code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# This script checks for FMRIPREP func files, identifies them and writes them to a file called "MISSING_FMRIPREP".
# This script also uses the confound derivatives from fmriprep output and extracts the
# Framewise Displacement column. Then it thresholds the values for both 0.2 (conservative)
# and 0.4 (lenient) for determining motion outlier runs for final analyses.
# It generates the INCLUDE.csv file which contains all information about each run
# FD outliers, binary exclusion criteria based on 0.2 and 0.4 threshold).
# -
import numpy as np
import csv
from glob import glob
import re
import os
from statistics import mean
def remove_junk(main_list, junk):
counter = 0
for elements in main_list:
if elements in junk:
main_list[counter] = ''
counter = counter + 1
main_list = list(filter(None, main_list))
return main_list
# subject look up table conversion (IGNORING undescores)
def Convert_Subname(Oldname):
# this may no longer be a dynamic file
# if need to update this, can use the vertical concatenation of the first two columns of these files:
# /om/user/rezzo/TOMLOC_info/tomloc_subject_info_internal.csv
# /om/group/saxelab/OpenAutism/data/Subject_Task_Info_Dima/subject_info_internal.csv
tmp_root = '/om/user/rezzo/Subject_Conversion_Table.csv'
with open(tmp_root, "r") as tsv:
for line in csv.reader(tsv, delimiter = ","):
if Oldname == line[1].replace("_",""):
Newname = line[0]
else:
continue
return Newname
def remove_underscores(subname):
newname = subname.replace("_","")
return newname
def find_between( s, first, last ):
try:
start = s.index( first ) + len( first )
end = s.index( last, start )
return s[start:end]
except ValueError:
return ""
def substring_after(s, delim):
return s.partition(delim)[2]
# +
# read in site folder:
open('INCLUDE.csv', 'w').close()
root = '/om/group/saxelab/OpenAutism/data/'
all_sites = os.listdir("/om/group/saxelab/OpenAutism/data/")
site_other_folders = ['subjectlists', 'TextFiles','Subject_Task_Info_Dima','.DS_Store', '._.DS_Store']
subject_other_folders = ['mriqc_output', 'BIDS', '_OLD_', 'SPM_firstlevel','.DS_Store', '._.DS_Store','SPOverview.xlsx']
all_sites = remove_junk(all_sites, site_other_folders)
all_sites.sort()
# for each site look at how many subjects;
for site in all_sites:
all_subjects = os.listdir(root+site)
all_subjects = remove_junk(all_subjects, subject_other_folders)
all_subjects.sort()
# to look at a specific subject's tasks
for subject in all_subjects:
tempsub = subject
print(tempsub)
subject = remove_underscores(subject)
try:
fmriprep = root+site+'/BIDS/derivatives/fmriprep/sub-'+subject
location = root+site+'/BIDS/derivatives/fmriprep/sub-'+subject+'/func/'
if os.path.isdir(location):
inner_dir = os.listdir(location)
#print(inner_dir)
# for the elements in directory list, make an exact list of tasks defined by preproc. file created:
task_list = []
for elements in inner_dir:
cur_task = find_between(elements, 'sub-'+subject+'_', '_bold_space-MNI152NLin2009cAsym_variant-smoothAROMAnonaggr_preproc.nii.gz')
if cur_task != '':
task_list.append(cur_task)
for tasks in task_list:
print(subject + tasks)
if os.path.isfile(location+'sub-'+subject+'_'+tasks+'_bold_confounds.tsv') is False:
#print('missing confound file for '+subject+' '+cur_task)
fo = open("MISSING_FMRIPREP.csv", "a")
line = fo.writelines('sub-'+subject+tasks+'\t'+'no_confound_file'+'\n')
fo.close()
else:
#print("ready to go")
ls = [] #array of valid values
cor = [] #array with 0s and 1s (for >= 0.2)
cor2 = [] #array with 0s and 1s (for >= 0.4)
cor_int = []
cor2_int = []
# Only interested in the 6th column (FD)
with open(location+'sub-'+subject+"_"+tasks+'_bold_confounds.tsv') as tsv:
for line in csv.reader(tsv, dialect="excel-tab"):
array = line[6]
#print(array)
try:
array = float(array)
ls.append(array)
cor.append(array >= 0.2)
cor_int.append(array >= 0.2)
cor2.append(array >= 0.4)
cor2_int.append(array >= 0.4)
except ValueError: # Catch the error if user input is not a number
#print('Could not read number for sub-' +subject+tasks)
continue
## converting boolean true/false to zeros/ones
for element in range (0,len(cor)):
if cor[element] == 0:
cor[element] = int(0)
cor_int[element] = cor[element]
cor[element] = str(cor[element])
else:
cor[element] = int(1)
cor_int[element] = cor[element]
cor[element] = str(cor[element])
## converting boolean true/false to zeros/ones
for element in range (0,len(cor2)):
if cor2[element] == 0:
cor2[element] = int(0)
cor2_int[element] = cor2[element]
cor2[element] = str(cor2[element])
else:
cor2[element] = int(1)
cor2_int[element] = cor2[element]
cor2[element] = str(cor2[element])
motion_mean = mean(ls)
# if 25% of the trial is motion, exclude
temp_len = len(cor) # in case certain runs have different lengths
exclude = (sum(cor_int) >= (0.25*temp_len)) #changed this so be careful it is right
exclude2 = (sum(cor2_int) >= (0.25*temp_len))
out0 = Convert_Subname(subject)
out1 = site
out2 = 'sub-'+subject
out3 = find_between(tasks, "task-", "_run") # task label
out4 = substring_after(tasks, "run-") # run lavel
out5 = str(sum(cor_int))
out6 = str(int(exclude))
out7 = str(sum(cor2_int))
out8 = str(int(exclude2))
out9 = str(motion_mean)
# save to a text file
fo = open("INCLUDE.csv", "a")
for element in range (0,1):
# Write sequence of lines at the end of the file.
fo.write(out0 + "\t" + out1 + "\t" + out2 + "\t" + out3 + "\t" + out4 + "\t" + out5 + "\t"
+ out6+ "\t" + out7+ "\t" + out8 + "\t" + out9 + "\n")
fo.close()
except UnboundLocalError:
print("Check for "+subject+"in subject conversion file and for no counfound file in bids-func.")
# -
| Motion_Threshold.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="fT5TU5IL7PlG"
# ### Imports
# + id="b-Xgp5qZ6rsB"
import os
import numpy as np
import tensorflow as tf
from tensorflow import keras
import csv, json, time
import pandas as pd
# + [markdown] id="o_R85uaq9Lc1"
# ### Mounting the google drive
# + colab={"base_uri": "https://localhost:8080/"} id="yi9xrHhg9KjX" outputId="8b458f57-97a8-4b94-b5f0-710773267614"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="J9c5dvVn6rpl" outputId="7d659cdf-80e2-4c9b-b8c7-3bda2aec0815"
data_path = '/content/drive/MyDrive/NLP Data/corona-tweets'
os.path.exists(data_path)
# + id="5GgcG6uztxsM"
data_frame = pd.read_csv(os.path.join(data_path, 'Corona_NLP_test.csv'))
sentiments = data_frame.Sentiment.values
tweets = data_frame.OriginalTweet.values
# + [markdown] id="cowJr9Oy7wIN"
# ### Data preparation.
# We are going to prepare our data so that labels will be one hot encoded vectors and we will remove some punctuation marks hashtags for all the tweets.
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="WyhNek3m6rnM" outputId="260b4d33-49a1-4fec-d46a-c9c878fe5e51"
tweets[2]
# + id="z8-EWKVNoQ6Q"
import re
# + colab={"base_uri": "https://localhost:8080/", "height": 52} id="PQxDzx23oUwy" outputId="2f5c2743-6078-4de5-ba05-18a0f4861cdc"
def process_clean_text(text:str)->str:
text = text.lower()
text = re.sub(r'http[s]?://(?:[a-zA-Z]|[0-9]|[$-_@.&+]|[!*\(\),]|(?:%[0-9a-fA-F][0-9a-fA-F]))+', ' ', text)
text = re.sub(r"[^a-z0-9.,?;']", ' ', text)
text = re.sub(r'\s+', ' ', text)
return text
process_clean_text(tweets[1])
# + [markdown] id="dl2UUIVYvKTt"
# ### Label Processing.
#
# The following are labels that we have in our dataset.
# + colab={"base_uri": "https://localhost:8080/"} id="w2JlbECiuErx" outputId="2e59a698-5d0b-46d5-fbc5-2f733125d39b"
from collections import Counter
counts = Counter(sentiments)
counts
# + [markdown] id="nrDnvbzSvpBq"
# ### Visualizing labels using PrettyTable.
# + id="SwUh8xCy6rkk"
from prettytable import PrettyTable
def tabulate(column_names, data, title):
table = PrettyTable(column_names)
table.title = title
for row in data:
table.add_row(row)
print(table)
# + colab={"base_uri": "https://localhost:8080/"} id="tmXv8vgYv8DK" outputId="ee75c3ec-be23-408f-eb47-1d083bf94440"
data_rows = []
for label, count in counts.items():
data_rows.append([label.upper(), count])
data_columns = ["LABEL", "COUNTS"]
title = "LABELS COUNTS"
tabulate(data_columns, data_rows, title )
# + [markdown] id="REtgZVWixCQg"
# ### Let's process the labels.
# * We are going to convert labels to numeric
# * We are also going to `one_hot` encode labels using `scikit-learn`.
# + id="32scxMKDxgpm"
from sklearn.preprocessing import OneHotEncoder, LabelEncoder
# + id="v_aoGKaXxnUC"
label_encoder = LabelEncoder()
encoded_labels = label_encoder.fit_transform(sentiments)
# + [markdown] id="EG0NRZ7wzCGV"
# ### Now the labels are looking as follows:
#
# ```
# ['Extremely Negative', 'Extremely Positive', 'Negative', 'Neutral', "Positive"] == [0, 1, 2, 3, 4]
#
# ```
# + id="mXUHgsLyz71u"
def one_hot_encode(index, depth=5):
return np.eye(depth, dtype="float32")[index]
# + id="dFd8Lypm0OOO"
labels_one_hot = np.array(list(map(one_hot_encode, encoded_labels)))
# + [markdown] id="LenKwoKC2DVK"
# ### Text (tweets).
# Now let's map for all the features and get the cleaned version of data.
# + id="8yQWqnYF12n2"
tweets_cleaned = list(map(process_clean_text, tweets))
# + [markdown] id="Rw3Nvp2B2o1G"
# ### Spliting datasets.
#
# We are going to split the data into 3 sets:
# * train `90%` (validation 10% + training 80%)
# * test `10%`
# * validation (validation during training) using the `validation_split`.
# + colab={"base_uri": "https://localhost:8080/"} id="aaCkaeTD12lU" outputId="75fec33f-685c-4048-97a9-56964f116688"
test_size = int(.1 * len(tweets_cleaned))
test_features = tweets_cleaned[:test_size]
test_labels = labels_one_hot[:test_size]
train_features = tweets_cleaned[test_size:]
train_labels = labels_one_hot[test_size:]
data_columns = ["SET", "EXAMPLE(s)"]
title = "LABELS COUNTS"
data_rows = ["TESTING", len(test_labels)], ["TRAINING", len(train_labels)]
tabulate(data_columns, data_rows, title )
# + [markdown] id="3NAlF4iwC_gG"
# ### Processing the text (features).
# * Create a word vocabulary.
# * Create `stoi` from each sentence.
# * pad the sentences so that they will have the same size.
#
# * We are going to join the `train` and `validation` features and labels, and then we will split them during training.
#
# **We are not going to touch the test data.**
# + id="615YPxEWC_b8" colab={"base_uri": "https://localhost:8080/"} outputId="915da81d-7233-480a-bbe6-887b2359f75f"
from nltk.tokenize import word_tokenize
import nltk
nltk.download('punkt')
# + colab={"base_uri": "https://localhost:8080/"} id="Ve0gN3_VC_Zj" outputId="8420ee9a-ab21-42fa-9000-40eacb78f0fe"
counter = Counter()
for sent in train_features:
words = word_tokenize(sent)
for word in words:
counter[word] += 1
counter.most_common(9)
# + [markdown] id="A4O30HxWOMKy"
# ### Vocabulary size (aka) number of unique words.
# + colab={"base_uri": "https://localhost:8080/"} id="p9sbL8tdC_Wd" outputId="e128f363-49b5-4566-c4c5-3edff89db581"
vocab_size = len(counter)
print(f"Vocabulary size: {vocab_size:,}")
# + [markdown] id="OAZQl6w2OhL0"
# ### Creating word vectors.
# + id="WuPLINFdC_Ti"
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
# + id="PKLNiFInC_QU"
tokenizer = Tokenizer(num_words=vocab_size)
tokenizer.fit_on_texts(train_features)
# + id="aEBCExtqC_NM"
word_indices = tokenizer.word_index
word_indices_reversed = dict([(v, k) for (k, v) in word_indices.items()])
# + [markdown] id="IscmPgMpSFoM"
# ### Helper functions.
#
# We are going to create two helper function. One will convert the text given to sequences and the other will take sequences and convert them to text.
#
# + id="Em8lTdRm_ypM"
def sequence_to_text(sequences):
return " ".join(word_indices_reversed[i] for i in sequences)
def text_to_sequence(sent):
words = word_tokenize(sent.lower())
sequences = []
for word in words:
try:
sequences.append(word_indices[word])
except:
sequences.append(0)
return sequences
# + [markdown] id="zx8TgBAiTdLT"
# ### Loading pretrainned weights glove.6B.
# We are going to load this pretrained weights from our google drive. I've uploaded them on my google drive.
# + id="Zek9CrB1TOBO"
embedding_path = "/content/drive/MyDrive/NLP Data/glove.6B/glove.6B.100d.txt"
# + id="MoC7Xlly7pO6"
def hms_string(sec_elapsed):
h = int(sec_elapsed / (60 * 60))
m = int((sec_elapsed % (60 * 60)) / 60)
s = sec_elapsed % 60
return "{}:{:>02}:{:>05.2f}".format(h, m, s)
# + id="i6LrBcKsTcuL" colab={"base_uri": "https://localhost:8080/"} outputId="bdd5493b-3908-44a2-b654-89c2194698d7"
embeddings_dictionary = dict()
start = time.time()
with open(embedding_path, encoding='utf8') as glove_file:
for line in glove_file:
records = line.split()
word = records[0]
vectors = np.asarray(records[1:], dtype='float32')
embeddings_dictionary[word] = vectors
print(f"ETA: {hms_string(time.time() - start)}")
# + [markdown] id="q5Vs2gHHUXOF"
# > Creating an `embedding matrix` that suits our data.
# + id="muB0PIgiTcr1" colab={"base_uri": "https://localhost:8080/"} outputId="399a7dbe-a7a5-49fd-a418-ac54cc5fb08a"
start = time.time()
embedding_matrix = np.zeros((vocab_size, 100))
for word, index in tokenizer.word_index.items():
vector = embeddings_dictionary.get(word)
if vector is not None:
try:
embedding_matrix[index] = vector
except:
pass
print(f"ETA: {hms_string(time.time() - start)}")
# + [markdown] id="G_My9Wf8WoUQ"
# ### Creating sequences.
# + id="DFA_nBkTTcoM"
sequence_tokens = tokenizer.texts_to_sequences(train_features)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="yGvx2PZeTcjt" outputId="0833ced0-cda2-4a1c-aaaf-3a65240b5d4a"
sequence_to_text(sequence_tokens[0])
# + [markdown] id="wtB8FDOgW2Mv"
# ### Padding sequences.
# We now want our sequences to have the same size.
# + id="VHisHjeuV69a"
max_words = 100
tokens_sequence_padded = pad_sequences(sequence_tokens, maxlen=max_words, padding="post", truncating="post")
# + [markdown] id="lW_mPvD2XVCN"
# ### Building the model.
#
# ### Model achitecture.
#
# ```
# [ Embedding Layer]
# |
# |
# [ LSTM ] <---- [Bidirectional Layer] ----> [GRU] (forward_layer)
# (backward_layer) |
# |
# [ Gated Recurrent Unit (GRU) ]
# |
# |
# [ Long Short Term Memory (LSTM) ]
# |
# |
# [ Flatten Layer]
# |
# |
# [Dense Layer 1]
# |
# |
# [ Dropout ]
# |
# |
# [Dense Layer 2]
# |
# |
# [Dense Layer 3] (output [6 classes])
# ```
# + colab={"base_uri": "https://localhost:8080/"} id="k2_IPav4XBUm" outputId="b6d7ac5e-f415-4684-e861-b132a5534675"
forward_layer = keras.layers.GRU(128, return_sequences=True, dropout=.5 )
backward_layer = keras.layers.LSTM(128, activation='tanh', return_sequences=True,
go_backwards=True, dropout=.5)
input_layer = keras.layers.Input(shape=(100, ), name="input_layer")
embedding_layer = keras.layers.Embedding(
vocab_size,
100,
input_length=max_words,
weights=[embedding_matrix],
trainable=True,
name = "embedding_layer"
)(input_layer)
bidirectional_layer = keras.layers.Bidirectional(
forward_layer,
backward_layer = backward_layer,
name= "bidirectional_layer"
)(embedding_layer)
gru_layer = keras.layers.GRU(
512, return_sequences=True,
dropout=.5,
name= "gru_layer"
)(bidirectional_layer)
lstm_layer = keras.layers.LSTM(
512, return_sequences=True,
dropout=.5,
name="lstm_layer"
)(gru_layer)
conv_layer_1 = keras.layers.Conv1D(64, 3, activation='relu')(gru_layer)
conv_layer_2 = keras.layers.Conv1D(512, 3, activation='relu')(conv_layer_1)
flatten_layer = keras.layers.Flatten(name="flatten_layer")(conv_layer_2)
fc_1 = keras.layers.Dense(64, activation='relu', name="dense_layer_1")(flatten_layer)
dropout_layer = keras.layers.Dropout(rate=0.5, name="dropout_layer")(fc_1)
fc_2 = keras.layers.Dense(512, activation='relu', name="dense_layer_2")(dropout_layer)
output_layer = keras.layers.Dense(5, activation='softmax')(fc_2)
covid_tweets_model = keras.Model(inputs=input_layer, outputs=output_layer, name="covid_tweets_model")
covid_tweets_model.summary()
# + [markdown] id="8E_tEILobB-l"
# ### Compiling and training the model.
# + id="5VaKv9tObB2I"
early_stoping = keras.callbacks.EarlyStopping(
monitor='val_loss',
min_delta=0,
patience=5,
verbose=1,
mode='auto',
baseline=None,
restore_best_weights=False,
)
covid_tweets_model.compile(
loss = keras.losses.CategoricalCrossentropy(from_logits=False),
optimizer = keras.optimizers.Adam(1e-3, 0.5),
metrics = ['accuracy']
)
# + colab={"base_uri": "https://localhost:8080/"} id="fH8T6U0ibBxi" outputId="8907d52d-6d92-40aa-cf72-a2a4ca878330"
covid_tweets_model.fit(
tokens_sequence_padded,
train_labels,
epochs = 10,
verbose = 1,
validation_split = .2,
shuffle=True,
batch_size= 32,
validation_batch_size = 16,
callbacks = [early_stoping]
)
# + [markdown] id="RMjQhhzqB8NT"
# ### BERT - TEXT CLASSIFICATION
# As we can see that our model is not performing well, it is not improving from the achitecture that works perfectly from the `emotionals-nlp-notebook`. We are going to use transfare learning to get reasonable accuracy for this task. Specifically we are going to use the `BERT` model.
#
# + id="vFGxLEJsDSg6"
# + [markdown] id="ji7Jjeuafxov"
# ### Evaluating the model.
#
# + colab={"base_uri": "https://localhost:8080/"} id="nVJCnBxRf0VL" outputId="1a946def-b66a-458a-d339-c1eb38385091"
def text_to_padded_sequences(sent):
tokens = text_to_sequence(sent)
padded_tokens = pad_sequences([tokens], maxlen=max_words, padding="post", truncating="post")
return tf.squeeze(padded_tokens)
X_test = np.array(list(map(text_to_padded_sequences, X_test_values)))
emotion_model.evaluate(X_test, y_test_labels_one_hot, verbose=1, batch_size=32)
# + [markdown] id="9JGdHvW1cCB1"
# ### Inference.
# + id="vo8lPGu2qrbN"
def tabulate(column_names, data):
table = PrettyTable(column_names)
table.align[column_names[0]] = "l"
table.align[column_names[1]] = "l"
for row in data:
table.add_row(row)
print(table.get_string(title="EMOTION PREDICTIONS TABLE"))
# + id="xfTD8ynWbBuX"
def predict(model, sent):
classes = ['anger', 'fear', 'joy', 'love', 'sadness', 'surprise' ]
tokens = text_to_sequence(sent)
padded_tokens = pad_sequences([tokens], maxlen=max_words, padding="post", truncating="post")
probabilities = model.predict(padded_tokens)
prediction = tf.argmax(probabilities, axis=1).numpy()[0]
class_name = classes[prediction]
emoji_text = emoji.emojize(emotions_emojis[class_name], language='en', use_aliases=True)
table_headers =["KEY", "VALUE"]
table_data = [
["PREDICTED CLASS", prediction],
["PREDICTED CLASS NAME", class_name],
["PREDICTED CLASS EMOJI", emoji_text],
["CONFIDENCE OVER OTHER CLASSES", f'{probabilities[0][prediction] * 100:.2f}%']
]
tabulate(table_headers, table_data)
# + [markdown] id="lMmn3HgchODO"
# ### Sadness
# + colab={"base_uri": "https://localhost:8080/"} id="di_Xaqp0bBrf" outputId="9c838888-864f-43d4-b818-c9ca875614b9"
predict(emotion_model, "im updating my blog because i feel shitty.")
# + [markdown] id="vDoKLPbrhShU"
# ### Fear
# + colab={"base_uri": "https://localhost:8080/"} id="nWLTsgoWbBp5" outputId="7e8d1acd-28d1-41a2-d0e5-4fadf72755ea"
predict(emotion_model, "i am feeling apprehensive about it but also wildly excited")
# + [markdown] id="YGmhhc2hhe2E"
# ### Joy
# + colab={"base_uri": "https://localhost:8080/"} id="ROGMsOAkbBlW" outputId="e5e5d80b-535a-4d05-a1e7-b7bd7c604394"
predict(emotion_model, "i feel a little mellow today.")
# + [markdown] id="ldyKnuyCh81u"
# ### Surprise
# + colab={"base_uri": "https://localhost:8080/"} id="jxiWcolKheke" outputId="0b6d85c4-4325-40da-e83b-8534a6aac8f0"
predict(emotion_model, "i feel shocked and sad at the fact that there are so many sick people.")
# + [markdown] id="NMRVPZckiDQH"
# ### Love
# + colab={"base_uri": "https://localhost:8080/"} id="S7ZR6hRybBiH" outputId="47721995-2b6f-45fd-9c92-b172368fbd85"
predict(emotion_model, "i want each of you to feel my gentle embrace.")
# + [markdown] id="uxLlCtn-iUPr"
# ### Anger.
# + colab={"base_uri": "https://localhost:8080/"} id="nBL9swhHhpM1" outputId="9321bcdc-97b5-498b-dc5d-4c58edd00b25"
predict(emotion_model, "i feel like my irritable sensitive combination skin has finally met it s match.")
# + [markdown] id="YE_obFubiRzg"
# ### Saving the model.
# + colab={"base_uri": "https://localhost:8080/"} id="Abyb05jjixlW" outputId="bb153f80-c5c7-4c18-ee21-b1e562302c23"
emotion_model.save(os.path.join(data_path, "emotional_model.h5"))
print("Model Saved!!")
| 04_Corona_Tweets/01_CovidTweets_NLP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## What is Pandas?
# Pandas is an expansive data-analysis and manipulation package for Python, built on numpy and matplotlib. It can be installed several ways, depending on your operating system.
#
# It can be installed directly with PyPi and functions with both Python2 and Python3: ```pip install pandas```
#
# However, perhaps the easiest way is to install Anaconda, which includes dozens of packages for data science, data analysis, and the like: https://www.anaconda.com/
#
# Anaconda includes the Jupyter Notebook, which is a web interface into IPython, or Interactive Python, and is more or less a mixture between a graphical Python interpreter and an IDE, allowing correction of errors, display of charts and graphics, and a mix of code and Markdown text. There are dozens of good tutorials on IPython/Jupyter available for free online. It allows saving all your work in "notebooks," which are files with an ```.ipynb``` extension for sharing with others or running later. Your work can also be exported in normal Python files, HTML, Markdown, etc.
#
# Anaconda is available for Windows, Linux, and macOS. It installs all required dependencies in what is similar to a virtual Python environment, installed in its own folder structure.
#
#
# ## Advantages of using Jupyter or IPython Notebooks
# As mentioned, code can be run live in Jupyter, but unlike a normal interpreter, corrections can be made, state can be saved, multiple sessions/notebooks can be opened, comments and graphics can be added in order to make a very clean experience. No messy errors cluttering up the screen. The kernel can be halted and re-run as well if you wish.
#
# All examples will be done in the Jupyter Notebook in this tutorial.
#
#
# ## Why Pandas?
# Besides the numerous functions and methods Pandas has for manipulating and analyzing data, it also contains methods to read and write Excel files, CSV/TSV, XML, JSON, SQL/SQLite into and out of a Pandas "DataFrame." The DataFrame is somewhere between a 2D array and a live spreadsheet, able to be indexed, searched, filtered, and otherwise played with however you want.
#
# Each DataFrame, like a table in a database, will have columns and entries. A DataFrame may have an index, similar to the primary key in an SQLite DB. The index may be whatever you want, such as a timestamp or a MAC Address, depending how you want to look at the data.
#
# Data can be filtered, sorted, indexed, and whatnot on any of the columns. DataFrames can be deduplicated based on any one or all of the columns. They can be ordered and grouped by one or more of the column entries, depending on your goals.
#
# I will cover some of the simple but useful ways Kismet database data can be manipulated in Pandas.
#
#
# ## Getting Started in the Jupyter Notebook
# I will assume you already know how to run the Jupyter Notebook and these examples won't explain much background but should demonstrate some easy ways to use Kismet data.
#
# Note: notebooks contain "cells" which can be run by hitting ```Shift-Enter``` or the Run button in the menu bar. This allows single lines or code blocks to be run at once.
# +
#these are some necessary imports for what we're going to do
#pandas is typically imported as 'pd' because... you know... save typing 4 letters
import pandas as pd
import sqlite3
import json
#these are some other libraries that are typically used, but we won't here
#import numpy as np
#import matplotlib.pyplot as plt
# +
#define a sql query, which is just text, then open connection to db
#we could also just pull the entries we want, but I'll do * for now
sql = 'SELECT * FROM devices;'
conn = sqlite3.connect('guate.kismet')
#read data from sqlite db into pandas dataframe, called 'guate'
guate = pd.read_sql_query(sql, conn)
#close database
conn.close()
# -
#the .head() method by default shows the first five rows, as you would expect
#there is also .tail()
guate.head()
#columns can be referenced (as a Pandas Series) like a Python dictionary
guate['first_time']
#or, if they have no illegal characters (like spaces), as an attribute
guate.first_time
#one of the most-useful methods on a dataframe is the value_counts method
#here it simply tells us all values of the 'type' column and how many of each
guate.type.value_counts()
#and... we can plot that as a bar plot or whatever
#note: figsize is a tuple defining width/height of plot
#for a bar plot, 15,5 is decent
guate.type.value_counts().plot.bar(figsize=(15,5))
guate.head()
#converting unix timestamps to datetime objects is dead simple
pd.to_datetime(1578757585, unit='s')
#and... we can use the same exact method to convert an entire series/column at once
#use the pandas to_datetime method to convert the first_time and last_time to actual datetime objects
guate.first_time = pd.to_datetime(guate.first_time, unit='s')
guate.last_time = pd.to_datetime(guate.last_time, unit='s')
#looking at the dataframe now, you'll see first_time and last_time are actual datetime objects
guate.head()
#we can drop any columns we don't need/want. In this case, no gps, so no purpose to these columns
#the axis=1 refers to the columns... the index is 0 and the columns are 1
#inplace=True modifies the existing dataframe in place without copying to another
#although you could leave out inplace and assign this to a different dataframe
guate.drop(['min_lat', 'min_lon', 'max_lat', 'max_lon', 'avg_lat', 'avg_lon'],
axis=1,
inplace=True)
guate.head()
#simple method to strip a mac down to its OUI
'2C:60:0C:AA:36:7A'.replace(':', '')[0:6]
#and we can write a function to do the same thing to the entire devmac column
def get_oui(mac):
oui = mac.replace(':', '')[0:6]
return oui
# +
#and create a new column called 'oui' out of the resulting values
#using the apply() method
#so... apply will do something to an entire column and assign the result to the same or different column
guate['oui'] = guate.devmac.apply(get_oui)
# -
guate.head()
#then we can do some meaningful analysis of the OUIs
#in this case, the DAA119 is the winner. No surprise there.
guate.oui.value_counts()
#here we take the same value_counts() method, get the top 10 of them, then plot a barplot (vertical)
guate.oui.value_counts()[0:10].plot.bar(figsize=(15,5),
width=0.9,
logy=False,
facecolor='green',
edgecolor='blue',
title='Dragorn Does Kismet',
rot=20,
grid=True
)
guate.head()
#using iloc to look at entry 0 (first entry) of the dataframe
#iloc is the standard way to look at the nth element in a dataframe
guate.iloc[0]
#and pulling just the device column, which is the json record of that device
#technically it's a bytes object, but that's basically a string with different encodings possible
guate.iloc[0].device
#a simple function to return just the manufacturer from the json data
#this will be used to create a new column in our dataframe
def get_manu(device):
devdata = json.loads(device)
manu = devdata['kismet.device.base.manuf']
return manu
#use the apply method to create the manuf column from the json data
guate['manuf'] = guate.device.apply(get_manu)
guate.head()
#take a look at which manufacturers are most prevalent
guate.manuf.value_counts()
#and plot them in a horizontal bar plot, ignoring the 'Unknown' devices by starting at item 1, not 0
#I'm using sort_values to sort them in descending order. Leave it out to get them in ascending.
guate.manuf.value_counts()[1:15].sort_values().plot.barh(figsize=(12,6),
width=0.9,
grid=True,
title='Most Common Manufacturers')
#same sort of thing as before, but this time get the commonname of the device
def get_name(device):
devdata = json.loads(device)
name = devdata['kismet.device.base.commonname']
return name
#and apply that function to the device's json data to create a new column (devname)
guate['devname'] = guate.device.apply(get_name)
guate.head()
#then use value_counts to see what names are most prevalent
guate.devname.value_counts()
#and plot a pie plot of the first 10
guate.devname.value_counts()[0:10].plot.pie(figsize=(10,10))
#a lot of standard Python functions work
#here it's just the length of the dataframe (# of records)
len(guate)
#min method to get the smallest value
guate.strongest_signal.min()
#max method to get the largest
guate.strongest_signal.max()
#same thing, but with the timestamps
guate.first_time.min()
#get max timestamp
guate.last_time.max()
# +
#connect to new database, pull data
conn2 = sqlite3.connect('tapachula_hotel.kismet')
tap_hotel = pd.read_sql_query(sql, conn2)
conn2.close()
# -
tap_hotel.head()
#standard Python set notation works too
#here we "convert" the devmac column to a set, called g1
g1 = set(guate.devmac)
#and the same for the tap_hotel's devmac column
g2 = set(tap_hotel.devmac)
#and use Python's set intersection to find the devices that show up in both
g1.intersection(g2)
#connect to 3rd db and pull data into dataframe with same sql query as before
conn3 = sqlite3.connect('tapachula_airport.kismet')
tap_ap = pd.read_sql_query(sql, conn3)
tap_ap.head()
#get device macs into a set
g3 = set(tap_ap.devmac)
#and get their intersection
g1.intersection(g3)
#let's see what was in the hotel and the airport
g2.intersection(g3)
#let's get their manufacturers by running the get_manu function we wrote earlier
tap_hotel['manuf'] = tap_hotel.device.apply(get_manu)
tap_ap['manuf'] = tap_ap.device.apply(get_manu)
#put the common devices in a list
common_devices = list(g2.intersection(g3))
common_devices
#create a new dataframe from the devices with devmacs that are in our list
#note the isin method and the structure of this 'filter'
#note, since they occur in both tap_ap and tap_hotel, you could use either dataframe to get this list
#but... one or the other might have additional data in the device's json field
interesting_devices = tap_ap[tap_ap.devmac.isin(common_devices)]
#These appear to mostly be mobile devices except 00:25:00:FF:94:73, which is Apple P2p Airplay (adhoc)
#yeah, yeah, still mobile, but not specific to one device
#also 02:00:00:00:00:00, which is an Android mac when you don't have permissions to get the mac
interesting_devices
#lastly, one thing I didn't do on the live stream is a basic filter
#the mask actually creates a truth table of True/False for the entire Series of values
#then the guate[mask] applies it to the dataframe
mask = guate.type=='Wi-Fi Client'
guate[mask]
#you can also use some string methods with these, like .contains, .startswith, .endswith, etc.
mask2 = (guate.type.str.contains('Wi-Fi'))
guate[mask2]
| kismet_db_guat_mex/kismet_pandas_basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Dependencies
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _kg_hide-input=true _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import json, warnings, shutil
from tweet_utility_scripts import *
from tweet_utility_preprocess_roberta_scripts import *
from transformers import TFRobertaModel, RobertaConfig
from tokenizers import ByteLevelBPETokenizer
from tensorflow.keras.models import Model
from tensorflow.keras import optimizers, metrics, losses, layers
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint, LearningRateScheduler
SEED = 0
seed_everything(SEED)
warnings.filterwarnings("ignore")
# -
# # Load data
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _kg_hide-input=true _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
database_base_path = '/kaggle/input/tweet-dataset-split-roberta-base-96/'
k_fold = pd.read_csv(database_base_path + '5-fold.csv')
display(k_fold.head())
# Unzip files
# !tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_1.tar.gz
# !tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_2.tar.gz
# !tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_3.tar.gz
# !tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_4.tar.gz
# !tar -xvf /kaggle/input/tweet-dataset-split-roberta-base-96/fold_5.tar.gz
# -
# # Model parameters
# +
vocab_path = database_base_path + 'vocab.json'
merges_path = database_base_path + 'merges.txt'
base_path = '/kaggle/input/qa-transformers/roberta/'
config = {
"MAX_LEN": 96,
"BATCH_SIZE": 32,
"EPOCHS": 6,
"LEARNING_RATE": 3e-5,
"ES_PATIENCE": 1,
"question_size": 4,
"N_FOLDS": 5,
"base_model_path": base_path + 'roberta-base-tf_model.h5',
"config_path": base_path + 'roberta-base-config.json'
}
with open('config.json', 'w') as json_file:
json.dump(json.loads(json.dumps(config)), json_file)
# -
# # Tokenizer
# + _kg_hide-output=true
tokenizer = ByteLevelBPETokenizer(vocab_file=vocab_path, merges_file=merges_path,
lowercase=True, add_prefix_space=True)
tokenizer.save('./')
# -
# ## Learning rate schedule
# + _kg_hide-input=true
LR_MIN = 1e-6
LR_MAX = config['LEARNING_RATE']
LR_EXP_DECAY = .5
@tf.function
def lrfn(epoch):
lr = LR_MAX * LR_EXP_DECAY**epoch
if lr < LR_MIN:
lr = LR_MIN
return lr
rng = [i for i in range(config['EPOCHS'])]
y = [lrfn(x) for x in rng]
fig, ax = plt.subplots(figsize=(20, 6))
plt.plot(rng, y)
print("Learning rate schedule: {:.3g} to {:.3g} to {:.3g}".format(y[0], max(y), y[-1]))
# -
# # Model
# +
module_config = RobertaConfig.from_pretrained(config['config_path'], output_hidden_states=True)
def model_fn(MAX_LEN):
input_ids = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='input_ids')
attention_mask = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='attention_mask')
base_model = TFRobertaModel.from_pretrained(config['base_model_path'], config=module_config, name="base_model")
_, _, hidden_states = base_model({'input_ids': input_ids, 'attention_mask': attention_mask})
h12 = hidden_states[-1]
h11 = hidden_states[-2]
h10 = hidden_states[-3]
h09 = hidden_states[-4]
x = layers.Average()([h12,h11, h10, h09])
x_start = layers.Dropout(.1)(x)
x_start = layers.Dense(1)(x_start)
x_start = layers.Flatten()(x_start)
y_start = layers.Activation('softmax', name='y_start')(x_start)
x_end = layers.Dropout(.1)(x)
x_end = layers.Dense(1)(x_end)
x_end = layers.Flatten()(x_end)
y_end = layers.Activation('softmax', name='y_end')(x_end)
model = Model(inputs=[input_ids, attention_mask], outputs=[y_start, y_end])
optimizer = optimizers.Adam(lr=config['LEARNING_RATE'])
model.compile(optimizer, loss={'y_start': losses.CategoricalCrossentropy(label_smoothing=0.2),
'y_end': losses.CategoricalCrossentropy(label_smoothing=0.2)},
metrics={'y_start': metrics.CategoricalAccuracy(),
'y_end': metrics.CategoricalAccuracy()})
return model
# -
# # Train
# + _kg_hide-input=true _kg_hide-output=true
history_list = []
for n_fold in range(config['N_FOLDS']):
n_fold +=1
print('\nFOLD: %d' % (n_fold))
# Load data
base_data_path = 'fold_%d/' % (n_fold)
x_train = np.load(base_data_path + 'x_train.npy')
y_train = np.load(base_data_path + 'y_train.npy')
x_valid = np.load(base_data_path + 'x_valid.npy')
y_valid = np.load(base_data_path + 'y_valid.npy')
### Delete data dir
shutil.rmtree(base_data_path)
# Train model
model_path = 'model_fold_%d.h5' % (n_fold)
model = model_fn(config['MAX_LEN'])
es = EarlyStopping(monitor='val_loss', mode='min', patience=config['ES_PATIENCE'],
restore_best_weights=True, verbose=1)
checkpoint = ModelCheckpoint(model_path, monitor='val_loss', mode='min',
save_best_only=True, save_weights_only=True)
lr_schedule = LearningRateScheduler(lrfn)
history = model.fit(list(x_train), list(y_train),
validation_data=(list(x_valid), list(y_valid)),
batch_size=config['BATCH_SIZE'],
callbacks=[checkpoint, es, lr_schedule],
epochs=config['EPOCHS'],
verbose=2).history
history_list.append(history)
# Make predictions
train_preds = model.predict(list(x_train))
valid_preds = model.predict(list(x_valid))
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'train', 'start_fold_%d' % (n_fold)] = train_preds[0].argmax(axis=-1)
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'train', 'end_fold_%d' % (n_fold)] = train_preds[1].argmax(axis=-1)
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'validation', 'start_fold_%d' % (n_fold)] = valid_preds[0].argmax(axis=-1)
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'validation', 'end_fold_%d' % (n_fold)] = valid_preds[1].argmax(axis=-1)
k_fold['end_fold_%d' % (n_fold)] = k_fold['end_fold_%d' % (n_fold)].astype(int)
k_fold['start_fold_%d' % (n_fold)] = k_fold['start_fold_%d' % (n_fold)].astype(int)
k_fold['end_fold_%d' % (n_fold)].clip(0, k_fold['text_len'], inplace=True)
k_fold['start_fold_%d' % (n_fold)].clip(0, k_fold['end_fold_%d' % (n_fold)], inplace=True)
k_fold['prediction_fold_%d' % (n_fold)] = k_fold.apply(lambda x: decode(x['start_fold_%d' % (n_fold)], x['end_fold_%d' % (n_fold)], x['text'], config['question_size'], tokenizer), axis=1)
k_fold['prediction_fold_%d' % (n_fold)].fillna(k_fold["text"], inplace=True)
k_fold['jaccard_fold_%d' % (n_fold)] = k_fold.apply(lambda x: jaccard(x['selected_text'], x['prediction_fold_%d' % (n_fold)]), axis=1)
# -
# # Model loss graph
# + _kg_hide-input=true
sns.set(style="whitegrid")
for n_fold in range(config['N_FOLDS']):
print('Fold: %d' % (n_fold+1))
plot_metrics(history_list[n_fold])
# -
# # Model evaluation
# + _kg_hide-input=true
display(evaluate_model_kfold(k_fold, config['N_FOLDS']).style.applymap(color_map))
# -
# # Visualize predictions
# + _kg_hide-input=true
display(k_fold[[c for c in k_fold.columns if not (c.startswith('textID') or
c.startswith('text_len') or
c.startswith('selected_text_len') or
c.startswith('text_wordCnt') or
c.startswith('selected_text_wordCnt') or
c.startswith('fold_') or
c.startswith('start_fold_') or
c.startswith('end_fold_'))]].head(15))
| Model backlog/Train/99-tweet-train-5fold-roberta-base-lbl-smth02-avg4.ipynb |
# ##### Copyright 2021 Google LLC.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # assignment_with_constraints_sat
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/examples/assignment_with_constraints_sat.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/examples/python/assignment_with_constraints_sat.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
# Copyright 2010-2021 Google LLC
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""Solve an assignment problem with combination constraints on workers."""
from ortools.sat.python import cp_model
def solve_assignment():
"""Solve the assignment problem."""
# Data.
cost = [[90, 76, 75, 70, 50, 74], [35, 85, 55, 65, 48,
101], [125, 95, 90, 105, 59, 120],
[45, 110, 95, 115, 104, 83], [60, 105, 80, 75, 59, 62], [
45, 65, 110, 95, 47, 31
], [38, 51, 107, 41, 69, 99], [47, 85, 57, 71,
92, 77], [39, 63, 97, 49, 118, 56],
[47, 101, 71, 60, 88, 109], [17, 39, 103, 64, 61,
92], [101, 45, 83, 59, 92, 27]]
group1 = [
[0, 0, 1, 1], # Workers 2, 3
[0, 1, 0, 1], # Workers 1, 3
[0, 1, 1, 0], # Workers 1, 2
[1, 1, 0, 0], # Workers 0, 1
[1, 0, 1, 0]
] # Workers 0, 2
group2 = [
[0, 0, 1, 1], # Workers 6, 7
[0, 1, 0, 1], # Workers 5, 7
[0, 1, 1, 0], # Workers 5, 6
[1, 1, 0, 0], # Workers 4, 5
[1, 0, 0, 1]
] # Workers 4, 7
group3 = [
[0, 0, 1, 1], # Workers 10, 11
[0, 1, 0, 1], # Workers 9, 11
[0, 1, 1, 0], # Workers 9, 10
[1, 0, 1, 0], # Workers 8, 10
[1, 0, 0, 1]
] # Workers 8, 11
sizes = [10, 7, 3, 12, 15, 4, 11, 5]
total_size_max = 15
num_workers = len(cost)
num_tasks = len(cost[1])
all_workers = range(num_workers)
all_tasks = range(num_tasks)
# Model.
model = cp_model.CpModel()
# Variables
selected = [[model.NewBoolVar('x[%i,%i]' % (i, j)) for j in all_tasks]
for i in all_workers]
works = [model.NewBoolVar('works[%i]' % i) for i in all_workers]
# Constraints
# Link selected and workers.
for i in range(num_workers):
model.AddMaxEquality(works[i], selected[i])
# Each task is assigned to at least one worker.
for j in all_tasks:
model.Add(sum(selected[i][j] for i in all_workers) >= 1)
# Total task size for each worker is at most total_size_max
for i in all_workers:
model.Add(
sum(sizes[j] * selected[i][j] for j in all_tasks) <= total_size_max)
# Group constraints.
model.AddAllowedAssignments([works[0], works[1], works[2], works[3]],
group1)
model.AddAllowedAssignments([works[4], works[5], works[6], works[7]],
group2)
model.AddAllowedAssignments([works[8], works[9], works[10], works[11]],
group3)
# Objective
model.Minimize(
sum(selected[i][j] * cost[i][j] for j in all_tasks
for i in all_workers))
# Solve and output solution.
solver = cp_model.CpSolver()
status = solver.Solve(model)
if status == cp_model.OPTIMAL:
print('Total cost = %i' % solver.ObjectiveValue())
print()
for i in all_workers:
for j in all_tasks:
if solver.BooleanValue(selected[i][j]):
print('Worker ', i, ' assigned to task ', j, ' Cost = ',
cost[i][j])
print()
print('Statistics')
print(' - conflicts : %i' % solver.NumConflicts())
print(' - branches : %i' % solver.NumBranches())
print(' - wall time : %f s' % solver.WallTime())
solve_assignment()
| examples/notebook/examples/assignment_with_constraints_sat.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Erasmus+ ICCT project (2018-1-SI01-KA203-047081)
# Toggle cell visibility
from IPython.display import HTML
tag = HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide()
} else {
$('div.input').show()
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
Toggle cell visibility <a href="javascript:code_toggle()">here</a>.''')
display(tag)
# Hide the code completely
# from IPython.display import HTML
# tag = HTML('''<style>
# div.input {
# display:none;
# }
# </style>''')
# display(tag)
# + [markdown] lang="it"
# ## Osservatore dello stato: sistemi non osservabili
# Questo esempio mostra cosa succede quando un osservatore è progettato per un sistema lineare non osservabile.
#
# Ricordando la forma standard di osservabilità:
#
# \begin{cases}
# \underbrace{\begin{bmatrix} \dot{x}_o \\ \dot{x}_{\bar{o}} \end{bmatrix}}_{\dot{\textbf{x}}} = \begin{bmatrix} A_o & 0 \\ A_{o\bar{o}} & A_{\bar{o}} \end{bmatrix} \underbrace{\begin{bmatrix} x_o \\ x_{\bar{o}} \end{bmatrix}}_{\textbf{x}} + \begin{bmatrix} B_o \\ B_{\bar{o}} \end{bmatrix} \textbf{u} \\
# \textbf{y} = \begin{bmatrix} C_o & 0 \end{bmatrix} \textbf{x},
# \end{cases}
#
# costruire:
# - un sistema non osservabile la cui dinamica non osservabile è stabile;
# - un sistema non osservabile la cui dinamica non osservabile è instabile;
# - un sistema non osservabile la cui dinamica non osservabile ha dimensione 1 e autovalore 0. Cosa cambia in questo caso?
# +
#Preparatory Cell
# %matplotlib inline
import control as control
import numpy
import sympy as sym
from IPython.display import display, Markdown
import ipywidgets as widgets
import matplotlib.pyplot as plt
#print a matrix latex-like
def bmatrix(a):
"""Returns a LaTeX bmatrix - by <NAME> (ICCT project)
:a: numpy array
:returns: LaTeX bmatrix as a string
"""
if len(a.shape) > 2:
raise ValueError('bmatrix can at most display two dimensions')
lines = str(a).replace('[', '').replace(']', '').splitlines()
rv = [r'\begin{bmatrix}']
rv += [' ' + ' & '.join(l.split()) + r'\\' for l in lines]
rv += [r'\end{bmatrix}']
return '\n'.join(rv)
# Display formatted matrix:
def vmatrix(a):
if len(a.shape) > 2:
raise ValueError('bmatrix can at most display two dimensions')
lines = str(a).replace('[', '').replace(']', '').splitlines()
rv = [r'\begin{vmatrix}']
rv += [' ' + ' & '.join(l.split()) + r'\\' for l in lines]
rv += [r'\end{vmatrix}']
return '\n'.join(rv)
#matrixWidget is a matrix looking widget built with a VBox of HBox(es) that returns a numPy array as value !
class matrixWidget(widgets.VBox):
def updateM(self,change):
for irow in range(0,self.n):
for icol in range(0,self.m):
self.M_[irow,icol] = self.children[irow].children[icol].value
#print(self.M_[irow,icol])
self.value = self.M_
def dummychangecallback(self,change):
pass
def __init__(self,n,m):
self.n = n
self.m = m
self.M_ = numpy.matrix(numpy.zeros((self.n,self.m)))
self.value = self.M_
widgets.VBox.__init__(self,
children = [
widgets.HBox(children =
[widgets.FloatText(value=0.0, layout=widgets.Layout(width='90px')) for i in range(m)]
)
for j in range(n)
])
#fill in widgets and tell interact to call updateM each time a children changes value
for irow in range(0,self.n):
for icol in range(0,self.m):
self.children[irow].children[icol].value = self.M_[irow,icol]
self.children[irow].children[icol].observe(self.updateM, names='value')
#value = Unicode('<EMAIL>', help="The email value.").tag(sync=True)
self.observe(self.updateM, names='value', type= 'All')
def setM(self, newM):
#disable callbacks, change values, and reenable
self.unobserve(self.updateM, names='value', type= 'All')
for irow in range(0,self.n):
for icol in range(0,self.m):
self.children[irow].children[icol].unobserve(self.updateM, names='value')
self.M_ = newM
self.value = self.M_
for irow in range(0,self.n):
for icol in range(0,self.m):
self.children[irow].children[icol].value = self.M_[irow,icol]
for irow in range(0,self.n):
for icol in range(0,self.m):
self.children[irow].children[icol].observe(self.updateM, names='value')
self.observe(self.updateM, names='value', type= 'All')
#self.children[irow].children[icol].observe(self.updateM, names='value')
#overlaod class for state space systems that DO NOT remove "useless" states (what "professor" of automatic control would do this?)
class sss(control.StateSpace):
def __init__(self,*args):
#call base class init constructor
control.StateSpace.__init__(self,*args)
#disable function below in base class
def _remove_useless_states(self):
pass
# +
# Preparatory cell
A = numpy.matrix('-1 1 0; 0 -2 0; 0 1 2')
B = numpy.matrix('0.7; 0.5; 1')
C = numpy.matrix('1 0 0')
X0 = numpy.matrix('2; 2; 2')
F = numpy.matrix('-1 0 3; 0 -2 -1; 0 1 -2')
G = numpy.matrix('0; 0; 1')
L = numpy.matrix([[4],[-2/3],[2/3]])
sol1 = numpy.linalg.eig(A)
Adefault = [A,
numpy.matrix('-1 1 0; 0 -2 0; 0 1 -2')]
Bdefault = B
Cdefault = C
Aw = matrixWidget(3,3)
Aw.setM(A)
Bw = matrixWidget(3,1)
Bw.setM(B)
Cw = matrixWidget(1,3)
Cw.setM(C)
X0w = matrixWidget(3,1)
X0w.setM(X0)
Fw = matrixWidget(3,3)
Fw.setM(F)
Gw = matrixWidget(3,1)
Gw.setM(G)
Lw = matrixWidget(3,1)
Lw.setM(L)
eig1o = matrixWidget(1,1)
eig2o = matrixWidget(2,1)
eig3o = matrixWidget(1,1)
eig1o.setM(numpy.matrix([-3]))
eig2o.setM(numpy.matrix([[-3],[0]]))
eig3o.setM(numpy.matrix([-3]))
# +
# Misc
#create dummy widget
DW = widgets.FloatText(layout=widgets.Layout(width='0px', height='0px'))
#create button widget
START = widgets.Button(
description='Test',
disabled=False,
button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Test',
icon='check'
)
def on_start_button_clicked(b):
#This is a workaround to have intreactive_output call the callback:
# force the value of the dummy widget to change
if DW.value> 0 :
DW.value = -1
else:
DW.value = 1
pass
START.on_click(on_start_button_clicked)
# default systems
sel_system = widgets.Dropdown(
options=[('Manuale','Manual'), ('Stato instabile e inosservabile','Unstable unobservable state'), ('Stato stabile e inosservabile','Stable unobservable state')],
value='Unstable unobservable state',
description='',
disabled=False
)
#define type of ipout
selu = widgets.Dropdown(
options=[('impulso','impulse'), ('gradino','step'), ('sinusoide','sinusoid'), ('onda quadra','square wave')],
value='impulse',
description='Input:',
disabled=False
)
# Define the values of the input
u = widgets.FloatSlider(
value=1,
min=0,
max=20.0,
step=0.1,
description='input u:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.1f',
)
period = widgets.FloatSlider(
value=0.5,
min=0.0,
max=10,
step=0.01,
description='Periodo: ',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.2f',
)
# +
def main_callback(A, B, C, X0w, L, u, period, sel_system, selu, DW):
global Adefault, Bdefault, Cdefault, Aw, Bw, Cw, Lw
if sel_system == 'Manual':
sol = numpy.linalg.eig(A-L*C)
elif sel_system == 'Unstable unobservable state':
A = Adefault[0]
B = Bdefault
C = Cdefault
Aw.setM(A)
Bw.setM(B)
Cw.setM(C)
L = numpy.matrix([[4],[2],[2]])
Lw.setM(L)
sol = numpy.linalg.eig(A-L*C)
elif sel_system == 'Stable unobservable state':
A = Adefault[1]
B = Bdefault
C = Cdefault
Aw.setM(A)
Bw.setM(B)
Cw.setM(C)
L = numpy.matrix([[4],[2],[10]])
Lw.setM(L)
sol = numpy.linalg.eig(A-L*C)
sol1 = numpy.linalg.eig(A)
print('Gli autovalori del sistema sono:',round(sol1[0][0],4),',',round(sol1[0][1],4),'e',round(sol1[0][2],4))
print('Gli autovalori dell\'osservatore sono:',round(sol[0][0],4),',',round(sol[0][1],4),'e',round(sol[0][2],4))
sys = sss(A,B,C,0)
syso = sss(A-L*C, numpy.concatenate((B,L),axis=1), numpy.eye(3), numpy.zeros(6).reshape((3,2)))
if (numpy.real([sol[0][0],sol[0][1],sol[0][2]]) == [0, 0, 0]).all():
T = numpy.linspace(0,20,1000)
else:
if min(numpy.abs(numpy.real([sol[0][0],sol[0][1],sol[0][2]]))) != 0:
T = numpy.linspace(0,10*1/min(numpy.abs(numpy.real([sol[0][0],sol[0][1],sol[0][2]]))),1000)
else:
T = numpy.linspace(0,10*1/max(numpy.abs(numpy.real([sol[0][0],sol[0][1],sol[0][2]]))),1000)
if selu == 'impulse': #selu
U = [0 for t in range(0,len(T))]
U[0] = u
T, yout, xout = control.forced_response(sys,T,U,X0w)
T, youto, xouto = control.forced_response(syso,T,numpy.matrix([U,yout]),[[0],[0],[0]])
if selu == 'step':
U = [u for t in range(0,len(T))]
T, yout, xout = control.forced_response(sys,T,U,X0w)
T, youto, xouto = control.forced_response(syso,T,numpy.matrix([U,yout]),[[0],[0],[0]])
if selu == 'sinusoid':
U = u*numpy.sin(2*numpy.pi/period*T)
T, yout, xout = control.forced_response(sys,T,U,X0w)
T, youto, xouto = control.forced_response(syso,T,numpy.matrix([U,yout]),[[0],[0],[0]])
if selu == 'square wave':
U = u*numpy.sign(numpy.sin(2*numpy.pi/period*T))
T, yout, xout = control.forced_response(sys,T,U,X0w)
T, youto, xouto = control.forced_response(syso,T,numpy.matrix([U,yout]),[[0],[0],[0]])
fig = plt.figure(num='Simulation', figsize=(16,10))
fig.add_subplot(321)
plt.ylabel('$X_1$ vs $X_{1est}$ (uscita del sistema)')
plt.plot(T,xout[0])
plt.plot(T,xouto[0])
plt.xlabel('t [s]')
plt.legend(['Reale','Stimato'])
plt.axvline(x=0,color='black',linewidth=0.8)
plt.axhline(y=0,color='black',linewidth=0.8)
plt.grid()
fig.add_subplot(323)
plt.ylabel('$X_2$ vs $X_{2est}$')
plt.plot(T,xout[1])
plt.plot(T,xouto[1])
plt.xlabel('t [s]')
plt.legend(['Reale','Stimato'])
plt.axvline(x=0,color='black',linewidth=0.8)
plt.axhline(y=0,color='black',linewidth=0.8)
plt.grid()
fig.add_subplot(325)
plt.ylabel('$X_3$ vs $X_{3est}$')
plt.plot(T,xout[2])
plt.plot(T,xouto[2])
plt.xlabel('t [s]')
plt.legend(['Reale','Stimato'])
plt.axvline(x=0,color='black',linewidth=0.8)
plt.axhline(y=0,color='black',linewidth=0.8)
plt.grid()
fig.add_subplot(322)
plt.ylabel('$e_1$')
plt.plot(T,xouto[0]-xout[0])
plt.xlabel('t [s]')
plt.grid()
fig.add_subplot(324)
plt.ylabel('$e_1$')
plt.plot(T,xouto[1]-xout[1])
plt.xlabel('t [s]')
plt.grid()
fig.add_subplot(326)
plt.ylabel('$e_1$')
plt.plot(T,xouto[2]-xout[2])
plt.xlabel('t [s]')
plt.grid()
fig1 = plt.figure(num='Output simulation', figsize=(16,5))
fig1.add_subplot(111)
plt.ylabel('$y$')
plt.plot(T,yout)
plt.xlabel('t [s]')
plt.grid()
alltogether = widgets.VBox([widgets.HBox([sel_system, widgets.Label(' ',border=3), selu]),
widgets.Label(' ',border=3),
widgets.HBox([widgets.Label('A:',border=3), Aw,
widgets.Label(' ',border=3),
widgets.Label(' ',border=3),
widgets.Label('B:',border=3), Bw,
widgets.Label(' ',border=3),
widgets.Label(' ',border=3),
widgets.Label('X0:',border=3), X0w,
widgets.Label(' ',border=3),
widgets.Label(' ',border=3),
widgets.Label('L:',border=3), Lw]),
widgets.Label(' ',border=3),
widgets.HBox([widgets.Label('C:',border=3), Cw]),
widgets.Label(' ',border=3),
widgets.HBox([u,
period,
START])])
out = widgets.interactive_output(main_callback, {'A':Aw, 'B':Bw, 'C':Cw, 'X0w':X0w, 'L':Lw,
'u':u, 'period':period, 'sel_system':sel_system,
'selu':selu, 'DW':DW})
out.layout.height = '980px'
display(out, alltogether)
| ICCT_it/examples/04/SS-29-Osservatore_per_sistemi_non_osservabili.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split, KFold, cross_val_score
from sklearn.metrics import accuracy_score, roc_auc_score, average_precision_score
# +
def fitted_svm(params):
svm = SVC(**params)
fitted_svm = svm.fit(X_train, Y_train)
return fitted_svm
def ClassifierMetrics (X_train, Y_train, X_test, Y_test, fitted_model):
Y_pred = fitted_model.predict(X_test)
metrics = {'train_accuracy': fitted_model.score(X_train, Y_train)
'test_accuracy': fitted_model.score(X_test, Y_test)
'test_auc': roc_auc_score(Y_test, Y_pred)
'test_ap': average_precision_score(Y_test, Y_pred)}
return metrics
# -
svm_model_params = {'kernel':[],'C':[], 'train_accuracy':[],'test_accuracy':[],'test_auc':[], 'test_r2':[]}
# +
svm_pd = pd.DataFrame(svm_model_params)
results_path = '../data/svm_svc_results_hj.csv'
svm_pd.to_csv(results_path, sep=',')
# svm_pd = pd.read_csv(results_path, sep=',', index_col=0)
| notebooks/ac4119/.ipynb_checkpoints/03-svm-model-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Evaluate features
#
# This notebook demonstrate how to evaluate the features stored in the anndata.obsm.
# The task we are interested in is to predict the gene expression based on the cell_type label and the covariates.
#
# I the next notebook (notebook3_all.ipynb) we loop over all the features and compare them to each other.
# +
# TO REMOVE when notebook is stable
# %load_ext autoreload
# %autoreload 2
# -
# ### Common Imports
# +
import numpy
import torch
import seaborn
import tarfile
import os
import matplotlib
import matplotlib.pyplot as plt
from anndata import read_h5ad
# tissue_purifier import
import tissue_purifier as tp
# -
# ### Download the annotated anndata object
#
# Altenatively you can use the anndata files generated by running notebook2_all.
# +
import tissue_purifier.io
bucket_name = "ld-data-bucket"
annotated_anndata_source_path = "tissue-purifier/annotated_slideseq_testis_anndata_h5ad.tar.gz"
annotated_anndata_dest_path = "./annotated_slideseq_testis_anndata_h5ad.tar.gz"
annotated_anndata_dest_folder = "./testis_anndata_annotated"
#tp.io.download_from_bucket(bucket_name, annotated_anndata_source_path, annotated_anndata_dest_path)
#with tarfile.open(annotated_anndata_dest_path, "r:gz") as fp:
# fp.extractall(path=annotated_anndata_dest_folder)
# Make a list of all the h5ad files in the annotated_anndata_dest_folder
fname_list = []
for f in os.listdir(annotated_anndata_dest_folder):
if f.endswith('.h5ad'):
fname_list.append(f)
print(fname_list)
# -
# ### Decide how to filter the anndata object
# +
# filter cells parameters
fc_bc_min_umi = 200 # filter cells with too few UMI
fc_bc_max_umi = 3000 # filter cells with too many UMI
fc_bc_min_n_genes_by_counts = 10 # filter cells with too few GENES
fc_bc_max_n_genes_by_counts = 2500 # filter cells with too many GENES
fc_bc_max_pct_counts_mt = 5 # filter cells with mitocrondial fraction too high
# filter genes parameters
fg_bc_min_cells_by_counts = 3000 # filter genes which appear in too few CELLS
# filter rare cell types parameters
fctype_bc_min_cells_absolute = 100 # filter cell-types which are too RARE in absolute number
fctype_bc_min_cells_frequency = 0.01 # filter cell-types which are too RARE in relative abundance
# -
# ### Open the first annotated anndata
adata = read_h5ad(filename=os.path.join(annotated_anndata_dest_folder, fname_list[0]))
adata
# ### compute few metrics
# +
import scanpy as sc
cell_type_key = "cell_type"
# mitocondria metrics
adata.var['mt'] = adata.var_names.str.startswith('mt-') # annotate the group of mitochondrial genes as 'mt'
sc.pp.calculate_qc_metrics(adata, qc_vars=['mt'], percent_top=None, log1p=False, inplace=True)
# counts cells frequency
tmp = adata.obs[cell_type_key].values.describe()
print(tmp)
mask1 = (tmp["counts"] > fctype_bc_min_cells_absolute)
mask2 = (tmp["freqs"] > fctype_bc_min_cells_frequency)
mask = mask1 * mask2
cell_type_keep = set(tmp[mask].index.values)
adata.obs["keep_ctype"] = adata.obs["cell_type"].apply(lambda x: x in cell_type_keep)
# Note that adata has extra annotation now
adata
# -
# ### Filter out cells, genes and cell-type
adata = adata[adata.obs["total_counts"] > fc_bc_min_umi, :]
adata = adata[adata.obs["total_counts"] < fc_bc_max_umi, :]
adata = adata[adata.obs["n_genes_by_counts"] > fc_bc_min_n_genes_by_counts, :]
adata = adata[adata.obs["n_genes_by_counts"] < fc_bc_max_n_genes_by_counts, :]
adata = adata[adata.obs["pct_counts_mt"] < fc_bc_max_pct_counts_mt, :]
adata = adata[adata.obs["keep_ctype"] == True, :]
adata = adata[:, adata.var["n_cells_by_counts"] > fg_bc_min_cells_by_counts]
# ### Show the cell-type count after filtering
tmp = adata.obs[cell_type_key].values.describe()
print(tmp)
adata
# ### Make a gene dataset from the anndata
# Here we choose the Neighboorhood Composition Vector computed using k=10 nearest neighbours.
# This serves as a reasonable baseline behavior.
# +
from tissue_purifier.genex import *
covariate_key = "ncv_k10"
gene_dataset = make_gene_dataset_from_anndata(
anndata=adata,
cell_type_key='cell_type',
covariate_key=covariate_key,
preprocess_strategy='raw',
apply_pca=False)
# -
gene_dataset.describe()
# ### Split the dataset into train/test/val
train_dataset, test_dataset, val_dataset = next(iter(train_test_val_split(gene_dataset, random_state=0)))
print("---------")
train_dataset.describe()
print("---------")
test_dataset.describe()
print("---------")
val_dataset.describe()
# ### Note that the split is tratified by cell_types
# +
ncols=3
fig, axes = plt.subplots(ncols=ncols, figsize=(6*ncols,6))
for c, (dataset, title) in enumerate(zip([train_dataset, test_dataset, val_dataset], ["train dataset", "test dataset", "val dataset"])):
x = list(dataset.cell_type_mapping.keys())
y = torch.bincount(dataset.cell_type_ids).numpy()
_ = seaborn.barplot(x=x, y=y, ax=axes[c])
x_labels_raw = axes[c].get_xticklabels()
axes[c].set_xticklabels(labels=x_labels_raw, rotation=90)
_ = axes[c].set_title(title)
# -
# ### Example training the model with no covariate (i.e. cell-type only)
gr = GeneRegression()
gr.configure_optimizer(optimizer_type='adam', lr=5E-4)
# +
# this will take about 20 minutes on a NVIDIA T4 GPU
# adjust subsample_size_cells and subsample_size_genes to make sure:
# 1. not to run out of GPU memory
# 2. have high GPU utilization
gr.train(
dataset=train_dataset,
n_steps=10000,
print_frequency=100,
use_covariates=True,
l1_regularization_strength=None,
l2_regularization_strength=None,
eps_range=(1.0E-5, 1.0E-1),
subsample_size_cells=2000,
subsample_size_genes=None,
initialization_type="scratch")
gr.save_ckpt("gr_ncv_k10_from_scratch.pt")
# -
# ### check the loss to see if the model has converged
fig, axes = plt.subplots(ncols=2, figsize=(8,4))
gr.show_loss(ax=axes[0])
gr.show_loss(ax=axes[1], logy=True, logx=True)
# _ = axes[0].set_ylim(2E7,2.5E7)
# ### if not satisfied, extend the training and save a new checkpoint
# +
# Here we:
# 1. extend the training for 5000 additional steps
# 2. recheck the loss function
# 3. save the ckpt
# gr.extend_train(
# dataset=train_dataset,
# n_steps=5000,
# print_frequency=100)
#
# fig, axes = plt.subplots(ncols=2, figsize=(8,4))
# gr.show_loss(ax=axes[0])
# gr.show_loss(ax=axes[1], logy=True)
#
# gr.save_ckpt("gr_ncv_k10_extended.pt")
# + [markdown] tags=[]
# # Visualization
# -
# get the fitted parameters
df_param = gr.get_params()
df_param
# Recall that in the LogNormalPoisson observation model the average count is given by:
#
# c_av = N_umi * exp[log_mu + 0.5 * eps]
#
# and that, in our linear model,:
#
# log_mu = beta0 + (beta dot X)
#
# Therefore:
#
# log(c_av) = log N_umi + beta0 + (beta X) + 0.5 * eps
#
# where:
# 1. N_umi of shape (n) is the total number of counts in a cell.
# 2. beta0 has shape (k,g) where K is the number of cell-types
# 3. beta has shape (k,l,g) where L is the number of covariates
# 4. X is the fixed covariate matrix of shape (n,l)
# 5. eps is the overdispersion of shape (k, g)
# ### Plot gene overdispersion
g = seaborn.displot(data=df_param, x="eps", hue="cell_type", col="cell_type", kind='hist', bins=200, common_bins=True, col_wrap=3)
_ = g.fig.suptitle("Distribution of eps, i.e. overdispersion", fontsize=16)
# g = g.set(xlim=gr._train_kargs["eps_range"])
fig, ax = plt.subplots(figsize=(12,6))
_ = seaborn.histplot(data=df_param, x="eps", hue="cell_type", bins=200, ax=ax, multiple="layer")
_ = ax.set_title("Distribution on gene-overdispersion params, i.e. eps")
#_ = ax.set_xlim(gr._train_kargs["eps_range"])
# We observe that:
# 1. the distribution of eps is well contained into the allowed_range. Therefore we do not need to tune/change the eps_range hyperparameter.
# 2. ES cells have higher overdispersion
# ### Plot distribution of beta0.
g = seaborn.displot(data=df_param, x="beta_0", hue="cell_type", col="cell_type", kind='hist', bins=200, common_bins=True, col_wrap=3)
g.fig.subplots_adjust(top=0.9)
_ = g.fig.suptitle("Distribution of beta_0, i.e. log_rate", fontsize=16)
fig, ax = plt.subplots(figsize=(12,6))
_ = seaborn.histplot(data=df_param, x="beta_0", hue="cell_type", bins=200, ax=ax)
_ = ax.set_title("Distribution of beta0, i.e. log of rate")
# ## Distribution of beta
# +
beta_names = []
for item in df_param.columns:
if item.startswith("beta") and item != "beta_0":
beta_names.append(item)
print(beta_names)
nmax = len(beta_names)
ncols = 3
nrows = int(numpy.ceil(float(nmax)/ncols))
fig, axes = plt.subplots(ncols=ncols, nrows=nrows, figsize=(6*ncols, 6*nrows))
fig.suptitle("Distribution of betas")
for n, beta_name in enumerate(beta_names):
r,c = n//ncols, n%ncols
ax_cur = axes[r,c]
_ = seaborn.histplot(data=df_param, x=beta_name, hue="cell_type", ax=ax_cur)
_ = ax_cur.set_title(beta_name)
# -
# From these plots we few interesting facts.
# For example:
# 1. the distribution of beta8 (which is the coefficients 'SPG' fraction) is wider for SPG cells than other cell_types. This seems to suggest that SPG cells are strongly influenced by the presence of other SPG cells in their neighborhood.
# 2. Leydig cells seems very affected by Endothelial cells (see beta2 plot)
#
# We look at beta_8 distribution in more details
g = seaborn.displot(data=df_param, x="beta_8", hue="cell_type", col="cell_type", kind='hist', bins=200, common_bins=True, col_wrap=3)
g.fig.subplots_adjust(top=0.9)
_ = g.fig.suptitle("Distribution of beta_8", fontsize=16)
# ## Distribution of highly expressed genes
# find the top k highly expressed (he) gene
he_gene_index = torch.topk(test_dataset.counts.sum(axis=0), axis=-1, k=10)[1]
he_gene_names = numpy.array(test_dataset.gene_names)[he_gene_index]
print(he_gene_index)
print(he_gene_names)
# run the predict method on the test dataset
df_metric, df_counts = gr.predict(
dataset=test_dataset,
num_samples=100,
subsample_size_cells=400,
subsample_size_genes=None)
df_just_one_gene = df_counts[df_counts["gene"] == he_gene_names[0]]
df_just_one_gene.head()
# +
unique_cell_types = numpy.unique(df_just_one_gene["cell_type"].values)
nmax = len(unique_cell_types)
ncols = 3
nrows = int(numpy.ceil(float(nmax)/ncols))
fig, axes = plt.subplots(ncols=ncols, nrows=nrows, figsize=(6*ncols, 6*nrows))
fig.suptitle("Distribution of gene={}".format(he_gene_names[0]))
for n, cell_type in enumerate(unique_cell_types):
r,c = n//ncols, n%ncols
ax_cur = axes[r,c]
df_tmp = df_just_one_gene[ df_just_one_gene["cell_type"] == cell_type]
_ = seaborn.histplot(data=df_tmp, x="counts_obs", color="skyblue", binwidth=1, alpha=0.5, discrete=True, ax=ax_cur)
_ = seaborn.histplot(data=df_tmp, x="counts_pred", color="red", binwidth=1, alpha=0.5, discrete=True, ax=ax_cur)
_ = ax_cur.set_title("Cell type = {}".format(cell_type))
# -
# ## Q_dist metrics
df_metric.head()
# +
g = seaborn.displot(data=df_metric, x="q_dist", hue="cell_type", col="cell_type",
kind='hist', stat="probability", discrete=True, col_wrap=3)
_ = g.fig.suptitle("Distribution of Q (i.e. absolute difference between obs and predicted)", fontsize=16)
# this is calling the matplotlib.axes.Axes.set() method
g = g.set(xlim=(0,5), ylim=(0,0.15), xticks=range(0,5), xticklabels=range(0,5))
# -
# same data as above but shown as a table
df_metric.groupby("cell_type").describe()["q_dist"]
# ## log_score metric
# +
g = seaborn.displot(data=df_metric, x="log_score", hue="cell_type", col="cell_type",
kind='hist', col_wrap=3)
#stat="probability", discrete=True, col_wrap=3)
_ = g.fig.suptitle("Distribution of log_p of observed counts under the predicted distribution", fontsize=16)
# this is calling the matplotlib.axes.Axes.set() method
#g = g.set(xlim=(0,4))
# -
# same data but as a table
df_metric.groupby("cell_type").describe()["log_score"]
| notebooks/notebook3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
#Misc
import scipy.io as sio
import os #to access files from the directories
import math
import time #to check total time took for running the script or function
#Visualization
import matplotlib.pyplot as plt
#Analysis
import pandas as pd
import numpy as np
from sklearn import svm
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import permutation_test_score
from sklearn.model_selection import StratifiedKFold
from sklearn.utils import shuffle
from sklearn.naive_bayes import GaussianNB
from sklearn import tree
# -
dataFolder = "imp/Data/"
columns = ["means","label"]
#df = pd.DataFrame({'A':1,'B':2}, index = None)
data_list = []
for matFile in os.listdir(dataFolder):
if matFile.startswith("Faces") and not matFile.endswith("389.mat"):
data = sio.loadmat(dataFolder + matFile)
for i in range (len(data["means"])):
#data_list.append(list(zip(data["means"],data["label"][0]))) # Check this how to do it
d = data["means"][i],data["label"][0][i]
data_list.append(d)
df = pd.DataFrame(data_list, columns = columns)
RoiNames = (data["RoiName"][:,0])
colRoi = []
for roi in RoiNames:
colRoi.append(roi[0])
df[colRoi] = pd.DataFrame(df.means.values.tolist(), index = df.index)
df.drop(['means'], axis=1, inplace=True)
#Extracting different labels entries
df1 = df[df.label == 1]
df2 = df[df.label == 2]
df3 = df[df.label == 3]
# +
#Combining two pairs off all combination
df12 = shuffle(df1.append(df2))
df23 = shuffle(df2.append(df3))
df31 = shuffle(df3.append(df1))
print(df12.label)
# -
# !conda install -c conda-forge pysurfer --yes
| .ipynb_checkpoints/trying-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/MJVelilla/CS488S21/blob/main/python_materials/learn-python3/notebooks/beginner/exercises/numbers_exercise.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="-7zRPe5OJr1x"
# # 1. Creating formulas
# Write the following mathematical formula in Python:
#
# \begin{align}
# result = 6a^3 - \frac{8b^2 }{4c} + 11
# \end{align}
#
# + editable=false id="On_PTXP_Jr15"
a = 2
b = 3
c = 2
# + id="FiwplRymJr15"
# Your formula here:
result = (6*(a**3)) - ((8*(b**2))/(4*c)) + 11
# + editable=false id="gDcsZJK6Jr16"
assert result == 50
# + [markdown] id="n0m2CyB0Jr16"
# # 2. Floating point pitfalls
# Show that `0.1 + 0.2 == 0.3`
# + id="YgFYJfM9Jr16" outputId="77719392-ef38-404c-916a-483d5eaa5d7d" colab={"base_uri": "https://localhost:8080/", "height": 214}
# Your solution here
from decimal import Decimal
str1 = Decimal("0.1")
str2 = Decimal("0.2")
print(str1 + str2)
# This won't work:
#assert 0.1 + 0.2 == 0.3
| python_materials/learn-python3/notebooks/beginner/exercises/numbers_exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/MOTURUPRAVEENBHARGAV/bert_models/blob/main/BERT(ktrain)_Sentiment_Analysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="UcQbleiLQOdo"
# Step 1: Installing ktrain
# + id="JunyltpzEZNs" colab={"base_uri": "https://localhost:8080/"} outputId="9de0b5f8-395f-4637-9547-ea19af8ff555"
#installing ktrain
# !pip install ktrain
# + [markdown] id="Pjrc-hhkQKeg"
# Step 2: Importing modules and reading the training file
# + id="hv59UcpRTECJ" colab={"base_uri": "https://localhost:8080/"} outputId="d984cd89-d794-42cf-de60-712a385c4190"
# %%time
import time
import ktrain
from ktrain import text
import pandas as pd
from sklearn.model_selection import train_test_split
import numpy as np
# + id="NeiyRuVHTSGL" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="b23fa708-8df9-40f0-b1ef-dde06a90fe42"
data=pd.read_csv("/content/drive/My Drive/Sentiment Analysis_Preskale/Cleaned_reviews.csv",index_col="Unnamed: 0")
data.head()
# + id="O1pZvvuwTtHe" colab={"base_uri": "https://localhost:8080/"} outputId="61a5c4cd-8a94-4ef5-e063-8f6605d308be"
data=data.loc[:,["content","Polarity","clean_text"]]
data.columns
# + id="LhPyzWOSNOtd" colab={"base_uri": "https://localhost:8080/"} outputId="d7ba22cf-f76e-45b2-cb3f-a764eb006b8f"
data.Polarity.value_counts()
# + id="X7mg2iCZNUsr" colab={"base_uri": "https://localhost:8080/", "height": 124} outputId="5d4e8741-5b0e-49be-9f69-152cae9a6ffc"
# cond=[df.Polarity=="Positive",df.Polarity=="Neutral",df.Polarity=="Negative"]
# choice=[2,1,0]
# df.Polarity=np.select(cond,choice)
# df.Polarity.head()
# + id="p9eXi5dwMKfv"
# #Splitting the data into X,y
# X=df["content"]
# y=df["Polarity"]
# X_train,X_test,y_train,y_test=train_test_split(X,y,shuffle=True,test_size=0.30, random_state=1)
# + id="_-UZy7ejrnuC" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="df5df71e-c8b2-4432-c898-a1c9078d1eb5"
type(X_train)
# + id="HaEhg8wcMtpz" colab={"base_uri": "https://localhost:8080/", "height": 88} outputId="be283d2a-8192-42ad-cf4d-6ede150af8be"
# print(f"X_train: {X_train.shape}")
# print(f"y_train: {y_train.shape}")
# print(f"X_test: {X_test.shape}")
# print(f"y_test: {y_test.shape}")
# + [markdown] id="Kq0gTzXdPuiB"
# ####Step 3: Convert data to Features for BERT
# ktrain provides a very convenient feature of directly converting the text
# data directly into features for the model that is needed. All the text
# preprocessing steps do not need to be manually performed and will be
# taken care of by the library itself. Since we will be reading the data from
# pandas series objects, we will use the function texts_from_array .
#
# + id="6Wa6Trx5Ptwe" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="2c2ecd31-af0a-4ad5-a013-77e06ead440c"
(X_train_bert,y_train_bert), (X_val_bert,y_val_bert), preproc= text.texts_from_df(train_df=data,
text_column="content",
label_columns="Polarity",
maxlen=150,
preprocess_mode='bert')
# + id="mf6445Kdua5A" colab={"base_uri": "https://localhost:8080/", "height": 266} outputId="18df5eea-e25a-4f07-a651-efc5d48f4098"
X_val_bert
# + id="2rKTGlE2uyDt" colab={"base_uri": "https://localhost:8080/"} outputId="d0eda6e5-3867-4800-e95c-7975bfafe3fc"
print(len(X_val_bert))
print("Total length of the data:" ,data.shape)
print("Shape of Validation Data :" , X_val_bert[0].shape)
print("Shape of Train Data:", X_train_bert[0].shape)
# + id="91OKMNL2IRvb"
# + id="c1ok047iISDa"
# + [markdown] id="6aIuZlnqXggh"
# Step 4: Load Bert in a learner object
# + id="FN-fhXv1XV7K" colab={"base_uri": "https://localhost:8080/"} outputId="72e702b2-9473-4292-9174-0dd4e378cb08"
model = text.text_classifier(name = 'bert',
train_data = (X_train_bert, y_train_bert),
preproc = preproc)
# + [markdown] id="4SmA7SC6YPh0"
# The function " text_classifier " loads the pre-trained BERT model with a randomly initialized final Dense layer. It is worthwhile to mention that although the final Dense layer is randomly initialized, it will not be only one getting updated during the training process. Since we have not frozen any layers and all the layers of the model are trainable, the weights of all the layers of the model will be updated during backpropagation.
# + id="H5urDluXYOai"
learner = ktrain.get_learner(model=model, train_data=(X_train_bert, y_train_bert),
val_data = (X_val_bert, y_val_bert),
batch_size =32)
# + [markdown] id="OUJEO5UoY37L"
# The function " get_learner " creates a learner object with train and validation data which can be used to fine-tune the classifier. The last argument of get_learner is the batch size. We use a small batch size of 10.
# + [markdown] id="0i2i6jGdY_kN"
# Step 5: Training (Finetuning the BERT Classifier)
# + id="h5cEoWNxYzvx"
#learner.lr_find() to find the best learning rate
# + id="9P_al4edbtsA" colab={"base_uri": "https://localhost:8080/"} outputId="31823158-4400-4681-dab2-33fd6d5fbb52"
#Essentially fit is a very basic training loop, whereas fit one cycle uses the one cycle policy callback
history=learner.fit_onecycle(lr = 2e-5, epochs = 4)
# + id="HwRj9JW4FMNg" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="2d71d102-229e-49ef-8cfe-bf38a7b39b4d"
history.history
# + id="6JKqu9oCvljE" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="27478d1b-24f9-41af-fbb1-e6b2117c2060"
type(history.history)
# + id="qz3XwfdOFmOy" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="082cc47f-0834-4563-fa67-2c393cce60c4"
history.history.keys()
# + id="RqhXas-OGa_U" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="add539d5-02e0-490a-caba-d43149103c2e"
report=learner.validate(val_data=(X_val_bert, y_val_bert),class_names=['Negative', 'Neutral','Positive'])
# + id="YjIbwTkyTV5I" colab={"base_uri": "https://localhost:8080/", "height": 70} outputId="84c53de9-5c8c-4ea1-d01f-8aa2ed593f24"
report
# + id="NJDREQM3UyY2"
#plotting confusion matrix
import matplotlib.pyplot as plt
def plot_confusion_matrix(cm, classes,normalize=False,title='BERT Confusion matrix',cmap=plt.cm.YlOrRd):
"""
See full source and example:
http://scikit-learn.org/stable/auto_examples/model_selection/plot_confusion_matrix.html
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=0)
plt.yticks(tick_marks, classes)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, cm[i, j],
horizontalalignment="right",
color="White" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# + id="cjfg5xrBVICo" colab={"base_uri": "https://localhost:8080/", "height": 329} outputId="67f89728-f91b-47ef-9ca0-259843f4c5c2"
import itertools
from google.colab import files
# plt.figure(figsize=(5,5))
plot_confusion_matrix(report, classes=['Negative', 'Neutral','Positive'])
# files.download( "bert confusion matrix.jpg" )
# + id="HViRwR7hMJlT" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="583dd3ea-f269-4130-8239-a999fe864b94"
learner.print_layers()
# + [markdown] id="pP5XuTeIG8Mo"
# ### Getting predictor variable
# + id="OEc2qsdkFpAF"
predictor = ktrain.get_predictor(learner.model, preproc)
# + [markdown] id="nQ3-FCwHHSQR"
# #### Saving the Model
# + id="Tj0c7mhgHVHd"
predictor.save('/content/drive/My Drive/Sentiment Analysis_Preskale')
# + id="0-XK1QWhH3XE"
#sample dataset to test on
data = ['this movie was horrible, the plot was really boring. acting was okay',
'the fild is really sucked. there is not plot and acting was bad',
'what a beautiful movie. great plot. acting was good. will see it again']
# + id="FL-4MAh4H7hs" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="76ee4c00-b385-4183-c5e7-cc6982345e51"
predictor.predict(data)
# + id="l4vZ5IAoIU9V"
#loading the model
predictor_load = ktrain.load_predictor('/content/drive/My Drive/Sentiment Analysis_Preskale')
# + id="_RoLvY0oT1rg" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="3a0030e4-a1c5-498b-b712-e03f02312777"
predictor_load.predict(data)
# + id="ALxCIKvrD0qN"
| BERT(ktrain)_Sentiment_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Time Series Forecasting w/ Facebook's Prophet
#
# Adopted from https://facebook.github.io/prophet/docs/quick_start.html.
#
# Other good resources include:
# https://xang1234.github.io/prophet/.
#
# And this textbook on Forecasting https://otexts.com/fpp2/.
#
# By default, Prophet uses an additive model
#
# \begin{align}
# y(t) = g(t) + s(t) + h(t) + \epsilon_t \\
# \end{align}
#
# where
# g(t) represents the trend, s(t) the periodic (i.e. seasonal) component,
# h(t) holiday related events and ϵ_t the error.
# The periodic component can be broken into monthly seasonality and weekly seasonality components.
# The model can also be switched to use a multiplicative process.
#
# If you want to look into Bayesian time series modeling or dynamic linear modeling, checkout this repo: https://github.com/wwrechard/pydlm.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import git
repo = git.Repo("./", search_parent_directories=True)
homedir = repo.working_dir
# Facebook's Open Source Time Series Analysis Tool
# pip install pystan
# pip install fbprophet
# If above ^ command doesn't install, see below issue
# https://github.com/facebook/prophet/issues/775
# pip install fbprophet --no-cache-dir --no-binary :all:
# Also for interactive visualizations you can install Plotly
# pip install plotly
import datetime
from fbprophet import Prophet
from fbprophet.plot import add_changepoints_to_plot
from fbprophet.diagnostics import cross_validation
from fbprophet.diagnostics import performance_metrics
from fbprophet.plot import plot_cross_validation_metric
df = pd.read_csv(f"{homedir}/data/us/flu/cases/ILI_Ages.csv")
df.head()
# Prophet requires daily data, so we will just transform from week to day
def get_first_day_of_week(yr, wk):
d = "%s-W%s" % (yr, wk)
# The -1 and -%w pattern tells the parser to pick the Monday in that week
firstdayofweek = datetime.datetime.strptime(d + '-1', "%Y-W%W-%w")
return firstdayofweek
# Prophet also requires dates to be stored in a column called 'ds'
# And that the values to predict will be stored in a column called 'y'
y_name = "ILITOTAL"
df["ds"] = df.apply(lambda row: get_first_day_of_week(row.YEAR, row.WEEK), axis=1)
df["y"] = df[y_name]
df.tail()
# Possible setting changes
# changepoint_prior_scale=0.5 (1 default)
# seasonality_mode='multiplicative' ('additive' default)
# Adding separate regressors to the model
m = Prophet()
m.fit(df)
future = m.make_future_dataframe(periods=90)
future.tail()
forecast = m.predict(future)
forecast.tail()
forecast[['ds', 'yhat', 'yhat_lower', 'yhat_upper']].tail()
# Note that H1N1 outbreak and increased volumn of patients break the forecast
fig1 = m.plot(forecast, xlabel="Time", ylabel=y_name)
# Uncomment to see where the trend line changes
# a=add_changepoints_to_plot(fig1.gca(), m, forecast)
fig1 = m.plot(forecast, xlabel="Time", ylabel=y_name)
# Uncomment to see where the trend line changes
a=add_changepoints_to_plot(fig1.gca(), m, forecast)
# Plot the yearly trends and monthly seasonality component
fig2 = m.plot_components(forecast)
# Cross Validation to measure forecast error
# Note: This takes a couple min (3-5)
# %time df_cv = cross_validation(m, initial='%i days' % (20 * 365), period='30 days', horizon = '365 days')
df_p = performance_metrics(df_cv)
df_p.head()
| exploratory/connor_time_series_flu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
#the datasheets are loaded
playstore=pd.read_csv('googleplaystore.csv')
review=pd.read_csv('googleplaystore_user_reviews.csv')
playstore.head()
playstore['Reviews'].replace('3.0M','3000000',inplace=True)
playstore['Reviews']=playstore['Reviews'].astype('int')
x=playstore[playstore['Reviews']>5000]
x=x[x['Type']=='Free']
x.shape
x.drop(columns=['Type', 'Price', 'Genres', 'Last Updated'],inplace=True)
x=x[(x['Content Rating']=='Everyone')|(x['Content Rating']=='Teen')]
for i in x['Size']:
if(i[-1]=='M'):
x['Size'].replace(i,1024*float(i[:-1]),inplace=True)
elif(i[-1]=='k'):
x['Size'].replace(i,float(i[:-1]),inplace=True)
playstore=x
playstore=playstore[playstore['Size']!='Varies with device']
playstore.head()
playstore['Size']=playstore['Size'].astype('int')
D={'1,000,000+':1000000, '10,000,000+':10000000, '5,000,000+':5000000, '500,000+':500000, '100,000,000+':100000000, '50,000,000+':50000000, '100,000+':100000, '500,000,000+':500000000,
'1,000,000,000+':1000000000, '50,000+':50000}
for i in D.keys():
playstore['Installs'].replace(i,D[i],inplace=True)
traffic=playstore.groupby('App')['Size'].mean()*playstore.groupby('App')['Installs'].mean()/(1024**3)
traffic=traffic.reset_index()
traffic.rename(columns={0:'TB Downloaded'},inplace=True)
playstore=playstore.merge(traffic,on='App')
playstore['TB Downloaded']=playstore['TB Downloaded'].astype('int')
z=playstore.groupby('App')['Reviews'].mean()*playstore.groupby('App')['Rating'].mean()/10000
z=z.reset_index()
playstore=playstore.merge(z,on='App')
playstore.rename(columns={0:'Popularity'},inplace=True)
playstore['Popularity']=playstore['Popularity'].astype('int')
current=playstore.groupby('Current Ver')['App'].count()
current=current.reset_index()
playstore=playstore.merge(current,on='Current Ver')
android=playstore.groupby('Android Ver')['App_x'].count()
android=android.reset_index()
playstore=playstore.merge(android,on='Android Ver')
playstore.drop(columns=['Android Ver','Current Ver'],inplace=True)
playstore.rename(columns={'App_x_x':'App','App_y':'Current Ver','App_x_y':'Android Ver'},inplace=True)
category=playstore.groupby('Category')['Rating'].mean()
category=category.reset_index()
playstore=playstore.merge(category,on='Category')
playstore.drop(columns=['Category'],inplace=True)
playstore.rename(columns={'Rating_x':'Rating','Rating_y':'Category'},inplace=True)
playstore.head()
playstore.drop(columns=['Installs','Size'],inplace=True)
review=review[review['Sentiment_Polarity']!=0]
review=review[review['Sentiment_Subjectivity']>0.60]
sentiment=review.groupby('App')['Sentiment_Polarity'].mean()*review.groupby('App')['Sentiment_Subjectivity'].mean()
sentiment=sentiment.reset_index()
playstore=playstore.merge(sentiment,on='App')
playstore.rename(columns={0:'Sentiment'},inplace=True)
playstore.corr()['Rating']
playstore.drop(columns=['App'],inplace=True)
playstore=playstore[['Reviews', 'TB Downloaded','Content Rating','Category','Popularity','Current Ver','Android Ver', 'Sentiment','Rating']]
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import OneHotEncoder
X=playstore.iloc[:,:-1].values
y=playstore.iloc[:,-1].values
encoder=LabelEncoder()
X[:,2]=encoder.fit_transform(X[:,2])
oencoder=OneHotEncoder(categorical_features=[2])
X=oencoder.fit_transform(X).toarray()
X.shape
X=X[:,1:]
X.shape
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestRegressor
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.2,random_state=9)
regressor=RandomForestRegressor(n_estimators=300)
regressor.fit(X_train,y_train)
y_pred=regressor.predict(X_test)
from sklearn.metrics import r2_score
from sklearn.metrics import mean_absolute_error
r2_score(y_test,y_pred)
| Playstore Rating.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.2
# language: julia
# name: julia-0.6
# ---
# 
# ### Author: [<NAME>](https://www.linkedin.com/in/getmykhan)
# --------
#importing dependencies
using GLM #Generalized Linear Models
using DataFrames # Data Manipulation and Stats
using Gadfly # Plotting
alpha = readtable("SalaryData.csv") # Read csv
data = DataFrame(alpha); # Convert it to a DataFrame
# ## EDA
# Quick EDA
head(alpha)
describe(alpha)
# ## Visualization
head(data)
#Plotting the Training result
plot(layer(x=data[1], y=data[2], Geom.point),
Guide.XLabel("Years of Experience"),
Guide.YLabel("Salary"))
# ## Fitting GLM models
# ### To fit a Generalized Linear Model (GLM), use the function, glm(formula, data, family, link), where,
#
# * formula: uses column symbols from the DataFrame data, for example, if names(data)=[:Y,:X1,:X2], then a valid formula is @formula(Y ~ X1 + X2)
# * data: a DataFrame which may contain NA values, any rows with NA values are ignored
# * family: chosen from Bernoulli(), Binomial(), Gamma(), Normal(), or Poisson()
# * link: chosen from the list below, for example, LogitLink() is a valid link for the Binomial() family
#
# An intercept is included in any GLM by default.
OLS = glm(@formula(Salary ~ YearsExperience), data, Normal(), IdentityLink())
coef(OLS) # Coefficients
stderr(OLS)
# ### Prediction
predict(OLS);
DataFrame([predict(OLS),data[2]])
new_pred_year = DataFrame(YearsExperience = [1,12,2.1,11]);
# Predicting new values
predict(OLS, new_pred_year)
#Plotting the Regression Line result
plot(layer(x=data[1], y=data[2], Geom.point),
layer(x = data[1], y= predict(OLS), Geom.line),
Guide.XLabel("Years of Experience"),
Guide.YLabel("Salary"))
