code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv_multimodal
# language: python
# name: venv_multimodal
# ---
# +
import numpy as np
import torch
import torch.distributions as dist
from torch import optim
from torch.utils.data import DataLoader
import models
#import objectives
import objectives_dev as objectives
from utils import Logger, Timer, save_model, save_vars, unpack_data
from utils import log_mean_exp, is_multidata, kl_divergence, get_mean
from datasets_dev import ATAC_Dataset, RNA_Dataset
import torch
import torch.distributions as dist
import torch.nn as nn
import torch.nn.functional as F
from numpy import prod, sqrt
from torch.utils.data import DataLoader
from torchvision import datasets, transforms
from torchvision.utils import save_image, make_grid
from utils import Constants, get_mean
from vis import plot_embeddings, plot_kls_df, embed_umap
from sklearn.manifold import TSNE
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.cluster.hierarchy import dendrogram, linkage
from sklearn.model_selection import ParameterGrid
# +
#args
experiment = 'hyperparam'
model = 'rna_atac_dev' #VAE試しに使う
obj = 'elbo'
K = 10
looser = False
llik_scaling = 0
batch_size = 128
epochs = 100
latent_dim = 32
num_hidden_layers = 2
hidden_dim =1024
learn_prior = False
logp = False
print_freq = 0
no_analytics = False
seed = 1
class params():
def __init__(self,
experiment,
model,
obj,
K,
looser,
llik_scaling,
batch_size,
epochs,
latent_dim,
num_hidden_layers,
hidden_dim,
learn_prior,
logp,
print_freq,
no_analytics,
seed):
self.experiment = experiment
self.model = model
self.obj = obj
self.K = K
self.looser = looser
self.llik_scaling = llik_scaling
self.batch_size = batch_size
self.epochs = epochs
self.latent_dim = latent_dim
self.num_hidden_layers = num_hidden_layers
self.hidden_dim = hidden_dim
self.learn_prior = learn_prior
self.logp = logp
self.print_freq = print_freq
self.no_analytics = no_analytics
self.seed = seed
args = params(experiment,
model,
obj,
K,
looser,
llik_scaling,
batch_size,
epochs,
latent_dim,
num_hidden_layers,
hidden_dim,
learn_prior,
logp,
print_freq,
no_analytics,
seed)
# -
test_losses = []
for i in range(40):
runId = str(i)
pretrained_path = '../experiments/' + args.experiment + '/' + runId
losses = torch.load(pretrained_path + '/losses.rar', map_location=torch.device('cpu') )
#train_loss = losses['train_loss']
val_loss = losses['val_loss']
test_loss = losses['test_loss']
id = val_loss.index(min(val_loss))
test_losses.append(test_loss[id])
test_losses
model_id = test_losses.index(min(test_losses))
print(model_id)
print('test_loss ' + str(test_losses[model_id]))
runId = str(model_id)
pretrained_path = '../experiments/' + args.experiment + '/' + runId
losses = torch.load(pretrained_path + '/losses.rar', map_location=torch.device('cpu') )
train_loss = losses['train_loss']
val_loss = losses['val_loss']
test_loss = losses['test_loss']
# +
#Select pretrained model
runId = '2020-03-20T15/58/02.732379xvk9jb3p'
pretrained_path = '../experiments/' + args.experiment + '/' + runId
print('Loading model {} from {}'.format(model.modelName, pretrained_path))
model.load_state_dict(torch.load(pretrained_path + '/model.rar', map_location=torch.device('cpu') ))
model._pz_params = model._pz_params
runPath = pretrained_path
|
src/.ipynb_checkpoints/analyze_supcomp-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Ruby 2.3.3
# language: ruby
# name: ruby
# ---
# # Combinational Circuits | Decoder
# All Combinational circuit components are in the module **Combinational**. All methods available can be found in the documentation.
# load gemfile ruby_circuits.rb
require '../../../../lib/ruby_ciruits'
# +
# Creating a Decoder instance
decoder = Combinational::Decoder.new(0, 1)
# output of the Decoder
print(decoder.output())
# +
# Changing the number of inputs - Input must be a power of 2
decoder.set_inputs(1, 0, 0)
# To get the input states
puts(decoder.get_input_states())
# -
# New output of the encoder
puts(decoder.output())
# +
# Creating a connector instance
conn = Connector.new
# setting the output of the decoder to the connector
decoder.set_output(1, conn)
# Passing the connector as an input to a gate
gate = LogicGates::AND.new(conn, 1)
# Output of the gate
print(gate.output())
# -
# Information about the instance
print(decoder)
|
examples/jupyter_notebook/digital_circuits/combinational/Decoder.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"is_executing": false, "name": "#%%\n"}
import numpy as np
import pandas as pd
from datetime import datetime as dt
# + pycharm={"is_executing": false, "name": "#%%\n"}
csv_path = "../output/repositories.csv"
df = pd.read_csv(csv_path)
df.head()
df_rq = pd.DataFrame(
data={
"age": [],
}
)
# + pycharm={"is_executing": false, "name": "#%%\n"}
"""
RQ 01. Sistemas populares são maduros/antigos?
Métrica: idade do repositório (calculado a partir da data de sua criação)
"""
created_list = []
for i, date_str in enumerate(df["createdAt"]):
date = dt.strptime(date_str, "%Y-%m-%dT%H:%M:%SZ")
age = (dt.today() - date)
created_list.append(age.total_seconds())
# if age.days < 180:
# print(i, age, df["nameWithOwner"][i])
print(f"Min:\t\t{np.min(created_list)/60/60/24/30} months")
print(f"Max:\t\t{np.max(created_list)/60/60/24/365} years")
print(f"Avg:\t\t{np.average(created_list)/60/60/24/365} years")
cl = created_list
cl.sort()
print(f"Avg 0~99:\t{np.average(cl[0:99])/60/60/24/365} years")
print(f"Avg 450~549:\t{np.average(cl[450:549])/60/60/24/365} years")
print(f"Avg 900~999:\t{np.average(cl[900:999])/60/60/24/365} years")
print(f"Median:\t\t{np.median(created_list)/60/60/24/365} years")
# + pycharm={"name": "#%%\n"}
"""
RQ 02. Sistemas populares recebem muita contribuição externa?
Métrica: total de pull requests aceitas
"""
# + pycharm={"name": "#%%\n"}
"""
RQ 03. Sistemas populares lançam releases com frequência?
Métrica: total de releases
"""
# + pycharm={"is_executing": false, "name": "#%%\n"}
"""
RQ 04. Sistemas populares são atualizados com frequência?
Métrica: tempo até a última atualização (calculado a partir da data de última atualização)
"""
updated_list = []
for i, date_str in enumerate(df["updatedAt"]):
date = dt.strptime(date_str, "%Y-%m-%dT%H:%M:%SZ")
age = (dt(2020, 3, 9, 23, 23, 0) - date)
updated_list.append(age.total_seconds())
# if age.days > 1:
# print(i, age, df["nameWithOwner"][i])
print(f"Min:\t\t{np.min(updated_list)/60} minutos")
print(f"Max:\t\t{np.max(updated_list)/60} minutos")
print(f"Avg:\t\t{np.average(updated_list)/60} minutos")
cl = updated_list
cl.sort()
print(f"Median:\t\t{np.median(updated_list)/60} minutos")
# + pycharm={"is_executing": false, "name": "#%%\n"}
"""
RQ 05.
"""
lang_list = []
lang_list_ = []
for lang in df["primaryLanguage"]:
lang_list_.append(lang)
if lang not in lang_list:
lang_list.append(lang)
for lang in lang_list:
print(f"{lang}, {lang_list_.count(lang)}")
df.groupby("primaryLanguage").count()
|
Lab01/sprints/Lab01S03.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multi-Layer Perceptron, MNIST
# ---
# In this notebook, we will train an MLP to classify images from the [MNIST database](http://yann.lecun.com/exdb/mnist/) hand-written digit database.
#
# The process will be broken down into the following steps:
# >1. Load and visualize the data
# 2. Define a neural network
# 3. Train the model
# 4. Evaluate the performance of our trained model on a test dataset!
#
# Before we begin, we have to import the necessary libraries for working with data and PyTorch.
# import libraries
import torch
import numpy as np
# ---
# ## Load and Visualize the [Data](http://pytorch.org/docs/stable/torchvision/datasets.html)
#
# Downloading may take a few moments, and you should see your progress as the data is loading. You may also choose to change the `batch_size` if you want to load more data at a time.
#
# This cell will create DataLoaders for each of our datasets.
# +
from torchvision import datasets
import torchvision.transforms as transforms
# number of subprocesses to use for data loading
num_workers = 0
# how many samples per batch to load
batch_size = 20
# convert data to torch.FloatTensor
transform = transforms.ToTensor()
# choose the training and test datasets
train_data = datasets.MNIST(root='data', train=True,
download=True, transform=transform)
test_data = datasets.MNIST(root='data', train=False,
download=True, transform=transform)
# prepare data loaders
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
num_workers=num_workers)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=batch_size,
num_workers=num_workers)
# -
# ### Visualize a Batch of Training Data
#
# The first step in a classification task is to take a look at the data, make sure it is loaded in correctly, then make any initial observations about patterns in that data.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy()
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
# print out the correct label for each image
# .item() gets the value contained in a Tensor
ax.set_title(str(labels[idx].item()))
# -
# ### View an Image in More Detail
# +
img = np.squeeze(images[1])
fig = plt.figure(figsize = (12,12))
ax = fig.add_subplot(111)
ax.imshow(img, cmap='gray')
width, height = img.shape
thresh = img.max()/2.5
for x in range(width):
for y in range(height):
val = round(img[x][y],2) if img[x][y] !=0 else 0
ax.annotate(str(val), xy=(y,x),
horizontalalignment='center',
verticalalignment='center',
color='white' if img[x][y]<thresh else 'black')
# -
# ---
# ## Define the Network [Architecture](http://pytorch.org/docs/stable/nn.html)
#
# The architecture will be responsible for seeing as input a 784-dim Tensor of pixel values for each image, and producing a Tensor of length 10 (our number of classes) that indicates the class scores for an input image. This particular example uses two hidden layers and dropout to avoid overfitting.
# +
import torch.nn as nn
import torch.nn.functional as F
## TODO: Define the NN architecture
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# linear layer (784 -> 1 hidden node)
self.input_nodes = 28 * 28
hidden_nodes1 = 512
hidden_nodes2 = 512
output_nodes = 10
self.fc1 = nn.Linear(self.input_nodes, hidden_nodes1)
self.fc2 = nn.Linear(hidden_nodes1, hidden_nodes2)
self.fc3 = nn.Linear(hidden_nodes2, output_nodes)
self.dropout = nn.Dropout(0.2)
def forward(self, x):
# flatten image input
x = x.view(-1, self.input_nodes)
# add hidden layer, with relu activation function
x = F.relu(self.fc1(x))
x = self.dropout(x)
x = F.relu(self.fc2(x))
x = self.dropout(x)
x = self.fc3(x)
return x
# initialize the NN
model = Net()
print(model)
# -
# ### Specify [Loss Function](http://pytorch.org/docs/stable/nn.html#loss-functions) and [Optimizer](http://pytorch.org/docs/stable/optim.html)
#
# It's recommended that you use cross-entropy loss for classification. If you look at the documentation (linked above), you can see that PyTorch's cross entropy function applies a softmax funtion to the output layer *and* then calculates the log loss.
# +
## TODO: Specify loss and optimization functions
# specify loss function
criterion = nn.CrossEntropyLoss()
# specify optimizer
optimizer = torch.optim.SGD(model.parameters(), lr=0.01)
# -
# ---
# ## Train the Network
#
# The steps for training/learning from a batch of data are described in the comments below:
# 1. Clear the gradients of all optimized variables
# 2. Forward pass: compute predicted outputs by passing inputs to the model
# 3. Calculate the loss
# 4. Backward pass: compute gradient of the loss with respect to model parameters
# 5. Perform a single optimization step (parameter update)
# 6. Update average training loss
#
# The following loop trains for 30 epochs; feel free to change this number. For now, we suggest somewhere between 20-50 epochs. As you train, take a look at how the values for the training loss decrease over time. We want it to decrease while also avoiding overfitting the training data.
# +
# number of epochs to train the model
n_epochs = 30 # suggest training between 20-50 epochs
model.train() # prep model for training
for epoch in range(n_epochs):
# monitor training loss
train_loss = 0.0
###################
# train the model #
###################
for data, target in train_loader:
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update running training loss
train_loss += loss.item()*data.size(0)
# print training statistics
# calculate average loss over an epoch
train_loss = train_loss/len(train_loader.sampler)
print('Epoch: {} \tTraining Loss: {:.6f}'.format(
epoch+1,
train_loss
))
# -
# ---
# ## Test the Trained Network
#
# Finally, we test our best model on previously unseen **test data** and evaluate it's performance. Testing on unseen data is a good way to check that our model generalizes well. It may also be useful to be granular in this analysis and take a look at how this model performs on each class as well as looking at its overall loss and accuracy.
#
# #### `model.eval()`
#
# `model.eval(`) will set all the layers in your model to evaluation mode. This affects layers like dropout layers that turn "off" nodes during training with some probability, but should allow every node to be "on" for evaluation!
# +
# initialize lists to monitor test loss and accuracy
test_loss = 0.0
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
model.eval() # prep model for *evaluation*
for data, target in test_loader:
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# update test loss
test_loss += loss.item()*data.size(0)
# convert output probabilities to predicted class
_, pred = torch.max(output, 1)
# compare predictions to true label
correct = np.squeeze(pred.eq(target.data.view_as(pred)))
# calculate test accuracy for each object class
for i in range(len(target)):
label = target.data[i]
class_correct[label] += correct[i].item()
class_total[label] += 1
# calculate and print avg test loss
test_loss = test_loss/len(test_loader.sampler)
print('Test Loss: {:.6f}\n'.format(test_loss))
for i in range(10):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
str(i), 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
# -
# ### Visualize Sample Test Results
#
# This cell displays test images and their labels in this format: `predicted (ground-truth)`. The text will be green for accurately classified examples and red for incorrect predictions.
# +
# obtain one batch of test images
dataiter = iter(test_loader)
images, labels = dataiter.next()
# get sample outputs
output = model(images)
# convert output probabilities to predicted class
_, preds = torch.max(output, 1)
# prep images for display
images = images.numpy()
# plot the images in the batch, along with predicted and true labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
ax.set_title("{} ({})".format(str(preds[idx].item()), str(labels[idx].item())),
color=("green" if preds[idx]==labels[idx] else "red"))
# -
|
convolutional-neural-networks/mnist-mlp/mnist_mlp_exercise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import gc
import time
import pyreadr as py
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from tqdm.notebook import tqdm
import torch
import torch.nn.functional as F
from torch import nn
from torch.optim import Adam, RMSprop
from torch.optim.lr_scheduler import StepLR, ReduceLROnPlateau
from torch.utils.data import Dataset, DataLoader
from torch.utils.tensorboard import SummaryWriter
from torch.nn.utils.rnn import pad_sequence, pack_padded_sequence, pad_packed_sequence
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import confusion_matrix
device = torch.device("cuda:5" if torch.cuda.is_available() else "cpu")
# device = torch.device("cpu")
print(device)
# -
# > ### Data downloading
# Data link
# https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/6C3JR1
# ##### Data description
# Here we consoder dataset of "Additional Tennessee Eastman Process Simulation Data for Anomaly Detection Evaluation"
# This dataverse contains the data referenced in Rieth et al. (2017). Issues and Advances in Anomaly Detection Evaluation for Joint Human-Automated Systems. To be presented at Applied Human Factors and Ergonomics 2017.
# ##### Columns description
# * **faultNumber** ranges from 1 to 20 in the “Faulty” datasets and represents the fault type in the TEP. The “FaultFree” datasets only contain fault 0 (i.e. normal operating conditions).
# * **simulationRun** ranges from 1 to 500 and represents a different random number generator state from which a full TEP dataset was generated (Note: the actual seeds used to generate training and testing datasets were non-overlapping).
# * **sample** ranges either from 1 to 500 (“Training” datasets) or 1 to 960 (“Testing” datasets). The TEP variables (columns 4 to 55) were sampled every 3 minutes for a total duration of 25 hours and 48 hours respectively. Note that the faults were introduced 1 and 8 hours into the Faulty Training and Faulty Testing datasets, respectively.
# * **columns 4-55** contain the process variables; the column names retain the original variable names.
# +
# # ! unzip ../../data/raw/dataverse_files.zip -d ../../data/raw/dataverse_files
# -
#reading train data in .R format
a1 = py.read_r("../../data/raw/dataverse_files/TEP_FaultFree_Training.RData")
a2 = py.read_r("../../data/raw/dataverse_files/TEP_Faulty_Training.RData")
# +
#reading test data in .R format
# a3 = py.read_r("../../data/raw/dataverse_files/TEP_FaultFree_Testing.RData")
# a4 = py.read_r("../../data/raw/dataverse_files/TEP_Faulty_Testing.RData")
# -
print("Objects that are present in a1 :", a1.keys())
print("Objects that are present in a2 :", a2.keys())
# print("Objects that are present in a3 :", a3.keys())
# print("Objects that are present in a4 :", a4.keys())
# +
# concatinating the train and the test dataset
raw_train = pd.concat([a1['fault_free_training'], a2['faulty_training']])
# raw_test = pd.concat([a3['fault_free_testing'], a4['faulty_testing']])
# +
# 5.250.000, 10.080.000
# len(raw_train), len(raw_test)
# -
# > ### Train-test-split
features = [
'xmeas_1', 'xmeas_2', 'xmeas_3', 'xmeas_4', 'xmeas_5', 'xmeas_6', 'xmeas_7', 'xmeas_8',
'xmeas_9', 'xmeas_10', 'xmeas_11', 'xmeas_12', 'xmeas_13', 'xmeas_14', 'xmeas_15', 'xmeas_16',
'xmeas_17', 'xmeas_18', 'xmeas_19', 'xmeas_20', 'xmeas_21', 'xmeas_22', 'xmeas_23', 'xmeas_24',
'xmeas_25', 'xmeas_26', 'xmeas_27', 'xmeas_28', 'xmeas_29', 'xmeas_30', 'xmeas_31', 'xmeas_32',
'xmeas_33', 'xmeas_34', 'xmeas_35', 'xmeas_36', 'xmeas_37', 'xmeas_38', 'xmeas_39', 'xmeas_40',
'xmeas_41', 'xmv_1', 'xmv_2', 'xmv_3', 'xmv_4', 'xmv_5', 'xmv_6', 'xmv_7', 'xmv_8', 'xmv_9',
'xmv_10', 'xmv_11'
]
raw_train = raw_train[~raw_train['faultNumber'].isin([0])]
len(raw_train), raw_train['faultNumber'].nunique()
raw_train['index'] = raw_train['faultNumber'] * 500 + raw_train['simulationRun'] - 1
# raw_test['index'] = raw_test['faultNumber'] * 500 + raw_test['simulationRun'] - 1
simulation_idx = raw_train[['index', 'faultNumber']].drop_duplicates()
X_train_idx, X_val_idx = train_test_split(simulation_idx['index'],
stratify=simulation_idx['faultNumber'],
test_size=0.2,
random_state=42)
X_train = raw_train[raw_train['index'].isin(X_train_idx)].drop('index', axis=1)
X_val = raw_train[raw_train['index'].isin(X_val_idx)].drop('index', axis=1)
# X_test = raw_test.drop('index', axis=1)
# > ### Scaling
# +
scaler = StandardScaler()
scaler.fit(X_train[features])
X_train[features] = scaler.transform(X_train[features])
X_val[features] = scaler.transform(X_val[features])
# X_test[features] = scaler.transform(X_test[features])
# -
# > ### Dataset and dataloader
def correct(y_pred, target):
y_pred = torch.softmax(y_pred, dim=1)
y_pred = torch.max(y_pred, dim=1)[1]
return torch.eq(y_pred, target).sum().item()
# +
# s_list = [0, 100, 200, 300, 400] #len = 5
# l_list = [1, 5, 10, 25, 50, 100] #len = 6
# for idx in range(239, 271):
# fault_sim_idx = idx // (len(s_list) * len(l_list))
# start_length_idxs = idx % (len(s_list) * len(l_list))
# print(fault_sim_idx, start_length_idxs)
# start_idx = s_list[start_length_idxs // len(l_list)]
# seq_length = l_list[start_length_idxs % len(l_list)]
# print("start:", start_idx)
# print("length:", seq_length)
# X_train_tmp = X_train[X_train['index'] == fault_sim_idx].values[start_idx:start_idx+seq_length, :]
# print(X_train_tmp.shape)
# +
class DataTEP(Dataset):
def __init__(self, X):
self.X = X
self.X = self.X.sort_values(['faultNumber', 'simulationRun', 'sample'])
self.X['index'] = self.X.groupby(['faultNumber', 'simulationRun']).ngroup()
self.X = self.X.set_index('index')
self.s_list = [20, 100, 200, 300, 400]
# self.s_list = [450]
# self.l_list = [1, 5, 10, 25, 50, 100]
# self.l_list = [10, 50]
self.l_list = [50]
self.features = [
'xmeas_1', 'xmeas_2', 'xmeas_3', 'xmeas_4', 'xmeas_5', 'xmeas_6', 'xmeas_7', 'xmeas_8', 'xmeas_9',
'xmeas_10', 'xmeas_11', 'xmeas_12', 'xmeas_13', 'xmeas_14', 'xmeas_15', 'xmeas_16', 'xmeas_17',
'xmeas_18', 'xmeas_19', 'xmeas_20', 'xmeas_21', 'xmeas_22', 'xmeas_23', 'xmeas_24', 'xmeas_25',
'xmeas_26', 'xmeas_27', 'xmeas_28', 'xmeas_29', 'xmeas_30', 'xmeas_31', 'xmeas_32', 'xmeas_33',
'xmeas_34', 'xmeas_35', 'xmeas_36', 'xmeas_37', 'xmeas_38', 'xmeas_39', 'xmeas_40', 'xmeas_41',
'xmv_1', 'xmv_2', 'xmv_3', 'xmv_4', 'xmv_5', 'xmv_6', 'xmv_7', 'xmv_8', 'xmv_9', 'xmv_10', 'xmv_11'
]
def __len__(self):
return self.X.index.nunique() * len(self.s_list) * len(self.l_list)
def __getitem__(self, idx):
fault_sim_idx = idx // (len(self.s_list) * len(self.l_list))
start_length_idxs = idx % (len(self.s_list) * len(self.l_list))
start_idx = self.s_list[start_length_idxs // len(self.l_list)]
seq_length = self.l_list[start_length_idxs % len(self.l_list)]
# print(start_idx, start_idx+seq_length)
features = self.X.loc[fault_sim_idx][self.features].values[start_idx : (start_idx+seq_length), :]
target = self.X.loc[fault_sim_idx]['faultNumber'].unique()[0]
features = torch.tensor(features, dtype=torch.float)
target = torch.tensor(target, dtype=torch.long)
return features, target
# -
BATCH_SIZE = 64
NUM_CLASSES = 20
def collate_fn(batch):
sequences = [x[0] for x in batch]
labels = [x[1] for x in batch]
lengths = torch.LongTensor([len(x) for x in sequences])
lengths, idx = lengths.sort(0, descending=True)
sequences = [sequences[i] for i in idx]
labels = torch.tensor(labels, dtype=torch.long)[idx]
sequences_padded = pad_sequence(sequences, batch_first=True)
return sequences_padded, lengths, labels
# +
train_ds = DataTEP(X_train)
train_dl = DataLoader(train_ds, batch_size=BATCH_SIZE, collate_fn=collate_fn, shuffle=True)
val_ds = DataTEP(X_val)
val_dl = DataLoader(val_ds, batch_size=BATCH_SIZE*4, collate_fn=collate_fn)
# test_ds = DataTEP(X_test)
# test_dl = DataLoader(test_ds, batch_size=BATCH_SIZE*4, collate_fn=collate_fn)
# -
len(train_ds), len(val_ds)
gc.collect()
# > ### Model
class TwinModel(torch.nn.Module) :
def __init__(self, NUM_LAYERS, INPUT_SIZE, HIDDEN_SIZE, LINEAR_SIZE, OUTPUT_SIZE, BIDIRECTIONAL, DEVICE):
super().__init__()
self.hidden_size = HIDDEN_SIZE
self.num_layers = NUM_LAYERS
self.input_size = INPUT_SIZE
self.linear_size = LINEAR_SIZE
self.output_size = OUTPUT_SIZE
self.bidirectional = BIDIRECTIONAL
self.lstm_1 = nn.LSTM(
input_size=self.input_size[0],
hidden_size=self.hidden_size,
num_layers=self.num_layers,
bidirectional=self.bidirectional,
batch_first=True,
dropout=0.4
)
self.lstm_2 = nn.LSTM(
input_size=self.input_size[1],
hidden_size=self.hidden_size,
num_layers=self.num_layers,
bidirectional=self.bidirectional,
batch_first=True,
dropout=0.4
)
self.head = nn.Sequential(
nn.Linear(in_features=2*self.hidden_size*(self.bidirectional+1), out_features=self.linear_size),
nn.ReLU(),
nn.Dropout(p=0.4),
nn.Linear(in_features=self.linear_size, out_features=OUTPUT_SIZE),
)
def forward(self, x):
x_1 = x[:, :, :41]
x_2 = x[:, :, 41:]
x_1, _ = self.lstm_1(x_1)
x_2, __ = self.lstm_2(x_2)
x_3 = torch.cat((x_1[:, -1], x_2[:, -1]), dim=-1)
x = self.head(x_3)
return x
class UniModel(torch.nn.Module) :
def __init__(self, NUM_LAYERS, INPUT_SIZE, HIDDEN_SIZE, LINEAR_SIZE, OUTPUT_SIZE, BIDIRECTIONAL, DEVICE):
super().__init__()
self.hidden_size = HIDDEN_SIZE
self.num_layers = NUM_LAYERS
self.input_size = INPUT_SIZE
self.linear_size = LINEAR_SIZE
self.output_size = OUTPUT_SIZE
self.bidirectional = BIDIRECTIONAL
self.lstm = nn.LSTM(
input_size=self.input_size,
hidden_size=self.hidden_size,
num_layers=self.num_layers,
bidirectional=self.bidirectional,
batch_first=True,
dropout=0.4
)
self.head = nn.Sequential(
nn.Linear(in_features=self.hidden_size*(self.bidirectional+1), out_features=self.linear_size),
nn.ReLU(),
nn.Dropout(p=0.4),
nn.Linear(in_features=self.linear_size, out_features=OUTPUT_SIZE),
)
def forward(self, x, x_length):
x_packed = pack_padded_sequence(x, x_length, batch_first=True)
x_lstm_out, _ = self.lstm(x_packed)
x_unpacked, _ = pad_packed_sequence(x_lstm_out, batch_first=True)
x = self.head(x_unpacked[:, -1])
return x
# +
NUM_EPOCHS = 100
LEARNING_RATE = 0.001
NUM_LAYERS = 2
HIDDEN_SIZE = 256
LINEAR_SIZE = 128
BIDIRECTIONAL = True
# +
model = UniModel(NUM_LAYERS=NUM_LAYERS, INPUT_SIZE=52, HIDDEN_SIZE=HIDDEN_SIZE,
LINEAR_SIZE=LINEAR_SIZE, OUTPUT_SIZE=NUM_CLASSES, BIDIRECTIONAL=BIDIRECTIONAL, DEVICE=device)
# model = TwinModel(NUM_LAYERS=NUM_LAYERS, INPUT_SIZE=[41, 11], HIDDEN_SIZE=HIDDEN_SIZE,
# LINEAR_SIZE=LINEAR_SIZE, OUTPUT_SIZE=NUM_CLASSES, BIDIRECTIONAL=BIDIRECTIONAL, DEVICE=device)
model = model.to(device)
optimizer = torch.optim.Adam(model.parameters(), lr=LEARNING_RATE)
criterion = nn.CrossEntropyLoss()
scheduler = StepLR(optimizer, step_size=25, gamma=0.5)
# scheduler = ReduceLROnPlateau(optimizer, mode='min', verbose=True)
torch.manual_seed(42)
# -
for i, (X_batch, X_lengths, y_batch) in enumerate(train_dl):
if i < 1:
print(type(X_batch), type(X_lengths), type(y_batch))
print(len(X_batch), len(X_lengths), len(y_batch))
X_batch, y_batch_train = X_batch.to(device), y_batch.to(device)
y_pred_train = model(X_batch, X_lengths)
print("y_batch_train.size()", y_batch.size())
print("y_pred_train.size()", y_pred_train.size(), '\n')
else:
break
# +
# # !rm -r runs
# -
# > ### Training
# +
# writer = SummaryWriter(comment=f'NL{NUM_LAYERS}_H{HIDDEN_SIZE}_L{LINEAR_SIZE}_B{BIDIRECTIONAL}_S{-100}')
writer = SummaryWriter(comment=f'window_50_only_faults')
# +
loss_train_all, loss_val_all = [], []
accuracy_train_all, accuracy_val_all = [], []
for epoch in range(NUM_EPOCHS):
start = time.time()
print(f'Epoch: {epoch}, Learning Rate: {scheduler.get_last_lr()[0]}\n')
# print(f'Epoch: {epoch}\n')
loss_train_epoch, loss_val_epoch = 0, 0
correct_train_epoch, correct_val_epoch = 0, 0
n_train, n_val = 0, 0
model.train()
for (X_batch_train, X_batch_lengths_train, y_batch_train) in tqdm(train_dl):
X_batch_train, X_batch_lengths_train, y_batch_train =\
X_batch_train.to(device), X_batch_lengths_train.to(device), y_batch_train.to(device)
optimizer.zero_grad()
y_pred_train = model(X_batch_train, X_batch_lengths_train)
loss_train = criterion(y_pred_train, y_batch_train)
loss_train.backward()
optimizer.step()
loss_train_epoch += loss_train.item() * y_batch_train.size()[0]
correct_train_epoch += correct(y_pred_train, y_batch_train)
n_train += y_batch_train.size()[0]
scheduler.step()
model.eval()
with torch.no_grad():
for item in model.parameters():
print(item.grad.mean())
for (X_batch_val, X_batch_lengths_val, y_batch_val) in tqdm(val_dl):
X_batch_val, X_batch_lengths_val, y_batch_val =\
X_batch_val.to(device), X_batch_lengths_val.to(device), y_batch_val.to(device)
y_pred_val = model(X_batch_val, X_batch_lengths_val)
loss_val = criterion(y_pred_val, y_batch_val)
loss_val_epoch += loss_val.item() * y_batch_val.size()[0]
correct_val_epoch += correct(y_pred_val, y_batch_val)
n_val += y_batch_val.size()[0]
loss_mean_train_epoch = loss_train_epoch / n_train
loss_mean_val_epoch = loss_val_epoch / n_val
loss_train_all.append(loss_mean_train_epoch)
loss_val_all.append(loss_mean_val_epoch)
accuracy_train_epoch = correct_train_epoch / n_train
accuracy_val_epoch = correct_val_epoch / n_val
accuracy_train_all.append(accuracy_train_epoch)
accuracy_val_all.append(accuracy_val_epoch)
writer.add_scalars('LOSS per epoch', {"train": loss_mean_train_epoch, "val": loss_mean_val_epoch}, epoch)
writer.add_scalars('ACCURACY per epoch', {"train": accuracy_train_epoch, "val": accuracy_val_epoch}, epoch)
# scheduler.step(loss_mean_val_epoch)
end = time.time()
print(f"epoch time: {end - start}")
print(f"mean loss train: {loss_mean_train_epoch}, mean loss val: {loss_mean_val_epoch}")
print(f"accuracy train: {accuracy_train_epoch}, accuracy val: {accuracy_val_epoch}")
print("---------------------------------------------------------------------------------------------------")
# -
# +
model.eval()
y_ans_val, y_true_val = [], []
with torch.no_grad():
for (X_batch_val, y_batch_val) in tqdm(val_dl):
X_batch_val, y_batch_val = X_batch_val.to(device), y_batch_val.to(device)
y_pred_val = model(X_batch_val)
y_pred_prob = F.softmax(y_pred_val.cpu(), dim=-1)
y_pred_class = y_pred_prob.max(dim=-1)[1]
y_ans_val += y_pred_class.tolist()
y_true_val += y_batch_val.tolist()
# -
plt.figure(figsize=(15, 10))
plt.title("FDR")
sns.heatmap(confusion_matrix(y_true_val, y_ans_val, normalize='pred'), annot=True, cmap=sns.cm.rocket_r)
plt.xlabel('predicted class')
plt.ylabel('true class')
plt.show()
plt.figure(figsize=(20, 8))
plt.title("loss")
plt.plot(np.arange(len(loss_train_all)), loss_train_all, '-o', marker='.', label='train')
plt.plot(np.arange(len(loss_val_all)), loss_val_all, '-o', marker='.', label='val')
plt.legend()
plt.show()
plt.figure(figsize=(20, 8))
plt.title("accuracy")
plt.plot(np.arange(len(accuracy_train_all)), accuracy_train_all, '-o', marker='.', label='train')
plt.plot(np.arange(len(accuracy_val_all)), accuracy_val_all, '-o', marker='.', label='val')
plt.legend()
plt.show()
# # ----------------------------------------------------------------------------------------
|
notebooks/experiments_lstm/medium_article_pytorch_tensorboard_lstm_window_50_cut_start_20_classes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: env_deep_fact
# language: python
# name: env_deep_fact
# ---
# +
import os, sys
import torch
from transformers import GPT2LMHeadModel, GPT2Config
from greenformer import auto_fact
from itertools import chain
from os import path
import sys
# -
def count_param(module, trainable=False):
if trainable:
return sum(p.numel() for p in module.parameters() if p.requires_grad)
else:
return sum(p.numel() for p in module.parameters())
# # Init Model
config = GPT2Config.from_pretrained('gpt2')
model = GPT2LMHeadModel(config=config)
model = GPT2LMHeadModel.from_pretrained('gpt2')
count_param(model)
# # Factorize Model
# ### Apply absolute rank
# %%time
fact_model = auto_fact(model, rank=256, deepcopy=True, solver='random', num_iter=20)
count_param(fact_model)
# %%time
fact_model = auto_fact(model, rank=256, deepcopy=True, solver='svd', num_iter=20)
count_param(fact_model)
# %%time
fact_model = auto_fact(model, rank=256, deepcopy=True, solver='snmf', num_iter=20)
count_param(fact_model)
# ### Apply percentage rank
# %%time
fact_model = auto_fact(model, rank=0.4, deepcopy=True, solver='random', num_iter=20)
count_param(fact_model)
# %%time
fact_model = auto_fact(model, rank=0.4, deepcopy=True, solver='svd', num_iter=20)
count_param(fact_model)
# %%time
fact_model = auto_fact(model, rank=0.4, deepcopy=True, solver='snmf', num_iter=20)
count_param(fact_model)
# + [markdown] tags=[]
# ### Apply factorization only on specific modules
# -
# Only factorize last 6 transformer layers and the pooler layer of the model
factorizable_submodules = list(model.transformer.h[6:])
# %%time
fact_model = auto_fact(model, rank=0.2, deepcopy=True, solver='random', num_iter=20, submodules=factorizable_submodules)
count_param(fact_model)
# %%time
fact_model = auto_fact(model, rank=0.2, deepcopy=True, solver='svd', num_iter=20, submodules=factorizable_submodules)
count_param(fact_model)
# %%time
fact_model = auto_fact(model, rank=0.2, deepcopy=True, solver='snmf', num_iter=20, submodules=factorizable_submodules)
count_param(fact_model)
# # Speed test on CPU
# ### Test Inference CPU
# %%timeit
with torch.no_grad():
y = model(torch.zeros(32,256, dtype=torch.long))
# %%timeit
with torch.no_grad():
y = fact_model(torch.zeros(32,256, dtype=torch.long))
# ### Test Forward-Backward CPU
# %%timeit
y = model(torch.zeros(8,256, dtype=torch.long))
y.logits.sum().backward()
# %%timeit
y = fact_model(torch.zeros(8,256, dtype=torch.long))
y.logits.sum().backward()
# # Speed test on GPU
# ### Move models to GPU
model = model.cuda()
fact_model = fact_model.cuda()
# ### Test Inference GPU
x = torch.zeros(16,256, dtype=torch.long).cuda()
# %%timeit
with torch.no_grad():
y = model(x)
# %%timeit
with torch.no_grad():
y = fact_model(x)
# ### Test Forward-Backward GPU
x = torch.zeros(8,256, dtype=torch.long).cuda()
# %%timeit
y = model(x)
y.logits.sum().backward()
# %%timeit
y = fact_model(x)
y.logits.sum().backward()
|
examples/greenformer_factorize_gpt2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Observations and Insights
#
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as st
import numpy as np
# Study data files
mouse_metadata_path = "data/Mouse_metadata.csv"
study_results_path = "data/Study_results.csv"
# Read the mouse data and the study results
mouse_metadata = pd.read_csv(mouse_metadata_path)
study_results = pd.read_csv(study_results_path)
# Combine the data into a single dataset
master_merge = pd.merge(mouse_metadata, study_results, on="Mouse ID", how="outer")
# Write this out to QA results in Excel
master_merge.to_csv("merge.csv", index=False, header=True)
# Display the data table for preview
master_merge
# -
# Checking the number of mice.
num_mice = master_merge["Mouse ID"].nunique()
num_mice
# Getting the duplicate mice by ID number that shows up for Mouse ID and Timepoint.
# Optional: Get all the data for the duplicate mouse ID.
duplicates = master_merge[master_merge.duplicated(["Mouse ID", "Timepoint"])]
duplicates
# Create a clean DataFrame by dropping the duplicate mouse by its ID.
master_merge.drop(master_merge.loc[master_merge['Mouse ID']=="g989"].index, inplace=True)
# Export to see what I've got
master_merge.to_csv("dropdupes.csv", index=False, header=True)
# Checking the number of mice in the clean DataFrame.
num_clean_mice = master_merge["Mouse ID"].nunique()
num_clean_mice
# Adding a check to see count of records - should be less than original count (we see it removed all 13 rows for g989)
master_merge
# ## Summary Statistics
# +
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume
# for each regimen
# Use groupby and summary statistical methods to calculate the following properties of each drug regimen:
# mean, median, variance, standard deviation, and SEM of the tumor volume.
# Groupby
regimen_group = master_merge.groupby(['Drug Regimen'])
# Calculate values
regimen_mean = regimen_group["Tumor Volume (mm3)"].mean()
regimen_median = regimen_group["Tumor Volume (mm3)"].median()
regimen_var = regimen_group["Tumor Volume (mm3)"].var()
regimen_stdev = regimen_group["Tumor Volume (mm3)"].std()
regimen_SEM = regimen_group["Tumor Volume (mm3)"].sem()
# Assemble the resulting series into a single summary dataframe.
regimen_stats_summary = pd.DataFrame({"Mean":regimen_mean,
"Median": regimen_median,
"Variance": regimen_var,
"Standard Deviation":regimen_stdev,
"SEM":regimen_SEM})
regimen_stats_summary
# +
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume
# for each regimen
# Using the aggregation method, produce the same summary statistics in a single line
regimen_group.agg({"Tumor Volume (mm3)": ["mean", "median", "var", "std", "sem"]})
# -
# ## Bar and Pie Charts
# +
# Generate a bar plot showing the total number of measurements taken on each drug regimen using pandas.
# I used Metastatic Sites field but not sure if there was another way?
reg_count_df = regimen_group["Metastatic Sites"].count().to_frame()
# Use DataFrame.plot() in order to create a bar chart of the data
reg_count_df.plot(kind="bar",figsize=(10,3))
plt.title("Total Number of Measurements by Drug Regimen")
plt.show()
# +
# Generate a bar plot showing the total number of measurements taken on each drug regimen using pyplot.
# Um definitely a more systematic way of doing this!?!?!
# And need to space out things better!
reg_count_df
#df = pd.DataFrame(Data,columns=['Country','GDP_Per_Capita'])
#New_Colors = ['green','blue','purple','brown','teal']
#plt.bar(reg_count_df['Drug Regimen'], reg_count_df['Metastatic Sites'], color=New_Colors)
#plt.title('Sites by drug regimen', fontsize=14)
#plt.xlabel('Regimen', fontsize=14)
#plt.ylabel('Num of sites', fontsize=14)
#plt.grid(True)
#plt.show()
# -
# Generate a pie plot showing the distribution of female versus male mice using pandas
regimen_group = master_merge.groupby(['Sex']).count().plot(kind="pie", y="Metastatic Sites",autopct='%1.1f%%')
# +
# Generate a pie plot showing the distribution of female versus male mice using pyplot
# -
# ## Quartiles, Outliers and Boxplots
# +
# Calculate the final tumor volume of each mouse across four of the treatment regimens:
# Capomulin, Ramicane, Infubinol, and Ceftamin
# Start by getting the last (greatest) timepoint for each mouse
# Merge this group df with the original dataframe to get the tumor volume at the last timepoint
# +
# Put treatments into a list for for loop (and later for plot labels)
# Create empty list to fill with tumor vol data (for plotting)
# Calculate the IQR and quantitatively determine if there are any potential outliers.
# Locate the rows which contain mice on each drug and get the tumor volumes
# add subset
# Determine outliers using upper and lower bounds
# -
# Generate a box plot of the final tumor volume of each mouse across four regimens of interest
# ## Line and Scatter Plots
# Generate a line plot of tumor volume vs. time point for a mouse treated with Capomulin
# Generate a scatter plot of average tumor volume vs. mouse weight for the Capomulin regimen
# ## Correlation and Regression
# Calculate the correlation coefficient and linear regression model
# for mouse weight and average tumor volume for the Capomulin regimen
|
Pymaceuticals/pymaceuticals_27Jan2021.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <a href="https://cognitiveclass.ai"><img src = "https://ibm.box.com/shared/static/9gegpsmnsoo25ikkbl4qzlvlyjbgxs5x.png" width = 400> </a>
#
# <h1 align=center><font size = 5>Learning FourSquare API with Python</font></h1>
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ## Introduction
#
# In this lab, you will learn in details how to make calls to the Foursquare API for different purposes. You will learn how to construct a URL to send a request to the API to search for a specific type of venues, to explore a particular venue, to explore a Foursquare user, to explore a geographical location, and to get trending venues around a location. Also, you will learn how to use the visualization library, Folium, to visualize the results.
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ## Table of Contents
#
# 1. <a href="#item1">Foursquare API Search Function</a>
# 2. <a href="#item2">Explore a Given Venue</a>
# 3. <a href="#item3">Explore a User</a>
# 4. <a href="#item4">Foursquare API Explore Function</a>
# 5. <a href="#item5">Get Trending Venues</a>
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ### Import necessary Libraries
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
import requests # library to handle requests
import pandas as pd # library for data analsysis
import numpy as np # library to handle data in a vectorized manner
import random # library for random number generation
# !conda install -c conda-forge geopy --yes
from geopy.geocoders import Nominatim # module to convert an address into latitude and longitude values
# libraries for displaying images
from IPython.display import Image
from IPython.core.display import HTML
# tranforming json file into a pandas dataframe library
from pandas.io.json import json_normalize
# !conda install -c conda-forge folium=0.5.0 --yes
import folium # plotting library
print('Folium installed')
print('Libraries imported.')
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ### Define Foursquare Credentials and Version
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ##### Make sure that you have created a Foursquare developer account and have your credentials handy
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
CLIENT_ID = 'your-client-ID' # your Foursquare ID
CLIENT_SECRET = 'your-client-secret' # your Foursquare Secret
VERSION = '20180604'
LIMIT = 30
print('Your credentails:')
print('CLIENT_ID: ' + CLIENT_ID)
print('CLIENT_SECRET:' + CLIENT_SECRET)
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Let's again assume that you are staying at the Conrad hotel. So let's start by converting the Contrad Hotel's address to its latitude and longitude coordinates.
# -
# In order to define an instance of the geocoder, we need to define a user_agent. We will name our agent <em>foursquare_agent</em>, as shown below.
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
address = '102 North End Ave, New York, NY'
geolocator = Nominatim(user_agent="foursquare_agent")
location = geolocator.geocode(address)
latitude = location.latitude
longitude = location.longitude
print(latitude, longitude)
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <a id="item1"></a>
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ## 1. Search for a specific venue category
# > `https://api.foursquare.com/v2/venues/`**search**`?client_id=`**CLIENT_ID**`&client_secret=`**CLIENT_SECRET**`&ll=`**LATITUDE**`,`**LONGITUDE**`&v=`**VERSION**`&query=`**QUERY**`&radius=`**RADIUS**`&limit=`**LIMIT**
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Now, let's assume that it is lunch time, and you are craving Italian food. So, let's define a query to search for Italian food that is within 500 metres from the Conrad Hotel.
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
search_query = 'Italian'
radius = 500
print(search_query + ' .... OK!')
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Define the corresponding URL
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
url = 'https://api.foursquare.com/v2/venues/search?client_id={}&client_secret={}&ll={},{}&v={}&query={}&radius={}&limit={}'.format(CLIENT_ID, CLIENT_SECRET, latitude, longitude, VERSION, search_query, radius, LIMIT)
url
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Send the GET Request and examine the results
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
results = requests.get(url).json()
results
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Get relevant part of JSON and transform it into a *pandas* dataframe
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# assign relevant part of JSON to venues
venues = results['response']['venues']
# tranform venues into a dataframe
dataframe = json_normalize(venues)
dataframe.head()
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Define information of interest and filter dataframe
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# keep only columns that include venue name, and anything that is associated with location
filtered_columns = ['name', 'categories'] + [col for col in dataframe.columns if col.startswith('location.')] + ['id']
dataframe_filtered = dataframe.loc[:, filtered_columns]
# function that extracts the category of the venue
def get_category_type(row):
try:
categories_list = row['categories']
except:
categories_list = row['venue.categories']
if len(categories_list) == 0:
return None
else:
return categories_list[0]['name']
# filter the category for each row
dataframe_filtered['categories'] = dataframe_filtered.apply(get_category_type, axis=1)
# clean column names by keeping only last term
dataframe_filtered.columns = [column.split('.')[-1] for column in dataframe_filtered.columns]
dataframe_filtered
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Let's visualize the Italian restaurants that are nearby
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
dataframe_filtered.name
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
venues_map = folium.Map(location=[latitude, longitude], zoom_start=13) # generate map centred around the Conrad Hotel
# add a red circle marker to represent the Conrad Hotel
folium.features.CircleMarker(
[latitude, longitude],
radius=10,
color='red',
popup='Conrad Hotel',
fill = True,
fill_color = 'red',
fill_opacity = 0.6
).add_to(venues_map)
# add the Italian restaurants as blue circle markers
for lat, lng, label in zip(dataframe_filtered.lat, dataframe_filtered.lng, dataframe_filtered.categories):
folium.features.CircleMarker(
[lat, lng],
radius=5,
color='blue',
popup=label,
fill = True,
fill_color='blue',
fill_opacity=0.6
).add_to(venues_map)
# display map
venues_map
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <a id="item2"></a>
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ## 2. Explore a Given Venue
# > `https://api.foursquare.com/v2/venues/`**VENUE_ID**`?client_id=`**CLIENT_ID**`&client_secret=`**CLIENT_SECRET**`&v=`**VERSION**
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ### A. Let's explore the closest Italian restaurant -- _Harry's Italian Pizza Bar_
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
venue_id = '4fa862b3e4b0ebff2f749f06' # ID of Harry's Italian Pizza Bar
url = 'https://api.foursquare.com/v2/venues/{}?client_id={}&client_secret={}&v={}'.format(venue_id, CLIENT_ID, CLIENT_SECRET, VERSION)
url
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Send GET request for result
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
result = requests.get(url).json()
print(result['response']['venue'].keys())
result['response']['venue']
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ### B. Get the venue's overall rating
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
try:
print(result['response']['venue']['rating'])
except:
print('This venue has not been rated yet.')
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# That is not a very good rating. Let's check the rating of the second closest Italian restaurant.
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
venue_id = '4f3232e219836c91c7bfde94' # ID of Conca Cucina Italian Restaurant
url = 'https://api.foursquare.com/v2/venues/{}?client_id={}&client_secret={}&v={}'.format(venue_id, CLIENT_ID, CLIENT_SECRET, VERSION)
result = requests.get(url).json()
try:
print(result['response']['venue']['rating'])
except:
print('This venue has not been rated yet.')
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Since this restaurant has no ratings, let's check the third restaurant.
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
venue_id = '3fd66200f964a520f4e41ee3' # ID of Ecco
url = 'https://api.foursquare.com/v2/venues/{}?client_id={}&client_secret={}&v={}'.format(venue_id, CLIENT_ID, CLIENT_SECRET, VERSION)
result = requests.get(url).json()
try:
print(result['response']['venue']['rating'])
except:
print('This venue has not been rated yet.')
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Since this restaurant has a slightly better rating, let's explore it further.
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ### C. Get the number of tips
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
result['response']['venue']['tips']['count']
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ### D. Get the venue's tips
# > `https://api.foursquare.com/v2/venues/`**VENUE_ID**`/tips?client_id=`**CLIENT_ID**`&client_secret=`**CLIENT_SECRET**`&v=`**VERSION**`&limit=`**LIMIT**
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Create URL and send GET request. Make sure to set limit to get all tips
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
## Ecco Tips
limit = 15 # set limit to be greater than or equal to the total number of tips
url = 'https://api.foursquare.com/v2/venues/{}/tips?client_id={}&client_secret={}&v={}&limit={}'.format(venue_id, CLIENT_ID, CLIENT_SECRET, VERSION, limit)
results = requests.get(url).json()
results
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Get tips and list of associated features
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
tips = results['response']['tips']['items']
tip = results['response']['tips']['items'][0]
tip.keys()
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Format column width and display all tips
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
pd.set_option('display.max_colwidth', -1)
tips_df = json_normalize(tips) # json normalize tips
# columns to keep
filtered_columns = ['text', 'agreeCount', 'disagreeCount', 'id', 'user.firstName', 'user.lastName', 'user.gender', 'user.id']
tips_filtered = tips_df.loc[:, filtered_columns]
# display tips
tips_filtered
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Now remember that because we are using a personal developer account, then we can access only 2 of the restaurant's tips, instead of all 15 tips.
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <a id="item3"></a>
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ## 3. Search a Foursquare User
# > `https://api.foursquare.com/v2/users/`**USER_ID**`?client_id=`**CLIENT_ID**`&client_secret=`**CLIENT_SECRET**`&v=`**VERSION**
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ### Define URL, send GET request and display features associated with user
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
user_id = '484542633' # user ID with most agree counts and complete profile
url = 'https://api.foursquare.com/v2/users/{}?client_id={}&client_secret={}&v={}'.format(user_id, CLIENT_ID, CLIENT_SECRET, VERSION) # define URL
# send GET request
results = requests.get(url).json()
user_data = results['response']['user']
# display features associated with user
user_data.keys()
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
print('First Name: ' + user_data['firstName'])
print('Last Name: ' + user_data['lastName'])
print('Home City: ' + user_data['homeCity'])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### How many tips has this user submitted?
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
user_data['tips']
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Wow! So it turns out that Nick is a very active Foursquare user, with more than 250 tips.
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ### Get User's tips
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# define tips URL
url = 'https://api.foursquare.com/v2/users/{}/tips?client_id={}&client_secret={}&v={}&limit={}'.format(user_id, CLIENT_ID, CLIENT_SECRET, VERSION, limit)
# send GET request and get user's tips
results = requests.get(url).json()
tips = results['response']['tips']['items']
# format column width
pd.set_option('display.max_colwidth', -1)
tips_df = json_normalize(tips)
# filter columns
filtered_columns = ['text', 'agreeCount', 'disagreeCount', 'id']
tips_filtered = tips_df.loc[:, filtered_columns]
# display user's tips
tips_filtered
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Let's get the venue for the tip with the greatest number of agree counts
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
tip_id = '5ab5575d73fe2516ad8f363b' # tip id
# define URL
url = 'http://api.foursquare.com/v2/tips/{}?client_id={}&client_secret={}&v={}'.format(tip_id, CLIENT_ID, CLIENT_SECRET, VERSION)
# send GET Request and examine results
result = requests.get(url).json()
print(result['response']['tip']['venue']['name'])
print(result['response']['tip']['venue']['location'])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ### Get User's friends
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
user_friends = json_normalize(user_data['friends']['groups'][0]['items'])
user_friends
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Interesting. Despite being very active, it turns out that Nick does not have any friends on Foursquare. This might definitely change in the future.
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ### Retrieve the User's Profile Image
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
user_data
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# 1. grab prefix of photo
# 2. grab suffix of photo
# 3. concatenate them using the image size
Image(url='https://igx.4sqi.net/img/user/300x300/484542633_mK2Yum7T_7Tn9fWpndidJsmw2Hof_6T5vJBKCHPLMK5OL-U5ZiJGj51iwBstcpDLYa3Zvhvis.jpg')
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <a id="item4"></a>
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ## 4. Explore a location
# > `https://api.foursquare.com/v2/venues/`**explore**`?client_id=`**CLIENT_ID**`&client_secret=`**CLIENT_SECRET**`&ll=`**LATITUDE**`,`**LONGITUDE**`&v=`**VERSION**`&limit=`**LIMIT**
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### So, you just finished your gourmet dish at Ecco, and are just curious about the popular spots around the restaurant. In order to explore the area, let's start by getting the latitude and longitude values of Ecco Restaurant.
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
latitude = 40.715337
longitude = -74.008848
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Define URL
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
url = 'https://api.foursquare.com/v2/venues/explore?client_id={}&client_secret={}&ll={},{}&v={}&radius={}&limit={}'.format(CLIENT_ID, CLIENT_SECRET, latitude, longitude, VERSION, radius, LIMIT)
url
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Send GET request and examine results
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
import requests
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
results = requests.get(url).json()
'There are {} around Ecco restaurant.'.format(len(results['response']['groups'][0]['items']))
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Get relevant part of JSON
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
items = results['response']['groups'][0]['items']
items[0]
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Process JSON and convert it to a clean dataframe
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
dataframe = json_normalize(items) # flatten JSON
# filter columns
filtered_columns = ['venue.name', 'venue.categories'] + [col for col in dataframe.columns if col.startswith('venue.location.')] + ['venue.id']
dataframe_filtered = dataframe.loc[:, filtered_columns]
# filter the category for each row
dataframe_filtered['venue.categories'] = dataframe_filtered.apply(get_category_type, axis=1)
# clean columns
dataframe_filtered.columns = [col.split('.')[-1] for col in dataframe_filtered.columns]
dataframe_filtered.head(10)
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Let's visualize these items on the map around our location
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
venues_map = folium.Map(location=[latitude, longitude], zoom_start=15) # generate map centred around Ecco
# add Ecco as a red circle mark
folium.features.CircleMarker(
[latitude, longitude],
radius=10,
popup='Ecco',
fill=True,
color='red',
fill_color='red',
fill_opacity=0.6
).add_to(venues_map)
# add popular spots to the map as blue circle markers
for lat, lng, label in zip(dataframe_filtered.lat, dataframe_filtered.lng, dataframe_filtered.categories):
folium.features.CircleMarker(
[lat, lng],
radius=5,
popup=label,
fill=True,
color='blue',
fill_color='blue',
fill_opacity=0.6
).add_to(venues_map)
# display map
venues_map
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <a id="item5"></a>
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ## 5. Explore Trending Venues
# > `https://api.foursquare.com/v2/venues/`**trending**`?client_id=`**CLIENT_ID**`&client_secret=`**CLIENT_SECRET**`&ll=`**LATITUDE**`,`**LONGITUDE**`&v=`**VERSION**
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# #### Now, instead of simply exploring the area around Ecco, you are interested in knowing the venues that are trending at the time you are done with your lunch, meaning the places with the highest foot traffic. So let's do that and get the trending venues around Ecco.
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# define URL
url = 'https://api.foursquare.com/v2/venues/trending?client_id={}&client_secret={}&ll={},{}&v={}'.format(CLIENT_ID, CLIENT_SECRET, latitude, longitude, VERSION)
# send GET request and get trending venues
results = requests.get(url).json()
results
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ### Check if any venues are trending at this time
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
if len(results['response']['venues']) == 0:
trending_venues_df = 'No trending venues are available at the moment!'
else:
trending_venues = results['response']['venues']
trending_venues_df = json_normalize(trending_venues)
# filter columns
columns_filtered = ['name', 'categories'] + ['location.distance', 'location.city', 'location.postalCode', 'location.state', 'location.country', 'location.lat', 'location.lng']
trending_venues_df = trending_venues_df.loc[:, columns_filtered]
# filter the category for each row
trending_venues_df['categories'] = trending_venues_df.apply(get_category_type, axis=1)
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# display trending venues
trending_venues_df
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Now, depending on when you run the above code, you might get different venues since the venues with the highest foot traffic are fetched live.
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ### Visualize trending venues
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
if len(results['response']['venues']) == 0:
venues_map = 'Cannot generate visual as no trending venues are available at the moment!'
else:
venues_map = folium.Map(location=[latitude, longitude], zoom_start=15) # generate map centred around Ecco
# add Ecco as a red circle mark
folium.features.CircleMarker(
[latitude, longitude],
radius=10,
popup='Ecco',
fill=True,
color='red',
fill_color='red',
fill_opacity=0.6
).add_to(venues_map)
# add the trending venues as blue circle markers
for lat, lng, label in zip(trending_venues_df['location.lat'], trending_venues_df['location.lng'], trending_venues_df['name']):
folium.features.CircleMarker(
[lat, lng],
radius=5,
poup=label,
fill=True,
color='blue',
fill_color='blue',
fill_opacity=0.6
).add_to(venues_map)
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
# display map
venues_map
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <a id="item6"></a>
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ### Thank you for completing this lab!
#
# This notebook was created by [<NAME>](https://www.linkedin.com/in/aklson/). I hope you found this lab interesting and educational. Feel free to contact me if you have any questions!
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# This notebook is part of a course on **Coursera** called *Applied Data Science Capstone*. If you accessed this notebook outside the course, you can take this course online by clicking [here](http://cocl.us/DP0701EN_Coursera_Week2_LAB1).
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# <hr>
# Copyright © 2018 [Cognitive Class](https://cognitiveclass.ai/?utm_source=bducopyrightlink&utm_medium=dswb&utm_campaign=bdu). This notebook and its source code are released under the terms of the [MIT License](https://bigdatauniversity.com/mit-license/).
|
Course-9 Applied Data Science Capstone/Labs/DP0701EN-2-2-1-Foursquare-API-py-v1.0.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Torch)
# language: python
# name: torch
# ---
import sys
sys.path.insert(0, "../..")
# +
import numpy as np
import torch
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from logistic_regression import LogisticRegressionGD
from extrapolation import *
from experiments import Experiment
# +
device = "cpu"
X = []
with open("../../../madelon_train.data") as f:
for line in f:
X.append([float(x) for x in line.split()])
X = np.array(X)
X /= X.max()
X = np.concatenate([np.ones((X.shape[0], 1)), X], axis=1)
X = torch.tensor(X, device=device)
with open("../../../madelon_train.labels") as f:
y = [int(x) for x in f]
y = torch.tensor(y, device=device)
X.shape, y.shape
# -
np.random.seed(2020)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.1)
# $\tau = 10^7$
model = LogisticRegressionGD(X_train, y_train, 1e7, device=device)
model.run_steps(5000)
preds = model.predict(X_test)
torch.mean((preds == y_test).double())
len(model.log)
k = 6
experiment = Experiment(model.log, model.obj, values=model.value_log, device=device)
experiment.run_method("RRE+QR", RRE, k, method_kwargs={"qr": True})
experiment.run_method("RNA", RNA, k, method_kwargs={"lambda_range": (1e-15, 1e-2), "linesearch": False, "norm": False})
experiment.run_method("RNA+norm", RNA, k, method_kwargs={"lambda_range": (1e-15, 1e-2), "linesearch": False})
experiment.run_method("RNA+ls", RNA, k, method_kwargs={"lambda_range": (1e-15, 1e-2), "linesearch": True})
plt.figure(figsize=(14, 8))
experiment.plot_values(n=2000)
plt.legend()
plt.figure(figsize=(14, 8))
experiment.plot_log_diff(n=2000)
plt.legend()
model.theta = experiment.best_x
preds = model.predict(X_test)
torch.mean((preds == y_test).double())
df = experiment.value_df()
df.to_csv(f"results/madelon:tau=1e7.csv")
# $\tau = 10^2$
model = LogisticRegressionGD(X_train, y_train, 1e2, device=device)
model.fit(1e-8, max_iter=100500)
len(model.log)
k = 6
experiment = Experiment(model.log, model.obj, values=model.value_log, device=device)
experiment.run_method("RRE+QR", RRE, k, method_kwargs={"qr": True})
experiment.run_method("Regularized RRE", regularized_RRE, k, method_kwargs={"lambda_": 1e-15})
experiment.run_method("RNA", RNA, k, method_kwargs={"lambda_range": (1e-15, 1e-2), "linesearch": False, "norm": False})
experiment.run_method("RNA+norm", RNA, k, method_kwargs={"lambda_range": (1e-15, 1e-2), "linesearch": False})
experiment.run_method("RNA+ls", RNA, k, method_kwargs={"lambda_range": (1e-15, 1e-2), "linesearch": True})
plt.figure(figsize=(14, 8))
experiment.plot_values(n=1000)
plt.ylim(1240, 1250)
plt.legend()
plt.figure(figsize=(14, 8))
experiment.plot_log_diff(n=1000)
plt.legend()
model.theta = experiment.best_x
preds = model.predict(X_test)
torch.mean((preds == y_test).double())
df = experiment.value_df()
df.to_csv(f"results/madelon:tau=1e2.csv")
# $\tau = 10^{-3}$
model = LogisticRegressionGD(X_train, y_train, 1e-3, device=device)
model.fit(1e-8, max_iter=100500)
len(model.log)
k = 6
experiment = Experiment(model.log, model.obj, values=model.value_log, device=device)
experiment.run_method("RRE+QR", RRE, k, method_kwargs={"qr": True})
experiment.run_method("Regularized RRE", regularized_RRE, k, method_kwargs={"lambda_": 1e-10})
experiment.run_method("RNA", RNA, k, method_kwargs={"lambda_range": (1e-15, 1e-2), "linesearch": False, "norm": False})
experiment.run_method("RNA+norm", RNA, k, method_kwargs={"lambda_range": (1e-15, 1e-2), "linesearch": False})
experiment.run_method("RNA+ls", RNA, k, method_kwargs={"lambda_range": (1e-15, 1e-2), "linesearch": True})
plt.figure(figsize=(14, 8))
experiment.plot_values(n=10000)
plt.ylim(1000, 1300)
plt.legend()
plt.figure(figsize=(14, 8))
experiment.plot_log_diff(n=10000)
plt.ylim(None, 3)
plt.legend()
model.theta = experiment.best_x
preds = model.predict(X_test)
torch.mean((preds == y_test).double())
df = experiment.value_df()
df.to_csv(f"results/madelon:tau=1e-3.csv")
|
notebooks/logistic regression/Madelon.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small><small><i>
# All the IPython Notebooks in this lecture series by Dr. <NAME> are available @ **[GitHub](https://github.com/milaan9/10_Python_Pandas_Module)**
# </i></small></small>
# # Python Pandas
#
# **[Pandas](https://pandas.pydata.org/)** is an open-source, BSD-licensed Python library. Pandas is a handy and useful data-structure tool for analyzing large and complex data.
# In this exercise, we are using **[pokemon_data.csv](https://github.com/milaan9/10_Python_Pandas_Module/blob/main/pokemon_data.csv)** for data analysis. This Dataset has different characteristics of an auto such as body-style, wheel-base, engine-type, price, mileage, horsepower, etc.
# ## Loading data into Pandas
# +
import pandas as pd
df = pd.read_csv('pokemon_data.csv') # just give the name of file only if the file is in the same folder.
