from sklearn.neighbors import KNeighborsRegressor
models = [KNeighborsRegressor(n_neighbors=k) for k in range(1,50)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X_tmp ,y ,scale=True,regr=True, plotta=True, plottaTrain=True,
plottaTest=True, xvalues=range(1,50), xlabel="k")
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(1,50))[best_model])
# -
# Provare standard scaler
# ## 2) DECISION TREE REGRESSOR
# +
from sklearn.tree import DecisionTreeRegressor
models = [DecisionTreeRegressor(max_leaf_nodes=k) for k in range(2,52)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X_tmp, y, scale=False,
regr=True, plotta=True, plottaTrain=True, plottaTest=True,
xvalues=range(2,52), xlabel="Numero massimo foglie")
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(2,52))[best_model])
# -
# ## 3) LINEAR REGRESSION
# +
from sklearn.linear_model import LinearRegression
models = [LinearRegression(fit_intercept=True)]
lista_trainValTest , best_k = model_selection_TrainValTest(models, X_tmp, y, regr=True, plotta=False, plottaTrain=True)
print("MSE : ",lista_trainValTest[best_k][1])
# -
# ## 4) RANDOM FOREST
# +
from sklearn.ensemble import RandomForestRegressor
models = [RandomForestRegressor(n_estimators=k) for k in range(1,51)]
list_trainValTest , best_model = model_selection_TrainValTest(models, X_tmp, y, scale=False,
regr=True, plotta=True, plottaTrain=True, plottaTest=True,
xvalues=range(1,51), xlabel="Numero alberi")
print("MSE migliore: ",list_trainValTest[best_model][1]," | k: ",list(range(1,51))[best_model])
# -
X = X_tmp
| src/.ipynb_checkpoints/Progetto_BanNumCatData_COPIA-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Implementation of Word2Vec and FastText Word Embedding with Gensim
# # Libraries
# +
import pandas as pd
import numpy as np
import re
from re import sub
import multiprocessing
from unidecode import unidecode
from gensim.models.phrases import Phrases, Phraser
from gensim.models import Word2Vec
from gensim.test.utils import get_tmpfile
from gensim.models import KeyedVectors
import string
from string import digits
from time import time
from collections import defaultdict
import logging # Setting up the loggings to monitor gensim
logging.basicConfig(format="%(levelname)s - %(asctime)s: %(message)s", datefmt= '%H:%M:%S', level=logging.INFO)
# -
# # Load data source
source_dir = '../data/yorubaDS2020/'
df = pd.read_table(source_dir + 'yoruba_on_tweets.txt', names=['text'], encoding='utf-8-sig')
def text_to_word_list(text, remove_polish_letters):
''' Pre process and convert texts to a list of words
method inspired by method from eliorc github repo: https://github.com/eliorc/Medium/blob/master/MaLSTM.ipynb'''
# Remove all numbers from text
remove_digits = str.maketrans('', '', digits)
text = sub("[0123456789]", "", text)
text = sub(" +", " ", text)
text = str(text)
text = text.lower()
return text
df.text = df.text.apply(lambda x: text_to_word_list(x, unidecode))
df[:2]
yoruba_model = df.copy()
yoruba_model = yoruba_model[yoruba_model.text.str.len()>1]
sent = [row for row in yoruba_model.text]
phrases = Phrases(sent, min_count=1, progress_per=50000)
bigram = Phraser(phrases)
sentences = bigram[sent]
sentences[0:5]
model_word2vec = Word2Vec(min_count=3,window=4,size=300,sample=1e-5,
alpha=0.03, min_alpha=0.0007, negative=20,
sg=0, workers=multiprocessing.cpu_count()-1)
# size: The number of dimensions of the embeddings and the default is 100.
# window: The maximum distance between a target word and words around the target word. The default window is 5.
# min_count: The minimum count of words to consider when training the model;
# words with occurrence less than this count will be ignored. The default for min_count is 5.
# sg: it is used to indicate skip-gram or CBOW but when CBOW=0 or skip gram=1
start = time()
model_word2vec.build_vocab(sentences, progress_per=50000)
print('Time to build vocab: {} mins'.format(round((time() - start) / 60, 2)))
start = time()
model_word2vec.train(sentences, total_examples=model_word2vec.corpus_count, epochs=30, report_delay=1)
print('Time to train the model: {} mins'.format(round((time() - start) / 60, 2)))
model_word2vec.init_sims(replace=True)
model_word2vec.save("yoruba_word2vec.model")
file_export = yoruba_model.copy()
file_export['old_title'] = file_export.text
file_export.old_title = file_export.old_title.str.join('')
file_export.text = file_export.text.apply(lambda x: ''.join(bigram[x]))
file_export[['text']].to_csv('cleaned_yoruba_on_tweet.csv', index=False)
file_export
# # T-SNE Visualizations
def display_closestwords_tsnescatterplot(model, word, size):
arr = np.empty((0,size), dtype='f')
word_labels = [word]
close_words = model.similar_by_word(word)
arr = np.append(arr, np.array([model[word]]), axis=0)
for wrd_score in close_words:
wrd_vector = model[wrd_score[0]]
word_labels.append(wrd_score[0])
arr = np.append(arr, np.array([wrd_vector]), axis=0)
tsne = TSNE(n_components=2, random_state=0)
np.set_printoptions(suppress=True)
Y = tsne.fit_transform(arr)
x_coords = Y[:, 0]
y_coords = Y[:, 1]
plt.scatter(x_coords, y_coords)
for label, x, y in zip(word_labels, x_coords, y_coords):
plt.annotate(label, xy=(x, y), xytext=(0, 0), textcoords='offset points')
plt.xlim(x_coords.min()+0.00005, x_coords.max()+0.00005)
plt.ylim(y_coords.min()+0.00005, y_coords.max()+0.00005)
plt.show()
display_closestwords_tsnescatterplot(model_word2vec, 'gba', 50)
# # FastText
from gensim.models import FastText
model_fastText = FastText(sentences, size=100, window=5, min_count=5, workers=4,sg=1)
| Word2Vect_yoruba.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (herschelhelp_internal)
# language: python
# name: helpint
# ---
# # AKARI-NEP master catalogue
# ## Preparation of Spitzer datafusion SERVS data
#
# The Spitzer catalogues are available in `dmu0_NEP-Spitzer`.
#
# In the catalouge, we keep:
#
# - The internal identifier (this one is only in HeDaM data);
# - The position;
# - The fluxes in aperture 2 (1.9 arcsec); CHECK!
# - The “auto” flux (which seems to be the Kron flux);
# - The stellarity in each band
#
#
from herschelhelp_internal import git_version
print("This notebook was run with herschelhelp_internal version: \n{}".format(git_version()))
import datetime
print("This notebook was executed on: \n{}".format(datetime.datetime.now()))
# +
# %matplotlib inline
# #%config InlineBackend.figure_format = 'svg'
import matplotlib.pyplot as plt
plt.rc('figure', figsize=(10, 6))
from collections import OrderedDict
import os
from astropy import units as u
from astropy.coordinates import SkyCoord
from astropy.table import Column, Table
import numpy as np
from herschelhelp_internal.flagging import gaia_flag_column
from herschelhelp_internal.masterlist import nb_astcor_diag_plot, remove_duplicates
from herschelhelp_internal.utils import astrometric_correction, mag_to_flux
# +
OUT_DIR = os.environ.get('TMP_DIR', "./data_tmp")
try:
os.makedirs(OUT_DIR)
except FileExistsError:
pass
RA_COL = "nep_ra"
DEC_COL = "nep_dec"
# -
# ## I - Column selection
# +
imported_columns = OrderedDict({
'nep_id': "nep_id",
'ra': "nep_ra",
'dec': "nep_dec",
'm_irac_i1': "m_irac_i1",
'merr_irac_i1': "merr_irac_i1",
'm_ap2_irac_i1': "m_ap_irac_i1",
'merr_ap2_irac_i1': "merr_ap_irac_i1",
'm_irac_i2': "m_irac_i2",
'merr_irac_i2': "merr_irac_i2",
'm_ap2_irac_i2': "m_ap_irac_i2",
'merr_ap2_irac_i2': "merr_ap_irac_i2",
'irac_stellarity': "irac_stellarity",
})
catalogue = Table.read("../../dmu0/dmu0_NEP-Spitzer/data/NEP-Spitzer-APJ.fits")[list(imported_columns)]
for column in imported_columns:
catalogue[column].name = imported_columns[column]
epoch = 2017 #Paper year
# Clean table metadata
catalogue.meta = None
# -
# Adding magnitude and band-flag columns
for col in catalogue.colnames:
if col.startswith('m_'):
errcol = "merr{}".format(col[1:])
catalogue[col][catalogue[col] > 90.] = np.nan
catalogue[errcol][catalogue[errcol] > 90.] = np.nan
flux, error = mag_to_flux(
np.array(catalogue[col]), np.array(catalogue[errcol]))
# Note that some fluxes are 0.
catalogue.add_column(Column(flux*1.e6, name="f{}".format(col[1:])))
catalogue.add_column(Column(error*1.e6, name="f{}".format(errcol[1:])))
# Band-flag column
if "ap" not in col:
catalogue.add_column(Column(np.zeros(len(catalogue), dtype=bool), name="flag{}".format(col[1:])))
catalogue[:10].show_in_notebook()
# ## II - Removal of duplicated sources
# We remove duplicated objects from the input catalogues.
# +
SORT_COLS = ['ferr_ap_irac_i1', 'ferr_ap_irac_i2']
FLAG_NAME = "nep_flag_cleaned"
nb_orig_sources = len(catalogue)
catalogue = remove_duplicates(catalogue, RA_COL, DEC_COL, sort_col=SORT_COLS,flag_name=FLAG_NAME)
nb_sources = len(catalogue)
print("The initial catalogue had {} sources.".format(nb_orig_sources))
print("The cleaned catalogue has {} sources ({} removed).".format(nb_sources, nb_orig_sources - nb_sources))
print("The cleaned catalogue has {} sources flagged as having been cleaned".format(np.sum(catalogue[FLAG_NAME])))
# -
# ## III - Astrometry correction
#
# We match the astrometry to the Gaia one. We limit the Gaia catalogue to sources with a g band flux between the 30th and the 70th percentile. Some quick tests show that this give the lower dispersion in the results.