# ### ends here!
| Julia for Machine Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
import numpy as np
import sympy as sym
# $$
# \Psi_{k,l}(t) = - \frac{\partial^{l - 1}}{\partial t ^{l - 1}} \left( \prod_{j = 0, j \neq k}^a (\lambda_j + t)^{-r_j} \right)
# =
# \begin{cases}
# \frac{(-1)^{l} (l-1)!}{t^l (t + \lambda_2)^l}
# \times \sum_{j=1}^{l} {l \choose j} t^{l - j} \lambda_2 ^ {j - 1}, & k=1 \\
# - \frac{1}{t (t + \lambda_1)^{r_1}}, & k=2
# \end{cases}
# $$
#
# +
def general_psi_function(arg, k, l, exp_rates, freq, a):
t = sym.symbols("t")
product = np.math.prod([(exp_rates[j] + t) ** (- freq[j]) for j in range(a + 1) if j != k])
psi_val = - sym.diff(product, t, l - 1)
psi_val = psi_val.subs({t: arg})
return psi_val
def specific_psi_function(arg, k, l, exp_rates, freq, a):
psi_val = 0
if k == 1:
for term in range(1, l + 1):
psi_val += math.comb(l, term) * arg ** (l - term) * exp_rates[2] ** (term - 1)
psi_val *= (-1) ** l * math.factorial(l - 1) / (arg ** l * (arg + exp_rates[2]) ** l)
if k == 2:
psi_val = -1 / (arg * (arg + exp_rates[1]) ** freq[1])
return psi_val
# +
lambda_1 = sym.Symbol("lambda_1")
lambda_2 = sym.Symbol("lambda_2")
r_2 = sym.Symbol("r_2")
t = sym.Symbol("t")
exp_rates = [sym.S(0), lambda_1, lambda_2]
freq = [1, r_2, 1]
a = len(exp_rates) - 1
# -
sym.factor(general_psi_function(arg=9, k=1, l=5, exp_rates=exp_rates, freq=freq, a=a))
sym.factor(specific_psi_function(arg=9, k=1, l=5, exp_rates=exp_rates, freq=freq, a=a))
# # $\Large{\textbf{Need to show that}}$:
# $$
# \Psi_{k,l}(t) = - \frac{\partial^{l - 1}}{\partial t ^{l - 1}} \left( \prod_{j = 0, j \neq k}^a (\lambda_j + t)^{-r_j} \right)
# =
# \begin{cases}
# \frac{(-1)^{l} (l-1)!}{t^l (t + \lambda_2)^l}
# \times \sum_{j=1}^{l} {l \choose j} t^{l - j} \lambda_2 ^ {j - 1}, & k=1 \\
# - \frac{1}{t (t + \lambda_1)^{r_1}}, & k=2
# \end{cases}
# $$
#
#
# $$
# \vec{\lambda} = (0, \lambda_1, \lambda_2) \\
# \vec{r} = (1, r_1, 1)
# $$
# Given our specific problem, we know that $k = 1,2$ and:
# \begin{align}
# & k = 1 \quad \Rightarrow \quad l \in [1, n] \\
# & k = 2 \quad \Rightarrow \quad l = 1
# \end{align}
#
# Therefore by applying this to $\Psi_{k,l}(t)$ we can get the values of $\Psi(t)$ for $k=1$ and $k=2$:
# # $\Large{\textbf{Proof for } k = 2}$:
# \begin{align}
# k=2, l=1 \qquad \Rightarrow \Psi_{2,1}(t) &= - \frac{\partial^0}{\partial t ^ 0} \left( \prod_{j = 0, j \neq k}^2 (\lambda_j + t)^{-r_j} \right) \\
# & = - (\lambda_0 + t)^{-r_0} (\lambda_1 + t)^{-r_1} \\
# & = - \frac{1}{t(t + \lambda_1)^{r_1}} \\
# & \hspace{4cm} \square
# \end{align}
#
# # $\Large{\textbf{Proof for } k = 1}$:
#
# $$
# - \frac{\partial^{l - 1}}{\partial t ^{l - 1}} \left( \prod_{j = 0, j \neq k}^a (\lambda_j + t)^{-r_j} \right)
# = \frac{(-1)^{l} (l-1)!}{t^l (t + \lambda_2)^l} \times \sum_{j=1}^{l} {l \choose j} t^{l - j} \lambda_2 ^ {j - 1}
# $$
#
# \begin{align}
# LHS &= - \frac{\partial^{l-1}}{\partial t ^ {l-1}} \left( \prod_{j = 0, j \neq k}^2 (\lambda_j + t)^{-r_j} \right) \\
# & = - \frac{\partial^{l-1}}{\partial t ^ {l-1}} \left( (\lambda_0 + t)^{-r_0} (\lambda_2 + t)^{-r_2} \right) \\
# & = - \frac{\partial^{l-1}}{\partial t ^ {l-1}} \left( \frac{1}{t (t + \lambda_2)} \right) \\
# \\
# \\
# RHS &= \frac{(-1)^{l} (l-1)!}{t^l (t + \lambda_2)^l} \times \sum_{j=1}^{l} {l \choose j} t^{l - j} \lambda_2 ^ {j - 1} \\
# & = \frac{(-1)^{l} (l-1)!}{t^l \lambda_2 (t + \lambda_2)^l} \times \sum_{j=1}^{l} {l \choose j} \lambda_2^j t^{l - j} \\
# & = \frac{(-1)^{l} (l-1)!}{t^l \lambda_2 (t + \lambda_2)^l} \times \left( \left[ \sum_{j=0}^{l} {l \choose j} \lambda_2^j t^{l - j} \right] -t^l \right) \\
# * & = \frac{(-1)^{l} (l-1)!}{t^l \lambda_2 (t + \lambda_2)^l} \times \left((t + \lambda_2)^l -t^l \right) \\
# & = \frac{(-1)^{l} (l-1)!}{\lambda_2} \left[\frac{1}{t^l} - \frac{1}{(t + \lambda_2)^l}\right] \\
# \end{align}
#
# ### *Binomial Theorem:
#
# $$
# (x + y)^n = \sum_{j=0}^n {n \choose j} x^{n-j} y^j
# $$
# ## Need to show that:
# $$
# -\frac{\partial^{l-1}}{\partial t ^ {l-1}} \left( \frac{1}{t (t + \lambda_2)} \right) =
# \frac{(-1)^{l} (l-1)!}{\lambda_2} \left[\frac{1}{t^l} - \frac{1}{(t + \lambda_2)^l}\right] \\
# $$
# ## Proof by induction:
#
# - Base Case: $l=1$:
# $$
# LHS = -\frac{\partial^{1-1}}{\partial t ^ {1-1}} \left( \frac{1}{t (t + \lambda_2)} \right) = - \frac{1}{t (t + \lambda_2)} \\
# RHS = \frac{(-1)^{1} (1-1)!}{\lambda_2} \left[\frac{1}{t^1} - \frac{1}{(t + \lambda_2)^1}\right] =
# - \frac{1}{\lambda_2} \left[\frac{t + \lambda_2 - t}{t (t + \lambda_2)}\right] = - \frac{\lambda_2}{\lambda_2 t (t + \lambda_2)} = - \frac{1}{t (t + \lambda_2)}
# $$
#
#
# - Assume true for $l=x$:
# $$
# -\frac{\partial^{x-1}}{\partial t ^ {x-1}} \left( \frac{1}{t (t + \lambda_2)} \right) =
# \frac{(-1)^{x} (x-1)!}{\lambda_2} \left[\frac{1}{t^x} - \frac{1}{(t + \lambda_2)^x}\right] \\
# $$
#
#
# - Prove true for $l=x+1$:
#
# (Show that: $ -\frac{\partial^{x}}{\partial t^{x}} \left( \frac{1}{t (t + \lambda_2)} \right) = \frac{(-1)^{x+1} (x)!}{\lambda_2} \left( \frac{1}{t^{x+1}} - \frac{1}{(t + \lambda_2)^{x+1}}\right)$)
#
# \begin{align}
# LHS &= -\frac{\partial^{x}}{\partial t^{x}} \left( \frac{1}{t (t + \lambda_2)} \right) \\
# &= \frac{\partial}{\partial t} \left[ -\frac{\partial^{x-1}}{\partial t^{x-1}} \left( \frac{1}{t (t + \lambda_2)} \right) \right] \\
# &= \frac{\partial}{\partial t} \left[ \frac{(-1)^{x} (x-1)!}{\lambda_2} \left( \frac{1}{t^x} - \frac{1}{(t + \lambda_2)^x}\right) \right] \\
# &= \frac{(-1)^{x} (x-1)!}{\lambda_2} \frac{\partial}{\partial t} \left( \frac{1}{t^x} - \frac{1}{(t + \lambda_2)^x}\right) \\
# &= \frac{(-1)^{x} (x-1)!}{\lambda_2} \left( \frac{-x}{t^{x+1}} - \frac{-x}{(t + \lambda_2)^{x+1}}\right) \\
# &= \frac{(-1)^{x} (x-1)!}{\lambda_2} (-x) \left( \frac{1}{t^{x+1}} - \frac{1}{(t + \lambda_2)^{x+1}}\right) \\
# &= \frac{(-1)^{x+1} (x)!}{\lambda_2} \left( \frac{1}{t^{x+1}} - \frac{1}{(t + \lambda_2)^{x+1}}\right) \\
# &= RHS \\
# & \hspace{8cm} \square
# \end{align}
# - # Proof $k=1$ case by induction:
# $$
# - \frac{\partial^{l-1}}{\partial t ^ {l-1}} \left( \frac{1}{t (t + \lambda_2)} \right) = \frac{(-1)^{l} (l-1)!}{t^l (t + \lambda_2)^l}
# \times \sum_{j=1}^{l} {l \choose j} t^{l - j} \lambda_2 ^ {j - 1}
# $$
#
# - Base Case: $l = 1$
# $$
# LHS = - \frac{\partial^{1-1}}{\partial t ^ {1-1}} \left( \frac{1}{t (t + \lambda_2)} \right) = \frac{-1}{t(t + \lambda_2)} \\
# RHS = \frac{(-1)^1 (1-1)!}{t(t+\lambda_2)} \times {1 \choose 1} t^0 \lambda_2^0 = \frac{-1}{t(t + \lambda_2)}
# $$
# - Assume true for $l = x$:
# $$
# - \frac{\partial^{x-1}}{\partial t ^ {x-1}} \left( \frac{1}{t (t + \lambda_2)} \right) = \frac{(-1)^{x} (x-1)!}{t^x (t + \lambda_2)^x} \times \sum_{j=1}^{x} {x \choose j} t^{x - j} \lambda_2 ^ {j - 1}
# $$
# - Prove true for $l = x + 1$:
# $$
# - \frac{\partial^{x}}{\partial t ^ {x}} \left( \frac{1}{t (t + \lambda_2)} \right) = \frac{\partial}{\partial t} \left[ \frac{\partial^{x-1}}{\partial t ^ {x-1}} \left( \frac{1}{t (t + \lambda_2)} \right) \right]
# = \frac{(-1)^{x+1} x!}{t^{x+1} (t + \lambda_2)^{x+1}} \times \sum_{j=1}^{x+1} \, {{x + 1} \choose j} \, t^{x+1-j} \, \lambda_2^{j-1}
# $$
# - $l = x+1$:
# $$
# - \frac{\partial^{x}}{\partial t ^ {x}} \left( \frac{1}{t (t + \lambda_2)} \right) = \frac{\partial}{\partial t} \left[ \frac{\partial^{x-1}}{\partial t ^ {x-1}} \left( \frac{1}{t (t + \lambda_2)} \right) \right] \\
# = \frac{\partial}{\partial t} \left[ \frac{(-1)^{x} (x-1)!}{t^x (t + \lambda_2)^x} \times \sum_{j=1}^{x} {x \choose j} t^{x - j} \lambda_2 ^ {j - 1} \right] \\
# = \frac{\partial}{\partial t} \left[ u \times v \right] \\
# = \frac{\partial u}{\partial t} v + u \frac{\partial v}{\partial t}
# $$
# ## Derivative of u
#
# \begin{align}
# \frac{\partial u}{\partial t} &= \frac{\partial}{\partial t} \left[ \frac{(-1)^{x} (x-1)!}{t^x (t + \lambda_2)^x} \right] \\
# &= (-1)^x (x-1)! (-x) t^{-x-1} (t + \lambda_2)^{-x} + (-1)^x (x-1)! t^{-x} (-x) (t + \lambda_2)^{-x+1} \\
# &= (-1)^{x+1} x! \frac{2t + \lambda_2}{t^{x+1} (t + \lambda_2)^{x+1}}
# \end{align}
#
# ## Derivative of v
#
# \begin{align}
# \frac{\partial v}{\partial t} &= \frac{\partial}{\partial t} \left[ \sum_{j=1}^{x} \, {x \choose j} \, t^{x - j} \, \lambda_2 ^ {j - 1} \right] \\
# &= \frac{\partial}{\partial t} \left[ \sum_{j=1}^{x} \, \frac{x!}{(x-j)! j!} \, t^{x - j} \, \lambda_2 ^ {j - 1} \right] \\
# &= \sum_{j=1}^{x} \, \frac{x! (x - j)}{(x-j)! \, j!} \, t^{x - j - 1} \, \lambda_2 ^ {j - 1} \\
# \end{align}
# ## Proof (cont'd)
#
# \begin{align}
# =& \frac{\partial u}{\partial t} v + u \frac{\partial v}{\partial t} \\
# =& \frac{(-1)^{x+1} x!}{t^{x+1} (t + \lambda_2)^{x+1}} (2t + \lambda_2) \sum_{j=1}^{x} \frac{x!}{(x-j)! j!} t^{x-j} \lambda_2^{j - 1} \\
# & \qquad \qquad + \frac{(-1)^{x} (x-1)!}{t^x(t + \lambda_2)^{x}} \sum_{j=1}^x \frac{x!}{(x-j)! j!} t^{x-j-1} \lambda_2^{j-1} \\
# =& \frac{(-1)^{x+1} \, x!}{t^{x+1} (t + \lambda_2)^{x+1}} \left[ (2t+\lambda_2) \, \sum_{j=1}^x \, \frac{x!}{(x-j)! \, j!} \, t^{x-j} \, \lambda_2^{j-1}
# - \frac{t(t+\lambda_2)}{x} \, \sum_{j=1}^{x} \, \frac{x!(x-j)}{(x-j)! j!} \, t^{x-j-1} \, \lambda_2^{j-1} \right]
# \end{align}
# #### ONLY LEFT TO SHOW THAT
#
# $$
# (2t+\lambda_2) \, \sum_{j=1}^x \, \frac{x!}{(x-j)! \, j!} \, t^{x-j} \, \lambda_2^{j-1} - \frac{t(t+\lambda_2)}{x} \, \sum_{j=1}^{x} \, \frac{x!(x-j)}{(x-j)! j!} \, t^{x-j-1} \, \lambda_2^{j-1} = \sum_{j=1}^{x+1} \, {{x + 1} \choose j} \, t^{x+1-j} \, \lambda_2^{j-1}
# $$
# +
def LHS_demo(x, t, lambda_2):
sum_1 = sum((math.factorial(x) * t ** (x - j) * lambda_2 ** (j - 1)) / (math.factorial(x - j) * math.factorial(j)) for j in range(1, x+1))
sum_2 = sum((math.factorial(x) * (x - j) * t ** (x - j - 1) * lambda_2 ** (j - 1)) / (math.factorial(x - j) * math.factorial(j)) for j in range(1, x+1))
sum_1 *= (2 * t + lambda_2)
sum_2 *= (-t * (t + lambda_2)) / x
return sum_1 + sum_2
def RHS_demo(x, t, lambda_2):
sum_1 = sum((math.factorial(x + 1) * t ** (x + 1 - j) * lambda_2 ** (j - 1)) / (math.factorial(x + 1 - j) * math.factorial(j)) for j in range(1, x+2))
return sum_1
# -
sym.expand(LHS_demo(x=10, t=t, lambda_2=lambda_2))
sym.expand(RHS_demo(x=10, t=t, lambda_2=lambda_2))
for x in range(1,50):
are_equal = sym.expand(LHS_demo(x=x, t=t, lambda_2=lambda_2)) == sym.expand(RHS_demo(x=x, t=t, lambda_2=lambda_2))
print(are_equal, x)
# ## Attempt 1:
#
# $$
# \sum_{j=1}^{x} \frac{x!}{(x-j)! j!} t^{x-j} \lambda_2^{j-1} = \left[ \sum_{j=1}^{x+1} \frac{x! j}{(x-j+1)! j!} t^{x-j+1} \lambda_2^{j-2} \right] - \frac{t^x}{\lambda_2}
# $$
def RHS_new_demo(x, t, lambda_2):
sum_1 = sum((math.factorial(x + 1) * t ** (x + 1 - j) * lambda_2 ** (j - 1)) / (math.factorial(x + 1 - j) * math.factorial(j)) for j in range(1, x+2))
return sum_1
# +
def LHS_new_demo(x, t, lambda_2):
sum_1 = sum((math.factorial(x) * t ** (x - j) * lambda_2 ** (j - 1)) / (math.factorial(x - j) * math.factorial(j)) for j in range(1, x+1))
return sum_1
def RHS_new_demo(x, t, lambda_2):
sum_1 = 0
for j in range(1, x + 2):
to_add = j*(math.factorial(x) * t ** (x + 1 - j) * lambda_2 ** (j - 2)) / (math.factorial(x + 1 - j) * math.factorial(j))
sum_1 += to_add
sum_1 -= t ** x / lambda_2
return sum_1
# -
sym.simplify(LHS_new_demo(5, t, lambda_2))
sym.expand(RHS_new_demo(5, t, lambda_2))
# ## Attempt 2:
#
# $$
#
#
# $$
# +
def LHS_demo_3(x, t, lambda_2):
sum_1 = sum((math.factorial(x + 1) * t ** (x + 1 - j) * lambda_2 ** (j - 1)) / (math.factorial(x + 1 - j) * math.factorial(j)) for j in range(1, x+2))
return sum_1
def RHS_demo_3(x, t, lambda_2):
sum_1 = 0
for j in range(1, x + 1):
to_add = (math.factorial(x) * t ** (x - j) * lambda_2 ** (j - 1) * (2 * t + lambda_2)) / (math.factorial(x - j) * math.factorial(j))
to_add -= (math.factorial(x) * (x - j) * t ** (x - j - 1) * lambda_2 ** (j - 1) * (t ** 2 + t * lambda_2) / x) / (math.factorial(x - j) * math.factorial(j))
sum_1 += to_add
return sum_1
def RHS_demo_4(x, t, lambda_2):
sum_1 = sum((math.factorial(x - 1) * t ** (x - j) * lambda_2 ** (j - 1)) * (x * t + j * t + lambda_2 * j) / (math.factorial(x - j) * math.factorial(j)) for j in range(1, x+1))
return sum_1
# -
sym.simplify(LHS_demo_3(7, t, lambda_2))
sym.simplify(RHS_demo_4(7, t, lambda_2))
# ## Attempt 3:
#
# $$
# - \frac{t^{x+1}}{\lambda_2} + \sum_{j=1}^{x+1} \, \frac{x!}{(x-j+1)! j!} \, t^{x+1-j} \, \lambda_2^{j-1} \left( \frac{xjt + tj^2 -tj +\lambda_2 j^2 - \lambda_2 j}{x \lambda_2} \right)= \sum_{j=1}^{x+1} \, {{x + 1} \choose j} \, t^{x+1-j} \, \lambda_2^{j-1}
# $$
# +
def LHS_demo_5(x, t, lambda_2):
sum_1 = sum((math.factorial(x + 1) * t ** (x + 1 - j) * lambda_2 ** (j - 1)) / (math.factorial(x + 1 - j) * math.factorial(j)) for j in range(1, x+2))
return sum_1
def RHS_demo_5(x, t, lambda_2):
sum_1 = sum((math.factorial(x) * t ** (x + 1 - j) * lambda_2 ** (j - 1) * (x*j*t + t*j**2 - t*j + lambda_2 * j**2 - lambda_2 * j)) / (x * lambda_2 * math.factorial(x + 1 - j) * math.factorial(j)) for j in range(2, x+2))
return sum_1 #- t ** (x+1) / lambda_2
# def RHS_demo_5(x, t, lambda_2):
# sum_1 = sum((math.factorial(x) * t ** (x + 1 - j) * lambda_2 ** (j - 1)) / (math.factorial(x - j) * math.factorial(j)) for j in range(1, x+1))
# sum_2 = sum((math.factorial(x) * j * t ** (x + 1 - j) * lambda_2 ** (j - 1)) / (math.factorial(x - j) * math.factorial(j)) for j in range(1, x+1))
# return sum_1 + sum_2 * (t + lambda_2) / (t * x)
# -
sym.expand(LHS_demo_5(7, t, lambda_2))
sym.expand(RHS_demo_5(7, t, lambda_2))
# ## Attempt 4
#
# $$
#
# \frac{xjt + tj^2 - tj + \lambda_2 j^2 - \lambda_2 j}{x \lambda_2 (x + 1)} - \frac{(x-j+1)! j!}{(x+1)!} \frac{t^j}{\lambda_2^j} = 1
#
# $$
from sympy.printing.latex import LatexPrinter, print_latex
# +
def LHS_demo_5(x, t, lambda_2):
sum_1 = 0
for j in range(1, x+2):
term = (math.factorial(x + 1) * t ** (x + 1 - j) * lambda_2 ** (j - 1)) / (math.factorial(x + 1 - j) * math.factorial(j))
print(term)
sum_1 += term
return sum_1
def RHS_demo_5(x, t, lambda_2):
sum_1 = 0
for j in range(1, x+2):
extra_term = (x*j*t + t*j**2 - t*j + lambda_2 * j**2 - lambda_2 * j) / (x * lambda_2)
extra_term -= (math.factorial(x - j + 1) * math.factorial(j) * t ** (x + 1)) / (math.factorial(x+1) * t ** (x + 1 - j) * lambda_2 ** j)
extra_term /= (x + 1)
term = (math.factorial(x + 1) * t ** (x + 1 - j) * lambda_2 ** (j - 1) * extra_term) / (math.factorial(x + 1 - j) * math.factorial(j))
print(term)
sum_1 += term
return sum_1
# -
sym.expand(LHS_demo_5(7, t, lambda_2))
sym.expand(RHS_demo_5(7, t, lambda_2))
# # Going back:
# ## Assuming true for $l=x$ prove true for $l=x+1$
# \begin{align}
# & - \frac{\partial^{x}}{\partial t ^ {x}} \left( \frac{1}{t (t + \lambda_2)} \right) \\
# & = \frac{\partial}{\partial t} \left[ \frac{\partial^{x-1}}{\partial t ^ {x-1}} \left( \frac{1}{t (t + \lambda_2)} \right) \right] \\
# & = \frac{\partial}{\partial t} \left[ \frac{(-1)^{x} (x-1)!}{t^x (t + \lambda_2)^x} \times \sum_{j=1}^{x} {x \choose j} t^{x - j} \lambda_2 ^ {j - 1} \right] \\
# & = \frac{\partial}{\partial t} \left[ \sum_{j=1}^{x} \frac{(-1)^{x} (x-1)!}{t^x (t + \lambda_2)^x} \times {x \choose j} t^{x - j} \lambda_2 ^ {j - 1} \right] \\
# & = \sum_{j=1}^{x} \left[ \frac{\partial}{\partial t} \frac{(-1)^{x} (x-1)!}{(t + \lambda_2)^x} \times {x \choose j} t^{- j} \lambda_2 ^ {j - 1} \right] \\
# & = \sum_{j=1}^{x} \left[ (-1)^{x} (x-1)! {x \choose j} \lambda_2 ^ {j - 1} \frac{\partial}{\partial t} \left( \frac{t^{- j}}{(t + \lambda_2)^x} \right) \right] \\
# & = \sum_{j=1}^{x} \left[ (-1)^{x} (x-1)! {x \choose j} \lambda_2 ^ {j - 1} \left( \frac{(-1)(jt + j \lambda_2 + xt)}{t^{j+1}(t + \lambda_2)^{x+1}} \right) \right] \\
# & = \frac{(-1)^{x+1} (x-1)!}{(t + \lambda_2)^{x+1}} \sum_{j=1}^{x} \left[ {x \choose j} \lambda_2 ^ {j - 1} t^{-j-1} (jt + j \lambda_2 + xt) \right] \\
# \end{align}
# +
def LHS_demo(x, t, lambda_2):
sum_1 = sum((math.factorial(x) * j * t ** (- j - 1) * lambda_2 ** (j - 1)) / (math.factorial(x - j) * math.factorial(j)) for j in range(1, x+1))
sum_2 = sum((math.factorial(x) * t ** (- j) * lambda_2 ** (j - 1)) / (math.factorial(x - j) * math.factorial(j)) for j in range(1, x+1))
sum_1 *= (-1) ** (x+1) * math.factorial(x - 1) / (t + lambda_2) ** x
sum_2 *= (-1) ** (x+1) * math.factorial(x) / (t + lambda_2) ** (x + 1)
return sum_1 + sum_2
def LHS_demo(x, t, lambda_2):
sum_1 = 0
for j in range(1, x+2):
term = math.factorial(x) * t ** (- j) * lambda_2 ** (j - 2) * ((j - 1) * (t + lambda_2) + x*t) / (math.factorial(x + 1 - j) * math.factorial(j - 1))
# if j > 1:
print(term)
sum_1 += term
sum_1 -= x / lambda_2
sum_1 *= (-1) ** (x+1) * math.factorial(x - 1) / (t + lambda_2) ** (x + 1)
return sum_1
def RHS_demo(x, t, lambda_2):
sum_1 = 0
for j in range(1, x+2):
term = (math.factorial(x + 1) * t ** (x + 1 - j) * lambda_2 ** (j - 1)) / (math.factorial(x + 1 - j) * math.factorial(j))
print(term)
sum_1 += term
sum_1 *= ((-1)**(x+1) * math.factorial(x)) / (t ** (x+1) * (t + lambda_2) ** (x + 1))
return sum_1
# -
sym.factor(RHS_demo(2, t, lambda_2))
sym.factor(LHS_demo(2, t, lambda_2))
# $$
# \frac{(-1)^{x+1} (x-1)!}{(t + \lambda_2)^{x+1}} \sum_{j=1}^{x} \left[ {x \choose j} \lambda_2 ^ {j - 1} t^{-j-1} (jt + j \lambda_2 + xt) \right] = \frac{(-1)^{x+1} x!}{(t + \lambda_2)^{x+1}} \times \sum_{j=1}^{x+1} \, {{x + 1} \choose j} \, t^{-j} \, \lambda_2^{j-1} \\
# \frac{(-1)^{x+1} x!}{(t + \lambda_2)^{x+1}} \times \sum_{j=1}^{x+1} \, {{x + 1} \choose j} \, t^{-j} \, \lambda_2^{j-1} - \frac{(-1)^{x+1} (x-1)!}{(t + \lambda_2)^{x+1}} \sum_{j=1}^{x} \left[ {x \choose j} t^{-j-1} \lambda_2 ^ {j - 1} (jt + j \lambda_2 + xt) \right] = 0 \\
#
# \vdots \\
# \vdots \\
#
# \sum_{j=1}^{x} \left[ \frac{(x-1)! \, t^{-j} \, \lambda_2^{j-1} \, (j-1)}{(x+1-j)! \, (j-1)!} - \frac{(x-1)! \, t^{-j-1} \lambda_2^j}{(x-j)! (j-1)!} \right] + \frac{\lambda_2^x}{t^{x+1}} = 0 \\
#
# \vdots \\
# \vdots \\
#
# (t + \lambda_2) \sum_{j=2}^{x} j \, \frac{(x-1)! \, t^{-j-1} \, \lambda_2^{j-1}}{(x+1-j)! \, (j-1)!} - (t + \lambda_2 + \lambda_2 x) \sum_{j=2}^{x} \, \frac{(x-1)! \, t^{-j-1} \, \lambda_2^{j-1}}{(x+1-j)! \, (j-1)!} = \frac{\lambda_2}{t^2} - \frac{\lambda_2^x}{t^{x+1}}
# $$
# +
def LHS_demo(x, t, lambda_2):
sum_1 = 0
for j in range(1, x+2):
term = (math.factorial(x + 1) * t ** (- j) * lambda_2 ** (j - 1)) / (math.factorial(x + 1 - j) * math.factorial(j))
sum_1 += term
sum_1 *= ((-1)**(x+1) * math.factorial(x)) / ((t + lambda_2) ** (x + 1))
return sum_1
def RHS_demo(x, t, lambda_2):
sum_1 = 0
for j in range(1, x+1):
term = (math.factorial(x) * t ** (- j - 1) * lambda_2 ** (j - 1) * (j*t + j*lambda_2 + x*t)) / (math.factorial(x - j) * math.factorial(j))
sum_1 += term
sum_1 *= ((-1)**(x+1) * math.factorial(x - 1)) / ((t + lambda_2) ** (x + 1))
return sum_1
# +
def diff_demo_2(x, t, lambda_2):
summ = 0
for j in range(2, x+1):
summ += (math.factorial(x - 1) * t ** (-j) * lambda_2 ** (j - 1)) / (math.factorial(x + 1 - j) * math.factorial(j - 2))
summ -= math.factorial(x-1) * (t ** (- j - 1) * lambda_2 ** j) / (math.factorial(x-j) * math.factorial(j - 1))
summ -= lambda_2 / (t ** 2)
summ += lambda_2 ** x / (t ** (x + 1))
return summ
def diff_demo(x, t, lambda_2):
return LHS_demo(x, t, lambda_2) - RHS_demo(x, t, lambda_2)
def diff_demo(x, t, lambda_2):
summ = 0
for j in range(1, x+1):
term = math.factorial(x - 1) * t ** (-j - 1) * lambda_2 ** (j - 1)
# term *= (x*(x+1)*t - (x+1-j)*(j*(t+lambda_2) + t*x))
term *= (j*t*(j-1) - j*lambda_2*(x+1-j))
term /= (math.factorial(x + 1 - j) * math.factorial(j))
summ += term
summ += lambda_2 ** x / (t ** (x + 1))
return summ
def diff_demo(x, t, lambda_2):
summ = 0
for j in range(1, x+1):
term_1 = math.factorial(x - 1) * t ** (-j) * lambda_2 ** (j - 1)
term_1 *= (j-1)
term_1 /= (math.factorial(x + 1 - j) * math.factorial(j - 1))
term_2 = math.factorial(x - 1) * t ** (-j - 1) * lambda_2 ** j
term_2 *= 1
term_2 /= (math.factorial(x - j) * math.factorial(j - 1))
summ += term_1 - term_2
summ += lambda_2 ** x / (t ** (x + 1))
return summ
# -
sym.expand(diff_demo(x=5, t=t, lambda_2=lambda_2))
sym.expand(diff_demo_2(x=10, t=t, lambda_2=lambda_2))
# ## Attempt Using Taylor Series
#
# $$
# \frac{-1}{t (t + \lambda_2)} = \sum_{n=l-1}^{\infty} \left[ \frac{-1}{t_0 (t_0 + \lambda_2)} \right]^{(n)} \frac{(t - t_0)^n}{n!}
# $$
#
# $$
# \begin{align}
# \Psi_{1,l}(t) & = \frac{\partial^{l-1}}{\partial t ^ {l-1}} \left( \frac{-1}{t (t + \lambda_2)} \right) \\
# & = \frac{\partial^{l-1}}{\partial t ^ {l-1}} \left( \sum_{n=0}^{\infty} \left[ \frac{-1}{t_0 (t_0 + \lambda_2)} \right]^{(n)} \frac{(t - t_0)^n}{n!} \right) \\
# & = \frac{\partial^{l-1}}{\partial t ^ {l-1}} \left( \sum_{n=l-1}^{\infty} \left[ \frac{-1}{t_0 (t_0 + \lambda_2)} \right]^{(n)} \frac{(t - t_0)^n}{n!} \right) \\
# \end{align}
# $$
#
# +
lambda_1 = sym.Symbol("lambda_1")
lambda_2 = sym.Symbol("lambda_2")
r_2 = sym.Symbol("r_2")
t = sym.Symbol("t")
exp_rates = [sym.S(0), lambda_1, lambda_2]
freq = [1, r_2, 1]
a = len(exp_rates) - 1
# -
t = 3
lambda_2 = 5
# # To prove:
# $$
# - \frac{\partial^{l-1}}{\partial t ^ {l-1}} \left( \frac{1}{t (t + \lambda_2)} \right) = \frac{(-1)^{l} (l-1)!}{(t + \lambda_2)^{l}} \times \sum_{j=1}^{l} \, {l \choose j} \, t^{-j} \, \lambda_2^{j-1}
# $$
# # Attempt by computing the $n^{th}$ derivative
# ### Leibniz formula for $n^{th}$ derivative:
# $$
# (u(x)v(x))^{(n)} = \sum_{i=1}^{n} {n \choose i} u^{i} v^{n-i}
# $$
# +
def RHS_demo(arg, k, l, exp_rates, freq, a):
psi_val = 0
for term in range(1, l + 1):
psi_val += math.comb(l, term) * arg ** (- term) * exp_rates[2] ** (term - 1)
psi_val *= (-1) ** l * math.factorial(l - 1) / ((arg + exp_rates[2]) ** l)
return psi_val
def LHS_demo(arg, k, l, exp_rates, freq, a):
psi_val = 0
for term in range(1, l + 1):
psi_val += (arg + exp_rates[2]) ** (term - 1) / (arg ** (term))
psi_val *= (-1) ** l * math.factorial(l - 1) / ((arg + exp_rates[2]) ** l)
return psi_val
# def LHS_demo(arg, k, l, exp_rates, freq, a):
# psi_val = 0
# for term in range(1, l + 1):
# for r in range(term):
# psi_val += math.comb(term-1, r) * arg ** (-r-1) * exp_rates[2] ** r
# psi_val *= (-1) ** l * math.factorial(l - 1) / ((arg + exp_rates[2]) ** l)
# return psi_val
# -
sym.factor(RHS_demo(arg=t, k=1, l=5, exp_rates=[sym.S(0), lambda_1, lambda_2], freq=[1, r_2, 1], a=2))
sym.factor(LHS_demo(arg=t, k=1, l=5, exp_rates=[sym.S(0), lambda_1, lambda_2], freq=[1, r_2, 1], a=2))
# ## More specific
# +
def RHS_demo(t, l, lambda_2):
summ = 0
for i in range(1, l + 1):
summ += math.comb(l, i) * t ** (-i) * lambda_2 ** (i-1)
return summ
def LHS_demo(t, l, lambda_2):
summ = 0
for i in range(1, l + 1):
summ += (t + lambda_2) ** (i - 1) / (t ** i)
return summ
def LHS_demo(t, l, lambda_2):
return ((t + lambda_2) ** l - t ** l) / (t**l * lambda_2)
# -
sym.factor(RHS_demo(t, 4, lambda_2))
sym.factor(LHS_demo(t, 4, lambda_2))
| nbs/src/Markov/markov-proportion-formula/proof of hypoexponential closed-form.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-2:429704687514:image/datascience-1.0
# ---
# # Data exploration/visualization
#
# **SageMaker Studio Kernel**: Data Science
#
# The challenge we're trying to address here is to detect anomalies in the components of a Wind Turbine. Each wind turbine has many sensors that reads data like:
# - Internal & external temperature
# - Wind speed
# - Rotor speed
# - Air pressure
# - Voltage (or current) in the generator
# - Vibration in the GearBox (using an IMU -> Accelerometer + Gyroscope)
#
# So, depending on the types of the anomalies we want to detect, we need to select one or more features and then prepare a dataset that 'explains' the anomalies. We are interested in three types of anomalies:
# - Rotor speed (when the rotor is not in an expected speed)
# - Produced voltage (when the generator is not producing the expected voltage)
# - Gearbox vibration (when the vibration of the gearbox is far from the expected)
#
# All these three anomalies (or violations) depend on many variables while the turbine is working. Thus, in order to address that, let's use a ML model called [Autoencoder](https://en.wikipedia.org/wiki/Autoencoder), with correlated features. This model is unsupervised. It learns the latent representation of the dataset and tries to predict (regression) the same tensor given as input. The strategy then is to use a dataset collected from a normal turbine (without anomalies). The model will then learn **'what is a normal turbine'**. When the sensors readings of a malfunctioning turbine is used as input, the model will not be able to rebuild the input, predicting something with a high error and detected as an anomaly.
#
# The sequence of the sensors readings can be seen as a time-series dataset and therefore we observe a high correlation between neighbour samples. We can explore this by reformatting the data as a multidimensional tensor. We'll create a temporal encoding of six features in 10x10 steps of 250ms each. 250ms is the interval computed using 5 samples (the time interval between each sample is ~50ms). It means that we will create a tensor with a shape of 6x10x10.
#
# 
#
# In the tensor above, each color is a different feature, encoded in 100 (10x10) timesteps (from the current reading to the past in a sliding window).
#
# Let's start preparing our dataset, then.
# ### Install this lib to improve data visualization
# !pip install -U matplotlib==3.4.1 seaborn==0.11.1
# ### And download the sensors (raw) data
# This dataset was created by mini wind turbines, 3D printed and assembled for experimenting with ML@Edge. If you're interested on building your own 3D printed mini wind turbines, please check the link to this project in the home of this workshop.
# !mkdir -p ../data
# !curl https://aws-ml-blog.s3.amazonaws.com/artifacts/monitor-manage-anomaly-detection-model-wind-turbine-fleet-sagemaker-neo/dataset_wind_turbine.csv.gz -o ../data/dataset_wind.csv.gz
# ## Let's take a look on the data
# Loading the dataset using Pandas...
# %matplotlib inline
# %config InlineBackend.figure_format='retina'
import pandas as pd
import matplotlib.pyplot as plt
from datetime import datetime
## preprocessing is the data preparation script we'll use in our automated ML Pipeline
## here, it will be just a loaded library
import preprocessing as dataprep
# +
parser = lambda date: datetime.strptime(date, '%Y-%m-%dT%H:%M:%S.%f+00:00')
df = pd.read_csv('../data/dataset_wind.csv.gz', compression="gzip", sep=',', low_memory=False, parse_dates=[ 'eventTime'], date_parser=parser)
df.head()
# -
# Features:
# - **nanoId**: id of the edge device that collected the data
# - **turbineId**: id of the turbine that produced this data
# - **arduino_timestamp**: timestamp of the arduino that was operating this turbine
# - **nanoFreemem**: amount of free memory in bytes
# - **eventTime**: timestamp of the row
# - **rps**: rotation of the rotor in Rotations Per Second
# - **voltage**: voltage produced by the generator in milivolts
# - **qw, qx, qy, qz**: quaternion angular acceleration
# - **gx, gy, gz**: gravity acceleration
# - **ax, ay, az**: linear acceleration
# - **gearboxtemp**: internal temperature
# - **ambtemp**: external temperature
# - **humidity**: air humidity
# - **pressure**: air pressure
# - **gas**: air quality
# - **wind_speed_rps**: wind speed in Rotations Per Second
## we will select the following features to prepare our dataset
## with these features we have parameters for vibration, rotation and voltage
quat=['qx', 'qy', 'qz', 'qw']
rot=['wind_speed_rps', 'rps']
volt=['voltage']
features = quat + rot + volt
# ### Ploting the vibration data, just to have an idea
df[quat[:3]].iloc[1910:2000].plot(figsize=(20,10))
# ### Now, plot the rotation of the turbine and the wind speed in RPS
df[rot].iloc[1910:2000].plot(figsize=(20,10))
# ### Finally, plot the voltage readings
df[volt].iloc[1910:2000].plot(figsize=(20,10))
# ## Data preparation
# The raw data for rotation is formated as angular acceleration using a Quaternion representation. We can convert it to Euler angles to make it easier to understand.
print('now converting quat to euler...')
roll,pitch,yaw = [], [], []
for idx, row in df.iterrows():
r,p,y = dataprep.euler_from_quaternion(row['qx'], row['qy'], row['qz'], row['qw'])
roll.append(r)
pitch.append(p)
yaw.append(y)
df['roll'] = roll
df['pitch'] = pitch
df['yaw'] = yaw
# ## Then, we can denoise and normalize the data to complete the process
# +
df_train = df.copy()
# select the features
features = ['roll', 'pitch', 'yaw', 'wind_speed_rps', 'rps', 'voltage']
# get the std for denoising
raw_std = df_train[features].std()
for f in features:
df_train[f] = dataprep.wavelet_denoise(df_train[f].values, 'db6', raw_std[f])#[:-1]
# normalize
training_std = df_train[features].std()
training_mean = df_train[features].mean()
df_train = (df_train[features] - training_mean) / training_std
df_train.head()
# -
# ### Alright, this is our dataset. Let's just plot the original vs the prepared data
# **Original Data**
df[features][:2000].plot(figsize=(20,10))
# **Denoised & Normalized Data**
df_train[:2000].plot(figsize=(20,10))
# > There is too much noise in the raw data, specially in the accelerometer + gyroscope readings
# > This process is important to remove the impurity and make the model more efficient
# +
import seaborn as sns
corr = df[features].corr()
fig, ax = plt.subplots(figsize=(15, 8))
sns.heatmap(corr, annot=True, fmt="f",
xticklabels=corr.columns.values,
yticklabels=corr.columns.values,
ax=ax)
# -
# As expected, the linear correlation between **rps (rotation speed)** and **voltage** is high. We need to keep both, given the model needs to understand what is a normal relationship between these two + other features.
#
# Alright! Now you can start exercise #2: create a ML pipeline to train your model and then deploy it to the edge devices.
#
# > [Exercise 02](../02-Training/02-Training-with-Pytorch.ipynb)
| lab/01-Data-Visualization/01-Data-Visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# +
import numpy as np
import k3d
vector_length = 0.3
x, y, z = np.meshgrid(np.arange(-0.8, 1, 0.2),
np.arange(-0.8, 1, 0.2),
np.arange(-0.8, 1, 0.8))
u = np.sin(np.pi * x) * np.cos(np.pi * y) * np.cos(np.pi * z)
v = -np.cos(np.pi * x) * np.sin(np.pi * y) * np.cos(np.pi * z)
w = (np.sqrt(2.0 / 3.0) * np.cos(np.pi * x) * np.cos(np.pi * y) *
np.sin(np.pi * z))
plot = k3d.plot()
vectors = k3d.vectors(origins=np.transpose([x,y,z]).astype(np.float32),
vectors=np.transpose([u,v,w]).astype(np.float32)*vector_length, head_size=0.6)
plot += vectors
plot.display()
# -
| K3D+MPL/quiver3d_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from jupyterthemes import get_themes
from jupyterthemes.stylefx import set_nb_theme
themes = get_themes()
set_nb_theme(themes[1])
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# 1. magic for inline plot
# 2. magic to print version
# 3. magic so that the notebook will reload external python modules
# %matplotlib inline
# %load_ext watermark
# %load_ext autoreload
# %autoreload 2
from sklearn.decomposition import PCA
from sklearn.pipeline import Pipeline
from sklearn.datasets import load_iris
from sklearn.metrics import accuracy_score
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split
# %watermark -a 'Ethen' -d -t -v -p numpy,pandas,matplotlib,sklearn
# -
# # Dimensionality Reduction
#
# The starting point most data analysis problems is to perform some sort of exploratory data analysis on our raw data. This step is important because until we have basic understanding of the structure of our raw data, it might be hard to know whether the data is suitable for the task at hand or even derive insights that we can later share. During this exploratoy process, unsupervised methods such as dimensionality reduction can help us identify simpler and more compact representations of the original raw data to either aid our understanding or provide useful input to other stages of analysis. Here, we'll be focusing on a specific dimensionality reduction technique called **Principal Component Analysis (PCA)**.
#
# Imagine that we're trying to understand some underlying phenomenon, in order to do so we measure various quantities potentially related to it. If we knew exactly what to measure in advance, we might be able to find some simple relationships in our data. But we typically don't, so we often measure anything that might be relevant and end up having irrelevant or redundant signals in our measurement. To make this a bit more concrete, we'll generate a toy 2-dimensional dataset to work with using the equation below.
#
# $$x_2 = 500 + 20 \times x_1 + \epsilon$$
# +
np.random.seed(123)
x1 = np.arange(12, 56, 0.5)
e = np.random.normal(0, 100, x1.shape[0])
x2 = 500 + 20 * x1 + e
X = np.c_[x1, x2]
# change default style figure and font size
plt.style.use('fivethirtyeight')
plt.rcParams['figure.figsize'] = 8, 6
plt.rcParams['font.size'] = 12
def plot2var(m):
plt.scatter(m[:, 0], m[:, 1], s = 40, alpha = 0.8)
plt.xlabel('x1')
plt.ylabel('x2')
plot2var(X)
# -
# We can clearly see from the plot that there's linear relationship between the two variables, thus we probably don't need to include both of them as $x_1$ can be easily explained by $x_2$ and vice versa.
#
# ## PCA
#
# If it is the case that one of the features is considered redundant, we should be able to summarize the data with less characteristics (features). So, the way PCA tackles this problem is: Instead of simply picking out the useful features and discarding the others, it uses a linear combination of the existing features and constructs some new features that are good alternative representation of the original data. In our 2D toy dataset, PCA will try to pick the best single direction, or often referred to as **first principal components** in 2D, and project our points onto that single direction. So the next question becomes, out of the many possible lines in 2D, what line should we pick?
#
# It turns out, there are two different answers to this question. First answer is that we are looking for some features that strongly differ across data points, thus, PCA looks for features that captures as much variation across data points as possible. The second answer is that we are looking for the features that would allow us to "reconstruct" the original features. Imagine that we come up with a feature that has no relation to the original ones; If we were to use this new feature, there is no way we can relate this to the original ones. So PCA looks for features that minimizes the reconstruction error. These two notions can be depicted in the graph below, where the black dots represents the original data point, the black line represents the projected line, the red dot on the left shows the points on the projected line and the red line on the right shows the reconstruction error.
#
# <img src='img/objective1.png' width='80%'>
#
# Surprisingly, it turns out that these two aims are equivalent and PCA can kill two birds with one stone. To see why minimizing squared residuals is equivalent to maximizing variance consider the 2 dimensions visualization below.
#
# <img src='img/objective2.png' width='80%'>
#
# Consider a datapoint $a_i$. The contribution of this specific data point to the total variance is $a_i^Ta_i$, or equivalently the squared Euclidean length $\lVert \mathbf{a}_i \lVert^2$. Applying the Pythagorean theorem shows that this total variance equals the sum of variance lost (the squared residual) and variance remaining. Thus, it is equivalent to either maximize remaining variance or minimize lost variance to find the principal components.
#
# Before we go another deeper, let's build some intuition using the scikit-learn library. The following section standardizes the data, fits the PCA model and prints out some of the important informations.
# +
# we start by standardizing our dataset
X_std = StandardScaler().fit_transform(X)
# call PCA specifying we only want the
# first two principal components (since
# we only have a 2d datset)
pca = PCA(n_components = 2)
pca.fit(X_std)
# important information
print('Components:\n ', pca.components_)
print('Explained Variance Ratio:\n ', pca.explained_variance_ratio_)
# -
# After fitting the PCA on the dataset, the fit learns some quantities from the data, most importantly the "components", which is the principal components (the new direction that our data points will be projected upon) and "explained variance ratio", which corresponds to the percentage of variance explained by each of the principal components. To get a better sense of what these numbers mean, let's visualize them over our standardized input data.
# +
def draw_vector(v0, v1):
"""draw principal components as vectors"""
ax = plt.gca()
arrowprops = dict(arrowstyle = '->',
linewidth = 2, edgecolor = 'black',
shrinkA = 0, shrinkB = 0)
plt.annotate('', v1, v0, arrowprops = arrowprops)
# plot data
plt.scatter(X_std[:, 0], X_std[:, 1], s = 40, alpha = 0.8)
for length, vector in zip(pca.explained_variance_, pca.components_):
# the larger the explained variance, the longer the
# annotated line will be
v = vector * 2 * np.sqrt(length)
draw_vector(pca.mean_, pca.mean_ + v)
plt.axis('equal')
plt.show()
# -
# From the plot, we see that our PCA model gives us two principal components (the black line) and the length of the lines indicate of how important that principal component is in describing the variation of the original data. So if we were to choose 1 principal component to summarize our 2d dataset, it will be line that has the longest length (largest explained variance ratio). There're two things worth noting:
#
# - The number of principal components matches the total number of features
# - The first and second principal components are orthogonal to each other
#
# The reason for this is that principal components transform the data into a new set of dimensions, and these new dimensions have to be equal to the original amount of dimensions. And just like the original $x$, $y$ axis that we're used to, they have to be orthogonal to each other. Let's now reduce the dimension of our 2d dataset into 1d by transforming our data onto the most important principal component and plot it along with the original data.
# +
# dimensionality reduction, keeping only
# the first principal component
pca = PCA(n_components = 1)
X_pca = pca.fit_transform(X_std)
print("original shape: ", X.shape)
print("transformed shape:", X_pca.shape)
# inverse transform to obtain the projected data
# and compare with the original
X_new = pca.inverse_transform(X_pca)
plt.scatter(X_std[:, 0], X_std[:, 1], s = 40, alpha = 0.2)
plt.scatter(X_new[:, 0], X_new[:, 1], s = 40, alpha = 0.8)
plt.show()
# -
# In the plot above, the lighter points are the original data and the darker points are the projected version. Looking at the projections we see that the points projected onto the first principal component all seem close to their initial representations and at the same time, it also captures most of the variations in our original data points.
#
# Hopefully, this elucidate what a PCA dimensionality reduction is doing. In a nutshell, PCA aims to find the directions or so called the principal components of maximum variance in high-dimensional data and project it onto a smaller dimensional subspace while retaining most of the information. In our 2d example, the linear relationship between $x_1$ and $x_2$ is mostly preserved using only 1 feature instead of 2.
# ## PCA From Scratch
#
# Now that we have a high level understanding of what PCA is accomplishing, let's now formalize this with some notations. Recall that PCA is trying the find the direction such that the projection of the data on it has the highest variance. So given $X$, the centered data matrix, the projection, $Xw$ (dot product between the data point, $X$ and the projection weight, $w$), its variance can be computed as follows:
#
# \begin{align}
# \frac{1}{n-1}(\mathbf{Xw})^\top \mathbf{Xw} = \mathbf w^\top (\frac{1}{n-1}\mathbf X^\top\mathbf X) \mathbf w = \mathbf w^\top \mathbf{Cw}
# \end{align}
#
# Where $\mathbf{C}$ is the covariance matrix of the data $\mathbf{X}$. In case this looks unfamiliar, the covariance matrix is a $d \times d$ matrix ($d$ is the total number of dimensions, features) where each element represents the covariance between two features. The covariance between two features $x_j$ and $x_k$ is calculated as follows:
#
# \begin{align}
# \sigma_{jk} = \frac{1}{n-1} \sum_{i=1}^{n} \left( x_{ij} - \bar{x}_j \right) \left( x_{ik} - \bar{x}_k \right)
# \end{align}
#
# Where $\bar{x}_j$ is simply the mean of vector (feature) $\bar{x}_j = \sum \limits_{i=1}^n x_{ij}$.
#
# We can also express the calculation of the covariance matrix via the following matrix equation:
#
# \begin{align}
# \mathbf{C} = \frac{1}{n-1} \big( (X - \bar{x})^\top( X - \bar{x}) \big)
# \end{align}
#
# Because we already assume to data point $X$ has been centered, thus the $X - \bar{x}$ part can be simplified into just $X$. And the formula above simply becomes:
#
# \begin{align}
# \mathbf{C} = \frac{1}{n-1} X^\top X
# \end{align}
#
# Next, apart from the original objective function $\mathbf w^\top \mathbf{Cw}$, we also introduce an additional constraint $\|\mathbf w\|=\mathbf w^\top \mathbf w=1$, saying our weight vector should have unit length. The intuition behind this is that: If we were to simply maximize the formula $\mathbf w^\top \mathbf{Cw}$, we can multiply $w$ by any number and the objective function will increase by the square of this number. So the problem becomes ill-defined since the maximum of this expression is infinite. Given these two pieces of information, our objective function for PCA becomes:
#
# \begin{align}
# & \underset{\mathbf{w}}{\text{maximize}}
# && \mathbf w^\top \mathbf{Cw} \nonumber \\
# & \text{subject to}
# && \mathbf w^\top \mathbf w=1
# \end{align}
#
# This objective function can be solved by the Lagrange multiplier, minimizing the loss function:
#
# \begin{align}
# L &= \mathbf w^\top \mathbf{Cw}-\lambda(\mathbf w^\top \mathbf w-1)
# \end{align}
#
# If this looks unfamiliar, check out the following tutorial on Lagrange multiplier. [Khan Academy: Lagrange multipliers, examples](https://www.khanacademy.org/math/multivariable-calculus/applications-of-multivariable-derivatives/constrained-optimization/a/lagrange-multipliers-examples). Next, to solve for $w$, we set the partial derivative of $L$ with respect to $w$ to 0.
#
# \begin{align}
# \frac{\partial L}{\partial \mathbf w}
# & \implies \mathbf{Cw} - \lambda \mathbf w = 0 \\
# & \implies \mathbf{Cw} = \lambda \mathbf w
# \end{align}
#
# Hopefully the formula above looks familar, since it's essentially an eigendecomposition problem. The notion of eigendecomposition is basically trying to solve the equation:
#
# \begin{align}
# Ax &= \lambda x
# \end{align}
#
# In our case, $A$ is our covariance matrix correponding to $\mathbf C$; $x$ is our eigenvector, correponding to $\mathbf w$ and $\lambda$ is our eigenvalue correponding to explained variance. After solving the equation above, we'll obtain eigenvector and eigenvalue pairs, where very eigenvector has a corresponding eigenvalue. An eigenvector is essentially the direction of each principal component and the eigenvalue is a number, telling us how much variance there is in the data in that direction, in other words, how spread out the data is on the line. For those interested, the following blog walks through the calculating eigenvalue and eigenvectors from scratch. [Blog: What are eigenvectors and eigenvalues?](http://www.visiondummy.com/2014/03/eigenvalues-eigenvectors/)
# Given all of that, let's see how this works in code. Just to summarize, the general framework for computing PCA is as follows:
#
# - Standardize the data
# - Obtain the Eigenvectors and Eigenvalues from the covariance matrix
# - Sort eigenvalues in descending order and choose the $k$ eigenvectors that correspond to the $k$ largest eigenvalues where $k$ is the number of dimensions of the new feature subspace
# - Projection onto the new feature space. During this step we will take the top $k$ eigenvectors and use it to transform the original dataset $X$ to obtain a k-dimensional feature subspace $X'$
#
# For the section below, we will be working with the famous "Iris" dataset. The iris dataset is a 150×4 matrix where the columns are the different features (sepal length in cm, sepal width in cm, petal length in cm, petal width in cm) and every every row represents a separate flower sample. The three classes in the Iris dataset are: Iris-setosa (n=50), Iris-versicolor (n=50), Iris-virginica (n=50). We'll use histograms to get a feeling of how the 3 different flower classes are distributes along the 4 different features.
# +
iris = load_iris()
X = iris['data']
y = iris['target']
label_dict = {0: 'Iris-Setosa',
1: 'Iris-Versicolor',
2: 'Iris-Virgnica'}
feature_dict = {0: 'sepal length [cm]',
1: 'sepal width [cm]',
2: 'petal length [cm]',
3: 'petal width [cm]'}
fig = plt.figure(figsize = (10, 6))
for feature in range(len(feature_dict)):
plt.subplot(2, 2, feature + 1)
for index, label in label_dict.items():
plt.hist(X[y == index, feature], label = label,
bins = 10, alpha = 0.3)
plt.xlabel(feature_dict[feature])
plt.legend(loc = 'upper right', fancybox = True, fontsize = 12)
plt.tight_layout()
plt.show()
# -
# ### Standardize
#
# In general, it's important to standardize the data prior to a PCA on the covariance matrix depends on the measurement scales of the original features. Since PCA yields a feature subspace that maximizes the variance along the axes, it makes sense to standardize the data, especially if it was measured on different scales so different features will have equal contribution in terms of their scales.
# subtract the mean and divide the
# standard deviation of each features
mean = np.mean(X, axis = 0)
scale = np.std(X, axis = 0)
X_std = (X - mean) / scale
# ### Eigendecomposition - Computing Eigenvectors and Eigenvalues
#
# The eigenvectors and eigenvalues of a covariance matrix is the core of PCA: The eigenvectors (principal components) determine the directions of the new feature space, and the eigenvalues determine their magnitude (variance explained along the principal components).
# +
# for those unfamiliar with the @ syntax it
# is equivalent of .dot, the dot product
vec_mean = np.mean(X_std, axis = 0)
vec_diff = X_std - vec_mean
cov_mat = vec_diff.T @ vec_diff / (X_std.shape[0] - 1)
print('Covariance matrix \n {}'.format(cov_mat))
# note that since we already standardize the data,
# meaning the mean vector for each features, vec_mean
# will be 0 (really small numbers if you were to print them out),
# hence we don't have to substract the mean to standardize them
cov_mat = X_std.T @ X_std / (X_std.shape[0] - 1)
print('Covariance matrix \n {}'.format(cov_mat))
# equivalently, we could have used the np.cov function:
# since each row represents a variable for np.cov,
# we'll need to transpose the matrix
print('NumPy covariance matrix: \n {}'.format(np.cov(X_std.T)))
# -
# After obtaining the covariance matrix, we perform an eigendecomposition on it to obtain the eigenvalues and eigenvectors.
# since computing this by hand can be quite tedious
# we'll simply use the pre-implemented function
eig_vals, eig_vecs = np.linalg.eig(cov_mat)
print('Eigenvectors \n%s' % eig_vecs)
print('\nEigenvalues \n%s' % eig_vals)
# ### Choosing Top K Eigenvectors
#
# Recall that the typical goal of a PCA is to reduce the dimensionality of the original feature space by projecting it onto a smaller subspace. In order to decide which eigenvector(s) can dropped without losing too much information for the construction of lower-dimensional subspace, we need to inspect the corresponding eigenvalues. The idea is: eigenvectors with the lowest eigenvalues bear the least information about the distribution of the data and those are the ones can be dropped. In order to do so, the common approach is to rank the eigenvalues from highest to lowest in order choose the top $k$ eigenvectors.
#
# After sorting the eigenpairs, the next question is "how many principal components are we going to choose for our new feature subspace?" A useful measure is the so-called “explained variance,” which can be calculated from the eigenvalues. The explained variance tells us how much extra information (variance) does each of the principal components contribute.
# +
eig_vals_total = np.sum(eig_vals)
var_exp = eig_vals / eig_vals_total
cum_var_exp = np.cumsum(var_exp)
print('Variance Explained: ', var_exp)
print('Cumulative Variance Explained: ', cum_var_exp)
plt.bar(range(var_exp.shape[0]), var_exp, alpha = 0.5,
align = 'center', label = 'individual explained variance')
plt.step(range(var_exp.shape[0]), cum_var_exp,
where = 'mid', label = 'cumulative explained variance')
plt.ylabel('Explained variance ratio')
plt.xlabel('Principal components')
plt.ylim(0, 1.1)
plt.legend(loc = 'best')
plt.tight_layout()
# -
# The plot above clearly shows that most of the variance (72.77% of the variance to be precise) can be explained by the first principal component alone. The second principal component still bears some information (23.03%) while the third and fourth principal components can safely be dropped without losing to much information. Together, the first two principal components explains 95.8% of the variance. Or in other words, it contain 95.8% of the information (95% or 99% are comon threshold that people use).
# ### Projection Onto the New Feature Space
#
# In the last step, we will use the 4×2-dimensional eigenvectors to transform our samples onto the new subspace via the equation $Y=XW$, where Y will be our 150×2 matrix of transformed samples (we reduce the dimension from the original 4 down to 2).
def plot_iris_pca(X_pca, y):
"""a scatter plot of the 2-dimensional iris data"""
markers = 's', 'x', 'o'
colors = list(plt.rcParams['axes.prop_cycle'])
target = np.unique(y)
for idx, (t, m) in enumerate( zip(target, markers) ):
subset = X_pca[y == t]
plt.scatter(subset[:, 0], subset[:, 1], s = 50,
c = colors[idx]['color'], label = t, marker = m)
plt.xlabel('PC 1')
plt.ylabel('PC 2')
plt.legend(loc = 'lower left')
plt.tight_layout()
plt.show()
X_std_pca = X_std.dot(eig_vecs[:, 0:2])
plot_iris_pca(X_std_pca, y)
# We'll put all of this into a single class, train it and confirm the result with scikit-learn's PCA model by printing out the explained variance ratio.
class PCAModel:
"""
Principal component analysis (PCA)
Parameters
----------
n_components : int
top number of principal components to keep
"""
def __init__(self, n_components):
self.n_components = n_components
def fit(self, X):
# standardize
X = X.copy()
self.mean = np.mean(X, axis = 0)
self.scale = np.std(X, axis = 0)
X = (X - self.mean) / self.scale
# eigendecomposition
eig_vals, eig_vecs = np.linalg.eig(np.cov(X.T))
self.components = eig_vecs[:, :self.n_components]
var_exp = eig_vals / np.sum(eig_vals)
self.explained_variance_ratio = var_exp[:self.n_components]
return self
def transform(self, X):
X = X.copy()
X = (X - self.mean) / self.scale
X_pca = X @ self.components
return X_pca
# +
# implementation from scratch
pca_model = PCAModel(n_components = 2).fit(X)
# using library to confirm results
scaler = StandardScaler()
X_std = scaler.fit_transform(X)
pca = PCA(n_components = 2).fit(X_std)
# print explained ratio to see if the matches
print('library: ', pca.explained_variance_ratio_)
print('from scratch: ', pca_model.explained_variance_ratio)
# -
# ## Takeaway
#
# Let's wrap this up with a summary of PCA and some of its applications. In order to identify patterns in our data, we often look for variation across observations to distinguish them from one another. Hence it seems reasonable to be able to find a succinct representation that best captures the variation in our initial data. PCA, in particular, look to explain our data via its maximum directions of variance. By compressing a higher dimensional dataset into lower one, while still retaining most of the variance this allows us to:
#
# - Perform Visualization: PCA summarizes our data along the principal components (or eigenvectors), which explains most the variance. Thus we can reduce the dataset to 2 or 3 dimensions and visualize our data's distribution. This can be helpful when we're performing clustering algorithm that needs to choose the cluster number beforehand.
# - Speed up machine learning algorithms: When dealing with Big Data, we might want to decompose the features into a lower dimension, without a significant loss in variance. By performing dimensionality reduction methods, we can reduce the amount of features we have to speed up the training algorithm and save memory. For this part, we'll use the Iris dataset again and show that by keeping the data's top 3 principal components we can obtain the same level of accuracy as keeping all 4 features.
# +
# split 30% of the iris data into a test set for evaluation
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.3,
random_state = 1)
# create the pipeline, where we'll
# standardize the data, perform PCA and
# fit the logistic regression
pipe1 = Pipeline([
('standardize', StandardScaler()),
('pca', PCA(n_components = 3)),
('logistic', LogisticRegression(random_state = 1))
])
pipe1.fit(X_train, y_train)
y_pred1 = pipe1.predict(X_test)
# pipeline without PCA
pipe2 = Pipeline([
('standardize', StandardScaler()),
('logistic', LogisticRegression(random_state = 1))
])
pipe2.fit(X_train, y_train)
y_pred2 = pipe2.predict(X_test)
# access the prediction accuracy
print('PCA Accuracy %.3f' % accuracy_score(y_test, y_pred1))
print('Accuracy %.3f' % accuracy_score(y_test, y_pred2))
# -
# # Reference
#
# - [Youtube: Dimensionality Reduction](https://www.youtube.com/playlist?list=PLnnr1O8OWc6aVexn2BY0qjklobY6TUEIy)
# - [Blog: What are eigenvectors and eigenvalues?](http://www.visiondummy.com/2014/03/eigenvalues-eigenvectors/)
# - [Blog: Principal Component Analysis 4 Dummies](https://georgemdallas.wordpress.com/2013/10/30/principal-component-analysis-4-dummies-eigenvectors-eigenvalues-and-dimension-reduction/)
# - [Blog: Everything you did and didn't know about PCA](http://alexhwilliams.info/itsneuronalblog/2016/03/27/pca/)
# - [Blog: Principal Component Analysis in 3 Simple Steps](http://sebastianraschka.com/Articles/2015_pca_in_3_steps.html)
# - [Notebook: In Depth: Principal Component Analysis](http://nbviewer.jupyter.org/github/jakevdp/PythonDataScienceHandbook/blob/95664b978439df948b2cd2f5f5b4e31f28b30394/notebooks/05.09-Principal-Component-Analysis.ipynb)
# - [StackExchange: Making sense of principal component analysis, eigenvectors & eigenvalues](http://stats.stackexchange.com/questions/2691/making-sense-of-principal-component-analysis-eigenvectors-eigenvalues)
# - [StackExchange: What is an intuitive explanation for how PCA turns from a geometric problem (with distances) to a linear algebra problem (with eigenvectors)?](http://stats.stackexchange.com/questions/217995/what-is-an-intuitive-explanation-for-how-pca-turns-from-a-geometric-problem-wit)
| dim_reduct/PCA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="W_tvPdyfA-BL"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="0O_LFhwSBCjm"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="PWUmcKKjtwXL"
# # Hub with Keras
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r1/tutorials/images/hub_with_keras.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r1/tutorials/images/hub_with_keras.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="crU-iluJIEzw"
# [TensorFlow Hub](http://tensorflow.org/hub) is a way to share pretrained model components. See the [TensorFlow Module Hub](https://tfhub.dev/) for a searchable listing of pre-trained models.
#
# This tutorial demonstrates:
#
# 1. How to use TensorFlow Hub with `tf.keras`.
# 1. How to do image classification using TensorFlow Hub.
# 1. How to do simple transfer learning.
# + [markdown] colab_type="text" id="CKFUvuEho9Th"
# ## Setup
# + [markdown] colab_type="text" id="7RVsYZLEpEWs"
# ### Imports
# + colab={} colab_type="code" id="nSiOCtv_Rwi_"
# !pip install -U tensorflow_hub
# + colab={} colab_type="code" id="OGNpmn43C0O6"
from __future__ import absolute_import, division, print_function, unicode_literals
import matplotlib.pylab as plt
try:
# # %tensorflow_version only exists in Colab.
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow.compat.v1 as tf
# + colab={} colab_type="code" id="-V4l8oN8Lw2q"
import tensorflow_hub as hub
from tensorflow.keras import layers
# + [markdown] colab_type="text" id="s4YuF5HvpM1W"
# ## An ImageNet classifier
# + [markdown] colab_type="text" id="xEY_Ow5loN6q"
# ### Download the classifier
#
# Use `hub.module` to load a mobilenet, and `tf.keras.layers.Lambda` to wrap it up as a keras layer.
#
# The URL of any [TF2-compatible image classification module](https://tfhub.dev/s?module-type=image-classification&q=tf2) from tfhub.dev will work here.
# + cellView="both" colab={} colab_type="code" id="feiXojVXAbI9"
classifier_url ="https://tfhub.dev/google/tf2-preview/mobilenet_v2/classification/2" #@param {type:"string"}
# + colab={} colab_type="code" id="y_6bGjoPtzau"
IMAGE_SHAPE = (224, 224)
classifier = tf.keras.Sequential([
hub.KerasLayer(classifier_url, input_shape=IMAGE_SHAPE+(3,))
])
# + [markdown] colab_type="text" id="pwZXaoV0uXp2"
# ### Run it on a single image
# + [markdown] colab_type="text" id="TQItP1i55-di"
# Download a single image to try the model on.
# + colab={} colab_type="code" id="w5wDjXNjuXGD"
import numpy as np
import PIL.Image as Image
grace_hopper = tf.keras.utils.get_file('image.jpg','https://storage.googleapis.com/download.tensorflow.org/example_images/grace_hopper.jpg')
grace_hopper = Image.open(grace_hopper).resize(IMAGE_SHAPE)
grace_hopper
# + colab={} colab_type="code" id="BEmmBnGbLxPp"
grace_hopper = np.array(grace_hopper)/255.0
grace_hopper.shape
# + [markdown] colab_type="text" id="0Ic8OEEo2b73"
# Add a batch dimension, and pass the image to the model.
# + colab={} colab_type="code" id="EMquyn29v8q3"
result = classifier.predict(grace_hopper[np.newaxis, ...])
result.shape
# + [markdown] colab_type="text" id="NKzjqENF6jDF"
# The result is a 1001 element vector of logits, rating the probability of each class for the image.
#
# So the top class ID can be found with argmax:
# + colab={} colab_type="code" id="rgXb44vt6goJ"
predicted_class = np.argmax(result[0], axis=-1)
predicted_class
# + [markdown] colab_type="text" id="YrxLMajMoxkf"
# ### Decode the predictions
#
# We have the predicted class ID,
# Fetch the `ImageNet` labels, and decode the predictions
# + colab={} colab_type="code" id="ij6SrDxcxzry"
labels_path = tf.keras.utils.get_file('ImageNetLabels.txt','https://storage.googleapis.com/download.tensorflow.org/data/ImageNetLabels.txt')
imagenet_labels = np.array(open(labels_path).read().splitlines())
# + colab={} colab_type="code" id="uzziRK3Z2VQo"
plt.imshow(grace_hopper)
plt.axis('off')
predicted_class_name = imagenet_labels[predicted_class]
_ = plt.title("Prediction: " + predicted_class_name.title())
# + [markdown] colab_type="text" id="amfzqn1Oo7Om"
# ## Simple transfer learning
# + [markdown] colab_type="text" id="K-nIpVJ94xrw"
# Using TF Hub it is simple to retrain the top layer of the model to recognize the classes in our dataset.
# + [markdown] colab_type="text" id="Z93vvAdGxDMD"
# ### Dataset
#
# For this example you will use the TensorFlow flowers dataset:
# + colab={} colab_type="code" id="DrIUV3V0xDL_"
data_root = tf.keras.utils.get_file(
'flower_photos','https://storage.googleapis.com/download.tensorflow.org/example_images/flower_photos.tgz',
untar=True)
# + [markdown] colab_type="text" id="jFHdp18ccah7"
# The simplest way to load this data into our model is using `tf.keras.preprocessing.image.ImageDataGenerator`,
#
# All of TensorFlow Hub's image modules expect float inputs in the `[0, 1]` range. Use the `ImageDataGenerator`'s `rescale` parameter to achieve this.
#
# The image size will be handled later.
# + colab={} colab_type="code" id="2PwQ_wYDcii9"
image_generator = tf.keras.preprocessing.image.ImageDataGenerator(rescale=1/255)
image_data = image_generator.flow_from_directory(str(data_root), target_size=IMAGE_SHAPE)
# + [markdown] colab_type="text" id="0p7iDOhIcqY2"
# The resulting object is an iterator that returns `image_batch, label_batch` pairs.
# + colab={} colab_type="code" id="W4lDPkn2cpWZ"
for image_batch, label_batch in image_data:
print("Image batch shape: ", image_batch.shape)
print("Label batch shape: ", label_batch.shape)
break
# + [markdown] colab_type="text" id="0gTN7M_GxDLx"
# ### Run the classifier on a batch of images
# + [markdown] colab_type="text" id="O3fvrZR8xDLv"
# Now run the classifier on the image batch.
# + colab={} colab_type="code" id="nbyg6tcyxDLh"
result_batch = classifier.predict(image_batch)
result_batch.shape
# + colab={} colab_type="code" id="Kv7ZwuR4xDLc"
predicted_class_names = imagenet_labels[np.argmax(result_batch, axis=-1)]
predicted_class_names
# + [markdown] colab_type="text" id="QmvSWg9nxDLa"
# Now check how these predictions line up with the images:
# + colab={} colab_type="code" id="IXTB22SpxDLP"
plt.figure(figsize=(10,9))
plt.subplots_adjust(hspace=0.5)
for n in range(30):
plt.subplot(6,5,n+1)
plt.imshow(image_batch[n])
plt.title(predicted_class_names[n])
plt.axis('off')
_ = plt.suptitle("ImageNet predictions")
# + [markdown] colab_type="text" id="FUa3YkvhxDLM"
# See the `LICENSE.txt` file for image attributions.
#
# The results are far from perfect, but reasonable considering that these are not the classes the model was trained for (except "daisy").
# + [markdown] colab_type="text" id="JzV457OXreQP"
# ### Download the headless model
#
# TensorFlow Hub also distributes models without the top classification layer. These can be used to easily do transfer learning.
#
# The URL of any [TF2-compatible image feature vector module](https://tfhub.dev/s?module-type=image-feature-vector&q=tf2) from tfhub.dev will work here.
# + cellView="both" colab={} colab_type="code" id="4bw8Jf94DSnP"
feature_extractor_url = "https://tfhub.dev/google/tf2-preview/mobilenet_v2/feature_vector/2" #@param {type:"string"}
# + [markdown] colab_type="text" id="sgwmHugQF-PD"
# Create the module, and check the expected image size:
# + colab={} colab_type="code" id="5wB030nezBwI"
feature_extractor_layer = hub.KerasLayer(feature_extractor_url,
input_shape=(224,224,3))
# + [markdown] colab_type="text" id="GUY-5Eyzuzlu"
# The feature extractor returns a 1280-element vector for each image.
# + colab={} colab_type="code" id="Of7i-35F09ls"
feature_batch = feature_extractor_layer(image_batch)
print(feature_batch.shape)
# + [markdown] colab_type="text" id="CtFmF7A5E4tk"
# Freeze the variables in the feature extractor layer, so that the training only modifies the new classifier layer.
# + colab={} colab_type="code" id="Jg5ar6rcE4H-"
feature_extractor_layer.trainable = False
# + [markdown] colab_type="text" id="RPVeouTksO9q"
# ### Attach a classification head
#
# Now wrap the hub layer in a `tf.keras.Sequential` model, and add a new classification layer.
# + colab={} colab_type="code" id="mGcY27fY1q3Q"
model = tf.keras.Sequential([
feature_extractor_layer,
layers.Dense(image_data.num_classes, activation='softmax')
])
model.summary()
# + colab={} colab_type="code" id="G9VkAz00HOJx"
predictions = model(image_batch)
# + colab={} colab_type="code" id="sB7sVGJ23vrY"
predictions.shape
# + [markdown] colab_type="text" id="OHbXQqIquFxQ"
# ### Train the model
#
# Use compile to configure the training process:
# + colab={} colab_type="code" id="3n0Wb9ylKd8R"
model.compile(
optimizer=tf.keras.optimizers.Adam(),
loss='categorical_crossentropy',
metrics=['acc'])
# + [markdown] colab_type="text" id="58-BLV7dupJA"
# Now use the `.fit` method to train the model.
#
# To keep this example short train just 2 epochs. To visualize the training progress, use a custom callback to log the loss and accuracy of each batch individually, instead of the epoch average.
# + colab={} colab_type="code" id="jZ54Gubac4Lu"
class CollectBatchStats(tf.keras.callbacks.Callback):
def __init__(self):
self.batch_losses = []
self.batch_acc = []
def on_train_batch_end(self, batch, logs=None):
self.batch_losses.append(logs['loss'])
self.batch_acc.append(logs['acc'])
self.model.reset_metrics()
# + colab={} colab_type="code" id="EyMDJxt2HdHr"
steps_per_epoch = np.ceil(image_data.samples/image_data.batch_size)
batch_stats_callback = CollectBatchStats()
history = model.fit(image_data, epochs=2,
steps_per_epoch=steps_per_epoch,
callbacks = [batch_stats_callback])
# + [markdown] colab_type="text" id="Kd0N272B9Q0b"
# Now after, even just a few training iterations, we can already see that the model is making progress on the task.
# + colab={} colab_type="code" id="A5RfS1QIIP-P"
plt.figure()
plt.ylabel("Loss")
plt.xlabel("Training Steps")
plt.ylim([0,2])
plt.plot(batch_stats_callback.batch_losses)
# + colab={} colab_type="code" id="3uvX11avTiDg"
plt.figure()
plt.ylabel("Accuracy")
plt.xlabel("Training Steps")
plt.ylim([0,1])
plt.plot(batch_stats_callback.batch_acc)
# + [markdown] colab_type="text" id="kb__ZN8uFn-D"
# ### Check the predictions
#
# To redo the plot from before, first get the ordered list of class names:
# + colab={} colab_type="code" id="JGbEf5l1I4jz"
class_names = sorted(image_data.class_indices.items(), key=lambda pair:pair[1])
class_names = np.array([key.title() for key, value in class_names])
class_names
# + [markdown] colab_type="text" id="4Olg6MsNGJTL"
# Run the image batch through the model and convert the indices to class names.
# + colab={} colab_type="code" id="fCLVCpEjJ_VP"
predicted_batch = model.predict(image_batch)
predicted_id = np.argmax(predicted_batch, axis=-1)
predicted_label_batch = class_names[predicted_id]
# + [markdown] colab_type="text" id="CkGbZxl9GZs-"
# Plot the result
# + colab={} colab_type="code" id="rpFQR1MPMtT1"
label_id = np.argmax(label_batch, axis=-1)
# + colab={} colab_type="code" id="wC_AYRJU9NQe"
plt.figure(figsize=(10,9))
plt.subplots_adjust(hspace=0.5)
for n in range(30):
plt.subplot(6,5,n+1)
plt.imshow(image_batch[n])
color = "green" if predicted_id[n] == label_id[n] else "red"
plt.title(predicted_label_batch[n].title(), color=color)
plt.axis('off')
_ = plt.suptitle("Model predictions (green: correct, red: incorrect)")
# + [markdown] colab_type="text" id="uRcJnAABr22x"
# ## Export your model
#
# Now that you've trained the model, export it as a saved model:
# + colab={} colab_type="code" id="PLcqg-RmsLno"
import time
t = time.time()
export_path = "/tmp/saved_models/{}".format(int(t))
tf.keras.experimental.export_saved_model(model, export_path)
export_path
# + [markdown] colab_type="text" id="AhQ9liIUsPsi"
# Now confirm that we can reload it, and it still gives the same results:
# + colab={} colab_type="code" id="7nI5fvkAQvbS"
reloaded = tf.keras.experimental.load_from_saved_model(export_path, custom_objects={'KerasLayer':hub.KerasLayer})
# + colab={} colab_type="code" id="jor83-LqI8xW"
result_batch = model.predict(image_batch)
reloaded_result_batch = reloaded.predict(image_batch)
# + colab={} colab_type="code" id="dnZO14taYPH6"
abs(reloaded_result_batch - result_batch).max()
# + [markdown] colab_type="text" id="TYZd4MNiV3Rc"
# This saved model can be loaded for inference later, or converted to [TFLite](https://www.tensorflow.org/lite/convert/) or [TFjs](https://github.com/tensorflow/tfjs-converter).
#
| site/en/r1/tutorials/images/hub_with_keras.ipynb |
;; -*- coding: utf-8 -*-
;; ---
;; jupyter:
;; jupytext:
;; text_representation:
;; extension: .scm
;; format_name: light
;; format_version: '1.5'
;; jupytext_version: 1.14.4
;; kernelspec:
;; display_name: Calysto Scheme 3
;; language: scheme
;; name: calysto_scheme
;; ---
;; ### 練習問題1.4
;; 我々の評価モデルでは、演算⼦が複合式であるような組み合わせが作れるということを観察せよ。
;; この観察結果を使って、次の⼿続きのふるまいを説明せよ。
;;
;; (define (a-plus-abs-b a b)
;; ((if (> b 0) + -) a b)
;; )
;;
(define (a-plus-abs-b a b)
((if (> b 0) + -) a b))
(a-plus-abs-b -5 3)
(a-plus-abs-b 3 -5)
(a-plus-abs-b 3 5)
;; 引数bの符号によって、演算方法を切り替えることができる。
| exercises/1.04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ### 20181013 flow data
# +
import fcsparser
import pandas as pd
import math
import os
#import all data, and merge into a single datatframe, with sample names in extra columns
def log_trans(x):
try:
return math.log(x, 10)
except:
return float('NaN')
cell_lines = ['54_UGAC-blast_x',
'55_UGAC-blast_x',
'56_UGAC_x',
'57_UGAC_a',
'58_AQP4_x',
'59_AQP4_a',
'60_OPRL1_x',
'61_OPRL1_a']
treatments = ['none', '100nM SMG1i', '500ug/ml G418']
data_folder = '20181013_G418_smg1i'
#will only look at data gated on good scatter
dfs = []
index = 0
for treatment in treatments:
for cell_line in cell_lines:
index += 1
data_file_name = 'Specimen_001_BFP_GFP_tdtomato_%03d_%03d.fcs' % (index, index)
data_file_path = os.path.join(data_folder, data_file_name)
meta, data = fcsparser.parse(data_file_path, meta_data_only=False, reformat_meta=True)
data['cell line'] = cell_line
data['treatment'] = treatment
data['sample'] = '%s %s' % (cell_line, treatment)
#print '%s %s' % (cell_line, treatment)
#print meta['_channels_']
#meta_dfs.append(meta)
#dfs.append(data.sample(n=1000))
dfs.append(data[:5000]) #to subsample and speed up plotting, use the above line instead of this one
all_data = pd.concat(dfs)
# +
#first just look at the forward and side scatter
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.stats import gaussian_kde
import numpy as np
sns.set(style="white", color_codes=True)
def density_colored_scatter(x, y, color, **kwargs):
# Calculate the point density
xy = np.vstack([x,y])
z = gaussian_kde(xy)(xy)
plt.scatter(x, y, c=z, **kwargs)
#plt.yscale('log')
#plt.xscale('log')
cutoff = 80000
pos = [yval for yval in y if yval>=cutoff]
percent_pos = 100.*len(pos)/len(y)
plt.hlines(cutoff,0,40000, linestyle='dashed')
plt.annotate("%.3f" % percent_pos, xy=(30000, cutoff+1))
g = sns.FacetGrid(all_data, col="sample", col_wrap=4)
cmap = 'viridis'
g= g.map(density_colored_scatter, "FSC-H", "FSC-W", cmap=cmap, s=1, linewidth=0)
plt.savefig('20181013_FSC_WH.pdf', transparent=True)
plt.savefig('20181013_FSC_WH.png', transparent=True)
# -
fsc_filter = all_data[all_data['FSC-W']<80000]
# +
#first just look at the forward and side scatter
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.stats import gaussian_kde
import numpy as np
sns.set(style="white", color_codes=True)
def density_colored_scatter(x, y, color, **kwargs):
# Calculate the point density
xy = np.vstack([x,y])
z = gaussian_kde(xy)(xy)
plt.scatter(x, y, c=z, **kwargs)
#plt.yscale('log')
#plt.xscale('log')
cutoff = 100000
pos = [yval for yval in y if yval>=cutoff]
percent_pos = 100.*len(pos)/len(y)
plt.hlines(cutoff,0,20000, linestyle='dashed')
plt.annotate("%.3f" % percent_pos, xy=(2000, cutoff+1))
#,ylim=(1,1000000)
g = sns.FacetGrid(all_data, col="sample", col_wrap=4, xlim=(0,20000), ylim=(0,200000))
cmap = 'viridis'
g= g.map(density_colored_scatter, "SSC-H", "SSC-W", cmap=cmap, s=1, linewidth=0)
plt.savefig('20181013_SSC_WH.pdf', transparent=True)
plt.savefig('20181013_SSC_WH.png', transparent=True)
# -
fsc_ssc_filter = fsc_filter[fsc_filter['SSC-W']<100000]
# +
#first just look at the forward and side scatter
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.stats import gaussian_kde
import numpy as np
sns.set(style="white", color_codes=True)
def density_colored_scatter(x, y, color, **kwargs):
# Calculate the point density
xy = np.vstack([x,y])
z = gaussian_kde(xy)(xy)
plt.scatter(x, y, c=z, **kwargs)
plt.yscale('log')
plt.xscale('log')
#cutoff = 400
#pos = [xval for xval in x if xval>=cutoff]
#percent_pos = 100.*len(pos)/len(x)
#plt.vlines(cutoff,1,1000000, linestyle='dashed')
#plt.annotate("%.1f" % percent_pos, xy=(cutoff+1, 500000))
#xlim=(1,1000000),ylim=(1,1000000)
g = sns.FacetGrid(fsc_ssc_filter, col="sample", col_wrap=4)
cmap = 'viridis'
g= g.map(density_colored_scatter, "FSC-A", "SSC-A", cmap=cmap, s=1, linewidth=0)
plt.savefig('20181013_fsc_ssc_filtered_FSC_SSC_A.pdf', transparent=True)
plt.savefig('20181013_fsc_ssc_filtered_FSC_SSC_A.png', transparent=True)
# -
good_scatter = fsc_ssc_filter.rename(index=str, columns={'PE-A':'tdTomato-A', 'Pacific Blue-A':'tagBFP-A'})
good_scatter['tdTomato/tagBFP'] = good_scatter['tdTomato-A']/good_scatter['tagBFP-A']
good_scatter['log tdTomato'] = good_scatter['tdTomato-A'].apply(log_trans)
good_scatter['log tagBFP'] = good_scatter['tagBFP-A'].apply(log_trans)
good_scatter['log GFP'] = good_scatter['GFP-A'].apply(log_trans)
good_scatter['log tdTomato/tagBFP'] = good_scatter['tdTomato/tagBFP'].apply(log_trans)
# +
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(style="white", color_codes=True)
fluor_reporters = ['56_UGAC_x',
'57_UGAC_a',
'58_AQP4_x',
'59_AQP4_a',
'60_OPRL1_x',
'61_OPRL1_a']
blast_reporters = ['54_UGAC-blast_x',
'55_UGAC-blast_x']
fig, ax = plt.subplots(1,3, figsize=(16, 8), sharey=False)
sns.boxplot(x="cell line", y="log tagBFP", hue='treatment', data=good_scatter[good_scatter['cell line'].isin(fluor_reporters)], ax=ax[0], notch=True, linewidth=1)
sns.boxplot(x="cell line", y="log tdTomato", hue='treatment', data=good_scatter[good_scatter['cell line'].isin(fluor_reporters)], ax=ax[1], notch=True, linewidth=1)
sns.boxplot(x="cell line", y="log tdTomato/tagBFP", hue='treatment', data=good_scatter[good_scatter['cell line'].isin(fluor_reporters)], ax=ax[2], notch=True, linewidth=1)
for ind_axis in ax:
for label in ind_axis.get_xticklabels():
label.set_rotation(90)
plt.savefig('20181013_log_tagBFP_tdTomato.pdf', transparent=True)
# +
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(style="white", color_codes=True)
fluor_reporters = ['56_UGAC_x',
'57_UGAC_a',
'58_AQP4_x',
'59_AQP4_a',
'60_OPRL1_x',
'61_OPRL1_a']
blast_reporters = ['56_UGAC_x', '54_UGAC-blast_x',
'55_UGAC-blast_x']
fig, ax = plt.subplots(1,2, figsize=(16, 8), sharey=False)
sns.boxplot(x="cell line", y="log tagBFP", hue='treatment', data=good_scatter[good_scatter['cell line'].isin(blast_reporters)], ax=ax[0], notch=True, linewidth=1)
sns.boxplot(x="cell line", y="log GFP", hue='treatment', data=good_scatter[good_scatter['cell line'].isin(blast_reporters)], ax=ax[1], notch=True, linewidth=1)
for ind_axis in ax:
for label in ind_axis.get_xticklabels():
label.set_rotation(90)
plt.savefig('20181013_log_tagBFP_eGFP.pdf', transparent=True)
# +
# %matplotlib inline
import matplotlib as mpl
mpl.rcParams['pdf.fonttype'] = 42
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.stats import gaussian_kde
import numpy as np
sns.set(style="white", color_codes=True)
colors = ["greyish", "windows blue", "amber", "faded green", "dusty purple"]
sns.set_palette(sns.xkcd_palette(colors))
g = sns.FacetGrid(good_scatter, col="cell line", hue="treatment", col_wrap=4,
legend_out=True)
#xlim=(0,5000),ylim=(0,1000)
g= g.map(plt.scatter, "log tagBFP", "log tdTomato", cmap=cmap, s=5, linewidth=0, alpha=0.5)
g = g.add_legend()
#for i, ax in enumerate(g.axes.flat):
# ax.set_yscale('log')
# ax.set_xscale('log')
plt.savefig('201801013_tagBFP_tdTomato_log_scatter_hue.pdf', transparent=True)
plt.savefig('201801013_tagBFP_tdTomato_log_scatter_hue.png', transparent=True)
#plt.legend()
# +
# %matplotlib inline
import matplotlib as mpl
mpl.rcParams['pdf.fonttype'] = 42
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.stats import gaussian_kde
import numpy as np
sns.set(style="white", color_codes=True)
colors = ["greyish", "windows blue", "amber", "faded green", "dusty purple"]
sns.set_palette(sns.xkcd_palette(colors))
g = sns.FacetGrid(good_scatter, col="cell line", hue="treatment", col_wrap=4,
legend_out=True)
#xlim=(0,5000),ylim=(0,1000)
g= g.map(plt.scatter, "log tagBFP", "log GFP", cmap=cmap, s=5, linewidth=0, alpha=0.5)
g = g.add_legend()
#for i, ax in enumerate(g.axes.flat):
# ax.set_yscale('log')
# ax.set_xscale('log')
plt.savefig('201801013_tagBFP_GFP_log_scatter_hue.pdf', transparent=True)
plt.savefig('201801013_tagBFP_GFP_log_scatter_hue.png', transparent=True)
#plt.legend()
# -
# ### 20181104 for lab meeting
# Make KDE histograms of data and theoretical 5% gating estimates for ratios
subset_samples = ['54_UGAC-blast_x', '56_UGAC_x','58_AQP4_x','60_OPRL1_x']
subset_data = good_scatter[good_scatter['cell line'].isin(subset_samples)]
from scipy.stats import gaussian_kde
import numpy as np
sns.set(style="white", color_codes=True, font_scale=3)
g = sns.FacetGrid(subset_data, hue='treatment', col='cell line', col_wrap=4, height=8)
g.map(sns.kdeplot, 'log tagBFP', linewidth = 5)
#g.fig.suptitle("filtered for td", size=16)
g.add_legend();
plt.savefig('20181104_log_tagBFP_kde.pdf', transparent=True)
from scipy.stats import gaussian_kde
import numpy as np
sns.set(style="white", color_codes=True, font_scale=3)
g = sns.FacetGrid(subset_data, hue='treatment', col='cell line', col_wrap=4, height=8)
g.map(sns.kdeplot, 'log tdTomato', linewidth = 5)
#g.fig.suptitle("filtered for td", size=16)
g.add_legend();
plt.savefig('20181104_log_tdTomato_kde.pdf', transparent=True)
from scipy.stats import gaussian_kde
import numpy as np
sns.set(style="white", color_codes=True, font_scale=3)
g = sns.FacetGrid(subset_data, hue='treatment', col='cell line', col_wrap=4, height=8)
g.map(sns.kdeplot, "log tdTomato/tagBFP", linewidth = 5)
#g.fig.suptitle("filtered for td", size=16)
g.add_legend();
plt.savefig('20181104_log_tdTomato_tagBFP_ratio_kde.pdf', transparent=True)
# +
#Figure out top and bottom 5% cutoff for controls, then compute gate %
from scipy.stats import gaussian_kde
import numpy as np
sns.set(style="white", color_codes=True, font_scale=3)
g = sns.FacetGrid(subset_data, col_order=subset_samples, hue='treatment', col='cell line', col_wrap=4, height=8)
g.map(sns.kdeplot, "log tdTomato/tagBFP", linewidth = 5)
#g.fig.suptitle("filtered for td", size=16)
g.add_legend();
axes = g.axes
#axes[0].vlines((bottom5_56, top5_56),0,3, linestyle='dashed', lw=3)
#axes[1].vlines((bottom5_58, top5_58),0,3, linestyle='dashed', lw=3)
ax_index = 0
treatments = ['none', '100nM SMG1i', '500ug/ml G418']
for cell_line in subset_samples:
untreated_distribution = good_scatter[good_scatter['sample'] == '%s none' % (cell_line)].sort_values(by='log tdTomato/tagBFP')
bottom5 = untreated_distribution.iloc[int(.05*len(untreated_distribution))]['log tdTomato/tagBFP']
top5 = untreated_distribution.iloc[int(.95*len(untreated_distribution))]['log tdTomato/tagBFP']
axes[ax_index].vlines((bottom5, top5),0,5, linestyle='dashed', lw=3)
axes[ax_index].set_ylim(0,5.5)
for treatment in treatments:
sample = '%s %s' % (cell_line, treatment)
sample_data = good_scatter[good_scatter['sample']==sample]
print '%s bottom 5 gate: %.2f' % (sample, 100.*len(sample_data[sample_data['log tdTomato/tagBFP']<bottom5])/float(len(sample_data)))
print '%s top 5 gate: %.2f' % (sample, 100.*len(sample_data[sample_data['log tdTomato/tagBFP']>top5])/float(len(sample_data)))
ax_index+=1
plt.savefig('20181104_log_tdTomato_tagBFP_ratio_kde_gating.pdf', transparent=True)
# -
# ## 20190624 - Figures for RNA club talk
grouped_data = good_scatter.groupby(['cell line', 'treatment'])
#colors - colorblind safe
black = (0,0,0)
gray = (0.6,0.6,0.6)
orange = (230/255.0,159/255.0,0)
skyBlue = (86/255.0,180/255.0,233/255.0)
bluishGreen = (0,158/255.0,115/255.0)
yellow = (240/255.0,228/255.0,66/255.0)
blue = (0,114/255.0,178/255.0)
vermillion = (213/255.0,94/255.0,0)
reddishPurple = (204/255.0,121/255.0,167/255.0)
colors = [black, vermillion, orange, skyBlue, bluishGreen, blue, reddishPurple, yellow, gray]
# +
# %matplotlib inline
import warnings
import matplotlib as mpl
warnings.filterwarnings("ignore")
mpl.rcParams['pdf.fonttype'] = 42
import matplotlib.pyplot as plt
import scipy.stats as stats
import numpy as np
#xmin=0
#xmax=100
#ymin=-0.1
#ymax=4
plots=[]
num_plots_wide = 1
num_plots_high = 1
plotting_cell_lines = ['56_UGAC_x','58_AQP4_x','60_OPRL1_x']
plotting_treatments = ['none']
x_var='tagBFP-A'
y_var='tdTomato-A'
plot_index =1
fig = plt.figure(figsize=(6*num_plots_wide, 6*num_plots_high))
colors=[black, skyBlue, bluishGreen]
for treatment in plotting_treatments:
color_index=0
plot = fig.add_subplot(num_plots_high, num_plots_wide, plot_index)
for cell_line in plotting_cell_lines:
plots.append(plot)
data = grouped_data.get_group((cell_line, treatment))
data.plot.scatter(x=x_var,y=y_var, color=colors[color_index], alpha=0.2, ax=plot, lw=0, s=6)
#DMSO_data.plot.scatter(x=x_var, y=y_var, color=black, alpha=0.2, ax=plot, lw=0, s=6)
#plot.set_title('%s %s'% (stop_codon, gene), fontsize=24)
plot.set_xlabel(x_var, fontsize=30)
plot.set_ylabel(y_var, fontsize=30)
plot.set_xlim(0, 5000)
plot.set_ylim(0, 10000)
color_index+=1
plot_index += 1
plt.box(False)
#plt.savefig('20190611_scatter_smg1i_vs_dmso.pdf', transparent=True)
#plt.savefig('20190611_scatter_smg1i_vs_dmso.png', transparent=True)
# +
# %matplotlib inline
import warnings
import matplotlib as mpl
warnings.filterwarnings("ignore")
mpl.rcParams['pdf.fonttype'] = 42
import matplotlib.pyplot as plt
import scipy.stats as stats
import numpy as np
from matplotlib.ticker import NullFormatter
plotting_cell_lines = ['56_UGAC_x','58_AQP4_x','60_OPRL1_x']
plotting_treatments = ['none']
nullfmt = NullFormatter() # no labels
# definitions for the axes
left, width = 0.1, 0.65
bottom, height = 0.1, 0.65
bottom_h = left_h = left + width + 0.02
rect_scatter = [left, bottom, width, height]
rect_histx = [left, bottom_h, width, 0.2]
rect_histy = [left_h, bottom, 0.2, height]
# start with a rectangular Figure
plt.figure(1, figsize=(8, 8))
axScatter = plt.axes(rect_scatter)
axHistx = plt.axes(rect_histx)
axHisty = plt.axes(rect_histy)
# no labels
axHistx.xaxis.set_major_formatter(nullfmt)
axHisty.yaxis.set_major_formatter(nullfmt)
axes = [axScatter, axHistx, axHisty]
hist_axes = [axHistx, axHisty]
x_var='log tagBFP'
y_var='log tdTomato'
colors=[black, skyBlue, bluishGreen]
for treatment in plotting_treatments:
color_index=0
for cell_line in plotting_cell_lines:
data = grouped_data.get_group((cell_line, treatment))
data.plot.scatter(x=x_var,y=y_var, color=colors[color_index], alpha=0.5, ax=axScatter, lw=0, s=6, label = cell_line)
data[x_var].plot.hist(histtype='step', bins=100, normed=True, color=colors[color_index], alpha=1, ax=axHistx, lw=3, label = cell_line)
data[y_var].plot.hist(histtype='step', bins=100, normed=True, color=colors[color_index], alpha=1, ax=axHisty, lw=3, orientation='horizontal', label = cell_line)
color_index+=1
for axis in axes:
for dir in axis.spines:
axis.spines[dir].set_visible(False)
for axis in hist_axes:
axis.get_xaxis().set_ticks([])
axis.get_yaxis().set_ticks([])
axis.set_ylabel('')
axScatter.set_xlabel(x_var, fontsize=30)
axScatter.set_ylabel(y_var, fontsize=30)
axScatter.set_xlim(1.5, 4)
axScatter.set_ylim(2.2, 4.3)
axHistx.set_xlim(axScatter.get_xlim())
axHisty.set_ylim(axScatter.get_ylim())
plt.legend()
plt.savefig('E4.036_20190624_compare_contexts_log.pdf', transparent=True)
#plt.savefig('E4.036_20190624_compare_contexts_log.pdf', transparent=True)
# +
# %matplotlib inline
import warnings
import matplotlib as mpl
warnings.filterwarnings("ignore")
mpl.rcParams['pdf.fonttype'] = 42
import matplotlib.pyplot as plt
import scipy.stats as stats
import numpy as np
from matplotlib.ticker import NullFormatter
plotting_cell_lines = ['56_UGAC_x']
plotting_treatments = ['none', '100nM SMG1i', '500ug/ml G418']
nullfmt = NullFormatter() # no labels
# definitions for the axes
left, width = 0.1, 0.65
bottom, height = 0.1, 0.65
bottom_h = left_h = left + width + 0.02
rect_scatter = [left, bottom, width, height]
rect_histx = [left, bottom_h, width, 0.2]
rect_histy = [left_h, bottom, 0.2, height]
# start with a rectangular Figure
plt.figure(1, figsize=(8, 8))
axScatter = plt.axes(rect_scatter)
axHistx = plt.axes(rect_histx)
axHisty = plt.axes(rect_histy)
# no labels
axHistx.xaxis.set_major_formatter(nullfmt)
axHisty.yaxis.set_major_formatter(nullfmt)
axes = [axScatter, axHistx, axHisty]
hist_axes = [axHistx, axHisty]
x_var='log tagBFP'
y_var='log tdTomato'
colors=[black, blue, vermillion]
color_index=0
for treatment in plotting_treatments:
for cell_line in plotting_cell_lines:
data = grouped_data.get_group((cell_line, treatment))
data.plot.scatter(x=x_var,y=y_var, color=colors[color_index], alpha=0.5, ax=axScatter, lw=0, s=6, label = treatment)
data[x_var].plot.hist(histtype='step', bins=100, normed=True, color=colors[color_index], alpha=1, ax=axHistx, lw=3, label = treatment)
data[y_var].plot.hist(histtype='step', bins=100, normed=True, color=colors[color_index], alpha=1, ax=axHisty, lw=3, orientation='horizontal', label = treatment)
color_index+=1
for axis in axes:
for dir in axis.spines:
axis.spines[dir].set_visible(False)
for axis in hist_axes:
axis.get_xaxis().set_ticks([])
axis.get_yaxis().set_ticks([])
axis.set_ylabel('')
axScatter.set_xlabel(x_var, fontsize=30)
axScatter.set_ylabel(y_var, fontsize=30)
axScatter.set_xlim(1.7, 3.5)
axScatter.set_ylim(2.2, 4.6)
axHistx.set_xlim(axScatter.get_xlim())
axHisty.set_ylim(axScatter.get_ylim())
plt.legend()
plt.savefig('E4.036_20190624_UGAC_compare_treatments_log.pdf', transparent=True)
#plt.savefig('E4.036_20190624_compare_contexts_log.pdf', transparent=True)
# +
# %matplotlib inline
import warnings
import matplotlib as mpl
warnings.filterwarnings("ignore")
mpl.rcParams['pdf.fonttype'] = 42
import matplotlib.pyplot as plt
import scipy.stats as stats
import numpy as np
from matplotlib.ticker import NullFormatter
plotting_cell_lines = ['58_AQP4_x']
plotting_treatments = ['none', '100nM SMG1i', '500ug/ml G418']
nullfmt = NullFormatter() # no labels
# definitions for the axes
left, width = 0.1, 0.65
bottom, height = 0.1, 0.65
bottom_h = left_h = left + width + 0.02
rect_scatter = [left, bottom, width, height]
rect_histx = [left, bottom_h, width, 0.2]
rect_histy = [left_h, bottom, 0.2, height]
# start with a rectangular Figure
plt.figure(1, figsize=(8, 8))
axScatter = plt.axes(rect_scatter)
axHistx = plt.axes(rect_histx)
axHisty = plt.axes(rect_histy)
# no labels
axHistx.xaxis.set_major_formatter(nullfmt)
axHisty.yaxis.set_major_formatter(nullfmt)
axes = [axScatter, axHistx, axHisty]
hist_axes = [axHistx, axHisty]
x_var='log tagBFP'
y_var='log tdTomato'
colors=[black, blue, vermillion]
color_index=0
for treatment in plotting_treatments:
for cell_line in plotting_cell_lines:
data = grouped_data.get_group((cell_line, treatment))
data.plot.scatter(x=x_var,y=y_var, color=colors[color_index], alpha=0.5, ax=axScatter, lw=0, s=6, label = treatment)
data[x_var].plot.hist(histtype='step', bins=100, normed=True, color=colors[color_index], alpha=1, ax=axHistx, lw=3, label = treatment)
data[y_var].plot.hist(histtype='step', bins=100, normed=True, color=colors[color_index], alpha=1, ax=axHisty, lw=3, orientation='horizontal', label = treatment)
color_index+=1
for axis in axes:
for dir in axis.spines:
axis.spines[dir].set_visible(False)
for axis in hist_axes:
axis.get_xaxis().set_ticks([])
axis.get_yaxis().set_ticks([])
axis.set_ylabel('')
axScatter.set_xlabel(x_var, fontsize=30)
axScatter.set_ylabel(y_var, fontsize=30)
axScatter.set_xlim(2.5, 4.)
axScatter.set_ylim(2.5, 4.)
axHistx.set_xlim(axScatter.get_xlim())
axHisty.set_ylim(axScatter.get_ylim())
plt.legend()
plt.savefig('E4.036_20190624_AQP4_compare_treatments_log.pdf', transparent=True)
#plt.savefig('E4.036_20190624_compare_contexts_log.pdf', transparent=True)
# +
# %matplotlib inline
import warnings
import matplotlib as mpl
warnings.filterwarnings("ignore")
mpl.rcParams['pdf.fonttype'] = 42
import matplotlib.pyplot as plt
import scipy.stats as stats
import numpy as np
from matplotlib.ticker import NullFormatter
plotting_cell_lines = ['60_OPRL1_x']
plotting_treatments = ['none', '100nM SMG1i', '500ug/ml G418']
nullfmt = NullFormatter() # no labels
# definitions for the axes
left, width = 0.1, 0.65
bottom, height = 0.1, 0.65
bottom_h = left_h = left + width + 0.02
rect_scatter = [left, bottom, width, height]
rect_histx = [left, bottom_h, width, 0.2]
rect_histy = [left_h, bottom, 0.2, height]
# start with a rectangular Figure
plt.figure(1, figsize=(8, 8))
axScatter = plt.axes(rect_scatter)
axHistx = plt.axes(rect_histx)
axHisty = plt.axes(rect_histy)
# no labels
axHistx.xaxis.set_major_formatter(nullfmt)
axHisty.yaxis.set_major_formatter(nullfmt)
axes = [axScatter, axHistx, axHisty]
hist_axes = [axHistx, axHisty]
x_var='log tagBFP'
y_var='log tdTomato'
colors=[black, blue, vermillion]
color_index=0
for treatment in plotting_treatments:
for cell_line in plotting_cell_lines:
data = grouped_data.get_group((cell_line, treatment))
data.plot.scatter(x=x_var,y=y_var, color=colors[color_index], alpha=0.5, ax=axScatter, lw=0, s=6, label = treatment)
data[x_var].plot.hist(histtype='step', bins=100, normed=True, color=colors[color_index], alpha=1, ax=axHistx, lw=3, label = treatment)
data[y_var].plot.hist(histtype='step', bins=100, normed=True, color=colors[color_index], alpha=1, ax=axHisty, lw=3, orientation='horizontal', label = treatment)
color_index+=1
for axis in axes:
for dir in axis.spines:
axis.spines[dir].set_visible(False)
for axis in hist_axes:
axis.get_xaxis().set_ticks([])
axis.get_yaxis().set_ticks([])
axis.set_ylabel('')
axScatter.set_xlabel(x_var, fontsize=30)
axScatter.set_ylabel(y_var, fontsize=30)
axScatter.set_xlim(2.5, 4.)
axScatter.set_ylim(2.8, 4.5)
axHistx.set_xlim(axScatter.get_xlim())
axHisty.set_ylim(axScatter.get_ylim())
plt.legend()
plt.savefig('E4.036_20190624_OPRL1_compare_treatments_log.pdf', transparent=True)
#plt.savefig('E4.036_20190624_compare_contexts_log.pdf', transparent=True)
# -
| F1_S1_S2_S3_flow_analysis/20181013_G418_smg1i-old/fcs/.ipynb_checkpoints/20181013_smg1i_G418_flow_analysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Python
#
# ## [Classes](https://docs.python.org/3/tutorial/classes.html) and OOP
#
#
# + Classes provide means of bundling data and functionality together.
# + Creating a new class is equivalent to creating a new type of object, allowing new instances of that type to be made.
# + Each class instance can have attributes attached to it for maintaining its state.
# + Class instances can also have methods for modifying its state.
#
# #### The class signature is:
#
# class ClassName(Superclass - optional):
# <statement-1>
# .
# .
# .
# <statement-N>
# ### Let's start with an example
class MyClass:
"""
class documentation string
"""
class_attribute1 = 1234
class_attribute2 = 5678
def class_method(self):
return 'Hello!'
print(type(MyClass))
x = MyClass()
y = MyClass()
print(type(x))
print(type(y))
dir(x)
print(x.__hash__())
print(y.__hash__())
x.class_attribute1
x.class_attribute2
x.class_method()
x.class_attribute1 = 4321
x.class_attribute1
y.class_attribute1
x.new_attribute = "any new attribute"
z = [1,2,3]
dir(x)
dir(y)
# ### Another example, with __init__() method:
class MyComplex:
def __init__(self, realpart, imagpart):
self.r = realpart
self.i = imagpart
print("__init__ has run")
z = MyComplex(2,-4)
j = MyComplex(13,12)
z.r
z.i
dir(j)
# ### Adding methods:
class Rectangle:
def __init__(self, width, height):
self.width = width
self.height = height
self.area = width * height
self.perimeter = 2 * (width+height)
if width < height:
self.invert_sides()
def invert_sides(self):
self.width, self.height = self.height, self.width
print('inverted height and width')
my_rectangle = Rectangle(50,10)
print(my_rectangle.width)
print(my_rectangle.height)
print(my_rectangle.area)
print(my_rectangle.perimeter)
my_rectangle.invert_sides()
print(my_rectangle.width)
print(my_rectangle.height)
dir(my_rectangle)
# #### What is printed when we call the "print" primitive in an object?
my_rectangle.__repr__()
# #### _print()_ calls the internal __repr__() method.
print(my_rectangle)
# ### Another class example:
class Triangle:
def __init__(self, side1, side2, side3):
self.side1 = side1
self.side2 = side2
self.side3 = side3
print('running __init__\n')
self.type_of_triangle()
def __repr__(self):
return f"I am a Triangle with sides {self.side1}, {self.side2} and {self.side3}"
#def __repr__(self):
# return "42"
def type_of_triangle(self):
if self.side1 == self.side2 and self.side1 == self.side3:
print('I am equilateral')
self.mytype = 'equilateral'
elif self.side1 == self.side2 or \
self.side1 == self.side3 or \
self.side2 == self.side3:
print('I am isosceles')
self.mytype = 'isosceles'
else:
print('I am scalene')
self.mytype = 'scalene'
tri = Triangle(5,5,5)
tri.__init__(5,4,5)
print(tri.mytype)
print(tri.side1)
tri.type_of_triangle()
print(tri)
dir(tri)
# ### [Special Methods for Classes](https://docs.python.org/3/reference/datamodel.html#special-method-names)
#
# [examples](https://www.pythonlikeyoumeanit.com/Module4_OOP/Special_Methods.html)
# x[key] = item<table class="docutils align-default">
# <colgroup>
# <col style="width: 33%">
# <col style="width: 33%">
# <col style="width: 33%">
# </colgroup>
# <thead>
# <tr class="row-odd"><th class="head"><p>Method</p></th>
# <th class="head"><p>Signature</p></th>
# <th class="head"><p>Explanation</p></th>
# </tr>
# </thead>
# <tbody>
# <tr class="row-even"><td><p>Returns string for a printable representation of object</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">__repr__(self)</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">repr(x)</span></code> invokes <code class="docutils literal notranslate"><span class="pre">x.__repr__()</span></code>, this is also invoked when an object is returned by a console</p></td>
# </tr>
# <tr class="row-odd"><td><p>Returns string representation of an object</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">__str__(self)</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">str(x)</span></code> invokes <code class="docutils literal notranslate"><span class="pre">x.__str__()</span></code></p></td>
# </tr>
# </tbody>
# </table>
# <table class="docutils align-default">
# <colgroup>
# <col style="width: 13%">
# <col style="width: 38%">
# <col style="width: 50%">
# </colgroup>
# <thead>
# <tr class="row-odd"><th class="head"><p>Method</p></th>
# <th class="head"><p>Signature</p></th>
# <th class="head"><p>Explanation</p></th>
# </tr>
# </thead>
# <tbody>
# <tr class="row-even"><td><p>Add</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">__add__(self,</span> <span class="pre">other)</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">x</span> <span class="pre">+</span> <span class="pre">y</span></code> invokes <code class="docutils literal notranslate"><span class="pre">x.__add__(y)</span></code></p></td>
# </tr>
# <tr class="row-odd"><td><p>Subtract</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">__sub__(self,</span> <span class="pre">other)</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">x</span> <span class="pre">-</span> <span class="pre">y</span></code> invokes <code class="docutils literal notranslate"><span class="pre">x.__sub__(y)</span></code></p></td>
# </tr>
# <tr class="row-even"><td><p>Multiply</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">__mul__(self,</span> <span class="pre">other)</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">x</span> <span class="pre">*</span> <span class="pre">y</span></code> invokes <code class="docutils literal notranslate"><span class="pre">x.__mul__(y)</span></code></p></td>
# </tr>
# <tr class="row-odd"><td><p>Divide</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">__truediv__(self,</span> <span class="pre">other)</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">x</span> <span class="pre">/</span> <span class="pre">y</span></code> invokes <code class="docutils literal notranslate"><span class="pre">x.__truediv__(y)</span></code></p></td>
# </tr>
# <tr class="row-even"><td><p>Power</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">__pow__(self,</span> <span class="pre">other)</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">x</span> <span class="pre">**</span> <span class="pre">y</span></code> invokes <code class="docutils literal notranslate"><span class="pre">x.__pow__(y)</span></code></p></td>
# </tr>
# </tbody>
# </table>
# <table class="docutils align-default">
# <colgroup>
# <col style="width: 33%">
# <col style="width: 33%">
# <col style="width: 33%">
# </colgroup>
# <thead>
# <tr class="row-odd"><th class="head"><p>Method</p></th>
# <th class="head"><p>Signature</p></th>
# <th class="head"><p>Explanation</p></th>
# </tr>
# </thead>
# <tbody>
# <tr class="row-even"><td><p>Length</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">__len__(self)</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">len(x)</span></code> invokes <code class="docutils literal notranslate"><span class="pre">x.__len__()</span></code></p></td>
# </tr>
# <tr class="row-odd"><td><p>Get Item</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">__getitem__(self,</span> <span class="pre">key)</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">x[key]</span></code> invokes <code class="docutils literal notranslate"><span class="pre">x.__getitem__(key)</span></code></p></td>
# </tr>
# <tr class="row-even"><td><p>Set Item</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">__setitem__(self,</span> <span class="pre">key,</span> <span class="pre">item)</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">x[key]</span> <span class="pre">=</span> <span class="pre">item</span></code> invokes <code class="docutils literal notranslate"><span class="pre">x.__setitem__(key,</span> <span class="pre">item)</span></code></p></td>
# </tr>
# <tr class="row-odd"><td><p>Contains</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">__contains__(self,</span> <span class="pre">item)</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">item</span> <span class="pre">in</span> <span class="pre">x</span></code> invokes <code class="docutils literal notranslate"><span class="pre">x.__contains__(item)</span></code></p></td>
# </tr>
# <tr class="row-even"><td><p>Iterator</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">__iter__(self)</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">iter(x)</span></code> invokes <code class="docutils literal notranslate"><span class="pre">x.__iter__()</span></code></p></td>
# </tr>
# <tr class="row-odd"><td><p>Next</p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">__next__(self)</span></code></p></td>
# <td><p><code class="docutils literal notranslate"><span class="pre">next(x)</span></code> invokes <code class="docutils literal notranslate"><span class="pre">x.__next__()</span></code></p></td>
# </tr>
# </tbody>
# </table>
# ### Operator overloading __gt__ (>) and __lt__ (<)
# +
class Thing:
def __init__(self, value):
self.__value = value
def __gt__(self, other):
return self.__value > other.__value
def __lt__(self, other):
return self.__value < other.__value
something = Thing(100)
nothing = Thing(10)
# -
something > nothing
something < nothing
something + nothing # erro
# +
class Thing:
def __init__(self, value):
self.__value = value
def __gt__(self, other):
return self.__value > other.__value
def __lt__(self, other):
return self.__value < other.__value
def __add__(self, other):
return self.__value + other.__value
something = Thing(100)
nothing = Thing(10)
# -
something + nothing
nothing + something
# ### [Class inheritance](https://medium.com/swlh/classes-subclasses-in-python-12b6013d9f3)
class Dog:
def __init__(self, name, race='Royal Street Dog'):
self.name = name
self.race = race
self.tricks = []
def add_trick(self, trick):
if not trick in self.tricks:
self.tricks.append(trick.title())
else:
print('I have learnt that already!')
d = Dog('Fido','<NAME>')
e = Dog('Buddy','Cocker')
f = Dog('Rex')
print(d.name)
print(d.race)
print(e.name)
print(e.race)
print(f.name)
print(f.race)
d.add_trick('Roll')
d.add_trick('Pretending dead')
d.tricks
d.add_trick('Ask for food')
d.tricks
e.tricks
f.add_trick('Bark')
f.tricks
# ### Creating a subclass using inheritance
# +
import time
class SuperDog(Dog):
def __init__(self, name, race):
Dog.__init__(self, name, race)
self.food = False
self.trained = False
self.last_meal = time.time()
def is_hungry(self):
if time.time() - self.last_meal < 20:
print('Not hungry')
else:
print(f'Yes, my last meal was {(time.time() - self.last_meal)/60:.2f} min. ago')
def train(self):
self.trained = True
def feed(self):
self.last_meal = time.time()
print('The food was delicious. Thanks!')
# -
f = SuperDog('Raghu','Labrador')
f.is_hungry()
f.is_hungry() #calling after some time...
f.feed()
f.is_hungry()
f.tricks
f.add_trick('Give Five')
f.tricks
f.trained
f.train()
f.trained
# ### [Multiple inheritance](https://www.techbeamers.com/python-multiple-inheritance/)
# +
class A:
def rk(self):
print(" In class A")
class B(A):
def rk(self):
print(" In class B")
class C(A):
def rk(self):
print("In class C")
# classes ordering
class D(B, C):
pass
r = D()
r.rk()
# -
# ### Another example:
class Animal:
def __init__(self, weight=0, height=0):
self.weight = weight
self.height = height
class Carnivore(Animal):
def __init__(self, weight=0, height=0):
super().__init__(weight, height)
def say(self):
raise NotImplementedError
TRex = Carnivore(2000)
TRex.weight
TRex.height
class Wolf(Carnivore):
def __init__(self, weight, height):
super().__init__(weight, height)
def say(self):
#print("Bark! Bark!")
return "Bark! Bark!"
Jack = Wolf(8, 45)
print(Jack.weight)
print(Jack.height)
print(Jack.say())
class Pet(Animal):
def __init__(self, tutor, weight=0, height=0):
super().__init__(weight, height)
self.tutor = tutor
# +
fish = Pet('John', 35)
print(fish.tutor)
print(fish.weight)
print(fish.height)
# -
class Cat(Carnivore, Pet):
def __init__(self, weight, height, tutor):
super(Carnivore, self).__init__(self, weight, height)
print(self.weight)
print(self.height)
Pet.__init__(self, tutor, self.weight, self.height)
print(self.tutor)
def say(self):
print("Meaw!")
# [C3 & MRO](https://en.wikipedia.org/wiki/C3_linearization)
Cat.__mro__
Garfield = Cat(4, 25, 'John')
print(f"Weight: {Garfield.weight}\nHeight: {Garfield.height}\nTutor: {Garfield.tutor}")
Garfield.say()
# ### Modifying an existing class
class MyInteger(int):
def __init__(self,number):
super().__init__()
def __add__(self, other):
return self * other
def __sub__(self, other):
return self + other
def square(self):
return self * self
a = MyInteger(2)
b = MyInteger(5)
dir(a)
a * b
a + b #?????
a - b
a.square()
b.square()
help(MyInteger.__mul__)
help(MyInteger.__str__)
# ### A more complete example:
# +
def strike(text):
""" Renders string with strike-through characters through it.
`strike('hello world')` -> '̶h̶e̶l̶l̶o̶ ̶w̶o̶r̶l̶d'
Notes
-----
\u0336 is a special strike-through unicode character; it
is not unique to Python."""
return ''.join('\u0336{}'.format(c) for c in text)
class ShoppingList:
def __init__(self, items):
self._needed = set(items)
self._purchased = set()
def __repr__(self):
""" Returns formatted shopping list as a string with
purchased items being crossed out.
Returns
-------
str"""
if self._needed or self._purchased:
remaining_items = [str(i) for i in self._needed]
purchased_items = [strike(str(i)) for i in self._purchased]
# You wont find the • character on your keyboard. I simply
# googled "unicode bullet point" and copied/pasted it here.
return "• " + "\n• ".join(remaining_items + purchased_items)
def add_new_items(self, items):
self._needed.update(items)
def mark_purchased_items(self, items):
self._purchased.update(set(items) & self._needed)
self._needed.difference_update(self._purchased)
# -
food = ShoppingList(["milk", "flour", "salt", "eggs"])
print(food)
food.mark_purchased_items(["flour", "salt"])
print(food)
# +
def __add__(self, other):
""" Add the unpurchased and purchased items from another shopping
list to the present one.
Parameters
----------
other : ShoppingList
The shopping list whose items we will add to the present one.
Returns
-------
ShoppingList
The present shopping list, with items added to it."""
new_list = ShoppingList([])
# populate new_list with items from `self` and `other`
for l in [self, other]:
new_list.add_new_items(l._needed)
# add purchased items to list, then mark as purchased
new_list.add_new_items(l._purchased)
new_list.mark_purchased_items(l._purchased)
return new_list
# set `__add__` as a method of `ShoppingList`
setattr(ShoppingList, "__add__", __add__)
# +
food = ShoppingList(["milk", "flour", "salt", "eggs"])
food.mark_purchased_items(["flour", "salt"])
office_supplies = ShoppingList(["staples", "pens", "pencils"])
office_supplies.mark_purchased_items(["pencils"])
clothes = ShoppingList(["t-shirts", "socks"])
# combine all three shopping lists
food + office_supplies + clothes
# -
class MyList:
def __init__(self, *args):
if len(args) == 1 and hasattr(args[0], '__iter__'):
# handles `MyList([1, 2, 3])
self._data = list(args[0])
else:
# handles `MyList(1, 2, 3)`
self._data = list(args)
def __getitem__(self, index):
out = self._data[index]
# slicing should return a `MyList` instance
# otherwise, the individual element should be returned as-is
return MyList(out) if isinstance(index, slice) else out
def __setitem__(self, key, value):
self._data[key] = value
def __len__(self):
return len(self._data)
def __repr__(self):
""" Use the character | as the delimiter for our list"""
# `self._data.__repr__()` returns '[ ... ]',
# thus we can slice to get the contents of the string
# and exclude the square-brackets, and add our own
# delimiters in their place
return "|" + self._data.__repr__()[1:-1] + "|"
def __contains__(self, item):
return item in self._data
def append(self, item):
self._data.append(item)
# MyList can accept any iterable as its
# first (and only) input argument
x = MyList("hello")
x
# MyList accepts an arbitrary number of arguments
x = MyList(1, 2, 3, 4, 5)
x
print(len(x))
# getting an item
x[0]
# slicing returns a MyList instance
x[2:4]
# setting an item
x[0] = -1
x
# checking membership
10 in x
MyList()
| Notebooks/09_Classes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
import pandas as pd
realdic=pd.read_csv("wordcountsreal5000.csv")#, usecols=["word", "count"])#index_col='Symbol', usecols=[0, 2, 3, 7])
#print(realdic)
list(realdic.columns.values)
#realdic['the']
#realdic['9418']
realdic['9418'].sum()
fakedic=pd.read_csv("wordcountsfake5000.csv")#, usecols=["word", "count"])#index_col='Symbol', usecols=[0, 2, 3, 7])
#print(fakedic)
#list(fakedic.columns.values)
fakedic['158487'].sum()
realdic.loc[:,'9418'] /= realdic['9418'].sum()
fakedic.loc[:,'158487'] /= fakedic['158487'].sum()
print(realdic.head())
print(realdic.head())
print(fakedic.head())
rld=realdic
fkd=fakedic
print(rld.head())
print(fkd.head())
rld=rld.rename(columns={'the': 'word', '9418': 'proportionreal'})
fkd=fkd.rename(columns={'the': 'word', '158487': 'proportionfake'})
print(rld.head())
print(fkd.head())
#rld.merge(fkd, on='word', how='left')
#join_df = rld.join(fkd,on='word',how='left')
#rld.join(fkd)
result = pd.merge(rld, fkd, on='word')
print(result.head())
result['diff']=(result['proportionreal']-result['proportionfake'])*1000
result
result=result.sort_values('diff')
print(result)
# +
import matplotlib.pyplot as plt
import matplotlib
#import numpy as np
bins = np.linspace(0, 10, 10)
plt.hist([1,2,1,1,1,9,9,9,8,8], bins, alpha = 0.5, color = 'b' , label = 'good')
plt.show()
# +
import matplotlib.pyplot as plt
newdic={'Clinton':2.290833,'Donald':0.910428,'political':0.729208,'American':0.708259,'FBI':0.667642,'war':0.630120,'election':0.586268}
# -
from matplotlib.pyplot import figure
figure(num=None, figsize=(8, 12), dpi=80, facecolor='w', edgecolor='k')
# +
import matplotlib.pyplot as plt
from matplotlib import pyplot as plt
from matplotlib.pyplot import figure
figure(num=None, figsize=(90, 12), dpi=280, facecolor='w', edgecolor='k')
counts=newdic
#data
data = [2.290833,0.910428,0.729208,0.708259,0.667642,0.630120,0.586268]
names = ['Clinton','Donald','political','American','FBI','war','election']
labels, heights = zip(*sorted(((k, v) for k, v in counts.items()), reverse=True))
fig, ax = plt.subplots(1, 1)
ax.bar(left, heights, 1)
ax.set_xticks(left + 0.5)
ax.set_xticklabels(labels, fontsize='large')
# +
from collections import Counter
x = ['spam', 'ham', 'eggs', 'ham', 'chips', 'eggs', 'spam', 'spam', 'spam']
counts = Counter(newdic)
print(counts)
from matplotlib import pyplot as plt
import numpy as np
# sort counts in descending order
labels, heights = zip(*sorted(((k, v) for k, v in counts.items()), reverse=True))
# lefthand edge of each bar
left = np.arange(len(heights))
fig, ax = plt.subplots(1, 1)
ax.bar(left, heights, 1)
ax.set_xticks(left + 0.5)
ax.set_xticklabels(labels, fontsize='large')
# +
from matplotlib import pyplot as plt
import numpy as np
# sort counts in descending order
labels, heights = zip(*sorted(((k, v) for k, v in counts.items()), reverse=True))
# lefthand edge of each bar
left = np.arange(len(heights))
fig, ax = plt.subplots(1, 1)
ax.bar(left, heights, 1)
ax.set_xticks(left + 0.5)
ax.set_xticklabels(labels, fontsize='large')
# +
import matplotlib.pyplot as plt
def combined_label(perc, tot):
"""
Format a label to include by Euros and %.
"""
return "{0:,.0f}k EUR, {1:.0f}%".format(perc * tot / 1000, perc * 100)
def cost_cum(data, focus, subject):
"""
Accumulate the stats.
- data is a DataFrame,
- focus is the colum to group by
- subject is the column to aggregate.