# print(df.head(5))
# print(df.tail(5))
# df_xlsx = pd.read_excel('pokemon_data.xlsx')
# print(df_xlsx.head(3))
# df = pd.read_csv('pokemon_data.txt', delimiter='\t') # without delimiter the data will look like mess.
# print(df.head(5))
df['HP']
# -
# ## Reading Data in Pandas
# +
#### Read Headers
df.columns
## Read each Column
#print(df[['Name', 'Type 1', 'HP']])
## Read Each Row
#print(df.iloc[0:4])
# for index, row in df.iterrows():
# print(index, row['Name'])
#df.loc[df['Type 1'] == "Grass"]
## Read a specific location (R,C)
#print(df.iloc[2,1])
# -
# ## Sorting/Describing Data
# +
df
df.sort_values('Name') #arrange acco to Name starting with 'A'
#df.sort_values('Name', ascending=False) #arrange acco to Name starting with 'Z'
# -
df.sort_values(['Type 1', 'HP'], ascending=[1,0]) # 'Type1' is ascending and 'HP' is decending
# ## Making changes to the data
df.head(5)
# +
df['Total'] = df['HP'] + df['Attack'] + df['Defense'] + df['Sp. Atk'] + df['Sp. Def'] + df['Speed']
df.head(5)
# -
45+49+49+65+65+45
# +
#df['Total'] = df['HP'] + df['Attack'] + df['Defense'] + df['Sp. Atk'] + df['Sp. Def'] + df['Speed']
df = df.drop(columns=['Total'])
df['Total'] = df.iloc[:, 4:10].sum(axis=1) # axis=0 column wise and axis=1 row wise
df
# +
cols = list(df.columns)
df = df[cols[0:4] + [cols[-1]]+cols[4:12]]
df.head(5)
# -
# ## Saving our Data (Exporting into Desired Format)
df
# +
df.to_csv('modified.csv', index=False)
#df.to_excel('modified.xlsx', index=False)
df.to_csv('modified.txt', index=False, sep='\t')
# -
# ## Filtering Data
# +
new_df = df.loc[(df['Type 1'] == 'Grass') & (df['Type 2'] == 'Poison') & (df['HP'] > 70)]
new_df
# -
df.loc[df['Name'].str.contains('Mega')]
df.loc[~df['Name'].str.contains('Mega')]
# +
import re
# df.loc[df['Type 1'].str.contains('Fire|Grass', regex = True)]
df.loc[df['Type 1'].str.contains('fire|grass', flags=re.I, regex=True)]
# +
# To list all the names starating wirh 'pi'
import re
df.loc[df['Name'].str.contains('pi[a-z]*', flags=re.I, regex=True)] # '*' means zero or more
#df.loc[df['Name'].str.contains('^pi[a-z]*', flags=re.I, regex=True)] # '^' means beginning
# +
new_df.reset_index(drop=True, inplace=True)
new_df
# -
new_df.to_csv('filtered.csv')
#
# ## Conditional Changes
# +
# TO change 'Fire' category of 'Type 1' to 'Flamer'
df.loc[df['Type 1'] == 'Fire','Type 1' ] = 'Flamer'
df
# +
# To chaange it back
df.loc[df['Type 1'] == 'Flamer','Type 1' ] = 'Fire'
df
# +
# To make all 'Fire' pokemon 'Legandary'
df.loc[df['Type 1'] == 'Fire','Legandary' ] = True
df
# -
df.loc[df['Total'] > 500, ['Generation','Legendary']] = 'TEST VALUE'
df
df.loc[df['Total'] > 500, ['Generation','Legendary']] = ['Test 1', 'Test 2']
df
# So far we have messed up our dataframe so lets go back to our checkpoint 'modified.csv'
# +
df = pd.read_csv('modified.csv')
df
# -
# ## Aggregate Statistics (Groupby)
#
# +
df = pd.read_csv('modified.csv')
df.groupby(['Type 1']).mean() # Computing avg attack of 'Type 1' pokemon
#df.groupby(['Type 1']).mean().sort_values('Defense', ascending=False) # computing avg defense
# +
df = pd.read_csv('modified.csv')
df['count'] = 1
df.groupby(['Type 1']).count()['count']
#df.groupby(['Type 1', 'Type 2']).count()['count']
# -
# ## Working with large amounts of data
#
#
for df in pd.read_csv('modified.csv', chunksize=5):
print("CHUNK DF")
print(df)
# +
new_df = pd.DataFrame(columns=df.columns)
for df in pd.read_csv('modified.csv', chunksize=5):
results = df.groupby(['Type 1']).count()
new_df = pd.concat([new_df, results])
# -
|
002_Python_Pandas_Exercise_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.11 ('gurobi')
# language: python
# name: python3
# ---
import pandas as pd
import os
import geopandas as gpd
import geoplot
import geoplot.crs as gcrs
import matplotlib.pyplot as plt
from shapely.geometry import Point
import folium
import rasterio
os.chdir('c:\\Users\\Jesse\\OneDrive\\Documenten\\Master BAOR\\Thesis\\GitHub\\dicra\\analytics\\geospatial_internship\\datasets')
fire_data = pd.read_csv('telangana_fires.csv')
fire_data
os.chdir('c:\\Users\\Jesse\\OneDrive\\Documenten\\Master BAOR\\Thesis\\GitHub\\dicra\\analytics\\notebooks\\relative_wealth_index\\data')
rwi_data = pd.read_csv('ind_pak_relative_wealth_index.csv')
rwi_data.head()
os.chdir('c:\\Users\\Jesse\\OneDrive\\Documenten\\Master BAOR\\Thesis\\GitHub\\dicra\\analytics\\notebooks\\relative_wealth_index\\data\\gadm40_IND_shp')
shape = gpd.read_file('gadm40_IND_2.shp')
shape.plot()
plt.show()
os.chdir('c:\\Users\\Jesse\\OneDrive\\Documenten\\Master BAOR\\Thesis\\GitHub\\dicra\\analytics\\notebooks\\relative_wealth_index\\result')
pd.read_excel('rwi_median.xlsx').head()
os.chdir('c:\\Users\\Jesse\\OneDrive\\Documenten\\Master BAOR\\Thesis\\GitHub\\dicra\\analytics\\notebooks\\infrastructure')
warehouse_data = pd.read_csv('ts-warehouse-data_march2022.csv')
warehouse_data
os.chdir('c:\\Users\\Jesse\\OneDrive\\Documenten\\Master BAOR\\Thesis\\GitHub\\dicra\\analytics\\notebooks\\market_yards')
market_yards_data = pd.read_excel('list_market_yards_2021.xlsx')
market_yards_data.head()
market_yards_list_data = pd.read_excel('market_yard_list.xlsx')
market_yards_list_data.head()
market_yard_data = pd.read_csv('market_yard_data.csv')
market_yard_data.head()
day_prices = pd.read_csv('day_prices_between_01-12-2021_31-12-2021.csv')
day_prices.head()
market_prices = pd.read_csv('market_yard_prices_01012019_26062019.csv')
market_prices.head()
os.chdir('c:\\Users\\Jesse\\OneDrive\\Documenten\\Master BAOR\\Thesis\\GitHub\\dicra\\analytics\\notebooks\\crop_fires')
gpd.read_file('telanganafire.geojson')
# Function that defines the pixel color in the TIF file
def getCoordinatePixel(map,lon,lat):
# open map
dataset = rasterio.open(map)
a = dataset.read(1)
# get pixel x+y of the coordinate
py, px = dataset.index(lon, lat)
return a[px,py]
os.chdir('c:\\Users\\Jesse\\OneDrive\\Documenten\\Master BAOR\\Thesis\\GitHub\\dicra\\analytics\\notebooks\\crop_fires')
getCoordinatePixel('01-01-2017.tif', fire_data['longitude'].iloc[0], fire_data['latitude'].iloc[0])
dataset = rasterio.open('01-01-2017.tif')
dataset.read().shape
# +
data = gpd.read_file('telangana_shapefile.geojson')
#geoplot.polyplot(data, projection=gcrs.AlbersEqualArea(), edgecolor='darkgrey', facecolor='lightgrey', linewidth=.3, figsize=(12, 8))
#plt.show()
# +
#geoplot.polyplot(data, projection=gcrs.AlbersEqualArea(), edgecolor='darkgrey', facecolor='lightgrey', linewidth=.3, figsize=(12, 8))
#plt.scatter(fires_data.longitude, fires_data.latitude)
#plt.xlabel('Longitude')
#plt.ylabel('Latitude')
#plt.show()
# +
#data.loc[0, 'geometry']
# -
data.plot(column='NAME_1', legend=True)
plt.show()
data.plot()
plt.scatter(fires_data.longitude, fires_data.latitude, c='red', edgecolor = 'black')
plt.xlabel('Longitude')
plt.ylabel('Latitude')
plt.show()
fires_data['geometry'] = fires_data.apply(lambda x: Point((x.longitude, x.latitude)), axis=1)
fires_data.head()
fires_data_crs = {'init': 'epsg:4326'}
fires_data_geo = gpd.GeoDataFrame(fires_data, crs = fires_data_crs, geometry = fires_data.geometry)
fires_data_geo.head(3)
#Convert geometry from decimal degrees to meters
fires_data_geo.geometry = fires_data_geo.geometry.to_crs(epsg = 3857)
fires_data_geo.head()
# +
#Interested arguments:
# gpdsjoin( op = 'contains' 'within' or 'intersects') & within_gdf.shape[0])
# +
#Centroid:
print(data.geometry[0].area)
data_m = data.geometry.to_crs(epsg = 3857)
print(data_m.geometry[0].area/10000, 'squared km')
#Distances:
fire1 = fires_data.geometry[0]
fire2 = fires_data.geometry[1]
print(fire1.distance(other = fire2))
# -
fire_map = folium.Map(location = [fires_data['latitude'].iloc[0], fires_data['longitude'].iloc[0]], zoom_start = 5)
folium.GeoJson(data.geometry).add_to(fire_map)
popup = '<strong> Fire 1 </strong>'
marker = folium.Marker(location = [fires_data['latitude'].iloc[0], fires_data['longitude'].iloc[0]], popup = popup)
marker.add_to(fire_map)
display(fire_map)
# +
os.chdir('c:\\Users\\Jesse\\OneDrive\\Documenten\\Master BAOR\\Thesis\\GitHub\\dicra\\src\\data_preprocessing\\tsdm\\')
data2 = gpd.read_file('District_Boundary.shp')
data3 = gpd.read_file('Mandal_Boundary.shp')
data2.head(3)
# +
#Choropleth
#gpd.GeoDataFrame(, crs = , geometry = )
mandal_counts= data3.groupby(['Dist_Name']).size()
mandal_counts_df = mandal_counts.to_frame()
mandal_counts_df = mandal_counts_df.reset_index()
mandal_counts_df.columns = ['Dist_Name', 'Amount of Mandals']
new_geo = pd.merge(data3, mandal_counts_df, on = 'Dist_Name')
new_geo['area'] = new_geo.geometry.area
new_geo['normalized value'] = new_geo.apply(lambda row: row['Amount of Mandals']/row['area'], axis = 1)
new_geo.plot(column = 'normalized value', cmap = 'BuGn', edgecolor = 'black', legend = True)
plt.xlabel('Longitude')
plt.ylabel('latitude')
plt.show()
|
analytics/notebooks/crop_fires/Exploration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/NoCodeProgram/CodingTest/blob/main/heap/findMedianStream.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="UcoH6ZgSE4SR"
# Title : Find Median from Data Stream
#
# Chapter : Heap
#
# Link :
#
# ChapterLink :
#
# 문제 : Data Stream에서 median 을 찾는 class를 만들어라
# + id="RuHjOlxME3IW"
import heapq
class MedianFinder:
def __init__(self):
self._maxHeap = []
self._minHeap = []
def addNum(self, num: int) -> None:
if len(self._minHeap) == 0 and len(self._maxHeap) == 0:
heapq.heappush(self._minHeap,num)
return
median = self.findMedian()
if median < num:
heapq.heappush(self._minHeap,num)
else:
heapq.heappush(self._maxHeap,-1 * num)
if len(self._maxHeap) + 1 < len(self._minHeap):
pop_num = heapq.heappop(self._minHeap)
heapq.heappush(self._maxHeap,-1 * pop_num) #python does not support max heap
elif len(self._minHeap) + 1 < len(self._maxHeap):
pop_num = -1 * heapq.heappop(self._maxHeap)
heapq.heappush(self._minHeap,pop_num)
def findMedian(self) -> float:
if len(self._minHeap)<len(self._maxHeap):
small_med = -1 * self._maxHeap[0]
return small_med
elif len(self._maxHeap)<len(self._minHeap):
large_med = self._minHeap[0]
return large_med
else:
small_med = -1 * self._maxHeap[0]
large_med = self._minHeap[0]
med = (small_med + large_med)/2
return med
def clear(self) -> None:
self._maxHeap.clear()
self._minHeap.clear()
median_finder = MedianFinder()
# + colab={"base_uri": "https://localhost:8080/"} id="eMKGqzB0FKNG" outputId="fec99bcf-afaa-4561-d5f8-05987eba635c"
median_finder.clear()
median_finder.addNum(1)
print("[1]", "median: ", median_finder.findMedian())
median_finder.addNum(3)
print("[1,3]", "median: ", median_finder.findMedian())
median_finder.addNum(5)
print("[1,3,5]", "median: ", median_finder.findMedian())
median_finder.addNum(5)
print("[1,3,5,5]", "median: ", median_finder.findMedian())
median_finder.addNum(4)
print("[1,3,5,5,4]", "median: ", median_finder.findMedian())
median_finder.addNum(8)
print("[1,3,5,5,4,8]", "median: ", median_finder.findMedian())
median_finder.addNum(5)
print("[1,3,5,5,4,8,5]", "median: ", median_finder.findMedian())
median_finder.addNum(1)
print("[1,3,5,5,4,8,5,1]", "median: ", median_finder.findMedian())
median_finder.addNum(2)
print("[1,3,5,5,4,8,5,1,2]", "median: ", median_finder.findMedian())
# + id="dhqdWXhIFZ62"
|
heap/findMedianStream.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-1:081325390199:image/datascience-1.0
# ---
# # GPU-Accelerated Tree SHAP on AWS
#
# With the release of XGBoost 1.3 comes an exciting new feature for model interpretability — GPU accelerated SHAP values. SHAP values are a technique for local explainability of model predictions. That is, they give you the ability to examine the impact of various features on model output in a principled way. SHAP at its core describes the average impact from adding a feature to a model, but does it in a way that attempts to account for all possible subsets of the other features as well. See [GPU-Accelerated SHAP values with XGBoost 1.3 and RAPIDS](https://medium.com/rapids-ai/gpu-accelerated-shap-values-with-xgboost-1-3-and-rapids-587fad6822) for more details.
#
# In this notebook, we provide an example of training an XGBoost model with AWS SageMaker's [XGBoost estimator](https://docs.aws.amazon.com/sagemaker/latest/dg/xgboost.html), and then use SHAP values to identify key features and feature interactions in our dataset. SHAP values have been available in XGBoost for several versions already, but 1.3 brings GPU acceleration, reducing computation time by up to 20x for SHAP values and 340x for SHAP interaction values. This is powered under the hood by RAPIDS GPUTreeShap, which offers portable CUDA C++ implementations of SHAP algorithms for decision tree models.
#
# We will be using the NYC Taxi dataset, which captures yellow cab trip details in New York in January 2020, stored in CSV format without any compression. The machine learning objective with this dataset is to predict whether a trip had an above average tip (>$2.20).
#
# We also provide functionality with the California housing dataset. This is a famous dataset of house prices and attributes in California from the 1990 Census, available via [scikit-learn](https://scikit-learn.org/stable/modules/generated/sklearn.datasets.fetch_california_housing.html). In order to use this dataset, the user must adjust (comment/uncomment) sections of the code accordingly.
#
# ## 0. Preliminary Setup
#
# This notebook was tested in an Amazon [SageMaker Studio](https://docs.aws.amazon.com/sagemaker/latest/dg/studio.html) notebook, on a ml.t3.medium instance with Python 3 (Data Science) kernel. As a preliminary step, we first ensure that the latest version of SageMaker is installed:
# !pip install -U sagemaker
# Let's start by specifying:
# 1. The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the Notebook Instance, training, and hosting.
# 2. The IAM role arn used to give training and hosting access to your data. See the [AWS documentation](https://docs.aws.amazon.com/glue/latest/dg/create-an-iam-role-sagemaker-notebook.html) regarding these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the boto regex with a the appropriate full IAM role arn string(s).
# +
import io
import os
import boto3
import sagemaker
import time
role = sagemaker.get_execution_role()
region = boto3.Session().region_name
# S3 bucket for saving code and model artifacts.
# Feel free to specify a different bucket here if you wish.
bucket = sagemaker.Session().default_bucket()
prefix = "sagemaker/DEMO-xgboost-inference-script-mode"
# -
# ## 1. Training the XGBoost model
#
# SageMaker can now run an XGboost script using the [XGBoost estimator](https://docs.aws.amazon.com/sagemaker/latest/dg/xgboost.html). A typical training script loads data from the input channels, configures training with hyperparameters, trains a model, and saves a model to `model_dir` so that it can be hosted later. In this notebook, we use the training script [train.py](train.py).
#
# After setting training parameters, we kick off training.
#
# To run our training script on SageMaker, we construct a `sagemaker.xgboost.estimator.XGBoost` estimator, which accepts several constructor arguments:
#
# * __entry_point__: The path to the Python script SageMaker runs for training and prediction.
# * __hyperparameters__ *(optional)*: A dictionary passed to the train function as hyperparameters. For the XGBoost estimator, the list of possible hyperparameters can be found [here](https://docs.aws.amazon.com/sagemaker/latest/dg/xgboost_hyperparameters.html). By default, within our training script we have set "tree_method" to "gpu_hist" and "predictor" to "gpu_predictor" to enable GPU-accelerated training and SHAP values. For the CPU version, set "tree_method" to "hist" and "predictor" to "cpu_predictor."
# * __role__: Role ARN
# * __instance_type__: The type of SageMaker instance to be used. A list of instance types can be found [here](https://aws.amazon.com/ec2/instance-types/). The `ml.g4dn.xlarge` is a GPU instance; other GPUs can be specified, or a CPU instance type for the CPU run.
# * __framework_version__: SageMaker XGBoost version you want to use for executing your model training code, e.g., `0.90-1`, `0.90-2`, `1.0-1`, or `1.3-1`. We must use `1.3-1` for GPU accelerated SHAP values.
# * __sagemaker_session__ *(optional)*: The session used to train on Sagemaker.
# +
from sagemaker.inputs import TrainingInput
from sagemaker.xgboost.estimator import XGBoost
job_name = "DEMO-xgboost-inference-script-mode-" + time.strftime("%Y-%m-%d-%H-%M-%S", time.gmtime())
print("Training job", job_name)
hyperparameters = {
"max_depth": "6",
"eta": "0.3",
"gamma": "0",
"min_child_weight": "1",
"subsample": "1",
"objective": "reg:squarederror",
"num_round": "500",
"verbosity": "1",
# "tree_method": "hist", "predictor": "cpu_predictor", # for CPU version
# dataset-specific params
# "sklearn_dataset": "sklearn.datasets.fetch_california_housing()", # uncomment to use California housing dataset
"content_type": "csv", # comment out when using California housing dataset
"label_column": "17", # comment out when using California housing dataset
}
instance_type = "ml.g4dn.xlarge" # "ml.c5.xlarge" for CPU, "ml.g4dn.xlarge" for GPU
xgb_script_mode_estimator = XGBoost(
entry_point="train.py",
hyperparameters=hyperparameters,
role=role,
instance_count=1,
instance_type=instance_type,
framework_version="1.3-1",
output_path="s3://{}/{}/{}/output".format(bucket, prefix, job_name),
)
# +
"""
Since the estimator requires a valid file type but we are specifying a sklearn_dataset,
we pass in a path to a tiny csv file which will not be used.
"""
content_type = "text/csv" # MIME type
train_input = TrainingInput(
"s3://sagemaker-rapids-hpo-us-east-1/dummy_data.csv", content_type=content_type
)
# Example of using a public CSV dataset - remember to remove "sklearn_dataset" hyperparameter
# Comment out when using California housing dataset
train_input = TrainingInput(
"s3://sagemaker-rapids-hpo-us-east-1/NYC_taxi/NYC_taxi_tripdata_2020-01.csv", content_type="text/csv"
)
# -
# ### 1.1 Train XGBoost Estimator on California Housing Data
#
# Training is as simple as calling `fit` on the Estimator. This will start a SageMaker Training job that will download the data, invoke the entry point code (in the provided script file), and save any model artifacts that the script creates.
#
# Note in the cell above that we can specify "sklearn_dataset": "sklearn.datasets.fetch_california_housing()" as a parameter in order to use the California housing dataset from scikit-learn. However, calling `fit` on the Estimator requires a valid filepath, but we do not want to have to download the scikit-learn data. Instead, we refer to a small (only a couple of bytes large) existing dummy CSV file, which is immediately discarded by `train.py` upon receiving a valid "sklearn_dataset" input.
# %%time
xgb_script_mode_estimator.fit({"train": train_input}, job_name=job_name)
# ## 2. Deploying the XGBoost endpoint
#
# After training, we can host the newly created model in SageMaker, and create an Amazon SageMaker endpoint – a hosted and managed prediction service that we can use to perform inference. If you call `deploy` after you call `fit` on an XGBoost estimator, it will create a SageMaker endpoint using the training script (i.e., `entry_point`).
#
# You can optionally specify other functions to customize the behavior of deserialization of the input request (`input_fn()`), serialization of the predictions (`output_fn()`), and how predictions are made (`predict_fn()`). First `input_fn()` is called, then its output is fed into `predict_fn()`, and finally `output_fn()` returns the predictions. Rather than defining three separate functions, you can also combine them all into a function `transform_fn()`, which is what we did for this example in **inference.py**. If any of these functions are not specified, the endpoint will use the default functions in the SageMaker XGBoost container. See the [SageMaker Python SDK documentation](https://sagemaker.readthedocs.io/en/stable/frameworks/xgboost/using_xgboost.html#sagemaker-xgboost-model-server) for details.
#
# In this notebook, we will run a separate inference script and customize the endpoint to return [SHAP](https://github.com/slundberg/shap) values and interactions in addition to predictions. The inference script that we will run in this notebook is provided as the accompanying file `inference.py`.
#
# ### 2.1 Deploy to an endpoint
#
# Since the inference script is separate from the training script, here we use `XGBoostModel` to create a model from s3 artifacts and specify `inference.py` as the `entry_point`.
# +
from sagemaker.xgboost.model import XGBoostModel
model_data = xgb_script_mode_estimator.model_data
print(model_data)
xgb_inference_model = XGBoostModel(
model_data=model_data,
role=role,
entry_point="inference.py",
framework_version="1.3-1",
)
# -
predictor = xgb_inference_model.deploy(
initial_instance_count=1,
instance_type=instance_type,
serializer=None, deserializer=None,
)
# ### 2.2 Make predictions
#
# Now that we have fetched the dataset and trained an XGBoost regression model with 500 trees (using GPU acceleration), we can generate predictions on the training set.
print(predictor.serializer)
predictor.serializer = sagemaker.serializers.CSVSerializer() # for NYC_taxi predictions. Comment out for sklearn predictions
# In order to calculate predictions for the NYC Taxi dataset, we must read in the CSV file itself. We make predictions on 20,000 rows of the data. The following two cells may be skipped if you are generating predictions for the California housing dataset instead.
# +
import pandas as pd
data = pd.read_csv('s3://sagemaker-rapids-hpo-us-east-1/NYC_taxi/NYC_taxi_tripdata_2020-01.csv')
X = data.iloc[:,:-1]
# +
cutoff = 0
input_data = []
for _, row in X.iterrows():
cutoff += 1
if cutoff > 20000:
break
to_predict = []
for i in range(row.shape[0]):
to_predict.append(row[i])
input_data.append(to_predict)
# -
# Here we make our predictions:
# input_data = "sklearn.datasets.fetch_california_housing()" # uncomment to make predictions on California housing dataset
predictor_input = str(input_data) + ", predict"
predictions = predictor.predict(predictor_input)
# Because the output is a string, we define a method to clean it up and re-cast it as a NumPy array.
# +
import numpy as np
def clean_array(arr, three_dim=False):
cleaned_list = []
arr_count = 0
for num in arr:
if '[' in num:
arr_count += 1
num = num.replace('[', '')
cleaned_list.append(float(num))
elif ']' in num:
num = num.replace(']', '')
cleaned_list.append(float(num))
else:
cleaned_list.append(float(num))
array = np.array(cleaned_list, dtype='float32')
if three_dim: # shap_interactions will be 3D
y = int( len(array) / arr_count )
x = int( arr_count / y )
array = array.reshape(x, y, y)
elif(arr_count > 1):
y = int( len(array) / arr_count )
array = array.reshape(arr_count, y)
return array
predictions = clean_array(predictions[0])
# -
# ### 2.3 Generate SHAP values
#
# Here we generate the SHAP values of the training set. The time it took to compute all the SHAP values was:
# +
predictor_input = str(input_data) + ", pred_contribs"
start = time.time()
shap_values = predictor.predict(predictor_input)
print("SHAP time {}".format(time.time() - start))
shap_values = clean_array(shap_values[0])
# -
# `shap_values` now contains a matrix where each row is a training instance from X and the columns contain the feature attributions (i.e. the amount that each feature contributed to the prediction). The last column in the output shap_values contains the ‘bias’ or the expected output of the model if no features were used. Each row always adds up exactly to the model prediction — this is a unique advantage of SHAP values compared to other model explanation techniques.
#
# Model predictions can be inspected individually using this output, or we can aggregate the SHAP values to gain insight into global feature importance. Here we take the mean absolute contribution of each feature and plot their magnitude.
# ### 2.4 Compute SHAP interactions
#
# Now with GPUTreeShap we can compute these interaction effects in a matter of seconds, even for large datasets with many features.
# +
predictor_input = str(input_data) + ", pred_interactions"
start = time.time()
shap_interactions = predictor.predict(predictor_input)
print("SHAP interactions time {}".format(time.time() - start))
shap_interactions = clean_array(shap_interactions[0], three_dim=True)
# -
# The NYC Taxi and California housing datasets are relatively small, with 17x17 and 8x8 possible feature interactions, respectively. For larger datasets, as shown in our paper, GPUTreeShap can reduce feature interaction computations from days to a matter of minutes.
#
# The output `shap_interactions` contains a symmetric matrix of interaction terms for each row, where the element-wise sum evaluates to the model prediction. The diagonal terms represent the main effects for each feature or the impact of that feature excluding second-order interactions.
#
# As before we can aggregate interactions to examine the most significant effects over the training set.
# ### 2.5 Delete the Endpoint
#
# If you're done with this exercise, please run the `delete_endpoint` line in the cell below. This will remove the hosted endpoint and avoid any charges from a stray instance being left on.
predictor.delete_endpoint()
# ## 3. References
#
# - [GPU-Accelerated SHAP values with XGBoost 1.3 and RAPIDS](https://medium.com/rapids-ai/gpu-accelerated-shap-values-with-xgboost-1-3-and-rapids-587fad6822)
#
# - [SageMaker XGBoost Abalone example](https://github.com/aws/amazon-sagemaker-examples/tree/master/introduction_to_amazon_algorithms/xgboost_abalone)
#
# - [SageMaker XGBoost docs](https://sagemaker.readthedocs.io/en/stable/frameworks/xgboost/index.html)
|
aws/gpu_tree_shap/gpu_tree_shap.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="GK7eZ1GohLjK" colab={"base_uri": "https://localhost:8080/", "height": 321} outputId="48402cbf-1b26-410a-8a25-e9ee94af2fe0"
import matplotlib.pyplot as plt
import numpy as np
from scipy.special import factorial
# Create the vectors X and Y
x = np.array(range(100))
y = (x ** -1.5)
fig, ax = plt.subplots()
# Create the plot
plt.plot(x,y,label=r"Power Law, $\alpha=1.5 $ ")
xs = np.arange(0, 100, 0.1)
#y = [(((30**x) * math.exp(-30))/math.factorial(x)) for x in xs]
y = np.exp(-20)*np.power(20, xs)/factorial(xs)
# Create the plot
plt.plot(xs,y,label="Poisson, $\lambda=20$")
plt.ylim(0,0.2)
plt.xlim(0,50)
plt.xlabel("$k$, node degree")
plt.ylabel("frequency")
# Show the plot
leg = ax.legend();
plt.show()
# + id="PLhS2ANDhjNN"
|
fig_sources/notebooks/distributions2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The numpy.random package
# <h2>Assignment 2018 for Programming for Data Analysis - Investigation into numpy.random</h2>
#
# There are four distinct tasks to be carried out in this investigation.
# <ol><li>Explain the overall purpose of the numpy.random package.</li>
# <li>Explain the use of the “Simple random data” and “Permutations” functions.</li>
# <li>Explain the use and purpose of at least five “Distributions” functions.</li>
# <li>Explain the use of seeds in generating pseudorandom numbers.</li></ol>
#
# <hr>
#
# <h4>1. Explanation of the overall purpose of the numpy.random package</h4>
# Numpy.random is a sub package of the NumPy library for Python. NumPy, which is short for Numerical Python, is a library which contains large, multidimensional arrays and matrices as well as a collection of high-level mathematical routines to process these arrays. NumPy enriches Python and ensures efficient calculation. To attempt a NumPy function through core Python would require extensive amounts of custom code.
#
# Random numbers - and the ability to generate random numbers - are an important part of data analysis. The numpy.random sub package can generate random, or to be more accurate, pseudorandom numbers - pseudorandom because numpy.random arrays are determined by an initial value known as a seed. As a result, pseudorandom number generators can never be considered truely random, but that is more a theoretical point. In terms of statistics and probability, pseudorandom number generators are more than sufficient.
#
# The purpose of the numpy.random package is to provide us with a way to generate and analyze pseudorandom number arrays and matrices.
#
# <h4>2. Explanation of the use of "Simple random data" and "Permutations" functions.</h4>
#
# <h5>Simple Random Data</h5>
#
# Simple random data is a collection of sub-routines within the numpy.random routine. At it's most basic, the simple random data routines are variations on the essential premise of numpy.random which is concerned with the creation of values from probabiliity distributions.
#
# Take the numpy.random.rand function as an example. numpy.random.rand is a convenience function that creates an array of specified shape and fills it with random values.
# Import Libraries
import numpy as np #import numpy
import matplotlib.pyplot as plt
np.random.rand(4) #create a basic 1D array of 4 rows and 1 column
np.random.rand(4) #re-running same line of code gives a random result
np.random.rand(4,5) #to run 2d array of random values
# The second sub routine in the simple random data collection we can look at is numpy.random.sample - which returns random floats in the half open interval [0.0, 1.0).
np.random.random_sample(4)
# Many of the sub-routines in numpy.random perform similar tasks, but random.random_sample takes it one step further and is just an alias for the functions random.random, random.ranf, and random.rand.
np.random.random_sample is np.random.ranf is np.random.random
# The only difference between random.random_sample and random.rand is that the shapes of the output array in random.rand are separate arguments compared to a signle tupel in random.random_sample.
# <h5>Permutations Functions</h5>
#
# A permutation is an arrangement or ordering of a number of different objects. Numpy.random contains two permutation sub-functions namely "shuffle" and "permutation".
#
# Shuffle, as the name suggests, modifies a sequence by shuffling or re-arranging its contents. Permutation is similar and can be used to randonly permute a sequence or range. The difference between the two routines is subtle. The shuffle function requires a list or array to be input as a paramater. The permutation function, on the other hand, can be used with an integer or an array. The permutation function will randomly permute np.arange(x) if x=integer.
m = np.array([1,2,3,4,5,6,7,8,9,10]) #define array
np.random.shuffle(m)
m
np.random.permutation(10) #function randomly permutes np.arrange(10) in this instance
# %matplotlib inline
m = np.array([1,2,3,4,5,6,7,8,9,10]) #define array
np.random.shuffle(m)
m
plt.hist(m) #show array on histogram
# <h4>Explain the use and purpose of at least five “Distributions” functions.</h4>
#
# Distributions are a visual representation of all possible values that your random variable can be plus the probabilities for each of those values. Distributions can take the form of tables, graphs or charts.
#
# The numpy.random function contains a number of distribution sub-functions to generate and visualize random variables from specific probability distribution.
#
#
# The five distributions we will examine in this section are:
# <ul><li><strong>beta(a, b[, size])</strong> Draw samples from a Beta distribution.</li>
# <li><strong>binomial(n, p[, size])</strong> Draw samples from a binomial distribution.</li>
# <li><strong>chisquare(df[, size])</strong> Draw samples from a chi-square distribution.</li>
# <li><strong>dirichlet(alpha[, size])</strong> Draw samples from the Dirichlet distribution.</li>
# <li><strong>exponential([scale, size])</strong> Draw samples from an exponential distribution.</li></ul>
#
# <strong>Beta</strong>
#
np.random.beta(4, 5, size=5)
x = np.random.beta(4, 5, size=5)#simple example
plt.hist(x)
# <strong>Binomial</strong>
#
# This function lets us draw samples from a binomial distribution. Samples are drawn from a Binomial distribution with specified parameters. These paramaters are n=trials and p=probability of success where n an integer > 0 and p is in the interval [0,1].
#
x = np.random.binomial(17, .5, size=1000)#17 samples, 50% probability of success, and 1000 experiments
plt.hist(x)
# In the example above the histogram shows that the majority of values are returned are between 8-10 which is consistent with our 50% probability of success when calculated out of n (n=17)
# <strong>Chisquare</strong>
#
# In probability, a chi-squared distribution is widely used in hypotheses testing to determine if there is a difference between an expected distribution and actual distribution.
#
# This distribution uses the parameters <i>df</i> for degrees of freedom and <i>size</i> for the sample size.
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
x = np.random.chisquare(2,4)
plt.hist(x)
# <strong>Dirichlet</strong><br>
# The Dirichlet distribution shows the probability distribution over a sample of numbers that add up to 1 also known as a probability simplex.
# <strong>Exponential</strong>
#
# An exponential distribution is closely related to the Poisson distribution - which allows us to answer question like the number of events that occur in an interval of time. Exponential, on the other hand, deals with the time taken between 2 events occuring.
np.random.exponential(scale=1.0, size=None)
# <h4>4. Explain the use of seeds in generating pseudorandom numbers</h4>
#
# Pseudorandom numbers have all the outward trappings of random numbers but they can never be truely random because they are set by a seed. A seed is the starting point from which software programs can generate pseudorandom numbers. Because of the presence of the seed, the generated numbers can not be considered truely random. A human who rolls a dice and records each result can create a truely random sequence because there is no set starting point.
#
# By seeding the numpy.random generator, you can make the generated numbers predictable.
np.random.seed(1) ; np.random.rand(6)
np.random.seed(1) ; np.random.rand(6)
#
# <strong>References:</strong>
#
# https://www.packtpub.com/mapt/book/application_development/9781783985128/3/ch03lvl1sec24/benefits-and-characteristics-of-numpy-arrays - numpy arrays overview
# <br>https://www.python-course.eu/numpy.php - numpy overview
# <br>https://stackabuse.com/numpy-tutorial-a-simple-example-based-guide/#advantagesofnumpy - numpy tutotial
# <br>https://www.packtpub.com/mapt/book/big_data_and_business_intelligence/9781785285110/2/ch02lvl1sec16/numpy-random-numbers - numpy and random number generation
# <br>https://www.numpy.org/devdocs/reference/generated/numpy.random.randn.html - numpy.random.rand info
# <br>https://en.wikipedia.org/wiki/Random_number_generation - info on random number generation
# <br>https://whatis.techtarget.com/definition/pseudo-random-number-generator-PRNG - Intro to pseudorandom number generation
# <br>https://www.python-course.eu/python_numpy_probability.php - Intro to how python is used for probability
# <br>https://www.geeksforgeeks.org/numpy-random-rand-python/ - explanation of numpy.random.rand sub function
# <br>https://study.com/academy/lesson/permutation-definition-formula-examples.html - permutation definition
# <br>https://www.youtube.com/watch?v=r7lVVeU834o - distributions definition
# <br>http://www.mathnstuff.com/math/spoken/here/2class/90/binom2.htm - binomial defintion
# <br>https://towardsdatascience.com/running-chi-square-tests-in-python-with-die-roll-data-b9903817c51b - good blog post on chi square distribution
# <br>https://www.quora.com/What-is-an-intuitive-explanation-of-the-Dirichlet-distribution - dirichlet distribution explainer
# <br>https://www.youtube.com/watch?v=bKkLYSi5XNE - exponential distribution definition
# https://stackoverflow.com/questions/21494489/what-does-numpy-random-seed0-do - numpy random seed explainer
#
#
#
|
Numpy-random.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="ItXfxkxvosLH"
# # TensorFlow and TextAttack
# -
# [](https://colab.research.google.com/drive/1cBRUj2l0m8o81vJGGFgO-o_zDLj24M5Y?usp=sharing)
#
# [](https://github.com/QData/TextAttack/blob/master/docs/examples/1_Introduction_and_Transformations.ipynb)
# + [markdown] colab_type="text" id="WooZ9pGnNJbv"
# ## Training
#
#
#
# The following is code for training a text classification model using TensorFlow (and on top of it, the Keras API). This comes from the Tensorflow documentation ([see here](https://www.tensorflow.org/tutorials/keras/text_classification_with_hub)).
#
# This cell loads the IMDB dataset (using `tensorflow_datasets`, not `nlp`), initializes a simple classifier, and trains it using Keras.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="2ew7HTbPpCJH" outputId="1c1711e1-cf82-4b09-899f-db7c9bb68513"
import numpy as np
import tensorflow as tf
import tensorflow_hub as hub
import tensorflow_datasets as tfds
import matplotlib.pyplot as plt
print("Version: ", tf.__version__)
print("Eager mode: ", tf.executing_eagerly())
print("Hub version: ", hub.__version__)
print("GPU is", "available" if tf.config.list_physical_devices('GPU') else "NOT AVAILABLE")
train_data, test_data = tfds.load(name="imdb_reviews", split=["train", "test"],
batch_size=-1, as_supervised=True)
train_examples, train_labels = tfds.as_numpy(train_data)
test_examples, test_labels = tfds.as_numpy(test_data)
model = "https://tfhub.dev/google/tf2-preview/gnews-swivel-20dim/1"
hub_layer = hub.KerasLayer(model, output_shape=[20], input_shape=[],
dtype=tf.string, trainable=True)
hub_layer(train_examples[:3])
model = tf.keras.Sequential()
model.add(hub_layer)
model.add(tf.keras.layers.Dense(16, activation='relu'))
model.add(tf.keras.layers.Dense(1))
model.summary()
x_val = train_examples[:10000]
partial_x_train = train_examples[10000:]
y_val = train_labels[:10000]
partial_y_train = train_labels[10000:]
model.compile(optimizer='adam',
loss=tf.losses.BinaryCrossentropy(from_logits=True),
metrics=['accuracy'])
history = model.fit(partial_x_train,
partial_y_train,
epochs=40,
batch_size=512,
validation_data=(x_val, y_val),
verbose=1)
# + [markdown] colab_type="text" id="3varlQvrnHqV"
# ## Attacking
#
# For each input, our classifier outputs a single number that indicates how positive or negative the model finds the input. For binary classification, TextAttack expects two numbers for each input (a score for each class, positive and negative). We have to post-process each output to fit this TextAttack format. To add this post-processing we need to implement a custom model wrapper class (instead of using the built-in `textattack.models.wrappers.TensorFlowModelWrapper`).
#
# Each `ModelWrapper` must implement a single method, `__call__`, which takes a list of strings and returns a `List`, `np.ndarray`, or `torch.Tensor` of predictions.
# + colab={} colab_type="code" id="fHX3Lo7wU2LM"
import numpy as np
import torch
from textattack.models.wrappers import ModelWrapper
class CustomTensorFlowModelWrapper(ModelWrapper):
def __init__(self, model):
self.model = model
def __call__(self, text_input_list):
text_array = np.array(text_input_list)
preds = self.model(text_array).numpy()
logits = torch.exp(-torch.tensor(preds))
logits = 1 / (1 + logits)
logits = logits.squeeze(dim=-1)
# Since this model only has a single output (between 0 or 1),
# we have to add the second dimension.
final_preds = torch.stack((1-logits, logits), dim=1)
return final_preds
# + [markdown] colab_type="text" id="Ku71HuZ4n7ih"
# Let's test our model wrapper out to make sure it can use our model to return predictions in the correct format.
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="9hgiLQC4ejmM" outputId="132c3be5-fe5e-4be4-ef98-5c2efedc0dfd"
CustomTensorFlowModelWrapper(model)(['I hate you so much', 'I love you'])
# + [markdown] colab_type="text" id="-Bs14Hr4n_Sp"
# Looks good! Now we can initialize our model wrapper with the model we trained and pass it to an instance of `textattack.attack.Attack`.
#
# We'll use the `PWWSRen2019` recipe as our attack, and attack 10 samples.
# + colab={"base_uri": "https://localhost:8080/", "height": 780} colab_type="code" id="07mOE-wLVQDR" outputId="e47a099e-c0f6-4c21-8e52-1a437741bc16"
model_wrapper = CustomTensorFlowModelWrapper(model)
from textattack.datasets import HuggingFaceNlpDataset
from textattack.attack_recipes import PWWSRen2019
dataset = HuggingFaceNlpDataset("rotten_tomatoes", None, "test", shuffle=True)
attack = PWWSRen2019.build(model_wrapper)
results_iterable = attack.attack_dataset(dataset, indices=range(10))
for result in results_iterable:
print(result.__str__(color_method='ansi'))
# + [markdown] colab_type="text" id="P3L9ccqGoS-J"
# ## Conclusion
#
# Looks good! We successfully loaded a model, adapted it for TextAttack's `ModelWrapper`, and used that object in an attack. This is basically how you would adapt any model, using TensorFlow or any other library, for use with TextAttack.
|
docs/datasets_models/Example_0_tensorflow.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Objects
# Just about everything we write or interact with in Python is an **object**. We already know how to make **String** objects, and today we will learn to make **List** and **Dictionary** objects. Str, List and Dict are all examples of **classes** of objects, and each class has specific properties and a type of data that it can store.
#
# The **String** class is defined to store a sequence of charaters in a specific order. We can check the class of an object using the `type()` function.
type('Hi, my name is...')
introduction = 'Hi, my name is...'
type(introduction)
# Recall that we can index str type objects
introduction[0:3]
# ## Object-oriented
# Object-oriented programming (OOP) is a design philosophy of certain programming languages, Python included. Objects are convenient ways to think about and organize data while we are programming. We will take a short detour to understand the power of object-oriented programming by making some example **classes**.
# ## Designing solar systems
# Say we want to represent (simple) solar systems in our code in an object-oriented way. Which solar system objects can we think about using?
#
# ### Hard-coded approach
# If we start from the perspective of our solar system, we may start with a `Sun` object, and then add a `Mercury`, `Venus`, `Earth`, and `Mars` object. But this doesn't generalize well any other solar system. Instead, we can think of what **classes** of objects all solar systems have in common.
#
# ### Hard-coded approach (with classes)
# All solar systems should have a star, so let's start with the `Star` class. Then we might have a `Planet` class for all of our planet objects. But there might also be things called `DwarfPlanets` so we should have a class for that too. But our inner planets are quite different from the outer planets so maybe we should actually split `Planet` into the `TerrestrialPlanet` and `JovianPlanet` classes. But now how do we make it clear that `TerrestrialPlanet` and `JovianPlanet` are planets, but `DwarfPlanet` is not? *Ahhhhhhh*.
#
# Let's try once more. The object-oriented way.
#
# ### Object-oriented approach (with inheritance)
# One of the critical features of an object oriented programming language is the ability to make a hierarchy of objects which are related to each other in some way. Say we start with the one thing all large objects in a solar system have in common: *being round*.
# ```Python
# class CelestialOrb:
# """A solar system object large enough to be rounded by its own gravity."""
# def isround(self):
# return True
# ```
#
# We don't actually call any specific object in the sky a "celestial orb", but this class can help us organize our other classes which all have the *isround* property. Now we can make a **subclass** of CelestialOrb, that automatically **inherits** the *isround* property, but then adds its own distinguishing feature. Let's again start with the `Star` class.
# ```Python
# # Star inherits from CelestialOrb, meaning it automatically gets the isround() method.
# class Star(CelestialOrb):
# """The central star of a solar system."""
# # We don't need to define isround because it was inherited
# def has_fusion(self):
# return True
# ```
#
# Now let's add a `Planet` class. This class does not have fusion, so it would not make sense to subclass `Star`. Since it is round, we can once again subclass `CelestialOrb`. There is *no limit to how many subclasses a class can have*.
#
# ```Python
# # Planet inherits from CelestialOrb
# class Planet(CelestialOrb):
# """A CelestialOrb which orbits the Star and clears its orbit."""
# def in_orbit_around(self):
# return self.star
#
# def clears_orbit(self):
# return True
# ```
#
# Now let's add `DwarfPlanets`. Since dwarf planets are not technically planets because they do not clear their orbit, we once again subclass `CelestialOrb`.
#
# ```Python
# # DwarfPlanet inherits from CelestialOrb
# class DwarfPlanet(CelestialOrb):
# """A CelestialOrb which orbits the Star and does not clear its orbit."""
# def in_orbit_around(self):
# return self.star
#
# def clears_orbit(self):
# return False
# ```
#
# If we wanted to distinguish between Jovian and terrestrial planets, we can subclass again! We would want to keep all of the features of Planet (`isround, in_orbit_around, clears_orbit`), so this time we subclass `Planet`.
#
# ```Python
# # TerrestrialPlanet inherits from Planet
# class TerrestrialPlanet(Planet):
# """A Planet which has a solid surface."""
# def has_solid_surface(self):
# return True
#
# # JovianPlanet inherits from Planet
# class JovianPlanet(Planet):
# """A Planet which does not have a solid surface."""
# def has_solid_surface(self):
# return False
# ```
#
# Now we can make our solar system out of our classes, but these classes are *flexible* enough to be used on other solar systems as well.
#
# ```Python
# Sun = Star()
# Mercury = TerrestrialPlanet(Sun)
# Venus = TerrestrialPlanet(Sun)
# Earth = TerrestrialPlanet(Sun)
# Mars = TerrestrialPlanet(Sun)
# Ceres = DwarfPlanet(Sun)
# Jupiter = JovianPlanet(Sun)
# Saturn = JovianPlanet(Sun)
# Uranus = JovianPlanet(Sun)
# Neptune = JovianPlanet(Sun)
# Pluto = DwarfPlanet(Sun)
#
# print(Sun.isround(), Mercury.isround(), Ceres.isround(), Jupiter.isround())
# # True True True True
#
# print(Sun.has_fusion())
# # True
#
# print(Mercury.in_orbit_around(), Ceres.in_orbit_around(), Jupiter.in_orbit_around())
# # sun sun sun
#
# print(Mercury.clears_orbit(), Ceres.clears_orbit(), Jupiter.clears_orbit())
# # True False True
#
# print(Mercury.has_solid_surface(), Jupiter.has_solid_surface())
# # True False
# ```
#
# The powerful thing about object-oriented design is that it is *extensible*. After you have written your object-oriented solar system code and we decide to add moon classes, we can always return to our code and add a `Moon` subclass without disrupting anything else about our model.
#
# Try to think about what a `Moon` subclass would look like in this picture. What would it inherit from, what would be distinguishing features about it? Without worrying much about the syntax, try to fill in the blanks below.
class Moon(____):
"""____ """
def in_orbit_around(self):
return self.____
# # Methods
# Classes have special functions defined only for their own members called class methods, or simply **methods**. Methods are called on an object by following that object with `.methodname()`. We saw methods like `.isround()` in action above. Built-in classes like `str` have methods too!
# The upper() method changes all characters in a string to uppercase
introduction = 'Hi, my name is...'
introduction.upper()
# The isdigit() method checks if all characters in a string are digits
'12345'.isdigit()
# Using the `help()` function on a class shows all methods available to instances of that class. The `__methods__` are private methods used internally by Python. Skip down to `capitalize(...)` to see the methods available to us.
help(str)
# The capitalize method capitalizes the fist letter of the str
'united states'.capitalize()
# # Functions
# Functions are like methods but are independent of a specific class. Any objects that they act on must be passed in as arguments. Let's break down the anatomy of a function.
#
# ```Python
# def funcname(arg1, arg2):
# """Docstring"""
# # Do stuff here
# return output
# ```
#
# All functions start with a `def` or **define** statement, followed by the name of the function and a list of arguments in parentheses.
#
# Below the `def` statement is the **docstring** in triple quotes `""" """`. Docstrings are important for humans (including you) who need to read / use your code. The docstring explains what the function does, what arguments the function needs to work properly, and can even suggest example usage. The docsting is what is shown when you call `help(funcname)` on your function.
#
# As we saw with the if blocks, Python uses indentation to organize code. All code indented in the function definition will be run when the function is called.
#
# Finally, if your function produces an output, it must be **returned** with a `return` statement. This signals the end of the function. Python will pick up where it left off before running the function.
#
# Let's work through an example. Say we encounter the following function `stuff()`. We may not know what it does initially if we don't know where it is defined. Let's try calling `help()`.
def stuff(a):
return a**2
help(stuff)
# Hmm. Not very descriptive. And the name of the function is not exactly helpful. I guess we need to try some examples to figure it out.
stuff('hello?')
# Well it didn't like the `str`, so let's try an `int` instead.
stuff(1)
# Now we're getting somewhere, maybe `stuff` returns the number it is given!
stuff(2)
# There goes that idea. But this looks like it could be pattern.
print(stuff(1),stuff(2),stuff(3),stuff(4),stuff(5))
# Cool it looks like `stuff` takes the square of the number it is given! Now to do the same trial and error with the function `allxsonthelistorunderbutnotboth(x1, x2, x3, x4, x5, x6, list1, list2)`. Uhh...
#
# That was an example of writing a function with poor *style*. The function worked as intended, but was frustrating to use if you didn't remember what `stuff()` did. I hope this highlights the importance of readable code. Python comes built-in with features like the **docstring** to avoid situations like the one above. Python won't force you to use docstrings, but it is highly encouraged to get into the habit, especially if you are working with others. And if not for others, do it for future you who won't remember what `stuff()` is in 6 months.
#
# So how do we improve our `stuff()` function for squaring numbers? The first step is giving it a self-evident name, e.g. `square(num)`. Next, we can add a docstring with a description `"""Return the square of num"""`. Finally, we can describe the parameters, return values and provide a couple examples of how to use it. Altogether, it might look like this.
def square(num):
"""Return the square of num.
Parameters
----------
num: int, float
The number to square.
Returns
-------
int, float
The square of num.
Examples
--------
>>> square(2)
4
>>> square(2.5)
6.25
"""
return num**2
# Say we encounter `square()` in the wild and want to know how to use it. Now we can call `help(square)` and see a nicely formatted docstring.
help(square)
# We can even try the examples it provides to ensure the function is working properly.
print(square(2), square(2.5))
# Much less frustrating!
#
# *Style* is an aspect of writing code that is often overlooked in sciences. Just like writing good `git commit` messages, it is very important to write code in a way that future you and future collaborators will be able to read and use.
#
# Another aspect of style is knowing when to break your code down into functions that perform small tasks. This is one of the hardest, but most useful programming skills to master. If you can define function(s) for complex / repetitive code and give those functions good names and good docstrings, you are on your way to writing readable, re-usable code!
#
# Your turn!
# ## Breaking code down into functions
# The following example is long and repetitive. See if you can define functions to shorten and simplify the code, and get the same result.
#
# In this example, we want to see if 3 people like apples, oranges, and are above the age of 20. The data is formatted as such:
#
# person = 'likesapples likesoranges age'
# ```Python
# person1 = 'yes yes 13'
# person2 = 'yes nah 21'
# person3 = 'nah nah 80'
# ```
#
# We want to `print('It's a match!')` if all 3 people like apples and oranges and are older than 20. Otherwise, `print('It's not a match!')`.
# +
person1 = 'yes yes 42'
person2 = 'yes yes 64'
person3 = 'yes yes 80'
# Uncomment these three for an example of not a match
# person1 = 'yes yes 13'
# person2 = 'yes nah 21'
# person3 = 'nah nah 80'
if person1[0:3] == 'yes' and person1[4:7] == 'yes' and int(person1[8:]) > 20:
if person2[0:3] == 'yes' and person2[4:7] == 'yes' and int(person2[8:]) > 20:
if person3[0:3] == 'yes' and person3[4:7] == 'yes' and int(person3[8:]) > 20:
print("It's a match!")
else:
print("It's not a match!")
else:
print("It's not a match!")
else:
print("It's not a match!")
# +
# Put your function version of the above code here
# Don't forget the docstrings!
# -
# There are many possible ways to break up code into functions. How specific you make your functions depends on your particular use case. If you want to see my solution, you can copy and paste it from [here](https://github.com/cjtu/sci_coding/tree/master/lessons/lesson3/data/function_solution.py) and compare with yours!
#
#
# Great job! You made it to the end of the crash course on objects, methods and functions (oh my). Next, we will be working with `Lists and Tuples`.
|
lessons/lesson3/data/1_Objects_Methods_and_Functions_oh_my!.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Predict Sports Scores with Regression
# ========================================
#
# Background: Who doesn't want to know if their favorite team will win on gameday? This notebook attempts to predict the scores of various sporting events using regression models.
#
# All data is courtesy of FiveThirtyEight, one of my favorite websites.
import pandas as pd
# Get data into dataframes
# I'm going to start out with Soccer
matches = pd.read_csv("../data/raw/soccer/spi_matches.csv")
rankings = pd.read_csv("../data/raw/soccer/spi_global_rankings.csv")
intl_rankings = pd.read_csv("../data/raw/soccer/spi_global_rankings_intl.csv")
# Take a look at each Data Frame
matches.head(5)
rankings.head(5)
intl_rankings.head(5)
# From the above dataframes, we can begin to determine what may be the most useful pieces of information. It seems like the "matches" dataframe has just about everything we could want...there's no need to match team names and rankings from the rankings df into the matches dataframe.
#
# One potentially useful feature would be to consider a team's al-time average score. This could be calculated by looking at all matches played by a team and taking a simple average. To go evern a step further, we could break this down by season.
# Take a look at our columns
list(matches.columns.values)
# From this list of columns, it seems that we may be able to eliminate some of the features. For instance, 'league_id' takes care of the 'league' attribute, so both are not needed.
#
# Additionally, it seems from this that the adjusted score and score are not needed as features in our model, since these are what we are looking to predict.
# +
# Run a simple regression test on the barebones dataframe with no
# feature engineering other than dropping some features.
from sklearn.preprocessing import LabelEncoder
e = LabelEncoder()
e.fit(matches['team1'])
matches['team1'] = e.transform(matches['team1'])
matches['team2'] = e.transform(matches['team2'])
e.fit(matches['date'])
matches['date'] = e.transform(matches['date'])
matches = matches.drop('league', 1)
matches = matches.dropna()
matches.head(5)
# -
# set up model by choosing labels and features
decision = matches[['score1']]
features = matches[['date',
'league_id',
'team1',
'team2',
'prob1',
'prob2',
'spi1',
'spi2',
'probtie',
'proj_score1',
'proj_score2',
'importance1',
'importance2',
'xg1',
'xg2']]
# +
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
train, test, train_d, test_d = train_test_split(features,
decision,
test_size = 0.2,
random_state = 14)
cls = RandomForestClassifier()
mdl = cls.fit(train, train_d.values.ravel())
# -
# Time to Make some Predictions
results = cls.predict(test)
# +
from sklearn.metrics import accuracy_score
accuracy_score(test_d, results)
# -
# determine importance of labels for a refactoring
import json
imp = {}
for feature, importance in zip(train.columns, cls.feature_importances_):
imp[feature] = importance
df = pd.Series(imp)
df = df.sort_values(ascending=False).head(5)
df
# By examining these importances, it is possible that eliminating some features may give better results. For example, it seems that the league_id may have a much smaller impact on the predicted score than previously thought.
#
# I am going to select the top 5 features and run the classifier again and see if it becomes more accurate.
# these are the top 5 features
keys = df.keys()
keys
# # copy the selection process from earlier, but pick the best 5 features
decision = matches[['score1']]
features = matches[keys]
# +
train, test, train_d, test_d = train_test_split(features,
decision,
test_size = 0.2,
random_state = 7)
cls = RandomForestClassifier()
mdl = cls.fit(train, train_d.values.ravel())
# -
results = cls.predict(test)
accuracy_score(test_d, results)
imp = {}
for feature, importance in zip(train.columns, cls.feature_importances_):
imp[feature] = importance
df = pd.Series(imp)
df.sort_values(ascending=False)
# This predictive model seems to have essentially the same accuracy as the original model, and in some cases is even worse.
#
# It seems like the best approach to making a better model may be to do some feature engineering to create some new features that may be of interest
|
notebooks/Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="YqTKIYLooHsq"
# # PixelCNN blind spots
# + [markdown] colab={} colab_type="code" id="wM-m3Z8CiLXU"
# *Note: Here we are using float64 to get more precise values of the gradients and avoid false values.
# + colab={} colab_type="code" id="gf5wwqP3ozaN"
import random as rn
import matplotlib
import matplotlib.pyplot as plt
from matplotlib.ticker import FixedLocator
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow import nn
from tensorflow.keras import initializers
from tensorflow.keras.utils import Progbar
tf.keras.backend.set_floatx('float64')
# + colab={} colab_type="code" id="yJ_JlzWco7ci"
class MaskedConv2D(keras.layers.Layer):
"""Convolutional layers with masks.
Convolutional layers with simple implementation of masks type A and B for
autoregressive models.
Arguments:
mask_type: one of `"A"` or `"B".`
filters: Integer, the dimensionality of the output space
(i.e. the number of output filters in the convolution).
kernel_size: An integer or tuple/list of 2 integers, specifying the
height and width of the 2D convolution window.
Can be a single integer to specify the same value for
all spatial dimensions.
strides: An integer or tuple/list of 2 integers,
specifying the strides of the convolution along the height and width.
Can be a single integer to specify the same value for
all spatial dimensions.
Specifying any stride value != 1 is incompatible with specifying
any `dilation_rate` value != 1.
padding: one of `"valid"` or `"same"` (case-insensitive).
kernel_initializer: Initializer for the `kernel` weights matrix.
bias_initializer: Initializer for the bias vector.
"""
def __init__(self,
mask_type,
filters,
kernel_size,
strides=1,
padding='same',
kernel_initializer='glorot_uniform',
bias_initializer='zeros'):
super(MaskedConv2D, self).__init__()
assert mask_type in {'A', 'B'}
self.mask_type = mask_type
self.filters = filters
self.kernel_size = kernel_size
self.strides = strides
self.padding = padding.upper()
self.kernel_initializer = initializers.get(kernel_initializer)
self.bias_initializer = initializers.get(bias_initializer)
def build(self, input_shape):
self.kernel = self.add_weight('kernel',
shape=(self.kernel_size,
self.kernel_size,
int(input_shape[-1]),
self.filters),
initializer=self.kernel_initializer,
trainable=True)
self.bias = self.add_weight('bias',
shape=(self.filters,),
initializer=self.bias_initializer,
trainable=True)
center = self.kernel_size // 2
mask = np.ones(self.kernel.shape, dtype=np.float64)
mask[center, center + (self.mask_type == 'B'):, :, :] = 0.
mask[center + 1:, :, :, :] = 0.
self.mask = tf.constant(mask, dtype=tf.float64, name='mask')
def call(self, input):
masked_kernel = tf.math.multiply(self.mask, self.kernel)
x = nn.conv2d(input,
masked_kernel,
strides=[1, self.strides, self.strides, 1],
padding=self.padding)
x = nn.bias_add(x, self.bias)
return x
# -
# And now, we define the residual block.
#
# *Note: Here we removed the ReLU activations to not mess with the gradients while we are investigating them.
# +
class ResidualBlock(keras.Model):
"""Residual blocks that compose pixelCNN
Blocks of layers with 3 convolutional layers and one residual connection.
Based on Figure 5 from [1] where h indicates number of filters.
Refs:
[1] - <NAME>., <NAME>., & <NAME>. (2016). Pixel recurrent
neural networks. arXiv preprint arXiv:1601.06759.
"""
def __init__(self, h):
super(ResidualBlock, self).__init__(name='')
self.conv2a = keras.layers.Conv2D(filters=h, kernel_size=1, strides=1)
self.conv2b = MaskedConv2D(mask_type='B', filters=h, kernel_size=3, strides=1)
self.conv2c = keras.layers.Conv2D(filters=2 * h, kernel_size=1, strides=1)
def call(self, input_tensor):
# x = nn.relu(input_tensor)
# x = self.conv2a(x)
x = self.conv2a(input_tensor)
# x = nn.relu(x)
x = self.conv2b(x)
# x = nn.relu(x)
x = self.conv2c(x)
x += input_tensor
return x
# +
height = 10
width = 10
n_channel = 1
data = tf.random.normal((1, height, width, n_channel))
inputs = keras.layers.Input(shape=(height, width, n_channel))
x = MaskedConv2D(mask_type='A', filters=1, kernel_size=3, strides=1)(inputs)
model = tf.keras.Model(inputs=inputs, outputs=x)
# + colab={} colab_type="code" id="jxCLMYc-FxdJ"
def plot_receptive_field(model, data):
with tf.GradientTape() as tape:
tape.watch(data)
prediction = model(data)
loss = prediction[:,5,5,0]
gradients = tape.gradient(loss, data)
gradients = np.abs(gradients.numpy().squeeze())
gradients = (gradients > 0).astype('float64')
gradients[5, 5] = 0.5
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
plt.xticks(np.arange(0, 10, step=1))
plt.yticks(np.arange(0, 10, step=1))
ax.xaxis.set_minor_locator(FixedLocator(np.arange(0.5, 10.5, step=1)))
ax.yaxis.set_minor_locator(FixedLocator(np.arange(0.5, 10.5, step=1)))
plt.grid(which="minor")
plt.imshow(gradients, vmin=0, vmax=1)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 281} colab_type="code" id="0qpDtNuvo9NL" outputId="926434e2-44bf-40d3-a39f-6c80ebc880b6"
plot_receptive_field(model, data)
# + colab={} colab_type="code" id="I2APaCzDGeqP"
inputs = keras.layers.Input(shape=(height, width, n_channel))
x = MaskedConv2D(mask_type='A', filters=1, kernel_size=3, strides=1)(inputs)
x = ResidualBlock(h=1)(x)
model = tf.keras.Model(inputs=inputs, outputs=x)
plot_receptive_field(model, data)
# +
inputs = keras.layers.Input(shape=(height, width, n_channel))
x = MaskedConv2D(mask_type='A', filters=1, kernel_size=3, strides=1)(inputs)
x = ResidualBlock(h=1)(x)
x = ResidualBlock(h=1)(x)
model = tf.keras.Model(inputs=inputs, outputs=x)
plot_receptive_field(model, data)
# +
inputs = keras.layers.Input(shape=(height, width, n_channel))
x = MaskedConv2D(mask_type='A', filters=1, kernel_size=3, strides=1)(inputs)
x = ResidualBlock(h=1)(x)
x = ResidualBlock(h=1)(x)
x = ResidualBlock(h=1)(x)
model = tf.keras.Model(inputs=inputs, outputs=x)
plot_receptive_field(model, data)
# +
inputs = keras.layers.Input(shape=(height, width, n_channel))
x = MaskedConv2D(mask_type='A', filters=1, kernel_size=3, strides=1)(inputs)
x = ResidualBlock(h=1)(x)
x = ResidualBlock(h=1)(x)
x = ResidualBlock(h=1)(x)
x = ResidualBlock(h=1)(x)
model = tf.keras.Model(inputs=inputs, outputs=x)
plot_receptive_field(model, data)
|
WIP/3 - PixelCNNs blind spot in the receptive field/pixelcnn_receptive_field.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
import networkx as nx
import numpy as np
import pandas as pd
# import projx as px
import matplotlib.pyplot as plt
import seaborn as sns
plt.rcParams['figure.figsize'] = (12, 7)
# +
def prob_dist(itrbl):
count = {}
for i in itrbl:
count.setdefault(i, 0)
count[i] += 1
sr = pd.Series(count)
prob = sr.apply(lambda x: float(x) / len(itrbl))
return prob
def basic_graph_stats(g):
stats = {
"num_nodes": len(g),
"num_edges": len(g.edges()),
"density": nx.density(g),
"diameter": nx.diameter(g),
"avg_short_path": nx.average_shortest_path_length(g),
"avg_clust": nx.average_clustering(g)
}
return pd.Series(stats)
# -
graph = nx.read_gexf("projections/onemode.gexf")
subgraphs = list(nx.connected_component_subgraphs(graph))
print([len(sub) for sub in subgraphs])
g = subgraphs[0]
g1 = subgraphs[1]
g2 = subgraphs[2]
# for js graphviz
import json
from networkx.readwrite import json_graph
peru = json_graph.node_link_data(g1)
china = json_graph.node_link_data(g2)
with open("js_viz/graphs/peru.json", "w") as f:
json.dump(peru, f)
with open("js_viz/graphs/china.json", "w") as f:
json.dump(china, f)
nx.write_gexf(g, "projections/subgraph_onemode_gc.gexf")
nx.write_gexf(g1, "projections/subgraph_onemode_sub1.gexf")
nx.write_gexf(g2, "projections/subgraph_onemode_sub2.gexf")
print(basic_graph_stats(g))
roles = [a.get("role", "") for (n, a) in g.nodes(data=True)] # Hmm.
places = [a["top_place"] for (n, a) in g.nodes(data=True)]
genres = [a["top_genre"] for (n, a) in g.nodes(data=True)]
sr = pd.Series({"role": prob_dist(roles), "place": prob_dist(places), "genre": prob_dist(genres)})
prob_dist(roles).plot(kind="bar")
plt.savefig("img/roles.png")
prob_dist(places).plot(kind="bar")
plt.savefig("img/places.png")
prob_dist(genres).plot(kind="bar")
plt.savefig("img/genres.png")
px.draw_simple_graph(g1)
#plt.savefig("img/china_component.png")
px.draw_simple_graph(g2)
#plt.savefig("img/peru_component.png")
print(len(g), len(g.edges()), nx.density(g))
# ### Centrality
bc = nx.betweenness_centrality(g, weight="weight")
ec = nx.eigenvector_centrality(g, weight="weight", max_iter=500)
cc = nx.closeness_centrality(g)
deg = nx.degree(g)
pr = nx.pagerank(g, max_iter=500, weight="weight")
cent_10_df = pd.DataFrame({
"bc": [(k, g.node[k]["label"], bc[k]) for k in sorted(bc, key=bc.get, reverse=True)[0:10]],
"ec": [(k, g.node[k]["label"], ec[k]) for k in sorted(ec, key=ec.get, reverse=True)[0:10]],
"cc": [(k, g.node[k]["label"], cc[k]) for k in sorted(cc, key=cc.get, reverse=True)[0:10]],
"dc": [(k, g.node[k]["label"], deg[k]) for k in sorted(deg, key=deg.get, reverse=True)[0:10]],
#"pr": [(k, g.node[k]["label"], pr[k]) for k in sorted(pr, key=pr.get, reverse=True)[0:10]]
})
print(cent_10_df)
pd.Series(deg.values()).hist()
deg_prob = prob_dist(deg.values())
plt.scatter(deg_prob.index, deg_prob)
plt.savefig("img/deg_dist.png")
# ### Degree by role
authors = pd.Series(nx.degree(g, nbunch=[n for (n, a) in g.nodes(data=True) if a.get("role") == "author"]).values())
patrons = pd.Series(nx.degree(g, nbunch=[n for (n, a) in g.nodes(data=True) if a.get("role") == "patron"]).values())
printers = pd.Series(nx.degree(g, nbunch=[n for (n, a) in g.nodes(data=True) if a.get("role") == "printer/editor"]).values())
signatories = pd.Series(nx.degree(g, nbunch=[n for (n, a) in g.nodes(data=True) if a.get("role") == "signatory"]).values())
# plt.plot(authors.index, authors, color="r")
# plt.plot(patrons.index, patrons, color="b")
# plt.plot(printers.index, printers, color="y")
# plt.plot(signatories.index, signatories, color="g")
df = pd.concat([authors, patrons, printers, signatories], axis=1, keys=["author", "patron", "printer", "signatory"])
desc = df.describe()
desc.to_clipboard()
a = [auth for auth in df["author"] if not np.isnan(auth)]
p = [auth for auth in df["patron"] if not np.isnan(auth)]
pr = [auth for auth in df["printer"] if not np.isnan(auth)]
s= [auth for auth in df["signatory"] if not np.isnan(auth)]
sns.boxplot([a,p,pr,s], names=["author", "patron", "printer", "signatory"])
authors = pd.Series(nx.betweenness_centrality(g, nbunch=[n for (n, a) in g.nodes(data=True) if a.get("role") == "author"]).values())
patrons = pd.Series(nx.betweenness_centrality(g, nbunch=[n for (n, a) in g.nodes(data=True) if a.get("role") == "patron"]).values())
printers = pd.Series(nx.betweenness_centrality(g, nbunch=[n for (n, a) in g.nodes(data=True) if a.get("role") == "printer/editor"]).values())
signatories = pd.Series(nx.betweenness_centrality(g, nbunch=[n for (n, a) in g.nodes(data=True) if a.get("role") == "signatory"]).values())
df = pd.concat([authors, patrons, printers, signatories], axis=1, keys=["author", "patron", "printer", "signatory"])
desc = df.describe()
a = [auth for auth in df["author"] if not np.isnan(auth)]
p = [auth for auth in df["patron"] if not np.isnan(auth)]
pr = [auth for auth in df["printer"] if not np.isnan(auth)]
s= [auth for auth in df["signatory"] if not np.isnan(auth)]
sns.boxplot([a,p,pr,s], names=["author", "patron", "printer", "signatory"])
g.nodes(data=True)
cent_df = pd.DataFrame({
"dc": nx.degree_centrality(g).values(),
"bc": bc.values(),
"cc": cc.values(),
"ec": ec.values(),
})
cent_df.hist(bins=20)
pd.Series(bc.values()).hist()
pd.Series(cc.values()).hist()
pd.Series(ec.values()).hist()
pd.Series(pr.values()).hist()
# ### Assortativity
# #### Degree
nx.degree_assortativity_coefficient(g)
r = nx.degree_assortativity_coefficient(g)
print("%3.1f"%r)
nodes_by_deg = sorted(deg, key=deg.get, reverse=True)
mtrx = nx.to_numpy_matrix(g, nodelist=nodes_by_deg)
sns.heatmap(mtrx)
weight_sr = pd.Series([attrs["weight"] for s, t, attrs in g.edges(data=True)])
weight_sr.describe()
quant = weight_sr.quantile(.75)
plt.rcParams['figure.figsize'] = (17, 12)
heatmap = plt.imshow(mtrx)
heatmap.set_clim(0.0, quant)
plt.colorbar()
stripmtrx = mtrx[:, :100]
heatmap = plt.imshow(stripmtrx)
heatmap.set_clim(0.0, quant)
plt.colorbar()
zoommtrx = nx.to_numpy_matrix(g, nodelist=nodes_by_deg)[:50, :50]
sns.heatmap(zoommtrx)
zoomquant = pd.Series(zoommtrx.flatten().tolist()[0]).quantile(0.9)
# +
# heatmap = plt.imshow(zoommtrx)
# heatmap.set_clim(0.0, zoomquant)
# plt.colorbar()
# -
# #### Attribute
nx.attribute_assortativity_coefficient(g, "top_place")
nx.attribute_assortativity_coefficient(g, "top_genre")
nx.attribute_assortativity_coefficient(g, "role")
len(g.edges())
# +
def edge_types(g):
tps = {}
for s, t, attrs in g.edges(data=True):
srole = g.node[s].get("role", "")
trole = g.node[t].get("role", "")
if srole and trole:
if (trole, srole) in tps:
tps[(trole, srole)].append(attrs["weight"])
else:
tps.setdefault((srole, trole), [])
tps[(srole, trole)].append(attrs["weight"])
return tps
def edge_aggs(tps):
aggs = {}
for k, v in tps.items():
aggs[k] = (len(v) / 2174.0, sum(v) / len(v))
return aggs
# -
etps = edge_types(g)
aggs = edge_aggs(etps)
aggs
2174 * 0.025
pd.Series(etps[(u'patron', u'author')]).plot(kind="box")
# ### Clustering
nx.average_clustering(g)
pd.Series(nx.clustering(g).values()).hist()
nx.transitivity(g)
# ### Paths
nx.diameter(g)
nx.average_shortest_path_length(g)
|
onemode.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MAT281 - Laboratorio N°03
#
#
# Nombre: <NAME>
#
# Rol: 201610519-0
# <a id='p1'></a>
# ## I.- Problema 01
#
#
# <img src="https://freedesignfile.com/upload/2013/06/Car-logos-1.jpg" width="360" height="360" align="center"/>
#
#
# El conjunto de datos se denomina `vehiculos_procesado_con_grupos.csv`, el cual contine algunas de las características más importante de un vehículo.