gaia = Table.read("../../dmu0/dmu0_GAIA/data/GAIA_AKARI-NEP.fits")
gaia_coords = SkyCoord(gaia['ra'], gaia['dec'])
nb_astcor_diag_plot(catalogue[RA_COL], catalogue[DEC_COL],
gaia_coords.ra, gaia_coords.dec)
# +
delta_ra, delta_dec = astrometric_correction(
SkyCoord(catalogue[RA_COL], catalogue[DEC_COL]),
gaia_coords
)
print("RA correction: {}".format(delta_ra))
print("Dec correction: {}".format(delta_dec))
# -
catalogue[RA_COL] = catalogue[RA_COL] + delta_ra.to(u.deg)
catalogue[DEC_COL] = catalogue[DEC_COL] + delta_dec.to(u.deg)
nb_astcor_diag_plot(catalogue[RA_COL], catalogue[DEC_COL],
gaia_coords.ra, gaia_coords.dec)
# ## IV - Flagging Gaia objects
catalogue.add_column(
gaia_flag_column(SkyCoord(catalogue[RA_COL], catalogue[DEC_COL]), epoch, gaia)
)
# +
GAIA_FLAG_NAME = "nep_flag_gaia"
catalogue['flag_gaia'].name = GAIA_FLAG_NAME
print("{} sources flagged.".format(np.sum(catalogue[GAIA_FLAG_NAME] > 0)))
# -
# ## V - Saving to disk
catalogue.write("{}/NEP-Spitzer.fits".format(OUT_DIR), overwrite=True)
| dmu1/dmu1_ml_AKARI-NEP/1.3_NEP-Spitzer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
df = pd.read_csv(r'C:\Users\chosun\Desktop\AN\taxi\train.csv',nrows=2_000_000, usecols=[1,2,3,4,5,6,7])
df['pickup_datetime'] = df['pickup_datetime'].str.slice(0, 16)
df['pickup_datetime'] = pd.to_datetime(df['pickup_datetime'], utc=True, format='%Y-%m-%d %H:%M')
# +
df.dropna(how='any', axis='rows', inplace=True)
mask = df['pickup_longitude'].between(-75, -73)
mask &= df['dropoff_longitude'].between(-75, -73)
mask &= df['pickup_latitude'].between(40, 42)
mask &= df['dropoff_latitude'].between(40, 42)
mask &= df['passenger_count'].between(0, 8)
mask &= df['fare_amount'].between(0, 250)
df = df[mask]
# -
def dist(pickup_lat, pickup_long, dropoff_lat, dropoff_long):
distance = np.abs(dropoff_lat - pickup_lat) + np.abs(dropoff_long - pickup_long)
return distance
# +
def transform(data):
# Extract date attributes and then drop the pickup_datetime column
data['hour'] = data['pickup_datetime'].dt.hour
data['day'] = data['pickup_datetime'].dt.day
data['month'] = data['pickup_datetime'].dt.month
data['year'] = data['pickup_datetime'].dt.year
data = data.drop('pickup_datetime', axis=1)
# Distances to nearby airports, and city center
# By reporting distances to these points, the model can somewhat triangulate other locations of interest
nyc = (-74.0063889, 40.7141667)
jfk = (-73.7822222222, 40.6441666667)
ewr = (-74.175, 40.69)
lgr = (-73.87, 40.77)
data['distance_to_center'] = dist(nyc[1], nyc[0],
data['pickup_latitude'], data['pickup_longitude'])
data['pickup_distance_to_jfk'] = dist(jfk[1], jfk[0],
data['pickup_latitude'], data['pickup_longitude'])
data['dropoff_distance_to_jfk'] = dist(jfk[1], jfk[0],
data['dropoff_latitude'], data['dropoff_longitude'])
data['pickup_distance_to_ewr'] = dist(ewr[1], ewr[0],
data['pickup_latitude'], data['pickup_longitude'])
data['dropoff_distance_to_ewr'] = dist(ewr[1], ewr[0],
data['dropoff_latitude'], data['dropoff_longitude'])
data['pickup_distance_to_lgr'] = dist(lgr[1], lgr[0],
data['pickup_latitude'], data['pickup_longitude'])
data['dropoff_distance_to_lgr'] = dist(lgr[1], lgr[0],
data['dropoff_latitude'], data['dropoff_longitude'])
data['long_dist'] = data['pickup_longitude'] - data['dropoff_longitude']
data['lat_dist'] = data['pickup_latitude'] - data['dropoff_latitude']
data['dist'] = dist(data['pickup_latitude'], data['pickup_longitude'],
data['dropoff_latitude'], data['dropoff_longitude'])
return data
df = transform(df)
# -
import xgboost as xgb
from bayes_opt import BayesianOptimization
from sklearn.metrics import mean_squared_error
| Untitled1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/vinaworld2/let-supgrate/blob/master/Assignment_no_8.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="-FUWuFYfuQ5Y" colab_type="code" colab={}
Assignment no 8
# + id="eXOcNj8GuU3x" colab_type="code" colab={}
def recur_cache(func):
cache = {}
def wrapper(arg):
if arg not in cache:
cache[arg] = func(arg)
return cache[arg]
return wrapper
@recur_cache
def fib(N):
if N < 0 or N == None :
return None
if N < 2:
return N
else:
return fib(N-1) + fib(N-2)
# + id="KMwXn19UusyH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="216624cc-e20b-4ccc-a58e-d20e1eabca61"
fib(3)
# + id="1Yc_zYXGu-zC" colab_type="code" colab={}
Que 2
# + id="pPl26NiXvAFY" colab_type="code" colab={}
file=open("vinaya.txt","w")
file.write("Hi, i am <NAME> learning python")
file.close()
# + id="L2wJQTAYvSvZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="45f32ff9-1d33-4918-8af2-e5c37b251617"
file = open("vinaya.txt","r")
fileData = file.read()
print(fileData)
file.close()
# + id="b3xQsSBWvdOn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="06d01f92-2977-4c9d-a9b6-304fb5e3b546"
try :
file = open("vinaya.txt","r")
file.write("Hellow")
file.close()
except Exception as e:
print(e)
| Assignment_no_8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Big data analysis homework 2
# ## Name: liusheng
# ## ID:1801212891
# # Problem 1
# Loading data first
# +
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
import warnings
warnings.filterwarnings("ignore")
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
data = pd.read_csv("climate_change_1.csv", index_col=0)
train_data = data.loc[:2006,:]
test_data = data.loc[2007:,:]
train_data.shape
data.head()
# -
# ### 1. Implement a function closed_form_1 that computes this closed form solution given the features 𝐗, labels Y (using Python or Matlab).
#
# Apparently, the some variables in data set are highly correlated, so the correlation coefficient of the correlated variables are unstable.For example, CO2 and N2O, CH4 and CFC-12, is highly correlated, the correlation coefficeints of those variables are not trusted
# +
temp = train_data.iloc[:,1:-1].corr()
temp[temp>0.9]
def closed_form_1(x,y, constant = True):
if constant:
x["Constant"]=1
columns = x.columns
x = np.matrix(x)
y = np.matrix(y).T
coeff = (x.T@x).I@(x.T@y)
result = pd.Series(coeff.A1, index=columns,
name="correlation coefficient")
return result
coeff = closed_form_1(train_data.iloc[:,1:-1], train_data.Temp)
coeff
# -
# ### 2.Write down the mathematical formula for the linear model and evaluate the model R2 on the training set and the testing set.
# The mathematical formula for the linear model is
# * $$y=X\beta+\epsilon$$
#
# The formula of $R^2$ is
# * $$R^2 = \frac{\Sigma_i(\hat y_i-\bar y_i)^2}{\Sigma_i(y_i-\bar y_i)^2}$$
# +
def r_square(coeff,x,y,constant = True):
if constant:
x["Constant"]=1
coeff = np.matrix(coeff)
x = np.matrix(x)
y = y.values
y_hat = (x@coeff.T).A1
var1 = ((y_hat-y_hat.mean())**2).sum()
var2 = ((y-y.mean())**2).sum()
r2 = var1/var2
return r2
print("r2 of training_data is", r_square(coeff,train_data.iloc[:,1:-1],train_data.Temp))
print("r2 of testing_data is", r_square(coeff,test_data.iloc[:,1:-1],test_data.Temp))
# -
# ### 3.Which variables are significant in the model?
# We evaluate the t-test on the model
# * The t-test based on the assumption $\epsilon|X\sim N(0,\sigma^2I_n)$
# * So, $({\hat\beta}-\beta)|X\sim N(0,\sigma^2(X'X)^{-1})$
# * $\sigma^2$ is unknown which should be replace by the estimator $s^2 = \frac1{n-k}\Sigma e_i^2$
#
# As we see in the result, the t-value of CH4 is only 0.2404, less than the critical value $t_{200,0.9}=1.653$,so it's not significant.<br>But,as we mentioned before,CO2 and N2O, CH4 and CFC-12, is highly correlated, the t value of those variables are not trusted.
# +
def t_value(coeff,x,y,constant=True):
if constant:
x["Constant"]=1
coeff = np.matrix(coeff)
index = x.columns
x = np.matrix(x)
y = np.matrix(y).T
y_hat = (x@coeff.T)
s2 = np.power(y-y_hat,2).sum()/(x.shape[0]-x.shape[1])
t = coeff/np.power(s2*(x.T@x).I.diagonal(),0.5)
result = pd.Series(t.A1, index=index, name="t value")
return result
t_result = t_value(coeff,train_data.iloc[:,1:-1],train_data.Temp)
t_result
t_result[t_result.abs()<1.653]
# -
# ### 4. Write down the necessary conditions for using the closed form solution. And you can apply it to the dataset climate_change_2.csv, explain the solution is unreasonable.
# The condition for the OLS is
# 1. $E(\epsilon|X)=0$
# 2. $(XX')^{-1} existed$
# 3. $Var(X^4)<\infty,Var(y^4)<\infty$
# 4. $E(\epsilon'\epsilon)=\sigma^2I$
#
# As we see, in data set 2, the covariance matrix shows the correlation coefficient between CO2 and N2O, CFC-12 and CH4 is pretty high, and N2O and CH4 is identical in trend, which violate the rule 2. So, the analytical solution is sensetive to the original value and unreasonable.
data2 = pd.read_csv("climate_change_2.csv", index_col=0)
corr = data2.loc[:,"MEI":"NO"].corr()
corr[corr>0.9]
train_data2 = data2.loc[:2006,:]
test_data2 = data2.loc[2007:,:]
coeff2 = closed_form_1(train_data2.iloc[:,1:-1], train_data2.Temp)
coeff2
r_result2 = r_square(coeff2,train_data2.iloc[:,1:-1],train_data2.Temp)
r_result2
t_result2 = t_value(coeff2,train_data2.iloc[:,1:-1],train_data2.Temp)
t_result2
# #### we compare the result with third-party packages
import statsmodels.api as sm
model = sm.OLS(train_data.Temp, sm.add_constant(train_data.iloc[:,1:-1])).fit()
model.summary()
# # Problem2
# ### 1. Please write down the loss function for linear model with L1 regularization, L2 regularization, respectively.**
# * The loss function with $L_1$ regularization $$(Y-X\beta)'(Y-X\beta)+\alpha e'|\beta|$$
# * The loss function with $L_2$ regularization $$(Y-X\beta)'(Y-X\beta)+\lambda \beta'\beta$$
# ### 2.The closed form solution for linear model with L2 regularization:
# $$\theta = (X^TX+\lambda I)^{-1}X^TY$$
# ### where I is the identity matrix. Write a function closed_form_2 that computes this closed form solution given the features X, labels Y and the regularization parameter $\lambda$
# +
def closed_form_2(x,y,lam=0,constant = True):
if constant:
x["Constant"]=1
columns = x.columns
x = np.matrix(x)
y = np.matrix(y).T
coeff = (x.T@x + lam*np.eye(x.shape[1])).I@(x.T@y)
result = pd.Series(coeff.A1, index=columns,
name="correlation coefficient")
return result
#we test the function when \lambda = 1e-3
coeff_l2 = closed_form_2(train_data.iloc[:,1:-1], train_data.Temp,1e-3)
coeff_l2
# -
# ### 3.Compare the two solutions in problem 1 and problem 2 and explain the reason why linear model with L2 regularization is robust. (using climate_change_1.csv)
# The all correlation coefficent value of Ridge regression shrinked compared with the OLS regression.When we add punishment on the correlation coefficent, the absolute value of them will decrease, so it's more stable compared to the original one.
print("correlation coefficient of OLS is \n", coeff)
print("\n")
print("correlation coefficient of Ridge regresion is \n", coeff_l2)
# ### 4. You can change the regularization parameter λ to get different solutions for this problem. Suppose we set λ = 10, 1, 0.1, 0.01, 0.001, and please evaluate the model R2 on the training set and the testing set. Finally, please decide the best regularization parameter $\lambda$.
#
# * At first, we assume the we have the constant term.We should choose the $\lambda$ at 1e-6, as the $R^2$ is stable after 1e-6
# +
lam_set = [10**i for i in range(-15,2)]
lam_set.reverse()
r_result = []
for lam in lam_set:
coeff = closed_form_2(train_data.iloc[:,1:-1], train_data.Temp, lam=lam)
r2_test = r_square(coeff,test_data.iloc[:,1:-1],test_data.Temp)
r2_train = r_square(coeff,train_data.iloc[:,1:-1],train_data.Temp)
r_result.append((r2_test,r2_train))
temp = np.array(r_result)
plt.xscale('log')
plt.plot(lam_set,temp[:,0], label = "train_data")
plt.plot(lam_set,temp[:,1], label = "test_data")
plt.ylabel("$R^2$")
plt.xlabel("$\lambda$")
plt.legend(loc='best')
plt.show()
# -
# Then, we assume the we don't have the constant term.We should choose the 𝜆 at 1e-2, as the 𝑅2 is stable after 1e-6
# +
lam_set = [10**i for i in range(-5,2)]
lam_set.reverse()
r_result = []
for lam in lam_set:
coeff = closed_form_2(train_data.iloc[:,1:-1], train_data.Temp, lam=lam, constant = False)
r2_test = r_square(coeff,test_data.iloc[:,1:-1],test_data.Temp, constant = False)
r2_train = r_square(coeff,train_data.iloc[:,1:-1],train_data.Temp, constant = False)
r_result.append((r2_test,r2_train))
temp = np.array(r_result)
plt.xscale('log')
plt.plot(lam_set,temp[:,0], label = "train_data")
plt.plot(lam_set,temp[:,1], label = "test_data")
plt.ylabel("$R^2$")
plt.xlabel("$\lambda$")
plt.legend(loc='best')
plt.show()
# -
# # Problem3
# ### 1. From Problem 1, you can know which variables are significant, therefore you can use less variables to train model. For example, remove highly correlated and redundant features. You can propose a workflow to select feature.