"""
# Setup data frame
parts = data[[focus, 'cost']].groupby(focus).sum().sort(subject, ascending=False)
parts['percent'] = parts['cost'] / parts.cost.sum()
parts['cum_percent'] = parts['percent'].cumsum()
return parts
def cost_pareto(data, focus_name, limit_percent = 0.75):
# Filter and organize the data frame
top_parts = data[data['cum_percent'] < limit_percent]
top_parts.set_index(top_parts['percent'])
# Draw the plots
fig = plt.figure(figsize=(10,7))
fig.subplots_adjust(bottom=0.4, left=0.15)
ax = fig.add_subplot(1,1,1)
top_parts['cum_percent'].plot(ax=ax, color="k", drawstyle="steps-post")
top_parts['percent'].plot(ax=ax, kind="bar", color="k", alpha=0.5)
ax.set_ylim(bottom=0, top=1)
tick_nums = [x/float(100) for x in range(0,101,20)]
ax.set_yticks(tick_nums)
tot_cost = top_parts['cost'].sum()
ax.set_yticklabels([combined_label(x, tot_cost) for x in tick_nums])
ax.set_title("Top %s%% of Cost Split By %s" % (int(limit_percent * 100), focus))
ax.set_xlabel("")
return ax
accumulated = cost_cum(data, 'Part', 'Cost')
chart = cost_pareto(accumulated, 'Part', 0.9)
# -
from paretochart import pareto
| mergedict.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import torch as t
import sys
sys.path.append('../')
from utils import get_score
import glob
import torch as t
import numpy as np
import json
import time
label_path = '/home/a/code/pytorch/zhihu/ddd/labels.json'
test_data_path='/home/a/code/pytorch/zhihu/ddd/test.npz'
index2qid = np.load(test_data_path)['index2qid'].item()
with open(label_path) as f:
labels_info = json.load(f)
qid2label = labels_info['d']
label2qid = labels_info['id2label']
files = glob.glob('../checkpoints/result/*test.pth')
r2=0
for file in files:
if 'MultiModel' not in file:
print file
r2+=t.load(file)
files = glob.glob('../checkpoints/result/tmp/*test.pth')
r_aug=0
for file in files:
if 'MultiModel' not in file and 'weight5' not in file:
print file
r_aug+=t.load(file)
files = glob.glob('../checkpoints/result/tmp/*test.pth')
r_multi=0
for file in files:
if 'MultiModel' in file :
print file
r_multi+=t.load(file)
files = glob.glob('../checkpoints/result/tmp/*test.pth')
r_5=0
for file in files:
if 'weight5' in file :
print file
r_5+=t.load(file)
tmp = (r2+r_aug*1.15+r_multi*13+r_5*0.15)
result=(tmp).topk(5,1)[1]
# +
# 手动调整权重
# tmp = tmp-t.load('../checkpoints/result/DeepText0.4103_word_test.pth')
# result=(tmp-t.load('../checkpoints/result/tmp/RCNN_0.41226_weight5_test.pth')*0.005).topk(5,1)[1]
# tmp = tmp+t.load('../checkpoints/result/DeepText0.4103_word_test.pth')*0.2
# tmp = tmp-t.load('../checkpoints/result/tmp/LSTMText0.41368_aug_word_test.pth')*0.1
# tmp = tmp-t.load('../checkpoints/result/CNNText_tmp0.4024_char_test.pth')*0.05
# tmp = tmp-t.load('../checkpoints/result/tmp/MultiCNNTextBNDeep_word_weight5_0.409409_test.pth')*0.009
# -
## 写csv 提交结果
rows = range(result.size(0))
for ii,item in enumerate(result):
rows[ii] = [index2qid[ii]] + [label2qid[str(_)] for _ in item ]
import csv
with open('final-0.43081.csv','w') as f:
writer = csv.writer(f)
writer.writerows(rows)
| notebooks/test_ensemble.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %pylab inline
# input image dimensions
img_rows, img_cols = 28, 28
# Uncomment the following lines if you have keras installed. Otherwise you can
# use the file I uploaded: mnist.npz
import keras
from keras.datasets import mnist
from keras import backend as K
# the data, split between train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
if K.image_data_format() == 'channels_first':
x_train = x_train.reshape(x_train.shape[0], 1, img_rows, img_cols)
x_test = x_test.reshape(x_test.shape[0], 1, img_rows, img_cols)
input_shape = (img_rows, img_cols)
else:
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols)
input_shape = (img_rows, img_cols)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
np.savez_compressed('mnist.npz', x_train, y_train, x_test, y_test)
arc = load('mnist.npz')
x_train = arc['arr_0']
y_train = arc['arr_1']
x_test = arc['arr_2']
y_test = arc['arr_3']
print(x_train.shape, y_train.shape)
print(x_test.shape, y_test.shape)
| final_project/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 ('tf')
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from sklearn.tree import DecisionTreeClassifier
from sklearn.tree import plot_tree
import matplotlib.pyplot as plt
# %matplotlib inline
# # Reading Data
#Reading our data
df=pd.read_csv('https://raw.githubusercontent.com/aamir09/Machine-Learning-/main/data/dtree.csv')
df.head()
df.shape
# # Seperating Data
#Seperate data in feature and target variable
X=df.drop(['Diabetic'],axis=1)
y=df.Diabetic
# # Creating & Plotting Decision Tree
#Creating a decision tree classifier with max-depth 2
dtree=DecisionTreeClassifier(max_depth=2)
dtree.fit(X,y)
#Plotting our tree
fig,ax=plt.subplots(dpi=100,figsize=(20,6))
plot_tree(dtree,max_depth=2,feature_names=['DBP','Age'],class_names=['Non-Diabetic','Diabetic'])
fig.savefig('DecisionTree.jpeg')
| DecisonTree.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Motivation
# + [markdown] slideshow={"slide_type": "slide"}
# ## Why computer modelling?
# Because it is **cheaper** than real-life experiment, or in the case when real-life experiment **is not possible**.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Typical steps of computer modelling
#
# 1. Formulate the mathematical problem as an **equation** for some **quantities**
#
# 2. Replace the continious problem by a discrete one (**discretization**)
#
# 3. Solve the resulting discrete problem
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# The simplest cycle:
# Mathematical model $\to$ Discretization $\to$ Solve
# + [markdown] slideshow={"slide_type": "slide"}
# ## Discretization
# The discretization is replacement of the region by discrete elements:
# <img src="pic/airplane2.jpeg">
# + [markdown] slideshow={"slide_type": "slide"}
# ## Random notes
# - Discretization and solve can be connected
# - Fast solvers are needed
# - Only a subproblem in **design and optimization**
# - Many physical problems are still too complex for math (turbulence!)
# + [markdown] slideshow={"slide_type": "slide"}
# Consider
# <img src="pic/weather.jpeg">
#
# It takes a lot to create
#
# 1. A model
# 2. Discretization
# 3. Solvers
#
# What if the computer time to compute a **forecast for 1 day is more
# than 1 day**?
# + [markdown] slideshow={"slide_type": "slide"}
# Many **process in physics** are modelled as PDEs.
#
# - Diffusion processes (heat transfer), electrostatics (circuit design) Poisson equation
# - Sound propagation (noise on the streets, buildings) – Helmholtz
# equation
# - Electromagnetics – MRI (magnetic resonance imaging) –
# Maxwell equations
# - Fluid flows – Stokes / Navier Stokes equations
#
# These are all partial differential equations!
# + [markdown] slideshow={"slide_type": "slide"}
# PDEs appear in many areas, including
# - Modelling of physical processes (heat, elasticity, fluid flows)
# - Financial math (Black Scholes equation)
# - Chemical engineering (Smoluchowsky equation)
# - Nanoworld (Schrodinger equation)
# - Optimal control of robots (Hamilton-Jacobi-Bellman equation)
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Why do we need fast methods?
# Because the growth of the computer power, being exponential, is still not enough!
#
# The growth due to the **algorithm improvements** is comparable (and the human brain is still the most energy-efficient computing element)
#
# <img src='pic/moore.png'>
# [Source](http://www.variousconsequences.com/2014/01/algorithmic-improvements-important-as-moores-law.html)
# + [markdown] slideshow={"slide_type": "slide"}
# ## What do we mean by fast methods?
# By fast methods we mean **improving the asymptotics** with respect to the problem size.
#
# Consider solution of linear system with a **sparse matrix** $A$:
#
# $$Au = b,$$
#
# where $A$ is a 5-point Laplacian discretization:
#
# $$\frac{u_{i+1, j} + u_{i-1, j} + u_{i, j-1} + u_{i, j+1} - 4 u_{ij} }{h^2} = f_{ij}.$$
#
# What are the complexities (next slide, but let us guess).
# + [markdown] slideshow={"slide_type": "slide"}
# ## Complexity (<font color='red'> essentials </font>)
#
# - Dense Gaussian elimination: $\mathcal{O}(N^3)$, works up to $10^4$
# - Sparse Gaussian elimination: $\mathcal{O}(N^{\frac{3}{2}})$, works up to $10^6$
# - FFT methods: $\mathcal{O}(N \log N)$, up to $10^8$
# - Multigrid method: $\mathcal{O}(N)$, up to $10^8$
# - Tensor methods for **particular** right-hand sides (for example, $f=1$), works up to **astronomically large sizes** ($N = 2^{50}$).
# + [markdown] slideshow={"slide_type": "slide"}
# ## Integral equations
# Now, to integral equations!
#
# -
# ## Physics is described by PDEs
#
# The physics of our world is typically described by local conservation laws, expressed in terms of partial differential equations.
#
# The **Poisson** equation writes as
#
# $$\Delta u \equiv \text{div} \nabla u = f,$$
#
# plus boundary conditions.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Model problem (electrostatics)
# Suppose physical setting: we have an ideally conducting surface $\Omega$ (for example, surface of a cube), which is attached to a battery.
#
# The charges can appear only at the surface, i.e.
#
# $$\Delta V(x) = 0, \quad x\not\in \partial \Omega$$
# but at the surface the potential should be constant:
#
# $$V(x) = V_0, \quad x\in \partial\Omega$$
#
# - This is a classical example of **external problem**.
#
# - The potential has to be defined in the **full** $\mathbb{R}^3$ space.
#
# - It is quite expensive, boundary conditions on the outer boundary are not straightforward.
# + [markdown] slideshow={"slide_type": "slide"}
# ## From electrostatics to integral equations
#
# The concept of **equivalent sources** leads to the **boundary integral formulation** of the problem.
#
# The charges can appear only at the boundary. The charge creates the field $\frac{1}{r}$.
#
# In $\mathbb{R}^3$ function $G(x,y) =\frac{1}{4\pi\|x-y\|}$ is **fundamental solution** of the operator $\Delta$, since it satisfies
#
# $$\Delta G (x,y) = \delta(x-y),$$
#
# where $\delta$ is a **delta-function.**
# + [markdown] slideshow={"slide_type": "slide"}
# ## BEM
#
# The boundary integral equation comes from the idea to **look for** the solution as
#
# $$V(x) = \int_{\partial \Omega} \frac{q(y)}{\Vert x - y\Vert} dy.$$
#
# (it is also called **single-layer potential**).
# + [markdown] slideshow={"slide_type": "slide"}
# ## Properties
# We have
# $$\Delta V(x) = \int_{\partial \Omega} q(y) \Delta_x\left(\frac{1}{\Vert x - y\Vert}\right) dy = 4\pi\int_{\partial \Omega} q(y) \delta (x-y) dy = (\text{why?}) = 0, \quad x\not\in\partial\Omega$$
#
# therefore it is sufficient to find the unknown function $q$ that satisfies the **Dirichlet boundary condition**
#
# $$\int_{\partial \Omega} \frac{q(y)}{\Vert x - y\Vert} dy = V_0, \quad x \in \partial \Omega$$
#
# That is the **first kind integral equation with singular kernel.**
#
# The main benefit is that the unknown function is defined only on $\partial \Omega$!
#
# However, the operator is "non-local" compared with the PDE formulation.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Model problem: acoustics
#
# Room acoustics (for opera houses), noise assessment (for roads, building construction, railways) can be modelled in the **same fashion**
#
# <img src='pic/stravinsky.jpg' width=550>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Model problem: acoustics
#
# The underlying equation is the **Helmholtz equation**
#
# $$\Delta p + k^2 p = f, $$
#
# plus boundary conditions (typically, **Neumann boundary conditions**), and $f$ are **sound sources** (typically, point sources).
#
# The fundamental solution is
#
# $$G(x,y) = \frac{\exp(i k \Vert x - y \Vert)}{\Vert x - y \Vert}.$$
# + [markdown] slideshow={"slide_type": "slide"}
# ## Summary
#
# - Intro lecture
# - First integral equations
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Next lecture
# - How to discretize IE (Nystrom, collocation, Galerkin method, other type of kernels)
# - What are the problems with such discretization.
# + slideshow={"slide_type": "skip"}
from IPython.core.display import HTML
def css_styling():
styles = open("./styles/custom.css", "r").read()
return HTML(styles)
css_styling()
| lectures/Lecture-1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
cpi = pd.read_excel('Core CPI, seas. adj..xlsx', sheet = 1)
sa_cpi = cpi['South Africa'] #south afrian consumer price index
xrate = pd.read_excel('Real Effective Exchange Rate.xlsx', sheet =1)
sa_xrate = xrate['South Africa'] #south african real effective exchange rate
xport = pd.read_excel('Exports Merchandise, Customs, current US$, millions, seas. adj..xlsx', sheet =1)
sa_xport = xport['South Africa'] #south african exports
fres = pd.read_excel('Foreign Reserves, Months Import Cover, Goods.xlsx', sheet =1)
sa_fres = fres['South Africa'] #SA foreign reserves
imports = pd.read_excel('Imports Merchandise, Customs, current US$, millions, seas. adj..xlsx', sheet =1)
sa_imports = imports['South Africa'] #SA imports
industrial = pd.read_excel('Industrial Production, constant 2010 US$, seas. adj..xlsx', sheet =1)
sa_industrials = industrial['South Africa'] # SA industrial production
stocks = pd.read_excel('Stock Markets, US$.xlsx', sheet =1)
sa_stocks = stocks['South Africa'] #SA stock markets
tres = pd.read_excel('Total Reserves.xlsx', sheet =1)
sa_tres = tres['South Africa'] #SA total reserves
nomxrate = pd.read_excel('Nominal Effecive Exchange Rate.xlsx', sheet=1)
sa_nomxrate = nomxrate['South Africa'] # SA nominal effective exchange rate
xchange = sa_xrate.drop(sa_xrate.iloc[0])
cpindex = sa_cpi.drop(sa_cpi.iloc[0])
cpindex
exports = sa_xport.drop(sa_xport.iloc[0])
exports
final_imports = sa_imports.drop(sa_imports.iloc[0])
final_imports
industry = sa_industrials.drop(sa_industrials.iloc[0])
industry
stock_market = sa_stocks.drop(sa_stocks.iloc[0])
stock_market
total_reserves = sa_tres.drop(sa_tres.iloc[0])
total_reserves
full_data = pd.DataFrame({'Total Reserves':total_reserves, 'Stock Market':stock_market,'Imports': final_imports,'Exports':exports, 'Consumer Price Index':cpindex, 'Industrial Production':industry, 'Real Effective Exchange rate':xchange})
full_data
#from 2002 to 2017
data = full_data.drop(full_data.index[[0,1,2,3,4,5,6,7,8,9,10,11,12]])
data
final_data = data.drop(data.index[16])
final_data
from sklearn.linear_model import LinearRegression as lr
import sklearn
X = final_data.drop('Real Effective Exchange rate', axis = 1)
y = final_data['Real Effective Exchange rate']
X_train, X_test, y_train, y_test = sklearn.model_selection.train_test_split(X, y, test_size = 0.25, random_state = 7)
X_train.shape
X_test.shape
y_train.shape
y_test.shape
md = lr()
md.fit(X_train, y_train)
y_pred = md.predict(X_test)
md.coef_
sklearn.metrics.r2_score(y_test, y_pred)
| Economic_Indicators_SA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generating counterfactuals for multi-class classification and regression models
# This notebook will demonstrate how the DiCE library can be used for multiclass classification and regression for scikit-learn models.
# You can use any method ("random", "kdtree", "genetic"), just specific it in the method argument in the initialization step. The rest of the code is completely identical.
# For demonstration, we will be using the genetic algorithm for CFs.
# +
import dice_ml
from dice_ml import Dice
from sklearn.datasets import load_iris, fetch_california_housing
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from sklearn.model_selection import train_test_split
from sklearn.compose import ColumnTransformer
from sklearn.ensemble import RandomForestClassifier, RandomForestRegressor
import pandas as pd
# -
# %load_ext autoreload
# %autoreload 2
# We will use sklearn's internal datasets to demonstrate DiCE's features in this notebook
# ## Multiclass Classification
# For multiclass classification, we will use sklearn's Iris dataset. This data set consists of 3 different types of irises’ (Setosa, Versicolour, and Virginica) petal and sepal length. More information at https://scikit-learn.org/stable/datasets/toy_dataset.html#iris-plants-dataset
df_iris = load_iris(as_frame=True).frame
df_iris.head()
df_iris.info()
outcome_name = "target"
continuous_features_iris = df_iris.drop(outcome_name, axis=1).columns.tolist()
target = df_iris[outcome_name]
# +
# Split data into train and test
datasetX = df_iris.drop(outcome_name, axis=1)
x_train, x_test, y_train, y_test = train_test_split(datasetX,
target,
test_size=0.2,
random_state=0,
stratify=target)
categorical_features = x_train.columns.difference(continuous_features_iris)
# We create the preprocessing pipelines for both numeric and categorical data.
numeric_transformer = Pipeline(steps=[
('scaler', StandardScaler())])
categorical_transformer = Pipeline(steps=[
('onehot', OneHotEncoder(handle_unknown='ignore'))])
transformations = ColumnTransformer(
transformers=[
('num', numeric_transformer, continuous_features_iris),
('cat', categorical_transformer, categorical_features)])
# Append classifier to preprocessing pipeline.
# Now we have a full prediction pipeline.
clf_iris = Pipeline(steps=[('preprocessor', transformations),
('classifier', RandomForestClassifier())])
model_iris = clf_iris.fit(x_train, y_train)
# +
d_iris = dice_ml.Data(dataframe=df_iris,
continuous_features=continuous_features_iris,
outcome_name=outcome_name)
# We provide the type of model as a parameter (model_type)
m_iris = dice_ml.Model(model=model_iris, backend="sklearn", model_type='classifier')
# -
exp_genetic_iris = Dice(d_iris, m_iris, method="genetic")
# As we can see below, all the target values will lie in the desired class
# Single input
query_instances_iris = x_test[2:3]
genetic_iris = exp_genetic_iris.generate_counterfactuals(query_instances_iris, total_CFs=7, desired_class=2)
genetic_iris.visualize_as_dataframe()
# Multiple queries can be given as input at once
query_instances_iris = x_test[17:19]
genetic_iris = exp_genetic_iris.generate_counterfactuals(query_instances_iris, total_CFs=7, desired_class=2)
genetic_iris.visualize_as_dataframe(show_only_changes=True)
# # Regression
# For regression, we will use sklearn's California Housing dataset. This dataset contains California house prices. More information at https://scikit-learn.org/stable/modules/generated/sklearn.datasets.fetch_california_housing.html
housing_data = fetch_california_housing()
df_housing = pd.DataFrame(housing_data.data, columns=housing_data.feature_names)
df_housing[outcome_name] = pd.Series(housing_data.target)
df_housing.head()
df_housing.info()
continuous_features_housing = df_housing.drop(outcome_name, axis=1).columns.tolist()
target = df_housing[outcome_name]
# +
# Split data into train and test
datasetX = df_housing.drop(outcome_name, axis=1)
x_train, x_test, y_train, y_test = train_test_split(datasetX,
target,
test_size=0.2,
random_state=0)
categorical_features = x_train.columns.difference(continuous_features_housing)
# We create the preprocessing pipelines for both numeric and categorical data.
numeric_transformer = Pipeline(steps=[
('scaler', StandardScaler())])
categorical_transformer = Pipeline(steps=[
('onehot', OneHotEncoder(handle_unknown='ignore'))])
transformations = ColumnTransformer(
transformers=[
('num', numeric_transformer, continuous_features_housing),
('cat', categorical_transformer, categorical_features)])
# Append classifier to preprocessing pipeline.
# Now we have a full prediction pipeline.
regr_housing = Pipeline(steps=[('preprocessor', transformations),
('regressor', RandomForestRegressor())])
model_housing = regr_housing.fit(x_train, y_train)
# -
d_housing = dice_ml.Data(dataframe=df_housing, continuous_features=continuous_features_housing, outcome_name=outcome_name)
# We provide the type of model as a parameter (model_type)
m_housing = dice_ml.Model(model=model_housing, backend="sklearn", model_type='regressor')
exp_genetic_housing = Dice(d_housing, m_housing, method="genetic")
# As we can see below, all the target values will lie in the desired range
# Multiple queries can be given as input at once
query_instances_housing = x_test[2:4]
genetic_housing = exp_genetic_housing.generate_counterfactuals(query_instances_housing,
total_CFs=2,
desired_range=[3.0, 5.0])
genetic_housing.visualize_as_dataframe(show_only_changes=True)
| docs/source/notebooks/DiCE_multiclass_classification_and_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="dUeKVCYTbcyT"
# #### Copyright 2019 The TensorFlow Authors.
# + cellView="form" id="4ellrPx7tdxq"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="7JfLUlawto_D"
# # 불균형 데이터 분류
# + [markdown] id="DwdpaTKJOoPu"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.google.com/"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />TensorFlow.org에서 보기</a>
# </td>
# <td>
# <a target="_blank" href="https://www.google.com/"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Google Colab에서 실행하기</a>
# </td>
# <td>
# <a target="_blank" href="https://www.google.com/"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />GitHub에서 소스 </a>
# </td>
# <td>
# <a href="https://www.google.com/"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="mthoSGBAOoX-"
# 이 튜토리얼에서는 한 클래스의 예시의 수가 다른 클래스보다 훨씬 많은 불균형 데이터세트를 분류하는 방법을 소개합니다. Kaggle에서 호스팅 되는 [신용 카드 부정 행위 탐지 데이터세트](https://www.kaggle.com/mlg-ulb/creditcardfraud)를 사용하여 작업할 것입니다. 총 284,807건의 거래에서 492건의 부정거래만 적발하는 것이 목적입니다. [Keras](../../guide/keras/overview.ipynb)를 사용하여 모델 및 [클래스 가중치](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/keras/Model)를 정의하여 불균형 데이터로부터 모델을 학습할 수 있도록 할 것입니다.
#
# 이 튜토리얼에서는 다음의 완전한 코드가 포함되어있습니다.:
#
# * Pandas를 사용하여 CSV 파일 로드.
# * 학습, 검증 및 테스트세트 작성.
# * Keras를 사용하여 모델을 정의하고 학습시키기(클래스 가중치 설정 포함).
# * 다양한 측정 기준(정밀도 및 재현 율 포함)을 사용하여 모델을 평가한다.
# * 불균형 데이터를 처리하기 위한 다음과 같은 기술을 사용해보십시오:
# * 클래스 가중치
# * 오버샘플링
#
# + [markdown] id="kRHmSyHxEIhN"
# ## 설정
# + id="JM7hDSNClfoK"
import tensorflow as tf
from tensorflow import keras
import os
import tempfile
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import sklearn
from sklearn.metrics import confusion_matrix
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
# + id="c8o1FHzD-_y_"
mpl.rcParams['figure.figsize'] = (12, 10)
colors = plt.rcParams['axes.prop_cycle'].by_key()['color']
# + [markdown] id="Z3iZVjziKHmX"
# ## 데이터 처리 및 탐색
# + [markdown] id="4sA9WOcmzH2D"
# ### Kaggle 신용 카드 부정 행위 데이터 세트
#
# Pandas는 구조화된 데이터를 로드하고 작업하는데 유용한 유틸리티가 많이 있는 Python 라이브러리로서 CSV를 데이터 프레임으로 다운로드 하는데 사용할 수 있다.
#
# 참고: 이 데이터세트는 큰 데이터 마이닝 및 부정 행위 감지에 대한 Worldline과 ULB의 [Machine Learning Group](http://mlg.ulb.ac.be) (Université Libre de Bruxelles)의 연구 협업을 통해 수집 및 분석 되었다. 관련 주제에 대한 현재 및 과거의 프로젝트에 대한 자세한 내용은 [여기](https://www.researchgate.net/project/Fraud-detection-5) and the page of the [DefeatFraud](https://mlg.ulb.ac.be/wordpress/portfolio_page/defeatfraud-assessment-and-validation-of-deep-feature-engineering-and-learning-solutions-for-fraud-detection/)에서 확인할 수 있으며 DefeatFraud 프로젝트 페이지도 참조하십시오
# + id="pR_SnbMArXr7"
file = tf.keras.utils
raw_df = pd.read_csv('https://storage.googleapis.com/download.tensorflow.org/data/creditcard.csv')
raw_df.head()
# + id="-fgdQgmwUFuj"
raw_df[['Time', 'V1', 'V2', 'V3', 'V4', 'V5', 'V26', 'V27', 'V28', 'Amount', 'Class']].describe()
# + [markdown] id="xWKB_CVZFLpB"
# ### 클래스 레이블 불균형 조사
#
# 데이터세트 불균형을 살펴보겠습니다.:
# + id="HCJFrtuY2iLF"
neg, pos = np.bincount(raw_df['Class'])
total = neg + pos
print('Examples:\n Total: {}\n Positive: {} ({:.2f}% of total)\n'.format(
total, pos, 100 * pos / total))
# + [markdown] id="KnLKFQDsCBUg"
# 이것은 양성 샘플의 작은 부분을 보여줍니다.
# + [markdown] id="6qox6ryyzwdr"
# ### 데이터 정리, 분할 및 정규화
#
# 원시 데이터에는 몇 가지 문제가 있습니다. `Time` 과 `Amount` 열은 너무 가변적이기 때문에 직접적으로 사용할 수 없습니다. 우선 `Time` 열을 삭제한 뒤에 (의미가 명확하지 않아서) `amount` 열의 로그를 가져와서 범위를 줄입니다.
# + id="Ef42jTuxEjnj"
cleaned_df = raw_df.copy()
# You don't want the `Time` column.
cleaned_df.pop('Time')
# The `Amount` column covers a huge range. Convert to log-space.
eps = 0.001 # 0 => 0.1¢
cleaned_df['Log Ammount'] = np.log(cleaned_df.pop('Amount')+eps)
# + [markdown] id="uSNgdQFFFQ6u"
# 데이터 세트를 학습, 검증 및 테스트 세트로 분할합니다. 검증 세트는 모델 피팅 중에 손실 및 메트릭을 평가하는 데 사용되지만 모델이 이 데이터에 적합하지 않습니다. 테스트 세트는 훈련 단계에서 완전히 사용되지 않으며 모델이 새 데이터로 얼마나 잘 일반화되는지 평가하기 위해 마지막에만 사용됩니다. 이는 훈련 데이터 부족으로 인하여 [오버피팅](https://developers.google.com/machine-learning/crash-course/generalization/peril-of-overfitting) 이 중요한 문제인 데이터 세트에서 특히 더 중요합니다.
# + id="xfxhKg7Yr1-b"
# Use a utility from sklearn to split and shuffle our dataset.
train_df, test_df = train_test_split(cleaned_df, test_size=0.2)
train_df, val_df = train_test_split(train_df, test_size=0.2)
# Form np arrays of labels and features.
train_labels = np.array(train_df.pop('Class'))
bool_train_labels = train_labels != 0
val_labels = np.array(val_df.pop('Class'))
test_labels = np.array(test_df.pop('Class'))
train_features = np.array(train_df)
val_features = np.array(val_df)
test_features = np.array(test_df)
# + [markdown] id="8a_Z_kBmr7Oh"
# sklearn StandardScaler를 사용하여 입력 기능을 정규화 합니다.
# 이것은 평균은 0으로, 표준 편차는 1로 설정합니다.
#
# 참고: StandardScaler는 오직 모델이 validation 또는 test set를 peeking 하지는 않았는지 확인하기 위해 train_feature를 사용할 때 적합합니다.
# + id="IO-qEUmJ5JQg"
scaler = StandardScaler()
train_features = scaler.fit_transform(train_features)
val_features = scaler.transform(val_features)
test_features = scaler.transform(test_features)
train_features = np.clip(train_features, -5, 5)
val_features = np.clip(val_features, -5, 5)
test_features = np.clip(test_features, -5, 5)
print('Training labels shape:', train_labels.shape)
print('Validation labels shape:', val_labels.shape)
print('Test labels shape:', test_labels.shape)
print('Training features shape:', train_features.shape)
print('Validation features shape:', val_features.shape)
print('Test features shape:', test_features.shape)
# + [markdown] id="XF2nNfWKJ33w"
# 주의: 모델을 배포하려면 전처리 계산을 유지하는 것이 중요합니다. 레이어로 구현하고 내보내기 전에 모델에 연결하는 것이 가장 쉬운 방법입니다.
#
# + [markdown] id="uQ7m9nqDC3W6"
# ### 데이터 분포 살펴보기
#
# 다음으로 몇 가지 기능에 대한 긍정 및 부정 예제의 분포를 비교하십시오. 이 때 스스로에게 물어볼 좋은 질문은 다음과 같습니다.:
#
# * 이러한 분포가 의미가 있습니까?
# * 예. 이미 입력을 정규화했으며 대부분 `+/- 2` 범위에 밀집되어 있습니다.
# * 분포의 차이를 볼 수 있습니까?
# * 예. 긍정적인 예는 그렇지 않은 것 보다 훨씬 더 높은 극단적인 값을 포함합니다.
# + id="raK7hyjd_vf6"
pos_df = pd.DataFrame(train_features[ bool_train_labels], columns=train_df.columns)
neg_df = pd.DataFrame(train_features[~bool_train_labels], columns=train_df.columns)
sns.jointplot(pos_df['V5'], pos_df['V6'],
kind='hex', xlim=(-5,5), ylim=(-5,5))
plt.suptitle("Positive distribution")
sns.jointplot(neg_df['V5'], neg_df['V6'],
kind='hex', xlim=(-5,5), ylim=(-5,5))
_ = plt.suptitle("Negative distribution")
# + [markdown] id="qFK1u4JX16D8"
# ## 모델 및 메트릭 정의
#
# 촘촘하게 연결된 히든 레이어, 과적합을 줄이기 위한 [drop out](https://developers.google.com/machine-learning/glossary/#dropout_regularization) 레이어, 거래 사기 가능성을 반환하는 출력 sigmoid 레이어로 간단한 신경망을 생성하는 함수를 정의합니다. :
# + id="3JQDzUqT3UYG"
METRICS = [
keras.metrics.TruePositives(name='tp'),
keras.metrics.FalsePositives(name='fp'),
keras.metrics.TrueNegatives(name='tn'),
keras.metrics.FalseNegatives(name='fn'),
keras.metrics.BinaryAccuracy(name='accuracy'),
keras.metrics.Precision(name='precision'),
keras.metrics.Recall(name='recall'),
keras.metrics.AUC(name='auc'),
]
def make_model(metrics=METRICS, output_bias=None):
if output_bias is not None:
output_bias = tf.keras.initializers.Constant(output_bias)
model = keras.Sequential([
keras.layers.Dense(
16, activation='relu',
input_shape=(train_features.shape[-1],)),
keras.layers.Dropout(0.5),
keras.layers.Dense(1, activation='sigmoid',
bias_initializer=output_bias),
])
model.compile(
optimizer=keras.optimizers.Adam(lr=1e-3),
loss=keras.losses.BinaryCrossentropy(),
metrics=metrics)
return model
# + [markdown] id="SU0GX6E6mieP"
# ### 유용한 메트릭 이해
#
# 위에서 정의한 몇 가지 지표는 성능을 평가할 때 도움이 될 모델에 의해 계산될 수 있다는 점에 유의하십시오.
#
#
#
# * **거짓** 음성 그리고 **거짓** 양성은 **잘못** 분류된 샘플입니다.
# * **참** 음성 그리고 **참** 양성은 **제대로** 분류된 샘플입니다.
# * **정확도** 는 올바르게 분류된 예제의 비율입니다.
# > $\frac{\text{true samples}}{\text{total samples}}$
# * **정밀도** 는 올바르게 분류된 **예측** 긍정 비율입니다.
# > $\frac{\text{true positives}}{\text{true positives + false positives}}$
# * **재현 율** 은 올바르게 분류된 **실제** 긍정 비율입니다.
# > $\frac{\text{true positives}}{\text{true positives + false negatives}}$
# * **AUC** 는 수신자 조작 특성 곡선 아래 영역(ROC-AUC)을 나타냅니다. 이 메트릭은 분류기가 무작위 음성 샘플보다 무작위 양성 샘플의 순위를 매길 확률과 동일합니다.
#
# 참고: 정확도는 이 작업에 유용한 측정 항목이 아닙니다. 항상 False를 예측해야 이 작업에서 99.8% 이상의 정확도를 얻을 수 있습니다.
#
# Read more:
# * [참 vs. 거짓 and 긍정 vs. 부정](https://developers.google.com/machine-learning/crash-course/classification/true-false-positive-negative)
# * [정확성](https://developers.google.com/machine-learning/crash-course/classification/accuracy)
# * [정밀도와 재현율](https://developers.google.com/machine-learning/crash-course/classification/precision-and-recall)
# * [ROC-AUC](https://developers.google.com/machine-learning/crash-course/classification/roc-and-auc)
# + [markdown] id="FYdhSAoaF_TK"
# ## 기준 모델
# + [markdown] id="IDbltVPg2m2q"
# ### 모델 구축
#
# 이제 이전에 정의한 함수를 통해서 모델을 만들고 학습시키십시오. 모델의 크기가 기본 배치 크기인 2048보다 큰 배치 크기를 사용하여야 적합한 것을 유의하십시오. 이는 각 배치에서 몇 개의 양성 샘플을 포함할 수 있는 적절한 기회를 확보하는데 있어서 중요하다.
#
#
# 참고: 이 모델은 클래스의 불균형을 잘 다루지 못합니다. 이를 이 튜토리얼의 뒷부분에서 개선하게 될 겁니다.
# + id="ouUkwPcGQsy3"
EPOCHS = 100
BATCH_SIZE = 2048
early_stopping = tf.keras.callbacks.EarlyStopping(
monitor='val_auc',
verbose=1,
patience=10,
mode='max',
restore_best_weights=True)
# + id="1xlR_dekzw7C"
model = make_model()
model.summary()
# + [markdown] id="Wx7ND3_SqckO"
# Test run the model:
# + id="LopSd-yQqO3a"
model.predict(train_features[:10])
# + [markdown] id="YKIgWqHms_03"
# ### 선택사항: 올바른 초기 바이어스를 설정합니다.
# + [markdown] id="qk_3Ry6EoYDq"
# 이러한 초기 추측은 좋지 못합니다. 데이터 세트가 불균형 하다는 것을 알고 있습니다. 그렇다면 이를 반영하도록 출력 계층의 바이어스를 설정합니다. (참조: [신경망 훈련을 위한 레시피: "init well"](http://karpathy.github.io/2019/04/25/recipe/#2-set-up-the-end-to-end-trainingevaluation-skeleton--get-dumb-baselines)). 이것은 초기 수렴에 도움이 될 수 있습니다.
# + [markdown] id="PdbfWDuVpo6k"
# 기본 바이어스 초기화를 사용하면 손실은 약 `math.log(2) = 0.69314`
# + id="H-oPqh3SoGXk"
results = model.evaluate(train_features, train_labels, batch_size=BATCH_SIZE, verbose=0)
print("Loss: {:0.4f}".format(results[0]))
# + [markdown] id="hE-JRzfKqfhB"
# 설정할 올바른 바이어스는 다음에서 파생 가능합니다.:
#
# $$ p_0 = pos/(pos + neg) = 1/(1+e^{-b_0}) $$
# $$ b_0 = -log_e(1/p_0 - 1) $$
# $$ b_0 = log_e(pos/neg)$$
# + id="F5KWPSjjstUS"
initial_bias = np.log([pos/neg])
initial_bias
# + [markdown] id="d1juXI9yY1KD"
# 이를 초기 바이어스로 설정하면 모델이 훨씬 더 합리적인 초기 추측을 제공합니다.
#
# 가까워 야합니다.: `pos/total = 0.0018`
# + id="50oyu1uss0i-"
model = make_model(output_bias=initial_bias)
model.predict(train_features[:10])
# + [markdown] id="4xqFYb2KqRHQ"
# 이 초기화를 통해서 초기 손실은 대략 다음과 같아야합니다.:
#
# $$-p_0log(p_0)-(1-p_0)log(1-p_0) = 0.01317$$
# + id="xVDqCWXDqHSc"
results = model.evaluate(train_features, train_labels, batch_size=BATCH_SIZE, verbose=0)
print("Loss: {:0.4f}".format(results[0]))
# + [markdown] id="FrDC8hvNr9yw"
# 이 초기 손실은 단순한 상태의 초기화에서 발생했을 때 보다 약 50배 적습니다.
#
# 이런 식으로 모델은 긍정적인 예시의 가능성이 낮다는 것을 배우면서 처음 몇 epoch를 보낼 필요는 없습니다. 이것은 또한 훈련 중 plot의 손실을 더 쉽게 읽어낼 수 있게 해줍니다.
# + [markdown] id="0EJj9ixKVBMT"
# ### 초기 가중치 체크 포인트
#
# 다양한 훈련 실행을 더욱 비교 가능하도록 하고 싶다면 초기 모델의 가중치를 체크 포인트 파일에 보관하고 훈련 전에 각 모델에 로드 하십시오
# + id="_tSUm4yAVIif"
initial_weights = os.path.join(tempfile.mkdtemp(), 'initial_weights')
model.save_weights(initial_weights)
# + [markdown] id="EVXiLyqyZ8AX"
# ### 바이어스 수정이 도움이 되는지 확인
#
# 계속 진행하기 전에 조심스러운 바이어스 초기화가 실제로 도움이 되었는지 빠르게 확인하십시오
#
# 조심스럽게 초기화를 한 것과 사용하지 않은 것의 20 epoch 동안 모델을 훈련하고 손실을 비교합니다.:
# + id="Dm4-4K5RZ63Q"
model = make_model()
model.load_weights(initial_weights)
model.layers[-1].bias.assign([0.0])
zero_bias_history = model.fit(
train_features,
train_labels,
batch_size=BATCH_SIZE,
epochs=20,
validation_data=(val_features, val_labels),
verbose=0)
# + id="j8DsLXHQaSql"
model = make_model()
model.load_weights(initial_weights)
careful_bias_history = model.fit(
train_features,
train_labels,
batch_size=BATCH_SIZE,
epochs=20,
validation_data=(val_features, val_labels),
verbose=0)
# + id="E3XsMBjhauFV"
def plot_loss(history, label, n):
# Use a log scale to show the wide range of values.
plt.semilogy(history.epoch, history.history['loss'],
color=colors[n], label='Train '+label)
plt.semilogy(history.epoch, history.history['val_loss'],
color=colors[n], label='Val '+label,
linestyle="--")
plt.xlabel('Epoch')
plt.ylabel('Loss')
plt.legend()
# + id="dxFaskm7beC7"
plot_loss(zero_bias_history, "Zero Bias", 0)
plot_loss(careful_bias_history, "Careful Bias", 1)
# + [markdown] id="fKMioV0ddG3R"
# 위의 그림은 이를 명확하게 보여줍니다. 유효성 검사 손실 측면에서 이 문제에 대해 조심스러운 초기화는 명확한 이점을 제공합니다.
# + [markdown] id="RsA_7SEntRaV"
# ### 모델 훈련
# + id="yZKAc8NCDnoR"
model = make_model()
model.load_weights(initial_weights)
baseline_history = model.fit(
train_features,
train_labels,
batch_size=BATCH_SIZE,
epochs=EPOCHS,
callbacks=[early_stopping],
validation_data=(val_features, val_labels))
# + [markdown] id="iSaDBYU9xtP6"
# ### 학습 이력 확인
# 이 섹션에서는 훈련 및 검증 세트에 대한 모델의 정확도와 손실에 대한 plot을 생성합니다. 이는 과적합을 확인하는데 유용하며 이 [튜토리얼](https://www.tensorflow.org/tutorials/keras/overfit_and_underfit)에서 자세한 내용을 확인할 수 있습니다.
#
# 추가적으로, 위에서 만든 모든 메트릭에 대해 이러한 plot을 생성할 수 있습니다. 거짓 음성이 포함되는 경우가 예시입니다.
# + id="WTSkhT1jyGu6"
def plot_metrics(history):
metrics = ['loss', 'auc', 'precision', 'recall']
for n, metric in enumerate(metrics):
name = metric.replace("_"," ").capitalize()
plt.subplot(2,2,n+1)
plt.plot(history.epoch, history.history[metric], color=colors[0], label='Train')
plt.plot(history.epoch, history.history['val_'+metric],
color=colors[0], linestyle="--", label='Val')
plt.xlabel('Epoch')
plt.ylabel(name)
if metric == 'loss':
plt.ylim([0, plt.ylim()[1]])
elif metric == 'auc':
plt.ylim([0.8,1])
else:
plt.ylim([0,1])
plt.legend()
# + id="u6LReDsqlZlk"
plot_metrics(baseline_history)
# + [markdown] id="UCa4iWo6WDKR"
# 참고: 검증 곡선은 일반적으로 훈련 곡선보다 성능이 좋습니다. 이는 주로 모델을 평가할 때 drop out 레이어가 활성화 되지 않았기 때문에 발생합니다.
# + [markdown] id="aJC1booryouo"
# ### 메트릭 평가
#
# [혼동 행렬](https://developers.google.com/machine-learning/glossary/#confusion_matrix) 을 사용하여 X축이 예측 레이블이고 Y축이 실제 레이블인 실제 레이블과 예측 레이블을 요약할 수 있습니다.
# + id="aNS796IJKrev"
train_predictions_baseline = model.predict(train_features, batch_size=BATCH_SIZE)
test_predictions_baseline = model.predict(test_features, batch_size=BATCH_SIZE)
# + id="MVWBGfADwbWI"
def plot_cm(labels, predictions, p=0.5):
cm = confusion_matrix(labels, predictions > p)
plt.figure(figsize=(5,5))
sns.heatmap(cm, annot=True, fmt="d")
plt.title('Confusion matrix @{:.2f}'.format(p))
plt.ylabel('Actual label')
plt.xlabel('Predicted label')
print('Legitimate Transactions Detected (True Negatives): ', cm[0][0])
print('Legitimate Transactions Incorrectly Detected (False Positives): ', cm[0][1])
print('Fraudulent Transactions Missed (False Negatives): ', cm[1][0])
print('Fraudulent Transactions Detected (True Positives): ', cm[1][1])
print('Total Fraudulent Transactions: ', np.sum(cm[1]))
# + [markdown] id="nOTjD5Z5Wp1U"
# 테스트 데이터 세트에서 모델을 평가하고 위에서 만든 측정 항목의 결과를 표시합니다.
# + id="poh_hZngt2_9"
baseline_results = model.evaluate(test_features, test_labels,
batch_size=BATCH_SIZE, verbose=0)
for name, value in zip(model.metrics_names, baseline_results):
print(name, ': ', value)
print()
plot_cm(test_labels, test_predictions_baseline)
# + [markdown] id="PyZtSr1v6L4t"
# 만약 모델이 모든 것을 완벽하게 예측했다면 이것은 잘못된 예측을 나타내는 주 대각선의 값이 0이 되는 [대각행렬](https://en.wikipedia.org/wiki/Diagonal_matrix) 이 됩니다. 이러한 경우에 매트릭스가 잘못 탐지한 경우가 상대적으로 적다는 것을 보여줍니다. 즉 잘못 플래그가 지정된 합법적인 거래가 상대적으로 적은 것을 의미합니다. 그러나 거짓 양성 수를 늘릴 때 드는 비용에도 불구하고 더 적은 수의 거짓 음성을 원할 수 있습니다. 거짓 음성 판정이 부정 거래를 통과할 수 있는 반면, 거짓 긍정 판정이 고객에게 이메일을 보내 카드 활동을 확인하도록 요청할 수 있기 때문에 이러한 거래 중단이 더 바람 직 할 수 있습니다.
# + [markdown] id="P-QpQsip_F2Q"
# ### ROC 플로팅
#
# 이제 [ROC](https://developers.google.com/machine-learning/glossary#ROC)을 플로팅 하십시오. 이 그래프는 출력 임계값을 조정하기만 해도 모델이 도달할 수 있는 성능 범위를 한눈에 보여주기 때문에 유용합니다.
# + id="lhaxsLSvANF9"
def plot_roc(name, labels, predictions, **kwargs):
fp, tp, _ = sklearn.metrics.roc_curve(labels, predictions)
plt.plot(100*fp, 100*tp, label=name, linewidth=2, **kwargs)
plt.xlabel('False positives [%]')
plt.ylabel('True positives [%]')
plt.xlim([-0.5,20])
plt.ylim([80,100.5])
plt.grid(True)
ax = plt.gca()
ax.set_aspect('equal')
# + id="DfHHspttKJE0"
plot_roc("Train Baseline", train_labels, train_predictions_baseline, color=colors[0])
plot_roc("Test Baseline", test_labels, test_predictions_baseline, color=colors[0], linestyle='--')
plt.legend(loc='lower right')
# + [markdown] id="gpdsFyp64DhY"
# 비교적 정밀도가 높은 것 같지만 회수율과 ROC 곡선(AUC) 밑 면적이 마음에 들 만큼 높지 않습니다. 분류자는 정밀도와 리콜을 모두 최대화 하려고 할 때 종종 도전해야 할 문제에 직면하는데, 이는 불균형 데이터세트로 작업할 떄 특히 그러합니다. 당신이 신경쓰는 문제의 맥락에서 다른 유형의 오류의 비용을 고려하는 것이 중요합니다. 이 예시에서 거짓음성(부정 거래를 놓친 경우)은 금전적인 비용을 초래하지만 , 거짓 양성(거래가 사기 행위로 잘못 표시됨)은 사용자들의 만족도를 감소시킬 수 있습니다.
# + [markdown] id="cveQoiMyGQCo"
# ## 클래스 가중치
# + [markdown] id="ePGp6GUE1WfH"
# ### 클래스 가중치 계산
#
# 목표는 부정 거래를 식별하는 것이지만, 여러분은 작업할 수 있는 긍정적인 샘플이 많지 않기 깨문에 분류자가 이용할 수 있는 몇 가지 예에 가중치를 두고자 할 것입니다. 매개 변수를 통해 각 클래스에 대한 Keras 가중치를 전달한다면 이 과정을 할 수 있습니다. 이로 인해 모델이 덜 표현된 클래스의 예에 "더 많은 주의를 기울이십시오"라고 할 수도 있습니다.
# + id="qjGWErngGny7"
# Scaling by total/2 helps keep the loss to a similar magnitude.
# The sum of the weights of all examples stays the same.
weight_for_0 = (1 / neg)*(total)/2.0
weight_for_1 = (1 / pos)*(total)/2.0
class_weight = {0: weight_for_0, 1: weight_for_1}
print('Weight for class 0: {:.2f}'.format(weight_for_0))
print('Weight for class 1: {:.2f}'.format(weight_for_1))
# + [markdown] id="Mk1OOE2ZSHzy"
# ### 클래스 가중치로 모델 교육
#
# 이제 해당 모델이 예측에 어떤 영향을 미치는지 확인하기 위하여 클래스 가중치로 모델을 재 교육하고 평가해 보십시오.
#
# 참고: `class_weights` 를 사용하면 손실 범위가 바뀝니다. 이는 최적기에 따라 학습의 안정성에 영향을 미칠 수 있습니다. 단계 크기가 그라데이션의 크기에 따라 달라지는 `optimizers.SGD` 와 같은 최적화 도구는 실패할 수 있습니다. 여기서 사용되는 최적화기인 `optimizers.Adam` 은 스케일링 변화에 영향을 받지 않습니다. 또한 가중치 때문에 전체 손실은 두 모델 간에 비교할 수 없습니다.
# + id="UJ589fn8ST3x"
weighted_model = make_model()
weighted_model.load_weights(initial_weights)
weighted_history = weighted_model.fit(
train_features,
train_labels,
batch_size=BATCH_SIZE,
epochs=EPOCHS,
callbacks=[early_stopping],
validation_data=(val_features, val_labels),
# The class weights go here
class_weight=class_weight)
# + [markdown] id="R0ynYRO0G3Lx"
# ### 학습 이력 조회
# + id="BBe9FMO5ucTC"
plot_metrics(weighted_history)
# + [markdown] id="REy6WClTZIwQ"
# ### 매트릭 평가
# + id="nifqscPGw-5w"
train_predictions_weighted = weighted_model.predict(train_features, batch_size=BATCH_SIZE)
test_predictions_weighted = weighted_model.predict(test_features, batch_size=BATCH_SIZE)
# + id="owKL2vdMBJr6"
weighted_results = weighted_model.evaluate(test_features, test_labels,
batch_size=BATCH_SIZE, verbose=0)
for name, value in zip(weighted_model.metrics_names, weighted_results):
print(name, ': ', value)
print()
plot_cm(test_labels, test_predictions_weighted)
# + [markdown] id="PTh1rtDn8r4-"
# 여기서 클래스 가중치를 사용하면 거짓 긍정이 더 많기 때문에 정확도와 정밀도가 낮다는 것을 알 수 있지만, 반대로 리콜과 AUC는 참 긍정이 더 많은 모델입니다. 정확도가 낮음에도 불구하고 이 모델은 리콜이 더 높습니다.(그리고 더 많은 부정 거래를 식별한다.) 물론 두 가지 유형의 오류에는 모두 비용이 발생합니다.(너무 많은 합법적인 거래를 사기로 표시하여 사용자를 괴롭히는 것을 원하지 않을 것입니다.) 응용 프로그램에 대하여 이러한 다양한 유형의 오류 간의 절충을 신중하게 고려하십시오.
# + [markdown] id="hXDAwyr0HYdX"
# ### ROC 플로팅
# + id="3hzScIVZS1Xm"
plot_roc("Train Baseline", train_labels, train_predictions_baseline, color=colors[0])
plot_roc("Test Baseline", test_labels, test_predictions_baseline, color=colors[0], linestyle='--')
plot_roc("Train Weighted", train_labels, train_predictions_weighted, color=colors[1])
plot_roc("Test Weighted", test_labels, test_predictions_weighted, color=colors[1], linestyle='--')
plt.legend(loc='lower right')
# + [markdown] id="5ysRtr6xHnXP"
# ## 오버샘플링
# + [markdown] id="18VUHNc-UF5w"
# ### 소수 계급 과대 표본
#
# 관련된 접근 방식은 소수 클래스를 오버 샘플링 하여 데이터 세트를 리 샘플링 하는 것입니다.
# + id="sHirNp6u7OWp"
pos_features = train_features[bool_train_labels]
neg_features = train_features[~bool_train_labels]
pos_labels = train_labels[bool_train_labels]
neg_labels = train_labels[~bool_train_labels]
# + [markdown] id="WgBVbX7P7QrL"
# #### NumPy 사용
#
# 긍정적인 예에서 적절한 수의 임의 인덱스를 선택하여 데이터 세트의 균형을
# 수동으로 조정할 수 있습니다.:
# + id="BUzGjSkwqT88"
ids = np.arange(len(pos_features))
choices = np.random.choice(ids, len(neg_features))
res_pos_features = pos_features[choices]
res_pos_labels = pos_labels[choices]
res_pos_features.shape
# + id="7ie_FFet6cep"
resampled_features = np.concatenate([res_pos_features, neg_features], axis=0)
resampled_labels = np.concatenate([res_pos_labels, neg_labels], axis=0)
order = np.arange(len(resampled_labels))
np.random.shuffle(order)
resampled_features = resampled_features[order]
resampled_labels = resampled_labels[order]
resampled_features.shape
# + [markdown] id="IYfJe2Kc-FAz"
# #### `tf.data` 사용
# + [markdown] id="usyixaST8v5P"
# `tf.data` 사용하는 경우 균형 잡힌 예제를 생성하는 가장 쉬운 방법은 `positive` 그리고 `negative` 데이터 세트로 시작하여 병합하는 것입니다. 더 많은 예는 [tf.data guide](../../guide/data.ipynb) 를 참조하세요.
# + id="yF4OZ-rI6xb6"
BUFFER_SIZE = 100000
def make_ds(features, labels):
ds = tf.data.Dataset.from_tensor_slices((features, labels))#.cache()
ds = ds.shuffle(BUFFER_SIZE).repeat()
return ds
pos_ds = make_ds(pos_features, pos_labels)
neg_ds = make_ds(neg_features, neg_labels)
# + [markdown] id="RNQUx-OA-oJc"
# 각 데이터 세트는 `(feature, label)` 쌍을 제공합니다.:
# + id="llXc9rNH7Fbz"
for features, label in pos_ds.take(1):
print("Features:\n", features.numpy())
print()
print("Label: ", label.numpy())
# + [markdown] id="sLEfjZO0-vbN"
# `experimental.sample_from_datasets` 를 사용하여 두 가지를 병합합니다.:
# + id="e7w9UQPT9wzE"
resampled_ds = tf.data.experimental.sample_from_datasets([pos_ds, neg_ds], weights=[0.5, 0.5])
resampled_ds = resampled_ds.batch(BATCH_SIZE).prefetch(2)
# + id="EWXARdTdAuQK"
for features, label in resampled_ds.take(1):
print(label.numpy().mean())
# + [markdown] id="irgqf3YxAyN0"
# 이 데이터 세트를 사용하려면 epoch 당 단계 수가 필요합니다.
#
# 이 경우 "epoch" 의 정의는 명확하지 않습니다. 각 부정적인 예를 한번 볼 때 필요한 배치 수라고 가정합니다.:
# + id="xH-7K46AAxpq"
resampled_steps_per_epoch = np.ceil(2.0*neg/BATCH_SIZE)
resampled_steps_per_epoch
# + [markdown] id="XZ1BvEpcBVHP"
# ### 오버 샘플링 된 데이터에 대한 학습
#
# 이제 클래스 가중치를 사용하는 대신 리 샘플링 된 데이터 세트로 모델을 학습하여 이러한 방법이 어떻게 비교되는지 확인하십시오.
#
# 참고: 긍정적인 예를 복제하여 데이터가 균형을 이루었기 때문에 총 데이터 세트 크기가 더 크고 각 세대가 더 많은 학습 단계를 위해 실행됩니다.
# + id="soRQ89JYqd6b"
resampled_model = make_model()
resampled_model.load_weights(initial_weights)
# Reset the bias to zero, since this dataset is balanced.
output_layer = resampled_model.layers[-1]
output_layer.bias.assign([0])
val_ds = tf.data.Dataset.from_tensor_slices((val_features, val_labels)).cache()
val_ds = val_ds.batch(BATCH_SIZE).prefetch(2)
resampled_history = resampled_model.fit(
resampled_ds,
epochs=EPOCHS,
steps_per_epoch=resampled_steps_per_epoch,
callbacks=[early_stopping],
validation_data=val_ds)
# + [markdown] id="avALvzUp3T_c"
# 만약 훈련 프로세스가 각 기울기 업데이트에서 전체 데이터 세트를 고려하는 경우, 이 오버 샘플링은 기본적으로 클래스 가중치와 동일합니다.
#
# 그러나 여기에서 한 것처럼 모델을 배치 방식으로 훈련 할 때 오버 샘플링 된 데이터는 더 부드러운 기울기 신호를 제공합니다. 각각의 긍정적인 예가 큰 가중치를 가진 하나의 배치로 표시되는 대신, 그것들은 작은 가중치로 매 회 많은 다른 배치로 보여집니다.
#
# 이 부드러운 기울기 신호는 모델을 더 쉽게 훈련 할 수 있습니다.
# + [markdown] id="klHZ0HV76VC5"
# ### 교육 이력 확인
#
# 학습 데이터의 분포가 검증 및 테스트 데이터와 완전히 다르기 때문에 여기서 측정 항목의 분포가 다를 수 있습니다.
# + id="YoUGfr1vuivl"
plot_metrics(resampled_history)
# + [markdown] id="1PuH3A2vnwrh"
# ### 재교육
#
# + [markdown] id="KFLxRL8eoDE5"
# 균형 잡힌 데이터에 대한 훈련이 더 쉽기 때문에 위의 훈련 절차가 빠르게 과적합 될 수 있습니다.
#
# 따라서 epochs를 분리하여 `callbacks.EarlyStopping`을 제공하십시오.
# + id="e_yn9I26qAHU"
resampled_model = make_model()
resampled_model.load_weights(initial_weights)
# Reset the bias to zero, since this dataset is balanced.
output_layer = resampled_model.layers[-1]
output_layer.bias.assign([0])
resampled_history = resampled_model.fit(
resampled_ds,
# These are not real epochs
steps_per_epoch=20,
epochs=10*EPOCHS,
callbacks=[early_stopping],
validation_data=(val_ds))
# + [markdown] id="UuJYKv0gpBK1"
# ### 훈련 이력 재확인
# + id="FMycrpJwn39w"
plot_metrics(resampled_history)
# + [markdown] id="bUuE5HOWZiwP"
# ### 메트릭 평가
# + id="C0fmHSgXxFdW"
train_predictions_resampled = resampled_model.predict(train_features, batch_size=BATCH_SIZE)
test_predictions_resampled = resampled_model.predict(test_features, batch_size=BATCH_SIZE)
# + id="FO0mMOYUDWFk"
resampled_results = resampled_model.evaluate(test_features, test_labels,
batch_size=BATCH_SIZE, verbose=0)
for name, value in zip(resampled_model.metrics_names, resampled_results):
print(name, ': ', value)
print()
plot_cm(test_labels, test_predictions_resampled)
# + [markdown] id="_xYozM1IIITq"
# ### ROC 플로팅
# + id="fye_CiuYrZ1U"
plot_roc("Train Baseline", train_labels, train_predictions_baseline, color=colors[0])
plot_roc("Test Baseline", test_labels, test_predictions_baseline, color=colors[0], linestyle='--')
plot_roc("Train Weighted", train_labels, train_predictions_weighted, color=colors[1])
plot_roc("Test Weighted", test_labels, test_predictions_weighted, color=colors[1], linestyle='--')
plot_roc("Train Resampled", train_labels, train_predictions_resampled, color=colors[2])
plot_roc("Test Resampled", test_labels, test_predictions_resampled, color=colors[2], linestyle='--')
plt.legend(loc='lower right')
# + [markdown] id="3o3f0ywl8uqW"
# ## 튜토리얼을 이 문제에 적용
#
# 불균형 데이터 분류는 학습 할 샘플이 너무 적기 때문에 본질적으로 어려운 작업입니다. 항상 데이터부터 시작하여 가능한 한 많은 샘플을 수집하고 모델이 소수 클래스를 최대한 활용할 수 있도록 어떤 기능이 관련 될 수 있는지에 대해 실질적인 생각을 하도록 최선을 다해야 합니다. 어떤 시점에서 모델은 원하는 결과를 개선하고 산출하는데 어려움을 겪을 수 있으므로 문제의 컨텍스트와 다양한 유형의 오류 간의 균형을 염두에 두는 것이 중요합니다.
| site/en/tutorials/structured_data/imbalanced_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from glob import glob
import os
import sys
sys.path.append(f'{os.environ["HOME"]}/Projects/planckClusters/catalogs')
from load_catalogs import load_PSZcatalog
# parallel processor
from utilities import parallel_process, system_call_env, system_call
# -
def combineXRT(name, outpath):
''' This combines the individual observations
IMPORTANT! The scripts that this (and other functions) creat are designed
to be run from the same directory as this notebook. They WILL NOT work
if you try to run them from the individual data directories.
'''
if not os.path.isdir(f'{outpath}/{name}'):
return
# find the x-ray files
files = glob(f'{outpath}/{name}/reduced/**/*xpcw[1-4]po_cl.evt', recursive=True)
# sort the files so we can control their input order
# this is used to make sure all of the data products are the same
# when we are combining observations
files = np.sort(files)
if len(files) < 1:
return
# write xsel.in
with open(f'{outpath}/{name}/{name}_xsel.in', 'w') as f:
for i, f_in in enumerate(files):
f_parts = f_in.split('/')
if i == 0:
# this is the session name... specify random letters to run many at once.
f.writelines(f'{name}\n')
f.writelines('read events\n')
# set the data directory
f.writelines('/'.join(f_parts[:3]) + '\n')
# first entry
f.writelines('/'.join(f_parts[3:]) + '\n')
f.writelines('yes\n')
continue
f.writelines('read events\n')
f.writelines('/'.join(f_parts[3:]) + '\n')
# if you try to read more than 20 exposures, it says "more?"
if i >= 19:
f.writelines('\n')
if i >= 42:
f.writelines('\n')
if i >= 65:
f.writelines('\n')
if i >= 88:
f.writelines('\n')
if i >= 111:
f.writelines('\n')
if i >= 134:
f.writelines('\n')
if i >= 157:
f.writelines('\n')
if i >= 180:
f.writelines('\n')
if i >= 203:
f.writelines('\n')
if i >= 226:
f.writelines('\n')
if i >= 249:
f.writelines('\n')
f.writelines('extract events\n')
f.writelines(f'save events {outpath}/{name}/{name}_events.fits\n')
if os.path.isfile(f'{outpath}/{name}/{name}_events.fits'):
f.writelines('yes\n')
f.writelines('yes\n')
f.writelines('set phaname PI\n')
# here we are going to make a few binned images for a few different energy ranges
# energies in loop
for eng in [200, 300, 400, 500, 600]:
f.writelines(f'filter pha_cutoff 50 {eng}\n')
# save non-binned image -- the yes's are to overwrite if file is already there
f.writelines('set xybinsize 1\n')
f.writelines('extract image\n')
f.writelines(f'save image {"/".join(f_parts[:3])}/{name}_img_50-{eng}.fits\n')
if os.path.isfile(f'{outpath}/{name}/{name}_img_50-{eng}.fits'):
f.writelines('yes\n')
# save binned image -- see above
f.writelines('set xybinsize 8\n')
f.writelines('extract image\n')
f.writelines(f'save image {"/".join(f_parts[:3])}/{name}_img_50-{eng}_bl8.fits\n')
if os.path.isfile(f'{outpath}/{name}/{name}_img_50-{eng}_bl8.fits'):
f.writelines('yes\n')
f.writelines('set xybinsize 4\n')
f.writelines('extract image\n')
f.writelines(f'save image {"/".join(f_parts[:3])}/{name}_img_50-{eng}_bl4.fits\n')
if os.path.isfile(f'{outpath}/{name}/{name}_img_50-{eng}_bl4.fits'):
f.writelines('yes\n')
f.writelines('exit\n')
f.writelines('no\n')
# call
cmd = f'xselect @{outpath}/{name}/{name}_xsel.in'
# stdout, stderr = system_call_env(cmd)
# log the output
log_file = f'{outpath}/{name}/{name}_xsel.log'
# with open(log_file, 'w') as f:
# f.writelines(stdout)
os.system(f'{cmd} > {log_file}')
return
# +
def combineXRT_exp(name, outpath):
''' This combines the exposure maps'''
if not os.path.isdir(f'{outpath}/{name}'):
return
# find the x-ray files
files = glob(f'{outpath}/{name}/reduced/**/*xpcw[1-4]po_ex.img', recursive=True)
# sort the observations -- see above
files = np.sort(files)
if len(files) < 1:
return name
# remove the old file if it is there
if os.path.isfile(f'{outpath}/{name}/{name}_exp.fits'):
os.remove(f'{outpath}/{name}/{name}_exp.fits')
# write xsel.in
with open(f'{outpath}/{name}/{name}_ximg_exp.in', 'w') as f:
for i, f_in in enumerate(files):
f_parts = f_in.split('/')
f.writelines(f'read {f_in}\n')
if i == 0:
continue
f.writelines('sum\n')
f.writelines('save\n')
f.writelines(f'write/fits {"/".join(f_parts[:3])}/{name}_exp.fits\n')
f.writelines('exit\n')
# # call
cmd = f'ximage @{outpath}/{name}/{name}_ximg_exp.in'
stdout, stderr = system_call_env(cmd)
print(stdout)
# log the output
log_file = f'{outpath}/{name}/{name}_ximg_exp.log'
with open(log_file, 'w') as f:
f.writelines(stdout)
# os.system(f'{cmd} > {log_file}')
return
# +
# get file data
data = load_PSZcatalog()
data = data.sort_index(axis=1)
outpath = './data_full_new'
arr = [{'name':n.replace(' ', '_'), 'outpath':outpath} for n in data['NAME']]
parallel_process(arr, combineXRT, use_kwargs=True, n_jobs=7)
parallel_process(arr, combineXRT_exp, use_kwargs=True, n_jobs=7) #4.5 hours
# +
# outpath = './data_full'
# name = 'PSZ2_G313.33+61.13'
# combineXRT(name, outpath)
# combineXRT_exp(name, outpath)
| 03. Combine Reduced Products.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
urlpage='https://www.yelp.com/search?find_desc=Coworking+Space&find_loc=Los+Angeles Webscraping'
# +
from openpyxl import load_workbook
from bs4 import BeautifulSoup
from selenium import webdriver
from time import sleep
import csv
from random import randint
import json, io
from selenium.webdriver.common.by import By
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.support.ui import Select
from selenium.webdriver.support.ui import WebDriverWait
from selenium.common.exceptions import TimeoutException
from selenium.webdriver.support import expected_conditions as EC
from selenium.common.exceptions import NoSuchElementException
from selenium.common.exceptions import NoAlertPresentException
from selenium.webdriver.common.action_chains import ActionChains
import urllib
import urllib3
import requests
import json, io
from bs4 import BeautifulSoup
urllib3.disable_warnings()
header = {'User-Agent':'Mozilla/5.0'}
chrome_options = webdriver.ChromeOptions()
chrome_options.add_argument('--user-agent="Mozilla/5.0')
chrome_options.add_argument("user-data-dir=selenium")
driver = webdriver.Chrome(chrome_options=chrome_options, executable_path=r'/Users/Name/Downloads/Compressed/chromedrives/chromedriver.exe')
cookies = json.load(open('cookiesdict.txt'))
for cookie in cookies:
driver.add_cookie(cookie)
# -
ul0='https://www.yelp.com/search?find_desc=Home+Design&find_loc=90036&start=220&l=g:-118.401374817,34.0253477381,-118.299064636,34.1106675388'
def extractpage():
so1=BeautifulSoup(driver.page_source, 'lxml')
data_Stor0=['-']*14
title0=so1.find_all('h1')[0].text
title1=title0.replace(' ','').replace('\n','')
# print title1
data_Stor0[0]=title1
biz_rating=so1.find_all('div',class_="biz-rating biz-rating-very-large clearfix")
if len(biz_rating)>0:
rating=biz_rating[0].find(class_='offscreen').get('alt')
# print rating
data_Stor0[1]=rating
reviews0=biz_rating[0].find('span').text
reviews1=reviews0.replace(' ','').replace('\n','')
# print reviews1
data_Stor0[2]=reviews1
mapbox=so1.find_all('div',class_="mapbox-text")[0]
street_address0=mapbox.find_all('strong',class_="street-address")[0]
street_address1=street_address0.next_element.next_element.next_element
street_address2=street_address1.next_element.next_element
try:
street_address11=street_address1.replace(' ','').replace('\n','')
# print street_address11
data_Stor0[3]=street_address11
except:
street_address11=street_address1
# print street_address11
data_Stor0[3]=street_address11
try:
street_address22=street_address2.replace(' ','').replace('\n','')
# print street_address22
data_Stor0[4]=street_address22
except:
street_address22=street_address2
# print street_address22
data_Stor0[4]=street_address22
biz_phone0=mapbox.find_all('span',class_="biz-phone")
if len(biz_phone0)>0:
biz_phone1=biz_phone0[0].text
biz_phone2=biz_phone1.replace(' ','').replace('\n','')
# print biz_phone2
data_Stor0[5]=biz_phone2
biz_website=mapbox.find_all('span',class_="biz-website js-biz-website js-add-url-tagging")
print len(biz_website)
if len(biz_website)>0:
website0=biz_website[0]
website=website0.find_all('a')[0].text
# print website
data_Stor0[6]=website
data_Stor0[7]=driver.current_url
biz_main_info=so1.find_all('div',class_="biz-main-info embossed-text-white")[0]
category_str_list=biz_main_info.find_all('span',class_="category-str-list")[0]
acategory=category_str_list.find_all('a')
category_stor=[]
for ii in range(len(acategory)):
# print acategory[ii].text
category_stor.append(acategory[ii].text)
for hg in range(len(category_stor)):
data_Stor0[8+hg]=category_stor[hg]
print data_Stor0
data_Stor1.append(data_Stor0)
driver.get(urlpage)
driver.switch_to.window(driver.window_handles[0])
search_result=driver.find_elements_by_class_name('regular-search-result')
len(search_result)
for kl in range(0,1):
search_result=driver.find_elements_by_class_name('regular-search-result')
for jk in range(0,len(search_result)):
driver.switch_to_window(driver.window_handles[0])
sleep(5)
actions = ActionChains(driver)
about=search_result[jk].find_element_by_class_name('biz-name')
actions.key_down(Keys.CONTROL).click(about).key_up(Keys.CONTROL).perform()
sleep(10)
driver.switch_to.window(driver.window_handles[1])
sleep(3)
extractpage()
driver.close()
driver.switch_to_window(driver.window_handles[0])
sleep(2)
driver.find_element_by_link_text('Next').click()
sleep(15)
jk
driver.switch_to.window(driver.window_handles[1])
extractpage()
jk
# +
250,-7
# -
driver.close()
len(data_Stor1)
data_Stor1
# +
import warnings
from openpyxl import Workbook
wb = Workbook(write_only=True)
ws = wb.create_sheet()
p=0
# now we'll fill it with 100 rows x 200 columns
for irow in data_Stor1:
print irow
p=p+1
print p
try:
ws.append(irow)
except:
try:
data_Stor1[p-1][3]=data_Stor1[p-1][3].text
except:
pass
try:
data_Stor1[p-1][4]=data_Stor1[p-1][4].text
except:
pass
print p
# save the file
wb.save('af456.xlsx')
# +
import warnings
from openpyxl import Workbook
wb = Workbook(write_only=True)
ws = wb.create_sheet()
# now we'll fill it with 100 rows x 200 columns
for irow in data_Stor1:
ws.append(irow)
# save the file
wb.save('home_des1--20.xlsx')
# -
cookiesdict=driver.get_cookies()
cookiesdict
import json, io
with io.open('cookiesdict.txt', 'w', encoding='utf8') as json_file:
data3 = json.dumps(cookiesdict, ensure_ascii=False, encoding='utf8',indent=4, sort_keys=True)
json_file.write(unicode(data3))
| streeteasy.com/streeteasy.com.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Example 3: Necessary and Sufficiency for containment in V-polytopes
# We investigate unitbox in V-polytope containment. First, we import `numpy` and `pypolycontain`.
import pypolycontain as pp
import numpy as np
np.random.seed(0)
n_range=[3,5,7,9]
Table={}
N_vertices_range=[1,2,4,6,8,10]
for n in n_range:
for N_vertices in N_vertices_range:
print(n,N_vertices)
X=pp.box(n)
Y= pp.V_polytope([np.random.normal(size=(n,1)) for i in range(N_vertices+n)])
Table[n,N_vertices+n]=pp.necessity_gap_k(X,Y,only_final=True)
import matplotlib.pyplot as plt
a={}
text={n: '%d'%n for n in n_range}
Color={2:'cyan', 3:'orange', 5:'red',7:'green',9:'blue'}
N_v= N_vertices_range
for n in n_range:
a[n]=plt.plot( N_v, [Table[n,k+n][0] for k in N_v],label=r'$n=%s$'%(text[n]),color=Color[n])
a[n]=plt.plot( N_v, [Table[n,k+n][0] for k in N_v],'o',color=Color[n])
plt.legend(loc="upper left")
plt.grid('on')
plt.xlabel(r'$n_v$',FontSize=20)
plt.ylabel(r'# of rows in $\Theta^*$ (Theorem 1)',FontSize=15)
import matplotlib.pyplot as plt
a={}
text={n: '%d'%n for n in n_range}
Color={2:'cyan', 3:'orange', 5:'red',7:'green',9:'blue'}
N_v= N_vertices_range
for n in n_range:
a[n]=plt.plot( N_v, [Table[n,k+n][1] for k in N_v],label=r'$n=%s$'%(text[n]),color=Color[n])
a[n]=plt.plot( N_v, [Table[n,k+n][1] for k in N_v],'o',color=Color[n])
plt.legend(loc="upper left")
plt.grid('on')
plt.xlabel(r'$n_v$',FontSize=20)
plt.ylabel(r'$\delta(\mathbb{X},\mathbb{Y},\mathbb{R}_+^{q_y})$ (Corollary 2)',FontSize=15)
| doc/source/Example 3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="SV7gaADiicnV"
# # Lambda School Data Science - Quantile Regression
#
# Regressing towards the median - or any quantile - as a way to mitigate outliers and control risk.
# + [markdown] colab_type="text" id="6klMj4q3iqMh"
# ## Lecture
#
# Let's look at data that has a bit of a skew to it:
#
# http://archive.ics.uci.edu/ml/datasets/Beijing+PM2.5+Data
# + colab={} colab_type="code" id="yw1AD_z9O0xL"
import pandas as pd
df = pd.read_csv('http://archive.ics.uci.edu/ml/machine-learning-databases/'
'00381/PRSA_data_2010.1.1-2014.12.31.csv')
# + colab={"base_uri": "https://localhost:8080/", "height": 206} colab_type="code" id="RTlH1lJ8PDv5" outputId="e073db49-81bd-4ebd-f43b-69c92aea8467"
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 320} colab_type="code" id="m-yC9OSPPFo8" outputId="d5602fe7-31ad-458e-d466-212c99e51cf4"
df.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 347} colab_type="code" id="hfV3WisFP_O6" outputId="a9809666-6c33-4778-fe1c-f3030f89d431"
df['pm2.5'].plot.hist();
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="OgbMTAHzQJB8" outputId="15e18384-ede7-4ccd-8961-280b35f66f85"
# How does linear regression handle it?
from sklearn.linear_model import LinearRegression
# Let's drop NAs and limit to numeric values
df = df._get_numeric_data().dropna()
X = df.drop('pm2.5', axis='columns')
y = df['pm2.5']
linear_reg = LinearRegression().fit(X, y)
linear_reg.score(X, y)
# + colab={"base_uri": "https://localhost:8080/", "height": 462} colab_type="code" id="-viFFtm0RizM" outputId="256d7680-1a43-4958-c74c-31aaef917906"
# Not bad - but what if we wanted to model the distribution more conservatively?
# Let's try quantile
import statsmodels.formula.api as smf
# Different jargon/API in StatsModel documentation
# "endogenous" response var is dependent (y), it is "inside"
# "exogenous" variables are independent (X), it is "outside"
# Bonus points - talk about "exogenous shocks" and you're a bona fide economist
# ~ style formulas look like what R uses
# y ~ x1 + x2 + ...
# Also, these formulas break with . in variable name, so lets change that
df = df.rename(index=str, columns={'pm2.5': 'pm25'})
# Now let's construct the formula string using all columns
quant_formula = 'pm25 ~ ' + ' + '.join(df.drop('pm25', axis='columns').columns)
print(quant_formula)
quant_mod = smf.quantreg(quant_formula, data=df)
quant_reg = quant_mod.fit(q=.5)
quant_reg.summary() # "summary" is another very R-thing
# + [markdown] colab_type="text" id="ZBkP4bewd-HT"
# That fit to the median (q=0.5), also called "Least Absolute Deviation." The pseudo-R^2 isn't really directly comparable to the R^2 from linear regression, but it clearly isn't dramatically improved. Can we make it better?
# + colab={"base_uri": "https://localhost:8080/", "height": 593} colab_type="code" id="BgvYeHg3bL4g" outputId="bf4547a0-7739-45d8-bf5a-26ab1684f7f6"
help(quant_mod.fit)
# + colab={"base_uri": "https://localhost:8080/", "height": 1424} colab_type="code" id="lpNPioZTei4U" outputId="40fc70a6-43c5-44a0-a012-923bd3f826a8"
quantiles = (.05, .96, .1)
for quantile in quantiles:
print(quant_mod.fit(q=quantile).summary())
# + [markdown] colab_type="text" id="Xqh4Jp1XgjrE"
# "Strong multicollinearity", eh? In other words - maybe we shouldn't throw every variable in our formula. Let's hand-craft a smaller one, picking the features with the largest magnitude t-statistics for their coefficients. Let's also search for more quantile cutoffs to see what's most effective.
# + colab={"base_uri": "https://localhost:8080/", "height": 975} colab_type="code" id="NmoELnXwgpXd" outputId="1865f1b1-778a-4e73-91b7-d30ad29b2ee2"
quant_formula = 'pm25 ~ DEWP + TEMP + Ir + hour + Iws'
quant_mod = smf.quantreg(quant_formula, data=df)
for quantile in range(50, 100):
quantile /= 100
quant_reg = quant_mod.fit(q=quantile)
print((quantile, quant_reg.prsquared))
# + colab={"base_uri": "https://localhost:8080/", "height": 444} colab_type="code" id="Bz0GmE5kuwQY" outputId="d139eca6-fa58-4f4c-a051-18b3e2d7ee13"
# Okay, this data seems *extremely* skewed
# Let's trying logging
import numpy as np
df['pm25'] = np.log(1 + df['pm25'])
quant_mod = smf.quantreg(quant_formula, data=df)
quant_reg = quant_mod.fit(q=.25)
quant_reg.summary() # "summary" is another very R-thing
# + [markdown] colab_type="text" id="8kXcxnNBgizX"
# Overall - in this case, quantile regression is not *necessarily* superior to linear regression. But it does give us extra flexibility and another thing to tune - what the center of what we're actually fitting in the dependent variable.
#
# The basic case of `q=0.5` (the median) minimizes the absolute value of residuals, while OLS minimizes the squared value. By selecting `q=0.25`, we're targeting a lower quantile and are effectively saying that we only want to over-estimate at most 25% of the time - we're being *risk averse*.
#
# Depending on the data you're looking at, and the cost of making a false positive versus a false negative, this sort of flexibility can be extremely useful.
#
# Live - let's consider another dataset! Specifically, "SkillCraft" (data on competitive StarCraft players): http://archive.ics.uci.edu/ml/datasets/SkillCraft1+Master+Table+Dataset
# + colab={} colab_type="code" id="ofvwSAZUhWDw"
# TODO Live!
# Hint - we may only care about the *top* quantiles here
# Another hint - there are missing values, but Pandas won't see them right away
skill = pd.read_csv('http://archive.ics.uci.edu/ml/machine-learning-databases/00272/SkillCraft1_Dataset.csv')
# -
skill.replace('?', np.nan, inplace=True)
skill.isnull().sum()
# replacing missing values for Age, HoursPerWeek, TotalHours
skill['Age'] = skill['Age'].fillna(skill['Age'].median())
skill['HoursPerWeek'] = skill['HoursPerWeek'].fillna(skill['HoursPerWeek'].median())
skill['TotalHours'] = skill['TotalHours'].fillna(skill['TotalHours'].median())
print ('Total nulls', skill.isnull().sum().sum())
# check dtypes
skill.dtypes
# correcting object types
skill['Age'] = pd.to_numeric(skill['Age'])
skill['HoursPerWeek'] = pd.to_numeric(skill['HoursPerWeek'])
skill['TotalHours'] = pd.to_numeric(skill['TotalHours'])
# +
# predicting top 10% of players in APM
target = 'APM'
features = skill.columns.drop(target)
X = skill[features]
y = skill[target]
# +
quant_formula_skill = 'APM ~ ' + ' + '.join(X.columns)
print(quant_formula_skill)
quant_mod = smf.quantreg(quant_formula_skill, data=skill)
quant_reg = quant_mod.fit(q=.9)
quant_reg.summary() # "summary" is another very R-thing
# -
quant_reg.predict(X.iloc[0, :])
# # Assignment - birth weight data¶
# Birth weight is a situation where, while the data itself is actually fairly normal and symmetric, our main goal is actually not to model mean weight (via OLS), but rather to identify mothers at risk of having children below a certain "at-risk" threshold weight.
#
# Quantile regression gives us just the tool we need. For the data we are using, see: http://people.reed.edu/~jones/141/BirthWgt.html
#
# bwt: baby's weight in ounces at birth
# gestation: duration of pregnancy in days
# parity: parity indicator (first born = 1, later birth = 0)
# age: mother's age in years
# height: mother's height in inches
# weight: mother's weight in pounds (during pregnancy)
# smoke: indicator for whether mother smokes (1=yes, 0=no)
#
# Use this data and statsmodels to fit a quantile regression, predicting bwt (birth weight) as a function of the other covariates. First, identify an appropriate q (quantile) to target a cutoff of 90 ounces - babies above that birth weight are generally healthy/safe, babies below are at-risk.
#
# Then, fit and iterate your model. Be creative! You may want to engineer features. Hint - mother's age likely is not simply linear in its impact, and the other features may interact as well.
#
# At the end, create at least 2 tables and 1 visualization to summarize your best model. Then (in writing) answer the following questions:
#
# What characteristics of a mother indicate the highest likelihood of an at-risk (low weight) baby?
# What can expectant mothers be told to help mitigate this risk?
# Note that second question is not exactly a data science question - and that's okay! You're not expected to be a medical expert, but it is a good exercise to do a little bit of digging into a particular domain and offer informal but informed opinions.
# !pip install -U numpy
# !pip install -U matplotlib
# +
import pandas as pd
import numpy as np
import scipy.stats
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
import statsmodels.formula.api as smf
# + colab={"base_uri": "https://localhost:8080/", "height": 206} colab_type="code" id="HUWKv16FjZsY" outputId="11f1ecab-4058-4e48-ac0f-cd7cf488a2f7"
newborn = pd.read_csv('http://people.reed.edu/~jones/141/Bwt.dat')
print(newborn.shape)
newborn.head()
# -
# newborn.isnull().sum()
newborn.dtypes
# + colab={"base_uri": "https://localhost:8080/", "height": 300} colab_type="code" id="dy5FkUZpkJT_" outputId="d4f46328-8e25-4fa5-e5b5-6ffad654c65c"
newborn.describe()
# + colab={} colab_type="code" id="Ez8qPLojjlFf"
newborn['bwt'].plot.hist(); # why do we need a semicolon at the end?
# +
# How does linear regression handle it?
# Let's drop NAs and limit to numeric values
newborn = newborn._get_numeric_data().dropna()
X = newborn.drop('bwt', axis = 'columns')
y = newborn['bwt']
linear_reg = LinearRegression().fit(X, y)
linear_reg.score(X, y)
# -
scipy.stats.percentileofscore(newborn.bwt, 90)/100
df = newborn.copy()
df.head()
df['less_than_250'] = df['gestation'].apply(lambda g: 1 if g < 250 else 0)
df[df['less_than_250'] == 1].head()
df['less_than_90'] = df['bwt'].apply(lambda b: 1 if b < 90 else 0)
df[df['less_than_90'] == 1].head()
df.head()
# smokers with early births
df['smokers_premie'] = (df['gestation'] < 260) * df['smoke']
df.head()
# early births in general
df['premie'] = df['gestation'] < 260
df.head()
# BMI
df['bmi'] = newborn['weight'] / (newborn['height']**2)
# age squared
df['age_squared'] = newborn['age'] ** 2
# age squared and smoke interaction term
df['smoker_age_squared'] = df['age_squared'] * df['smoke']
df.head()
df_group = df.groupby(["age", 'gestation']).count().sort_values(by = ['less_than_90'], ascending = False)
df_group = pd.DataFrame(df_group, columns = ['less_than_90'])
df_group.head()
df.columns
# +
# quantile
import statsmodels.formula.api as smf
# using all columns
quant_formula = 'bwt ~ ' + ' + '.join(df.drop('bwt', axis='columns').columns)
print(quant_formula)
quant_mod = smf.quantreg(quant_formula, data = df)
quant_reg = quant_mod.fit(q = 0.05664)
quant_reg.summary()
# +
target = 'bwt'
features = df.columns.drop([target, 'weight', 'height'])
X = df[features]
y = df[target]
quant_formula_bwt = 'bwt ~ ' + ' + '.join(X.columns)
print(quant_formula)
quant_mod = smf.quantreg(quant_formula, data = df)
quant_reg = quant_mod.fit(q=.05)
quant_reg.summary()
# +
target = 'bwt'
features = df.columns.drop([target, 'parity', 'age', 'height', 'weight', 'bmi',
'age_squared', 'smoker_age_squared', 'smokers_premie'])
X = df[features]
y = df[target]
quant_formula = 'bwt ~ ' + ' + '.join(X.columns)
print(quant_formula)
quant_mod = smf.quantreg(quant_formula, data = df)
quant_reg = quant_mod.fit(q = 0.05664)
quant_reg.summary() # "summary" is another very R-thing
# +
sns.set(style = "ticks", color_codes = True)
x_columns = df.columns.drop('bwt')
# Only plot the scatterplot of x variables with our y variable
sns.pairplot(data = df, y_vars = ['bwt'], x_vars = x_columns)
# -
quant_formula = 'bwt ~ gestation + parity + age + height + weight + smoke'
quant_mod = smf.quantreg(quant_formula, data = df)
for quantile in range(1, 6):
quantile /= 100
quant_reg = quant_mod.fit(q = quantile)
print((quantile, quant_reg.prsquared))
quant_formula = 'bwt ~ age'
quant_mod = smf.quantreg(quant_formula, data = df)
for quantile in range(1, 6):
quantile /= 100
quant_reg = quant_mod.fit(q = quantile)
print((quantile, quant_reg.prsquared))
quant_formula = 'bwt ~ smoke'
quant_mod = smf.quantreg(quant_formula, data = df)
for quantile in range(1, 6):
quantile /= 100
quant_reg = quant_mod.fit(q = quantile)
print((quantile, quant_reg.prsquared))
quant_formula = 'bwt ~ gestation'
quant_mod = smf.quantreg(quant_formula, data = df)
for quantile in range(1, 6):
quantile /= 100
quant_reg = quant_mod.fit(q = quantile)
print((quantile, quant_reg.prsquared))
quant_formula = 'bwt ~ parity'
quant_mod = smf.quantreg(quant_formula, data = df)
for quantile in range(1, 6):
quantile /= 100
quant_reg = quant_mod.fit(q = quantile)
print((quantile, quant_reg.prsquared))
# pairplot with smoker as hue
sns.pairplot(data = df, x_vars = df.columns.drop(['bwt', 'parity', 'smoke']), y_vars = 'bwt', hue = 'smoke');
# What characteristics of a mother indicate the highest likelihood of an at-risk (low weight) baby?
# - Smoking appears to be a significant characteristic contributing to a low weight baby.
# - Gestation is also a significant factor leading to low birth weight.
# What can expectant mothers be told to help mitigate this risk?
# - Smoking is bad for your health regardless of pregnancy. However, smoking while pregnant should be
# avoided.
# - Gestation is not generally something a woman can control. It is possible for certain activities, like
# over doing things later in pregnancy, can contribute to early labor. Risky behavior can also lead to
# early labor. The earlier the labor or shorter the gestation, the more likely it is the newborn will be
# smaller and/or have issues
# + [markdown] colab_type="text" id="XY9JGAnJisdB"
# ## Resources and stretch goals
# + [markdown] colab_type="text" id="inFWXSpqmND5"
# Resources:
# - [statsmodels QuantReg example](http://www.statsmodels.org/dev/examples/notebooks/generated/quantile_regression.html)
# - [How Shopify used Quantile Regression in modeling risk](https://medium.com/data-shopify/how-shopify-capital-uses-quantile-regression-to-help-merchants-succeed-10ee1b36b17d)
#
# Stretch goals:
# - Find a dataset where you think quantile regression may be appropriate, and try both it and linear regression - compare/contrast their strengths/weaknesses, and write a summary for which you think is better for the situation and why
# - Check out [deep quantile regression](https://www.kdnuggets.com/2018/07/deep-quantile-regression.html), an approach that uses a custom quantile loss function and Keras to train a quantile model
| module3-quantile-regression/LS_DS_233_Quantile_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# change the river discharge source point of Fraser and create new Fraser River flow file
# # Re-write monthly & yearly rivers file with different freshwater grid cell
from __future__ import division
from salishsea_tools import rivertools
from salishsea_tools import nc_tools
import numpy as np
import matplotlib.pyplot as plt
import netCDF4 as nc
import arrow
import numpy.ma as ma
import sys
sys.path.append('/ocean/klesouef/meopar/tools/I_ForcingFiles/Rivers')
# %matplotlib inline
filename = '/ocean/jieliu/research/meopar/nemo-forcing/rivers/rivers_month.nc'
clim_rivers = nc.Dataset(filename, 'r')
nc_tools.show_dimensions(clim_rivers)
nc_tools.show_variables(clim_rivers)
criverflow = clim_rivers.variables['rorunoff']
# get other variables so we can put them in new files
lat = clim_rivers.variables['nav_lat']
lon = clim_rivers.variables['nav_lon']
riverdepth = clim_rivers.variables['rodepth']
rivertype = 'constant' ## monthly or constant(yearly)
if rivertype == 'monthly':
fluxfile = nc.Dataset('/ocean/sallen/allen/research/MEOPAR/nemo-forcing/rivers/Salish_allrivers_monthly.nc','r')
#inialise the runoff and run_depth arrays
runoff, run_depth, run_temp = rivertools.init_runoff_array_monthly()
#get river fluxes from netcdf file
if rivertype == 'constant':
fluxfile = nc.Dataset('/ocean/sallen/allen/research/MEOPAR/nemo-forcing/rivers/Salish_allrivers_cnst.nc','r')
#inialise the runoff and run_depth arrays
runoff, run_depth, run_temp = rivertools.init_runoff_array()
#list of watersheds we are including
names = ['skagit','fraser','evi_n','howe','bute','puget','jdf','evi_s','jervis','toba']
for name in range(0,len(names)):
watershedname = names[name]
Flux = fluxfile.variables[watershedname][:]
if rivertype == 'constant':
Flux = float(Flux)
runoff_orig = np.copy(runoff)
runoff, run_depth, run_temp = rivertools.put_watershed_into_runoff(rivertype,
watershedname, Flux, runoff, run_depth, run_temp)
if rivertype == 'monthly':
rivertools.check_sum_monthly(runoff_orig, runoff, Flux)
if rivertype == 'constant':
rivertools.check_sum(runoff_orig, runoff, Flux)
print run_depth[414,334]
if rivertype == 'monthly':
nemo = nc.Dataset('/ocean/jieliu/research/meopar/river-treatment/rivers_month_edit.nc', 'w')
nemo.description = 'Monthly Averages, All Rivers, modify on depth and runoff grid point'
# dimensions
nemo.createDimension('x', 398)
nemo.createDimension('y', 898)
nemo.createDimension('time_counter', None)
# variables
# latitude and longitude
nav_lat = nemo.createVariable('nav_lat','float32',('y','x'),zlib=True)
nav_lat = lat
x = nemo.createVariable('nav_lon','float32',('y','x'),zlib=True)
nav_lon = lon
# time
time_counter = nemo.createVariable('time_counter', 'float32', ('time_counter'),zlib=True)
time_counter.units = 'non-dim'
time_counter[0:12] = range(1,13)
# runoff
rorunoff = nemo.createVariable('rorunoff', 'float32', ('time_counter','y','x'), zlib=True)
rorunoff._Fillvalue = 0.
rorunoff._missing_value = 0.
rorunoff._units = 'kg m-2 s-1'
rorunoff[0:12,:] = runoff
# depth
rodepth = nemo.createVariable('rodepth','float32',('y','x'),zlib=True)
rodepth._Fillvalue = -1.
rodepth.missing_value = -1.
rodepth.units = 'm'
rodepth[:] = run_depth[0,:,:]
# temperature
rotemper = nemo.createVariable('rotemper','float32',('time_counter','y','x'),zlib=True)
rotemper._Fillvalue = -99.
rotemper.missing_value = -99.
rotemper.units = 'deg C'
rotemper[0:12,:] = run_temp
nemo.close()
if rivertype == 'constant':
nemo = nc.Dataset('/ocean/jieliu/research/meopar/river-treatment/rivers_cnst_edit.nc', 'w')
nemo.description = 'Constant Yearly Average, All Rivers, modify on depth and runoff grid point'
# dimensions
nemo.createDimension('x', 398)
nemo.createDimension('y', 898)
nemo.createDimension('time_counter', None)
# variables
# latitude and longitude
nav_lat = nemo.createVariable('nav_lat','float32',('y','x'),zlib=True)
nav_lat = lat
x = nemo.createVariable('nav_lon','float32',('y','x'),zlib=True)
nav_lon = lon
# time
time_counter = nemo.createVariable('time_counter', 'float32', ('time_counter'),zlib=True)
time_counter.units = 'non-dim'
time_counter[0] = 1
# runoff
rorunoff = nemo.createVariable('rorunoff', 'float32', ('time_counter','y','x'), zlib=True)
rorunoff._Fillvalue = 0.
rorunoff._missing_value = 0.
rorunoff._units = 'kg m-2 s-1'
rorunoff[0,:] = runoff
# depth
rodepth = nemo.createVariable('rodepth','float32',('y','x'),zlib=True)
rodepth._Fillvalue = -1.
rodepth.missing_value = -1.
rodepth.units = 'm'
rodepth[:] = run_depth
nemo.close()
# # Re-write daily Fraser flow file from May 14, 2015-June 14, 2015
# Constant and data ranges etc
year = 2015
smonth = 06
emonth = 06
startdate = arrow.get(year,smonth,14)
enddate = arrow.get(year,emonth,14)
print startdate, enddate
# get Fraser Flow data
filename = '/data/dlatorne/SOG-projects/SOG-forcing/ECget/Fraser_flow'
fraserflow = np.loadtxt(filename)
print fraserflow
#Fraser watershed
pd = rivertools.get_watershed_prop_dict('fraser')
totalfraser = (pd['Fraser1']['prop'] + pd['Fraser2']['prop'] +
pd['Fraser3']['prop'] + pd['Fraser4']['prop'])
# +
# Climatology, Fraser Watershed
fluxfile = nc.Dataset('/ocean/jieliu/research/meopar/nemo-forcing/rivers/Salish_allrivers_monthly.nc','r')
climFraserWaterShed = fluxfile.variables['fraser'][:]
# Fraser River at Hope Seasonal Climatology (found in matlab using Mark's mean daily data)
climFraseratHope = (931, 878, 866, 1814, 4097, 6970, 5538, 3539, 2372, 1937, 1595, 1119)
NonHope = climFraserWaterShed - climFraseratHope
otherratio = 0.016
fraserratio = 1-otherratio
nonFraser = (otherratio * climFraserWaterShed.sum()/NonHope.sum()) * NonHope
afterHope = NonHope - nonFraser
print pd['Fraser1']['i'],pd['Fraser1']['j']
# -
def calculate_daily_flow(r,criverflow):
'''interpolate the daily values from the monthly values'''
print r.day, r.month
if r.day < 16:
prevmonth = r.month-1
if prevmonth == 0:
prevmonth = 12
nextmonth = r.month
else:
prevmonth = r.month
nextmonth = r.month + 1
if nextmonth == 13:
nextmonth = 1
fp = r - arrow.get(year,prevmonth,15)
fn = arrow.get(year,nextmonth,15) - r
ft = fp+fn
fp = fp.days/ft.days
fn = fn.days/ft.days
print ft, fp, fn
driverflow = fn*criverflow[prevmonth-1] + fp*criverflow[nextmonth-1]
return driverflow
def write_file(r,flow,lat,lon,riverdepth):
''' given the flow and the riverdepth and the date, write the nc file'''
directory = '.'
# set up filename to follow NEMO conventions
filename = 'RFraserCElse_y'+str(year)+'m'+'{:0=2}'.format(r.month)+'d'+'{:0=2}'.format(r.day)+'.nc'
# print directory+'/'+filename
nemo = nc.Dataset(directory+'/'+filename, 'w')
nemo.description = 'Real Fraser Values, Daily Climatology for Other Rivers'
# dimensions
ymax, xmax = lat.shape
nemo.createDimension('x', xmax)
nemo.createDimension('y', ymax)
nemo.createDimension('time_counter', None)
# variables
# latitude and longitude
nav_lat = nemo.createVariable('nav_lat','float32',('y','x'),zlib=True)
nav_lat = lat
x = nemo.createVariable('nav_lon','float32',('y','x'),zlib=True)
nav_lon = lon
# time
time_counter = nemo.createVariable('time_counter', 'float32', ('time_counter'),zlib=True)
time_counter.units = 'non-dim'
time_counter[0:1] = range(1,2)
# runoff
rorunoff = nemo.createVariable('rorunoff', 'float32', ('time_counter','y','x'), zlib=True)
rorunoff._Fillvalue = 0.
rorunoff._missing_value = 0.
rorunoff._units = 'kg m-2 s-1'
rorunoff[0,:] = flow
# depth
rodepth = nemo.createVariable('rodepth','float32',('y','x'),zlib=True)
rodepth._Fillvalue = -1.
rodepth.missing_value = -1.
rodepth.units = 'm'
rodepth = riverdepth
nemo.close()
return
def fraser_correction(pd, fraserflux, r, afterHope, NonFraser, fraserratio, otherratio,
runoff):
''' for the Fraser Basin only, replace basic values with the new climatology after Hope and the
observed values for Hope. Note, we are changing runoff only and not using/changing river
depth '''
for key in pd:
if "Fraser" in key:
flux = calculate_daily_flow(r,afterHope) + fraserflux
subarea = fraserratio
else:
flux = calculate_daily_flow(r,NonFraser)
subarea = otherratio
river = pd[key]
runoff = rivertools.fill_runoff_array(flux*river['prop']/subarea,river['i'],
river['di'],river['j'],river['dj'],river['depth'],
runoff,np.empty_like(runoff))[0]
return runoff
# # load the re-written climatology river files
##open climatolgy file with modified fresh water point source
clim_rivers_edit = nc.Dataset('rivers_month_edit.nc','r' )
criverflow_edit = clim_rivers_edit.variables['rorunoff']
# +
for r in arrow.Arrow.range('day', startdate, enddate):
print r
driverflow = calculate_daily_flow(r, criverflow_edit)
storeflow = calculate_daily_flow(r, criverflow_edit)
step1 = fraserflow[fraserflow[:,0] == r.year]
step2 = step1[step1[:,1] == r.month]
step3 = step2[step2[:,2] == r.day]
# print r.year, r.month, r.day, step3[0,3]
runoff = fraser_correction(pd, step3[0,3] , r, afterHope, nonFraser, fraserratio, otherratio,
driverflow)
write_file(r,runoff,lat,lon,riverdepth)
ig = 418
jg = 397
print criverflow_edit[7:10,418,397], driverflow[ig,jg]
print storeflow[ig,jg], driverflow[ig,jg]
ig = 351; jg = 345
print storeflow[ig,jg], driverflow[ig,jg]
ig = 749; jg=123
print storeflow[ig,jg], driverflow[ig,jg]
# jan 0, feb 1, mar 2, apr 3, may 4, jun 5
# jul 6, aug 7, sep 8
# -
| jie/New river discharge file.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# coding: utf-8
import sys, os
sys.path.append(os.pardir) # 親ディレクトリのファイルをインポートするための設定
import pickle
import numpy as np
from collections import OrderedDict
from common.layers import *
from common.gradient import numerical_gradient
class SimpleConvNet:
"""単純なConvNet
conv - relu - pool - affine - relu - affine - softmax
Parameters
----------
input_size : 入力サイズ(MNISTの場合は784)
hidden_size_list : 隠れ層のニューロンの数のリスト(e.g. [100, 100, 100])
output_size : 出力サイズ(MNISTの場合は10)
activation : 'relu' or 'sigmoid'
weight_init_std : 重みの標準偏差を指定(e.g. 0.01)
'relu'または'he'を指定した場合は「Heの初期値」を設定
'sigmoid'または'xavier'を指定した場合は「Xavierの初期値」を設定
"""
def __init__(self, input_dim=(1, 28, 28),
conv_param={'filter_num':30, 'filter_size':5, 'pad':0, 'stride':1},
hidden_size=100, output_size=10, weight_init_std=0.01):
filter_num = conv_param['filter_num']
filter_size = conv_param['filter_size']
filter_pad = conv_param['pad']
filter_stride = conv_param['stride']
input_size = input_dim[1]
conv_output_size = (input_size - filter_size + 2*filter_pad) / filter_stride + 1
pool_output_size = int(filter_num * (conv_output_size/2) * (conv_output_size/2))
# 重みの初期化
self.params = {}
self.params['W1'] = weight_init_std * \
np.random.randn(filter_num, input_dim[0], filter_size, filter_size)
self.params['b1'] = np.zeros(filter_num)
self.params['W2'] = weight_init_std * \
np.random.randn(pool_output_size, hidden_size)
self.params['b2'] = np.zeros(hidden_size)
self.params['W3'] = weight_init_std * \
np.random.randn(hidden_size, output_size)
self.params['b3'] = np.zeros(output_size)
# レイヤの生成
self.layers = OrderedDict()
self.layers['Conv1'] = Convolution(self.params['W1'], self.params['b1'],
conv_param['stride'], conv_param['pad'])
self.layers['Relu1'] = Relu()
self.layers['Pool1'] = Pooling(pool_h=2, pool_w=2, stride=2)
self.layers['Affine1'] = Affine(self.params['W2'], self.params['b2'])
self.layers['Relu2'] = Relu()
self.layers['Affine2'] = Affine(self.params['W3'], self.params['b3'])
self.last_layer = SoftmaxWithLoss()
def predict(self, x):
for layer in self.layers.values():
x = layer.forward(x)
return x
def loss(self, x, t):
"""損失関数を求める
引数のxは入力データ、tは教師ラベル
"""
y = self.predict(x)
return self.last_layer.forward(y, t)
def accuracy(self, x, t, batch_size=100):
if t.ndim != 1 : t = np.argmax(t, axis=1)
acc = 0.0
for i in range(int(x.shape[0] / batch_size)):
tx = x[i*batch_size:(i+1)*batch_size]
tt = t[i*batch_size:(i+1)*batch_size]
y = self.predict(tx)
y = np.argmax(y, axis=1)
acc += np.sum(y == tt)
return acc / x.shape[0]
def numerical_gradient(self, x, t):
"""勾配を求める(数値微分)
Parameters
----------
x : 入力データ
t : 教師ラベル
Returns
-------
各層の勾配を持ったディクショナリ変数
grads['W1']、grads['W2']、...は各層の重み
grads['b1']、grads['b2']、...は各層のバイアス
"""
loss_w = lambda w: self.loss(x, t)
grads = {}
for idx in (1, 2, 3):
grads['W' + str(idx)] = numerical_gradient(loss_w, self.params['W' + str(idx)])
grads['b' + str(idx)] = numerical_gradient(loss_w, self.params['b' + str(idx)])
return grads
def gradient(self, x, t):
"""勾配を求める(誤差逆伝搬法)
Parameters
----------
x : 入力データ
t : 教師ラベル
Returns
-------
各層の勾配を持ったディクショナリ変数
grads['W1']、grads['W2']、...は各層の重み
grads['b1']、grads['b2']、...は各層のバイアス
"""
# forward
self.loss(x, t)
# backward
dout = 1
dout = self.last_layer.backward(dout)
layers = list(self.layers.values())
layers.reverse()
for layer in layers:
dout = layer.backward(dout)
# 設定
grads = {}
grads['W1'], grads['b1'] = self.layers['Conv1'].dW, self.layers['Conv1'].db
grads['W2'], grads['b2'] = self.layers['Affine1'].dW, self.layers['Affine1'].db
grads['W3'], grads['b3'] = self.layers['Affine2'].dW, self.layers['Affine2'].db
return grads
def save_params(self, file_name="params.pkl"):
params = {}
for key, val in self.params.items():
params[key] = val
with open(file_name, 'wb') as f:
pickle.dump(params, f)
def load_params(self, file_name="params.pkl"):
with open(file_name, 'rb') as f:
params = pickle.load(f)
for key, val in params.items():
self.params[key] = val
for i, key in enumerate(['Conv1', 'Affine1', 'Affine2']):
self.layers[key].W = self.params['W' + str(i+1)]
self.layers[key].b = self.params['b' + str(i+1)]
| ch7/simple_convnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="ISubpr_SSsiM"
# ##### Copyright 2019 The TensorFlow Authors.
#
# + cellView="form" colab={} colab_type="code" id="3jTMb1dySr3V"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="6DWfyNThSziV"
# # tf.function
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/beta/tutorials/eager/tf_function"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r2/tutorials/eager/tf_function.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r2/tutorials/eager/tf_function.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/r2/tutorials/eager/tf_function.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="J122XQYG7W6w"
#
# In TensorFlow 2.0, eager execution is turned on by default. The user interface is intuitive and flexible (running one-off operations is much easier
# and faster), but this can come at the expense of performance and deployability.
#
# To get peak performance and to make your model deployable anywhere, use
# `tf.function` to make graphs out of your programs.
# Thanks to AutoGraph, a surprising amount of Python code just works with
# tf.function, but there are still pitfalls to be wary of.
#
# The main takeaways and recommendations are:
#
# - Don't rely on Python side effects like object mutation or list appends.
# - tf.function works best with TensorFlow ops, rather than NumPy ops or Python primitives.
# - When in doubt, use the `for x in y` idiom.
# + colab={} colab_type="code" id="otIdN1TS8N7S"
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# %tensorflow_version 2.x # Colab only.
except Exception:
pass
import tensorflow as tf
# + colab={} colab_type="code" id="D25apou9IOXa"
import contextlib
# Some helper code to demonstrate the kinds of errors you might encounter.
@contextlib.contextmanager
def assert_raises(error_class):
try:
yield
except error_class as e:
print('Caught expected exception \n {}: {}'.format(error_class, e))
except Exception as e:
print('Got unexpected exception \n {}: {}'.format(type(e), e))
else:
raise Exception('Expected {} to be raised but no error was raised!'.format(
error_class))
# + [markdown] colab_type="text" id="rfayNj-ZIkIB"
# A `tf.function` you define is just like a core TensorFlow operation: You can execute it eagerly; you can use it in a graph; it has gradients; and so on.
# + colab={} colab_type="code" id="SbtT1-Wm70F2"
# A function is like an op
@tf.function
def add(a, b):
return a + b
add(tf.ones([2, 2]), tf.ones([2, 2])) # [[2., 2.], [2., 2.]]
# + colab={} colab_type="code" id="uP-zUelB8DbX"
# Functions have gradients
@tf.function
def add(a, b):
return a + b
v = tf.Variable(1.0)
with tf.GradientTape() as tape:
result = add(v, 1.0)
tape.gradient(result, v)
# + colab={} colab_type="code" id="l5qRjdbBVdU6"
# You can use functions inside functions
@tf.function
def dense_layer(x, w, b):
return add(tf.matmul(x, w), b)
dense_layer(tf.ones([3, 2]), tf.ones([2, 2]), tf.ones([2]))
# + [markdown] colab_type="text" id="uZ4Do2AV80cO"
# ## Tracing and polymorphism
#
# Python's dynamic typing means that you can call functions with a variety of argument types, and Python will do something different in each scenario.
#
# On the other hand, TensorFlow graphs require static dtypes and shape dimensions. `tf.function` bridges this gap by retracing the function when necessary to generate the correct graphs. Most of the subtlety of `tf.function` usage stems from this retracing behavior.
#
# You can call a function with arguments of different types to see what is happening.
# + colab={} colab_type="code" id="kojmJrgq8U9v"
# Functions are polymorphic
@tf.function
def double(a):
print("Tracing with", a)
return a + a
print(double(tf.constant(1)))
print()
print(double(tf.constant(1.1)))
print()
print(double(tf.constant("a")))
print()
# + [markdown] colab_type="text" id="4pJqkDR_Q2wz"
# To control the tracing behavior, use the following techniques:
#
# - Create a new `tf.function`. Separate `tf.function` objects are guaranteed not to share traces.
# - Use the `get_concrete_function` method to get a specific trace
# - Specify `input_signature` when calling `tf.function` to trace only once per calling graph.
# + colab={} colab_type="code" id="mHg2CGtPQ3Hz"
print("Obtaining concrete trace")
double_strings = double.get_concrete_function(tf.TensorSpec(shape=None, dtype=tf.string))
print("Executing traced function")
print(double_strings(tf.constant("a")))
print(double_strings(a=tf.constant("b")))
print("Using a concrete trace with incompatible types will throw an error")
with assert_raises(tf.errors.InvalidArgumentError):
double_strings(tf.constant(1))
# + colab={} colab_type="code" id="_BDMIRmu1RGB"
@tf.function(input_signature=(tf.TensorSpec(shape=[None], dtype=tf.int32),))
def next_collatz(x):
print("Tracing with", x)
return tf.where(tf.equal(x % 2, 0), x // 2, 3 * x + 1)
print(next_collatz(tf.constant([1, 2])))
# We specified a 1-D tensor in the input signature, so this should fail.
with assert_raises(ValueError):
next_collatz(tf.constant([[1, 2], [3, 4]]))
# + [markdown] colab_type="text" id="Es0WZkLIUSdu"
# ## When to retrace?
#
# A polymorphic `tf.function` keeps a cache of concrete functions generated by tracing. The cache keys are effectively tuples of keys generated from the function args and kwargs. The key generated for a `tf.Tensor` argument is its shape and type. The key generated for a Python primitive is its value. For all other Python types, the keys are based on the object `id()` so that methods are traced independently for each instance of a class. In the future, TensorFlow may add more sophisticated caching for Python objects that can be safely converted to tensors.
# + [markdown] colab_type="text" id="AY5oiQN0XIyA"
# ## Python or Tensor args?
#
# Often, Python arguments are used to control hyperparameters and graph constructions - for example, `num_layers=10` or `training=True` or `nonlinearity='relu'`. So if the Python argument changes, it makes sense that you'd have to retrace the graph.
#
# However, it's possible that a Python argument is not being used to control graph construction. In these cases, a change in the Python value can trigger needless retracing. Take, for example, this training loop, which AutoGraph will dynamically unroll. Despite the multiple traces, the generated graph is actually identical, so this is a bit inefficient.
# + colab={} colab_type="code" id="uydzR5JYUU8H"
def train_one_step():
pass
@tf.function
def train(num_steps):
print("Tracing with num_steps = {}".format(num_steps))
for _ in tf.range(num_steps):
train_one_step()
train(num_steps=10)
train(num_steps=20)
# + [markdown] colab_type="text" id="f6pjnylLUW8P"
# The simple workaround here is to cast your arguments to Tensors if they do not affect the shape of the generated graph.
# + colab={} colab_type="code" id="TmL8T-w3UYes"
train(num_steps=tf.constant(10))
train(num_steps=tf.constant(20))
# + [markdown] colab_type="text" id="129-iRsPS-gY"
# ## Side effects in `tf.function`
#
# In general, Python side effects (like printing or mutating objects) only happen during tracing. So how can you reliably trigger side effects from `tf.function`?
#
# The general rule of thumb is to only use Python side effects to debug your traces. Otherwise, TensorFlow ops like `tf.Variable.assign`, `tf.print`, and `tf.summary` are the best way to ensure your code will be traced and executed by the TensorFlow runtime with each call. In general using a functional style will yield the best results.
# + colab={} colab_type="code" id="w2sACuZ9TTRk"
@tf.function
def f(x):
print("Traced with", x)
tf.print("Executed with", x)
f(1)
f(1)
f(2)
# + [markdown] colab_type="text" id="e1I0dPiqTV8H"
# If you would like to execute Python code during each invocation of a `tf.function`, `tf.py_function` is an exit hatch. The drawback of `tf.py_function` is that it's not portable or particularly performant, nor does it work well in distributed (multi-GPU, TPU) setups. Also, since `tf.py_function` has to be wired into the graph, it casts all inputs/outputs to tensors.
# + colab={} colab_type="code" id="7aJD--9qTWmg"
external_list = []
def side_effect(x):
print('Python side effect')
external_list.append(x)
@tf.function
def f(x):
tf.py_function(side_effect, inp=[x], Tout=[])
f(1)
f(1)
f(1)
assert len(external_list) == 3
# .numpy() call required because py_function casts 1 to tf.constant(1)
assert external_list[0].numpy() == 1
# + [markdown] colab_type="text" id="msTmv-oyUNaf"
# ## Beware of Python state
#
# Many Python features, such as generators and iterators, rely on the Python runtime to keep track of state. In general, while these constructs work as expected in Eager mode, many unexpected things can happen inside a `tf.function` due to tracing behavior.