#
# En este ejercicio se tiene como objetivo, es poder clasificar los distintos vehículos basados en las cracterísticas que se presentan a continuación. La dificultad de este ejercicio radíca en que ahora tenemos variables numéricas y variables categóricas.
#
# Lo primero será cargar el conjunto de datos:
# +
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import MinMaxScaler
from sklearn.dummy import DummyClassifier
from sklearn.cluster import KMeans
import warnings
warnings.filterwarnings("ignore")
# %matplotlib inline
sns.set_palette("deep", desat=.6)
sns.set(rc={'figure.figsize':(11.7,8.27)})
# +
# cargar datos
df = pd.read_csv(os.path.join("data","vehiculos_procesado_con_grupos.csv"), sep=",")\
.drop(
["fabricante",
"modelo",
"transmision",
"traccion",
"clase",
"combustible",
"consumo"],
axis=1)
df
# -
# En este caso, no solo se tienen datos numéricos, sino que también categóricos. Además, tenemos problemas de datos **vacíos (Nan)**. Así que para resolver este problema, seguiremos varios pasos:
# ### 1.- Normalizar datos
#
# 1. Cree un conjunto de datos con las variables numéricas, además, para cada dato vacía, rellene con el promedio asociado a esa columna. Finalmente, normalize los datos mediante el procesamiento **MinMaxScaler** de **sklearn**.
#
# 2.- Cree un conjunto de datos con las variables categóricas , además, transforme de variables categoricas a numericas ocupando el comando **get_dummies** de pandas ([referencia](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.get_dummies.html)). Explique a grande rasgo como se realiza la codificación de variables numéricas a categóricas.
#
# 3.- Junte ambos dataset en uno, llamado **df_procesado**.
#
# +
########Parte 1.A
df_num = df[['year','desplazamiento','cilindros','co2','consumo_litros_milla']] #solo columnas con valores numericos
for col in df_num.columns: #iteracion sobre columnas
df_num[col] = df_num[col].fillna(df_num[col].mean()) #Reemplazar los NaN por el promedio de la columna
df_num.isna().sum() #Verificacion de si quedan valores Nan
# +
##########Parte 1.B
scaler = MinMaxScaler()
df_num[df_num.columns] = scaler.fit_transform(df_num[df_num.columns])
df_num.head()
# +
#############Parte 2
df_cat = df.drop(['year','desplazamiento','cilindros','co2','consumo_litros_milla'], axis=1)
df_cat = pd.get_dummies(df_cat)
df_cat.head()
# +
#####Parte 3
df_procesado = pd.concat([df_num, df_cat], axis=1) #Se juntan ambos dataframe creados en la parte 1 y 2
df_procesado.head()
# -
# ### 2.- Realizar ajuste mediante kmeans
#
# Una vez depurado el conjunto de datos, es momento de aplicar el algoritmo de **kmeans**.
#
# 1. Ajuste el modelo de **kmeans** sobre el conjunto de datos, con un total de **8 clusters**.
# 2. Asociar a cada individuo el correspondiente cluster y calcular valor de los centroides de cada cluster.
# 3. Realizar un resumen de las principales cualidades de cada cluster. Para esto debe calcular (para cluster) las siguientes medidas de resumen:
# * Valor promedio de las variables numérica
# * Moda para las variables numericas
#
#
# +
#######Parte 1
X = np.array(df_procesado)
kmeans = KMeans(n_clusters= 8,n_init=25, random_state=123)
kmeans.fit(X)
# +
#########Parte 2
centroids = kmeans.cluster_centers_ # centros
clusters = kmeans.labels_ # clusters
df_procesado2 = df_procesado
df_procesado2["cluster"] = clusters
df_procesado2["cluster"] = df_procesado["cluster"].astype('category') #Tenemos el dataframe final con el cluster asociado a cada item
df_procesado2.head()
# -
for i in range(1,9):
df2 = df_procesado2.loc[df_procesado2['cluster'] == i]
df2 = df_procesado2[['year','desplazamiento','cilindros','co2','consumo_litros_milla']] #Solo variables numericas
df2[['year','desplazamiento','cilindros','co2','consumo_litros_milla']] = df[['year','desplazamiento','cilindros','co2','consumo_litros_milla']]
##Regresamos los valores originales antes de escalar para tener observaciones más interpretables
print('----------------------O----------------------')
print('Cluster' + str(i))
print('Promedio')
print(df2.mean())
print(' ')
print('Moda')
print(df.mean())
print('----------------------O----------------------')
# +
#######Centroides
centroids_df = pd.DataFrame(centroids)
centroids_df["cluster"] = [1,2,3,4,5,6,7,8]
centroids_df.head()
centroids_final = centroids_df.set_index('cluster')
centroids_final #Vemos el centroide de cada cluster, con el cluster como indice del dataframe
# -
# ### 3.- Elegir Número de cluster
#
# Estime mediante la **regla del codo**, el número de cluster apropiados para el caso.
# Para efectos prácticos, eliga la siguiente secuencia como número de clusters a comparar:
#
# $$[5, 10, 20, 30, 50, 75, 100, 200, 300]$$
#
# Una ve realizado el gráfico, saque sus propias conclusiones del caso.
#
# +
# implementación de la regla del codo
Nc = [5,10,20,30,50,75,100,200,300]
kmeans = [KMeans(n_clusters=i) for i in Nc]
score = [kmeans[i].fit(df_procesado).inertia_ for i in range(len(kmeans))]
df_Elbow = pd.DataFrame({'Number of Clusters':Nc,
'Score':score})
df_Elbow.head()
# -
# graficar los datos etiquetados con k-means
fig, ax = plt.subplots(figsize=(11, 8.5))
plt.title('Elbow Curve')
sns.lineplot(x="Number of Clusters",
y="Score",
data=df_Elbow)
sns.scatterplot(x="Number of Clusters",
y="Score",
data=df_Elbow)
plt.show()
# A partir de 200 clusters se ve que el Score baja y empieza a estabilizarse.
# ## II.- Problema 02
#
# <img src="https://live.staticflickr.com/7866/47075467621_85ab810139_c.jpg" align="center"/>
#
# Para el conjunto de datos de **Iris**, se pide realizar una reducción de dimensionalidad ocupando las técnicas de PCA y TSNE (vistas en clases).
#
# El objetivo es aplicar ambos algoritmos de la siguiente manera:
#
# * Análisis detallado algoritma PCA (tablas, gráficos, etc.)
# * Análisis detallado algoritma TSNE (tablas, gráficos, etc.)
# * Comparar ambos algoritmos (conclusiones del caso)
import pandas as pd
from sklearn.datasets import load_iris
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
from sklearn.pipeline import make_pipeline
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
from mpl_toolkits.mplot3d import Axes3D
import seaborn as sns
from sklearn.preprocessing import StandardScaler
# +
dataset = load_iris()
features = ['sepal_length', 'sepal_width', 'petal_length', 'petal_width']
target = 'species'
iris = pd.DataFrame(
dataset.data,
columns=features)
iris[target] = dataset.target
iris.head()
# -
# ### PCA
# +
# Entrenamiento de PCA con escalamiento de datos
pca_pipe = make_pipeline(StandardScaler(), PCA())
pca_pipe.fit(iris)
# Se extrae el modelo entrenado del pipeline
modelo_pca = pca_pipe.named_steps['pca']
df = pd.DataFrame(
data = modelo_pca.components_,
columns = iris.columns,
index = ['PC1', 'PC2', 'PC3', 'PC4', 'PC5']
)
df ##Datos despues de aplicarles PCA y escalamiento
# -
# Mapa de calor para ver la influencia de variables sobre otras
# ==============================================================================
plt.figure(figsize=(12,4))
componentes = modelo_pca.components_
plt.imshow(componentes.T, cmap='viridis', aspect='auto')
plt.yticks(range(len(iris.columns)), iris.columns)
plt.xticks(range(len(iris.columns)), np.arange(modelo_pca.n_components_) + 1)
plt.grid(False)
plt.colorbar();
# +
# graficar varianza por componente
percent_variance = np.round(modelo_pca.explained_variance_ratio_* 100, decimals =2)
columns = ['PC1', 'PC2', 'PC3', 'PC4','PC5']
plt.figure(figsize=(12,4))
plt.bar(x= range(1,6), height=percent_variance, tick_label=columns)
plt.xticks(np.arange(modelo_pca.n_components_) + 1)
plt.ylabel('Componente principal')
plt.xlabel('Por. varianza explicada')
plt.title('Porcentaje de varianza explicada por cada componente')
plt.show()
# +
# graficar varianza por la suma acumulada de los componente
percent_variance_cum = np.cumsum(percent_variance)
columns = ['PC1', 'PC1+PC2', 'PC1+PC2+PC3', 'PC1+PC2+PC3+PC4', 'PC1+PC2+PC3+PC4+PC5']
plt.figure(figsize=(12,4))
plt.bar(x= range(1,6), height=percent_variance_cum, tick_label=columns)
plt.ylabel('Percentate of Variance Explained')
plt.xlabel('Principal Component Cumsum')
plt.title('PCA Scree Plot')
plt.show()
# -
# Se puede ver que si se utilizan solo las 3 primeras componentes se tiene casi cerca de un 100% de la varianza explicada.
|
labs/lab_09_Cristobal_Lobos.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (tunnel)
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/wesleybeckner/technology_fundamentals/blob/main/C3%20Machine%20Learning%20I/LABS_PROJECT/Tech_Fun_C3_P4_Game_AI%2C_Heuristical_Agents.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="vhe1yX4AMckE"
# # Technology Fundamentals Course 3, Project Part 4: Heuristical Agents (Symbolic AI)
#
# **Instructor**: <NAME>
#
# **Contact**: <EMAIL>
#
# **Teaching Assitants**: <NAME>, <NAME>
#
# **Contact**: <EMAIL>, <EMAIL>
# <br>
#
# ---
#
# <br>
#
# We makin' some wack AI today
#
# <br>
#
# ---
#
# <br>
#
# <a name='top'></a>
#
# + [markdown] id="mNtJitcRW51Y"
# <a name='x.0'></a>
#
# ## 4.0 Preparing Environment and Importing Data
#
# [back to top](#top)
# + [markdown] id="chdcBoBL8SNm"
# <a name='x.0.1'></a>
#
# ### 4.0.1 Import Packages
#
# [back to top](#top)
# + id="eJ7-FDq3JhhI"
import random
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
class TicTacToe:
# can preset winner and starting player
def __init__(self, winner='', start_player=''):
self.winner = winner
self.start_player = start_player
self.board = {1: ' ',
2: ' ',
3: ' ',
4: ' ',
5: ' ',
6: ' ',
7: ' ',
8: ' ',
9: ' ',}
self.win_patterns = [[1,2,3], [4,5,6], [7,8,9],
[1,4,7], [2,5,8], [3,6,9],
[1,5,9], [7,5,3]]
# the other functions are now passed self
def visualize_board(self):
print(
"|{}|{}|{}|\n|{}|{}|{}|\n|{}|{}|{}|\n".format(*self.board.values())
)
def check_winning(self):
for pattern in self.win_patterns:
values = [self.board[i] for i in pattern]
if values == ['X', 'X', 'X']:
self.winner = 'X' # we update the winner status
return "'X' Won!"
elif values == ['O', 'O', 'O']:
self.winner = 'O'
return "'O' Won!"
return ''
def check_stalemate(self):
if (' ' not in self.board.values()) and (self.check_winning() == ''):
self.winner = 'Stalemate'
return "It's a stalemate!"
class GameEngine(TicTacToe):
def __init__(self, setup='auto'):
super().__init__()
self.setup = setup
def setup_game(self):
if self.setup == 'user':
players = int(input("How many Players? (type 0, 1, or 2)"))
self.player_meta = {'first': {'label': 'X',
'type': 'human'},
'second': {'label': 'O',
'type': 'human'}}
if players == 1:
first = input("who will go first? (X, (AI), or O (Player))")
if first == 'O':
self.player_meta = {'second': {'label': 'X',
'type': 'ai'},
'first': {'label': 'O',
'type': 'human'}}
else:
self.player_meta = {'first': {'label': 'X',
'type': 'ai'},
'second': {'label': 'O',
'type': 'human'}}
elif players == 0:
first = random.choice(['X', 'O'])
if first == 'O':
self.player_meta = {'second': {'label': 'X',
'type': 'ai'},
'first': {'label': 'O',
'type': 'ai'}}
else:
self.player_meta = {'first': {'label': 'X',
'type': 'ai'},
'second': {'label': 'O',
'type': 'ai'}}
elif self.setup == 'auto':
first = random.choice(['X', 'O'])
if first == 'O':
self.start_player = 'O'
self.player_meta = {'second': {'label': 'X',
'type': 'ai'},
'first': {'label': 'O',
'type': 'ai'}}
else:
self.start_player = 'X'
self.player_meta = {'first': {'label': 'X',
'type': 'ai'},
'second': {'label': 'O',
'type': 'ai'}}
def play_game(self):
while True:
for player in ['first', 'second']:
self.visualize_board()
player_label = self.player_meta[player]['label']
player_type = self.player_meta[player]['type']
if player_type == 'human':
move = input("{}, what's your move?".format(player_label))
# we're going to allow the user to quit the game from the input line
if move in ['q', 'quit']:
self.winner = 'F'
print('quiting the game')
break
move = int(move)
if self.board[move] != ' ':
while True:
move = input("{}, that position is already taken! "\
"What's your move?".format(player))
move = int(move)
if self.board[move] != ' ':
continue
else:
break
else:
while True:
move = random.randint(1,9)
if self.board[move] != ' ':
continue
print('test')
else:
break
self.board[move] = player_label
# the winner varaible will now be check within the board object
self.check_winning()
self.check_stalemate()
if self.winner == '':
continue
elif self.winner == 'Stalemate':
print(self.check_stalemate())
self.visualize_board()
break
else:
print(self.check_winning())
self.visualize_board()
break
if self.winner != '':
return self
# + [markdown] id="R7vDY-8U8SWd"
# <a name='x.0.1'></a>
#
# ### 4.0.2 Load Dataset
#
# [back to top](#top)
# + [markdown] id="h1YRk95hvZcH"
# ## 4.3 AI Heuristics
#
# Develop a better AI based on your analyses of game play so far.
# + [markdown] id="X742CiYgvKhX"
# ### Q1
#
# In our groups, let's discuss what rules we would like to hard code in. Harsha, Varsha and I will help you with the flow control to program these rules
# + id="igGzSx79BT47"
# we will define some variables to help us define the types of positions
middle = 5
side = [2, 4, 6, 8]
corner = [1, 3, 7, 9]
# + id="7cu_oGLOBQTT" colab={"base_uri": "https://localhost:8080/"} outputId="6eb253ee-b505-463f-9bcc-35e0a07b12d5"
# recall that our board is a dictionary
tictactoe = TicTacToe()
tictactoe.board
# + colab={"base_uri": "https://localhost:8080/"} id="Ee8gu3oU6Kqn" outputId="269bb60b-994f-4d27-972e-7cbea70d5407"
# and we have a win_patterns object to help us with the algorithm
tictactoe.win_patterns
# + [markdown] id="itBZnCGZvevG"
# for example, if we want to check if the middle piece is available, and play it if it is. How do we do that?
# + id="aidA_v6qvjU6"
# set some key variables
player = 'X'
opponent = 'O'
avail_moves = [i for i in tictactoe.board.keys() if tictactoe.board[i] == ' ']
# a variable that will keep track if we've found a move we like or not
move_found = False
# <- some other moves we might want to make would go here -> #
# and now for our middle piece play
if move_found == False: # if no other move has been found yet
if middle in avail_moves: # if middle is available
move_found = True # then change our move_found status
move = middle # update our move
# + [markdown] id="GhAM9sBnx-FS"
# Our standard approach will be to always ***return a move by the agent***. Whether the agent is heruistical or from some other ML framework we ***always want to return a move***
# + [markdown] id="nPvvfKhzyKed"
# Repeate after me: ***ALWAYS RETURN A MOVE***. Make sure you know what move is. Make sure you know what it is. And return it. Return a move. The purpose of the next lines of code we will write is to return a move.
#
# Make sure your code returns a move.
# + [markdown] id="0frNCaL6yX0S"
# ### Q2
#
# Write down your algorithm steps in markdown. i.e.
#
# 1. play a corner piece
# 2. play to opposite corner from the opponent, etc.
# 3. ....etc.
# + [markdown] id="MOaNULYLymHn"
# ### Q3
#
# Begin to codify your algorithm from Q3. Make sure that no matter what, you ***return a move***
# + id="DDgrKuKk7szs"
# some starting variables for you
player_label = 'X'
opponent = 'O'
avail_moves = [i for i in tictactoe.board.keys() if tictactoe.board[i] == ' ']
# temp board will allow us to play hypothetical moves and see where they get us
# in case you need it
temp_board = tictactoe.board.copy()
# + [markdown] id="xLgSm04oGXBI"
# ## 4.4 Wrapping our Agent
#
# Now that we've created a conditional tree for our AI to make a decision, we need to integrate this within the gaming framework we've made so far. How should we do this? Let's define this thought pattern or tree as an agent.
#
# Recall our play_game function within `GameEngine`
#
# + id="tPUpRGx-GWS0"
def play_game(self):
while True:
for player in ['first', 'second']:
self.visualize_board()
player_label = self.player_meta[player]['label']
player_type = self.player_meta[player]['type']
if player_type == 'human':
move = input("{}, what's your move?".format(player_label))
# we're going to allow the user to quit the game from the input line
if move in ['q', 'quit']:
self.winner = 'F'
print('quiting the game')
break
move = int(move)
if self.board[move] != ' ':
while True:
move = input("{}, that position is already taken! "\
"What's your move?".format(player))
move = int(move)
if self.board[move] != ' ':
continue
else:
break
########################################################################
##################### WE WANT TO CHANGE THESE LINES ####################
########################################################################
else:
while True:
move = random.randint(1,9)
if self.board[move] != ' ':
continue
print('test')
else:
break
self.board[move] = player_label
# the winner varaible will now be check within the board object
self.check_winning()
self.check_stalemate()
if self.winner == '':
continue
elif self.winner == 'Stalemate':
print(self.check_stalemate())
self.visualize_board()
break
else:
print(self.check_winning())
self.visualize_board()
break
if self.winner != '':
return self
# + [markdown] id="h93OZ_kaH68y"
# ### 4.4.1 Redefining the Random Agent
#
# In particular, we want to change lines 30-37 to take our gaming agent in as a parameter to make decisions. Let's try this.
#
# In `setup_game` we want to have the option to set the AI type/level. In `play_game` we want to make a call to that AI to make the move. For instance, our random AI will go from:
#
# ```
# while True:
# move = random.randint(1,9)
# if self.board[move] != ' ':
# continue
# else:
# break
# ```
#
# to:
#
# ```
# def random_ai(self):
# while True:
# move = random.randint(1,9)
# if self.board[move] != ' ':
# continue
# else:
# break
# return move
# ```
#
# + id="BrzdLPP2IA18"
class GameEngine(TicTacToe):
def __init__(self, setup='auto'):
super().__init__()
self.setup = setup
##############################################################################
########## our fresh off the assembly line tictactoe playing robot ###########
##############################################################################
def random_ai(self):
while True:
move = random.randint(1,9)
if self.board[move] != ' ':
continue
else:
break
return move
def setup_game(self):
if self.setup == 'user':
players = int(input("How many Players? (type 0, 1, or 2)"))
self.player_meta = {'first': {'label': 'X',
'type': 'human'},
'second': {'label': 'O',
'type': 'human'}}
if players != 2:
########################################################################
################# Allow the user to set the ai level ###################
########################################################################
level = int(input("select AI level (1, 2)"))
if level == 1:
self.ai_level = 1
elif level == 2:
self.ai_level = 2
else:
print("Unknown AI level entered, this will cause problems")
if players == 1:
first = input("who will go first? (X, (AI), or O (Player))")
if first == 'O':
self.player_meta = {'second': {'label': 'X',
'type': 'ai'},
'first': {'label': 'O',
'type': 'human'}}
else:
self.player_meta = {'first': {'label': 'X',
'type': 'ai'},
'second': {'label': 'O',
'type': 'human'}}
elif players == 0:
first = random.choice(['X', 'O'])
if first == 'O':
self.player_meta = {'second': {'label': 'X',
'type': 'ai'},
'first': {'label': 'O',
'type': 'ai'}}
else:
self.player_meta = {'first': {'label': 'X',
'type': 'ai'},
'second': {'label': 'O',
'type': 'ai'}}
elif self.setup == 'auto':
first = random.choice(['X', 'O'])
if first == 'O':
self.start_player = 'O'
self.player_meta = {'second': {'label': 'X',
'type': 'ai'},
'first': {'label': 'O',
'type': 'ai'}}
else:
self.start_player = 'X'
self.player_meta = {'first': {'label': 'X',
'type': 'ai'},
'second': {'label': 'O',
'type': 'ai'}}
##########################################################################
############## and automatically set the ai level otherwise ##############
##########################################################################
self.ai_level = 1
def play_game(self):
while True:
for player in ['first', 'second']:
self.visualize_board()
player_label = self.player_meta[player]['label']
player_type = self.player_meta[player]['type']
if player_type == 'human':
move = input("{}, what's your move?".format(player_label))
if move in ['q', 'quit']:
self.winner = 'F'
print('quiting the game')
break
move = int(move)
if self.board[move] != ' ':
while True:
move = input("{}, that position is already taken! "\
"What's your move?".format(player))
move = int(move)
if self.board[move] != ' ':
continue
else:
break
else:
if self.ai_level == 1:
move = self.random_ai()
######################################################################
############## we will leave this setting empty for now ##############
######################################################################
elif self.ai_level == 2:
pass
self.board[move] = player_label
self.check_winning()
self.check_stalemate()
if self.winner == '':
continue
elif self.winner == 'Stalemate':
print(self.check_stalemate())
self.visualize_board()
break
else:
print(self.check_winning())
self.visualize_board()
break
if self.winner != '':
return self
# + [markdown] id="d1-yoZ9BKHcl"
# Let's test that our random ai works now in this format
# + colab={"base_uri": "https://localhost:8080/"} id="p2o0N_MFJ0ha" outputId="056768f2-4ed6-400c-b94d-a33e93d6ef67"
random.seed(12)
game = GameEngine(setup='auto')
game.setup_game()
game.play_game()
# + [markdown] id="5_VJYE6ct5ug"
# Let's try it with a user player:
# + colab={"base_uri": "https://localhost:8080/"} id="L-u6SV9Qt7FL" outputId="63a980c3-56fb-4001-8946-e4b1a5ed407a"
random.seed(12)
game = GameEngine(setup='user')
game.setup_game()
game.play_game()
# + [markdown] id="m8C0hN4swbr8"
# ### Q4
#
# Now let's fold in our specialized AI agent. Add your code under the `heurstic_ai` function. Note that the `player_label` is passed as an input parameter now
# + id="9O1ZjPz0w4GP"
class GameEngine(TicTacToe):
def __init__(self, setup='auto'):
super().__init__()
self.setup = setup
##############################################################################
################### YOUR BADASS HEURISTIC AGENT GOES HERE ####################
##############################################################################
def heuristic_ai(self, player_label):
# SOME HELPER VARIABLES IF YOU NEED THEM
opponent = ['X', 'O']
opponent.remove(player_label)
opponent = opponent[0]
avail_moves = [i for i in self.board.keys() if self.board[i] == ' ']
temp_board = self.board.copy()
################## YOUR CODE GOES HERE, RETURN THAT MOVE! ##################
while True: # DELETE LINES 20 - 25, USED FOR TESTING PURPOSES ONLY
move = random.randint(1,9)
if self.board[move] != ' ':
continue
else:
break
############################################################################
return move
def random_ai(self):
while True:
move = random.randint(1,9)
if self.board[move] != ' ':
continue
else:
break
return move
def setup_game(self):
if self.setup == 'user':
players = int(input("How many Players? (type 0, 1, or 2)"))
self.player_meta = {'first': {'label': 'X',
'type': 'human'},
'second': {'label': 'O',
'type': 'human'}}
if players != 2:
########################################################################
################# Allow the user to set the ai level ###################
########################################################################
level = int(input("select AI level (1, 2)"))
if level == 1:
self.ai_level = 1
elif level == 2:
self.ai_level = 2
else:
print("Unknown AI level entered, this will cause problems")
if players == 1:
first = input("who will go first? (X, (AI), or O (Player))")
if first == 'O':
self.player_meta = {'second': {'label': 'X',
'type': 'ai'},
'first': {'label': 'O',
'type': 'human'}}
else:
self.player_meta = {'first': {'label': 'X',
'type': 'ai'},
'second': {'label': 'O',
'type': 'human'}}
elif players == 0:
first = random.choice(['X', 'O'])
if first == 'O':
self.player_meta = {'second': {'label': 'X',
'type': 'ai'},
'first': {'label': 'O',
'type': 'ai'}}
else:
self.player_meta = {'first': {'label': 'X',
'type': 'ai'},
'second': {'label': 'O',
'type': 'ai'}}
elif self.setup == 'auto':
first = random.choice(['X', 'O'])
if first == 'O':
self.start_player = 'O'
self.player_meta = {'second': {'label': 'X',
'type': 'ai'},
'first': {'label': 'O',
'type': 'ai'}}
else:
self.start_player = 'X'
self.player_meta = {'first': {'label': 'X',
'type': 'ai'},
'second': {'label': 'O',
'type': 'ai'}}
##########################################################################
############## and automatically set the ai level otherwise ##############
##########################################################################
self.ai_level = 1
def play_game(self):
while True:
for player in ['first', 'second']:
self.visualize_board()
player_label = self.player_meta[player]['label']
player_type = self.player_meta[player]['type']
if player_type == 'human':
move = input("{}, what's your move?".format(player_label))
if move in ['q', 'quit']:
self.winner = 'F'
print('quiting the game')
break
move = int(move)
if self.board[move] != ' ':
while True:
move = input("{}, that position is already taken! "\
"What's your move?".format(player))
move = int(move)
if self.board[move] != ' ':
continue
else:
break
else:
if self.ai_level == 1:
move = self.random_ai()
######################################################################
############## we will leave this setting empty for now ##############
######################################################################
elif self.ai_level == 2:
move = self.heuristic_ai(player_label)
self.board[move] = player_label
self.check_winning()
self.check_stalemate()
if self.winner == '':
continue
elif self.winner == 'Stalemate':
print(self.check_stalemate())
self.visualize_board()
break
else:
print(self.check_winning())
self.visualize_board()
break
if self.winner != '':
return self
# + [markdown] id="h-wBe5SZ0plM"
# ### Q5
#
# And we'll test that it works!
# + colab={"base_uri": "https://localhost:8080/"} id="3spYweFLxn7K" outputId="f6c2ba09-659f-49a9-b966-c9a7ecef5ba7"
random.seed(12)
game = GameEngine(setup='user')
game.setup_game()
game.play_game()
# + [markdown] id="wWlBsMNt02L2"
# ### Q6
#
# Test the autorun feature!
# + id="lGCjHo1f0GLq" colab={"base_uri": "https://localhost:8080/"} outputId="b3ac717e-5e47-4503-b27b-5b96b580389b"
game = GameEngine(setup='auto')
game.setup_game()
game.play_game()
|
C3 Machine Learning I/LABS_PROJECT/Tech_Fun_C3_P4_Game_AI,_Heuristical_Agents.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Summary
#
# In this chapter, we looked at the key concepts related to images. Images constitute rich information that's necessary to build your computer vision projects. OpenCV uses the `BGR` color format instead of `RGB`, but some Python packages (for example, Matplotlib) use the latter format. Therefore, we have covered how to convert the image from one color format into the other.
#
# Additionally, we have summarized the main functions and options to work with images:
#
# **Additionally, we have summarized the main functions and options to work with images:**
#
# * To access image properties
# * Some OpenCV functions, such as:-
# >cv2.imread(),<br>
# cv2.split(),<br>
# cv2.merge(),<br>
# cv2.imshow(),<br>
# cv2.waitKey(), and<br>
# cv2.destroyAllWindows()
#
# How to get and set image pixels in both BGR and grayscale images
# #### Q1: What are the main image-processing steps?
#
# * Get the image to work with. This process usually involves some functions so that you can read the image from different sources (camera, video stream, disk, online resources).
# * Process the image by applying image-processing techniques to achieve the required functionality (for example, detecting a cat in an image).
# * Show the result of the processing step (for example, drawing a bounding box in the image and then saving it to disk).
# #### Q2 What are the three processing levels?
#
# * Low-level
# * Mid-level
# * High-level
# #### Q3 What is the difference between a grayscale image and a black and white image?
#
# * The grayscale image has just one channel wchich is not displayed when we run .shape on it
# * Gray-scale has just 2 dimensions, rows and columns
# * Flipping the channel of the GrayScale image basically flips the image vertically
# * Gray and Black and White are pretty much the same
import cv2
import numpy as np
import matplotlib.pyplot as plt
gray = cv2.imread('logo.png', cv2.IMREAD_GRAYSCALE)
plt.imshow(gray)
plt.show()
# +
# Convert to Binary
(thresh, im_bw) = cv2.threshold(gray, 128, 255, cv2.THRESH_BINARY | cv2.THRESH_OTSU)
# -
plt.imshow(im_bw)
plt.show()
im_bw.shape == gray.shape
im_bw.ndim == gray.ndim
#
#
#
#
#
|
Ch.2/image_basics_in_OpenCV/.ipynb_checkpoints/05.summary_and_questions-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# orphan: true
# ---
# # Running Tune experiments with Skopt
#
# In this tutorial we introduce Skopt, while running a simple Ray Tune experiment. Tune’s Search Algorithms integrate with Skopt and, as a result, allow you to seamlessly scale up a Skopt optimization process - without sacrificing performance.
#
# Scikit-Optimize, or skopt, is a simple and efficient library to optimize expensive and noisy black-box functions, e.g. large-scale ML experiments. It implements several methods for sequential model-based optimization. Noteably, skopt does not perform gradient-based optimization, and instead uses computationally cheap surrogate models to
# approximate the expensive function. In this example we minimize a simple objective to briefly demonstrate the usage of Skopt with Ray Tune via `SkOptSearch`. It's useful to keep in mind that despite the emphasis on machine learning experiments, Ray Tune optimizes any implicit or explicit objective. Here we assume `scikit-opitmize==0.8.1` library is installed. To learn more, please refer to the [Scikit-Optimize website](https://scikit-optimize.github.io).
#
# + tags=["remove-cell"]
# # !pip install ray[tune]
# !pip install scikit-optimize==0.8.1
# !pip install sklearn==0.18.2
# -
# Click below to see all the imports we need for this example.
# You can also launch directly into a Binder instance to run this notebook yourself.
# Just click on the rocket symbol at the top of the navigation.
# + tags=["hide-input"]
import time
from typing import Dict, Optional, Any
import ray
import skopt
from ray import tune
from ray.tune.suggest import ConcurrencyLimiter
from ray.tune.suggest.skopt import SkOptSearch
# + tags=["remove-cell"]
ray.init(configure_logging=False)
# -
# Let's start by defining a simple evaluation function. Again, an explicit math formula is queried here for demonstration, yet in practice this is typically a black-box function-- e.g. the performance results after training an ML model. We artificially sleep for a bit (`0.1` seconds) to simulate a long-running ML experiment. This setup assumes that we're running multiple `step`s of an experiment while tuning three hyperparameters, namely `width`, `height`, and `activation`.
def evaluate(step, width, height, activation):
time.sleep(0.1)
activation_boost = 10 if activation=="relu" else 0
return (0.1 + width * step / 100) ** (-1) + height * 0.1 + activation_boost
# Next, our `objective` function to be optimized takes a Tune `config`, evaluates the `score` of your experiment in a training loop,
# and uses `tune.report` to report the `score` back to Tune.
def objective(config):
for step in range(config["steps"]):
score = evaluate(step, config["width"], config["height"], config["activation"])
tune.report(iterations=step, mean_loss=score)
# Next we define a search space. The critical assumption is that the optimal hyperparamters live within this space. Yet, if the space is very large, then those hyperparameters may be difficult to find in a short amount of time.
search_space = {
"steps": 100,
"width": tune.uniform(0, 20),
"height": tune.uniform(-100, 100),
"activation": tune.choice(["relu", "tanh"]),
}
# The search algorithm is instantiated from the `SkOptSearch` class. We also constrain the the number of concurrent trials to `4` with a `ConcurrencyLimiter`.
algo = SkOptSearch()
algo = ConcurrencyLimiter(algo, max_concurrent=4)
# The number of samples is the number of hyperparameter combinations that will be tried out. This Tune run is set to `1000` samples.
# (you can decrease this if it takes too long on your machine).
num_samples = 1000
# + tags=["remove-cell"]
# We override here for our smoke tests.
num_samples = 10
# -
# Finally, we run the experiment to `"min"`imize the "mean_loss" of the `objective` by searching `search_config` via `algo`, `num_samples` times. This previous sentence is fully characterizes the search problem we aim to solve. With this in mind, notice how efficient it is to execute `tune.run()`.
analysis = tune.run(
objective,
search_alg=algo,
metric="mean_loss",
mode="min",
name="skopt_exp",
num_samples=num_samples,
config=search_space
)
# We now have hyperparameters found to minimize the mean loss.
print("Best hyperparameters found were: ", analysis.best_config)
# ## Providing an initial set of hyperparameters
#
# While defining the search algorithm, we may choose to provide an initial set of hyperparameters that we believe are especially promising or informative, and
# pass this information as a helpful starting point for the `SkOptSearch` object. We also can pass the known rewards for these initial params to save on unnecessary computation.
initial_params = [
{"width": 10, "height": 0, "activation": "relu"},
{"width": 15, "height": -20, "activation": "tanh"}
]
known_rewards = [-189, -1144]
# Now the `search_alg` built using `SkOptSearch` takes `points_to_evaluate`.
algo = SkOptSearch(points_to_evaluate=initial_params)
algo = ConcurrencyLimiter(algo, max_concurrent=4)
# And again run the experiment, this time with initial hyperparameter evaluations:
analysis = tune.run(
objective,
search_alg=algo,
metric="mean_loss",
mode="min",
name="skopt_exp_with_warmstart",
num_samples=num_samples,
config=search_space
)
# And we again show the ideal hyperparameters.
print("Best hyperparameters found were: ", analysis.best_config)
# + tags=["remove-cell"]
ray.shutdown()
|
doc/source/tune/examples/skopt_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
# %pylab inline
import glob
import fitsne
from sklearn.decomposition import PCA, FastICA
from skimage.color import rgb2gray
from skimage.io import imread
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.manifold import TSNE
import umap
import seaborn as sns
sns.set_style('whitegrid')
sns.set_context('paper', font_scale=2)
from tqdm import tqdm
# +
#tumor_patches_dir = '../../../histopath_data/CAMELYON16/training/extracted_tumor_patches/level_5/'
#normal_patches_dir = '../../../histopath_data/CAMELYON16/training/extracted_normal_patches/level_5/'
normal_patches_dir = '../normal_patches_test/level_0/'
tumor_patches_dir = '../tumor_patches_test/level_0/'
np.random.seed(42)
master_matrix = []
label_matrix = []
y = []
list_of_tumor_files = list(glob.glob('{}*.png'.format(tumor_patches_dir)))
list_of_tumor_files = np.random.choice(list_of_tumor_files, 5000)
for f in tqdm(list_of_tumor_files):
master_matrix.append(imread(f))
label_matrix.append('tumor')
y.append(1)
tumor_count = len(label_matrix)
list_of_normal_files = list(glob.glob('{}*.png'.format(normal_patches_dir)))
list_of_normal_files = np.random.choice(list_of_normal_files, 5000)
#np.random.shuffle(list_of_normal_files)
for f in tqdm(list_of_normal_files):
master_matrix.append(imread(f))
label_matrix.append('normal')
y.append(0)
master_matrix = np.array(master_matrix)
y=np.array(y)
# -
shape = master_matrix.shape
master_matrix_reshaped = master_matrix.flatten().reshape(shape[0], shape[1]*shape[2]*shape[3])
# # PCA
# +
pca = PCA(n_components=2)
X_r = pca.fit(master_matrix_reshaped).transform(master_matrix_reshaped)
# Percentage of variance explained for each components
print('explained variance ratio (first two components): %s'
% str(pca.explained_variance_ratio_))
fig = plt.figure(figsize=(10, 10))
colors = ['navy', 'darkorange']
lw = 0.2
label_matrix = ['tumor', 'normal']
for color, i, target_name in zip(colors, [0, 1], label_matrix):
plt.scatter(X_r[y == i, 0], X_r[y == i, 1], color=color, alpha=.4, lw=lw,
label=target_name)
plt.legend(loc='best', shadow=False, scatterpoints=1)
plt.title('PCA ')
# -
# # LDA
# +
lda = LinearDiscriminantAnalysis(n_components=2)
X_r2 = lda.fit(master_matrix_reshaped, y).transform(master_matrix_reshaped)
fig = plt.figure(figsize=(10, 10))
for color, i, target_name in zip(colors, [0, 1], label_matrix):
plt.scatter(X_r2[y == i, 0], X_r2[y == i, 0], color=color, alpha=.8, lw=lw,
label=target_name)
plt.legend(loc='best', shadow=False, scatterpoints=1)
plt.title('LDA ')
# -
# # ICA
# +
ica = FastICA(n_components=2)
X_r = ica.fit(master_matrix_reshaped).transform(master_matrix_reshaped)
fig = plt.figure(figsize=(10, 10))
for color, i, target_name in zip(colors, [0, 1], label_matrix):
plt.scatter(X_r[y == i, 0], X_r[y == i, 1], color=color, alpha=.8, lw=lw,
label=target_name)
plt.legend(loc='best', shadow=False, scatterpoints=1)
plt.title('ICA ')
# -
# # UMAP
# +
embedding = umap.UMAP(n_neighbors=20,
min_dist=0.3,
metric='correlation').fit_transform(master_matrix_reshaped)
fig = plt.figure(figsize=(10, 10))
for color, i, target_name in zip(colors, [0, 1], label_matrix):
plt.scatter(embedding[y == i, 0], embedding[y == i, 1], color=color, alpha=.8, lw=lw,
label=target_name)
fig.tight_layout()
plt.title('UMAP')
# -
# # t-sne (slow)
# +
X_embedded = TSNE(n_components=2).fit_transform(master_matrix_reshaped)
fig = plt.figure(figsize=(10, 10))
for color, i, target_name in zip(colors, [0, 1], label_matrix):
plt.scatter(X_embedded[y == i, 0], X_embedded[y == i, 1], color=color, alpha=.8, lw=lw,
label=target_name)
fig.tight_layout()
plt.title('tSNE')
# -
# # FIt-SNE
# +
Y = fitsne.FItSNE(master_matrix_reshaped.astype(float))# max_iter=500)
fig = plt.figure(figsize=(10, 10))
for color, i, target_name in zip(colors, [0, 1], label_matrix):
plt.scatter(Y[y == i, 0], Y[y == i, 1], color=color, alpha=.8, lw=lw,
label=target_name)
plt.legend(loc='best', shadow=False, scatterpoints=1)
fig.tight_layout()
plt.title('FIt-SNE')
# -
|
notebooks/05.colorPCA-for-normal-tumor-images.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
class Employee:
raise_amt = 1.04
def __init__(self, first, last, pay):
self.first = first
self.last = last
self.email = first + '.' + last + '@email.<EMAIL>'
self.pay = pay
def fullname(self):
return '{} {}'.format(self.first, self.last)
def apply_raise(self):
self.pay = int(self.pay * self.raise_amt)
class Developer(Employee):
raise_amt = 1.10
def __init__(self, first, last, pay, prog_lang):
super().__init__(first, last, pay)
self.prog_lang = prog_lang
class Manager(Employee):
def __init__(self, first, last, pay, employees=None):
super().__init__(first, last, pay)
if employees is None:
self.employees = []
else:
self.employees = employees
def add_emp(self, emp):
if emp not in self.employees:
self.employees.append(emp)
def remove_emp(self, emp):
if emp in self.employees:
self.employees.remove(emp)
def print_emps(self):
for emp in self.employees:
print('-->', emp.fullname())
dev_1 = Developer('Corey', 'Schafer', 50000, 'Python')
dev_2 = Developer('Test', 'Employee', 60000, 'Java')
mgr_1 = Manager('Sue', 'Smith', 90000, [dev_1])
mgr_2 = Manager('John', 'Clark', 90000, [dev_1,dev_2])
print(mgr_2.email)
mgr_2.print_emps()
print(mgr_1.email)
mgr_1.add_emp(dev_2)
mgr_1.remove_emp(dev_2)
mgr_1.print_emps()
# this method helps to check the methods information, inherited as well as its own.
#print(help(Developer))
# -
|
OOP/Inheritance.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
import sys
import pydocx
import codecs
import pymysql
import pypandoc
import os
import base64
import imghdr
from wand.image import Image
from tqdm import tqdm
from docx import Document
from docx.shared import Inches, Cm
from bs4 import BeautifulSoup
def convert_to_jpeg(picture):
'''
convert non-support image into jpg form
'''
img = Image(filename=picture)
img.format = 'jpeg'
img.save(filename='tmp.jpg')
return 'tmp.jpg'
convert_to_jpeg('img.webp')
def p_discriminate(p):
'''
judge whether the line input in a picture
'''
if str(p).find("img") != -1:
return (0, "pic")
else:
return (1, "book")
def base64_to_img(b64_data, image_type):
'''
Convert the base64 data back to regular binary image data
and figure out the image type (png, gif, jpg, etc)
'''
image_data = base64.b64decode(b64_data)
# image_type = imghdr.what('', image_data)
# Create the image file and tell the user about it
destination_file_name = 'tmp_converted.' + str(image_type)
try:
destination = open(destination_file_name, 'wb')
except IOError:
print ("Unable to create image file. You might not have permission to create a file in this location.")
exit()
else:
destination.write(image_data)
destination.close()
return destination_file_name
# print ("New image file: {}".format(destination_file_name))
def re_the_src(img_src):
# print(img_src)
this_img = BeautifulSoup(img_src, features="lxml").find('img')
src = str(this_img.get('src'))
height = str(this_img.get('height'))
width = str(this_img.get('width'))
# print(src)
b64_data = src.split("base64,")[1]
image_type = src.split("base64,")[0].split("/")[1][:-1]
return b64_data, image_type, height, width
class tuban():
'''
a tuban class represents a tuban's basic structural information including imgs and the number of tuban
'''
def __init__(self, input_file, output_file):
'''
init the class
para: input_file: the file need to be rearrange
pare: output_file: the file after rearrange
'''
self.tuban_html = pydocx.PyDocX.to_html(input_file)
self.tuban_soup = BeautifulSoup(self.tuban_html, features="lxml")
self.input = input_file
self.output = output_file
self.errors = []
self.character = {}
self.ch_dict = {}
self.p_list = self.tuban_soup.select('p')
def parse(self):
'''
parse the file (html form)
para: self
'''
p_list = self.p_list
flag = 0
# if the pre_stage = 1 means that the situation is booknum, 0 means the pictures
pre_stage = 0
pre_tag = 0
this_booknum = ""
this_character = []
pre_num = ""
for p in tqdm(p_list):
if flag%2 == 0:
this_booknum = ""
else:
this_character = []
if flag%2 == 0:
stage = p_discriminate(p)[0]
if stage == 1:
if stage != pre_stage:
this_booknum = p.text
pre_stage = stage
else:
self.errors.append(("error 1 多重简号或者换行符有误", pre_num, p.text, "flag="+str(flag)))
return flag
else:
self.errors.append(("error 2 缺省简号或者字体截图中换行符有误", pre_num, p.text, "flag="+str(flag)))
return flag
else:
stage = p_discriminate(p)[0]
if stage == 0:
children = p.children
tag = 1 # 0: character, 1:image
cflag = 0
for child in children:
# if str(child)
if cflag%2 ==0:
if str(child).find("img") == -1:
tag = 0
pre_tag = tag
tmp_pair_character = str(child).strip()
else:
self.errors.append(("error 3 出现连续文字结点", this_booknum, p.text, "flag="+str(flag)))
return flag
else:
if str(child).find("img") != -1:
tag = 1
if tag != pre_tag:
tmp_pair_image = str(child)
else:
self.errors.append(("error 4 出现连续图片", this_booknum, p.text, "flag="+str(flag)))
return flag
else:
self.errors.append(("error 5 出现连续文字结点", this_booknum, str(child), "flag="+str(flag)))
return flag
if len(tmp_pair_character)!=0 and len(tmp_pair_image)!=0:
this_character.append((tmp_pair_character, tmp_pair_image))
cflag+=1
pre_stage = stage
else:
self.errors.append(("error 6 多重简号或者换行符有误", this_booknum, p.text, "flag=",flag))
return flag
# flag-=1
if len(this_booknum)!=0 and len(this_character)!=0 and flag%2==1:
# print(flag, len(this_booknum),len(this_character))
self.character[this_booknum] = this_character
pre_num = this_booknum
flag += 1
def array2dict(self):
array = self.character
self.ch_dict = {}
for k in array.keys():
for item in array[k]:
if item[0] not in self.ch_dict.keys():
self.ch_dict[item[0]] = []
self.ch_dict[item[0]].append((item[1], k))
else:
self.ch_dict[item[0]].append((item[1], k))
def get_error(self):
return self.errors
def get_output_html(self):
f = open(self.output, "w", encoding="utf8")
for k in self.ch_dict.keys():
print("<p>"+k+"</p>", file=f)
print("<p>", end="", file=f)
for item in self.ch_dict[k]:
print(item[0], end="", file=f) # img
print(item[1], end=",", file=f) # book
print("</p>", file=f)
f.close()
def get_output_docx_by_pandoc(self):
output = pypandoc.convert_file(self.output, 'docx', outputfile="out59.docx", extra_args=["-M8GB", "+RTS", "-K4096m", "-RTS"])
def get_output_docx_by_docx(self):
document = Document()
for k in tqdm(self.ch_dict.keys()):
paragraph_character = document.add_paragraph(k)
paragraph_image = document.add_paragraph()
for item in self.ch_dict[k]:
fig = re_the_src(item[0])
convert_to_jpeg(base64_to_img(fig[0], fig[1]))
paragraph_image.add_run().add_picture("tmp.jpg", width=Cm(1.5))
paragraph_image.add_run(item[1]) # book
document.save('demo.docx')
tb = tuban('../data/59.docx', 'res02.html')
tb.parse()
tb.array2dict()
tb.errors
tb.get_output_html()
tb.get_output_docx_by_docx()
# # python-docx example
document = Document()
paragraph = document.add_paragraph()
# paragraph = document.add_paragraph('first item in unordered list')
paragraph.add_run('dolor sit amet.')
paragraph.add_run().add_picture("tmp.jpg",width=Cm(2))
document.save('demo.docx')
|
app/utils/shuowen_base64.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import warnings
warnings.simplefilter('ignore')
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.cluster import KMeans
from sklearn.decomposition import PCA
from sklearn.metrics import silhouette_score
# %matplotlib inline
# -
# # Load Dataset
item_to_id = pd.read_csv('./data/grocery/item_to_id.csv')
item_to_id.head()
item_to_id.info()
# Unique ids
print(sorted(item_to_id['Item_id'].unique()))
history = pd.read_csv('./data/grocery/purchase_history.csv')
history.head()
history.info()
# # Data Processing
# ref: https://github.com/stasi009/TakeHomeDataChallenges/blob/master/09.ClusterGrocery/cluster_grocery.ipynb
def id_to_item(df):
""" function to convert id into counts """
# 'sum' here is adding two lists into one big list
ids = df['id'].str.split(',').sum()
id_list = [0 for i in range(1, 49)]
for i in ids:
id_list[int(i) - 1] += 1
return pd.Series(id_list, index=list(range(1, 49)))
# +
# get user_item count matrix
user_item_count = history.groupby('user_id').apply(id_to_item)
user_item_count.head()
# -
# # Question 1
# ### The customer who bought the most items overall in her lifetime
user_count = user_item_count.sum(axis=1).reset_index().rename(columns={0: 'count'})
user_count.sort_values(by='count', ascending=False).head()
# So the customer who bought the most items is 269335 with 72 items
# ### For each item, the customer who bought that product the most
# +
item_user_most = user_item_count.apply(np.argmax, axis=0).reset_index()
item_user_most = item_user_most.rename(columns={'index': 'Item_id', 0: 'User_id'})
df = pd.merge(left=item_user_most, right=item_to_id, on='Item_id', how='left')
df[['Item_id', 'Item_name', 'User_id']]
# -
# # Question 2
# define the feature matrix: item_id, user_id
feature = user_item_count.T
feature.head()
# +
# determine the best number of clusters
clusters = range(2, 30)
inertias = []
silhouettes = []
for n_clusters in clusters:
kmeans = KMeans(n_clusters=n_clusters, init='k-means++', random_state=42, n_jobs=-1)
kmeans = kmeans.fit(feature)
label = kmeans.predict(feature)
inertias.append(kmeans.inertia_)
silhouettes.append(silhouette_score(feature, label))
# -
# visualization
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(18, 6))
ax[0].plot(clusters, inertias, 'o-', label='Sum of Squared Distances')
ax[0].grid(True)
ax[1].plot(clusters, silhouettes, 'o-', label='Silhouette Coefficient')
ax[1].grid(True)
plt.legend(fontsize=12)
plt.tight_layout()
plt.show()
# It seems 21 clusters is the best choice. Now, let's apply PCA on the data and visualize the data.
# build K-Means model
kmeans = KMeans(n_clusters=21, init='k-means++', random_state=42, n_jobs=-1)
kmeans = kmeans.fit(feature)
label = kmeans.predict(feature)
# PCA for dimension reduction
pca = PCA(n_components=2, random_state=42)
pca = pca.fit(feature)
pca_feature = pca.transform(feature)
# +
# visualization in 2D
colors = ['b', 'g', 'r', 'c', 'm', 'y', 'k', 'w']
fig, ax = plt.subplots(figsize=(16, 16))
for j in range(len(pca_feature)):
x = pca_feature[j][0]
y = pca_feature[j][1]
ax.plot(x, y, 'o', color=colors[label[j] % 8])
name = item_to_id[item_to_id['Item_id'] == j + 1]['Item_name']
ax.annotate(xy=(x+0.2, y+0.2), s=name.values[0], fontsize=12)
plt.show()
# +
# print useful information
df = pd.DataFrame({'Item_id': list(range(1, 49)), 'label': label})
df = pd.merge(left=df, right=item_to_id, on='Item_id', how='left')
for i in range(0, 21):
print(i, '\t', list(df[df['label'] == i]['Item_name']))
# -
# Above clustering is based on the raw feature, namely the user purchase for each item. There are other methods for this problem. For example, this notebook contains another method: [notebook link](https://github.com/stasi009/TakeHomeDataChallenges/blob/master/09.ClusterGrocery/cluster_grocery.ipynb).