#
# 1. Remove the highly correlated variable first.
# 2. Run regression, remove the insignificant variables step by step
corr = train_data.iloc[:,1:-1].corr()
corr[corr>0.9]
# As we see the CO2 highly correlated with N2O, CH4 highly correlated with CFC-12.<br>
# We exam the variance of each pair variables and remove the less flactuated one.<br>
# So, we need to delete CH4 and N2O.
train_data.iloc[:,1:].std()
# ### 2.Train a better model than the model in Problem 2.
# As the explaining variables will be reduced in the feature selection part, we need to evaluate the $adj-R^2$
# +
def adj_r2(coeff,x,y,constant = True):
if constant:
x["Constant"]=1
coeff = np.matrix(coeff)
x = np.matrix(x)
y = y.values
y_hat = (x@coeff.T).A1
k = x.shape[1]
n = x.shape[0]
var1 = ((y_hat-y_hat.mean())**2).sum()
var2 = ((y-y.mean())**2).sum()
r2 = var1/var2
r2 = 1- (1-r2)*(n-1)/(n-k-1)
return r2
temp = train_data.iloc[:,1:-1].copy()
temp.drop(columns = ["CFC-12", "CH4", "Aerosols"], inplace = True)
coeff = closed_form_1(temp.iloc[:,1:-1], train_data.Temp)
test_temp = test_data.iloc[:,1:-1].copy()
test_temp.drop(columns = ["CFC-12", "CH4","Aerosols"], inplace = True)
print ("The adj_r2 of the new nodel is", adj_r2(coeff,temp.iloc[:,1:-1],train_data.Temp))
print ("The r2 of the test set is", r_square(coeff,test_temp.iloc[:,1:-1],test_data.Temp))
coeff = closed_form_1(train_data.iloc[:,1:-1], train_data.Temp)
print ("The adj_r2 of the origial model is", adj_r2(coeff,train_data.iloc[:,1:-1],train_data.Temp))
print ("The r2 of the test set is", r_square(coeff,test_data.iloc[:,1:-1],test_data.Temp))
# -
# # Problem 4
# **Gradient descent algorithm is an iterative process that takes us to the minimum of a function. Please write down the iterative expression for updating the solution of linear model and implement it using Python or Matlab in gradientDescent function.**
# * Batch based gradient descent iterative formula is$$\beta_n = \beta_{n-1} -\frac{\alpha}{m} X'(X\beta -y)$$
# The X is a batch of random choosed observation
# +
from sklearn import preprocessing
def loss(coeff,x,y):
return (x@coeff-y).T@(x@coeff-y)
def gradientDescent(x,y,iter_max=200000, batch=100,alpha=1e-5,
min_gap=1e-1,constant=True):
record = []
if constant:
x["Constant"]=1
columns = x.columns
x = np.matrix(x)
y = np.matrix(y).T
coeff = np.matrix(np.arange(x.shape[1])/x.shape[1]).T
for i in range(iter_max):
idx = np.random.choice(x.shape[0],batch)
x_batch = x[idx,:]
y_batch = y[idx,:]
iteration = alpha/batch*(x_batch.T@(x_batch@coeff-y_batch))
if np.abs(iteration.A1).all() < min_gap:
print("iteration terminal at %s times"%i)
break
else:
coeff = coeff - iteration
record.append(loss(coeff,x,y)[0,0])
plt.plot(record, label = "loss_function")
plt.ylabel("total_loss")
plt.xlabel("iter_times")
plt.legend()
plt.show()
print("The final loss is ", record[-1])
result = pd.Series(coeff.A1, index=columns, name="correlation coefficient")
return result
min_max_scaler = preprocessing.MinMaxScaler()
scaled_train = min_max_scaler.fit_transform(train_data.iloc[:,1:].values)
scaled_train = pd.DataFrame(scaled_train, columns = train_data.columns[1:])
result_gd = gradientDescent(scaled_train.iloc[:,:-1], scaled_train.Temp)
result_gd
# -
# #### We compare the result with the OLS analytical solution.The result of the gradient descent is better.
coeff_ols = closed_form_1(scaled_train.iloc[:,1:-1], scaled_train.Temp)
coeff_ols
scaled_train_x = scaled_train.iloc[:,1:-1].copy()
scaled_train_x["Constant"]=1
loss(coeff_ols,np.matrix(scaled_train_x),np.matrix(scaled_train.Temp).T)[0,0]
| homework2/Big_Data_Homework2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 1
# +
amount = 1000
interest_rate = 0.1
final_amount = 0
# -
# Use the base amount and interest rate to calculate the final amount after compound interest over 5 years.
# +
# Do your calculations here
# ...
# -
print('The final amoutn is ', final_amount)
# # Exercise 2
# Calculate how many years you need to obtain a certain amount (e.g. 2000) by starting with an initial amount (e.g. 1000) for a given interest rate (e.g. 2%)
# +
amount = 1000
target = 2000
interest_rate = 0.02
number_of_years = 0
# +
# Do your calculations here
# ...
# -
print('The number of years needed is ', number_of_years)
# # Exercise 3
# Remove the duplicates from this list:
name_list = ["Johannes", "Jamal", "Jamal", "Johannes", "Galina"]
# +
# Enter your code here
# ...
# -
# Check the output:
print(name_list)
# # Exercise 4
# Make a list of all the 4th and 6th letters of a word:
words = ["Johannes", "Jamal", "Jamal", "Johannes", "Galina"]
new_list = []
# +
# Enter your code here
# ...
# -
print(new_list)
# # Exercise 5
# Create a list of temperatures in Fahrenheit from another list measured in Celsius using comprehensions:
# +
celsius = [15, 18, 32, 19]
# Fill in the right comprehension
fahrenheit = []
# -
# Check the output:
print(fahrenheit)
# # Exercise 6
# Calculate how many years you need to obtain a certain amount by starting with an initial amount for a given interest rate
amounts = [1000,1200,1300]
target = 2000
interest_rates = [0.02,0.05,0.1]
# You can use your previous code from exercise 2 and reuse it in a function:
# +
# Define function
# ...
# +
# Loop over amounts and interest rates and call function
# Print an informative line of code
# ...
# -
# # Exercise 7
# Calculate the number of ECTS taught by every member in the dictionary
courses = {"Johannes" : "Web Analytics", "Jamal" : "DEA", "Maurizio" : "Simulation"}
ects = {"Web Analytics": 15, "DEA": 15, "Simulation":15}
# Enter your code here
# ...
# # Exercise 8
# Find the most common letter at the third place in a word:
words = ["Johannes", "Jamal", "Jamal", "Adme", "Galina"]
# +
# Enter your code here
# ...
# -
# # Exercise 9
# In this exercise we are going to build a decision tree using scikit-learn and pandas. Code some stuff yourself, or find the available modules that can do it for you:
# +
import pandas as pd
import numpy as np
# Read csv with pandas, index indicates whether the first column is an index
data = pd.read_csv("data/attrition.csv", index_col=0)
print(data.head())
# -
# Attrition is our label, so we can split our data into X and y:
y = data['Attrition']
X = data.drop('Attrition', axis=1)
# Write the pre-processing functions:
# +
# Create training and test set with function
# def splitDataset(X, y ratio):
# ...
# +
# Convert all attributes in data dataframe
# def convertAttribute(column):
# ...
# -
# The pre-processsing:
# +
# # convert all attributes at once
# for column in X.columns:
# X[column] = convertAttribute(X[column])
# y = convertAttribute(y)
# # Use our new function to split according to a certain ratio
# train_X, test_X, train_y, test_y = splitDataset(X, y, 0.7)
# -
# Build decision tree and calculate accuracy:
# +
# # Build tree
# #tree = ...
# tree.fit(train_X, train_y)
# ## Use tree for prediction
# y_pred = tree.predict(test_X)
# #accuracy = ...
# print(accuracy)
| Week0/Week0-Exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import logging
import seaborn as sns
sns.set(style="darkgrid")
# ### Logger
# +
#logging.basicConfig(level=logging.DEBUG, format='%(asctime)s - %(levelname)s - %(message)s')
logger = logging.getLogger()
logger.setLevel(logging.DEBUG)
formatter = logging.Formatter('%(asctime)s - %(levelname)s - %(message)s')
#logging.basicConfig
fh = logging.FileHandler('logger_lineplot.txt')
fh.setLevel(logging.DEBUG)
fh.setFormatter(formatter)
logger.addHandler(fh)
'''
#For streaming only
ch = logging.StreamHandler()
ch.setLevel(logging.DEBUG)
ch.setFormatter(formatter)
logger.addHandler(ch)
'''
logger.info('This is a test log message.')