#
# To give one example, advancing iterator state is a Python side effect and therefore only happens during tracing.
# + colab={} colab_type="code" id="FNPD4unZUedH"
external_var = tf.Variable(0)
@tf.function
def buggy_consume_next(iterator):
external_var.assign_add(next(iterator))
tf.print("Value of external_var:", external_var)
iterator = iter([0, 1, 2, 3])
buggy_consume_next(iterator)
# This reuses the first value from the iterator, rather than consuming the next value.
buggy_consume_next(iterator)
buggy_consume_next(iterator)
# + [markdown] colab_type="text" id="5XMGXMu-Ufjm"
# If an iterator is generated and consumed entirely within the tf.function, then it should work correctly. However, the entire iterator is probably being traced, which can lead to a giant graph. This may be what you want. But if you're training on an large in-memory dataset represented as a Python list, then this can generate a very large graph, and `tf.function` is unlikely to yield a speedup.
#
# If you want to iterate over Python data, the safest way is to wrap it in a tf.data.Dataset and use the `for x in y` idiom. AutoGraph has special support for safely converting `for` loops when `y` is a tensor or tf.data.Dataset.
#
# + colab={} colab_type="code" id="ms7f1o_QUiHE"
def measure_graph_size(f, *args):
g = f.get_concrete_function(*args).graph
print("{}({}) contains {} nodes in its graph".format(
f.__name__, ', '.join(map(str, args)), len(g.as_graph_def().node)))
@tf.function
def train(dataset):
loss = tf.constant(0)
for x, y in dataset:
loss += tf.abs(y - x) # Some dummy computation.
return loss
small_data = [(1, 1)] * 2
big_data = [(1, 1)] * 10
measure_graph_size(train, small_data)
measure_graph_size(train, big_data)
measure_graph_size(train, tf.data.Dataset.from_generator(
lambda: small_data, (tf.int32, tf.int32)))
measure_graph_size(train, tf.data.Dataset.from_generator(
lambda: big_data, (tf.int32, tf.int32)))
# + [markdown] colab_type="text" id="dGDstsFpWHEI"
#
# When wrapping Python/Numpy data in a Dataset, be mindful of `tf.data.Dataset.from_generator` versus ` tf.data.Dataset.from_tensors`. The former will keep the data in Python and fetch it via `tf.py_function` which can have performance implications, whereas the latter will bundle a copy of the data as one large `tf.constant()` node in the graph, which can have memory implications.
#
# Reading data from files via TFRecordDataset/CsvDataset/etc. is the most effective way to consume data, as then TensorFlow itself can manage the asynchronous loading and prefetching of data, without having to involve Python.
# + [markdown] colab_type="text" id="tRdlnCfV_UTn"
# ## Automatic Control Dependencies
#
# A very appealing property of functions as the programming model, over a general dataflow graph, is that functions can give the runtime more information about what was the intended behavior of the code.
#
# For example, when writing code which has multiple reads and writes to the same variables, a dataflow graph might not naturally encode the originally intended order of operations. In `tf.function`, we resolve ambiguities in execution order by referring to the execution order of statements in the original Python code. This way, ordering of stateful operations in a `tf.function` replicates the semantics of Eager mode.
#
# This means there's no need to add manual control dependencies; `tf.function` is smart enough to add the minimal set of necessary and sufficient control dependencies for your code to run correctly.
# + colab={} colab_type="code" id="SASm0ss8erVX"
# Automatic control dependencies
a = tf.Variable(1.0)
b = tf.Variable(2.0)
@tf.function
def f(x, y):
a.assign(y * b)
b.assign_add(x * a)
return a + b
f(1.0, 2.0) # 10.0
# + [markdown] colab_type="text" id="lPr_6mK_AQWL"
# ## Variables
#
# We can use the same idea of leveraging the intended execution order of the code to make variable creation and utilization very easy in `tf.function`. There is one very important caveat, though, which is that with variables it's possible to write code which behaves differently in eager mode and graph mode.
#
# Specifically, this will happen when you create a new Variable with each call. Due to tracing semantics, `tf.function` will reuse the same variable each call, but eager mode will create a new variable with each call. To guard against this mistake, `tf.function` will raise an error if it detects dangerous variable creation behavior.
# + colab={} colab_type="code" id="Tx0Vvnb_9OB-"
@tf.function
def f(x):
v = tf.Variable(1.0)
v.assign_add(x)
return v
with assert_raises(ValueError):
f(1.0)
# + colab={} colab_type="code" id="DKzNjVg8h4ao"
# Non-ambiguous code is ok though
v = tf.Variable(1.0)
@tf.function
def f(x):
return v.assign_add(x)
print(f(1.0)) # 2.0
print(f(2.0)) # 4.0
# + colab={} colab_type="code" id="HQrG5_kOiKl_"
# You can also create variables inside a tf.function as long as we can prove
# that those variables are created only the first time the function is executed.
class C: pass
obj = C(); obj.v = None
@tf.function
def g(x):
if obj.v is None:
obj.v = tf.Variable(1.0)
return obj.v.assign_add(x)
print(g(1.0)) # 2.0
print(g(2.0)) # 4.0
# + colab={} colab_type="code" id="_IOVc1eujMH2"
# Variable initializers can depend on function arguments and on values of other
# variables. We can figure out the right initialization order using the same
# method we use to generate control dependencies.
state = []
@tf.function
def fn(x):
if not state:
state.append(tf.Variable(2.0 * x))
state.append(tf.Variable(state[0] * 3.0))
return state[0] * x * state[1]
print(fn(tf.constant(1.0)))
print(fn(tf.constant(3.0)))
# + [markdown] colab_type="text" id="5f05Vr_YBUCz"
# # Using AutoGraph
#
# The [autograph](https://www.tensorflow.org/guide/autograph) library is fully integrated with `tf.function`, and it will rewrite conditionals and loops which depend on Tensors to run dynamically in the graph.
#
# `tf.cond` and `tf.while_loop` continue to work with `tf.function`, but code with control flow is often easier to write and understand when written in imperative style.
# + colab={} colab_type="code" id="yCQTtTPTW3WF"
# Simple loop
@tf.function
def f(x):
while tf.reduce_sum(x) > 1:
tf.print(x)
x = tf.tanh(x)
return x
f(tf.random.uniform([5]))
# + colab={} colab_type="code" id="jlQD1ffRXJhl"
# If you're curious you can inspect the code autograph generates.
# It feels like reading assembly language, though.
def f(x):
while tf.reduce_sum(x) > 1:
tf.print(x)
x = tf.tanh(x)
return x
print(tf.autograph.to_code(f))
# + [markdown] colab_type="text" id="xgKmkrNTZSyz"
# ## AutoGraph: Conditionals
#
# AutoGraph will convert `if` statements into the equivalent `tf.cond` calls.
#
# This substitution is made if the condition is a Tensor. Otherwise, the conditional is executed during tracing.
# + colab={} colab_type="code" id="E-7KllizZYsy"
def test_tf_cond(f, *args):
g = f.get_concrete_function(*args).graph
if any(node.name == 'cond' for node in g.as_graph_def().node):
print("{}({}) uses tf.cond.".format(
f.__name__, ', '.join(map(str, args))))
else:
print("{}({}) executes normally.".format(
f.__name__, ', '.join(map(str, args))))
# + colab={} colab_type="code" id="o86paGR-Zadi"
@tf.function
def hyperparam_cond(x, training=True):
if training:
x = tf.nn.dropout(x, rate=0.5)
return x
@tf.function
def maybe_tensor_cond(x):
if x < 0:
x = -x
return x
test_tf_cond(hyperparam_cond, tf.ones([1], dtype=tf.float32))
test_tf_cond(maybe_tensor_cond, tf.constant(-1))
test_tf_cond(maybe_tensor_cond, -1)
# + [markdown] colab_type="text" id="5xFLfdApZh8q"
# `tf.cond` has a number of subtleties.
# - it works by tracing both sides of the conditional, and then choosing the appropriate branch at runtime, depending on the condition. Tracing both sides can result in unexpected execution of Python code
# - it requires that if one branch creates a tensor used downstream, the other branch must also create that tensor.
#
# + colab={} colab_type="code" id="VTMoZEVaZiwk"
@tf.function
def f():
x = tf.constant(0)
if tf.constant(True):
x = x + 1
print("Tracing `then` branch")
else:
x = x - 1
print("Tracing `else` branch")
return x
f()
# + colab={} colab_type="code" id="k_dxWHeFZlaQ"
@tf.function
def f():
if tf.constant(True):
x = tf.ones([3, 3])
return x
# Throws an error because both branches need to define `x`.
with assert_raises(ValueError):
f()
# + [markdown] colab_type="text" id="yho4J0a0ZkQS"
# ## AutoGraph and loops
#
# AutoGraph has a few simple rules for converting loops.
#
# - `for`: Convert if the iterable is a tensor
# - `while`: Convert if the while condition depends on a tensor
#
# If a loop is converted, it will be dynamically unrolled with `tf.while_loop`, or in the special case of a `for x in tf.data.Dataset`, transformed into `tf.data.Dataset.reduce`.
#
# If a loop is _not_ converted, it will be statically unrolled
# + colab={} colab_type="code" id="OyzGNQAuZsky"
def test_dynamically_unrolled(f, *args):
g = f.get_concrete_function(*args).graph
if any(node.name == 'while' for node in g.as_graph_def().node):
print("{}({}) uses tf.while_loop.".format(
f.__name__, ', '.join(map(str, args))))
elif any(node.name == 'ReduceDataset' for node in g.as_graph_def().node):
print("{}({}) uses tf.data.Dataset.reduce.".format(
f.__name__, ', '.join(map(str, args))))
else:
print("{}({}) gets unrolled.".format(
f.__name__, ', '.join(map(str, args))))
# + colab={} colab_type="code" id="Q7tmncQTZt6_"
@tf.function
def for_in_range():
x = 0
for i in range(5):
x += i
return x
@tf.function
def for_in_tfrange():
x = tf.constant(0, dtype=tf.int32)
for i in tf.range(5):
x += i
return x
@tf.function
def for_in_tfdataset():
x = tf.constant(0, dtype=tf.int64)
for i in tf.data.Dataset.range(5):
x += i
return x
test_dynamically_unrolled(for_in_range)
test_dynamically_unrolled(for_in_tfrange)
test_dynamically_unrolled(for_in_tfdataset)
# + colab={} colab_type="code" id="l6s7aU-padY5"
@tf.function
def while_py_cond():
x = 5
while x > 0:
x -= 1
return x
@tf.function
def while_tf_cond():
x = tf.constant(5)
while x > 0:
x -= 1
return x
test_dynamically_unrolled(while_py_cond)
test_dynamically_unrolled(while_tf_cond)
# + [markdown] colab_type="text" id="dSr64Xn6ap-S"
# If you have a `break` or early `return` clause that depends on a tensor, the top-level condition or iterable should also be a tensor.
# + colab={} colab_type="code" id="Q-VirD-5avdZ"
@tf.function
def buggy_while_py_true_tf_break(x):
while True:
if tf.equal(x, 0):
break
x -= 1
return x
@tf.function
def while_tf_true_tf_break(x):
while tf.constant(True):
if tf.equal(x, 0):
break
x -= 1
return x
with assert_raises(TypeError):
test_dynamically_unrolled(buggy_while_py_true_tf_break, 5)
test_dynamically_unrolled(while_tf_true_tf_break, 5)
@tf.function
def buggy_py_for_tf_break():
x = 0
for i in range(5):
if tf.equal(i, 3):
break
x += i
return x
@tf.function
def tf_for_tf_break():
x = 0
for i in tf.range(5):
if tf.equal(i, 3):
break
x += i
return x
with assert_raises(TypeError):
test_dynamically_unrolled(buggy_py_for_tf_break)
test_dynamically_unrolled(tf_for_tf_break)
# + [markdown] colab_type="text" id="hyksHW9TCukR"
# In order to accumulate results from a dynamically unrolled loop, you'll want to use `tf.TensorArray`.
#
# + colab={} colab_type="code" id="HJ3Vb3dXfefN"
batch_size = 2
seq_len = 3
feature_size = 4
def rnn_step(inp, state):
return inp + state
@tf.function
def dynamic_rnn(rnn_step, input_data, initial_state):
# [batch, time, features] -> [time, batch, features]
input_data = tf.transpose(input_data, [1, 0, 2])
max_seq_len = input_data.shape[0]
states = tf.TensorArray(tf.float32, size=max_seq_len)
state = initial_state
for i in tf.range(max_seq_len):
state = rnn_step(input_data[i], state)
states = states.write(i, state)
return tf.transpose(states.stack(), [1, 0, 2])
dynamic_rnn(rnn_step,
tf.random.uniform([batch_size, seq_len, feature_size]),
tf.zeros([batch_size, feature_size]))
# + [markdown] colab_type="text" id="9gmLpHY-bkly"
# As with `tf.cond`, `tf.while_loop` also comes with a number of subtleties.
# - Since a loop can execute 0 times, all tensors used downstream of the while_loop must be initialized above the loop
# - The shape/dtypes of all loop variables must stay consistent with each iteration
# + colab={} colab_type="code" id="CocT5RHwblrQ"
@tf.function
def buggy_loop_var_uninitialized():
for i in tf.range(3):
x = i
return x
@tf.function
def f():
x = tf.constant(0)
for i in tf.range(3):
x = i
return x
with assert_raises(ValueError):
buggy_loop_var_uninitialized()
f()
# + colab={} colab_type="code" id="FSftc9cCbpAo"
@tf.function
def buggy_loop_type_changes():
x = tf.constant(0, dtype=tf.float32)
for i in tf.range(3): # Yields tensors of type tf.int32...
x = i
return x
with assert_raises(tf.errors.InvalidArgumentError):
buggy_loop_type_changes()
# + colab={} colab_type="code" id="kWF189prbuK0"
@tf.function
def buggy_concat():
x = tf.ones([0, 10])
for i in tf.range(5):
x = tf.concat([x, tf.ones([1, 10])], axis=0)
return x
with assert_raises(ValueError):
buggy_concat()
@tf.function
def concat_with_padding():
x = tf.zeros([5, 10])
for i in tf.range(5):
x = tf.concat([x[:i], tf.ones([1, 10]), tf.zeros([4-i, 10])], axis=0)
x.set_shape([5, 10])
return x
concat_with_padding()
| site/en/r2/tutorials/eager/tf_function.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from chainer import datasets, serializers
from cvae import IAFCVAE
import matplotlib.pyplot as plt
# +
result_dir = "result/cae-mnist"
_, test = datasets.get_mnist(withlabel=False, ndim=3)
h_channel = 2
params = dict(
in_channel=1,
h_channel=h_channel,
depth=1,
n_iaf_block=1,
iaf_params=dict(
in_dim=h_channel,
z_dim=2,
h_dim=h_channel,
ksize=3,
pad=1,
)
)
model = IAFCVAE(**params)
serializers.load_npz(result_dir + "/model_weights.npz", model)
# -
recon, _, _ = model(test[0:10])
# +
nplot = 10
figs = plt.figure(figsize=(10, 4))
for n in range(nplot):
ax = figs.add_subplot(2, nplot, n+1)
ax.imshow(recon.data[n][0])
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
ax.set_title("recon")
ax = figs.add_subplot(2, nplot, nplot+n+1)
gt_result = test[n][0]
ax.imshow(gt_result)
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
ax.set_title("gt")
plt.show()
# -
import numpy as np
img = model.generate(np.array([0.0,0.0], dtype="float32"), batch_size=10)
# +
import numpy as np
img = model.generate(np.array([0.0,0.0], dtype="float32"), batch_size=10)
nplot = 10
figs = plt.figure(figsize=(10, 4))
for n in range(nplot):
ax = figs.add_subplot(2, nplot, n+1)
ax.imshow(img.data[n][0])
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
ax.set_title("gen")
plt.show()
# -
model.iaf_layers[0].forward_down(np.array([0.,0.], dtype="float32"),sample=True)
| examples/test_cvae_iaf_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Day 5
# 7, 100
# ### Import modules
# %matplotlib inline
from __future__ import division
import sys
import os
os.environ['MKL_THREADING_LAYER']='GNU'
sys.path.append('../')
from Modules.Basics import *
from Modules.Class_Basics import *
# ## Options
# +
classTrainFeatures = ['DER_mass_MMC', 'DER_mass_transverse_met_lep', 'DER_mass_vis', 'DER_pt_h', 'DER_deltaeta_jet_jet', 'DER_mass_jet_jet', 'DER_prodeta_jet_jet', 'DER_deltar_tau_lep', 'DER_pt_tot', 'DER_sum_pt', 'DER_pt_ratio_lep_tau', 'DER_met_phi_centrality', 'DER_lep_eta_centrality', 'PRI_met_pt', 'PRI_met_sumet', 'PRI_jet_num', 'PRI_jet_all_pt', 'PRI_tau_px', 'PRI_tau_py', 'PRI_tau_pz', 'PRI_lep_px', 'PRI_lep_pz', 'PRI_jet_leading_px', 'PRI_jet_leading_py', 'PRI_jet_leading_pz', 'PRI_jet_subleading_px', 'PRI_jet_subleading_py', 'PRI_jet_subleading_pz', 'PRI_met_px', 'PRI_met_py']
inputPipe, outputPipe = getPreProcPipes(normIn=True)
classModel = 'modelSwish-7-100'
varSet = "basic_rot_features"
nSplits = 10
ensembleSize = 10
ensembleMode = 'loss'
maxEpochs = 200
compileArgs = {'loss':'binary_crossentropy', 'optimizer':'adam', 'depth':7, 'width':50}
trainParams = {'epochs' : 1, 'batch_size' : 256, 'verbose' : 0}
modelParams = {'version':classModel, 'nIn':len(classTrainFeatures), 'compileArgs':compileArgs}
print "\nTraining on", len(classTrainFeatures), "features:", [var for var in classTrainFeatures]
# -
# ## Import data
trainData = h5py.File(dirLoc + 'train.hdf5', "r+")
valData = h5py.File(dirLoc + 'val.hdf5', "r+")
# ## Determine LR
lrFinder = batchLRFindClassifier(trainData, nSplits, getClassifier, modelParams, trainParams, lrBounds=[1e-5,1e-1], trainOnWeights=True, verbose=0)
compileArgs['lr'] = 5e-3
# ## Train classifier
results, histories = batchTrainClassifier(trainData, nSplits, getClassifier, modelParams, trainParams, patience=100, cosAnnealMult=2, trainOnWeights=True, maxEpochs=maxEpochs, verbose=1)
# ## Construct ensemble
with open('train_weights/resultsFile.pkl', 'r') as fin:
results = pickle.load(fin)
ensemble, weights = assembleEnsemble(results, ensembleSize, ensembleMode, compileArgs)
# ## Response on development data
batchEnsemblePredict(ensemble, weights, trainData, ensembleSize=10, verbose=1)
print 'Training ROC AUC: unweighted {}, weighted {}'.format(roc_auc_score(getFeature('targets', trainData), getFeature('pred', trainData)),
roc_auc_score(getFeature('targets', trainData), getFeature('pred', trainData), sample_weight=getFeature('weights', trainData)))
# ## Response on val data
batchEnsemblePredict(ensemble, weights, valData, ensembleSize=10, verbose=1)
print 'Testing ROC AUC: unweighted {}, weighted {}'.format(roc_auc_score(getFeature('targets', valData), getFeature('pred', valData)),
roc_auc_score(getFeature('targets', valData), getFeature('pred', valData), sample_weight=getFeature('weights', valData)))
# ## Evaluation
# ### Import in dataframe
valframe = convertToDF(valData)
sigVal = (valframe.gen_target == 1)
bkgVal = (valframe.gen_target == 0)
# ### MVA distributions
getClassPredPlot([valframe[bkgVal], valframe[sigVal]], weightName='gen_weight')
valframe['ams'] = amsScan(valframe)[0]
bests = foldAMSScan(valframe, 10)
# +
def scoreTest(ensemble, weights):
testData = h5py.File(dirLoc + 'testing.hdf5', "r+")
batchEnsemblePredict(ensemble, weights, testData, ensembleSize=10, verbose=1)
def saveTest(cut, name):
testData = h5py.File(dirLoc + 'testing.hdf5', "r+")
data = pandas.DataFrame()
data['EventId'] = getFeature('EventId', testData)
data['pred_class'] = getFeature('pred', testData)
data['Class'] = 'b'
data.loc[data.pred_class >= cut, 'Class'] = 's'
data.sort_values(by=['pred_class'], inplace=True)
data['RankOrder']=range(1, len(data)+1)
data.sort_values(by=['EventId'], inplace=True)
print dirLoc + name + '_test.csv'
data.to_csv(dirLoc + name + '_test.csv', columns=['EventId', 'RankOrder', 'Class'], index=False)
# -
scoreTest(ensemble, weights)
def saveTest(cut, name):
testData = h5py.File(dirLoc + 'testing.hdf5', "r+")
data = pandas.DataFrame()
data['EventId'] = getFeature('EventId', testData)
data['pred_class'] = getFeature('pred', testData)
data['Class'] = 'b'
data.loc[data.pred_class >= cut, 'Class'] = 's'
data.sort_values(by=['pred_class'], inplace=True)
data['RankOrder']=range(1, len(data)+1)
data.sort_values(by=['EventId'], inplace=True)
print dirLoc + name + '_test.csv'
data.to_csv(dirLoc + name + '_test.csv', columns=['EventId', 'RankOrder', 'Class'], index=False)
saveTest(0.957172385359, 'Day_5')
# !kaggle competitions submit -c higgs-boson -f ../Data/Day_5_test.csv -m"7-50"
| Classifiers/Day5/Day_5_7-50.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hello GCN
import numpy as np
import networkx
from matplotlib import pyplot as plt
def ReLU(x):
return (abs(x) + x) / 2
# ## GCN 的输入
#
# 与传统网络不同的是,$GCN$ 的输入并不只是的特征,它额外包含了图中各个结点之间的关系,那么关系用什么表征呢?显而易见,邻接矩阵是一个非常简单易行的表示方法,这里**使用邻接矩阵 $A$ 来表示各个顶点之间的关系**,显然 $A.shape = (N, N)$
#
# 相应地,各个顶点的特征使用矩阵组织起来就好啦,这里使用 $X$ 表示输入层特征矩阵, $H^i$ 表示第 $i$ 隐藏层特征矩阵,当然 $X = H^0$ ,显然 $H^i.shape = (N, F^i)$
#
# > $F^i$ 表示第 $i$ 层的特征维度,$N$ 表示顶点个数
#
# ### 与 CNN 的相似性
#
# 我们知道,普通 $CNN$ 的卷积是同时对空间与通道进行卷积,而 $Xception$ 提出的深度可分离卷积是对空间卷积与通道的全连接进行的分离,而且深度可分离卷积确实比普通卷积有着更少的参数,在等量参数下,深度可分离卷积有着更好的效果
#
# 深度可分离卷积将一个卷积层分成两个子层,第一步是对每个通道进行空间上的卷积,各个通道独立操作,第二步是对通道进行线性组合,也就是传统神经网络所做的事,那么对于一个图数据,是否也可以这样做呢?
#
# 假如一张图也是可以卷积的,那么我们首先对它进行卷积,之后对特征进行全连接,这便是 $GCN$ 的基本结构了
#
# 相应的,特征的维度便是传统网络的某一层结点单元数,特征维度的变换便是各层结点之间的全连接,当然这也说明了为何 $W^i.shape = (F^i, F^{i+1})$ ,当然,$W^i$ 的所有参数都用在了两层之间全连接上了,图卷积并无参数(区别于 $CNN$ 卷积核需要参数)
# ## 图卷积网络层的构建
#
# 首先考虑一种非常非常简单的传播方式
#
# $$
# H^{i+1} = g(A H^i W^i)
# $$
#
# > $g$ 为激活函数,$W^i$ 为第 $i$ 层的参数,$W^i.shape = (F^i, F^{i+1})$,另外值得注意的是,这里的乘法均为矩阵乘法,仅仅看 $shape$ 便可以知道这种变幻的合理性了, $(N, N) \cdot (N, F^i) \cdot (F^i, F^{i+1}) = (N, F^{i+1})$ ,但是究竟为什么这样会有效呢?
#
# 下面考虑一个非常简单的示例,图结构如下($N = 4$),
#
# 
#
# 首先写出邻接矩阵 $A$ ($shape=(4, 4)$)
A = np.matrix([
[0, 1, 0, 0],
[0, 0, 1, 1],
[0, 1, 0, 0],
[1, 0, 1, 0]
],dtype=np.float64
)
# 然后表示输入特征 $X$ ($shape=(4, 2)$) (这里输入特征维度为 $2$)
X = np.matrix([
[i, -i] for i in range(A.shape[0])
], dtype=np.float64)
X
# 应用图卷积
A * X
# 我们可以发现,原始特征在经过第一步图卷积之后在图上发生了传播,比如顶点 1 ,聚合了邻居 2 和 3 的特征
#
# > 如果存在从 $u$ 到 $v$ 的边,则 $v$ 是 $u$ 的邻居,也即有向图沿着箭头的反方向传播,无向图沿着边传播
#
# 但是这样就会产生两个问题
#
# - 首先,每个顶点特征在传播后自身的信息会丢失,为了避免这一问题,可以通过**在图上增加自环**来解决,具体方法就是在 $A$ 的基础上增加单位矩阵 $I$ ,得到的修正结果为 $\hat{A}$
I = np.matrix(np.eye(A.shape[0]))
I
A_hat = A + I
A_hat * X
# - 另外还有一个问题就是,各个特征在聚合的过程中传播的次数取决于顶点度的大小,度大的顶点会使得特征的在整体表征所占权重更大,这可能会引发梯度消失或梯度爆炸,一种简单的想法就是**对度进行归一化**,至于归一化的方法,可以使用 $D^{-1} A$ ,论文中使用 $D^{-\frac{1}{2}} A D^{-\frac{1}{2}}$,道理都一样,本文暂时不涉及后者
#
# > $D$ 为矩阵 $A$ 的度矩阵
#
# 首先写出获取一个矩阵的度矩阵的函数,并求得矩阵 $A$ 的度矩阵 $D$
def get_degree_matrix(A):
D = np.array(np.sum(A, axis=0))[0]
D = np.matrix(np.diag(D))
return D
D = get_degree_matrix(A)
D
# 下面求出归一化的邻接矩阵
D**-1 * A
# 很明显,邻接矩阵在度大的方向减小了链接权重(除以对应的度)
D**-1 * A * X
# 相应地,聚合时传播效果也是归一化的结果
#
# 下面,将上述两个问题结合起来,首先求得 $\hat{A}$ ,之后对 $\hat{A}$ 进行归一化,当然,$\hat{D}$ 是 $\hat{A}$ 的度矩阵
D_hat = get_degree_matrix(A_hat)
D_hat**-1 * A_hat * X
# 至于后面的参数,乘一下就好了,然后加上激活函数便完成一层图卷积网络层了
W = np.matrix([
[1, -1],
[-1, 1]
])
ReLU(D_hat**-1 * A_hat * X * W)
# ## 何谓卷积
#
# 回顾传统卷积,可以看做是每个卷积核处,不同像素点的特征聚合于卷积核中心处而已
#
# 
#
# 而图卷积,是沿着图的边,将邻居的特征聚合于自身
#
# 
#
# > 本图为无向图,若为有向图,聚合方向沿着箭头的反方向
#
# 当然,前者发生在欧式空间,后者是在拓扑结构上,所谓卷积,可以说是相邻结点的一次信息聚合,而信息的聚合,[Ref3](https://www.zhihu.com/question/54504471)一文中首先使用了温度场模型的热量传播进行比拟,之后推到图模型,由浅及深地进行了解释
#
# 对于一个连续的模型,$t$ 时刻的结果就是 $f(x, y, z, t)$ ,而一个离散的结构,每一时刻的结果都与前一时刻相关联,每一位置的结果都与周围位置相关联,在求得了前一时刻各位置的结果后,下一时刻任何一个位置都可以求得,每一个位置的结果取决于其相邻结点,具体关系可对原来的连续模型下的公式进行离散化,化微分为差分,便可得到相邻结点传播公式
#
# > 聚合针对某一结点,传播针对整个结构
# > 另外,由于本学期有一门专业课恰好学习并实践温度场模型的有限差分模拟,所以看到这篇文章倍感亲切~
# ## 图卷积的优点
#
# 图卷积是在传统网络的基础上增加了图的结构,增加的信息当然不是毫无用处,因为图的存在,使用随机初始化的参数便可完成初步的聚类,而且只需要较少的标签便可完成学习,因此 $GCN$ 也被称为是一种**半监督学习**方式
# ## 总结
#
# 整个图卷积网络层,可以分为两步,第一步是图卷积,第二步是层与层之间的连接
#
# 前者使用邻接矩阵 $A$ ,使得特征沿着图的边进行传播,得到 $A H^i$ ,考虑到自环问题和归一化问题的话,改为 $\hat{D}^{-1} \hat{A} H^i$ 或者 $D^{-\frac{1}{2}} A D^{-\frac{1}{2}} \hat{A} H^i$ 即可
#
# 后者使用链接权重 $W^i$ ,与传统网络并无不同,其 $shape$ 依然为 $(f^{in}, f^{out})$ (这里 $in$ 和 $out$ 用来表示层的输入与输出),之后激活一下就好啦
#
# 总的来说,图卷积不过是在前层特征计算完之后再整张图上传播一下(第一步),之后和传统网络并无区别,所以说, $GCN$ 也没啥难的嘛~
# > 下面是基于空手道俱乐部数据搭建的网络结构
zkc = networkx.karate_club_graph()
order = sorted(list(zkc.nodes()))
A = networkx.to_numpy_matrix(zkc, nodelist=order)
I = np.eye(zkc.number_of_nodes())
A_hat = A + I
D_hat = np.array(np.sum(A_hat, axis=0))[0]
D_hat = np.matrix(np.diag(D_hat))
W_1 = np.random.normal(
loc=0, scale=1, size=(zkc.number_of_nodes(), 4))
W_2 = np.random.normal(
loc=0, size=(W_1.shape[1], 2))
def gcn_layer(A_hat, D_hat, X, W):
return ReLU(D_hat**-1 * A_hat * X * W)
H_1 = gcn_layer(A_hat, D_hat, I, W_1)
H_2 = gcn_layer(A_hat, D_hat, H_1, W_2)
output = H_2
feature_representations = {
node: np.array(output)[node]
for node in zkc.nodes()}
# # Reference
#
# 1. [图卷积网络到底怎么做,这是一份极简的 Numpy 实现](https://mp.weixin.qq.com/s/sg9O761F0KHAmCPOfMW_kQ)
# 2. [何时能懂你的心——图卷积神经网络(GCN)](https://mp.weixin.qq.com/s/I3MsVSR0SNIKe-a9WRhGPQ)
# 3. [如何理解 Graph Convolutional Network(GCN)?](https://www.zhihu.com/question/54504471)
# 4. [GCN graph convolutional networks 详解](https://blog.csdn.net/guotong1988/article/details/82628156)
| Codes/DeepLearning/GCN/hello_gcn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Images et matrices
#
# Quelques manipulations d'images avec deux modules [Pillow](https://pillow.readthedocs.io/en/5.1.x/) et [scikit-image](http://scikit-image.org/). Le premier module implémente les entrées sorties et quelques effet spéciaux, le second est pratique quand il faut travailler numériquement avec les images.
from jyquickhelper import add_notebook_menu
add_notebook_menu()
# ## PIL : operations simples
# ### Open
from PIL import Image
img = Image.open("images1.jpg")
img
img.size
img.resize((50, 50))
# ### Combiner
new_img = Image.new('RGB', (img.size[0]*2, img.size[1]))
new_img.paste(img, (0,0))
new_img.paste(img, (img.size[0],0))
new_img
# +
def combine(*imgs, mode='RGB', vert=False):
if vert:
sizesx = [im.size[0] for im in imgs]
sizesy = [im.size[1] for im in imgs]
new_img = Image.new(mode, (max(sizesx), sum(sizesy)))
y = 0
for im in imgs:
new_img.paste(im, (0, y))
y += im.size[1]
else:
sizesx = [im.size[0] for im in imgs]
sizesy = [im.size[1] for im in imgs]
new_img = Image.new(mode, (sum(sizesx), max(sizesy)))
x = 0
for im in imgs:
new_img.paste(im, (x, 0))
x += im.size[0]
return new_img
combine(img, img)
# -
combine(img, img, vert=True)
# ## PIL to array
#
# Une image en couleur contient trois images, une pour chaque couleur primaire.
import numpy
array = numpy.array(img.getdata(), dtype=numpy.uint8).reshape(img.size[1], img.size[0], 3)
array.shape
array.dtype
# ## D'une matrice à sa transposée
array.transpose((2, 1, 0)).shape
# ## Matrice à PIL
from PIL import Image
img2 = Image.fromarray(array)
img2
# ## Séparer les couleurs
im_r, im_b, im_g = img.split()
combine(im_r, im_b, im_g, mode="L")
# ### YCbCr
img_ycbcr = img.convert('YCbCr')
img_ycbcr.size
img_y, img_cb, img_cr = img_ycbcr.split()
img_y.size
combine(img_y, img_cb, img_cr, mode="L")
| _doc/notebooks/cheat_sheet/chsh_images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
pip install keras-vggface
pip install keras_applications==1.0.4 --no-deps
pip install keras_preprocessing==1.0.2 --no-deps
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
import glob
import random as rn
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import cv2
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Conv2D, MaxPooling2D, Flatten, Dropout, BatchNormalization
from tensorflow.keras.callbacks import EarlyStopping, ReduceLROnPlateau
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.optimizers import RMSprop, Adam
from tensorflow.keras.utils import to_categorical
from sklearn.metrics import classification_report, confusion_matrix
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import os
import cv2
import matplotlib.pyplot as plt
# %matplotlib inline
import operator
import tensorflow as tf
import random
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Model
import numpy as np
import os
import cv2
import matplotlib.pyplot as plt
from keras.models import load_model
import numpy as np
import pandas as pd
import os
import glob
import cv2
import random
import matplotlib.pyplot as plt
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential
from keras.optimizers import RMSprop
from keras.layers import Conv2D, MaxPooling2D
from keras.layers import Activation, Dropout, Flatten, Dense
from keras import regularizers
from keras.callbacks import CSVLogger
#from livelossplot import PlotLossesKeras
import os
import numpy as np
#from imgaug import augmenters as iaa
#import cv2
from keras.layers.normalization import BatchNormalization
#import seaborn as sns
import pandas as pd
from keras import initializers
from keras import optimizers
import keras.backend as K
import tensorflow as tf
from keras.models import Model
import cv2
import numpy as np
from tensorflow.keras import layers
from tensorflow.keras.applications import DenseNet121
from tensorflow.keras.callbacks import Callback, ModelCheckpoint
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.models import Sequential
from tensorflow.keras.optimizers import Adam
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn import metrics
import tensorflow as tf
from tqdm import tqdm
# -
def build_model(pretrained):
model = Sequential([
pretrained,
layers.GlobalAveragePooling2D(),
layers.Dense(1, activation='sigmoid')
])
model.compile(
loss='binary_crossentropy',
optimizer=Adam(),
metrics=['accuracy']
)
return model
# +
base_path = '/kaggle/input/140k-real-and-fake-faces/real_vs_fake/real-vs-fake/'
image_gen = ImageDataGenerator(rescale=1./255.)
train_flow = image_gen.flow_from_directory(
base_path + 'train/',
target_size=(224, 224),
batch_size=64,
class_mode='binary'
)
image_gen1 = ImageDataGenerator(rescale=1./255.)
valid_flow = image_gen1.flow_from_directory(
base_path + 'valid/',
target_size=(224, 224),
batch_size=64,
class_mode='binary'
)
# -
densenet = DenseNet121(
weights=None,
include_top=False,
input_shape=(224,224,3)
)
model = build_model(densenet)
model.summary()
# +
train_steps = 100000//64
valid_steps = 20000//64
history = model.fit_generator(
train_flow,
epochs = 10,
steps_per_epoch =train_steps,
validation_data =valid_flow,
validation_steps = valid_steps
)
# -
train_flow.class_indices
valid_flow.class_indices
test_flow.class_indices
def plot_loss(epochs, loss, val_loss):
plt.plot(epochs, loss, 'bo', label='Training Loss')
plt.plot(epochs, val_loss, 'orange', label = 'Validation Loss')
plt.title('Training and Validation Loss')
plt.legend()
plt.show()
"""
Plot the training and validation accuracy
epochs - list of epoch numbers
acc - training accuracy for each epoch
val_acc - validation accuracy for each epoch
"""
def plot_accuracy(epochs, acc, val_acc):
plt.plot(epochs, acc, 'bo', label='Training accuracy')
plt.plot(epochs, val_acc, 'orange', label = 'Validation accuracy')
plt.title('Training and Validation Accuracy')
plt.legend()
plt.show()
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
plot_loss(range(1, len(loss) + 1), loss, val_loss)
plot_accuracy(range(1, len(loss) + 1), acc, val_acc)
test_flow = image_gen1.flow_from_directory(
base_path + 'test/',
target_size=(224, 224),
batch_size=1,
shuffle = False,
class_mode='binary'
)
y_pred = model.predict(test_flow)
y_test = test_flow.classes
# +
print("ROC AUC Score:", metrics.roc_auc_score(y_test, y_pred))
print("AP Score:", metrics.average_precision_score(y_test, y_pred))
print()
print(metrics.classification_report(y_test, y_pred > 0.5))
| deepfake.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.1 64-bit
# language: python
# name: python38164bitdbd76ed984a5488496eb976b9e8b3b8e
# ---
# +
# dataset: tf_idf_part (333, 688)
# algorithms need to test: SVC,DecisionTreeClassifier,RidgeClassifierCV, SGDClassifier, MLPClassifier
# generator : input_shape:688(features dims)+20(noise dims) hidden layer nodes:256 output layer nodes:128
# subsititude detector: 128 - 256 - 1
# +
from tensorflow.keras.layers import Input
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Activation
from tensorflow.keras.layers import Maximum, Concatenate
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import Adam
from sklearn.svm import SVC
from sklearn.tree import DecisionTreeClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import RidgeClassifierCV, SGDClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import numpy as np
from pathlib import Path
import pandas as pd
global seed
# define the random_state seed
seed = 43
# -
# load dataset
# get the data from partial TF-IDF features
tf_idf_part = pd.read_pickle('../dataset/training_data/features_ran_part.pkl')
# bypass the label column
tf_idf_part.cov()
# build the ACGAN Model for Ransomware
class ACGAN_RAN():
def __init__(self, blackbox='RF', same_train_data=1, data_path='../dataset/training_data/features_ran_part.pkl'):
self.apifeature_dims = 128
self.z_dims = 20
self.hide_layers = 256
self.generator_layers = [self.apifeature_dims+self.z_dims, self.hide_layers, self.apifeature_dims]
self.substitute_detector_layers = [self.apifeature_dims, self.hide_layers, 1]
self.blackbox = blackbox # RF LR DT SVM MLP VOTE
optimizer = Adam(lr=0.001, 0.5)
self.data = pd.read_pickle(data_path)
# Build and Train blackbox_detector
self.blackbox_detector = self.build_blackbox_detector()
# Build and compile the substitute_detector
self.substitute_detector = self.build_substitute_detector()
self.substitute_detector.compile(loss='binary_crossentropy', optimizer=optimizer, metrics=['accuracy'])
# Build the generator
self.generator = self.build_generator()
# The generator takes malware and noise as input and generates adversarial malware examples
example = Input(shape=(self.apifeature_dims,))
noise = Input(shape=(self.z_dims,))
input = [example, noise]
malware_examples = self.generator(input)
# The discriminator takes generated images as input and determines validity
validity = self.substitute_detector(malware_examples)
# The combined model (stacked generator and substitute_detector)
# Trains the generator to fool the discriminator
self.combined = Model(input, validity)
self.combined.compile(loss='binary_crossentropy', optimizer=optimizer)
# For the combined model we will only train the generator
self.substitute_detector.trainable = False
def build_blackbox_detector(self):
if self.blackbox is 'SVM':
blackbox_detector = svm.SVC()
elif self.blackbox is 'DT':
blackbox_detector = RandomForestClassifier(n_estimators=100, max_depth=3, random_state=seed)
elif self.blackbox is 'SGD':
blackbox_detector = SGDClassifier(random_state=seed)
elif self.blackbox is 'RC':
blackbox_detector = RidgeClassifierCV()
elif self.blackbox is 'MLP':
blackbox_detector = MLPClassifier(hidden_layer_sizes=(50,), max_iter=50, alpha=1e-4,
solver='sgd', verbose=0, tol=1e-4, random_state=seed,
learning_rate_init=.1)
return blackbox_detector
def build_generator(self):
example = Input(shape=(self.apifeature_dims,))
noise = Input(shape=(self.z_dims,))
x = Concatenate(axis=1)([example, noise])
for dim in self.generator_layers[1:]:
x = Dense(dim)(x)
x = Activation(activation='sigmoid')(x)
x = Maximum()([example, x])
generator = Model([example, noise], x, name='generator')
generator.summary()
return generator
def build_substitute_detector(self):
input = Input(shape=(self.substitute_detector_layers[0],))
x = input
for dim in self.substitute_detector_layers[1:]:
x = Dense(dim)(x)
x = Activation(activation='sigmoid')(x)
substitute_detector = Model(input, x, name='substitute_detector')
substitute_detector.summary()
return substitute_detector
def load_data(self):
x_ran, y_ran, x_ben, y_ben = self.data[self.data['label']==1].iloc[:,:-1],
self.data[self.data['label']==1].iloc[:,-1],
self.data[self.data['label']==0].iloc[:,:-1],
self.data[self.data['label']==0].iloc[:,-1]
return (x_ran, y_ran), (x_ben, y_ben)
def train(self, epochs, batch_size=32, is_first=1):
# Load and Split the dataset
(x_ran, y_ran), (x_ben, y_ben) = self.load_data()
xtrain_mal, xtest_mal, ytrain_mal, ytest_mal = train_test_split(x_ran, y_ran, test_size=0.20)
xtrain_ben, xtest_ben, ytrain_ben, ytest_ben = train_test_split(xben, yben, test_size=0.20)
if self.same_train_data:
bl_xtrain_mal, bl_ytrain_mal, bl_xtrain_ben, bl_ytrain_ben = xtrain_mal, ytrain_mal, xtrain_ben, ytrain_ben
else:
xtrain_mal, bl_xtrain_mal, ytrain_mal, bl_ytrain_mal = train_test_split(xtrain_mal, ytrain_mal, test_size=0.50)
xtrain_ben, bl_xtrain_ben, ytrain_ben, bl_ytrain_ben = train_test_split(xtrain_ben, ytrain_ben, test_size=0.50)
# if is_first is Ture, Train the blackbox_detctor
if is_first:
self.blackbox_detector.fit(np.concatenate([xmal, xben]),
np.concatenate([ymal, yben]))
ytrain_ben_blackbox = self.blackbox_detector.predict(bl_xtrain_ben)
Original_Train_TPR = self.blackbox_detector.score(bl_xtrain_mal, bl_ytrain_mal)
Original_Test_TPR = self.blackbox_detector.score(xtest_mal, ytest_mal)
Train_TPR, Test_TPR = [Original_Train_TPR], [Original_Test_TPR]
best_TPR = 1.0
for epoch in range(epochs):
for step in range(xtrain_mal.shape[0] // batch_size):
# ---------------------
# Train substitute_detector
# ---------------------
# Select a random batch of malware examples
idx = np.random.randint(0, xtrain_mal.shape[0], batch_size)
xmal_batch = xtrain_mal[idx]
noise = np.random.uniform(0, 1, (batch_size, self.z_dims))
idx = np.random.randint(0, xmal_batch.shape[0], batch_size)
xben_batch = xtrain_ben[idx]
yben_batch = ytrain_ben_blackbox[idx]
# Generate a batch of new malware examples
gen_examples = self.generator.predict([xmal_batch, noise])
ymal_batch = self.blackbox_detector.predict(np.ones(gen_examples.shape)*(gen_examples > 0.5))
# Train the substitute_detector
d_loss_real = self.substitute_detector.train_on_batch(gen_examples, ymal_batch)
d_loss_fake = self.substitute_detector.train_on_batch(xben_batch, yben_batch)
d_loss = 0.5 * np.add(d_loss_real, d_loss_fake)
# ---------------------
# Train Generator
# ---------------------
idx = np.random.randint(0, xtrain_mal.shape[0], batch_size)
xmal_batch = xtrain_mal[idx]
noise = np.random.uniform(0, 1, (batch_size, self.z_dims))
# Train the generator
g_loss = self.combined.train_on_batch([xmal_batch, noise], np.zeros((batch_size, 1)))
# Compute Train TPR
noise = np.random.uniform(0, 1, (xtrain_mal.shape[0], self.z_dims))
gen_examples = self.generator.predict([xtrain_mal, noise])
TPR = self.blackbox_detector.score(np.ones(gen_examples.shape) * (gen_examples > 0.5), ytrain_mal)
Train_TPR.append(TPR)
# Compute Test TPR
noise = np.random.uniform(0, 1, (xtest_mal.shape[0], self.z_dims))
gen_examples = self.generator.predict([xtest_mal, noise])
TPR = self.blackbox_detector.score(np.ones(gen_examples.shape) * (gen_examples > 0.5), ytest_mal)
Test_TPR.append(TPR)
# Save best model
if TPR < best_TPR:
self.combined.save_weights('saves/malgan.h5')
best_TPR = TPR
# Plot the progress
if is_first:
print("%d [D loss: %f, acc.: %.2f%%] [G loss: %f]" % (epoch, d_loss[0], 100*d_loss[1], g_loss))
flag = ['DiffTrainData', 'SameTrainData']
print('\n\n---{0} {1}'.format(self.blackbox, flag[self.same_train_data]))
print('\nOriginal_Train_TPR: {0}, Adver_Train_TPR: {1}'.format(Original_Train_TPR, Train_TPR[-1]))
print('\nOriginal_Test_TPR: {0}, Adver_Test_TPR: {1}'.format(Original_Test_TPR, Test_TPR[-1]))
# Plot TPR
plt.figure()
plt.plot(range(len(Train_TPR)), Train_TPR, c='r', label='Training Set', linewidth=2)
plt.plot(range(len(Test_TPR)), Test_TPR, c='g', linestyle='--', label='Validation Set', linewidth=2)
plt.xlabel('Epoch')
plt.ylabel('TPR')
plt.legend()
plt.savefig('saves/Epoch_TPR({0}, {1}).png'.format(self.blackbox, flag[self.same_train_data]))
plt.show()
def retrain_blackbox_detector(self):
(xmal, ymal), (xben, yben) = self.load_data()
xtrain_mal, xtest_mal, ytrain_mal, ytest_mal = train_test_split(xmal, ymal, test_size=0.20)
xtrain_ben, xtest_ben, ytrain_ben, ytest_ben = train_test_split(xben, yben, test_size=0.20)
# Generate Train Adversarial Examples
noise = np.random.uniform(0, 1, (xtrain_mal.shape[0], self.z_dims))
gen_examples = self.generator.predict([xtrain_mal, noise])
gen_examples = np.ones(gen_examples.shape) * (gen_examples > 0.5)
self.blackbox_detector.fit(np.concatenate([xtrain_mal, xtrain_ben, gen_examples]),
np.concatenate([ytrain_mal, ytrain_ben, ytrain_mal]))
# Compute Train TPR
train_TPR = self.blackbox_detector.score(gen_examples, ytrain_mal)
# Compute Test TPR
noise = np.random.uniform(0, 1, (xtest_mal.shape[0], self.z_dims))
gen_examples = self.generator.predict([xtest_mal, noise])
gen_examples = np.ones(gen_examples.shape) * (gen_examples > 0.5)
test_TPR = self.blackbox_detector.score(gen_examples, ytest_mal)
print('\n---TPR after the black-box detector is retrained(Before Retraining MalGAN).')
print('\nTrain_TPR: {0}, Test_TPR: {1}'.format(train_TPR, test_TPR))
if __name__ == '__main__':
malgan = ACGAN_RAN(blackbox='MLP')
malgan.train(epochs=50, batch_size=32)
malgan.retrain_blackbox_detector()
malgan.train(epochs=50, batch_size=32, is_first=False)
| Classifier/ACGAN_RAN_Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tf15
# language: python
# name: tf15
# ---
# #### Import neccessary libraries and set paths
# +
import tensorflow as tf
config_tf = tf.ConfigProto()
config_tf.gpu_options.allow_growth=True
sess = tf.Session(config=config_tf)
import json
from keras.models import model_from_json
import sys
from pathlib import Path
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from keras.utils import multi_gpu_model
from sklearn.ensemble import RandomForestRegressor
from importlib import reload
from pygifsicle import optimize
import imageio
import os
import matplotlib.animation as animation
# -
params = {
"legend.fontsize": "x-large",
"axes.labelsize": "x-large",
"axes.titlesize": "x-large",
"xtick.labelsize": "x-large",
"ytick.labelsize": "x-large",
"figure.facecolor": "w",
"xtick.top": True,
"ytick.right": True,
"xtick.direction": "in",
"ytick.direction": "in",
"font.family": "serif",
"mathtext.fontset": "dejavuserif"
}
plt.rcParams.update(params)
# +
# Path where your software library is saved
# Clone the latest version of morphCaps branch from github
path_photoz = '/home/bid13/code/photozCapsNet'
sys.path.insert(1, path_photoz)
path_photoz = Path(path_photoz)
# -
# #### Import custom modules
from encapzulate.data_loader.data_loader import load_data
from encapzulate.utils.fileio import load_model, load_config
from encapzulate.utils import metrics
from encapzulate.utils.utils import import_model
from encapzulate.utils.metrics import Metrics, probs_to_redshifts, bins_to_redshifts
reload(metrics)
# #### Specify the results to be explored
# Parameters for the exploration
run_name = "paper1_regression_80perc_0"
checkpoint_eval = 100
#Create and set different paths
# path_output = "/data/bid13/photoZ/results"
path_output = "/home/bid13/code/photozCapsNet/results"
path_output = Path(path_output)
path_results = path_output / run_name.split("_")[0] / run_name / "results"
path_config = path_results / "config.yml"
# #### Load Config, Model and Data
config = load_config(path_config)
scale= config['image_scale']
log = pd.read_csv(path_results/ "logs" /"log.csv")
max_acc = log[log.val_decoder_model_loss==log.val_decoder_model_loss.min()]
max_acc
#with tf.device('/cpu:0'):
model = load_model( path_results / "eval_model.json", path_results/ "weights" / f"weights-{checkpoint_eval:02d}.h5")
# model = multi_gpu_model(model,gpus=2)
model.summary()
(
(x_train, y_train, vals_train, z_spec_train, cat_train),
(x_dev, y_dev, vals_dev, z_spec_dev, cat_dev),
(x_test, y_test, vals_test, z_spec_test, cat_test),
) = load_data(load_cat=True, **config)
# #### Run Predictions
y_caps_dev, y_caps_all_dev, y_prob_dev, x_recon_dev, z_phot_dev = model.predict(x_dev,batch_size=1024)
del x_train
del x_test
# del x_dev
# del x_recon_test
# del x_recon_dev
# #### Plot images
# +
# https://github.com/legacysurvey/imagine/blob/acac773c6a43c7e6d6ea0c128d5e963ad8295229/map/views.py#L3881
def sdss_rgb(imgs, bands, scales=None, m=0.02, Q=20, alpha=1, p=0.7):
import numpy as np
rgbscales = {
"u": (2, 1.5), # 1.0,
"g": (2, 2.8),
"r": (1, 1.4),
"i": (0, 1.1),
"z": (0, 0.4), # 0.3
}
if scales is not None:
rgbscales.update(scales)
I = 0
for img, band in zip(imgs, bands):
plane, scale = rgbscales[band]
img = np.maximum(0, img * scale + m)
I = I + img
I /= len(bands)
# Q = 20
# alpha = 1
# p =0.7
# # fI = np.arcsinh(Q * I) / np.sqrt(Q)
fI = np.arcsinh(alpha * Q * I) / (Q**p)
I += (I == 0.0) * 1e-6
H, W = I.shape
rgb = np.zeros((H, W, 3), np.float32)
for img, band in zip(imgs, bands):
plane, scale = rgbscales[band]
rgb[:, :, plane] = (img * scale + m) * fI / I
# R = fI * r / I
# G = fI * g / I
# B = fI * b / I
# # maxrgb = reduce(np.maximum, [R,G,B])
# # J = (maxrgb > 1.)
# # R[J] = R[J]/maxrgb[J]
# # G[J] = G[J]/maxrgb[J]
# # B[J] = B[J]/maxrgb[J]
# rgb = np.dstack((R,G,B))
rgb = np.clip(rgb, 0, 1)
return rgb
# + tags=[]
# from astropy.visualization import make_lupton_rgb
# # The function below has not yet been finalized. Can be fine tuned before incorporating into the main code
# def plot_image(image, band, scaling="linear", ax=None, show=False, input_bands=None):
# """Plot different colored images of galaxies
# Args:
# image (array): five colored sdss image
# band (str): u, g, r, i or z band or gri composite image (also works with 0,1,2,3,4,5 codes)
# scaling: linear or asinh for the single band images. gri images are always asinh scaled
# ax (object): Matplotlib object to plot on
# show (bool): Whether or not to show the plot
# input_bands: use gri if input image has only three colors
# Returns:
# Matplotlib axis object
# """
# bands = {"u":0, "g":1, "r":2, "i":3, "z":4, "gri":5}
# assert (band in bands) or (band in bands.values()) , "Choose from u, g, r, i, z bands or gri composite image"
# assert (scaling in ["linear", "asinh"]), "scaling should be either linear or asinh for the single band images"
# if ax == None:
# fig, ax = plt.subplots()
# if (band == "gri") or (band==5):
# if input_bands == "gri":
# stretch = 1
# Q=8
# scale =1.3
# rgb = make_lupton_rgb(scale*1*image[:,:,2], scale*1.8*image[:,:,1], scale*2.3*image[:,:,0], stretch=stretch, Q=Q)
# else:
# stretch = 1.5
# Q=5
# scale = 1
# rgb = make_lupton_rgb(scale*1*image[:,:,3], scale*1.5*image[:,:,2], scale*2.5*image[:,:,1], stretch=stretch, Q=Q, minimum=-0.02)
# ax.imshow(rgb, aspect="equal", origin="lower")
# ax.axes.get_xaxis().set_ticks([])
# ax.axes.get_yaxis().set_ticks([])
# else:
# if band in bands:
# band = bands[band]
# if scaling == "linear":
# ax.imshow(image[:,:,band], aspect="equal", origin="lower", cmap="Greys_r")
# if scaling == "asinh":
# img = make_lupton_rgb(image[:,:,band], image[:,:,band], image[:,:,band], stretch=stretch, Q=Q)
# ax.imshow(img[:,:,0], aspect="equal", origin="lower", cmap="Greys_r")
# ax.axis("off")
# if show:
# plt.show()
# return ax
# -
def plot_image(image, band="gri", ax=None, m=0., Q=20, alpha=0.8, p=0.7):
rgb = sdss_rgb(np.moveaxis(image, -1,0)[1:4], [ "g", "r", "i"],m=m, Q=Q, alpha=alpha, p=p)
if ax == None:
fig, ax = plt.subplots()
ax.imshow(rgb, aspect="equal", origin="lower")
ax.axes.get_xaxis().set_ticks([])
ax.axes.get_yaxis().set_ticks([])
return ax
# +
# index = 0
# from scipy import ndimage
# fig, ax = plt.subplots(1,2)
# ax = ax.ravel()
# rgb_obs = sdss_rgb(np.moveaxis(scale*x_dev[index], -1,0)[1:4], [ "g", "r", "i"],m=-0.02)
# # rgb_obs = ndimage.median_filter(rgb_obs, 2)
# ax[0].imshow(rgb_obs, aspect="equal", origin="lower")
# ax[0].set_xlabel("Observed", fontsize=20)
# rgb_recon = sdss_rgb(np.moveaxis(scale*x_recon_dev[index], -1,0)[1:4], [ "g", "r", "i"],m =-0.02)
# ax[1].imshow(rgb_recon, aspect="equal", origin="lower")
# ax[1].set_xlabel("Reconstructed", fontsize=20)
# +
index = 0
fig, ax = plt.subplots(1,2)
ax = ax.ravel()
plot_image(scale*x_dev[index], "gri", ax=ax[0])
ax[0].set_xlabel("Observed", fontsize=20)
plot_image(scale*x_recon_dev[index], "gri", ax=ax[1])
ax[1].set_xlabel("Reconstructed", fontsize=20)
# -
for i in [1,2,3]:
plt.hist(np.ravel(scale*x_dev[index][i]), histtype="step", label="observed")
plt.hist(np.ravel(scale*x_recon_dev[index][i]), histtype="step", label="recon")
plt.legend()
plt.show()
mean_o = np.mean(scale*x_dev[index], axis=(0,1))
std_o = np.std(scale*x_dev[index], axis=(0,1))
mean_r = np.mean(scale*x_recon_dev[index], axis=(0,1))
std_r = np.std(scale*x_recon_dev[index], axis=(0,1))
# +
index = 0
fig, ax = plt.subplots(1,2)
ax = ax.ravel()
plot_image(scale*x_dev[index], "gri", ax=ax[0])
ax[0].set_xlabel("Observed", fontsize=20)
abcd = (((scale*x_recon_dev[index]-mean_r)/std_r))*std_o + mean_r
plot_image(abcd, "gri", ax=ax[1])
ax[1].set_xlabel("Reconstructed", fontsize=20)
# -
fig, axs = plt.subplots(3,2, figsize=(7.8,12))
axs =axs.flatten()
selected_spirals = [0, 14, 13]
for i in range(3):
plot_image(scale*x_dev[selected_spirals[i]], "gri", ax = axs[2*(i)])
plot_image(scale*x_recon_dev[selected_spirals[i]], "gri", ax =axs[2*(i)+1])
t = fig.suptitle("Spirals", fontsize=40, y=1.0)
axs[-2].set_xlabel("Observed", fontsize=30)
axs[-1].set_xlabel("Reconstructed", fontsize=30)
plt.tight_layout()
fig.savefig("./figs/disks.pdf",bbox_inches='tight',bbox_extra_artists=[t],dpi=300)
fig, axs = plt.subplots(3,2, figsize=(7.8,12))
axs =axs.flatten()
selected_spirals = [20, 57, 80]
for i in range(3):
plot_image(scale*x_dev[selected_spirals[i]], "gri", ax = axs[2*(i)])
plot_image(scale*x_recon_dev[selected_spirals[i]], "gri", ax =axs[2*(i)+1])
t = fig.suptitle("Ellipticals", fontsize=40, y=1.)
axs[-2].set_xlabel("Observed", fontsize=30)
axs[-1].set_xlabel("Reconstructed", fontsize=30)
plt.tight_layout()
fig.savefig("./figs/spheroids.pdf",bbox_inches='tight',bbox_extra_artists=[t],dpi=300)
#with tf.device('/cpu:0'):
model = load_model( path_results / "train_model.json", path_results/ "weights" / f"weights-{checkpoint_eval:02d}.h5")
model = multi_gpu_model(model,gpus=2)
model.summary()
config["input_shape"] = config["image_shape"]
CapsNet = import_model(model_name=config["model_name"])
train_model, eval_model,manipulate_model,decoder_model,redshift_model, = CapsNet(**config)
manipulate_model.load_weights(
path_results / "weights" / f"weights-{checkpoint_eval:02d}.h5", by_name=True
)
# # Tinker All disk
# +
config["input_shape"] = config["image_shape"]
CapsNet = import_model(model_name=config["model_name"])
train_model, eval_model,manipulate_model,decoder_model,redshift_model, = CapsNet(**config)
manipulate_model.load_weights(
path_results / "weights" / f"weights-{checkpoint_eval:02d}.h5", by_name=True
)
img_indx =0 # 20 and 0
sigma_arr = np.std(y_caps_dev, axis=0)
caps_gal = y_caps_dev[img_indx].copy()
change_grid = [-3,-2,-1,0,1,2,3]
num_caps=16
fig, axs = plt.subplots(num_caps,len(change_grid), figsize=(1.4*8.3,2.1*11.7))
for caps_index in range(num_caps):
for j in range(len(change_grid)):
tinkered_dim = caps_gal[caps_index] + change_grid[j] * sigma_arr[caps_index]
tinkered_caps = caps_gal.copy()
tinkered_caps[caps_index] = tinkered_dim
tinkered_recon = decoder_model.predict(np.expand_dims(tinkered_caps, axis=0))[0]
plot_image(scale*tinkered_recon, "gri", ax =axs[caps_index][j] )
cols = ['{}$\sigma$'.format(col) for col in change_grid]
rows = ['Dim: {}'.format(row) for row in np.arange(1,num_caps+1).astype(str) ]
for ax, col in zip(axs[0], cols):
ax.set_title(col, size=25)
for ax, row in zip(axs[:,0], rows):
ax.set_ylabel(row, size=23)
# fig.suptitle("Redshift: "+ str(z_spec_dev[img_indx]), y =1.01, size=20)
plt.tight_layout()
fig.savefig("./figs/tinker_disk_appendix.pdf",bbox_inches='tight',dpi=300)
# -
# # Tinker all spheroid
# +
config["input_shape"] = config["image_shape"]
CapsNet = import_model(model_name=config["model_name"])
train_model, eval_model,manipulate_model,decoder_model,redshift_model, = CapsNet(**config)
manipulate_model.load_weights(
path_results / "weights" / f"weights-{checkpoint_eval:02d}.h5", by_name=True
)
img_indx =20 # 20 and 0
sigma_arr = np.std(y_caps_dev, axis=0)
caps_gal = y_caps_dev[img_indx].copy()
change_grid = [-3,-2,-1,0,1,2,3]
num_caps=16
fig, axs = plt.subplots(num_caps,len(change_grid), figsize=(1.4*8.3,2.1*11.7))
for caps_index in range(num_caps):
for j in range(len(change_grid)):
tinkered_dim = caps_gal[caps_index] + change_grid[j] * sigma_arr[caps_index]
tinkered_caps = caps_gal.copy()
tinkered_caps[caps_index] = tinkered_dim
tinkered_recon = decoder_model.predict(np.expand_dims(tinkered_caps, axis=0))[0]
plot_image(scale*tinkered_recon, "gri", ax =axs[caps_index][j] )
cols = ['{}$\sigma$'.format(col) for col in change_grid]
rows = ['Dim: {}'.format(row) for row in np.arange(1,num_caps+1).astype(str) ]
for ax, col in zip(axs[0], cols):
ax.set_title(col, size=25)
for ax, row in zip(axs[:,0], rows):
ax.set_ylabel(row, size=23)
# fig.suptitle("Redshift: "+ str(z_spec_dev[img_indx]), y =1.01, size=20)
plt.tight_layout()
fig.savefig("./figs/tinker_spheroid_appendix.pdf",bbox_inches='tight',dpi=300)
# -
# # Tinker some
# +
config["input_shape"] = config["image_shape"]
CapsNet = import_model(model_name=config["model_name"])
train_model, eval_model,manipulate_model,decoder_model,redshift_model, = CapsNet(**config)
manipulate_model.load_weights(
path_results / "weights" / f"weights-{checkpoint_eval:02d}.h5", by_name=True
)
img_indx = 20
sigma_arr = np.std(y_caps_dev, axis=0)
caps_gal = y_caps_dev[img_indx].copy()
change_grid = [-3, -2, -1, 0, 1, 2, 3]
num_caps=[1,8,12,13]
fig, axs = plt.subplots(len(num_caps),len(change_grid), figsize=(15,9))
for caps_count, caps_dim in enumerate(num_caps):
for j in range(len(change_grid)):
tinkered_dim = caps_gal[caps_dim] + change_grid[j] * sigma_arr[caps_dim]
tinkered_caps = caps_gal.copy()
tinkered_caps[caps_dim] = tinkered_dim
tinkered_recon = decoder_model.predict(np.expand_dims(tinkered_caps, axis=0))[0]
plot_image(scale*tinkered_recon, "gri", ax =axs[caps_count][j] )
col_names = [ r"$-3\sigma$", r"$-2\sigma$", r"$-1\sigma$", r"$0\sigma$", r"$1\sigma$", r"$2\sigma$", r"$3\sigma$"]
row_names = ["Size\n(Dim: 2)","Orientation\n(Dim: 9)", "Bulge\n(Dim: 13)", "Surface\nBrightness\n(Dim: 14)" ]
for ax, col in zip(axs[0], col_names):
ax.set_title(col, fontsize=30)
for ax, row in zip(axs[:,0], row_names):
ax.set_ylabel(row, fontsize=25)
# t=fig.suptitle("Spirals", size=30, y=1.01)
plt.tight_layout()
fig.savefig("./figs/tinker_spheroid.pdf",bbox_inches="tight")#,bbox_extra_artists=[t])
# +
config["input_shape"] = config["image_shape"]
CapsNet = import_model(model_name=config["model_name"])
train_model, eval_model,manipulate_model,decoder_model,redshift_model, = CapsNet(**config)
manipulate_model.load_weights(
path_results / "weights" / f"weights-{checkpoint_eval:02d}.h5", by_name=True
)
img_indx =0
sigma_arr = np.std(y_caps_dev, axis=0)
caps_gal = y_caps_dev[img_indx].copy()
change_grid = [-3, -2, -1, 0, 1, 2, 3]
num_caps=[1,8,12,13]
fig, axs = plt.subplots(len(num_caps),len(change_grid), figsize=(15,9))
for caps_count, caps_dim in enumerate(num_caps):
for j in range(len(change_grid)):
tinkered_dim = caps_gal[caps_dim] + change_grid[j] * sigma_arr[caps_dim]
tinkered_caps = caps_gal.copy()
tinkered_caps[caps_dim] = tinkered_dim
tinkered_recon = decoder_model.predict(np.expand_dims(tinkered_caps, axis=0))[0]
plot_image(scale*tinkered_recon, "gri", ax =axs[caps_count][j] )
col_names = [ r"$-3\sigma$", r"$-2\sigma$", r"$-1\sigma$", r"$0\sigma$", r"$1\sigma$", r"$2\sigma$", r"$3\sigma$"]
row_names = ["Size\n(Dim: 2)","Orientation\n(Dim: 9)", "Bulge\n(Dim: 13)", "Surface\nBrightness\n(Dim: 14)" ]
for ax, col in zip(axs[0], col_names):
ax.set_title(col, fontsize=30)
for ax, row in zip(axs[:,0], row_names):
ax.set_ylabel(row, fontsize=25)
# t=fig.suptitle("Spirals", size=30, y=1.01)
plt.tight_layout()
fig.savefig("./figs/tinker_disk.pdf",bbox_inches="tight")#,bbox_extra_artists=[t])
# -
# # GIF for presentation
# +
img_indx =[0, 20]
config["input_shape"] = config["image_shape"]
CapsNet = import_model(model_name=config["model_name"])
train_model, eval_model,manipulate_model,decoder_model,redshift_model, = CapsNet(**config)
manipulate_model.load_weights(
path_results / "weights" / f"weights-{checkpoint_eval:02d}.h5", by_name=True
)
# -
# # MAke GIF
# + tags=[]
sigma_arr = np.std(y_caps_dev, axis=0)
step =0.2
pause = 10
change_grid = np.concatenate([np.arange(0,3+step,step),
3*np.ones(pause),
np.arange(3,-3-step,-1*step),
-3*np.ones(pause),
np.arange(-3-step,0,step),
0*np.ones(pause),])
num_caps=[1,8,12,13]
filenames = []
for frame, tinker in enumerate(change_grid):
fig, axs = plt.subplots(2,6, figsize=(18,15*1080/1920))
for img_count, i in enumerate(img_indx):
plot_image(scale*x_dev[i], "gri", ax =axs[img_count][0])
plot_image(scale*x_recon_dev[i], "gri", ax =axs[img_count][1])
caps_gal = y_caps_dev[i].copy()
for caps_count, caps_dim in enumerate(num_caps):
tinkered_dim = caps_gal[caps_dim] + tinker * sigma_arr[caps_dim]
tinkered_caps = caps_gal.copy()
tinkered_caps[caps_dim] = tinkered_dim
tinkered_recon = decoder_model.predict(np.expand_dims(tinkered_caps, axis=0))[0]
plot_image(scale*tinkered_recon, "gri", ax =axs[img_count][caps_count+2] )
row_names = ["Disk", "Spheroid"]
col_names = ["Observed","Reconstructed", f"Size\n ({round(tinker,3)}$\sigma$)",
f"Orientation\n ({round(tinker,3)}$\sigma$)",
f"Central Bulge\n ({round(tinker,3)}$\sigma$)",
f"Surface\nBrightness\n ({round(tinker,3)}$\sigma$)",]
for ax, col in zip(axs[0], col_names):
ax.set_title(col, fontsize=30, y = 1.1)
for ax, row in zip(axs[:,0], row_names):
ax.set_ylabel(row, fontsize=40)
# save frame
filename = f'{frame}.png'
filenames.append(filename)
plt.tight_layout()
plt.savefig(filename, bbox_inches="tight", dpi=100)
plt.close()
# build gif
with imageio.get_writer('./figs/tinker_gif.gif', mode='I', fps=5) as writer:
for filename in filenames:
image = imageio.imread(filename)
writer.append_data(image)
# frames = []
# for filename in filenames:
# image = imageio.imread(filename)
# frames.append(image)
# imageio.mimsave('./figs/tinker_gif.gif', frames)
# optimize('./figs/tinker_gif.gif')
# Remove files
for filename in set(filenames):
os.remove(filename)
# -
from IPython.display import Image
Image(filename="./figs/tinker_gif.gif")
# # Make video
# +
sigma_arr = np.std(y_caps_dev, axis=0)
step =0.2
pause = 10
change_grid = np.concatenate([np.arange(0,3+step,step),
3*np.ones(pause),
np.arange(3,-3-step,-1*step),
-3*np.ones(pause),
np.arange(-3-step,0,step),
0*np.ones(pause),])
num_caps=[1,8,12,13]
imgs = []
for frame, tinker in enumerate(change_grid):
fig, axs = plt.subplots(2,6, figsize=(18,15*1080/1920))
for img_count, i in enumerate(img_indx):
plot_image(scale*x_dev[i], "gri", ax =axs[img_count][0])
plot_image(scale*x_recon_dev[i], "gri", ax =axs[img_count][1])
caps_gal = y_caps_dev[i].copy()
for caps_count, caps_dim in enumerate(num_caps):
tinkered_dim = caps_gal[caps_dim] + tinker * sigma_arr[caps_dim]
tinkered_caps = caps_gal.copy()
tinkered_caps[caps_dim] = tinkered_dim
tinkered_recon = decoder_model.predict(np.expand_dims(tinkered_caps, axis=0))[0]
plot_image(scale*tinkered_recon, "gri", ax =axs[img_count][caps_count+2] )
row_names = ["Disk", "Spheroid"]
col_names = ["Observed","Reconstructed", f"Size\n ({round(tinker,3)}$\sigma$)",
f"Orientation\n ({round(tinker,3)}$\sigma$)",
f"Central Bulge\n ({round(tinker,3)}$\sigma$)",
f"Surface\nBrightness\n ({round(tinker,3)}$\sigma$)",]
for ax, col in zip(axs[0], col_names):
ax.set_title(col, fontsize=30, y = 1.1)
for ax, row in zip(axs[:,0], row_names):
ax.set_ylabel(row, fontsize=40)
# save frame
filename = f'{frame}.png'
filenames.append(filename)
plt.tight_layout()
plt.savefig(filename, bbox_inches="tight", dpi=100)
plt.close()
# build gif
with imageio.get_writer('./figs/tinker_vid.mp4', mode='I', fps=5) as writer:
for filename in filenames:
image = imageio.imread(filename)
writer.append_data(image)
# frames = []
# for filename in filenames:
# image = imageio.imread(filename)
# frames.append(image)
# imageio.mimsave('./figs/tinker_gif.gif', frames)
# optimize('./figs/tinker_gif.gif')
# Remove files
for filename in set(filenames):
os.remove(filename)
# -
from IPython.display import Video
Video(data="./figs/tinker_vid.mp4")
# # Check Dim 10
threshold = np.percentile(y_caps_dev[:,9], 95.5)
threshold2 = np.percentile(y_caps_dev[:,9], 95.6)
mask = (y_caps_dev[:,9]>threshold) & (y_caps_dev[:,9]<threshold2)
# +
config["input_shape"] = config["image_shape"]
CapsNet = import_model(model_name=config["model_name"])
train_model, eval_model,manipulate_model,decoder_model,redshift_model, = CapsNet(**config)
manipulate_model.load_weights(
path_results / "weights" / f"weights-{checkpoint_eval:02d}.h5", by_name=True
)
sigma_arr = np.std(y_caps_dev, axis=0)
caps_gal = y_caps_dev.copy()
change_grid = [-3,-2,-1,0,1,2,3]
fig, axs = plt.subplots(np.sum(mask),len(change_grid), figsize=(30,200))
for j in range(len(change_grid)):
tinkered_dim = caps_gal[mask][:,9] + change_grid[j] * sigma_arr[9]
tinkered_caps = (caps_gal[mask]).copy()
tinkered_caps[:,9] = tinkered_dim
tinkered_recon = decoder_model.predict(tinkered_caps)
for k in range(np.sum(mask)):
plot_image(scale*tinkered_recon[k], "gri", ax =axs[k][j] )
cols = ['{}$\sigma$'.format(col) for col in change_grid]
# rows = ['Dim: {}'.format(row) for row in np.arange(1,num_caps+1).astype(str) ]
for ax, col in zip(axs[0], cols):
ax.set_title(col, size=25)
# for ax, row in zip(axs[:,0], rows):
# ax.set_ylabel(row, size=23)
# # fig.suptitle("Redshift: "+ str(z_spec_dev[img_indx]), y =1.01, size=20)
# plt.tight_layout()
# fig.savefig("./figs/tinker_spheroid_appendix.pdf",bbox_inches='tight',dpi=300)
# -
| notebooks/plot_galaxy_pictures.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import bmt
import requests
import json
from pprint import pprint
import csv
tk = bmt.Toolkit()
# -
association_slots = tk.get_all_edge_properties(True)
#pprint(association_slots)
geno_class = tk.get_all_elements_by_mapping('GENO:0000536', True)
#pprint(geno_class)
all_classes = tk.get_all_classes()
#pprint(all_classes)
all_associations = tk.get_all_associations()
predicate_descendents = tk.get_descendants("related to")
#geno_ancestors = tk.get_ancestors("Genotype", False)
predicate_map = {}
mapping_list = []
with open('/tmp/predicates_full.csv', 'w') as csv_file:
writer = csv.writer(csv_file)
for descendent in predicate_descendents:
p_element = tk.get_element(descendent)
for mapping in p_element.exact_mappings:
mapping_list.append(mapping)
for mapping_n in p_element.narrow_mappings:
mapping_list.append(mapping_n)
for mapping_b in p_element.broad_mappings:
mapping_list.append(mapping_b)
if 'RO' not in mapping_list:
#if tk.has_inverse(descendent):
# print(descendent)
# continue
#else:
for item in mapping_list:
#print(descendent + "," + item)
writer.writerow([descendent, mapping])
print(descendent)
#with open('/tmp/predicates_full.csv', 'w') as csv_file:
# writer = csv.writer(csv_file)
# for key, value in predicate_map.items():
# writer.writerow([key, value])
# pprint(p_element)
#pprint(geno_descendents)
#pprint(geno_ancestors)
#pprint(all_associations)
| examples/alliance-demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from tqdm import tqdm
import timeit
from matplotlib import pyplot as plt
from Code import ULA
from Code import Algo1
from Code import ZVnbrosse
# ### Gaussian mixture
#
# ## $\pi (x) = \frac{1}{2(2\pi)^{d/2}} \left( e ^{\frac{-| x-a|^2}{2}} + e ^{\frac{-| x+a|^2}{2}} \right), \quad x \in\mathbb R^d$
#
# $U(x) = \frac{1}{2} \|x - a\|_2^2 - \text{log}(1 + e^{-2x^\top a})$
#
# $\nabla U(x) = x-a +2a(1 + e^{2 x^\top a})^{-1}$
#
# $ m = 1 - \|a \|_2^2 \quad $ (strongly convex function)
#
# $M = 1 \quad$ (Lipschitz continuous gradient)
#
# $a = (\frac{1}{\sqrt{2d}}, \dots, \frac{1}{\sqrt{2d}})$
# ## Setup
# +
dim = 2
h = 0.2
n = 10000
N_train = 10
N_test = 100
deg = 1
def f_grad(x):
a = np.ones(dim) / np.sqrt(2*dim)
return x - a + 2 * a/(1 + np.exp(2* (x.T @ a)))
def local_weighted_estimator(X):
return X.sum(axis=1).mean()
def estimator(XX):
Pi = np.empty(XX.shape[0])
for i in tqdm(range(XX.shape[0])):
Pi[i] = local_weighted_estimator(XX[i])
return Pi
# -
f_target = 'sum'
# ### Generate sample and plot
np.random.seed(2342)
X, G, Z = ULA.ULA_with_burnin(d=dim, step=h, burn_in=10000, n=100000,f_grad=f_grad)
ULA.plot_distr(X,Z,G,0)
ULA.plot_distr(X,Z,G,1)
# ### Generate train ant test trajectories
# +
np.random.seed(123)
XX, GG, ZZ = ULA.generate_train_trajectories(X, N_train=N_train,d = dim, step = h, n = n, f_grad=f_grad)
XX_test, GG_test, ZZ_test = ULA.generate_test_trajetories(N_test=N_test, d =dim, step=h,
burn_in=10000, n = n, f_grad=f_grad)
# -
# ### Algorithm 1
Betas, degrees = Algo1.G_pml_fit_mean(XX,f_target, max_deg = deg)
k_comb = np.zeros((dim,dim), dtype=np.int16)
for i in range(dim):
k_comb[i,i] = 1
# +
# k_comb = np.array([[1,0], [0,1], [1,1], [2,0], [0,2]], dtype=np.int16)
# -
# ### Check and time
i_test = 5
time_1 = timeit.default_timer()
p = local_weighted_estimator(XX_test[i_test])
print ("mean of target function =", p)
for i in range(len(k_comb)):
p = p - Algo1.M_bias(k_comb[i],XX_test[i_test],GG_test[i_test], ZZ_test[i_test], h, degrees,Betas,50)
print (" (k = {}) = {} [Time:{}]".format(k_comb[i],p,timeit.default_timer()-time_1))
# ## Check results on test trajectories
P_test = estimator(XX_test)
# +
res_zv_1 = []
for i in range (XX_test.shape[0]):
res_zv_1.append(ZVnbrosse.ZVpolyOne(XX_test[i].reshape(-1,dim), GG_test[i].reshape(-1,dim), f_target))
res_zv_1 = np.array(res_zv_1).reshape(-1)
res_zv_2 = []
for i in range (XX_test.shape[0]):
res_zv_2.append(ZVnbrosse.ZVpolyTwo(XX_test[i].reshape(-1,dim), GG_test[i].reshape(-1,dim), f_target))
res_zv_2 = np.array(res_zv_2).reshape(-1)
res_cv_1 = []
for i in range (XX_test.shape[0]):
res_cv_1.append(ZVnbrosse.CVpolyOne(XX_test[i].reshape(-1,dim), GG_test[i].reshape(-1,dim), f_target))
res_cv_1 = np.array(res_cv_1).reshape(-1)
res_cv_2 = []
for i in range (XX_test.shape[0]):
res_cv_2.append(ZVnbrosse.CVpolyTwo(XX_test[i].reshape(-1,dim), GG_test[i].reshape(-1,dim), f_target))
res_cv_2 = np.array(res_cv_2).reshape(-1)
# -
def save_plot(M_test, n_tilde):
all_data = [P_test]
all_data.append(P_test - M_test.sum(axis = 0))
all_data.append(res_zv_1)
all_data.append(res_cv_1)
plt.figure(figsize=(10,10))
plt.violinplot(all_data, showmeans=True, showmedians=False)
plt.xticks(np.arange(1,5), ('O', 'CV_B', 'ZV', 'CV'))
plt.tick_params(labelsize = 15)
plt.grid()
plt.savefig("Algo1_logs/GM_{}d_violin_(h = {}, n = {}, K = 1, n_tilde={}, f_target = {}).png".format(dim, h, n, n_tilde,f_target))
all_data.append(res_zv_2)
all_data.append(res_cv_2)
plt.figure(figsize=(10,10))
plt.violinplot(all_data, showmeans=True, showmedians=False)
plt.xticks(np.arange(1,7), ('O', 'CV_B', 'ZV-1', 'CV-1', 'ZV-2', 'CV-2'))
plt.tick_params(labelsize = 15)
plt.grid()
plt.savefig("Algo1_logs/GM_{}d_violin_full_(h = {}, n = {}, K = 1, n_tilde={}, f_target = {}).png".format(dim, h, n, n_tilde,f_target))
for n_tilde in [30, 50, 70]:
M_test = np.empty((len(k_comb),P_test.shape[0]))
for i in range(len(k_comb)):
M_test[i] = Algo1.estimator_bias(k_comb[i], XX_test, GG_test, ZZ_test, h, degrees,Betas, n_tilde, n_jobs=3)
save_plot(M_test, n_tilde)
np.save("Algo1_logs/GM_{}d_M_test(h = {}, n = {}, K = 1, n_tilde={}, f_target = {}).npy".
format(dim, h, n, n_tilde,f_target), M_test)
print("plots_saved for n_tilde = {}".format(n_tilde))
# +
plt.figure(figsize=(10,10))
plt.hist(P_test,15, facecolor='r', density=True, alpha=1, label=r"$\pi$")
plt.hist((P_test-M_test.sum(axis=0)),15, density=True, facecolor='g', alpha=0.8, label="Distribution of $\pi - M^2_N$")
plt.hist(res_zv_1,15, facecolor='y', density=True, alpha=0.5, label="ZV 1")
# plt.hist(res_zv_2,15, facecolor='b', density=True, alpha=0.5, label="ZV 2")
plt.hist(res_cv_1,15, facecolor='pink', density=True, alpha=0.5, label="CV 1")
# plt.hist(res_cv_2,15, facecolor='black', density=True, alpha=0.5, label="CV 2")
plt.legend(loc=2, prop={'size': 17})
plt.grid(linestyle='-', linewidth=0.2, color='black')
plt.tick_params(axis='y',color = 'w', labelcolor = 'w', which='both')
plt.xlim(-0.65, 0.65)
plt.show()
# +
all_data = []
all_data.append(P_test - M_test.sum(axis = 0))
all_data.append(res_zv_1)
# all_data.append(res_zv_2)
all_data.append(res_cv_1)
# all_data.append(res_cv_2)
plt.figure(figsize=(10,10))
plt.violinplot(all_data, showmeans=True, showmedians=False)
# plt.title('violin plot')
plt.xticks(np.arange(1,5), ('pi', 'CV_B', 'ZV_1', 'CV_1'))
plt.grid()
plt.show()
# +
all_data = [P_test]
all_data.append(P_test - M_test.sum(axis = 0))
all_data.append(res_zv_1)
# all_data.append(res_zv_2)
all_data.append(res_cv_1)
# all_data.append(res_cv_2)
plt.figure(figsize=(10,10))
plt.violinplot(all_data, showmeans=True, showmedians=False)
# plt.title('violin plot')
plt.xticks(np.arange(1,5), ('pi', 'CV_B', 'ZV_1', 'CV_1'))
plt.grid()
plt.show()
# -
print (' Variance of Pi = ',P_test.var(ddof = 1))
for i in range(len(k_comb)):
print ('Variance of new estimator =',(P_test-M_test[:i+1].sum(axis = 0)).var(ddof = 1))
print (' Variance of CV 1 = ',res_cv_1.var(ddof=1))
print (' Variance of CV 2 = ',res_cv_2.var(ddof=1))
print (' Variance of ZV 1 = ',res_zv_1.var(ddof=1))
print (' Variance of ZV 2 = ',res_zv_2.var(ddof=1))
print ('VRF CVB = ',P_test.var(ddof = 1)/ (P_test - M_test.sum(axis = 0)).var(ddof=1))
print ('VRF CV1 = ',P_test.var(ddof = 1)/res_cv_1.var(ddof = 1))
print ('VRF CV2 = ',P_test.var(ddof = 1)/res_cv_2.var(ddof = 1))
print ('VRF ZV1 = ',P_test.var(ddof = 1)/res_zv_1.var(ddof = 1))
print ('VRF ZV2 = ',P_test.var(ddof = 1)/res_zv_2.var(ddof = 1))
| GM_2d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
#
#
# # Background information on filtering
#
#
# Here we give some background information on filtering in general,
# and how it is done in MNE-Python in particular.
# Recommended reading for practical applications of digital
# filter design can be found in Parks & Burrus [1]_ and
# Ifeachor and Jervis [2]_, and for filtering in an
# M/EEG context we recommend reading Widmann *et al.* 2015 [7]_.
# To see how to use the default filters in MNE-Python on actual data, see
# the `tut_artifacts_filter` tutorial.
#
# Problem statement
# =================
#
# The practical issues with filtering electrophysiological data are covered
# well by Widmann *et al.* in [7]_, in a follow-up to an article where they
# conclude with this statement:
#
# Filtering can result in considerable distortions of the time course
# (and amplitude) of a signal as demonstrated by VanRullen (2011) [[3]_].
# Thus, filtering should not be used lightly. However, if effects of
# filtering are cautiously considered and filter artifacts are minimized,
# a valid interpretation of the temporal dynamics of filtered
# electrophysiological data is possible and signals missed otherwise
# can be detected with filtering.
#
# In other words, filtering can increase SNR, but if it is not used carefully,
# it can distort data. Here we hope to cover some filtering basics so
# users can better understand filtering tradeoffs, and why MNE-Python has
# chosen particular defaults.
#
#
# Filtering basics
# ================
#
# Let's get some of the basic math down. In the frequency domain, digital
# filters have a transfer function that is given by:
#
# \begin{align}H(z) &= \frac{b_0 + b_1 z^{-1} + b_2 z^{-2} + ... + b_M z^{-M}}
# {1 + a_1 z^{-1} + a_2 z^{-2} + ... + a_N z^{-M}} \\
# &= \frac{\sum_0^Mb_kz^{-k}}{\sum_1^Na_kz^{-k}}\end{align}
#
# In the time domain, the numerator coefficients $b_k$ and denominator
# coefficients $a_k$ can be used to obtain our output data
# $y(n)$ in terms of our input data $x(n)$ as:
#
# \begin{align}:label: summations
#
# y(n) &= b_0 x(n) + b_1 x(n-1) + ... + b_M x(n-M)
# - a_1 y(n-1) - a_2 y(n - 2) - ... - a_N y(n - N)\\
# &= \sum_0^M b_k x(n-k) - \sum_1^N a_k y(n-k)\end{align}
#
# In other words, the output at time $n$ is determined by a sum over:
#
# 1. The numerator coefficients $b_k$, which get multiplied by
# the previous input $x(n-k)$ values, and
# 2. The denominator coefficients $a_k$, which get multiplied by
# the previous output $y(n-k)$ values.
#
# Note that these summations in :eq:`summations` correspond nicely to
# (1) a weighted `moving average`_ and (2) an autoregression_.
#
# Filters are broken into two classes: FIR_ (finite impulse response) and
# IIR_ (infinite impulse response) based on these coefficients.
# FIR filters use a finite number of numerator
# coefficients $b_k$ ($\forall k, a_k=0$), and thus each output
# value of $y(n)$ depends only on the $M$ previous input values.
# IIR filters depend on the previous input and output values, and thus can have
# effectively infinite impulse responses.
#
# As outlined in [1]_, FIR and IIR have different tradeoffs:
#
# * A causal FIR filter can be linear-phase -- i.e., the same time delay
# across all frequencies -- whereas a causal IIR filter cannot. The phase
# and group delay characteristics are also usually better for FIR filters.
# * IIR filters can generally have a steeper cutoff than an FIR filter of
# equivalent order.
# * IIR filters are generally less numerically stable, in part due to
# accumulating error (due to its recursive calculations).
#
# In MNE-Python we default to using FIR filtering. As noted in Widmann *et al.*
# 2015 [7]_:
#
# Despite IIR filters often being considered as computationally more
# efficient, they are recommended only when high throughput and sharp
# cutoffs are required (Ifeachor and Jervis, 2002 [2]_, p. 321),
# ...FIR filters are easier to control, are always stable, have a
# well-defined passband, can be corrected to zero-phase without
# additional computations, and can be converted to minimum-phase.
# We therefore recommend FIR filters for most purposes in
# electrophysiological data analysis.
#
# When designing a filter (FIR or IIR), there are always tradeoffs that
# need to be considered, including but not limited to:
#
# 1. Ripple in the pass-band
# 2. Attenuation of the stop-band
# 3. Steepness of roll-off
# 4. Filter order (i.e., length for FIR filters)
# 5. Time-domain ringing
#
# In general, the sharper something is in frequency, the broader it is in time,
# and vice-versa. This is a fundamental time-frequency tradeoff, and it will
# show up below.
#
# FIR Filters
# ===========
#
# First we will focus first on FIR filters, which are the default filters used by
# MNE-Python.
#
#
# Designing FIR filters
# ---------------------
# Here we'll try designing a low-pass filter, and look at trade-offs in terms
# of time- and frequency-domain filter characteristics. Later, in
# `tut_effect_on_signals`, we'll look at how such filters can affect
# signals when they are used.
#
# First let's import some useful tools for filtering, and set some default
# values for our data that are reasonable for M/EEG data.
#
#
# +
import numpy as np
from scipy import signal, fftpack
import matplotlib.pyplot as plt
from mne.time_frequency.tfr import morlet
from mne.viz import plot_filter, plot_ideal_filter
import mne
sfreq = 1000.
f_p = 40.
flim = (1., sfreq / 2.) # limits for plotting
# -
# Take for example an ideal low-pass filter, which would give a value of 1 in
# the pass-band (up to frequency $f_p$) and a value of 0 in the stop-band
# (down to frequency $f_s$) such that $f_p=f_s=40$ Hz here
# (shown to a lower limit of -60 dB for simplicity):
#
#
# +
nyq = sfreq / 2. # the Nyquist frequency is half our sample rate
freq = [0, f_p, f_p, nyq]
gain = [1, 1, 0, 0]
third_height = np.array(plt.rcParams['figure.figsize']) * [1, 1. / 3.]
ax = plt.subplots(1, figsize=third_height)[1]
plot_ideal_filter(freq, gain, ax, title='Ideal %s Hz lowpass' % f_p, flim=flim)
# -
# This filter hypothetically achieves zero ripple in the frequency domain,
# perfect attenuation, and perfect steepness. However, due to the discontunity
# in the frequency response, the filter would require infinite ringing in the
# time domain (i.e., infinite order) to be realized. Another way to think of
# this is that a rectangular window in frequency is actually sinc_ function
# in time, which requires an infinite number of samples, and thus infinite
# time, to represent. So although this filter has ideal frequency suppression,
# it has poor time-domain characteristics.
#
# Let's try to naïvely make a brick-wall filter of length 0.1 sec, and look
# at the filter itself in the time domain and the frequency domain:
#
#
n = int(round(0.1 * sfreq)) + 1
t = np.arange(-n // 2, n // 2) / sfreq # center our sinc
h = np.sinc(2 * f_p * t) / (4 * np.pi)
plot_filter(h, sfreq, freq, gain, 'Sinc (0.1 sec)', flim=flim)
# This is not so good! Making the filter 10 times longer (1 sec) gets us a
# bit better stop-band suppression, but still has a lot of ringing in
# the time domain. Note the x-axis is an order of magnitude longer here,
# and the filter has a correspondingly much longer group delay (again equal
# to half the filter length, or 0.5 seconds):
#
#
n = int(round(1. * sfreq)) + 1
t = np.arange(-n // 2, n // 2) / sfreq
h = np.sinc(2 * f_p * t) / (4 * np.pi)
plot_filter(h, sfreq, freq, gain, 'Sinc (1.0 sec)', flim=flim)
# Let's make the stop-band tighter still with a longer filter (10 sec),
# with a resulting larger x-axis:
#
#
n = int(round(10. * sfreq)) + 1
t = np.arange(-n // 2, n // 2) / sfreq
h = np.sinc(2 * f_p * t) / (4 * np.pi)
plot_filter(h, sfreq, freq, gain, 'Sinc (10.0 sec)', flim=flim)
# Now we have very sharp frequency suppression, but our filter rings for the
# entire second. So this naïve method is probably not a good way to build
# our low-pass filter.
#
# Fortunately, there are multiple established methods to design FIR filters
# based on desired response characteristics. These include:
#
# 1. The Remez_ algorithm (:func:`scipy.signal.remez`, `MATLAB firpm`_)
# 2. Windowed FIR design (:func:`scipy.signal.firwin2`, `MATLAB fir2`_
# and :func:`scipy.signal.firwin`)
# 3. Least squares designs (:func:`scipy.signal.firls`, `MATLAB firls`_)
# 4. Frequency-domain design (construct filter in Fourier
# domain and use an :func:`IFFT <scipy.fftpack.ifft>` to invert it)
#
# <div class="alert alert-info"><h4>Note</h4><p>Remez and least squares designs have advantages when there are
# "do not care" regions in our frequency response. However, we want
# well controlled responses in all frequency regions.
# Frequency-domain construction is good when an arbitrary response
# is desired, but generally less clean (due to sampling issues) than
# a windowed approach for more straightfroward filter applications.
# Since our filters (low-pass, high-pass, band-pass, band-stop)
# are fairly simple and we require precisel control of all frequency
# regions, here we will use and explore primarily windowed FIR
# design.</p></div>
#
# If we relax our frequency-domain filter requirements a little bit, we can
# use these functions to construct a lowpass filter that instead has a
# *transition band*, or a region between the pass frequency $f_p$
# and stop frequency $f_s$, e.g.:
#
#
# +
trans_bandwidth = 10 # 10 Hz transition band
f_s = f_p + trans_bandwidth # = 50 Hz
freq = [0., f_p, f_s, nyq]
gain = [1., 1., 0., 0.]
ax = plt.subplots(1, figsize=third_height)[1]
title = '%s Hz lowpass with a %s Hz transition' % (f_p, trans_bandwidth)
plot_ideal_filter(freq, gain, ax, title=title, flim=flim)
# -
# Accepting a shallower roll-off of the filter in the frequency domain makes
# our time-domain response potentially much better. We end up with a
# smoother slope through the transition region, but a *much* cleaner time
# domain signal. Here again for the 1 sec filter:
#
#
h = signal.firwin2(n, freq, gain, nyq=nyq)
plot_filter(h, sfreq, freq, gain, 'Windowed 10-Hz transition (1.0 sec)',
flim=flim)
# Since our lowpass is around 40 Hz with a 10 Hz transition, we can actually
# use a shorter filter (5 cycles at 10 Hz = 0.5 sec) and still get okay
# stop-band attenuation:
#
#
n = int(round(sfreq * 0.5)) + 1
h = signal.firwin2(n, freq, gain, nyq=nyq)
plot_filter(h, sfreq, freq, gain, 'Windowed 10-Hz transition (0.5 sec)',
flim=flim)
# But then if we shorten the filter too much (2 cycles of 10 Hz = 0.2 sec),
# our effective stop frequency gets pushed out past 60 Hz:
#
#
n = int(round(sfreq * 0.2)) + 1
h = signal.firwin2(n, freq, gain, nyq=nyq)
plot_filter(h, sfreq, freq, gain, 'Windowed 10-Hz transition (0.2 sec)',
flim=flim)
# If we want a filter that is only 0.1 seconds long, we should probably use
# something more like a 25 Hz transition band (0.2 sec = 5 cycles @ 25 Hz):
#
#
trans_bandwidth = 25
f_s = f_p + trans_bandwidth
freq = [0, f_p, f_s, nyq]
h = signal.firwin2(n, freq, gain, nyq=nyq)
plot_filter(h, sfreq, freq, gain, 'Windowed 50-Hz transition (0.2 sec)',
flim=flim)
# So far we have only discussed *acausal* filtering, which means that each
# sample at each time point $t$ is filtered using samples that come
# after ($t + \Delta t$) *and* before ($t - \Delta t$) $t$.
# In this sense, each sample is influenced by samples that come both before
# and after it. This is useful in many cases, espcially because it does not
# delay the timing of events.
#
# However, sometimes it can be beneficial to use *causal* filtering,
# whereby each sample $t$ is filtered only using time points that came
# after it.
#
# Note that the delay is variable (whereas for linear/zero-phase filters it
# is constant) but small in the pass-band. Unlike zero-phase filters, which
# require time-shifting backward the output of a linear-phase filtering stage
# (and thus becoming acausal), minimum-phase filters do not require any
# compensation to achieve small delays in the passband. Note that as an
# artifact of the minimum phase filter construction step, the filter does
# not end up being as steep as the linear/zero-phase version.
#
# We can construct a minimum-phase filter from our existing linear-phase
# filter with the ``minimum_phase`` function (that will be in SciPy 0.19's
# :mod:`scipy.signal`), and note that the falloff is not as steep:
#
#
h_min = mne.fixes.minimum_phase(h)
plot_filter(h_min, sfreq, freq, gain, 'Minimum-phase', flim=flim)
#
# Applying FIR filters
# --------------------
#
# Now lets look at some practical effects of these filters by applying
# them to some data.
#
# Let's construct a Gaussian-windowed sinusoid (i.e., Morlet imaginary part)
# plus noise (random + line). Note that the original, clean signal contains
# frequency content in both the pass band and transition bands of our
# low-pass filter.
#
#
# +
dur = 10.
center = 2.
morlet_freq = f_p
tlim = [center - 0.2, center + 0.2]
tticks = [tlim[0], center, tlim[1]]
flim = [20, 70]
x = np.zeros(int(sfreq * dur) + 1)
blip = morlet(sfreq, [morlet_freq], n_cycles=7)[0].imag / 20.
n_onset = int(center * sfreq) - len(blip) // 2
x[n_onset:n_onset + len(blip)] += blip
x_orig = x.copy()
rng = np.random.RandomState(0)
x += rng.randn(len(x)) / 1000.
x += np.sin(2. * np.pi * 60. * np.arange(len(x)) / sfreq) / 2000.
# -
# Filter it with a shallow cutoff, linear-phase FIR (which allows us to
# compensate for the constant filter delay):
#
#
# +
transition_band = 0.25 * f_p
f_s = f_p + transition_band
filter_dur = 6.6 / transition_band / 2. # sec
n = int(sfreq * filter_dur)
freq = [0., f_p, f_s, sfreq / 2.]
gain = [1., 1., 0., 0.]
# This would be equivalent:
h = mne.filter.create_filter(x, sfreq, l_freq=None, h_freq=f_p,
fir_design='firwin')
x_v16 = np.convolve(h, x)[len(h) // 2:]
plot_filter(h, sfreq, freq, gain, 'MNE-Python 0.16 default', flim=flim)
# -
# Filter it with a different design mode ``fir_design="firwin2"``, and also
# compensate for the constant filter delay. This method does not produce
# quite as sharp a transition compared to ``fir_design="firwin"``, despite
# being twice as long:
#
#
# +
transition_band = 0.25 * f_p
f_s = f_p + transition_band
filter_dur = 6.6 / transition_band # sec
n = int(sfreq * filter_dur)
freq = [0., f_p, f_s, sfreq / 2.]
gain = [1., 1., 0., 0.]
# This would be equivalent:
# h = signal.firwin2(n, freq, gain, nyq=sfreq / 2.)
h = mne.filter.create_filter(x, sfreq, l_freq=None, h_freq=f_p,
fir_design='firwin2')
x_v14 = np.convolve(h, x)[len(h) // 2:]
plot_filter(h, sfreq, freq, gain, 'MNE-Python 0.14 default', flim=flim)
# -
# This is actually set to become the default type of filter used in MNE-Python
# in 0.14 (see `tut_filtering_in_python`).
#
# Let's also filter with the MNE-Python 0.13 default, which is a
# long-duration, steep cutoff FIR that gets applied twice:
#
#
# +
transition_band = 0.5 # Hz
f_s = f_p + transition_band
filter_dur = 10. # sec
n = int(sfreq * filter_dur)
freq = [0., f_p, f_s, sfreq / 2.]
gain = [1., 1., 0., 0.]