#
# With 21 clusters, some cluster looks very weird. Since there are only 48 different items, with one business expert in related area, it only take a few time to manually determine the best cluster numbers.
|
09. Clustering Grocery Items.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## This script loads the current model and performs an evaluation of it
# ### Initialize
# First, initialize the model with all parameters
#
# +
from data_source import DataSource
from visualize import Visualize
from sphere import Sphere
from model import Model
from loss import TripletLoss, ImprovedTripletLoss
from training_set import TrainingSet
from average_meter import AverageMeter
from data_splitter import DataSplitter
from mission_indices import MissionIndices
from database_parser import DatabaseParser
import torch
import torch.nn.functional as F
import torchvision
import torchvision.transforms as transforms
from torch.autograd import Variable
from torch.utils.tensorboard import SummaryWriter
from torchsummary import summary
import pyshtools
from pyshtools import spectralanalysis
from pyshtools import shio
from pyshtools import expand
import sys
import time
import math
import operator
import numpy as np
import pandas as pd
import open3d as o3d
import matplotlib.pyplot as plt
from matplotlib.backends.backend_pdf import PdfPages
from tqdm.auto import tqdm
import scipy.stats as st
from scipy import spatial
# %reload_ext autoreload
# %autoreload 2
# -
torch.cuda.set_device(0)
torch.backends.cudnn.benchmark = True
n_features = 2
bandwidth = 100
from model_relu_old import ModelOld
net = ModelOld(n_features, bandwidth).cuda()
restore = False
optimizer = torch.optim.SGD(net.parameters(), lr=5e-3, momentum=0.9)
batch_size = 12
num_workers = 12
descriptor_size = 256
net_input_size = 2*bandwidth
cache = 50
criterion = ImprovedTripletLoss(margin=2, alpha=0.5, margin2=0.2)
writer = SummaryWriter()
stored_model = './net_params_arche_low_res_small_lidar_only.pkl'
net.load_state_dict(torch.load(stored_model))
#summary(net, input_size=[(2, 200, 200), (2, 200, 200), (2, 200, 200)])
# Initialize the data source
# +
#dataset_path = "/media/scratch/berlukas/spherical/"
dataset_path = "/home/berlukas/data/arche_low_res2/"
db_parser = DatabaseParser(dataset_path)
training_missions, test_missions = MissionIndices.get_arche_low_res()
training_indices, test_indices = db_parser.extract_training_and_test_indices(
training_missions, test_missions)
print(f'Found {len(test_missions)} test indices.')
n_test_data = 2500
n_test_cache = n_test_data
ds_test = DataSource(dataset_path, n_test_cache, -1)
idx = np.array(test_indices['idx'].tolist())
ds_test.load(n_test_data, idx, filter_clusters=True)
n_test_data = len(ds_test.anchors)
# -
ds_test.rotate_all_positives('z', 20)
test_set = TrainingSet(restore, bandwidth)
test_set.generateAll(ds_test)
# +
# hack for removing the images
test_set.anchor_features = test_set.anchor_features[:,0:2,:,:]
test_set.positive_features = test_set.positive_features[:,0:2,:,:]
test_set.negative_features = test_set.negative_features[:,0:2,:,:]
n_test_set = len(test_set)
print("Total size: ", n_test_set)
test_loader = torch.utils.data.DataLoader(test_set, batch_size=10, shuffle=False, num_workers=1, pin_memory=True, drop_last=False)
# -
# ## Generate the descriptors for anchor and positive
# +
def accuracy(dista, distb):
margin = 0
pred = (dista - distb - margin).cpu().data
acc = ((pred < 0).sum()).float()/dista.size(0)
return acc
net.eval()
n_iter = 0
anchor_embeddings = np.empty(1)
positive_embeddings = np.empty(1)
with torch.no_grad():
test_accs = AverageMeter()
test_pos_dist = AverageMeter()
test_neg_dist = AverageMeter()
for batch_idx, (data1, data2, data3) in enumerate(test_loader):
embedded_a, embedded_p, embedded_n = net(data1.cuda().float(), data2.cuda().float(), data3.cuda().float())
dist_to_pos, dist_to_neg, loss, loss_total = criterion(embedded_a, embedded_p, embedded_n)
writer.add_scalar('Ext_Test/Loss', loss, n_iter)
acc = accuracy(dist_to_pos, dist_to_neg)
test_accs.update(acc, data1.size(0))
test_pos_dist.update(dist_to_pos.cpu().data.numpy().sum())
test_neg_dist.update(dist_to_neg.cpu().data.numpy().sum())
writer.add_scalar('Ext_Test/Accuracy', test_accs.avg, n_iter)
writer.add_scalar('Ext_Test/Distance/Positive', test_pos_dist.avg, n_iter)
writer.add_scalar('Ext_Test/Distance/Negative', test_neg_dist.avg, n_iter)
anchor_embeddings = np.append(anchor_embeddings, embedded_a.cpu().data.numpy().reshape([1,-1]))
positive_embeddings = np.append(positive_embeddings, embedded_p.cpu().data.numpy().reshape([1,-1]))
n_iter = n_iter + 1
desc_anchors = anchor_embeddings[1:].reshape([n_test_set, descriptor_size])
desc_positives = positive_embeddings[1:].reshape([n_test_set, descriptor_size])
# -
# ## Simple old testing pipeline (index based)
sys.setrecursionlimit(50000)
tree = spatial.KDTree(desc_positives)
p_norm = 2
max_pos_dist = 0.05
max_anchor_dist = 1
for n_nearest_neighbors in tqdm(range(1,21)):
pos_count = 0
anchor_count = 0
idx_count = 0
for idx in range(n_test_set):
nn_dists, nn_indices = tree.query(desc_anchors[idx,:], p = p_norm, k = n_nearest_neighbors)
nn_indices = [nn_indices] if n_nearest_neighbors == 1 else nn_indices
for nn_i in nn_indices:
if (nn_i >= n_test_set):
break;
dist = spatial.distance.euclidean(desc_positives[nn_i,:], desc_positives[idx,:])
if (dist <= max_pos_dist):
pos_count = pos_count + 1;
break
for nn_i in nn_indices:
if (nn_i >= n_test_set):
break;
dist = spatial.distance.euclidean(desc_positives[nn_i,:], desc_anchors[idx,:])
if (dist <= max_anchor_dist):
anchor_count = anchor_count + 1;
break
for nn_i in nn_indices:
if (nn_i == idx):
idx_count = idx_count + 1;
break
pos_precision = (pos_count*1.0) / n_test_set
anchor_precision = (anchor_count*1.0) / n_test_set
idx_precision = (idx_count*1.0) / n_test_set
print(f'recall {idx_precision} for {n_nearest_neighbors} neighbors')
writer.add_scalar('Ext_Test/Precision/Positive_Distance', pos_precision, n_nearest_neighbors)
writer.add_scalar('Ext_Test/Precision/Anchor_Distance', anchor_precision, n_nearest_neighbors)
writer.add_scalar('Ext_Test/Precision/Index_Count', idx_precision, n_nearest_neighbors)
# ## New testing pipeline (location based)
# +
print(f'Running test pipeline for a map size of {len(desc_positives)} descriptors.')
sys.setrecursionlimit(50000)
tree = spatial.KDTree(desc_positives)
p_norm = 2
max_pos_dist = 5.0
max_anchor_dist = 1
anchor_poses = ds_test.anchor_poses
positive_poses = ds_test.positive_poses
assert len(anchor_poses) == len(positive_poses)
for n_nearest_neighbors in tqdm(range(1,21)):
loc_count = 0
for idx in range(n_test_set):
nn_dists, nn_indices = tree.query(desc_anchors[idx,:], p = p_norm, k = n_nearest_neighbors)
nn_indices = [nn_indices] if n_nearest_neighbors == 1 else nn_indices
for nn_i in nn_indices:
if (nn_i >= n_test_set):
break;
dist = spatial.distance.euclidean(positive_poses[nn_i,5:8], anchor_poses[idx,5:8])
if (dist <= max_pos_dist):
loc_count = loc_count + 1;
break
loc_precision = (loc_count*1.0) / n_test_set
print(f'recall {loc_precision} for {n_nearest_neighbors} neighbors')
#print(f'{loc_precision}')
#writer.add_scalar('Ext_Test/Precision/Location', loc_precision, n_nearest_neighbors)
# -
# ## Place Voting using Global Spectral Analysis
#
# +
print(f'Running test pipeline for a map size of {len(desc_positives)} descriptors.')
sys.setrecursionlimit(50000)
tree = spatial.KDTree(desc_positives)
p_norm = 2
max_pos_dist = 5.0
anchor_poses = ds_test.anchor_poses
anchor_clouds = ds_test.anchors
anchor_features = test_set.anchor_features
positive_poses = ds_test.positive_poses
positive_clouds = ds_test.positives
positive_features = test_set.anchor_features
for n_nearest_neighbors in tqdm(range(1,21)):
n_matches = 0
loc_count = 0
for idx in range(0, n_test_set):
nn_dists, nn_indices = tree.query(desc_anchors[idx,:], p = p_norm, k = n_nearest_neighbors)
nn_indices = [nn_indices] if n_nearest_neighbors == 1 else nn_indices
z_scores = [0] * n_nearest_neighbors
contains_match = False
true_match_idx = 0
for i in range(0, n_nearest_neighbors):
nn_i = nn_indices[i]
if (nn_i >= n_test_set):
print(f'ERROR: index {nn_i} is outside of {n_data}')
break;
dist = spatial.distance.euclidean(positive_poses[nn_i,5:8], anchor_poses[idx,5:8])
if (dist <= max_pos_dist):
contains_match = True
true_match_idx = i
a_range = anchor_features[idx][0,:,:]
p_range = positive_features[nn_i][0,:,:]
a_intensity = anchor_features[idx][1,:,:]
p_intensity = positive_features[nn_i][1,:,:]
#a_img = anchor_features[idx][2,:,:]
#p_img = positive_features[nn_i][2,:,:]
a_range_coeffs = pyshtools.expand.SHExpandDH(a_range, sampling=1)
p_range_coeffs = pyshtools.expand.SHExpandDH(p_range, sampling=1)
a_intensity_coeffs = pyshtools.expand.SHExpandDH(a_intensity, sampling=1)
p_intensity_coeffs = pyshtools.expand.SHExpandDH(p_intensity, sampling=1)
#a_img_coeffs = pyshtools.expand.SHExpandDH(a_img, sampling=1)
#p_img_coeffs = pyshtools.expand.SHExpandDH(p_img, sampling=1)
#a_fused = np.empty([3, a_range_coeffs.shape[0], a_range_coeffs.shape[1]])
#p_fused = np.empty([3, p_range_coeffs.shape[0], p_range_coeffs.shape[1]])
#print(a_range_coeffs.shape)
#a_fused[0,:] = a_range_coeffs
admit, error, corr = spectralanalysis.SHAdmitCorr(a_range_coeffs, p_range_coeffs)
for l in range(0, 4):
prob = spectralanalysis.SHConfidence(l, corr[l])
score = st.norm.ppf(1-(1-prob)/2) if prob < 0.99 else 4.0
z_scores[i] = z_scores[i] + score
#if math.isinf(z_scores[i]):
#print(f'z-score is inf: prob = {prob}, z-score {st.norm.ppf(1-(1-prob)/2)}')
#if (contains_match is not True):
#print(f'Match not found for index {idx} and {n_nearest_neighbors} neighbors')
#continue
n_matches = n_matches + 1
max_index, max_z_score = max(enumerate(z_scores), key=operator.itemgetter(1))
matching_index = nn_indices[max_index]
dist = spatial.distance.euclidean(positive_poses[matching_index,5:8], anchor_poses[idx,5:8])
if (dist <= max_pos_dist):
loc_count = loc_count + 1;
else:
#print(f'Place invalid: distance anchor <-> positive: {dist} with score {max_z_score}.')
matching_index = nn_indices[true_match_idx]
dist = spatial.distance.euclidean(positive_poses[matching_index,5:8], positive_poses[true_match_idx,5:8])
#print(f'Distance positive <-> true_match: {dist}, true_match score: {z_scores[true_match_idx]}')
loc_precision = (loc_count*1.0) / n_matches
#print(f'Recall {loc_precision} for {n_nearest_neighbors} neighbors with {n_matches}/{n_data} correct matches.')
print(f'{loc_precision}')
writer.add_scalar('Ext_Test/Precision/Voting', loc_precision, n_nearest_neighbors)
# -
# ## Place Voting using Global Spectral Analysis
#
# +
print(f'Running test pipeline for a map size of {len(desc_positives)} descriptors.')
sys.setrecursionlimit(50000)
tree = spatial.KDTree(desc_positives)
p_norm = 2
max_pos_dist = 5.0
anchor_poses = ds_test.anchor_poses
anchor_clouds = ds_test.anchors
anchor_features = test_set.anchor_features
positive_poses = ds_test.positive_poses
positive_clouds = ds_test.positives
positive_features = test_set.anchor_features
for n_nearest_neighbors in tqdm(range(1,21)):
n_matches = 0
loc_count = 0
final_count = 0
for idx in range(0, n_test_set):
nn_dists, nn_indices = tree.query(desc_anchors[idx,:], p = p_norm, k = n_nearest_neighbors)
nn_indices = [nn_indices] if n_nearest_neighbors == 1 else nn_indices
z_scores_range = [0] * n_nearest_neighbors
z_scores_intensity = [0] * n_nearest_neighbors
z_scores_image = [0] * n_nearest_neighbors
contains_match = False
true_match_idx = 0
for i in range(0, n_nearest_neighbors):
nn_i = nn_indices[i]
if (nn_i >= n_test_set):
print(f'ERROR: index {nn_i} is outside of {n_data}')
break;
dist = spatial.distance.euclidean(positive_poses[nn_i,5:8], anchor_poses[idx,5:8])
if (dist <= max_pos_dist):
contains_match = True
true_match_idx = i
a_range = anchor_features[idx][0,:,:]
p_range = positive_features[nn_i][0,:,:]
a_intensity = anchor_features[idx][1,:,:]
p_intensity = positive_features[nn_i][1,:,:]
#a_img = anchor_features[idx][2,:,:]
#p_img = positive_features[nn_i][2,:,:]
a_range_coeffs = pyshtools.expand.SHExpandDH(a_range, sampling=1)
p_range_coeffs = pyshtools.expand.SHExpandDH(p_range, sampling=1)
a_intensity_coeffs = pyshtools.expand.SHExpandDH(a_intensity, sampling=1)
p_intensity_coeffs = pyshtools.expand.SHExpandDH(p_intensity, sampling=1)
#a_img_coeffs = pyshtools.expand.SHExpandDH(a_img, sampling=1)
#p_img_coeffs = pyshtools.expand.SHExpandDH(p_img, sampling=1)
tapers, eigenvalues, taper_order = spectralanalysis.SHReturnTapers(2.01, 1)
saa_range = spectralanalysis.spectrum(a_range_coeffs)
saa_intensity = spectralanalysis.spectrum(a_intensity_coeffs)
#saa_img = spectralanalysis.spectrum(a_img_coeffs)
saa = np.empty([n_features, saa_range.shape[0]])
saa[0,:] = saa_range
saa[1,:] = saa_intensity
#saa[2,:] = saa_img
#saa = np.mean(saa, axis=0)
saa = np.amax(saa, axis=0)
spp_range = spectralanalysis.spectrum(p_range_coeffs)
spp_intensity = spectralanalysis.spectrum(p_intensity_coeffs)
#spp_img = spectralanalysis.spectrum(p_img_coeffs)
spp = np.empty([n_features, spp_range.shape[0]])
spp[0,:] = saa_range
spp[1,:] = saa_intensity
#spp[2,:] = saa_img
#spp = np.mean(spp, axis=0)
spp = np.amax(spp, axis=0)
sap_range = spectralanalysis.cross_spectrum(a_range_coeffs, p_range_coeffs)
sap_intensity = spectralanalysis.cross_spectrum(a_intensity_coeffs, p_intensity_coeffs)
#sap_img = spectralanalysis.cross_spectrum(a_img_coeffs, p_img_coeffs)
sap = np.empty([n_features, sap_range.shape[0]])
sap[0,:] = saa_range
sap[1,:] = saa_intensity
#sap[2,:] = saa_img
#sap = np.mean(sap, axis=0)
sap = np.amax(sap, axis=0)
#saa = spectralanalysis.spectrum(a_coeffs)
#spp = spectralanalysis.spectrum(p_coeffs)
#sap = spectralanalysis.cross_spectrum(a_coeffs, p_coeffs)
#admit, corr = spectralanalysis.SHBiasAdmitCorr(sap_img, saa_img, spp_img, tapers)
#admit, corr = spectralanalysis.SHBiasAdmitCorr(sap, saa, spp, tapers)
admit, corr = spectralanalysis.SHBiasAdmitCorr(sap_range, saa_range, spp_range, tapers)
for l in range(0, 10):
prob = spectralanalysis.SHConfidence(l, corr[l])
score = st.norm.ppf(1-(1-prob)/2) if prob < 0.99 else 4.0
z_scores_range[i] = z_scores_range[i] + score
admit, corr = spectralanalysis.SHBiasAdmitCorr(sap_intensity, saa_intensity, spp_intensity, tapers)
for l in range(0, 10):
prob = spectralanalysis.SHConfidence(l, corr[l])
score = st.norm.ppf(1-(1-prob)/2) if prob < 0.99 else 4.0
z_scores_intensity[i] = z_scores_intensity[i] + score
#if (contains_match is not True):
#print(f'Match not found for index {idx} and {n_nearest_neighbors} neighbors')
#continue
n_matches = n_matches + 1
max_index_range, max_z_score_range = max(enumerate(z_scores_range), key=operator.itemgetter(1))
max_index_intensity, max_z_score_intensity = max(enumerate(z_scores_intensity), key=operator.itemgetter(1))
#print(f'max range: {max_z_score_range}, max intensity: {max_z_score_intensity}')
max_index = max_index_range if max_z_score_range > max_z_score_intensity else max_index_intensity
matching_index = nn_indices[max_index]
dist = spatial.distance.euclidean(positive_poses[matching_index,5:8], anchor_poses[idx,5:8])
if (dist <= max_pos_dist):
loc_count = loc_count + 1;
else:
#print(f'Place invalid: distance anchor <-> positive: {dist} with score {max_z_score}.')
matching_index = nn_indices[true_match_idx]
dist = spatial.distance.euclidean(positive_poses[matching_index,5:8], positive_poses[true_match_idx,5:8])
#print(f'Distance positive <-> true_match: {dist}, true_match score: {z_scores[true_match_idx]}')
loc_precision = (loc_count*1.0) / n_matches
#print(f'Recall {loc_precision} for {n_nearest_neighbors} neighbors with {n_matches}/{n_data} correct matches.')
print(f'{loc_precision}')
writer.add_scalar('Ext_Test/Precision/WindowedVoting', loc_precision, n_nearest_neighbors)
# -
anchor_poses[0:100,5:8]
a = np.matrix('1 2; 3 4')
b = np.matrix('4 5; 6 7')
c = np.empty([2,2,2])
c[0,:,:] = a
c[1,:,:] = b
np.mean(c, axis=0)
a.shape[0]
|
script/playground/ext_evaluation_rotated.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # <center>RGBIF<center>
# ## <center>working with GBIF occurrence data in R<center>
#
# <center><img src="..\images\GBIF&R.png"><center>
# + [markdown] slideshow={"slide_type": "subslide"}
# ## <center>What is RGBIF?<center>
# + [markdown] slideshow={"slide_type": "fragment"}
# RGBIF is a **R wrapper** around the [GBIF API](https://www.gbif.org/developer/summary "GBIF API") to allow you to talk to GBIF from R.
# + [markdown] slideshow={"slide_type": "slide"}
# ## <center>RGBIF installation and loading libraries<center>
# + [markdown] slideshow={"slide_type": "fragment"}
# ### <center>Installation<center>
# We can install RGBIF directly from CRAN as any other R package:
# ```r
# install.packages("rgbif")
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ### <center>Load libraries<center>
# + slideshow={"slide_type": "skip"}
# Basic packages
library(assertthat)
library(stringr)
# + slideshow={"slide_type": "skip"}
# Tidyverse packages (not shown for brevity, still important to load!)
library(purrr)
library(dplyr)
library(magrittr)
library(tidyr)
# + slideshow={"slide_type": "skip"}
# Graphic packages (not shown for brevity, still important to load!)
library(ggplot2)
library(sp)
library(ggmap)
library(rgdal)
library(leaflet)
library(IRdisplay)
# + slideshow={"slide_type": "subslide"}
# RGBIF package
library(rgbif)
# + [markdown] slideshow={"slide_type": "slide"}
# ## <center>Download GBIF occurrences: `occ_data`<center>
#
# <center>download GBIF occurrence data *on the fly*<center>
# + [markdown] slideshow={"slide_type": "subslide"}
# ### <center>Download GBIF occurrences: `occ_data`<center>
#
# * search species occurrences by
# * scientific name (`scientificName` or `taxonKey` from the GBIF backbone),
# * geographical areas (`country`, `continent`, `decimalLatitude` `decimalLongitude`, ...),
# * temporal windows (`year`, `month`, ...),
# * datasets, `datasetKey`
#
# Check [RGBIF documentation](https://cran.r-project.org/web/packages/rgbif/rgbif.pdf "RGBIF") for the full list of options.
#
# + [markdown] slideshow={"slide_type": "slide"}
# In this tutorial we search occurrences data of:
# * *Vanessa atalanta Linnaeus, 1758*, a butterfly, best known as red admiral.
# 
# + slideshow={"slide_type": "fragment"}
species <- "Vanessa atalanta Linnaeus, 1758"
# + [markdown] slideshow={"slide_type": "subslide"}
# * during 2016
# + slideshow={"slide_type": "fragment"}
year <- 2016
# + [markdown] slideshow={"slide_type": "fragment"}
# * in Belgium
# + slideshow={"slide_type": "fragment"}
country <- "BE"
# + [markdown] slideshow={"slide_type": "subslide"}
# ```r
# occurrences <- rgbif::occ_data(
# scientificName = species,
# country = country,
# year = year,
# limit = 200000)
# ```
# + slideshow={"slide_type": "skip"}
load("../data/interim/occurrences.rda") # used during presentation to not wait for getting occurrences via occ_data
# + slideshow={"slide_type": "subslide"}
head(occurrences)
# + [markdown] slideshow={"slide_type": "subslide"}
# The interesting part is the `data` itself:
# + slideshow={"slide_type": "fragment"}
occ_Vanessa_BE <- occurrences$data
head(occ_Vanessa_BE[c("decimalLongitude", "decimalLatitude", "eventDate",
"scientificName")])
# + [markdown] slideshow={"slide_type": "slide"}
# # <center>Temporal occurrence distribution<center>
# Retrieving the monthly occurrence distribution:
# + slideshow={"slide_type": "fragment"}
counts_per_month <- count(occ_Vanessa_BE, month)
gg_month <- ggplot(counts_per_month, aes(y = n, x = month)) +
geom_bar(stat="identity", fill = "#0072B2") +
scale_x_continuous(breaks=seq(1,12,1),
labels = c("Jan", "Feb", "Mar", "Apr", "May", "Jun",
"Jul", "Aug", "Sep", "Oct", "Nov", "Dec"))
# + slideshow={"slide_type": "skip"}
options(repr.plot.width=8, repr.plot.height=3)
# + slideshow={"slide_type": "subslide"}
gg_month
# + [markdown] slideshow={"slide_type": "slide"}
# # <center>Geographical occurrence distribution<center>
# + [markdown] slideshow={"slide_type": "fragment"}
# ## <center>`gbifmap`<center>
# Plot function in RGBIF: *`gbifmap`*
# + slideshow={"slide_type": "skip"}
options(repr.plot.width=6, repr.plot.height=4)
# + slideshow={"slide_type": "subslide"}
rgbif::gbifmap(input = occ_Vanessa_BE, mapdatabase = "world", region = "Belgium")
# + [markdown] slideshow={"slide_type": "subslide"}
# Basic output. Perfectly for quick screening, but we should get this cleaner...
# + [markdown] slideshow={"slide_type": "subslide"}
# ### <center>HOW?<center>
# We have a dataframe with geographical coordinates (*latitude* and *longitude*) of the occurrences.
#
# We could:
# + [markdown] slideshow={"slide_type": "fragment"}
#
# - divide Belgium in square cells
# - __count__ the number of occurrences with coordinates within each grid cell
# * plot grid and counts (*heatmap*)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### <center>Occurrence geographical distribution based on UTM-10 grid<center>
# Let's start from the UTM-10 grid for Belgium, i.e. a set of cells covering all Belgium by means of squares with a side length of 10 km.
# <center><center>
# + [markdown] slideshow={"slide_type": "subslide"}
# Quickly screen the UTM-10 grid for Belgium:
# + slideshow={"slide_type": "fragment"}
eu_10grid <- readOGR("../data/external/EUgrid10.geojson")
plot(eu_10grid)
# + [markdown] slideshow={"slide_type": "subslide"}
# we need this regularly...
# so we've written a little function for it:
# <center>`counts_pts_in_polygons`<center>
# + slideshow={"slide_type": "subslide"}
source("counts_pts_in_polygons.R")
polygons_with_counts_df <- counts_pts_in_polygons(occ_Vanessa_BE, eu_10grid)
# + [markdown] slideshow={"slide_type": "skip"}
# Remove squares contanining no occurrences
# + slideshow={"slide_type": "skip"}
polygons_with_counts_df_subset <- subset(polygons_with_counts_df, value != 0)
# + [markdown] slideshow={"slide_type": "skip"}
# Choose a color palette
# ```r
# pal <- colorNumeric("viridis", domain = NULL)
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# And make a leaflet map, `n_occ_grid_Vanessa_BE_leaflet`
# + [markdown] slideshow={"slide_type": "subslide"}
# ```r
# n_occ_grid_Vanessa_BE_leaflet <- leaflet() %>%
# addTiles() %>%
# addPolygons(
# data = polygons_with_counts_df_subset,
# stroke = FALSE,
# fillColor = ~pal(value),
# fillOpacity = 0.7,
# smoothFactor = 0.1,
# label = ~paste(value,
# species,
# sep = ": "))
# ```
# + [markdown] slideshow={"slide_type": "skip"}
# ```r
# addLegend(pal = pal,
# values = polygons_with_counts_df_subset$value,
# opacity = 1.0,
# title = "OCCURRENCES")
# ```
# + [markdown] slideshow={"slide_type": "skip"}
# Saving the map as HTML file
# ```r
# library(htmlwidgets)
# saveWidget(widget = n_occ_grid_Vanessa_BE_leaflet,
# file = "EUgrid10_occ_Vanessa_BE_2016.html")
# file.rename(from = "./EUgrid10_occ_Vanessa_BE_2016.html",
# to = "../data/processed/EUgrid10_occ_Vanessa_BE_2016.html")
# ```
# + slideshow={"slide_type": "subslide"}
IRdisplay::display_html('<iframe src="../data/processed/EUgrid10_occ_Vanessa_BE_2016.html" width=1000, height=500></iframe> ')
# + [markdown] slideshow={"slide_type": "slide"}
# ## <center>More data to download?<center>
# * `occ_data` has a hard limit of 200000!
# * use asynchronuous download function: `occ_download`
#
# But that's another story...
#
# * Do you want to know more about `occ_download` and in general about the potential of using GBIF in R?
# * Would you like to produce maps like this one?
# + slideshow={"slide_type": "subslide"}
IRdisplay::display_html('<iframe src="../data/external/EUgrid10_occ_Vanessa&Phylloscopus_SE_2016.html" width=1000, height=450></iframe> ')
# + [markdown] slideshow={"slide_type": "subslide"}
# ### <center>Coming soon: a blog post at [LifeWatch INBO](https://lifewatch.inbo.be/blog/)!<center>
#
# <center>Download all INBO tutorials [here](https://github.com/inbo/lifewatch-meeting-2018) and try yourself!<center>
#
# <center><center>
# <center>**Open source for open science**<center>
#
# <center>[@LifeWatchINBO](https://twitter.com/LifeWatchINBO)<center>
# <center>[@INBOVlaanderen](https://twitter.com/INBOVlaanderen)<center>
#
# <center>[INBO](https://www.inbo.be/)<center>
#
|
package_tutorials/src/tutorial_rgbif_package.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from IPython.display import display_html
stocks_2016=pd.read_csv('data/stocks_2016.csv',index_col='Symbol')
stocks_2017=pd.read_csv('data/stocks_2017.csv',index_col='Symbol')
s_list=[stocks_2016,stocks_2017]
# +
# concatenating two DataFrames with index aligning function pd.concat()
# -
pd.concat(s_list)
pd.concat(s_list,keys=['2016','2017'],names=['Year','Symbol'])
pd.concat(s_list,keys=['2016','2017'],axis=1,sort=False).style.highlight_null()
pd.concat(s_list,axis='columns',keys='2016 2017'.split(),join='inner').stack(0).swaplevel(0,1)
# +
# pd.concat() vs join() vs merge()
# -
years=2016,2017,2018
years
stock_tables=[pd.read_csv('data/stocks_{}.csv'.format(year),index_col='Symbol') for year in years]
def display_frames(frames,num_spaces=10):
t_style = '<table style="display: inline;"'
tables_html = [df.to_html().replace('<table', t_style)
for df in frames]
space = ' ' * num_spaces
display_html(space.join(tables_html), raw=True)
display_frames(stock_tables,30)
stocks_2016,stocks_2017,stocks_2018=stock_tables
pd.concat(stock_tables,keys=[2016,2017,2018],names=['Year','Symbol'])
pd.concat(dict(zip(years,stock_tables)),axis=1,sort=False).style.highlight_null()
stocks_2016.join(stocks_2017,lsuffix='_2016',rsuffix='_2017',how='outer')
other=[stocks_2017.add_suffix('_2017'),stocks_2018.add_suffix('_2018')]
stocks_2016.add_suffix('_2016').join(other,how='outer')
# +
# checking for equality
# -
stock_join=stocks_2016.add_suffix('_2016').join(other,how='outer')
stock_concat=pd.concat([eval("stocks{0}.add_suffix('{1}')".format('_'+str(year),'_'+str(year))) for year in years]
,axis='columns')
stock_concat
# +
# asserting an equality
# -
stock_join.equals(stock_concat)
# +
# merge()
# -
stocks_2016.merge(stocks_2017,left_index=True,right_index=True,how='outer')
stocks_2016.merge(stocks_2017,left_index=True,right_index=True,suffixes=('_2016','_2017'),how="outer")
stock_merge=stocks_2016.merge(stocks_2017,
left_index=True,right_index=True,suffixes=('_2016','_2017'),how="outer").merge(stocks_2018.add_suffix('_2018'),
left_index=True,right_index=True,how='outer')
# +
# asserting equality once again
# -
stock_merge.equals(stock_concat)
|
concatenating multiple DataFrames together.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
import etherscan.tokens as tokens
import json
# -
# ## Import API key from file
with open('../../api_key.json', mode='r') as key_file:
key = json.loads(key_file.read())['key']
# ## Initiate the Etherscan API
# Start by declaring the ERC20 token contract address
# These can be found by searching Etherscan.io for the coin you want
contract_address = '0x57d90b64a1a57749b0f932f1a3395792e12e7055'
api = tokens.Tokens(contract_address=contract_address, api_key=key)
# ## Get token balance of an address
address = '0xe04f27eb70e025b78871a2ad7eabe85e61212761'
api.get_token_balance(address=address)
# ## Get total supply of tokens
api.get_total_supply()
|
examples/tokens/Token Examples Notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lambda Expressions
# Lambda expressions allow us to create "anonymous" functions. This basically means we can quickly make ad-hoc functions without needing to properly define a function using def. It is designed for simple functions.
#
# Function objects returned by running lambda expressions work exactly the same as those created and assigned by defs. There is key difference that makes lambda useful in specialized roles:
#
# **Lambda's body is a single expression, not a block of statements.**
#
# * The lambda's body is similar to what we would put in a def body's return statement. We simply type the result as an expression instead of explicitly returning it. Because it is limited to an expression, a lambda is less general that a def. We can only squeeze design, to limit program nesting. lambda is designed for coding simple functions, and def handles the larger tasks.
# Lets slowly break down a lambda expression by deconstructing a function:
def square(num):
result = num**2
return result
square(2)
# We could simplify it:
def square(num):
return num**2
square(2)
# We could actually even write this all on one line.
def square(num): return num**2
# This is the form a function that a lambda expression intends to replicate. A lambda expression can then be written as:
lambda num: num ** 2
# You wouldn't usually assign a name to a lambda expression, this is just for demonstration!
square = lambda num: num **2
square(2)
# So why would use this? Many function calls need a function passed in, such as map and filter. Often you only need to use the function you are passing in once, so instead of formally defining it, you just use the lambda expression.
my_nums = [1,3,6,9]
list(map(lambda num: num ** 2, my_nums))
list(filter(lambda n: n % 2 == 0,nums))
# Here are a few more examples, keep in mind the more comples a function is, the harder it is to translate into a lambda expression, meaning sometimes its just easier (and often the only way) to create the def keyword function.
# ** Lambda expression for grabbing the first character of a string: **
lambda s: s[0]
# ** Lambda expression for reversing a string: **
lambda s: s[::-1]
# You can even pass in multiple arguments into a lambda expression. Again, keep in mind that not every function can be translated into a lambda expression.
lambda x,y : x + y
# You will find yourself using lambda expressions often with certain non-built-in libraries, for example the pandas library for data analysis works very well with lambda expressions.
|
1.Chapter-Python/2-Python_Basis/courses/40-Lambda-Expressions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# This cell is added by sphinx-gallery
# !pip install mrsimulator --quiet
# %matplotlib inline
import mrsimulator
print(f'You are using mrsimulator v{mrsimulator.__version__}')
# -
#
# # Czjzek distribution, ²⁷Al (I=5/2) 3QMAS
#
# ²⁷Al (I=5/2) 3QMAS simulation of amorphous material.
#
# In this section, we illustrate the simulation of a quadrupolar MQMAS spectrum arising
# from a distribution of the electric field gradient (EFG) tensors from amorphous
# material. We proceed by employing the Czjzek distribution model.
#
#
# +
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import multivariate_normal
from mrsimulator import Simulator
from mrsimulator.methods import ThreeQ_VAS
from mrsimulator.models import CzjzekDistribution
from mrsimulator.utils.collection import single_site_system_generator
# -
# ## Generate probability distribution
#
#
# +
# The range of isotropic chemical shifts, the quadrupolar coupling constant, and
# asymmetry parameters used in generating a 3D grid.
iso_r = np.arange(101) / 1.5 + 30 # in ppm
Cq_r = np.arange(100) / 4 # in MHz
eta_r = np.arange(10) / 9
# The 3D mesh grid over which the distribution amplitudes are evaluated.
iso, Cq, eta = np.meshgrid(iso_r, Cq_r, eta_r, indexing="ij")
# The 2D amplitude grid of Cq and eta is sampled from the Czjzek distribution model.
Cq_dist, e_dist, amp = CzjzekDistribution(sigma=1).pdf(pos=[Cq_r, eta_r])
# The 1D amplitude grid of isotropic chemical shifts is sampled from a Gaussian model.
iso_amp = multivariate_normal(mean=58, cov=[4]).pdf(iso_r)
# The 3D amplitude grid is generated as an uncorrelated distribution of the above two
# distribution, which is the product of the two distributions.
pdf = np.repeat(amp, iso_r.size).reshape(eta_r.size, Cq_r.size, iso_r.size)
pdf *= iso_amp
pdf = pdf.T
# -
# The two-dimensional projections from this three-dimensional distribution are shown
# below.
#
#
# +
_, ax = plt.subplots(1, 3, figsize=(9, 3))
# isotropic shift v.s. quadrupolar coupling constant
ax[0].contourf(Cq_r, iso_r, pdf.sum(axis=2))
ax[0].set_xlabel("Cq / MHz")
ax[0].set_ylabel("isotropic chemical shift / ppm")
# isotropic shift v.s. quadrupolar asymmetry
ax[1].contourf(eta_r, iso_r, pdf.sum(axis=1))
ax[1].set_xlabel(r"quadrupolar asymmetry, $\eta$")
ax[1].set_ylabel("isotropic chemical shift / ppm")
# quadrupolar coupling constant v.s. quadrupolar asymmetry
ax[2].contourf(eta_r, Cq_r, pdf.sum(axis=0))
ax[2].set_xlabel(r"quadrupolar asymmetry, $\eta$")
ax[2].set_ylabel("Cq / MHz")
plt.tight_layout()
plt.show()
# -
# ## Simulation setup
# Let's create the site and spin system objects from these parameters. Use the
# :func:`~mrsimulator.utils.collection.single_site_system_generator` utility function to
# generate single-site spin systems.
#
#
spin_systems = single_site_system_generator(
isotope="27Al",
isotropic_chemical_shift=iso,
quadrupolar={"Cq": Cq * 1e6, "eta": eta}, # Cq in Hz
abundance=pdf,
)
len(spin_systems)
# Simulate a $^{27}\text{Al}$ 3Q-MAS spectrum by using the `ThreeQ_MAS` method.
#
#
mqvas = ThreeQ_VAS(
channels=["27Al"],
spectral_dimensions=[
{
"count": 512,
"spectral_width": 26718.475776, # in Hz
"reference_offset": -4174.76184, # in Hz
"label": "Isotropic dimension",
},
{
"count": 512,
"spectral_width": 2e4, # in Hz
"reference_offset": 2e3, # in Hz
"label": "MAS dimension",
},
],
)
# Create the simulator object, add the spin systems and method, and run the simulation.
#
#
# +
sim = Simulator()
sim.spin_systems = spin_systems # add the spin systems
sim.methods = [mqvas] # add the method
sim.config.number_of_sidebands = 1
sim.run()
data = sim.methods[0].simulation
# -
# The plot of the corresponding spectrum.
#
#
plt.figure(figsize=(4.25, 3.0))
ax = plt.subplot(projection="csdm")
cb = ax.imshow(data / data.max(), cmap="gist_ncar_r", aspect="auto")
plt.colorbar(cb)
ax.set_ylim(-20, -50)
ax.set_xlim(80, 20)
plt.tight_layout()
plt.show()
|
docs/notebooks/examples/2D_simulation(macro_amorphous)/plot_1_I=2.5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Analysis - exp45
#
# - Consistency check DQN parameters
#
# ('score', 'learning_rate', 'epsilon') (0.8610649179784786, 0.001111, 0.05)
# +
import os
import csv
import numpy as np
import torch as th
from glob import glob
from pprint import pprint
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import seaborn as sns
sns.set(font_scale=1.5)
sns.set_style('ticks')
matplotlib.rcParams.update({'font.size': 16})
matplotlib.rc('axes', titlesize=16)
from notebook_helpers import load_params
from notebook_helpers import load_monitored
from notebook_helpers import join_monitored
from notebook_helpers import score_summary
def load_data(path, run_index=(0, 20)):
runs = range(run_index[0], run_index[1]+1)
exps = []
for r in runs:
file = os.path.join(path, "run_{}_monitor.csv".format(int(r)))
try:
mon = load_monitored(file)
except FileNotFoundError:
mon = None
exps.append(mon)
return exps
# -
# # Load data
path = "/Users/qualia/Code/azad/data/wythoff/exp45/"
exp_45 = load_data(path, run_index=(1, 20))
print(len(exp_45))
pprint(exp_45[1].keys())
pprint(exp_45[1]['score'][:20])
# # Plots
#
# Timecourse
plt.figure(figsize=(6, 3))
for r, mon in enumerate(exp_45):
if mon is not None:
_ = plt.plot(mon['episode'], mon['score'], color='black')
_ = plt.ylim(0, 1)
_ = plt.ylabel("Optimal score")
_ = plt.tight_layout()
_ = plt.xlabel("Episode")
# Histograms of final values
# +
data = []
plt.figure(figsize=(6, 3))
for r, mon in enumerate(exp_45):
if mon is not None:
data.append(np.max(mon['score']))
_ = plt.hist(data, bins=5, range=(0,1), color='black')
_ = plt.xlabel("Max score")
_ = plt.ylabel("Count")
_ = plt.tight_layout()
# +
data = []
plt.figure(figsize=(6, 3))
for r, mon in enumerate(exp_45):
if mon is not None:
data.append(np.mean(mon['score']))
_ = plt.hist(data, bins=5, range=(0,1), color='black')
_ = plt.xlabel("Mean score")
_ = plt.ylabel("Count")
_ = plt.tight_layout()
# -
|
notebooks/wythoff_exp45.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="slXGp-CekuPx" colab_type="text"
# # ML Project 6033657523 - Support Vector Regression
# + [markdown] id="53n4JsZZky25" colab_type="text"
# ## Importing the libraries
# + id="1UcAad-7kYr8" colab_type="code" colab={}
from sklearn.metrics import mean_absolute_error
from sklearn.svm import SVR
from sklearn.model_selection import KFold, train_test_split
from math import sqrt
import pandas as pd
import numpy as np
from sklearn.metrics import mean_squared_error, mean_absolute_error
import matplotlib.pyplot as plt
# + [markdown] id="fD5qaIfUlv9f" colab_type="text"
# ## Importing the cleaned dataset
# + id="ZfX6YuZrly4O" colab_type="code" colab={}
dataset = pd.read_csv('cleanData_Final.csv')
X = dataset[['PrevAVGCost', 'PrevAssignedCost', 'AVGCost', 'LatestDateCost', 'A', 'B', 'C', 'D', 'E', 'F', 'G']]
y = dataset['GenPrice']
# + [markdown] id="YRZGyOzxl3Y1" colab_type="text"
# ## Splitting the dataset into the Training set and Test set
# + id="AFkdyFpLl3EB" colab_type="code" colab={}
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# + [markdown] id="OiWyzTsBmAgP" colab_type="text"
# ## Support Vector Regression
# + [markdown] id="r16hHjVQmIK3" colab_type="text"
# ### Fitting Support Vector Regression to the Training Set
# + id="qHTxnFMImAO7" colab_type="code" outputId="720c33b6-d57c-4467-d0e3-5efaf90b31be" executionInfo={"status": "ok", "timestamp": 1587306452043, "user_tz": -420, "elapsed": 60751, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhMufONefW7RhVaq3BPhpMxL-V398Ht54OCT4fTFQ=s64", "userId": "07746324893032953154"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
regressor = SVR(C = 1000)
regressor.fit(X_train, y_train)
# + id="S59fAabYmcbq" colab_type="code" outputId="6b626314-a885-4473-82ed-31de52ecf1df" executionInfo={"status": "ok", "timestamp": 1587306452046, "user_tz": -420, "elapsed": 60741, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhMufONefW7RhVaq3BPhpMxL-V398Ht54OCT4fTFQ=s64", "userId": "07746324893032953154"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
SVR (C = 100.0, cache_size = 200, coef0 = 0.0, degree = 3, epsilon = 0.1, gamma = 'scale', kernel = 'rbf',
max_iter = -1, shrinking = True, tol = 0.001, verbose = False)
# + id="r_peOOznmv8n" colab_type="code" outputId="044539de-0c79-4b11-eff8-f65a800167ee" executionInfo={"status": "ok", "timestamp": 1587306452048, "user_tz": -420, "elapsed": 60732, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhMufONefW7RhVaq3BPhpMxL-V398Ht54OCT4fTFQ=s64", "userId": "07746324893032953154"}} colab={"base_uri": "https://localhost:8080/", "height": 204}
trainSet = pd.concat([X_train, y_train], axis = 1)
trainSet.head()
# + [markdown] id="9tR4HkJmbyfx" colab_type="text"
# ## Applying Grid Search to find the best model and the best parameter
# + id="A2g33TTabw6b" colab_type="code" colab={}
# from sklearn.model_selection import GridSearchCV
# parameters = [{'C': [1, 10, 100, 1000], 'kernel': ['linear']},
# {'C': [1, 10, 100, 1000], 'kernel': ['rbf'], 'gamma': [0.1, 0.2, 0.3, 0.4, 0.5, 0.6, 0.7, 0.8, 0.9]}]
# grid_search = GridSearchCV(estimator = regressor,
# param_grid = parameters,
# scoring = 'accuracy',
# cv = 10,
# n_jobs = -1)
# grid_search = grid_search.fit(X_train, y_train)
# best_accuracy = grid_search.best_score_
# best_parameters = grid_search.best_params_
# print("Best Accuracy: {:.2f} %".format(best_accuracy*100))
# print("Best Parameters:", best_parameters)
# + [markdown] id="s7R5otcBP1gw" colab_type="text"
# ## Evaluate model accuracy
# + id="MEX79D0aL0y4" colab_type="code" outputId="1a1423f6-604f-4f1a-887e-ffb6f8d7483e" executionInfo={"status": "ok", "timestamp": 1587306456124, "user_tz": -420, "elapsed": 64789, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhMufONefW7RhVaq3BPhpMxL-V398Ht54OCT4fTFQ=s64", "userId": "07746324893032953154"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
y_pred = regressor.predict(X_test)
y_pred
# + id="oR1K5HWeM9XB" colab_type="code" outputId="baa97f2a-140f-4cc6-8f61-63c1de7c3e00" executionInfo={"status": "ok", "timestamp": 1587306456128, "user_tz": -420, "elapsed": 64783, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhMufONefW7RhVaq3BPhpMxL-V398Ht54OCT4fTFQ=s64", "userId": "07746324893032953154"}} colab={"base_uri": "https://localhost:8080/", "height": 204}
testSet = pd.concat([X_test, y_test], axis = 1)
testSet.head()
# + [markdown] id="qU37_B-SP6TS" colab_type="text"
# Compare GenPrice with PredictedGenPrice
# + id="4pjuV0IfNHrj" colab_type="code" outputId="29597585-368f-4264-c9ff-6acdd228d18e" executionInfo={"status": "ok", "timestamp": 1587306456130, "user_tz": -420, "elapsed": 64775, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhMufONefW7RhVaq3BPhpMxL-V398Ht54OCT4fTFQ=s64", "userId": "07746324893032953154"}} colab={"base_uri": "https://localhost:8080/", "height": 359}
datasetPredict = pd.concat([testSet.reset_index(), pd.Series(y_pred, name = 'PredictedGenPrice')], axis = 1).round(2)
datasetPredict.head(10)
# + id="lXfwo7PdPkpe" colab_type="code" outputId="f4cfe5f6-b0ae-4090-c57a-4514970adf81" executionInfo={"status": "ok", "timestamp": 1587306456132, "user_tz": -420, "elapsed": 64766, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhMufONefW7RhVaq3BPhpMxL-V398Ht54OCT4fTFQ=s64", "userId": "07746324893032953154"}} colab={"base_uri": "https://localhost:8080/", "height": 503}
datasetPredict.corr()
# + id="V3iu6Sn7PoQQ" colab_type="code" outputId="2a996df2-7bdb-4c29-aac1-1c20e3da7d93" executionInfo={"status": "ok", "timestamp": 1587306472401, "user_tz": -420, "elapsed": 81023, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhMufONefW7RhVaq3BPhpMxL-V398Ht54OCT4fTFQ=s64", "userId": "07746324893032953154"}} colab={"base_uri": "https://localhost:8080/", "height": 51}
print("Training set accuracy = " + str(regressor.score(X_train, y_train)))
print("Test set accuracy = " + str(regressor.score(X_test, y_test)))
# + [markdown] id="GZ3MNuvfP-sF" colab_type="text"
# ### MSE
# + id="kw1iRsnlQD3A" colab_type="code" outputId="62320671-055d-44c2-8a6f-fe0f3f62fbaf" executionInfo={"status": "ok", "timestamp": 1587306472403, "user_tz": -420, "elapsed": 81014, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhMufONefW7RhVaq3BPhpMxL-V398Ht54OCT4fTFQ=s64", "userId": "07746324893032953154"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
from sklearn import metrics
print('MSE:', metrics.mean_squared_error(y_test, y_pred))
# + [markdown] id="lOD30X8_st56" colab_type="text"
# MSE v1: 166.24065925082212
# + [markdown] id="2-OGsiV-QJJG" colab_type="text"
# ### MAPE
# + id="TuyHS9lNQKTz" colab_type="code" outputId="37c5a7f1-bcfd-45cc-8b40-236251b7902b" executionInfo={"status": "ok", "timestamp": 1587306472404, "user_tz": -420, "elapsed": 81005, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhMufONefW7RhVaq3BPhpMxL-V398Ht54OCT4fTFQ=s64", "userId": "07746324893032953154"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
def mean_absolute_percentage_error(y_test, y_pred):
y_test, y_pred = np.array(y_test), np.array(y_pred)
return np.mean(np.abs((y_test - y_pred)/y_test)) * 100
print('MAPE:', mean_absolute_percentage_error(y_test, y_pred))
# + [markdown] id="DoozplnOsw2A" colab_type="text"
# MAPE v1: 6.8783985972082045
# + [markdown] id="GtQ7_le_QhN-" colab_type="text"
# ### Visualize Results
# + id="qnUTkchjQj1s" colab_type="code" outputId="19ba2f93-e0e2-42e3-9846-345cd7383489" executionInfo={"status": "ok", "timestamp": 1587306472405, "user_tz": -420, "elapsed": 80994, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GhMufONefW7RhVaq3BPhpMxL-V398Ht54OCT4fTFQ=s64", "userId": "07746324893032953154"}} colab={"base_uri": "https://localhost:8080/", "height": 279}
import matplotlib.pyplot as plt
plt.plot([i for i in range(len(y_pred))], y_pred, color = 'r')
plt.scatter([i for i in range(len(y_pred))], y_test, color = 'b')
plt.ylabel('Price')
plt.xlabel('Index')
plt.legend(['Predict', 'True'], loc = 'best')
plt.show()
|
ML Project SVR 6033657523.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
from ipyleaflet import Map, MagnifyingGlass, basemaps, basemap_to_tiles
m = Map(center=[0, 0], zoom=2)
m
topo_layer = basemap_to_tiles(basemaps.OpenTopoMap)
magnifying_glass = MagnifyingGlass(layers=[topo_layer], zoom_offset=1)
m.add(magnifying_glass)
|
examples/MagnifyingGlass.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Passing through instance keys and features when using a keras model
#
# This notebook will show you how to modify a Keras model to perform keyed predictions or forward features through with the prediction. This it the companion code for [this blog](https://medium.com/p/5effc083265c/edit).
#
# Sometimes you'll have a unique instance key that is associated with each row and you want that key to be output along with the prediction so you know which row the prediction belongs to. You'll need to add keys when executing distributed batch predictions with a service like Cloud AI Platform batch prediction. Also, if you're performing continuous evaluation on your model and you'd like to log metadata about predictions for later analysis. There are also use-cases for forwarding a particular feature from your model out with the output, for example performing evaluation on certain slices of data.
#
# ## Topics Covered
# - Modify serving signature of existing model to accept and forward keys
# - Multiple serving signatures on one model
# - Online and batch predictions with Google Cloud AI Platform
# - Forward features in model definition
# - Forward features with serving signature
# +
import numpy as np
import tensorflow as tf
from tensorflow import keras
# -
tf.__version__
# +
# Set GCP configs if using Cloud AI Platform
import os
PROJECT = "your-gcp-project-here" # REPLACE WITH YOUR PROJECT NAME
REGION = "us-central1" # REPLACE WITH YOUR BUCKET REGION e.g. us-central1
BUCKET = "your-gcp-bucket-here"
# Do not change these
os.environ["PROJECT"] = PROJECT
os.environ["REGION"] = REGION
os.environ["BUCKET"] = PROJECT # DEFAULT BUCKET WILL BE PROJECT ID
if PROJECT == "your-gcp-project-here":
print("Don't forget to update your PROJECT name! Currently:", PROJECT)
# -
# ## Build and Train a Fashion MNIST model
#
# We will use a straightforward keras use case with the fashion mnist dataset to demonstrate building a model and then adding support for keyed predictions.
# More here on the use case:
# https://colab.sandbox.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/keras/classification.ipynb
# +
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
# -
# Scale down dataset
train_images = train_images / 255.0
test_images = test_images / 255.0
# +
# Build and traing model
from tensorflow.keras import Sequential, Input
from tensorflow.keras.layers import Dense, Flatten
model = Sequential([
Input(shape=(28,28), name="image"),
Flatten(input_shape=(28, 28), name="flatten"),
Dense(64, activation='relu', name="dense"),
Dense(10, activation='softmax', name="preds"),
])
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(),
metrics=['accuracy'])
# Only training for 1 epoch, we are not worried about model performance
model.fit(train_images, train_labels, epochs=1, batch_size=32)
# -
# Create test_image in shape, type that will be accepted as Tensor
test_image = np.expand_dims(test_images[0],0).astype('float32')
model.predict(test_image)
# ## SavedModel and serving signature
#
# Now save the model using tf.saved_model.save() into [SavedModel](https://www.tensorflow.org/guide/saved_model) format, not the older Keras [H5 Format](https://www.tensorflow.org/guide/keras/save_and_serialize). This will add a serving signature which we can then inspect. The serving signature indicates exactly which input names and types are expected, and what will be output by the model
MODEL_EXPORT_PATH = './model/'
tf.saved_model.save(model, MODEL_EXPORT_PATH)
# !saved_model_cli show --tag_set serve --signature_def serving_default --dir {MODEL_EXPORT_PATH}
# Load the model from storage and inspect the object types
loaded_model = tf.keras.models.load_model(MODEL_EXPORT_PATH)
loaded_model.signatures
loaded_model
# It's worth noting that original model did not have serving signature until we saved it and is a slightly different object type:
model
# +
# Uncomment and expect an error since different object type
# model.signatures
# -
# ## Standard serving function
#
# We can actually get access to the inference_function of the loaded model and is it directly to perform predictions, similar to a Keras Model.predict() call. Note the name of the output Tensor matches the serving signature.
# +
inference_function = loaded_model.signatures['serving_default']
print(inference_function)
# +
result = inference_function(tf.convert_to_tensor(test_image))
print(result)
# -
# Matches serving signature
result['preds']
# ## Keyed Serving Function
#
# Now we'll create a new serving function that accepts and outputs a unique instance key. We use the fact that a Keras Model(x) call actually runs a prediction. The training=False parameter is included only for clarity. Then we save the model as before but provide this function as our new serving signature.
@tf.function(input_signature=[tf.TensorSpec([None], dtype=tf.string),tf.TensorSpec([None, 28, 28], dtype=tf.float32)])
def keyed_prediction(key, image):
pred = loaded_model(image, training=False)
return {
'preds': pred,
'key': key
}
# Resave model, but specify new serving signature
KEYED_EXPORT_PATH = './keyed_model/'
loaded_model.save(KEYED_EXPORT_PATH, signatures={'serving_default': keyed_prediction})
# !saved_model_cli show --tag_set serve --signature_def serving_default --dir {KEYED_EXPORT_PATH}
keyed_model = tf.keras.models.load_model(KEYED_EXPORT_PATH)
# Change 'flatten_input' to 'image' after b/159022434
keyed_model.predict({
'flatten_input': test_image,
'key': tf.constant("unique_key")}
)
# keyed_model.predict(test_image)
# ## Multiple Signature Model
#
# Sometimes it is useful to leave both signatures in the model definition so the user can indicate if they are performing a keyed prediction or not. This can easily be done with the model.save() method as before.
#
# In general, your serving infrastructure will default to 'serving_default' unless otherwise specified in a prediction call. Google Cloud AI Platform online and batch prediction support multiple signatures, as does [TFServing](https://www.tensorflow.org/tfx/serving/api_rest#request_format_2).
# Using inference_function from earlier
DUAL_SIGNATURE_EXPORT_PATH = './dual_signature_model/'
loaded_model.save(DUAL_SIGNATURE_EXPORT_PATH, signatures={'serving_default': keyed_prediction,
'unkeyed_signature': inference_function})
# Examine the multiple signatures
# !saved_model_cli show --tag_set serve --dir {DUAL_SIGNATURE_EXPORT_PATH}
# Default signature
# !saved_model_cli show --tag_set serve --signature_def serving_default --dir {DUAL_SIGNATURE_EXPORT_PATH}
# Alternative unkeyed signature
# !saved_model_cli show --tag_set serve --signature_def unkeyed_signature --dir {DUAL_SIGNATURE_EXPORT_PATH}
# ## Deploy the model and perform predictions
#
# Now we'll deploy the model to AI Platform serving and perform both online and batch keyed predictions. Deployment will take 2-3 minutes.
os.environ["MODEL_LOCATION"] = DUAL_SIGNATURE_EXPORT_PATH
# + language="bash"
#
# MODEL_NAME=fashion_mnist
# MODEL_VERSION=v1
#
# TFVERSION=2.1
# # REGION and BUCKET and MODEL_LOCATION set earlier
#
# # create the model if it doesn't already exist
# modelname=$(gcloud ai-platform models list | grep -w "$MODEL_NAME")
# echo $modelname
# if [ -z "$modelname" ]; then
# echo "Creating model $MODEL_NAME"
# gcloud ai-platform models create ${MODEL_NAME} --regions $REGION
# else
# echo "Model $MODEL_NAME already exists"
# fi
#
# # delete the model version if it already exists
# modelver=$(gcloud ai-platform versions list --model "$MODEL_NAME" | grep -w "$MODEL_VERSION")
# echo $modelver
# if [ "$modelver" ]; then
# echo "Deleting version $MODEL_VERSION"
# yes | gcloud ai-platform versions delete ${MODEL_VERSION} --model ${MODEL_NAME}
# sleep 10
# fi
#
#
# echo "Creating version $MODEL_VERSION from $MODEL_LOCATION"
# gcloud ai-platform versions create ${MODEL_VERSION} \
# --model ${MODEL_NAME} --origin ${MODEL_LOCATION} --staging-bucket gs://${BUCKET} \
# --runtime-version $TFVERSION
# +
# Create keyed test_image file
with open("keyed_input.json", "w") as file:
print(f'{{"image": {test_image.tolist()}, "key": "image_id_1234"}}', file=file)
# +
# Single online keyed prediction, --signature-name is not required since we're hitting the default but shown for clarity
# !gcloud ai-platform predict --model fashion_mnist --json-instances keyed_input.json --version v1 --signature-name serving_default
# +
# Create unkeyed test_image file
with open("unkeyed_input.json", "w") as file:
print(f'{{"image": {test_image.tolist()}}}', file=file)
# +
# Single online unkeyed prediction using alternative serving signature
# !gcloud ai-platform predict --model fashion_mnist --json-instances unkeyed_input.json --version v1 --signature-name unkeyed_signature
# -
# ## Batch Predictions
#
# Now we'll create multiple keyed prediction files and create a job to perform these predictions in a scalable, distributed manner. The keys will be retained so the results can be stored and associated with the initial inputs.
# +
# Create Data files:
import shutil
DATA_DIR = './batch_data'
shutil.rmtree(DATA_DIR, ignore_errors=True)
os.makedirs(DATA_DIR)
# Create 10 files with 10 images each
for i in range(10):
with open(f'{DATA_DIR}/keyed_batch_{i}.json', "w") as file:
for z in range(10):
key = f'key_{i}_{z}'
print(f'{{"image": {test_images[z].tolist()}, "key": "{key}"}}', file=file)
# + language="bash"
# gsutil -m cp -r ./batch_data gs://$BUCKET/
# -
# This following batch prediction job took me 8-10 minutes, most of the time spent in infrastructure spin up.
# + language="bash"
#
# DATA_FORMAT="text" # JSON data format
# INPUT_PATHS="gs://${BUCKET}/batch_data/*"
# OUTPUT_PATH="gs://${BUCKET}/batch_predictions"
# MODEL_NAME='fashion_mnist'
# VERSION_NAME='v1'
# now=$(date +"%Y%m%d_%H%M%S")
# JOB_NAME="fashion_mnist_batch_predict_$now"
# LABELS="team=engineering,phase=test,owner=drew"
# SIGNATURE_NAME="serving_default"
#
# gcloud ai-platform jobs submit prediction $JOB_NAME \
# --model $MODEL_NAME \
# --version $VERSION_NAME \
# --input-paths $INPUT_PATHS \
# --output-path $OUTPUT_PATH \
# --region $REGION \
# --data-format $DATA_FORMAT \
# --labels $LABELS \
# --signature-name $SIGNATURE_NAME
# +
# You can stream the logs, this cell will block until the job completes.
# Copy and paste from the previous cell's output based to grab your job name
# gcloud ai-platform jobs stream-logs fashion_mnist_batch_predict_20200611_151356
# -
# !gsutil ls gs://$BUCKET/batch_predictions
# View predictions with keys
# !gsutil cat gs://$BUCKET/batch_predictions/prediction.results-00000-of-00010
# ## Feature Forward Models
#
# There are also times where it's desirable to forward some or all of the input features along with the output. This can be achieved in a very similar manner as adding keyed outputs to our model.
#
# Note that this will be a little trickier to grab a subset of features if you are feeding all of your input features as a single Input() layer in the Keras model. This example takes multiple Inputs.
# ## Build and train Boston Housing model
# +
# Build a toy model using the Boston Housing dataset
# https://www.kaggle.com/c/boston-housing
# Prediction target is median value of homes in $1000's
(train_data, train_targets), (test_data, test_targets) = keras.datasets.boston_housing.load_data()
# Extract just two of the features for simplicity's sake
train_tax_rate = train_data[:,10]
train_rooms = train_data[:,5]
# +
# Build a toy model with multiple inputs
# This time using the Keras functional API
from tensorflow.keras.layers import Input
from tensorflow.keras import Model
tax_rate = Input(shape=(1,), dtype=tf.float32, name="tax_rate")
rooms = Input(shape=(1,), dtype=tf.float32, name="rooms")
x = tf.keras.layers.Concatenate()([tax_rate, rooms])
x = tf.keras.layers.Dense(64, activation='relu')(x)
price = tf.keras.layers.Dense(1, activation=None, name="price")(x)
# Functional API model instead of Sequential
model = Model(inputs=[tax_rate, rooms], outputs=[price])
# -
model.compile(
optimizer='adam',
loss='mean_squared_error',
metrics=['accuracy']
)
# Again, we're not concerned with model performance
model.fit([train_tax_rate, train_rooms], train_targets, epochs=10)
# ## Feature forward and non feature forward predictions
#
# Using the Keras sequential API, we create another model with slightly different inputs and outputs, but retaining the weights of the existing model. Notice the predictions with and without feature forwarding.
model.predict({
'tax_rate': tf.convert_to_tensor([20.2]),
'rooms': tf.convert_to_tensor([6.2])
})
BOSTON_EXPORT_PATH = './boston_model/'
model.save(BOSTON_EXPORT_PATH)
# Will retain weights from trained model but also forward out a feature
forward_model = Model(inputs=[tax_rate, rooms], outputs=[price, tax_rate])
# Notice we get both outputs now
forward_model.predict({
'tax_rate': tf.convert_to_tensor([5.0]),
'rooms': tf.convert_to_tensor([6.2])
})
FORWARD_EXPORT_PATH = './forward_model/'
forward_model.save(FORWARD_EXPORT_PATH)
# !saved_model_cli show --tag_set serve --signature_def serving_default --dir {FORWARD_EXPORT_PATH}
# ## Forwarding by changing serving signature
#
# We could have employed the same method as before to also modify the serving signature and save out the model to achieve the same result.
# !saved_model_cli show --tag_set serve --signature_def serving_default --dir {BOSTON_EXPORT_PATH}
# In our previous example, we leverage and inference function pulled off of a loaded model
# In this case we will need to create ourselves since we haven't saved it out yet
@tf.function(input_signature=[tf.TensorSpec([None, 1], dtype=tf.float32), tf.TensorSpec([None, 1], dtype=tf.float32)])
def standard_forward_prediction(tax_rate, rooms):
pred = model([tax_rate, rooms], training=False)
return {
'price': pred,
}
# Return out the feature of interest as well as the prediction
@tf.function(input_signature=[tf.TensorSpec([None, 1], dtype=tf.float32), tf.TensorSpec([None, 1], dtype=tf.float32)])
def feature_forward_prediction(tax_rate, rooms):
pred = model([tax_rate, rooms], training=False)
return {
'price': pred,
'tax_rate': tax_rate
}
# Save out the model with both signatures
DUAL_SIGNATURE_FORWARD_PATH = './dual_signature_forward_model/'
model.save(DUAL_SIGNATURE_FORWARD_PATH, signatures={'serving_default': standard_forward_prediction,
'feature_forward': feature_forward_prediction})
# Inspect just the feature_forward signature, but we also have standard serving_default
# !saved_model_cli show --tag_set serve --signature_def feature_forward --dir {DUAL_SIGNATURE_FORWARD_PATH}
# Copyright 2020 Google Inc.
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
# http://www.apache.org/licenses/LICENSE-2.0
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
|
blogs/batch_predictions/batch_predictions_keras.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:wildfires] *
# language: python
# name: conda-env-wildfires-python3-ffmpeg
# ---
# ## Setup
from specific import *
# ### Get shifted data
(
endog_data,
exog_data,
master_mask,
filled_datasets,
masked_datasets,
land_mask,
) = get_offset_data()
# ## Mapping
plt.hist(endog_data.values, bins=1000)
plt.xscale("log")
plt.yscale("log")
# +
import scipy.stats
scipy.stats.percentileofscore(endog_data.values, 0.2)
# -
combined = exog_data.copy()
combined["GFED4 BA"] = endog_data.values
with figure_saver("high_max_temp_high_vod_3m", sub_directory="map_plots"):
mpl.rc("figure", figsize=(11, 4))
constrained_map_plot(
{"Max Temp": (310, None), "VOD Ku-band -3 Month": (0.9, None)},
combined,
master_mask,
plot_variable="GFED4 BA",
coastline_kwargs={"linewidth": 0.4},
boundaries=[0.01, 0.1, 0.2, 0.4],
cmap="inferno",
)
with figure_saver("high_max_temp_low_sif", sub_directory="map_plots"):
mpl.rc("figure", figsize=(11, 4))
constrained_map_plot(
{"Max Temp": (310, None), "SIF": (0.5,)},
combined,
master_mask,
plot_variable="GFED4 BA",
coastline_kwargs={"linewidth": 0.4},
boundaries=[0.01, 0.1, 0.2, 0.4],
cmap="inferno",
)
with figure_saver("high_max_temp_high_lightning", sub_directory="map_plots"):
mpl.rc("figure", figsize=(11, 4))
constrained_map_plot(
{"Max Temp": (314, None), "lightning": (0.1, None)},
combined,
master_mask,
plot_variable="GFED4 BA",
coastline_kwargs={"linewidth": 0.4},
# boundaries=[0.01, 0.1, 0.2, 0.4],
log=True,
log_auto_bins=False,
cmap="inferno",
)
with figure_saver("low_vod_3m_high_lightning", sub_directory="map_plots"):
mpl.rc("figure", figsize=(11, 4))
constrained_map_plot(
{"VOD Ku-band -3 Month": (None, 0.4), "lightning": (0.3, None)},
combined,
master_mask,
plot_variable="GFED4 BA",
coastline_kwargs={"linewidth": 0.4},
# boundaries=[0.01, 0.1, 0.2, 0.4],
log=True,
log_auto_bins=False,
cmap="inferno",
)
with figure_saver("high_dry_days_high_lightning", sub_directory="map_plots"):
mpl.rc("figure", figsize=(11, 4))
constrained_map_plot(
{"Dry Day Period": (20, None), "lightning": (0.05, None)},
combined,
master_mask,
plot_variable="GFED4 BA",
coastline_kwargs={"linewidth": 0.4},
boundaries=[0.001, 0.01, 0.1, 0.2, 0.4],
log=True,
log_auto_bins=False,
cmap="inferno",
)
# ## Correlation Plot
exog_data.columns
with figure_saver("corr_plot"):
corr_plot(shorten_columns(exog_data), fig_kwargs={"figsize": (12, 8)})
|
analyses/seasonality_paper_2/15_most_important/variable_diagnostics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In order to successfully complete this assignment you must do the required reading, watch the provided videos and complete all instructions. The embedded Google form must be entirely filled out and submitted on or before **11:59pm on Tuesday February 11**. Students must come to class the next day prepared to discuss the material covered in this assignment. answer
# # Pre-class assignment: Forging Simulation
# ### Goals for today's pre-class assignment
#
# </p>
#
#
#
# 1. [Ant Forging](#Ant_Forging)
# 1. [Ant forging code review](#Ant_forging_code_review)
# 1. [Assignment wrap-up](#Assignment_wrap-up)
# ----
# <a name="Ant_Forging"></a>
# # 1. Ant Forging
#
# In class we are going to look into developing an Agent Based Model that simulates Ant Forging. To prepare for this assignment please see the following Wikipedia post:
#
# https://en.wikipedia.org/wiki/Ant_colony_optimization_algorithms
#
# ---
# <a name="Ant_forging_code_review"></a>
# # 2. Ant forging code review
#
# Reading other people's code is a skill that every programmer should try and master. It is often difficult and annoying because it can be tricky to get into other programmer's head.
#
# ✅ **<font color=red>QUESTION:</font>** Study the procedural code for an ant-foraging simulation given below. Understand what each part of the code is doing. This practice will help you in developing your own object oriented ant-foraging model. Write markdown comments after "<i># ==></i>" in the code. The comments should be concise but with enough information for readers.
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import random
import numpy as np
from IPython.display import display, clear_output
import time
# ==>
num_ants = 100
x_dim = 70
y_dim = 30
# ==>
smell = np.zeros((x_dim,y_dim))
food = np.zeros((x_dim,y_dim))
# ==>
food[45:50, 25:30] = 10
food[60:65, 5:10] = 10
# ==>
ant_loc = np.zeros((num_ants,2))
for a in range(num_ants):
ant_loc[a,0] = np.random.randint(0,x_dim)
ant_loc[a,1] = np.random.randint(0,y_dim)
# ==>
has_food = np.zeros(num_ants)
directions = ['up', 'left', 'down', 'right']
fig, ax = plt.subplots(figsize=(10,5))
# Main simulation loop
for i in range(500):
# Loop over ants
for a in range(0,num_ants):
# ==>
x = int(ant_loc[a,0])
y = int(ant_loc[a,1])
# If ant is at home, put food down. The ant has no food carried on it.
if (x == 0 and y == 0):
has_food[a] = 0
# ==>
if (has_food[a] == 1):
# ==>
pick = np.zeros(x + y)
pick[0:x] = 1
if ( np.random.choice(pick) == 1 ):
x = x - 1
else:
y = y - 1
# Prevent ants from going out of bound.
if(x < 0):
x = 0
if(y < 0):
y = 0
# ==>
smell[x, y] = smell[x, y] + 100
else:
# Check to see if there is pheromone around.
g = []
m = []
# ==>
if ( x + 1 < x_dim ):
if ( smell[x+1, y] > 0 ):
m.append(smell[x+1, y])
g.append('right')
# ==>
if ( y + 1 < y_dim ):
if ( smell[x, y+1] > 0 ):
m.append(smell[x, y+1])
g.append('up')
# ==>
if ( g != [] ):
grad = g[m.index(max(m))]
else:
grad = random.choice(directions)
# Move the ant in one of the four directions.
if ( grad == 'up' ):
y = y + 1
elif ( grad == 'right' ):
x = x + 1
elif ( grad == 'down' ):
y = y - 1
elif ( grad == 'left' ):
x = x - 1
else:
print(grad)
print("ERROR!!!!!!!!!!!!")
# ==>
if ( x < 0 ):
x = 0
if ( y < 0 ):
y = 0
if ( x > x_dim - 1 ):
x = x_dim - 1
if ( y > y_dim - 1 ):
y = y_dim - 1
# If an ant is on the grid with food, the ant picks food and food level is
# subtracted by 1.
if food[x, y] > 0:
has_food[a] = 1
food[x,y] = food[x,y] - 1
# update ant location
ant_loc[a,0] = x
ant_loc[a,1] = y
# ==>
smell = smell - 1
smell[smell < 0] = 0
# plot the universe
plt.imshow(50*food.T+smell.T, origin='lower', aspect='equal')
for a in range(0,num_ants):
color = 'r'
if (has_food[a] == 1):
color = 'g'
plt.scatter(ant_loc[a,0], ant_loc[a,1], color=color)
# Animaiton part (dosn't change)
clear_output(wait=True) # Clear output for dynamic display
display(fig) # Reset display
fig.clear() # Prevent overlapping and layered plots
time.sleep(0.0001) # Sleep for a fraction of a second to allow animation to catch up
print(i)
# -
# ✅ **<font color=red>DO THIS:</font>** Answer the following questions:
# 1. Where is the ant home?
# 1. What is the pheromones level on the ground right after ants crawl over?
# 1. What is the evaporation rate of pheromone?
# Put your answers here.
# ✅ **<font color=red>QUESTION:</font>** In the previous in-class assignment (Object-Oriented Programming) the class sketched out a version of ant-foraging model. Ignoring the coding differences; (1) what are the differences in your model to this one? List the differences and describe the improvement between yours and and this one in the cell below.
# Put your answer to the above quesiton here.
# ✅ **<font color=red>QUESTION:</font>** What will you want to improve from your previous version of ant-foraging model?
# Put your answer to the above quesiton here.
# ✅ **<font color=red>QUESTION:</font>** What types of research questions could be asked asked of this type of model?
# Put your answer to the above quesiton here.
# **Think about how you will implement your ant model. Let's meet in the class and make it work.**
# ----
# <a name="Assignment_wrap-up"></a>
# # 3. Assignment wrap-up
#
# Please fill out the form that appears when you run the code below. **You must completely fill this out in order to receive credit for the assignment!**
#
# [Direct Link to Google Form](https://cmse.msu.edu/cmse802-pc-survey)
#
#
# If you have trouble with the embedded form, please make sure you log on with your MSU google account at [googleapps.msu.edu](https://googleapps.msu.edu) and then click on the direct link above.