# -
# ## Render graph functions
# +
def renderGraph(db, workload):
new_db = setCriteria(db, workload)
graph = setLinePlot(new_db,workload)
exportGraph(graph, workload)
def setCriteria(db, workload):
return db[db.workload == workload]
def setLinePlot(db, workload):
dbgraph = sns.lineplot(x=db.record_nbr,
y=db.value,
hue=db.database,
style=db.database,
sort=False,
legend='full',
markers=True,
dashes=True)
handles, labels = dbgraph.get_legend_handles_labels()
dbgraph.legend(handles=handles[1:],
labels=labels[1:])
dbgraph.set(xlabel='Number of Records',
ylabel="Time (seconds)")
dbgraph.set_title(workload.upper())
dbgraph
return dbgraph
def setFileName(workload):
name = workload.replace(" ", "")
filename = "".join((name,'.png'))
return filename
def exportGraph(graph, workload):
name = setFileName(workload)
fig = graph.get_figure()
fig.savefig(name)
fig.show
return
# -
class LinePlot():
def __init__(self, db, workload):
self.db = db
self.workload = workload
#TODO refractor x y graph axis
def graph_ops_vs_latency(self):
new_db = self.set_criteria(self.db, self.workload)
graph = self.set_ops_latency(new_db, self.workload)
self.exportGraph(graph, self.workload)
return
def graph_records_vs_latency(self):
new_db = self.set_criteria(self.db, self.workload)
graph = self.set_records_latency(new_db, self.workload)
self.exportGraph(graph, self.workload)
return
@staticmethod
def set_criteria(db, workload):
return db[db.workload == workload]
@staticmethod
def set_ops_latency(db, workload):
dbgraph = sns.lineplot(x=db.throughput_value,
y=db.runtime_value,
hue=db.database,
style=db.database,
sort=True,
legend='full',
markers=True,
dashes=True)
handles, labels = dbgraph.get_legend_handles_labels()
dbgraph.legend(loc=2,
handles=handles[1:],
labels=labels[1:])
dbgraph.set(xlabel='Throughput(ops/sec)',
ylabel="Read latency(seconds)")
dbgraph.set_title(workload.upper())
return dbgraph
@staticmethod
def set_records_latency(db, workload):
dbgraph = sns.lineplot(x=db.records,
y=db.runtime_value,
hue=db.database,
style=db.database,
sort=False,
legend='full',
markers=True,
dashes=True)
handles, labels = dbgraph.get_legend_handles_labels()
dbgraph.legend(loc=2,
handles=handles[1:],
labels=labels[1:])
dbgraph.set(xlabel='Number of Records',
ylabel="Time (seconds)")
dbgraph.set_title(workload.upper())
return dbgraph
@staticmethod
def setFileName(workload):
extension = ".png"
name = workload.replace(" ", "")
filename = "".join((name,extension))
return filename
@staticmethod
def exportGraph(graph, workload):
graph_directory = 'graphs/' + workload
name = setFileName(graph_directory)
fig = graph.get_figure()
fig.savefig(name)
fig.show
return
# ## Iterate thru the output folder and get root path
# +
import re
from os import path
from os import walk
class FileIterator:
def __init__(self):
self.path_root = 'output'
def iterate_filenames(self):
databases = self.__iterate_database(self.path_root)
files = self.__iterate_file(databases)
return files
def __iterate_database(self, root_dir):
path_dbs = []
databases = ['redis','mongodb','cassandra-cql']
#databases = ['cassandra-cql']
for database in databases:
path_dbs.append(root_dir + '/' + database)
return path_dbs
def __iterate_file(self, path_dbs):
path_db_workload_files = []
for path_db in path_dbs:
for root, dirs, files in walk(path_db):
for filename in files:
if ".ipynb_checkpoints" in dirs:
pass
else:
if ".swp" not in filename and ".ipynb_checkpoints" not in filename and ".DS_Store" not in filename and ".txt~" not in filename:
path_db_workload_files.append(root + '/' + filename)
#logger.debug(root + '/' + filename)
return path_db_workload_files
# -
# ## Set dataframe values
class BuildDataFrame():
def __init__(self, output, filename_path):
self.output = output
self.filename_path = filename_path
self.file = filename_path.split(sep='/')
self.filename = filename_path.split(sep='/')[len(filename_path.split(sep='/'))-1]
def set_data_row(self):
output_data = {}
output_data['run'] = self.__set_run(self.filename)
output_data['database'] = self.__set_database(self.filename_path)
output_data['workload'] = self.__set_workload(self.filename)
output_data['record_nbr'] = self.__set_record_nbr(self.filename)
output_data['records'] = self.__set_records(self.filename)
output_data['type'] = self.__set_type(self.output)
#output_data['label'] = self.__set_label(self.output)
#output_data['value'] = self.__set_value(self.output)
output_data['runtime'] = self.__set_runtime(self.output)
output_data['runtime_value'] = self.__set_runtime_value(self.output)
output_data['throughput'] = self.__set_throughput(self.output)
output_data['throughput_value'] = self.__set_throughput_value(self.output)
return output_data
def __set_runs(self,file):
filename = (file[len(file)-1])
col_run = filename.split(sep='.')[1]
return col_run
def __set_run(self,filename):
col_run = filename.split(sep='.')[1]
return col_run
def __set_database(self, filename_path):
database_name = ''
col_database = filename_path.split(sep='/')[1]
if(col_database == 'cassandra-cql'):
database_name = 'Cassandra'
else:
database_name = col_database.title()
return database_name
def __set_workload(self, filename):
col_workload = filename.split(sep='.')[2]
w_type = col_workload[len(col_workload)-1]
w_base = col_workload[:-1]
workload_name = w_base + ' ' + w_type
return workload_name
def __set_record_nbr(self, filename):
record_nbr = ''
col_record = filename.split(sep='.')[0]
if(col_record == '1k'):
record_nbr = 1000
elif(col_record == '2k'):
record_nbr = 2000
elif(col_record == '3k'):
record_nbr = 3000
elif(col_record == '10k'):
record_nbr = 10000
elif(col_record == '100k'):
record_nbr = 100000
#pass
elif(col_record == '1M'):
record_nbr = 1000000
else:
record_nbr = 100
return record_nbr
def __set_records(self, filename):
record_nbr = ''
col_record = filename.split(sep='.')[0]
if(col_record == '1k'):
record_nbr = '1k'
elif(col_record == '2k'):
record_nbr = '2000'
elif(col_record == '3k'):
record_nbr = '3000'
elif(col_record == '10k'):
record_nbr = '10k'
elif(col_record == '100k'):
record_nbr = '100K'
#pass
elif(col_record == '1M'):
record_nbr = '1M'
else:
record_nbr = '100'
return record_nbr
def __set_type(self, output):
task_type = output.loc[0][0]
return task_type[1:-1]
def __set_label(self, output):
#runtime
#task_label = output.loc[0][1].lstrip()
#throughput
task_label = output.loc[1][1].lstrip()
return task_label
def __set_value(self, output):
#runtime
#seconds = float(output.loc[0][2]) / 1000
#throughput
seconds = float(output.loc[1][2])
return seconds
def __set_runtime(self, output):
#runtime
runtime_label = output.loc[0][1].lstrip()
return runtime_label
def __set_runtime_value(self, output):
#runtime
runtime_value = float(output.loc[0][2]) / 1000
return runtime_value
def __set_throughput(self, output):
throughput_label = output.loc[1][1].lstrip()
return throughput_label
def __set_throughput_value(self, output):
#runtime
#runtime_value = float(output.loc[0][2]) / 1000
#throughput
#throughput_value = float(output.loc[1][2])
throughput_value = float(output.loc[1][2])
return throughput_value
# ### Read from csv files
class CSVReader():
def __init__(self, filename_paths):
self.filename_paths = filename_paths
#print(filename_paths)
def get_data(self):
data = []
for filename_path in self.filename_paths:
#print(filename_path)
file = filename_path.split(sep='/')
filename = (file[len(file)-1])
database = file[1]
if(database == 'mongodb'):
csv_output = pd.read_csv(filename_path, skiprows=1, header=None)
else:
#TODO filter doTransactionReadModifyWrite
csv_output = pd.read_csv(filename_path, header=None)
#TODO refractor
df_output = BuildDataFrame(csv_output, filename_path)
data.append(df_output.set_data_row())
return data
# ## __Main__
def get_latest_data():
files = FileIterator()
file_paths = files.iterate_filenames()
#print(list(file_paths))
csv_files = CSVReader(file_paths)
csv_data = []
csv_data = csv_files.get_data()
return csv_data
# ### Convert source file to dataFrame
# +
def output_df(csv_data):
data_df = pd.DataFrame(csv_data, columns=['run','database','workload','record_nbr','records','type','throughput','throughput_value','runtime','runtime_value'])
data_df = data_df.sort_values(by=['database','workload','record_nbr'])
data_df.head(2)
return data_df
def export_output_df(data_df):
data_df.to_csv('output_df.csv',sep=',')
return true
# -
# ### Add zero based graph rendering
def zero_base():
zero_df = pd.read_csv("zero_df.csv")
zero_df['records'].astype('str')
zero_df['records'] = zero_df['records'].apply(str)
zero_df.head(2)
return zero_df
# ### Concat all output csv
def concat_output(zero_df, data_df):
db3 = pd.concat([zero_df,data_df], ignore_index=True)
db3.to_csv('concat_df.csv',sep=',')
return db3
# ## BUILD LATEST DATA
def build_latest(csv_data):
zero_df = zero_base()
data_df = output_df(csv_data)
db3 = concat_output(zero_df, data_df)
return db3
# ## INITIALIZE FUNCTION
csv_data = get_latest_data()
db3 = build_latest(csv_data)
db3 = db3.query("record_nbr != '100'")
#db3 = db3.query("workload=='workload a'")
# ### Render Graphs
# ## WORKLOAD INSERT
workload = 'workload a'
load_1 = db3.query('run=="run-8"')
graph_i = LinePlot(load_1,workload)
graph_i.graph_records_vs_latency()
# ## WORKLOAD I
workload = 'workload i'
graph_i = LinePlot(db3,workload)
graph_i.graph_records_vs_latency()
# +
def chart_ops_latency(db, workload):
dbgraph = sns.lineplot(x=db.throughput_value,
y=db.runtime_value,
hue=db.database,
style=db.database,
sort=False,
legend='full',
markers=True,
dashes=True)
handles, labels = dbgraph.get_legend_handles_labels()
dbgraph.legend(handles=handles[1:],
labels=labels[1:])
dbgraph.set(xlabel='Throughput(ops/sec)',
ylabel="Latency(ms)")
dbgraph.set_title(workload.upper())
return dbgraph
workload = 'workload i'
db5 = db3[db3.workload == workload]
db5 = db5.sort_values(by=['record_nbr'])
db5 = db5.query("run=='load-1'")
#db5 = db5.query("database=='Redis'")
chart_ops_latency(db5,workload)
db5
# -
# ## WORKLOAD A
workload = 'workload a'
graph_a = LinePlot(db3,workload)
graph_a.graph_records_vs_latency()
# ## WORKLOAD B
workload = 'workload b'
graph_b = LinePlot(db3,workload)
graph_b.graph_records_vs_latency()
# ## WORKLOAD C
workload = 'workload c'
graph_c = LinePlot(db3,workload)
graph_c.graph_records_vs_latency()
# ## WORKLOAD D
workload = 'workload d'
graph_d = LinePlot(db3,workload)
graph_d.graph_records_vs_latency()
# ## WORKLOAD E
workload = 'workload e'
graph_e = LinePlot(db3,workload)
graph_e.graph_records_vs_latency()
workload = 'workload f'
graph_f = LinePlot(db3,workload)
graph_f.graph_records_vs_latency()
| visualization/.ipynb_checkpoints/lineplot-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Objective and Outline
#
# Main goal is to have some fun with the PPP dataset [located here](https://home.treasury.gov/policy-issues/cares-act/assistance-for-small-businesses/sba-paycheck-protection-program-loan-level-data). Inspired by [this tweet](https://twitter.com/dataeditor/status/1280278987797942272) we'll look at city misspellings in the PPP data, starting with Philadelphia. Specifically we'll look at calculating a histogram of the [Levenshtein distance](https://en.wikipedia.org/wiki/Levenshtein_distance) across all misspellings in the dataset.
#
# Some Prefect features we hope to highlight:
# - Prefect's dynamic mapping feature, and how this places nicely with Dask
# - running Prefect Flows on Dask with a `LocalCluster`
#
# ## The Data
#
# - download the data
# - look at it
# - eyeball the Philadelphia mispellings
# +
import pandas as pd
df = pd.read_csv("PPPData.csv")
df.head()
# -
len(df.State.value_counts())
df[df['BusinessName'].str.lower().str.contains('yeez') == True]
philly_mask = (df['City'].str.lower().str.startswith('phil') == True) & (df['State'] == 'PA')
df[philly_mask].City.value_counts()
df2 = pd.read_csv("PPPDataSmallLoan-PA.csv")
philly_mask2 = (df2['City'].str.lower().str.startswith('phil') == True) & (df2['State'] == 'PA')
df2[philly_mask2].City.value_counts()
# ## Build a Prefect Flow for reproducibility + efficiency
#
# Now we'll build a more programmatic set of tasks for downloading / processing our files. Some Prefect features this is intended to highlight:
# - Prefect "mapping"
#
# +
import collections
import datetime
import glob
import io
import os
import pandas as pd
import tempfile
import zipfile
from Levenshtein import distance
from prefect import task, Flow, Parameter, unmapped
config = Parameter("config", default=dict(prefix="phil",
city="Philadelphia"))
@task
def get_PPP_dataframe():
return pd.read_csv("PPPData.csv")
@task
def extract_city_spellings(config, data):
"""
Given config w/ prefix and dataset, returns the unique
set of cities beginning with that prefix.
"""
prefix = config['prefix'].lower()
mask = data['City'].str.lower().str.startswith(prefix) == True
return list(set(data[mask].City.unique()))
@task
def compute_dist(string, truth):
"""
Returns the Levenshtein distance between the two strings
"""
return distance(string, truth)
@task
def aggregate_dists(distances):
"""
Returns a 'histogram' of the counts
"""
return collections.Counter(distances)
# compile our tasks into a Flow object
with Flow("PPP Coiled Demo") as flow:
data = get_PPP_dataframe()
spellings = extract_city_spellings(config, data)
# more interesting dynamic fan out + reduce step
distances = compute_dist.map(spellings, unmapped(config['city']))
results = aggregate_dists(distances)
flow.visualize()
# -
# ## Various ways of running the Flow
# purely local sequential run
flow_state = flow.run()
flow_state.result[results].result # final tally
# +
# local dask run
from prefect.engine.executors import DaskExecutor
flow_state = flow.run(executor=DaskExecutor())
flow_state.result[results].result # final tally
| advanced-tutorials/prefect_PPP_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # BootstrapOutOfBag
# An implementation of the out-of-bag bootstrap to evaluate supervised learning algorithms.
# > `from mlxtend.evaluate import BootstrapOutOfBag`
# ## Overview
# Originally, the bootstrap method aims to determine the statistical properties of an estimator when the underlying distribution was unknown and additional samples are not available. Now, in order to exploit this method for the evaluation of predictive models, such as hypotheses for classification and regression, we may prefer a slightly different approach to bootstrapping using the so-called Out-Of-Bag (OOB) or Leave-One-Out Bootstrap (LOOB) technique. Here, we use out-of-bag samples as test sets for evaluation instead of evaluating the model on the training data. Out-of-bag samples are the unique sets of instances that are not used for model fitting as shown in the figure below [1].