# This would be equivalent
# h = signal.firwin2(n, freq, gain, nyq=sfreq / 2.)
h = mne.filter.create_filter(x, sfreq, l_freq=None, h_freq=f_p,
h_trans_bandwidth=transition_band,
filter_length='%ss' % filter_dur,
fir_design='firwin2')
x_v13 = np.convolve(np.convolve(h, x)[::-1], h)[::-1][len(h) - 1:-len(h) - 1]
plot_filter(h, sfreq, freq, gain, 'MNE-Python 0.13 default', flim=flim)
# -
# Let's also filter it with the MNE-C default, which is a long-duration
# steep-slope FIR filter designed using frequency-domain techniques:
#
#
# +
h = mne.filter.design_mne_c_filter(sfreq, l_freq=None, h_freq=f_p + 2.5)
x_mne_c = np.convolve(h, x)[len(h) // 2:]
transition_band = 5 # Hz (default in MNE-C)
f_s = f_p + transition_band
freq = [0., f_p, f_s, sfreq / 2.]
gain = [1., 1., 0., 0.]
plot_filter(h, sfreq, freq, gain, 'MNE-C default', flim=flim)
# -
# And now an example of a minimum-phase filter:
#
#
h = mne.filter.create_filter(x, sfreq, l_freq=None, h_freq=f_p,
phase='minimum', fir_design='firwin')
x_min = np.convolve(h, x)
transition_band = 0.25 * f_p
f_s = f_p + transition_band
filter_dur = 6.6 / transition_band # sec
n = int(sfreq * filter_dur)
freq = [0., f_p, f_s, sfreq / 2.]
gain = [1., 1., 0., 0.]
plot_filter(h, sfreq, freq, gain, 'Minimum-phase filter', flim=flim)
# Both the MNE-Python 0.13 and MNE-C filhters have excellent frequency
# attenuation, but it comes at a cost of potential
# ringing (long-lasting ripples) in the time domain. Ringing can occur with
# steep filters, especially on signals with frequency content around the
# transition band. Our Morlet wavelet signal has power in our transition band,
# and the time-domain ringing is thus more pronounced for the steep-slope,
# long-duration filter than the shorter, shallower-slope filter:
#
#
# +
axes = plt.subplots(1, 2)[1]
def plot_signal(x, offset):
t = np.arange(len(x)) / sfreq
axes[0].plot(t, x + offset)
axes[0].set(xlabel='Time (sec)', xlim=t[[0, -1]])
X = fftpack.fft(x)
freqs = fftpack.fftfreq(len(x), 1. / sfreq)
mask = freqs >= 0
X = X[mask]
freqs = freqs[mask]
axes[1].plot(freqs, 20 * np.log10(np.abs(X)))
axes[1].set(xlim=flim)
yticks = np.arange(7) / -30.
yticklabels = ['Original', 'Noisy', 'FIR-firwin (0.16)', 'FIR-firwin2 (0.14)',
'FIR-steep (0.13)', 'FIR-steep (MNE-C)', 'Minimum-phase']
plot_signal(x_orig, offset=yticks[0])
plot_signal(x, offset=yticks[1])
plot_signal(x_v16, offset=yticks[2])
plot_signal(x_v14, offset=yticks[3])
plot_signal(x_v13, offset=yticks[4])
plot_signal(x_mne_c, offset=yticks[5])
plot_signal(x_min, offset=yticks[6])
axes[0].set(xlim=tlim, title='FIR, Lowpass=%d Hz' % f_p, xticks=tticks,
ylim=[-0.200, 0.025], yticks=yticks, yticklabels=yticklabels,)
for text in axes[0].get_yticklabels():
text.set(rotation=45, size=8)
axes[1].set(xlim=flim, ylim=(-60, 10), xlabel='Frequency (Hz)',
ylabel='Magnitude (dB)')
mne.viz.tight_layout()
plt.show()
# -
# IIR filters
# ===========
#
# MNE-Python also offers IIR filtering functionality that is based on the
# methods from :mod:`scipy.signal`. Specifically, we use the general-purpose
# functions :func:`scipy.signal.iirfilter` and :func:`scipy.signal.iirdesign`,
# which provide unified interfaces to IIR filter design.
#
# Designing IIR filters
# ---------------------
#
# Let's continue with our design of a 40 Hz low-pass filter, and look at
# some trade-offs of different IIR filters.
#
# Often the default IIR filter is a `Butterworth filter`_, which is designed
# to have a *maximally flat pass-band*. Let's look at a few orders of filter,
# i.e., a few different number of coefficients used and therefore steepness
# of the filter:
#
# <div class="alert alert-info"><h4>Note</h4><p>Notice that the group delay (which is related to the phase) of
# the IIR filters below are not constant. In the FIR case, we can
# design so-called linear-phase filters that have a constant group
# delay, and thus compensate for the delay (making the filter
# acausal) if necessary. This cannot be done with IIR filters, as
# they have a non-linear phase (non-constant group delay). As the
# filter order increases, the phase distortion near and in the
# transition band worsens. However, if acausal (forward-backward)
# filtering can be used, e.g. with :func:`scipy.signal.filtfilt`,
# these phase issues can theoretically be mitigated.</p></div>
#
#
# +
sos = signal.iirfilter(2, f_p / nyq, btype='low', ftype='butter', output='sos')
plot_filter(dict(sos=sos), sfreq, freq, gain, 'Butterworth order=2', flim=flim)
# Eventually this will just be from scipy signal.sosfiltfilt, but 0.18 is
# not widely adopted yet (as of June 2016), so we use our wrapper...
sosfiltfilt = mne.fixes.get_sosfiltfilt()
x_shallow = sosfiltfilt(sos, x)
# -
# The falloff of this filter is not very steep.
#
# <div class="alert alert-info"><h4>Note</h4><p>Here we have made use of second-order sections (SOS)
# by using :func:`scipy.signal.sosfilt` and, under the
# hood, :func:`scipy.signal.zpk2sos` when passing the
# ``output='sos'`` keyword argument to
# :func:`scipy.signal.iirfilter`. The filter definitions
# given in tut_filtering_basics_ use the polynomial
# numerator/denominator (sometimes called "tf") form ``(b, a)``,
# which are theoretically equivalent to the SOS form used here.
# In practice, however, the SOS form can give much better results
# due to issues with numerical precision (see
# :func:`scipy.signal.sosfilt` for an example), so SOS should be
# used when possible to do IIR filtering.</p></div>
#
# Let's increase the order, and note that now we have better attenuation,
# with a longer impulse response:
#
#
sos = signal.iirfilter(8, f_p / nyq, btype='low', ftype='butter', output='sos')
plot_filter(dict(sos=sos), sfreq, freq, gain, 'Butterworth order=8', flim=flim)
x_steep = sosfiltfilt(sos, x)
# There are other types of IIR filters that we can use. For a complete list,
# check out the documentation for :func:`scipy.signal.iirdesign`. Let's
# try a Chebychev (type I) filter, which trades off ripple in the pass-band
# to get better attenuation in the stop-band:
#
#
sos = signal.iirfilter(8, f_p / nyq, btype='low', ftype='cheby1', output='sos',
rp=1) # dB of acceptable pass-band ripple
plot_filter(dict(sos=sos), sfreq, freq, gain,
'Chebychev-1 order=8, ripple=1 dB', flim=flim)
# And if we can live with even more ripple, we can get it slightly steeper,
# but the impulse response begins to ring substantially longer (note the
# different x-axis scale):
#
#
sos = signal.iirfilter(8, f_p / nyq, btype='low', ftype='cheby1', output='sos',
rp=6)
plot_filter(dict(sos=sos), sfreq, freq, gain,
'Chebychev-1 order=8, ripple=6 dB', flim=flim)
# Applying IIR filters
# --------------------
#
# Now let's look at how our shallow and steep Butterworth IIR filters
# perform on our Morlet signal from before:
#
#
axes = plt.subplots(1, 2)[1]
yticks = np.arange(4) / -30.
yticklabels = ['Original', 'Noisy', 'Butterworth-2', 'Butterworth-8']
plot_signal(x_orig, offset=yticks[0])
plot_signal(x, offset=yticks[1])
plot_signal(x_shallow, offset=yticks[2])
plot_signal(x_steep, offset=yticks[3])
axes[0].set(xlim=tlim, title='IIR, Lowpass=%d Hz' % f_p, xticks=tticks,
ylim=[-0.125, 0.025], yticks=yticks, yticklabels=yticklabels,)
for text in axes[0].get_yticklabels():
text.set(rotation=45, size=8)
axes[1].set(xlim=flim, ylim=(-60, 10), xlabel='Frequency (Hz)',
ylabel='Magnitude (dB)')
mne.viz.adjust_axes(axes)
mne.viz.tight_layout()
plt.show()
# Some pitfalls of filtering
# ==========================
#
# Multiple recent papers have noted potential risks of drawing
# errant inferences due to misapplication of filters.
#
# Low-pass problems
# -----------------
#
# Filters in general, especially those that are acausal (zero-phase), can make
# activity appear to occur earlier or later than it truly did. As
# mentioned in VanRullen 2011 [3]_, investigations of commonly (at the time)
# used low-pass filters created artifacts when they were applied to smulated
# data. However, such deleterious effects were minimal in many real-world
# examples in Rousselet 2012 [5]_.
#
# Perhaps more revealing, it was noted in Widmann & Schröger 2012 [6]_ that
# the problematic low-pass filters from VanRullen 2011 [3]_:
#
# 1. Used a least-squares design (like :func:`scipy.signal.firls`) that
# included "do-not-care" transition regions, which can lead to
# uncontrolled behavior.
# 2. Had a filter length that was independent of the transition bandwidth,
# which can cause excessive ringing and signal distortion.
#
#
# High-pass problems
# ------------------
#
# When it comes to high-pass filtering, using corner frequencies above 0.1 Hz
# were found in Acunzo *et al.* 2012 [4]_ to:
#
# "...generate a systematic bias easily leading to misinterpretations of
# neural activity.”
#
# In a related paper, Widmann *et al.* 2015 [7]_ also came to suggest a 0.1 Hz
# highpass. And more evidence followed in Tanner *et al.* 2015 [8]_ of such
# distortions. Using data from language ERP studies of semantic and syntactic
# processing (i.e., N400 and P600), using a high-pass above 0.3 Hz caused
# significant effects to be introduced implausibly early when compared to the
# unfiltered data. From this, the authors suggested the optimal high-pass
# value for language processing to be 0.1 Hz.
#
# We can recreate a problematic simulation from Tanner *et al.* 2015 [8]_:
#
# "The simulated component is a single-cycle cosine wave with an amplitude
# of 5µV, onset of 500 ms poststimulus, and duration of 800 ms. The
# simulated component was embedded in 20 s of zero values to avoid
# filtering edge effects... Distortions [were] caused by 2 Hz low-pass and
# high-pass filters... No visible distortion to the original waveform
# [occurred] with 30 Hz low-pass and 0.01 Hz high-pass filters...
# Filter frequencies correspond to the half-amplitude (-6 dB) cutoff
# (12 dB/octave roll-off)."
#
# <div class="alert alert-info"><h4>Note</h4><p>This simulated signal contains energy not just within the
# pass-band, but also within the transition and stop-bands -- perhaps
# most easily understood because the signal has a non-zero DC value,
# but also because it is a shifted cosine that has been
# *windowed* (here multiplied by a rectangular window), which
# makes the cosine and DC frequencies spread to other frequencies
# (multiplication in time is convolution in frequency, so multiplying
# by a rectangular window in the time domain means convolving a sinc
# function with the impulses at DC and the cosine frequency in the
# frequency domain).</p></div>
#
#
#
# +
x = np.zeros(int(2 * sfreq))
t = np.arange(0, len(x)) / sfreq - 0.2
onset = np.where(t >= 0.5)[0][0]
cos_t = np.arange(0, int(sfreq * 0.8)) / sfreq
sig = 2.5 - 2.5 * np.cos(2 * np.pi * (1. / 0.8) * cos_t)
x[onset:onset + len(sig)] = sig
iir_lp_30 = signal.iirfilter(2, 30. / sfreq, btype='lowpass')
iir_hp_p1 = signal.iirfilter(2, 0.1 / sfreq, btype='highpass')
iir_lp_2 = signal.iirfilter(2, 2. / sfreq, btype='lowpass')
iir_hp_2 = signal.iirfilter(2, 2. / sfreq, btype='highpass')
x_lp_30 = signal.filtfilt(iir_lp_30[0], iir_lp_30[1], x, padlen=0)
x_hp_p1 = signal.filtfilt(iir_hp_p1[0], iir_hp_p1[1], x, padlen=0)
x_lp_2 = signal.filtfilt(iir_lp_2[0], iir_lp_2[1], x, padlen=0)
x_hp_2 = signal.filtfilt(iir_hp_2[0], iir_hp_2[1], x, padlen=0)
xlim = t[[0, -1]]
ylim = [-2, 6]
xlabel = 'Time (sec)'
ylabel = 'Amplitude ($\mu$V)'
tticks = [0, 0.5, 1.3, t[-1]]
axes = plt.subplots(2, 2)[1].ravel()
for ax, x_f, title in zip(axes, [x_lp_2, x_lp_30, x_hp_2, x_hp_p1],
['LP$_2$', 'LP$_{30}$', 'HP$_2$', 'LP$_{0.1}$']):
ax.plot(t, x, color='0.5')
ax.plot(t, x_f, color='k', linestyle='--')
ax.set(ylim=ylim, xlim=xlim, xticks=tticks,
title=title, xlabel=xlabel, ylabel=ylabel)
mne.viz.adjust_axes(axes)
mne.viz.tight_layout()
plt.show()
# -
# Similarly, in a P300 paradigm reported by Kappenman & Luck 2010 [12]_,
# they found that applying a 1 Hz high-pass decreased the probaility of
# finding a significant difference in the N100 response, likely because
# the P300 response was smeared (and inverted) in time by the high-pass
# filter such that it tended to cancel out the increased N100. However,
# they nonetheless note that some high-passing can still be useful to deal
# with drifts in the data.
#
# Even though these papers generally advise a 0.1 HZ or lower frequency for
# a high-pass, it is important to keep in mind (as most authors note) that
# filtering choices should depend on the frequency content of both the
# signal(s) of interest and the noise to be suppressed. For example, in
# some of the MNE-Python examples involving `ch_sample_data`,
# high-pass values of around 1 Hz are used when looking at auditory
# or visual N100 responses, because we analyze standard (not deviant) trials
# and thus expect that contamination by later or slower components will
# be limited.
#
# Baseline problems (or solutions?)
# ---------------------------------
#
# In an evolving discussion, Tanner *et al.* 2015 [8]_ suggest using baseline
# correction to remove slow drifts in data. However, Maess *et al.* 2016 [9]_
# suggest that baseline correction, which is a form of high-passing, does
# not offer substantial advantages over standard high-pass filtering.
# Tanner *et al.* [10]_ rebutted that baseline correction can correct for
# problems with filtering.
#
# To see what they mean, consider again our old simulated signal ``x`` from
# before:
#
#
# +
def baseline_plot(x):
all_axes = plt.subplots(3, 2)[1]
for ri, (axes, freq) in enumerate(zip(all_axes, [0.1, 0.3, 0.5])):
for ci, ax in enumerate(axes):
if ci == 0:
iir_hp = signal.iirfilter(4, freq / sfreq, btype='highpass',
output='sos')
x_hp = sosfiltfilt(iir_hp, x, padlen=0)
else:
x_hp -= x_hp[t < 0].mean()
ax.plot(t, x, color='0.5')
ax.plot(t, x_hp, color='k', linestyle='--')
if ri == 0:
ax.set(title=('No ' if ci == 0 else '') +
'Baseline Correction')
ax.set(xticks=tticks, ylim=ylim, xlim=xlim, xlabel=xlabel)
ax.set_ylabel('%0.1f Hz' % freq, rotation=0,
horizontalalignment='right')
mne.viz.adjust_axes(axes)
mne.viz.tight_layout()
plt.suptitle(title)
plt.show()
baseline_plot(x)
# -
# In respose, Maess *et al.* 2016 [11]_ note that these simulations do not
# address cases of pre-stimulus activity that is shared across conditions, as
# applying baseline correction will effectively copy the topology outside the
# baseline period. We can see this if we give our signal ``x`` with some
# consistent pre-stimulus activity, which makes everything look bad.
#
# <div class="alert alert-info"><h4>Note</h4><p>An important thing to keep in mind with these plots is that they
# are for a single simulated sensor. In multielectrode recordings
# the topology (i.e., spatial pattiern) of the pre-stimulus activity
# will leak into the post-stimulus period. This will likely create a
# spatially varying distortion of the time-domain signals, as the
# averaged pre-stimulus spatial pattern gets subtracted from the
# sensor time courses.</p></div>
#
# Putting some activity in the baseline period:
#
#
n_pre = (t < 0).sum()
sig_pre = 1 - np.cos(2 * np.pi * np.arange(n_pre) / (0.5 * n_pre))
x[:n_pre] += sig_pre
baseline_plot(x)
# Both groups seem to acknowledge that the choices of filtering cutoffs, and
# perhaps even the application of baseline correction, depend on the
# characteristics of the data being investigated, especially when it comes to:
#
# 1. The frequency content of the underlying evoked activity relative
# to the filtering parameters.
# 2. The validity of the assumption of no consistent evoked activity
# in the baseline period.
#
# We thus recommend carefully applying baseline correction and/or high-pass
# values based on the characteristics of the data to be analyzed.
#
#
# Filtering defaults
# ==================
#
#
# Defaults in MNE-Python
# ----------------------
#
# Most often, filtering in MNE-Python is done at the :class:`mne.io.Raw` level,
# and thus :func:`mne.io.Raw.filter` is used. This function under the hood
# (among other things) calls :func:`mne.filter.filter_data` to actually
# filter the data, which by default applies a zero-phase FIR filter designed
# using :func:`scipy.signal.firwin`. In Widmann *et al.* 2015 [7]_, they
# suggest a specific set of parameters to use for high-pass filtering,
# including:
#
# "... providing a transition bandwidth of 25% of the lower passband
# edge but, where possible, not lower than 2 Hz and otherwise the
# distance from the passband edge to the critical frequency.”
#
# In practice, this means that for each high-pass value ``l_freq`` or
# low-pass value ``h_freq`` below, you would get this corresponding
# ``l_trans_bandwidth`` or ``h_trans_bandwidth``, respectively,
# if the sample rate were 100 Hz (i.e., Nyquist frequency of 50 Hz):
#
# +------------------+-------------------+-------------------+
# | l_freq or h_freq | l_trans_bandwidth | h_trans_bandwidth |
# +==================+===================+===================+
# | 0.01 | 0.01 | 2.0 |
# +------------------+-------------------+-------------------+
# | 0.1 | 0.1 | 2.0 |
# +------------------+-------------------+-------------------+
# | 1.0 | 1.0 | 2.0 |
# +------------------+-------------------+-------------------+
# | 2.0 | 2.0 | 2.0 |
# +------------------+-------------------+-------------------+
# | 4.0 | 2.0 | 2.0 |
# +------------------+-------------------+-------------------+
# | 8.0 | 2.0 | 2.0 |
# +------------------+-------------------+-------------------+
# | 10.0 | 2.5 | 2.5 |
# +------------------+-------------------+-------------------+
# | 20.0 | 5.0 | 5.0 |
# +------------------+-------------------+-------------------+
# | 40.0 | 10.0 | 10.0 |
# +------------------+-------------------+-------------------+
# | 45.0 | 11.25 | 5.0 |
# +------------------+-------------------+-------------------+
# | 48.0 | 12.0 | 2.0 |
# +------------------+-------------------+-------------------+
#
# MNE-Python has adopted this definition for its high-pass (and low-pass)
# transition bandwidth choices when using ``l_trans_bandwidth='auto'`` and
# ``h_trans_bandwidth='auto'``.
#
# To choose the filter length automatically with ``filter_length='auto'``,
# the reciprocal of the shortest transition bandwidth is used to ensure
# decent attenuation at the stop frequency. Specifically, the reciprocal
# (in samples) is multiplied by 3.1, 3.3, or 5.0 for the Hann, Hamming,
# or Blackman windows, respectively as selected by the ``fir_window``
# argument for ``fir_design='firwin'``, and double these for
# ``fir_design='firwin2'`` mode.
#
# <div class="alert alert-info"><h4>Note</h4><p>For ``fir_design='firwin2'``, the multiplicative factors are
# doubled compared to what is given in Ifeachor and Jervis [2]_
# (p. 357), as :func:`scipy.signal.firwin2` has a smearing effect
# on the frequency response, which we compensate for by
# increasing the filter length. This is why
# ``fir_desgin='firwin'`` is preferred to ``fir_design='firwin2'``.</p></div>
#
# In 0.14, we default to using a Hamming window in filter design, as it
# provides up to 53 dB of stop-band attenuation with small pass-band ripple.
#
# <div class="alert alert-info"><h4>Note</h4><p>In band-pass applications, often a low-pass filter can operate
# effectively with fewer samples than the high-pass filter, so
# it is advisable to apply the high-pass and low-pass separately
# when using ``fir_design='firwin2'``. For design mode
# ``fir_design='firwin'``, there is no need to separate the
# operations, as the lowpass and highpass elements are constructed
# separately to meet the transition band requirements.</p></div>
#
# For more information on how to use the
# MNE-Python filtering functions with real data, consult the preprocessing
# tutorial on `tut_artifacts_filter`.
#
# Defaults in MNE-C
# -----------------
# MNE-C by default uses:
#
# 1. 5 Hz transition band for low-pass filters.
# 2. 3-sample transition band for high-pass filters.
# 3. Filter length of 8197 samples.
#
# The filter is designed in the frequency domain, creating a linear-phase
# filter such that the delay is compensated for as is done with the MNE-Python
# ``phase='zero'`` filtering option.
#
# Squared-cosine ramps are used in the transition regions. Because these
# are used in place of more gradual (e.g., linear) transitions,
# a given transition width will result in more temporal ringing but also more
# rapid attenuation than the same transition width in windowed FIR designs.
#
# The default filter length will generally have excellent attenuation
# but long ringing for the sample rates typically encountered in M-EEG data
# (e.g. 500-2000 Hz).
#
# Defaults in other software
# --------------------------
# A good but possibly outdated comparison of filtering in various software
# packages is available in [7]_. Briefly:
#
# * EEGLAB
# MNE-Python in 0.14 defaults to behavior very similar to that of EEGLAB,
# see the `EEGLAB filtering FAQ`_ for more information.
# * Fieldrip
# By default FieldTrip applies a forward-backward Butterworth IIR filter
# of order 4 (band-pass and band-stop filters) or 2 (for low-pass and
# high-pass filters). Similar filters can be achieved in MNE-Python when
# filtering with :meth:`raw.filter(..., method='iir') <mne.io.Raw.filter>`
# (see also :func:`mne.filter.construct_iir_filter` for options).
# For more inforamtion, see e.g. `FieldTrip band-pass documentation`_.
#
# Summary
# =======
#
# When filtering, there are always tradeoffs that should be considered.
# One important tradeoff is between time-domain characteristics (like ringing)
# and frequency-domain attenuation characteristics (like effective transition
# bandwidth). Filters with sharp frequency cutoffs can produce outputs that
# ring for a long time when they operate on signals with frequency content
# in the transition band. In general, therefore, the wider a transition band
# that can be tolerated, the better behaved the filter will be in the time
# domain.
#
# References
# ==========
#
# .. [1] Parks TW, Burrus CS (1987). Digital Filter Design.
# New York: Wiley-Interscience.
# .. [2] <NAME>., & <NAME>. (2002). Digital Signal Processing:
# A Practical Approach. Prentice Hall.
# .. [3] <NAME>. (2011). Four common conceptual fallacies in mapping
# the time course of recognition. Perception Science, 2, 365.
# .. [4] <NAME>., <NAME>., & <NAME>, <NAME>. (2012).
# Systematic biases in early ERP and ERF components as a result
# of high-pass filtering. Journal of Neuroscience Methods,
# 209(1), 212–218. http://doi.org/10.1016/j.jneumeth.2012.06.011
# .. [5] <NAME>. (2012). Does filtering preclude us from studying
# ERP time-courses? Frontiers in Psychology, 3(131)
# .. [6] <NAME>., & <NAME>. (2012). Filter effects and filter
# artifacts in the analysis of electrophysiological data.
# Perception Science, 233.
# .. [7] <NAME>., <NAME>., & <NAME>. (2015). Digital filter
# design for electrophysiological data – a practical approach.
# Journal of Neuroscience Methods, 250, 34–46.
# .. [8] <NAME>., <NAME>., & <NAME>. (2015).
# How inappropriate high-pass filters can produce artifactual effects
# and incorrect conclusions in ERP studies of language and cognition.
# Psychophysiology, 52(8), 997–1009. http://doi.org/10.1111/psyp.12437
# .. [9] <NAME>., <NAME>., & <NAME>. (2016).
# High-pass filters and baseline correction in M/EEG analysis.
# Commentary on: “How inappropriate high-pass filters can produce
# artefacts and incorrect conclusions in ERP studies of language
# and cognition.” Journal of Neuroscience Methods, 266, 164–165.
# .. [10] <NAME>., <NAME>., <NAME>., & <NAME>. (2016).
# On high-pass filter artifacts (they’re real) and baseline correction
# (it’s a good idea) in ERP/ERMF analysis.
# .. [11] <NAME>., <NAME>., & <NAME>. (2016).
# High-pass filters and baseline correction in M/EEG analysis-continued
# discussion. Journal of Neuroscience Methods, 266, 171–172.
# Journal of Neuroscience Methods, 266, 166–170.
# .. [12] <NAME>. & <NAME>. (2010). The effects of impedance on data
# quality and statistical significance in ERP recordings.
# Psychophysiology, 47, 888-904.
#
#
#
| 0.15/_downloads/plot_background_filtering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## RNN Overview
#
# <img src="http://colah.github.io/posts/2015-08-Understanding-LSTMs/img/RNN-unrolled.png" alt="nn" style="width: 600px;"/>
#
# References:
# - [Long Short Term Memory](http://deeplearning.cs.cmu.edu/pdfs/Hochreiter97_lstm.pdf), <NAME> & <NAME>, Neural Computation 9(8): 1735-1780, 1997.
# +
from __future__ import absolute_import, division, print_function
# Import TensorFlow v2.
import tensorflow as tf
from tensorflow.keras import Model, layers
import numpy as np
import random
# +
# Dataset parameters.
num_classes = 2 # linear sequence or not.
seq_max_len = 20 # Maximum sequence length.
seq_min_len = 5 # Minimum sequence length (before padding).
masking_val = -1 # -1 will represents the mask and be used to pad sequences to a common max length.
max_value = 10000 # Maximum int value.
# Training Parameters
learning_rate = 0.001
training_steps = 2000
batch_size = 64
display_step = 100
# Network Parameters
num_units = 32 # number of neurons for the LSTM layer.
# +
# ====================
# TOY DATA GENERATOR
# ====================
def toy_sequence_data():
""" Generate sequence of data with dynamic length.
This function generates toy samples for training:
- Class 0: linear sequences (i.e. [1, 2, 3, 4, ...])
- Class 1: random sequences (i.e. [9, 3, 10, 7,...])
NOTICE:
We have to pad each sequence to reach 'seq_max_len' for TensorFlow
consistency (we cannot feed a numpy array with inconsistent
dimensions). The dynamic calculation will then be perform and ignore
the masked value (here -1).
"""
while True:
# Set variable sequence length.
seq_len = random.randint(seq_min_len, seq_max_len)
rand_start = random.randint(0, max_value - seq_len)
# Add a random or linear int sequence (50% prob).
if random.random() < .5:
# Generate a linear sequence.
seq = np.arange(start=rand_start, stop=rand_start+seq_len)
# Rescale values to [0., 1.].
seq = seq / max_value
# Pad sequence until the maximum length for dimension consistency.
# Masking value: -1.
seq = np.pad(seq, mode='constant', pad_width=(0, seq_max_len-seq_len), constant_values=masking_val)
label = 0
else:
# Generate a random sequence.
seq = np.random.randint(max_value, size=seq_len)
# Rescale values to [0., 1.].
seq = seq / max_value
# Pad sequence until the maximum length for dimension consistency.
# Masking value: -1.
seq = np.pad(seq, mode='constant', pad_width=(0, seq_max_len-seq_len), constant_values=masking_val)
label = 1
yield np.array(seq, dtype=np.float32), np.array(label, dtype=np.float32)
# -
# Use tf.data API to shuffle and batch data.
train_data = tf.data.Dataset.from_generator(toy_sequence_data, output_types=(tf.float32, tf.float32))
train_data = train_data.repeat().shuffle(5000).batch(batch_size).prefetch(1)
# +
# Create LSTM Model.
class LSTM(Model):
# Set layers.
def __init__(self):
super(LSTM, self).__init__()
# Define a Masking Layer with -1 as mask.
self.masking = layers.Masking(mask_value=masking_val)
# Define a LSTM layer to be applied over the Masking layer.
# Dynamic computation will automatically be performed to ignore -1 values.
self.lstm = layers.LSTM(units=num_units)
# Output fully connected layer (2 classes: linear or random seq).
self.out = layers.Dense(num_classes)
# Set forward pass.
def call(self, x, is_training=False):
# A RNN Layer expects a 3-dim input (batch_size, seq_len, num_features).
x = tf.reshape(x, shape=[-1, seq_max_len, 1])
# Apply Masking layer.
x = self.masking(x)
# Apply LSTM layer.
x = self.lstm(x)
# Apply output layer.
x = self.out(x)
if not is_training:
# tf cross entropy expect logits without softmax, so only
# apply softmax when not training.
x = tf.nn.softmax(x)
return x
# Build LSTM model.
lstm_net = LSTM()
# +
# Cross-Entropy Loss.
# Note that this will apply 'softmax' to the logits.
def cross_entropy_loss(x, y):
# Convert labels to int 64 for tf cross-entropy function.
y = tf.cast(y, tf.int64)
# Apply softmax to logits and compute cross-entropy.
loss = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=x)
# Average loss across the batch.
return tf.reduce_mean(loss)
# Accuracy metric.
def accuracy(y_pred, y_true):
# Predicted class is the index of highest score in prediction vector (i.e. argmax).
correct_prediction = tf.equal(tf.argmax(y_pred, 1), tf.cast(y_true, tf.int64))
return tf.reduce_mean(tf.cast(correct_prediction, tf.float32), axis=-1)
# Adam optimizer.
optimizer = tf.optimizers.Adam(learning_rate)
# -
# Optimization process.
def run_optimization(x, y):
# Wrap computation inside a GradientTape for automatic differentiation.
with tf.GradientTape() as g:
# Forward pass.
pred = lstm_net(x, is_training=True)
# Compute loss.
loss = cross_entropy_loss(pred, y)
# Variables to update, i.e. trainable variables.
trainable_variables = lstm_net.trainable_variables
# Compute gradients.
gradients = g.gradient(loss, trainable_variables)
# Update weights following gradients.
optimizer.apply_gradients(zip(gradients, trainable_variables))
# Run training for the given number of steps.
for step, (batch_x, batch_y) in enumerate(train_data.take(training_steps), 1):
# Run the optimization to update W and b values.
run_optimization(batch_x, batch_y)
if step % display_step == 0 or step == 1:
pred = lstm_net(batch_x, is_training=True)
loss = cross_entropy_loss(pred, batch_y)
acc = accuracy(pred, batch_y)
print("step: %i, loss: %f, accuracy: %f" % (step, loss, acc))
| Tutorials/TensorFlow_V2/notebooks/3_NeuralNetworks/dynamic_rnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Indicator Tutorial
#
# The purpose of this spreadsheet is to demonstrate the custom indicators that pinkfish provides.
# + language="javascript"
# IPython.OutputArea.prototype._should_scroll = function(lines) {
# return false;
# }
# +
import datetime
import pandas as pd
import pinkfish as pf
# Format price data
pd.options.display.float_format = '{:0.3f}'.format
# -
# Some global data
symbol = 'SPY'
start = datetime.datetime(2000, 1, 1)
end = datetime.datetime(2020, 1, 1)
# Fetch symbol data from cache, if available.
ts = pf.fetch_timeseries(symbol)
ts.tail()
# Select timeseries between start and end.
ts = pf.select_tradeperiod(ts, start, end)
ts.head()
# ### CROSSOVER Indicator
print(pf.CROSSOVER.__doc__)
# We see that SPY has been in a bull market regime for nearly 200 days.
ts['regime'] = pf.CROSSOVER(ts, timeperiod_fast=50, timeperiod_slow=200)
ts.tail(10)
# ### MOMENTUM Indicator
print(pf.MOMENTUM.__doc__)
# We see that SPY has positive momentum. Over the last 6 months
# the price has increased by about 8%.
ts['mom'] = pf.MOMENTUM(ts, lookback=6, time_frame='monthly')
ts.tail(10)
# ### VOLATILITY Indicator
print(pf.VOLATILITY.__doc__)
# We compute the annualized 20 days volatility. Notice that the
# volatility is decreasing.
ts['vola'] = pf.VOLATILITY(ts, lookback=20, time_frame='yearly')
ts.tail(10)
# ### ANNUALIZED_RETURNS Indicator
print(pf.ANNUALIZED_RETURNS.__doc__)
# The 3 month annualized returns are about 18%.
ts['rets'] = pf.ANNUALIZED_RETURNS(ts, lookback=3/12)
ts.tail(10)
# ## ANNUALIZED_STANDARD_DEVIATION Indicator
print(pf.ANNUALIZED_STANDARD_DEVIATION.__doc__)
# The 3 year annualized standard deviation is about 13%.
ts['std_dev'] = pf.ANNUALIZED_STANDARD_DEVIATION(ts, lookback=3)
ts.tail(10)
# ## ANNUALIZED_SHARPE_RATIO Indicator
print(pf.ANNUALIZED_SHARPE_RATIO.__doc__)
# The 3 year annualized sharpe ratio is about 1.7.
ts['sharpe_ratio'] = pf.ANNUALIZED_SHARPE_RATIO(ts, lookback=3)
ts.tail(10)
# ## COMBINATION of INDICATORS
#
# Here's how to combine 2 indicators to create a new one.
# SMA_MOMENTUM = SMA(MOMENTUM)
#
#
# We first calculate the MOM, then apply a SMA to it.
# +
from talib.abstract import *
def SMA_MOMENTUM(ts, mom_lookback=1, sma_timeperiod=20, price='close'):
""" Returns a series which is an SMA with of a daily MOM. """
mom = pf.MOMENTUM(ts, lookback=mom_lookback, time_frame='daily', price=price)
sma_mom = SMA(mom, timeperiod=sma_timeperiod)
return sma_mom
# -
# We see that SPY has positive sma momentum. Over the last 20 days
# the average 5 day momenteum (averaged over 20 samples) has been positive
# at about 0.7%.
ts['sma_mom'] = SMA_MOMENTUM(ts, mom_lookback=5, sma_timeperiod=20)
ts.tail(10)
| examples/071.indicator-tutorial/indicator-tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab inline
import numpy as np
import math
# Condiciones iniciales del problema.
q=-1.602176565E-19
v_0=3.0E5
theta_0=0
B=-10**(-4)
m=9.1093829E-31
N=1000
# Cálculo del tiempo que tarda la partícula en volver a cruzar el eje $x$ teóricamente t definición del intervalo de tiempo a observar.
def tiempo(q,B,theta_0, m,N):
t_max=(m/(q*B))*(2*theta_0+math.pi)
dt=t_max/N
t=np.arange(0,t_max+dt,dt)
return t, t_max
time,t_max=tiempo(q,B,theta_0, m,N)
print("El tiempo total del recorrido teóricamente hasta llegar al detector es {}.".format(t_max))
# Ecuaciones de posición teóricas respecto al tiempo $t$ para $\theta_0$ arbitrario y na velocidad $v_0$ de entrada.
def posicion(q,B,v_0,theta_0,m,t):
omega=q*B/m
x=-v_0*np.cos(theta_0-omega*t)/omega+v_0*np.cos(theta_0)/omega
y=-v_0*np.sin(theta_0-omega*t)/omega+v_0*np.sin(theta_0)/omega
return x,y
# Gráfica del recorrido circular de una partícula de carga $q$ que incide en el eje $x$ con rapidez $v_0$ y ángulo de incidencia $\theta_0=0,$debido a un campo $\mathbf{B}$ perpendicular.
plt.figure(figsize=(50/6,5))
xTeo,yTeo=posicion(q,B,v_0,theta_0,m,time)
plt.plot(xTeo,yTeo, label="Trayectoria circular")
plt.plot(xTeo,np.zeros(len(xTeo)), c="black")
plt.legend()
plt.xlabel("Posición en x (m)")
plt.ylabel("Posición en y (m)")
plt.savefig("recorridoTeorico.jpg")
# Cálculo de la posición final de la partícula al llegar al detector, es decir el punto en el que la trayectoria cruza el eje $x$ nuevamente.
x_max, y_max=posicion(q,B,v_0,theta_0,m,t_max)
print("Teóricamente a partícula alcanza el detector cuando este se encuentra en x={}m y y={}m.".format(x_max,y_max))
# Cálculo del momento inicial $p_0=mv_0$, final $p_f=\frac{1}{2}qBx$ y la diferencia de momento que comprueba la conservación del momento lineal.
p_0=m*v_0
p_f=0.5*q*B*x_max
dp=np.abs(p_f-p_0)
print("El momento inicial de la partícula es {} kg m/s, el momento final es {} kg m/s y la diferencia de momento es {} kg m/s.".format(p_0,p_f,dp))
# Definición de la función de trayectoria de la partícula que incide con rapidez $v_0$ y ángulo $\theta_0$ a una región de campo magnético erpendicular $\mathbf{B}$; siguiendo el paso a paso de Feynmann.
def pasoApaso(q,B,v_0,theta_0,m,N):
t=0.0
omega=q*B/m
dt=1/(omega*N)
x=[0]
y=[0]
v_x=-v_0*np.sin(theta_0)
v_y=v_0*np.cos(theta_0)
while y[-1]>=0:
a_x=omega*v_y
a_y=-omega*v_x
x_new=x[-1]+v_x*dt
y_new=y[-1]+v_y*dt
x.append(x_new)
y.append(y_new)
v_x=v_x+a_x*dt
v_y=v_y+a_y*dt
t=t+dt
x=np.array(x)
y=np.array(y)
return x,y,t
# Gráfica de la trayectoria circular por medio del Método de Feynmann de una partícula de carga $q$ que incide en el eje $x$ con rapidez $v_0$ y ángulo de incidencia $\theta_0=0,$debido a un campo $\mathbf{B}$ perpendicular.
plt.figure(figsize=(50/6,5))
xF,yF,t_maxF=pasoApaso(q,B,v_0,theta_0,m,N)
plt.plot(xF,yF, label="Trayectoria circular")
plt.plot(xF,np.zeros(len(xF)), c="black")
plt.legend()
plt.xlabel("Posición en x (m)")
plt.ylabel("Posición en y (m)")
# Cálculo numérico de la posición final de la partícula al llegar al detector.
xF_max=xF[-1]
yF_max=yF[-1]
print("Mediante el Método de Feynmann la partícula alcanza el detector cuando este se encuentra en x={}m y y={}m.".format(xF_max,yF_max))
# Cálculo del momento inicial $p_0=mv_0$, final $p_f=\frac{1}{2}qBx$ y la diferencia de momento que comprueba la conservación del momento lineal.
pF_0=m*v_0
pF_f=0.5*q*B*xF_max
dpF=np.abs(pF_f-pF_0)
print("El momento inicial de la partícula es {} kg m/s, el momento final es {} kg m/s y la diferencia de momento es {} kg m/s.".format(pF_0,pF_f,dpF))
# Definición de cambio en la velocidad y la función de trayectoria de la partícula que incide con rapidez $v_0$ y ángulo $\theta_0$ a una región de campo magnético erpendicular $\mathbf{B}$; siguiendo el paso a paso del Método de Runge Kutta de cuarto orden.
# +
def delta(omega,v_x,v_y,dt):
delta11=dt*omega*v_y
delta12=dt*omega*(v_y+delta11/2)
delta13=dt*omega*(v_y+delta12/2)
delta14=dt*omega*(v_y+delta13)
delta1=(delta11+2*delta12+2*delta13+delta14)/6
delta21=-dt*omega*v_x
delta22=-dt*omega*(v_x+delta21/2)
delta23=-dt*omega*(v_x+delta22/2)
delta24=-dt*omega*(v_x+delta23)
delta2=(delta21+2*delta22+2*delta23+delta24)/6
return delta1, delta2
def rungePaso(q,B,v_0,theta_0,m,N):
t=0.0
omega=q*B/m
dt=1/(omega*N)
x=[0]
y=[0]
v_x=-v_0*np.sin(theta_0)
v_y=v_0*np.cos(theta_0)
while y[-1]>=0:
x_new=x[-1]+v_x*dt
y_new=y[-1]+v_y*dt
x.append(x_new)
y.append(y_new)
v_x=v_x+delta(omega,v_x,v_y,dt)[0]
v_y=v_y+delta(omega,v_x,v_y,dt)[1]
t=t+dt
x=np.array(x)
y=np.array(y)
return x,y,t
# -
# Gráfica de la trayectoria circular por medio del Método de Runge Kutta 4 de una partícula de carga $q$ que incide en el eje $x$ con rapidez $v_0$ y ángulo de incidencia $\theta_0=0,$debido a un campo $\mathbf{B}$ perpendicular.
plt.figure(figsize=(50/6,5))
xR,yR,t_maxR=rungePaso(q,B,v_0,theta_0,m,N)
plt.plot(xR,yR, label="Trayectoria circular")
plt.plot(xR,np.zeros(len(xR)), c="black")
plt.legend()
plt.xlabel("Posición en x (m)")
plt.ylabel("Posición en y (m)")
# Cálculo numérico de la posición final de la partícula al llegar al detector.
xR_max=xR[-1]
yR_max=yR[-1]
print("Mediante el Método de Runge Kutta 4 la partícula alcanza el detector cuando este se encuentra en x={}m y y={}m.".format(xR_max,yR_max))
# Cálculo del momento inicial $p_0=mv_0$, final $p_f=\frac{1}{2}qBx$ y la diferencia de momento que comprueba la conservación del momento lineal.
pR_0=m*v_0
pR_f=0.5*q*B*xR_max
dpR=np.abs(pR_f-pR_0)
print("El momento inicial de la partícula es {} kg m/s, el momento final es {} kg m/s y la diferencia de momento es {} kg m/s.".format(pR_0,pR_f,dpR))
# Comparación gráfica de los métodos numéricos para $\theta_0=0$.
plt.figure(figsize=(14,4.6))
plt.title("Recorridos de partículas con carga {}C, masa {}kg, velocidad {}m/s, ángulo de entrada {}rad, debidas a un campo perpendicular B={}T.".format(q,m,v_0,theta_0,B))
plt.subplot(1,2,1)
plt.plot(xF,yF, label="PasoFeynmann")
plt.legend()
plt.xlabel("Posición x (m)")
plt.xlabel("Posición y (m)")
plt.subplot(1,2,2)
plt.plot(xR,yR, label="PasoRunge",c="orange")
plt.legend()
plt.xlabel("Posición en x (m)")
plt.xlabel("Posición en y (m)")
plt.savefig("recorridos.jpg")
# Comparación del error en el cálculo del momento final $Error=\frac{|p_f-p_0|}{p_0}*100%$ de las partículas mediante el método de Feynmann y de Runge Kutta 4 para $N=1000$.
errorF=dpF*100/p_0
errorR=dpR*100/p_0
print("El error porcentual en la conservación para el método de Feynmann fue de {}%, mientras que para el método de Runge Kutta 4 fue de {}%".format(errorF,errorR))
# Comportameinto del momento final para los métodos de Feynmann y Runge Kutta 4 a medida que aumenta $N$
N=np.array([100,200,500,1000,1500])
def momentoVsN(q,B,v_0,theta_0,m,N):
xF=np.zeros(len(N))
xR=np.zeros(len(N))
for i in range(len(N)):
xF[i]=pasoApaso(q,B,v_0,theta_0,m,N[i])[0][-1]
xR[i]=rungePaso(q,B,v_0,theta_0,m,N[i])[0][-1]
pF=0.5*q*B*xF
pR=0.5*q*B*xR
p0=m*v_0
errF=np.abs(pF-p0)*100/p0
errR=np.abs(pR-p0)*100/p0
return pF,pR,errF,errR
pF,pR,errF,errR=momentoVsN(q,B,v_0,theta_0,m,N)
print("Para los N={} los momentos finales obtenidos fueron: para el Método de Feynmann {} y para el Método de Runge Kutta 4 {}, respectivamente a cada N. Con error en el cálculo respectivo de {} para el método de Feynmann y de {} para el método de Runge Kutta 4.".format(N,pF,pR,errF,errR))
# Gráficas del comportamiento del momento final de las partículas al llegar al detector y el error en términos del momento de los métodos numéricos.
plt.figure(figsize=(14,5))
plt.subplot(1,2,1)
plt.plot(N,pF, label="Método de Feymann")
plt.plot(N,pR, label="Método de Runge Kutta 4")
#plt.title("Momento final de las partículas en función del número de repeticiones N")
plt.xlabel("Número de divisiones en el tiempo N")
plt.ylabel("Momento final (kg m/s)")
plt.legend()
plt.subplot(1,2,2)
plt.plot(N,errF, label="Método de Feymann")
plt.plot(N,errR, label="Método de Runge Kutta 4")
#plt.title("Error en el momento de las partículas en función del número de repeticiones N")
plt.xlabel("Número de divisiones en el tiempo N")
plt.ylabel("Error en el momento (%)")
plt.legend()
plt.savefig("analisisMomentosN.jpg")
# Comportamiento teórico del alcance máximo o posición en x final de la partícula con respecto a $\theta_0.$
def xFinalVsTheta(q,B,theta0, m,N):
xFinal=np.zeros(len(theta0))
for i in range(len(theta0)):
tmax=tiempo(q,B,theta0[i], m,N)[1]
xFinal[i]=posicion(q,B,v_0,theta0[i],m,tmax)[0]
return xFinal
# Gráfica de la posición en x final en función del ángulo de incidencia $\theta_0$.
N=1000
plt.figure()
theta0=np.arange(-np.pi/3,np.pi/3,2*np.pi/N)
xFinal=xFinalVsTheta(q,B,theta0, m,N)
plt.plot(theta0,xFinal)
plt.xlabel("Ángulo de incidencia "+ r"$\theta_0(rad)$")
plt.ylabel("Posición final en x (m)")
# Trayectorias de enfoque con Runge Kutta 4 y $N=1000$ para los ángulos $\theta_0=0$, $\theta_0=\frac{\pi}{8}$ y $\theta_0=-\frac{\pi}{8}$
plt.figure(figsize=(50/6,5))
theta=[0,-np.pi/8,np.pi/8]
xR0,yR0,t_maxR0=rungePaso(q,B,v_0,theta[0],m,N)
xR1,yR1,t_maxR1=rungePaso(q,B,v_0,theta[1],m,N)
xR2,yR2,t_maxR2=rungePaso(q,B,v_0,theta[2],m,N)
plt.plot(xR0,yR0, label="Trayectoria para "+r"$\theta_0={} rad$".format(theta[0]))
plt.plot(xR1,yR1, label="Trayectoria para "+r"$\theta_0=-\frac{\pi}{8} rad$")
plt.plot(xR2,yR2, label="Trayectoria para "+r"$\theta_0=\frac{\pi}{8} rad $")
plt.plot(xR0,np.zeros(len(xR0)), c="black")
plt.legend()
plt.xlabel("Posición en x (m)")
plt.ylabel("Posición en y (m)")
plt.savefig("propiedadEnfoque.jpg")
# Definición de la función de alcance en x final en función de $\theta_0$ para el Método de Runge Kutta 4 con $N=1000$
def xFinalVsThetaRK4(q,B,theta0, m,N):
xFinal=np.zeros(len(theta0))
for i in range(len(theta0)):
xFinal[i]=rungePaso(q,B,v_0,theta0[i],m,N)[0][-1]
return xFinal
# Gráfica de la posición en x final en función del ángulo de incidencia $\theta_0$ con el Método Runge Kutta 4.
plt.figure()
theta0=np.arange(-np.pi/3,np.pi/3,2*np.pi/N)
xFinalRK4=xFinalVsThetaRK4(q,B,theta0, m,N)
plt.plot(theta0,xFinalRK4)
plt.xlabel("Ángulo de incidencia "+ r"$\theta_0(rad)$")
plt.ylabel("Posición final en x (m)")
plt.savefig("xFinalVSTheta0.jpg")
# Datos para algunos ángulos de incidencia para ver la igualdad en la propiedad de enfoque
thetaPos=np.arange(0,1,0.1)
xFinalPos=xFinalVsThetaRK4(q,B,thetaPos, m,N)
thetaNeg=-thetaPos
xFinalNeg=xFinalVsThetaRK4(q,B,thetaNeg, m,N)
print(xFinalPos,xFinalNeg)
print(thetaPos,thetaNeg)
diff=np.zeros(len(xFinalPos))
for i in range(len(xFinalPos)):
diff[i]=np.abs(xFinalPos[i]-xFinalNeg[i])
diff
| TrayectoriaParticulas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Title
# MCMC from Scratch for Linear Regression
#
# ## Description :
# The aim of this exercise is to perform Monte Carlo Markov Chain (MCMC) from scratch for linear regression. For this, we will be using our old friend the Advertising dataset.
#
# On completing the exercise you should be able to see the following distribution. One for each of the beta value:
#
# <img src="../fig/fig.png" style="width: 500px;">
#
# ## Instructions:
# - Read the data file Advertising.csv and set the predictor and response variables.
# - Fit a linear regression model on the advertising data and take a look at the beta values.
# - Create 2 lists to store the beta values and initialize the beta values.
# - Define a function get_prior to compute the prior value given the beta values.
# - Compute the likelihood, prior and posterior for the initial beta values.
# - For a selected number of sampling "epochs":
# - Compute new beta values
# - Compute the corresponding likelihood, prior and posterior.
# - Compute the exponential ratio of the current and previous posterior.
# - Based on the ratio, select or reject the new beta values.
# - Choose a burn rate.
# - Plot the histogram of the beta values.
#
# ## Hints:
#
# <a href="https://numpy.org/doc/stable/reference/generated/numpy.log.html" target="_blank">np.log()</a> Computes the natural logarithm, element-wise.
#
# <a href="https://numpy.org/doc/stable/reference/generated/numpy.exp.html?highlight=exp#numpy.exp" target="_blank">np.exp()</a> Calculates the exponential of all elements in the input array.
#
# <a href="https://www.google.com/search?q=sklearn+linear+gressiogn&rlz=1C5CHFA_enIN777IN777&oq=sklearn+linear+gressiogn&aqs=chrome..69i57j69i59l2j0i271j69i60.3137j0j7&sourceid=chrome&ie=UTF-8" target="_blank">LinearRegression()</a> Initiates an ordinary least squares Linear Regression.
#
# <a href="https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html?highlight=linear%20regression#sklearn.linear_model.LinearRegression.fit" target="_blank">.fit()</a> Fits the linear model to the data.
#
# <a href="https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html" target="_blank">model.coef_</a> Estimated coefficients for the linear regression problem
#
# <a href="https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html" target="_blank">model.intercept_</a> Independent term in the linear model.
#
# <a href="https://numpy.org/doc/stable/reference/random/generated/numpy.random.normal.html?highlight=random%20normal#numpy.random.normal" target="_blank">np.random.normal()</a> Draw random samples from a normal (Gaussian) distribution.
#
# <a href="https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.norm.html" target="_blank">norm.pdf()</a> A normal continuous random variable.
#
# <a href="https://numpy.org/doc/stable/reference/generated/numpy.sum.html?highlight=sum" target="_blank">np.sum()</a> Sum of array elements over a given axis.
#
# <a href="https://numpy.org/doc/stable/reference/random/generated/numpy.random.uniform.html?highlight=random%20uniform" target="_blank">np.random.uniform()</a> Draw samples from a uniform distribution.
# Import necessary libraries
from sklearn.linear_model import LinearRegression
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
# %matplotlib inline
from scipy import stats
from scipy.stats import norm
# +
# Read the data file 'Advertising.csv'
df = pd.read_csv("Advertising.csv")
# Use the column "tv" as the predictor
x = df[['tv']]
# Use the column "sales" as the response
y = df.sales.values
# -
# Take a quick look at the data
df.head()
# +
# Initiate a Linear Regression model
model = ___
# Fit the model on the predictor and response data
___
# -
# Take a quick look at the beta values got after model fitting
# Use the model.intercept_ and model.coef_ for this
b0 = ___
b1 = ___
print("Beta0 is",b0)
print("Beta1 is", b1)
# Helper code to plot the true and predicted data
plt.plot(x,y,'o', label="True Data", color='darkblue')
plt.plot(x,model.predict(df[['tv']]), label="Prediction", color='coral')
plt.xlabel("TV")
plt.ylabel("Sales")
plt.legend()
# Define 2 empty lists to store the accepted beta values in a list
beta0_list = []
beta1_list = []
# +
# Initialize beta0 to a resonable value based on the model parameter seen above
beta0 = ___
# Initialize beta1 to a resonable value based on the model parameter seen above
beta1 = ___
# -
# Function to get the prior given the beta0 and beta1 values
# NOTE - All the computations are done in the log space so that the numbers are managable.
def get_log_prior(beta0,beta1):
# The prior of beta0 is a value from a normal PDF of beta0 with mean as 100 and standard deviation as 50
# Take the log of these value
log_prior_b0 = ___
# The prior of beta1 is a value from a normal PDF of beta1 with mean as 1 and standard deviation as 1
# Take the log of this value
log_prior_b1 = ___
# Compute the prior as the sum of the log priors of beta0 and beta1
log_prior = ___
# Return the prior value
return log_prior
# +
# Compute the log-likelihood for the initial beta values
# pay attention to the dimensions of y and x.
log_likelihood = - np.sum( np.log( (y.reshape(-1,1) - np.array( beta1*x + beta0))**2) )
# Get the prior of the intial beta values by calling the get_log_prior function
log_prior = ___
# Compute the log posterior of the initial beta values
# The posterior is the sum of the log_likelihood and log_prior
log_posterior = ___
# +
# Save the initial posterior value as prev_posterior for comparision later
prev_logposterior = log_posterior
# Append the initial beta values i.e. beta0 and beta1 to the list
beta0_list.append(beta0)
beta1_list.append(beta1)
# -
# Specify the number of sampling "epochs" (less than 500k)
epochs = ___
# ### ⏸ How does the number of samples generated affect results of MCMC?
#
# #### A. As the number of samples are increased the beta values chosen grow increasing random.
# #### B. For a smaller number of samples the beta values are closer to the true value with reduced randomness.
# #### C. The number of samples does not affect the beta values, it only depends on the prior.
# #### D. As the number of samples increase, the beta values slowly converge to their true values.
### edTest(test_chow1) ###
# Submit an answer choice as a string below (eg. if you choose option C, put 'C')
answer1 = '___'
# Loop over the range of sampling "epochs"
for i in range(epochs):
# Get a new beta1 value with mean as the latest element beta1 and scale as 0.1
beta0 = ___
# Get a new beta0 value with mean as the latest element beta0 and scale as 0.5
beta1 = ___
# Get the prior values for the new beta values by calling the get_log_prior function
log_prior = ___
# Compute P(data|w) i.e. the log-likelihood for all the data points
log_likelihood = ___
# To compute the posterior given the likelihood and prior
# The posterior is the sum of the likelihood and prior
log_posterior = ___
# Compute the the exponential of the ratio of the posterior given its previous value
# Since it is the log, the ratio is computed as the difference between the values
exp_ratio = ___
# If the ratio is greater than 1 then accept the new beta values in this case
if exp_ratio>1:
# Append the beta0 and beta1 to the beta list values
beta0_list.append(beta0)
beta1_list.append(beta1)
# Save the accepted posterior as the previous posterior
prev_logposterior = log_posterior
# If the ratio is less than 1 then get a random value between 0 and 1
else:
coin = ___
# Set a threshold value
threshold = ___
# Check if the random value is higher than the threshold
# Append the beta values to the list and update the previous posterior
if coin > threshold:
beta0_list.append(beta0)
beta1_list.append(beta1)
prev_logposterior = log_posterior
# ### ⏸ If the threshold is set to a higher value, new beta values are rejected more often if they do not improve the convergence to the true value
#
# ### The statement above is:
#
# #### A. True for all cases
# #### B. False for all cases
# #### C. True only when the number of samples is less
# #### D. True only when prior is extremely far from the real value
### edTest(test_chow2) ###
# Submit an answer choice as a string below (eg. if you choose option C, put 'C')
answer2 = '___'
# The number of data points to consider after the beta list has been populated
burn_rate = int(len(beta0_list)*0.3)
### edTest(test_chow3) ###
# Check posterior mean for beta0 and beta1
print(np.mean(beta0_list[burn_rate:]), np.mean(beta1_list[burn_rate:]))
# +
# Helper code to plot the histogram of the beta values
# Plot the histogram of the beta0 values
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(15, 6))
ax1.hist(beta0_list[burn_rate:], color='#B2D7D0',edgecolor="black", linewidth=1)
ax1.set_xlabel("BETA 0", fontsize=14)
ax1.set_ylabel("FREQUENCY", fontsize=14);
# Plot the histogram of the beta1 values
ax2.hist(beta1_list[burn_rate:], color='#EFAEA4',edgecolor="black", linewidth=1)
ax2.set_xlabel("BETA 1", fontsize=14)
ax2.set_ylabel("FREQUENCY", fontsize=14);
| content/lectures/lecture27/notebook/MCMC_FromScratch_Scaffold.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8
# language: python
# name: python3.8
# ---
# # Event Prediction
# Observations come from 2 data streams (people flow in and out of the building), over 15 weeks, 48 time slices per day (half hour count aggregates). The purpose is to predict the presence of an event such as a conference in the building that is reflected by unusually high people counts for that day/time period.
#
#
# **Source**: https://archive.ics.uci.edu/ml/datasets/CalIt2+Building+People+Counts
# <img src="https://novotel.accor.com/imagerie/business-meeting-hotel/seminars-picture.jpg">
# ## Goals:
#
# ### Implement a predictive task
# - Define and prepare features to be used
# - Define the validation approach
# - Define the algorithms to use
# - Define the evaluation metric
# - Analyse the results and repeat the process
# +
import numpy as np
import pandas as pd
import seaborn as sns
from matplotlib import pyplot as plt
# %matplotlib inline
# -
pd.set_option("display.max_columns", None)
# +
dodgers_events_cols = ["date", "begin_event_time", "end_event_time", "game_attendance", "away team", "win_lose_score"]
dodgers_counts_cols = ["datetime", "count"]
dodgers_counts = pd.read_csv("data/Dodgers.data", header=None, names=dodgers_counts_cols)
dodgers_events = pd.read_csv("data/Dodgers.events", header=None, names=dodgers_events_cols)
# +
calit2_events_cols = ["date", "begin_event_time", "end_event_time", "event_name"]
calit2_counts_cols = ["date", "time", "count"]
calit2_counts = pd.read_csv("data/CalIt2.data", header=None, names=calit2_counts_cols)
calit2_events = pd.read_csv("data/CalIt2.events", header=None, names=calit2_events_cols)
# -
| event/Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Executive Summary
# +
# This analysis looks at the frequency of flights on radar tracks at KHPN. The question being reviewed is "Has the frequency of flights changed?"
# +
# Data used here is 3D radar track data from August 2005, 2010, 2015, and 2018
# -
# # Import Required Packages
# +
from __future__ import print_function
import json
import time
import datetime
from urllib.request import urlopen
import os
import math
import matplotlib
import pandas as pd
import io
import numpy as np
import numpy.random
import matplotlib.pyplot as plt
from matplotlib import cm as CM
from matplotlib import pyplot as PLT
from bs4 import BeautifulSoup
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:90% !important; }</style>"))
# -
# # Define Functions
def trackScope(df,southernLat,northernLat,westernLong,easternLong,maxAlt):
df = df[(df['Latitude']>=southernLat) & (df['Latitude']<=northernLat) & (df['Longitude']<=easternLong) & (df['Longitude']>=westernLong)]
return df
def trackPrep(df):
df['Actual Date/Time'] = pd.to_datetime(df['Actual Date/Time'])
df['Track Point Time'] = pd.to_datetime(df['Track Point Time'])
return df
def trackDiffs(df):
df = df.groupby('Operation No')['Track Point Time'].min().reset_index()
df = df.sort_values(by=['Track Point Time'])
df['Diff'] = df['Track Point Time'].shift(-1) - df['Track Point Time']
df['DiffSeconds'] = df['Diff'].dt.total_seconds()
df['DiffSecondsClipped'] = df['DiffSeconds']
df['DiffSecondsClipped'] = np.clip(df['DiffSeconds'], a_max=900, a_min=None)
return df
def trackHistAnalysis(Lat1, Lat2, Long1, Long2, Alt, Title):
df2005Scope = trackDiffs(trackScope(df[df['Dataset']=='2005-08'],Lat1, Lat2, Long1, Long2, Alt))
df2010Scope = trackDiffs(trackScope(df[df['Dataset']=='2010-08'],Lat1, Lat2, Long1, Long2, Alt))
df2015Scope = trackDiffs(trackScope(df[df['Dataset']=='2015-08'],Lat1, Lat2, Long1, Long2, Alt))
df2018Scope = trackDiffs(trackScope(df[df['Dataset']=='2018-08'],Lat1, Lat2, Long1, Long2, Alt))
dataScope = [df2005Scope["DiffSecondsClipped"], df2010Scope["DiffSecondsClipped"], df2015Scope["DiffSecondsClipped"], df2018Scope["DiffSecondsClipped"]]
headers = ["2005", "2010", "2015", "2018"]
dataScope = pd.concat(dataScope, axis=1, keys=headers)
MainMap = df[df['Dataset']=='2018-08']
MainMap['Color'] = 'DarkBlue'
FocusMap = trackScope(df[df['Dataset']=='2018-08'],Lat1, Lat2, Long1, Long2, Alt)
FocusMap['Color'] = 'Red'
MapData = pd.concat([MainMap, FocusMap])
MapData.plot.scatter(x='Longitude',
y='Latitude',
c=MapData['Color'],
xlim=(-73.77,-73.65), ylim=(41.000,41.120),
alpha=0.05, s=3, figsize=(10,10), title=Title);
dataScope[["2005", "2010", "2015", "2018"]].plot.hist(bins=60, alpha=0.2, title="Histogram of Flight Frequency by Seconds", density=True)
dataScope.hist(color="k", alpha=0.5, bins=60, figsize=(10, 10), density=True);
return dataScope
# # ETL Data
df2018 = pd.read_csv('data/August 2018 HPN traffic.raw.csv')
df2018 = trackPrep(df2018)
df2010 = pd.read_csv('data/August 2010 HPN traffic smoothed.csv')
df2010 = trackPrep(df2010)
df2005 = pd.read_csv('data/August 2005 HPN traffic smoothed.csv')
df2005 = trackPrep(df2005)
df2015 = pd.read_csv('data/August 2015 HPN traffic.raw.csv')
df2015 = trackPrep(df2015)
df2005['Dataset'] = '2005-08'
df2010['Dataset'] = '2010-08'
df2015['Dataset'] = '2015-08'
df2018['Dataset'] = '2018-08'
df = pd.concat([df2005, df2010, df2015, df2018])
# # Analysis
# ## Set 1 - Approach End of 34
set1 = trackHistAnalysis(41.051,41.054,-73.702,-73.692,3500,"KHPN Traffic Frequency Analysis - Approach End of 34")
set1.describe()
# ## Set 2 - HPN7 Departure off Runway 16 (Purchase St. and Anderson Hill Road)
set2 = trackHistAnalysis(41.031,41.041,-73.715,-73.701,3500,"KHPN Traffic Frequency Analysis - HPN7 Departure off 16 in Purchase")
set2.describe()
# ## Set 3 - HPN7 Approach End of 16
set3 = trackHistAnalysis(41.077,41.085,-73.728,-73.708,3500,"KHPN Traffic Frequency Analysis - HPN7 Departure off 16 in Purchase")
set3.describe()
# ## Set 4 - Runway 34 Arrivals
set4 = trackHistAnalysis(41.01,41.02,-73.67,-73.66,3500,"KHPN Traffic Frequency Analysis - Runway 34 Approach")
set4.describe()
# ## Set 5 - Runway 16 Arrivals
set5 = trackHistAnalysis(41.10,41.11,-73.74,-73.73,3500,"KHPN Traffic Frequency Analysis - Runway 16 Approach")
set5.describe()
# ## Set 6 - Runway 29 Arrivals
set6 = trackHistAnalysis(41.05,41.07,-73.69,-73.68,3500,"KHPN Traffic Frequency Analysis - Runway 29 Approach")
set6.describe()
| KHPN 16 Departures Radar Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# !python --version
# !pwd
CONFIG = 'basic_rnn'
BUNDLE_PATH = '/root/magenta/magenta/models/melody_rnn/mag/basic_rnn.mag'
OUTPUT_DIR = '/root/magenta/magenta/models/melody_rnn/output/2020.3.7-01'
# # 用 pre-trained model 生成序列
# !melody_rnn_generate \
# --config='attention_rnn' \
# --bundle_file='/root/magenta/magenta/models/melody_rnn/mag/attention_rnn.mag' \
# --output_dir='/root/magenta/magenta/models/melody_rnn/output/2020.3.30' \
# --num_outputs=5 \
# --num_steps=256 \
# --primer_midi='/root/magenta/magenta/models/melody_rnn/primer.mid'
# # 制作数据集
# **注:这一节的脚本只是记录下来的,由于输出过多,VS Code容易与服务器断开连接,所以脚本需要在终端运行。**
# ## Building your Dataset
# +
INPUT_DIRECTORY=/download/clean_midi
# TFRecord file that will contain NoteSequence protocol buffers.
SEQUENCES_TFRECORD=/root/notesequences.tfrecord
convert_dir_to_note_sequences \
--input_dir=$INPUT_DIRECTORY \
--output_file=$SEQUENCES_TFRECORD \
--recursive
# -
# ## Create SequenceExamples
melody_rnn_create_dataset \
--config=attention_rnn \
--input=/root/notesequences.tfrecord \
--output_dir=/download/melody_rnn/sequence_examples \
--eval_ratio=0.10
# # 训练模型
# ## 第一次训练(3.30-3.31): run1
melody_rnn_train \
--config=attention_rnn \
--run_dir=/root/magenta/magenta/models/melody_rnn/logdir/run1 \
--sequence_example_file=/download/melody_rnn/sequence_examples/training_melodies.tfrecord \
--hparams="batch_size=64,rnn_layer_sizes=[64,64]" \
--num_training_steps=20000
# run1因为未知原因被中止。生成旋律:
# !melody_rnn_generate \
# --config=attention_rnn \
# --run_dir=/root/magenta/magenta/models/melody_rnn/logdir/run1 \
# --output_dir=/root/magenta/magenta/models/melody_rnn/generated/run1 \
# --num_outputs=10 \
# --num_steps=128 \
# --hparams="batch_size=64,rnn_layer_sizes=[64,64]" \
# --primer_melody="[60]"
# ## 生成bundle file(.mag文件)
# !melody_rnn_generate \
# --config=attention_rnn \
# --run_dir=/root/magenta/magenta/models/melody_rnn/logdir/run1 \
# --hparams="batch_size=64,rnn_layer_sizes=[64,64]" \
# --bundle_file=/root/magenta/magenta/models/melody_rnn/generated/run1/attention_rnn.mag \
# --save_generator_bundle
# ## 2020.4.9 继续训练 & eval
melody_rnn_train \
--config=attention_rnn \
--run_dir=/root/magenta/magenta/models/melody_rnn/logdir/run1 \
--sequence_example_file=/download/basic_rnn/sequence_examples/eval_melodies.tfrecord \
--hparams="batch_size=64,rnn_layer_sizes=[64,64]" \
--num_training_steps=20000 \
--eval
# ## 第二次训练(4.4): run2
melody_rnn_create_dataset \
--config=basic_rnn \
--input=/root/notesequences.tfrecord \
--output_dir=/download/basic_rnn/sequence_examples \
--eval_ratio=0.10
# **制作数据集出错:**
#
# > Traceback (most recent call last):
# File "/root/anaconda3/envs/magenta/bin/melody_rnn_create_dataset", line 11, in <module>
# load_entry_point('magenta', 'console_scripts', 'melody_rnn_create_dataset')() File "/root/magenta/magenta/models/melody_rnn/melody_rnn_create_dataset.py", line 63, in console_entry_point
# tf.app.run(main)
# File "/root/anaconda3/envs/magenta/lib/python3.7/site-packages/tensorflow_core/python/platform/app.py", line 40, in run
# _run(main=main, argv=argv, flags_parser=_parse_flags_tolerate_undef)
# File "/root/anaconda3/envs/magenta/lib/python3.7/site-packages/absl/app.py", line 299, in run
# _run_main(main, args)
# File "/root/anaconda3/envs/magenta/lib/python3.7/site-packages/absl/app.py", line 250, in _run_main
# sys.exit(main(argv))
# File "/root/magenta/magenta/models/melody_rnn/melody_rnn_create_dataset.py", line 59, in main
# FLAGS.output_dir)
# File "/root/magenta/magenta/pipelines/pipeline.py", line 374, in run_pipeline_serial
# for input_ in input_iterator:
# File "/root/magenta/magenta/pipelines/pipeline.py", line 310, in tf_record_iterator
# for raw_bytes in tf.python_io.tf_record_iterator(tfrecord_file):
# File "/root/anaconda3/envs/magenta/lib/python3.7/site-packages/tensorflow_core/python/lib/io/tf_record.py", line 181, in tf_record_iterator
# reader.GetNext()
# File "/root/anaconda3/envs/magenta/lib/python3.7/site-packages/tensorflow_core/python/pywrap_tensorflow_internal.py", line 1034, in GetNext return _pywrap_tensorflow_internal.PyRecordReader_GetNext(self)tensorflow.python.framework.errors_impl.DataLossError: truncated record at 1412079946
melody_rnn_train \
--config=basic_rnn \
--run_dir=/root/magenta/magenta/models/melody_rnn/logdir/run2 \
--sequence_example_file=/download/basic_rnn/sequence_examples/training_melodies.tfrecord \
--hparams="batch_size=64,rnn_layer_sizes=[64,64]" \
--num_training_steps=20000
# 训练成功,测试生成旋律。
# **eval**
melody_rnn_train \
--config=basic_rnn \
--run_dir=/root/magenta/magenta/models/melody_rnn/logdir/run2 \
--sequence_example_file=/download/basic_rnn/sequence_examples/eval_melodies.tfrecord \
--hparams="batch_size=64,rnn_layer_sizes=[64,64]" \
--num_training_steps=20000 \
--eval
# !melody_rnn_generate \
# --config=basic_rnn \
# --run_dir=/root/magenta/magenta/models/melody_rnn/logdir/run2 \
# --output_dir=/root/magenta/magenta/models/melody_rnn/generated/run2 \
# --num_outputs=10 \
# --num_steps=128 \
# --hparams="batch_size=64,rnn_layer_sizes=[64,64]" \
# --primer_melody="[60]"
| magenta/models/melody_rnn/melody_rnn_train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 5. CREATE PYLDAVIS BROWSER
#
# [pyLDAvis](https://github.com/bmabey/pyLDAvis) is a port of the R LDAvis package for interactive topic model visualization by <NAME> and <NAME>.
#
# pyLDAvis is designed to help users interpret the topics in a topic model by examining the relevance and salience of terms in topics. Along the way, it displays tabular data which can be used to examine the model.
#
# pyLDAvis is not designed to use Mallet data out of the box. This notebook transforms the Mallet state file into the appropriate data formats before generating the visualisation. The code is based on Jeri Wieringa's blog post [Using pyLDAvis with Mallet](http://jeriwieringa.com/2018/07/17/pyLDAviz-and-Mallet/) and has been slightly altered and commented.
#
# ### INFO
#
# __author__ = '<NAME>'
# __copyright__ = 'copyright 2019, The WE1S Project'
# __license__ = 'GPL'
# __version__ = '2.0'
# __email__ = '<EMAIL>'
# ## Settings
# +
import gzip
import json
import os
from IPython.display import display, HTML
from pathlib import Path
# current_dir = %pwd
current_pathobj = Path(current_dir)
project_dir = str(current_pathobj.parent.parent)
print(project_dir)
published_site_folder_name = os.path.basename(project_dir)
data_dir = project_dir + '/project_data'
model_dir = data_dir + '/models'
pyldavis_script_path = current_dir + '/' + 'pyldavis_scripts/PyLDAvis.py'
output_path = current_dir
output_file = 'index.html'
json_dir = data_dir + '/json'
# %run {pyldavis_script_path}
# -
# ## Configuration
#
# Select models to create pyldavis visualizations for. **Please run the next cell regardless of whether you change anything.**
#
# By default, this notebook is set to create a pyldavis for all of the models you produced in Notebook 2 (`02_model_topics.ipynb`). If you would like to select only certain models to produce a pyldavis for, make those selections in the next cell (see next paragraph). Otherwise leave the value in the next cell set to `None`, which is the default.
#
# **To produce pyldavis for a selection of the models you created, but not all:** Navigate to the `your_project_name/project_data/models` directory in your project. Note the name of each subdirectory in that folder. Each subdirectory should be called `topicsn1`, where `n1` is the number of topics you chose to model. You should see a subdirectory for each model you produced. To choose which subdirectory/ies you would like to produce browsers for, change the value of `selection` in the cell below to a list of subdirectory names. For example, if you wanted to produce browsers for only the 50- and 75-topic models you created, change the value of `selection` below to this:
#
# Example:
#
# `selection = ['topics50','topics75']`
#
# Please follow this format exactly.
selection = ['topics10']
# Get names of model subdirectories to visualize and their state files.
# +
models = get_models(model_dir, selection)
# Display all model sub-directories with index numbers
for index, item in enumerate(models):
print(str(index) + ': ' + item['model'])
# -
# ### Add metadata and labels for the user interface (Optional).
#
# To do this, identify the index number for each model in the list above, and add the necessary information using the following lines in the next cell.
#
# ```python
# models[0]['metadata'] = 'pub'
# models[0]['ui_labels'] = [
# 'Intertopic Distance Map (via multidimensional scaling)',
# 'topic',
# 'publication',
# 'publications',
# 'tokens'
# ]
# ```
# Additional models would be `models[1]`, `models[2]`, etc.
#
# The `ui_labels` must be given in the following order:
#
# 1. The title of the multidimensional scaling graph
# 2. The type of unit represented by the graph circles
# 3. The singular form of the unit represented in the bar graphs on the right
# 4. The plural form of the unit represented in the bar graph on the right.
# 5. The unit represented by the percentage in the Relevance display.
#
# The example above indicates that the model will represent a map of intertopic distances in which each topic will show the distribution of publications, as represented by the percentage of topic tokens in the publication.
#
# **If you are unsure what to put, you do not have to assign `ui_labels`. A visualization will still be generated but may not have appropriate labels for the type of metadata you are using.**
# +
# Uncomment and modify these lines to run the cell
models[0]['metadata'] = 'pub'
models[0]['ui_labels'] = [
'Intertopic Distance Map (AKA the 10 Circles of Hell)',
'topic',
'publication',
'publications',
'tokens'
]
# models[1]['metadata'] = 'pub'
# models[1]['ui_labels'] = [
# 'Intertopic Distance Map (via multidimensional scaling)',
# 'topic',
# 'publication',
# 'publications',
# 'tokens'
# ]
display(HTML('<h4>Here is a summary of the information you will be using to generate your visualization(s).</h4>'))
print(json.dumps(models, indent=2))
# -
# ## Generate the Visualizations
# %run {pyldavis_script_path}
generate(model_dir, models, output_path, output_file, json_dir)
| src/templates/v0.1.9/modules/pyldavis/.ipynb_checkpoints/new_pyldavis-checkpoint.ipynb |
import sys
sys.stdout.write('hello world\n')
sys.stdout.flush()
for i in range(3):
sys.stdout.write('%s\n' % i)
sys.stdout.flush()
sys.stderr.write('output to stderr\n')
sys.stderr.flush()
sys.stdout.write('some more stdout text\n')
sys.stdout.flush()
# # Markdown Cell
#
# $ e^{ \pm i\theta } = \cos \theta \pm i\sin \theta + \beta $
#
# *It* **really** is!
this is a syntax error
print('test')
from IPython.display import Latex
Latex('''The mass-energy equivalence is described by the famous equation
$$E=mc^2$$
discovered in 1905 by <NAME>.
In natural units ($c$ = 1), the formula expresses the identity
\\begin{equation}
E=m
\\end{equation}''')
from IPython.display import display
import ipywidgets as w
# s = w.IntSlider(0, 10)£
# s
a = w.IntSlider()
b = w.IntText()
w.jslink((a, 'value'), (b, 'value'))
display(a, b)
import plotly.express as px
df = px.data.tips()
fig = px.histogram(df, x="total_bill", y="tip", color="sex", marginal="rug",
hover_data=df.columns)
fig.show()
import numpy as np
size = 100
np.random.seed(0)
x_data = np.arange(size)
y_data = np.cumsum(np.random.randn(size) * 100.0)
from bqplot import pyplot as plt
plt.figure(title="My First Plot")
plt.plot(x_data, y_data)
plt.show()
# +
from pyecharts.charts import Bar
from pyecharts import options as opts
bar = (
Bar()
.add_xaxis(["衬衫", "毛衣", "领带", "裤子", "风衣", "高跟鞋", "袜子"])
.add_yaxis("商家A", [114, 55, 27, 101, 125, 27, 105])
.add_yaxis("商家B", [57, 134, 137, 129, 145, 60, 49])
.set_global_opts(title_opts=opts.TitleOpts(title="某商场销售情况"))
)
# for jupyterlab
# bar.load_javascript()
bar.render_notebook()
# -
bar
| dev/notebooks/ping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Generating OpenFoam 6DoF Files
#
# This Jupyter notebook will show you how to generate a simple 6 degree of freedom kinematic file for OpenFoam. The data file we will generate specifies the magnitude and direction of motion for a series of time points.
#
# It is a combination of a linear displacement vector and a rotation vector about the specified center of gravity (CofG) of the object, which is defined on the dynamicMeshDict file.
#
# This data file is later linearly interpolated by OpenFoam when running your CFD simulation, so your time points do not need to match exactly the timesteps you have set in OpenFoam. However, more time points will smooth your dynamics, increasing the accuracy of your simulation, specially if, as seen in the case below, you have a sinusoidal or other non-linear motion.
#
# The format of the data file we want to create for OpenFoam is shown below:
#
# 4 //number of data points in the file
# //Position formatting is not important. File is based on the character sequence only.
# //Vectors are not relative. Each vector is total displacement and total rotation.
#
# (
#
# //(time_point ( (linear displacement vector) (rotation vector roll-yaw-pitch) ) )
#
# //(seconds ( (following unit system, usually meters) (degrees) ) )
#
# (0 ( (0.25 0.50 1.0) (0.220 0.30 0.40) ) )
# (0.25 ( (0.50 1.0 2.0) (0.60 0.60 0.60) ) )
# (0.75 ( (0.75 5.0 1.0) (1.2 2.4 5.0) ) )
# (10.0 ( (0.1 6.0 1.0) (5.0 3.0 5.5) ) )
# )
#
# Each time point first specifies the reference time, and the following bracket groups contains the motion, first linear and then angular displacement in their own lists.
#
# We will begin by importing the libraries we need: Numpy, Pandas, MatPlotLib to visualize our output, OS to import paths, and IPython to clean up our display.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import os
from IPython.display import clear_output
# We will now add our parameters for the kinematics. In this case we are making a sinusoidal pitch-plunge movement, for 5 periods. The geometric angle of attack (our pitch angle) will be 15 degrees, and our plunge amplitude will be 0.05m. It is a 2D motion, so most of our degrees of freedom are actually just going to be 0.
numberofperiods = 5
alphaGeoAmp = 15
plungeamplitude = 0.05
period = 3.801279221
resolution = 1000 #this is our time resolution, the number of data points our file will have
# We are going to be saving this file as "pitchplunge.dat"
#outputfile
outputfilename = os.getcwd()
outputfilename += '/pitchplunge.dat'
# We generate our time vector by using the numpy function linspace, with our set resolution
#time
time = np.linspace(0,period*numberofperiods,resolution)
# And now it's just a question of creating our X,Y and Z vectors for our linear and rotational displacement. We will store them in a Pandas DataFrame (our motion matrix).
# +
#linear displacement
linearDisplacement = pd.DataFrame(data=time,columns={'Time'})
linearDisplacement['X'] = 0
linearDisplacement['Y'] = plungeamplitude*np.sin(time)
linearDisplacement['Z'] = 0
#rotational displacement
rotationalDisplacement = pd.DataFrame(data=time,columns={'Time'})
rotationalDisplacement['phi'] = 0
rotationalDisplacement['psi'] = 0
rotationalDisplacement['theta'] = -alphaGeoAmp * np.sin(time)
# -
# Finally we can save our motion file, adding our different brackets and required formatting as per OpenFoam's requirements. We will use the f.write file writer, and build each line with a for loop.
# +
#saving motion file
f= open(outputfilename,"w+")
f.write(str(resolution) + '\n(\n')
for i in range(0,resolution,1):
line = '('
line += str(linearDisplacement['Time'].iloc[i])
line += '(('
line += str(linearDisplacement['X'].iloc[i])
line += ' '
line += str(linearDisplacement['Y'].iloc[i])
line += ' '
line += str(linearDisplacement['Z'].iloc[i])
line += ')('
line += str(rotationalDisplacement['phi'].iloc[i])
line += ' '
line += str(rotationalDisplacement['psi'].iloc[i])
line += ' '
line += str(rotationalDisplacement['theta'].iloc[i])
line += ')))\n'
f.write(line)
if i%1000==0: #this is a tiny add-on I like to use as a test and map progress
print('Completing save ',round(i/resolution*100,2), '%')
clear_output(wait=True)
f.write(')')
# -
# We will now plot our linear displacement to double check our motion:
plt.plot(linearDisplacement['Time']/period, linearDisplacement['Y'])
plt.xlim(0,numberofperiods)
plt.ylabel("Linear Displacement /m")
plt.xlabel("t/T")
plt.title("Linear Displacement against non-dimensional time")
# And now our rotational displacement:
plt.plot(rotationalDisplacement['Time']/period,rotationalDisplacement['theta'])
plt.xlim(0,numberofperiods)
plt.ylabel("Angular displacement /rad")
plt.xlabel("t/T")
plt.title("Angular displacement against non-dimensional time")
| 6DoFOpenFoam/6DoFOpenFoam.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ismailyou/ner/blob/main/Tebyan_Arabic_NER_Model_Testing.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="2S6yjmkar6LG"
# !pip3 install simpletransformers==0.61.5 nltk==3.5
# + id="ziEifcGzscCe"
from simpletransformers.ner import NERModel, NERArgs
import logging
import re
import nltk
nltk.download('punkt')
from nltk.tokenize import word_tokenize
logging.basicConfig(level=logging.INFO)
transformers_logger = logging.getLogger("transformers")
transformers_logger.setLevel(logging.WARNING)
# + [markdown] id="y74C07iaswRS"
# ## Load the Model
# + id="7SJBvtDxsl9_"
# Load the Model
custom_labels = ["O", "B-job", "I-job", "B-nationality", "B-person", "I-person", "B-location",
"B-time", "I-time", "B-event", "I-event", "B-organization", "I-organization",
"I-location", "I-nationality", "B-product", "I-product", "B-artwork", "I-artwork"]
model_args = NERArgs()
model_args.labels_list=custom_labels
model_args.silent = True
ner_model = NERModel(
"xlmroberta", "marefa-nlp/marefa-ner",
args=model_args,
use_cuda=True # set to False to use CPU
)
# + [markdown] id="ieIjAavbs36p"
# ## Model Inference
# + id="3j-PAEYntShR"
# Model Inference
samples = [
"تلقى تعليمه في الكتاب ثم انضم الى الأزهر عام 1873م. تعلم على يد السيد جمال الدين الأفغاني والشيخ محمد عبده",
"بعد عودته إلى القاهرة، التحق نجيب الريحاني فرقة جورج أبيض، الذي كان قد ضمَّ - قُبيل ذلك - فرقته إلى فرقة سلامة حجازي . و منها ذاع صيته",
"امبارح اتفرجت على مباراة مانشستر يونايتد مع ريال مدريد في غياب الدون كرستيانو رونالدو",
"Government extends flight ban from India, Pakistan until June 21"
]
# Preprocess
samples = [ " ".join(word_tokenize(sample.strip())) for sample in samples if sample.strip() != "" ]
# Predict
predictions, raw_outputs = ner_model.predict(samples)
# + colab={"base_uri": "https://localhost:8080/"} id="GMFUtNmLtZEw" outputId="487082ee-8102-4fca-f031-a8ac4a0413ac"
# Group the Predicted Entities
entities = []
for pred in predictions:
grouped_entities = []
for rec in pred:
token = list(rec.keys())[0]
label = rec[token]
if label == "O":
continue
if "B-" in label:
grouped_entities.append({"token": token, "label": label.replace("B-","")})
elif "I-" in label and len(grouped_entities) > 0:
grouped_entities[-1]["token"] += f" {token}"
entities.append(grouped_entities)
# Print the model outputs
for sample, results in zip(samples, entities):
print(sample)
for res in results:
print("\t", res["token"], "=>", res["label"])
print("==================")
| Tebyan_Arabic_NER_Model_Testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.4 64-bit (''.venv'': venv)'
# language: python
# name: python3
# ---
# # Female headed households with low income in South Africa
# ### - Prediction of a keyindicator
# -----------------------------
#
# __*Summary*__:
#
# We developed a machine learning model to predict the percentage of female headed households per ward in South Africa. The best result was archieved by XGBoost with a RMSE of 3.92. Further, we found, that the best predictors are school attendance and the ownership of a car and/or land ownership.
#
# -------------
#
# __*Objective*__:
#
# In South Africa, the number of female headed households is increasing. Low income and social challenges puts women at high risk for poverty. Non-Profits are interested to estimate the aid and support for those low income households.
#
# In this project, we develop a machine learning model to predict the percentage of female headed households with income under 19,600 R per year per ward.
#
# __*Value of the product*__:
#
# Monitor the percentage of female headed households between census years. Since female-headed households are prone to poverty, the stakeholder (a NGO) uses it as a keyindicator to estimate the amount of relief supplies in certain regions of South Africa.
#
# __*Data*__:
# + derived from Zindi ([link](https://zindi.africa/competitions/womxn-in-big-data-south-africa-female-headed-households-in-south-africa))
# + data are aggregated from south-african census 2011
# + 2822 observations, 64 variables
# + variables from survey: housing forms, school attendance, access to water etc
# + all variables are expressed as percentage
#
# __*Methods*__:
# + Regression methods (linear regression, random forest, gradient boosting)
# + identification of good predictors (features)
# + predict per ward
#
# __*Evaluation metrics*__:
# R2 and RMSE
# # Data Exploration and cleaning
# ------------------------------------
#
# +
# Import libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
import statsmodels.api as sm
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.metrics import r2_score, mean_squared_error
RSEED = 42
# -
# ## Data import
# Load the data
df = pd.read_csv("data/Train.csv")
df.head()
# Check the datatypes
df.info()
# Check for missing values
df.isnull().sum()
# Check for duplicates
df.duplicated().sum()
# Findings:
#
# + Data are mostly numerical
# + casted in the right datatype
# + no missing values found
# + no duplicates found
#
# -> no cleaning is needed at this point
# ## Distribution of the target
# Histogram of the target
sns.histplot(df.target).set_title("Distribution of the target")
sns.set_theme(style="darkgrid");
# QQ- plot of the data
sm.qqplot(df.target, line="q")
sns.set_theme(style="darkgrid")
plt.show()
# Conclusion: The target is normally distributed.
# ## Relationships and correlations
#
# The strength of a relationship between target and variable can be expressed as correlation. To identify features that are correlated with the target, the correlation is visualized in a heatmap and the highest correlations will be checked. The highest correlated features are selected on how allegeable they are for the model. Colinear features are excluded.
#
# +
# Setting style for the heatmap
sns.set_theme(style="white")
# Calculate a correlation matrix
corr = df.corr()
# Generate a mask for the upper triangle
mask = np.triu(np.ones_like(corr, dtype=bool))
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(30, 25))
# Generate a custom diverging colormap
cmap = sns.diverging_palette(230, 20, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(corr, mask=mask, cmap=cmap, vmax=.3, center=0,
square=True, linewidths=.5, cbar_kws={"shrink": .5});
# -
# __Findings on the heatmap:___
#
# + High positiv correlations can be found with NL, pw_00, pg_03, lan_00, lan_01, lln_00, car_00, stv_00, psa_01
# + High negativ correlation can be found with pw_02 - pw_06, pg_00, lan_04, lan_05, lln_01, car_01, stv_01, psa_00, dw_01
# + The features dw_12, dw_13, lan_13, pw_07, pw_08 do not contain values. The columns have to be checked.
# + The features lln_00/lln_01, car_00/car_01, stv_00/stv_01, psa_00/psa_01 are highly correlated with each other, because they contain yes/no answers.
# Check empty features
df[["dw_12", "dw_13", "lan_13", "pw_07", "pw_08"]].head()
#check for zero-values
def identify_zeros(dataframe):
'''counts all values for each feature that are not zero'''
for col, data in dataframe.iteritems():
non_zeros = np.count_nonzero(data.values)
zeros = len(data.values)- non_zeros
percent = round((zeros*100)/len(data.values), 1)
print("The feature", col, "contains", zeros, "Nulls. Percentage:", percent, "%")
# Calculate number of zeros in the features
identify_zeros(df[["dw_12", "dw_13", "lan_13", "pw_07", "pw_08"]])
# The features dw_12, dw_13, lan_13, pw_07, pw_08 contain only zeros and can be dropped.
# Drop columns that contain zeros only
df.drop(["dw_12", "dw_13", "lan_13", "pw_07", "pw_08"], axis=1, inplace=True)
df.head()
# ### Identify features with a correlation higher than 0.5
# Create a list of all features that are correlated
corr_high = corr[abs(corr["target"]) > 0.5]
corr_high = corr_high.sort_values("target").index.values.tolist()
corr_high
# ### Visual control of features with a correlation higher than 0.5
#Plot target and feature with correlation > 0.5
for col, data in df[corr_high].iteritems():
f, ax = plt.subplots()
ax.scatter(data.values, df["target"])
ax.set_xlabel(str(col), fontsize=15)
ax.set_ylabel("Target", fontsize=15)
ax.set_title("Correlation with target", fontsize=15)
sns.set_theme(style="darkgrid")
plt.show()
# Findings: All features are visually correlated, whereby psa_00 and psa_01 seem to be the most promising. However, these features contain the percentage listing present school attendance, psa_00: "Yes", psa_01: "No", and are therefore complementory.
# ## Conclusions
#
#
# + Correlations can be found with 23 features (NL, pw_00, pg_03, lan_00, lan_01, lln_00, car_00, stv_00, psa_01, pw_02 - pw_06, pg_00, lan_04, lan_05, lln_01, car_01, stv_01, psa_00, dw_01).
#
# + Collinearity: The features lln_00/lln_01, car_00/car_01, stv_00/stv_01, psa_00/psa_01 are highly correlated with each other, because they contain yes/no answers.
#
# + The features dw_12, dw_13, lan_13, pw_07, pw_08 do not contain values and were dropped.
#
#
| EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial N: Title
#
# **Filled notebook:**
# [](https://github.com/phlippe/uvadlc_notebooks/blob/master/docs/tutorial_notebooks/DL2/template/TemplateNotebook.ipynb)
# [](https://colab.research.google.com/github/phlippe/uvadlc_notebooks/blob/master/docs/tutorial_notebooks/DL2/template/TemplateNotebook.ipynb)
# **Pre-trained models:**
# [](https://github.com/phlippe/saved_models/tree/main/DL2/template/)
# **Recordings:**
# [](https://youtu.be/waVZDFR-06U)
# **Authors:**
# Your name here
# __TODOs:__
#
# * Update the links for the filled notebook (both github and collab) to your new notebook
# * Update the link for the saved models
# * Update the link for the YouTube recording if you have any. If you want to upload one to the UvA DLC YouTube account, you can contact Phillip.
# * Fill in the author names
# Here, you are supposed to add some intro about the topic. Give a short abstract motivating the tutorial, and then detail what will be done. It is good to have pictures here as well. If you add images, make sure to use SVGs for best resolution, and put them in the same folder as your notebook. An example is given below (use any HTML editing you like).
#
# <center width="100%"><img src="example_image.svg" width="350px" style="padding: 20px"></center>
#
# The next cell is where you import all your packages that you need. In case you have non-standard packages, make sure to install them to make it executable on GoogleColab (see for instance the PyTorch Lightning install).
# +
## Standard libraries
import os
import numpy as np
## Imports for plotting
import matplotlib.pyplot as plt
plt.set_cmap('cividis')
# %matplotlib inline
from IPython.display import set_matplotlib_formats
set_matplotlib_formats('svg', 'pdf') # For export
import matplotlib
matplotlib.rcParams['lines.linewidth'] = 2.0
import seaborn as sns
sns.set()
## tqdm for loading bars
from tqdm.notebook import tqdm
## PyTorch
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.utils.data as data
import torch.optim as optim
## Torchvision (TODO: ONLY NEEDED FOR VISION-BASED DATASETS)
import torchvision
from torchvision import transforms
# PyTorch Lightning
try:
import pytorch_lightning as pl
except ModuleNotFoundError: # Google Colab does not have PyTorch Lightning installed by default. Hence, we do it here if necessary
# !pip install --quiet pytorch-lightning>=1.4
import pytorch_lightning as pl
from pytorch_lightning.callbacks import LearningRateMonitor, ModelCheckpoint
# Import tensorboard (TODO: REMOVE IF YOU DO NOT WANT TO RUN TENSORBOARDS INTERACTIVELY)
# %load_ext tensorboard
# Path to the folder where the datasets are/should be downloaded (e.g. CIFAR10)
DATASET_PATH = "../data"
# Path to the folder where the pretrained models are saved (TODO: UPDATE LINK BELOW TO A FOLDER WITH THE NAME OF YOUR NOTEBOOK FOLDER)
CHECKPOINT_PATH = "../../saved_models/DL2/template"
# Setting the seed
pl.seed_everything(42)
# Ensure that all operations are deterministic on GPU (if used) for reproducibility
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
device = torch.device("cuda:0") if torch.cuda.is_available() else torch.device("cpu")
print("Device:", device)
# -
# You will likely have some pretrained models that you want to share with the students, and download when running on GoogleColab. You can do this with the cell below. If you don't have any pretrained models, you can remove the cell.
# +
import urllib.request
from urllib.error import HTTPError
# Github URL where saved models are stored for this tutorial (TODO: UPDATE URL BELOW TO YOUR NOTEBOOK FOLDER)
base_url = "https://raw.githubusercontent.com/phlippe/saved_models/main/DL2/template/"
# Files to download
pretrained_files = [] # (TODO: ADD A LIST OF STRINGS THAT ARE THE FILES YOU WANT TO DOWNLOAD. PATHS WITH RESPECT TO BASE_URL)
# Create checkpoint path if it doesn't exist yet
os.makedirs(CHECKPOINT_PATH, exist_ok=True)
# For each file, check whether it already exists. If not, try downloading it.
for file_name in pretrained_files:
file_path = os.path.join(CHECKPOINT_PATH, file_name)
if "/" in file_name:
os.makedirs(file_path.rsplit("/",1)[0], exist_ok=True)
if not os.path.isfile(file_path):
file_url = base_url + file_name
print(f"Downloading {file_url}...")
try:
urllib.request.urlretrieve(file_url, file_path)
except HTTPError as e:
print("Something went wrong. Please try later again, or contact the author with the full output including the following error:\n", e)
# -
# ## My Tutorial Topic
#
# Start your notebook from here. Introduce the topics, go step by step, don't forget to explain the code, etc.
#
# You can make use of different heading levels, they will be shown as tabs on the RTD website.
# ## Conclusion
#
# Give a conclusion and summary of the notebook. Give a retroview: what have the students learned from this notebook, what is there to further explore in this topic, anything critical to keep in mind?
#
# ### References
#
# Give a list of references, especially the papers that introduce the methods you implemented in this notebook.
| docs/tutorial_notebooks/DL2/template/TemplateNotebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import scipy.io, os
import numpy as np
import matplotlib.pyplot as plt
from netCDF4 import Dataset
from fastjmd95 import rho
from matplotlib.colors import ListedColormap
import seaborn as sns; sns.set()
sns.set()
import seawater as sw
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
import matplotlib as mpl
colours=sns.color_palette('colorblind', 10)
my_cmap = ListedColormap(colours)
color_list=colours
# -
# ## Code to plot the meridional overturning and density structure from the North Atlantic from Sonnewald and Lguensat (2021).
#
# Data used are from the ECCOv4 State Estimate available: https://ecco-group.org/products-ECCO-V4r4.html
#
# Note: Data is generated for the North Atlantic, also including the Southern Ocean and Artcic basin. Data fro the Paciic and Indian ocean are also generated, and the bleow code can be adjusted to plot this also.
#
# +
gridInfo=np.load('latLonDepthLevelECCOv4.npz')
zLev=gridInfo['depthLevel'][:]
depthPlot=zLev.cumsum()
lat=gridInfo['lat'][:]
lon=gridInfo['lon'][:]
zMat=np.repeat(zLev,720*360).reshape((50,360,720))
dvx=np.rot90(0.5*111000*np.cos(lat*(np.pi/180)),1)
masks=np.load('regimeMasks.npz')
maskMD=masks['maskMD']
maskSSV=masks['maskSSV']
maskNSV=masks['maskNSV']
maskTR=masks['maskTR']
maskSO=masks['maskSO']
maskNL=masks['maskNL']
# -
def getData(NR):
arr = os.listdir('/home/maike/Documents/ECCO_BV/NVELSTAR/.')
f =Dataset('/home/maike/Documents/ECCO_BV/NVELSTAR/'+arr[NR])
nvelS =f.variables['NVELSTAR'][:]
arr = os.listdir('/home/maike/Documents/ECCO_BV/NVELMASS/.')
f =Dataset('/home/maike/Documents/ECCO_BV/NVELMASS/'+arr[NR])
nvelM =f.variables['NVELMASS'][:]
return(nvelS+nvelM)
# ## Creating the basin masks
nvel= getData(1) #To get the shape
# +
globalMask=np.ones(nvel[0].shape)
maskArea=np.zeros(nvel[0].shape)*np.nan
maskArea[:,65:360,0:222]=1
maskArea[:,65:360,500:720]=1
maskArea[:,310:360,:]=np.nan
maskArea[:,210:350,160:250]=np.nan
maskArea[:,0:140,500:650]=np.nan
maskArea[:,0:165,500:620]=np.nan
maskArea[:,0:255,500:560]=np.nan
maskArea[:,0:210,500:570]=np.nan
maskArea[:,0:185,500:590]=np.nan
pacificMask=maskArea
maskArea=np.zeros(nvel[0].shape)*np.nan
maskArea[:,:,221:400]=1
maskArea[:,200:360,160:400]=1
maskArea[:,0:65,:]=1
maskArea[:,310:360,:]=1
maskArea[:,199:215,160:180]=np.nan
maskArea[:,199:210,160:190]=np.nan
atlanticMask=maskArea
maskArea=np.ones(nvel[0].shape)
indA=np.where(atlanticMask==1)
indP=np.where(pacificMask==1)
maskArea[indA]=np.nan
maskArea[indP]=np.nan
maskArea[:,100:250,100:250]=np.nan
indianMask=maskArea
plt.figure()
plt.imshow(np.flipud(globalMask[0]*nvel[0,0]))
plt.figure()
plt.imshow(np.flipud(atlanticMask[0]*nvel[0,0]))
plt.figure()
plt.imshow(np.flipud(pacificMask[0]*nvel[0,0]))
plt.figure()
plt.imshow(np.flipud(indianMask[0]*nvel[0,0]))
# -
# ## Calculating the streamfunction
#
# The overall meridional overturning ($\Psi_{z\theta}$) from Fig. 3 in Sonnewald and Lguensat (2021) is defined as:
#
# $$\Psi_{z\theta}(\theta,z)=- \int^z_{-H} \int_{\phi_2}^{\phi_1} v(\phi,\theta,z')d\phi dz',$$
#
# \noindent where $z$ is the relative level depth and $v$ is the meridional (north-south) component of velocity. For the regimes, the relevant velocity fields were then used. A positive $\Psi_{z\theta}$ signifies a clockwise circulation, while a negative $\Psi_{z\theta}$ signifies an anticlockwise circulation.
def psiZ(NVEL_IN, mask):
'''Function to calculate overturning in depth space as described in Sonnewald and Lguensat (2021).'''
ntrans=np.zeros(NVEL_IN[:,:,0].shape);
gmoc=np.zeros(NVEL_IN[:,:,0].shape);
NVEL=NVEL_IN*mask
# zonal transport integral
for zz in np.arange(0,50):
ntrans[zz,:]=np.nansum(NVEL[zz,:,:]*dvx,axis=1);
for zz in np.flipud(np.arange(0,49)):
gmoc[zz,:]=gmoc[zz+1,:]+ntrans[zz+1,:]*zLev[zz+1];
gmoc=gmoc/1e6;
return(gmoc)
def psiMaskedCalc(mask):
'''Calculating the overturning in depth space for the different regimes, as plotted in Fig. 3 in Sonnewald and Lguensat (2021).'''
yrs, months=20,12
PSI_all = np.zeros((yrs*months, 50, 360))*np.nan
PSI_NL = np.zeros((yrs*months, 50, 360))*np.nan
PSI_SO = np.zeros((yrs*months, 50, 360))*np.nan
PSI_SSV = np.zeros((yrs*months, 50, 360))*np.nan
PSI_NSV = np.zeros((yrs*months, 50, 360))*np.nan
PSI_MD = np.zeros((yrs*months, 50, 360))*np.nan
PSI_TR = np.zeros((yrs*months, 50, 360))*np.nan
ITTER=0
for NR in np.arange(0,yrs):
nvel= getData(NR)
# print('Got data')
for MM in np.arange(0,months):
PSI_all[ITTER]=psiZ(nvel[MM], np.ones(maskSO.shape)*mask)
PSI_NL[ITTER]=psiZ(nvel[MM], maskNL*mask)
PSI_SO[ITTER]=psiZ(nvel[MM], maskSO*mask)
PSI_SSV[ITTER]=psiZ(nvel[MM], maskSSV*mask)
PSI_NSV[ITTER]=psiZ(nvel[MM], maskNSV*mask)
PSI_MD[ITTER]=psiZ(nvel[MM], maskMD*mask)
PSI_TR[ITTER]=psiZ(nvel[MM], maskTR*mask)
ITTER+=1
return PSI_all, PSI_NL, PSI_SO, PSI_SSV, PSI_NSV, PSI_MD, PSI_TR
# +
PSI_all_A, PSI_NL_A, PSI_SO_A, PSI_SSV_A, PSI_NSV_A, PSI_MD_A, PSI_TR_A = psiMaskedCalc(atlanticMask)
PSI_all_P, PSI_NL_P, PSI_SO_P, PSI_SSV_P, PSI_NSV_P, PSI_MD_P, PSI_TR_P = psiMaskedCalc(pacificMask)
PSI_all_I, PSI_NL_I, PSI_SO_I, PSI_SSV_I, PSI_NSV_I, PSI_MD_I, PSI_TR_I = psiMaskedCalc(indianMask)
PSI_all_G, PSI_NL_G, PSI_SO_G, PSI_SSV_G, PSI_NSV_G, PSI_MD_G, PSI_TR_G = psiMaskedCalc(globalMask)
# +
#Save the data
np.savez('PSI_global', PSI_all_G=PSI_all_G, PSI_NL_G=PSI_NL_G, PSI_SO_G=PSI_SO_G, PSI_SSV_G=PSI_SSV_G, PSI_NSV_G=PSI_NSV_G, PSI_MD_G=PSI_MD_G, PSI_TR_G=PSI_TR_G)
np.savez('PSI_atlantic', PSI_all_A=PSI_all_A, PSI_NL_A=PSI_NL_A, PSI_SO_A=PSI_SO_A, PSI_SSV_A=PSI_SSV_A, PSI_NSV_A=PSI_NSV_A, PSI_MD_A=PSI_MD_A, PSI_TR_A=PSI_TR_A)
np.savez('PSI_pacific', PSI_all_P=PSI_all_P, PSI_NL_P=PSI_NL_P, PSI_SO_P=PSI_SO_P, PSI_SSV_P=PSI_SSV_P, PSI_NSV_P=PSI_NSV_P, PSI_MD_P=PSI_MD_P, PSI_TR_P=PSI_TR_P)
np.savez('PSI_indian', PSI_all_I=PSI_all_I, PSI_NL_I=PSI_NL_I, PSI_SO_I=PSI_SO_I, PSI_SSV_I=PSI_SSV_I, PSI_NSV_I=PSI_NSV_I, PSI_MD_I=PSI_MD_I, PSI_TR_I=PSI_TR_I)
# -
# ## Calculate the density in $\sigma_2$
def getDataTS(NR):
'''Retrieve the T and S data. Data from the ECCOv4 state estimate.'''
arr = os.listdir('/home/maike/Documents/ECCO_BV/THETA/.')
f =Dataset('/home/maike/Documents/ECCO_BV/THETA/'+arr[NR])
T =f.variables['THETA'][:]
arr = os.listdir('/home/maike/Documents/ECCO_BV/SALT/.')
f =Dataset('/home/maike/Documents/ECCO_BV/SALT/'+arr[NR])
S =f.variables['SALT'][:]
return(T, S)
dens=np.zeros((50,360,720))
ITTER=1
yrs=20
months=12
for NR in np.arange(0,yrs):
T,S = getDataTS(NR)
print('Got data', NR)
#Tin=sw.eos80.temp(S, T, -np.cumsum(zMat, axis=0), pr=np.cumsum(zMat, axis=0))
for MM in np.arange(0,months):
dens = dens+rho(S[MM], T[MM], 2000) - 1000
ITTER+=1
dens=dens/ITTER
#Save the density data.
np.save('density20yr', np.array(dens))
# # Finally, we plot the data.
#
# The plot is a composite of different subplots.
#
# +
levs=[32,33,34, 34.5, 35, 35.5,36,36.5,37,37.25,37.5,37.75,38]
cols=plt.cm.viridis([300,250, 200,150, 125, 100, 50,30, 10,15,10,9,1])
Land=np.ones(np.nansum(PSI_all_A, axis=0).shape)*np.nan
Land[np.nansum(PSI_all_A, axis=0)==0.0]=0
land3D=np.ones(dens.shape)
land3D[dens==0]=np.nan
def zPlotSurf(ax, data,zMin, zMax,label,mm,latMin,latMax,RGB,Ticks,saveName='test'):
land=np.ones(np.nanmean(data, axis=0).shape)*np.nan
land[np.nansum(data, axis=0)==0.0]=0
n=50
levels = np.linspace(-20, 20, n+1)
ax.contourf(lat[0,latMin:latMax],-depthPlot[zMin:zMax],-np.nanmean(data, axis=0)[zMin:zMax,latMin:latMax], levels=np.linspace(-20, 20, n+1),cmap=plt.cm.seismic, extend='both')
n2=30
densityPlot=np.nanmean((dens*land3D*mm), axis=2)
assert(len(levs)==len(cols))
CS=ax.contour(lat[0,latMin:latMax],-depthPlot[zMin:zMax],densityPlot[zMin:zMax,latMin:latMax],
levels=levs,
linewidths=3,colors=cols, extend='both')
ax.tick_params(axis='y', labelsize=20)
if Ticks == 0:
ax.set_xticklabels( () )
elif Ticks == 1:
ax.set_xticklabels( () )
ax.set_yticklabels( () )
ax.contourf(lat[0,latMin:latMax],-depthPlot[zMin:zMax],land[zMin:zMax,latMin:latMax], 1,cmap=plt.cm.Set2)
ax.contourf(lat[0,latMin:latMax],-depthPlot[zMin:zMax],Land[zMin:zMax,latMin:latMax], 50,cmap=plt.cm.bone)
yL=ax.get_ylim()
xL=ax.get_xlim()
plt.text(xL[0]+0.02*np.ptp(xL), yL[0]+0.4*np.ptp(yL), label, fontsize=20, size=30,
weight='bold', bbox={'facecolor':'white', 'alpha':0.7}, va='bottom')
def zPlotDepth(ax, data,zMin, zMax,label,mm,latMin,latMax,RGB,Ticks,saveName='test'):
land=np.ones(np.nanmean(data, axis=0).shape)*np.nan
land[np.nansum(data, axis=0)==0.0]=0
n=50
levels = np.linspace(-20, 20, n+1)
ax.contourf(lat[0,latMin:latMax],-depthPlot[zMin:zMax],-np.nanmean(data, axis=0)[zMin:zMax,latMin:latMax], levels=np.linspace(-20, 20, n+1),cmap=plt.cm.seismic, extend='both')
n2=30
densityPlot=np.nanmean((dens*land3D*mm), axis=2)
ax.contour(lat[0,latMin:latMax],-depthPlot[zMin:zMax],densityPlot[zMin:zMax,latMin:latMax], colors=cols,
levels=levs,
linewidths=3, extend='both')
if Ticks == 0:
ax.tick_params(axis='y', labelsize=20)
#ax.set_xticklabels( () )
elif Ticks== 1:
#ax.set_xticklabels( () )
ax.set_yticklabels( () )
plt.tick_params(axis='both', labelsize=20)
#plt.clim(cmin, cmax)
ax.contourf(lat[0,latMin:latMax],-depthPlot[zMin:zMax],land[zMin:zMax,latMin:latMax], 1,cmap=plt.cm.Set2)
ax.contourf(lat[0,latMin:latMax],-depthPlot[zMin:zMax],Land[zMin:zMax,latMin:latMax], 50,cmap=plt.cm.bone)
yL=ax.get_ylim()
xL=ax.get_xlim()
plt.text(xL[0]+0.03*np.ptp(xL), yL[0]+0.03*np.ptp(yL), label, fontsize=20, size=30,
weight='bold', bbox={'facecolor':RGB, 'alpha':1}, va='bottom')
# +
# Set general figure options
# figure layout
xs = 15.5 # figure width in inches
nx = 2 # number of axes in x dimension
ny = 3 # number of sub-figures in y dimension (each sub-figure has two axes)
nya = 2 # number of axes per sub-figure
idy = [2.0, 1.0] # size of the figures in the y dimension
xm = [0.07, 0.07,0.9, 0.07] # x margins of the figure (left to right)
ym = [1.5] + ny*[0.07, 0.1] + [0.3] # y margins of the figure (bottom to top)
# pre-calculate some things
xcm = np.cumsum(xm) # cumulative margins
ycm = np.cumsum(ym) # cumulative margins
idx = (xs - np.sum(xm))/nx
idy_off = [0] + idy
ys = np.sum(idy)*ny + np.sum(ym) # size of figure in y dimension
# make the figure!
fig = plt.figure(figsize=(xs, ys))
# loop through sub-figures
ix,iy=0,0
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
#ax = plt.axes(loc)
for iys in range(nya):
# (bottom left corner x, bottom left corner y, width, height)
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
ax = plt.axes(loc)
# split between your two figure types
if iys == 0:
zPlotDepth(ax, PSI_TR_A,1,50,'TR', maskTR,200, 310, color_list[1],'')
# if not the bottom figure remove x ticks
if iy > 0:
ax.set_xticks([])
else:
xticks = ax.get_xticks()
ax.set_xticklabels(['{:0.0f}$^\circ$N'.format(xtick) for xtick in xticks])
elif iys == 1:
zPlotSurf(ax, PSI_TR_A,0,10,'', maskTR,200, 310, color_list[1],'')
# remove x ticks
ax.set_xticks([])
ix,iy=0,1
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
#ax = plt.axes(loc)
for iys in range(nya):
# (bottom left corner x, bottom left corner y, width, height)
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
ax = plt.axes(loc)
# split between your two figure types
if iys == 0:
zPlotDepth(ax, PSI_NL_A,1,50,'NL', maskNL,200, 310, color_list[-1],'')
# if not the bottom figure remove x ticks
if iy > 0:
ax.set_xticks([])
elif iys == 1:
zPlotSurf(ax, PSI_NL_A,0,10,'', maskNL,200, 310, color_list[4],'')
# remove x ticks
ax.set_xticks([])
############### n-SV
ix,iy=0,2
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
#ax = plt.axes(loc)
for iys in range(nya):
# (bottom left corner x, bottom left corner y, width, height)
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
ax = plt.axes(loc)
# split between your two figure types
if iys == 0:
zPlotDepth(ax, PSI_NSV_A,1,50,'N-SV', maskNSV,200, 310, color_list[4],'')
# if not the bottom figure remove x ticks
if iy > 0:
ax.set_xticks([])
elif iys == 1:
zPlotSurf(ax, PSI_NSV_A,0,10,'', maskNSV,200, 310, color_list[-1],'')
# remove x ticks
ax.set_xticks([])
#
#_______________________________________________________________________
# S-SV
ix,iy=1,2
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
# ax = plt.axes(loc)
for iys in range(nya):
# (bottom left corner x, bottom left corner y, width, height)
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
ax = plt.axes(loc)
# split between your two figure types
if iys == 0:
zPlotDepth(ax, PSI_SSV_A,1,50,'S-SV', maskSSV,200, 310, color_list[2],1,'')
# if not the bottom figure remove x ticks
if iy > 0:
ax.set_xticks([])
elif iys == 1:
zPlotSurf(ax, PSI_SSV_A,0,10,'', maskSSV,200, 310, color_list[-3],1,'')
# remove x ticks
ax.set_xticks([])
#%%%%%%%%%%%%%%%%%%%%%%%%% SO
ix,iy=1,1
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
#ax = plt.axes(loc)
for iys in range(nya):
# (bottom left corner x, bottom left corner y, width, height)
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
ax = plt.axes(loc)
# split between your two figure types
if iys == 0:
zPlotDepth(ax, PSI_SO_A,1,50,'SO', maskSO,200, 310, color_list[-3],1,'')
# if not the bottom figure remove x ticks
if iy > 0:
ax.set_xticks([])
elif iys == 1:
zPlotSurf(ax, PSI_SO_A,0,10,'', maskSO,200, 310, color_list[-3],1,'')
# remove x ticks
ax.set_xticks([])
#%%%%%%%MD
ix,iy=1,0
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
#ax = plt.axes(loc)
for iys in range(nya):
# (bottom left corner x, bottom left corner y, width, height)
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
ax = plt.axes(loc)
# split between your two figure types
if iys == 0:
zPlotDepth(ax, PSI_MD_A,1,50,'MD', maskMD,200, 310, color_list[0],1,'')
# if not the bottom figure remove x ticks
if iy > 0:
ax.set_xticks([])
else:
xticks = ax.get_xticks()
ax.set_xticklabels(['{:0.0f}$^\circ$N'.format(xtick) for xtick in xticks])
elif iys == 1:
zPlotSurf(ax, PSI_MD_A,0,10,'', maskMD,200, 310, color_list[-3],1,'')
# remove x ticks
ax.set_xticks([])
cmap = plt.get_cmap('viridis')
cmap = mpl.colors.ListedColormap(cols)
ncol = len(levs)
axes = plt.axes([(xcm[0])/(xs), (ym[0]-0.6)/ys, (2*idx + xm[1])/(xs*2), (0.2)/ys])
cb = fig.colorbar(plt.cm.ScalarMappable(norm=mpl.colors.Normalize(-0.5, ncol - 0.5), cmap=cmap),
cax=axes, orientation='horizontal')
cb.ax.set_xticks(np.arange(ncol))
cb.ax.set_xticklabels(['{:0.2f}'.format(lev) for lev in levs])
cb.ax.tick_params(labelsize=20)
cb.set_label(label=r'Density, $\sigma_2$',weight='bold', fontsize=20)
cmap = plt.get_cmap('seismic')
ncol = len(cols)
axes = plt.axes([(xcm[2]+2*idx)/(xs*2), (ym[0]-0.6)/ys, (2*idx+xm[3])/(xs*2), (0.2)/ys])
cb = fig.colorbar(plt.cm.ScalarMappable(norm=mpl.colors.Normalize(-20,20), cmap=cmap),
cax=axes, label='title', orientation='horizontal', extend='both',format='%.0f',
boundaries=np.linspace(-20, 20, 41))
cb.ax.tick_params(labelsize=20)
cb.set_label(label=r'SV ($10^{6}m^{2}s^{-2}$)',weight='bold', fontsize=20)
# save as a png
#fig.savefig('psiRho_NAtl_sigma2.png', dpi=200, bbox_inches='tight')
# -
| figures/.ipynb_checkpoints/MOC_z_wBasinCalc-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Pn0Ub9k5M0Sw"
# # **Curso Python - Curso em Vídeo - Mundo 3**
# + [markdown] id="loUUdM1k1KPK"
# # **Tuplas**
# Utiliza-se '( )' e não podem ser modificadas
# + id="MjjW-RZE1NtY" colab={"base_uri": "https://localhost:8080/"} outputId="1f320640-6047-4608-8db9-13e48ff9eeaf"
#As tuplas sao imutaveis
lanche=('Hamburguer', 'Suco', 'Pizza', 'Pudim')
print(lanche)
print(lanche[1])
print(lanche[-2:])
for cont in range(0, len(lanche)):
print(lanche[cont])
for pos, comida in enumerate(lanche):
print(f'Vou comer {comida} na posição {pos}')
print(sorted(lanche))
# + colab={"base_uri": "https://localhost:8080/"} id="OheN7zv66cbA" outputId="67fb9e46-b818-4e67-d9e8-069338e040df"
a=(2,5,4)
b=(5,8,1,2)
#Nao vai somar. Vai juntar
c=a+b
#Quantas vezes aparece o numero 5
print(c.count(5))
# + colab={"base_uri": "https://localhost:8080/"} id="zMmxKOVF9rci" outputId="76efb886-0a95-42dc-c37c-881d19391aac"
#Desafio 072 - Numero por extenso
t=('zero', 'um', 'dois', 'tres', 'quatro', 'cinco', 'seis', 'sete', 'oito', 'nove', 'dez',
'onze', 'doze', 'treze', 'quatorze', 'quinze', 'dezesseis', 'dezessete', 'dezoito', 'dezenove', 'vinte')
while True:
d=(int(input('Digite um número: ')))
if 0<=d<=20:
break
print('Tente novamente. ', end='')
print(f'Você digitou o número {t[d]}')
# + id="GiFxYmy52UKc" colab={"base_uri": "https://localhost:8080/"} outputId="47bf2cf5-e1ff-4730-c42f-156f572978f8"
#Desafio 073 - 20 primeiros colocados do brasileirao. Mostrar: os 5 primeiros, os 4 ultimos, ordem alfabetica e a posicao do chapecoense (ele nao saiu da serie A?)
a=('Atlético-MG', 'Palmeiras', 'Fortaleza', 'Bragantino', 'Flamengo', 'Corinthians', 'Fluminense',
'Atlético-GO', 'Athlético-PR', 'Ceará', 'Cuiabá', 'Internacional', 'Juventude', 'Santos', 'São Paulo', 'Bahia', 'América-MG', 'Sport', 'Grêmio', 'Chapecoense')
#Contar quantos times
#len(a)
while True:
p=0
print('Os cinco primeiros são: \n')
while p<5:
print(f'{a[p]}')
p+=1
u=1
print('-='*10)
print('Os quatro últimos são: \n')
while u<5:
print(f'{a[-u]}')
u+=1
print('-='*10)
print('Ordem alfabética: \n')
print(sorted(a))
print(f'O chapecoense está na {a.index("Chapecoense")+1}ª posição')
break
# + colab={"base_uri": "https://localhost:8080/"} id="nziaueRTaZUc" outputId="b8a1b0ca-fea2-4a2f-d5cf-6224672e90a3"
#Desafio 074 - Gerar 5 numeros aleatorios numa tupla. Indique o menor e o maior
from random import randint
numeros=(randint(1,10), randint(1, 10), randint(1, 10), randint(1, 10), randint(1,10))
print('Os valores sorteados são: ')
for n in numeros:
print(f'{n} ', end='')
print(f'\n o maior número foi {max(numeros)}')
print(f' o menor número foi {min(numeros)}')
# + colab={"base_uri": "https://localhost:8080/"} id="ZbzU1hF3iB2R" outputId="230df18f-cb13-4a6a-fe89-b2b70b955cf2"
#Desafio 075 - Ler quatro valores. Mostrar: quantas vezes apareceu o numero 9, a posicao do primeiro valor 3 e quais foram os numeros pares
num=(int(input('Digite o primeiro número: ')),
int(input('Digite o segundo número: ')),
int(input('Digite o terceiro número: ')),
int(input('Digite o quarto número: ')))
print(f'Você digitou: {num}')
print(f'O valor 9 apareceu {num.count(9)} vezes')
if 3 in num:
print(f'O número 3 apareceu na {num.index(3)+1}ª posição')
else:
print(f'O número 3 não foi digitado')
print('Os valores pares obtidos foram: ')
for i in num:
if i%2 == 0:
print(i, end='')
# + id="6EMaH7F6CfB5" colab={"base_uri": "https://localhost:8080/"} outputId="e295f8d0-9513-4401-e9dd-f2bbfb5374b0"
#Desafio 076 - Tupla com produto e preco
listagem=('Lápis', 1.75, 'Borracha', 2, 'Caderno', 15.90, 'Estojo', 25, 'Transferidor', 4.20, 'Compasso', 9.99, 'Mochila', 120.32, 'Canetas', 22.30, 'Livro', 34.90)
print('-' * 40)
print(f'{"LISTAGEM DE PREÇOS":^40}')
print('-' * 40)
for pos in range(0, len(listagem)):
if pos%2 == 0:
print(f'{listagem[pos]:.<30}', end='')
else:
#7.2f para mostrar com 2 casas decimais
print(f'R${listagem[pos]:>7.2f}')
print('-' * 40)
# + [markdown] id="B6IxMV6FreS0"
# # **Listas**
# Utiliza-se '[ ]' e podem ser modificadas
# + colab={"base_uri": "https://localhost:8080/"} id="gEGdFeUSrhXT" outputId="5fc9ae99-6a42-478d-fcdc-8a2edc895635"
#.append() para adicionar um elemento novo no final da lista
#.insert(0, ) para adicionar um elemento na posição zero
lanche=['Cachorro-quente', 'Hambúrguer', 'Suco', 'Pizza']
#del lanche[3]
#lanche.pop(3)
lanche.remove('Pizza')
lanche
# + colab={"base_uri": "https://localhost:8080/"} id="NjLl7cVw9Szn" outputId="46cfdad8-8ea2-430f-f45d-e7ff43aba73b"
valores=list(range(4,11))
valores
# + colab={"base_uri": "https://localhost:8080/"} id="-yy-IQdj9o3F" outputId="554c92d4-cfcc-4761-e2c7-b901c5b40a75"
valores=[8,2,5,4,9,3,0]
valores
valores.sort()
valores.sort(reverse=True)
valores
# + colab={"base_uri": "https://localhost:8080/"} id="J01QopFf99uk" outputId="b6cbcf99-d9d9-4bba-ca6d-4df6023ac4b3"
len(valores)
# + colab={"base_uri": "https://localhost:8080/"} id="LLb1lkTl-JVf" outputId="d5315f15-cfae-463d-e873-5ba0ae24b5ea"
#Tupla
#num=(2,5,9,1)
#lista
num=[2,5,9,1]
num[2]=3
num.append(7)
num
# + colab={"base_uri": "https://localhost:8080/"} id="nYPcIOi3vErx" outputId="9ac946a9-ec24-412f-ebb0-cb7e2fe1f2a0"
#Desafio 078 - lista com 5 valores numericos. Mostrar o maior e o menor e suas posicoes
num=[int(input('Digite um número para posição 0: ')),
int(input('Digite um número para posição 1: ')),
int(input('Digite um número para posição 2: ')),
int(input('Digite um número para posição 3: ')),
int(input('Digite um número para posição 4: '))]
menor=num[0]
maior=num[0]
a=0
b=0
print(f'Você digitou: {num}')
for pos in range(0, len(num)):
if num[pos]>maior:
maior=num[pos]
a=num.index(num[pos])
if num[pos]<menor:
menor=num[pos]
b=num.index(num[pos])
print(f'O maior número foi o {maior} digitado na posição número:\n.')
for i, v in enumerate(num):
if v == maior:
print(f'{i}...', end='')
print()
print(f'Já o menor número foi o {menor} na posição número\n')
for i, v in enumerate(num):
if v == menor:
print(f'{i}...', end='')
print()
# + colab={"base_uri": "https://localhost:8080/"} id="-Y0TjXMI45i7" outputId="755c2c42-3054-4d06-dcd1-5d69f30bc0e9"
#Desafio 079 - Cadastro de numeros. Quando houver repeticao, nao registrar. No final mostrar toda a lista
num=[]
while True:
a=int(input('Digite um número: '))
if a not in num:
num.append(a)
r=' '
while r not in 'SN':
r=str(input('Quer continuar? (s/n) ')).strip().upper()[0]
if r == 'N':
print('end')
break
num.sort()
print(num)
# + colab={"base_uri": "https://localhost:8080/"} id="yEozlOlLKCsY" outputId="e17a45ad-8c12-4bdc-d041-0bc9936b7ca2"
#Desafio 080 - Cadastrar 5 numeros em ordem crescente sem usar a funcao sort()
num=[]
for i in range(0,4):
a=(int(input('Digite um número: ')))
if i==0 or a >= num[len(num)-1]:
num.append(a)
print('Adicionado no final da lista')
else:
pos=0
while pos < len(num):
if a <= num[pos]:
num.insert(pos, a)
print(f'Adicionado na posição {pos} da lista...')
break
pos+=1
print('-='*30)
print(f'Os valores adicionados em ordem foram {num}')
# + colab={"base_uri": "https://localhost:8080/"} id="NE8FGWCxeLwH" outputId="1f0e3947-10ed-446d-95a2-9b4f017070df"
#Desafio 081 - Lista com varios numeros. Mostrar quantos numeros foram digitados,
#a lista em ordem decrescente e se o valor '5' esta na lista ou nao
num=[]
while True:
a=int(input('Digite um número: '))
num.append(a)
r=' '
while r not in 'SN':
r=str(input('Quer continuar (s/n)? ')).strip().upper()[0]
if r=='N':
print('Fim')
break
if 5 in num:
print('O número 5 está presente')
num.sort(reverse=True)
print(len(num))
print(num)
# + colab={"base_uri": "https://localhost:8080/"} id="z8ZUOpnUjOa9" outputId="c5e8bc19-e3d1-433f-d954-497e3371d38e"
#Desafio 082 - Criar uma lista. Criar uma lista que contem os numeros pares e outra que contem os impares. Mostre as tres listas no final
num=[]
p=[]
i=[]
while True:
a=int(input('Digite um número: '))
num.append(a)
if a%2 == 0:
p.append(a)
else:
i.append(a)
r=' '
while r not in 'SN':
r=str(input('Quer continuar (s/n)? ')).strip().upper()[0]
if r=='N':
print('Fim')
break
print(f'A lista original: {num}\n')
print(f'A lista dos pares: {p}\n')
print(f'A lista dos ímpares: {i}\n')
# + colab={"base_uri": "https://localhost:8080/"} id="OOE4UXCSmmbA" outputId="9445ab83-e5b0-4d97-e345-2d3dc4bf1c3a"
pessoas=[['Pedro', '25'], ['Maria', '19'], ['João', '32']]
print(pessoas[0][0])
print(pessoas[1][1])
print(pessoas[1])
# + colab={"base_uri": "https://localhost:8080/"} id="UBFt4lXhnlcp" outputId="43d2886a-04ab-4045-a8b3-a705ff5ba0d0"
teste=list()
teste.append('Gustavo')
teste.append(40)
galera=list()
galera.append(teste[:])
teste[0]='Maria'
teste[1]=22
galera.append(teste[:])
print(galera)
# + colab={"base_uri": "https://localhost:8080/"} id="LSZoC7AJqVET" outputId="7615f70d-d4a2-4423-e0a2-54e1f51a39c4"
#Desafio 084 - Lista com nome e peso de pessoas. Mostrar quantas pessoas foram cadastradas.