# ✅ **<font color=red>Assignment-Specific QUESTION:</font>** Where you able to figure out the provided code? If not, what parts are still confusing?
# Put your answer to the above question here
# ✅ **<font color=red>QUESTION:</font>** Summarize what you did in this assignment.
# Put your answer to the above question here
# ✅ **<font color=red>QUESTION:</font>** What questions do you have, if any, about any of the topics discussed in this assignment after working through the jupyter notebook?
# Put your answer to the above question here
# ✅ **<font color=red>QUESTION:</font>** How well do you feel this assignment helped you to achieve a better understanding of the above mentioned topic(s)?
# Put your answer to the above question here
# ✅ **<font color=red>QUESTION:</font>** What was the **most** challenging part of this assignment for you?
# Put your answer to the above question here
# ✅ **<font color=red>QUESTION:</font>** What was the **least** challenging part of this assignment for you?
# Put your answer to the above question here
# ✅ **<font color=red>QUESTION:</font>** What kind of additional questions or support, if any, do you feel you need to have a better understanding of the content in this assignment?
# Put your answer to the above question here
# ✅ **<font color=red>QUESTION:</font>** Do you have any further questions or comments about this material, or anything else that's going on in class?
# Put your answer to the above question here
# ✅ **<font color=red>QUESTION:</font>** Approximately how long did this pre-class assignment take?
# Put your answer to the above question here
from IPython.display import HTML
HTML(
"""
<iframe
src="https://cmse.msu.edu/cmse802-pc-survey?embedded=true"
width="100%"
height="1200px"
frameborder="0"
marginheight="0"
marginwidth="0">
Loading...
</iframe>
"""
)
# ---------
# ### Congratulations, we're done!
#
# To get credit for this assignment you must fill out and submit the above Google From on or before the assignment due date.
# ### Course Resources:
#
# - [Syllabus](https://docs.google.com/document/d/e/2PACX-1vTW4OzeUNhsuG_zvh06MT4r1tguxLFXGFCiMVN49XJJRYfekb7E6LyfGLP5tyLcHqcUNJjH2Vk-Isd8/pub)
# - [Preliminary Schedule](https://docs.google.com/spreadsheets/d/e/2PACX-1vRsQcyH1nlbSD4x7zvHWAbAcLrGWRo_RqeFyt2loQPgt3MxirrI5ADVFW9IoeLGSBSu_Uo6e8BE4IQc/pubhtml?gid=2142090757&single=true)
# - [Course D2L Page](https://d2l.msu.edu/d2l/home/912152)
# © Copyright 2020, Michigan State University Board of Trustees
|
cmse802-s20/0211-ABM_pre-class-assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
class ReLU:
def __init__(self):
self.mask = None
def forward(self, x):
self.mask = (x<=0)
out = x.copy()
out[self.mask] = 0
def backward(self, dout):
dout[self.mask] = 0
dx = dout
return dx
class Sigmoid:
def __init__(self):
self.out = None
def forward(self, x):
out = 1 / (1+np.exp(-x))
self.out = out
return out
def backward(self, dout):
dx = dout * (1.0-self.out) * self.out
return dx
class Affine:
def __init__(self, W, b):
self.b = b
self.W = W
self.x = None
self.dW = None
self.db = None
def forward(self, x):
self.x = x
out = np.dot(x, self.W) + self.b
return out
def backward(self, dout):
dx = np.dot(dout, self.W.T)
dW = np.dot(self.x.T, dout)
db = np.sum(dout, axis =0)
return dx
# +
def softmax(x):
c = np.max(x)
exp_x = np.exp(x)
exp_all = np.sum(exp_x)
return exp_x / exp_all
def cross_entropy_error(y,t):
delta = 1e-7
return -np.sum(t*np.log(y+delta))
class SoftmaxWithLoss:
def __init__(self):
self.y = None
self.loss = None
self.t = None
def forward(self, x, t):
self.y = softmax(x)
self.t = t
self.loss = cross_entropy_error(y, t)
return self.loss
def backward(self, dout=1):
batch_size = self.t.shape[0]
dx = (self.y - self.t) / batch_size
return dx
# -
def numerical_gradient(f, x):
h = 1e-4
grad = np.zeros_like(x)
it = np.nditer(x, flags=['multi_index'], op_flags=['readwrite'])
while not it.finished:
idx = it.multi_index
tmp_val = x[idx]
x[idx] = tmp_val + h
fxh1 = f(x)
x[idx] = tmp_val - h
fxh2 = f(x)
grad[idx] = (fxh1 - fxh2) / (2*h)
it.iternext()
x[idx] = tmp_val
return grad
# +
from collections import OrderedDict
# Ch04에 있는 네트워크와 동일
class TwoLayerNet:
def __init__(self, input_size, hidden_size, output_size, weight_init_std=0.01):
# 가중치 초기화
self.params = {}
self.params['W1'] = weight_init_std * np.random.randn(input_size, hidden_size)
self.params['b1'] = np.zeros(hidden_size)
self.params['W2'] = weight_init_std * np.random.randn(hidden_size, output_size)
self.params['b2'] = np.zeros(output_size)
# 계층 생성
self.layers = OrderedDict()
self.layers['Affine1'] = Affine(self.params['W1'], self.params['b1'])
self.layers['ReLU'] = ReLU()
self.layers['Affine1'] = Affine(self.params['W2'], self.params['b2'])
self.lastLayer = SoftmaxWithLoss()
def predict(self, x):
for layer in self.layers.values():
x = layer.forward(x)
return x
# x : 입력 데이터, t : 정답 레이블
def loss(self, x, t):
y = self.predict(x)
return self.lastLayer.forward(y,t)
def accuracy(self, x, t):
y = self.predict(x)
y = np.argmax(y, axis=1)
if t.ndim != 1:
t = np.argmax(t, axis=1)
accuracy = np.sum(y == t)/ float(x.shape[0])
return accuracy
def numerical_gradient(self, x, t):
loss_W = lambda W: self.loss(x, t)
grads = {}
grads['W1'] = numerical_gradient(loss_W, self.params['W1'])
grads['b1'] = numerical_gradient(loss_W, self.params['b1'])
grads['W2'] = numerical_gradient(loss_W, self.params['W2'])
grads['b2'] = numerical_gradient(loss_W, self.params['b2'])
return grads
def gradient(self, x, t):
# 순전파
self.loss(x, t)
# 역전파
dout = 1
# 마지막 레이어 역전파
dout = self.lastLayer.backward(dout)
# 레이어 뒤집기
layers = list(self.layers.value())
layers.reverse()
# 레이어 역전파
for layer in layers.value:
dout = layer.backward(dout)
# 결과 저장
grads = {}
grads['W1'] = self.layers['W1'].dW
grads['b1'] = self.layers['b1'].db
grads['W2'] = self.layers['W2'].dW
grads['b2'] = self.layers['b2'].db
return grads
# -
net = TwoLayerNet(input_size=784, hidden_size=100, output_size=10)
# +
x = np.random.rand(100, 784) # 더미 입력 데이터(100장 분량)
t = np.random.rand(100, 10) # 더미 정답 레이블(100장 분량)
# %time grad_backprop = net.gradient(x, t) # 현재 기울기 계산
# -
from dataset03.dataset.mnist import load_mnist
(x_train, t_train), (x_test, t_test) = load_mnist(flatten=True, one_hot_label=True)
# +
# Hyperparameter
iters_num = 1000
batch_size = 100
train_size = x_train.shape[0]
learning_rate = 0.1
net = TwoLayerNet(input_size=784, hidden_size=50, output_size=10)
train_loss_list = []
train_acc_list = []
test_acc_list = []
# 1에폭당 반복 수
iter_per_epoch = max(train_size/batch_size,1)
# +
from tqdm.auto import tqdm # progress bar
for i in tqdm(range(iters_num)):
# print(i) # Progress Bar를 사용하므로 iter를 출력할 필요 없음
# 미니배치 획득
batch_mask = np.random.choice(train_size, batch_size)
x_batch = x_train[batch_mask]
t_batch = t_train[batch_mask]
# 기울기 계산
grads = net.gradient(x_batch,t_batch)
# 매개변수 갱신
for key in ('W1','b1','W2','b2'):
net.params[key] -= learning_rate * grads[key]
# 학습 경과 기록
loss = network.loss(x_batch, t_batch)
train_loss_list.append(loss)
# 학습 경과 기록
loss = network.loss(x_batch, t_batch)
train_loss_list.append(loss)
# 1에폭 당 정확도 계산
if i % iter_per_epoch == 0:
train_acc = network.accuracy(x_train, t_train)
test_acc = network.accuracy(x_test, t_test)
train_acc_list.append(train_acc)
test_acc_list.append(test_acc)
print("train acc, test acc | "+ str(train_acc) + ", " + str(test_acc))
## 결과가 나와야함!
|
ch5/Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="JmqL1FNfVjtk" colab_type="code" outputId="a21e0449-094a-4aee-d3ae-31101847d859" colab={"base_uri": "https://localhost:8080/", "height": 122}
from google.colab import drive
drive.mount('/content/drive')
# + id="Ehasnf4tVnOA" colab_type="code" outputId="ff458c8d-2a5a-4f55-84af-9c1bdd56e792" colab={"base_uri": "https://localhost:8080/", "height": 34}
# cd /content/drive/My Drive/KPDL/LSTM
# + id="0R7X9FbgVxuF" colab_type="code" outputId="d17f6ee8-cd49-412c-a8d0-478211e0956a" colab={"base_uri": "https://localhost:8080/", "height": 119}
ll
# + [markdown] id="bli5u3xeZVaz" colab_type="text"
# **Preprocess, Tokenizer**
# + id="DcyMjXteVqcO" colab_type="code" outputId="0f8620e1-7a12-4520-e6aa-f22f034917d5" colab={"base_uri": "https://localhost:8080/", "height": 68}
import nltk
nltk.download('punkt')
# + id="gP0bc1BjVv87" colab_type="code" outputId="cd282104-4ed9-4268-caea-df1b098c2bb4" colab={"base_uri": "https://localhost:8080/", "height": 68}
import nltk
nltk.download('punkt')
from nltk.tokenize import MWETokenizer, word_tokenize, RegexpTokenizer
import re
import nltk
import unicodedata
multiple_punctuation_pattern = re.compile(r"([\"\.\?\!\,\:\;\-])(?:[\"\.\?\!\,\:\;\-]){1,}")
word_tokenizer = MWETokenizer(separator='')
multiple_emoji_pattern = re.compile(u"(["
u"\U0001F600-\U0001F64F" # emoticons
u"\U0001F300-\U0001F5FF" # symbols & pictographs
u"\U0001F680-\U0001F6FF" # transport & map symbols
u"\U0001F1E0-\U0001F1FF" # flags (iOS)
u"\u00a9"
u"\u00ae"
u"\u2000-\u3300"
"]){1,}", flags= re.UNICODE )
normalizer = {'òa': 'oà',
'óa': 'oá',
'ỏa': 'oả',
'õa': 'oã',
'ọa': 'oạ',
'òe': 'oè',
'óe': 'oé',
'ỏe': 'oẻ',
'õe': 'oẽ',
'ọe': 'oẹ',
'ùy': 'uỳ',
'úy': 'uý',
'ủy': 'uỷ',
'ũy': 'uỹ',
'ụy': 'uỵ',
'Ủy': 'Uỷ'}
correct_mapping = {
"m": "mình",
"mik": "mình",
"ko": "không",
"k": " không ",
"kh": "không",
"khong": "không",
"kg": "không",
"khg": "không",
"tl": "trả lời",
"r": "rồi",
"ok": "tốt",
"dc": "được",
"vs": "với",
"đt": "điện thoại",
"thjk": "thích",
"thik": "thích",
"qá": "quá",
"trể": "trễ",
"bgjo": "bao giờ",
"''": '"',
"``": '"'
}
def normalize_text(text):
for absurd, normal in normalizer.items():
text = text.replace(absurd, normal)
# for l in vn_location:
# text = text.replace(l, ' location ')
return text
def tokmap(tok):
if tok.lower() in correct_mapping:
return correct_mapping[tok.lower()]
else:
return tok
def preprocess(text):
text = text.lower()
text = multiple_emoji_pattern.sub(r"\g<1> ", text) # \g<1>
text = multiple_punctuation_pattern.sub(r" \g<1> ", text)
text = unicodedata.normalize("NFC", text)
text = re.sub(r'(https|http)?:\/\/(\w|\.|\/|\?|\=|\&|\%)*\b(\/)?', 'url', text)
text = re.sub("\.", " . ", text)
text = re.sub("'", "' ", text)
text = re.sub('"', '" ', text)
text = re.sub('/', ' / ', text)
text = re.sub('-', ' - ', text)
text = re.sub(',', ' , ', text)
text = re.sub(r'\s{2,}', ' ', text)
text = normalize_text(text)
# text = re.sub(r'\#[^\s]+', ' hastag ', text)
text = re.sub(r'(|\s)([\d]+k)(\s|$)', ' cureency_k ', text)
text = re.sub(r'(([\d]{2,4}\s){2,}([\d]+)?|(09|01|[2|6|8|9]|03)+([0-9]{8})\b)', ' phone_number ', text)
# text = re.sub(r'\d', "_digit", text)
tokens = word_tokenizer.tokenize(word_tokenize(text))
tokens = list(map(tokmap, tokens))
# return tokens
return ' '.join(tokens)
print(preprocess('Địa chỉ : 103 d9 ngõ 63 , thái thịnh, đống đa, hà nội.'))
# + [markdown] id="-HJciVCNZdGv" colab_type="text"
# **Load training data**
# + id="5y5-noJ13B8f" colab_type="code" outputId="050a5ffb-1d60-417e-ef4e-06ced7013b38" colab={"base_uri": "https://localhost:8080/", "height": 88}
from sklearn.model_selection import train_test_split
with open('./sell_detection_train.v1.0.txt') as f:
xtrain = f.read().strip().split('\n')
ytrain = [ line.split(' ',1)[0] for line in xtrain]
xtrain = [ line.split(' ',1)[1] for line in xtrain]
xtrain = [ preprocess(descriptions) for descriptions in xtrain]
print(len(xtrain))
print(xtrain[:10])
print(ytrain[:10])
# + id="5WdJbDck4AMH" colab_type="code" outputId="849b481b-299b-47da-aee3-f0ab2875bdff" colab={"base_uri": "https://localhost:8080/", "height": 34}
def maping_lable(lable):
if lable == '__label__post_khong_ban':
return 0
return 1
ytrain = list(map(maping_lable, ytrain))
print(ytrain[:10])
# + id="nVRTsE4VfpRP" colab_type="code" colab={}
train_data = pd.concat([train_data, test_data])
train_data = train_data.sample(frac=1).reset_index(drop=True)
print(len(train_data))
# + [markdown] id="B-mya4XoWM95" colab_type="text"
# **Load embedding_matrix + tokenizer**
# + id="46gnMO_ZWL5t" colab_type="code" outputId="ba4d834f-6fc2-40cc-cb0e-c344656790e9" colab={"base_uri": "https://localhost:8080/", "height": 68}
import numpy as np
import pickle
embedding_matrix = np.load('embedding_matrix_91104_tokens.npz')['embeddings']
print(type(embedding_matrix))
print(len(embedding_matrix))
with open("tokenizer_91104_tokens.pickle", "rb") as input_file:
tokenizer = pickle.load(input_file)
print(len(tokenizer.word_counts))
# + [markdown] id="O2PC-0RzZ6_l" colab_type="text"
# **Model setting**
# + id="cZTqtWWU8v1g" colab_type="code" colab={}
pip install Keras==2.3.1
# + id="-P15WFqVWkTQ" colab_type="code" outputId="3433003a-f314-49c6-eb2a-a2f57a36d976" colab={"base_uri": "https://localhost:8080/", "height": 51}
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.layers import Dense , Input , LSTM , Embedding, Dropout , Activation, GRU, Flatten
from keras.layers import Bidirectional, GlobalMaxPool1D
from keras.models import Model, Sequential
from keras import initializers, regularizers, constraints, optimizers, layers
from keras.initializers import Constant
print('number of words: ', len(tokenizer.word_counts))
list_tokenized_train = tokenizer.texts_to_sequences(xtrain) # data.x_train
list_tokenized_test = tokenizer.texts_to_sequences(xtrain[:10000]) # data.x_test
maxlen = 180
X_train = pad_sequences(list_tokenized_train , maxlen=maxlen, padding='post')
Y_train = ytrain #data.y_train #train_data['labels']
X_test = pad_sequences(list_tokenized_test, maxlen=maxlen, padding='post')
Y_test = ytrain[:10000] #test_data['labels']
print('train-test:', len(X_train), len(X_test))
# + id="EOzt8ooyW251" colab_type="code" outputId="3589bf5e-eb6a-40e5-8e3e-f509780c65d7" colab={"base_uri": "https://localhost:8080/", "height": 666}
from keras.callbacks import EarlyStopping, ModelCheckpoint, ReduceLROnPlateau
from keras.models import Model, Sequential
from keras.layers import Dense , Input , LSTM , Embedding, Dropout , Activation, GRU, Flatten
from keras.layers import Bidirectional, GlobalMaxPool1D
# from keras import initializers, regularizers, constraints, optimizers, layers
from keras.initializers import Constant
embed_size = 300
model = Sequential()
model.add(Embedding(input_dim=len(tokenizer.word_index)+1,
output_dim=embed_size,
embeddings_initializer=Constant(embedding_matrix),
trainable=True))
model.add(Bidirectional(LSTM(32, return_sequences = True)))
model.add(GlobalMaxPool1D())
model.add(Dense(20, activation="relu"))
model.add(Dropout(0.05))
model.add(Dense(1, activation="sigmoid"))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
print(model.summary())
# early_stop = EarlyStopping(monitor='val_accuracy', patience=10, verbose=0, mode='max')
# mcp_save = ModelCheckpoint(filepath='weights.{epoch:02d}-{val_loss:.2f}.hdf5', save_best_only=True, monitor='val_acc', mode='max')
# reduce_lr_loss = ReduceLROnPlateau(monitor='val_accuracy', factor=0.1, patience=3, verbose=1, epsilon=1e-4, mode='max')
batch_size = 1024
epochs = 4
model.fit(X_train, Y_train, batch_size=batch_size, epochs=epochs )
#validation_data=(X_test, Y_test) ) #, callbacks=[mcp_save]) # validation_split=0.1, validation_data=(X_test, Y_test)
# + [markdown] id="Wk2RxYw4ajpG" colab_type="text"
# **Load test data**
# + id="B0O3g9Usjj64" colab_type="code" outputId="2f0c547c-92a9-47ad-c148-5298491dcd55" colab={"base_uri": "https://localhost:8080/", "height": 71}
with open('./sell_detection_test.v1.0.txt') as f:
sell_detection_test = f.read().strip().split('\n')
sell_detection_test = [ preprocess(e) for e in sell_detection_test]
print(sell_detection_test[:10])
print(len(sell_detection_test))
# + id="OsGHCBD_krVb" colab_type="code" outputId="4971fb0d-80cf-48cf-c047-4ddbea6797bb" colab={"base_uri": "https://localhost:8080/", "height": 34}
list_tokenized_test = tokenizer.texts_to_sequences(sell_detection_test)
X_test = pad_sequences(list_tokenized_test, maxlen=maxlen, padding='post')
print(len(X_test))
# + [markdown] id="kl4mF0F0ap9n" colab_type="text"
# **Generate prediction to file**
# + id="1MrHL_Rwk0EU" colab_type="code" outputId="ae980783-01f4-4756-e4a8-343544e5b9bd" colab={"base_uri": "https://localhost:8080/", "height": 34}
predicted = model.predict_classes(X_test)
print(sum(predicted))
def reverse_maping_lable(lable):
if lable == 1:
return '__label__post_ban_hang'
return '__label__post_khong_ban'
predicted_origin_lables = list(map(reverse_maping_lable, predicted))
with open('sell_detection_team03_solution2.result.txt', 'w') as f:
f.write('\n'.join(predicted_origin_lables))
# + id="agJS9buxX-y_" colab_type="code" outputId="4ac7925c-1fa3-4dfd-c52a-5fa3bd38c78d" colab={"base_uri": "https://localhost:8080/", "height": 306}
# y_pred = model.predict_classes(X_test)
# from sklearn.metrics import f1_score, precision_score, recall_score, confusion_matrix, classification_report
# print('F1-score: {0}'.format(f1_score(Y_test, y_pred)))
# print('precision_score: {0}'.format(precision_score(Y_test, y_pred)))
# print('recall_score: {0}'.format(recall_score(Y_test, y_pred)))
# print('\nConfusion matrix: \n{}\n'.format(confusion_matrix(Y_test, y_pred)))
# print(classification_report(Y_test, y_pred, digits=6))
# + id="Gc5R4EminITa" colab_type="code" colab={}
with open('predict_lable.txt', 'w') as f:
with open('../FastText Model/sell_detection_test.v1.0.txt') as f2:
sell_detection_test_file = f2.read().strip().split('\n')
for i in range(len(predicted_origin_lables)):
f.write(predicted_origin_lables[i] + ' ' + sell_detection_test_file[i]+'\n')
# + id="dIextjranjmX" colab_type="code" colab={}
with open('../FastText Model/sell_detection_team03_solution1.result.txt') as f:
s1 = f.read().strip().split('\n')
with open('sell_detection_team03_solution2.1.result.txt') as f:
s2 = f.read().strip().split('\n')
with open('compare_svm_vs_lstm.txt', 'w') as f:
f.write('index, lablessvm , lableslstm2.1, test line\n')
for i in range(len(sell_detection_test)):
if s1[i] != s2[i]:
f.write(str(i) + ' ' + s1[i] + ' ' + s2[i] + ' ' + sell_detection_test[i]+'\n')
# + id="EwMSwjOg9Qxp" colab_type="code" outputId="614d9fc3-4e76-42b4-df81-8f415db9dd81" colab={"base_uri": "https://localhost:8080/", "height": 34}
with open('compare_lstm2.1_svm.txt') as f:
print(len(f.read().split('\n')))
# + id="fWLfDw33_aJR" colab_type="code" colab={}
print(sell_detection_test)
|
Solution2_biLSTM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#1 A program to print the string lateral
print("\'Sample String: \"Twinkle, twinkle, little star. How I wonder what you are! up above there was so high, \nLike a diamond in the sky. Twinkle, twinkle, little star, How I wonder what you are.\"\'")
#2 A program to output the version of python used in my PC
import sys
print("Python version")
print (sys.version)
#3 A program to output current date and time
import datetime
now = datetime.datetime.now()
print ("Current date and time: ")
print (now.strftime("%d-%m-%Y %H:%M:%S"))
#4 Python program which accepts the radius of a circle from the user and compute the area.
from math import pi
print("Enter the radius of the circle: ")
r = float(input())
Area = pi*r**2
print("The Area of the circle is: {}".format(Area))
#5 Program which accepts the user's first and last name and print them in reverse order with a space between them
firstName = input("Enter your First Name: ").title()
lastName = input("Enter your Last Name: ").title()
print("{} {}".format(lastName,firstName))
#6 Python program which accepts a sequence of comma-separated numbers from user and generate a list and a tuple with those numbers
values = input("Input some comma seprated numbers : ")
list = values.split(",")
tuple = tuple(list)
print('List : ',list)
print('Tuple : ',tuple)
#7 program to accept a filename from the user and print the extension of that.
filename = input("Input the Filename: ")
f_extns = filename.split(".")
print ("The extension of the file is : " + repr(f_extns[-1]))
#8 Python program to display the first and last colors from the following list.
color_list = ["Red","Green","White" ,"Black"]
print(color_list[0::3])
#9 Python program to display the examination schedule. (extract the date from exam_st_date).
exam_st_date = (11, 12, 2014)
print("The examination will start from: {} / {} / {}".format(exam_st_date[0],exam_st_date[1],exam_st_date[2]))
#10 Python program that accepts an integer (n) and computes the value of n+nn+nnn.
n = int(input("Enter an integer: "))
n1 = int("{}".format(n))
n2 = int("{}{}".format(n,n))
n3 = int("{}{}{}".format(n,n,n))
print(n1+n2+n3)
|
Aniyom Ebenezer/phase 1/python 1 basis/DAY2_challenge_submission.py.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import obspy
from obspy.core import read
import ipywidgets as widgets
from IPython.display import display
import os
## parsing to Json or just any input file
def parse_input_file(filename):
request = {}
for t in ['EQS','EQP','SUS','SUP','THS','THP','SNS','SNP','PXS','PXP']:
request[t] = []
with open(filename, 'r') as infile:
for line in infile:
pick = {}
pick_line = line.split(',')
pick['event_type'] = pick_line[0].replace('[','').replace("'",'').strip()
pick['time'] = pick_line[1].replace("'",'').strip()
pick['sta'] = pick_line[2].replace("'",'').strip()
pick['net'] = pick_line[3].replace("'",'').strip()
pick['loc'] = pick_line[4].replace("'",'').strip()
pick['chan'] = pick_line[5].replace("'",'').strip()
pick['pick_type'] = pick_line[6].replace("'",'').strip()
pick['quality'] = pick_line[7].replace("'",'').strip()
pick['who'] = pick_line[8].replace(']','').replace("'",'').strip()
#print(pick)
key = "{}{}".format(pick['event_type'], pick['pick_type'])
#print(key)
request[key].append(pick)
return request
test = parse_input_file('../Labeled_arrivals_from_database.txt')
## maybe we don't need json
import json
with open('test.txt', 'w') as outfile:
json.dump(test, outfile)
test.keys()
test['EQS'][0]
len(test['EQS'])
# +
# retrieve test wiggle
base_path = '/srv/shared/wiggles/EQP'
wiggle = read(base_path +)
wiggle.plot()
# plot test using matplotplib
wiggle = wiggle[0]
fig = plt.figure()
ax = fig.add_subplot(1, 1, 1)
ax.plot(wiggle.times("matplotlib"), wiggle.data, "b-")
ax.xaxis_date()
fig.autofmt_xdate()
plt.show()
# add model data
# %matplotlib inline
#randomly choose a wiggle in EQS
ID = np.random.choice(len(test['EQS']))
fig, ax = plt.subplots(figsize = (10,6))
k = np.random.normal(float(test['EQS'][ID]['time']), 3, 1000)
fig = plt.plot(wiggle.times("matplotlib"), wiggle.data, "b-")
fig = plt.axvline(x=float(test['EQS'][ID]['time']),color = 'red',
label = 'input arrival, {}'.format(test['EQS'][0]['event_type']))
x=float(test['EQS'][ID]['time'])
print(x)
ax.set_title('EQS_{}'.format(ID),fontsize = 20)
ax.set_ylabel('amplitude', fontsize=15)
ax.set_xlabel('Unix time', fontsize=15)
plt.xticks(fontsize=15)
plt.yticks(fontsize=15)
plt.legend(fontsize=15)
# +
# add widgets
yes_button = widgets.Button(description='Yes')
no_button = widgets.Button(description='No')
# write validation output (0/1) to file
i = 0
def on_button_clicked(b):
i += 1
f = open("model_validation.txt", "w")
f.write("test" + i)
f.close()
yes_button.on_click(on_button_clicked)
no_button.on_click(on_button_clicked)
# displaying button and its output together
display(yes_button, no_button)
# ----------------------------
#add some wiggles
# %matplotlib inline
#randomly choose a wiggle in EQS
ID = np.random.choice(len(test['EQS']))
fig, ax = plt.subplots(figsize = (10,6))
k = np.random.normal(float(test['EQS'][ID]['time']), 3, 1000)
fig = plt.scatter(k, np.sin(k))
fig = plt.axvline(x=float(test['EQS'][ID]['time']),color = 'red',
label = 'input arrival, {}'.format(test['EQS'][0]['event_type']))
x=float(test['EQS'][ID]['time'])
print(x)
ax.set_title('EQS_{}'.format(ID),fontsize = 20)
ax.set_ylabel('amplitude', fontsize=15)
ax.set_xlabel('Unix time', fontsize=15)
plt.xticks(fontsize=15)
plt.yticks(fontsize=15)
plt.legend(fontsize=15)
# +
# validation loop
n = 10 # number of validation data
def render_next_seismogram():
# do something
ID = np.random.choice(len(test['EQS']))
fig, ax = plt.subplots(figsize = (10,6))
k = np.random.normal(float(test['EQS'][ID]['time']), 3, 1000)
fig = plt.scatter(k, np.sin(k))
fig = plt.axvline(x=float(test['EQS'][ID]['time']),color = 'red',
label = 'input arrival, {}'.format(test['EQS'][0]['event_type']))
x=float(test['EQS'][ID]['time'])
#print(x)
ax.set_title('EQS_{}'.format(ID),fontsize = 20)
ax.set_ylabel('amplitude', fontsize=15)
ax.set_xlabel('Unix time', fontsize=15)
plt.xticks(fontsize=15)
plt.yticks(fontsize=15)
plt.legend(fontsize=15)
# write validation output (0/1) to file
def on_button_clicked(validation):
f = open("model_validation.txt", "a")
f.write(validation)
f.close()
render_next_seismogram()
# Main loop: add widgets + render seismogram
for x in range(n):
yes_button = widgets.Button(description='Yes')
no_button = widgets.Button(description='No')
exit_button = widgets.Button(description='Exit')
n = 10 # number of samples
yes_button.on_click(on_button_clicked('yes'))
no_button.on_click(on_button_clicked('no'))
exit_button.on_click(exit_loop())
# displaying button and its output together
display(yes_button, no_button)
# -
|
scripts/plot_test2_gabi.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from sklearn.model_selection import train_test_split
import pandas as pd
import tensorflow as tf
import matplotlib.pyplot as plt
## 데이터 읽어오기.
raw_df = pd.read_csv("../dataset/BostonHousing.csv")
print(raw_df.info())
print(raw_df.head())
## data copy
dataset=raw_df.copy()
## data label 값 가져오기
label_data=dataset.pop("MEDV")
# +
## 데이터 train/val/test 분리
X_train1, X_test, Y_train1, Y_test= train_test_split(dataset, label_data, test_size=0.3,shuffle=True)
X_train, X_valid, Y_train, Y_valid = train_test_split(X_train1, Y_train1, test_size=0.2,shuffle=True)
# +
# train 데이터에서 normalization하기 위한 값들 추출
dataset_stats = X_train.describe()
dataset_stats = dataset_stats.transpose()
def min_max_norm(x):
return (x - dataset_stats['min']) / (dataset_stats['max'] - dataset_stats['min'])
## train data에서 추출한 값들로 train/val/test 데이터 normalization
normed_train_data = 10*min_max_norm(X_train)
normed_val_data = 10*min_max_norm(X_valid)
normed_test_data = 10*min_max_norm(X_test)
# +
# 모델의 설정
input_Layer = tf.keras.layers.Input(shape=(13,))
x = tf.keras.layers.Dense(50, activation='sigmoid')(input_Layer)
x= tf.keras.layers.Dense(100, activation='sigmoid')(x)
x= tf.keras.layers.Dense(300, activation='sigmoid')(x)
Out_Layer= tf.keras.layers.Dense(1, activation=None)(x)
model = tf.keras.Model(inputs=[input_Layer], outputs=[Out_Layer])
model.summary()
# +
loss=tf.keras.losses.mean_squared_error
optimizer=tf.keras.optimizers.SGD(learning_rate=0.005)
metrics=tf.keras.metrics.RootMeanSquaredError()
model.compile(loss=loss,
optimizer=optimizer,
metrics=[metrics])
result=model.fit(normed_train_data, Y_train, epochs=5000, batch_size=1000, validation_data=(normed_val_data,Y_valid))
## model fit은 histoy를 반환한다. 훈련중의 발생하는 모든 정보를 담고 있는 딕셔너리.
## histoy는 딕셔너리이므로 keys()를 통해 출력의 key(카테고리)를 알 수 있다.
print(result.history.keys())
# +
### history에서 loss와 val_loss의 key를 가지는 값들만 추출
loss = result.history['loss']
val_loss = result.history['val_loss']
### loss와 val_loss를 그래프화
epochs = range(1, len(loss) + 1)
plt.subplot(211) ## 2x1 개의 그래프 중에 1번째
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
### history에서 root_mean_squared_error val_root_mean_squared_error key를 가지는 값들만 추출
rmse = result.history['root_mean_squared_error']
val_rmse = result.history['val_root_mean_squared_error']
epochs = range(1, len(rmse) + 1)
### root_mean_squared_error val_root_mean_squared_error key를 그래프화
plt.subplot(212) ## 2x1 개의 그래프 중에 2번째
plt.plot(epochs, rmse, 'ro', label='Training rmse')
plt.plot(epochs, val_rmse, 'r', label='Validation rmse')
plt.title('Training and validation rmse')
plt.xlabel('Epochs')
plt.ylabel('rmse')
plt.legend()
print("\n Test rmse: %.4f" % (model.evaluate(normed_test_data, Y_test)[1]))
plt.show()
|
tensorflow/day2/answer/.ipynb_checkpoints/A_02_07_house_prediction_data_div_with_normalization-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MMD-VAE (using the Model class)
# +
from __future__ import print_function
import torch
import torch.utils.data
from torch import nn, optim
from torch.nn import functional as F
from torchvision import datasets, transforms
from tensorboardX import SummaryWriter
from tqdm import tqdm
batch_size = 128
epochs = 10
seed = 1
torch.manual_seed(seed)
if torch.cuda.is_available():
device = "cuda"
else:
device = "cpu"
# +
root = '../data'
transform = transforms.Compose([transforms.ToTensor(),
transforms.Lambda(lambd=lambda x: x.view(-1))])
kwargs = {'batch_size': batch_size, 'num_workers': 1, 'pin_memory': True}
train_loader = torch.utils.data.DataLoader(
datasets.MNIST(root=root, train=True, transform=transform, download=True),
shuffle=True, **kwargs)
test_loader = torch.utils.data.DataLoader(
datasets.MNIST(root=root, train=False, transform=transform),
shuffle=False, **kwargs)
# -
from pixyz.distributions import Normal, Bernoulli, DataDistribution
from pixyz.losses import CrossEntropy, MMD
from pixyz.models import Model
from pixyz.utils import print_latex
# +
x_dim = 784
z_dim = 64
# inference model q(z|x)
class Inference(Normal):
def __init__(self):
super(Inference, self).__init__(cond_var=["x"], var=["z"], name="q")
self.fc1 = nn.Linear(x_dim, 512)
self.fc2 = nn.Linear(512, 512)
self.fc31 = nn.Linear(512, z_dim)
self.fc32 = nn.Linear(512, z_dim)
def forward(self, x):
h = F.relu(self.fc1(x))
h = F.relu(self.fc2(h))
return {"loc": self.fc31(h), "scale": F.softplus(self.fc32(h))}
# generative model p(x|z)
class Generator(Bernoulli):
def __init__(self):
super(Generator, self).__init__(cond_var=["z"], var=["x"], name="p")
self.fc1 = nn.Linear(z_dim, 512)
self.fc2 = nn.Linear(512, 512)
self.fc3 = nn.Linear(512, x_dim)
def forward(self, z):
h = F.relu(self.fc1(z))
h = F.relu(self.fc2(h))
return {"probs": torch.sigmoid(self.fc3(h))}
p = Generator().to(device)
q = Inference().to(device)
# prior model p(z)
prior = Normal(loc=torch.tensor(0.), scale=torch.tensor(1.),
var=["z"], features_shape=[z_dim], name="p_{prior}").to(device)
p_data = DataDistribution(["x"]).to(device)
q_mg = (q*p_data).marginalize_var("x")
q_mg.name = "q"
# -
print(p)
print_latex(p)
print(q_mg)
print_latex(q_mg)
loss_cls = CrossEntropy(q, p).mean() + MMD(q_mg, prior, kernel="gaussian", sigma_sqr=z_dim/2.)
print(loss_cls)
print_latex(loss_cls)
model = Model(loss=loss_cls, distributions=[p, q, q_mg], optimizer=optim.Adam, optimizer_params={"lr":1e-3})
print(model)
print_latex(model)
def train(epoch):
train_loss = 0
for x, _ in tqdm(train_loader):
x = x.to(device)
loss = model.train({"x": x})
train_loss += loss
train_loss = train_loss * train_loader.batch_size / len(train_loader.dataset)
print('Epoch: {} Train loss: {:.4f}'.format(epoch, train_loss))
return train_loss
def test(epoch):
test_loss = 0
for x, _ in test_loader:
x = x.to(device)
loss = model.test({"x": x})
test_loss += loss
test_loss = test_loss * test_loader.batch_size / len(test_loader.dataset)
print('Test loss: {:.4f}'.format(test_loss))
return test_loss
# +
def plot_reconstrunction(x):
with torch.no_grad():
z = q.sample({"x": x}, return_all=False)
recon_batch = p.sample_mean(z).view(-1, 1, 28, 28)
comparison = torch.cat([x.view(-1, 1, 28, 28), recon_batch]).cpu()
return comparison
def plot_image_from_latent(z_sample):
with torch.no_grad():
sample = p.sample_mean({"z": z_sample}).view(-1, 1, 28, 28).cpu()
return sample
# +
writer = SummaryWriter()
z_sample = 0.5 * torch.randn(64, z_dim).to(device)
_x, _ = iter(test_loader).next()
_x = _x.to(device)
for epoch in range(1, epochs + 1):
train_loss = train(epoch)
test_loss = test(epoch)
recon = plot_reconstrunction(_x[:8])
sample = plot_image_from_latent(z_sample)
writer.add_scalar('train_loss', train_loss.item(), epoch)
writer.add_scalar('test_loss', test_loss.item(), epoch)
writer.add_images('Image_from_latent', sample, epoch)
writer.add_images('Image_reconstrunction', recon, epoch)
writer.close()
# -
|
examples/mmd_vae.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <small><i>This notebook was prepared by [<NAME>](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).</i></small>
# # Solution Notebook
# ## Problem: Determine if a tree is a valid binary search tree.
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# ## Constraints
#
# * Can the tree have duplicates?
# * Yes
# * Can we assume we already have a Node class?
# * Yes
# ## Test Cases
#
# <pre>
# Valid:
# 5
# / \
# 5 8
# / \ /
# 4 6 7
#
# Invalid:
# 5
# / \
# 5 8
# \
# 20
# </pre>
# ## Algorithm
#
# We'll use a recursive solution that valides left <= current < right, passing down the min and max values as we do a depth-first traversal.
#
# * If the node is None, return False
# * If min is set and the node's value <= min, return False
# * if max is set and the node's value > max, return False
# * Recursively call the validate function on node.left, updating max
# * Recursively call the validate function on node.right, updating min
#
# Complexity:
# * Time: O(n)
# * Space: O(h), where h is the height of the tree
# ## Code
# %run ../bst/bst.py
# +
def validate_bst(node):
return __validate_bst__(node, None, None)
def __validate_bst__(node, mininum, maximum):
if node is None:
return True
if mininum is not None and node.data <= mininum:
return False
if maximum is not None and node.data > maximum:
return False
if not __validate_bst__(node.left, mininum, node.data):
return False
if not __validate_bst__(node.right, node.data, maximum):
return False
return True
# -
# ## Unit Test
# +
# %%writefile test_bst_validate.py
from nose.tools import assert_equal
class TestBstValidate(object):
def test_bst_validate(self):
node = Node(5)
insert(node, 8)
insert(node, 5)
insert(node, 6)
insert(node, 4)
insert(node, 7)
assert_equal(validate_bst(node), True)
root = Node(5)
left = Node(5)
right = Node(8)
invalid = Node(20)
root.left = left
root.right = right
root.left.right = invalid
assert_equal(validate_bst(root), False)
print('Success: test_bst_validate')
def main():
test = TestBstValidate()
test.test_bst_validate()
if __name__ == '__main__':
main()
# -
# %run -i test_bst_validate.py
|
graphs_trees/bst_validate/bst_validate_solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Median of Two Sorted Arrays
# + active=""
# There are two sorted arrays nums1 and nums2 of size m and n respectively.
# Find the median of the two sorted arrays. The overall run time complexity should be O(log (m+n)).
# You may assume nums1 and nums2 cannot be both empty.
#
# Example 1:
# nums1 = [1, 3]
# nums2 = [2]
# The median is 2.0
#
# Example 2:
# nums1 = [1, 2]
# nums2 = [3, 4]
# The median is (2 + 3)/2 = 2.5
# -
class Solution: # 81.45%
def findMedianSortedArrays(self, nums1, nums2): # 99.91%
lst = nums1 + nums2
lst.sort()
n = len(lst)
if n % 2 == 1:
return lst[n//2]
else:
h = n//2
return (lst[h-1] + lst[h]) / 2
nums1 = list(range(0, 1000, 2))
nums2 = list(range(1, 1001, 2))
ans = Solution()
ans.findMedianSortedArrays(nums1, nums2)
# %%timeit
ans.findMedianSortedArrays(nums1, nums2)
|
004. Median of Two Sorted Arrays.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install --quiet climetlab
# # Downloading from an URL
import climetlab as cml
URL = "https://www.ncei.noaa.gov/data/international-best-track-archive-for-climate-stewardship-ibtracs/v04r00/access/csv/ibtracs.SP.list.v04r00.csv"
# + tags=[]
data = cml.load_source("url", URL)
# -
pd = data.to_pandas()
uma = pd[pd.NAME == "UMA:VELI"]
cml.plot_map(uma, style="cyclone-track")
|
docs/examples/02-source-url.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="vGUYG0tRzoHU" colab_type="text"
# This file contains
#
# 1. Conditions using Logical Operators(and or not)
# + [markdown] id="zLPw7akV05-6" colab_type="text"
# Logical Operators ကို **if/elif statement** မှာဘာကြောင့်သုံးလဲ?
#
# Eg : `if (age >= 18) (age <=40)`
# Logical operator တွေဖြစ်တဲ့ and or နဲ့ သုံးလို့ရတယ်
#
# Eg: `if(condition) and (condition)`
# `if(condition) or (condition)`
#
#
# + id="lrWvywfTzjz0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} executionInfo={"status": "ok", "timestamp": 1594889502388, "user_tz": -390, "elapsed": 993, "user": {"displayName": "AVAIRDS AV, AI, R & DS", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggh4U2DIE3BiNSBSVPM5e6QQKnQOTyp7A5mGHY=s64", "userId": "14938682159450874323"}} outputId="66c03aa4-ea53-4028-bcdf-3d3071244301"
# and operator
age = 17
if(age <18) and age > 12:
print('True')
# or operator
count = 10
if(count == 10) or (count != 2):
print('True')
# not operator
if not 2 + 2 == 5:
print('True')
# + id="IoMHIN3D0Orn" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} executionInfo={"status": "ok", "timestamp": 1594889558007, "user_tz": -390, "elapsed": 590, "user": {"displayName": "AVAIRDS AV, AI, R & DS", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggh4U2DIE3BiNSBSVPM5e6QQKnQOTyp7A5mGHY=s64", "userId": "14938682159450874323"}} outputId="6c4b9740-0cb5-41c0-b89d-388addb7648d"
# Mixed operator
if 2 + 2 == 4 and not 2 + 2 == 5 and 2 * 2 == 2 + 2:
print(True)
|
coding-exercises-avairds/week3/part1-dictionaries-multiple-conditions/multiple-conditions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
import molsysmt
# # Help
from molsysmt import info_forms, info_convert, info_select, info_viewers
info_forms()
info_forms(form_type='file')
info_convert()
info_convert(from_form='mdtraj.Trajectory', to_form_type='string')
info_convert(from_form='pytraj.Trajectory', to_form_type='file', as_rows='to')
from_list=['pytraj.Trajectory','mdanalysis.Universe']
to_list=['mdtraj.Trajectory', 'openmm.Topology']
info_convert(from_form=from_list, to_form=to_list)
to_list=['pytraj.Trajectory','nglview.NGLWidget']
info_convert(from_form_type='file', to_form=to_list)
info_convert(from_form_type='file', to_form_type='viewer')
info_viewers(from_form_type='file')
|
docs/contents/help/help.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Пример использования библиотеки gensim для тематического моделирования
# Такая полезная теорема Байеса! :)
#
# 
from gensim import corpora, models
# Импортируем данные в формте UCI Bag of Words
data = corpora.UciCorpus("docword.xkcd.txt", "vocab.xkcd.txt")
dictionary = data.create_dictionary()
# обучение модель
# %time ldamodel = models.ldamodel.LdaModel(data, id2word=dictionary, num_topics=5, passes=20, alpha=1.25, eta=1.25)
# Сохранение модели
ldamodel.save("ldamodel_xkcd")
# Загрузка модели
ldamodel = models.ldamodel.LdaModel.load("ldamodel_xkcd")
# выводим топы слов
for t, top_words in ldamodel.print_topics(num_topics=10, num_words=10):
print("Topic", t, ":", top_words)
# Вычисляем логарифм перплексии и немного преобразуем, чтобы привести к общепринятому виду
perplexity = ldamodel.log_perplexity(list(data))
print(2**(-perplexity))
perp = ldamodel.bound(data)
2**(-perp/float(87409))
# Добавление в модель новых документов, содержащихся в новом корупсе data2
ldamodel.update(data2, passes=10)
# Получение распределения тем для конкретного документа
doc = list(data)[0]
ldamodel.get_document_topics(doc)
# Эти люди не знают про тематические модели:
#
# 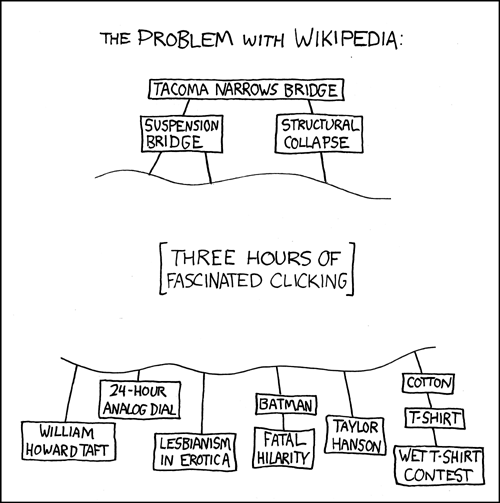 | 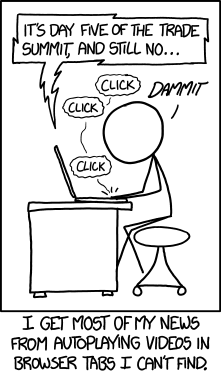
|
SearchOfDataStructure/week4/gensim.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="ebUMqK9mGIDm"
# ## The basics: interactive NumPy on GPU and TPU
#
# ---
#
#
# + colab={} colab_type="code" id="27TqNtiQF97X"
import jax
import jax.numpy as jnp
from jax import random
# + colab={} colab_type="code" id="cRWoxSCNGU4o"
key = random.PRNGKey(0)
key, subkey = random.split(key)
x = random.normal(key, (5000, 5000))
print(x.shape)
print(x.dtype)
# + colab={} colab_type="code" id="diPllsvgGfSA"
y = jnp.dot(x, x)
print(y[0, 0])
# + colab={} colab_type="code" id="8-psauxnGiRk"
x
# + colab={} colab_type="code" id="-2FMQ8UeoTJ8"
import matplotlib.pyplot as plt
plt.plot(x[0])
# -
print(jnp.dot(x, x.T))
# + colab={} colab_type="code" id="z4VX5PkMHJIu"
print(jnp.dot(x, 2 * x)[[0, 2, 1, 0], ..., None, ::-1])
# + colab={} colab_type="code" id="ORZ9Odu85BCJ"
import numpy as np
x_cpu = np.array(x)
# %timeit -n 5 -r 2 np.dot(x_cpu, x_cpu)
# + colab={} colab_type="code" id="5BKh0eeAGvO5"
# %timeit -n 5 -r 5 jnp.dot(x, x).block_until_ready()
# + [markdown] colab_type="text" id="fm4Q2zpFHUAu"
# ## Automatic differentiation
# + colab={} colab_type="code" id="MCIQbyUYHWn1"
from jax import grad
# + colab={} colab_type="code" id="kfqZpKYsHo4j"
def f(x):
if x > 0:
return 2 * x ** 3
else:
return 3 * x
# + colab={} colab_type="code" id="K_26_odPHqLJ"
key = random.PRNGKey(0)
x = random.normal(key, ())
print(grad(f)(x))
print(grad(f)(-x))
# + colab={} colab_type="code" id="q5V3A6loHrhS"
print(grad(grad(f))(-x))
print(grad(grad(grad(f)))(-x))
# + [markdown] colab_type="text" id="bmxAPFC0I8b0"
# Other JAX autodiff highlights:
#
# * Forward- and reverse-mode, totally composable
# * Fast Jacobians and Hessians
# * Complex number support (holomorphic and non-holomorphic)
# * Jacobian pre-accumulation for elementwise operations (like `gelu`)
#
#
# For much more, see the [JAX Autodiff Cookbook (Part 1)](https://jax.readthedocs.io/en/latest/notebooks/autodiff_cookbook.html).
# + [markdown] colab_type="text" id="TRkxaVLJKNre"
# ## End-to-end compilation with XLA with `jit`
# + colab={} colab_type="code" id="bKo4rX9-KSW7"
from jax import jit
# + colab={} colab_type="code" id="94iIgZSfKWh8"
key = random.PRNGKey(0)
x = random.normal(key, (5000, 5000))
# + colab={} colab_type="code" id="Ybuz8Ag9KXMd"
def f(x):
y = x
for _ in range(10):
y = y - 0.1 * y + 3.
return y[:100, :100]
f(x)
# + colab={} colab_type="code" id="Y9dx5ifSKaGJ"
g = jit(f)
g(x)
# + colab={} colab_type="code" id="UtsS67BvKYkC"
# %timeit f(x).block_until_ready()
# + colab={} colab_type="code" id="-vfcaSo9KbvR"
# %timeit -n 100 g(x).block_until_ready()
# + colab={} colab_type="code" id="E3BQF1_AKeLn"
grad(jit(grad(jit(grad(jnp.tanh)))))(1.0)
# + [markdown] colab_type="text" id="Tmf1NT2Wqv5p"
# ## Parallelization over multiple accelerators with pmap
# + colab={} colab_type="code" id="t6RRAFn1CEln"
jax.device_count()
# + colab={} colab_type="code" id="tEK1I6Duqunw"
from jax import pmap
# + colab={} colab_type="code" id="S-iCNfeGqzkY"
y = pmap(lambda x: x ** 2)(jnp.arange(8))
print(y)
# + colab={} colab_type="code" id="xgutf5JPP3wi"
y
# + colab={} colab_type="code" id="uvDL2_bCq7kq"
import matplotlib.pyplot as plt
plt.plot(y)
# + [markdown] colab_type="text" id="xf5N9ZRirJhL"
# ### Collective communication operations
# +
from functools import partial
from jax.lax import psum
@partial(pmap, axis_name='i')
def f(x):
total = psum(x, 'i')
return x / total, total
normalized, total = f(jnp.arange(8.))
print(f"normalized:\n{normalized}\n")
print("total:", total)
# + [markdown] colab_type="text" id="jC-KIMQ1q-lK"
# For more, see the [`pmap` cookbook](https://colab.research.google.com/github/google/jax/blob/main/cloud_tpu_colabs/Pmap_Cookbook.ipynb).
# -
# ## Automatic parallelization with sharded_jit (new!)
from jax.experimental import sharded_jit, PartitionSpec as P
# +
from jax import lax
conv = lambda image, kernel: lax.conv(image, kernel, (1, 1), 'SAME')
# +
image = jnp.ones((1, 8, 2000, 1000)).astype(np.float32)
kernel = jnp.array(np.random.random((8, 8, 5, 5)).astype(np.float32))
np.set_printoptions(edgeitems=1)
conv(image, kernel)
# -
# %timeit conv(image, kernel).block_until_ready()
# +
image_partitions = P(1, 1, 4, 2)
sharded_conv = sharded_jit(conv,
in_parts=(image_partitions, None),
out_parts=image_partitions)
sharded_conv(image, kernel)
# -
# %timeit -n 10 sharded_conv(image, kernel).block_until_ready()
|
cloud_tpu_colabs/JAX_NeurIPS_2020_demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Support for entering your Multiple Choice questions and answers into Blackboard
# The class mc_question does not do much more than collecting the question (string)
# and the multiple answers , and which answer is the correct one
# (only one allowed to be correct).
# -
# # This cell is defining the class for the MC questions
# +
class mc_question:
"""
Support class for creating mulitple choice question for Blackboard
Attributes
----------
default_points : int
default points for the question (for future use)
status : dictionary
key value pairs for error and status and validation of the objects
Methods
-------
enter_answer()
enter_question()
export_to(fmt="blackboard")
mark_answer()
reset()
validate()
"""
version=0.1
max_answers=5
default_points=2
status = {'error':-9,'empty':0,'question_complete':1,'answers_complete':2,
'validated':4}
def __init__(self):
"""create a new multiple choice question"""
self.label="" # a label id for the question
self.topic="" # enter a topic or leave blank"
self.question="" #insert your text question here"
self.answers = []
self.correct = [ False, False, False, False, False]
self.points=self.default_points
self.number=-1 # for random order in blackboard
self.istatus=self.status['empty']
def __str__(self):
result="Question: "+self.question+"\n"
result+='\n'
for i,a in enumerate(self.answers):
if self.correct[i]:
flag="(T)"
else:
flag="(F)"
result+="Answer "+flag+" : "+a+"\n"
return result
def enter_answer(self):
"""Allows user to enter an answer (string)."""
if len(self.answers)<self.max_answers:
self.answers.append(input("Add an(other) answer:"))
else:
print("You already provided "+
"the maximum number of answers ({0}) ".format(len(self.answers)))
if len(self.answers)==self.max_answers:
self.istatus+=self.status['answers_complete']
def enter_question(self):
"""Allows user to enter a question (string)."""
self.question=input("Enter your question (hit enter to complete):")
self.istatus+=self.status['question_complete']
## this is the important function: formatting for Blackboard
## http://blackboardsupport.calpoly.edu/content/faculty/tips_upload.html#questions
def export_to(self,fmt='blackboard'):
"""
Puts the question and answers into a single string.
Parameters:
fmt : str, optional
Determines the format of the text (default is for Blackboard MC questions).
Currently, only one format is supported.
"""
if fmt=="blackboard" or fmt=="Blackboard":
line=""
delim='\t' # use a tab to separate entries (columns)
content_type="MC"
question=self.question
line=line+content_type+delim+question+delim
for i, a in enumerate(self.answers):
if self.correct[i]:
flag='Correct'
else:
flag='Incorrect'
line=line+a+delim+flag+delim
line=line[:-1]
return line
def mark_answer(self):
"""Allows user to mark the correct answer (only one correct answer is allowed)."""
print("Enter 'a' and hit enter to mark the answer"+
"as the correct answer, else just hit enter")
for i, a in enumerate(self.answers):
res=input(a+" <<< ")
if len(res) > 0:
if res[0]=='a' or res[0] == 'A':
self.correct[i]=True
print("set correct "+str(i))
else:
self.correct[i]=False
else:
self.correct[i]=False
def reset(self,what="all"):
"""
Clear parts or all of the content
Parameters:
-----------
what : str, optional
Determines what attributes of the object are reset to default values.
Currenly, default and only option is 'all': a full reset via __init__.
"""
if what=='all':
self.__init__()
def validate(self):
"""
Validate the multiple choice question that it is formally correct.
This function does a check of the answers: are there sufficient answers,
and is one of the questions marked as the correct answer.
It also does a check if a question had been entered.
It does not check for logic or context relations between questions and answers!
"""
if self.istatus<0:
result=False
print ("Multiple choice question could "+
"not be validated (status={0})".format(self.istatus))
elif self.istatus<self.status['validated']:
icheck=0
result=False
for i, a in enumerate(self.answers):
if self.correct[i]:
icheck+=1
if icheck==1:
result=True
else:
print ("Check your answer solution guide (self.correct)!")
if self.istatus!=3 and self.istatus!=1:
print ("Check your question text. Was the question entered already?")
result=False
return result
# -
# ## The code below is designed to create questions
#
# you can change the variable
# +
# USERS SHOULD ADJUST THESE PARAMETERS
# FOR TEST PURPOSE: set variable nquestion to 1 or 2
# (Later you can set it to 10
# if you are ready to enter 10 question with 5 answers for each question.)
nquestion = 2
# add to list of questions
# file name for the export of the content of list quiz to a simple text file.
# this can be uploaded to Blackboard.
outfile="test.txt"
append_to_file=False
# -
# ### Here is the main loop
# ### Notes:
# When asked for the correct answer, type 'a + ENTER' for the correct answer. Hit just ENTER for the other false answers.
# +
# new list to collect the MC questions
quiz = []
test=mc_question() # a new test question
for n in range(nquestion):
# fill in your test question
test.enter_question()
# fill in the answers
for i in range(mc_question.max_answers):
test.enter_answer()
# mark the correct answer
test.mark_answer()
if test.validate():
quiz.append(test.export_to(fmt='blackboard'))
print("Passed validation test. Export and append to list of MC questions.")
print(".... ")
test.reset()
# -
# ### Writing all the contents to a file
# +
# export all questions
append_to_file=False
if (append_to_file):
fout=open(outfile,'a')
else:
fout=open(outfile,'w')
for q in quiz:
fout.write(q+'\n')
fout.close()
print("wrote text to file "+outfile)
print("done.")
# -
print(quiz[0])
print(quiz[1])
# References:
#
# [Link to Blackboard text formatting information](http://blackboardsupport.calpoly.edu/content/faculty/tips_upload.html#questions)
|
multiple_choice_questions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/chrismarkella/Kaggle-access-from-Google-Colab/blob/master/generators_starter.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="5pYQ5TSvG_YE" colab_type="text"
# ###Generators
# `eagerness` vs `laziness`
#
# ###Eagerness
# - Return the `entire list` even if I want to process the entries `one by one`.
# - It is also use `memory` for the `whole list`.
#
# ###It is wastfull in two ways
# - `timewise`
# - `memory wise`
# + id="PeUTH6M9Fmes" colab_type="code" colab={}
from time import sleep
# + id="utajTj2wGn6O" colab_type="code" outputId="0a75ffb1-94a2-4bba-c607-e41bc64d36d7" colab={"base_uri": "https://localhost:8080/", "height": 121}
def compute():
rv = []
for i in range(5):
print('waiting...')
sleep(0.5)
rv.append(i)
return rv
compute()
# + [markdown] id="8AGEx7o0I2B6" colab_type="text"
#
# ###Laziness
# - return the elements `one by one`
# - takes `memory only for one` element
# ```python
# for x in xs:
# pass
# ```
# corresponds to the underlying methods `iter` and `next`
#
#
# ```python
# xi = iter(xs)
# while True:
# x = next(xi)
# ```
# We could implement a `class` with the following `underscore-underscore` mentods:
# - `__iter__`
# - `__next__`
#
#
#
#
# + id="D2bz4OxfIY7d" colab_type="code" outputId="95ecccb7-1f07-4ef9-d23b-79d82ea28a34" colab={"base_uri": "https://localhost:8080/", "height": 191}
class Compute:
def __iter__(self):
self.last = 0
return self
def __next__(self):
rv = self.last
self.last = self.last + 1
if self.last > 5:
raise StopIteration()
else:
print('waiting...')
sleep(0.5)
return rv
for value in Compute():
print(value)
# + [markdown] id="vEqd-M-jMuqi" colab_type="text"
# ###This solved the `eagerness` problem.
# However this code looks `ugly`.
#
#
# ```python
# class Compute:
# def __iter__(self):
# self.last = 0
# return self
# def __next__(self):
# rv = self.last
# self.last = self.last + 1
# if self.last > 5:
# raise StopIteration()
# else:
# print('waiting...')
# sleep(0.5)
# return rv
#
# ```
# There is a simplier way to write this.
#
# Using the generator syntax with `yield`:
#
#
# ```python
# def compute_gen():
# for i in range(5):
# print('waiting...')
# sleep(0.5)
# yield i
# ```
#
#
#
# + id="rBMrQJFeLsOm" colab_type="code" outputId="af521277-1841-43ce-dae1-39fa6a11e343" colab={"base_uri": "https://localhost:8080/", "height": 191}
def compute_gen():
for i in range(5):
print('waiting...')
sleep(0.5)
yield i
for value in compute_gen():
print(value)
|
generators_starter.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %pylab inline
import numpy as np
import matplotlib.pyplot as plt
# PyTorch imports
import torch
# This has neural network layer primitives that you can use to build things quickly
import torch.nn as nn
# This has things like activation functions and other useful nonlinearities
from torch.nn import functional as F
# This has various gradient descent algorithms
import torch.optim
# In order to take derivatives, we have to wrap things as a Variable or a Parameter.
# Variables are things like inputs to the model
# Parameters are things like weights
# If you make a child class of nn.Module, it automatically keeps tracks of all parameters declared during
# __init__ for you - really handy!
from torch.autograd import Variable
from torch.nn import Parameter
# -
# ## Linear regression example
#
# In modern neural network libraries, one of the big things is that the library handles calculating derivatives of expressions analytically for you without having to do things by hand. These derivatives can be used for a lot of things, not just training a standard neural network.
#
# In this example, we'll set up a linear regression on a simple dataset and use PyTorch's automatic differentiation to optimize the parameters via gradient descent.
#
# For this example, we'll just use fake data, since the main point is to get the idea of how PyTorch works.
#
# ### Some things to play with:
#
# - Use nn.Linear to set up the regression rather than doing it directly
# - Try changing the model/data to some arbitrary nonlinear regression problem
# - Use a different objective function for the regression
# - Use the derivatives for something else
# - Implement a proper train/test split
# ## Data
#
# First we'll make some fake data. This will just be some noisy points on a line/plane/etc.
def makeData(features = 1, outputs = 1, N=50):
A = np.random.randn(features, outputs)
b = np.random.randn(outputs)
x = np.random.randn(N, features)*5
return x, np.matmul(x,A)+b + np.random.randn(N, outputs)
# +
# Example generated dataset
xdata, ydata = makeData()
plt.scatter(xdata,ydata)
plt.show()
# -
# ## Regression model
#
# We'll now make a child class of nn.Module to take advantage of PyTorch automatically keeping track of parameters for us.
class LinearRegression(nn.Module):
# This function is called when you make an instance of this class
def __init__(self, features = 1, outputs = 1):
# Inherit stuff from nn.Module
super(LinearRegression,self).__init__()
# Initialize parameters of the regression
# In this case, the regression will look like: y = A*x + b
# We will want to run this on all the data in parallel, so
# 'x' will be a matrix of size BATCH x FEATURES
# 'A' should be a matrix of size FEATURES x OUTPUTS
# 'b' should be a vector of size OUTPUTS
#
# We'll initialize these to Gaussian distributed random values
#
# For stuff to work with PyTorch, it has to be converted to FloatTensor, LongTensor, etc
# rather than being NumPy arrays. We can then wrap it in Variable or Parameter as needed.
self.A = Parameter(torch.FloatTensor(np.random.randn(features, outputs)))
self.b = Parameter(torch.FloatTensor(np.random.randn(outputs)))
# This initializes a gradient descent algorithm - here, Stochastic Gradient Descent,
# with a step size of 1e-2. The first argument is a list of parameters to do
# gradient descent over - conveniently, PyTorch is keeping track of that for us so
# we can just call self.parameters() to get the ones we've defined so far
#
# A commonly used optimizer that is a bit more sophisticated is torch.optim.Adam(),
# which you could just swap out here
self.optimizer = torch.optim.SGD(self.parameters(), lr = 1e-2)
# This function takes an input batch and returns the output of the regression
def forward(self,x):
# torch.matmul does a matrix multiplication
# If you do *, it means an element-wise multiplication
#
# Note that even though b is a vector, we can add it and PyTorch automatically reshapes
# it and broadcasts it appropriately to each example
return torch.matmul(x, self.A) + self.b
# Here we'll put the error function we want to minimize for the regression problem.
# In machine learning this is usually called the 'loss'.
#
# In this case, we want to minimize the mean squared error between the true values (y)
# and the predictions (p)
#
# Writing .sum(1) means summing over axis 1. Writing .mean(0) means taking the average over
# axis 0 (the batch dimension)
# We can also write .mean() or .sum() as a short-hand to do that over all axes at once
def loss(self, y, p):
return ((y-p)**2).sum(1).mean(0)
# Here we'll put the code to:
# - Make predictions given the inputs
# - Calculate the derivative of the error
# - Perform gradient descent on the parameters
def optimize(self, x, y):
# PyTorch stores derivatives on the variables of interest directly, so
# before we do anything we should zero those out.
self.zero_grad()
# We now calculate the predictions
p = self.forward(x)
# Now we calculate the error
loss = self.loss(y,p)
# Now, we calculate the derivative of all of the Parameters with respect to the error
# If you call backward() multiple times, it accumulates all the derivatives additively
# until you clear them.
#
# '.backward()' has to be called on a scalar. To do higher order derivatives, you need to use
# functions from torch.autograd, so we won't get into that here.
#
# If you want to look at the derivatives directly, you can access for example:
# self.A.grad
# self.b.grad
loss.backward()
# Now, we will take one step of gradient descent
self.optimizer.step()
# Return the error so we can watch the model converge
# We're projecting back to standard NumPy formats here so we
# don't carry around excess PyTorch-specific information
return loss.data.numpy()
# ## Fitting
#
# Now lets fit the model. I've included a bit of code to show the results in real time as it trains.
# +
from IPython import display
import time
xdata, ydata = makeData()
linear = LinearRegression(xdata.shape[1], ydata.shape[1])
errors = []
grada = []
gradb = []
for epoch in range(20):
# This trains the model. We have to wrap the inputs as Variables.
# For more extensive work, its useful to define a function to do this mapping
# since you end up doing it quite a lot.
# Adding requires_grad=False for Variables you won't need derivatives for speeds things up a bit
err = linear.optimize(Variable(torch.FloatTensor(xdata), requires_grad=False),
Variable(torch.FloatTensor(ydata), requires_grad=False))
errors.append(err)
# Storing gradients for visualization
# The whole .data.numpy().copy() thing is to ensure that these are not pointers to the
# PyTorch gradients but are just the actual numbers
grada.append(linear.A.grad.data.numpy().copy()[0,0])
gradb.append(linear.b.grad.data.numpy().copy()[0])
# This is all plotting stuff
plt.clf()
plt.subplot(1,3,1)
plt.title("Error")
plt.plot(errors)
plt.subplot(1,3,2)
plt.title("Gradients")
plt.plot(grada,label="dL/dA")
plt.plot(gradb,label="dL/db")
plt.legend()
plt.subplot(1,3,3)
x = np.arange(-10,10,0.1)
x = x.reshape((x.shape[0],1))
y = linear.forward(Variable(torch.FloatTensor(x), requires_grad=False)).data.numpy()
plt.title("Fit")
plt.scatter(xdata,ydata)
plt.plot(x,y,'sienna')
plt.gcf().set_size_inches((18,6))
display.clear_output(wait=True)
display.display(plt.gcf())
time.sleep(0.01)
plt.clf()
# -
# ## 2d example
# +
xdata, ydata = makeData(features=2, outputs=1, N=400)
linear = LinearRegression(xdata.shape[1], ydata.shape[1])
errors = []
for epoch in range(20):
# This trains the model. We have to wrap the inputs as Variables.
# For more extensive work, its useful to define a function to do this mapping
# since you end up doing it quite a lot.
# Adding requires_grad=False for Variables you won't need derivatives for speeds things up a bit
err = linear.optimize(Variable(torch.FloatTensor(xdata), requires_grad=False),
Variable(torch.FloatTensor(ydata), requires_grad=False))
errors.append(err)
# This is all plotting stuff
plt.clf()
plt.subplot(1,2,1)
plt.title("Error")
plt.plot(errors)
x1,x2 = np.meshgrid( np.arange(-20,20,2.0), np.arange(-20,20,2.0) )
x1 = x1.reshape((400,1))
x2 = x2.reshape((400,1))
x = np.hstack([x1,x2])
y = linear.forward(Variable(torch.FloatTensor(x), requires_grad=False)).data.numpy()
plt.subplot(1,2,2)
plt.contour(x[:,0].reshape((20,20)),
x[:,1].reshape((20,20)),
y[:,0].reshape((20,20)))
plt.scatter(xdata[:,0],xdata[:,1], c=ydata[:,0])
plt.gcf().set_size_inches((12,6))
display.clear_output(wait=True)
display.display(plt.gcf())
time.sleep(0.01)
plt.clf()
# -
|
LinearRegression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="bOChJSNXtC9g"
# # Introduction to Python
# + [markdown] colab_type="text" id="OLIxEDq6VhvZ"
# In this lesson we will learn the basics of the Python programming language (version 3). We won't learn everything about Python but enough to do some basic machine learning.