#
# 
#
#
# The figure above illustrates how three random bootstrap samples drawn from an exemplary ten-sample dataset ($X_1,X_2, ..., X_{10}$) and their out-of-bag sample for testing may look like. In practice, <NAME> and <NAME> recommend drawing 50 to 200 bootstrap samples as being sufficient for reliable estimates [2].
# ### References
#
# - [1] https://sebastianraschka.com/blog/2016/model-evaluation-selection-part2.html
# - [2] Efron, Bradley, and <NAME>. An introduction to the bootstrap. CRC press, 1994. Management of Data (ACM SIGMOD '97), pages 265-276, 1997.
# ## Example 1 -- Evaluating the predictive performance of a model
# The `BootstrapOutOfBag` class mimics the behavior of scikit-learn's cross-validation classes, e.g., `KFold`:
# +
from mlxtend.evaluate import BootstrapOutOfBag
import numpy as np
oob = BootstrapOutOfBag(n_splits=3)
for train, test in oob.split(np.array([1, 2, 3, 4, 5])):
print(train, test)
# -
# Consequently, we can use `BootstrapOutOfBag` objects via the `cross_val_score` method:
# +
from sklearn.datasets import load_iris
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import cross_val_score
iris = load_iris()
X = iris.data
y = iris.target
lr = LogisticRegression()
print(cross_val_score(lr, X, y))
# -
print(cross_val_score(lr, X, y, cv=BootstrapOutOfBag(n_splits=3, random_seed=456)))
# In practice, it is recommended to run at least 200 iterations, though:
print('Mean accuracy: %.1f%%' % np.mean(100*cross_val_score(
lr, X, y, cv=BootstrapOutOfBag(n_splits=200, random_seed=456))))
# Using the bootstrap, we can use the percentile method to compute the confidence bounds of the performance estimate. We pick our lower and upper confidence bounds as follows:
#
# - $ACC_{lower}$ = $\alpha_1th$ percentile of the $ACC_{boot}$ distribution
# - $ACC_{lower}$ = $\alpha_2th$ percentile of the $ACC_{boot}$ distribution
#
# where $\alpha_1 = \alpha$ and $\alpha_2 = 1-\alpha$, and the degree of confidence to compute the $100 \times (1-2 \times \alpha)$ confidence interval. For instance, to compute a 95% confidence interval, we pick $\alpha=0.025$ to obtain the 2.5th and 97.5th percentiles of the *b* bootstrap samples distribution as the upper and lower confidence bounds.
import matplotlib.pyplot as plt
# %matplotlib inline
# +
accuracies = cross_val_score(lr, X, y, cv=BootstrapOutOfBag(n_splits=1000, random_seed=456))
mean = np.mean(accuracies)
lower = np.percentile(accuracies, 2.5)
upper = np.percentile(accuracies, 97.5)
fig, ax = plt.subplots(figsize=(8, 4))
ax.vlines(mean, [0], 40, lw=2.5, linestyle='-', label='mean')
ax.vlines(lower, [0], 15, lw=2.5, linestyle='-.', label='CI95 percentile')
ax.vlines(upper, [0], 15, lw=2.5, linestyle='-.')
ax.hist(accuracies, bins=11,
color='#0080ff', edgecolor="none",
alpha=0.3)
plt.legend(loc='upper left')
plt.show()
# -
# ## API
with open('../../api_modules/mlxtend.evaluate/BootstrapOutOfBag.md', 'r') as f:
s = f.read()
print(s)
| mlxtend/docs/sources/user_guide/evaluate/BootstrapOutOfBag.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="s9c-vMyjq7-c"
# ____
# __Universidad Tecnológica Nacional, Buenos Aires__\
# __Ingeniería Industrial__\
# __Autor: <NAME>__\
# __Cátedra de Investigación Operativa - Curso I4051 - Turno Miércoles Noche__
# ____
# + id="bX4wWD1aqtFw"
# importamos librerias
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
sns.set_context("talk",font_scale=0.6)
# + [markdown] id="vSK_ru0RrqpW"
# Queremos encontrar el vector $Q = [ q_1, q_2, q_3, ..., q_m ] $ de cantidades a ordenar de multiples productos que minimiza el Costo Total Esperado.
#
# $$ min Z = f(Q) = CTE(Q) = CTE(q_1, q_2, ..., q_m)$$
#
# Este problema de optimizacion esta sujeto a las siguientes restricciones de volumen S
#
# $$
# q_1 s_1 + q_2 s_2 + ... + q_m s_m \leq S
# $$
#
# donde cada item $q_i$ ocupa un volumen $s_i$ en el almacen. A las restricciones se las puede replantear como
#
# $$
# g(q_1, q_2, ..., q_m,S) = q_1 s_1 + q_2 s_2 + ... + q_m s_m - S
# $$
#
#
#
# Para resolver el problema de optimizacion sujeto a restricciones podemos utilizar los multiplicadores de lagrange
#
# $$
# L(Q, \lambda) = f(Q) + \lambda g(Q)
# $$
#
# donde el valor de $\lambda$ penaliza el volumen que ocupa cada item quedando la nueva funcion objetivo como
#
# $$
# L = f(q_1,...,q_m) + \lambda [(s_1 q_1 + ...+ s_m q_m) -S]
# $$
#
# entonces si queremos minimizar la funcion L podemos derivar en funcion de $Q$ e igualar a cero
#
# $$
# \frac{\partial L}{\partial Q} = 0
# $$
#
# para arribar a la expresion que determina el valor de la cantidad a pedir del item $i$ penalizado por el volumen que ocupa
#
# $$
# \frac{\partial K}{\partial q_i} = \frac{1}{2}C_{1i} T - \frac{K_i D_i}{q^{2}_i} + \lambda S_i \rightarrow q_i = \sqrt{\frac{2 K_i D_i}{T C_{1i} + 2 \lambda S_i }}
# $$
#
#
# + id="UN3p9C9Ur5Py"
# definimos la funcion para calcular el q_i de cada item
def q_opt_restrict(k, d, C1, lambd, Si):
'calcula el Qi para cada item considerando el volumen que ocupa'
qi = np.sqrt((2*k*d)/(C1 + 2*lambd*Si))
return qi
# definimos la funcion que calcula la superficie total
def sup_total(q_all_items , s_all_items):
'calculamos la superficie total dado un vector de Qi donde cada posicion del vector es la cantidad a ordenar por item'
# hacemos la combinacion lineal entre el vector q y el vector sup_all_items
sup_tot = np.dot(q_all_items, s_all_items)
return sup_tot
# + [markdown] id="DnCxBPJlyMqu"
# Cargamos los datos del problema
# + id="jjNw04exxmNf"
k1 = 10
D1 = 200
c1_1 = 0.3
s1 = 1
k2 = 5
D2 = 400
c1_2 = 0.1
s2 = 1
k3 = 15
D3 = 400
c1_3 = 0.2
s3 = 1
k4 = 6
D4 = 300
c1_4 = 0.1
s4 = 2
k5 = 9
D5 = 700
c1_5 = 0.15
s5 = 1.5
s_all = np.array([s1,s2,s3,s4,s5])
n_skus = 5
# + id="LYganlGB35PC"
# si lambda = 0 entonces estamos en un problema sin restricciones
lambd_test = 0
q1_0 = q_opt_restrict(k1, D1, c1_1, lambd_test, s1)
q2_0 = q_opt_restrict(k2, D2, c1_2, lambd_test, s2)
q3_0 = q_opt_restrict(k3, D3, c1_3, lambd_test, s3)
q4_0 = q_opt_restrict(k4, D4, c1_4, lambd_test, s4)
q5_0 = q_opt_restrict(k5, D5, c1_5, lambd_test, s5)
q_all_0 = np.array([q1_0, q2_0, q3_0, q4_0, q5_0])
# + id="suhgnH7X4OyW" colab={"base_uri": "https://localhost:8080/"} outputId="211e1e41-17c6-4655-e131-fc8ce63e8827"
# imprimimos en pantalla la superficie total sin considerar las restricciones de espacio
sup_total(q_all_0, s_all)
# + id="IZILmvyJ5M32"
# hacemos una grilla de 100 valores de lambda entre 0 y 1 y para cada valor de la grilla
# calculamos la superficie total
# definimos las variables y vectores iniciales antes de las iteraciones
iteraciones = 100
q1_i = np.zeros(iteraciones)
q2_i = np.zeros(iteraciones)
q3_i = np.zeros(iteraciones)
q4_i = np.zeros(iteraciones)
q5_i = np.zeros(iteraciones)
q_all_i = np.zeros(iteraciones)
q_vec_i = np.zeros((iteraciones,n_skus))
s_total_i = np.zeros(iteraciones)
# con el for loop computamos para distintos valores de lambda
for lambd_i in range(0, iteraciones):
q1_i[lambd_i] = q_opt_restrict(k1, D1, c1_1, lambd_i/100, s1)
q2_i[lambd_i] = q_opt_restrict(k2, D2, c1_2, lambd_i/100, s2)
q3_i[lambd_i] = q_opt_restrict(k3, D3, c1_3, lambd_i/100, s3)
q4_i[lambd_i] = q_opt_restrict(k4, D4, c1_4, lambd_i/100, s4)
q5_i[lambd_i] = q_opt_restrict(k5, D5, c1_5, lambd_i/100, s5)
# guardamos el vector de Qi para cada iteracion
q_vec_i[lambd_i, :] = np.array([q1_i[lambd_i], q2_i[lambd_i], q3_i[lambd_i], q4_i[lambd_i], q5_i[lambd_i]])
# calculamos la superficie total para cada iteracion de distintos valores de lambda
s_total_i[lambd_i] = sup_total(q_vec_i[lambd_i, :], s_all)
#q_all_i[lambd_i] = sup_total()
# + id="tB4SPFqj9PWz" colab={"base_uri": "https://localhost:8080/", "height": 299} outputId="66e5cd9e-22a9-4252-8de3-166aedaeb9de"
# imprimimos en pantalla la superficie resultante de cada iteracion en funcion de los valores de lambda
plt.plot(np.arange(0.0, 1, 0.01),s_total_i, label = 'superficie resultante')
plt.title('Superficie total a ocupar en funcion de la penalizacion')
plt.xlabel('Grilla de lambdas')
plt.ylabel('Superficie total a ocupar')
plt.plot(np.arange(0.0, 1, 0.01),np.full((100),600), label = 'superficie limite')
plt.vlines(0.23, ymin = 10, ymax = 1300, linestyles= '--',colors = 'r', alpha = 0.3)
plt.legend()
plt.show()
# + id="scYSpOjL7Icn" colab={"base_uri": "https://localhost:8080/", "height": 299} outputId="c4aa6502-4e37-47eb-e61c-edea081362b0"
plt.plot(np.arange(0.0, 1, 0.01),q_vec_i[:,0], label = 'sku 1')
plt.plot(np.arange(0.0, 1, 0.01),q_vec_i[:,1], label = 'sku 2')
plt.plot(np.arange(0.0, 1, 0.01),q_vec_i[:,2], label = 'sku 3')
plt.plot(np.arange(0.0, 1, 0.01),q_vec_i[:,3], label = 'sku 4')
plt.plot(np.arange(0.0, 1, 0.01),q_vec_i[:,4], label = 'sku 5')
plt.vlines(0.23, ymin = 10, ymax = 280, linestyles= '--',colors = 'r')
plt.legend()
plt.title('Cantidad a pedir por item en funcion de la penalizacion')
plt.xlabel('Penalizacion lambda')
plt.ylabel('cantidad a pedir Q_i por item i')
#plt.xticks(np.arange(100), np.arange(100)/10, rotation=90)
plt.locator_params(nbins=4)
plt.show()
# + id="F4DH--ai7XaM" colab={"base_uri": "https://localhost:8080/", "height": 299} outputId="78e83ee9-5730-4ed3-d78b-769e24444769"
sns.heatmap(q_vec_i)
plt.xlabel('items')
plt.ylabel('Iteraciones de penalizacion')
plt.title('Cantidad a pedir por item')
plt.show()
# + id="xzaUPz4y8DYm"
lambda_final = 0.37
# + id="S8tXZgxrAmtT"
| 07_programacion_matematica/casos_codigo/ioperativ_clase25_multiproducto_restriccion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <br><br><br><br><br>
#
# # Jailbreaking Python
#
# <br><br><br><br><br>
# ### Tale of two VMs
#
# **Java** and **Python** are both high-level programming environments based on virtual machines. Your programs compile to virtual machine instructions that are emulated at runtime. This limits the consequences of mistakes, making it easier to develop interactively.