#Uma lista com a pessoa mais pesada. Uma com a pessoa mais leve
temp=[]
princ=[]
mai=men=0
while True:
temp.append(str(input('Nome: ')))
temp.append(float(input('Peso: ')))
if len(princ) == 0:
mai=men=temp[1]
else:
if temp[1] > mai:
mai = temp[1]
if temp[1] < men:
men = temp[1]
princ.append(temp[:])
temp.clear()
resp=str(input('Quer continuar? [S/N] '))
if resp in 'Nn':
break
print('-='*30)
print(f'Ao todo, você cadastrou {len(princ)} pessoas. ')
print(f'O maior peso foi de {mai} kg. Peso de ', end='')
for p in princ:
if p[1] == mai:
print(f'[{p[0]}] ', end='')
print()
print(f'O menor peso foi de {men} kg. Peso de ', end='')
for p in princ:
if p[1] == men:
print(f'[{p[0]}] ', end='')
print()
# + colab={"base_uri": "https://localhost:8080/"} id="CtU32LFNIdu9" outputId="df0d4d5f-88c6-42ae-cc5b-259dccbf4e5c"
#Desafio 085 - Usuario digita 7 valores numa lista. Nessa lista, ha separacao dos pares e dos impares.
#No final, mostre os valores pares e impares em ordem crescente
num=[[],[]]
for i in range(1,8):
a=int(input('Digite um valor: '))
if a%2 == 0:
num[1].append(a)
i+=1
else:
num[0].append(a)
i+=1
print(f'Os valores ímpares foram: {num[0]} ')
print(f'Os valores pares foram: {num[1]} ')
# + colab={"base_uri": "https://localhost:8080/"} id="GnL3qcQcPOrQ" outputId="71a44f5c-9f9f-4c8a-c0c3-98afe3dded24"
#Desafio 086 - Matriz 3x3 preenchida casa por casa. Mostrar a matriz inteira com formatacao correta
matriz=[[0,0,0],[0,0,0],[0,0,0]]
for i in range(0,3):
for j in range(0,3):
matriz[i][j] = int(input(f'Digite um valor para {i}{j}: '))
print('-='*30)
for i in range(0,3):
for j in range(0,3):
print(f'[{matriz[i][j]:^5}]', end='')
print()
# + colab={"base_uri": "https://localhost:8080/"} id="yoMee23tVUHm" outputId="72d09652-a47b-4cb1-a634-ecf681ec3975"
#Desafio 087 - A mesma matriz do desafio 086. A soma dos valores pares digitados,
#a soma dos valores da terceira coluna, o maior valor da segunda linha.
matriz=[[0,0,0],[0,0,0],[0,0,0]]
sum=0
col=0
lin=0
for i in range(0,3):
for j in range(0,3):
matriz[i][j] = int(input(f'Digite um valor para {i}{j}: '))
lin=matriz[i][1]
if matriz[i][j]%2 == 0:
sum=sum+matriz[i][j]
if j==2:
col=col+matriz[i][j]
if i==1:
if matriz[i][j] > lin:
lin=matriz[i][j]
print('-='*30)
for i in range(0,3):
for j in range(0,3):
print(f'[{matriz[i][j]:^5}]', end='')
print()
print(f'A soma de todos os valores pares: {sum}\n')
print(f'A soma dos valores da terceira coluna: {col}\n')
print(f'O maior valor da segunda linha: {lin}\n')
# + colab={"base_uri": "https://localhost:8080/"} id="8LaVdTLIe_Ij" outputId="95959f28-c862-40e4-cf96-ada34a41d872"
#Desafio 088 - Mega sena. Digitar quantos jogos serão feitos e o programa gera os numeros.
from random import randint
lista=[]
jogo=[]
a=int(input('Quantos jogos você quer? '))
i=1
while i<=a:
cont=0
while True:
b=randint(1,60)
if b not in lista:
lista.append(b)
cont+=1
if cont>6:
break
lista.sort()
jogo.append(lista[:])
lista.clear()
i+=1
print(f'Os jogos sorteados foram: \n')
for e, f in enumerate(jogo):
print(f'jogo {e+1}: {f}')
# + [markdown] id="KX9Is-zqnud1"
# # **Dicionários**
# Identificado por '{}' ou 'dict()'
# + colab={"base_uri": "https://localhost:8080/"} id="78nY_ETtp5mf" outputId="5e34fd7a-1967-48cd-c0ed-8a537acc65f1"
dados={'nome':'Pedro', 'idade':25}
print(dados['nome'])
dados['sexo']='M'
print(dados)
# + colab={"base_uri": "https://localhost:8080/"} id="pEnqxwKEqa5L" outputId="59bcd4a7-5ea1-4a44-f596-d48c8ae6423c"
dados={'nome':'Pedro', 'idade':25}
del dados['idade']
dados
# + colab={"base_uri": "https://localhost:8080/"} id="3vfXrmI4rIIu" outputId="10a7ff8d-8147-4cb2-d2c1-29123294e9e8"
dados={'nome':'Pedro', 'idade':25}
print(dados['nome'])
dados['sexo']='M'
print(dados.keys())
print(dados.items())
print(dados.values())
# + colab={"base_uri": "https://localhost:8080/"} id="kiBK6ErprfgW" outputId="b900a330-37e7-4224-aaba-1356f9c409a3"
filme={'titulo':'Star Wars', 'ano':1977, 'diretor':'<NAME>'}
print(filme)
for k, v in filme.items():
print(f'o {k} é {v}')
# + colab={"base_uri": "https://localhost:8080/"} id="uPp0ZETovRNb" outputId="d90c2ee9-06ed-490e-fe72-2ad64c1abd60"
estado=dict()
brasil=list()
for c in range(0,3):
estado['uf']=str(input('Unidade Federativa: '))
estado['sigla']=str(input('Sigla do estado: '))
brasil.append(estado.copy())
for e in brasil:
for k, v in e.items():
print(f'O campo {k} tem valor {v}.')
# + colab={"base_uri": "https://localhost:8080/"} id="xlJktjLyxRZ0" outputId="97d184e5-9d0a-4b55-fad3-8170c597f93c"
#Desafio 090 - Nome, media e situacao do aluno. Mostrar tudo
aluno={}
aluno['Nome']=str(input('Digite o nome: '))
aluno['Media']=float(input('Digite a media: '))
if aluno['Media']>= 7:
aluno['Situacao']='Aprovado'
else:
aluno['Situacao']='Reprovado'
print(aluno[])
# + colab={"base_uri": "https://localhost:8080/"} id="EcBr0eYtbV1R" outputId="e0a24d69-c6ce-49c7-fda0-bcd237176001"
#Desafio 091 - 4 jogadores jogando um dado. Mostrar a ordem de ranking
from random import randint
#itemgetter para ordenar os valores do ranking
from operator import itemgetter
jogo={'jogador1':randint(1,6), 'jogador2':randint(1,6),
'jogador3': randint(1,6), 'jogador4':randint(1,6),
'jogador5':randint(1,6), 'jogador6': randint(1,6)}
ranking=list()
print('Valores sorteados')
for k, v in jogo.items():
print(f'{k} tirou {v}')
ranking = sorted(jogo.items(), key=itemgetter(1), reverse=True)
print('-='*30)
for i, v in enumerate(ranking):
print(f'{i+1}º lugar: {v[0]} com {v[1]}.')
# + colab={"base_uri": "https://localhost:8080/"} id="rvnpFOGOgHGT" outputId="96a10898-44e6-428c-a44f-26449ea4a972"
#Desafio 092 - Programa que leia nome, ano de nascimento, carteira de trabalho (se for diferente de zero, ler o ano de contratacao e salario).
#Calcular a idade e com quantos anos ela vai se aposentar
pessoa={}
pessoa['nome']=str(input('Digite o nome: '))
pessoa['idade']=2021-int(input('Ano de nascimento: '))
pessoa['CTPS']=int(input('Número de CTPS: (0 para não)'))
if pessoa['CTPS']!=0:
pessoa['contratacao']=int(input('Qual o ano de contratação? '))
pessoa['salario']=int(input('Qual é o salário? '))
pessoa['aposentadoria']=pessoa['contratacao']+35
print(pessoa)
# + colab={"base_uri": "https://localhost:8080/"} id="Fqwvz3D3lF6K" outputId="b9d2d8b8-57ab-4bbf-d7df-95f97cf253f6"
#Desafio 094 - Ler nome, sexo e idade de varias pessoas e salvar tudo numa lista.
#Mostrara media de idade do grupo, uma lista com as mulheres e uma lista com todas as pessoas com idade acima da media
pessoas={}
galera=[]
soma=media=0
while True:
pessoas.clear()
pessoas['nome']=str(input('Digite o nome: '))
while True:
pessoas['sexo']=str(input('Digite o sexo (M/F): ')).strip().upper()[0]
if pessoas['sexo'] in 'MF':
break
print('Por favor digite apenas M ou F')
pessoas['idade']=int(input('Digite a idade: '))
soma+=pessoas['idade']
galera.append(pessoas.copy())
while True:
r=str(input('Quer continuar? (S/N)')).strip().upper()[0]
if r in 'SN':
break
print('Digite apenas S ou N')
if r=='N':
break
print('-='*30)
print(f'Ao todo temos {len(galera)} cadastrados.')
media=soma/len(galera)
print(f'A média de idade é {media:5.2f} anos')
print(f'As mulheres cadastradas foram ', end='')
for p in galera:
if p['sexo'] in 'Ff':
print(f'{p["nome"]} ', end='')
print()
print('Lista de pessoas que estão acima da média: ')
for p in galera:
if p['idade']>media:
print(' ')
for k, v in p.items():
print(f'{k} = {v}; ', end='')
print()
print('Encerrado')
# + [markdown] id="2jdwIwurwufO"
# # **Funções**
# Uma rotina. def xxx():
# + colab={"base_uri": "https://localhost:8080/"} id="yx4wMaRC3a6I" outputId="73783404-20ec-4607-8e3e-4b091dd57120"
#Desempacotamento
def contador(*num):
print(num)
contador(2,3,4)
contador(8,65,3,5,78,9)
# + colab={"base_uri": "https://localhost:8080/"} id="TYL2oWNQ7lQl" outputId="cc996a46-5999-4c0a-a371-d6d605457c53"
#Desafio 096 - Ler largura e comprimento, calcular a area
def calculo(a, b):
area=a*b
print(f'A área do terreno {a}x{b} é {area} metros quadradados.')
l=int(input('Digite a largura: '))
c=int(input('Digite o comprimento: '))
calculo(l,c)
# + colab={"base_uri": "https://localhost:8080/"} id="pnCO_sn36Lyj" outputId="374c1ea9-6a6e-41d7-ed6c-892f51be275e"
#Desafio 097 - Um print especial
def escreva(msg):
tam= len(msg) +4
print('~' * tam)
print(f' {msg} ')
print('~' * tam)
escreva('<NAME>')
escreva('Curso em Vídeo')
# + colab={"base_uri": "https://localhost:8080/"} id="BsCYYCZM8GlN" outputId="ad58f1e7-b955-4edd-b916-0a97a4ed6872"
#Desafio 098 - Contador
from time import sleep
def contador(i, f, p):
print('-='*30)
print(f'Contagem de {i} até {f} ao passo de {p}')
if p<0:
p*=-1
if p==0:
p=1
if i<f:
cont=i
while cont <= f:
print(f'{cont} ', end='', flush=True)
sleep(0.5)
cont +=p
print('FIM')
else:
cont=i
while cont>= f:
print(f'{cont} ', end='', flush=True)
sleep(0.5)
cont -=p
print('FIM')
contador(1,10,1)
contador(10,0,2)
print('-='*30)
ini=int(input('Digite o início: '))
fim=int(input('Digite o final: '))
pas=int(input('Digite o passo: '))
contador(ini, fim, pas)
# + colab={"base_uri": "https://localhost:8080/"} id="oecufLavOzhG" outputId="2938c93a-09c1-40df-b208-ddebcf3d27ff"
#Desafio 099 - Chamar o maior numero
from time import sleep
def maior(*num):
cont=maior=0
print('-='*30)
print('Analisando os valores passados...')
for val in num:
print(f'{val} ', end='', flush=True)
sleep(0.1)
if cont ==0:
maior=val
else:
if val>maior:
maior=val
cont += 1
print(f'\nO maior valor foi {maior} ')
maior(3,9,88,5,1,4,3,5)
# + colab={"base_uri": "https://localhost:8080/"} id="GYgHvXqCTQQH" outputId="f68433c1-7cb4-4cfd-cf93-405c5c474741"
#Desafio 100 - Sortear e somar os pares
from random import randint
num=[]
def sorteia(lista):
for i in range(0,5):
a=randint(1,10)
lista.append(a)
print(lista)
def somapar(lista):
i=0
soma=0
while i<len(lista):
if lista[i]%2 ==0:
soma=soma+lista[i]
i+=1
print(f'\nA soma dos valores pares resulta em: {soma}')
sorteia(num)
somapar(num)
# + colab={"base_uri": "https://localhost:8080/"} id="4ucW4uN6aMih" outputId="15203f9c-19fe-4e02-89d6-62753606583a"
#Descritivo da funcao
help(print)
# + colab={"base_uri": "https://localhost:8080/"} id="VKjqINzOak9s" outputId="4a3a9b49-c5e6-44a3-f147-167c16206f25"
#Outro descritivo de funcao
print(input.__doc__)
# + colab={"base_uri": "https://localhost:8080/"} id="94mmS75XdLxF" outputId="54ae9d56-44c1-4ddd-9976-030deef67c83"
def somar(a=0, b=0, c=0):
s=a+b+c
return s
r1=somar(1,2,3)
r2=somar(1)
r3=somar(20,1)
print(f'Os resultados foram: {r1}, {r2}, {r3}')
# + id="vE8J4hQsfdOp"
#declarar uma variavel global dentro de uma funcao
#global a
# + id="c_JFGaT_gS2E"
def somar(a=0, b=0, c=0):
s=a+b+c
print(f'A soma vale {s} ')
somar(1,2,3)
somar(1)
# + colab={"base_uri": "https://localhost:8080/"} id="S6D1Vp_fjc_p" outputId="007ae772-e482-4378-813d-e2f3232c3d6a"
def fatorial(num=1):
f=1
for c in range(num, 0, -1):
f*=c
return f
f1=fatorial(5)
f2=fatorial(4)
f3=fatorial()
print(f'{f1}, {f2}, {f3}')
# + colab={"base_uri": "https://localhost:8080/", "height": 52} id="0FVSVbYIk5bq" outputId="cd71e7dd-7fa7-4e57-cafb-9846b7274e6f"
#Desafio 101 - Votacao negado, opcional ou obrigatorio
def voto(idade):
if idade>16:
if idade>17 and idade<=70:
t='Voto obrigatório'
return t
else:
t='Voto opcional'
return t
else:
t='Voto negado'
return t
i=int(input('Digite a idade: '))
voto(i)
# + colab={"base_uri": "https://localhost:8080/"} id="Z4ORS7Puo4t1" outputId="8c537054-5b98-488d-d5ec-6d4071e4f5a9"
#Desafio 102 - Calculo de fatorial e paramentro show (mostrar o calculo)
def fatorial(num, show=False):
"""
Calcula o fatorial de um número. Se show for verdadeiro, mostrar o calculo
"""
f=1
for c in range(num, 0, -1):
f*=c
if show:
print(c, end='')
if c>1:
print(' X ', end='')
else:
print(' = ', end='')
return f
print(fatorial(5, show=True))
# + colab={"base_uri": "https://localhost:8080/"} id="ccXAMZ7BRdcW" outputId="96a801cf-fd90-44c6-ed1e-4c2f577ada07"
#Desafio 103 - Jogador (nome e gols)
def ficha(nome='<desconhecido>', gol=0):
print(f'O jogador {nome} fez {gol} gol(s) no campeonato!')
n=str(input('Digite o nome do jogador: '))
g=str(input('Digite o número de gols: '))
if g.isnumeric():
g=int(g)
else:
g=0
if n.strip()=='':
ficha(gol=g)
else:
ficha(n,g)
# + colab={"base_uri": "https://localhost:8080/"} id="9fZNFEfVa2Pm" outputId="47f5aa70-36d3-4cbf-dd6b-03c078012898"
#Desafio 104 - criar uma funcao leiaint() que aceita apenas numeros inteiros
def leiaInt(msg):
"""
Similar a funcao 'input'
"""
ok = False
valor = 0
while True:
n=str(input(msg))
if n.isnumeric():
valor = int(n)
ok = True
else:
print('Digite um número inteiro! ')
if ok:
break
return valor
n=leiaInt('Digite um número: ')
print(f'Você digitou {n}')
# + colab={"base_uri": "https://localhost:8080/"} id="QRUn8rCBlCuR" outputId="28a58b6b-7fb0-4edb-c646-f10d2bf8d7a1"
#Desafio 105 - Dicionario de notas
def notas(*n, sit=False):
"""
sit para mostrar a situacao
"""
r=dict()
r['total']=len(n)
r['maior']=max(n)
r['menor']=min(n)
r['media']=sum(n)/len(n)
if sit:
if r['media']>=7:
r['situacao']='Boa'
elif r['media']>=5:
r['situacao']='Razoavel'
else:
r['situacao']='Ruim'
return r
resp=notas(5.5,9.5,10,6.5, sit=True)
print(resp)
#help(notas)
| Curso_de_Python_3_Mundo_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Facial Recognition Using Vgg Face
#
#
# The VGG-Face is a deep-CNN created by Department of Engineering Science, University of Oxford. [HOMEPAGE CLICK HERE](http://www.robots.ox.ac.uk/~vgg/research/very_deep/)
#
# This deep-CNN has been trained using Softmax loss to recognize faces of 2,622 celebrity identities. It implements VGG-Very-Deep-16 CNN architecture as described in a [paper](https://www.robots.ox.ac.uk/~vgg/publications/2015/Parkhi15/parkhi15.pdf) published in 2015.
#
# The model architecture is a linear sequence of layer transformations of the following types:
# * Convolution + ReLU activations
# * MaxPooling
# * softmax
# <img src="https://image.slidesharecdn.com/tfmandreavideoretrievalofspecificpersonsinspecificlocations-160920130647/95/video-retrieval-of-specific-persons-in-specific-locations-23-638.jpg?cb=1474377027" height="311" width="533">
#
# ## Vgg16 vs Vgg Face
#
# Briefly, the VGG-Face model is the same NeuralNet architecture as the VGG16 model used to identity 1000 classes of object in the ImageNet competition.
# The VGG16 name simply states the model originated from the Visual Geometry Group and that it was 16 trainable layers.
# The main difference between the VGG16-ImageNet and VGG-Face model is the set of calibrated weights as the training sets were different.
#
# ## In this jupyter notebook, we did:
# * Load vgg-face
# * Detect and crop faces from input picture using OpenCV
# * Predict name from crpped faces
# * Remove image mean and predict again
# Load we load some important standard packages.
import matplotlib.pyplot as plt
import matplotlib.image
import skimage.io
import skimage.transform
from PIL import Image
import numpy as np
import copy
# %matplotlib inline
# Then we load the appropriate Keras packages.
import keras.backend as K
from keras_vggface.vggface import VGGFace
from keras.models import Sequential, Model
from keras.layers import Input, Dense, Flatten, Dropout, Activation, Lambda, Permute, Reshape
from keras.layers import Convolution2D, ZeroPadding2D, MaxPooling2D
# Clear the Keras session.
K.clear_session()
# Set the dimensions of the input image. Vgg-face only takes input size as (244,244,3).
nrow = 224
ncol = 224
# ## Using pre trained vgg face network
#
# Create a model using VGG-Face and its pre-trained weight matrix.
#
# When you use it for the first time ,weights are downloaded and stored in ~/.keras folder.
#
# It's very important to set input shape.
model = VGGFace(model='vgg16',
weights='vggface',
input_shape=(nrow,ncol,3))
model.summary()
# We import the weight matrix by ourself and save the names in description.
# +
from scipy.io import loadmat
data = loadmat('vgg_face_matconvnet/data/vgg_face.mat',
matlab_compatible=False,
struct_as_record=False)
net = data['net'][0,0]
l = net.layers
description = net.classes[0,0].description
print(description)
# -
# Let use a photo of <NAME> to test how Vgg Face predicts.
# What should be known is that this photo is well cropped and resized and centralized to satify the input requirements of vgg face. Most of the pictures found on the Internet will not be this ideal.
# +
imagePath = 'ak.png'
image = Image.open(imagePath)
plt.imshow(image)
plt.xticks([])
plt.yticks([])
plt.show()
# -
# Before predict, we should transform this image into a (224,224,3) numpy array with float values. Also, check the summary of the model, we could find that the output shape of first input layer is (None, 224, 224, 3), so this numpy array should expand to four dimensions to fit in this model.
#
# After prediction, we find the final output is a 2622 array, which contains the posibilities of whether this photo should be someone. For example, if the third value of output is 0.004, which means that this face is less likely to be the person labeled no.3. And if the fifth value is 0.974, then the fifth name in the dataset is most likely to have this face.
# +
imarr = np.array(image).astype(np.float32)
imarr = np.expand_dims(imarr, axis=0)
out = model.predict(imarr)
print(out.shape)
print(out)
# -
# So we need to find the max value of the predict output. Since it is probablity, we need to save the index of this max value, and look for the name according to this index in description, the data we extract from weight matrix.
#
# The result shows it is 96.2% to be Aamia Khan, which is quite nice.
max_index = np.argmax(out, axis=1)[0]
max_name = description[max_index,0]
print(max_index, max_name[0], out[0,max_index])
# ## Detect and crop faces from normal photos
# ## Using OpenCV
#
# Vgg Face is trained with well prepared photos. Let's look at the very first picture in vgg face dataset. It is picture of A.C.Burkey. But in its dataset, the detail of this picture is 'http://www.contactmusic.com/pics/ld/active_for_life_arrivals_090110/a.j_buckley_2706152.jpg 165.21 105.50 298.57 238.86 4.00 3.59 1
# '. The numbers after url point to the location of his face. And when training, they only use his face as the input, instead of the whole picture.
# <img src="http://www.contactmusic.com/pics/ld/active_for_life_arrivals_090110/a.j_buckley_2706152.jpg" height="500" width="160">
#
# So the next thing we should do is to prepare input photos download from internet to only have the face. Of course we can't just reshape the picture because it might twist it. And we can;t just the same method in MINST lab, because we are finding faces this time, and faces are found by facial features. Also, most of the pictures of celebrities would includes other celebrities as well. Thinking of finding your favorite actor in a group photo of oscar. We need some other method to find faces.
# <img src="https://www.nanrenwo.net/uploads/allimg/151120/8388-1511200Z418-51.jpg" height="240" width="260">
#
# We considered using OpenCV to find faces.OpenCV is the most popular library for computer vision. It uses machine learning algorithms to search for faces within a picture. For something as complicated as a face, there isn’t one simple test that will tell you if it found a face or not. Instead, there are thousands of small patterns/features that must be matched. The algorithms break the task of identifying the face into thousands of smaller, bite-sized tasks, each of which is easy to solve. These tasks are also called classifiers. OpenCV uses cascades to detect these features.
#
# Casade is like a series of waterfalls, the OpenCV cascade breaks the problem of detecting faces into multiple stages. For each block, it does a very rough and quick test. If that passes, it does a slightly more detailed test, and so on. The algorithm may have 30-50 of these stages or cascades, and it will only detect a face if all stages pass. The advantage is that the majority of the pictures will return negative during the first few stages, which means the algorithm won’t waste too much time detect thounands of features.
#
# The cascades themselves are just a bunch of XML files that contain OpenCV data used to detect objects.
# Now we create the cascade and initialize it with our face cascade. We use haarcascade_frontalface_default.xml. It can be downloaded from [here](http:https://github.com/opencv/opencv/blob/master/data/haarcascades/haarcascade_frontalface_default.xml).
#
# This loads the face cascade into memory so it’s ready for use.
# +
import cv2
imagePath = 'cf.jpg'
faceCascade = cv2.CascadeClassifier("haarcascade_frontalface_default.xml")
# Read the image
# Convert it to grayscale. Many operations in OpenCv are done in grayscale.
image = cv2.imread(imagePath)
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
# -
# Then we use the function detectMultiScale to detect faces.
#
# * The detectMultiScale function is a general function that detects objects. Since we are calling it on the face cascade, that’s what it detects. The first option is the grayscale image.
#
# * The second is the scaleFactor. Since some faces may be closer to the camera, they would appear bigger than those faces in the back. The scale factor compensates for this.
#
# * The detection algorithm uses a moving window to detect objects. minNeighbors defines how many objects are detected near the current one before it declares the face found. minSize, meanwhile, gives the size of each window.
#
# The function returns a list of rectangles where it believes it found a face.
# We use the picture of <NAME> and <NAME>.
# +
# Detect faces in the image
faces = faceCascade.detectMultiScale(
gray,
scaleFactor=1.2,
minNeighbors=5,
minSize=(30, 30)
)
faces = faceCascade.detectMultiScale(gray, 1.2, 5)
print("Found {0} faces!".format(len(faces)))
print(faces)
# +
# Draw a rectangle around the faces
for (x, y, w, h) in faces:
cv2.rectangle(image, (x, y), (x+w, y+h), (255, 0, 0), 3)
plt.imshow(image)
plt.show()
# -
# Then we created a function called pred to predict the name.
def pred(model, crpimg):
imarr = np.array(crpimg).astype(np.float32)
imarr = np.expand_dims(imarr, axis=0)
out = model.predict(imarr)
max_index = np.argmax(out, axis=1)[0]
max_name = description[max_index,0]
print(max_index, max_name[0], out[0,max_index])
# ## Predict name from crpped faces
#
# Now we crop the faces from the rectangles and use cropped pictures to predict.
#
# First make a function to predict face by face:
def face_recon (im, n, faces, model):
for i in range(len(faces)):
(x, y, w, h) = faces[i]
center_x = x+w/2
center_y = y+h/2
b_dim = min(max(w,h)*1.1,im.width, im.height)
box = (center_x-b_dim/2, center_y-b_dim/2, center_x+b_dim/2, center_y+b_dim/2)
# Crop Image
crpim = im.crop(box).resize((224,224))
plt.imshow(np.asarray(crpim))
plt.show()
pred(model, crpim)
im = Image.open(imagePath)
face_recon(im, len(faces), faces, model)
# Well the prediction works well on Bingbing, but not on Jackie Chan.
#
# Beause the training set of Jackie Chan does not includes pictures of him .
#
# But <NAME> shows around with his glasses every time, and they indeed look like each other. So the prediction is Clark Duke. Which makes sense.
# <img src="https://www.aceshowbiz.com/images/photo/clark_duke.jpg">
#
# So let try with another picture with multiple faces, and with only bare faces with no glasses.
#
# The photo below is the poster of movie *Twilight*. It incldes the three main charaters of it.
#
# The result is good. However, <NAME> has the least possilibility to be himself. This is because he plays a vampire in this movie, and it slightly changes his appearance from real life. And the other two actors are still human in this movie, so the probabilities are higher.
#
# Which indicates that the result is closely related to the training dataset.
# +
imagePath = 'mg.jpg'
faceCascade = cv2.CascadeClassifier("haarcascade_frontalface_default.xml")
image = cv2.imread(imagePath)
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
faces = faceCascade.detectMultiScale(
gray,
scaleFactor=1.2,
minNeighbors=5,
minSize=(30, 30))
faces = faceCascade.detectMultiScale(gray, 1.2, 5)
for (x, y, w, h) in faces:
cv2.rectangle(image, (x, y), (x+w, y+h), (255, 0, 0), 3)
plt.imshow(image)
plt.show()
im = Image.open(imagePath)
face_recon(im, len(faces), faces, model)
# -
# ## Remove image mean and predict again
#
# During our research, we found that when people are using imagenet, they tend to extract mean of the photo before prediction.
#
# So we tried to modified our *predict* function to zero-center by mean pixel before prediction.
#
# Based on [this](https://github.com/rcmalli/keras-vggface) and [this] (https://github.com/keras-team/keras/blob/master/keras/applications/imagenet_utils.py) page.
#
# This function did:
# * Remove image mean
# * im[:, :, 0] -= 93.5940
# * im[:, :, 1] -= 104.7624
# * im[:, :, 2] -= 129.1863
#
# +
from keras_vggface import utils
def modified_pred(model, crpimg):
imarr = np.array(crpimg).astype(np.float32)
imarr = np.expand_dims(imarr, axis=0)
# Extract mean frome picture
imarr = utils.preprocess_input(imarr, version=1)
out = model.predict(imarr)
max_index = np.argmax(out, axis=1)[0]
max_name = description[max_index,0]
print(max_index, max_name[0], out[0,max_index])
def modified_face_recon (im, n, faces, model):
for i in range(len(faces)):
(x, y, w, h) = faces[i]
center_x = x+w/2
center_y = y+h/2
b_dim = min(max(w,h)*1.1,im.width, im.height)
box = (center_x-b_dim/2, center_y-b_dim/2, center_x+b_dim/2, center_y+b_dim/2)
# Crop Image
crpim = im.crop(box).resize((224,224))
plt.imshow(np.asarray(crpim))
plt.show()
modified_pred(model, crpim)
# -
# Let's do the prediction on *Twilight* again,
modified_face_recon(im, len(faces), faces, model)
# Comparing to the result without mean extracting, this result is slightly worse.
#
# But [this](http://www.pythonexample.com/code/a-basic-interpersonal-communication-model/) code paper says this would be more robust.
| Recognition-Algorithms/Face Recognition using VGG/Training VGG Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="bo0eLE4v4IYE"
# # Removing noise from *K2* and *TESS* light curves using Pixel Level Decorrelation (`PLDCorrector`)
# + [markdown] id="A9eLo3kLAn9E"
# ## Learning Goals
#
# By the end of this tutorial, you will:
#
# * Understand how to apply the [Lightkurve](https://docs.lightkurve.org) [PLDCorrector](https://docs.lightkurve.org/reference/api/lightkurve.correctors.PLDCorrector.html?highlight=pldcorrector) tool to remove instrumental noise from *K2* and *TESS* light curves.
# * Be able to create an exoplanet transit mask and use it to improve PLD.
# * Be aware of common issues, caveats, and potential biases associated with the use of PLD.
# + [markdown] id="wMnkYd8EA40B"
# ## Introduction
#
# The [*K2*](https://archive.stsci.edu/k2) and [*TESS*](https://archive.stsci.edu/tess) missions both provide high-precision photometry for thousands of exoplanet candidates. However, observations by both telescopes can be muddled by instrumental systematic trends, making exoplanet detection or stellar characterization difficult.
#
# Pixel Level Decorrelation (PLD) is a method that has primarily been used to remove systematic trends introduced by small spacecraft motions during observations, and has been shown to be successful at improving the precision of data taken by the *Spitzer* space telescope ([Deming et al. 2015](https://ui.adsabs.harvard.edu/abs/2015ApJ...805..132D/abstract)) and the *K2* mission ([Luger et al. 2016](https://ui.adsabs.harvard.edu/abs/2016AJ....152..100L/abstract); [2018](https://ui.adsabs.harvard.edu/abs/2018AJ....156...99L/abstract)). PLD works by identifying a set of trends in the pixels surrounding the target star, and performing linear regression to create a combination of these trends that effectively models the systematic noise introduced by spacecraft motion. This noise model is then subtracted from the uncorrected light curve.
#
# This method has been shown to be very effective at removing the periodic systematic trends in *K2*, and can also help remove the scattered light background signal in *TESS* observations. This tutorial will demonstrate how to use the [Lightkurve](https://docs.lightkurve.org) [PLDCorrector](https://docs.lightkurve.org/reference/api/lightkurve.correctors.PLDCorrector.html?highlight=pldcorrector) for each mission, and will give advice on how to best implement PLD. The [PLDCorrector](https://docs.lightkurve.org/reference/api/lightkurve.correctors.PLDCorrector.html?highlight=pldcorrector) is a special case of the [Lightkurve](https://docs.lightkurve.org) [RegressionCorrector](https://docs.lightkurve.org/reference/api/lightkurve.correctors.RegressionCorrector.html?highlight=regressioncorrector). For more information on how to use [RegressionCorrector](https://docs.lightkurve.org/reference/api/lightkurve.correctors.RegressionCorrector.html?highlight=regressioncorrector) to choose custom regressors and remove scattered light from *TESS*, please see the tutorial specifically on removing scattered light from *TESS* data using the Lightkurve [RegressionCorrector](https://docs.lightkurve.org/reference/api/lightkurve.correctors.RegressionCorrector.html?highlight=regressioncorrector).
#
# Before reading this tutorial, it is recommended to first familiarize yourself with using target pixel file (TPF) products and light curve products with Lightkurve.
# + [markdown] id="5wOFCyG74-fA"
# ## Imports
#
# We only need to import the **[Lightkurve](https://docs.lightkurve.org)** package for this tutorial, which in turn uses **[Matplotlib](https://matplotlib.org/)** for plotting.
# + id="d-idwZ_c5GJp"
import lightkurve as lk
# %matplotlib inline
# + [markdown] id="O9KCiDg2sCmc"
# ---
# + [markdown] id="VqvB7m3c1wbF"
# ## 1. Applying PLD to a *K2* Light Curve
#
# The dominant source of noise in *K2* data is created by the motion of the *Kepler* spacecraft due to periodic thruster firings. This causes stars to drift across different pixels on the detector, which have varied sensitivity. There are two classes of sensitivity variation on a charge-couped device (CCD) detector — variation between pixels (inter-pixel) and variation within each pixel (intra-pixel). Both inter- and intra-pixel sensitivity variations are present on the *Kepler* detector, which ultimately causes different flux levels to be detected as the target's Point Spread Function (PSF) drifts across the variations, introducing the systematic trends.
#
# The [PLDCorrector](https://docs.lightkurve.org/reference/api/lightkurve.correctors.PLDCorrector.html?highlight=pldcorrector) uses information from nearby pixels to create a noise model, so we need to use the [TargetPixelFile](https://docs.lightkurve.org/reference/targetpixelfile.html?highlight=targetpixelfile) data product (for more information, see the tutorial on using *Kepler* target pixel file products). We can use the [search_targetpixelfile](https://docs.lightkurve.org/reference/api/lightkurve.search_targetpixelfile.html?highlight=search_targetpixelfile) method to identify available observations for the desired target, and the [download](https://docs.lightkurve.org/reference/search.html?highlight=download) method to access the data.
#
# In what follows below, we will demonstrate PLD on the exoplanet system K2-199, which was observed during *K2* Campaign 6. We can download the pixel data as follows:
# + colab={"base_uri": "https://localhost:8080/", "height": 420} executionInfo={"elapsed": 11452, "status": "ok", "timestamp": 1601325033072, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="hOFE_Ab19QXL" outputId="1651145e-c9c3-4542-cb2d-2c6a99d56ea8"
tpf = lk.search_targetpixelfile('K2-199', author='K2', campaign=6).download()
tpf.plot();
# + [markdown] id="Re-Wj1Bn6wuZ"
# There are two ways to create a `PLDCorrector` object. The first is to create an instance of the class directly and pass in the `TargetPixelFile`.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"elapsed": 1246, "status": "ok", "timestamp": 1601325069125, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="0yOLHCKJ1qJc" outputId="dcd72580-46bd-495b-b4cb-468a8c6c5bff"
from lightkurve.correctors import PLDCorrector
pld = PLDCorrector(tpf)
print(pld)
# + [markdown] id="4n_9tbJA7DsT"
# For convenience, you can also use the [to_corrector](https://docs.lightkurve.org/reference/api/lightkurve.LightCurve.to_corrector.html?highlight=to_corrector#lightkurve.LightCurve.to_corrector) method of the `TargetPixelFile` object, and pass in the string `'pld'` to specify the desired corrector type.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} executionInfo={"elapsed": 952, "status": "ok", "timestamp": 1601325073513, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="9NcNvXn4AN-w" outputId="72746534-8df3-4aa0-a6db-c28d5c3b5bcf"
pld = tpf.to_corrector('pld')
print(pld)
# + [markdown] id="IeiKtfNZ7f5G"
# Both of these approaches return an identical `PLDCorrector` object. From here, getting a corrected light curve is possible: call the `correct` method.
# + id="SI8OAsVn8Cg4"
corrected_lc = pld.correct()
# + [markdown] id="Elc6Dlyt8I_T"
# Now we can compare the output of PLD to an uncorrected light curve. To create the uncorrected light curve, we can use the `to_lightcurve` method of the [KeplerTargetPixelFile](https://docs.lightkurve.org/reference/api/lightkurve.KeplerTargetPixelFile.html?highlight=keplertargetpixelfile) object, which performs simple aperture photometry (SAP) to create a light curve from the pixel data.
#
# Below, the uncorrected light curve is shown in red and the PLD-corrected light curve is plotted in black.
# + colab={"base_uri": "https://localhost:8080/", "height": 389} executionInfo={"elapsed": 1791, "status": "ok", "timestamp": 1601325094416, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="dlCmbjzF8OlH" outputId="8e3e0893-9d39-4596-bf40-712a40de86b9"
# Create and plot an uncorrected light curve using Simple Aperture Photometry
uncorrected_lc = tpf.to_lightcurve()
ax = uncorrected_lc.normalize().scatter(color='red', label='Uncorrected Light Curve');
# Plot the PLD-corrected light curve in black on top
corrected_lc.normalize().remove_outliers().scatter(ax=ax, color='black', label='PLD-corrected Light Curve');
# + [markdown] id="aa_H_3yIpeub"
# The uncorrected light curve is dominated by a short period (about six hours) sawtooth-shaped pattern caused by the *Kepler* spacecraft thruster firings. PLD captures this trend in the noise model and has subtracted it, leaving the much more accurate light curve in black.
#
# We can quantify the improvement by comparing the Combined Differential Photometric Precision (CDPP) values for each light curve.
# + colab={"base_uri": "https://localhost:8080/", "height": 51} executionInfo={"elapsed": 884, "status": "ok", "timestamp": 1601325115841, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="cCN34aicNkkS" outputId="3dd21702-af7f-480a-ad4b-cec3749422ae"
uncorrected_cdpp = uncorrected_lc.estimate_cdpp()
corrected_cdpp = corrected_lc.estimate_cdpp()
print(f"Uncorrected CDPP = {uncorrected_cdpp:.0f}")
print(f"Corrected CDPP = {corrected_cdpp:.0f}")
# + [markdown] id="i6EwwaxuNk1b"
# By this metric, the photometric precision improved by more than a factor of 25 after applying PLD.
#
# Another important trait of PLD is that long-term variability trends in the light curve are preserved. In this example, we can see the stellar rotation of K2-199 as the sinusoidal signal left in the light curve after correction. This is done by fitting a polynomial spline model to the light curve while simultaneously fitting the noise model, because the uncorrected observation is a combination of the signals.
# + [markdown] id="DjiRK0mHkEAS"
# ## 2. Diagnosing the Success of the Correction
#
# The success of PLD depends on a number of factors including the brightness of the object, the choice of pixels used to create the light curve and the noise model, and whether or not there exists a correlation between the instrumental noise and the astrophysical signals. For these reasons, it is important to carefully review the correct operation of the algorithm each time you use it, and tune the optional parameters of the algorithm if necessary.
#
# The most convenient way to diagnose the performance of PLD is to use the [diagnose()](https://docs.lightkurve.org/reference/api/lightkurve.correctors.PLDCorrector.diagnose.html?highlight=diagnose#lightkurve.correctors.PLDCorrector.diagnose) method, which generates a set of diagnostic plots which we will explain below the graph.
# + colab={"base_uri": "https://localhost:8080/", "height": 764} executionInfo={"elapsed": 3724, "status": "ok", "timestamp": 1601325169792, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="RlPUcH_8pbpT" outputId="16533b42-d387-4e4c-8b5d-f9343d045f54"
pld.diagnose();
# + [markdown] id="Jhe5lRIksuAs"
# The diagnostic plot is composed of the following panels:
# - **Top panel**: The uncorrected light curve (`original`) is shown in black, with the Combined Differential Photometric Precision (CDPP), a measurement of the light curve's scatter, noted in the legend.
# - **Middle panel**: The PLD-corrected light curve (`corrected`) is plotted in gray with the noise model created by the combination of pixel values (`pixel_series`) in blue, the estimated background trend (`background`) in red, and the polynomial spline fit to the stellar variability (`spline`) in yellow. For *K2*, the background signal is minimal compared to the systematic trends due to motion, but it is much more significant for *TESS* observations. Notice that the corrected light curve closely matches the spline, which is tracing the preserved stellar variability.
# - **Bottom panel**: A direct comparison between the uncorrected light curve (`original`) and the PLD-corrected light curve (`corrected`), again noting the CDPP of each light curve in the legend. This panel also indicates which cadences were flagged as outliers (`outlier_mask`), which lie greater than five standard deviations above or below the light curve, as well as which cadences are excluded from the spline fit (`~cadence_mask`). As we will see below, `cadence_mask` is a Boolean array with the same length as the `TargetPixelFile`'s time array, where `True` is included in the spline fit and `False` is excluded. The tilde (`~`) indicates that the inverse of the mask is marked in this plot, that is, the excluded cadences will be crossed out in blue.
# + [markdown] id="lo-1ipZkkPyD"
# The plot above makes it convenient to review the components of the noise removal algorithm. The performance of the algorithm is strongly affected by the choice of pixels which go into the model components. To diagnose this part of the algorithm, we can use the `diagnose_masks` method:
# + colab={"base_uri": "https://localhost:8080/", "height": 391} executionInfo={"elapsed": 3398, "status": "ok", "timestamp": 1601325212709, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="Z9vnchy_kN3q" outputId="5db05284-7563-4a27-a9e9-85221f801c64"
pld.diagnose_masks();
# + [markdown] id="eH2Lh5LswENV"
# The panels in the figure visualize the following pixel masks:
#
# * `aperture_mask`: Pixels used to create the SAP flux light curve from which the noise model is subtracted.
#
# * `pld_aperture_mask`: Pixels used to create the correlated noise model.
#
# * `background_aperture_mask`: Pixels used to create the background model.
#
#
# You can alter the `pld_aperture_mask` and `background_aperture_mask` by passing them as optional arguments to the `correct()` method we used earlier. For more information about these masks, and how to use them effectively, please see the full list of optional parameters and FAQ at the end of this tutorial.
# + [markdown] id="MrIdImCPp8OL"
# ## 3. How to Avoid Overfitting Exoplanet Transits
#
# In the example above, it looks like PLD did a great job of removing the instrumental noise introduced by the *K2* detector drift, but the bottom panel of the diagnostic plot seems to indicate that we've falsely labeled in-transit data points as outliers. To alleviate this, we can introduce the `cadence_mask`.
#
# ### 3.1 Overfitting
#
# It's necessary to mask out transits and flares when fitting the spline. The spline term fits a polynomial to the long-term trend of the light curve, and the highest likelihood solution generally follows the median of the corrected light curve. The presence of transits or flares can pull the median of the light curve down or up, respectively, causing the spline to deviate from the underlying stellar trend.
#
# In practice, this causes transits to be partially "fit out" by the spline, reducing their depth and giving an incorrect estimate of the planet's radius.
#
# You can use a custom `cadence_mask` by creating a Boolean array with one value per cadence, where `True` indicates a cadence you wish to include and `False` means that the cadence is masked out.
#
# For this example, we want to be sure that the in-transit cadences are not marked as outliers or used in the spline fit, as they can cause the spline to erroneously deviate from the stellar signal. To accomplish this, we can create a transit mask using the `create_transit_mask` method of the `LightCurve` object, using the known parameters of the planet system:
# + id="FXIoU0Qfp7oN"
transit_mask = corrected_lc.create_transit_mask(period=[3.2277, 7.3745],
duration=[0.25, 0.25],
transit_time=[2385.6635, 2389.9635])
# + [markdown] id="OxY7dgn1IgYK"
# We can double-check to make sure the transit mask looks good by plotting it on top of the corrected light curve in red:
# + colab={"base_uri": "https://localhost:8080/", "height": 387} executionInfo={"elapsed": 1610, "status": "ok", "timestamp": 1601325273989, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="Tm_ISx4PrAze" outputId="17705db1-6bd7-49c9-b598-459982a57da4"
ax = corrected_lc.scatter(label='Corrected Light Curve')
corrected_lc[transit_mask].scatter(ax=ax, c='r', label='transit_mask');
# + [markdown] id="RdVcFjDZUMDh"
# The red points appear to match the in-transit cadences for both of the planets around K2-199.
#
# An additional option for the `PLDCorrector` is the ability to remove the long-term trend. This is ideal for planet candidates, which can be more difficult to detect in the presence of stellar variability. Here, we can set the `restore_trend` parameter to `False` in order to return a light curve with the long-period trend removed.
#
# Now, we can call the `correct` method of the `PLDCorrector` again, this time passing in our `cadence_mask`.
# + colab={"base_uri": "https://localhost:8080/", "height": 764} executionInfo={"elapsed": 5807, "status": "ok", "timestamp": 1601325293513, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="m52H3ZWIrCg7" outputId="46de7312-77cb-432f-cfe6-4261b9c22b78"
corrected_lc = pld.correct(cadence_mask=~transit_mask, restore_trend=False)
pld.diagnose();
# + [markdown] id="irADOmNjb1G8"
# Now, only points that are greater than five standard deviations above or below the light curve but not in-transit will be marked as outliers, and the in-transit points (marked in blue in the bottom panel) will not be used to fit the spline.
#
# If we examine the light curve and its cadence mask again, we will see that the long-period trend has been removed, and the transits are clearly visible by eye.
# + colab={"base_uri": "https://localhost:8080/", "height": 387} executionInfo={"elapsed": 2127, "status": "ok", "timestamp": 1601325306335, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="abiEX8zWRLrx" outputId="bf915a95-f692-4b0a-a70f-81e4fecc7c60"
ax = corrected_lc.scatter(label='Corrected Light Curve')
corrected_lc[transit_mask].scatter(ax=ax, c='r', label='transit_mask');
# + [markdown] id="i1GKgzCARaOA"
# This correction looks great! We have demonstrated that PLD is effective at removing systematic trends in *K2* data. Now, let's apply PLD to *TESS* observations.
# + [markdown] id="ldY8hLjlKJ-r"
# ## 4. Applying PLD to a *TESS* Light Curve
#
#
# + [markdown] id="ZwG6JkcnVG0a"
# *TESS* has multiple observing modes. For example, there are two-minute cadence observations, which were retrieved for specific targeted objects, and there are 30-minute cadence Full Frame Images (FFIs), which capture the entire *TESS* field of view.
#
# In this example, we will examine a target using the FFI observation. The *TESS* FFIs are dominated by the scattered light background signal on the *TESS* detector, which creates high-amplitude, periodic variation. This background can make planet detection difficult, but it can be removed using PLD.
#
# To access FFI data, we will use the [TESScut](https://mast.stsci.edu/tesscut/) tool on the Mikulski Archive for Space Telescopes (MAST), developed by [Brasseur et al. 2019](https://ui.adsabs.harvard.edu/abs/2019ascl.soft05007B/abstract). Lightkurve has a built-in search method for creating cutouts called [search_tesscut](https://docs.lightkurve.org/reference/api/lightkurve.search_tesscut.html?highlight=search_tesscut), which uses the same syntax as the [search_targetpixelfile](https://docs.lightkurve.org/reference/api/lightkurve.search_targetpixelfile.html?highlight=search_targetpixelfile) method above. We will search for the Wolf-Rayet star WR 40, which was observed by *TESS* in Sector 10.
# + colab={"base_uri": "https://localhost:8080/", "height": 78} executionInfo={"elapsed": 3522, "status": "ok", "timestamp": 1601325326993, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="m0Q4Pweqb2b3" outputId="9964dbc9-65ab-44eb-9470-8706b7990ac8"
search_result = lk.search_tesscut('WR40', sector=10)
search_result
# + [markdown] id="5YrXnxhnXRrH"
# We can download the pixel data with the [download](https://docs.lightkurve.org/reference/search.html?highlight=download) method. When using TESScut, this method takes the additional parameter `cutout_size`, which determines how many pixels each side length of the cutout target pixel file should have.
#
# Here, we use seven pixels on each side, which strikes a good balance between downloading enough pixels to create a good background model, and not downloading too many pixels, which can result in the corrector running slowly or including neighboring stars in the noise model.
# + colab={"base_uri": "https://localhost:8080/", "height": 420} executionInfo={"elapsed": 14922, "status": "ok", "timestamp": 1601325388777, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="HzVkJdboxNbx" outputId="3f214311-9196-4202-b9ba-73ef72eee8e3"
tpf = search_result.download(cutout_size=12)
tpf.plot();
# + [markdown] id="XjbfbE46zVmW"
# We can create an uncorrected SAP light curve from this target pixel file using a threshold mask.
# + colab={"base_uri": "https://localhost:8080/", "height": 387} executionInfo={"elapsed": 1028, "status": "ok", "timestamp": 1601325429733, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="zxTuP09cdS0H" outputId="d4550c45-0278-498b-86f5-0eaa4ab269dc"
uncorrected_lc = tpf.to_lightcurve(aperture_mask='threshold')
uncorrected_lc.plot();
# + [markdown] id="zK7mRJkAzmc-"
# The dominant trend in the SAP light curve above is the dramatic ramp up in flux due to the scattered light background on the *TESS* detector. The pulsation signal of WR 40 can also be seen clearly, with some additional long-period variability.
#
# We will create a [PLDCorrector](https://docs.lightkurve.org/reference/api/lightkurve.correctors.PLDCorrector.html?highlight=pldcorrector) object, and use the default values for [PLDCorrector.correct](https://docs.lightkurve.org/reference/api/lightkurve.correctors.PLDCorrector.correct.html?highlight=pldcorrector%20correct#lightkurve.correctors.PLDCorrector.correct) to remove this scattered light background.
# + colab={"base_uri": "https://localhost:8080/", "height": 764} executionInfo={"elapsed": 3206, "status": "ok", "timestamp": 1601325435795, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="x6q-cAPfDvUu" outputId="bdd0c002-183a-4ce0-9393-18538d6ec6d4"
pld = PLDCorrector(tpf)
corrected_lc = pld.correct()
pld.diagnose();
# + [markdown] id="ytZmzD0wRxAj"
# The `background` component of the PLD model (shown in blue in the middle panel) has successfully isolated the large spikes without fitting out the pulsations of WR 40.
#
# We can also examine the apertures used to perform this correction. For *TESS*, the dominant source of noise is the scattered light background, so by default only those pixels will be used. In the third panel, we can see that the `background_aperture_mask` contains only background pixels, reducing the risk of contamination by neighboring stars.
# + colab={"base_uri": "https://localhost:8080/", "height": 391} executionInfo={"elapsed": 2450, "status": "ok", "timestamp": 1601325443065, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="GxDcg8Cew4TX" outputId="bd03fe73-7553-4490-911a-84944652564f"
pld.diagnose_masks();
# + [markdown] id="SpV0VzDB3l4m"
# ## 5. Tuning `PLDCorrector` Using Optional Parameters
# + [markdown] id="L82MTs7WHPXT"
# In this section, we explore the optional parameters available when using the PLD method.
# + [markdown] id="Ym5J2kr0cPh9"
# These keywords can be used to influence the performance of PLD. The PLD design matrix is constructed from three distinct submatrices, shown below with their corresponding keywords:
# * **`background_model`**
# * `background_aperture_mask`
# * **`pixel_series`**
# * `pld_order`
# * `pca_components`
# * `pld_aperture_mask`
# * **`spline`**
# * `spline_n_knots`
# * `spline_degree`
# * `restore_trend`
#
# + [markdown] id="bZXztmqDNHjn"
# ### 5.1 Definitions of all additional parameters
#
# Full descriptions of each of these keywords can be found below.
# + [markdown] id="LVwU0SR62xO5"
# **`pld_order`** (`int`):
# * The order of Pixel Level Decorrelation to be performed. First order (`n=1`) uses only the pixel fluxes to construct the design matrix. Higher order populates the design matrix with columns constructed from the products of pixel fluxes. Default 3 for *K2* and 1 for *TESS*.
#
# **`pca_components`** (`int` or tuple of `int`):
# * Number of terms added to the design matrix for each order of PLD
# pixel fluxes. Increasing this value may provide higher precision
# at the expense of slower speed and/or overfitting. If performing PLD with `pld_order > 1`, `pca_components` can be a tuple containing the number of terms for each order of PLD. If a single `int` is passed, the same number of terms will be used for each order. If zero is passed, Principle Component Analysis (PCA) will not be performed.
#
# **`background_aperture_mask`** (`array-like` or `None`):
# * A Boolean array flagging the background pixels such that `True` means
# that the pixel will be used to generate the background systematics model.
# If `None`, all pixels which are fainter than 1-sigma above the median flux will be used.
#
# **`pld_aperture_mask`** (`array-like`, `'pipeline'`, `'all'`, `'threshold'`, or `None`):
#
# * A Boolean array describing the aperture such that `True` means
# that the pixel will be used when selecting the PLD basis vectors. If `None` or `all` are passed in, all pixels will be used. If `'pipeline'` is passed, the mask suggested by the official pipeline will be returned. If `'threshold'` is passed, all pixels brighter than 3-sigma above the median flux will be used.
#
# **`spline_n_knots`** (`int`):
# * Number of knots in spline.
#
# **`spline_degree`** (`int`)
# * Polynomial degree of spline.
#
# **`restore_trend`** (`bool`):
# * Whether to restore the long-term spline trend to the light curve.
#
# **`sparse`** (`bool`):
# * Whether to create `SparseDesignMatrix`.
#
# **`cadence_mask`** (`np.ndarray` of `bool`)
# * (optional) Mask, where `True` indicates a cadence that should be used.
#
# **`sigma`** (`int`):
# * Standard deviation at which to remove outliers from fitting (default 5).
#
# **`niters`** (`int`)
# * Number of iterations to fit and remove outliers (default 5).
#
# **`propagate_errors`** (`bool`):
# * Whether to propagate the uncertainties from the regression. Default is `False`. Setting to `True` will increase run time, but will sample from multivariate normal distribution of weights.
# + [markdown] id="3qBFe8TY_lp9"
# ## 6. Frequently Asked Questions
#
# **How should I select the pixels to use?**
#
# As shown earlier, there are three aperture masks used in the `PLDCorrector`.
#
# * `aperture_mask`: Used to create the SAP flux light curve from which the noise model is subtracted. For this aperture, you should select as many pixels as possible that only contain flux from the target star. This is done automatically using a threshold mask, but it is a good idea to examine that mask with the [diagnose_masks](https://docs.lightkurve.org/reference/api/lightkurve.correctors.PLDCorrector.diagnose_masks.html?highlight=diagnose_masks#lightkurve.correctors.PLDCorrector.diagnose_masks) method to ensure it does not include background targets.
#
# * `background_aperture_mask`: Pixels used to create the background model. These pixels should not contain flux from the target star whose light curve you are attempting to correct, nor should it contain flux from background targets.
#
# * `pld_aperture_mask`: Pixels used to create the correlated noise model. This aperture mask is more difficult to define, and may change on a case-by-case basis. For *K2*, this mask should contain as many pixels as possible in order to best capture the persistent motion-generated noise. For *TESS*, this mask should have less of an impact than the background mask, but should include all pixels in the cutout that do not contain background stars.
#
# **How can I speed up the correction?**
#
# The spline [DesignMatrix](https://docs.lightkurve.org/reference/api/lightkurve.correctors.DesignMatrix.html?highlight=designmatrix#lightkurve.correctors.DesignMatrix) used in [PLDCorrector](https://docs.lightkurve.org/reference/api/lightkurve.correctors.PLDCorrector.html?highlight=pldcorrector) can be substituted for a [SparseDesignMatrix](https://docs.lightkurve.org/reference/api/lightkurve.correctors.SparseDesignMatrix.html?highlight=sparsedesignmatrix). These behave identically to traditional `DesignMatrix` objects, but use `scipy.sparse` to speed up calculation and reduce memory. This can be done by passing in `sparse=True` to the `correct` method.
#
# **What do I do if I get a singular matrix error?**
#
# A singular matrix error occurs when a matrix used in [RegressionCorrector](https://docs.lightkurve.org/reference/api/lightkurve.correctors.RegressionCorrector.html?highlight=regressioncorrector) cannot be inverted, a step necessary for optimizing the coefficients. The primary reason this occurs is that the input `DesignMatrix` has low rank relative to the number of column vectors. There are two suggested solutions to this issue:
# * Limit the number of input column vectors by performing Principle Component Analysis (PCA). This is done automatically in `PLDCorrector`, but the number of output PCA vectors can be reduced using the `pca_components` keyword in the `correct` method from its default value of 16.
# * Ensure you are not masking out too much of your data. For instance, if you are using the `cadence_mask`, make sure that the values you want to include in your detrending are labeled as `True`. Using the inverse `cadence_mask` will often raise a singular matrix error.
# + [markdown] id="g6HorDwoUBQs"
# ## About this Notebook
# + [markdown] id="0cJVL81oTcGn"
# **Authors**: <NAME> (<EMAIL>), <NAME>
#
# **Updated**: September 29, 2020
# + [markdown] id="iQdt2aj0eJG3"
# ## Citing Lightkurve and Astropy
#
# If you use `lightkurve` or its dependencies in your published research, please cite the authors. Click the buttons below to copy BibTeX entries to your clipboard.
# + colab={"base_uri": "https://localhost:8080/", "height": 144} executionInfo={"elapsed": 845, "status": "ok", "timestamp": 1601325533294, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj8sjdnDeqdejfe7OoouYPIclAQV0KSTpsU469Jyeo=s64", "userId": "05704237875861987058"}, "user_tz": 420} id="UgxFosOkNgal" outputId="1a9b03a7-ec62-4362-9409-888480ddb49a"
lk.show_citation_instructions()
# + [markdown] id="1i_uXbTNuYxF"
# <img style="float: right;" src="https://raw.githubusercontent.com/spacetelescope/notebooks/master/assets/stsci_pri_combo_mark_horizonal_white_bkgd.png" alt="Space Telescope Logo" width="200px"/>
#
| docs/source/tutorials/2-creating-light-curves/2-3-k2-pldcorrector.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.0
# language: julia
# name: julia-1.1
# ---
#using Pkg
#Pkg.add("HTTP");
#Pkg.add("DataFrames");
#Pkg.add("CSV");
#Pkg.add("Plots")
using Plots;
using HTTP;
using CSV;
# + inputHidden=false outputHidden=false
using Pkg
#Pkg.add("HTTP");
#Pkg.add("DataFrames");
#Pkg.add("CSV");
#Pkg.add("Plots")
using Plots;
using HTTP;
using CSV;
res = HTTP.request("GET",
"https://api.daf.teamdigitale.it/dataset-manager/v1/dataset/daf%3A%2F%2Fopendata%2Flecce_o_quoziente_i_immigrazione_ed_emigrazione_1?format=csv",
[("Authorization", "Basic ZF9hbGVAZGFmLml0OlNpbHZpQWxlNzg4MQ==")]);
mycsv = CSV.read(IOBuffer(res.body));
mycsv
# + inputHidden=false outputHidden=false
mycsv
kernel not supported yet
# + inputHidden=false outputHidden=false
using Pkg
Pkg.add("HTTP");
Pkg.add("DataFrames");
Pkg.add("CSV");
Pkg.add("Plots")
using Plots;
using HTTP;
using CSV;
res = HTTP.request("GET",
"https://api.daf.teamdigitale.it/dataset-manager/v1/dataset/daf%3A%2F%2Fopendata%2Flecce_o_quoziente_i_immigrazione_ed_emigrazione_1?format=csv",
[("Authorization", "Bearer YOU_MUST_BE_LOGGEDIN")]);
lecce_o_quoziente_i_immigrazione_ed_emigrazione_1 = CSV.read(IOBuffer(res.body));
lecce_o_quoziente_i_immigrazione_ed_emigrazione_1
# + inputHidden=false outputHidden=false
# define the Lorenz attractor
mutable struct Lorenz
dt; σ; ρ; β; x; y; z
end
function step!(l::Lorenz)
dx = l.σ*(l.y - l.x) ; l.x += l.dt * dx
dy = l.x*(l.ρ - l.z) - l.y ; l.y += l.dt * dy
dz = l.x*l.y - l.β*l.z ; l.z += l.dt * dz
end
attractor = Lorenz((dt = 0.02, σ = 10., ρ = 28., β = 8//3, x = 1., y = 1., z = 1.)...)
# initialize a 3D plot with 1 empty series
plt = plot3d(1, xlim=(-25,25), ylim=(-25,25), zlim=(0,50),
title = "Lorenz Attractor", marker = 2)
# build an animated gif by pushing new points to the plot, saving every 10th frame
@gif for i=1:1500
step!(attractor)
push!(plt, attractor.x, attractor.y, attractor.z)
end every 10
| open-notebooks-example/julia-kernel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/AureliaWambui19/Home-team-Away-team-scores-prediction/blob/main/wambui_aurelia_core_week_6_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="e3Se4tiPVPg8"
# # 1.1 Introduction
# * Football is a family of team sports that involve, to varying degrees, kicking a ball to score a goal.
#
# * The men's FIFA World Ranking is a ranking system for men's national teams in association football.
# * Currently led by Belgium
#
# * A points system is used, with points being awarded based on the results of all FIFA-recognised full international matches.
#
#
# + [markdown] id="6jLcnEGyPmsq"
# ## 1.1 Defining the Question
#
# * I have been recruited as a football analyst in a company - Mchezopesa Ltd and tasked to accomplish the task below :
#
# * Predict the result of a game between team 1 and team 2, based on who's home and who's away, and on whether or not the game is friendly (including rank of the respective team).
# + [markdown] id="0KRU2WCnPmpx"
# ## 1.2 Metrics for Success
#
# * Obtaining about 80% Accuracy score and above
# * Correctly identifying status of results(win,loss,draw)
#
#
# + [markdown] id="T27Fr4V4Pmi7"
# ## 1.3 The Context
#
# The new model for calculating the FIFA/Coca-Cola World Ranking (FWR) was developed over two years
# during which time a large number of different algorithms was tested and extensively discussed.
# Throughout this review and consultation process, the main aim was to identify an algorithm that is not
# only intuitive, easy to understand and improves overall accuracy of the formula, but also addresses
# feedback received about the previous model and provides fair and equal opportunities for all teams
# across all confederations to ascend the FWR
#
# The Elo method of calculation adds/subtracts points (as opposed to averaging points) for individual
# matches to/from a team’s existing point total. The points which are added or subtracted are partially
# determined by the relative strength of the two opponents, including the logical expectation that teams
# higher in the ranking should fare better against teams lower in the ranking.
#
#
#
# + [markdown] id="1P7ENPLrPmfT"
# ## 1.4 Experimental design taken
#
# - Perform your EDA
# - Perform any necessary feature engineering
# - Check of multicollinearity
# - Building a model
# * Approach 1: Polynomial regression model
#
# * Model 1: Predict how many goals the home team scores
# * Model 2: Predict how many goals the away team scores
#
# * Approach 2: Logistic regression model
#
# * Figure out from the home team’s perspective if the game is a Win, Lose or Draw (W, L, D)
#
# - Cross-validate the model
# - Compute RMSE
# - Create residual plots for the model
# - Assess Heteroscedasticity using Bartlett’s test
#
# + [markdown] id="ZvJ6FlIRUrZ7"
# ## 1.5 Appropriateness of the available Data
#
# This project has two datasets:
#
# * Ranking dataset: contains the team ranks from 1993 to 2018
#
# * Results dataset: contains matches and the team scores since 1892 to 2019
#
# The link to the dataset is:
#
# * https://drive.google.com/open?id=1BYUqaEEnFtAe5lvzJh9lpVpR2MAvERUc
#
# The data is relevant for this project
#
# + [markdown] id="bJNEdPrKVGsm"
# # 2 Data Understanding
# + id="_RgEG9oSAfMb"
# Importing Libraries we use for our analysis
import pandas as pd
import numpy as np
import scipy as sp
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LinearRegression, LogisticRegression
from sklearn import metrics
from sklearn.model_selection import KFold, LeaveOneOut
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import PolynomialFeatures
from sklearn.preprocessing import StandardScaler
# + [markdown] id="EG14C7aYV8Xa"
# # 2.1 Viewing our Datasets
# + id="bRfLIuVMBimz"
# Reading the datasets
FifaRank = pd.read_csv('fifa_ranking.csv')
FifaResult = pd.read_csv('results.csv')
# + id="roHou8vIBijx" outputId="f3da3e77-300e-47e6-e30d-8bf9345fce27" colab={"base_uri": "https://localhost:8080/", "height": 213}
# Viewing the top 3 observation in the fifa ranking dataset
FifaRank.head(3)
# + id="_SbcOGbOePsN" outputId="8c13c3cd-bdef-4619-89d5-c9295419c056" colab={"base_uri": "https://localhost:8080/", "height": 213}
# Viewing the last 3 observation in the fifa ranking dataset
FifaRank.tail(3)
# + id="p7ksjP4KBihG" outputId="43fb282d-1b21-4533-e3cf-00a50310713d" colab={"base_uri": "https://localhost:8080/", "height": 193}
# Viewing the last 3 observation in the result dataset
FifaResult.tail(3)
# + [markdown] id="eHs-LjqYLUEw"
# # 2.2 Checking data
# + id="EjVy7zUOBid9" outputId="53f72a3d-5305-4156-f0ed-b28e179a33e7" colab={"base_uri": "https://localhost:8080/"}
# Checking the size of the fifa ranking dataset
FifaRank.shape
# + [markdown] id="jxb9PlRrcdXJ"
# This dataset has 57993 rows and 16 columns
# + id="irnDPocPBicE" outputId="695200fa-3451-4fb6-9e9c-2662a290a6be" colab={"base_uri": "https://localhost:8080/"}
# checking the size of the results dataset
FifaResult.shape
# + [markdown] id="ZWrYLfYEcmLo"
# This dataset has 40839 rows and 9 columns
# + id="0cXsZFMaBkk_" outputId="89a3082e-a5e8-409f-befa-9b15cb3567c3" colab={"base_uri": "https://localhost:8080/"}
# Checking the ranking dataset information
FifaRank.info()
# + id="fIOCLdxSBkh5" outputId="cb3b02d9-1799-4b1b-b7f9-4b58471cefc3" colab={"base_uri": "https://localhost:8080/"}
# Checking the result dataset information
FifaResult.info()
# + id="ScaKqyfdBkge" outputId="0c682cdb-f21c-4b15-9f2b-0b8d99e8bf0c" colab={"base_uri": "https://localhost:8080/"}
# Viewing the column names of the ranking dataset
FifaRank.columns
# + id="XOK5BVWjBbKW" outputId="9d6ada7a-f81d-4953-d807-63a39065f0b2" colab={"base_uri": "https://localhost:8080/"}
# Viewing the column names of the result dataset
FifaResult.columns
# + [markdown] id="z4F90dHq_ISM"
# # 3 Feature Engineering
# + id="nKJySHSzaRIZ" outputId="548d2060-00e2-4aed-db93-a6ff49e3f0ea" colab={"base_uri": "https://localhost:8080/", "height": 313}
# finding the difference of scores and storing them in a new column called game_result
FifaResult['game_results'] = FifaResult['home_score'] -FifaResult['away_score']
FifaResult.head(4)
# + id="PGmEbcfomffN"
# Creating a function to specify whether its a win , loss or a draw based on a home team perspective
# the results (its a win , loss or a draw ) are stored in a new column called status
def home_team(game_results):
if game_results > 0:
return 'win'
elif game_results < 0:
return 'loss'
else:
return 'draw'
FifaResult['status'] = FifaResult.game_results.apply(lambda w: home_team(w))
# + id="ZYRxyIoYp5PS" outputId="280121fa-de45-4db2-d9e1-dffeb2c94827" colab={"base_uri": "https://localhost:8080/", "height": 182}
FifaResult.head(2)
# + id="maMIAfRPmfVr" outputId="507fad6d-7818-48f1-9910-f40409ce80df" colab={"base_uri": "https://localhost:8080/"}
# finding unique values in tournament column
FifaResult.tournament.unique()
# + id="aqDI76PPeTbo"
# Changing the tournament type into three categories
# Tournament type (World cup, Friendly, Other)
# The 3 respective category will be stored in a new column named tournament_type
def ton(tournament_type):
if tournament_type == 'Friendly':
return 'Friendly'
elif tournament_type == 'FIFA World Cup':
return 'World cup'
else:
return 'Other'
FifaResult['tournament_type'] = FifaResult.tournament.apply(lambda t: ton(t))
# + id="PG1AqXSJrqjF" outputId="3d824e73-d4fa-4c87-8d64-59fb3e1d7054" colab={"base_uri": "https://localhost:8080/", "height": 182}
FifaResult.head(2)
# + id="bndZPcrU2H_j"
# Changing the dates column in both datasets into datetime format
FifaResult['date'] = pd.to_datetime(FifaResult['date'])
FifaRank['rank_date'] = pd.to_datetime(FifaRank['rank_date'])
# + id="R-qz8Wezhl01" outputId="34e4ba04-02ef-433a-9326-2e5993d31990" colab={"base_uri": "https://localhost:8080/"}
# Confirming that we have changed the date columns into datetime datatypes
print(FifaRank.rank_date.dtypes)
print(' ')
print(FifaResult.date.dtypes)
# + id="jN5tk1uYjKOR"
# Extracting the year and month from the date column;
# Here we will create a new column for each
FifaResult['year'] = pd.DatetimeIndex(FifaResult['date']).year
FifaResult['month'] = pd.DatetimeIndex(FifaResult['date']).month
FifaRank['year'] = FifaRank['rank_date'].dt.year
FifaRank['month'] =FifaRank['rank_date'].dt.month
# + id="YOmyvK3AjKL0" outputId="643c2a8c-db94-41fb-ba5e-fa16a5987110" colab={"base_uri": "https://localhost:8080/", "height": 247}
# confirming the changes
FifaResult.head(3)
# + id="dqaHKZEejKJI" outputId="c45fa5bb-0852-4d42-949d-8f98fad825a2" colab={"base_uri": "https://localhost:8080/", "height": 165}
# confirming changes
FifaRank.head(2)
# + id="dzz1cf8k8IdT"
# changing the full country column name in ranking dataset to home_team so as to ease manipulation of the datasets when merging them later
FifaRank= FifaRank.rename({'country_full': 'home_team'}, axis = 1)
# + id="ffnz1oyx8IaZ" outputId="96b74a39-3cd8-4d57-c3ea-14d9cdbf33bc" colab={"base_uri": "https://localhost:8080/", "height": 165}
# confirming changes
FifaRank.head(2)
# + id="UC7r3XpA9rdI"
# Dropping unnecessary columns in result dataset
FifaResult.drop(['date', 'game_results'], axis = 1, inplace = True)
# + id="Libno3OMtGEz" outputId="9890cd2b-860e-4fdb-b100-e18f8be2a25d" colab={"base_uri": "https://localhost:8080/"}
FifaResult.columns
# + id="y_BPjVDa9rYP"
# Dropping unnecessary columns in rank dataset
FifaRank.drop(['country_abrv','rank_date', 'total_points', 'previous_points','cur_year_avg', 'cur_year_avg_weighted' ,'last_year_avg' , 'last_year_avg_weighted' , 'two_year_ago_avg', 'two_year_ago_weighted', 'three_year_ago_avg' ,'three_year_ago_weighted', 'confederation'], axis =1, inplace = True)
# + id="Zecu1i2ZL_lq"
# Merging datasets
# Based on home_team, year, month
home_me= pd.merge(FifaResult,FifaRank, how="left", on = ['home_team', 'year', 'month'])
# + id="mNiN2aepAbXw" outputId="1bab2509-4552-4f09-f292-9d9871a38441" colab={"base_uri": "https://localhost:8080/", "height": 191}
# viewing our merged dataset 4 top observations
home_me.head(4)
# + id="JM7XIpkdIfii" outputId="0fa2a0c4-9947-4279-e9ab-2bc16d3a9325" colab={"base_uri": "https://localhost:8080/", "height": 213}
# viewing our merged dataset lastb 3 observations
home_me.tail(3)
# + id="AHQfPlLpm4_A"
# renaming the rank column name to home_rank so as to get the respective rank of the home team
home_me = home_me.rename({'rank': 'home_rank'}, axis = 1)
# + id="SWRW6q17nH9b" outputId="c02e3afa-3c13-44bb-ed9a-c1f89416d32c" colab={"base_uri": "https://localhost:8080/", "height": 130}
# Confirming changes
home_me.head(2)
# + id="kqsOgG2ooXpK" outputId="c88640b7-a932-445b-824e-73134f4b3971" colab={"base_uri": "https://localhost:8080/", "height": 110}
# renaming the column home_team (originally called country full) as away team so that we get their individual ranks of away teams
FRankone= FifaRank.rename({'home_team': 'away_team'}, axis = 1)
FRankone.head(2)
# + id="Mpm4Yexars1J"
# Merging the home_merged dataset with the
# Based on away_team, year, month
Fiifa = pd.merge(home_me,FRankone, how="left", on = ['away_team', 'year', 'month'])
# + id="KWh33rUUsKRY" outputId="9bd268f6-9bb3-4142-a578-baa69d00310d" colab={"base_uri": "https://localhost:8080/", "height": 130}
# Checking the first two observations of the merged dataset
Fiifa.head(2)
# + id="soLvzzWyseww" outputId="8c8aefbe-831e-4ba7-be56-6cbc60719b7d" colab={"base_uri": "https://localhost:8080/", "height": 222}
# renaming the rank column as away rank in the new dataframe
Fiifa = Fiifa.rename({'rank': 'away_rank'}, axis = 1)
Fiifa.head()
# + [markdown] id="-Z75RJ3QLx0G"
# # 4 Tyding the dataset
# + id="PJPYg1iW8w19" outputId="0e0316a1-efbc-47e2-a48c-267daae0594f" colab={"base_uri": "https://localhost:8080/"}
# checking for unique year rankings
FifaRank.year.unique()
# + [markdown] id="0VnbrYJ846jX"
# Rankings are from 1993 t0 2018 .after merging a lot of missing values were noted especialy in years before 1993 and after 2018.therefore i will drop the data where this was observed as there are no rankings available
# + id="s4VbsB4B4aDl"
Fiifa.dropna(inplace = True)
# + id="vzblMaFJD096" outputId="70152208-2494-4856-bf18-6604e700a969" colab={"base_uri": "https://localhost:8080/"}
# confirming that there are no null values
Fiifa.isnull().sum()
# + id="Mt7IeSzEJc_9" outputId="2373d14b-2f27-4e3f-86ac-1931bf70568d" colab={"base_uri": "https://localhost:8080/"}
# checking for duplicates
Fiifa.duplicated().sum()
# + id="X7U5QxkbJc5U"
# dropping the duplicates
Fiifa.drop_duplicates(inplace = True)
# + id="ogOjz3hQLHSQ" outputId="3e3c11a1-0169-4fb4-a660-3ea7385acf54" colab={"base_uri": "https://localhost:8080/"}
# Checking that we have no duplicates in the data
Fiifa.duplicated().sum()
# + id="IOQXcviOL6fb" outputId="07968395-1a03-4cb7-8439-118923763fa9" colab={"base_uri": "https://localhost:8080/"}
# checking columns of merged dataset
Fiifa.columns
# + id="10U7Hw4tD07G" outputId="5319a1c7-156c-4e1a-f7c1-2be00303006a" colab={"base_uri": "https://localhost:8080/", "height": 395}
# viewing our dataset after cleaning
Fiifa.head()
# + id="CHziyD9NAbUj" outputId="ed305db0-5619-47cb-91b7-e62c3eef1142" colab={"base_uri": "https://localhost:8080/"}
# checking the shape of the cleaned data
Fiifa.shape
# + [markdown] id="m4O6IQowlpB4"
# This dataset has 16889 rows and 16 columns
# + id="aM8UmLFsSmj9"
# Encoding the categorical columns so as to manage perform operations such as correlation check
#
le = LabelEncoder()
Fiifa= Fiifa.apply(le.fit_transform)
# + id="gTxyu786Td2C" outputId="809b7c29-7ef5-4ae1-a971-81925eb05a18" colab={"base_uri": "https://localhost:8080/", "height": 222}
# Confirming the changes
Fiifa.head(5)
# + id="24GK1soO1wUA" outputId="3db42d99-7934-4f02-dc2b-3dcb7b2400eb" colab={"base_uri": "https://localhost:8080/", "height": 404}
# checking for outliers in our dataset
# Using boxplots
# Labeling the title of our chart
# Displaying chart
plt.figure(dpi = 100)
ax = sns.boxplot(data = Fiifa,orient='h')
plt.title(' Outliers in Fifa dataset', color = 'red')
plt.xlabel(' Frequency')
plt.show()
# + [markdown] id="ch3NXLQh_cdk"
# # 5 Exploratory data analysis
# + id="-qBs4diILGqD" outputId="dc7c476a-3351-43ec-ca1e-f06268fef3cb" colab={"base_uri": "https://localhost:8080/", "height": 295}
h=Fiifa['home_score']
plt.hist(h, histtype='bar', rwidth=0.9)
plt.xlabel('No. of home scores')
plt.ylabel('Quantity')
plt.title('number of home scores',color='red')
plt.show()
# + [markdown] id="X3Fmm_Q5VeO8"
# home teams scored mostly one goal
# + id="EyaWJFdrVHIk" outputId="7322b5a4-8e1a-4efc-b9b1-6652d8e02f4a" colab={"base_uri": "https://localhost:8080/", "height": 295}
a=Fiifa['away_score']
plt.hist(h, histtype='bar', rwidth=0.9)
plt.xlabel('No. of away scores')
plt.ylabel('Quantity')
plt.title('number of away scores',color='red')
plt.show()
# + [markdown] id="jF8oo1CzCiWQ"
# Most away teams score atleast one goal
#
# Both histograms are positively skewed .This shape indicates that there are a number of data points, perhaps outliers, that are greater than the mode
#
# + id="Tvp6_QQzPgpC" outputId="94e11c4f-bbe8-455c-c5b0-0c572690a562" colab={"base_uri": "https://localhost:8080/", "height": 499}
# status of game results in respect tothe home team(draw = 0, lose =1, win = 2)
# Using a countplot to visualize these results
# Using Seaborn
# Labeling the x and y axis
# Giving a title to our chart
# Displaying our chart
plt.figure(figsize = (6,6), dpi = 80)
sns.countplot(Fiifa['status'])
plt.xlabel('status (draw = 0, lose =1, win = 2)')
plt.ylabel('Count')
plt.title('status of games results', color = 'red')
plt.show()
# + [markdown] id="FfxZH43SDn_7"
# The above bar chart above shows that wins by the home teams are high as compared to loss/draws
# + [markdown] id="kFIeh2HWMWji"
# # 6 Multicollinearity
# * Checking for multicollinearity
# * Solving multicollinearity
# + id="1f3ve3HwLolc" outputId="adb4ae45-752d-4378-f50b-ca61b7db52b4" colab={"base_uri": "https://localhost:8080/", "height": 559}
# Before we build a model we shall check if
# the independent variables are collinear/ correlated to each other
# Getting the pearson correation coefficient for each of the variables
correlation = Fiifa.corr()
correlation
# + [markdown] id="7qbJj_aCFEQf"
# The correlation matrix indicates that most variables are moderately or weakly correlated.*(both positively and negatively)
#
# This is very beneficial when creating a model, as collinear variables reduce the power of the model to identify independent
# variables that are statistically significant.
#
# We will use the correlation matrix to calculate the vif (Variance Inflation Factor).
# Variance inflation factor (VIF) is a measure of the amount of multicollinearity in a set of multiple regression variables. Mathematically, the VIF for a regression model variable is equal to the ratio of the overall model variance to the variance of a model that includes only that single independent variable. This ratio is calculated for each independent variable. A high VIF indicates that the associated independent variable is highly collinear with the other variables in the model.
#
# + id="2cOFejwkMcIZ" outputId="71003218-0640-4479-ad62-09449febb0d7" colab={"base_uri": "https://localhost:8080/", "height": 559}
# checking for multicollinearity
# Using the variance Inflation Factor (VIF)
#
# This is calculated using linear algebra inverse function
pd.DataFrame(np.linalg.inv(correlation.values), index = correlation.index, columns = correlation.columns)
# From the correlation matrix below there are no correlated independent variables as all have VIF below 5, which is the threshold
# + [markdown] id="tRGDvSKnHiu6"
# We check VIFs along the diagonal.
#
# VIFs Values greater than 5 indicate that the presence of multicollinearity.
#
# If present we remove the variable with the greatest VIF value.
#
# Typically, a VIF value around 5 is a potential problem, and value around 10 is considered seriously problematic and suggests that the related variable should be dropped from the model.
#
# From the correlation matrix there are no correlated independent variables as all have VIF values are below 5, which is the threshold and therefore no variable will be dropped in this project
# + [markdown] id="zKd8cdlTMgNw"
# # 8 Building a Model
# + [markdown] id="KnM1vinqS3aV"
# ## 8.1 Polynomial Regression Model
# + id="-xy3JU150U_v"
# Approach 1: Polynomial approach
# What to train given:
# Rank of home team
# Rank of away team
# Tournament type
# Model 1: Predict how many goals the home team scores
# Model 2: Predict how many goals the away team scores
# + [markdown] id="bO3cFM0LJIUh"
# ### Model 1
#
# Predict how many goals the home team scores
# + id="K9HguHrd1Kle" outputId="c628db2e-424a-41f4-dc7b-4624be499ec0" colab={"base_uri": "https://localhost:8080/", "height": 130}
# Viewing our dataset before splitting
Fiifa.head(2)
# + id="GgvltokP0U82"
# Model 1
# Predict how many goals the home team scores given home rank
X = Fiifa['home_rank'].values.reshape(-1, 1)
y = Fiifa['home_score'].values.reshape(-1, 1)
# + id="I8b05Gp60c1Z" outputId="18fddea9-0289-48b0-c335-0a6be146c471" colab={"base_uri": "https://localhost:8080/", "height": 295}
# showing relationship between home rank and home score
plt.scatter(X,y)
plt.title('Home team performance', color = 'red')
plt.xlabel('homerank')
plt.ylabel('home score')
plt.show()
# + [markdown] id="d05Ncv0cJdDF"
# There are more points on the lower side of the scatter plot.
# home team scores are mostly between 0 and 5 goals.
# + id="MzYYEOl6qRhj" outputId="87860350-6436-413d-b4dd-4a9054ad9d5f" colab={"base_uri": "https://localhost:8080/"}
X.shape
# + id="4DAniykeqROx" outputId="94917b32-85db-4bb7-b611-dab35ebee3db" colab={"base_uri": "https://localhost:8080/"}
y.shape
# + id="sJ3eIPNl0cyH"
# Split the dataset into train and test sets
# this means training data is 80% while test size is 20%
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2, random_state=2)
# + id="0ejsXrjB49vC"
# Fitting the polynomial features to the X the independent variable
poly_reg = PolynomialFeatures(degree =4)
X_poly = poly_reg.fit_transform(X)
# + id="Dn4cK4xy49ry" outputId="c9e39946-1b97-40ad-d94a-04460bf9f85a" colab={"base_uri": "https://localhost:8080/"}
# Fitting a polynomial Regression to the dataset.
pol_reg = LinearRegression()
pol_reg.fit(X_poly, y)
# + id="mi5uyVD362nC" outputId="f0666924-78e6-4b1f-c18e-1af4358aa5ba" colab={"base_uri": "https://localhost:8080/", "height": 295}
# Visualizing the polynomial Regression results
plt.scatter(X, y, color='blue')
plt.plot(X, pol_reg.predict(X_poly),color='red')
plt.title('home score prediction')
plt.xlabel('home rank')
plt.ylabel('home score')
plt.show()
# + [markdown] id="KxKrTq2RLgUL"
#
# Using the polynomial regression model of degree 4,
#
# most data points have been omitted
# the visualization as appears makes it difficult to analyze and makes use of this model difficult to use for predictions
# + id="NzFbxKzi62je" outputId="48757c16-7647-4891-fe88-fdb3f8e07005" colab={"base_uri": "https://localhost:8080/"}
# Making predictions using our model
poly_pred = pol_reg.predict(poly_reg.fit_transform([[20]]))
print('Polynomial prediction when home rank is 20 the home team score is: %d' %poly_pred)
# + [markdown] id="44QFXeJvSL6l"
# ### Model 2
#
# Predict how many goals the away team scores
#
#
# + id="ICrAyuqQTsdO"
# Model 2: Predict how many goals the away team scores given the away team rank
#
X = Fiifa['away_rank'].values.reshape(-1, 1)
y = Fiifa['away_score'].values.reshape(-1, 1)
# + id="sPy4zAZcTsaL" outputId="ad1b8208-e53d-4e11-8e21-df9857cf6c66" colab={"base_uri": "https://localhost:8080/", "height": 295}
# Visualizing the dependent vs independent variable using a scatter plot
plt.scatter(X,y)
plt.title('away team performance', color = 'red')
plt.xlabel('away rank')
plt.ylabel('away score')
plt.show()
# + [markdown] id="uFb0YrX905lh"
#
# most cases the away team scores between 0 and 4 goals.
# + id="s7erOmy1TsXF"
## Split the dataset into train and test sets
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2, random_state=0)
# + id="dPiSaUQ5TsT1"
# Fitting the polynomial features to the X
poly_reg = PolynomialFeatures(degree = 4)
X_poly = poly_reg.fit_transform(X)
# + id="rWxogUxyTsQu" outputId="645eb890-5da4-4609-b604-4e124c83c281" colab={"base_uri": "https://localhost:8080/"}
# Fitting a polynomial Regression to the dataset
pol_reg = LinearRegression()
pol_reg.fit(X_poly, y)
# + id="IMejuCmHTsNZ" outputId="5678abb7-1045-4ebf-cdc6-7b170784cc05" colab={"base_uri": "https://localhost:8080/", "height": 295}
# Visualizing the polynomial Regression results using a scatter plot
plt.scatter(X, y, color='blue')
plt.plot(X, pol_reg.predict(X_poly),color='red')
plt.title('away team prediction')
plt.xlabel('away rank')
plt.ylabel('away score')
plt.show()
# + [markdown] id="tcN2QtUH1Lyz"
# Using the polynomial regression model of degree 4,
# most data points have not been highlighted
# This is underfitting.
# The polynomial regression is not a good model to predict how many goals the away team scores given the away team rank.
# + id="kPI7KjlrVpsH" outputId="45b45afd-0684-4f98-ea52-1c11fce6c9bf" colab={"base_uri": "https://localhost:8080/"}
# Making predictions using our model
poly_pred = pol_reg.predict(poly_reg.fit_transform([[58]]))
print('Polynomial prediction when home away rank is 58 the away team score is: %d' %poly_pred)
# + [markdown] id="Q43NBsK_TcaF"
# ## 8.2 Logistic Regression Model
#
# - Logistic regression is a predictive analysis.
#
# - Logistic regression is used to describe data and to explain the relationship between one dependent binary variable and one or more nominal, ordinal, interval or ratio-level independent variables.
# - Logistic Regression is used when the dependent variable(target) is categorical.
# - In this model, we will be predicting whether the home team (Wins, Losses or Draws) in a match.
#
# + id="MZwFVOFTWfGO" outputId="d095b721-c4e2-4fa2-9fc6-7110c69330ac" colab={"base_uri": "https://localhost:8080/", "height": 130}
# Viewing the first two observations before splittig our dataset
Fiifa.head(2)
# + id="qSASSfbZWfCT"
# Splitting our dataset
# X: independent variables
# y: dependent variable
# Splitting the data into train and test sets
X = Fiifa.drop(['status'], axis = 1)
y = Fiifa.status
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = .2, random_state=0)
# + id="SqPbsNqpMtdd" outputId="257d6728-5234-445b-e6c6-e4dbcb4e215a" colab={"base_uri": "https://localhost:8080/"}
# Fitting our model to our train sets
# Logistic Regression model in this case
#
LogReg = LogisticRegression()
LogReg.fit(X_train, y_train)
# + id="OiZgIDgNS2jY"
# Using our model to make a prediction
y_pred = LogReg.predict(X_test)
# + id="-HoMZBFYUdYJ" outputId="3e40229c-bde1-4a64-fb02-f912b1b172de" colab={"base_uri": "https://localhost:8080/"}
# Evalauting the model
print(accuracy_score(y_test, y_pred))
# + [markdown] id="IVj4m-lJZlHt"
# The model has an accuracy score of 62.72%
# + [markdown] id="4LFaXRjUMun5"
# # 9 Cross-Validation
# + id="A9GHmqX8MtaF" outputId="7906947f-45f3-449b-cc55-02c44aab2198" colab={"base_uri": "https://localhost:8080/"}
# Using KFolds
# Splitting our dataset
# independet variables as X
# dependent variable as y
X = Fiifa.drop(['status'], axis = 1).values
y = Fiifa.status.values
# specifying the number of folds
folds = KFold(n_splits = 10)
# We now create and assess 10 models based on the folds we created.
RMSES = [] # An array of RMSEs to keep track of the RSME of each model
count = 1 # starting point # helps to keep track of the model number in training
for train_index, test_index in folds.split(X):
# Setting up the train and test based on the split determined by KFold
# With 10 folds we split our data into training and test sets
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
# fitting a Logistic regression model
LogReg = LogisticRegression()
LogReg.fit(X_train, y_train)
# Assess the accuracy of the model
y_pred = LogReg.predict(X_test)
# Calculating the RMSES of each model
# Appending each RMSE into the list earlier created
rmse_value = np.sqrt(metrics.mean_squared_error(y_test, y_pred))
RMSES.append(rmse_value)
# printing each model RMSE
print('Model ' + str(count) + ' Root Mean Squared Error:',rmse_value)
count = count + 1
# + id="2vvJlA5kd5sB" outputId="a7036bff-f6d2-4de2-c373-c00a8ecf91ba" colab={"base_uri": "https://localhost:8080/"}
# Printing the mean of the RMSES in all the 10 models
print(np.mean(RMSES))
# + id="IcEVrQHYOwOY" outputId="a4d43a2f-2fd1-4361-8ae7-98be1f4edfc3" colab={"base_uri": "https://localhost:8080/", "height": 295}
# Visualizing the 10-folds RMSES using a scatter plot
plt.plot(RMSES)
plt.ylabel('RMSE value')
plt.title("RMSE line plot", color = 'red')
plt.xlabel('model ID')
plt.show()
# + [markdown] id="7FZt4x0qA_4U"
# # 10.Heteroskedisity
# + [markdown] id="rkGa2u3uBGR_"
# Heteroscedasticity means unequal scatter. In regression analysis, we talk about heteroscedasticity in the context of the residuals or error term. Specifically, heteroscedasticity is a systematic change in the spread of the residuals over the range of measured values.
# + id="HS3dpNGaiUiu" outputId="2fd36dc5-5985-4cde-9b43-0f81f3f276ec" colab={"base_uri": "https://localhost:8080/"}
# First: splitting our dataset
# Into the feature set and the target variable
X = Fiifa.drop(['status'], axis = 1)
y = Fiifa.status
# Split the dataset into train and test sets
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size = 0.2, random_state=0)
# Fitting a Logistic model
LogReg = LogisticRegression()
LogReg.fit(X_train, y_train)
# Using our model to make a prediction
y_pred = LogReg.predict(X_test)
# We now create the residual by subtracting the test value from the predicted
# value for each row in our dataset
residuals = np.subtract(y_pred, y_test)
# Creating a summary description of the residuals:
pd.DataFrame(residuals).describe()
residuals.mean()
# + [markdown] id="Hjm9yJepPe20"
# ## 10..1 Residual Plots
# + id="suZljruKPd0F" outputId="2b41efa4-d12f-4cca-824a-348e7e5a4f3b" colab={"base_uri": "https://localhost:8080/", "height": 279}
# Visualizing the residuals using a scatter plot
plt.scatter(y_pred, residuals, color='black')
plt.ylabel('residual')
plt.xlabel('predicted values')
plt.axhline(y= residuals.mean(), color='red', linewidth=1)
plt.show()
# + [markdown] id="Q08my9X1qRfO"
# Residuals are centered around a mean of appx 0.43
# …positive values for the residual (on the y-axis) mean the prediction was too low, and negative values mean the prediction was too high; 0 means the guess was exactly correct
#
# + [markdown] id="ikzQTH1rPptl"
# ## 10.2 Barlett's test
# + id="rKy_8vbUPdyA" outputId="022d8e56-f88c-4346-89b0-4cc7830be7c1" colab={"base_uri": "https://localhost:8080/"}
# Carrying out Barlett's test
# It is a more thorough heteroskedasticity test.
test_result, p_value = sp.stats.bartlett(y_pred, residuals)
# To interpret the results we must also compute a critical value of the chi squared distribution
degree_of_freedom = len(y_pred)-1
probability = 1 - p_value
critical_value = sp.stats.chi2.ppf(probability, degree_of_freedom)
print(p_value)
# If the test_result is greater than the critical value, then we reject our null
# hypothesis. This would mean that there are patterns to the variance of the data
# Otherwise, we can identify no patterns, and we accept the null hypothesis that
# the variance is homogeneous across our data
if (test_result > critical_value):
print('the variances are unequal, and the model should be reassessed')
else:
print('The variances are Homogeneous!')
| wambui_aurelia_core_week_6_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# +
import astropy.coordinates as coord
import astropy.table as at
import astropy.units as u
import matplotlib as mpl
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
from pyia import GaiaData
# -
tbl = at.Table.read('../data/RRL-with-prob.fits')
mask = (tbl['member_prob'] > 0.5) & tbl['inside_stream_track']
mask.sum()
g = GaiaData(tbl[mask])
c = g.get_skycoord(distance=g.D_ps1*u.kpc).transform_to(coord.Galactocentric)
np.savetxt('../../visualizations/mwstreams/data/pal5.txt', c.data.xyz.to_value(u.kpc).T)
| notebooks/mwstreams-viz-prepare.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Philosophy
import this
# [PEP 8](https://www.python.org/dev/peps/pep-0008/) - Style Guide for Python Code
# # Pure python
#
# - Know what's in the [standard library](https://docs.python.org/3/library/index.html).
# - Decorators.
# - Generators.
# - Object oriented programming.
# # Pandas and data analysis
#
# pandas is huge! We didn't cover many things. Among the more important parts that you should be aware of there are:
#
# - Resampling the time units of timed data.
#
# After pandas you will probably need some [numpy](http://www.numpy.org/) and [matplotlib](https://matplotlib.org/). The [scipy lecture notes](http://www.scipy-lectures.org/) will help you with that. [See my blog for more details](http://blog.tomgurion.me/getting-started-with-python.html).
# There's also [scikit-learn](https://pip.pypa.io/en/stable/) for machine learning (not deep learning!).
| Further reading.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import itertools
# %matplotlib inline
from keras.layers import *
from keras.models import *
from keras.optimizers import *
from keras.callbacks import *
import keras
from keras import backend as K
import tensorflow as tf
from rl.agents import *
from rl.policy import *
from rl.memory import *
from rl.random import *
import gym
from gym import Env, Space, spaces
env = gym.make('CartPole-v0')
env.seed(123)
env.reset()
nb_actions = env.action_space.n
observation_shape = env.observation_space.shape
window_length = 1
model_x = Input((window_length, ) + observation_shape)
model_y = Flatten()(model_x)
model_y = Dense(4, activation='elu')(model_y)
model_y = Dense(4, activation='elu')(model_y)
model_y = Dense(4, activation='elu')(model_y)
model_y = Dense(nb_actions)(model_y)
model = Model(model_x, model_y)
layer = model.layers[2]
layer.output
policy = EpsGreedyQPolicy()
memory = SequentialMemory(100000, window_length=window_length)
dqn = DQNAgent(model, nb_actions=nb_actions, policy=policy, memory=memory,
nb_steps_warmup=100, target_model_update=0.9)
dqn.compile(Adam(), metrics=['mse'])
hist = dqn.fit(env, nb_steps=100000, visualize=False, verbose=2,
callbacks=[LambdaCallback(on_epoch_end=lambda epoch, logs:print(epoch, logs))])
rewards=[]
rewards.extend(hist.history.get('episode_reward'))
plt.plot(rewards)
env.render(close=True)
dqn.test(env, nb_episodes=5)
| testzie/keras_rl_CartPole.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # XGBoost Regression
# XGBoost stands for eXtreme Gradient Boosting.
#
# It is an implementation of gradient boosting machines.XGBoost is a software library that you can download and
# install on your machine, then access from a variety of interfaces.The implementation of the model supports the
# features of the scikit-learn with new additions like regularization
# Import libraries
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import pandas as pd
from sklearn import metrics
from sklearn.metrics import r2_score
import xgboost as xgb
# Import dataset
dataset = pd.read_csv('/home/webtunix/Desktop/Regression/random.csv')
print(len(dataset))
# Split dataset into x and y sets
x = dataset.iloc[:,1:4].values
y = dataset.iloc[:,4].values
# Apply the training and testing on data
X_train, X_test, y_train, y_test = train_test_split(x, y, test_size = 0.3)
# Apply the XGB Regression
model = xgb.XGBRegressor(max_depth=3, learning_rate=0.1, n_estimators=100,
verbosity=1, objective='reg:linear',
booster='gbtree', n_jobs=1, gamma=0, min_child_weight=1, max_delta_step=0)
# max_depth (int) – Maximum tree depth for base learners.
#
# learning_rate (float) – Boosting learning rate (xgb’s “eta”)
#
# n_estimators (int) – Number of trees to fit.
#
# verbosity (int) – The degree of verbosity. Valid values are 0 (silent) - 3 (debug).
#
# booster (string) – Specify which booster to use: gbtree, gblinear or dart.
#
# n_jobs (int) – Number of parallel threads used to run xgboost. (replaces nthread)
#
# gamma (float) – Minimum loss reduction required to make a further partition on a leaf node of the tree.
#
# min_child_weight (int) – Minimum sum of instance weight(hessian) needed in a child.
#
# max_delta_step (int) – Maximum delta step we allow each tree’s weight estimation to be
# Fitting and score of model
model.fit(X_train,y_train)
model.score(X_train,y_train)
# predict the values
pred = model.predict(X_test)
print(pred)
# Accuracy of model
print("Accuracy:",r2_score(y_test,pred))
# Plotting the scatter graph of actual values and predicting values
# +
colors = np.random.rand(72)
#plot target and predicted values
plt.scatter(colors,y_test, c='orange',label='target')
plt.scatter(colors,pred, c='black',label='predicted')
#plot x and y lables
plt.xlabel('x')
plt.ylabel('y')
#plot title
plt.title('XGBoost Regression')
plt.legend()
plt.show()
# -
# # Research Infinite Solutions LLP
# by Research Infinite Solutions (https://www.ris-ai.com//)
#
# Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE
| Regression_models/xgboost_implement.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: deeplearning
# language: python
# name: deeplearning
# ---
# + pycharm={"is_executing": false}
import cv2
import numpy as np
from sklearn.metrics import pairwise
# + pycharm={"name": "#%%\n", "is_executing": false}
background = None
accumulated_weight = 0.5
roi_top = 20
roi_bottom = 300
roi_right = 300
roi_left = 600
# + pycharm={"name": "#%%\n", "is_executing": false}
def calc_accum_avg(frame, accumulated_weight):
global background
if background is None:
background = frame.copy().astype("float")
return None
cv2.accumulateWeighted(frame, background, accumulated_weight)
# + pycharm={"name": "#%%\n", "is_executing": false}
def segment(frame, threshold=25):
global background
diff = cv2.absdiff(background.astype("uint8"), frame)
ret, thresholded = cv2.threshold(diff, threshold, 255, cv2.THRESH_BINARY)
image, contours, hierarchy = cv2.findContours(thresholded.copy(), cv2.RETR_EXTERNAL, cv2.CHAIN_APPROX_SIMPLE)
if len(contours) == 0:
return None
else:
hand_segment = max(contours, key=cv2.contourArea)
return (thresholded, hand_segment)
# + pycharm={"name": "#%%\n", "is_executing": false}
def count_fingers(thresholded, hand_segment):
conv_hull = cv2.convexHull(hand_segment)
top = tuple(conv_hull[conv_hull[:, :, 1].argmin()][0])
bottom = tuple(conv_hull[conv_hull[:, :, 1].argmax()][0])
left = tuple(conv_hull[conv_hull[:, :, 0].argmin()][0])
right = tuple(conv_hull[conv_hull[:, :, 0].argmax()][0])
cX = (left[0] + right[0]) // 2
cY = (top[1] + bottom[1]) // 2
distance = pairwise.euclidean_distances([(cX, cY)], Y=[left, right, top, bottom])[0]
max_distance = distance.max()
radius = int(0.8 * max_distance)
circumference = (2 * np.pi * radius)
circular_roi = np.zeros(thresholded.shape[:2], dtype="uint8")
cv2.circle(circular_roi, (cX, cY), radius, 255, 10)
circular_roi = cv2.bitwise_and(thresholded, thresholded, mask=circular_roi)
image, contours, hierarchy = cv2.findContours(circular_roi.copy(), cv2.RETR_EXTERNAL, cv2.CHAIN_APPROX_NONE)
count = 0
for cnt in contours:
(x, y, w, h) = cv2.boundingRect(cnt)
out_of_wrist = ((cY + (cY * 0.25)) > (y + h))
limit_points = ((circumference * 0.25) > cnt.shape[0])
if out_of_wrist and limit_points:
count += 1
return count
# + pycharm={"name": "#%%\n", "is_executing": false}
cam = cv2.VideoCapture(0)
num_frames = 0
while True:
ret, frame = cam.read()
frame = cv2.flip(frame, 1)
frame_copy = frame.copy()
roi = frame[roi_top:roi_bottom, roi_right:roi_left]
gray = cv2.cvtColor(roi, cv2.COLOR_BGR2GRAY)
gray = cv2.GaussianBlur(gray, (7, 7), 0)
if num_frames < 60:
calc_accum_avg(gray, accumulated_weight)
if num_frames <= 59:
cv2.putText(frame_copy, "WAIT! GETTING BACKGROUND AVG.", (200, 400), cv2.FONT_HERSHEY_SIMPLEX, 1, (0,0,255), 2)
cv2.imshow("Finger Count",frame_copy)
else:
hand = segment(gray)
if hand is not None:
thresholded, hand_segment = hand
cv2.drawContours(frame_copy, [hand_segment + (roi_right, roi_top)], -1, (255, 0, 0),1)
fingers = count_fingers(thresholded, hand_segment)
cv2.putText(frame_copy, str(fingers), (70, 45), cv2.FONT_HERSHEY_SIMPLEX, 1, (0,0,255), 2)
cv2.imshow("Thesholded", thresholded)
cv2.rectangle(frame_copy, (roi_left, roi_top), (roi_right, roi_bottom), (0,0,255), 5)
num_frames += 1
cv2.imshow("Finger Count", frame_copy)
k = cv2.waitKey(1) & 0xFF
if k == 27:
break
cam.release()
cv2.destroyAllWindows()
# + pycharm={"name": "#%%\n", "is_executing": false}
# + pycharm={"name": "#%%\n"}
| OpenCV_Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.1
# language: julia
# name: julia-1.6
# ---
# ## <center> Plotly isosurface, slices and caps </center>
# Let F be a function defined on a parallelipiped. We illustrate how Plotly chart type, `isosurface`, defines an isosurface of equation F(x,y,z) = isoval.