#
# <img src="figures/python.png" width=350>
#
#
#
# + [markdown] colab_type="text" id="VoMq0eFRvugb"
# # Variables
# + [markdown] colab_type="text" id="qWro5T5qTJJL"
# Variables are objects in Python that can hold anything with numbers or text. Let's look at how to create some variables.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="0-dXQiLlTIgz" outputId="38d1f8a5-b067-416b-b042-38a373624a8b"
# Numerical example
x = 4 #change 5 to 4
print (x)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="5Ym0owFxTkjo" outputId="72c2781a-4435-4c21-b15a-4c070d47bd86"
# Text example
message = "hello, Tom" #changed var name and string value
print (message)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="1a4ZhMV1T1-0" outputId="0817e041-5f79-46d8-84cc-ee4aaea0eba2"
# Variables can be used with each other
a = 2.5
b = 4.5
c = a + b #changed a and b values to floats
print (c)
# + [markdown] colab_type="text" id="nbKV4aTdUC1_"
# Variables can come in lots of different types. Even within numerical variables, you can have integers (int), floats (float), etc. All text based variables are of type string (str). We can see what type a variable is by printing its type.
# + colab={"base_uri": "https://localhost:8080/", "height": 153} colab_type="code" id="c3NJmfO4Uc6V" outputId="04b91fa4-51af-48f4-e9ac-591b5bf3e714"
# int variable
num_int = 100
print (num_int)
print (type(num_int)) #changed var name and value
# float variable
num_float = 6.542
print (num_float)
print (type(num_float)) #changed var name and value
# text variable
x = "text string"
print (x)
print (type(x)) #changed string value
# boolean variable
boolx = False
print (boolx)
print (type(boolx)) #changed var name and value
# + [markdown] colab_type="text" id="6HPtavfdU8Ut"
# It's good practice to know what types your variables are. When you want to use numerical operations on them, they need to be compatible.
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="8pr1-i7IVD-h" outputId="c2bce48d-b69f-4aab-95c1-9e588f67a6c3"
# int variables
a = 10
b = 500
print (a + b) #changed a and b values
# string variables
a = "5555"
b = "3333"
print (a + b) #changed a and b values
# + [markdown] colab_type="text" id="q4R_UF6PVw4V"
# # Lists
# + [markdown] colab_type="text" id="LvGsQBj4VjMl"
# Lists are objects in Python that can hold a ordered sequence of numbers **and** text.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="9iPESkq9VvlX" outputId="67dfbe9f-d4cb-4a62-a812-7c5c8a01c2fa"
# Creating a list
list_x = ["apple", "banana", "orange"] #changed list values to strings
print (list_x)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="7lbajc-zV515" outputId="4345bbe0-0f0c-4f84-bcf2-a76130899f34"
# Adding to a list
list_x.append("pineapple") #changed append value to another fruit
print (list_x)
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="W0xpIryJWCN9" outputId="a7676615-aff1-402f-d41f-81d004728f94"
# Accessing items at specific location in a list
print ("first item: ", list_x[0])
print ("second item: ", list_x[1])
print ("third item: ", list_x[2])
print ("last item: ", list_x[-1]) # the last item
print ("second to last item: ", list_x[-2]) # the second to last item
##changed text to be more descriptive
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="VSu_HNrnc1WK" outputId="3c40cce2-9599-41aa-b01c-7c6f39329212"
# Slicing
print ("full list: ", list_x[:])
print ("third item on: ", list_x[2:])
print ("second and third item: ", list_x[1:3])
print ("everything but last item: ", list_x[:-1])
##changed text to be more descriptive
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="dImY-hVzWxB4" outputId="8394f232-aa11-4dbd-8580-70adb5adc807"
# Length of a list
len(list_x)
print("the list has {} items".format(len(list_x))) #prints out length of list using format
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="3-reXDniW_sm" outputId="382d1a40-ad1a-49f7-f70f-2c2a02ffd88d"
# Replacing items in a list
list_x[1] = "grape" #changed value to new fruit name
print (list_x)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="X8T5I3bjXJ0S" outputId="1ede1c5c-c6ea-452f-b13d-ff9efd3d53b0"
# Combining lists
list_y = ["cucumber", "pepper", "onion"] #changed list values
list_z = list_x + list_y
print (list_z)
# + [markdown] colab_type="text" id="ddpIO6LLVzh0"
# # Tuples
# + [markdown] colab_type="text" id="CAZblq7oXY3s"
# Tuples are also objects in Python that can hold data but you cannot replace their values (for this reason, tuples are called immutable, whereas lists are known as mutable).
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="G95lu8xWXY90" outputId="c23250e5-534a-48e6-ed52-f034859f73c2"
# Creating a tuple
tuple_x = ("red", "blue", "yellow") #change tuple values
print (tuple_x)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="kq23Bej1acAP" outputId="34edfbff-dbc0-4385-a118-7f1bcc49e84f"
# Adding values to a tuple
tuple_x = tuple_x + ("green",) #changed new item to add to tuple
print (tuple_x)
# + colab={"base_uri": "https://localhost:8080/", "height": 164} colab_type="code" id="vyTmOc6BXkge" outputId="dadeac9a-4bb4-43a3-ff40-e8ca6a05ba2c"
# Trying to change a tuples value (you can't, this should produce an error.)
tuple_x[1] = "purple" #changed name
# + [markdown] colab_type="text" id="UdlJHkwZV3Mz"
# # Dictionaries
# + [markdown] colab_type="text" id="azp3AoxYXS26"
# Dictionaries are Python objects that hold key-value pairs. In the example dictionary below, the keys are the "name" and "eye_color" variables. They each have a value associated with them. A dictionary cannot have two of the same keys.
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="pXhNLbzpXXSk" outputId="e4bb80e5-4e7b-4cbb-daa6-77490ab25145"
# Creating a dictionary
dog = {"name": "Sparky",
"eye_color": "black"} #changed name and color
print (dog)
print ("the dog's name is " + dog["name"]) #made more descriptive
print ("the dog's eye color is " + dog["eye_color"]) #made more descriptive
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="1HXtX8vQYjXa" outputId="ad8d1a0f-d134-4c87-99c1-0f77140f2de0"
# Changing the value for a key
dog["eye_color"] = "blue" #changed value
print (dog)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="qn33iB0MY5dT" outputId="bd89033e-e307-4739-8c1d-f957c32385b5"
# Adding new key-value pairs
dog["age"] = "7" #changed value
print (dog)
print ("the dog's age is " + dog["age"])
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="g9EYmzMKa9YV" outputId="4b9218b9-2f4d-4287-932a-caba430713aa"
# Length of a dictionary
print (len(dog))
print ("there are {} items in this dictionary".format(len(dog))) #prints out how many items using format
# + [markdown] colab_type="text" id="B-DInx_Xo2vJ"
# # If statements
# + [markdown] colab_type="text" id="ZG_ICGRGo4tY"
# You can use `if` statements to conditionally do something.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="uob9lQuKo4Pg" outputId="21d40476-ea6a-4149-f744-0119d0894d77"
# If statement
x = 8
if x < 6:
score = "C"
elif x <= 8:
score = "B"
else:
score = "A"
print (score) #changed score values
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="vwsQaZqIpfJ3" outputId="1f190875-b910-4e54-a58a-d4230b7c8169"
# If statment with a boolean
x = False
if x:
print ("it worked")
else:
print ("no it didn't") #added else statement
# + [markdown] colab_type="text" id="sJ7NPGEKV6Ik"
# # Loops
# + [markdown] colab_type="text" id="YRVxhVCkn0vc"
# In Python, you can use `for` loop to iterate over the elements of a sequence such as a list or tuple, or use `while` loop to do something repeatedly as long as a condition holds.
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="OB5PtyqAn8mj" outputId="b4595670-99d4-473e-b299-bf8cf47f1d81"
# For loop
x = 1
for i in range(10): # goes from i=0 to i=9 #changed range
x += 2 # same as x = x + 2 #changed value
print ("i={0}, x={1}".format(i, x)) # printing with multiple variables
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="6XyhCrFeoGj4" outputId="2427ae1f-85f7-4888-f47f-8de1992a84c3"
# Loop through items in a list
x = 10
for i in [0, 1, 2, 3, 4, 5]: #added to list
x += 10 #changed value
print ("i={0}, x={1}".format(i, x))
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="5Tf2x4okp3fH" outputId="1ac41665-2f35-4c7d-e9f5-22614d3ba35c"
# While loop
x = 1
while x < 10: #changed while statement
x += 1 # same as x = x + 1
print (x)
# + [markdown] colab_type="text" id="gJw-EDO9WBL_"
# # Functions
# + [markdown] colab_type="text" id="hDIOUdWCqBwa"
# Functions are a way to modularize reusable pieces of code.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="iin1ZXmMqA0y" outputId="3bfae4a7-482b-4d43-8350-f8bb5e8a35ac"
# Create a function
def multiply_two(x): #changed function
x = x * 2
return x
# Use the function
score = 5
score = multiply_two(x=score)
print (score)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="DC6x3DMrqlE3" outputId="8965bfab-3e20-41ae-9fc1-f22a7d4f3333"
# Function with multiple inputs
def join_name(first_name, last_name):
joined_name = first_name + " " + last_name
return joined_name
# Use the function
first_name = ["John", "Tom", "Brandon"] #changed string to a list with multiples values
last_name = ["Doe", "Pike", "Mohr"] #changed string to a list with multiple values
x = 0
for i in first_name:
joined_name = join_name(first_name=first_name[x], last_name=last_name[x])
x += 1
print (joined_name)
print ("~list complete~") #added for loop to print out each first and last name in the list
# + [markdown] colab_type="text" id="lBLa1n54WEd2"
# # Classes
# + [markdown] colab_type="text" id="mGua8QnArAZh"
# Classes are a fundamental piece of object oriented programming in Python.
# + colab={} colab_type="code" id="DXmPwI1frAAd"
# Creating the class
class Pets(object):
# Initialize the class
def __init__(self, breed, color, name):
self.breed = breed
self.color = color
self.name = name
# For printing
def __str__(self):
return "{0} {1} named {2}.".format(self.color, self.breed, self.name)
# Example function
def change_name(self, new_name):
self.name = new_name
#changed species to breed for below example
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="ezQq_Fhhrqrv" outputId="bf159745-99b1-4e33-af4d-f63924a1fe74"
# Creating an instance of a class
my_dog = Pets(breed="dalmation", color="black and white", name="Spots",)
print (my_dog)
print ("my dog's name is " + my_dog.name) #added descriptive text
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="qTinlRj1szc5" outputId="80939a31-0242-4465-95ff-da0e5caaa67c"
# Using a class's function
my_dog.change_name(new_name="Spots Jr.") #changed new name
print (my_dog)
print (my_dog.name)
# + [markdown] colab_type="text" id="kiWtd0aJtNtY"
# # Additional resources
# + [markdown] colab_type="text" id="cfLF4ktmtSC3"
# This was a very quick look at Python and we'll be learning more in future lessons. If you want to learn more right now before diving into machine learning, check out this free course: [Free Python Course](https://www.codecademy.com/learn/learn-python)
|
notebooks/Python-in-2-days/D1_L3_Python/__Python_Summary__.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Jackesgamero/TensorFlow-for-Deep-Learning/blob/master/Fashion-MNIST%20Clohing%20Classifier/Clothes_Image_Identifier.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Gg4RX_ebmdsw"
# ## Instalar e importar dependencias
#
#
# + [markdown] id="637RNl7-mpnT"
# Utilizamos la API TensorFlow Datasets para simplicar el acceso y descarga de conjuntos de datos
# + id="dyz8euFlmkur" outputId="15475fb8-a0d4-400f-acd1-e6a5dac4a326" colab={"base_uri": "https://localhost:8080/"}
# !pip install -U tensorflow_datasets
# + id="_FxXYSCXGQqQ"
import tensorflow as tf
# + id="1UbK0Uq7GWaO"
# Import TensorFlow Datasets
import tensorflow_datasets as tfds
tfds.disable_progress_bar()
# Helper libraries
import math
import numpy as np
import matplotlib.pyplot as plt
# + id="590z76KRGtKk"
import logging
logger = tf.get_logger()
logger.setLevel(logging.ERROR)
# + [markdown] id="yR0EdgrLCaWR"
# ## Importar el conjunto de datos Fashion MNIST
#
# + [markdown] id="DLdCchMdCaWQ"
# Utilizaremos el conjunto de datos [Fashion MNIST](https://github.com/zalandoresearch/fashion-mnist), que contiene 70.000 imágenes en escala de grises en 10 categorías. Las imágenes muestran prendas de vestir individuales a baja resolución (28 $\times$ 28 pixels), como se puede ver aquí:
#
# <table>
# <tr><td>
# <img src="https://tensorflow.org/images/fashion-mnist-sprite.png"
# alt="Fashion MNIST sprite" width="600">
# </td></tr>
# <tr><td align="center">
# <b>Figure 1.</b> <a href="https://github.com/zalandoresearch/fashion-mnist">Fashion-MNIST samples</a> (by Zalando, MIT License).<br/>
# </td></tr>
# </table>
#
# Usaremos 60.000 imágenes para entrenar la red y 10.000 imágenes para evaluar la precisión con la que la red aprendió a clasificar las imágenes.
# + id="7MqDQO0KCaWS" outputId="96cb1ab9-ec6f-430e-f194-b23b04cc80cb" colab={"base_uri": "https://localhost:8080/"}
dataset, metadata = tfds.load('fashion_mnist', as_supervised=True, with_info=True)
train_dataset, test_dataset = dataset['train'], dataset['test']
# + [markdown] id="OFvKtIOvohRb"
# La carga del conjunto de datos devuelve metadatos, así como un conjunto de *datos de entrenamiento* y un conjunto de *datos de prueba*.
#
# * El modelo se entrena usando `train_dataset`.
# * El modelo se prueba con `test_dataset`.
#
# Las imágenes son matrices de 28 $\times$ 28, con valores de píxeles en el rango `[0, 255]`. Las *etiquetas* son una matriz de números enteros, en el rango `[0, 9]`. TEstos corresponden a la *clase* de ropa que representa la imagen:
#
# <table>
# <tr>
# <th>Label</th>
# <th>Class</th>
# </tr>
# <tr>
# <td>0</td>
# <td>T-shirt/top</td>
# </tr>
# <tr>
# <td>1</td>
# <td>Trouser</td>
# </tr>
# <tr>
# <td>2</td>
# <td>Pullover</td>
# </tr>
# <tr>
# <td>3</td>
# <td>Dress</td>
# </tr>
# <tr>
# <td>4</td>
# <td>Coat</td>
# </tr>
# <tr>
# <td>5</td>
# <td>Sandal</td>
# </tr>
# <tr>
# <td>6</td>
# <td>Shirt</td>
# </tr>
# <tr>
# <td>7</td>
# <td>Sneaker</td>
# </tr>
# <tr>
# <td>8</td>
# <td>Bag</td>
# </tr>
# <tr>
# <td>9</td>
# <td>Ankle boot</td>
# </tr>
# </table>
#
# Cada imagen se asigna a una sola etiqueta. Dado que los nombres de las clases no se incluyen con el conjunto de datos, los guardaremos aquí para usarlos más adelante al trazar las imágenes:
# + id="IjnLH5S2CaWx" outputId="5b30aa67-2e32-4842-d8c1-6e269b70e91e" colab={"base_uri": "https://localhost:8080/"}
class_names = metadata.features['label'].names
print("Class names: {}".format(class_names))
# + [markdown] id="wuWheTRLpquA"
# ### Explorar datos
# + id="MaOTZxFzi48X" outputId="585ba72a-6e52-433d-ccc1-255ebd68c958" colab={"base_uri": "https://localhost:8080/"}
num_train_examples = metadata.splits['train'].num_examples
num_test_examples = metadata.splits['test'].num_examples
print("Number of training examples: {}".format(num_train_examples))
print("Number of test examples: {}".format(num_test_examples))
# + [markdown] id="ES6uQoLKCaWr"
# ## Preprocesar los datos
#
# El valor de cada píxel en los datos de la imagen es un número entero en el rango `[0,255]`. Para que el modelo funcione correctamente, estos valores deben normalizarse al rango `[0,1]`. Entonces, aquí creamos una función de normalización y luego la aplicamos a cada imagen en los conjuntos de datos de prueba y entrenamiento.
# + id="nAsH3Zm-76pB"
def normalize(images, labels):
images = tf.cast(images, tf.float32)
images /= 255
return images, labels
# The map function applies the normalize function to each element in the train
# and test datasets
train_dataset = train_dataset.map(normalize)
test_dataset = test_dataset.map(normalize)
# The first time you use the dataset, the images will be loaded from disk
# Caching will keep them in memory, making training faster
train_dataset = train_dataset.cache()
test_dataset = test_dataset.cache()
# + [markdown] id="lIQbEiJGXM-q"
# ### Explorar los datos procesados
#
# Trazamos una imagen para ver cómo se ve.
# + id="oSzE9l7PjHx0" outputId="aa76e368-e2d3-4acc-9010-b4026d309c15" colab={"base_uri": "https://localhost:8080/", "height": 269}
# Take a single image, and remove the color dimension by reshaping
for image, label in test_dataset.take(1):
break
image = image.numpy().reshape((28,28))
# Plot the image - voila a piece of fashion clothing
plt.figure()
plt.imshow(image, cmap=plt.cm.binary)
plt.colorbar()
plt.grid(False)
plt.show()
# + [markdown] id="Ee638AlnCaWz"
# Mostramos las primeras 25 imágenes del *conjunto de entrenamiento* y el nombre de la clase debajo de cada imagen.
# + id="oZTImqg_CaW1" outputId="3b4335ab-f6e7-46ca-e954-b33ef02354e3" colab={"base_uri": "https://localhost:8080/", "height": 589}
plt.figure(figsize=(10,10))
for i, (image, label) in enumerate(test_dataset.take(25)):
image = image.numpy().reshape((28,28))
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(image, cmap=plt.cm.binary)
plt.xlabel(class_names[label])
plt.show()
# + [markdown] id="59veuiEZCaW4"
# ## Construir modelo
#
# Configuramos las capas del modelo y lo compilamos
# + [markdown] id="Gxg1XGm0eOBy"
# ### Configuración de capas
#
#
# + id="9ODch-OFCaW4"
model = tf.keras.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28, 1)),
tf.keras.layers.Dense(128, activation=tf.nn.relu),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
# + [markdown] id="gut8A_7rCaW6"
# Esta red tiene tres capas:
#
# * **entrada** `tf.keras.layers.Flatten` — Esta capa transforma las imágenes de una matriz 2d de 28 $\times$ 28 pixeles, a una matriz 1d de 784 píxeles (28\*28).
#
# * **"oculta"** `tf.keras.layers.Dense`— Una capa densamente conectada de 128 neuronas. Cada neurona (o nodo) toma la entrada de los 784 nodos de la capa anterior, ponderando esa entrada de acuerdo con los parámetros ocultos que se aprenderán durante el entrenamiento, y envía un valor único a la siguiente capa.
#
# * **salida** `tf.keras.layers.Dense` — Una capa de 128 neuronas, seguida de una capa *softmax* de 10 nodos. ECada nodo representa una clase de ropa. Como en la capa anterior, la capa final toma la entrada de los 128 nodos de la capa anterior y genera un valor en el rango `[0, 1]`, rque representa la probabilidad de que la imagen pertenezca a esa clase. La suma de los 10 valores de los nodos es 1.
#
# > Nota: La funcion de activacion `softmax` y `SparseCategoricalCrossentropy()` tienen algunos problemas que son corregidos en al modelo `tf.keras`. Una aproximación mas segura, en general, es utilizar una salida lineal (sin función de activación) con `SparseCategoricalCrossentropy(from_logits=True)`.
#
#
# ### Compilar el modelo
#
# Añadimos las métricas, la función de pérdida y la función de optimización
#
# + id="Lhan11blCaW7"
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(),
metrics=['accuracy'])
# + [markdown] id="qKF6uW-BCaW-"
# ## Entrenar el modelo
#
# Primero, definimos el comportamiento de cada iteración para el conjunto de datos de entrenamiento:
# 1. Repetir siempre con `dataset.repeat()` (el parámetro `epochs` que aparece más adelante indica cuanto tiempo realizaremos el entrenamiento).
# 2. `dataset.shuffle(60000)` aleatoriza el orden por lo que nuestro modelo no se puede aprender nada de la orden de los ejemplos.
# 3. `dataset.batch(32)` indica a `model.fit` que use lotes de 32 imágenes y etiquetas al actualizar las variables del modelo.
#
# El entrenamiento se realiza llamando al método `model.fit`:
# 1. Utilizamos `train_dataset` como datos de entrenamiento.
# 2. El parámetro `epochs=5` limita el entrenamiento a 5 iteraciones completas del conjunto de datos de entrenamiento, por lo que un total de 5 * 60000 = 300000 ejemplos.
# + id="o_Dp8971McQ1"
BATCH_SIZE = 32
train_dataset = train_dataset.cache().repeat().shuffle(num_train_examples).batch(BATCH_SIZE)
test_dataset = test_dataset.cache().batch(BATCH_SIZE)
# + id="xvwvpA64CaW_" outputId="c87cab7f-32ae-4a4d-c73b-38958120b7bf" colab={"base_uri": "https://localhost:8080/"}
model.fit(train_dataset, epochs=5, steps_per_epoch=math.ceil(num_train_examples/BATCH_SIZE))
# + [markdown] id="oEw4bZgGCaXB"
# ## Evaluar la precisión
#
# Analizamos el rendimiento del modelo en el conjunto de datos de prueba
# + id="VflXLEeECaXC" outputId="f1037a90-c1fc-440a-b4fc-a130f71c3077" colab={"base_uri": "https://localhost:8080/"}
test_loss, test_accuracy = model.evaluate(test_dataset, steps=math.ceil(num_test_examples/32))
print('Accuracy on test dataset:', test_accuracy)
# + [markdown] id="xsoS7CPDCaXH"
# ## Realizar predicciones
#
# + id="Ccoz4conNCpl"
for test_images, test_labels in test_dataset.take(1):
test_images = test_images.numpy()
test_labels = test_labels.numpy()
predictions = model.predict(test_images)
# + id="Gl91RPhdCaXI" outputId="3bba3e46-cc9e-4760-95c1-81b4dd9d3ead" colab={"base_uri": "https://localhost:8080/"}
predictions.shape
# + [markdown] id="x9Kk1voUCaXJ"
# Aqui el modelo indica que la primera imagen del conjunto de prueba ha dado como resultado 32 predicciones con 10 valores cada una (el porcentaje de que sea cada prenda). Echemos un ojo a la primera predicción:
# + id="3DmJEUinCaXK" outputId="1a350548-a300-48fd-cc18-8cbf579ac127" colab={"base_uri": "https://localhost:8080/"}
predictions[0]
# + [markdown] id="-hw1hgeSCaXN"
# Una predicción es una matriz de 10 números. Estos describen la "confianza" del modelo en que la imagen corresponde a cada una de las 10 diferentes prendas de vestir. Podemos ver qué etiqueta tiene el valor de confianza más alto:
# + id="qsqenuPnCaXO" outputId="d4f8bf0f-52b9-4b99-f375-c2e6b3a4981b" colab={"base_uri": "https://localhost:8080/"}
np.argmax(predictions[0])
# + [markdown] id="ygh2yYC972ne"
# Podemos graficar esto para ver el conjunto completo de 10 predicciones.
# + id="DvYmmrpIy6Y1"
def plot_image(i, predictions_array, true_labels, images):
predictions_array, true_label, img = predictions_array[i], true_labels[i], images[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img[...,0], cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'blue'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
def plot_value_array(i, predictions_array, true_label):
predictions_array, true_label = predictions_array[i], true_label[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
thisplot = plt.bar(range(10), predictions_array, color="#777777")
plt.ylim([0, 1])
predicted_label = np.argmax(predictions_array)
thisplot[predicted_label].set_color('red')
thisplot[true_label].set_color('blue')
# + [markdown] id="d4Ov9OFDMmOD"
# Veamos la imagen 0, las predicciones y la matriz de predicciones.
# + id="HV5jw-5HwSmO" outputId="e951bb39-6986-4fa2-840e-64bf1542ace1" colab={"base_uri": "https://localhost:8080/", "height": 203}
i = 0
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(1,2,2)
plot_value_array(i, predictions, test_labels)
# + id="Ko-uzOufSCSe" outputId="42291ca9-d76e-46cd-a1cc-43492fc9360b" colab={"base_uri": "https://localhost:8080/", "height": 203}
i = 12
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(1,2,2)
plot_value_array(i, predictions, test_labels)
# + [markdown] id="kgdvGD52CaXR"
# Trazamos varias imágenes con sus predicciones. Las etiquetas de predicción correctas son azules y las etiquetas de predicción incorrectas son rojas. El número da el porcentaje (de 100) de la etiqueta predicha. Tenga en cuenta que puede estar mal incluso cuando se tiene mucha confianza.
# + id="hQlnbqaw2Qu_" outputId="c1bef7f1-bf18-425d-f3b1-4e7b255e85a4" colab={"base_uri": "https://localhost:8080/", "height": 589}
# Plot the first X test images, their predicted label, and the true label
# Color correct predictions in blue, incorrect predictions in red
num_rows = 5
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(2*2*num_cols, 2*num_rows))
for i in range(num_images):
plt.subplot(num_rows, 2*num_cols, 2*i+1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(num_rows, 2*num_cols, 2*i+2)
plot_value_array(i, predictions, test_labels)
# + [markdown] id="R32zteKHCaXT"
# Podemos usar el modelo entrenado para hacer una predicción sobre una sola imagen.
# + id="yRJ7JU7JCaXT" outputId="8249d6f5-8c5a-4237-b521-0e44cd9cd4d7" colab={"base_uri": "https://localhost:8080/"}
# Grab an image from the test dataset
img = test_images[0]
print(img.shape)
# + id="lDFh5yF_CaXW" outputId="b4e85eeb-8686-4775-b18f-cb05178c3719" colab={"base_uri": "https://localhost:8080/"}
# Add the image to a batch where it's the only member.
img = np.array([img])
print(img.shape)
# + id="o_rzNSdrCaXY" outputId="cf20840e-e954-47c6-f0b7-040db8ff0d60" colab={"base_uri": "https://localhost:8080/"}
predictions_single = model.predict(img)
print(predictions_single)
# + id="6Ai-cpLjO-3A" outputId="3dd560a4-a866-42dd-a4f8-76bb1c7142a7" colab={"base_uri": "https://localhost:8080/", "height": 300}
plot_value_array(0, predictions_single, test_labels)
_ = plt.xticks(range(10), class_names, rotation=45)
# + id="2tRmdq_8CaXb" outputId="a69bb23a-8b21-4405-c938-54286abab79e" colab={"base_uri": "https://localhost:8080/"}
np.argmax(predictions_single[0])
|
Fashion-MNIST Clothing Classifier/Clothes_Image_Identifier.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.4.6-pre
# language: julia
# name: julia-0.4
# ---
# ## Viscoplastic material
#
# Small example, how to create a material model with viscoplastic properties. Formulation is done in context of small strains
using ForwardDiff
using PyPlot
# +
"""
Hooke isotropic stiffness tensor
"""
function hookeStiffnessTensor(E, ν)
a = 1 - ν
b = 1 - 2*ν
c = 1 + ν
multiplier = E / (b * c)
return Float64[a ν ν 0 0 0;
ν a ν 0 0 0;
ν ν a 0 0 0;
0 0 0 b 0 0;
0 0 0 0 b 0;
0 0 0 0 0 b].*multiplier
end
# Pick material values
E = 152.0e3
ν = 0.3
C = hookeStiffnessTensor(E, ν)
# +
# Equations can be found from: http://mms2.ensmp.fr/msi_paris/transparents/Georges_Cailletaud/2013-GC-plas3D.pdf
"""
J2 plasticity
"""
function J(σ)
e1 = (σ[1] - σ[2])^2
e2 = (σ[2] - σ[3])^2
e3 = (σ[3] - σ[1])^2
e4 = σ[4]^2
e5 = σ[5]^2
e6 = σ[6]^2
return sqrt((e1 + e2 + e3 + 6 * (e4 + e5 + e6)) / 2.)
end
"""
Yield function
"""
function f(σ, σ_y)
return J(σ) - σ_y
end
"""
Viscoplastic potential
Parameters
----------
f_val: float
Yield function value
n: float
Material constant
K: float
Material constant
"""
function viscoplastic_potential(f_val, n, K)
n_1 = n+1
return K / n_1 * ( f_val./ K) ^ n_1
end
"""
Function for calculating stress
"""
function calculate_stress(σ_old, dϵ, C, dt, σ_y, n, K)
σ_new = σ_old + C * dϵ * dt
if f(σ_new, σ_y) > 0
# Viscoplastic potential derivated via ForwardDiff
visco_wrap(x) = viscoplastic_potential(f(x, σ_y), n, K)
dϵ_vp = ForwardDiff.gradient(visco_wrap)
σ_new = σ_old + C * (dϵ - dϵ_vp(σ_new)) * dt
end
return σ_new
end
# +
steps = 30
ϵ_hist = zeros(6)
max_strain = 0.0001
ϵ_hist[1] = max_strain
ϵ_hist[2] = max_strain*-0.3
ϵ_hist[3] = max_strain*-0.3;
# -
# Plotting with three different $dt$
# +
ϵ_tot_1 = zeros(6)
ϵ_tot_2 = zeros(6)
ϵ_tot_3 = zeros(6)
nσ_1 = zeros(6)
nσ_2 = zeros(6)
nσ_3 = zeros(6)
dt_1 = 0.01
dt_2 = 0.1
dt_3 = 0.4
σ_1 = []
σ_2 = []
σ_3 = []
ϵ_1 = []
ϵ_2 = []
ϵ_3 = []
σ_y = 150.0
nn = 0.92
K = 180.0e3
for n=1:steps
dϵ_1 = ϵ_hist / dt_1
dϵ_2 = ϵ_hist / dt_2
dϵ_3 = ϵ_hist / dt_3
nσ_1 = calculate_stress(nσ_1, dϵ_1, C, dt_1, σ_y, nn, K)
nσ_2 = calculate_stress(nσ_2, dϵ_2, C, dt_2, σ_y, nn, K)
nσ_3 = calculate_stress(nσ_3, dϵ_3, C, dt_3, σ_y, nn, K)
ϵ_tot_1 += dϵ_1 * dt_1
ϵ_tot_2 += dϵ_2 * dt_2
ϵ_tot_3 += dϵ_3 * dt_3
push!(σ_1, nσ_1[1])
push!(σ_2, nσ_2[1])
push!(σ_3, nσ_3[1])
push!(ϵ_1, ϵ_tot_1[1])
push!(ϵ_2, ϵ_tot_2[1])
push!(ϵ_3, ϵ_tot_3[1])
end
# -
PyPlot.plot(ϵ_1, σ_1)
PyPlot.plot(ϵ_2, σ_2)
PyPlot.plot(ϵ_1, σ_3)
PyPlot.xlabel("Total strain")
PyPlot.ylabel("Stress")
|
docs/tutorials/2016_5_2_viscoplastic_material_small_strain.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Created a merged corresponding authors dataset
#
# Identifies corresponding authors from PubMed Central author list if available. Otherwise, use PubMed author list.
import pathlib
import pandas
import pubmedpy
# ## Prepare country assignment
# read affiliations to country mapping
country_df = pandas.read_csv("data/affiliations/countries.tsv.xz", sep='\t', keep_default_na=False)
# be careful reading country codes: Namibia code is NA
assert country_df.notna().all(axis=None)
country_df.head()
source = 'pubmed'
directory = pathlib.Path('data') / source
affil_df = pandas.read_csv(directory / "affiliations.tsv.xz", sep="\t")
affil_df = affil_df.merge(country_df)
def read_authors(source: str) -> pandas.DataFrame:
"""
Read authors table for source, and add a countries column.
"""
directory = pathlib.Path('data') / source
# read authors
author_df = pandas.read_csv(directory / "authors.tsv.xz", sep="\t")
# read author affiliations
affil_df = pandas.read_csv(directory / "affiliations.tsv.xz", sep="\t")
affil_df = affil_df.merge(country_df)
# assign countries to authors by affiliations
primary_key = {"pubmed": "pmid", "pmc": "pmcid"}[source]
countries_df = (
affil_df
.groupby([primary_key, "position"])
.country
.apply(lambda x: ",".join(sorted(set(x))))
.reset_index()
.rename(columns={"country": "countries"})
)
author_df = author_df.merge(countries_df, how='left')
return author_df
# ## Read pubmed authors
pubmed_df = pandas.read_csv("data/pubmed/articles.tsv.xz", sep="\t")
pubmed_df.head(2)
pubmed_df.head()
pubmed_author_df = read_authors("pubmed")
pubmed_author_df.head(2)
# +
# pmc_author_df = read_authors("pmc")
# pmc_author_df.head(2)
# +
# def get_corresponding(df):
# # if df.corresponding.any():
# # df = df.query("corresponding == 1")
# # df['use_last'] = 0
# # else:
# df = df.query("reverse_position == 1")
# df['use_last'] = 1
# return df.assign(n_corresponding=len(df))
# pmc_corresp_df = (
# pubmed_df[['pmid', 'pmcid']]
# .merge(pmc_author_df)
# .assign(source="pmc")
# .groupby('pmid')
# .apply(get_corresponding)
# )
# pmc_corresp_df.head(2)
# +
# pmc_corresp_df.countries.notna().mean()
# -
pubmed_author_df.query('reverse_position == 1')
# +
# pubmed_corresp_df.countries.notna().mean()
# -
# corresp_df = pandas.concat([pmc_corresp_df, pubmed_corresp_df], sort=False).reset_index(drop=True)
corresp_df = (pubmed_author_df
.query('reverse_position == 1')
.assign(source='pubmed', use_last = 1))
fore_df = pandas.read_csv("data/names/fore-names.tsv.xz", sep="\t", keep_default_na=False)
last_df = pandas.read_csv("data/names/last-names.tsv.xz", sep="\t", keep_default_na=False)
corresp_df = (
corresp_df
.merge(fore_df[["fore_name", "fore_name_simple"]], how="left")
.merge(last_df[["last_name", "last_name_simple"]], how="left")
)
corresp_df.head(2)
# number of authors with 1+ assinged countries by source
# False indicates no assigned countries.
# True indicates one or more countries
tab = pandas.crosstab(
corresp_df.source,
corresp_df.countries.notnull(),
margins=True,
)
# percent of authors with 1+ assinged countries by source
pandas.crosstab(
corresp_df.source,
corresp_df.countries.notnull(),
margins=True, normalize="index"
).applymap("{:.1%}".format)
pmc_country = (
pubmed_df
.merge(corresp_df[['pmid', 'source', 'countries']].drop_duplicates())
)
pmc_country.head()
# If available, collect PMCID of articles with no countries to query more affiliations from:
pmcids_query = (pmc_country.query('(countries != countries) and (pmcid == pmcid)'))['pmcid']
pmcids_query
def get_frontmatter_etree_via_api(pmcid):
url = f"https://www.ncbi.nlm.nih.gov/pmc/oai/oai.cgi?verb=GetRecord&identifier=oai:pubmedcentral.nih.gov:{pmcid[3:]}&metadataPrefix=pmc_fm"
response = requests.get(url)
tree = etree.fromstring(response.content)
article = tree.find("{*}GetRecord/{*}record/{*}metadata/{*}article")
return article
from pubmedpy.pmc_oai import extract_authors_from_article
# from pubmedpy.tests.test_pmc_oai import get_frontmatter_etree_via_api
from lxml import etree
# from lxml.etree import tostring
import requests
art = get_frontmatter_etree_via_api('PMC6986237')
extract_authors_from_article(art)
art = get_frontmatter_etree_via_api('PMC6986235')
print(etree.tostring(art, encoding = 'unicode'))
extract_authors_from_article(art)
# ### Analyze sources for corresponding authors
source_df = (
pubmed_df
.merge(corresp_df[['pmid', 'source', 'use_last']].drop_duplicates())
)
source_df.head(2)
# number of articles by authorship source by journal
pandas.crosstab(source_df.journal, source_df.source, margins=True)
# number of articles by authorship source by year
pandas.crosstab(source_df.publication_date.str.slice(0, 4), source_df.source, margins=True)
# Number of articles where all corresponding authors have assigned countries by journal
articles_with_na_countries = set(corresp_df.loc[corresp_df.countries.isna()].pmid)
source_df["corresp_has_countries"] = ~source_df.pmid.isin(articles_with_na_countries)
pandas.crosstab(source_df.journal, source_df.corresp_has_countries, margins=True)
# Percent of articles where all corresponding authors have assigned countries by journal
pandas.crosstab(
source_df.journal, source_df.corresp_has_countries,
margins=True, normalize="index"
).applymap("{:.1%}".format)
corresp_df.to_csv('data/names/corresponding-authors.tsv.xz', sep='\t', index=False)
|
08.corresponding-authors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:pyvizenv] *
# language: python
# name: conda-env-pyvizenv-py
# ---
# # San Francisco Rental Prices Dashboard
#
# In this notebook, you will compile the visualizations from the previous analysis into functions that can be used for a Panel dashboard.
# imports
import panel as pn
pn.extension('plotly')
import plotly.express as px
import pandas as pd
import hvplot.pandas
import matplotlib.pyplot as plt
import os
from pathlib import Path
from dotenv import load_dotenv
from panel.interact import interact
# Read the Mapbox API key
load_dotenv()
map_box_api = os.getenv("MAP_BOX_API")
print(map_box_api)
px.set_mapbox_access_token(map_box_api)
# # Import Data
# Import the necessary CSVs to Pandas DataFrames
# YOUR CODE HERE!
file_path = Path("Data/neighborhoods_coordinates.csv")
coordinatesData = pd.read_csv(file_path).dropna()
coordinatesData.head(5)
file_path = Path("Data/sfo_neighborhoods_census_data.csv")
sfo_data = pd.read_csv(file_path,index_col="year")
#sfo_data.reset_index(inplace = True)
sfo_data.head()
# - - -
# ## Panel Visualizations
#
# In this section, you will copy the code for each plot type from your analysis notebook and place it into separate functions that Panel can use to create panes for the dashboard.
#
# These functions will convert the plot object to a Panel pane.
#
# Be sure to include any DataFrame transformation/manipulation code required along with the plotting code.
#
# Return a Panel pane object from each function that can be used to build the dashboard.
#
# Note: Remove any `.show()` lines from the code. We want to return the plots instead of showing them. The Panel dashboard will then display the plots.
# +
# Define Panel Visualization Functions
def housing_units_per_year():
"""Housing Units Per Year."""
# YOUR CODE HERE!
meanOfHousingUnitsPerYear = (sfo_data["housing_units"].groupby(['year']).mean())
barPlotOfMeanOfHousingUnitsPerYear = meanOfHousingUnitsPerYear.hvplot.bar(x='year', y= 'housing_units',ylim = (370000,390000)).opts(yformatter="%.0f", title = 'Housig Units in San Fransisco 2010-2016')
return barPlotOfMeanOfHousingUnitsPerYear
def average_gross_rent():
"""Average Gross Rent in San Francisco Per Year."""
# YOUR CODE HERE!
avgPricePerSqftAndGrossRent = (sfo_data.groupby(['year']).mean())
linePlotGrossRent = avgPricePerSqftAndGrossRent["gross_rent"].hvplot(label = 'Gross Rent' )
return linePlotGrossRent
def average_sales_price():
"""Average Sales Price Per Year."""
# YOUR CODE HERE!
avgPricePerSqftAndGrossRent = (sfo_data.groupby(['year']).mean())
linePlotSalePricePerSqft = avgPricePerSqftAndGrossRent['sale_price_sqr_foot'].hvplot(label = 'Sale Price Per Square Foot' )
return linePlotSalePricePerSqft
def average_price_by_neighborhood():
"""Average Prices by Neighborhood."""
# YOUR CODE HERE!
avgPricePerSqftByNeighborhood = (sfo_data.groupby(['year','neighborhood']).mean())
interactivePlotAvgPricePerSqftByNeighborhood = avgPricePerSqftByNeighborhood['gross_rent'].hvplot.line(x= 'year', y = 'gross_rent',title = 'Average Monthly Rent',groupby = 'neighborhood')
return interactivePlotAvgPricePerSqftByNeighborhood
def top_most_expensive_neighborhoods():
"""Top 10 Most Expensive Neighborhoods."""
# YOUR CODE HERE!
averageNeighborhood = (sfo_data.groupby(['neighborhood']).mean())
top10Neighborhood = averageNeighborhood.nlargest(10,'sale_price_sqr_foot')
top10Neighborhood.reset_index()
barPlotTop10Neighborhood = top10Neighborhood.hvplot.bar(x= 'neighborhood', y = 'sale_price_sqr_foot',title = 'Top 10 Neighbourhood' )
return barPlotTop10Neighborhood
def most_expensive_neighborhoods_rent_sales():
"""Comparison of Rent and Sales Prices of Most Expensive Neighborhoods."""
# YOUR CODE HERE!
avgPricePerSqftByNeighborhood = (sfo_data.groupby(['year','neighborhood']).mean())
avgPricePerSqftByNeighborhood.drop('housing_units', axis = 1, inplace = True )
compareBarPlot = avgPricePerSqftByNeighborhood.hvplot.bar(x= 'year',title = 'Sale Price Per Square Foot',groupby = 'neighborhood',rot = 90 )
return compareBarPlot
def parallel_coordinates():
"""Parallel Coordinates Plot."""
# YOUR CODE HERE!
averageNeighborhood = (sfo_data.groupby(['neighborhood']).mean())
top10Neighborhood = averageNeighborhood.nlargest(10,'sale_price_sqr_foot')
top10Neighborhood.reset_index(inplace = True)
parallelCoordinatesPlot = px.parallel_coordinates(top10Neighborhood, color='sale_price_sqr_foot',width = 1000, title = 'Parallel Coordinates Analysis of most expensive neighborhood in San Fransisco')
return parallelCoordinatesPlot
def parallel_categories():
"""Parallel Categories Plot."""
averageNeighborhood = (sfo_data.groupby(['neighborhood']).mean())
top10Neighborhood = averageNeighborhood.nlargest(10,'sale_price_sqr_foot')
top10Neighborhood.reset_index(inplace = True)
parallelCategoriesPlot = px.parallel_categories(
top10Neighborhood,
dimensions=['neighborhood', 'sale_price_sqr_foot', 'housing_units', 'gross_rent'],
color='sale_price_sqr_foot',width = 1500, title = 'Parallel Categoris Analysis of most expensive neighborhood in San Fransisco')
return parallelCategoriesPlot
def neighborhood_map():
"""Neighborhood Map."""
# YOUR CODE HERE!
meanPricePerSqftByNeighborhood = (sfo_data.groupby(['neighborhood']).mean())
meanPricePerSqftByNeighborhood.reset_index(inplace = True)
avgNeighborhoodLocation = pd.concat([coordinatesData,meanPricePerSqftByNeighborhood],axis = 1 , join = 'inner' )
avgNeighborhoodLocation.drop('neighborhood',axis =1 , inplace = True)
px.set_mapbox_access_token(map_box_api)
map_1 = px.scatter_mapbox(
avgNeighborhoodLocation,
lat="Lat",
lon="Lon",
color="gross_rent",
title = 'Neighborhood Map'
)
return map_1
def sunburst():
"""Sunburst Plot."""
# YOUR CODE HERE!
avgPricePerSqftByNeighborhood = (sfo_data.groupby(['year','neighborhood']).mean())
avgPricePerSqftByNeighborhood.reset_index(inplace = True)
fig = px.sunburst(avgPricePerSqftByNeighborhood ,path = ['year', 'neighborhood'],values = 'gross_rent',color = 'gross_rent', title = 'Cost Analysis of most expensive neighborhood in San Fransisco by Year')
return fig
# -
# ## Panel Dashboard
#
# In this section, you will combine all of the plots into a single dashboard view using Panel. Be creative with your dashboard design!
# +
# Create a Title for the Dashboard
# YOUR CODE HERE!
#row_of_bar = pn.Row(parallel_coordinates())
#row_of_bar.append(parallel_categories())
rowOfParallel = pn.Row(housing_units_per_year(), average_gross_rent(),average_sales_price())
colOfParallel = pn.Column(parallel_coordinates(), parallel_categories())
colOfParallelNeighborhood = pn.Column(average_price_by_neighborhood(),most_expensive_neighborhoods_rent_sales(),top_most_expensive_neighborhoods())
# Create a tab layout for the dashboard
# YOUR CODE HERE!
tabs = pn.Tabs(
("Welcome", neighborhood_map()),
("Yearly Market Analysis", rowOfParallel),
("Neighborhood Analysis", colOfParallelNeighborhood),
("Parallel Plot Analysis", colOfParallel),
("Suburst Plot Analysis", sunburst())
)
tabs
# Create the dashboard
# YOUR CODE HERE!
# -
# ## Serve the Panel Dashboard
# +
# Serve the# dashboard
# YOUR CODE HERE!
# -
# # Debugging
#
# Note: Some of the Plotly express plots may not render in the notebook through the panel functions.
#
# However, you can test each plot by uncommenting the following code
# +
#housing_units_per_year()
# +
#average_gross_rent()
# +
#average_sales_price()
# +
#average_price_by_neighborhood()
# +
#top_most_expensive_neighborhoods()
# +
#most_expensive_neighborhoods_rent_sales()
# +
#neighborhood_map()
# +
#parallel_categories()
# +
#parallel_coordinates()
# +
#sunburst()
# -
|
dashboard.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Spur Fitting
# +
import pathlib
import warnings
import warnings
warnings.filterwarnings('ignore')
import os
import sys
sys.path.append('../code/')
import pm_model_func as pmf
# Third-party
import astropy.coordinates as coord
import astropy.table as at
from astropy.table import Table, vstack
from astropy.io import fits
import astropy.units as u
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from numpy.lib.recfunctions import stack_arrays
from scipy.interpolate import InterpolatedUnivariateSpline, interp1d
from scipy.ndimage.filters import gaussian_filter
# %matplotlib inline
import gala.coordinates as gc
import gala.dynamics as gd
import gala.potential as gp
from pyia import GaiaData
from scipy.stats import binned_statistic
import arviz as az
import pymc3 as pm
import seaborn as sns
from tqdm import trange
from pymc3 import *
import theano.tensor as tt
import pymc3_ext as pmx
from patsy import dmatrix
print(f"Running on PyMC3 v{pm.__version__}")
# +
def plot_pretty(dpi=175, fontsize=15, labelsize=15, figsize=(10, 8), tex=True):
# import pyplot and set some parameters to make plots prettier
plt.rc('savefig', dpi=dpi)
plt.rc('text', usetex=tex)
plt.rc('font', size=fontsize)
plt.rc('xtick.major', pad=1)
plt.rc('xtick.minor', pad=1)
plt.rc('ytick.major', pad=1)
plt.rc('ytick.minor', pad=1)
plt.rc('figure', figsize=figsize)
mpl.rcParams['xtick.labelsize'] = labelsize
mpl.rcParams['ytick.labelsize'] = labelsize
mpl.rcParams.update({'figure.autolayout': False})
plot_pretty(fontsize=20, labelsize=20)
# +
import importlib
importlib.reload(pmf)
gaia = GaiaData('../data/gd1_ps1_with_basic_masks_thin.fits')
gaia = gaia[(gaia.phi1 > -45) & (gaia.phi1 < -25)]
stream_mask = gaia.gi_cmd_mask
g = gaia[(stream_mask)]# & (gaia.g_0 < 18)
dist = g.get_distance(min_parallax=1e-3*u.mas)
c = g.get_skycoord(distance=dist)
stream_coord = c.transform_to(gc.GD1)
phi1 = stream_coord.phi1.degree
phi2 = stream_coord.phi2.degree
pm1 = stream_coord.pm_phi1_cosphi2
pm2 = stream_coord.pm_phi2
after = GaiaData('../data/sorted_pm_member_prob_all_stars_8comp.fits')
after = after[(after.phi1 > -45) & (after.phi1 < -25)]
g_sorted, obs_pm_all, obs_pm_cov_all, phi1_stream_all, phi2_stream_all, bkg_ind = pmf.pre_model(gaia, g, after)
ln_bg_prob_all = after.pm_ln_bkg_prob.astype('float64')
ln_bg_prob = ln_bg_prob_all[bkg_ind]
# +
import importlib
importlib.reload(pmf)
with pm.Model() as model:
# mixture weight
alpha = pm.Uniform('alpha', lower = 0, upper = 1)
beta = pm.Uniform('beta', lower=0, upper = 1, testval=0.3)
loglike_fg_pm, loglike_fg_pm_all = pmf.short_pm_model_spur(model, obs_pm_all, obs_pm_cov_all,
phi1_stream_all, bkg_ind)
ll_fg_pm = pm.Deterministic('ll_fg_pm', tt.log(alpha) + loglike_fg_pm)
loglike_fg_phi2, loglike_fg_phi2_all = pmf.short_phi2_model_spur(model, phi1_stream_all,
phi2_stream_all, bkg_ind,)
loglike_fg_phi2 = loglike_fg_phi2.reshape(loglike_fg_pm.shape)
ll_fg_phi2 = pm.Deterministic('ll_fg_phi2', tt.log(beta) + loglike_fg_phi2)
loglike_fg_spur =pmf.short_spur_model(model, phi1_stream_all, phi2_stream_all,
obs_pm_all, obs_pm_cov_all, bkg_ind)
loglike_fg_spur = loglike_fg_spur.reshape(loglike_fg_pm.shape)
ll_fg_phi2_spur = pm.Deterministic('ll_fg_phi2_spur', tt.log(alpha) + tt.log(1-beta) + loglike_fg_spur)
#total track likelihood (including spur)
loglike_fg_phi2_total = pm.Deterministic('ll_fg_phi2_total', pm.logaddexp(loglike_fg_phi2, loglike_fg_spur))
#total foreground likelihood
loglike_fg = loglike_fg_pm + loglike_fg_phi2_total
ll_fg_full = pm.Deterministic('ll_fg_full', tt.log(alpha) + loglike_fg)
ll_bg_full = pm.Deterministic('ll_bg_full', tt.log(1 - alpha) + ln_bg_prob)
loglike = pm.logaddexp(ll_fg_full, ll_bg_full)
pm.Potential("loglike", loglike)
# -
with model:
res, logp = pmx.optimize(start={'b4': 0.45,
'std_phi2_spur': 0.15,
'beta': 0.3},
return_info = True)
# +
post_member_prob3 = np.exp(
res['ll_fg_full']
- np.logaddexp(res['ll_fg_full'], res['ll_bg_full']))
post_member_prob3_pm = np.exp(
res['ll_fg_pm']
- np.logaddexp(res['ll_fg_pm'], res['ll_bg_full']))
post_member_prob3_phi2 = np.exp(
res['ll_fg_phi2_total']
- np.logaddexp(res['ll_fg_phi2_total'], res['ll_bg_full']))
post_member_prob3_spur = np.exp(
res['ll_fg_phi2_spur']
- np.logaddexp(res['ll_fg_phi2_spur'], res['ll_bg_full']))
print('# among sel stars with total member prob > 0.1: {}'.format((post_member_prob3 > 0.1).sum()))
print('# among sel stars with PM member prob > 0.1: {}'.format((post_member_prob3_pm > 0.1).sum()))
print('# among sel stars with track member prob > 0.1: {}'.format((post_member_prob3_phi2 > 0.1).sum()))
# +
tbl = at.Table()
tbl['phi1'] = phi1_stream_all[bkg_ind]
tbl['phi2'] = phi2_stream_all[bkg_ind]
tbl['g_0'] = g_sorted.g_0[bkg_ind]
tbl['i_0'] = g_sorted.i_0[bkg_ind]
tbl['pm1'] = obs_pm_all[:,0][bkg_ind]
tbl['pm2'] = obs_pm_all[:,1][bkg_ind]
tbl['pm_cov'] = obs_pm_cov_all[bkg_ind]
tbl['ln_bg_prob'] = ln_bg_prob_all[bkg_ind]
tbl['post_member_prob'] = post_member_prob3
tbl['post_member_prob_pm'] = post_member_prob3_pm
tbl['post_member_prob_phi2'] = post_member_prob3_phi2
tbl['post_member_prob_spur'] = post_member_prob3_spur
tbl.write('../data/member_prob_3_spur.fits', overwrite=True)
# +
after3 = GaiaData('../data/member_prob_3_spur.fits')
high_memb_prob3_pm = after3[after3.post_member_prob_pm > 0.5]
high_memb_prob3_phi2 = after3[after3.post_member_prob_phi2 > 0.5]
high_memb_prob3_spur = after3[(after3.post_member_prob_spur > 0.5)]
high_memb_prob3 = after3[(after3.post_member_prob > 0.5)]
plt.figure(figsize=(10,3))
plt.scatter(high_memb_prob3_pm.phi1, high_memb_prob3_pm.phi2, c = high_memb_prob3_pm.post_member_prob_pm,
s = 5, cmap='plasma_r', vmax = 1)
plt.colorbar()
plt.xlim(-45, -25); plt.ylim(-5, 5);
plt.xlabel(r'$\phi_1$ [deg]'); plt.ylabel(r'$\phi_2$ [deg]')
plt.title(r'Proper Motion Memb Prob')
plt.figure(figsize=(10,3))
plt.scatter(high_memb_prob3_phi2.phi1, high_memb_prob3_phi2.phi2, c=high_memb_prob3_phi2.post_member_prob_phi2,
s = 5, cmap='plasma_r', vmin=0.5, vmax=1)
plt.colorbar()
plt.xlim(-45, -25); plt.ylim(-5, 5);
plt.xlabel(r'$\phi_1$ [deg]'); plt.ylabel(r'$\phi_2$ [deg]')
plt.title(r'Phi2 Membership Probability')
plt.figure(figsize=(10,3))
plt.scatter(high_memb_prob3.phi1, high_memb_prob3.phi2, c = high_memb_prob3.post_member_prob,
s = 5, cmap='plasma_r', vmax=1)
plt.colorbar()
plt.xlim(-45, -25); plt.ylim(-5, 5);
plt.xlabel(r'$\phi_1$ [deg]'); plt.ylabel(r'$\phi_2$ [deg]')
plt.title(r'Membership Probabilities Combined')
plt.figure(figsize=(10,3))
plt.scatter(high_memb_prob3_spur.phi1, high_memb_prob3_spur.phi2, c = high_memb_prob3_spur.post_member_prob,
s = 5, cmap='plasma_r', vmax=1)
plt.colorbar()
plt.xlim(-45, -25); plt.ylim(-5, 5);
plt.xlabel(r'$\phi_1$ [deg]'); plt.ylabel(r'$\phi_2$ [deg]')
plt.title(r'Membership Probabilities Combined')
#plt.savefig('../memb_probabilities_stream_with_spur.jpg')
# -
print(res['mean_pm_stream'], res['ln_std_pm_stream'])
print(res['mean_pm_spur'], res['ln_std_pm_spur'])
np.max(res['ll_fg_phi2_spur'])
plt.figure(figsize=(10,3))
plt.scatter(after3.phi1[res['ll_fg_phi2_spur']>-10], after3.phi2[res['ll_fg_phi2_spur']>-10],
c = res['ll_fg_phi2_spur'][res['ll_fg_phi2_spur']>-10],
s = 5, cmap='plasma_r', vmin=-10)
plt.colorbar()
plt.plot(np.linspace(-40, -25, 10), 0.45*np.sqrt(np.linspace(-40, -25, 10)+40))
plt.xlim(-45, -25); plt.ylim(-5, 5);
plt.xlabel(r'$\phi_1$ [deg]'); plt.ylabel(r'$\phi_2$ [deg]')
plt.title(r'Proper Motion Memb Prob')
# +
plt.figure(figsize = (10,7))
plt.scatter(high_memb_prob3.pm1, high_memb_prob3.pm2, c = high_memb_prob3.post_member_prob,
cmap='Blues',vmax=1, label = 'stream')
plt.colorbar(label = r'$\textrm{Membership} \: \textrm{Prob}$')
plt.scatter(high_memb_prob3_spur.pm1, high_memb_prob3_spur.pm2, c = high_memb_prob3_spur.post_member_prob_spur,
cmap='Reds', vmax=1, label = 'spur')
plt.colorbar()
plt.legend()
plt.xlim(-15, -11)
plt.ylim(-5,-1)
plt.xlabel(r'$\mu_{\phi_1}$')
plt.ylabel(r'$\mu_{\phi_2}$')
# -
print('sqrt scaling factor: {}'.format(res['spur_track_scale']))
print('width of spur: {}'.format(res['std_phi2_spur']))
|
notebooks/Spur_Fitting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="gtEoCCOIuhQj" outputId="5a528146-e1ba-4153-f604-5489a28163d8" colab={"base_uri": "https://localhost:8080/", "height": 539, "referenced_widgets": ["5668ce5993414626a6b87d0dd17ea095", "eedfba5d208e4351b5ed6fbf12121fe4", "e1753f80210a4058aae0687e43ea36c1", "<KEY>", "<KEY>", "b949858533d64398af3a7a7aa5ca12f0", "5d0758b9625244748978664c27909b82", "<KEY>", "37d14ca72e694c7294180e620b6aa117", "2384c584a2984c69ac6dede4034d6cca", "af9c020206df42649d788974e78284f5", "1d174cdddc2c4a82be1820cb215da117", "<KEY>", "<KEY>", "<KEY>", "29b262a78fe741c081ebaac798e196ec", "999d11eacfde466782795f39f20874c1", "6d33034875f34e3fae7e25218ed59a59", "42ccbe30c1804ed998a3779ef487afc1", "d56680a3dcc642608deddc46c7d3e8a6", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "60bf921dacdb40e49b24ef799db35c5a", "<KEY>", "d394642769db47fe8921ac20a7ad7362", "1fa89e6c45ae46b8a4e25a5916730849", "813a4eec80bc48dd9de3acedf0f1db02", "<KEY>", "<KEY>", "a09531a5db044978a43f1ab310b36aef"]}
from torchvision import datasets
import torch
data_folder = '../data/FMNIST' # This can be any directory you want to download FMNIST to
fmnist = datasets.FashionMNIST(data_folder, download=True, train=True)
# + id="-GzkCv0PuiMj"
tr_images = fmnist.data
tr_targets = fmnist.targets
# + id="oajhutCkujd1"
val_fmnist = datasets.FashionMNIST(data_folder, download=True, train=False)
val_images = val_fmnist.data
val_targets = val_fmnist.targets
# + id="uk8CcQkAulye"
from imgaug import augmenters as iaa
aug = iaa.Sequential([
iaa.Affine(translate_px={'x':(-10,10)}, mode='constant'),
])
# + id="PGNQZrRWw58o" colab={"base_uri": "https://localhost:8080/"} outputId="8f216c7b-f958-4557-e540-a8a72062b832"
# %%time
for i in range(32):
aug.augment_image(tr_images[i])
# + id="Tf6gYqrGxKkP" colab={"base_uri": "https://localhost:8080/"} outputId="6f035d96-1054-454a-e6da-cc1337f4b288"
# %%time
x = aug.augment_images(tr_images[:32])
# + id="ToJpsyQmuz1S"
|
Chapter04/Time_comparison_of_augmentation_scenario.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Semantische Segmentierung mittels U-Net / ResNet
# Jetzt seid ihr dran. Wir wollen nun ein U-Net oder ResNet nutzen, um eine semantische Segmentierung der Daten vorzunehmen. Den Code müsst ihr selbst schreiben oder zusammen kopieren. Wir gehen wieder wie in den anderen Notebooks vor...
# ## Imports und env setzen
# ## Daten
# ### Daten laden
# ### Labels in semantische label übertragen
# Zeigt den Unterschied der Labels zwischen *semantic* und *instance segmentation* mit einem Plot.
# ### Validation Set erstellen
# ### Daten Preprocessing
# #### Reskalieren und / oder normalisieren
# #### Was ist der Output eures Netzwerks?
# ## Die Netzwerke
# Erstellt jetzt ein U-Net oder ResNet. Wie ihr das macht ist freigestellt. Wenn ihr die Architektur dynamisch durch Variablen verändern könnt, könnt ihr ganz einfach verschiedene Netzwerke erstellen, trainieren und vergleichen.
# ### Idee:
# Kann das Netzwerk angeben, wie sicher es sich ist, dass eine Zelle vorhanden ist?
# ## Training
# ## Trainingshistories plotten
# ### Unterschiede der Netzwerke?
# ## Evaluation mit Test-Set
# ### Was sind die Metriken
# ### Wie sieht die Prediction aus
|
05_unet_semantic_segmentation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import seldon_mlmd_tools as mlmd_tools
print("version:", mlmd_tools.__version__)
print("file:", mlmd_tools.__file__)
# -
# # Prepare Model
from example_data import model_schema as model_example_dict
model = mlmd_tools.schemas.Model.parse_obj(model_example_dict)
[type(x) for x in model.input_schema.features]
from pprint import pprint; pprint(model.dict())
# # Put Model into Store
mlmd_store = mlmd_tools.connect_and_initialize("localhost:8080")
artifact = mlmd_tools.save_seldon_model_to_store(mlmd_store, model)
artifact.id
|
notebooks/put model into mlmd.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Get Techniques from Data Sources
# ## Import Libraries
# +
from attackcti import attack_client
import logging
logging.getLogger('taxii2client').setLevel(logging.CRITICAL)
# -
# ## Initialize Client
lift = attack_client()
# ## Retrieve Techniques
techniques = lift.get_techniques_by_data_sources(
"Network Traffic",
"Process"
)
len(techniques)
techniques[0]
techniques[0]['x_mitre_data_sources']
|
docs/playground/5-Collect_Techniques_by_Data_Sources.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# <h1 style="color:blue" align="center">Crosstab Tutorial</h1>
import pandas as pd
df = pd.read_excel("survey.xls")
df
pd.crosstab(df.Nationality,df.Handedness)
pd.crosstab(df.Sex,df.Handedness)
# <h2 style="color:purple">Margins</h2>
pd.crosstab(df.Sex,df.Handedness, margins=True)
# <h2 style="color:purple">Multi Index Column and Rows</h2>
pd.crosstab(df.Sex, [df.Handedness,df.Nationality], margins=True)
pd.crosstab([df.Nationality, df.Sex], [df.Handedness], margins=True)
# <h2 style="color:purple">Normalize</h2>
pd.crosstab(df.Sex, df.Handedness, normalize='index')
# <h2 style="color:purple">Aggfunc and Values</h2>
import numpy as np
pd.crosstab(df.Sex, df.Handedness, values=df.Age, aggfunc=np.average)
|
Program's_Contributed_By_Contributors/AI-Summer-Course/py-master/pandas/13_crosstab/pandas_crosstab.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="nCc3XZEyG3XV"
# Lambda School Data Science
#
# *Unit 2, Sprint 3, Module 1*
#
# ---
#
#
# # Define ML problems
#
# You will use your portfolio project dataset for all assignments this sprint.
#
# ## Assignment
#
# Complete these tasks for your project, and document your decisions.
#
# - [x] Choose your target. Which column in your tabular dataset will you predict?
# - [x] Choose which observations you will use to train, validate, and test your model. And which observations, if any, to exclude.