#
# However, **Java** and **Python**'s relationships to the outside world are very different.
#
# * **Java** lives almost entirely inside its virtual machine. There's a pure Java library for just about everything, while its C API is very hard to use. Consequently, there are many Java Runtime Environment implementations, all satisfying the same specification.
# * **Python** has extension modules for a lot of functionality. Its C API is relatively easy to use and there are dozens of wrappers. However, this makes it difficult for alterantive implementations, like PyPy, to compete with the standard CPython because most of those extension modules only work with CPython.
#
# **Python**, through its extension modules, is a leaky abstraction.
#
# <br>
# +
# Let's break Python!
hello = b"Hello, world!" # Python strings are immutable so they can be referenced, not copied.
try: # A lot of code depends on strings not changing after they're made.
hello[4:8] = b"????"
except TypeError as err:
print("Not allowed to change it: " + str(err))
import numpy # Wrap it with a Numpy array (not a copy).
a = numpy.frombuffer(hello, dtype=numpy.uint8)
print("a =", a)
print("a.view('S1') =", a.view('S1'))
a.flags.writeable = True # Set the writable flag to True,
a[4:8] = [69, 86, 73, 76] # and we can write to it.
print("hello =", repr(hello)) # And it is evil.
# +
# See how evil this is:
# Interpreted as it looks: we change one b"Hello, world!" but not the other.
hello = b"Hello, world!"
a = numpy.frombuffer(hello, dtype=numpy.uint8)
a.flags.writeable = True
a[4:8] = [69, 86, 73, 76]
print("case 1:", hello == b"Hello, world!")
# Interpreted as a single .pyc; literal b"Hello, world!" strings are replaced by a common object.
exec("""
hello = b"Hello, world!"
a = numpy.frombuffer(hello, dtype=numpy.uint8)
a.flags.writeable = True
a[4:8] = [69, 86, 73, 76]
print("case 2:", hello == b"Hello, world!")
""")
# -
# <br><br><br><br><br>
#
# ### Python is an environment in which you can poke individual bytes
#
# <br><br><br><br><br>
# +
x = 12345
import ctypes
import sys
ptr = ctypes.cast(id(x), ctypes.POINTER(ctypes.c_uint8))
a = numpy.ctypeslib.as_array(ptr, (sys.getsizeof(x),))
print("a =", a)
# We're looking at a Python object header, a pointer to the int type (also a Python object), and
# then the number itself: 12345 in little endian bytes is [57, 48, 0, 0]. Do you see it?
# +
# Now for a string.
y = "Hey there."
ptr = ctypes.cast(id(y), ctypes.POINTER(ctypes.c_uint8))
a = numpy.ctypeslib.as_array(ptr, (sys.getsizeof(y),))
print("a =", a)
print("\na[-11:] =", a[-11:])
print("\na[-11:].tostring() =", repr(a[-11:].tostring()))
# +
# The snake eats its own tail: reading PyObject refcount without changing it.
class PyObject(ctypes.Structure): pass
PyObject._fields_ = [("ob_refcnt", ctypes.c_size_t),
("ob_type", ctypes.POINTER(PyObject))]
hello = b"Hello, world!"
ptr = PyObject.from_address(id(hello))
print("ptr =", ptr)
print("\nBefore:")
print("ptr.ob_refcnt =", ptr.ob_refcnt, "sys.getrefcount(hello) =", sys.getrefcount(hello))
biglist = [hello] * 1000
print("\nAfter:")
print("ptr.ob_refcnt =", ptr.ob_refcnt, "sys.getrefcount(hello) =", sys.getrefcount(hello))
# -
# <br><br><br><br><br>
#
# ### Doing something _useful_ with this power
#
# <br><br><br><br><br>
# +
# Suppose you're on a supercomputer with Non-Uniform Memory Access (NUMA) and you want Numpy arrays.
import ctypes.util
libnuma = ctypes.cdll.LoadLibrary(ctypes.util.find_library("numa"))
libnuma.numa_alloc_local.argtypes = (ctypes.c_size_t,) # manually set the expected types (no .h file)
libnuma.numa_alloc_local.restype = ctypes.POINTER(ctypes.c_double)
ptr = libnuma.numa_alloc_local(4*1024) # allocate it!
a = numpy.ctypeslib.as_array(ptr, (1024,)) # wrap it as an array!
a[:] = numpy.linspace(-511.5, 511.5, 1024) # assign to it; use it!
print("a =", a)
# +
# Of course, if you're allocating CPU-local memory in Numpy arrays, you'll want to ensure that
# your thread doesn't switch to another CPU.
import psutil
psutil.Process().cpu_affinity([0])
# -
# <br><br><br><br>
#
# ### Interacting with C and C++
#
# With <tt>ctypes</tt>, we can call any function in any library that supports Foreign Function Interface (FFI). This includes almost every compiled language except C++.
#
# Both for C++ and for safety from low-level hacks, we should use a library.
#
# <br><br><br><br>
# <center><img src="img/history-of-bindings.png" width="85%"></center>
#
# **We've already seen that Cython is hard to use for performance.**
# +
# It's easy to get started with pybind11, and it uses a modern subset of C++.
import os
with open("tmp.cpp", "w") as file:
file.write("""
#include<pybind11/pybind11.h>
double add(double a, double b) {
return a + b;
}
PYBIND11_MODULE(tmp, m) {
m.def("add", &add, "a compiled function");
}""")
if os.system("""c++ -Wall -shared -std=c++11 -fPIC `python -m pybind11 --includes` \
tmp.cpp -o tmp`python3-config --extension-suffix`""") == 0:
import tmp
print(tmp.add(3.14, 99.9))
# -
# #### You can do the reverse: compile Python, pass as function pointer to C
#
# <center><img src="img/writing-c-functions-in-python.png" width="90%"></center>
# #### Numpy ufuncs are a C function protocol, overridden by compiled code
#
# <center><img src="img/creating-your-own-ufunc.png" width="90%"></center>
# <br><br><br><br>
#
# # Day 3 Homework
#
# Did you bring a project that you'd like to accelerate, rework in Numpy/awkward, or bind to C++?
#
# <br><br><br><br>
| 09-jailbreaking-python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <br>
# <hr>
# **“You don't have to be great to start, but you have to start to be great.”** ~ <NAME>
# <hr>
# <br>
#
# Hey friends,<br>
# this is a gentle introduction into Image Classification with the Python library fast.ai.
#
# What do you need to get started?
# - basic coding skills in Python
# - some familiarity with fast.ai (at least the first video of the course)
#
# [click if you don't know how to code at all](https://www.codecademy.com/learn/learn-python-3)<br>
# [click if you know some coding, but not Python](https://developers.google.com/edu/python/)<br>
# [click if you don't know the fast.ai course](https://course.fast.ai/videos/?lesson=1)
#
# Feel free to fork and tweak constants to improve accuracy. Just by doing so, I was able to get a score of 99.32% easily.
#
# Check out the [Q&A section](#Questions-and-Answers) of this notebook. If you have any questions or certain explanations weren't completely clear, let me know in the comments. I'm happy to help everyone :)
#
# 1. [Preparation](#Preparation)
# 2. [Training](#Training)
# 3. [Evaluation](#Evaluation)
# 4. [Prediction](#Prediction)
# 5. [Questions and Answers](#Questions-and-Answers)
#
# [click for fast.ai documentation](https://docs.fast.ai)
#
#
# ## Preparation
# first of all make sure you enabled GPU so the CNN trains faster
# #### Setup environment and import necessary modules
# + _kg_hide-input=true
# the following three lines are suggested by the fast.ai course
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
# + _kg_hide-input=true
# hide warnings
import warnings
warnings.simplefilter('ignore')
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# the fast.ai library
from fastai import *
from fastai.vision import *
# to inspect the directory
import os
from pathlib import Path
# for data manipulation (in this Kernel mainly used to read .csv files)
import pandas as pd
# for numerical analysis
import numpy as np
# to display imags
from PIL import Image
# -
# #### Inspect and understand input data
# The first step in most competitions is to check out the input data. Let's do this:
INPUT = Path("../input/digit-recognizer")
os.listdir(INPUT)
# We found 3 interesting files:
# - sample_submission.csv
# - train.csv
# - test.csv
#
# 'sample_submission.csv' will show us, how we have to structure our data at the end before we submit it to the competition. We will only need this file at the end.
#
# 'train.csv' is a file that contains all necessary information for training the CNN
#
# 'test.csv' is the file we later use to test how good our CNN is
#
# Let's look at 'train.csv' and 'test.csv' to see how the data looks
train_df = pd.read_csv(INPUT/"train.csv")
train_df.head(3)
test_df = pd.read_csv(INPUT/"test.csv")
test_df.head(3)
# The data looks very interesting. Let's summarize what we got here:
#
# What we know about 'train.csv':
# - Each row is one image
# - The first row of each image is the label. It tells us which digit is shown.
# - The other 784 rows are the pixel for each digit and should be read like this
#
# `000 001 002 003 ... 026 027
# 028 029 030 031 ... 054 055
# 056 057 058 059 ... 082 083
# | | | | ... | |
# 728 729 730 731 ... 754 755
# 756 757 758 759 ... 782 783`
#
# What we know about 'test.csv':
# - The structure is the same as in train.csv, but there are no labels because it's our task to predict the labels
#
# To read more about the data, read the ['Data' tab of the competition](https://www.kaggle.com/c/digit-recognizer/data)
#
# #### Getting the data into the right format
# Looking at the [fast.ai documentation](https://docs.fast.ai/vision.data.html#ImageDataBunch) we can quickly see, that only accepts image files for Computer Vision. In this competition we were not offered images, but csv files where each cell is one pixel. If we want to use fast.ai we have to create images from the data we have.
#
# Fast.ai accepts image data in different formats. We will use the from_folder function of the ImageDataBunch class to load in the data. To do this we need all images in the following structure:
#
# `path\
# train\
# 0\
# ___.jpg
# ___.jpg
# ___.jpg
# 1\
# ___.jpg
# ___.jpg
# 2\
# ...
# 3\
# ...
# ...
# test\
# ___.jpg
# ___.jpg
# ...
# `
#
# Let's first create the folder structure!
#
# (nice to know: the input folder of Kaggle Competitions is always read-only, so if we want to add data or create folders, we have to do so outside of the input folder)
TRAIN = Path("../train")
TEST = Path("../test")
# Create training directory
for index in range(10):
try:
os.makedirs(TRAIN/str(index))
except:
pass
# Test whether creating the training directory was successful
sorted(os.listdir(TRAIN))
#Create test directory
try:
os.makedirs(TEST)
except:
pass
# Okay, all folders are created! The next step is to create the images inside of the folders from 'train.csv' and 'test.csv'. We will use the Image module from PIL to do this.