#
# The example given below also shows a slice within the sub-volume bounded by the isosurface, and the caps produced by the bounding planes of the initial volume.
#
# Plotly `isosurface` has a special definition. It doesn't require an isovalue to get an isosurface, but two values, `isomin= m`, and `isomax = M`. If we are setting, `surface_show=true`, and `surface_counts=1`, then one generates the isosurface of equation $F(x,y,z)=(m+M)/2$.
#
# For a given colorscale, the interval [isomin, isomax] is mapped to that colorscale, and as a consequence the isosurface points
# are mapped to a unique color, corresponding to the isovalue, $(m+M)/2$, i.e. to the central color on the colorbar.
#
# To make visible the sub-volume bounded by the isosurface we can set a non-dense pattern.
#
# If `surface_show=true`, and `surface_count=2`, then two isosurfaces are drawn: F(x,y,z)=isomin, and F(x,y, z)=isomax. For two close values, `isomin`, and `isomax`, we get a surface with two faces (see the notebook: `15-Isosurface-with-two-faces.ipynb`).
using PlotlyJS, Rotations
include("src/plotlyju.jl")
include("src/PlotlyColorSchemes.jl");
# +
F(x,y,z) = x^4 + y^4 + z^4 - (x^2+y^2+z^2)^2 + 3*(x^2+y^2+z^2) - 4.35
# Define a 3D grid, and flatten each coordinate variable, because
# isosurface accepts only vectors for x, y, z, and value:
xl = yl = zl = LinRange(-2, 2, 50)
x = vec([xx for zz in zl, yy in yl, xx in xl])
y = vec([yy for zz in zl, yy in yl, xx in xl])
z = vec([zz for zz in zl, yy in yl, xx in xl])
Fvals = F.(x, y, z);
isosurf = isosurface(
x=x,
y=y,
z=z,
value=Fvals,
surface=attr(show=true, fill= 0.65, pattern= "all", count=1),
colorscale=pl_BrBG,
colorbar_len=0.75,
slices=attr(z= attr(show=true,
fill=1, #no patterns
locations=[-0.8])), #slice cut by plane z=-0.8
caps=attr(x= attr(show=true),
y= attr(show=true),
z= attr(show=true)),
isomin= -2,
isomax= 2,
)
black_bg = Dict( #black background
:showbackground=>true,
:backgroundcolor=>"rgb(20, 20, 20)",
:gridcolor=>"rgb(150, 150, 150)",
:zeroline=>false)
layout = Layout(width=600, height=600,
scene=attr(camera_eye=attr(x=1.55, y=1.55, z=0.98),
xaxis=black_bg, yaxis=black_bg, zaxis=black_bg))
pl = Plot(isosurf, layout, style=plotlyju)
# -
# 
# The same isosurface, but with pl_curl, as colorscale, default style background, and particular settings for lighting+lightposition, given in the last cell:
# 
lighting=attr(ambient=0.5,
diffuse=0.5,
fresnel=0.25,
specular=0.25,
roughness=0.25,
facenormalsepsilon=0,
vertexnormalsepsilon=0)
lightposition=attr(x=100,
y=100,
z=1000)
| 14-Plotly-isosurface-slices-and-caps.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""
Starter code for the regression mini-project of Udacity Intro to Machine Learning.
Loads up/formats a modified version of the dataset
(why modified? we've removed some trouble points
that you'll find yourself in the outliers mini-project).
Draws a little scatterplot of the training/testing data
"""
import sys
import pickle
sys.path.append("../tools/")
from feature_format import featureFormat, targetFeatureSplit
dictionary = pickle.load( open("../final_project/final_project_dataset_modified.pkl", "rb") )
# -
### list the features you want to look at--first item in the
### list will be the "target" feature
features_list = ["bonus", "long_term_incentive"]
data = featureFormat(dictionary, features_list, remove_any_zeroes=True)
target, features = targetFeatureSplit( data )
from sklearn.model_selection import train_test_split
feature_train, feature_test, target_train, target_test = train_test_split(features, target, test_size=0.5, random_state=42)
train_color = "b"
test_color = "r"
from sklearn.linear_model import LinearRegression
reg = LinearRegression().fit(feature_train, target_train)
reg.score(feature_train, target_train)
reg.coef_
reg.intercept_
# +
### draw the scatterplot, with color-coded training and testing points
import matplotlib.pyplot as plt
for feature, target in zip(feature_test, target_test):
plt.scatter( feature, target, color=test_color )
for feature, target in zip(feature_train, target_train):
plt.scatter( feature, target, color=train_color )
### labels for the legend
plt.scatter(feature_test[0], target_test[0], color=test_color, label="test")
plt.scatter(feature_test[0], target_test[0], color=train_color, label="train")
# -
### draw the regression line, once it's coded
try:
plt.plot(feature_test, reg.predict(feature_test) )
except NameError:
pass
plt.xlabel(features_list[1])
plt.ylabel(features_list[0])
plt.legend()
plt.show()
| Lesson07_Regression/finance_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:bench_env]
# language: python
# name: conda-env-bench_env-py
# ---
# + [markdown] papermill={"duration": 0.005069, "end_time": "2022-06-03T06:59:11.850235", "exception": false, "start_time": "2022-06-03T06:59:11.845166", "status": "completed"} tags=[]
# # TF CNN Classifier
#
# To run this notebook on an another benchmark, use
#
# ```
# papermill utils/tf_cnn_classifier.ipynb tf_cnn_experiments/[DATASET NAME].ipynb -p DATASET [DATASET NAME]
# ```
# + papermill={"duration": 0.010701, "end_time": "2022-06-03T06:59:11.865007", "exception": false, "start_time": "2022-06-03T06:59:11.854306", "status": "completed"} tags=["parameters"]
DATASET = 'demo_coding_vs_intergenomic_seqs'
VERSION = 0
BATCH_SIZE = 64
EPOCHS = 10
# + papermill={"duration": 0.007732, "end_time": "2022-06-03T06:59:11.876923", "exception": false, "start_time": "2022-06-03T06:59:11.869191", "status": "completed"} tags=["injected-parameters"]
# Parameters
DATASET = "human_nontata_promoters"
# + papermill={"duration": 0.009411, "end_time": "2022-06-03T06:59:11.891473", "exception": false, "start_time": "2022-06-03T06:59:11.882062", "status": "completed"} tags=[]
print(DATASET, VERSION, BATCH_SIZE, EPOCHS)
# + [markdown] papermill={"duration": 0.005788, "end_time": "2022-06-03T06:59:11.901119", "exception": false, "start_time": "2022-06-03T06:59:11.895331", "status": "completed"} tags=[]
# # Data download
# + papermill={"duration": 3.547173, "end_time": "2022-06-03T06:59:15.452503", "exception": false, "start_time": "2022-06-03T06:59:11.905330", "status": "completed"} tags=[]
from pathlib import Path
import tensorflow as tf
import tensorflow_addons as tfa
import numpy as np
from genomic_benchmarks.loc2seq import download_dataset
from genomic_benchmarks.data_check import is_downloaded, info
from genomic_benchmarks.models.tf import vectorize_layer
from genomic_benchmarks.models.tf import get_basic_cnn_model_v0 as get_model
if not is_downloaded(DATASET):
download_dataset(DATASET)
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "completed"} tags=[]
info(DATASET)
# + [markdown] papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "completed"} tags=[]
# ## TF Dataset object
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "completed"} tags=[]
SEQ_PATH = Path.home() / '.genomic_benchmarks' / DATASET
CLASSES = [x.stem for x in (SEQ_PATH/'train').iterdir() if x.is_dir()]
NUM_CLASSES = len(CLASSES)
train_dset = tf.keras.preprocessing.text_dataset_from_directory(
SEQ_PATH / 'train',
batch_size=BATCH_SIZE,
class_names=CLASSES)
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "completed"} tags=[]
if NUM_CLASSES > 2:
train_dset = train_dset.map(lambda x, y: (x, tf.one_hot(y, depth=NUM_CLASSES)))
# + [markdown] papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "completed"} tags=[]
# ## Text vectorization
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "completed"} tags=[]
vectorize_layer.adapt(train_dset.map(lambda x, y: x))
VOCAB_SIZE = len(vectorize_layer.get_vocabulary())
vectorize_layer.get_vocabulary()
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "completed"} tags=[]
def vectorize_text(text, label):
text = tf.expand_dims(text, -1)
return vectorize_layer(text)-2, label
train_ds = train_dset.map(vectorize_text)
# + [markdown] papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "completed"} tags=[]
# ## Model training
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "completed"} tags=[]
model = get_model(NUM_CLASSES, VOCAB_SIZE)
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "completed"} tags=[]
history = model.fit(
train_ds,
epochs=EPOCHS)
# + [markdown] papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "completed"} tags=[]
# ## Evaluation on the test set
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "completed"} tags=[]
test_dset = tf.keras.preprocessing.text_dataset_from_directory(
SEQ_PATH / 'test',
batch_size=BATCH_SIZE,
class_names=CLASSES)
if NUM_CLASSES > 2:
test_dset = test_dset.map(lambda x, y: (x, tf.one_hot(y, depth=NUM_CLASSES)))
test_ds = test_dset.map(vectorize_text)
# + papermill={"duration": null, "end_time": null, "exception": null, "start_time": null, "status": "completed"} tags=[]
model.evaluate(test_ds)
| experiments/tf_cnn_experiments/human_nontata_promoters.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:light
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from imp import reload
import sys
sys.path.insert(0, '..')
import utilities as ut
from skimage import img_as_float
hst = ut.read_hyperstack('../data/wt_gbe_20180110.h5')
ut.imshow(hst[0,350:500,100:900])
# # Segmentation to remove background
fhst = img_as_float(hst)
test = fhst[0,350:500,100:900]
ut.imshow(test)
from scipy.ndimage import gaussian_filter
gaus = gaussian_filter(test,4)
ut.imshow(gaus)
from skimage.feature import canny
edges = canny(gaus)
type(edges)
plt.imshow(edges)
from scipy import ndimage as ndi
fill = ndi.binary_fill_holes(edges)
plt.imshow(fill)
from skimage.filters import sobel
elevation_map = sobel(test)
ut.imshow(elevation_map)
markers = np.zeros_like(test)
markers[test<0.1] = 1
markers[test>0.9] = 2
plt.imshow(markers)
from skimage.morphology import watershed
segmentation = watershed(elevation_map,markers)
plt.imshow(segmentation)
| notebooks/20181006.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 딥러닝의 미래
# - 이미지 스타일(화풍) 변환
# - 딥러닝을 활용해 화가처럼 그림을 그리는 것
# - 콘텐츠 이미지와 스타일 이미를 조합해 새로운 그림을 그려준다.
# - 이 기술은 네트워크의 중간 데이터가 콘텐츠 이미지의 중간 데이터와 비슷해지도록 학습한다. 이렇게 하면 입력 이미지를 콘텐츠 이미지의 형태를 흉내낼 수 있다.
# - 또, 스타일 이미지의 화풍을 흡수하기 위해 스타일 행렬이라는 개념을 도입한다. 그 스타일 행렬의 오차를 줄이도록 학습하여 입력 이미지를 고흐의 화풍과 비슷해지게 만들 수 있는 것이다.
# - 이미지 생성
# - 이미지 스타일 변환은 새로운 그림을 생성하려면 이미지 두 장을 입력해야 했다.
# - 하지만 이 방법은 입력 이미지 없이도 새로운 이미지를 그려내는 연구이다.
# - DCGAN 기법이 있다.
# - 해당 기법의 핵심은 생성자와 식별자로 불리는 2개의 신경망을 이용한다는 점이다.
# - 생성자가 진짜와 똑같은 이미지를 생성하고 식별자는 그것이 진짜인지(생성자가 생성한 이미지인지, 아니면 실제로 촬영된이미지인지)를 판정한다. 이런 방식으로 둘을 겨루도록 학습시켜, 생성자는 더 정교한 가짜 이미지 생성 기술을 학습하고 식별자는 더 정확하게 간파할 수 있는 감정사로 성장하는 것이다. 이렇게 둘의 능력을 수련시킨다는 개념이 GAN이다.
# - 자율 주행
# - 자율 주행은 주행 경로를 정하는 경로 계획 기술과 카메라나 레이저 등의 탐사 기술 그리고 주위 환경을 올바르게 인식하는 기술이 필요하다 그 중 주위 환경을 올바르게 인식하는 기술이 가장 중요하다.
# - 예를 들어 SegNet이라는 CNN기반 신경망은 주변 환경을 정확하게 인식해낸다.
# - 입력 이미지를 분할(픽셀 수준에서 판정)하고 있다.
# - Deep Q-Network(강화학습)
# - 사람이 자전거를 배울 때 시행착오를 겪으며 배우듯, 컴퓨터도 시행착오 과정에서 스스로 학습하게 하려는 분야이다.
# - 가르침에 의존하는 지도 학습과는 다른 분야이다.
# - 에이전트라는 것이 환경에 맞게 행동을 선택하고, 그 행동에 의해서 환경이 변한다는 게 기본적인 틀이다. 환경이 변화하면 에이전트는 어떠한 보상을 얻는다. 강화학습의 목적은 더 나은 보상을 받는 쪽으로 에이전트의 행동 지침을 바로잡는 것이다.
# - 이때 보상은 정해진 것이 아니라 예상 보상이다. 예를 들어, 게임 캐릭터 마리오를 오른쪽으로 이동시켰을 때 얻는 보상이 항상 명확하진 않다. 상황에 따라 그것이 100원이 될 수도 혹은 장애물이 될 수도 혹은 1000원이 될 수도 있듯 어떤 상황에서 이동한 것이냐에 따라 보상은 천차만별이 될 수 있다. 이런 불명확한 상황에서는 게임 점수(동전을 먹거나 적을 쓰러뜨리는 등)나 게임 종료 등의 명확한 지표로부터 역산해서 예상 보상을 정해야 한다.
# - DQN
# - Q라는 강화학습 알고리즘을 기초로 한다. Q학습에서는 최적 행동 가치 함수로 최적인 행동을 정한다. 이 함수를 딥러닝으로 비슷하게 흉내 내어 사용하는 것이 DQN이다.
# ---
# - 간단히 1회독이 종료됐다.
# - 2회독부터는 진도 속도를 늦추되 좀 더 정밀히 공부해보자
| _notebooks/2022-02-23-dl.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++17
// language: C++17
// name: xcpp17
// ---
// + [markdown] toc=true
// <h1>Table of Contents<span class="tocSkip"></span></h1>
// <div class="toc"><ul class="toc-item"></ul></div>
// -
#include<iostream>
#include <vector>
using namespace std;
// +
vector<int> v = {1,2,3,4};
for (auto &x: v) cout << x << " ";
// -
for_each(v.begin(),v.end(),[](int &n) {return ++n;});
v
| Learn_cpp/practice.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Facial Keypoint Detection
#
# ## Load and Visualize Data
#
# #### Training and Testing Data
# +
import glob
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import cv2
# +
key_pts_frame = pd.read_csv('data/training_frames_keypoints.csv')
n = 0
image_name = key_pts_frame.iloc[n, 0]
key_pts = key_pts_frame.iloc[n, 1:].as_matrix()
key_pts = key_pts.astype('float').reshape(-1, 2)
print('Image name: ', image_name)
print('Landmarks shape: ', key_pts.shape)
print('First 4 key pts: {}'.format(key_pts[:4]))
# -
# print out some stats about the data
print('Number of images: ', key_pts_frame.shape[0])
# ## Look at some images
def show_keypoints(image, key_pts):
"""Show image with keypoints"""
plt.imshow(image)
plt.scatter(key_pts[:, 0], key_pts[:, 1], s=20, marker='.', c='m')
# +
# Display a few different types of images by changing the index n
# select an image by index in our data frame
n = 0
image_name = key_pts_frame.iloc[n, 0]
key_pts = key_pts_frame.iloc[n, 1:].as_matrix()
key_pts = key_pts.astype('float').reshape(-1, 2)
plt.figure(figsize=(5, 5))
show_keypoints(mpimg.imread(os.path.join('data/training/', image_name)), key_pts)
plt.show()
# -
# ## Dataset class and Transformations
#
# #### Dataset class
#
# +
from torch.utils.data import Dataset, DataLoader
class FacialKeypointsDataset(Dataset):
"""Face Landmarks dataset."""
def __init__(self, csv_file, root_dir, transform=None):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.key_pts_frame = pd.read_csv(csv_file)
self.root_dir = root_dir
self.transform = transform
def __len__(self):
return len(self.key_pts_frame)
def __getitem__(self, idx):
image_name = os.path.join(self.root_dir,
self.key_pts_frame.iloc[idx, 0])
image = mpimg.imread(image_name)
# if image has an alpha color channel, get rid of it
if(image.shape[2] == 4):
image = image[:,:,0:3]
key_pts = self.key_pts_frame.iloc[idx, 1:].as_matrix()
key_pts = key_pts.astype('float').reshape(-1, 2)
sample = {'image': image, 'keypoints': key_pts}
if self.transform:
sample = self.transform(sample)
return sample
# +
# Construct the dataset
face_dataset = FacialKeypointsDataset(csv_file='data/training_frames_keypoints.csv',
root_dir='data/training/')
# print some stats about the dataset
print('Length of dataset: ', len(face_dataset))
# +
# Display a few of the images from the dataset
num_to_display = 3
for i in range(num_to_display):
# define the size of images
fig = plt.figure(figsize=(20,10))
# randomly select a sample
rand_i = np.random.randint(0, len(face_dataset))
sample = face_dataset[rand_i]
# print the shape of the image and keypoints
print(i, sample['image'].shape, sample['keypoints'].shape)
ax = plt.subplot(1, num_to_display, i + 1)
ax.set_title('Sample #{}'.format(i))
# Using the same display function, defined earlier
show_keypoints(sample['image'], sample['keypoints'])
# -
# ## Transforms
# +
import torch
from torchvision import transforms, utils
# tranforms
class Normalize(object):
"""Convert a color image to grayscale and normalize the color range to [0,1]."""
def __call__(self, sample):
image, key_pts = sample['image'], sample['keypoints']
image_copy = np.copy(image)
key_pts_copy = np.copy(key_pts)
# convert image to grayscale
image_copy = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
# scale color range from [0, 255] to [0, 1]
image_copy= image_copy/255.0
# scale keypoints to be centered around 0 with a range of [-1, 1]
# mean = 100, sqrt = 50, so, pts should be (pts - 100)/50
key_pts_copy = (key_pts_copy - 100)/50.0
return {'image': image_copy, 'keypoints': key_pts_copy}
class Rescale(object):
"""Rescale the image in a sample to a given size.
Args:
output_size (tuple or int): Desired output size. If tuple, output is
matched to output_size. If int, smaller of image edges is matched
to output_size keeping aspect ratio the same.
"""
def __init__(self, output_size):
assert isinstance(output_size, (int, tuple))
self.output_size = output_size
def __call__(self, sample):
image, key_pts = sample['image'], sample['keypoints']
h, w = image.shape[:2]
if isinstance(self.output_size, int):
if h > w:
new_h, new_w = self.output_size * h / w, self.output_size
else:
new_h, new_w = self.output_size, self.output_size * w / h
else:
new_h, new_w = self.output_size
new_h, new_w = int(new_h), int(new_w)
img = cv2.resize(image, (new_w, new_h))
# scale the pts, too
key_pts = key_pts * [new_w / w, new_h / h]
return {'image': img, 'keypoints': key_pts}
class RandomCrop(object):
"""Crop randomly the image in a sample.
Args:
output_size (tuple or int): Desired output size. If int, square crop
is made.
"""
def __init__(self, output_size):
assert isinstance(output_size, (int, tuple))
if isinstance(output_size, int):
self.output_size = (output_size, output_size)
else:
assert len(output_size) == 2
self.output_size = output_size
def __call__(self, sample):
image, key_pts = sample['image'], sample['keypoints']
h, w = image.shape[:2]
new_h, new_w = self.output_size
top = np.random.randint(0, h - new_h)
left = np.random.randint(0, w - new_w)
image = image[top: top + new_h,
left: left + new_w]
key_pts = key_pts - [left, top]
return {'image': image, 'keypoints': key_pts}
class ToTensor(object):
"""Convert ndarrays in sample to Tensors."""
def __call__(self, sample):
image, key_pts = sample['image'], sample['keypoints']
# if image has no grayscale color channel, add one
if(len(image.shape) == 2):
# add that third color dim
image = image.reshape(image.shape[0], image.shape[1], 1)
# swap color axis because
# numpy image: H x W x C
# torch image: C X H X W
image = image.transpose((2, 0, 1))
return {'image': torch.from_numpy(image),
'keypoints': torch.from_numpy(key_pts)}
# -
# ## Test out the transforms
# +
# test out some of these transforms
rescale = Rescale(100)
crop = RandomCrop(50)
composed = transforms.Compose([Rescale(250),
RandomCrop(224)])
# apply the transforms to a sample image
test_num = 500
sample = face_dataset[test_num]
fig = plt.figure()
for i, tx in enumerate([rescale, crop, composed]):
transformed_sample = tx(sample)
ax = plt.subplot(1, 3, i + 1)
plt.tight_layout()
ax.set_title(type(tx).__name__)
show_keypoints(transformed_sample['image'], transformed_sample['keypoints'])
plt.show()
# -
# ## Create the transformed dataset
# +
# define the data tranform
# order matters! i.e. rescaling should come before a smaller crop
data_transform = transforms.Compose([Rescale(250),
RandomCrop(224),
Normalize(),
ToTensor()])
# create the transformed dataset
transformed_dataset = FacialKeypointsDataset(csv_file='data/training_frames_keypoints.csv',
root_dir='data/training/',
transform=data_transform)
# +
# print some stats about the transformed data
print('Number of images: ', len(transformed_dataset))
# make sure the sample tensors are the expected size
for i in range(5):
sample = transformed_dataset[i]
print(i, sample['image'].size(), sample['keypoints'].size())
| 1. Load and Visualize Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## In this demo, we will show how to use uq360 metrics for hyperparameters optimization using Lale and sklearn gridsearchCV.
# %matplotlib inline
import sys
sys.path.append("../../")
# !pip install lale
# !pip install 'liac-arff>=2.4.0'
from uq360.utils.misc import make_sklearn_compatible_scorer
# ### The uq360 metrics can be converted to a sklearn scorer usign `make_sklearn_compatible_scorer` utility function.
#
# #### We will now show to how to convert ECE and AURRRC metric in uq360 into scorer that can be used with LALE framework for hyperparameter optimization.
sklearn_aurrrc = make_sklearn_compatible_scorer(task_type="classification", metric="aurrrc", greater_is_better=False)
sklearn_ece = make_sklearn_compatible_scorer(task_type="classification", metric="ece", greater_is_better=False)
import lale
from lale.lib.lale import Hyperopt
lale.wrap_imported_operators()
from sklearn import datasets
X, y = datasets.load_breast_cancer(return_X_y=True)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
from lale.lib.sklearn import AdaBoostClassifier as Model
clf_ece = Hyperopt(estimator=Model, cv=3, max_evals=20, scoring=sklearn_ece, verbose=True)
clf_aurrrc = Hyperopt(estimator=Model, cv=3, max_evals=20, scoring=sklearn_aurrrc, verbose=True)
trained_with_ece = clf_ece.fit(X_train, y_train)
trained_with_aurrrc = clf_aurrrc.fit(X_train, y_train)
clf_accuracy = Hyperopt(estimator=Model, cv=3, max_evals=20, verbose=True)
trained_with_accuracy = clf_accuracy.fit(X_train, y_train)
from sklearn.metrics import classification_report
print(classification_report(y_test, trained_with_accuracy.predict(X_test)))
print(classification_report(y_test, trained_with_ece.predict(X_test)))
print(classification_report(y_test, trained_with_aurrrc.predict(X_test)))
# ## Lets us now see how to use `uq360` models with sklearn's `GridsearchCV`
#
# ### We use the boston housing dataset with `QuantileRegression` model while optimizing the `picp` metric.
from sklearn.model_selection import GridSearchCV
import pandas as pd
# +
house_prices_dataset = datasets.load_boston()
house_prices_df = pd.DataFrame(house_prices_dataset['data'])
house_prices_df.columns = house_prices_dataset['feature_names']
all_features = ['RM','CRIM','PTRATIO', 'DIS']
X = house_prices_df[all_features].values
y = house_prices_dataset['target']
# -
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
# +
base_config = {
"alpha":0.95,
"n_estimators":20,
"max_depth":3,
"learning_rate":0.1,
"min_samples_leaf":20,
"min_samples_split":20
}
configs = {"config": []}
for num_estimators in [1, 2, 5, 10, 20, 30, 40, 50]:
config = base_config.copy()
config["n_estimators"] = num_estimators
configs["config"].append(config)
# -
from uq360.algorithms.quantile_regression import QuantileRegression
sklearn_picp = make_sklearn_compatible_scorer(task_type="regression", metric="picp", greater_is_better=True)
clf = GridSearchCV(QuantileRegression(config=base_config), configs, scoring=sklearn_picp)
clf.fit(X_train, y_train)
df = pd.DataFrame(clf.cv_results_)
df
| examples/autoai/demo_autoai.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''udemyPro_py388'': venv)'
# name: python3
# ---
# +
# in einem Set darf jeder Wert nur 1x vorkommen!!
# sets müssen immutable sein!
my_set1 = {1, 2, 3}
print(my_set1, type(my_set1))
# doppelte Werte werden verworfen
my_set1 = {1, 2, 3, 1}
print(my_set1, type(my_set1))
# -
# Konstruktor
my_set2 = set([1, 2, 3, 1])
print(my_set2, type(my_set2))
# +
my_set3 = {4, 5, 6}
my_set4 = {6, 7, 8}
# Was haben beide Sets gemeinsam
print(my_set3.intersection(my_set4))
# beide Sets mergen
print(my_set3.union(my_set4))
# -
if 4 in my_set3:
print("yes")
print("Length", len(my_set3))
# +
my_set3 = {4, 5, 6}
my_set3.add(42)
print("set3:", my_set3)
# remove a random element
my_set3.pop()
print("set3:", my_set3)
# remove a specific element
my_set3.discard(4)
print("set3:", my_set3)
# -
my_idxs = [1, 2, 3, 3 , 4, 5 ,5]
my_set = set(my_idxs)
print(my_set)
| Python/zzz_training_challenge/UdemyPythonPro/Chapter4_Iterables/Sets/set.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import os
import pprint
import torch
import torch.nn.parallel
import torch.backends.cudnn as cudnn
import torch.optim
import torch.utils.data
import torch.utils.data.distributed
import torchvision.transforms as transforms
from tools import init_paths
from config import cfg
from config import update_config
from core.loss import JointsMSELoss
from core.function import validate
from utils.utils import create_logger
import dataset
import models
# +
# modifying validate function
# %load_ext autoreload
# %autoreload 2
from core.function import AverageMeter
from core.function import _print_name_value
import numpy as np
import time
import os
from utils.transforms import flip_back
from core.evaluate import accuracy
from core.inference import get_final_preds
from utils.vis import save_debug_images
def my_validate(val_loader, val_dataset, model, criterion, output_dir,
tb_log_dir, writer_dict=None):
# switch to evaluate mode
model.eval()
with torch.no_grad():
all_preds, all_boxes, losses, acc, image_path, filenames, imgnums = predict_batch(val_loader,val_dataset,model,criterion,output_dir)
name_values, perf_indicator = val_dataset.evaluate(
cfg, all_preds, output_dir, all_boxes, image_path,
filenames, imgnums
)
model_name = cfg.MODEL.NAME
if isinstance(name_values, list):
for name_value in name_values:
_print_name_value(name_value, model_name)
else:
_print_name_value(name_values, model_name)
if writer_dict:
writer = writer_dict['writer']
global_steps = writer_dict['valid_global_steps']
writer.add_scalar(
'valid_loss',
losses.avg,
global_steps
)
writer.add_scalar(
'valid_acc',
acc.avg,
global_steps
)
if isinstance(name_values, list):
for name_value in name_values:
writer.add_scalars(
'valid',
dict(name_value),
global_steps
)
else:
writer.add_scalars(
'valid',
dict(name_values),
global_steps
)
writer_dict['valid_global_steps'] = global_steps + 1
return perf_indicator
def predict_batch(val_loader, val_dataset, model, criterion, output_dir):
end = time.time()
batch_time = AverageMeter()
losses = AverageMeter()
acc = AverageMeter()
num_samples = len(val_dataset)
all_preds = np.zeros(
(num_samples, cfg.MODEL.NUM_JOINTS, 3),
dtype=np.float32
)
all_boxes = np.zeros((num_samples, 6))
image_path = []
filenames = []
imgnums = []
idx = 0
for i, (input, target, target_weight, meta) in enumerate(val_loader):
# compute output
outputs = model(input)
if isinstance(outputs, list):
output = outputs[-1]
else:
output = outputs
if cfg.TEST.FLIP_TEST:
# this part is ugly, because pytorch has not supported negative index
# input_flipped = model(input[:, :, :, ::-1])
input_flipped = np.flip(input.cpu().numpy(), 3).copy()
input_flipped = torch.from_numpy(input_flipped).cuda()
outputs_flipped = model(input_flipped)
if isinstance(outputs_flipped, list):
output_flipped = outputs_flipped[-1]
else:
output_flipped = outputs_flipped
output_flipped = flip_back(output_flipped.cpu().numpy(),
val_dataset.flip_pairs)
output_flipped = torch.from_numpy(output_flipped.copy()).cuda()
# feature is not aligned, shift flipped heatmap for higher accuracy
if cfg.TEST.SHIFT_HEATMAP:
output_flipped[:, :, :, 1:] = \
output_flipped.clone()[:, :, :, 0:-1]
output = (output + output_flipped) * 0.5
target = target.cuda(non_blocking=True)
target_weight = target_weight.cuda(non_blocking=True)
loss = criterion(output, target, target_weight)
num_images = input.size(0)
# measure accuracy and record loss
losses.update(loss.item(), num_images)
_, avg_acc, cnt, pred = accuracy(output.cpu().numpy(),
target.cpu().numpy())
acc.update(avg_acc, cnt)
# measure elapsed time
batch_time.update(time.time() - end)
end = time.time()
c = meta['center'].numpy()
s = meta['scale'].numpy()
score = meta['score'].numpy()
preds, maxvals = get_final_preds(
cfg, output.clone().cpu().numpy(), c, s)
all_preds[idx:idx + num_images, :, 0:2] = preds[:, :, 0:2]
all_preds[idx:idx + num_images, :, 2:3] = maxvals
# double check this all_boxes parts
all_boxes[idx:idx + num_images, 0:2] = c[:, 0:2]
all_boxes[idx:idx + num_images, 2:4] = s[:, 0:2]
all_boxes[idx:idx + num_images, 4] = np.prod(s*200, 1)
all_boxes[idx:idx + num_images, 5] = score
image_path.extend(meta['image'])
idx += num_images
if i % cfg.PRINT_FREQ == 0:
msg = 'Test: [{0}/{1}]\t' \
'Time {batch_time.val:.3f} ({batch_time.avg:.3f})\t' \
'Loss {loss.val:.4f} ({loss.avg:.4f})\t' \
'Accuracy {acc.val:.3f} ({acc.avg:.3f})'.format(
i, len(val_loader), batch_time=batch_time,
loss=losses, acc=acc)
logger.info(msg)
prefix = '{}_{}'.format(
os.path.join(output_dir, 'val'), i
)
save_debug_images(cfg, input, meta, target, pred*4, output,
prefix)
return all_preds, all_boxes, losses, acc, image_path, filenames, imgnums
# -
def update_my_config():
cfg.defrost()
cfg.merge_from_file('experiments/mpii/hrnet/w32_256x256_adam_lr1e-3.yaml')
opts = ["TEST.MODEL_FILE", "models/HRNet/pose_mpii/pose_hrnet_w32_256x256.pth"]
# opts = ["TEST.MODEL_FILE", "/mnt/models/HRNet/pose_mpii/pose_hrnet_w32_256x256.pth"]
cfg.merge_from_list(opts)
cfg.OUTPUT_DIR = "output_test"
cfg.LOG_DIR = "log_test"
cfg.freeze()
# +
update_my_config()
logger, output_dir, tb_log_dir = create_logger(
cfg, "experiments/mpii/hrnet/w32_256x256_adam_lr1e-3.yaml", 'valid')
#logger.info(pprint.pformat(args))
logger.info(cfg)
# cudnn related setting
cudnn.benchmark = cfg.CUDNN.BENCHMARK
torch.backends.cudnn.deterministic = cfg.CUDNN.DETERMINISTIC
torch.backends.cudnn.enabled = cfg.CUDNN.ENABLED
model = eval('models.'+cfg.MODEL.NAME+'.get_pose_net')(
cfg, is_train=False
)
if cfg.TEST.MODEL_FILE:
logger.info('=> loading model from {}'.format(cfg.TEST.MODEL_FILE))
model.load_state_dict(torch.load(cfg.TEST.MODEL_FILE), strict=False)
else:
model_state_file = os.path.join(
final_output_dir, 'final_state.pth'
)
logger.info('=> loading model from {}'.format(model_state_file))
model.load_state_dict(torch.load(model_state_file))
model = torch.nn.DataParallel(model, device_ids=[0]).cuda()
# model = torch.nn.DataParallel(model, device_ids=cfg.GPUS).cuda()
# define loss function (criterion) and optimizer
criterion = JointsMSELoss(
use_target_weight=cfg.LOSS.USE_TARGET_WEIGHT
).cuda()
# Data loading code
normalize = transforms.Normalize(
mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225]
)
val_dataset = eval('dataset.'+cfg.DATASET.DATASET)(
cfg, cfg.DATASET.ROOT, cfg.DATASET.TEST_SET, False,
transforms.Compose([
transforms.ToTensor(),
normalize,
])
)
val_loader = torch.utils.data.DataLoader(
val_dataset,
batch_size=cfg.TEST.BATCH_SIZE_PER_GPU*len(cfg.GPUS),
shuffle=False,
num_workers=cfg.WORKERS,
pin_memory=True
)
# evaluate on validation set
# my_validate(val_loader, val_dataset, model, criterion,
# output_dir, tb_log_dir)
# switch to evaluate mode
model.eval()
with torch.no_grad():
all_preds, all_boxes, losses, acc, image_path, filenames, imgnums = predict_batch(val_loader, val_dataset, model, criterion, output_dir)
name_values, perf_indicator = val_dataset.evaluate(
cfg, all_preds, output_dir, all_boxes, image_path,
filenames, imgnums
)
model_name = cfg.MODEL.NAME
if isinstance(name_values, list):
for name_value in name_values:
_print_name_value(name_value, model_name)
else:
_print_name_value(name_values, model_name)
# -
print(all_preds.shape)
print(all_boxes.shape)
print(len(image_path))
# ## Analyzing results
# `lib/dataset/mpii.py`
# +
import numpy as np
from scipy.io import loadmat, savemat
from dataset.JointsDataset import JointsDataset
from collections import OrderedDict
def my_evaluate(cfg, preds, output_dir, *args, **kwargs):
# convert 0-based index to 1-based index
preds = preds[:, :, 0:2] + 1.0
if output_dir:
pred_file = os.path.join(output_dir, 'pred.mat')
savemat(pred_file, mdict={'preds': preds})
if 'test' in cfg.DATASET.TEST_SET:
return {'Null': 0.0}, 0.0
SC_BIAS = 0.6
threshold = 0.5
gt_file = os.path.join(cfg.DATASET.ROOT,
'annot',
'gt_{}.mat'.format(cfg.DATASET.TEST_SET))
gt_dict = loadmat(gt_file)
dataset_joints = gt_dict['dataset_joints']
jnt_missing = gt_dict['jnt_missing']
pos_gt_src = gt_dict['pos_gt_src']
headboxes_src = gt_dict['headboxes_src']
pos_pred_src = np.transpose(preds, [1, 2, 0])
head = np.where(dataset_joints == 'head')[1][0]
lsho = np.where(dataset_joints == 'lsho')[1][0]
lelb = np.where(dataset_joints == 'lelb')[1][0]
lwri = np.where(dataset_joints == 'lwri')[1][0]
lhip = np.where(dataset_joints == 'lhip')[1][0]
lkne = np.where(dataset_joints == 'lkne')[1][0]
lank = np.where(dataset_joints == 'lank')[1][0]
rsho = np.where(dataset_joints == 'rsho')[1][0]
relb = np.where(dataset_joints == 'relb')[1][0]
rwri = np.where(dataset_joints == 'rwri')[1][0]
rkne = np.where(dataset_joints == 'rkne')[1][0]
rank = np.where(dataset_joints == 'rank')[1][0]
rhip = np.where(dataset_joints == 'rhip')[1][0]
jnt_visible = 1 - jnt_missing
uv_error = pos_pred_src - pos_gt_src
uv_err = np.linalg.norm(uv_error, axis=1)
headsizes = headboxes_src[1, :, :] - headboxes_src[0, :, :]
headsizes = np.linalg.norm(headsizes, axis=0)
headsizes *= SC_BIAS
scale = np.multiply(headsizes, np.ones((len(uv_err), 1)))
scaled_uv_err = np.divide(uv_err, scale)
scaled_uv_err = np.multiply(scaled_uv_err, jnt_visible)
jnt_count = np.sum(jnt_visible, axis=1)
less_than_threshold = np.multiply((scaled_uv_err <= threshold),
jnt_visible)
PCKh = np.divide(100.*np.sum(less_than_threshold, axis=1), jnt_count)
# save
rng = np.arange(0, 0.5+0.01, 0.01)
pckAll = np.zeros((len(rng), 16))
for r in range(len(rng)):
threshold = rng[r]
less_than_threshold = np.multiply(scaled_uv_err <= threshold,
jnt_visible)
pckAll[r, :] = np.divide(100.*np.sum(less_than_threshold, axis=1),
jnt_count)
PCKh = np.ma.array(PCKh, mask=False)
PCKh.mask[6:8] = True
jnt_count = np.ma.array(jnt_count, mask=False)
jnt_count.mask[6:8] = True
jnt_ratio = jnt_count / np.sum(jnt_count).astype(np.float64)
name_value = [
('Head', PCKh[head]),
('Shoulder', 0.5 * (PCKh[lsho] + PCKh[rsho])),
('Elbow', 0.5 * (PCKh[lelb] + PCKh[relb])),
('Wrist', 0.5 * (PCKh[lwri] + PCKh[rwri])),
('Hip', 0.5 * (PCKh[lhip] + PCKh[rhip])),
('Knee', 0.5 * (PCKh[lkne] + PCKh[rkne])),
('Ankle', 0.5 * (PCKh[lank] + PCKh[rank])),
('Mean', np.sum(PCKh * jnt_ratio)),
('Mean@0.1', np.sum(pckAll[11, :] * jnt_ratio))
]
name_value = OrderedDict(name_value)
return name_value, name_value['Mean']
# -
name_values, perf_indicator = my_evaluate(cfg, all_preds, output_dir, all_boxes, image_path, filenames, imgnums)
print(name_values, perf_indicator)
## get GT info
gt_file = os.path.join(cfg.DATASET.ROOT, 'annot', 'gt_{}.mat'.format(cfg.DATASET.TEST_SET))
gt_dict = loadmat(gt_file)
dataset_joints = gt_dict['dataset_joints']
jnt_missing = gt_dict['jnt_missing']
pos_gt_src = gt_dict['pos_gt_src']
print(pos_gt_src.shape)
headboxes_src = gt_dict['headboxes_src']
print(dataset_joints)
print(np.where(dataset_joints == 'head'))
# dataset_joints[0][9][1]
pos_pred_src = np.transpose(all_preds, [1, 2, 0])
pos_pred_src.shape
# +
import matplotlib.pyplot as plt
# %matplotlib inline
import cv2
# +
idx_sample = 1
# read and plot image
im = cv2.imread(image_path[idx_sample])
plt.figure(figsize=(15,10))
plt.subplot(211)
plt.imshow(im[:,:,::-1])
# plot estimates joints
plt.scatter(pos_pred_src[:,0,idx_sample], pos_pred_src[:,1,idx_sample])
plt.scatter(pos_gt_src[:,0,idx_sample], pos_gt_src[:,1,idx_sample])
plt.legend(['estimate', 'true'])
# # plot head box
# hbox = headboxes_src[:,:,idx_sample]
# plot estiamte vs. true
plt.subplot(223)
plt.scatter(pos_gt_src[:,0,idx_sample], pos_pred_src[:,0,idx_sample])
plt.xlabel('true x')
plt.ylabel('estimated x')
plt.axis('equal')
plt.subplot(224)
plt.scatter(pos_gt_src[:,1,idx_sample], pos_pred_src[:,1,idx_sample])
plt.xlabel('true y')
plt.ylabel('estimated y')
plt.axis('equal')
plt.savefig('result_{}.png'.format(idx_sample))
# -
# ## predict for one image
# +
# preapre a scaled image
from utils.transforms import get_affine_transform
#center = np.array([im.shape[0]/2, im.shape[1]/2])
center = np.array([320, 270])
scale = 1.8
rot = 0
affine = get_affine_transform(center, scale, rot, [256,256])
input = cv2.warpAffine(
im,
affine,
(256, 256),
flags=cv2.INTER_LINEAR)
print(input.shape)
plt.imshow(input)
# +
trans = transforms.Compose([
transforms.ToTensor(),
normalize,
])
im_tensor = trans(input)
print(im_tensor.shape)
with torch.no_grad():
output = model(im_tensor.reshape([1, 3, 256, 256]))
print(output.shape)
# +
from core.inference import get_final_preds
pred, maxval = get_final_preds(cfg, output.clone().cpu().numpy(), [center], [scale])
pred.shape
# -
plt.imshow(im[:,:,::-1])
plt.scatter(pred[0,:,0], pred[0,:,1])
| test_mpii.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Suppress warnings
import warnings
warnings.filterwarnings('ignore')
import gc
import math
import random
import matplotlib
# matplotlib and seaborn for plotting
import matplotlib.pyplot as plt
import seaborn as sns
import matplotlib.patches as patches
from plotly import tools, subplots
import plotly.offline as py
py.init_notebook_mode(connected=True)
import plotly.graph_objs as go
import plotly.express as px
pd.set_option('max_columns', 100)
pd.set_option('max_rows', 30)
from datetime import datetime
py.init_notebook_mode(connected=True)
from plotly.offline import init_notebook_mode, iplot
init_notebook_mode(connected=True)
import plotly.graph_objs as go
path='../input/ashrae-energy-prediction/'
path_new = '../input/lag-transform/'
path_new = '../input/transformer/'
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import StratifiedKFold, KFold
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
from tqdm import tqdm_notebook
from math import log
import lightgbm as lgb
import datetime
from sklearn.preprocessing import LabelEncoder, MinMaxScaler
pd.options.mode.chained_assignment = None
from pandas.tseries.holiday import USFederalHolidayCalendar as calendar
import holidays
import os
from scipy.sparse import csr_matrix,coo_matrix, hstack
from sklearn.linear_model import Ridge
import seaborn as sns
from sklearn.neighbors import KNeighborsRegressor
for dirname, _, filenames in os.walk(path):
for filename in filenames:
print(os.path.join(dirname, filename))
def ls():
print([v for v in globals().keys() if not v.startswith('_')])
def seed_everything(seed=0):
random.seed(seed)
np.random.seed(seed)
def free_space(del_list):
for name in del_list:
if not name.startswith('_'):
del globals()[name]
gc.collect()
def chunk_predictor(X,model,n=100,verbose=True):
predict_fold = KFold(n)
result = np.zeros(X.shape[0])
if verbose:
for trn_idx, val_idx in tqdm_notebook(predict_fold.split(X)):
result[val_idx] = model.predict(X.iloc[val_idx,:])
else:
for trn_idx, val_idx in predict_fold.split(X):
result[val_idx] = model.predict(X.iloc[val_idx,:])
return(result)
# def one_hot_encoder(train,col,tr_index):
# scaler = MinMaxScaler()
# temp = csr_matrix(scaler.fit_transform(pd.get_dummies(train,prefix_sep='_',columns=col,drop_first=True)))
# return(temp[:tr_index,:],temp[tr_index:,:])
def one_hot_encoder(train,col,tr_index):
temp = (pd.get_dummies(train,prefix_sep='_',columns=col,drop_first=True)).values #csr_matrix
return(temp[:tr_index,:],temp[tr_index:,:])
# +
path_load = '../input/train-transformer/'
path_dataset ='../input/rescale-knn/'
train = pd.read_hdf(f'{path_dataset}dataset_rescale.h5',key='rescale_train')
test = pd.read_hdf(f'{path_dataset}dataset_rescale.h5',key='rescale_test')
# train = pd.read_hdf(f'{path_load}dataset.h5',key='train')
# test = pd.read_hdf(f'{path_load}dataset.h5',key='test')
# column = pd.read_hdf(f'{path_load}dataset.h5',key='column')['column'].tolist()
# timestamp = pd.read_hdf(f'{path_load}dataset.h5',key='timestamp')
# train['month']=timestamp.dt.month.astype(int)
# category_column = pd.read_hdf(f'{path_load}dataset.h5',key='category_column')['cat_column'].tolist()
target = pd.read_hdf(f'{path_load}dataset.h5',key='target')
support = pd.read_hdf(f'{path_load}dataset.h5',key='train')[['building_id','meter']]
valid_build = support['building_id'].copy()
valid_meter = support['meter'].copy()
del support
gc.collect()
# -
SEED = 42
seed_everything(SEED)
gc.collect()
# +
# valid_build = train['building_id'].copy()
# valid_meter = train['meter'].copy()
# gc.collect()
# +
# replace = train.groupby('building_meter')['pm2k'].mean()
# train.loc[train.pm2k.isnull(),'pm2k'] = train['building_meter'].map(replace)[train.pm2k.isnull()]
# test.loc[test.pm2k.isnull(),'pm2k'] = test['building_meter'].map(replace)[test.pm2k.isnull()]
# strat_train = train['building_meter']
# strat_test = test['building_meter']
# train = train[['day','hour','weekend','air_temperature','dew_temperature',
# 'heat_index','feel','air_temperature_mean_lag24','dew_temperature_mean_lag24','feel_mean_lag24','heating_required','m2k','pm2k']]#'month',
# test = test[['day','hour','weekend','air_temperature','dew_temperature',
# 'heat_index','feel','air_temperature_mean_lag24','dew_temperature_mean_lag24','feel_mean_lag24','heating_required','m2k','pm2k']]#'month',
# +
strat_train = train.pop('building_meter')
strat_test = test.pop('building_meter')
to_dummy = ['month','day','hour','weekend']
# -
to_dummy = ['hour','weekend'] #'month','day',
train.drop(['month','day'],axis=1,inplace=True)
test.drop(['day','month'],axis=1,inplace=True)
# +
n_fold = 2
# models = {}
cv_scores = {"build_meter": [], "cv_score": []}
seed = 0
kf = KFold(n_splits=n_fold, random_state=seed,shuffle=True)
pred_test = np.zeros(test.shape[0])
pred_x = np.zeros(train.shape[0])
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# %%time
num_nearest = 10
for sel in tqdm_notebook(np.sort(strat_train.unique())):
temp = train[strat_train==sel]
train_row = temp.shape[0]
temp = temp.append(test[strat_test==sel],ignore_index =True)
temp_train, temp_test = one_hot_encoder(temp,to_dummy,train_row)
y = target.values[strat_train==sel]
score = 0
for fold, (train_index, valid_index) in enumerate(kf.split(temp_train, y)):
x_train, x_valid = temp_train[train_index,:], temp_train[valid_index,:]
y_train, y_valid = y[train_index], y[valid_index]
model = KNeighborsRegressor(n_neighbors = num_nearest,n_jobs = -1,algorithm ='kd_tree')
model.fit(x_train, y_train)
pred_test[strat_test==sel] += model.predict(temp_test)/n_fold
pred_x[np.where(strat_train==sel)[0][valid_index]] += model.predict(x_valid)
score += np.sqrt(mean_squared_error(y_valid, pred_x[np.where(strat_train==sel)[0][valid_index]]))/n_fold
# print(f'cv score: {score}')
cv_scores["build_meter"].append(sel)
cv_scores["cv_score"].append(score)
del temp, temp_train, y,x_train, x_valid, y_train, y_valid, model, score
gc.collect()
# -
fig = plt.figure(figsize=(12,8))
sns.distplot(cv_scores['cv_score'],label ='cv-error')
plt.legend()
# +
result = pd.DataFrame({'Building_ID':valid_build,'meter':valid_meter,'target_real':target,'target_pred':pred_x}).dropna()
result.reset_index(drop=True,inplace=True)
gc.collect()
free_space(['valid_build','valid_meter'])
gc.collect()
# -
gc.collect()
sample_submission = pd.read_csv(f'{path}sample_submission.csv')
sample_submission['meter_reading'] = np.expm1(pred_test)
# leak_df = pd.read_csv('../input/ashrae-leak-data-station/sample_leak.csv',index_col=0).dropna()
# sample_submission.loc[leak_df.index, 'meter_reading'] = leak_df['meter_reading']
# sample_submission.reset_index(inplace=True,drop=True)
sample_submission.loc[sample_submission['meter_reading']<0,'meter_reading'] = 0
sample_submission.to_csv('submission_linear.csv',index=False)
sample_submission.iloc[:50,:]
fig = plt.figure(figsize=(12,8))
sns.distplot(result.target_real,label='Real')
sns.distplot(result.target_pred,label='Forecast')
plt.legend()
fig = plt.figure(figsize=(12,8))
sns.distplot((result.target_real-result.target_pred),label ='Error')
plt.legend()
sns.scatterplot(x=result.target_real, y=result.target_pred)
| ASHRAE - Great Energy Predictor III/Model/knn-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Recurrent Group-Testing as an alternative to Social Distancing for containing the COVID-19 pandemic
# If social distancing orders are to be relaxed in the near future, it is imperative to have an alternative strategy for mitigating the spread of COVID-19.
#
# This notebook studies the extent to which recurrent group-testing can be used as an alternative to social distancing.
#
# The recurrent group-testing procedure works as follows:
# * We consider a city with $N$ households. Each household contains $M$ individuals. $M$ and $N$ are configurable parameters. There is no travel in or out of the city for the length of the simulation.
# * A test day occurs once every $D$ days. $D$ is a configurable parameter.
# * On each test day, every citizen participates in a group test with probability $P$. $P$ is a configurable parameter.
# * The (randomly selected) group-test participants are allocated into buckets of max-size $S$. $S$ is a configurable parameter. Household members are always allocated to the same group.
# * A single test is used to check for the presence of COVID-19 in each group.
# * A parameter $FNR$ models the false-negative rate for an individual. $FNR$ is configurable.
# * The probability that the test produces a false negative for a group with $S$ individuals is assumed to be
# $$ 1 - FNR^S. $$
# * When a group test does not detect the presence of COVID-19, the participants have social distancing restrictions lifted. The rate of spread of the disease across the population of non-social-distancing citizens is a configurable parameter.
# * When a group test does return positive then all of the citizens must **remain in complete isolation** until the next test day period. This is a much more severe assumption that current social distancing guidelines. We say these participants are quarantined. When an individual is quarantined we assume that all individuals in the household become quarantined. If a quarantined individual is infected, he or she can only spread the disease to members of his or her household. This spread occurs with a configurable probability $SAR$.
# * **Note:** The existence of this possibility would create a strong **disincentive** for citizens to participate in the test.
# * The disease ends in an individual after a configurable number of days.
# * The simulation assumes that every individual begins in the strict quarantine state
#
# Summary of parameters:
# * $N$: Number of households
# * $M$: Household size
# * $D$: Test day frequency
# * $P$: Probability each individual participates in a group test
# * $S$: Size of group for each group test
# * $FNR$: False-negative rate for an individual. If $K$ out of the $S$ individuals in the group are infected, then the group test produces a false-negative with probability $$1-FNR^K.$$
# * $SAR$: Secondary attack rate, i.e. probability disease spreads within a household on each day
# * $\alpha$: The rate-of-spread of the disease for the population of non-quarantined individuals
# * $d_0$: The length of the disease
# * $p$: fatality percent
#
#
# For a particular instantiation of each of the above parameters, the simulation looks at the following metrics.
# As a function of time:
# * what is the total fraction of the population that has been infected by the pandemic?
# * what is the total fraction of the population that has died from the pandemic?
# * what is the fraction of population that is not in quarantine?
# * what is the resource usage (in terms of total number of tests) required to supply the entire test
#
# +
import matplotlib.pyplot as plt
# %matplotlib inline
from population import Population
from group_testing import SymptomaticIndividualTest, ThreeStageHierarchicalTest
from simulation import Simulation
def initiate_simulation():
if not test_only_symptomatic_individuals:
test = ThreeStageHierarchicalTest(large_group_size,
small_group_size,
group_test_participation_rate,
outer_false_negative_rate,
inner_false_negative_rate,
)
else:
test = SymptomaticIndividualTest(inner_false_negative_rate)
population = Population(n_households,
household_size,
initial_prevalence,
disease_length,
time_until_symptomatic,
non_quarantine_alpha,
daily_secondary_attack_rate,
fatality_pct,
daily_outside_infection_pct,
outside_symptomatic_prob,
initial_quarantine)
simulation = Simulation(population, test, test_day_frequency, test_latency, halt_operations_if_case_detected)
return simulation
def summarize(simulation):
print("Total number of tests performed over {} days: {}".format(simulation.current_day,
simulation.cumulative_tests_to_date))
days = range(simulation.current_day)
cumulative_infected_pct = [simulation.recorded_data[day]['cumulative_infected_fraction'] for day in days]
within_population_infected_pct = [simulation.recorded_data[day]['cumulative_infected_within_population'] for day in days]
quarantine_pct = [simulation.recorded_data[day]['in_quarantine_fraction'] for day in days]
infected_pct = [simulation.recorded_data[day]['infected_fraction'] for day in days]
plt.figure(figsize=(10,6))
plt.ylim((-0.1,1.1))
plt.plot(days, cumulative_infected_pct, label="Cumulative Fraction of Population Infected")
plt.plot(days, quarantine_pct, label="Fraction of Population in Quarantine")
plt.plot(days, within_population_infected_pct, label="Cumulative Fraction of Population Infected due to Within-Community Spread")
plt.plot(days, infected_pct, label="Fraction of Population Infected")
plt.legend(loc='best')
plt.show()
cumulative_tests = [simulation.recorded_data[day]['cumulative_tests_to_date'] for day in days]
plt.figure(figsize=(10,6))
plt.plot(days, cumulative_tests, label='Cumulative Tests to Date')
plt.legend(loc='best')
plt.show()
if simulation.halt_operations_if_case_detected:
days_halted = [simulation.recorded_data[day]['cumulative_days_halted'] for day in days]
plt.figure(figsize=(10,6))
plt.plot(days, days_halted, label="Cumulative Days Halted due to Detected Cases")
plt.legend(loc='best')
plt.show()
def run(simulation, number_of_days):
for _ in range(number_of_days):
simulation.step()
# +
# Population parameters
n_households = 100
household_size = 1
daily_outside_infection_pct = 0.001
initial_prevalence = 0.05
disease_length = 14
R0 = 5.0
non_quarantine_alpha = R0 ** (1 / disease_length)
print("alpha = {:.2f}".format(non_quarantine_alpha))
daily_secondary_attack_rate = 0.5
fatality_pct = 0.02
initial_quarantine = False
# Group test parameters
large_group_size = 25
small_group_size = 5
group_test_participation_rate = 1
outer_false_negative_rate = 0.1
inner_false_negative_rate = 0.1
test_day_frequency = 5
test_latency = 2
halt_operations_if_case_detected = False
test_only_symptomatic_individuals = True
outside_symptomatic_prob = 0.0
time_until_symptomatic = 7
# -
from statistics import mean
def collect_multiple_trajectories(num_trajectories=1000, number_of_days=90):
trajectories = []
for _ in range(num_trajectories):
simulation = initiate_simulation()
run(simulation, number_of_days=number_of_days)
trajectory_data = {}
days = range(number_of_days)
trajectory_data['mean_quarantine_pct'] = mean([simulation.recorded_data[day]['in_quarantine_fraction']
for day in days])
final_day = days[-1]
trajectory_data['cumulative_infected_fraction'] = simulation.recorded_data[final_day]['cumulative_infected_fraction']
trajectory_data['cumulative_within_warehouse_fraction'] = simulation.recorded_data[final_day]['cumulative_infected_within_population']
trajectory_data['max_infected_unquarantined_fraction'] = \
max([simulation.recorded_data[day]['infected_unquarantined_fraction'] for day in days])
trajectory_data['cumulative_tests'] = simulation.recorded_data[final_day]['cumulative_tests_to_date']
trajectories.append(trajectory_data)
return trajectories
# +
trajectory_collection_baselines = []
trajectory_collection_grptests = []
magnitudes = [1e-06, 1e-05, 1e-04, 1e-03]
multiples = list(range(1,10,2))
external_infection_risks = []
for magnitude in magnitudes:
for multiple in multiples:
risk = magnitude * multiple
print("on risk = {}".format(risk))
external_infection_risks.append(risk)
daily_outside_infection_pct = risk
test_only_symptomatic_individuals = True
test_latency = 0
test_day_frequency = 1
trajectory_collection_baselines.append(collect_multiple_trajectories())
test_only_symptomatic_individuals = False
test_latency = 1
test_day_frequency = 5
trajectory_collection_grptests.append(collect_multiple_trajectories())
# -
import matplotlib
matplotlib.rcParams.update({'font.size': 14})
# +
from statistics import mean
quarantine_mean_baseline = []
quarantine_mean_grptest = []
for trajectories_baseline, trajectories_grptest in zip(trajectory_collection_baselines, trajectory_collection_grptests):
quarantine_mean_baseline.append(100 * mean([trajectory['mean_quarantine_pct'] for trajectory in trajectories_baseline]))
quarantine_mean_grptest.append(100 * mean([trajectory['mean_quarantine_pct'] for trajectory in trajectories_grptest]))
plt.figure(figsize=(10,6))
plt.plot([100 * r for r in external_infection_risks], quarantine_mean_baseline, label="Baseline", marker='o')
plt.plot([100 * r for r in external_infection_risks], quarantine_mean_grptest, label="Group Test", marker='o')
plt.legend(loc='best')
plt.xscale('log')
#plt.yscale('log')
plt.title('mean quarantine pct vs. outside infection pct.')
plt.show()
# +
from statistics import mean
unquarantined_infection_baseline = []
unquarantined_infection_grptest = []
for trajectories_baseline, trajectories_grptest in zip(trajectory_collection_baselines, trajectory_collection_grptests):
unquarantined_infection_baseline.append(100 * mean([trajectory['max_infected_unquarantined_fraction'] for trajectory in trajectories_baseline]))
unquarantined_infection_grptest.append(100 * mean([trajectory['max_infected_unquarantined_fraction'] for trajectory in trajectories_grptest]))
plt.figure(figsize=(10,6))
plt.plot([100 * r for r in external_infection_risks], unquarantined_infection_baseline, label="Baseline", marker='o')
plt.plot([100 * r for r in external_infection_risks], unquarantined_infection_grptest, label="Group Test", marker='o')
plt.legend(loc='best')
plt.xscale('log')
#plt.yscale('log')
plt.title('max infected unquarantined vs. outside infection pct.')
plt.show()
# -
list(range(1,10,2))
1e-03
range(1e-05, 0.001)
xs = []
ys_baseline = []
ys_grptest = []
for x in trajectory_collection_baseline.keys():
xs.append(0.0001 * x)
ys_baseline.append(mean([trajectory['mean_quarantine_pct'] for trajectory in trajectory_collection_baseline[x]]))
ys_grptest.append(mean([trajectory['mean_quarantine_pct'] for trajectory in trajectory_collection_grptest[x]]))
# +
plt.figure(figsize=(10,6))
plt.plot(xs, ys_baseline, label="Baseline")
plt.plot(xs, ys_grptest, label="Group Test")
plt.legend(loc='best')
plt.title('avg % of workforce in quarantine vs. outside infection pct.')
plt.show()
# -
xs = []
ys_baseline = []
ys_grptest = []
for x in trajectory_collection_baseline.keys():
xs.append(0.0001 * x)
ys_baseline.append(mean([trajectory['cumulative_infected_fraction'] for trajectory in trajectory_collection_baseline[x]]))
ys_grptest.append(mean([trajectory['cumulative_infected_fraction'] for trajectory in trajectory_collection_grptest[x]]))
# +
plt.figure(figsize=(10,6))
plt.plot(xs, ys_baseline, label="Baseline")
plt.plot(xs, ys_grptest, label="Group Test")
plt.legend(loc='best')
plt.title('Cumulative Infected Fraction vs. outside infection pct')
plt.show()
# -
trajectories = collect_multiple_trajectories()
test_only_symptomatic_individuals = False
trajectories_grptest = collect_multiple_trajectories()
plt.hist([trajectory['max_infected_unquarantined_fraction'] for trajectory in trajectories], alpha=0.7, label="Baseline")
plt.hist([trajectory['max_infected_unquarantined_fraction'] for trajectory in trajectories_grptest], alpha=0.7, label="Group test")
plt.legend(loc='best')
plt.title('Max infected unquarantined fraction')
plt.show()
plt.hist([trajectory['cumulative_infected_fraction'] for trajectory in trajectories], alpha=0.7, label="Baseline")
plt.hist([trajectory['cumulative_infected_fraction'] for trajectory in trajectories_grptest], alpha=0.7, label="Group test")
plt.legend(loc='best')
plt.title('90-day cumulative infection distribution')
plt.show()
plt.hist([trajectory['cumulative_tests'] for trajectory in trajectories], alpha=0.7, label="Baseline")
plt.hist([trajectory['cumulative_tests'] for trajectory in trajectories_grptest], alpha=0.7, label="Group test")
plt.legend(loc='best')
plt.title('90-day cumulative tests')
plt.show()
plt.hist([trajectory['cumulative_within_warehouse_fraction'] for trajectory in trajectories], alpha=0.7, label="Baseline")
plt.hist([trajectory['cumulative_within_warehouse_fraction'] for trajectory in trajectories_grptest], alpha=0.7, label="Group test")
plt.legend(loc='best')
plt.title('90-day cumulative within-warehouse infection distribution')
plt.show()
trajectories[0]
plt.hist([trajectory['cumulative_infected_within_population'] for trajectory in trajectories])
plt.hist([trajectory['cumulative_days_halted'] for trajectory in trajectories])
simulation = initiate_simulation()
run(simulation, number_of_days=90)
summarize(simulation)
test_only_symptomatic_individuals = False
simulation = initiate_simulation()
run(simulation, number_of_days=60)
summarize(simulation)
j
| boqn/group_testing/notebooks/RecurrentGroupTesting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"is_executing": true}
import numpy as np
import sklearn.datasets
import sklearn.model_selection
import matplotlib.pyplot as plt
from scipy.sparse import linalg
import random
# load dataset
data = sklearn.datasets.load_svmlight_file('dataset/housing_scale', n_features=13)
# split the dataset into traning set and validation set(80% for training set , 20% for validation set)
X_train, X_val, Y_train, Y_val = sklearn.model_selection.train_test_split(data[0], data[1], test_size=0.2, random_state=222)
# loss function (using least square loss)
def Loss(y, y_):
loss = 0.5*((y-y_)**2)
return loss
# initialize parameter w
w = np.random.normal(size=13)
# y = wx
predict_init = X_train.dot(w)
loss_init = Loss(predict_init, Y_train)
print('initial mean loss is:{}'.format(loss_init.mean()))
w_k = linalg.inv(X_train.T.dot(X_train)).dot(X_train.T).dot(Y_train)
print(w_k)
# some parameters
EPOCHS = 500
LR = 0.0008 # learning rate
# initialize
loss_train = []
loss_val = []
L2_norm = []
# SGD(Stochastic Gradient Descent)
for epoch in range(EPOCHS):
for i in range(X_train.shape[0]):
# pick a sample randomly
randnumber = random.randint(0, X_train.shape[0]-1)
X = X_train[randnumber]
Y = Y_train[randnumber]
# gradient
G = X.T.dot(X.dot(w)-Y)
D = -G
w += LR*D
L2_norm.append(np.linalg.norm(w - w_k, ord=2))
loss_train.append(Loss(X_train.dot(w), Y_train).mean())
loss_val.append(Loss(X_val.dot(w), Y_val).mean())
'''
# GD
for epoch in range(EPOCHS):
G = X_train.T.dot(X_train.dot(w)-Y_train)
D = -G
w += LR*D
loss_train.append(Loss(X_train.dot(w), Y_train).mean())
loss_val.append(Loss(X_val.dot(w), Y_val).mean())
L2_norm.append(np.linalg.norm(w-w_k, ord=2))
'''
print('mean loss_train is:{}'.format(loss_train[-1]))
print('mean loss_val is:{}'.format(loss_val[-1]))
# + pycharm={"is_executing": true, "name": "#%%\n"}
# plot img1
plt.figure(figsize=[15, 6])
plt.title('L2 norm optimization')
plt.xlabel('epoch')
plt.ylabel('||W_k - W*||2')
plt.plot(L2_norm, color='red', linewidth=1, label='L2 norm')
plt.legend()
plt.savefig('optimize')
plt.show()
# + pycharm={"is_executing": true, "name": "#%%\n"}
# plot img2
plt.figure(figsize=[15, 4])
plt.title('Validation Set Loss')
plt.xlabel('epoch')
plt.ylabel('loss')
plt.plot(loss_val, color='red', linewidth=1, label='valid')
plt.plot(loss_train, color='blue', linewidth=1, label='train')
plt.legend()
plt.savefig('SGD_Validation-Set-Loss.png')
plt.show()
# + pycharm={"is_executing": true, "name": "#%%\n"}
| 2019ML_Lab/Lab1/SGD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import print_function
import torch
from torch import nn
from torch import optim
from torch.utils.data import DataLoader, random_split, WeightedRandomSampler, SubsetRandomSampler
from torchvision import models, transforms
import os
import numpy as np
import sys
import matplotlib.pyplot as plt
from functools import partial
from few_shot_learning.datasets import FashionProductImages, FashionProductImagesSmall
from few_shot_learning.utils_evaluation import ToSize
from few_shot_learning.train_transfer import transfer
from few_shot_learning.utils_evaluation import evaluate_transfer
from few_shot_learning.utils import accuracy
from config import DATA_PATH
# -
# # 1. Data Preprocessing and Statistics
# Preprocessing and loading of image samples and label information is accomplished through the classes `few_shot_learning.dataset.FashionProductImages` and `few_shot_learning.datasets.FashionProductImagesSmall`. Important arguments are `split` and `classes`. For the purposes of transfer learning, the split should be one of `split='train'` or `split='test'` and the classes one of `classes='top'` or `classes='bottom'` selecting the top 20 classs with the most samples or the remaining 123 classes.
#
# The following code snippets illustrate the functionality:
all_data = FashionProductImagesSmall(DATA_PATH, split='all', classes=None)
# The complete dataset description looks like this:
all_data.df
# +
datasets = dict()
for classes in ['top', 'bottom', None]:
datasets[classes] = dict()
for split in ['train', 'test', 'all']:
dataset = FashionProductImages(DATA_PATH, split=split, classes=classes, transform=ToSize())
size = len(dataset)
loader = DataLoader(dataset, batch_size=size, shuffle=False, num_workers=8)
# use loader to efficiently load all images using multiple workers
#for widths_heights, labels in loader:
# break
#widths, heights = widths_heights
#widths = widths.numpy()
#heights = heights.numpy()
#labels = labels.numpy()
datasets[classes][split] = dict(data=dataset,
size=size,
loader=loader)#,
#widths=widths,
#heights=heights,
#labels=labels)
# -
# ## 1.1. Distribution of image sizes and aspect ratios
# We first have a look at the image format and sizes. It turns out they're half-way regular, mostly 1800-by-1200 pixels or 2400-by-1800 pixels, mostly RGB. Aspect ratio is almost always $4/3$. The resolution is quite high, I chose to convert all images of the full dataset to **400-by-300 pixels** (not shown here).
# +
from matplotlib.colors import LogNorm
unique_heights = np.unique(datasets['top']['train']['heights'])
unique_widths = np.unique(datasets['top']['train']['widths'])
print(unique_heights)
print(unique_widths)
plt.figure(figsize=(12,9))
plt.hist2d(
datasets['top']['train']['heights'],
datasets['top']['train']['widths'],
bins=[unique_heights, unique_widths],
norm=LogNorm(),
)
plt.xticks(unique_heights)
plt.yticks(unique_widths)
plt.colorbar()
plt.show()
# +
aspect_ratios = datasets['top']['train']['heights']/datasets['top']['train']['widths']
unique_aspect_ratios = np.unique(aspect_ratios)
print(unique_aspect_ratios)
print((aspect_ratios == 4/3).sum())
plt.figure(figsize=(8,6))
plt.hist(aspect_ratios)
plt.show()
# -
# ## 1.2. Distribution of Classes in Training and Test Set
# Importantly, the distribution of classes in the training and test set is quite different. For the chosen master split, all training samples are from **even** years and all test samples are from **odd** years.
#
# The histogram below illustrates the differences between training and test set. The most striking difference is the class `'Perfume and Body Mist'` which is **absent from the training set**. Additionally, although this is less concerning, some classes are almost absent from the test set (classes `'Watches'` and `'Sunglasses'`)
#
# The situation is worse for the remaining 123 classes. There we observe a substantial mismatch between the class distributions of training and test set, as shown below.
def get_mapped_class_dist(dataset, classes, normed=False):
class_dist = np.bincount(dataset.target_indices, minlength=len(classes))
if normed:
class_dist = class_dist / class_dist.sum()
class_indices = dataset.target_codec.transform(classes)
return class_dist[class_indices]
# +
# top 20 classes
top20 = FashionProductImages.top20_classes
plt.figure(figsize=(12,9))
plt.bar(top20, get_mapped_class_dist(datasets['top']['train']['data'], top20), label='training set', width=0.4, align='edge')
plt.bar(top20, get_mapped_class_dist(datasets['top']['test']['data'], top20), label='test set', width=-0.4, align='edge')
plt.xticks(rotation='vertical')
plt.legend()
plt.show()
# +
bottom123 = datasets['bottom']['train']['data'].classes
bottom_train_dist = get_mapped_class_dist(datasets['bottom']['train']['data'], bottom123)
bottom_test_dist =get_mapped_class_dist(datasets['bottom']['test']['data'], bottom123)
plt.figure(figsize=(10,50))
plt.barh(bottom123, bottom_train_dist, label='training set', height=0.4, align='edge')
plt.barh(bottom123, bottom_test_dist, label='test set', height=-0.4, align='edge')
# plt.xticks(rotation='vertical')
plt.legend()
plt.show()
# -
# Classes **not present in the training set** while present in the test set and their relative proportion in the test set:
# +
testonly_classes = bottom123[(bottom_test_dist > 0) & (bottom_train_dist == 0)]
print(testonly_classes)
# -
# proportion of test samples affected
print(bottom_test_dist[(bottom_test_dist > 0) & (bottom_train_dist == 0)].sum() / bottom_test_dist.sum())
# Furthermore, there are classes present in the training set which are **absent from the test set**.
# +
# classes not present in the test set while present in the training set
trainonly_classes = bottom123[(bottom_test_dist == 0) & (bottom_train_dist > 0)]
print(trainonly_classes)
# -
# Finally, there are classes which are missing from both training and test set. These are classes that nominally have samples in `styles.csv` but whose images are missing from folder `images/`. They won't affect neither training nor testing in any way.
# +
# classes neither in the training set nor test set
missing_classes = bottom123[(bottom_test_dist == 0) & (bottom_train_dist == 0)]
print(missing_classes)
# -
# # 2. Tests
#
# Run `pytest` in the root directory to run tests.
# # 3. Training
# A training run can also be started via e.g.
#
# ```
# python -m experiments.transfer_learning -a resnet50 -p 50 --distributed --date --epochs 100
# ```
#
# For a list of arguments, refer to the help message:
run "~/few-shot-learning/experiments/transfer_experiment.py" -h
# I focused on running experiments with ResNet50. These are the important decisions for the training procedure:
#
# - *n_epochs*: 100
# - *learning rate schedule*:
# - Top20 initial finetuning: `learning_rate=1e-3`, decayed by 0.7 every 5 epochs (ad-hoc choice)
# - Transfer learning: `learning_rate=1e-3`, decayed by 0.7 every 5 epochs (ad-hoc choice)
# - *optimizer*: `torch.optim.Adam`
# - *batch_size*: 64
# - *model architecture*: **ResNet50** (trained on 2 GPUs with `torch.nn.DataParallel`)
# - *validation*:
# - Top20 initial finetuning: selected the model with the best validation accuracy on a random 10% of training samples. Training and validation sets stratified across classes. Continued with this model for transfer learning.
# - Transfer learning: selected the model with the best validation accuracy on a random 10% of training samples again.
# Training and validation sets not stratified.
# - *data augmentation*: `torchvision.transforms.RandomResizedCrop`, `torchvision.transforms.ColorJitter`, `torchvision.transforms.RandomHorizontalFlip`.
# - *balanced training*: When sampling from the training set, I sampled uniformly from all classes (both finetuning and transfer). This means I undersampled classes with many samples and oversampled classes with very few samples. I could have alternatively implemented a weighted loss function or a focal loss to give higher weight to classes with fewer samples. Importantly, there is no a priori reason to believe that certain misclassifications are worse than others for our dataset (e.g. something like misclassifying suits is worse than misclassifying jeans.). For this reason, balanced training was employed only to combat overfitting on dominant classes and to thus improve overall test performance. Consequently, no balanced sampling was employed for the validation set (to be closer to the test set).
# Training can also be run with:
transfer(
data_dir=DATA_PATH,
architecture='resnet50',
num_workers=8,
epochs=100,
batch_size=64,
learning_rate=1e-3,
optimizer_cls=torch.optim.Adam,
print_freq=50,
seed=None,
distributed=True,
date_prefix=True,
model_dir=os.path.expanduser("~/few-shot-learning/models"),
log_dir=os.path.expanduser("~/few-shot-learning/logs")
)
# # 4. Evaluation
# +
LOG_DIR = os.path.expanduser("~/few-shot-learning/logs/")
MODEL_DIR = os.path.expanduser("~/few-shot-learning/models/")
folder = "19_10_08_1917"
# -
finetuning_results = torch.load(os.path.join(LOG_DIR, folder, "finetuning_training_log.json"))
transfer_results = torch.load(os.path.join(LOG_DIR, folder, "transfer_training_log.json"))
finetuning_best_model = torch.load(
os.path.join(MODEL_DIR, folder, "finetuning_model_best_unwrapped.pth.tar"),
map_location=torch.device('cuda', 1)
)
transfer_best_model = torch.load(
os.path.join(MODEL_DIR, folder, "transfer_model_best_unwrapped.pth.tar"),
map_location=torch.device('cuda', 1)
)
# ## 4.1 Monitoring Training
#
# The learning curves below show training and validation losses and accuracies over the course of training. In general, training seems to have been successful with the ad-hoc choice of initial learning rates and learning rate schedules.
#
# The initial finetuning learning curve for the validation loss hints at overfitting but the validation accuracy is best later in training. The best model is marked by a horizontal line. Top5 validation accuracy rises to almost 100% for the best model.
#
# The transfer learning seems to have been successful as well. The best top1 validation accuracy is lower than for the initial finetuning on the top-20 classes.
# +
plt.figure(figsize=(16,12))
plt.subplot(221)
plt.plot(finetuning_results["train_loss"], label="training loss")
plt.plot(finetuning_results["val_loss"], label="validation loss")
plt.title("Finetuning for top20 classes - Loss", fontsize=15)
plt.xlabel("Epoch")
plt.ylabel("Loss")
plt.legend()
plt.subplot(222)
plt.plot(finetuning_results["train_acc1"], label="top1 accuracy (training)", color='#1f77b4')
plt.plot(finetuning_results["train_acc5"], label="top5 accuracy (training)", color='#1f77b4', linestyle="--")
plt.plot(finetuning_results["val_acc1"], label="top1 accuracy (validation)", color='#ff7f0e')
plt.plot(finetuning_results["val_acc5"], label="top5 accuracy (validation)", color='#ff7f0e', linestyle="--")
plt.vlines(finetuning_best_model["epoch"]-1, 86, 100, label="best model")
plt.title("Finetuning for top20 classes - Accuracy", fontsize=15)
plt.xlabel("Epoch")
plt.ylabel("Classification Accuracy")
plt.legend()
plt.subplot(223)
plt.plot(transfer_results["train_loss"], label="training loss", color='#2ca02c')
plt.plot(transfer_results["val_loss"], label="validation loss", color='#d62728')
plt.title("Transfer for remaining 123 classes - Loss", fontsize=15)
plt.xlabel("Epoch")
plt.ylabel("Loss")
plt.legend()
plt.subplot(224)
plt.plot(transfer_results["train_acc1"], label="top1 accuracy (training)", color='#2ca02c')
plt.plot(transfer_results["train_acc5"], label="top5 accuracy (training)", color='#2ca02c', linestyle="--")
plt.plot(transfer_results["val_acc1"], label="top1 accuracy (validation)", color='#d62728')
plt.plot(transfer_results["val_acc5"], label="top5 accuracy (validation)", color='#d62728', linestyle="--")
plt.vlines(transfer_best_model["epoch"]-1, 50, 100, label="best model")
plt.title("Transfer for remaining 123 classes - Accuracy", fontsize=15)
plt.xlabel("Epoch")
plt.ylabel("Classification Accuracy")
plt.legend()
plt.show()
# -
# ## 4.2. Performance on Test Set
# For the initial finetuning on the top-20 classes the test set performance is given below. Importantly, test performance is substantially impacted by the class that was missing from the training set:
#
# | Finetuning (Top-20) - Test Set Accuracy | Top1 | Top5 |
# |-------------------------------------------------------|------|-----|
# | average across classes | 88.3 | 95.4|
# | average across classes (w/o 'Perfume and Body Mist') | 92.1 | 99.4|
#
# For transfer learning on the remaining classes the test set performance is. Here the problem of test-only classes is even more pronounced.
#
# | Transfer (Remaining classes) Test Set Accuracy | Top1 | Top5 |
# |-------------------------------------------------------|------|-----|
# | average across classes | 46.8 | 60.5|
# | average across classes (w/o test-only classes) | ? | ? |
#
#
# Detailed results for classwise accuracies are shown in the histograms below.
if torch.cuda.is_available():
device = torch.device('cuda', 1)
else:
device = torch.device('cpu')
acc1_avg, acc5_avg, acc1_classwise, acc5_classwise, outputs, targets = evaluate_transfer(
finetuning_best_model['state_dict'],
device=device,
architecture='resnet50',
classes='top',
small_dataset=False,
num_workers=4,
batch_size=128
)
print(acc1_avg, acc5_avg)
# +
perfume_class = datasets['top']['test']['data'].target_codec.transform(["Perfume and Body Mist"])
acc1_without_perfume, acc5_without_perfume = accuracy(
outputs[targets != perfume_class[0]], targets[targets != perfume_class[0]], topk=(1, 5))
# -
print(acc1_without_perfume.item(), acc5_without_perfume.item())
plt.figure(figsize=(12,9))
plt.bar(top20 + ["All classes"], list(acc1_classwise[datasets['top']['test']['data'].target_codec.transform(top20)]) + [0.0], label='Top 1 Accuracy', width=-0.4, align='edge')
plt.bar(top20 + ["All classes"], list(acc5_classwise[datasets['top']['test']['data'].target_codec.transform(top20)]) + [0.0], label='Top 5 Accuracy', width=0.4, align='edge')
plt.bar(top20 + ["All classes"], list(np.zeros_like(acc1_classwise)) + [acc1_avg], label='Top 1 Accuracy (All Classes)', width=-0.5, align='edge')
plt.bar(top20 + ["All classes"], list(np.zeros_like(acc5_classwise)) + [acc5_avg], label='Top 5 Accuracy (All Classes)', width=0.5, align='edge')
plt.xticks(rotation='vertical')
plt.legend(loc="lower right")
plt.show()
acc1_avg_transfer, acc5_avg_transfer, acc1_classwise_transfer, acc5_classwise_transfer, outputs_transfer, targets_transfer = evaluate_transfer(
transfer_best_model['state_dict'],
device=device,
architecture='resnet50',
classes='bottom',
small_dataset=False,
num_workers=4,
batch_size=128
)
print(acc1_avg_transfer, acc5_avg_transfer)
# TODO accuracy without test-only classes
testonly = torch.from_numpy(datasets['bottom']['test']['data'].target_codec.transform(testonly_classes)).to(device)
# +
bottom123 = list(datasets['bottom']['test']['data'].classes)
reordered_acc1_classwise = list(acc1_classwise_transfer[datasets['bottom']['test']['data'].target_codec.transform(bottom123)])
reordered_acc5_classwise = list(acc5_classwise_transfer[datasets['bottom']['test']['data'].target_codec.transform(bottom123)])
plt.figure(figsize=(10,50))
plt.barh(bottom123 + ["All classes"], reordered_acc1_classwise + [0.0], label='Top 1 Accuracy', height=0.4, align='edge')
plt.barh(bottom123 + ["All classes"], reordered_acc5_classwise + [0.0], label='Top 5 Accuracy', height=-0.4, align='edge')
plt.barh(bottom123 + ["All classes"], list(np.zeros_like(reordered_acc1_classwise)) + [acc1_avg], label='Top 1 Accuracy (All Classes)', height=0.5, align='edge')
plt.barh(bottom123 + ["All classes"], list(np.zeros_like(reordered_acc5_classwise)) + [acc5_avg], label='Top 5 Accuracy (All Classes)', height=-0.5, align='edge')
plt.legend(loc="upper left")
plt.show()
# -
# # 5. Outlook
#
# ## 5.1. Todos
#
# The training procedure could probably be improved by the following strategies:
#
# - *hyperparameter optimization*: The most sensitive hyperparameter is the learning rate, which was not optimized in any way for the experiments. Possible optimization strategies are a random search (in logspace) or a Bayesian optimiziation with validation accuracy is the criterion to search over.
# - *learning rate schedule*: the annealing of the learning rate was chosen ad-hoc. This could be improved by searching over the annealing hyperparameters together with the learning rate or by implementing a plateau detection to decrease learning rate whenever the validation accuracy plateaus.
# - *training/validation split*: currently it is not guaranteed that the training set for the remaining classes in the transfer learning contains samples from all classes, as some might be present in only in the validation set. Stratification was not possible due to the fact that for some classes there is just a single data point.
# - *image size*: the choice of 400-by-300 pixels was ad-hoc since it retains the dominant aspect ratio of the original images and is in the ballpark of ImageNet. There could be a better choice for this.
#
# ## 5.2. Extensions
#
# The remaining meta-data (fields 'masterCategory' and 'subCategory') could probably be used to improve learning and performance on the test set. The easiest approach would be to encode them as one-hot vectors and concatenate them with the output of the next-to-last layer of the pretrained ImageNet model, then learn a linear layer on top.
# For real-world purposes, this would mean that we deem it likely to encounter unseen data where the 'articleType' is not known but at least its 'masterCategory' and 'subCategory' are. There are potential caveats to this approach however, as some of the 'subCategory' labels are dead giveaways to the 'articleType' , e.g. the for 'articleType' 'Watches', the 'subCategory' is always 'Watches'. A model trained on this input might reasonably ignore the visual input entirely and just use the 'subCategory' label, which might not be desired.
#
# Using the field 'productDisplayName' has the same problems. Although I did not analyze this quantitatively, it seems that the 'prductDisplayName' often contains the class label directly or at least slight variations of it. Using this information via NLP techniques bears the risk of overfitting to the text features and ignoring the visual input.
| notebook/transfer_learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
from scipy.stats.stats import pearsonr
#from scipy.stats import linregress
sns.set_style('darkgrid')
import os
# ### What is Regression?
#
# Regression is a statistical method used to draw the relation between two variables. Often when data are collected there might be variables which are dependent on others. The exact relation between those variables can only be established by the regression methods. Determining this relationship helps to understand and predict the behaviour of one variable to the other.
#
# Most common application of the regression analysis is to estimate the value of the dependent variable for a given value or range of values of the independent variables. For example, using regression we can establish the relation between the commodity price and the consumption, based on the data collected from a random sample. Regression analysis produces the regression function of a data set, which is a mathematical model that best fits to the data available. This can easily be represented by a scatter plot. Graphically, regression is equivalent to finding the best fitting curve for the give data set. The function of the curve is the regression function. Using the mathematical model, the demand of a commodity can be predicted for a given price.
#
# Therefore, the regression analysis is widely used in predicting and forecasting. It is also used to establish relationships in experimental data, in the fields of physics, chemistry, and many natural sciences and engineering disciplines. If the relationship or the regression function is a linear function, then the process is known as a linear regression. In the scatter plot, it can be represented as a straight line. If the function is not a linear combination of the parameters, then the regression is non-linear.
# +
# linregress?
# -
# ### What is Correlation?
#
# Correlation is a measure of strength of the relationship between two variables. The correlation coefficient quantifies the degree of change in one variable based on the change in the other variable. In statistics, correlation is connected to the concept of dependence, which is the statistical relationship between two variables.
#
# The Pearsons’s correlation coefficient or just the correlation coefficient r is a value between -1 and 1 (-1≤r≤+1) . It is the most commonly used correlation coefficient and valid only for a linear relationship between the variables. If r=0, no relationship exist, and if r≥0, the relation is directly proportional; i.e. the value of one variable increases with the increase of the other. If r≤0, the relationship is inversely proportional; i.e. one variable decreases as the other increases.
#
# Because of the linearity condition, correlation coefficient r can also be used to establish the presence of a linear relationship between the variables.
# ### What is the difference between Regression and Correlation?
#
# Regression gives the form of the relationship between two random variables, and the correlation gives the degree of strength of the relationship.
#
# Regression analysis produces a regression function, which helps to extrapolate and predict results while correlation may only provide information on what direction it may change.
#
# The more accurate linear regression models are given by the analysis, if the correlation coefficient is higher. (|r|≥0.8)
# +
# pearsonr?
# -
# $$ r = \frac{\sum (x - m_x) (y - m_y)}
# {\sqrt{\sum (x - m_x)^2 \sum (y - m_y)^2}} $$
# $$ dot(x,y) = {\sum x_i y_i} $$
#
length = 100
x = np.random.normal(0, 10, length)
y = x + np.random.normal(0, 5, length)
plt.scatter(x, y, linewidths=0.02)
# +
# plt.scatter?
# -
np.dot(x - np.mean(x), y - np.mean(y)) / (np.std(x) * np.std(y)) / length
np.dot(x, y) / (np.std(x)* np.std(y)) / length
pearsonr(x, y)[0]
# +
# pearsonr?
# -
sns.regplot(x, y)
path = '../data/stock_dfs'
names = os.listdir(path)
names = names[:50]
df_main = pd.DataFrame()
for name in names:
add = os.path.join(path, name)
df = pd.read_csv(add, index_col=0)
df.drop(['High', 'Low', 'Open', 'Close', 'Volume'], axis=1, inplace=True)
df = df.rename(columns={'Adj Close': name[:-4]})
df_main = df_main.join(df, how='outer')
df_corr = df_main.corr()
plt.figure(figsize=(20,20))
sns.heatmap(df_corr, cmap='YlGnBu')
df_corr['UAL'].sort_values(ascending=False)
plt.figure(figsize=(15,10))
df_main['UAL'].plot()
df_main['BA'].plot()
# +
#sns.pairplot(df_main, kind="reg")
# -
df['x'] = x
df['y'] = y
df.corr()
path = '../data/stock_dfs'
names = os.listdir(path)[:5]
names[1]
os.path.join(path, names[2])
df1 = pd.read_csv('../data/stock_dfs/UAL.csv', index_col=0)
df1['Open']
df2 = pd.read_csv('../data/stock_dfs/TROW.csv', index_col=0)
df2.drop(['High', 'Low', 'Close', , 'Volume'], 1, inplace=True)
df2
names[0][:-4]
# +
main_df = pd.DataFrame()
for name in names:
adds = os.path.join(path, name)
df = pd.read_csv('{}'.format(adds))
df.set_index('Date', inplace=True)
df.rename(columns={'Adj Close':name[:-4]}, inplace=True)
df.drop(['Open','High','Low','Close','Volume'],1,inplace=True)
if main_df.empty:
main_df = df
else:
main_df = main_df.join(df, how='outer')
print(main_df.head())
#main_df.to_csv('sp500_joined_closes.csv')
# -
sns.pairplot(main_df.head(5), kind='reg')
sns.heatmap(main_df.corr(), cmap="YlGnBu")
# +
# sns.pairplot?
# +
# sns.heatmap?
# -
main_df.corr()['UAL'].sort_values(ascending=False)
| numericalPython/correlation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.12 ('NLU_env')
# language: python
# name: python3
# ---
# # Import libraries
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
# # Import results
df = pd.read_csv("Results/Total_score.csv", index_col=0)
df.head()
# # Create a dictionary for each Finetune dataset and Testing dataset
score_list = ['Rouge-1','Rouge-2','Rouge-l','BERTscore']
Sum_score_dict = {}
# Sum_score_dict.keys = score_list
for score in score_list:
df_mean = df.groupby(['Finetune', 'Category']).agg({score: ['mean']})
df_mean = df_mean.pivot_table(score, ['Finetune'], 'Category')
df_mean = df_mean.droplevel(0, axis=1)
Sum_score_dict[score] = df_mean
df_mean.head()
# # Heatmap plot
def plot_heatmap(dict, key):
sns.set(rc={'figure.figsize':(10, 8)})
ax = sns.heatmap(dict[key], annot=True, cmap="YlGnBu", fmt='.3g')
plt.yticks(rotation=0)
ax.xaxis.tick_top()
plt.tick_params(top = False)
ax.xaxis.set_label_position('top')
plt.title(key, y=-.08, fontsize=20)
# plt.savefig('Heatmap_'+key+'.png')
plt.show()
for score in score_list:
plot_heatmap(Sum_score_dict, score)
# +
# df_mean.style.background_gradient(cmap ='coolwarm').set_properties(**{'font-size': '20px'})
# -
# # Boxplot plot
for score in score_list:
sns.set(rc={'figure.figsize':(15, 8)})
sns.set_theme(style="ticks", palette="pastel")
sns.boxplot(x="Category", y=score,
hue="Finetune",
data=df)
sns.despine(offset=10, trim=True)
plt.title(score, fontsize=20)
plt.show()
# # Tokenize lenght
# +
model_params={
"MODEL":"t5-base", # model_type: t5-base/t5-large
"SEED": 42 # set seed for reproducibility
}
from transformers import T5Tokenizer
tokenizer = T5Tokenizer.from_pretrained(model_params["MODEL"])
import torch
# Set random seeds and deterministic pytorch for reproducibility
torch.manual_seed(model_params["SEED"]) # pytorch random seed
np.random.seed(model_params["SEED"]) # numpy random seed
torch.backends.cudnn.deterministic = True
# +
tokenizer = T5Tokenizer.from_pretrained('t5-base')
df_tkn = pd.DataFrame()
df_tkn['tokenized_sents'] = df.apply(lambda row: tokenizer(row['Actual Text']), axis=1)
df['Actual_tkn_len'] = df_tkn.apply(lambda row: len(row['tokenized_sents']['input_ids']), axis=1)
df_tkn['tokenized_sents'] = df.apply(lambda row: tokenizer(row['Generated Text']), axis=1)
df['Generate_tkn_len'] = df_tkn.apply(lambda row: len(row['tokenized_sents']['input_ids']), axis=1)
df.head()
# -
sns.displot(
data = df,
x = 'Actual_tkn_len',
kind = "kde",
color = 'darkblue',
hue = "Finetune",
height = 8,
aspect = 1.5
)
sns.displot(
data = df[df.Finetune == "entertainment"],
x = 'Actual_tkn_len',
kind = "kde",
color = 'darkblue',
hue = "Category",
height = 8,
aspect = 1.5
)
sns.displot(
data = df[df.Finetune == "politics"],
x = 'Actual_tkn_len',
kind = "kde",
color = 'darkblue',
hue = "Category",
height = 8,
aspect = 1.5
)
df_ = df[df.Finetune == "politics"]
df_ = df_[df.Category == "entertainment"]
sns.lineplot(
data = df_,
x = 'Actual_tkn_len',
y = 'BERTscore'
)
sns.lineplot(
data = df_,
x = 'Actual_tkn_len',
y = 'Rouge-l'
)
plt.axvline(x=181,
color='red',
ls='--',
lw=2)
Real_df = pd.read_csv('Dataset_Evaluate_Mixed.csv')
Real_df
for _, row in df.iterrows():
string = (row['Generated Text'])
break
string
Real_sum = list(Real_df["summaries"])
Real_sum[0]
| All python files/1_7_Visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import numpy, scipy, matplotlib.pyplot as plt, IPython.display as ipd
import librosa, librosa.display
import numpy as np
import math
os.chdir('/home/student/Downloads/new_violin_viola/all/viola/combined_train')
x1=[]
y_train=[]
mfcc_list_mean = []
mfcc_list_std = []
freqViolin = []
freqViola = []
for f_name in os.listdir('/home/student/Downloads/new_violin_viola/all/viola/combined_train'):
if f_name.endswith('.mp3') or f_name.endswith('.wav'):
print(f_name)
#temp = x/x.max() #normalization
#S = librosa.feature.melspectrogram(temp, sr=sr, n_mels=128) # 128 mel bands
#mfcc = librosa.feature.mfcc(S=librosa.power_to_db(S), n_mfcc=13)
#tempList = list(np.mean(mfcc,1))
#tempList1 = list(np.std(mfcc,1))
x, sr = librosa.load(f_name)
hop_length = 512
X = librosa.stft(x)
S = librosa.amplitude_to_db(X, ref=np.max)
l1 =[]
#by default, there are 1025 frequency bins in 0 to sr/2 (sr = 22050)
hist=[]
hist1=[]
for i in range(len(S)):
for j in range(len(S[0])):
if S[i][j]>=(-11):
temp = i *((22050/2)/1025)
if temp >0:
hist.append(math.log2(temp))
hist1.append(temp)
x1.append(hist)
freqViola.append(hist1)
y_train.append("viola")
os.chdir('/home/student/Downloads/new_violin_viola/all/violin/combined_train')
for f_name in os.listdir('/home/student/Downloads/new_violin_viola/all/violin/combined_train'):
if f_name.endswith('.mp3') or f_name.endswith('.wav'):
print(f_name)
x, sr = librosa.load(f_name)
hop_length = 512
X = librosa.stft(x)
S = librosa.amplitude_to_db(X, ref=np.max)
l1 =[]
#by default, there are 1025 frequency bins in 0 to sr/2 (sr = 22050)
hist=[]
hist1=[]
for i in range(len(S)):
for j in range(len(S[0])):
if S[i][j]>=(-11):
temp = i *((22050/2)/1025)
if temp >0:
hist.append(math.log2(temp))
hist1.append(temp)
x1.append(hist)
freqViolin.append(hist1)
y_train.append("violin")
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import statistics
X_train= [[statistics.mean(a), statistics.median(a), min(a), max(a),np.percentile(np.array(a), 25), np.percentile(np.array(a), 75)] for a in x1]
#y=["violin","violin","violin","violin","violin","viola","viola","viola","viola","viola"]
# +
os.chdir('/home/student/Downloads/new_violin_viola/all/viola/combined_test')
x1=[]
y_test=[]
for f_name in os.listdir('/home/student/Downloads/new_violin_viola/all/viola/combined_test'):
if f_name.endswith('.mp3') or f_name.endswith('.wav'):
print(f_name)
x, sr = librosa.load(f_name)
hop_length = 512
X = librosa.stft(x)
S = librosa.amplitude_to_db(X, ref=np.max)
l1 =[]
#by default, there are 1025 frequency bins in 0 to sr/2 (sr = 22050)
hist=[]
hist1=[]
for i in range(len(S)):
for j in range(len(S[0])):
if S[i][j]>=(-11):
temp = i *((22050/2)/1025)
if temp >0:
hist.append(math.log2(temp))
hist1.append(temp)
x1.append(hist)
freqViola.append(hist1)
y_test.append("viola")
os.chdir('/home/student/Downloads/new_violin_viola/all/violin/combined_test')
for f_name in os.listdir('/home/student/Downloads/new_violin_viola/all/violin/combined_test'):
if f_name.endswith('.mp3') or f_name.endswith('.wav'):
print(f_name)
x, sr = librosa.load(f_name)
hop_length = 512
X = librosa.stft(x)
S = librosa.amplitude_to_db(X, ref=np.max)
l1 =[]
#by default, there are 1025 frequency bins in 0 to sr/2 (sr = 22050)
hist=[]
hist1=[]
for i in range(len(S)):
for j in range(len(S[0])):
if S[i][j]>=(-11):
temp = i *((22050/2)/1025)
if temp >0:
hist.append(math.log2(temp))
hist1.append(temp)
x1.append(hist)
freqViolin.append(hist1)
y_test.append("violin")
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import statistics
X_test= [[statistics.mean(a), statistics.median(a), min(a), max(a),np.percentile(np.array(a), 25), np.percentile(np.array(a), 75)] for a in x1]
# +
from numpy.random import seed
seed(14)
from sklearn import metrics
from sklearn.metrics import classification_report, confusion_matrix
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
classifier = RandomForestClassifier(max_depth=10, random_state=0)
classifier.fit(X_train, y_train)
y_pred = classifier.predict(X_test)
print(confusion_matrix(y_test, y_pred))
print('Accuracy' + str(accuracy_score(y_test, y_pred)))
print(classification_report(y_test,y_pred))
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
# +
from sklearn.neighbors import KNeighborsClassifier
model = KNeighborsClassifier(n_neighbors=3)
# Train the model using the training sets
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
print(confusion_matrix(y_test, y_pred))
print('Accuracy' + str(accuracy_score(y_test, y_pred)))
print(classification_report(y_test,y_pred))
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
# +
from sklearn import svm
#Create a svm Classifier
clf = svm.SVC(random_state=42,kernel='linear') # Linear Kernel
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
print(confusion_matrix(y_test, y_pred))
print('Accuracy' + str(accuracy_score(y_test, y_pred)))
print(classification_report(y_test,y_pred))
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
# -
svm = svm.SVC(random_state=42, kernel='rbf')
svm.fit(X_train, y_train)
y_pred = svm.predict(X_test)
print(confusion_matrix(y_test, y_pred))
print('Accuracy' + str(accuracy_score(y_test, y_pred)))
print(classification_report(y_test,y_pred))
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
# +
#statistics.mean(freqViola), statistics.median(freqViola), np.percentile(np.array(freqViola), 25), np.percentile(np.array(freqViola), 75)
#viola1= [[statistics.mean(freqViola[i])] for i in range(len(freqViola))]
# +
#freqViola
#freqViolin
#mean, median, mode, 25th, 75th
from statistics import stdev
viola1= [[statistics.mean(a), statistics.median(a), np.percentile(np.array(a), 25), np.percentile(np.array(a), 75)] for a in freqViola]
violin1= [[statistics.mean(a), statistics.median(a), np.percentile(np.array(a), 25), np.percentile(np.array(a), 75)] for a in freqViolin]
meanstdViolaList = []
for i in viola1:
meanstdViolaList.append(i[0])
meanstdViola = stdev(meanstdViolaList)
print("Standard deviation of mean Viola frequency: " + str(meanstdViola))
meanstdViolinList = []
for i in violin1:
meanstdViolinList.append(i[0])
meanstdViolin = stdev(meanstdViolinList)
print("Standard deviation of mean Violin frequency: " + str(meanstdViolin))
# +
medianstdViolaList = []
for i in viola1:
medianstdViolaList.append(i[1])
medianstdViola = stdev(medianstdViolaList)
print("Standard deviation of median Viola frequency: " + str(medianstdViola))
medianstdViolinList = []
for i in violin1:
medianstdViolinList.append(i[1])
medianstdViolin = stdev(medianstdViolinList)
print("Standard deviation of median Violin frequency: " + str(medianstdViolin))
# +
#modestdViolaList = []
#for i in viola1:
# modestdViolaList.append(i[2])
#modestdViola = stdev(modestdViolaList)
#print("Standard deviation of mode Viola frequency: " + str(modestdViola))
#modestdViolinList = []
#for i in violin1:
# modestdViolinList.append(i[2])
#modestdViolin = stdev(modestdViolinList)
#print("Standard deviation of mode Violin frequency: " + str(modestdViolin))
# +
temp = []
for i in viola1:
temp.append(i[2])
firstQ1 = stdev(temp)
print("Standard deviation of 25th percentile Viola frequency: " + str(firstQ1))
temp1 = []
for i in violin1:
temp1.append(i[2])
firstQ2 = stdev(temp1)
print("Standard deviation of 25th percentile Violin frequency: " + str(firstQ2))
temp3 = []
for i in viola1:
temp3.append(i[3])
thirdQ1 = stdev(temp3)
print("Standard deviation of 75th percentile Viola frequency: " + str(thirdQ1))
temp4 = []
for i in violin1:
temp4.append(i[3])
thirdQ2 = stdev(temp4)
print("Standard deviation of 75th percentile Violin frequency: " + str(thirdQ2))
meanstdList= meanstdViolaList+meanstdViolinList
meanstd = stdev(meanstdList)
print("Standard deviation of mean frequency: " + str(meanstd))
medianstdList= medianstdViolaList+medianstdViolinList
medianstd = stdev(medianstdList)
print("Standard deviation of median frequency: " + str(medianstd))
temp0 = temp + temp1
firstQ = stdev(temp0)
print("Standard deviation of 25th percentile frequency: " + str(firstQ))
temp00 = temp3 + temp4
thirdQ = stdev(temp00)
print("Standard deviation of 75th percentile frequency: " + str(thirdQ))
# -
# +
import plotly.express as px
import plotly.io as pio
from sklearn.decomposition import PCA
pca = PCA(n_components=2)
components = pca.fit_transform(X_train+X_test)
fig = px.scatter(components, x=0, y=1, color=y_train+y_test)
fig.update_layout(
title="",
xaxis_title="First Component",
yaxis_title="Second Component",
font=dict(
family="Courier New, monospace",
size=18,
color="#7f7f7f"
),
legend=dict(
yanchor="top",
y=0.99,
xanchor="left",
x=0.00
)
)
fig.show()
#pio.write_image(fig, 'soloAll.png')
#fig.write_image("soloAll.eps")
# -
| frequencyHistogram_nonsolo_all.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.10.4 (''venv'': venv)'
# language: python
# name: python3
# ---
# # Boston Housing Classification Random Forest
import sys
sys.path.append("..")
from pyspark.sql.types import BooleanType
from pyspark.ml.feature import StringIndexer, VectorAssembler
from pyspark.ml.classification import RandomForestClassifier
from pyspark.sql.session import SparkSession
from pyspark.sql.functions import expr
from pyspark.ml.evaluation import BinaryClassificationEvaluator
from helpers.path_translation import translate_to_file_string
from helpers.data_prep_and_print import print_df
inputFile = translate_to_file_string("../data/Boston_Housing_Data.csv")
# Spark session creation
spark = (SparkSession
.builder
.appName("BostonHousingClassificationRandomForest")
.getOrCreate())
# DataFrame creation using an ifered Schema
df = spark.read.option("header", "true") \
.option("inferSchema", "true") \
.option("delimiter", ";") \
.csv(inputFile) \
.withColumn("CATBOOL", expr("CAT").cast(BooleanType()))
print(df.printSchema())
# Prepare training and test data.
# +
featureCols = df.columns.copy()
featureCols.remove("MEDV")
featureCols.remove("CAT")
featureCols.remove("CATBOOL")
print(featureCols)
assembler = VectorAssembler(outputCol="features", inputCols=featureCols)
# -
labledPointDataSet = assembler.transform(df)
splits = labledPointDataSet.randomSplit([0.9, 0.1 ], 12345)
training = splits[0]
test = splits[1]
# Random Forest Classifier
rf = RandomForestClassifier(labelCol="CAT", featuresCol="features",impurity="gini", \
minInstancesPerNode=10, featureSubsetStrategy='sqrt', subsamplingRate=0.95, seed= 12345)
# Train the model
rfModel = rf.fit(training)
# Test the model
predictions = rfModel.transform(test)
print_df(predictions)
evaluator = BinaryClassificationEvaluator(labelCol="CAT",rawPredictionCol="rawPrediction", metricName="areaUnderROC")
accuracy = evaluator.evaluate(predictions)
print("Test Error",(1.0 - accuracy))
| solutions/boston_housing_classification_random_forest_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.0
# language: julia
# name: julia-0.6
# ---
using PyPlot,Seismic
# +
#Create linear events
d,ext = SeisLinearEvents(p1 = [-.001, 0.0015],tau=[1., 1./3.],dx1=5);
#Randomly decimate
dec = SeisDecimate(d);
# -
param = Dict(:Niter=>100,:fmax=>60,:padt=>2,:padx=>2,:dt=>0.004)
dpocs = SeisPOCS(dec;param...);
figure(1, figsize=(10, 5))
subplot(121)
SeisPlot(dec,cmap="seismic",fignum=1,pclip=200,fignum=1)
subplot(122)
SeisPlot(dec,plot_type="FK", cmap="seismic", dy=0.004,fignum=1,hbox=5,pclip=200,fignum=1)
figure(2,figsize=(10, 5))
subplot(121)
SeisPlot(dpocs[:,:,1,1,1],cmap="seismic",fignum=2,pclip=200)
subplot(122)
SeisPlot(dpocs[:,:,1,1,1],plot_type="FK",cmap="seismic",dy=0.004,fignum=2,pclip=200)
| examples/POCS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] origin_pos=0
# # 来自变形金刚(BERT)的双向编码器表示
# :label:`sec_bert`
#
# 我们引入了几个单词嵌入模型来理解自然语言。训练前后,可以将输出视为矩阵,其中每一行都是表示预定义词汇的一个单词的矢量。事实上,这些单词嵌入模型都是 * 上下文无关 *。让我们首先说明这个财产。
#
# ## 从上下文独立到上下文敏感
#
# 回想一下 :numref:`sec_word2vec_pretraining` 和 :numref:`sec_synonyms` 中的实验。例如,word2vec 和 GLOVE 都将相同的预训练向量分配给同一个单词,而不管单词的上下文如何(如果有)。从形式上来说,任何词元 $x$ 的上下文无关表示是一个函数 $f(x)$,它只需要 $x$ 作为输入。鉴于自然语言中的多聚论和复杂语义的丰富性,与上下文无关的表示有明显的局限性。例如,上下文 “起重机正在飞行” 和 “起重机驾驶员来了” 中的 “起重机” 一词具有完全不同的含义;因此,根据上下文,同一个词可能被分配不同的表示形式。
#
# 这激励了 * 上下文敏感的 * 单词表示形式的开发,其中单词的表示取决于它们的上下文。因此,词元 $x$ 的上下文相关表示是函数 $f(x, c(x))$,具体取决于 $x$ 及其上下文 $c(x)$。受欢迎的上下文相关表示包括 tagLM(语言模型增强序列词元器):cite:`Peters.Ammar.Bhagavatula.ea.2017`、Cove(上下文向量):cite:`McCann.Bradbury.Xiong.ea.2017` 和 elMO(来自语言模型的嵌入):cite:`Peters.Neumann.Iyyer.ea.2018`。
#
# 例如,通过将整个序列作为输入,elMO 是一个函数,它为输入序列中的每个单词分配一个表示形式。具体来说,elMO 将预训练的双向 LSTM 中的所有中间图层表示法合并为输出表示法。然后,elMO 表示法将作为附加功能添加到下游任务的现有监督模型中,例如连接现有模型中的 elMO 表示法和原始表示法(例如 GLOVE)。一方面,在添加 elMO 表示之后,预训练的双向 LSTM 模型中的所有权重都会被冻结。另一方面,现有的受监督模型是专门针对给定任务定制的。当时利用不同的最佳模型来处理不同的任务,增加 ELMO 改善了六个自然语言处理任务的最新状态:情绪分析、自然语言推断、语义角色词元化、共引解析、命名实体识别和问题回答。
#
# ## 从特定于任务到不可知的任务
#
# 尽管 elMO 显著改进了针对各种自然语言处理任务的解决方案,但每个解决方案仍然取决于 * 任务特定的 * 架构。但是,为每个自然语言处理任务设计一个特定的架构实际上并不平凡。GPT(生成预训练)模型代表着为上下文相关表示 :cite:`Radford.Narasimhan.Salimans.ea.2018` 设计一个通用 *任务无关* 模型的努力。GPT 建立在变压器解码器之上,预先训练将用于表示文本序列的语言模型。将 GPT 应用于下游任务时,语言模型的输出将被输入添加的线性输出图层,以预测任务的标签。与冻结预训练模型参数的 elMO 形成鲜明对比,GPT 在监督学习下游任务期间对预训练的变压器解码器中的 * 所有参数进行了微调。GPT 在自然语言推断、问答、句子相似性和分类等十二项任务上进行了评估,并在对模型架构的改动最小的情况下改善了其中 9 项任务的最新状态。
#
# 但是,由于语言模型的自回归性质,GPT 只是向前(从左到右)。在 “我去银行存款现金” 和 “我去银行坐下来” 的情况下,由于 “银行” 对左边的情境敏感,GPT 将返回 “银行” 的相同表述,尽管它有不同的含义。
#
# ## BERT:结合两全其美
#
# 正如我们所看到的那样,elMO 以双向方式对上下文进行编码,但使用特定于任务的架构;虽然 GPT 与任务无关,但是对上下文进行了从左到右编码。BERT(来自变形金刚的双向编码器表示)结合了两全其美的结合,对于范围广泛的自然语言处理任务 :cite:`Devlin.Chang.Lee.ea.2018`,对于上下文的双向编码器表示法,只需最少的体系结构更改。使用预训练的变压器编码器,BERT 能够根据其双向上下文表示任何词元。在监督下游任务学习期间,BERT 在两个方面与 GPT 类似。首先,BERT 表示将被输入添加的输出层,根据任务的性质对模型架构进行最小的更改,例如对每个词元的预测与整个序列的预测。其次,预训练的变压器编码器的所有参数都经过微调,而额外的输出层将从头开始训练。:numref:`fig_elmo-gpt-bert` 描述了 elMO、GPT 和 BERT 之间的差异。
#
# 
# :label:`fig_elmo-gpt-bert`
#
# BERT 进一步改善了十一项自然语言处理任务的最新状态,这些类别包括:(i) 单一文本分类(例如情绪分析)、(ii)文本对分类(例如自然语言推理)、(iii)问答、(iv)文本词元化(例如,指定实体识别)。所有这些都在 2018 年提出,从上下文敏感的 elMO 到与任务无关的 GPT 和 BERT,概念上简单但经验强大的自然语言深度表示预训练,彻底改变了各种自然语言处理任务的解决方案。
#
# 在本章的其余部分,我们将深入研究 BERT 的预培训。当 :numref:`chap_nlp_app` 中解释自然语言处理应用程序时,我们将说明对下游应用程序的 BERT 的微调。
#
# + origin_pos=2 tab=["pytorch"]
import torch
from torch import nn
from d2l import torch as d2l
# + [markdown] origin_pos=3
# ## 输入表示法
# :label:`subsec_bert_input_rep`
#
# 在自然语言处理中,某些任务(例如情绪分析)将单个文本作为输入,而在其他一些任务(例如自然语言推断)中,输入是一对文本序列。BERT 输入序列明确表示单个文本对和文本对。在前者中,BERT 输入序列是特殊分类词元 “<cls>”、文本序列的词元和特殊分隔词元 “<sep>” 的串联。在后者中,BERT 输入序列是 “<cls>”、第一个文本序列的词元 “<sep>”、第二个文本序列的词元和 “<sep>” 的连接。我们将始终将术语 “BERT 输入序列” 与其他类型的 “序列” 区分开来。例如,一个 *BERT 输入序列 * 可能包含一个 * 文本序列 * 或两个 * 文本序列 *。
#
# 为了区分文本对,学习的细分嵌入 $\mathbf{e}_A$ 和 $\mathbf{e}_B$ 分别添加到第一个序列和第二序列的词元嵌入中。对于单个文本输入,只使用 $\mathbf{e}_A$。
#
# 以下 `get_tokens_and_segments` 以一句或两句话作为输入,然后返回 BERT 输入序列的词元及其对应的段 ID。
#
# + origin_pos=4 tab=["pytorch"]
#@save
def get_tokens_and_segments(tokens_a, tokens_b=None):
"""Get tokens of the BERT input sequence and their segment IDs."""