# - [x] Determine whether your problem is regression or classification.
# - [ ] Choose your evaluation metric.
# - [ ] Begin to clean and explore your data.
# - [ ] Begin to choose which features, if any, to exclude. Would some features "leak" information from the future?
#
# -
import pandas as pd
import numpy as np
import pickle
import pandas_profiling
# +
#import PKL Dataset ready for explortation
infile = open("/Users/user/Documents/GitHub/MIR/Spotify/master_list2.pkl",'rb')
df = pickle.load(infile)
# -
# The target I am choosing for my dataset is "not_skipped" which has a 34% recurrence (below the recommended 70%).
#
# **Keeping feature for indexing**
#
# * track_id
#
# **Target:**
#
# * not_skipped
#
# **Observations:**
#
# * session_position
# * session_length
# * skip_1
# * skip_2
# * skip_3
# * context_switch
# * no_pause_before_play
# * short_pause_before_play
# * long_pause_before_play
# * hist_user_behavior_n_seekfwd
# * hist_user_behavior_n_seekback
# * hist_user_behavior_is_shuffle
# * hour_of_day
# * context_type
# * hist_user_behavior_reason_start
# * hist_user_behavior_reason_end
# * duration
# * release_year
# * us_popularity_estimate
# * acousticness
# * beat_strength
# * bounciness
# * danceability
# * dyn_range_mean
# * energy
# * flatness
# * instrumentalness
# * key
# * liveness
# * loudness
# * mechanism
# * mode
# * organism
# * speechiness
# * tempo
# * time_signature
# * valence
profile = df.profile_report(title='Pandas Profiling Report')
profile.to_file(output_file="Spotify_data_profile.html")
|
module1/Zhenya_Warshavsky_assignment_applied_modeling_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Use a Symbolic Link
#
# The file [mymodule.py](mymodule.py) in the current directory is a symbolic link to [../module-subdirectory/mymodule.py](../module-subdirectory/mymodule.py).
#
# This may not work on all operating systems!
import mymodule
mymodule.hello()
|
importing-local-python-modules-from-jupyter-notebooks/symbolic-link/symlink.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Linear Regression Model
#
# We will proceed initially to use the last 3 years to model the data with a simple linear regression.
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression, Ridge
from sklearn.preprocessing import PolynomialFeatures
from sklearn.pipeline import Pipeline
from collections import defaultdict
import re
# +
year_from = 2005
def load_data(year_from):
assert isinstance(year_from,int)
# Imports
train = pd.read_csv('data/TrainingSet.csv',index_col=0)
submission = pd.read_csv('data/SubmissionRows.csv',index_col=0)
# Remove [YR****] and input '_' for last 3 cols
train.columns = list(map(lambda x: re.findall(r'\d+',x)[0],train.columns[:36])) + list(map(lambda x: '_'.join(x.split()),train.columns[36:]))
# Use last 3 years for predictions: This is subjected to change
train = train.loc[:,f'{year_from}':]
return train, submission
# -
# Load data:
train, submission = load_data(2005)
def submission_data_missing(train,submission):
mask = train.loc[submission.index,].T.isna().sum()>0
train_X = train.loc[submission.index,]
return train_X.loc[mask]
# 16 rows with missing values: Interpolate these values
submission_data_missing(train,submission)
def interpolate_data(train,submission):
train_X = train.loc[submission.index,:]
# Interpolate: Both directions are bfilled and ffilled respectively
train_X.loc[:,:'2007'] = train_X.loc[:,:'2007'].interpolate(limit_direction='both',axis=1)
return train_X
# +
# Data:
data = interpolate_data(train,submission)
# Func to split that dataframe to values [2005,2006,2007] and [country_name,series_code,series_name]
def split_dataframe(data):
raw_data = data.loc[:,:'2007']
description = data.loc[:,'Country_Name':]
return raw_data,description
# -
# Split:
raw_data,description = split_dataframe(data)
# Export CSV:
raw_data.to_csv('data/raw_data.csv')
description.to_csv('data/description.csv')
def format_dataframe(raw_data):
"""
Quick utility function to format dataframe into a X,y format with X = Numbers of years from the start, y = values.
Example:
Initial Dataframe being:
2005 0.4
2006 0.6
2007 0.8
The function transforms it into
X y
0 0.4
1 0.6
2 0.8
Returns: X,y
Note: If we have 10 different timeseries (features) X.shape = (n_years,n_features) so slicing will be needed to predict
individually.
"""
# Extract index from raw data before transforming:
raw_data_index = list(raw_data.index)
raw_data.columns = raw_data.columns.astype('int')
# Transponse to have time as index instead of columns
raw_data = raw_data.T
X = np.asarray(raw_data.index - raw_data.index[0]).reshape(-1,1)
y = raw_data.values
return X,y,raw_data_index
X,y,raw_data_index = format_dataframe(raw_data)
def linear_regression_poly(X,y,degree,year):
assert isinstance(X,np.ndarray)
assert isinstance(y,np.ndarray)
assert isinstance(degree,int) and degree > 0
assert year >= 2005
year_pred = np.array([[year % year_from]])
pipe = Pipeline([('poly',PolynomialFeatures(degree=degree)),('linear',LinearRegression())])
n_features = y.shape[1]
predictions_year = defaultdict(list)
# Fit:
for i in range(n_features):
# slice each series:
y_i = y[:,i]
pipe.fit(X,y_i)
# prediction value for year specified
y_pred = pipe.predict(year_pred)[0]
predictions_year[f'{year}[YR{year}]'].append(y_pred)
# To dataframe: with correctly indexed submission values
df = pd.DataFrame(predictions_year,index=raw_data_index)
return df
# +
# Predictions: Polynomial degree = 1:
_2008 = linear_regression_poly(X,y,1,2008).values
_2012 = linear_regression_poly(X,y,1,2012).values
# Into submission:
submission['2008 [YR2008]'] = _2008
submission['2012 [YR2012]'] = _2012
submission.head()
# +
# Index mapper for plotting:
index_mapper = {k:index for index,k in enumerate(raw_data_index)}
# Plotting:
def plot_predictions(raw_data,description,pred_2008,pred_2012,n_features):
pred_2008_values = pred_2008
pred_2012_values = pred_2012
pred_concat = np.concatenate([pred_2008_values,pred_2012_values],axis=1)
random_indicators = np.random.choice(raw_data_index,size=n_features)
# plot:
fig,ax = plt.subplots(1,1,figsize=(14,8))
ax.set_title(f'Values from {year_from} including pred: 2008, 2012')
for i in random_indicators:
values = raw_data.T.loc[:,i].values
ax.plot([str(x) for x in range(year_from,2008)],values,label=f'{description.Country_Name[i]} : {description.Series_Name[i]}',marker='o')
ax.plot(['2008','2012'],pred_concat[index_mapper[i],:],linestyle='--', label=f'{description.Country_Name[i]} : {description.Series_Name[i]} PRED',marker='o')
ax.legend()
# -
# Plot:
np.random.seed(2)
plot_predictions(raw_data,description,_2008,_2012,5)
# +
# Predictions: Polynomial degree = 2:
_2008 = linear_regression_poly(X,y,2,2008).values
_2012 = linear_regression_poly(X,y,2,2012).values
# Into submission:
submission['2008 [YR2008]'] = _2008
submission['2012 [YR2012]'] = _2012
submission.head()
# -
# Plot: degree = 2
np.random.seed(2)
plot_predictions(raw_data,description,_2008,_2012,5)
# ### Note:
#
# Values from linear regression with degree = 1 scored higher (RMSE = 0.0549). Degree = 2 scored RMSE = 0.2483.
# ### Find correlated indicators
# +
train, submission = load_data(1973)
# +
kenya_data = train[train.Country_Name == 'Kenya']
kenya_values = kenya_data[[str(element) for element in range(2003,2007)]].values
# get the total number of time series we have for Kenya
nseries = kenya_values.shape[0]
# -1 as default
lag_corr_mat = np.ones([nseries, nseries], dtype=np.float64)*-1
# create a matrix to hold our lagged correlations
for i in range(nseries):
for j in range(nseries):
# skip comparing a series with itself
if i!=j:
# get original (1972-2006) and shifted (1973-2007)
original = kenya_values[i,1:]
shifted = kenya_values[j,:-1]
# for just the indices where neither is nan
non_nan_mask = (~np.isnan(original) & ~np.isnan(shifted))
# if we have at least 2 data points
if non_nan_mask.sum() >= 2:
lag_corr_mat[i,j] = np.correlate(original[non_nan_mask], shifted[non_nan_mask])
# +
# let's look at one of the indicators we are suppoed to predict
to_predict_ix = 131042
# first, we get the index of that row in the correlation matrix
i = np.where(kenya_data.index.values == to_predict_ix)[0][0]
# then, we see which value in the matrix is the largest for that row
j_max = np.argmax(lag_corr_mat[i,:])
# finally, let's see what these correspond to
max_corr_ix = kenya_data.index.values[j_max]
# # now write out what we've found
# fmt_string = "In Kenya, the progress of '{}' is "\
# "most correlated with a change in '{}' during the year before."
# print(kenya_data["Series Name"][to_predict_ix], kenya_data["Series Name"][max_corr_ix])
print(kenya_data.loc[to_predict_ix])
print()
print(kenya_data.loc[max_corr_ix])
# +
from sklearn.preprocessing import MinMaxScaler
correlated_indicators=train[train.index.isin([to_predict_ix,max_corr_ix])].loc[:,'2000':'2007'].T
pd.DataFrame(
MinMaxScaler().fit_transform(correlated_indicators),
columns=correlated_indicators.columns,
index=correlated_indicators.index).plot()
# -
df=pd.DataFrame(data=lag_corr_mat[0:,0:],
index=kenya_data.index.values,
columns=kenya_data.index.values)
df.head()
# df.loc[to_predict_ix,max_corr_ix]
top_10_indicators=np.abs(df.loc[to_predict_ix,:]).sort_values(ascending=False).iloc[0:10]
top_10_indicators_and_to_predict = np.append(to_predict_ix, top_10_indicators.index.values)
# +
correlated_indicators=train[train.index.isin(top_10_indicators_and_to_predict)].loc[:,'2000':'2007'].T
pd.DataFrame(
MinMaxScaler().fit_transform(correlated_indicators),
columns=correlated_indicators.columns,
index=correlated_indicators.index).plot(figsize=(20,10))
# +
less_correlated_indicators=np.abs(df.loc[to_predict_ix,:]).sort_values(ascending=True).iloc[50:60].index.values
less_correlated_indicators=np.append(to_predict_ix, less_correlated_indicators)
less_correlated_indicators=train[train.index.isin(less_correlated_indicators)].loc[:,'2000':'2007'].T
pd.DataFrame(
MinMaxScaler().fit_transform(less_correlated_indicators),
columns=less_correlated_indicators.columns,
index=less_correlated_indicators.index).plot(figsize=(20,10))
# +
non_correlated_indicators=np.abs(df.loc[to_predict_ix,:]).sort_values(ascending=True).iloc[0:10].index.values
non_correlated_indicators=np.append(to_predict_ix, non_correlated_indicators)
non_correlated_indicators=train[train.index.isin(non_correlated_indicators)].loc[:,'2000':'2007'].T
pd.DataFrame(
MinMaxScaler().fit_transform(non_correlated_indicators),
columns=non_correlated_indicators.columns,
index=non_correlated_indicators.index).plot(figsize=(20,10))
# -
kenya_data.loc[correlated_indicators.columns]
# kenya_data.loc[to_predict_ix]
|
Linear_Regression_Model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ontol]
# language: python
# name: conda-env-ontol-py
# ---
# # Introduction
#
# This notebook compares the test set performance of classifiers using term list scores based on the data-driven, RDoC, and DSM frameworks.
#
# # Load the data
# +
import pandas as pd
import numpy as np
np.random.seed(42)
import sys
sys.path.append("..")
import utilities
# -
# ## Input parameters
metric = "proto" # Variable being compared between frameworks
n_iter = 1000 # Number of iterations for bootstrap distributions
frameworks = ["data-driven_k22", "data-driven", "rdoc", "dsm"]
clfs = ["_lr", "_lr", "", ""]
list_suffixes = ["", "", "_opsim", "_opsim"]
splits = ["discovery", "replication"]
suffix = "lr"
# ## Framework contents
lists, circuits = {}, {}
for fw, clf, list_suffix in zip(frameworks, clfs, list_suffixes):
lists[fw], circuits[fw] = utilities.load_framework(fw, suffix=list_suffix, clf=clf)
# ## Observed values
obs = {}
for fw, clf in zip(frameworks, clfs):
obs[fw] = pd.read_csv("data/{}_obs_{}{}.csv".format(metric, fw, clf), index_col=0, header=0)
obs["data-driven_k22"].loc[obs["data-driven_k22"]["SPLIT"] == "discovery"]["OBSERVED"].mean()
obs["data-driven_k22"].loc[obs["data-driven_k22"]["SPLIT"] == "replication"]["OBSERVED"].mean()
# ## Bootstrap distributions
boot, mean = {}, {}
for split in splits:
boot[split], mean[split] = {}, {}
for fw, clf in zip(frameworks, clfs):
boot[split][fw] = pd.read_csv("data/{}_boot_{}{}_{}_{}iter.csv".format(metric, fw, clf, split, n_iter),
index_col=0, header=0)
mean[split][fw] = pd.read_csv("data/{}_mean_{}{}_{}.csv".format(metric, fw, clf, split),
index_col=0, header=0)
# # Perform comparison tests
#
# Is the mean of fw<sub>i</sub> greater than the mean of fw<sub>j</sub>?
from statsmodels.stats.multitest import multipletests
p, fdr = {}, {}
for split in splits:
p[split] = np.empty((len(frameworks), len(frameworks)))
for i, fw_i in enumerate(frameworks):
for j, fw_j in enumerate(frameworks):
boot_i = np.mean(boot[split][fw_i], axis=0)
boot_j = np.mean(boot[split][fw_j], axis=0)
p[split][i,j] = np.sum((boot_i - boot_j) <= 0.0) / float(n_iter)
fdr[split] = multipletests(p[split].ravel(), method="fdr_bh")[1].reshape(p[split].shape)
fdr[split] = pd.DataFrame(fdr[split], index=frameworks, columns=frameworks)
print("-" * 65 + "\n" + split.upper() + "\n" + "-" * 65)
print(fdr[split])
print("")
# +
interval = 0.999
tail = (1.0 - interval) / 2.0 # Two-sided test
idx_lower = int(n_iter * tail)
idx_upper = int(n_iter * (1.0 - tail))
CI = {}
for split in splits:
CI[split] = {}
CI[split] = pd.DataFrame("", index=frameworks, columns=frameworks)
for i, fw_i in enumerate(frameworks):
for j, fw_j in enumerate(frameworks):
boot_i = np.mean(boot[split][fw_i], axis=0)
boot_j = np.mean(boot[split][fw_j], axis=0)
boot_dif = np.sort(boot_i - boot_j)
CI_lower = boot_dif[idx_lower]
CI_upper = boot_dif[idx_upper]
CI_sig = ""
if (CI_lower > 0 and CI_upper > 0) or (CI_lower < 0 and CI_upper < 0):
CI_sig = "*"
CI[split].loc[fw_i,fw_j] = "{:.3f}, {:.3f} {}".format(CI_lower, CI_upper, CI_sig)
print("-" * 65 + "\n" + split.upper() + "\n" + "-" * 65)
print(CI[split])
print("")
# -
# # Plot evaluation metrics
# %matplotlib inline
for split in splits:
utilities.plot_framework_comparison(boot[split], obs, mean[split], metric=metric, w=1.2,
n_iter=n_iter, suffix="k22_" + suffix + "_" + split,
ylim=[-0.25,1], yticks=[-0.25,0,0.25,0.5,0.75,1])
|
prototype/comp_frameworks_lr_k22.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import random
import math
import itertools
from copy import deepcopy
def CliffordUpdate(psi,gate,qubits,n):
"""psi is a stabilizer state in CH form
gate is 'S','CZ','CX',or 'H'
qubits=[q] or qubits=[q1 q2] are the qubits the gate acts on
n is the total number of qubits
psi is a dict containing np.arrays/integers {n,s,r,Uc and p}"""
if psi['n']!=n:
raise AssertionError("Inconsistent number of qubits, n ", n, psi['n'])
else:
if gate=='S':
psi['Uc']=LeftMultS(psi['Uc'],qubits,n)
elif gate=='CZ':
psi['Uc']=LeftMultCZ(psi['Uc'],qubits[0],qubits[1],n)
elif gate=='CX':
psi['Uc']=LeftMultCNOT(psi['Uc'],qubits[0],qubits[1],n)
elif gate=='X':
P=psi['Uc'][qubits[0],:].copy() #Pauli U_c^{\dagger} X_j U_c
# Now conjugate by H(r)
numy=np.mod(np.count_nonzero(P[0:n]*P[n:2*n]*psi['r']),2)#Count number of times Y-->-Y when applying H(r)
P[2*n]=np.mod(P[2*n]+2*numy,4)#adjust phase accordingly
inds=np.nonzero(psi['r'])[0] #Apply H(r): swap X and Z at appropriate indices
for j in inds:
holder=P[j].copy()
P[j]=P[n+j].copy()
P[n+j]=holder
# Now compute P|s>
v1=P[0:n]
w1=P[n:2*n]
alpha=np.mod(P[2*n]+np.sum(v1*w1*(1+2*psi['s']))+2*np.sum((1-v1)*w1*psi['s']),4)
psi['p']=np.mod(psi['p']+2*alpha,8)
psi['s']=np.mod(psi['s']+P[0:n],2)
if gate=='H':
#Takes a stabilizer state psi in CH form applies the single-qubit H gate to the
#specified qubit
#Define Paulis
# P1=H(r) U_c ^{\dagger} X_qubit U_c H(r)
# = [v1 | w1 | b1] (v1 is n bits, w1 is n bits, and b1 in the set 0,1,2,3)
# P2=H(r) U_c ^{\dagger} Z_qubit U_c H(r)
# = [v2 | w2 | b2]
# Then the state after applying H is
# |phi>= w^p U_c H(r) (P1|s>+P2|s>)/sqrt(2)
qubit=qubits[0]
P1=psi['Uc'][qubit,:].copy()
numy=np.mod(np.count_nonzero(P1[0:n]*P1[n:2*n]*psi['r']),2)
P1[2*n]=np.mod(P1[2*n]+2*numy,4)
P2=psi['Uc'][n+qubit,:].copy()
numy2=np.mod(np.count_nonzero(P2[0:n]*P2[n:2*n]*psi['r']),2)
P2[2*n]=np.mod(P2[2*n]+2*numy2,4)
inds=np.nonzero(psi['r'])[0] #Apply H(r): swap X and Z at appropriate indices
for j in inds:
holder=P1[j].copy()
P1[j]=P1[n+j].copy()
P1[n+j]=holder
holder=P2[j].copy()
P2[j]=P2[n+j].copy()
P2[n+j]=holder
b1=P1[2*n]
v1=P1[0:n]
w1=P1[n:2*n]
b2=P2[2*n]
v2=P2[0:n]
w2=P2[n:2*n]
# print(type(b1))
# print(type(v1))
# print(type(w1))
# print(type(psi['s']))
alpha=np.mod(b1+np.sum(v1*w1*(1+2*psi['s']))+2*np.sum((1-v1)*w1*psi['s']),4)
beta=np.mod(b2+np.sum(v2*w2*(1+2*psi['s']))+2*np.sum((1-v2)*w2*psi['s']),4)
s1=np.mod(psi['s']+v1,2)
# print('s1=',s1)
s2=np.mod(psi['s']+v2,2)
# print('s2=',s2)
a=np.mod(beta-alpha,4)
# print('a=',a)
y=np.mod(s1+s2,2)
# print(y)
# print("=y")
if np.count_nonzero(y)==0:
if a==1:
psi['p']=np.mod(psi['p']+2*alpha+1,8)
else:
psi['p']=np.mod(psi['p']+2*alpha-1,8)
psi['s']=s1
if np.count_nonzero(y)>0:
#Two cases
klist=np.nonzero(np.mod(psi['r']+1,2)*y)[0]#Indices where r_k=0 and y_k=1
# print(klist)
# print("=klist")
mlist=np.nonzero(psi['r']*y)[0]#Indices where r_k=1 and y_k=1
# print(mlist)
# print("=mlist")
if klist.size!=0:# if klist is nonempty
k=klist[0]
psi['p']=np.mod(psi['p']+2*alpha+2*s1[k]*a,8)
psi['r'][k]=1
if s1[k]==0:
psi['s']=s1.copy()
else:
psi['s']=s2.copy()
for j in klist[1::]: #All except k=klist[0]
psi['Uc']=RightMultCNOT(psi['Uc'],k,j,n)
for j in mlist:
psi['Uc']=RightMultCZ(psi['Uc'],k,j,n)
if s1[k]==0:
# print("a=")
# print(a)
for j in range(1,a+1):
# print("j=")
# print(j)
psi['Uc']=RightMultS(psi['Uc'],k,n)
else:
a2=np.mod(-a,4)
# print("a2=")
# print(a2)
for j in range(1,a2+1):
# print("j=")
# print(j)
psi['Uc']=RightMultS(psi['Uc'],k,n)
# print("psi['Uc']=")
# print(psi['Uc'])
if klist.size==0:# if klist is empty
k=mlist[0]
# Update phase p
if a==1:
psi['p']=np.mod(psi['p']+2*alpha+1,8)
if a==3:
psi['p']=np.mod(psi['p']+2*alpha-1,8)
if (a==0 or a==2):
psi['p']=np.mod(psi['p']+2*alpha,8)
B=np.nonzero(y)[0]#0-indexed in python
# Update Uc
if s1[k]==1:
psi['Uc']=RightMultS(psi['Uc'],k,n)#Apply Z_k gate
psi['Uc']=RightMultS(psi['Uc'],k,n)
for j in B:
# print('B=',B)
# print('j=',j)
# print('k=',k)
if j != k:
psi['Uc']=RightMultCNOT(psi['Uc'],j,k,n)
if (a==1 or a==3):
psi['Uc']=RightMultS(psi['Uc'],k,n)
# Update r
# print('r_in=',psi['r'])
# print('k=',k)
if (a==0 or a==2):
# print('r_to_update=',psi['r'])
# print('k-th=',[k])
# print('update r')
psi['r'][k]=np.mod(psi['r'][k]+1,2)
# print('r_out=',psi['r'])
# Update s
psi['s']=s1
if (s1[k]==1 and (a==1 or a==2) or (s1[k]==0 and (a==0 or a==3))):
pass
else:
psi['s'][k]=np.mod(psi['s'][k]+1,2)
return psi
# +
def AmplitudeUpdates(Pauli, psi, j, n):
"""Compute amplitude
amp=w^p <0|U_c^{\dagger} X_j U_c Pauli*H(r)|s> where j is a bit index
Here eps=0,1
m=-n,...0
p =0, 1,2, ...,7
w=exp(i*pi/4)
Note <x|psi>=<0|U_c^{\dagger} X(x) U_c H(r)|s>w^p
X=[x zeros(1,n) 0];
P=PauliImage(X,psi.Uc,n);"""
P = MultiplyPauli(psi['Uc'][j,:], Pauli, n)
v1 = P[0:n]
eps = 1
if np.count_nonzero(np.mod(v1*(1-psi['r'])+psi['s']*(1-psi['r']),2)) > 0:
#v1 and s should only differ where r is 1
eps = 0
m = 0
p = 0
amp = 0
else:
p = np.mod(psi['p']-2*np.count_nonzero(P[n:2*n]*P[0:n])+2*P[2*n], 8)
#Number of Y paulis in P is nnz(P(n+1:2*n).*P(1:n)), each contributes -i
m = np.count_nonzero(psi['r'])
# Now compute sign(<v1|H(r)|s>)
p = np.mod(p+4*np.mod(np.sum(v1*psi['s']*psi['r']), 2), 8)
#Look at places where r=v1=s=1. Each contributes a minus sign
amp = 2**(-m/2)*eps*np.exp(1j*p*np.pi/4)
return amp
# -
def Amplitude(x, psi, n):
"""Compute amplitude amp=<x|psi>=2^(-m/2)*eps* w^{p) where x is an n-bit string and psi is a
stabilizer state in CH form
Here eps=0,1
m=-n,...0
p =0, 1,2, ...,7
w=exp(i*pi/4)
Note <x|psi>=<0|U_c^{\dagger} X(x) U_c H(r)|s>w^p"""
X = np.hstack((x,np.zeros(n,dtype=int),0))
P = PauliImage(X, psi['Uc'], n)
v1 = P[0:n]
eps = 1
if np.count_nonzero(np.mod(v1*(1-psi['r'])+psi['s']*(1-psi['r']),2)) > 0:
#v1 and s should only differ where r is 1
eps = 0
m = 0
p = 0
amp = 0
else:
p = np.mod(psi['p']-2*np.count_nonzero(P[n:2*n]*P[0:n])+2*P[2*n], 8)
#Number of Y paulis in P is nnz(P(n+1:2*n).*P(1:n)), each contributes -i
m = np.count_nonzero(psi['r'])
# Now compute sign(<v1|H(r)|s>)
p = np.mod(p+4*np.mod(np.sum(v1*psi['s']*psi['r']), 2), 8)
#Look at places where r=v1=s=1. Each contributes a minus sign
amp = 2**(-m/2)*eps*np.exp(1j*p*np.pi/4)
return amp
def CompBasisVector(z):
"""constructs a computational basis vector in C-H form
|psi>= w^p U_c H(r)|s>
where
s is a computational basis vector
H(r) is hadamards applied to qubits in r
U_c is a Clifford such that U_c|0>=|0> (stored by its tableau).
tableau format: first n columns are U^{\dagger} X_j U plus phase bit, last
n columns are U^{\dagger} Z_j U plus phase bit
p is an integer, w=exp(i\pi/4).
Here z vector of length n, and basis is either 0 (z basis) or 1 (x basis)"""
if np.ndim(z)!=1:
z=np.squeeze(z) #Make z into a 1-d array by removing singletons
n = np.size(z)
psi={
'n':n,
's':np.asarray(z),
#'s':z,
'r':np.zeros(n,dtype=int),
'Uc':np.concatenate((np.eye(2*n,dtype=int), np.zeros([2*n,1],dtype=int)), axis=1),
'p':0,
'c':np.exp(0*1j*np.pi)
}
return psi
def XBasisVector(z):
"""constructs an X-basis vector in C-H form
|psi>= w^p U_c H(r)|s>
where
s is a computational basis vector
H(r) is hadamards applied to qubits in r
U_c is a Clifford such that U_c|0>=|0> (stored by its tableau).
tableau format: first n columns are U^{\dagger} X_j U plus phase bit, last
n columns are U^{\dagger} Z_j U plus phase bit
p is an integer, w=exp(i\pi/4).
Here z vector of length n, and basis is either 0 (z basis) or 1 (x basis)"""
if np.ndim(z)!=1:
z=np.squeeze(z) #Make z into a 1-d array by removing singletons
n = np.size(z)
psi={
'n':n,
's':np.asarray(z),
#'s':z,
'r':np.ones(n,dtype=int),
'Uc':np.concatenate((np.eye(2*n,dtype=int), np.zeros([2*n,1],dtype=int)), axis=1),
'p':0,
'c':np.exp(0*1j*np.pi)
}
return psi
def RightMultCNOT(tableau, control, target, n):
"""Takes a tableau for a Clifford U and outputs the tableau for
U*CNOT_{control,target}
XI --> XX
YI --> YX
IZ --> ZZ
IY--> ZY
XX --> XI
ZZ--> IZ
YY--> -XZ
XY--> YZ
YX--> YI
XZ--> -YY
YZ--> XY
ZY-->IY
tableau format: first n columns are U^{\dagger} X_j U plus phase bit,
last n columns are U^{\dagger} Z_j U plus phase bit"""
#Look at the columns of the tableau corresponding to qubits control/target
for j in range(2*n):
a = np.array([tableau[j, control], tableau[j, control + n], tableau[j, target], tableau[j, target + n]],dtype=int)
b = 1*a[0] + 2*a[1] + 4*a[2] + 8*a[3]
if b == 1: # [1 0 0 0] % XI-->XX
tableau[j, target] = 1
elif b == 3: #[1 1 0 0] % YI-->YX
tableau[j, target] = 1
elif b == 8: # [0 0 0 1] %IZ-->ZZ
tableau[j, control + n] = 1
elif b == 12: #[0 0 1 1] %IY-->ZY
tableau[j, control + n] = 1
elif b == 5: # [1 0 1 0] % XX-->XI
tableau[j, target] = 0
elif b == 10: #[0 1 0 1] %ZZ-->IZ
tableau[j, control + n] = 0
elif b == 15: #[1 1 1 1] %YY-->-XZ
tableau[j, control + n] = 0
tableau[j, target] = 0
tableau[j, -1] = np.mod(tableau[j,-1] + 2, 4)
elif b == 13: #[1 0 1 1] %XY-->YZ
tableau[j, control + n] = 1
tableau[j, target] = 0
elif b == 7: # [1 1 1 0] %YX-->YI
tableau[j, target] = 0
elif b == 9: # [1 0 0 1] %XZ-->-YY
tableau[j, control + n] = 1
tableau[j, target] = 1
tableau[j, -1] = np.mod(tableau[j, -1] + 2, 4)
elif b == 11: #[1 1 0 1] %YZ-->XY
tableau[j, control + n] = 0
tableau[j, target] = 1
elif b == 14: #[0 1 1 1] %ZY-->IY
tableau[j, control + n] = 0
else:
pass
return tableau
def RightMultCZ(tableau, control, target, n):
""" Takes a tableau for a Clifford U and outputs the tableau for
U*CZ_{control,target}
XI --> XZ
YI --> YZ
IY--> ZY
IX--> ZX
XX --> YY
YY--> XX
XY--> -YX
YX--> -XY
XZ--> XI
ZX-->IX
YZ--> YI
ZY-->IY
"""
#Look at the columns of the tableau corresponding to qubits control/target
for j in range(2*n):
a = np.array([tableau[j, control], tableau[j, control + n], tableau[j, target], tableau[j, target + n]],dtype=int)
b = 1*a[0] + 2*a[1] + 4*a[2] + 8*a[3]
if b == 1: # [1 0 0 0] % XI-->XZ
tableau[j,n + target] = 1
elif b == 3: #[1 1 0 0] % YI-->YZ
tableau[j,n + target] = 1
elif b == 12: # [0 0 1 1] %IY-->ZY
tableau[j, control + n] = 1
elif b == 4: #[0 0 1 0] %IX-->ZX
tableau[j, control + n] = 1
elif b == 5: # [1 0 1 0] % XX--> YY
tableau[j, control + n] = 1
tableau[j, target + n] = 1
elif b == 15: #[1 1 1 1] %YY-->XX
tableau[j, control + n] = 0
tableau[j, target + n] = 0
elif b == 13: #[1 0 1 1] %XY-->-YX
tableau[j, control + n] = 1
tableau[j, target + n] = 0
tableau[j, -1] = np.mod(tableau[j, -1] + 2, 4)
elif b == 7: #[1 1 1 0] %YX-->-XY
tableau[j, control + n] = 0
tableau[j, target + n] = 1
tableau[j, -1] = np.mod(tableau[j, -1] + 2, 4)
elif b == 9: #[1 0 0 1] %XZ-->XI
tableau[j, target + n] = 0
elif b == 6: #[0 1 1 0] %ZX-->IX
tableau[j, control + n] = 0
elif b == 11: #[1 1 0 1] %YZ-->YI
tableau[j, target + n] = 0
elif b == 14: #[0 1 1 1] %ZY-->IY
tableau[j, control + n] = 0
else:
pass
return tableau
def LeftMultCNOT(tableau, control, target, n):
"""Takes a tableau for a Clifford U and outputs the tableau for
CNOT_{control,target}*U
X_control --> X_control X_target
Z_target --> Z_control Z_target"""
newtableau = tableau.copy()
Xprod = np.zeros(2 * n + 1,dtype=int)
Xprod[control] = 1
Xprod[target] = 1 #Pauli X_control*X_target
Zprod = np.zeros( 2 * n + 1,dtype=int)
Zprod[n + control] = 1
Zprod[n + target] = 1 #Pauli Z_control*Z_target
newtableau[control, :] = PauliImage(Xprod, tableau, n)
newtableau[n + target, :] = PauliImage(Zprod, tableau, n)
return newtableau
def LeftMultCZ(tableau, control, target, n):
""" Takes a tableau for a Clifford U and outputs the tableau for
CZ_{control,target}*U
X_control --> X_control Z_target
X_target --> Z_control X_target"""
newtableau = tableau.copy()
XZ = np.zeros(2 * n + 1,dtype=int)
XZ[control] = 1
XZ[target + n] = 1 #Pauli X_control*Z_target
ZX = np.zeros( 2 * n + 1,dtype=int)
ZX[n + control] = 1
ZX[target] = 1 #Pauli Z_control*X_target
newtableau[control, :] = PauliImage(XZ, tableau, n) #X_control --> U^{\dagger} X_control Z_target U
newtableau[target, :] = PauliImage(ZX, tableau, n) #X_target -->U^{\dagger} Z_control X_target U
return newtableau
def LeftMultS(tableau, qubit, n):
""" Takes a stabilizer tableau for a Clifford U and left multiplies the single-qubit S gate to the
specified qubit : U--> S_qubit*U
S^{\dagger} X S=-Y
S^{\dagger} Z S=Z
So must replace row qubit of tableau with image of -Y under U"""
newtableau = tableau.copy()
minusYj = np.zeros(2 * n + 1,dtype=int)
minusYj[2*n] = 2 #phase is -1
minusYj[qubit] = 1
# print(minusYj)
# print(type(_))
# print(qubit)
minusYj[qubit[0] + n] = 1
newtableau[qubit, :] = PauliImage(minusYj, tableau, n)# outputs -U^{\dagger} Y_qubit U^{\dagger}
return newtableau
def RightMultS(tableau, qubit, n):
""" Takes a stabilizer tableau for a Clifford U and left multiplies the single-qubit S gate to the
specified qubit : U--> U*S
Look at the columns of the tableau corresponding to qubit"""
for j in range(2*n):
if tableau[j, qubit] == 1: #X_qubit--> -Y_qubit; %Y_qubit--> X_qubit
# print("j=")
# print(j)
tableau[j, qubit + n] = np.mod(tableau[j, qubit + n] + 1, 2)
# print("tableau[j, qubit]=")
# print(tableau[j, qubit])
tableau[j, 2*n] = np.mod(tableau[j, 2*n] + 2 * tableau[j, qubit + n], 4) #Flip phase if X-->-Y
# print("tableau[j, 2*n]=")
# print(tableau[j, 2*n])
return tableau
def PauliImage(P, tableau, n):
"""Takes as input a tableau for a Clifford U_c and a Pauli P and outputs
the pauli Uc^{\dagger} P U_c"""
Pout = np.zeros(2 * n + 1,dtype=int)
Pout[-1] = P[-1].copy()
for j in range(n):
if P[j] == 1: #Multiply by U_c X_j U_c
Pout = MultiplyPauli(Pout, tableau[j,:], n)
if P[n + j] == 1: #Multiply by U_c Z_j U_c
Pout = MultiplyPauli(Pout, tableau[n + j,:], n)
if P[j] == 1 and P[n + j] == 1: #Correct phase X_j*Z_j*(+i)=Y_j
Pout[-1] = np.mod(Pout[-1] + 1, 4)
return Pout
def MultiplyPauli(P1, P2, n):
#Multiplies Paulis P3=P1*P2 as in Eq. 15.17 of Kitaev/Vyalyi/Shen book
P3 = np.mod(P1 + P2, 2)#Gets everything correct except phase bit
tau1 = 0
tau2 = 0
tau3 = 0
kap = 0
for j in range(n):
tau1 = tau1 + P1[j] * P1[n + j]
tau2 = tau2 + P2[j] * P2[n + j]
tau3 = tau3 + P3[j] * P3[n + j]
kap = np.mod(kap + P2[j] * P1[j + n], 2)
P3[-1] = np.mod(P1[-1] + P2[-1] + tau1 + tau2 - tau3 + 2 * kap, 4)
return P3
def bin_array(num, m):
"""Convert a positive integer num into an m-bit bit vector"""
return np.array(list(np.binary_repr(num).zfill(m))).astype(np.int8)
def Indicator(nparray, *length):
if not length:
n_values = np.max(np.asarray(nparray))+1
mat=np.eye(n_values,dtype=int)[nparray]
else:
n_values = length[0]+1
mat=np.eye(n_values,dtype=int)[nparray]
return mat
# +
def U_H(psi,i,n):
i+=1 #zero-indexing
h=np.asarray([[1,1],[1,-1]])*2**-0.5
H=np.kron(np.kron(np.eye(2**(i-1)),h),np.eye(2**(n-i)))
#print(H)
return H@psi
def U_X(psi,i,n):
i+=1 #zero-indexing
x=np.asarray([[0,1],[1,0]])
X=np.kron(np.kron(np.eye(2**(i-1)),x),np.eye(2**(n-i)))
#print(X)
return X@psi
def U_S(psi,i,n):
i+=1 #zero-indexing
s=np.asarray([[1,0],[0,1j]],dtype=complex)
S=np.kron(np.kron(np.eye(2**(i-1)),s),np.eye(2**(n-i)))
#print(S)
return S@psi
def U_CX(psi,i,j,n):
i+=1 #zero-indexing
j+=1 #zero-indexing
# Makes local projector P_ij
swap=np.asarray([[1,0,0,0],[0,0,1,0],[0,1,0,0],[0,0,0,1]])
cnot=np.asarray([[1,0,0,0],[0,1,0,0],[0,0,0,1],[0,0,1,0]])
if (j<i):
cnot=swap@cnot@swap
i2=min([i,j])
j2=max([i,j])
j=j2
i=i2
CNOT=np.kron(np.kron(np.eye(2**(i-1)),cnot),np.eye(2**(n-i-1)))
for k in range(i+1,j):
swk=np.kron(np.kron(np.eye(2**(k-1)),swap),np.eye(2**(n-k-1)))
CNOT=swk@CNOT@swk
return CNOT@psi
def U_CZ(psi,i,j,n):
i+=1 #zero-indexing
j+=1 #zero-indexing
# Makes local projector P_ij
swap=np.asarray([[1,0,0,0],[0,0,1,0],[0,1,0,0],[0,0,0,1]])
cz=np.asarray([[1,0,0,0],[0,1,0,0],[0,0,1,0],[0,0,0,-1]])
if (j<i):
cz=swap@cz@swap
i2=min([i,j])
j2=max([i,j])
j=j2
i=i2
# print('i,j=',i,j)
CZ=np.kron(np.kron(np.eye(2**(i-1)),cz),np.eye(2**(n-i-1)))
# print(list(range(i+1,j)))
for k in range(i+1,j):
swk=np.kron(np.kron(np.eye(2**(k-1)),swap),np.eye(2**(n-k-1)))
CZ=swk@CZ@swk
# print(CZ)
return CZ@psi
# -
def UnitaryUpdate(psi,gate,qubits,n):
"""psi is a stabilizer state in CH form
gate is 'S','CZ','CX',or 'H'
qubits=[q] or qubits=[q1 q2] are the qubits the gate acts on
n is the total number of qubits
psi is a dict containing np.arrays/integers {n,s,r,Uc and p}"""
if gate=='S':
psi=U_S(psi,qubits,n)
elif gate=='CZ':
psi=U_CZ(psi,qubits[0],qubits[1],n)
elif gate=='CX':
psi=U_CX(psi,qubits[0],qubits[1],n)
elif gate=='X':
psi=U_X(psi,qubits,n)
elif gate=='H':
psi=U_H(psi,qubits,n)
return psi
def compUnitaryUpdate(psi,gate,qubits,n):
"""psi is a stabilizer state in CH form
gate is 'S','CZ','CX',or 'H'
qubits=[q] or qubits=[q1 q2] are the qubits the gate acts on
n is the total number of qubits
psi is a dict containing np.arrays/integers {n,s,r,Uc and p}"""
if gate=='S':
psi=U_S(psi,qubits[0],n)
elif gate=='CZ':
psi=U_CZ(psi,qubits[0],qubits[1],n)
elif gate=='CX':
psi=U_CX(psi,qubits[0],qubits[1],n)
elif gate=='X':
psi=U_X(psi,qubits[0],n)
elif gate=='H':
psi=U_H(psi,qubits[0],n)
return psi
def fullvec(psi):
"Write out psi in the computational basis using normal lexic ordering e.g. 000,001,...,111"
n=psi['n']
psivec=np.zeros((2**n,1),dtype=complex)
for j in range(2**n):
psivec[j]=Amplitude(bin_array(j, n),psi,n)
if 'c' in psi:
psivec=psi['c']*psivec
return psivec
def normalize(v):
return v / np.linalg.norm(v)
# +
n=3
#Make |100>.
# Here the leftmost bit is the least significant
# The qubits are labeled 0,1,2,3,...n-1 from left to right
psi=CompBasisVector([1, 0, 0]);
# Apply CNOT_{12} gate (1 is target, 2 is control)
psi=CliffordUpdate(psi,'CX',[0, 1],n);
# Apply CNOT_{23} gate
psi=CliffordUpdate(psi,'CX',[1, 2],n);
#State is now |111>
#Apply Hadamards to qubits 2,3
psi=CliffordUpdate(psi,'H',[1],n);
psi=CliffordUpdate(psi,'H',[2],n);
# State is |1-->
#Apply CZ_{12} and CZ_{13}
psi=CliffordUpdate(psi,'CZ',[0, 1],n);
psi=CliffordUpdate(psi,'CZ',[0, 2],n);
# State is |1++>
# Apply S gate to 1st qubit
psi=CliffordUpdate(psi,'S',[0],n);
# State is i*|1++>
#Note that the ordering of bits is opposite to the tensor product
#ordering. The state q is equal to
v=(1/2)*1j*np.kron(np.kron(np.asarray([[0],[1]]),np.asarray([[1],[1]])),np.asarray([[1],[1]]))
print(v)
print(fullvec(psi))
#Check q-v=0
# norm(q-v)
# -
def update_and_compare(psi_in,phi_in,gtype,gqubits,n):
psi = CliffordUpdate(psi_in, gtype, gqubits, n)
phi = compUnitaryUpdate(phi_in, gtype, gqubits, n)
return psi, phi, np.linalg.norm(fullvec(psi) - phi)
def displayboth(psi,phi):
print(np.linalg.norm(fullvec(psi) - phi) )
print(np.squeeze(np.asarray([fullvec(psi),phi]),axis=2).T)
def randomstabilizer(n):
psi=XBasisVector(np.zeros((n),dtype=int)); # Psi will be the state computed by the CH Clifford simulator
phi=np.ones((2**n,1),dtype=complex)*2**(-n/2)
m=100; #Number of randomly chosen gates
r=np.random.randint(5, size=m);
qubits=[];
nrm=0;
for j in range(m):
if r[j] == 0:
q = np.random.randint(n)
# print('S',[q])
qubits = np.append(qubits, q)
psi, phi, nrm = update_and_compare(psi,phi,'S',[q],n)
# print('S',[q],nrm)
if r[j] == 1 or r[j] == 2:
collision = 0
while collision == 0:
q1 = np.random.randint(n)
q2 = np.random.randint(n)
if q1 != q2:
collision = 1
qubits = np.append(qubits, [q1, q2])
# print(qubits)
if r[j] == 1:
# print('CZ', [q1, q2])
psi, phi, nrm = update_and_compare(psi,phi,'CZ', [q1, q2],n)
# print('CZ', [q1, q2],nrm)
if r[j] == 2:
# print('CX', [q1, q2])
psi, phi, nrm = update_and_compare(psi,phi,'CX', [q1, q2],n)
# print('CX', [q1, q2],nrm)
if r[j] == 3:
q = np.random.randint(n)
# print('H',[q])
qubits = np.append(qubits, q)
psi, phi, nrm = update_and_compare(psi,phi,'H',[q],n)
# print('H',[q],nrm)
if r[j] == 4:
q = np.random.randint(n)
# print('X',[q])
qubits = np.append(qubits, q)
psi, phi, nrm = update_and_compare(psi,phi,'X',[q],n)
# print('X',[q],nrm)
q=fullvec(psi)
return psi, phi
psi, phi = randomstabilizer(4)
displayboth(psi,phi)
def EquatorialA(n):
"Randomly generate the A matrix corresponding to |phi_A> in Eq 61 "
A=np.zeros((n,n),dtype=np.int8)
offdiag=np.random.randint(2,size=int(n*(n-1)/2))
A[np.triu_indices(n, 1)]=offdiag
A=A+A.T
np.fill_diagonal(A,np.random.randint(4,size=n))
return A
def ind2vec(ind, N=None):
ind = np.asarray(ind)
if N is None:
N = ind.max() + 1
return (np.arange(N) == ind[:,None]).astype(int)
def binvec2dec(vec):
return np.dot((2**np.arange(vec.shape[0], dtype = np.uint64)[::-1]),vec).astype(np.int)
def Equatorialfullvec(A):
"""Write out psi in the computational basis using normal lexic ordering e.g. 000,001,...,111.
See Sec IV C of BBCCGH"""
n=A.shape[0]
psivec=np.zeros((2**n,1),dtype=complex)
for j in range(2**n):
x=bin_array(j,n)
psivec=psivec+1j**((x@A)@x.T)*ind2vec([binvec2dec(x)],2**n).T
psivec=2**(-n/2)*psivec
return psivec
def InnerPsiA(psi,A):
"""Implementation of <phi|phi_A> as described in Lemma 3, Sec IV C of BBCCGH"""
n=psi['n']
r=psi['r']
s=psi['s']
gamma=psi['Uc'][0:n,-1]
F=psi['Uc'][0:n,0:n]
M=psi['Uc'][0:n,n:2*n]
G=psi['Uc'][n:2*n,n:2*n]
J=np.mod(M@F.T+np.diag(gamma),4)
mask=~np.eye(J.shape[0],dtype=bool)
J[mask]=np.mod(J[mask],2)
K=G.T@(A+J)@G
wtr=np.count_nonzero(r)
term1=2**(-(n+wtr)/2)
term2=1j**((s@K)@s.T)
term3=(-1)**np.dot(s,r)
phase=np.exp(-psi['p']*1j*np.pi/4)*psi['c'].conj()
locs=np.nonzero(r)[0]
sumtot=0
for j in range(2**(wtr)):
x=np.zeros(n,dtype=int)
x[locs]=bin_array(j, wtr)#n-bit strings x satisfying x_j <= r_j
sumtot=sumtot+1j**((x@K)@x.T+2*x@(s+s@K))
return phase*term1*term2*term3*sumtot
def checkInnerPsiA(psi,A):
aa=abs(InnerPsiA(psi,A))
bb= abs(np.vdot(fullvec(psi),Equatorialfullvec(A)))
return print([aa, bb, aa-bb])
def etaA(SUBSET,Amat):
n=Amat.shape[0]
olap=sum(list(map(lambda x: InnerPsiA(SUBSET[x],Amat), range(len(SUBSET)))))
return 2**n*abs(olap)**2
def eta_estimate(SUBSET,n,Lsamples):
return np.mean([etaA(SUBSET,EquatorialA(n)) for j in range(Lsamples)])
def explicitetaA(SUBSET,Amat):
tot=0
for j in range(len(SUBSET)):
tot=tot+2**6*np.vdot(fullvec(SUBSET[j]),Equatorialfullvec(Amat))
return tot
def HiddenShiftCircuitBuilderCH(numtof,numqub,*hidstring):
"""Build the circuit for implimenting Hidden Shift for Bent functions"""
n=numqub
if not hidstring:
s=np.random.randint(2, size=n)
else:
s=np.asarray(hidstring)[0]
q3=np.random.choice(int(n/2), 3, replace=False).copy()
Og=pd.DataFrame([{'H':[q3[2].copy()]},{'Toff':q3.copy()},{'H':[q3[2].copy()]}])
Og2=pd.DataFrame([{'H':[int(n/2)+q3[2].copy()]},{'Toff':int(n/2)+q3.copy()},{'H':[int(n/2)+q3[2].copy()]}])
for j in range(numtof-1):
for i in range(10):# The number of random Clifford gates Z or C-Z
r=np.random.randint(2)
if r==1:
#print('random Z')
q=np.random.randint(n/2)
Og=Og.append(pd.DataFrame([{'S':[q]},{'S':[q]}]),ignore_index=True) # Z gate applied to qubit q
Og2=Og2.append(pd.DataFrame([{'S':[int(n/2)+q]},{'S':[int(n/2)+q]}]),ignore_index=True)
elif r==0:
#print('random CZ')
q2=np.random.choice(int(60/2), 2, replace=False).copy()
Og=Og.append(pd.DataFrame([{'CZ':q2}]),ignore_index=True) # C-Z gate between qubits in q2
Og2=Og2.append(pd.DataFrame([{'CZ':int(n/2)+q2}]),ignore_index=True)
q3=np.random.choice(int(60/2), 3, replace=False).copy()
Og=Og.append(pd.DataFrame([{'H':[q3[2]]},{'Toff':q3},{'H':[q3[2]]}]),ignore_index=True)
Og2=Og2.append(pd.DataFrame([{'H':[int(n/2)+q3[2]]},{'Toff':int(n/2)+q3},{'H':[int(n/2)+q3[1]]}]),ignore_index=True)
Of=Og.copy()
Oftil=pd.DataFrame()
for j in range(int(n/2)):
Of=Of.append(pd.DataFrame([{'CZ':[j,j+int(n/2)]}]),ignore_index=True)
Oftil=Oftil.append(pd.DataFrame([{'CZ':[j,j+int(n/2)]}]),ignore_index=True)
Oftil=Oftil.append(Og2)
# This portion needs checking
Xs=pd.DataFrame()
FT=pd.DataFrame()
for i in range(n):
FT=FT.append(pd.DataFrame([{'H':[i]}]))
if s[i]==1:
Xs=Xs.append(pd.DataFrame(np.asarray([[[i],np.nan],[np.nan,[i]],[np.nan,[i]],[[i],np.nan]],dtype=object),columns=['H','S']),ignore_index=True)
U=FT.append([Of,Xs,FT,Oftil,FT],ignore_index=True)
return U, s
def SamplesToTake(circ,delta):
numtof=0
numt=0
if 'CCZ' in circ.columns:
numtof=numtof+circ.count()['CCZ']
if 'Toff' in circ.columns:
numtof=numtof+circ.count()['Toff']
if 'T' in circ.columns:
numt=numt+circ.count()['T']
print('numtof',numtof)
print('numt',numt)
c1T=np.cos(np.pi/8)**(-numt)
c1Toff=(3/4)**(-numtof)
ksamps=int(round((c1T*c1Toff/delta)**2))
#2**(-m/2)*eps*np.exp(1j*p*np.pi/4)
return ksamps
def CreateSubsetStates(circ,ksamps,psi_in):
cliffordgates={'H','S','CZ','CX'}
pathind=0
SUBSET=[]
for path in range(ksamps):
#print('path=',path)
#psi=XBasisVector([0,0,0])#create the |+++> state
psi=deepcopy(psi_in)
for ind, row in circ.iterrows():
tmp=row.dropna()
gtype=tmp.index[0]
# print(gtype)
gqubits=np.asarray(tmp.values[0])
if gtype in cliffordgates:
psi=CliffordUpdate(psi, gtype, gqubits, n)
elif gtype=='CCZ':
x=random.choices(list(range(0,8)), weights=np.ones(8)/6, k=1)[0]
#print('CCZ','x',x)
if x==0: # Identity
pass
elif x==1: # CZ_{12}
psi=CliffordUpdate(psi, 'CZ', gqubits[[0,1]],n)
elif x==2: # CZ_{13}
psi=CliffordUpdate(psi, 'CZ', gqubits[[0,2]],n)
elif x==3: # CZ_{12,13}Z_{1}
psi=CliffordUpdate(psi, 'CZ', gqubits[[0,1]], n)
psi=CliffordUpdate(psi, 'CZ', gqubits[[0,2]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[0]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[0]], n)
elif x==4: # CZ_{23}
psi=CliffordUpdate(psi, 'CZ', gqubits[[1,2]], n)
elif x==5: # CZ_{23,12}Z_{2}
psi=CliffordUpdate(psi, 'CZ', gqubits[[1,2]], n)
psi=CliffordUpdate(psi, 'CZ', gqubits[[0,1]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[1]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[1]], n)
elif x==6: # CZ_{13,23}Z_{3}
psi=CliffordUpdate(psi, 'CZ', gqubits[[0,2]], n)
psi=CliffordUpdate(psi, 'CZ', gqubits[[1,2]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[2]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[2]], n)
elif x==7: # -CZ_{12,13,23}Z_{1,2,3}
psi=CliffordUpdate(psi, 'CZ', gqubits[[0,1]], n)
psi=CliffordUpdate(psi, 'CZ', gqubits[[0,2]], n)
psi=CliffordUpdate(psi, 'CZ', gqubits[[1,2]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[0]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[0]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[1]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[1]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[2]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[2]], n)
psi['p']=np.mod(psi['p']+4,8) # This inserts the minus sign
elif gtype=='Toff':
psi=CliffordUpdate(psi, 'H', gqubits[[2]], n)
x=random.choices(list(range(0,8)), weights=np.ones(8)/6, k=1)[0]
# x=feynmanpaths[pathind]
# print('Toff','x',x)
pathind+=1
if x==0: # Identity
pass
elif x==1: # CZ_{12}
psi=CliffordUpdate(psi, 'CZ', gqubits[[0,1]],n)
elif x==2: # CZ_{13}
psi=CliffordUpdate(psi, 'CZ', gqubits[[0,2]],n)
elif x==3: # CZ_{12,13}Z_{1}
psi=CliffordUpdate(psi, 'CZ', gqubits[[0,1]], n)
psi=CliffordUpdate(psi, 'CZ', gqubits[[0,2]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[0]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[0]], n)
elif x==4: # CZ_{23}
psi=CliffordUpdate(psi, 'CZ', gqubits[[1,2]], n)
elif x==5: # CZ_{23,12}Z_{2}
psi=CliffordUpdate(psi, 'CZ', gqubits[[1,2]], n)
psi=CliffordUpdate(psi, 'CZ', gqubits[[0,1]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[1]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[1]], n)
elif x==6: # CZ_{13,23}Z_{3}
psi=CliffordUpdate(psi, 'CZ', gqubits[[0,2]], n)
psi=CliffordUpdate(psi, 'CZ', gqubits[[1,2]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[2]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[2]], n)
elif x==7: # -CZ_{12,13,23}Z_{1,2,3}
psi=CliffordUpdate(psi, 'CZ', gqubits[[0,1]], n)
psi=CliffordUpdate(psi, 'CZ', gqubits[[0,2]], n)
psi=CliffordUpdate(psi, 'CZ', gqubits[[1,2]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[0]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[0]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[1]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[1]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[2]], n)
psi=CliffordUpdate(psi, 'S', gqubits[[2]], n)
psi['p']=np.mod(psi['p']+4,8) # This inserts the minus sign
# psi['c']=(-1)*psi['c']
psi=CliffordUpdate(psi, 'H', gqubits[[2]], n)
elif gtype=='T':
x=random.choices(list(range(0,2)), weights=np.ones(2)/2, k=1)[0]
# T = c_0 Id + c_1 S where
# c_0 = 0.5*(1+1j)*(np.exp(1j*math.pi/4)-1j)
# c_1 = -0.5*(1+1j)*(np.exp(1j*math.pi/4)-1)
phi_0=np.angle(0.5*(1+1j)*(np.exp(1j*math.pi/4)-1j))
phi_1=np.angle(-0.5*(1+1j)*(np.exp(1j*math.pi/4)-1))
#print('T','x',x)
if x==0: # Identity
pass
psi['c']=np.exp(1j*phi_0)*psi['c']
elif x==1: # S
psi=CliffordUpdate(psi, 'S', gqubits, n)
psi['c']=np.exp(1j*phi_1)*psi['c']
SUBSET.append(psi)
return SUBSET
# +
(circ,shift)=HiddenShiftCircuitBuilderCH(1,6,[0,0,0,1,1,1])
print(shift)
# (circ,shift)=HiddenShiftCircuitBuilderCH(1,6)
# print(shift)
SamplesToTake(circ,0.1)
# -
n=shift.shape[0]
def SumOverCliffords(circ):
print('n=',n)
ksamps=SamplesToTake(circ,0.1)
print(ksamps)
return CreateSubsetStates(circ,ksamps,CompBasisVector(np.asarray(np.zeros(n),dtype=int)))
# +
n=3
circ=pd.DataFrame([{'CCZ':[0,1,2]}])
ksamps=SamplesToTake(circ,0.1)
print('ksamps',ksamps)
SUBSET=CreateSubsetStates(circ,ksamps,XBasisVector([0,0,0]))
print(normalize(sum(list(map(fullvec,SUBSET)))))
# +
circ=pd.DataFrame([{'Toff':[0,1,2]}])
ksamps=SamplesToTake(circ,0.1)
print('ksamps',ksamps)
SUBSET=CreateSubsetStates(circ,ksamps,XBasisVector([0,0,0]))
# SUBSET, phiSUBSET =DebugSubsetStates(circ,ksamps,CompBasisVector([1,1,1]))
print(normalize(sum(list(map(fullvec,SUBSET)))))
# +
n=3
circ=pd.DataFrame([{'CCZ':[0,1,2]},{'Toff':[0,1,2]},{'T':[0]},{'T':[1]},{'T':[2]}])
ksamps=SamplesToTake(circ,0.05)
print('ksamps',ksamps)
SUBSET=CreateSubsetStates(circ,ksamps,XBasisVector([0,0,0]))
# SUBSET, phiSUBSET =DebugSubsetStates(circ,ksamps,CompBasisVector([1,1,1]))
print(normalize(sum(list(map(fullvec,SUBSET)))))
# +
(circ,shift)=HiddenShiftCircuitBuilderCH(1,6,[0,0,0,1,1,1])
print(shift)
# (circ,shift)=HiddenShiftCircuitBuilderCH(1,6)
# print(shift)
SamplesToTake(circ,0.1)
# -
n=shift.shape[0]
def SumOverCliffords(circ):
print('n=',n)
ksamps=SamplesToTake(circ,0.1)
print(ksamps)
return CreateSubsetStates(circ,ksamps,CompBasisVector(np.asarray(np.zeros(n),dtype=int)))
SUBSET=SumOverCliffords(circ)
unnorm=sum(list(map(fullvec,SUBSET)))
answer=normalize(unnorm)
print(np.sum(np.round(answer)-fullvec(CompBasisVector(shift))))
A6= EquatorialA(6)
print(A6)
phiA=Equatorialfullvec(A6)
print('<phi_A|phi_A>=',np.vdot(phiA,phiA))
print('<phi_A|psi>=',np.vdot(unnorm,phiA))
print('<psi|psi>=nrm_sq=',np.vdot(unnorm,unnorm))
print('eta_estimate=',eta_estimate(SUBSET,6,100))
|
PythonCH_Clifford_simulator.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
# First assign the URL of Wikipedia home page to a strings
wiki_home = "https://en.wikipedia.org/wiki/Main_Page"
response = requests.get(wiki_home)
# +
def encoding_check(r):
return (r.encoding)
def decode_content(r,encoding):
return (r.content.decode(encoding))
contents = decode_content(response,encoding_check(response))
# -
type(contents)
len(contents)
contents[:10000]
|
Chapter07/Exercise 7.03/Exercise 7.03.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Variational Quantum Eigensolver
#
# <em> Copyright (c) 2021 Institute for Quantum Computing, Baidu Inc. All Rights Reserved. </em>
# ## Overview
#
# It is widely believed that one of the most promising applications of quantum computing in the near future is solving quantum chemistry problems [1-2]. **Variational Quantum Eigensolver** (VQE) is a strong proof to this possibility of studying quantum chemistry with **Noisy Intermediate-Scale Quantum** (NISQ) devices [1-4]. The core task is to solve the ground state of any molecular Hamiltonian $\hat{H}$ by preparing a parametrized wave function ansatz $|\Psi(\boldsymbol\theta)\rangle$ on a quantum computer and adopt classical optimization methods (e.g. gradient descent) to adjust the parameters $\boldsymbol\theta$ to minimize the expectation value $\langle \Psi(\boldsymbol\theta)|\hat{H}|\Psi(\boldsymbol\theta)\rangle$. This approach is based on the **Rayleigh-Ritz variational principle**.
#
# $$
# E_0 = \min_{\boldsymbol\theta} \langle \Psi(\boldsymbol\theta)|\hat{H}|\Psi(\boldsymbol\theta)\rangle.
# \tag{1}
# $$
#
# where $E_0$ denotes the ground state energy. Numerically, it can be understood as finding the smallest eigenvalue $\lambda_{\min}$ of a **discretized** Hamiltonian $H$ (hermitian matrix) and its corresponding eigenvector $|\Psi_0\rangle$. How such a discretization can be done on a classical computer belongs to the art of quantum chemistry and is far beyond the scope of this tutorial. We will discuss this part with a few words in the background section. In general, such a Hamiltonian $H$ is expressed as a weighted sum of Pauli spin operators $\{X,Y,Z\}$ (native to quantum devices) such that this information can be processed on a quantum computer.
#
# $$
# H = \sum_k c_k ~ \bigg( \bigotimes_{j=0}^{M-1} \sigma_j^{(k)} \bigg),
# \tag{2}
# $$
#
# where $\sigma_j^{(k)} \in \{I,X,Y,Z\}$ and $M$ stands for qubit number. We refer this form of Hamiltonian as **Pauli strings**. For example,
#
# $$
# H= 0.12~Y_0 \otimes I_1-0.04~X_0\otimes Z_1.
# \tag{3}
# $$
#
# In the next section, we will provide a brief review on the electronic structure problem which essentially tells us where does the Hamiltonian $H$ come from. For those who are already familiar with this topic or only interested in how to implement VQE on Paddle Quantum, please skip this part and jump into the illustrative example of hydrogen molecule $H_2$.
#
# ## Background: the electronic structure problem
#
# In this section, we focus on one of the fundamental problems in quantum chemistry -- **the electronic structure problem**. To be more specific, we are interested in the low lying energy eigenstates of any given molecule. These knowledge could help predict reaction rates and location of stable structures [5]. Suppose a molecule consists of $N_n$ nuclei and $N_e$ electrons, the first quantized (canonical quantization) Hamiltonian operator $\hat{H}_{mol}$ describing the total energy of this molecular system can be written as
#
# $$
# \begin{align}
# \hat{H}_{\text{mol}} & = -\sum_{i}\frac{\nabla_{R_i}^2}{2M_i} - \sum_{i} \frac{\nabla_{r_i}^2}{2} -\sum_{i,j}\frac{Z_i}{\lvert R_i - r_j\lvert} + \sum_{i,j>i}\frac{Z_iZ_j}{\lvert R_i - R_j\lvert} + \sum_{i, j>i}\frac{1}{\lvert r_i - r_j\lvert},
# \tag{4}
# \end{align}
# $$
#
# where $R_i, M_i,$ and $Z_i$ denote the position, mass and atomic number (the number of protons) of the $i^{th}$ nucleus, and the positions of electrons are $r_i$. The first two sums describe the kinetic energy of nuclei and electrons, respectively. The third sum describes the attractive Coulomb interaction between the positively charged nuclei and the negatively charged electrons. The last two terms represent the repulsive nuclei-nuclei and electron-electron interactions. Here, the molecular Hamiltonian $\hat{H}_\text{mol}$ is already in atomic units of energy, **Hartree**. 1 Hartree is $[\hbar^2/(m_ee^2a_0^2)] = 27.2$ eV or 630 kcal/mol, where $m_e, e,$ and $a_0$ stand for the mass of electron, charge of electron, and Bohr radius.
#
#
# **Note:** The spin-orbit interaction and hyperfine interaction are not considered in this picture. They can be treated as perturbations if necessary.