#
#
# we have to reshape each numpy array to have the desired dimensions of the image (28x28)
#
# `000 001 002 003 ... 026 027
# 028 029 030 031 ... 054 055
# 056 057 058 059 ... 082 083
# | | | | ... | |
# 728 729 730 731 ... 754 755
# 756 757 758 759 ... 782 783`
#
# then we use the fromarray function to create a .jpg image from the numpy array and save it into the desired folder
# save training images
for index, row in train_df.iterrows():
label,digit = row[0], row[1:]
filepath = TRAIN/str(label)
filename = f"{index}.jpg"
digit = digit.values
digit = digit.reshape(28,28)
digit = digit.astype(np.uint8)
img = Image.fromarray(digit)
img.save(filepath/filename)
# save testing images
for index, digit in test_df.iterrows():
filepath = TEST
filename = f"{index}.jpg"
digit = digit.values
digit = digit.reshape(28,28)
digit = digit.astype(np.uint8)
img = Image.fromarray(digit)
img.save(filepath/filename)
# Display some images
# + _kg_hide-input=true
def displayRandomImagesFromEveryFolder(directory=TRAIN, samplesPerDigit=5):
fig = plt.figure(figsize=(5,10))
for rowIndex in range(1, 10):
subdirectory = str(rowIndex)
path = directory/subdirectory
images = os.listdir(path)
for sampleIndex in range(1,samplesPerDigit+1):
randomNumber = random.randint(0, len(images)-1)
image = Image.open(path/images[randomNumber])
ax = fig.add_subplot(10, 5, samplesPerDigit*rowIndex + sampleIndex)
ax.axis("off")
plt.imshow(image, cmap='gray')
plt.show()
displayRandomImagesFromEveryFolder()
# -
# ### Load data into DataBunch
# Now that we have the right folder structure and images inside of the folders we can continue.
# Before training a model in fast.ai, we have to load the data into a [DataBunch](https://docs.fast.ai/basic_data.html#DataBunch), in this case, we use a ImageDataBunch, a special version of the DataBunch.
# Fast.ai offers different functions to create a DataBunch. We will use the from_folder method of the ImageDataBunch class to create the dataset.
# There are different hyperparameters we can tweak to make the model perform better:
# - [valid_pct](#What-are-Train,-Test-and-Validation-datasets?)
# - [bs (batch size)](#What-is-the-batch-size?)
# - [size](#What-image-size-should-I-choose?)
# - [num_workers](#What-is-multiprocessing?)
# - [ds_tfms](#What-are-transforms-and-which-transforms-should-I-use?)
# + _kg_hide-input=true
flip_tfm = RandTransform(tfm=TfmPixel (flip_lr), kwargs={}, p=1, resolved={}, do_run=True, is_random=True, use_on_y=True)
random_number = str(3)
random_filename = os.listdir(TRAIN/random_number)[0]
img = open_image(TRAIN/random_number/random_filename)
display(img)
display(img.apply_tfms(flip_tfm))
# -
# transforms
tfms = get_transforms(do_flip=False, max_zoom=1.2)
data = ImageDataBunch.from_folder(
path = TRAIN,
test = TEST,
valid_pct = 0.2,
bs = 32,
size = 28,
ds_tfms = tfms,
#num_workers = 0
)
# Let's perform normalization to make the CNN converge faster. fast.ai already defined the variable mnist_stats, that we can use to normalize our data. Alternatively, we can call normalize() without any paramters. In this case fast.ai simply calculates the exact stats needed for the dataset at hand.
mnist_stats
data.normalize(mnist_stats)
# all the classes in data
print(data.classes)
# ## Training
# The next step is to select and create a CNN. In fast.ai creating a CNN is really easy. You just have to select one of the models from the [Computer Vision models zoo](https://docs.fast.ai/vision.models.html#Computer-Vision-models-zoo). Parameters to tweak: dropout propability: ps (default: 0.5)
learn = cnn_learner(data, models.resnet18, metrics=accuracy, model_dir="/tmp/models", callback_fns=ShowGraph)
# Now it's time to train the neural network using the fit_one_cycle() function. <br>Parameters to modify: the number of epochs(cyc_len) to train, the learning rate(max_lr) and the momentum (moms)
learn.fit_one_cycle(cyc_len=5)
# ## Evaluation
# Create a ClassificationInterpretation object to evaluate your results.
interp = ClassificationInterpretation.from_learner(learn)
# Plot the 9 images with the highest loss. These are the images the CNN was most sure about, but still got wrong.
interp.plot_top_losses(9, figsize=(7, 7))
# A good way to summarize the performance of a classification algorithm is to create a confusion matrix. Confusion Matricies are used to understand which classes are most easily confused. As labeled on the axis, the x-axis shows the predicted classes and the y-axis the actual classes. So if (4/7)=10 it means that it happened 10 times that the CNN predicted a 7 but in reality if was a 4.
interp.plot_confusion_matrix()
# ## Prediction
#
# Get the predictions on the test set.<br>
# learn.get_preds() returns a propability distribution over all possible classes for every given image.
class_score, y = learn.get_preds(DatasetType.Test)
# That means that for every image in the test set it predicts how likely each class is. In this case the highest value is obviously 1
probabilities = class_score[0].tolist()
[f"{index}: {probabilities[index]}" for index in range(len(probabilities))]
# But we want the CNN to predict only one class. The class with the highest probability.
# Argmax returns the index of the highest value.
class_score = np.argmax(class_score, axis=1)
# This is exactly what we want.
class_score[0].item()
# The last step is creating the submission file.<br>"sample_submission.csv" is showing us the desired format
sample_submission = pd.read_csv(INPUT/"sample_submission.csv")
display(sample_submission.head(2))
display(sample_submission.tail(2))
# Columns the submission file has to have:
# - ImageId: index in the test set, starting from 1, going up to 28000
# - Label: a prediction which digit the image shows
# remove file extension from filename
ImageId = [os.path.splitext(path)[0] for path in os.listdir(TEST)]
# typecast to int so that file can be sorted by ImageId
ImageId = [int(path) for path in ImageId]
# # +1 because index starts at 1 in the submission file
ImageId = [ID+1 for ID in ImageId]
submission = pd.DataFrame({
"ImageId": ImageId,
"Label": class_score
})
# submission.sort_values(by=["ImageId"], inplace = True)
submission.to_csv("submission.csv", index=False)
display(submission.head(3))
display(submission.tail(3))
# ## Questions and Answers
# - [What are Train, Test and Validation datasets?](#What-are-Train,-Test-and-Validation-datasets?)
# - [What is the batch size?](#What-is-the-batch-size?)
# - [What image size should I choose?](#What-image-size-should-I-choose?)
# - [What is multiprocessing?](#What-is-multiprocessing?)
# - [What are transforms and which transforms should I use?](#What-are-transforms-and-which-transforms-should-I-use?)
# - [How to improve your score?](#How-to-improve-your-score?)
# - [What is the best CNN architecture?](#What-is-the-best-CNN-architecture?)
# - [How to submit?](#How-to-submit?)
#
# ### What are Train, Test and Validation datasets?
# Whenever we train a CNN we need to split the data into 3 parts:
# - training set: used to modify weights of neural network
# - validation set: prevent overfitting
# - test set: test accuracy of fully-trained model
#
# In this kernel we only have a training and a test set. That's why we split the test set to get a validation set. We do this with the 'valid_pct' parameter. This is one of the parameters you could tune to increase the accuracy. To learn more about this read [this stackoverflow post](https://stackoverflow.com/a/13623707)
# ### What is the batch size?
# The batch size refers to the number of images in one batch. Everytime an entire batch of images is passed through the neural network the weights of the neural network are updated
# <br><br>Why would you increase the batch size:
# - to improve accuracy of the model
#
# <br>Why would you decrease the batch size:
# - to train the network faster
# - to reduce the memory used
# ### What image size should I choose?
#
# (Most) CNNs need images of the same size. By setting the size parameter we tell fast.ai to make all images that size.
#
# Bigger images, result in more calculations and thus slower speed, but the accuracy improves. Smaller images on the other hand reduce accuracy but improve speed.
# Don't make the trainig images bigger than the original images, as this would only be a waste of time.
#
# Our data is already of shape 28x28.
#
# We could make the images smaller than 28x28. This would decrease the training time, but also decrease the accuracy. Because our CNN trains in a reasonable amount of time there is no reason to decrease the image size
#
# Never make the training image bigger than the original image.
# ### What is multiprocessing?
# In Computer Science there is this thing called multiprocessing. This means that we have two or more things happending at the same time. A computer typically has multiple CPUs and we make use of exactly that. Every CPU can run one process at a time. Ususally when we do multiprocessing every CPU gets its own task.
# Exactly that is the default for ImageDataBunch: number workers = number of CPUs. This works fine for Linux, but makes a lot of problems in Windows. If you're on Windows, set num_workers to 0, if you're on Linux, don't set anything, then it defaults to the number of CPUs
#
# If you use a cloud solution and are not sure which operating system is used, execute the following code
# > import platform; platform.system()
# ### What are transforms and which transforms should I use?
# To make models generalize better we can use so called transforms. They randomly change the image slightly. For example a bit of zoom or rotation. Fortunately, we don't really have to deal with transforms a lot in fast.ai. The package offers a convenient function called get_transforms() that returns pretty good values for transformations.
# In the case of digit recognition we want to tranform the data as much as possible so that it generalizes better, but only so much that the image would still be recognized by a human being.
# One parameter we definitely have to change for that reason is do_flip. If this is activated, random flips (with a probability of 0.5) would be applied to the images. We don't want that. The next two images are an image of the number three and another image of a number three that is flipped horizontally. This would confuse our CNN.
# ### How to improve your score?
# add special features
# - pooling layers
# - data augmentation
# - dropout
# - batch normalization
# - decaying learning rate
# - advanced optimization
#
# [source](https://www.kaggle.com/c/digit-recognizer/discussion/61480)
# ### What is the best CNN architecture?
# According to [this post](https://www.kaggle.com/cdeotte/how-to-choose-cnn-architecture-mnist) by [<NAME>](https://www.kaggle.com/cdeotte) the best architecture is:
# > 784 - [32C3-32C3-32C5S2] - [64C3-64C3-64C5S2] - 128 - 10
# > with 40% dropout, batch normalization, and data augmentation added
#
# 784 input nodes
#
# [32C3-32C3-32C5S2]
# - 2x (convolutional layer with 32 feature maps, 3x3 filter and stride 1)
# - convolutional layer with 32 feature maps, 5x5 filter and stride 2
#
# [64C3-64C3-64C5S2]
# - 2x (convolutional layer with 64 feature maps, 3x3 filter and stride 1)
# - convolutional layer with 64 feature maps, 5x5 filter and stride 2
#
# 128 fully connected dense layers
#
# 10 output nodes
#
#
# ### What is the best possible score?
# The best score inside of Kernels is [99.75%](https://www.kaggle.com/cdeotte/25-million-images-0-99757-mnist).
#
# The best score anywhere is [99.79%](http://yann.lecun.com/exdb/publis/pdf/wan-icml-13.pdf).
# ### How to submit?
# Now we're throught the entire process of how to create a CNN with fast.ai to recognize digits. To submit a file to the the competition, you have two different options.