tokens = ['<cls>'] + tokens_a + ['<sep>']
# 0 and 1 are marking segment A and B, respectively
segments = [0] * (len(tokens_a) + 2)
if tokens_b is not None:
tokens += tokens_b + ['<sep>']
segments += [1] * (len(tokens_b) + 1)
return tokens, segments
# + [markdown] origin_pos=5
# BERT 选择变压器编码器作为其双向架构。在变压器编码器中常见,位置嵌入在 BERT 输入序列的每个位置都添加。但是,与原来的变压器编码器不同,BERT 使用 * 可学习 * 位置嵌入。总而言之,:numref:`fig_bert-input` 显示,BERT 输入序列的嵌入是词元嵌入、区段嵌入和位置嵌入的总和。
#
#  :label:`fig_bert-input`
#
# 以下 `BERTEncoder` 类与 :numref:`sec_transformer` 中实施的 `TransformerEncoder` 类类似。与 `TransformerEncoder` 不同,`BERTEncoder` 使用细分嵌入和可学习的位置嵌入。
#
# + origin_pos=7 tab=["pytorch"]
#@save
class BERTEncoder(nn.Module):
"""BERT encoder."""
def __init__(self, vocab_size, num_hiddens, norm_shape, ffn_num_input,
ffn_num_hiddens, num_heads, num_layers, dropout,
max_len=1000, key_size=768, query_size=768, value_size=768,
**kwargs):
super(BERTEncoder, self).__init__(**kwargs)
self.token_embedding = nn.Embedding(vocab_size, num_hiddens)
self.segment_embedding = nn.Embedding(2, num_hiddens)
self.blks = nn.Sequential()
for i in range(num_layers):
self.blks.add_module(f"{i}", d2l.EncoderBlock(
key_size, query_size, value_size, num_hiddens, norm_shape,
ffn_num_input, ffn_num_hiddens, num_heads, dropout, True))
# In BERT, positional embeddings are learnable, thus we create a
# parameter of positional embeddings that are long enough
self.pos_embedding = nn.Parameter(torch.randn(1, max_len,
num_hiddens))
def forward(self, tokens, segments, valid_lens):
# Shape of `X` remains unchanged in the following code snippet:
# (batch size, max sequence length, `num_hiddens`)
X = self.token_embedding(tokens) + self.segment_embedding(segments)
X = X + self.pos_embedding.data[:, :X.shape[1], :]
for blk in self.blks:
X = blk(X, valid_lens)
return X
# + [markdown] origin_pos=8
# 假设词汇量大小是 10000。为了演示 `BERTEncoder` 的前向推理,让我们创建它的实例并初始化其参数。
#
# + origin_pos=10 tab=["pytorch"]
vocab_size, num_hiddens, ffn_num_hiddens, num_heads = 10000, 768, 1024, 4
norm_shape, ffn_num_input, num_layers, dropout = [768], 768, 2, 0.2
encoder = BERTEncoder(vocab_size, num_hiddens, norm_shape, ffn_num_input,
ffn_num_hiddens, num_heads, num_layers, dropout)
# + [markdown] origin_pos=11
# 我们将 `tokens` 定义为 2 个长度为 8 的 BERT 输入序列,其中每个词元都是词汇的索引。输入 `BERTEncoder` 的 `BERTEncoder` 和输入 `tokens` 返回编码结果,其中每个词元由超参数 `num_hiddens` 预定义的向量表示,其长度由超参数 `num_hiddens` 预定义。此超参数通常称为变压器编码器的 * 隐藏大小 *(隐藏单位数)。
#
# + origin_pos=13 tab=["pytorch"]
tokens = torch.randint(0, vocab_size, (2, 8))
segments = torch.tensor([[0, 0, 0, 0, 1, 1, 1, 1], [0, 0, 0, 1, 1, 1, 1, 1]])
encoded_X = encoder(tokens, segments, None)
encoded_X.shape
# + [markdown] origin_pos=14
# ## 培训前任务
# :label:`subsec_bert_pretraining_tasks`
#
# `BERTEncoder` 的前向推断给出了输入文本的每个词元的 BERT 表示以及插入的特殊词元 “<cls>” 和 “<seq>”。接下来,我们将使用这些表示法来计算预训练 BERT 的损失函数。预培训由以下两项任务组成:蒙版语言建模和下一句话预测。
#
# ### 蒙面语言建模
# :label:`subsec_mlm`
#
# 如 :numref:`sec_language_model` 所示,语言模型使用左侧的上下文来预测词元。为了对上下文进行双向编码以表示每个词元,BERT 会随机掩盖词元,并使用双向上下文中的词元以自我监督的方式预测被掩码的词元。此任务被称为 * 蒙面语言模型 *。
#
# 在此预训任务中,15% 的代币将随机选择作为预测的蒙面代币。要在不使用标签作弊的情况下预测蒙面的词元,一种简单的方法是始终在 <mask>BERT 输入序列中用特殊的 “” 词元替换它。但是,人为的特殊词元 “<mask>” 永远不会出现在微调中。为避免预训和微调之间的这种不匹配,如果词元被掩盖进行预测(例如,在 “这部电影很棒” 中选择了 “很棒” 来掩盖和预测),则在输入内容中将替换为:
#
# * <mask>80% 的时间里,一个特殊的 “” 词元(例如,“这部电影很棒” 变成 “这部电影是”<mask>);
# * 10% 的时间内随机词元(例如,“这部电影很棒” 变成 “这部电影很喝”);
# * 10% 的时间内不变的标签词元(例如,“这部电影很棒” 变成 “这部电影很棒”)。
#
# 请注意,在 15% 的时间里,插入随机词元的 10%。这种偶尔的噪音鼓励 BERT 在双向上下文编码中减少对蒙面词元的偏见(特别是当标签词元保持不变时)。
#
# 我们实施了以下 `MaskLM` 课程来预测 BERT 预训的蒙面语言模型任务中的蒙面词元。该预测使用一个隐藏层 MLP(`self.mlp`)。在前向推断中,它需要两个输入:`BERTEncoder` 的编码结果和用于预测的代币位置。输出是这些仓位的预测结果。
#
# + origin_pos=16 tab=["pytorch"]
#@save
class MaskLM(nn.Module):
"""The masked language model task of BERT."""
def __init__(self, vocab_size, num_hiddens, num_inputs=768, **kwargs):
super(MaskLM, self).__init__(**kwargs)
self.mlp = nn.Sequential(nn.Linear(num_inputs, num_hiddens),
nn.ReLU(),
nn.LayerNorm(num_hiddens),
nn.Linear(num_hiddens, vocab_size))
def forward(self, X, pred_positions):
num_pred_positions = pred_positions.shape[1]
pred_positions = pred_positions.reshape(-1)
batch_size = X.shape[0]
batch_idx = torch.arange(0, batch_size)
# Suppose that `batch_size` = 2, `num_pred_positions` = 3, then
# `batch_idx` is `torch.tensor([0, 0, 0, 1, 1, 1])`
batch_idx = torch.repeat_interleave(batch_idx, num_pred_positions)
masked_X = X[batch_idx, pred_positions]
masked_X = masked_X.reshape((batch_size, num_pred_positions, -1))
mlm_Y_hat = self.mlp(masked_X)
return mlm_Y_hat
# + [markdown] origin_pos=17
# 为了演示 `MaskLM` 的前向推断,我们创建了它的实例 `mlm` 并对其进行初始化。回想一下,`encoded_X` 从前向推断 `BERTEncoder` 代表 2 个 BERT 输入序列。我们将 `mlm_positions` 定义为在 `encoded_X` 的 BERT 输入序列中要预测的 3 个指数。`mlm` 的前瞻推断回报预测结果为 `mlm_Y_hat`,在 `encoded_X` 的所有蒙面仓位 `mlm_positions`。对于每个预测,结果的大小等于词汇量大小。
#
# + origin_pos=19 tab=["pytorch"]
mlm = MaskLM(vocab_size, num_hiddens)
mlm_positions = torch.tensor([[1, 5, 2], [6, 1, 5]])
mlm_Y_hat = mlm(encoded_X, mlm_positions)
mlm_Y_hat.shape
# + [markdown] origin_pos=20
# 通过掩码下的预测词元 `mlm_Y_hat` 的地面真相标签 `mlm_Y`,我们可以计算 BERT 预训练中蒙面语言模型任务的交叉熵损失。
#
# + origin_pos=22 tab=["pytorch"]
mlm_Y = torch.tensor([[7, 8, 9], [10, 20, 30]])
loss = nn.CrossEntropyLoss(reduction='none')
mlm_l = loss(mlm_Y_hat.reshape((-1, vocab_size)), mlm_Y.reshape(-1))
mlm_l.shape
# + [markdown] origin_pos=23
# ### 下一句预测
# :label:`subsec_nsp`
#
# 虽然蒙版语言建模能够对表示单词的双向上下文进行编码,但它并没有明确建模文本对之间的逻辑关系。为了帮助理解两个文本序列之间的关系,BERT 在其预训中考虑了二进制分类任务 * 下一句预测 *。在为预训生成句子对时,有一半时间它们确实是带有 “True” 标签的连续句子;而另一半时间,第二个句子是从标有 “False” 标签的语料库中随机抽取的。
#
# 接下来的 `NextSentencePred` 类使用一个隐藏层 MLP 来预测第二句是否是 BERT 输入序列中第一句的下一句。由于变压器编码器中的自我注意力,特殊词元 “<cls>” 的 BERT 表示对输入的两个句子进行了编码。因此,MLP 分类器的输出层 (`self.output`) 采用 `X` 作为输入,其中 `X` 是 MLP 隐藏层的输出,其输入是编码的 “<cls>” 词元。
#
# + origin_pos=25 tab=["pytorch"]
#@save
class NextSentencePred(nn.Module):
"""The next sentence prediction task of BERT."""
def __init__(self, num_inputs, **kwargs):
super(NextSentencePred, self).__init__(**kwargs)
self.output = nn.Linear(num_inputs, 2)
def forward(self, X):
# `X` shape: (batch size, `num_hiddens`)
return self.output(X)
# + [markdown] origin_pos=26
# 我们可以看到,`NextSentencePred` 实例的前向推断返回每个 BERT 输入序列的二进制预测。
#
# + origin_pos=28 tab=["pytorch"]
# PyTorch by default won't flatten the tensor as seen in mxnet where, if
# flatten=True, all but the first axis of input data are collapsed together
encoded_X = torch.flatten(encoded_X, start_dim=1)
# input_shape for NSP: (batch size, `num_hiddens`)
nsp = NextSentencePred(encoded_X.shape[-1])
nsp_Y_hat = nsp(encoded_X)
nsp_Y_hat.shape
# + [markdown] origin_pos=29
# 还可以计算两个二进制分类的交叉熵损失。
#
# + origin_pos=31 tab=["pytorch"]
nsp_y = torch.tensor([0, 1])
nsp_l = loss(nsp_Y_hat, nsp_y)
nsp_l.shape
# + [markdown] origin_pos=32
# 值得注意的是,上述两项预培训任务中的所有标签都可以在没有人工标签的情况下从培训前语料库中轻而易举地获得。原来的 BERT 已经在 Bookcorpus :cite:`Zhu.Kiros.Zemel.ea.2015` 和英语维基百科的连接方面进行了预培训。这两个文本语句是巨大的:它们分别有 8 亿个单词和 25 亿个单词。
#
# ## 把所有东西放在一起
#
# 在预训练 BERT 时,最终损失函数是掩码语言建模的损失函数和下一句预测的线性组合。现在我们可以通过实例化三个类 `BERTEncoder`、`MaskLM` 和 `NextSentencePred` 来定义 `BERTModel` 类。前向推理返回编码的 BERT 表示 `encoded_X`、对蒙面语言建模 `mlm_Y_hat` 的预测以及下一句预测 `nsp_Y_hat`。
#
# + origin_pos=34 tab=["pytorch"]
#@save
class BERTModel(nn.Module):
"""The BERT model."""
def __init__(self, vocab_size, num_hiddens, norm_shape, ffn_num_input,
ffn_num_hiddens, num_heads, num_layers, dropout,
max_len=1000, key_size=768, query_size=768, value_size=768,
hid_in_features=768, mlm_in_features=768,
nsp_in_features=768):
super(BERTModel, self).__init__()
self.encoder = BERTEncoder(vocab_size, num_hiddens, norm_shape,
ffn_num_input, ffn_num_hiddens, num_heads, num_layers,
dropout, max_len=max_len, key_size=key_size,
query_size=query_size, value_size=value_size)
self.hidden = nn.Sequential(nn.Linear(hid_in_features, num_hiddens),
nn.Tanh())
self.mlm = MaskLM(vocab_size, num_hiddens, mlm_in_features)
self.nsp = NextSentencePred(nsp_in_features)
def forward(self, tokens, segments, valid_lens=None, pred_positions=None):
encoded_X = self.encoder(tokens, segments, valid_lens)
if pred_positions is not None:
mlm_Y_hat = self.mlm(encoded_X, pred_positions)
else:
mlm_Y_hat = None
# The hidden layer of the MLP classifier for next sentence prediction.
# 0 is the index of the '<cls>' token
nsp_Y_hat = self.nsp(self.hidden(encoded_X[:, 0, :]))
return encoded_X, mlm_Y_hat, nsp_Y_hat
# + [markdown] origin_pos=35
# ## 摘要
#
# * Word2vec 和 Glove 等单词嵌入模型与上下文无关。无论单词的上下文如何(如果有),它们都会将相同的预训练向量分配给同一个单词。他们很难以很好地处理自然语言中的多聚结或复杂的语义。
# * 对于上下文相关的单词表示(例如 elMO 和 GPT),单词的表示取决于它们的上下文。
# * elMO 以双向方式对上下文进行编码,但使用特定于任务的架构(但是,为每个自然语言处理任务设计一个特定的架构实际上并不平凡);而 GPT 与任务无关,但是从左到右编码上下文。
# * BERT 结合了两全其美:它以双向方式编码上下文,对于各种自然语言处理任务,只需最少的体系结构更改。
# * BERT 输入序列的嵌入是词元嵌入、区段嵌入和位置嵌入的总和。
# * 培训前 BERT 由两项任务组成:蒙面语言建模和下一句话预测。前者能够对表示单词的双向上下文进行编码,而后者则明确建模文本对之间的逻辑关系。
#
# ## 练习
#
# 1. 为什么 BERT 会成功?
# 1. 所有其他事情都相同,蒙面语言模型是否需要比从左到右语言模型需要更多或更少的预训步骤才能收敛?为什么?
# 1. 在 BERT 的最初实现中,`BERTEncoder`(通过 `d2l.EncoderBlock`)中的定位前馈网络和 `MaskLM` 中的完全连接层都使用高斯误差线性单元 (GELU) :cite:`Hendrycks.Gimpel.2016` 作为激活函数。研究 GELU 和 RELU 之间的区别。
#
# + [markdown] origin_pos=37 tab=["pytorch"]
# [Discussions](https://discuss.d2l.ai/t/1490)
#
| d2l/chapter_natural-language-processing-pretraining/bert.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import jax.numpy as jnp
import matplotlib.pyplot as plt
import jax
from jax import lax
from gym.envs.classic_control import PendulumEnv
from deluca.envs import BalloonLung
from deluca.agents import PID
# + pycharm={"name": "#%%\n"}
def loop(context, x):
env, agent = context
agent_in, agent_out = agent
error = env.observation['target'] - env.observation['measured']
control_in = agent_in(error)
control_out = agent_out(error)
_, reward, _, _ = env.step((control_in, control_out))
return (env, (agent_in, agent_out)), reward
# -
# BalloonLung env
lung = BalloonLung(leak=False,
peep_valve=5.0,
PC=40.0,
P0=0.0,
C=10.0,
R=15.0,
dt=0.03,
waveform=None,
reward_fn=None)
# +
# for loop version
T = 10
xs = jnp.array(jnp.arange(T))
agent_in = PID([3.0, 4.0, 0.0])
agent_out = PID([3.0, 4.0, 0.0])
print(lung.reset())
reward = 0
for i in range(T):
(lung, (agent_in, agent_out)), r = loop((lung, (agent_in, agent_out)), 0)
reward += r
reward_forloop = reward
# scan version
agent_in = PID([3.0, 4.0, 0.0])
agent_out = PID([3.0, 4.0, 0.0])
print(lung.reset())
_,reward_scan = lax.scan(loop, (lung, (agent_in, agent_out)), xs)
# correctness test
print('reward_forloop = ' + str(reward_forloop))
print('reward_scan sum = ' + str(jnp.sum(reward_scan)))
# -
| examples/agents/balloon_lung_pid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Python for Bioinformatics
# -----------------------------
#
# 
#
# This Jupyter notebook is intented to be used alongside the book [Python for Bioinformatics](http://py3.us/)
# Chapter 21: Web Server for Multiple Alignment
# -----------------------------
# This program is not in this notebook because it is a web server. Follow chapter instructions to run it locally or in a web server.
| notebooks/Chapter 21 - Web Server for Multiple Alignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# + vscode={"languageId": "r"}
library(mclust)
library(flexmix)
library(ggplot2)
suppressPackageStartupMessages(library(dendextend))
library(caret)
library(mvtnorm)
library(rsvd)
# -
# # Load and Inspect the Data
# + vscode={"languageId": "r"}
data(banknote)
# + vscode={"languageId": "r"}
head(banknote)
# + vscode={"languageId": "r"}
banknote[sample.int(nrow(banknote), 5), ]
# + vscode={"languageId": "r"}
summary(banknote)
# -
# Apply PCA with two Components to the data to obtain a two-dimensional representation of the Swiss banknotes.
# + vscode={"languageId": "r"}
banknote_pca <- prcomp(banknote[,2:7], scale. = FALSE)
# + vscode={"languageId": "r"}
# first two for 2D
pcs_banknote <- banknote_pca$x[,1:2]
# + vscode={"languageId": "r"}
# Plot the pcs in 2D
plot_data <- data.frame('PC1' = pcs_banknote[,1], 'PC2' = pcs_banknote[,2] , 'Status' = banknote$Status)
ggplot(plot_data, aes(PC1, PC2, color=Status)) +
geom_point() +
theme_minimal() +
labs(title = 'Scatterplot of first two PCs colored by their class')
# -
# We can see that the dataset is well seperated in its first two principal componenents. Hence, it should be easy to cluster them.
# # Agglomerative Hierarchical Clustering
#
# Since Clustering is based on similarity measured e.g. by distance, we introduce functions to compute similarity of binary vectors.
# + vscode={"languageId": "r"}
# functions for binary distances
matching_coefficient <- function(x, y){
stopifnot(length(x) == length(y))
sum(x == y) / length(x)
}
similarity_coefficient <- function(x, y){
stopifnot(length(x) == length(y))
a11 <- sum(x * y)
a10 <- sum(x == 1 & y == 0)
a01 <- sum(x == 0 & y == 1)
a11 / (a11 + a10 + a01)
}
general_similarity_measurement <- function(x, y, alpha = 1, beta=1) {
a11 <- sum(x * y)
a00 <- sum(x + y == 0)
a10 <- sum(x == 1 & y == 0)
a01 <- sum(x == 0 & y == 1)
(a11 + alpha * a00) / (a11 + alpha * a00 + beta*(a10 + a01))
}
# -
# We test the binary measurements with two vectors where each case of match for a single digit is exactly contained once, i.e., for two vectors at position i the following cases can occur: both 0 (a00), both 1(a11), first 0 and second 1 (a01) or the other way around (a10).
# + vscode={"languageId": "r"}
x <- c(1, 0, 0, 1)
y <- c(1, 1, 0, 0)
# distances
matching_coefficient(x, y)
similarity_coefficient(x,y)
general_similarity_measurement(x,y)
# -
# Now, for the hierarchical clustering, we compute the euclidian distance on the first two principal components.
# + vscode={"languageId": "r"}
# compute the distance matrix
dist_banknote <- dist(pcs_banknote)
# + vscode={"languageId": "r"}
cl_single <- hclust(dist_banknote, method = 'single')
cl_complete <- hclust(dist_banknote, method = 'complete')
cl_mean <- hclust(dist_banknote, method = 'average')
cl_ward <- hclust(dist_banknote, method = 'ward.D2')
# + vscode={"languageId": "r"}
par(mfrow = c(2,2))
plot(cl_single)
plot(cl_complete)
plot(cl_mean)
plot(cl_ward)
# + vscode={"languageId": "r"}
# Now color the respective cluster
par(mfrow = c(1,2))
options(repr.plot.width = 7.5, repr.plot.height = 3.5)
plot(color_branches(as.dendrogram(cl_single), k = 3, col= c('red','black', 'green')))
plot(pcs_banknote, col = cutree(cl_single, k=3), pch = 16)
# + vscode={"languageId": "r"}
par(mfrow = c(1,2))
options(repr.plot.width = 7.5, repr.plot.height = 3.5)
plot(color_branches(as.dendrogram(cl_complete), k = 3, col= c('black', 'green','red')))
plot(pcs_banknote, col = cutree(cl_complete, k=3), pch = 16)
# + vscode={"languageId": "r"}
par(mfrow = c(1,2))
options(repr.plot.width = 7.5, repr.plot.height = 3.5)
plot(color_branches(as.dendrogram(cl_mean), k = 4, col=c('green', 'blue', 'red', 'black')))
plot(pcs_banknote, col = cutree(cl_mean, k=4), pch = 16)
# + vscode={"languageId": "r"}
par(mfrow = c(1,2))
options(repr.plot.width = 7.5, repr.plot.height = 3.5)
plot(color_branches(as.dendrogram(cl_ward), k = 2, col = c('red', 'black')))
plot(pcs_banknote, col = cutree(cl_ward, k=2), pch = 16)
# -
# In conclusion, we observe that the clustering with ward's method works best for our data.
# # K-Means
#
# In the following, we apply K-Means clustering to the Swiss banknote dataset. First on the full data and then on the first two principal components only.
# + vscode={"languageId": "r"}
set.seed(2022)
(kmeans_swiss <- kmeans(banknote[, -1], centers = 2, iter.max = 100, nstart = 1, algorithm = "Lloyd"))
# + vscode={"languageId": "r"}
options(repr.plot.width = 10.5, repr.plot.height = 5.5)
pairs(banknote[, -1], col=kmeans_swiss$cluster, pch=16)
# + vscode={"languageId": "r"}
# Accuracy
sum(diag(table(kmeans_swiss$cluster, banknote[, 1]))) / length(banknote[,1])
# + vscode={"languageId": "r"}
confusionMatrix(as.factor(ifelse(kmeans_swiss$cluster == 1, 'counterfeit', 'genuine')), banknote[,1])
# + vscode={"languageId": "r"}
# Repeat for only first two principal diagnoses
set.seed(2022)
(kmeans_swiss <- kmeans(pcs_banknote, centers = 2, iter.max = 100, nstart = 100, algorithm = "Lloyd"))
# + vscode={"languageId": "r"}
plot(pcs_banknote, col= as.factor(kmeans_swiss$cluster), pch=16)
# + vscode={"languageId": "r"}
confusionMatrix(as.factor(ifelse(kmeans_swiss$cluster == 1, 'counterfeit', 'genuine')), banknote[,1])
# -
# For this dataset, K-Means works well on the whole data as well as the first two principal components, since it works well if the cluster are spherical which is the case for the Swiss banknote dataset.
# # Mixture Models and the Expectation Maximization Algorithm
#
# First we will see how K-Means fails on the IRIS dataset, because the three clusters are not clearly distinguishable. Since the EM algorithm leverages an assumption about the data distribution it is able to better distinguish the clusters in IRIS. We assume the specieses to be normal distributed for the standart EM algorithm.
# + vscode={"languageId": "r"}
data(iris)
# + vscode={"languageId": "r"}
set.seed(33)
kmeans_iris <- kmeans(iris[, 1:4], centers = 3, iter.max = 10, nstart = 100)
em_iris <- Mclust(iris[, 1:4], G=3)
# + vscode={"languageId": "r"}
(kmeans_ct <- table(kmeans_iris$cluster, iris$Species))
# + vscode={"languageId": "r"}
# Accuracy of kmeans on Iris
sum(diag(kmeans_ct)) / length(iris$Species)
# + vscode={"languageId": "r"}
(em_ct <- table(em_iris$classification, iris$Species))
# + vscode={"languageId": "r"}
sum(diag(em_ct)) / length(iris$Species) # Much better than Kmeans
# + vscode={"languageId": "r"}
# visual comparison
par(mfrow = c(1,3))
options(repr.plot.width = 10.5, repr.plot.height = 3.5)
plot(iris[, 1:2], col=kmeans_iris$cluster, pch=16, main='K-Means')
plot(iris[, 1:2], col=iris$Species, pch=16, main='Ground Truth')
plot(iris[, 1:2], col=em_iris$classification, pch=16, main='EM')
# -
# On the Iris dataset, a gaussian mixture model is better than K-Means, since two clusters are not differential by a sphere. Now we show how we can use the EM algorithm to assign new data to a known cluster and how to sample from the gaussian mixture model.
# + vscode={"languageId": "r"}
new_iris <- c(5.5, 2.1, 4.0, 0.8)
# + vscode={"languageId": "r"}
# extract the parameters from the EM
prior_probabilities <- em_iris$parameters$pro
cluster_means <- em_iris$parameters$mean
cluster_variances <- em_iris$parameters$variance$sigma
# + vscode={"languageId": "r"}
# Compute the posterior probability
cluster_probability <- numeric(3)
for (i in 1:3) {
cluster_probability[i] <- prior_probabilities[i] * dmvnorm(new_iris, mean = cluster_means[, i], sigma =
cluster_variances[, , i])
}
cluster_probability / sum(cluster_probability)
# + vscode={"languageId": "r"}
# alternatively
predict.Mclust(em_iris, data.frame(X1= new_iris[1], X2= new_iris[2], X3= new_iris[3], X4=new_iris[4]))
# -
# Since the probability for the second EM cluster is the highest we would assign the new observation to the second cluster which corresponds to assuming the species to be "versicolor". Now, we can use the parameters to sample a set with the same distribution than iris.
# + vscode={"languageId": "r"}
set.seed(2022)
# sample first the species from the cluster priors
sample_species <- sample(1:3, size = 150, replace = TRUE, prob = prior_probabilities)
# -
# ## Sampling
# + vscode={"languageId": "r"}
# Now draw random datapoints from the respective cluster
sampled_iris <- sapply(sample_species, function(species){
rmvnorm(1, mean = cluster_means[, species], sigma =
cluster_variances[, , species])
})
sampled_iris <- t(sampled_iris)
# + vscode={"languageId": "r"}
# show the sampled dataset
options(repr.plot.width = 6.5, repr.plot.height = 4.5)
plot(sampled_iris[, 1:2], col = sample_species, pch = 16,
xlab="Sepal Length", ylab='Sepal Width' ,main = "Sampled Iris from EM model")
# -
# ## Bernoulli Mixture Models
#
# Finally, we will apply a bernoilli mixture model to a subset of the MNIST dataset. We start by preparing the data.
# + vscode={"languageId": "r"}
data(digits)
# + vscode={"languageId": "r"}
mnist_labels <- digits[, 1]
mnist <- digits[, -1]
# + vscode={"languageId": "r"}
# function for plotting
plot_digit <- function(row){
digit <- matrix(row, nrow=28, ncol=28)
digit[digit > 0 ] <- 1
image(digit, col=gray(255:0 / 255))
}
# + vscode={"languageId": "r"}
plot_digit(mnist[1,])
# -
# ## Compute the binomial model using the Expectation Maximization Algorithm
# + vscode={"languageId": "r"}
new_digit <- mnist[3500, ] #test case for subsequent demonstration of prediction
mnist <- mnist[-3500,]
# + vscode={"languageId": "r"}
# EM for binomial mixture model
number_cluster <- 4
set.seed(2022)
binomial_em <- flexmix(as.matrix(mnist)~1, k = number_cluster, model=FLXMCmvbinary(), control=list(iter =100))
# + vscode={"languageId": "r"}
binomial_em
# + vscode={"languageId": "r"}
cluster_parameters <- parameters(binomial_em)
df <- data.frame( PixelCol = as.vector(cluster_parameters), Cluster = rep(1:number_cluster, each = 784),
X = rep(rep(1:28, 28),number_cluster), Y = rep(rep(28:1, each = 28), number_cluster))
# We show the cluster that the model estimated
ggplot(df, aes( x= X, y = Y, fill = PixelCol))+
geom_tile()+
scale_fill_gradient(low = "white", high = "black")+
facet_wrap(~ Cluster)
# -
# ## Classify and Sample New Data
# + vscode={"languageId": "r"}
# now classify the new data
plot_digit(new_digit)
# + vscode={"languageId": "r"}
# compute the posterior probabilities for each cluster
cluster_probabilities <- numeric(4)
for (j in 1:4) {
cluster_probabilities[j] <- mean(prior(binomial_em)[j] * dbinom(new_digit, 784, parameters(binomial_em)[, j]))
}
which.max(cluster_probabilities / sum(cluster_probabilities))
# -
# We would assign the new datapoint to the fourth cluster which represents the number 1. Indeed, this is the correct cluster, however, this does not wor in general. Below, we calculate the accuracy for the estimated clusters that the binomial mixture model assigned the datapoints from mnist to.
# + vscode={"languageId": "r"}
estimated_cluster <- binomial_em@cluster
# relabeling
estimated_cluster[estimated_cluster == 3] <- 0
estimated_cluster[estimated_cluster == 1] <- 3
estimated_cluster[estimated_cluster == 4] <- 1
(ct <- table(mnist_labels[-3500], estimated_cluster))
sum(diag(ct)) / length(mnist_labels[-3500])
# -
# Accuracy is ~83% which is not too bad. With the model at hand, we are also able to sample new observations from the clusters. Below, this is demonstrated by sampling from cluster three, hence a zero.
# + vscode={"languageId": "r"}
set.seed(2022)
sample_zero <- rbinom(784, 1, parameters(binomial_em)[,3])
# + vscode={"languageId": "r"}
plot_digit(sample_zero)
| Cluster Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# [](https://colab.research.google.com/github/DJCordhose/ai/blob/master/notebooks/tensorflow/nn-reg.ipynb)
# + colab_type="code" id="XMQp66kq-X9Z" outputId="8d1e32a0-4f5e-477c-9e71-6b8de1998998" colab={"base_uri": "https://localhost:8080/", "height": 34}
import warnings
warnings.filterwarnings('ignore')
# %matplotlib inline
# %pylab inline
import matplotlib.pyplot as plt
import pandas as pd
import tensorflow as tf
from tensorflow import keras
# + colab_type="code" id="hNS5gA2I-X92" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="cff08b3e-5f1c-495f-b0d6-65b897a9dfb9"
# !curl -O https://raw.githubusercontent.com/DJCordhose/deep-learning-crash-course-notebooks/master/data/insurance-customers-1500.csv
# + colab_type="code" id="xpX9iD2c-X94" colab={}
df = pd.read_csv('./insurance-customers-1500.csv', sep=';')
y = df['group']
df.drop('group', axis='columns', inplace=True)
X = df.as_matrix()
# + colab_type="code" id="ivu4Ex_C-X-E" colab={}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42, stratify=y)
# + [markdown] id="14qOwFEovIzw" colab_type="text"
# ### We have several means of regularization
#
# _We use a combination of them:_
# - reduce capacity of model
# - dropout
# - batch normalization
# - change activation to relu for faster training
# - reduce amount of training cycles
# + [markdown] id="h3KaE8S9vKc-" colab_type="text"
# ## Dropout explained in a funny way
#
# <img src='https://raw.githubusercontent.com/DJCordhose/deep-learning-crash-course-notebooks/master/img/dropout-smerty.jpg'>
#
# https://twitter.com/Smerity/status/980175898119778304
# + [markdown] id="upyLOX_3vQRx" colab_type="text"
# ### An experimental approach:
# - keep adding regularization to make validation and train scores come closer to each other
# - this will come at the cost of train scores going down
# - if both values start going down you have gone too far
# - each experiment takes some time
# - for larger datasets and more complex models some people start by overfitting on a subsample of the data (because it trains much faster)
# - then you can be sure you have an architecture that at least has the capacity to solve the problem
# - then keep adding regularizations
# - eventually try using the complete data
# - if you want to use batch normalization place it between raw output of neuron and activation function
# + colab_type="code" id="ODSrdLfF-X-g" outputId="7e4d7a8a-fa71-4cb0-dcf4-d31c5d0e57da" colab={"base_uri": "https://localhost:8080/", "height": 306}
from tensorflow.keras.layers import Dense, Dropout, \
BatchNormalization, Activation
dropout = 0.6
model = keras.Sequential()
# reduce capacity by decreasing number of neurons
model.add(Dense(500, name='hidden1', input_dim=3))
# model.add(BatchNormalization())
model.add(Activation('relu'))
# model.add(Dropout(dropout))
model.add(Dense(500, name='hidden2'))
# model.add(BatchNormalization())
model.add(Activation('relu'))
# model.add(Dropout(dropout))
model.add(Dense(3, name='softmax', activation='softmax'))
model.compile(loss='sparse_categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.summary()
# + colab_type="code" id="A-JeNgyv-X-z" outputId="15468c6a-3df3-44d4-f3ee-6ab9b67e704d" colab={"base_uri": "https://localhost:8080/", "height": 51}
# reducing batch size might increase overfitting,
# but might be necessary to reduce memory requirements
BATCH_SIZE=1000
# reduce this based on what you see in the training history
EPOCHS = 10000
model.compile(loss='sparse_categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# %time history = model.fit(X_train, y_train, epochs=EPOCHS, batch_size=BATCH_SIZE, validation_split=0.2, verbose=0)
# + colab_type="code" id="ekAIERy_-X-5" outputId="fa86e412-16ac-4e8f-e8d0-4b80e5c74c4e" colab={"base_uri": "https://localhost:8080/", "height": 51}
train_loss, train_accuracy = model.evaluate(X_train, y_train, batch_size=BATCH_SIZE)
train_accuracy
# + colab_type="code" id="ZV-LZZGd-X--" outputId="95bce690-f68e-477d-beb9-c08da569153a" colab={"base_uri": "https://localhost:8080/", "height": 335}
def plot_history(history, samples=100, init_phase_samples=None):
epochs = history.params['epochs']
acc = history.history['acc']
val_acc = history.history['val_acc']
every_sample = int(epochs / samples)
acc = pd.DataFrame(acc).iloc[::every_sample, :]
val_acc = pd.DataFrame(val_acc).iloc[::every_sample, :]
fig, ax = plt.subplots(figsize=(20,5))
ax.plot(acc, 'bo', label='Training acc')
ax.plot(val_acc, 'b', label='Validation acc')
ax.set_title('Training and validation accuracy')
ax.legend()
plot_history(history)
# + id="RpJPY6rV8l53" colab_type="code" outputId="831a2f5a-34ff-4d8a-a37c-ca005644c726" colab={"base_uri": "https://localhost:8080/", "height": 34}
model.predict(np.array([[100, 47, 10]]))
# + id="NAdBi_a18l5_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="9876d56a-4e05-4817-a31a-088ec8232c12"
test_loss, test_accuracy = model.evaluate(X_test, y_test, batch_size=BATCH_SIZE)
test_accuracy
# + id="exbDxJ5xwe-u" colab_type="code" colab={}
| notebooks/tensorflow/nn-reg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="TKq7cP4Vx8TH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fd91a8da-17dc-4610-b82d-4d1aa86dc998" executionInfo={"status": "ok", "timestamp": 1576652334697, "user_tz": 300, "elapsed": 5753, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBOMYelIpWRDXKf5OHRCPHy4uvNYyEHD-uQofvOmw=s64", "userId": "05057968350501293125"}}
from zipfile import ZipFile
file_name = 'blood-cells-100-test.zip'
with ZipFile(file_name, 'r') as zip:
zip.extractall()
print("Done :)")
# + id="mE6aq3w_zVmR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fe9d2b73-1441-4c7c-dcb0-d0b453d57974" executionInfo={"status": "ok", "timestamp": 1576652340916, "user_tz": 300, "elapsed": 855, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBOMYelIpWRDXKf5OHRCPHy4uvNYyEHD-uQofvOmw=s64", "userId": "05057968350501293125"}}
# %cd blood-cells/
# + id="RmsEvdDYzXsZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 751} outputId="f4383491-a134-4e6b-a7ee-0f3daa21ebb9" executionInfo={"status": "ok", "timestamp": 1576653269889, "user_tz": 300, "elapsed": 920626, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mBOMYelIpWRDXKf5OHRCPHy4uvNYyEHD-uQofvOmw=s64", "userId": "05057968350501293125"}}
# !python3 main.py
# + id="UuRN35VrzZ8M" colab_type="code" colab={}
| test-100.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pycon Africa - Developing Maltego Transform Using Python
# ### By <NAME> - @marengz
# # Maltego Data Flow
#
# 
# # Maltego Request Example
#
# 
#
#
# # Maltego Response Example
#
# 
#
#
# ## Maltego-TRX Reference - [Link](https://github.com/paterva/maltego-trx/blob/master/README.md)
# ### Constants
# The following constants can be imported from `maltego_trx.maltego`.
#
# **Message Types:**
# - `UIM_FATAL`
# - `UIM_PARTIAL`
# - `UIM_INFORM`
# - `UIM_DEBUG`
#
# **Bookmark Colors:**
# - `BOOKMARK_COLOR_NONE`
# - `BOOKMARK_COLOR_BLUE`
# - `BOOKMARK_COLOR_GREEN`
# - `BOOKMARK_COLOR_YELLOW`
# - `BOOKMARK_COLOR_PURPLE`
# - `BOOKMARK_COLOR_RED`
#
# **Link Styles:**
# - `LINK_STYLE_NORMAL`
# - `LINK_STYLE_DASHED`
# - `LINK_STYLE_DOTTED`
# - `LINK_STYLE_DASHDOT`
#
#
#
# ### Request/MaltegoMsg
# The request/maltego msg object given to the transform contains the information about the input entity.
#
# **Attributes:**
# - `Value: str`: The display value of the input entity on the graph
# - `Weight: int`: The weight of the input entity
# - `Slider: int`: Results slider setting in the client
# - `Type: str`: The input entity type
# - `Properties: dict(str: str)`: A key-value dictionary of the input entity properties
# - `TransformSettings: dict(str: str)`: A key-value dictionary of the transform settings
#
# **Methods:**
# - `getProperty(name: str)`: get a property value of the input entity
# - `getTransformSetting(name: str)`: get a transform setting value
#
# ### Response/MaltegoTransform
#
# **Methods:**
# - `addEntity(type: str, value: str) -> Entity`: Add an entity to the transform response. Returns an Entity object created by the method.
# - `addUIMessagte(msg: str, messageType='Inform')`: Return a UI message to the user. For message type, use a message type constant.
#
# ### Entity
#
# **Methods:**
# - `setType(type: str)`: Set the entity type (e.g. "Phrase" for maltego.Phrase entity)
# - `setValue(value: str)`: Set the entity value
# - `setWeight(weight: int)`: Set the entity weight
# - `addDisplayInformation(content: str, title: str)`: Add display information for the entity.
# - `addProperty(fieldName: str, displayName: str, matchingRule: str, value: str)`: Add a property to the entity. Matching rule can be `strict` or `loose`.
# - `setIconURL(url: str)`: Set the entity icon URL
# - `setBookmark(bookmark: int)`: Set bookmark color index (e.g. -1 for BOOKMARK_COLOR_NONE, 3 for BOOKMARK_COLOR_PURPLE)
# - `setNote(note: str)`: Set note content
#
# **Link Methods:**
# - `setLinkColor(color: str)`: Set the link color (e.g. hex "#0000FF" for blue)
# - `setLinkStyle(style: int)`: Set the link style index (e.g. 0 for LINK_STYLE_NORMAL, 2 for LINK_STYLE_DOTTED)
# - `setLinkThickness(thick: int)`: Set link thickness (default is 1)
# - `setLinkLabel(label: str)`: Set the label of the link
# - `reverseLink()`: Reverse the link direction
#
| Getting Started.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Longitudinal Car Model
#
# > "Overview of longitudinal vehicle dynamics"
#
# - toc:true
# - branch: master
# - badges: false
# - comments: false
# - author: <NAME>
# - categories: [longitudinal-dynamics, autonomous-vehicles, mathematical-modelling, vehicle-dynamics]
# ## Longitudinal car model
# In this section, we will go over the concept of the <a href="https://www.sciencedirect.com/topics/engineering/longitudinal-dynamic">vehicle longitudinal dynamics</a>. The two major elements of the longitudinal vehicle model discussed in this section are
# - Vehicle dynamics
# - Powertrain dynamics
# The vehicle dynamics are influenced by longitudinal tire forces, aerodynamic drag forces, rolling
# resistance forces and gravitational forces. The longitudinal powertrain system of the vehicle
# consists of the internal combustion engine, the torque converter, the
# transmission and the wheels. This <a href="https://www.coursera.org/lecture/intro-self-driving-cars/lesson-4-longitudinal-vehicle-modeling-V8htX"> video</a> explains nicely the concepts.
# The longitudinal vehicle dynamic model is simply based on the dynamics of
# the vehicle that generate forward motion. The following figure shows a typical vehicle
# longitudinal motion force diagram on an inclined road.
# 
# *Figure 1. Schematics of vehicle logitudinal model on an inclined road. Image from [1].*
#
# We have the followig forces acting on the vehicle
# - The front tire forces $F_{xf}$
# - The rear tire forces $F_{xr}$
# - The <a href="https://www.sciencelearn.org.nz/resources/1346-causes-of-aerodynamic-drag">aerodynamic drag force</a> $F_{aero}$
# - The rolling resistance forces $R_{xf}$ and $R_{xr}$
# - The force due to gravity $F_g$
# According to <a href="https://en.wikipedia.org/wiki/Newton%27s_laws_of_motion">Newton’s laws of motion</a>, and in particular the second law, the longitudinal tire forces of the front and
# rear tyres, $F_{xf}$ and $F_{xr}$, should balance the resistance forces $F_{aero}$, the gravitational
# force $F_g$ , and the rolling resistance of the front and rear tires, $R_{xf}$ and $R_{xr}$.
# Any imbalance between these forces creates an acceleration of the vehicle in
# the longitudinal direction denoted by $\ddot{x}$. Thus, the basic logintudinal motion model is given by
# $$m\ddot{x} = F_{xf} + F_{xr} - F_{aero} - F_g - R_{xf} - R_{xr}$$
# where $m$ is the mass of the vehicle. The forces $F_{xf}$ and $F_{xr}$ come from the vehicle power train. We can express them collectively as $F_x$. Furthermore, we group together the rolling resistance forces under the symbol $R_x$. Thus, the reduced model is
# $$m\ddot{x} = F_x - F_{aero} - F_g - R_x $$
# We will need a way to express the involved quantities in order to be able to solve for $\ddot{x}$. Let's start with the gravitational force $F_g$.
# ### Gravitational froce
# We can express $F_g$ as [2]
# $$F_g = mg sin (\alpha)$$
# where $\alpha$ is the local road slope. For small slope angles, we can write
# $$sin (\alpha) \approx \alpha$$
# ### Aerodynamic drag
# A vehicles longitudinal motion is resisted by aerodynamic drag rolling resistance
# and the force due to gravity. The aerodynamic drag force $F_{aero}$ is typically
# modeled as dependent on air density $\rho$, frontal area of the vehicle $A$, the vehicles
# coefficient of friction $C_D$, and the current speed of the vehicle. The functional relationship of all these quantities is given in the equation beow
# $$F_{aero} = \frac{1}{2}C_D\rho A v^2$$
# ### Rolling resistance
# Tires are elastic materials that are subject to deformation in the patch which is in contact with the road surface. Let's neglect the the deformation of the road.
# The tire is subject to a normal load. Due to this load, the tire material will be deflected normally at the contact patch and then regaining its shape whilst leaving the patch neighborhood. However, internal damping of the material does not allow the energy lost during deforming the tire to be completely recovered when the material
# returns to its original shape [1]. It appears therefore, that some loss of energy occurs. This loss is represented by a force on the tires called the rolling resistance that acts in the opposite direction of the motion of the vehicle.
# Hence, the rolling resistance depends on the normal tire load, tire pressure and vehicle speed. A model is given below [1],
# $$R_x = N(c_{r, 0} + c_{r,1}|\dot{x}| + c_{r,2}|\dot{x}|^2)$$
# see also [2] for further modelling. If we assume nominal operating conditions and drop the second-order terms for simplicity, we can arrive at a linear rolling resistance model, where $c_{r,1}$ is the linear rolling resistance coefficient.
# $$R_x \approx c_{r,1}|\dot{x}|$$
# ### Tire forces
# We now discuss the longitudinal tire forces expressed under the term $F_x$. Longitudinal tire forces depend on the following factors [2]
# - Slip ratio
# - Normal load on the tires
# - Friction coefficient on the tire road interface
# Let's see these components
# #### Slip ratio
# For an effective wheel radius $R_{effective}$ and a wheel velocity $\omega_w$ the velocity is described by
# $$V_{wheel} = R_{effective}\omega_{wheel}$$
# However, the actual longitudinal velocity at the axle of the wheel, $V_x$ may be different than that. This is called longitudinal slip [2]. In other words, the longitudinal slip is defined as [2]
# $$\sigma = V_{wheel} - V_x$$
# Moreover, we define the longitudinal slip ratio during braking and acceleration as [2]
# $$\sigma_{xf} = \begin{cases} \frac{R_{effecive}\omega_{wf} - V_x}{V_{x}}, ~~\text{during breaking} \\ \frac{R_{effecive}\omega_{wf} - V_x}{R_{effecive}\omega_{wf}}, ~~\text{during acceleration} \end{cases}$$
# We have a similar expression for the rear wheels. Given the slip coefficients, we can express the longitudinal tire forces as
# $$F_{xf} = C_{\sigma f}\sigma_{xf}, ~~ F_{xr} = C_{\sigma r}\sigma_{xr}$$
# where $C_{\sigma f}$ and $C_{\sigma r}$ are called the longitudinal tire stiffness parameters of the front and rear tires respectively [2].
# ### Powertrain forces
# The longitudinal tire forces, denoted collectivelly above with $F_x$, acting on the driving wheels are the main forces that drive the vehicle forward [2]. These forces depend on the difference between the rotational wheel velocity $R_{effective}\omega_{w}$ and the vehicle longitudinal velocity $\dot{x}$. In particular, we saw that we can model the longitudinal tire forces as
# $$F_{xf} = C_{\sigma f}\sigma_{xf}, ~~ F_{xr} = C_{\sigma r}\sigma_{xr}$$
# where $C_{\sigma f}$ and $C_{\sigma r}$ are called the longitudinal tire stiffness parameters of the front and rear tires respectively [2]. However, $\omega_w$ is highly influence by the powertrain dynamics of the vehicle. The powertrain has the following major components [2]
# - Engine
# - Transmission or gearbox
# - Torque converter or clutch
# - Differential
# - Wheels
# 
# *Figure 2. Powertrain schematics. Image from [1].*
#
#
# Let's see each of the components separately
# ### Torque converter
# The torque cnverter connects the engine to the transmission. When the engine is turning slowly, e.g. when the car waits at a stoplight, the amount of torque passed through the torque converter is very small. Thus, maintaining the the car stopped requires only a light pressure on the brake pedal. Hence, we don't have to stall the engine in order to maintain the vehicle stopped. In contrast, when the vehicle accelerates the torque converter gives the car more torque [2].
# The torque converter has the following major components [2]
# - pump
# - turbine
# - transmission fluid
# The pump turns at the same speed as the engine whilst the turbine is connected to the
# transmission and causes the transmission to spin at the same speed as the
# turbine [2] This is what basically moves the vehicle. The coupling between the turbine and the pump is through the transmission fluid. Torque is transmitted from the
# pump to the turbine of the torque converter [2].
# Various models have been introduced to model the pump torque $T_{pump}$ and the turbine torque $T_{turbine}$ see [2 page 103].
# #### Transmission dynamics
# Let's denote with $GR$ the gear ratio of the transmission. In general, $GR < 1$ and increases as the gear shifts upwards. The input to the transmission module is the torbine torque $T_{turbine}$ [2]. The torque transmitted to the wheels is $T_{wheels}$. Then, at steady state, this torque is given by
# $$ T_{wheels} = \frac{1}{GR} T_{turbine}$$
# Furthermore, we have the following relaton between the transmission and the wheel speed [2]
# $$ \omega_{transmission} = \frac{1}{GR} \omega_{wheels}$$
# Note that these equations cannot be used during gear change. See [2 page 105] for a model based on first order equations.
# ### Engine dynamics
# A simplified engine dynamic model is
# $$J_{engine} \dot{\omega}_e = T_{engine} - T_{pump}$$
# In general, the engine torque $T_{engine}$ depends on the dynamics in the intake and
# exhaust manifold of the engine and on the accelerator input from the driver [2]. $T_{pump}$ is the torque from the pump is the load of the engine from the torque converter [2].
# ### Wheel dynamics
# The driving wheels rotational dynamics, e.g. for the rear wheels in a rear wheel driven vehicle, are dictated by [2]
# $$J_{wheel} \dot{\omega}_{wheel, r} = T_{wheel} - R_{effective}F_{xr}$$
# For the non-driven wheels the torque term is zero.
# ## Refernces
# 1. <a href="https://www.coursera.org/lecture/intro-self-driving-cars/lesson-4-longitudinal-vehicle-modeling-V8htX">Lesson 4: Longitudinal Vehicle Modeling</a>
# 2. <NAME>. ```Longitudinal Vehicle Dynamics. In: Vehicle Dynamics and Control.```, Mechanical Engineering Series. Springer 2012.
| _notebooks/2021-04-10-longitudinal-vehicle-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # In this notebook we will read in the model that was already trained by some other notebook.
# ## We will use that model to predict new results and produce corresponding plots.
# ### Standard imports
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import IPython
import pandas as pd
# ### Handy function to know how much RAM the notebook is taking
# +
#memory usage and release
#https://stackoverflow.com/questions/39100971/how-do-i-release-memory-used-by-a-pandas-dataframe
import os, psutil
def usage():
process = psutil.Process(os.getpid())
return process.memory_info()[0] / float(2 ** 20)
# -
usage()
# ### Read in the files of interest. I am using 'npy' format, which is great for ML. Other formats can be used. I am loading many files, but you need just one - the file to be used for prediction.
# +
x_train = np.load('x_train.npy')
y_train = np.load('y_train.npy')
x_test = np.load('x_test.npy')
y_test = np.load('y_test.npy')
x_val = np.load('x_val.npy')
y_val = np.load('y_val.npy')
# -
# ### Define loss functions used during the training.
# +
class HuberLoss(object):
def __init__(self,delta=1.):
self.delta = delta
self.n_params = 1
self.__name__ = 'HuberLoss'
def __call__(self,y_true,y_pred):
z = K.abs(y_true[:,0] - y_pred[:,0])
mask = K.cast(K.less(z,self.delta),K.floatx())
return K.mean( 0.5*mask*K.square(z) + (1.-mask)*(self.delta*z - 0.5*self.delta**2) )
# ---------------------------------------------------------------------------------------------------
class QuantileLoss(object):
def __init__(self,taus=[0.5,0.25,0.75],weights=[1.,1.2,0.9]):
self.taus = np.array(taus).reshape(1,-1)
self.weights = np.array(weights).reshape(1,-1)
self.n_params = len(taus)
self.__name__ = 'QuantileLoss'
def __call__(self,y_true,y_pred):
e = y_true - y_pred
print(e.shape)
return K.mean( self.weights*( self.taus*e + K.clip( -e, K.epsilon(), np.inf ) ) )
# ---------------------------------------------------------------------------------------------------
class HybridLoss(object):
def __init__(self,delta=1.,taus=[0.25,0.75],weights=[1.]):
self.__name__ = 'HybridLoss'
self.huber = HuberLoss(delta)
self.quantiles = QuantileLoss(taus,weights)
self.n_params = self.huber.n_params + self.quantiles.n_params
def __call__(self,y_true,y_pred):
return self.huber(y_true,y_pred) + self.quantiles(y_true,y_pred[:,1:])
# -
# ### Additional functions just for compliteness, can be skipped.
# +
# ---------------------------------------------------------------------------------------------------
def gauss_nll(y_true,y_pred):
mean = y_pred[:,0]
sigma2 = K.square(y_pred[:,0]) + 0.01**2
z2 = K.square( y_true[:,0] - mean) / sigma2
return 0.5*K.mean( z2 + 0.5*K.log(sigma2), axis=-1 )
## return K.mean(K.square(y_true-y_pred))
# ---------------------------------------------------------------------------------------------------
def mse0(y_true,y_pred):
return K.mean( K.square(y_true[:,0] - y_pred[:,0]) )
# ---------------------------------------------------------------------------------------------------
def mae0(y_true,y_pred):
return K.mean( K.abs(y_true[:,0] - y_pred[:,0]) )
# ---------------------------------------------------------------------------------------------------
def r2_score0(y_true,y_pred):
return 1. - K.sum( K.square(y_true[:,0] - y_pred[:,0]) ) / K.sum( K.square(y_true[:,0] - K.mean(y_true[:,0]) ) )
# -
# ### Create the function object to be passed to the Keras 'load_model', since we used a custom Loss function.
quantiles_and_Huber_Loss = HybridLoss()
# ### Import to load the already trained model, printed dimensions will be (?,2)
# +
from keras.models import load_model
import h5py
from keras import backend as K
model = load_model('my_model_4.h5', custom_objects={'HybridLoss': quantiles_and_Huber_Loss})
# -
# ### Using test data set, make prediction with our model.
y_pred = model.predict(x_test)
y_pred
# ### Plot the estimate (ratio) and other important ratios of interest.
# +
plt.title("Electron Emc/Emeasured")
plot_name = 'results.pdf'
# add extra 0 if want a finer binning.
bin_size = 0.001;
bins = 20000
min_edge = 0.95; max_edge = 1.15
N = (max_edge-min_edge)/bin_size; Nplus1 = N + 1
bin_list = np.linspace(min_edge, max_edge, Nplus1)
bins= bin_list
plt.grid(axis='y', alpha=0.75)
plt.grid(axis='x', alpha=0.75)
plt.xlim((min_edge, max_edge)) # adjust the top leaving bottom unchanged
# The convention is that 0th element is the main estimate, and 1th and 2nd elements in
# the y_pred array(array of 3 elements) are 25 and 75% quantiles correspondingly.
# To plot the estimate, use '0:1', for 25% quantile '1:2', for 75% quantile use '2:3'.
plt.hist(y_pred[:,0:1], bins, alpha=0.7, label='y_pred')
plt.hist(y_test, bins, alpha=0.7, label='y_test')
plt.rcParams["figure.figsize"] = [15,15]
plt.legend(prop={'size': 18}, loc='upper right')
#plot.legend(loc=2, prop={'size': 6})
plt.xlabel("Emc/Emeasured", fontsize=18)
plt.ylabel('Events', fontsize=18)#plt.show()
# May need to uncomment lines below if the picture is small.
#fig = plt.figure( figsize=(20, 20))
#fig.set_figheight(20)
#fig.set_figwidth(30)
plt.savefig(plot_name, dpi=400)
# -
# ### Plot the estimate and quantiles.
# +
plt.title("Electron Emc/Emeasured")
plot_name = 'Estimate_and_quantiles.pdf'
# add extra 0 if want a finer binning.
bin_size = 0.001;
bins = 20000
min_edge = 0.95; max_edge = 1.15
N = (max_edge-min_edge)/bin_size; Nplus1 = N + 1
bin_list = np.linspace(min_edge, max_edge, Nplus1)
bins= bin_list
plt.grid(axis='y', alpha=0.75)
plt.grid(axis='x', alpha=0.75)
plt.xlim((min_edge, max_edge)) # adjust the top leaving bottom unchanged
# The convention is that 0th element is the main estimate, and 1th and 2nd elements in
# the y_pred array(array of 3 elements) are 25 and 75% quantiles correspondingly.
# To plot the estimate, use '0:1', for 25% quantile '1:2', for 75% quantile use '2:3'.
plt.hist(y_pred[:,0:1], bins, alpha=0.7, label='y_pred')
plt.hist(y_pred[:,1:2], bins, alpha=0.7, label='y_pred_25')
plt.hist(y_pred[:,2:3], bins, alpha=0.7, label='y_pred_75')
plt.rcParams["figure.figsize"] = [15,15]
plt.legend(prop={'size': 18}, loc='upper right')
#plot.legend(loc=2, prop={'size': 6})
plt.xlabel("Emc/Emeasured", fontsize=18)
plt.ylabel('Events', fontsize=18)#plt.show()
# May need to uncomment lines below if the picture is small.
#fig = plt.figure( figsize=(20, 20))
#fig.set_figheight(20)
#fig.set_figwidth(30)
plt.savefig(plot_name, dpi=400)
# -
| notebooks/Egamma_5_ReadModelCheckResults.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="nibpbUnTsxTd"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" colab_type="code" id="tXAbWHtqs1Y2" colab={}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="HTgMAvQq-PU_"
# # Ragged tensors
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/guide/ragged_tensor"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/ragged_tensor.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/ragged_tensor.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/ragged_tensor.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="cDIUjj07-rQg"
# ## Setup
# + colab_type="code" id="KKvdSorS-pDD" colab={}
# !pip install -q tf_nightly
import math
import tensorflow as tf
# + [markdown] colab_type="text" id="pxi0m_yf-te5"
# ## Overview
#
# Your data comes in many shapes; your tensors should too.
# *Ragged tensors* are the TensorFlow equivalent of nested variable-length
# lists. They make it easy to store and process data with non-uniform shapes,
# including:
#
# * Variable-length features, such as the set of actors in a movie.
# * Batches of variable-length sequential inputs, such as sentences or video
# clips.
# * Hierarchical inputs, such as text documents that are subdivided into
# sections, paragraphs, sentences, and words.
# * Individual fields in structured inputs, such as protocol buffers.
#
# + [markdown] colab_type="text" id="1mhU_qY3_mla"
# ### What you can do with a ragged tensor
#
# Ragged tensors are supported by more than a hundred TensorFlow operations,
# including math operations (such as `tf.add` and `tf.reduce_mean`), array operations
# (such as `tf.concat` and `tf.tile`), string manipulation ops (such as
# `tf.substr`), control flow operations (such as `tf.while_loop` and `tf.map_fn`), and many others:
# + colab_type="code" id="vGmJGSf_-PVB" colab={}
digits = tf.ragged.constant([[3, 1, 4, 1], [], [5, 9, 2], [6], []])
words = tf.ragged.constant([["So", "long"], ["thanks", "for", "all", "the", "fish"]])
print(tf.add(digits, 3))
print(tf.reduce_mean(digits, axis=1))
print(tf.concat([digits, [[5, 3]]], axis=0))
print(tf.tile(digits, [1, 2]))
print(tf.strings.substr(words, 0, 2))
print(tf.map_fn(tf.math.square, digits))
# + [markdown] colab_type="text" id="Pt-5OIc8-PVG"
# There are also a number of methods and operations that are
# specific to ragged tensors, including factory methods, conversion methods,
# and value-mapping operations.
# For a list of supported ops, see the **`tf.ragged` package
# documentation**.
# + [markdown] colab_type="text" id="r8fjGgf3B_6z"
# Ragged tensors are supported by many TensorFlow APIs, including [Keras](https://www.tensorflow.org/guide/keras), [Datasets](https://www.tensorflow.org/guide/data), [tf.function](https://www.tensorflow.org/guide/function), [SavedModels](https://www.tensorflow.org/guide/saved_model), and [tf.Example](https://www.tensorflow.org/tutorials/load_data/tfrecord). For more information, see the section on **TensorFlow APIs** below.
# + [markdown] colab_type="text" id="aTXLjQlcHP8a"
# As with normal tensors, you can use Python-style indexing to access specific
# slices of a ragged tensor. For more information, see the section on
# **Indexing** below.
# + colab_type="code" id="n8YMKXpI-PVH" colab={}
print(digits[0]) # First row
# + colab_type="code" id="Awi8i9q5_DuX" colab={}
print(digits[:, :2]) # First two values in each row.
# + colab_type="code" id="sXgQtTcgHHMR" colab={}
print(digits[:, -2:]) # Last two values in each row.
# + [markdown] colab_type="text" id="6FU5T_-8-PVK"
# And just like normal tensors, you can use Python arithmetic and comparison
# operators to perform elementwise operations. For more information, see the section on
# **Overloaded Operators** below.
# + colab_type="code" id="2tdUEtb7-PVL" colab={}
print(digits + 3)
# + colab_type="code" id="X-bxG0nc_Nmf" colab={}
print(digits + tf.ragged.constant([[1, 2, 3, 4], [], [5, 6, 7], [8], []]))
# + [markdown] colab_type="text" id="2tsw8mN0ESIT"
# If you need to perform an elementwise transformation to the values of a `RaggedTensor`, you can use `tf.ragged.map_flat_values`, which takes a function plus one or more arguments, and applies the function to transform the `RaggedTensor`'s values.
# + colab_type="code" id="pvt5URbdEt-D" colab={}
times_two_plus_one = lambda x: x * 2 + 1
print(tf.ragged.map_flat_values(times_two_plus_one, digits))
# + [markdown] colab_type="text" id="HNxF6_QKAzkl"
# Ragged tensors can be converted to nested Python `list`s and numpy `array`s:
# + colab_type="code" id="A5NHb8ViA9dt" colab={}
digits.to_list()
# + colab_type="code" id="2o1wogVyA6Yp" colab={}
digits.numpy()
# + [markdown] colab_type="text" id="7M5RHOgp-PVN"
# ### Constructing a ragged tensor
#
# The simplest way to construct a ragged tensor is using
# `tf.ragged.constant`, which builds the
# `RaggedTensor` corresponding to a given nested Python `list` or numpy `array`:
# + colab_type="code" id="yhgKMozw-PVP" colab={}
sentences = tf.ragged.constant([
["Let's", "build", "some", "ragged", "tensors", "!"],
["We", "can", "use", "tf.ragged.constant", "."]])
print(sentences)
# + colab_type="code" id="TW1g7eE2ee8M" colab={}
paragraphs = tf.ragged.constant([
[['I', 'have', 'a', 'cat'], ['His', 'name', 'is', 'Mat']],
[['Do', 'you', 'want', 'to', 'come', 'visit'], ["I'm", 'free', 'tomorrow']],
])
print(paragraphs)
# + [markdown] colab_type="text" id="SPLn5xHn-PVR"
# Ragged tensors can also be constructed by pairing flat *values* tensors with
# *row-partitioning* tensors indicating how those values should be divided into
# rows, using factory classmethods such as `tf.RaggedTensor.from_value_rowids`,
# `tf.RaggedTensor.from_row_lengths`, and
# `tf.RaggedTensor.from_row_splits`.
#
# #### `tf.RaggedTensor.from_value_rowids`
# If you know which row each value belongs in, then you can build a `RaggedTensor` using a `value_rowids` row-partitioning tensor:
#
# 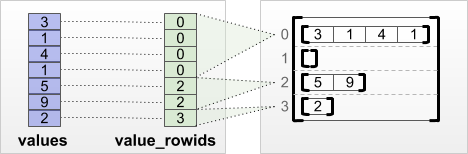
# + colab_type="code" id="SEvcPUcl-PVS" colab={}
print(tf.RaggedTensor.from_value_rowids(
values=[3, 1, 4, 1, 5, 9, 2],
value_rowids=[0, 0, 0, 0, 2, 2, 3]))
# + [markdown] colab_type="text" id="RBQh8sYc-PVV"
# #### `tf.RaggedTensor.from_row_lengths`
#
# If you know how long each row is, then you can use a `row_lengths` row-partitioning tensor:
#
# 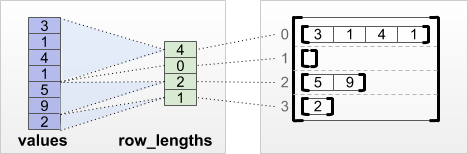
# + colab_type="code" id="LBY81WXl-PVW" colab={}
print(tf.RaggedTensor.from_row_lengths(
values=[3, 1, 4, 1, 5, 9, 2],
row_lengths=[4, 0, 2, 1]))
# + [markdown] colab_type="text" id="8p5V8_Iu-PVa"
# #### `tf.RaggedTensor.from_row_splits`
#
# If you know the index where each row starts and ends, then you can use a `row_splits` row-partitioning tensor:
#
# 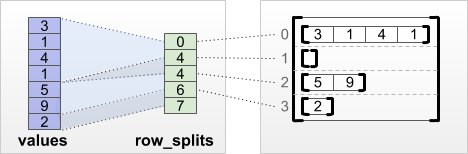
# + colab_type="code" id="FwizuqZI-PVb" colab={}
print(tf.RaggedTensor.from_row_splits(
values=[3, 1, 4, 1, 5, 9, 2],
row_splits=[0, 4, 4, 6, 7]))
# + [markdown] colab_type="text" id="E-9imo8DhwuA"
# See the `tf.RaggedTensor` class documentation for a full list of factory methods.
#
# Note: By default, these factory methods add assertions that the row partition tensor is well-formed and consistent with the number of values. The `validate=False` parameter can be used to skip these checks if you can guarantee that the inputs are well-formed and consistent.
# + [markdown] colab_type="text" id="YQAOsT1_-PVg"
# ### What you can store in a ragged tensor
#
# As with normal `Tensor`s, the values in a `RaggedTensor` must all have the same
# type; and the values must all be at the same nesting depth (the *rank* of the
# tensor):
# + colab_type="code" id="SqbPBd_w-PVi" colab={}
print(tf.ragged.constant([["Hi"], ["How", "are", "you"]])) # ok: type=string, rank=2
# + colab_type="code" id="83ZCSJnQAWAf" colab={}
print(tf.ragged.constant([[[1, 2], [3]], [[4, 5]]])) # ok: type=int32, rank=3
# + colab_type="code" id="ewA3cISdDfmP" colab={}
try:
tf.ragged.constant([["one", "two"], [3, 4]]) # bad: multiple types
except ValueError as exception:
print(exception)
# + colab_type="code" id="EOWIlVidDl-n" colab={}
try:
tf.ragged.constant(["A", ["B", "C"]]) # bad: multiple nesting depths
except ValueError as exception:
print(exception)
# + [markdown] colab_type="text" id="nhHMFhSp-PVq"
# ## Example use case
#
# The following example demonstrates how `RaggedTensor`s can be used to construct
# and combine unigram and bigram embeddings for a batch of variable-length
# queries, using special markers for the beginning and end of each sentence.
# For more details on the ops used in this example, see the `tf.ragged` package documentation.
# + colab_type="code" id="ZBs_V7e--PVr" colab={}
queries = tf.ragged.constant([['Who', 'is', 'Dan', 'Smith'],
['Pause'],
['Will', 'it', 'rain', 'later', 'today']])
# Create an embedding table.
num_buckets = 1024
embedding_size = 4
embedding_table = tf.Variable(
tf.random.truncated_normal([num_buckets, embedding_size],
stddev=1.0 / math.sqrt(embedding_size)))
# Look up the embedding for each word.
word_buckets = tf.strings.to_hash_bucket_fast(queries, num_buckets)
word_embeddings = tf.nn.embedding_lookup(embedding_table, word_buckets) # ①
# Add markers to the beginning and end of each sentence.
marker = tf.fill([queries.nrows(), 1], '#')
padded = tf.concat([marker, queries, marker], axis=1) # ②
# Build word bigrams & look up embeddings.
bigrams = tf.strings.join([padded[:, :-1], padded[:, 1:]], separator='+') # ③
bigram_buckets = tf.strings.to_hash_bucket_fast(bigrams, num_buckets)
bigram_embeddings = tf.nn.embedding_lookup(embedding_table, bigram_buckets) # ④
# Find the average embedding for each sentence
all_embeddings = tf.concat([word_embeddings, bigram_embeddings], axis=1) # ⑤
avg_embedding = tf.reduce_mean(all_embeddings, axis=1) # ⑥
print(avg_embedding)
# + [markdown] colab_type="text" id="Y_lE_LAVcWQH"
# 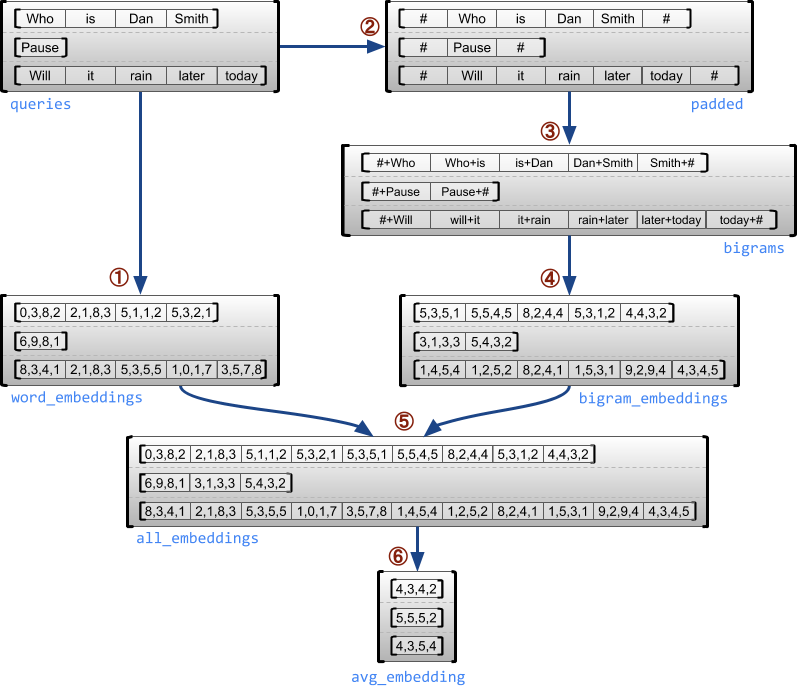
# + [markdown] colab_type="text" id="An_k0pX1-PVt"
# ## Ragged and uniform dimensions
#
# A ***ragged dimension*** is a dimension whose slices may have different lengths. For example, the
# inner (column) dimension of `rt=[[3, 1, 4, 1], [], [5, 9, 2], [6], []]` is
# ragged, since the column slices (`rt[0, :]`, ..., `rt[4, :]`) have different
# lengths. Dimensions whose slices all have the same length are called *uniform
# dimensions*.
#
# The outermost dimension of a ragged tensor is always uniform, since it consists
# of a single slice (and so there is no possibility for differing slice
# lengths). The remaining dimensions may be either ragged or uniform. For
# example, we might store the word embeddings for
# each word in a batch of sentences using a ragged tensor with shape
# `[num_sentences, (num_words), embedding_size]`, where the parentheses around
# `(num_words)` indicate that the dimension is ragged.
#
# 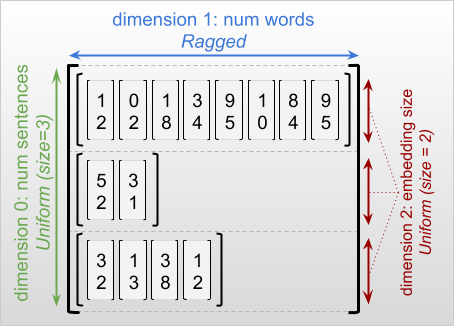
#
# Ragged tensors may have multiple ragged dimensions. For example, we could store
# a batch of structured text documents using a tensor with shape `[num_documents,
# (num_paragraphs), (num_sentences), (num_words)]` (where again parentheses are
# used to indicate ragged dimensions).
#
# As with `tf.Tensor`, the ***rank*** of a ragged tensor is its total number of dimensions (including both ragged and uniform dimensions).
# A ***potentially ragged tensor*** is a value that might be
# either a `tf.Tensor` or a `tf.RaggedTensor`.
#
# When describing the shape of a RaggedTensor, ragged dimensions are conventionally indicated by
# enclosing them in parentheses. For example, as we saw above, the shape of a 3-D
# RaggedTensor that stores word embeddings for each word in a batch of sentences
# can be written as `[num_sentences, (num_words), embedding_size]`.
#
# The `RaggedTensor.shape` attribute returns a `tf.TensorShape` for a
# ragged tensor, where ragged dimensions have size `None`:
#
# + colab_type="code" id="M2Wzx4JEIvmb" colab={}
tf.ragged.constant([["Hi"], ["How", "are", "you"]]).shape
# + [markdown] colab_type="text" id="G9tfJOeFlijE"
# The method `tf.RaggedTensor.bounding_shape` can be used to find a tight
# bounding shape for a given `RaggedTensor`:
# + colab_type="code" id="5DHaqXHxlWi0" colab={}
print(tf.ragged.constant([["Hi"], ["How", "are", "you"]]).bounding_shape())
# + [markdown] colab_type="text" id="V8e7x95UcLS6"
# ## Ragged vs. sparse
#
# A ragged tensor should *not* be thought of as a type of sparse tensor. In particular, sparse tensors are *efficient encodings for tf.Tensor*, that model the same data in a compact format; but ragged tensor is an *extension to tf.Tensor*, that models an expanded class of data. This difference is crucial when defining operations:
#
# * Applying an op to a sparse or dense tensor should always give the same result.
# * Applying an op to a ragged or sparse tensor may give different results.
#
# As an illustrative example, consider how array operations such as `concat`,
# `stack`, and `tile` are defined for ragged vs. sparse tensors. Concatenating
# ragged tensors joins each row to form a single row with the combined length:
#
# 
#
# + colab_type="code" id="ush7IGUWLXIn" colab={}
ragged_x = tf.ragged.constant([["John"], ["a", "big", "dog"], ["my", "cat"]])
ragged_y = tf.ragged.constant([["fell", "asleep"], ["barked"], ["is", "fuzzy"]])
print(tf.concat([ragged_x, ragged_y], axis=1))
# + [markdown] colab_type="text" id="pvQzZG8zMoWa"
# But concatenating sparse tensors is equivalent to concatenating the corresponding dense tensors,
# as illustrated by the following example (where Ø indicates missing values):
#
# 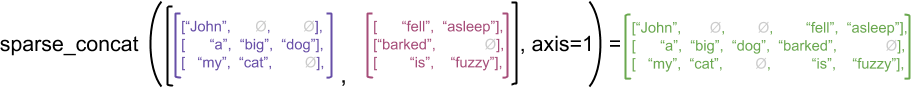
#
# + colab_type="code" id="eTIhGayQL0gI" colab={}
sparse_x = ragged_x.to_sparse()
sparse_y = ragged_y.to_sparse()
sparse_result = tf.sparse.concat(sp_inputs=[sparse_x, sparse_y], axis=1)
print(tf.sparse.to_dense(sparse_result, ''))
# + [markdown] colab_type="text" id="Vl8eQN8pMuYx"
# For another example of why this distinction is important, consider the
# definition of “the mean value of each row” for an op such as `tf.reduce_mean`.
# For a ragged tensor, the mean value for a row is the sum of the
# row’s values divided by the row’s width.
# But for a sparse tensor, the mean value for a row is the sum of the
# row’s values divided by the sparse tensor’s overall width (which is
# greater than or equal to the width of the longest row).
#
# + [markdown] colab_type="text" id="u4yjxcK7IPXc"
# ## TensorFlow APIs
# + [markdown] colab_type="text" id="VoZGwFQjIYU5"
# ### Keras
#
# [tf.keras](https://www.tensorflow.org/guide/keras) is TensorFlow's high-level API for building and training deep learning models. Ragged tensors may be passed as inputs to a Keras model by setting `ragged=True` on `tf.keras.Input` or `tf.keras.layers.InputLayer`. Ragged tensors may also be passed between Keras layers, and returned by Keras models. The following example shows a toy LSTM model that is trained using ragged tensors.
# + colab_type="code" id="pHls7hQVJlk5" colab={}
# Task: predict whether each sentence is a question or not.
sentences = tf.constant(
['What makes you think she is a witch?',
'She turned me into a newt.',
'A newt?',
'Well, I got better.'])
is_question = tf.constant([True, False, True, False])
# Preprocess the input strings.
hash_buckets = 1000
words = tf.strings.split(sentences, ' ')
hashed_words = tf.strings.to_hash_bucket_fast(words, hash_buckets)
# Build the Keras model.
keras_model = tf.keras.Sequential([
tf.keras.layers.Input(shape=[None], dtype=tf.int64, ragged=True),
tf.keras.layers.Embedding(hash_buckets, 16),
tf.keras.layers.LSTM(32, use_bias=False),
tf.keras.layers.Dense(32),
tf.keras.layers.Activation(tf.nn.relu),
tf.keras.layers.Dense(1)
])
keras_model.compile(loss='binary_crossentropy', optimizer='rmsprop')
keras_model.fit(hashed_words, is_question, epochs=5)
print(keras_model.predict(hashed_words))
# + [markdown] colab_type="text" id="8B_sdlt6Ij61"
# ### tf.Example
#
# [tf.Example](https://www.tensorflow.org/tutorials/load_data/tfrecord) is a standard [protobuf](https://developers.google.com/protocol-buffers/) encoding for TensorFlow data. Data encoded with `tf.Example`s often includes variable-length features. For example, the following code defines a batch of four `tf.Example` messages with different feature lengths:
# + colab_type="code" id="xsiglYM7TXGr" colab={}
import google.protobuf.text_format as pbtext
def build_tf_example(s):
return pbtext.Merge(s, tf.train.Example()).SerializeToString()
example_batch = [
build_tf_example(r'''
features {
feature {key: "colors" value {bytes_list {value: ["red", "blue"]} } }
feature {key: "lengths" value {int64_list {value: [7]} } } }'''),
build_tf_example(r'''
features {
feature {key: "colors" value {bytes_list {value: ["orange"]} } }
feature {key: "lengths" value {int64_list {value: []} } } }'''),
build_tf_example(r'''
features {
feature {key: "colors" value {bytes_list {value: ["black", "yellow"]} } }
feature {key: "lengths" value {int64_list {value: [1, 3]} } } }'''),
build_tf_example(r'''
features {
feature {key: "colors" value {bytes_list {value: ["green"]} } }
feature {key: "lengths" value {int64_list {value: [3, 5, 2]} } } }''')]
# + [markdown] colab_type="text" id="szUuXFvtUL2o"
# We can parse this encoded data using `tf.io.parse_example`, which takes a tensor of serialized strings and a feature specification dictionary, and returns a dictionary mapping feature names to tensors. To read the variable-length features into ragged tensors, we simply use `tf.io.RaggedFeature` in the feature specification dictionary:
# + colab_type="code" id="xcdaIbYVT4mo" colab={}
feature_specification = {
'colors': tf.io.RaggedFeature(tf.string),
'lengths': tf.io.RaggedFeature(tf.int64),
}
feature_tensors = tf.io.parse_example(example_batch, feature_specification)
for name, value in feature_tensors.items():
print("{}={}".format(name, value))
# + [markdown] colab_type="text" id="IK9X_8rXVr8h"
# `tf.io.RaggedFeature` can also be used to read features with multiple ragged dimensions. For details, see the [API documentation](https://www.tensorflow.org/api_docs/python/tf/io/RaggedFeature).
# + [markdown] colab_type="text" id="UJowRhlxIX0R"
# ### Datasets
#
# [tf.data](https://www.tensorflow.org/guide/data) is an API that enables you to build complex input pipelines from simple, reusable pieces. Its core data structure is `tf.data.Dataset`, which represents a sequence of elements, in which each element consists of one or more components.
# + colab_type="code" id="fBml1m2G2vO9" colab={}
# Helper function used to print datasets in the examples below.
def print_dictionary_dataset(dataset):
for i, element in enumerate(dataset):
print("Element {}:".format(i))
for (feature_name, feature_value) in element.items():
print('{:>14} = {}'.format(feature_name, feature_value))
# + [markdown] colab_type="text" id="gEu_H1Sp2jz1"
# #### Building Datasets with ragged tensors
#
# Datasets can be built from ragged tensors using the same methods that are used to build them from `tf.Tensor`s or numpy `array`s, such as `Dataset.from_tensor_slices`:
# + colab_type="code" id="BuelF_y2mEq9" colab={}
dataset = tf.data.Dataset.from_tensor_slices(feature_tensors)
print_dictionary_dataset(dataset)
# + [markdown] colab_type="text" id="mC-QNkJc56De"
# Note: `Dataset.from_generator` does not support ragged tensors yet, but support will be added soon.
# + [markdown] colab_type="text" id="K0UKvBLf1VMu"
# #### Batching and unbatching Datasets with ragged tensors
#
# Datasets with ragged tensors can be batched (which combines *n* consecutive elements into a single elements) using the `Dataset.batch` method.
# + colab_type="code" id="lk62aRz63IZn" colab={}
batched_dataset = dataset.batch(2)
print_dictionary_dataset(batched_dataset)
# + [markdown] colab_type="text" id="NLSGiYEQ5A8N"
# Conversely, a batched dataset can be transformed into a flat dataset using `Dataset.unbatch`.
# + colab_type="code" id="CxLlaPw_5Je4" colab={}
unbatched_dataset = batched_dataset.unbatch()
print_dictionary_dataset(unbatched_dataset)
# + [markdown] colab_type="text" id="YzpLQFh33q0N"
# #### Batching Datasets with variable-length non-ragged tensors
#
# If you have a Dataset that contains non-ragged tensors, and tensor lengths vary across elements, then you can batch those non-ragged tensors into ragged tensors by applying the `dense_to_ragged_batch` transformation:
# + colab_type="code" id="PYnhERwh3_mf" colab={}
non_ragged_dataset = tf.data.Dataset.from_tensor_slices([1, 5, 3, 2, 8])
non_ragged_dataset = non_ragged_dataset.map(tf.range)
batched_non_ragged_dataset = non_ragged_dataset.apply(
tf.data.experimental.dense_to_ragged_batch(2))
for element in batched_non_ragged_dataset:
print(element)
# + [markdown] colab_type="text" id="nXFPeE-CzJ-s"
# #### Transforming Datasets with ragged tensors
#
# Ragged tensors in Datasets can also be created or transformed using `Dataset.map`.
# + colab_type="code" id="Ios1GuG-pf9U" colab={}
def transform_lengths(features):
return {
'mean_length': tf.math.reduce_mean(features['lengths']),
'length_ranges': tf.ragged.range(features['lengths'])}
transformed_dataset = dataset.map(transform_lengths)
print_dictionary_dataset(transformed_dataset)
# + [markdown] colab_type="text" id="WD2lWw3fIXrg"
# ### tf.function
#
# [tf.function](https://www.tensorflow.org/guide/function) is a decorator that precomputes TensorFlow graphs for Python functions, which can substantially improve the performance of your TensorFlow code. Ragged tensors can be used transparently with `@tf.function`-decorated functions. For example, the following function works with both ragged and non-ragged tensors:
# + colab_type="code" id="PfyxgVaj_8tl" colab={}
@tf.function
def make_palindrome(x, axis):
return tf.concat([x, tf.reverse(x, [axis])], axis)
# + colab_type="code" id="vcZdzvEnDEt0" colab={}
make_palindrome(tf.constant([[1, 2], [3, 4], [5, 6]]), axis=1)
# + colab_type="code" id="4WfCMIgdDMxj" colab={}
make_palindrome(tf.ragged.constant([[1, 2], [3], [4, 5, 6]]), axis=1)
# + [markdown] colab_type="text" id="X2p69YPOBUz8"
# If you wish to explicitly specify the `input_signature` for the `tf.function`, then you can do so using `tf.RaggedTensorSpec`.
# + colab_type="code" id="k6-hkhdDBk6G" colab={}
@tf.function(
input_signature=[tf.RaggedTensorSpec(shape=[None, None], dtype=tf.int32)])
def max_and_min(rt):
return (tf.math.reduce_max(rt, axis=-1), tf.math.reduce_min(rt, axis=-1))
max_and_min(tf.ragged.constant([[1, 2], [3], [4, 5, 6]]))
# + [markdown] colab_type="text" id="fSs-7E0VD85q"
# #### Concrete functions
#
# [Concrete functions](https://www.tensorflow.org/guide/concrete_function) encapsulate individual traced graphs that are built by `tf.function`. Starting with TensorFlow 2.3 (and in `tf-nightly`), ragged tensors can be used transparently with concrete functions.
#
# + colab_type="code" id="yyJeXJ4wFWox" colab={}
# Preferred way to use ragged tensors with concrete functions (TF 2.3+):
try:
@tf.function
def increment(x):
return x + 1
rt = tf.ragged.constant([[1, 2], [3], [4, 5, 6]])
cf = increment.get_concrete_function(rt)
print(cf(rt))
except Exception as e:
print(f"Not supported before TF 2.3: {type(e)}: {e}")
# + [markdown] colab_type="text" id="cXWDzG5jFVro"
# If you need to use ragged tensors with concrete functions prior to TensorFlow 2.3, then we recommend decomposing ragged tensors into their components (`values` and `row_splits`), and passing them in as separate arguments.
# + colab_type="code" id="5dYzeI0FIdDV" colab={}
# Backwards-compatible way to use ragged tensors with concrete functions:
@tf.function
def decomposed_ragged_increment(x_values, x_splits):
x = tf.RaggedTensor.from_row_splits(x_values, x_splits)
return x + 1
rt = tf.ragged.constant([[1, 2], [3], [4, 5, 6]])
cf = decomposed_ragged_increment.get_concrete_function(rt.values, rt.row_splits)
print(cf(rt.values, rt.row_splits))
# + [markdown] colab_type="text" id="iYLyPlatIXhh"
# ### SavedModels
#
# A [SavedModel](https://www.tensorflow.org/guide/saved_model) is a serialized TensorFlow program, including both weights and computation. It can be built from a Keras model or from a custom model. In either case, ragged tensors can be used transparently with the functions and methods defined by a SavedModel.
#
# + [markdown] colab_type="text" id="98VpBSdOgWqL"
# #### Example: saving a Keras model
# + colab_type="code" id="D-Dg9w7Je5pU" colab={}
import tempfile
keras_module_path = tempfile.mkdtemp()
tf.saved_model.save(keras_model, keras_module_path)
imported_model = tf.saved_model.load(keras_module_path)
imported_model(hashed_words)
# + [markdown] colab_type="text" id="9-7k-E92gaoR"
# #### Example: saving a custom model
#
# + colab_type="code" id="Sfem1ESrdGzX" colab={}
class CustomModule(tf.Module):
def __init__(self, variable_value):
super(CustomModule, self).__init__()
self.v = tf.Variable(variable_value)
@tf.function
def grow(self, x):
return x * self.v
module = CustomModule(100.0)
# Before saving a custom model, we must ensure that concrete functions are
# built for each input signature that we will need.
module.grow.get_concrete_function(tf.RaggedTensorSpec(shape=[None, None],
dtype=tf.float32))
custom_module_path = tempfile.mkdtemp()
tf.saved_model.save(module, custom_module_path)
imported_model = tf.saved_model.load(custom_module_path)
imported_model.grow(tf.ragged.constant([[1.0, 4.0, 3.0], [2.0]]))
# + [markdown] colab_type="text" id="SAxis5KBhrBN"
# Note: SavedModel [signatures](https://www.tensorflow.org/guide/saved_model#specifying_signatures_during_export) are concrete functions. As discussed in the section on Concrete Functions above, ragged tensors are only handled correctly by concrete functions starting with TensorFlow 2.3 (and in `tf_nightly`). If you need to use SavedModel signatures in a previous version of TensorFlow, then we recommend decomposing the ragged tensor into its component tensors.
# + [markdown] colab_type="text" id="cRcHzS6pcHYC"
# ## Overloaded operators
#
# The `RaggedTensor` class overloads the standard Python arithmetic and comparison
# operators, making it easy to perform basic elementwise math:
# + colab_type="code" id="skScd37P-PVu" colab={}
x = tf.ragged.constant([[1, 2], [3], [4, 5, 6]])
y = tf.ragged.constant([[1, 1], [2], [3, 3, 3]])
print(x + y)
# + [markdown] colab_type="text" id="XEGgbZHV-PVw"
# Since the overloaded operators perform elementwise computations, the inputs to
# all binary operations must have the same shape, or be broadcastable to the same
# shape. In the simplest broadcasting case, a single scalar is combined
# elementwise with each value in a ragged tensor:
# + colab_type="code" id="IYybEEWc-PVx" colab={}
x = tf.ragged.constant([[1, 2], [3], [4, 5, 6]])
print(x + 3)
# + [markdown] colab_type="text" id="okGb9dIi-PVz"
# For a discussion of more advanced cases, see the section on
# **Broadcasting**.
#
# Ragged tensors overload the same set of operators as normal `Tensor`s: the unary
# operators `-`, `~`, and `abs()`; and the binary operators `+`, `-`, `*`, `/`,
# `//`, `%`, `**`, `&`, `|`, `^`, `==`, `<`, `<=`, `>`, and `>=`.
#
# + [markdown] colab_type="text" id="f2anbs6ZnFtl"
# ## Indexing
#
# Ragged tensors support Python-style indexing, including multidimensional
# indexing and slicing. The following examples demonstrate ragged tensor indexing
# with a 2-D and a 3-D ragged tensor.
# + [markdown] colab_type="text" id="XuEwmC3t_ITL"
# ### Indexing examples: 2D ragged tensor
# + colab_type="code" id="MbSRZRDz-PV1" colab={}
queries = tf.ragged.constant(
[['Who', 'is', 'George', 'Washington'],
['What', 'is', 'the', 'weather', 'tomorrow'],
['Goodnight']])
# + colab_type="code" id="2HRs2xhh-vZE" colab={}
print(queries[1]) # A single query
# + colab_type="code" id="EFfjZV7YA3UH" colab={}
print(queries[1, 2]) # A single word
# + colab_type="code" id="VISRPQSdA3xn" colab={}
print(queries[1:]) # Everything but the first row
# + colab_type="code" id="J1PpSyKQBMng" colab={}
print(queries[:, :3]) # The first 3 words of each query
# + colab_type="code" id="ixrhHmJBeidy" colab={}
print(queries[:, -2:]) # The last 2 words of each query
# + [markdown] colab_type="text" id="cnOP6Vza-PV4"
# ### Indexing examples 3D ragged tensor
# + colab_type="code" id="8VbqbKcE-PV6" colab={}
rt = tf.ragged.constant([[[1, 2, 3], [4]],
[[5], [], [6]],
[[7]],
[[8, 9], [10]]])
# + colab_type="code" id="f9WPVWf4grVp" colab={}
print(rt[1]) # Second row (2-D RaggedTensor)
# + colab_type="code" id="ad8FGJoABjQH" colab={}
print(rt[3, 0]) # First element of fourth row (1-D Tensor)
# + colab_type="code" id="MPPr-a-bBjFE" colab={}
print(rt[:, 1:3]) # Items 1-3 of each row (3-D RaggedTensor)
# + colab_type="code" id="6SIDeoIUBi4z" colab={}
print(rt[:, -1:]) # Last item of each row (3-D RaggedTensor)
# + [markdown] colab_type="text" id="_d3nBh1GnWvU"
# `RaggedTensor`s supports multidimensional indexing and slicing, with one
# restriction: indexing into a ragged dimension is not allowed. This case is
# problematic because the indicated value may exist in some rows but not others.
# In such cases, it's not obvious whether we should (1) raise an `IndexError`; (2)
# use a default value; or (3) skip that value and return a tensor with fewer rows
# than we started with. Following the
# [guiding principles of Python](https://www.python.org/dev/peps/pep-0020/)
# ("In the face
# of ambiguity, refuse the temptation to guess" ), we currently disallow this
# operation.
# + [markdown] colab_type="text" id="IsWKETULAJbN"
# ## Tensor type conversion
#
# The `RaggedTensor` class defines methods that can be used to convert
# between `RaggedTensor`s and `tf.Tensor`s or `tf.SparseTensor`s:
# + colab_type="code" id="INnfmZGcBoU_" colab={}
ragged_sentences = tf.ragged.constant([
['Hi'], ['Welcome', 'to', 'the', 'fair'], ['Have', 'fun']])
# + colab_type="code" id="__iJ4iXtkGOx" colab={}
# RaggedTensor -> Tensor
print(ragged_sentences.to_tensor(default_value='', shape=[None, 10]))
# + colab_type="code" id="-rfiyYqne8QN" colab={}
# Tensor -> RaggedTensor
x = [[1, 3, -1, -1], [2, -1, -1, -1], [4, 5, 8, 9]]
print(tf.RaggedTensor.from_tensor(x, padding=-1))
# + colab_type="code" id="41WAZLXNnbwH" colab={}
#RaggedTensor -> SparseTensor
print(ragged_sentences.to_sparse())
# + colab_type="code" id="S8MkYo2hfVhj" colab={}
# SparseTensor -> RaggedTensor
st = tf.SparseTensor(indices=[[0, 0], [2, 0], [2, 1]],
values=['a', 'b', 'c'],
dense_shape=[3, 3])
print(tf.RaggedTensor.from_sparse(st))
# + [markdown] colab_type="text" id="qx025sNMkAHH"
# ## Evaluating ragged tensors
#
# To access the values in a ragged tensor, you can:
#
# 1. Use `tf.RaggedTensor.to_list()` to convert the ragged tensor to a
# nested python list.
# 1. Use `tf.RaggedTensor.numpy()` to convert the ragged tensor to a numpy array
# whose values are nested numpy arrays.
# 1. Decompose the ragged tensor into its components, using the
# `tf.RaggedTensor.values` and `tf.RaggedTensor.row_splits`
# properties, or row-paritioning methods such as
# `tf.RaggedTensor.row_lengths()` and `tf.RaggedTensor.value_rowids()`.
# 1. Use Python indexing to select values from the ragged tensor.
#
# + colab_type="code" id="uMm1WMkc-PV_" colab={}
rt = tf.ragged.constant([[1, 2], [3, 4, 5], [6], [], [7]])
print("python list:", rt.to_list())
print("numpy array:", rt.numpy())
print("values:", rt.values.numpy())
print("splits:", rt.row_splits.numpy())
print("indexed value:", rt[1].numpy())
# + [markdown] colab_type="text" id="EdljbNPq-PWS"
# ## Broadcasting
#
# Broadcasting is the process of making tensors with different shapes have
# compatible shapes for elementwise operations. For more background on
# broadcasting, see:
#
# * [Numpy: Broadcasting](https://docs.scipy.org/doc/numpy/user/basics.broadcasting.html)
# * `tf.broadcast_dynamic_shape`
# * `tf.broadcast_to`
#
# The basic steps for broadcasting two inputs `x` and `y` to have compatible
# shapes are:
#
# 1. If `x` and `y` do not have the same number of dimensions, then add outer
# dimensions (with size 1) until they do.
#
# 2. For each dimension where `x` and `y` have different sizes:
#
# * If `x` or `y` have size `1` in dimension `d`, then repeat its values
# across dimension `d` to match the other input's size.
#
# * Otherwise, raise an exception (`x` and `y` are not broadcast
# compatible).
#
# Where the size of a tensor in a uniform dimension is a single number (the size
# of slices across that dimension); and the size of a tensor in a ragged dimension
# is a list of slice lengths (for all slices across that dimension).
# + [markdown] colab_type="text" id="-S2hOUWx-PWU"
# ### Broadcasting examples
# + colab_type="code" id="0n095XdR-PWU" colab={}
# x (2D ragged): 2 x (num_rows)
# y (scalar)
# result (2D ragged): 2 x (num_rows)
x = tf.ragged.constant([[1, 2], [3]])
y = 3
print(x + y)
# + colab_type="code" id="0SVYk5AP-PWW" colab={}
# x (2d ragged): 3 x (num_rows)
# y (2d tensor): 3 x 1
# Result (2d ragged): 3 x (num_rows)
x = tf.ragged.constant(
[[10, 87, 12],
[19, 53],
[12, 32]])
y = [[1000], [2000], [3000]]
print(x + y)
# + colab_type="code" id="MsfBMD80s8Ux" colab={}
# x (3d ragged): 2 x (r1) x 2
# y (2d ragged): 1 x 1
# Result (3d ragged): 2 x (r1) x 2
x = tf.ragged.constant(
[[[1, 2], [3, 4], [5, 6]],
[[7, 8]]],
ragged_rank=1)
y = tf.constant([[10]])
print(x + y)
# + colab_type="code" id="rEj5QVfnva0t" colab={}
# x (3d ragged): 2 x (r1) x (r2) x 1
# y (1d tensor): 3
# Result (3d ragged): 2 x (r1) x (r2) x 3
x = tf.ragged.constant(
[
[
[[1], [2]],
[],
[[3]],
[[4]],
],
[
[[5], [6]],
[[7]]
]
],
ragged_rank=2)
y = tf.constant([10, 20, 30])
print(x + y)
# + [markdown] colab_type="text" id="uennZ64Aqftb"
# Here are some examples of shapes that do not broadcast:
# + colab_type="code" id="UpI0FlfL4Eim" colab={}
# x (2d ragged): 3 x (r1)
# y (2d tensor): 3 x 4 # trailing dimensions do not match
x = tf.ragged.constant([[1, 2], [3, 4, 5, 6], [7]])
y = tf.constant([[1, 2, 3, 4], [5, 6, 7, 8], [9, 10, 11, 12]])
try:
x + y
except tf.errors.InvalidArgumentError as exception:
print(exception)
# + colab_type="code" id="qGq1zOT4zMoc" colab={}
# x (2d ragged): 3 x (r1)
# y (2d ragged): 3 x (r2) # ragged dimensions do not match.
x = tf.ragged.constant([[1, 2, 3], [4], [5, 6]])
y = tf.ragged.constant([[10, 20], [30, 40], [50]])
try:
x + y
except tf.errors.InvalidArgumentError as exception:
print(exception)
# + colab_type="code" id="CvLae5vMqeji" colab={}
# x (3d ragged): 3 x (r1) x 2
# y (3d ragged): 3 x (r1) x 3 # trailing dimensions do not match
x = tf.ragged.constant([[[1, 2], [3, 4], [5, 6]],
[[7, 8], [9, 10]]])
y = tf.ragged.constant([[[1, 2, 0], [3, 4, 0], [5, 6, 0]],
[[7, 8, 0], [9, 10, 0]]])
try:
x + y
except tf.errors.InvalidArgumentError as exception:
print(exception)
# + [markdown] colab_type="text" id="m0wQkLfV-PWa"
# ## RaggedTensor encoding
#
# Ragged tensors are encoded using the `RaggedTensor` class. Internally, each
# `RaggedTensor` consists of:
#
# * A `values` tensor, which concatenates the variable-length rows into a
# flattened list.
# * A `row_partition`, which indicates how those flattened values are divided
# into rows.
#
# 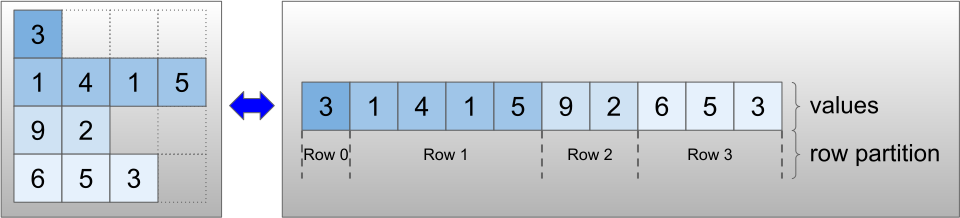
#
# The `row_partition` can be stored using four different encodings:
#
# * `row_splits` is an integer vector specifying the split points between rows.
# * `value_rowids` is an integer vector specifying the row index for each value.
# * `row_lengths` is an integer vector specifying the length of each row.
# * `uniform_row_length` is an integer scalar specifying a single length for
# all rows.
#
# 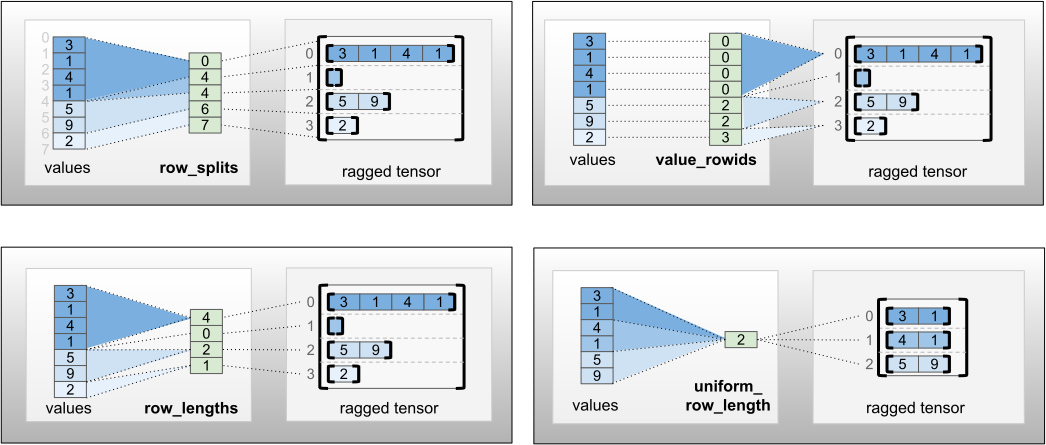
#
# An integer scalar `nrows` can also be included in the `row_partition` encoding, to account for empty trailing rows with `value_rowids`, or empty rows with `uniform_row_length`.
#
# + colab_type="code" id="MrLgMu0gPuo-" colab={}
rt = tf.RaggedTensor.from_row_splits(
values=[3, 1, 4, 1, 5, 9, 2],
row_splits=[0, 4, 4, 6, 7])
print(rt)
# + [markdown] colab_type="text" id="wEfZOKwN1Ra_"
# The choice of which encoding to use for row partitions is managed internally by ragged tensors, to improve efficiency in some contexts. In particular, some of the advantages and disadvantages of the different row-partitioning
# schemes are:
#
# # + **Efficient indexing**:
# The `row_splits` encoding enables
# constant-time indexing and slicing into ragged tensors.
#
# # + **Efficient concatenation**:
# The `row_lengths` encoding is more efficient when concatenating ragged
# tensors, since row lengths do not change when two tensors are concatenated
# together.
#
# # + **Small encoding size**:
# The `value_rowids` encoding is more efficient when storing ragged tensors
# that have a large number of empty rows, since the size of the tensor
# depends only on the total number of values. On the other hand, the
# `row_splits` and `row_lengths` encodings
# are more efficient when storing ragged tensors with longer rows, since they
# require only one scalar value for each row.
#
# # + **Compatibility**:
# The `value_rowids` scheme matches the
# [segmentation](https://www.tensorflow.org/api_docs/python/tf/math#about_segmentation)
# format used by operations such as `tf.segment_sum`. The `row_limits` scheme
# matches the format used by ops such as `tf.sequence_mask`.
#
# # + **Uniform dimensions**:
# As discussed below, the `uniform_row_length` encoding is used to encode
# ragged tensors with uniform dimensions.
# + [markdown] colab_type="text" id="bpB7xKoUPtU6"
# ### Multiple ragged dimensions
#
# A ragged tensor with multiple ragged dimensions is encoded by using a nested
# `RaggedTensor` for the `values` tensor. Each nested `RaggedTensor` adds a single
# ragged dimension.
#
# 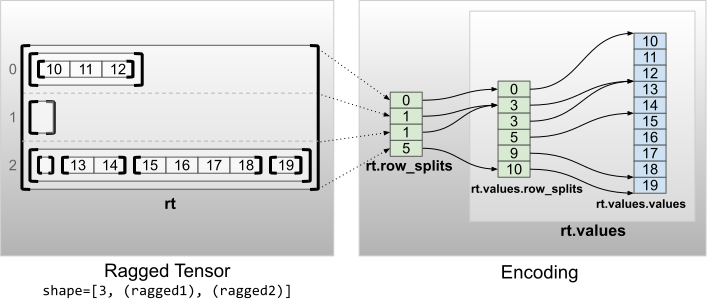
#
# + colab_type="code" id="yy3IGT2a-PWb" colab={}
rt = tf.RaggedTensor.from_row_splits(
values=tf.RaggedTensor.from_row_splits(
values=[10, 11, 12, 13, 14, 15, 16, 17, 18, 19],
row_splits=[0, 3, 3, 5, 9, 10]),
row_splits=[0, 1, 1, 5])
print(rt)
print("Shape: {}".format(rt.shape))
print("Number of partitioned dimensions: {}".format(rt.ragged_rank))
# + [markdown] colab_type="text" id="5HqEEDzk-PWc"
# The factory function `tf.RaggedTensor.from_nested_row_splits` may be used to construct a
# RaggedTensor with multiple ragged dimensions directly, by providing a list of
# `row_splits` tensors:
# + colab_type="code" id="AKYhtFcT-PWd" colab={}
rt = tf.RaggedTensor.from_nested_row_splits(
flat_values=[10, 11, 12, 13, 14, 15, 16, 17, 18, 19],
nested_row_splits=([0, 1, 1, 5], [0, 3, 3, 5, 9, 10]))
print(rt)
# + [markdown] colab_type="text" id="BqAfbkAC56m0"
# ### Ragged rank and flat values
#
# A ragged tensor's ***ragged rank*** is the number of times that the underlying
# `values` Tensor has been partitioned (i.e., the nesting depth of `RaggedTensor` objects). The innermost `values` tensor is known as its ***flat_values***. In the following example, `conversations` has ragged_rank=3, and its `flat_values` is a 1D `Tensor` with 24 strings:
#
# + colab_type="code" id="BXp-Tt2bClem" colab={}
# shape = [batch, (paragraph), (sentence), (word)]
conversations = tf.ragged.constant(
[[[["I", "like", "ragged", "tensors."]],
[["Oh", "yeah?"], ["What", "can", "you", "use", "them", "for?"]],
[["Processing", "variable", "length", "data!"]]],
[[["I", "like", "cheese."], ["Do", "you?"]],
[["Yes."], ["I", "do."]]]])
conversations.shape
# + colab_type="code" id="DZUMrgxXFd5s" colab={}
assert conversations.ragged_rank == len(conversations.nested_row_splits)
conversations.ragged_rank # Number of partitioned dimensions.
# + colab_type="code" id="xXLSNpS0Fdvp" colab={}
conversations.flat_values.numpy()
# + [markdown] colab_type="text" id="uba2EnAY-PWf"
# ### Uniform inner dimensions
#
# Ragged tensors with uniform inner dimensions are encoded by using a
# multidimensional `tf.Tensor` for the flat_values (i.e., the innermost `values`).
#
# 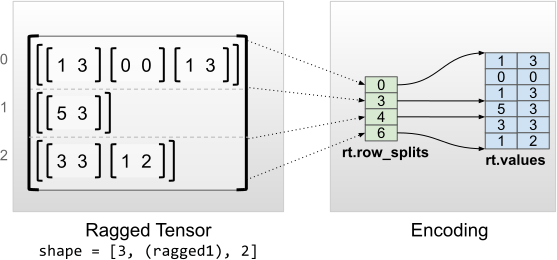
# + colab_type="code" id="z2sHwHdy-PWg" colab={}
rt = tf.RaggedTensor.from_row_splits(
values=[[1, 3], [0, 0], [1, 3], [5, 3], [3, 3], [1, 2]],
row_splits=[0, 3, 4, 6])
print(rt)
print("Shape: {}".format(rt.shape))
print("Number of partitioned dimensions: {}".format(rt.ragged_rank))
print("Flat values shape: {}".format(rt.flat_values.shape))
print("Flat values:\n{}".format(rt.flat_values))
# + [markdown] colab_type="text" id="WoGRKd50x_qz"
# ### Uniform non-inner dimensions
#
# Ragged tensors with uniform non-inner dimensions are encoded by partitioning rows with `uniform_row_length`.
#
# 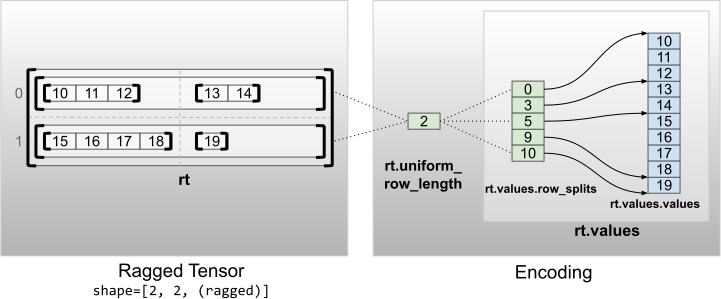
# + colab_type="code" id="70q1aCKwySgS" colab={}
rt = tf.RaggedTensor.from_uniform_row_length(
values=tf.RaggedTensor.from_row_splits(
values=[10, 11, 12, 13, 14, 15, 16, 17, 18, 19],
row_splits=[0, 3, 5, 9, 10]),
uniform_row_length=2)
print(rt)
print("Shape: {}".format(rt.shape))
print("Number of partitioned dimensions: {}".format(rt.ragged_rank))
| site/en/guide/ragged_tensor.ipynb |