# #### Code in Kaggle Kernel
# 1. go to the kernel
# 2. commit the kernel
# 3. go back to all of your kernels
# 4. select the kernel again
# 5. scroll down to Output
# 6. click on "Submit to Competition"
# #### Code locally on PC
# 1. go to [the competition](https://www.kaggle.com/c/digit-recognizer)
# 2. click on ["Submit Predictions"](https://www.kaggle.com/c/digit-recognizer/submit)
# 3. upload your submission file
# 4. add a description
# 5. click on submit
# TODO:
# - image augmentation
# https://www.kaggle.com/anisayari/generate-more-training-data-fastai
# - created images to zip file
# https://www.kaggle.com/anisayari/generate-more-training-data-fastai
# - better comments
# - add images to "How to Submit"
# - explain normalization better
| 2 digit recognizer/beginners-guide-to-mnist-with-fast-ai.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
# # Introduction
# Models are trained in google colab with batch_size of 200-700. For online prediction, we only need batch_size of 1.
# Here we recombined models in
# ### Get source folder and append to sys directory
from __future__ import print_function
import os
import sys
PROJ_ROOT = os.path.join(os.pardir)
print(os.path.abspath(PROJ_ROOT))
src_dir = os.path.join(PROJ_ROOT, "src")
sys.path.append(src_dir)
# Data path example
#pump_data_path = os.path.join(PROJ_ROOT,
# "data",
# "raw",
# "pumps_train_values.csv")
# ### Imports
# Import libraries and write settings here.
# +
# Data manipulation
import pandas as pd
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
import sklearn
import random
# Options for pandas
pd.options.display.max_columns = 50
pd.options.display.max_rows = 30
# Display all cell outputs
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = 'all'
from IPython import get_ipython
ipython = get_ipython()
# autoreload extension
if 'autoreload' not in ipython.extension_manager.loaded:
# %load_ext autoreload
# %autoreload 1
# Use %aimport module to reload each module
# %aimport data.read_data
from data.read_data import read_data_by_type
from data.create_load_transform_processed_data import load_reshaped_array, create_tensorflow_dataset
# Visualizations
import matplotlib.pyplot as plt
# -
data = read_data_by_type(PROJ_ROOT=PROJ_ROOT,data_type="interim", output_type="dataframe")
data
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler(feature_range=(-1,1))
scaler_PM25 = MinMaxScaler(feature_range=(-1,1))
scaler_PM25.fit(data['PM25'].values.reshape(-1, 1))
scaler_AQI = MinMaxScaler(feature_range=(-1,1))
scaler_AQI.fit(data['AQI_h'].values.reshape(-1, 1))
# # Analysis/Modeling
# +
# What I need to do: Read all models and their weights
# save them as a new model with included weights
# Check the performance of saved models with loaded models
# +
from models.create_and_load_model import create_model
# Read models and combine them with weights
_data_to_model_path = os.path.join(PROJ_ROOT,
"data",
"model_input",
"hanoi")
_hanoi_model_path = os.path.join(PROJ_ROOT,
"models",
"hcm")
_converted_model_path = os.path.join(PROJ_ROOT,
"models",
"combined")
def get_model_name(timesteps, target_hour):
return 'model_{}_{}.h5'.format(timesteps, target_hour)
def get_saved_model_name(timesteps, target_hour):
# Saved model but for gcp
return 'model_{}_{}'.format(timesteps, target_hour)
def get_model_weigts_name(timesteps, target_hour):
return 'weights_{}_{}.ckpt'.format(timesteps, target_hour)
def root_mean_squared_error(y_true, y_pred):
return K.sqrt(K.mean(K.square(y_pred - y_true), axis=0))
def mean_absolute_percentage_error(y_true, y_pred,
sample_weight=None,
multioutput='uniform_average'):
epsilon = np.finfo(np.float64).eps
mape = np.abs(y_pred - y_true) / np.maximum(np.abs(y_true), epsilon)
output_errors = np.average(mape,
weights=sample_weight, axis=0)
if isinstance(multioutput, str):
if multioutput == 'raw_values':
return output_errors
elif multioutput == 'uniform_average':
# pass None as weights to np.average: uniform mean
multioutput = None
return np.average(output_errors, weights=multioutput)
for hour in [1]:
rmse = []
r2 = []
mae = []
mape = []
for timesteps in [5]:
print(timesteps)
batch_size = 700
test, y_test = load_reshaped_array(timesteps, target_hour=hour, folder_path=_data_to_model_path, data_type="test")
test_data_tf, test_steps_per_epochs = create_tensorflow_dataset(test, y_test, batch_size)
colab_model = keras.models.load_model("{}/{}".format(_hanoi_model_path,get_model_name(timesteps=timesteps, target_hour=hour)),
custom_objects={'LeakyReLU': layers.LeakyReLU(alpha=0.01),
'root_mean_squared_error': root_mean_squared_error})
old_input_layer = colab_model.layers.pop(0)
timesteps, features = old_input_layer.output_shape[0][1:]
# Model with batch_size=1
converted_model = create_model(batch_size=1, timesteps=timesteps, features=features)
converted_model.load_weights('{}/{}'.format(_hanoi_model_path,
get_model_weigts_name(timesteps=timesteps, target_hour=hour))).expect_partial()
converted_model.save("{}/{}".format(_converted_model_path, get_saved_model_name(timesteps=timesteps, target_hour=hour)))
# np.testing.assert_allclose(
# model.predict(test_data_tf, steps=test_steps_per_epochs), reconstructed_model.predict(test_data_tf, steps=test_steps_per_epochs),
# )
# print(model.predict(test_data_tf, steps=test_steps_per_epochs))
# scaler_AQI = MinMaxScaler(feature_range=(-1,1))
# scaler_AQI.fit(thudohanoi_df['AQI_h'].values.reshape(-1, 1))
# y_test = scaler_AQI.inverse_transform(y_test.reshape(-1, 1))
# predict = scaler_AQI.inverse_transform(predict.reshape(-1, 1))
# print("=============================================\n")
# # print("Predict")
# # predict_vs_truth = pd.DataFrame({'predict': predict[rand:rand+20],
# # 'truth': y_test[rand:rand+20]})
# # print(predict_vs_truth)
# print("R2: {}".format(r2_score(predict, y_test)))
# print("Root mean squared error: {}".format(mean_squared_error(predict, y_test, squared=False)))
# print("Mean absolute percentage error: {}".format(mean_absolute_percentage_error(predict, y_test)))
# print("Mean absolute error: {}".format(mean_absolute_error(predict, y_test)))
# rmse.append(mean_squared_error(predict, y_test, squared=False))
# r2.append(r2_score(predict, y_test))
# mape.append(mean_absolute_percentage_error(predict, y_test))
# mae.append(mean_absolute_error(predict, y_test))
# -
# Try predict with converted models
converted_models.predict(test_data_tf, steps=test_steps_per_epochs)
# # Results
# Show graphs and stats here
# # Conclusions and Next Steps
# Summarize findings here
| notebooks/09_nam_convert_models_for_gcp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#데이터 로드
sales = pd.read_csv("sales_rate.csv", encoding='euc-kr' )
sales_ori = pd.read_csv("sales_2018.csv", encoding='euc-kr' )
df_merged = pd.read_csv("df_sejin.csv", encoding='euc-kr')
# #### 주중매출액/ 총매출액 컬럼 만들기(dayrate)
sales.tail(1)
sales.columns
sales['DW_total']= sales['day_sales']+sales['weekend_sale']
sales['day_rate']=sales['day_sales']/sales['DW_total']
# 주중 매출/ 총매출 컬럼있는 df 만들기
df_dayrate=sales[['기준_년_코드', '기준_분기_코드', '상권_코드', '서비스_업종_코드', '당월_매출_금액', 'day_rate']]
df_dayrate.columns=['year', 'quarter', 'district','code','sales','day_rate']
# #### 여성매출/총매출 컬럼있는 df 만들기
sales_ori.columns
# 'prop_2030s',
# 'prop_06_11', 'prop_11_14', 'prop_14_17', 'prop_17_21', 'prop_21_24',
sales_ori['time_total']= sales_ori['시간대_00~06_매출_금액']+sales_ori['시간대_06~11_매출_금액']+sales_ori['시간대_11~14_매출_금액']\
+sales_ori['시간대_14~17_매출_금액']+ sales_ori['시간대_17~21_매출_금액']+sales_ori['시간대_21~24_매출_금액']
# +
sales_ori['prop_06_11']= sales_ori['시간대_00~06_매출_금액']/sales_ori['time_total']
sales_ori['prop_11_14']= sales_ori['시간대_11~14_매출_금액']/sales_ori['time_total']
sales_ori['prop_14_17']= sales_ori['시간대_14~17_매출_금액']/sales_ori['time_total']
sales_ori['prop_17_21']= sales_ori['시간대_17~21_매출_금액']/sales_ori['time_total']
sales_ori['prop_21_24']= sales_ori['시간대_21~24_매출_금액']/sales_ori['time_total']
# -
sales_ori['gender_sale']= sales_ori['남성_매출_금액']+sales_ori['여성_매출_금액']
sales_ori['female_rate']= sales_ori['여성_매출_금액']/sales_ori['gender_sale']
sales_ori.columns
df_female_rate=sales_ori[['기준_년_코드', '기준_분기_코드', '상권_코드','서비스_업종_코드','당월_매출_금액', 'female_rate' ,'prop_06_11',
'prop_11_14', 'prop_14_17', 'prop_17_21', 'prop_21_24']]
df_female_rate.columns = ['year', 'quarter', 'district','code','sales', 'female_rate','prop_06_11',
'prop_11_14', 'prop_14_17', 'prop_17_21', 'prop_21_24' ]
# ### 다른 변수들이 있는 df랑 merge 하기
df_merged.tail()
df_result=pd.merge(df_merged,df_dayrate, on=['year', 'quarter', 'district', 'code', 'sales'])
df_result[df_result['sales'].isnull()] # sales가 없는(잘못 merge된) 행이 있는지 확인
df_result=pd.merge(df_result,df_female_rate, on=['year', 'quarter', 'district', 'code', 'sales'])
df_result[df_result['sales'].isnull()] # sales가 없는(잘못 merge된) 행이 있는지 확인
df_result = df_result.drop(columns=['sales_per_store.1']) #중복된 칼럼 제거
df_result.columns
#저장
df_result.to_csv("df_result.csv", sep=",", encoding = 'euc-kr', index=False)
| dayoung_trial1/11_add_variables_modified .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# +
from __future__ import print_function
import os
import sys
import numpy as np
from keras.optimizers import SGD, Nadam, RMSprop
from keras.callbacks import CSVLogger, ModelCheckpoint
from keras.regularizers import l1, l2
sys.path.append(os.path.join(os.getcwd(), os.pardir))
import config
from utils.dataset.data_generator import DataGenerator
from models.cnn3 import cnn, cnn_regularized
from utils.training.callbacks import Logger
# +
lr=0.005
l1 = 0.00001
l2 = 0.00001
dropout = 0.5
n_epochs=100
batch_size=32
input_shape=(140, 140, 3)
weights='cnn_140_rgb_corrected_lr_0.005000_sgd_he_normal__l1_0.000010_l2_0.000010_dropout_0.500000_r_training_weights_best.hdf5'
name = 'cnn_140_rgb_corrected_full_lr_%f_sgd_he_normal__l1_%f_l2_%f_dropout_%f_r' % (lr, l1, l2, dropout)
# +
print('loading model...')
# model = cnn(input_shape=input_shape, init='he_normal')
model = cnn_regularized(input_shape=input_shape, init='he_normal', l1=l1, l2=l2, weights=weights)
model.summary()
optimizer = SGD(lr=lr, clipnorm=4., nesterov=True)
# optimizer = Nadam(lr=lr)
# optimizer = RMSprop(lr=lr)
print('compiling model...')
model.compile(optimizer=optimizer, loss='binary_crossentropy', metrics=['accuracy'])
print('done.')
logger = Logger('%s_training_batch.log' % name, append=True)
csv_logger = CSVLogger('%s_training.log' % name, append=True)
best_model_checkpointer = ModelCheckpoint(filepath=("./%s_training_weights_best.hdf5" % name), verbose=1,
save_best_only=True)
current_model_checkpointer = ModelCheckpoint(filepath=("./%s_training_weights_current.hdf5" % name), verbose=0)
# +
print('Initializing data generators...')
train_set_file = '/home/tanuj/Workspace/power-grid-detection/dataset/corrected/19/train_data.csv'
validation_set_file = '/home/tanuj/Workspace/power-grid-detection/dataset/corrected/19/validation_data.csv'
test_set_file = '/home/tanuj/Workspace/power-grid-detection/dataset/corrected/19/test_data.csv'
train_data_gen = DataGenerator(dataset_file=train_set_file, batch_size=batch_size)
validation_data_gen = DataGenerator(dataset_file=validation_set_file, batch_size=batch_size)
test_data_gen = DataGenerator(dataset_file=test_set_file, batch_size=batch_size)
print('done.')
# -
print('Fitting model...')
history = model.fit_generator(train_data_gen,
nb_epoch=n_epochs,
samples_per_epoch=train_data_gen.n_batches * batch_size,
validation_data=validation_data_gen,
nb_val_samples=validation_data_gen.n_samples,
verbose=1,
callbacks=[csv_logger, logger, best_model_checkpointer, current_model_checkpointer],
nb_worker=2)
print('done.')
# +
print('Evaluating model...')
score = model.evaluate_generator(test_data_gen, val_samples=test_data_gen.n_samples)
print('done.')
print('Test score:', score[0])
print('Test accuracy:', score[1])
# -
| training/Training_rgb-r_r-corrected-full.ipynb |