#
# ### Born-Oppenheimer approximation
#
# Since the nuclei are much heavier than electrons, the electrons will move much faster than the nuclei. It is reasonable to treat the positions of nuclei as fixed, $R_i =$ constants. This is known as the Born-Oppenheimer approximation by decoupling the behavior of nuclei and electrons in time scale. Consequently, the kinetic energy term of nuclei will disappear and the nuclei-nuclei repulsive interaction term can be viewed as an energy shift (independent of electron positions $r_i$). We could derive the electronic Hamiltonian $\hat{H}_{\text{electron}}$ as
#
# $$
# \begin{align}
# \hat{H}_{\text{electron}} & = - \sum_{i} \frac{\nabla_{r_i}^2}{2} -\sum_{i,j}\frac{Z_i}{\lvert R_i - r_j\lvert} + \sum_{i, j>i}\frac{1}{\lvert r_i - r_j\lvert}
# \tag{5},
# \end{align}
# $$
#
# The energy levels of the electrons in the molecule can be found by solving the time independent Schrödinger equation
#
# $$
# \hat{H}_{\text{electron}} |\Psi_n \rangle = E_n |\Psi_n \rangle,
# \tag{6}
# $$
#
# where $n$ stands for the energy level. Notice the electron repulsion terms scale as $N_e(N_e-1)/2$ which means for the Oxygen molecule $O_2$ carrying 16 electrons there will be 120 electron repulsion terms in total! In general, this problem cannot be solved analytically. As Dirac concluded in [Quantum mechanics of many-electron systems](https://royalsocietypublishing.org/doi/10.1098/rspa.1929.0094) [6],
#
# > *The underlying physical laws necessary for the mathematical theory of a large part of physics and the whole of chemistry are thus completely known, and the difficulty is only that the exact application of these laws leads to equations much too complicated to be soluble.*
# >
# > -- <NAME> (1929)
#
# A straightforward numerical approach is discretizing the infinite-dimensional Hilbert space into equidistant grid points where linear algebra guides the whole calculation. Suppose each axis of space is discretized into $k$ points, the $N$-electron (drop the subscript e for simplicity) wave function can be written as [2]
#
# $$
# |\Psi \rangle = \sum_{\mathbf{x_1}, \ldots, \mathbf{x_N}} \psi(\mathbf{x_1}, \ldots, \mathbf{x_N}) \mathcal{A}(|\mathbf{x_1}, \ldots, \mathbf{x_N}\rangle).
# \tag{7}
# $$
#
# where coordinate $|\mathbf{x_j}\rangle = |r_j\rangle |\sigma_j\rangle$ records the spatial location and spin of the $j^{th}$ electron, $|r_j\rangle = |x_j,y_j,z_j\rangle$ for $j\in \{1,2,\cdots,N\}$, $x_j,y_j,z_j \in \{0,1,\cdots,k-1\}$ and $\sigma_j \in \{\downarrow,\uparrow\}$ for spin down or up. There will be $k^{3N}\times 2^{N}$ complex amplitudes in total. Here, $\mathcal{A}$ denotes anti-symmetrization and a consequence of the Pauli exclusion principle (electrons are fermion), and $\psi(\mathbf{x_1}, \mathbf{x_2}, \ldots, \mathbf{x_N})=\langle\mathbf{x_1}, \mathbf{x_2}, \ldots, \mathbf{x_N}|\Psi\rangle$. One can see that storing such a wave function already requires **exponentially growing memory** with respect to the number of electrons $N$. This would make classical simulation methods based on this naive numerical approach intractable for systems size larger than few tens of electrons. Now, the question becomes can we prepare such a wave function $|\Psi\rangle$ directly on a quantum computer and measure the expectation value $E_0$? In the next section, let's take the simplest molecular system -- hydrogen molecule $H_2$ as a concrete example.
#
#
#
# **Note:** A detailed review on quantum chemistry and existing classical computational methods are far beyond the scope of this tutorial, we refer the enthusiastic readers to the standard textbooks *'Molecular Electronic-Structure Theory'* [5] by Helgaker and *'Modern Quantum Chemistry: Introduction to Advanced Electronic Structure Theory'* [7] by Szabo & Ostlund. To bridge to knowledge gap between quantum chemistry and quantum computing, please check the following review papers [Quantum computational chemistry](https://journals.aps.org/rmp/abstract/10.1103/RevModPhys.92.015003) [2] and [Quantum Chemistry in the Age of Quantum Computing](https://pubs.acs.org/doi/10.1021/acs.chemrev.8b00803) [1].
#
# **Note:** For energy calculation, it is desired to reach the **chemical accuracy** of $1.6\times10^{-3}$ Hartree or 1 kcal/mol .
# ## Ground state of the hydrogen molecule $H_2$
#
# ### Building electronic Hamiltonian
#
# First of all, let us import the necessary libraries and packages.
#
#
# +
import os
import platform
import matplotlib.pyplot as plt
from IPython.display import clear_output
import numpy
from numpy import concatenate
from numpy import pi as PI
from numpy import savez, zeros
import paddle
from paddle_quantum.circuit import UAnsatz
from paddle_quantum.utils import pauli_str_to_matrix
from paddle_quantum.VQE.chemistrysub import H2_generator
# -
# To analyze specific molecules, we need several key information such as **geometry**, **basis set** (such as STO-3G), **multiplicity**, and **charge** to obtain the discretized Hamiltonian $H$. Specifically, through our built-in quantum chemistry toolkit, fermion-to-qubit mapping technology can be used to output the qubit Hamiltonian of hydrogen molecule $H_2$,
Hamiltonian, N = H2_generator()
# For more advanced users, we provide a simple tutorial on how to generate such a Hamiltonian. Install the following two packages first (**only available for Mac/Linux users, not available to Windows users temporarily**):
# !pip install openfermion
clear_output()
# !pip install openfermionpyscf
clear_output()
# +
# Operating system information
sysStr = platform.system()
# Decide which operating system the user is using
if sysStr in ('Linux', 'Darwin'):
import openfermion
import openfermionpyscf
# Please check whether the geometric configuration file of h2 is downloaded correctly
geometry = 'h2.xyz'
# geometry = [('H', (0.0, 0.0, 0.0)), ('H', (0.0, 0.0, 0.74))]
basis = 'sto-3g'
charge = 0
multiplicity = 1
# Generate Hamiltonian
molecular_hamiltonian = openfermionpyscf.generate_molecular_hamiltonian(geometry, basis, multiplicity, charge)
qubit_op = openfermion.transforms.jordan_wigner(molecular_hamiltonian)
# Print result
print("The generated h2 Hamiltonian is \n", qubit_op)
# -
# **Note:** This Hamiltonian is generated with an interatomic distance of $d = 74$ pm.
#
# In addition to hydrogen molecule $H_2$, we also provide the geometric configuration file of hydrogen fluoride (HF) molecule `hf.xyz`. If you need to test the geometric configuration of more molecules, please check out this [database](http://smart.sns.it/molecules/index.html). In addition, we also need to convert the Hamiltonian into the Pauli string format supported by Paddle Quantum. Here we provide this interface.
#
#
# +
def Hamiltonian_str_convert(qubit_op):
'''
Convert the Hamiltonian information provided above into Pauli strings supported by Paddle Quantum
H = [[1.0, "z0,x1"], [-1.0, "y0,z1"], ...]
'''
info_dic = qubit_op.terms
def process_tuple(tup):
if len(tup) == 0:
return 'i0'
else:
res = ''
for ele in tup:
res += ele[1].lower()
res += str(ele[0])
res += ','
return res[:-1]
H_info = []
for key, value in qubit_op.terms.items():
H_info.append([value.real, process_tuple(key)])
return H_info
if sysStr in ('Linux', 'Darwin'):
Hamiltonian = Hamiltonian_str_convert(qubit_op)
# -
# ### Building QNN and trial wave function
#
# To implement VQE, we firstly need to design a quantum neural network QNN to prepare the wave function ansatz $|\Psi(\boldsymbol\theta)\rangle$. Here, we provide a 4-qubit quantum circuit template with a depth of $D$ blocks. The dotted frame in the figure below denotes a single block:
#
#
# 
#
# Next, we use the `UAnsatz` class and the built-in `real_entangled_layer(theta, D)` circuit template in Paddle Quantum to realize this QNN.
#
#
def U_theta(theta, Hamiltonian, N, D):
"""
Quantum Neural Network
"""
# Initialize the quantum neural network according to the number of qubits N
cir = UAnsatz(N)
# Built-in {R_y + CNOT} circuit template
cir.real_entangled_layer(theta[:D], D)
# Lay R_y gates in the last row
for i in range(N):
cir.ry(theta=theta[D][i][0], which_qubit=i)
# The quantum neural network acts on the default initial state |0000>
cir.run_state_vector()
# Calculate the expected value of a given Hamiltonian
expectation_val = cir.expecval(Hamiltonian)
return expectation_val, cir
# ### Setting up the loss function and model
#
# Now that we have the target Hamiltonian and QNN, we will further define the training model and loss function. By applying the QNN $U(\theta)$ on the initial state $|0..0\rangle$, we get the output state $|\psi(\boldsymbol\theta)\rangle $. Then, the loss function to be minimized is the expectation value,
#
#
# $$
# \min_{\boldsymbol\theta} \mathcal{L}(\boldsymbol \theta) = \min_{\boldsymbol\theta} \langle \Psi(\boldsymbol\theta)|H |\Psi(\boldsymbol\theta)\rangle
# = \min_{\boldsymbol\theta} \sum_k c_k~\langle \Psi(\boldsymbol\theta)| \bigotimes_j \sigma_j^{(k)}|\Psi(\boldsymbol\theta)\rangle.
# \tag{8}
# $$
class StateNet(paddle.nn.Layer):
def __init__(self, shape, dtype="float64"):
super(StateNet, self).__init__()
# Initialize the theta parameter list and fill the initial value with a uniform distribution of [0, 2*pi]
self.theta = self.create_parameter(shape=shape,
default_initializer=paddle.nn.initializer.Uniform(low=0.0, high=2*PI),
dtype=dtype, is_bias=False)
# Define loss function and forward propagation mechanism
def forward(self, N, D):
# Calculate the loss function/expected value
loss, cir = U_theta(self.theta, Hamiltonian, N, D)
return loss, cir
# ### Hyper-parameters
#
# Before training the QNN, we also need to set some training hyper-parameters, mainly the learning rate (LR), the number of iterations (ITR), and the depth (D) of repeated blocks.
ITR = 80 # Set the number of optimization iterations
LR = 0.4 # Set the learning rate
D = 2 # Set the depth of the repetitive calculation module in QNN
# ### Training
#
# After all the training model parameters are set, we convert the data into Tensor in the Paddle, and then train the quantum neural network. The results of the training process is stored in the summary_data file.
#
# +
# Determine the parameter dimension of the network
net = StateNet(shape=[D + 1, N, 1])
# Generally speaking, we use Adam optimizer to obtain relatively good convergence,
# You can change it to SGD or RMS prop.
opt = paddle.optimizer.Adam(learning_rate=LR, parameters=net.parameters())
# Record optimization results
summary_iter, summary_loss = [], []
# Optimization loop
for itr in range(1, ITR + 1):
# Forward propagation to calculate loss function
loss, cir = net(N, D)
# Use back propagation to minimize the loss function
loss.backward()
opt.minimize(loss)
opt.clear_grad()
# Record optimization results
summary_loss.append(loss.numpy())
summary_iter.append(itr)
# Print result
if itr % 20 == 0:
print("iter:", itr, "loss:", "%.4f" % loss.numpy())
print("iter:", itr, "Ground state energy:", "%.4f Ha"
% loss.numpy())
if itr == ITR:
print("\nThe trained circuit:")
print(cir)
# Save the training results to the output folder
os.makedirs("output", exist_ok=True)
savez("./output/summary_data", iter = summary_iter,
energy=summary_loss)
# -
# ### Benchmarking
# We have now completed the training of the quantum neural network, and the estimated value of the ground state energy obtained is $E_0 \approx -1.1361$ Hartree, we compare it with the theoretical value $E_0 = -1.13618$ to benchmark our model. The estimation obtained with VQE agree with a full configuration-interaction (FCI) calculation within chemical accuracy $\varepsilon = 1.6 \times 10^{-3}$ Hartree.
#
#
# +
result = numpy.load('./output/summary_data.npz')
eig_val, eig_state = numpy.linalg.eig(
pauli_str_to_matrix(Hamiltonian, N))
min_eig_H = numpy.min(eig_val.real)
min_loss = numpy.ones([len(result['iter'])]) * min_eig_H
plt.figure(1)
func1, = plt.plot(result['iter'], result['energy'],
alpha=0.7, marker='', linestyle="-", color='r')
func_min, = plt.plot(result['iter'], min_loss,
alpha=0.7, marker='', linestyle=":", color='b')
plt.xlabel('Number of iteration')
plt.ylabel('Energy (Ha)')
plt.legend(handles=[
func1,
func_min
],
labels=[
r'$\left\langle {\psi \left( {\theta } \right)} '
r'\right|H\left| {\psi \left( {\theta } \right)} \right\rangle $',
'Ground-state energy',
], loc='best')
#plt.savefig("vqe.png", bbox_inches='tight', dpi=300)
plt.show()
# -
# ## Determining the interatomic distance
#
# Recall the above calculation is done with an interatomic distance $d = 74$ pm between two hydrogen atoms. Another interesting aspect we can try with VQE is determining the true interatomic distance by modifying the `h2.xyz` file. The results are summarize in figure below,
#
# 
#
# The lowest value is found around $d = 74$ pm (1 pm = $1\times 10^{-12}$m), which is consistent with the [experimental data](https://cccbdb.nist.gov/exp2x.asp?casno=1333740&charge=0) $d_{exp} (H_2) = 74.14$ pm.
# _______
#
# ## References
#
# [1] <NAME>, et al. Quantum Chemistry in the Age of Quantum Computing. [Chemical reviews 119.19 (2019): 10856-10915.](https://pubs.acs.org/doi/10.1021/acs.chemrev.8b00803)
#
# [2] <NAME>, et al. Quantum computational chemistry. [Reviews of Modern Physics 92.1 (2020): 015003.](https://journals.aps.org/rmp/abstract/10.1103/RevModPhys.92.015003)
#
#
# [3] <NAME>. et al. A variational eigenvalue solver on a photonic quantum processor. [Nat. Commun. 5, 4213 (2014).](https://www.nature.com/articles/ncomms5213)
#
# [4] Moll, Nikolaj, et al. Quantum optimization using variational algorithms on near-term quantum devices. [Quantum Science and Technology 3.3 (2018): 030503.](https://iopscience.iop.org/article/10.1088/2058-9565/aab822)
#
# [5] Helgaker, Trygve, <NAME>, and <NAME>. Molecular electronic-structure theory. <NAME> & Sons, 2014.
#
# [6] Dirac, <NAME>. Quantum mechanics of many-electron systems. [Proceedings of the Royal Society of London. Series A, Containing Papers of a Mathematical and Physical Character 123.792 (1929): 714-733.](https://royalsocietypublishing.org/doi/10.1098/rspa.1929.0094)
#
# [7] Szabo, Attila, and <NAME>. Modern quantum chemistry: introduction to advanced electronic structure theory. Courier Corporation, 2012.
|
tutorial/quantum_simulation/VQE_EN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + cellView="form" id="hQnpY5Kpm5rT"
#sets the project id
PROJECT_ID = "mlrh-330919" #@param {type:"string"}
# + id="v0L2i-Nnp7_R"
import os
from google.colab import auth
from IPython.display import display
#sets dateset
DATASET_PROJECT_ID = 'amsterdamumcdb'
DATASET_ID = 'version1_0_2'
LOCATION = 'eu'
#all libraries check this environment variable, so set it:
os.environ["GOOGLE_CLOUD_PROJECT"] = PROJECT_ID
auth.authenticate_user()
print('Authenticated')
# + id="izTeiKOs3F7p"
import pandas as pd
import matplotlib.pyplot as plt
from scipy import stats
import numpy as np
# + id="kmQ4q26l4bqS"
from google.colab import drive
drive.mount('/content/drive')
# + id="45PwzqgK4vEm"
os.chdir('/content/drive/MyDrive/MLRFH')
# + id="Nyva7EbsU_yt"
#Some preprocessing functions
def to_cols(data):
grouped = data.pivot_table(index=['admissionid', 'time'],
columns=['item'], values='value')
return grouped
def to_cols_action(data):
grouped = data.pivot_table(index=['admissionid', 'time'],
columns=['item'], values='administered')
return grouped
def remove_outliers(grouped):
#delete outliers
outliers = grouped.reset_index() #return to single index
#select outlier cols
all_cols = ['Kalium (bloed)', 'ABP gemiddeld', 'Kreatinine (bloed)', 'Natrium (bloed)', 'UrineCAD', 'UrineSupraPubis', 'UrineSpontaan', 'UrineUP', 'Kreatinine', 'Nefrodrain re Uit', 'Nefrodrain li Uit', 'UrineIncontinentie']
grouped['Kalium (bloed)'][grouped['Kalium (bloed)'] > 8.] = np.nan
grouped['ABP gemiddeld'][grouped['ABP gemiddeld'] > 200.] = np.nan
grouped['Kreatinine (bloed)'][grouped['Kreatinine (bloed)'] > 220.] = np.nan
grouped['Natrium (bloed)'][grouped['Natrium (bloed)'] > 180.] = np.nan
grouped['UrineCAD'][grouped['UrineCAD'] > 750.] = np.nan
#return grouped[grouped[all_cols] >= 0]
grouped[all_cols] = grouped[all_cols].applymap(lambda x: np.nan if x < 0 else x)
return grouped
def remove_outliers_action(grouped):
#delete outliers
outliers = grouped.reset_index() #return to single index
cols = ['Noradrenaline (Norepinefrine)', 'NaCl 0,45%/Glucose 2,5%']
#select outlier cols
grouped['Noradrenaline (Norepinefrine)'][grouped['Noradrenaline (Norepinefrine)'] > 10.] = np.nan
grouped['NaCl 0,45%/Glucose 2,5%'][grouped['NaCl 0,45%/Glucose 2,5%'] > 500.] = np.nan
grouped[cols] = grouped[cols].applymap(lambda x: np.nan if x < 0 else x)
return grouped
def aggregate(outliers_removed):
#per patient, average the values in 4h timeslots
outliers_removed = outliers_removed.sort_values('time')
data_agg = outliers_removed.groupby([pd.Grouper(level='admissionid'),
pd.Grouper(level='time', freq='4H')]
).mean()
return data_agg
def interpolate(data_agg):
#interpolate null values
return data_agg.interpolate(limit_direction='both')
def process_statespace(data):
data['time'] = pd.to_datetime(data['time'], unit='ms')
print("data", data.columns)
grouped = to_cols(data)
grouped = remove_outliers(grouped)
data_agg = aggregate(grouped)
data_filled = interpolate(data_agg)
return data_filled.reset_index()
def process_actionspace(data):
data['time'] = pd.to_datetime(data['time'], unit='ms')
print("data", data.columns)
grouped = to_cols_action(data)
grouped = remove_outliers_action(grouped)
data_agg = aggregate(grouped)
data_filled = interpolate(data_agg)
return data_filled.reset_index()
# + id="gnhJELl1t5cG"
#plot distribution of cols
def draw_histograms(df, variables, n_rows, n_cols):
colors = ["pink", "orange", "yellow", "green", "blue", "purple", "black", "darkgreen", "darkblue", "grey", "lightblue", "red"]
fig=plt.figure()
for i, var_name in enumerate(variables):
ax=fig.add_subplot(n_rows,n_cols,i+1)
df[var_name].hist(bins=100,ax=ax, color=colors[i])
ax.set_title(var_name)
fig.tight_layout() # Improves appearance a bit.
plt.show()
#normalization, nice for plotting
def minmax(df):
return (df - df.min()) / ( df.max() - df.min())
# + id="p9st0wgM3mmT"
#data = pd.read_csv('new_new_dataset.csv')
data = pd.read_csv('state_v2.csv')
# + id="_RBSmDVxVjqP"
data.head()
# + id="fI0Fcj41FopT"
#Plot before distributions
cols1 = ['Kalium (bloed)', 'ABP gemiddeld', 'Kreatinine (bloed)', 'Natrium (bloed)', 'UrineCAD', 'UrineSupraPubis']
cols2 = ['UrineSpontaan', 'UrineUP', 'Kreatinine', 'Nefrodrain re Uit', 'Nefrodrain li Uit', 'UrineIncontinentie']
colors = ["pink", "orange", "yellow", "green", "blue", "purple", "black", "darkgreen", "darkblue", "grey", "lightblue", "red"]
fig=plt.figure()
for i, var_name in enumerate(cols1):
ax=fig.add_subplot(3,3,i+1)
data['value'][data['item'] == var_name].hist(bins=100,ax=ax, color=colors[i])
ax.set_title(var_name)
fig.tight_layout() # Improves appearance a bit.
plt.show()
# + id="o7OV9OXlGQmU"
fig=plt.figure()
for i, var_name in enumerate(cols2):
ax=fig.add_subplot(3,3,i+1)
data['value'][data['item'] == var_name].hist(bins=100,ax=ax, color=colors[i])
ax.set_title(var_name)
fig.tight_layout() # Improves appearance a bit.
plt.show()
# + id="aBK09Lnpk0mO"
data['item'].value_counts()
# + id="nUx5hZymosGm"
#patient id=0 before preprocessing
pd.set_option('display.max_rows', 100)
data['time'] = pd.to_datetime(data['measuredat'], unit='ms')
data[data['admissionid'] == 0].sort_values(by = "measuredat")
# + id="lSsW-qCOiKXz"
statespace = process_statespace(data)
# + id="1rcfi7P6pheH"
#check for one patient whether the aggregations are correct
statespace[statespace['admissionid'] == 0]
# + id="SHkmTz8_zQqi"
from numpy.lib import histograms
variables = ['Kalium (bloed)', 'ABP gemiddeld', 'Kreatinine (bloed)', 'Natrium (bloed)', 'UrineCAD', 'UrineSupraPubis']
draw_histograms(statespace, variables, 3, 3)
# + id="P0nEwXnP6y8x"
variables2 = ['UrineSpontaan', 'UrineUP', 'Kreatinine', 'Nefrodrain re Uit', 'Nefrodrain li Uit', 'UrineIncontinentie']
draw_histograms(statespace, variables2, 3, 3)
# + id="qXtNtYo6vcPl"
statespace
# + [markdown] id="5XEClhZriUS5"
# **Action** **Space**
# + id="Sc8WgbHpiToR"
action = pd.read_csv('action_space.csv')
action['time'] = pd.to_datetime(action['stop'] - action['start'], unit='ms')
action = action.drop(columns = ['start', 'stop'])
# + id="GiesMTI-Hf4W"
action['administered'][action['item'] == 'Noradrenaline (Norepinefrine)'].mean()
# + id="UyjIyau8H_vf"
action['administered'][action['item'] == 'NaCl 0,45%/Glucose 2,5%'].max()
# + id="-zjNkLIU3G9D"
#check for 1 patient
pd.set_option('display.max_rows', 135)
action[action['admissionid'] == 4251]
# + id="BdyBGag9mjcv"
actionspace = process_actionspace(action)
# + id="AbrcT8-DJPoh"
actionspace['Noradrenaline (Norepinefrine)'].max()
# + id="UrQNYz_h3bZm"
actionspace[actionspace['admissionid'] == 4251]
# + [markdown] id="jOlJ3RKDw6NC"
# **Combine two dataframes**
# + id="GJkZnnWKBgy-"
#take first 48 hours per patient
statespace = statespace.sort_values(by=['admissionid', 'time'])
statespace48h = statespace.groupby('admissionid').head(12)
# + id="tHEld4PuDjo8"
action.isnull().sum()
# + id="zvYD2Xqj5His"
#we need to know the gender, otherwise we cannot compute AKI --> nans are dropped
genders = action[['admissionid', 'gender']].dropna()
# + id="JYi9SF3AB-sc"
#add gender to dataframe
def check_gender(admissionid):
try:
gender = genders['gender'][genders['admissionid'] == admissionid].head(1).item()
except ValueError:
gender = "Unknown"
return gender
# + id="Mj39PjglEQvh"
#add gender to dataframe
def check_age(admissionid):
age = action['agegroup'][action['admissionid'] == admissionid].head(1).item()
return age
# + id="R2tN3h7hrznU"
statespace48h['gender'] = [check_gender(x) for x in statespace48h['admissionid']]
# + id="2r_qYkyqrsin"
#remove unknowns, as we need to know the gender to compute the AKI
statespace48h = statespace48h[statespace48h.gender != 'Unknown']
# + id="tPXuj50mEmm0"
statespace48h['agegroup'] = [check_age(x) for x in statespace48h['admissionid']]
# + id="JFLXR2vtK_0L"
statespace48h['agegroup'].value_counts()
# + id="OjJfnrFHGVvf"
#Agegroup is categorical --> encode them into a numerical variable
# agegroups = pd.get_dummies(statespace48h['agegroup'])
# statespace48h = pd.concat([statespace48h, agegroups], axis=1)
ages = {"agegroup": {"18-39": 1, "40-49": 2, "50-59": 3, "60-69":4, "70-79":5, "80+":6}}
statespace48h = statespace48h.replace(ages)
statespace48h
# + id="pVTRWkqOIb5g"
#df with 1 row per patient for demographic stats
demo = statespace48h.drop_duplicates(subset=['admissionid'], keep='first')
# + id="SKQ3Rkb1th_q"
#add AKI feature
def AKI(kreatinine, gender):
if gender == 'Vrouw':
if kreatinine > 106 and kreatinine <= 134:
return 1
if kreatinine > 134 and kreatinine <= 205:
return 2
if kreatinine > 205:
return 3
else:
return 0
if gender == 'Man':
if kreatinine > 119 and kreatinine <= 151:
return 1
if kreatinine > 151 and kreatinine <= 231:
return 2
if kreatinine > 231:
return 3
else:
return 0
# + id="k3tDjoi8toAP"
#add AKI to every row in statespace
statespace48h['AKI'] = statespace48h.apply(lambda row: AKI(row['Kreatinine (bloed)'], row['gender']), axis=1)
# + id="sScanRFNuFvx"
statespace48h['AKI'].value_counts()
# + id="xpThDJCDuOPT"
#merge datasets --> left merge on actionspace, as states without actions are not useful for our model
space = actionspace.merge(statespace48h, on=["admissionid", "time"], how="left")
# + id="yafVTlFdungH"
#all the null values are patients that are not in the state space and only in the action space --> we cannot use them so they are dropped
space.isnull().sum()
space = space.dropna()
# + id="OuANHRacJ5Tl"
#one hot encode gender
space = pd.get_dummies(space, columns = ['gender'])
# + id="_eAjxLUkvTke"
#save final space in the drive
#space.to_csv("space_correct_demo7.csv")
# + id="Ls1soMJSwFJQ"
space[space['admissionid'] == 4251]
|
Colab Notebooks/MLRH_Preprocessing_Complete.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/suleymanVR/MachineLearning/blob/main/Week6_LogisticRegression.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="LOQGYyf6xknp"
# # LogisticRegression
# + id="gZxY68FDxpyh"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/"} id="aVw660WNxtVI" outputId="2bd4a3fe-b16e-4fb1-ffba-776824ab6b7c"
data = pd.read_csv("/content/data.csv") #İyi ve kötü huylu kanser hücreleri hakkında dataset // #M kotu huylu tumor B iyi huylu tumor
data.drop(["Unnamed: 32","id"],axis=1,inplace=True) #Verideki Unnamed: 32 ve id isimli kısımları kaldırdık
data.diagnosis = [1 if each =="M" else 0 for each in data.diagnosis]
print(data.info())
y=data.diagnosis.values
x_data = data.drop(["diagnosis"],axis=1) #data.diagnosis harici butun veriler x ekseninde olacak.
# + [markdown] id="gC9ze8LLx_Ls"
# Normalization
# + id="O1Rd2nGGx7fH"
# (x-min(x))/(max(x)-min(x))
x=(x_data - np.min(x_data))/(np.max(x_data)-np.min(x_data)).values
# + [markdown] id="30WVxU0zyH00"
# Train-test split
# + colab={"base_uri": "https://localhost:8080/"} id="8RamzuN4yFnp" outputId="033b6e9f-3d89-49a1-ec98-df7c918523f0"
from sklearn.model_selection import train_test_split
x_train,x_test,y_train,y_test = train_test_split(x,y,test_size=0.2,random_state=42) #%20lik kismi test icin ayirdik
#transpozlarını aldik
x_train = x_train.T
x_test = x_test.T
y_train = y_train.T
y_test = y_test.T
print("x_train:",x_train.shape)
print("x_test,:",x_test.shape)
print("y_train",y_train.shape)
print("y_test:",y_test.shape)
# + id="XQBPQ9H_yRA0"
# parameter initialize and sigmoid function
# dimension = 30
# w=weight(agirlik), b=bias
def initialize_weights_and_bias(dimension):
w = np.full((dimension,1),0.01) #0.01'lerden olusan bir matris olusturur. #En basta rastgele sectigimiz weight degerlerini buradan alıyoruz.
b = 0.0
return w,b
#w,b = initialize_weights_and_bias(30)
def sigmoid(z):
y_head =1/(1 + np.exp(-z)) #sigmoid f. formulu
return y_head
# + id="CTaAvdpRyZtV"
def forward_backward_propagation(w,b,x_train,train):
#forward propagation
z = np.dot(w.T,x_train) + b
y_head = sigmoid(z)
loss = -y_train*np.log(y_head)-(1-y_train)*np.log(1-y_head)
cost = (np.sum(loss))/x_train.shape[1]
#backward propagation
derivative_weight = (np.dot(x_train,((y_head-y_train).T)))/x_train.shape[1]
derivative_bias = np.sum(y_head-y_train)/x_train.shape[1]
gradients = {"derivative_weight": derivative_weight,"derivative_bias": derivative_bias}
return cost,gradients
# 1 forward propagation + 1 backward propagation = 1 number_of_iteration
# En basta rastgele 0.01 olarak aldıgımız weight degerlerini en verimli hale getirmek icin forward ve backward pro. uyguluyoruz.
# + id="RzosWofvyaSh"
#number_of_iteration = kac kez backward ve forward propagation yapacagimiz.(1back+1forwawrd=1number_of_iteration)
def update(w, b, x_train, y_train, learning_rate, number_of_iteration):
cost_list = []
cost_list2 = []
index = []
for i in range(number_of_iteration):
cost,gradients = forward_backward_propagation(w,b,x_train,y_train)
cost_list.append(cost) #tüm costlarin degerini tutuyoruz
#weight - learningrate * weigthin cost'a gore turevi
w = w - learning_rate * gradients["derivative_weight"]
b = b - learning_rate * gradients["derivative_bias"]
if i % 10 == 0: #her 10 adimda bir costlarin degerini tutar
cost_list2.append(cost)
index.append(i)
print("Cost after iteration %i: %f" %(i, cost))
parameters = {"weight":w,"bias":b}
plt.plot(index,cost_list2)
plt.xticks(index,cost_list2)
plt.xlabel("Number of Iteration")
plt.ylabel("Cost")
plt.show()
#cost ile gradient degerleri birbiri ile bağlantili degerlerdir birisi azalirsa digeri de azalir
return parameters,gradients,cost_list2
# + id="Iy244iL7ycpa"
def predict(w,b,x_test):
z = sigmoid(np.dot(w.T,x_test)+b)
Y_prediction = np.zeros((1,x_test.shape[1]))
for i in range(z.shape[1]):
if z[0,i]<=0.5:
Y_prediction[0,i] = 0
else:
Y_prediction[0,i] = 1
return Y_prediction
# + colab={"base_uri": "https://localhost:8080/", "height": 817} id="83jYs60MyeQB" outputId="c14e3daa-7649-41b0-eb57-0949df4eaae8"
def logistic_regression(x_train, y_train, x_test, y_test, learning_rate , num_iterations):
# initialize
dimension = x_train.shape[0] # that is 30
w,b = initialize_weights_and_bias(dimension)
# do not change learning rate
parameters, gradients, cost_list = update(w, b, x_train, y_train, learning_rate,num_iterations)
y_prediction_test = predict(parameters["weight"],parameters["bias"],x_test)
# Print train/test Errors
print("test accuracy: {} %".format(100 - np.mean(np.abs(y_prediction_test - y_test)) * 100))
logistic_regression(x_train, y_train, x_test, y_test,learning_rate = 1, num_iterations = 300)
# learning rate ve num_iterations degerlerini degistirerek accuracy degerini bir noktaya kadar arttırabiliriz.
# learning rate degerini fazla yükseltmek iyi egitilmemis bir makine olusturabilir.
# Test accuracy 96.49122807017544 %
# + [markdown] id="PCdlJwXO0f55"
# Learning rate = 3 num_iteration = 1000 oldugu durumda accuracy degerini kontrol edelim.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="8M34czfx0nvQ" outputId="b17f3d3a-1568-4b3b-b133-ab400e32e90d"
def logistic_regression(x_train, y_train, x_test, y_test, learning_rate , num_iterations):
# initialize
dimension = x_train.shape[0] # that is 30
w,b = initialize_weights_and_bias(dimension)
# do not change learning rate
parameters, gradients, cost_list = update(w, b, x_train, y_train, learning_rate,num_iterations)
y_prediction_test = predict(parameters["weight"],parameters["bias"],x_test)
# Print train/test Errors
print("test accuracy: {} %".format(100 - np.mean(np.abs(y_prediction_test - y_test)) * 100))
logistic_regression(x_train, y_train, x_test, y_test,learning_rate = 3, num_iterations = 1000)
# Test accuracy 97.36842105263158 %
# Accuracy degerinde %1'lik bir artıs gerceklesti
# + [markdown] id="RDl4Ylhf1iBM"
# Sklearn de verilen hazır kütüphane ve methodlar kullanılarak yapılan logisticregression analiz sonucları
# + colab={"base_uri": "https://localhost:8080/"} id="6fsUgRJU0tcY" outputId="af208ef9-a37a-4b74-bc34-8c8dafc5100c"
from sklearn.linear_model import LogisticRegression
lr = LogisticRegression()
lr.fit(x_train.T,y_train.T)
print("test accuracy {}".format(lr.score(x_test.T,y_test.T)))
# Accuracy degeri %98 seviyelerine ulastı.
|
Week6_LogisticRegression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Model Selection using RFE(HOUSING CASE STUDY)
# ### Importing and Understanding Data
import pandas as pd
import numpy as np
# Importing Housing.csv
housing = pd.read_csv('Housing.csv')
# Looking at the first five rows
housing.head()
# ### Data Preparation
# Converting Yes to 1 and No to 0
housing['mainroad'] = housing['mainroad'].map({'yes': 1, 'no': 0})
housing['guestroom'] = housing['guestroom'].map({'yes': 1, 'no': 0})
housing['basement'] = housing['basement'].map({'yes': 1, 'no': 0})
housing['hotwaterheating'] = housing['hotwaterheating'].map({'yes': 1, 'no': 0})
housing['airconditioning'] = housing['airconditioning'].map({'yes': 1, 'no': 0})
housing['prefarea'] = housing['prefarea'].map({'yes': 1, 'no': 0})
# Creating dummy variable for variable furnishingstatus and dropping the first one
status = pd.get_dummies(housing['furnishingstatus'],drop_first=True)
#Adding the results to the master dataframe
housing = pd.concat([housing,status],axis=1)
# Dropping the variable 'furnishingstatus'
housing.drop(['furnishingstatus'],axis=1,inplace=True)
# #### Creating a new variable
# Let us create the new metric and assign it to "areaperbedroom"
housing['areaperbedroom'] = housing['area']/housing['bedrooms']
# Metric: bathrooms per bedroom
housing['bbratio'] = housing['bathrooms']/housing['bedrooms']
# ### Rescaling the Features
# It is extremely important to rescale the variables so that they have a comparable scale.
# There are twocoon ways of rescaling
# 1. Normalisation (min-max scaling) and
# 2. standardisation (mean-o, sigma-1)
# Let's try normalisation
# +
#defining a normalisation function
def normalize (x):
return ( (x-np.mean(x))/ (max(x) - min(x)))
# applying normalize ( ) to all columns
housing = housing.apply(normalize)
# -
# ### Splitting data into training and testing sets
# +
# Putting feature variable to X
X = housing[['area', 'bedrooms', 'bathrooms', 'stories', 'mainroad',
'guestroom', 'basement', 'hotwaterheating', 'airconditioning',
'parking', 'prefarea', 'semi-furnished', 'unfurnished',
'areaperbedroom', 'bbratio']]
# Putting response variable to y
y = housing['price']
# -
#random_state is the seed used by the random number generator, it can be any integer.
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.7 ,test_size = 0.3, random_state=100)
# UDF for calculating vif value
def vif_cal(input_data, dependent_col):
vif_df = pd.DataFrame( columns = ['Var', 'Vif'])
x_vars=input_data.drop([dependent_col], axis=1)
xvar_names=x_vars.columns
for i in range(0,xvar_names.shape[0]):
y=x_vars[xvar_names[i]]
x=x_vars[xvar_names.drop(xvar_names[i])]
rsq=sm.OLS(y,x).fit().rsquared
vif=round(1/(1-rsq),2)
vif_df.loc[i] = [xvar_names[i], vif]
return vif_df.sort_values(by = 'Vif', axis=0, ascending=False, inplace=False)
# ### RFE
# Importing RFE and LinearRegression
from sklearn.feature_selection import RFE
from sklearn.linear_model import LinearRegression
# Running RFE with the output number of the variable equal to 9
lm = LinearRegression()
rfe = RFE(lm, 9) # running RFE
rfe = rfe.fit(X_train, y_train)
print(rfe.support_) # Printing the boolean results
print(rfe.ranking_)
col = X_train.columns[rfe.support_]
# ### Building model using sklearn
# Creating X_test dataframe with RFE selected variables
X_train_rfe = X_train[col]
# Adding a constant variable
import statsmodels.api as sm
X_train_rfe = sm.add_constant(X_train_rfe)
lm = sm.OLS(y_train,X_train_rfe).fit() # Running the linear model
#Let's see the summary of our linear model
print(lm.summary())
# Calculating Vif value
vif_cal(input_data=housing.drop(['area','bedrooms','stories','basement','semi-furnished','areaperbedroom'], axis=1), dependent_col="price")
# ## Making Predictions
# +
# Now let's use our model to make predictions.
# Creating X_test_6 dataframe by dropping variables from X_test
X_test_rfe = X_test[col]
# Adding a constant variable
X_test_rfe = sm.add_constant(X_test_rfe)
# Making predictions
y_pred = lm.predict(X_test_rfe)
# -
# ## Model Evaluation
# +
# Now let's check how well our model is able to make predictions.
# Importing the required libraries for plots.
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# -
# Actual and Predicted
import matplotlib.pyplot as plt
c = [i for i in range(1,165,1)] # generating index
fig = plt.figure()
plt.plot(c,y_test, color="blue", linewidth=2.5, linestyle="-") #Plotting Actual
plt.plot(c,y_pred, color="red", linewidth=2.5, linestyle="-") #Plotting predicted
fig.suptitle('Actual and Predicted', fontsize=20) # Plot heading
plt.xlabel('Index', fontsize=18) # X-label
plt.ylabel('Housing Price', fontsize=16) # Y-label
# Error terms
c = [i for i in range(1,165,1)]
fig = plt.figure()
plt.plot(c,y_test-y_pred, color="blue", linewidth=2.5, linestyle="-")
fig.suptitle('Error Terms', fontsize=20) # Plot heading
plt.xlabel('Index', fontsize=18) # X-label
plt.ylabel('ytest-ypred', fontsize=16) # Y-label
# Plotting y_test and y_pred to understand the spread.
fig = plt.figure()
plt.scatter(y_test,y_pred)
fig.suptitle('y_test vs y_pred', fontsize=20) # Plot heading
plt.xlabel('y_test', fontsize=18) # X-label
plt.ylabel('y_pred', fontsize=16) # Y-label
# Plotting the error terms to understand the distribution.
fig = plt.figure()
sns.distplot((y_test-y_pred),bins=50)
fig.suptitle('Error Terms', fontsize=20) # Plot heading
plt.xlabel('y_test-y_pred', fontsize=18) # X-label
plt.ylabel('Index', fontsize=16) # Y-label
# Now let's check the Root Mean Square Error of our model.
import numpy as np
from sklearn import metrics
print('RMSE :', np.sqrt(metrics.mean_squared_error(y_test, y_pred)))
|
Linear Regression/Housing Case Study RFE.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#importing the libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
#using pandas to read the data
loans=pd.read_csv('loan_data.csv')
#cheking out the head,info and description of our data
loans.info()
loans.head()
loans.describe()
# +
# Exploratory Data Analysis
# -
#Creating a histogram of two FICO distributions on top of each other, one for each credit.policy outcome.
fig,axes=plt.subplots(figsize=(10,6))
loans[loans['credit.policy']==1]['fico'].hist(alpha=0.4,bins=30,label='Credit.Policy=1')
loans[loans['credit.policy']==0]['fico'].hist(alpha=0.5,bins=30,label='Credit.Policy=0')
plt.legend()
# +
fig,axes=plt.subplots(figsize=(10,5))
loans[loans['not.fully.paid']==0]['fico'].hist(alpha=0.7,bins=40,label='not.fully.paid=0',color='red')
loans[loans['not.fully.paid']==1]['fico'].hist(alpha=0.5,bins=40,label='not.fully.paid=1',color='blue')
plt.legend()
# +
#Creating a countplot using seaborn showing the counts of loans by purpose, with the color hue defined
#by not.fully.paid.
# -
fig,axes=plt.subplots(figsize=(10,6))
sns.countplot('purpose',hue='not.fully.paid',data=loans)
#creating a jointplot between FICO score and interest rate
sns.jointplot(x='fico',y='int.rate',data=loans)
#Creating the following lmplots to see if the trend differed between not.fully.paid and credit.policy
sns.lmplot(x='fico',y='int.rate',col='not.fully.paid',data=loans,hue='credit.policy')
# +
#setting up the data
# -
#categorical features
#converting the categorical features into dummy variables using
#pd.getdummies()
cat_feats = ['purpose']
final_data = pd.get_dummies(loans,columns=cat_feats,drop_first=True)
final_data.head(2)
# +
#Train test split
# +
#using sklearn spilting the data into training set and testing set
# -
from sklearn.cross_validation import train_test_split
X=final_data.drop('not.fully.paid',axis=1)
y=final_data['not.fully.paid']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3,random_state=101)
# Traing A Decision Tree Model
# +
#Importing Decision Tree Classifier
# -
from sklearn.tree import DecisionTreeClassifier
tree=DecisionTreeClassifier()
#fittimg our training data to our model
tree.fit(X_train,y_train)
# ## Predictions and Evaluation of Decision Tree
# **Creating predictions from the test set and creating a classification report and a confusion matrix.**
# +
#Creating Predictions
# -
predictions=tree.predict(X_test)
#importing classification report and confusion matrix from sklearn
from sklearn.metrics import classification_report,confusion_matrix
#Printing the classification report and confusio matrix
print(classification_report(y_test,predictions))
print('\n')
print(confusion_matrix(y_test,predictions))
# ## Training the Random Forest model
#
#importing random forest tree from sklearn.ensemble
from sklearn.ensemble import RandomForestClassifier
rfc=RandomForestClassifier(n_estimators=200)
#fitting the data to our model
rfc.fit(X_train,y_train)
#making predictions from our model
rfc_pred=rfc.predict(X_test)
# +
#Printing the classification report and confusio matrix
# -
print(classification_report(y_test,rfc_pred))
print('\n')
print(confusion_matrix(y_test,rfc_pred))
|
Lending Club.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %env CUDA_DEVICE_ORDER=PCI_BUS_ID
# %env CUDA_VISIBLE_DEVICES=2
# %load_ext autoreload
# %autoreload 2
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from tqdm.autonotebook import tqdm
from joblib import Parallel, delayed
import umap
import pandas as pd
from avgn.utils.paths import DATA_DIR, most_recent_subdirectory, ensure_dir
from avgn.signalprocessing.create_spectrogram_dataset import flatten_spectrograms
from avgn.visualization.spectrogram import draw_spec_set
from avgn.visualization.quickplots import draw_projection_plots
from avgn.visualization.projections import (
scatter_projections,
draw_projection_transitions,
)
# ### Collect data
DATASET_ID = 'bengalese_finch_sober'
# ### create dataset
from avgn.utils.hparams import HParams
from avgn.dataset import DataSet
hparams = HParams(
num_mel_bins = 32,
mel_lower_edge_hertz=500,
mel_upper_edge_hertz=15000,
butter_lowcut = 500,
butter_highcut = 15000,
ref_level_db = 20,
min_level_db = -25,
mask_spec = True,
win_length_ms = 10,
hop_length_ms = 2,
nex=-1,
n_jobs=-1,
verbosity = 1,
)
# create a dataset object
dataset = DataSet(DATASET_ID, hparams = hparams)
# ### load syllable df
df_loc = DATA_DIR / 'syllable_dfs' / DATASET_ID / 'bf.pickle'
df_loc
syllable_df = pd.read_pickle(df_loc)
syllable_df[:3]
plt.hist(np.array(syllable_df.end_time - syllable_df.start_time), bins=50);
# ### Cluster
def norm(x):
return (x - np.min(x)) / (np.max(x) - np.min(x))
from cuml.manifold.umap import UMAP as cumlUMAP
import hdbscan
from avgn.umap import umap_reduce
# +
fig, ax = plt.subplots(nrows=2, ncols=len(syllable_df.indv.unique()), figsize=(10*len(syllable_df.indv.unique()), 20))
indv_dfs = {}
for indvi, indv in enumerate(tqdm(syllable_df.indv.unique())):
#if indv != 'Bird5': continue
indv_dfs[indv] = syllable_df[syllable_df.indv == indv]
indv_dfs[indv] = indv_dfs[indv].sort_values(by=["key", "start_time"])
print(indv, len(indv_dfs[indv]))
specs = [norm(i) for i in indv_dfs[indv].spectrogram.values]
# sequencing
indv_dfs[indv]["syllables_sequence_id"] = None
indv_dfs[indv]["syllables_sequence_pos"] = None
for ki, key in enumerate(indv_dfs[indv].key.unique()):
indv_dfs[indv].loc[indv_dfs[indv].key == key, "syllables_sequence_id"] = ki
indv_dfs[indv].loc[indv_dfs[indv].key == key, "syllables_sequence_pos"] = np.arange(
np.sum(indv_dfs[indv].key == key)
)
# umap
specs_flattened = flatten_spectrograms(specs)
#cuml_umap = cumlUMAP(min_dist=0.5)
#z = list(cuml_umap.fit_transform(specs_flattened))
z, _ = umap_reduce(specs_flattened, min_dist = 0.5)
indv_dfs[indv]["umap"] = list(z)
# HDBSCAN UMAP
clusterer = hdbscan.HDBSCAN(
min_cluster_size=int(len(z) * 0.01), # the smallest size we would expect a cluster to be
min_samples=1, # larger values = more conservative clustering
)
clusterer.fit(z);
indv_dfs[indv]['hdbscan_labels'] = clusterer.labels_
# -
len(indv_dfs)
# ### Plot spectrogram with labels
import seaborn as sns
def song_barcode(start_times, stop_times, labels, label_pal_dict, resolution = 0.01):
"""
"""
begin = np.min(start_times)
end = np.max(stop_times)
trans_list = np.zeros(
int((end - begin) / resolution)
).astype('uint8').astype("object")
#print(end, begin, end-begin, resolution, len(trans_list))
for start, stop, label in zip(start_times, stop_times, labels):
trans_list[int((start - begin)/resolution):int((stop-begin)/resolution)] = label
color_list = [label_pal_dict[i] if i in label_pal_dict else [1,1,1] for i in trans_list]
color_list = np.expand_dims(color_list, 1)
return trans_list, color_list
def indv_barcode(indv_df, time_resolution = 0.02, label = 'labels', pal = "tab20"):
unique_labels = indv_df[label].unique()
# song palette
label_pal = np.random.permutation(sns.color_palette(pal, len(unique_labels)))
label_pal_dict = {
lab: color
for lab, color in zip(
unique_labels,
label_pal,
)
}
sns.palplot(list(label_pal_dict.values()))
label_dict = {lab:int(i) for i, lab in enumerate(unique_labels)}
# get list of syllables by time
trans_lists = []
color_lists = []
for key in tqdm(indv_dfs[indv].key.unique()):
# dataframe of wavs
wav_df = indv_dfs[indv][indv_dfs[indv]['key'] == key]
labels = wav_df['labels'].values
start_times = wav_df.start_time.values
stop_times = wav_df.end_time.values
start_times[:3], stop_times[:3], labels[:3]
trans_list, color_list = song_barcode(start_times, stop_times, labels, label_pal_dict, resolution = resolution)
color_lists.append(color_list)
trans_lists.append(trans_list)
return color_lists, trans_lists, label_pal_dict, label_pal
# for each individual in the dataset
for indv in tqdm(indv_dfs.keys()):
color_lists, trans_lists, label_pal_dict, label_pal = indv_barcode(
indv_dfs[indv], time_resolution=0.02, label="labels", pal="tab20"
)
break
from scipy.cluster import hierarchy
from nltk.metrics.distance import edit_distance
from scipy.cluster.hierarchy import dendrogram, linkage
from scipy.spatial.distance import squareform
from matplotlib import gridspec
max_list_len = 600 # maximum length to visualize
seq_len = 100 # maximumim length to compute lev distance
nex = 50 # only show up to NEX examples
# +
# subset dataset
color_lists = color_lists[:nex]
trans_lists = trans_lists[:nex]
# get length of lists
list_lens = [len(i) for i in trans_lists]
# set max list length
if max_list_len is None:
max_list_len = np.max(list_lens)
# make a matrix for color representations of syllables
color_item = np.ones((max_list_len, len(list_lens), 3))
for li, _list in enumerate(tqdm(color_lists)):
color_item[:len(_list), li, :] = np.squeeze(_list[:max_list_len])
color_items = color_item.swapaxes(0,1)
# make a list of symbols padded to equal length
trans_lists = np.array(trans_lists)
cut_lists = [
list(i[:seq_len].astype("str"))
if len(i) >= seq_len
else list(i) + list(np.zeros(seq_len - len(i)).astype("str"))
for i in trans_lists
]
cut_lists = [''.join(np.array(i).astype('str')) for i in cut_lists]
# create a distance matrix (THIS COULD BE PARALLELIZED)
dist = np.zeros((len(cut_lists), len(cut_lists)))
for i in tqdm(range(1, len(cut_lists))):
for j in range(0, i):
dist[i,j] = edit_distance(cut_lists[i], cut_lists[j])
dist[j,i] = edit_distance(cut_lists[i], cut_lists[j])
# hierarchical clustering
dists = squareform(dist)
linkage_matrix = linkage(dists, "single")
# +
fig = plt.figure(figsize=(25, 6))
gs = gridspec.GridSpec(1, 2, width_ratios=[1, 4], wspace=0, hspace=0)
ax0 = plt.subplot(gs[0])
ax1 = plt.subplot(gs[1])
dn = dendrogram(
linkage_matrix,
p=6,
truncate_mode="none",
get_leaves=True,
orientation="left",
no_labels=True,
link_color_func=lambda k: "k",
ax=ax0,
show_contracted=False
)
plt.title("test")
ax0.axis("off")
ax1.imshow(
color_item.swapaxes(0, 1)[np.array(dn["leaves"])],
aspect="auto",
interpolation=None,
origin='lower'
)
ax1.axis("off")
plt.show()
|
notebooks/05.0-viz-transitions-continuous/.ipynb_checkpoints/transitions-bengalese-finch-sober-with-dendrogram-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.4
# language: python
# name: python-374
# ---
# # ipytest
# ## Setup
# +
# Set the file name (required)
__file__ = 'testing.ipynb'
# Add ipython magics
# Add ipython magics
import ipytest
import pytest
ipytest.autoconfig()
# -
# ## Test Case
# +
# %%ipytest
def test_sorted():
assert sorted([4, 2, 1, 3]) == [1, 2, 3, 4]
# -
# ## Test Fixture
# +
# %%ipytest
@pytest.fixture
def dict_list():
return [
dict(a='a', b=3),
dict(a='c', b=1),
dict(a='b', b=2),
]
def test_sorted__key_example_1(dict_list):
assert sorted(dict_list, key=lambda d: d['a']) == [
dict(a='a', b=3),
dict(a='b', b=2),
dict(a='c', b=1),
]
def test_sorted__key_example_2(dict_list):
assert sorted(dict_list, key=lambda d: d['b']) == [
dict(a='c', b=1),
dict(a='b', b=2),
dict(a='a', b=3),
]
# -
# ## Testparametrisierung
# +
# %%ipytest
@pytest.mark.parametrize('input,expected', [
([2, 1], [1, 2]),
('zasdqw', list('adqswz')),
])
def test_examples(input, expected):
actual = sorted(input)
assert actual == expected
# -
# ## Referenz
#
# ### `%%run_pytest …`
#
# IPython-Magic, die zuerst die Zelle und dann `run_pytest` ausführt. In der Zelle übergebene Argumente werden direkt an pytest weitergeleitet. Zuvor sollten mit `import ipytest.magics` die Magics importiert worden sein.
#
# ### `ipytest.run_pytest(module=None, filename=None, pytest_options=(), pytest_plugins=())`
#
# führt die Tests im bestehenden Modul (standardmäßig `main`) mit pytest aus.
#
# Argumente:
#
# * `module`: das Modul, das die Tests enthält. Wenn nicht angegeben wird, wird `__main__` verwendet.
# * `filename`: Dateiname der Datei, die die Tests enthält. Wenn nichts angegeben wird, wird das `__file__`-Attribut des übergebenen Moduls verwendet.
# * `pytest_options`: zusätzliche Optionen, die an pytest übergeben werden
# * `pytest_plugins`: zusätzliche pytest-Plugins
#
# ### `ipytest.run_tests(doctest=False, items=None)`
#
# Argumente:
#
# * `doctest`: Wenn als Wert `True` angegeben wird, wird nach Doctests gesucht.
# * `items`: Das *globals*-Objekt, das die Tests enthält. Wenn `None` angegeben wird, wird das *globals*-Objekt aus dem Call Stack ermittelt.
#
# ### `ipytest.clean_tests(pattern="test*", items=None)`
#
# löscht diejenigen Tests, deren Namen dem angegebenen Muster entsprechen.
#
# In IPython werden die Ergebnisse aller Auswertungen in globalen Variablen gespeichert, sofern sie nicht explizit gelöscht werden. Dieses Verhalten impliziert, dass beim Umbenennen von Tests die vorherigen Definitionen weiterhin gefunden werden, wenn sie nicht gelöscht werden. Diese Methode zielt darauf ab, diesen Prozess zu vereinfachen.
#
# Ein effektive Methode besteht darin, mit `clean_tests` eine Zelle zu beginnen, dann alle Testfälle zu definieren und schließlich `run_tests` aufzurufen. Auf diese Weise funktioniert das Umbenennen von Tests wie erwartet.
#
# Argumente:
#
# * `pattern`: Ein glob-Pattern, das verwendet wird, um die zu löschenden Tests zu finden.
# * `items`: Das *globals*-Objekt, das die Tests enthält. Wenn `None` angegeben wird, wird das *globals*-Objekt aus dem Call Stack ermittelt.
#
# ### `ipytest.collect_tests(doctest=False, items=None)`
#
# sammelt alle Testfälle und sendet sie an `unittest.TestSuite`.
#
# Die Argumente sind die gleichen wie für `ipytest.run_tests`.
#
# ### `ipytest.assert_equals(a, b, *args, **kwargs)`
#
# vergleicht zwei Objekte und wirft eine *Exception*, wenn sie nicht gleich sind.
#
# Die Methode `ipytest.get_assert_function` bestimmt die zu verwendende Assert-Implementierung in Abhängigkeit von den folgenden Argumenten:
#
# * `a, b`: die zwei zu vergleichenden Objekte.
# * `args, kwargs`: (Schlüsselwort)-Argumente, die an die zugrundeliegende Testfunktion übergeben werden.
#
# ### `ipytest.get_assert_function(a, b)`
#
# bestimmt die zu verwendende Assert-Funktion in Abhängigkeit von den Argumenten.
#
# Wenn eines der Objekte `numpy.ndarray`, `pandas.Series`, `pandas.DataFrame` oder `pandas.Panel` ist, werden die von `numpy` und `pandas` bereitgestellten Assert-Funktionen zurückgegeben.
#
# ### `ipytest.unittest_assert_equals(a, b)`
#
# vergleicht zwei Objekte mit der `assertEqual`-Methode von `unittest.TestCase`.
#
|
docs/productive/testing/ipytest.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import random
from random import randint
import pandas as pd
import numpy as np
import gym
import matplotlib.pyplot as plt
import seaborn as sns
# ---
# ## Helper functions
#
# ### Reward and reward per episode
def var(N,episodes,epsilon,alpha,gamma):
M = np.zeros((N,episodes))
Mre=np.zeros((N,episodes))
for n in range(N):
_,R,_,_,_,Rf,_ = FL_play(episodes=episodes,epsilon=epsilon,alpha=alpha,gamma=gamma)
M[n] = Rf
Mre[n] = R
return np.mean(M,axis=0),np.mean(Mre,axis=0)
def plot_average_rewards_per_episode(N,episodes,epsilon,alpha,gamma):
M,Mre = var(N,episodes=episodes,epsilon=epsilon,alpha=alpha,gamma=gamma)
Mr,Mrer = var(N,episodes=episodes,epsilon=1.0,alpha=alpha,gamma=gamma)
plt.figure(figsize=(15,8))
plt.plot(range(M.shape[0]),np.cumsum(M),label="$\epsilon$=0.1, std:{:.3f}, lr={:.1f}, $\gamma$={:.1f}".format(np.std(Mre),alpha,gamma))
plt.plot(range(Mr.shape[0]),np.cumsum(Mr),label="$\epsilon$=1.0, std:{:.3f}, lr={:.1f}, $\gamma$={:.1f}".format(np.std(Mrer),alpha,gamma))
plt.title('Learning curve - Max R: {:.0f}'.format(sum(Mre)),fontsize=16)
plt.xlabel('Episodes',fontsize=16)
plt.ylabel('Average Reward per episode',fontsize=16)
plt.legend(loc=0,fontsize=16)
plt.show()
def plot_average_rewards(N,episodes,epsilon,alpha,gamma):
M,Mre = var(N,episodes=episodes,epsilon=epsilon,alpha=alpha,gamma=gamma)
Mr,Mrer = var(N,episodes=episodes,epsilon=1.0,alpha=alpha,gamma=gamma)
plt.figure(figsize=(15,8))
plt.plot(range(M.shape[0]),np.cumsum(Mre),label="$\epsilon$=0.1, std:{:.3f}, lr={:.1f}, $\gamma$={:.1f}".format(np.std(Mre),alpha,gamma))
plt.plot(range(Mr.shape[0]),np.cumsum(Mrer),label="$\epsilon$=1.0, std:{:.3f}, lr={:.1f}, $\gamma$={:.1f}".format(np.std(Mrer),alpha,gamma))
plt.title('Learning Gain - Max R: {:.0f}'.format(sum(Mre)),fontsize=16)
plt.xlabel('Episodes',fontsize=16)
plt.ylabel('Average Reward',fontsize=16)
plt.legend(loc=0,fontsize=16)
plt.show()
# ### Q-Network
def var_q(N,episodes,epsilon,alpha,gamma):
M = np.zeros((env.observation_space.n, env.action_space.n))
for n in range(N):
Q,_,_,_,_,_,_ = FL_play(episodes=episodes,epsilon=epsilon,alpha=alpha,gamma=gamma)
M += Q
return M / N
def var_qnetwork(N,episodes,epsilon,alpha,gamma):
Q = var_q(N,episodes=episodes,epsilon=epsilon,alpha=alpha,gamma=gamma)
plt.figure(figsize=(20,5))
sns.heatmap(Q.T,annot=Q.T,linewidths=.75,cmap="YlGnBu",square=True, fmt=".2f")
plt.xlabel('States',fontsize=16)
plt.ylabel('Actions',fontsize=16)
plt.plot('Q-network value')
plt.yticks((0,1,2,3),('Left','Down','Right','Up'),rotation=0,fontsize=16)
plt.show()
# ---
# ## Gym - Frozen Lake
env = gym.make('FrozenLake-v0')
# ## SARSA
def policy(Q,state,epsilon):
if np.random.rand() < epsilon:
action = np.random.choice([0,1])
else:
action = np.argmax(Q[state,:])
return action
def FL_play(episodes,epsilon,alpha,gamma):
Q = np.random.rand(env.observation_space.n, env.action_space.n)*0.0001
R = []
Rf = []
for i in range(episodes):
state = env.reset()
r = 0
action = policy(Q,state,epsilon)
while True:
next_state,reward,done,_ = env.step(action)
next_action = policy(Q,next_state,epsilon)
r+=reward
if done:
Q[state,action] = (1-alpha)*Q[state,action] + alpha*reward
break
else:
Q[state,action] = (1-alpha)*Q[state,action] + alpha*(reward +gamma*Q[next_state,next_action])
state = next_state
action = next_action
R.append(r)
Rf.append(r/(i+1))
return Q,R,epsilon,alpha,gamma,Rf,L
# %%time
plot_average_rewards(N=50,episodes=150,epsilon=0.1,alpha=0.9,gamma=0.9)
# %%time
plot_average_rewards(N=50,episodes=1500,epsilon=0.1,alpha=0.9,gamma=0.9)
# %%time
plot_average_rewards(N=5,episodes=10000,epsilon=0.1,alpha=0.9,gamma=0.1)
# %%time
plot_average_rewards_per_episode(N=10,episodes=10000,epsilon=0.1,alpha=0.9,gamma=0.9)
# %%time
plot_average_rewards_per_episode(N=10,episodes=10000,epsilon=0.05,alpha=0.9,gamma=0.9)
# %%time
var_qnetwork(N=5,episodes=20000,epsilon=0.1,alpha=0.9,gamma=0.95)
|
TemporalDifference/SARSA/SARSA-FrozenLake.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Part 1: Preprocessing data
#
# Part 1: prepare training set and stores key information about the training set in MLRun (we wasn't able to use functions with our own naming to run the workflow, so this pipeline is heavily based on MLrun tutorial 1. Apologize in advance for the confusing naming)
#
# We have completed the following:
#
# - Created a MLRun function that automates data processing
# - Stored data artifacts in a central database
# - Ran the code on Kubernetes cluster
# <a id="gs-tutorial-1-mlrun-intro"></a>
# <a id="gs-tutorial-1-mlrun-envr-init"></a>
# ## Step1: Initializing MLRun Environment
# Run the following code to initialize MLRun environment to use a "getting-started-tutorial-<username>"
# project and store the project artifacts in the default artifacts path:
# +
from os import path
import mlrun
# Set the base project name
project_name_base = 'getting-started-tutorial'
# Initialize the MLRun environment and save the project name and artifacts path
project_name, artifact_path = mlrun.set_environment(project=project_name_base,
user_project=True)
# Display the current project name and artifacts path
print(f'Project name: {project_name}')
print(f'Artifacts path: {artifact_path}')
# -
# <a id="gs-tutorial-1-step-create-basic-function"></a>
# ## Step 2: Creating a Basic Function
#
# Develop a MLRun functions converted from a local function used for data preprocessing
# <a id="gs-tutorial-1-define-local-func"></a>
# +
import pandas as pd
# Ingest a data set
def prep_data(source_url, label_column):
df = pd.read_csv(source_url)
df[label_column] = df[label_column].astype('category').cat.codes
return df, df.shape[0]
# -
# <a id="gs-tutorial-1-create-and-run-an-mlrun-function"></a>
# +
# mlrun: start-code
# -
import mlrun
def prep_data(context, source_url: mlrun.DataItem, label_column='label'):
# Convert the DataItem to a pandas DataFrame
df = source_url.as_df()
df[label_column] = df[label_column].astype('category').cat.codes
# Record the DataFrane length after the run
context.log_result('num_rows', df.shape[0])
# Store the data set in your artifacts database
context.log_dataset('cleaned_data', df=df, index=False, format='csv')
# +
# mlrun: end-code
# -
# Convert the local prep_data function to an MLRun project function
data_prep_func = mlrun.code_to_function(name='prep_data', kind='job', image='mlrun/mlrun')
# <a id="gs-tutorial-1-run-mlrun-function-locally"></a>
# Set the source-data URL. We uploaded our dataset into the directory under data, which is the only directory
# that can save changes for users on docker
source_url = '/home/jovyan/data/preprocessed-2.csv'
# Run the `data_prep_func` MLRun function locally
prep_data_run = data_prep_func.run(name='prep_data',
handler=prep_data,
inputs={'source_url': source_url},
local=True)
# <a id="gs-tutorial-1-get-run-object-info"></a>
# make sure the MLrun function is run successfully
prep_data_run.state()
# make sure the cleaned data is generated
prep_data_run.outputs['cleaned_data']
# <a id="gs-tutorial-1-read-output"></a>
# ## Step 3: Reading and Storing the Output
#
# Investigate the output of cleaned dataset
dataset = mlrun.run.get_dataitem(prep_data_run.outputs['cleaned_data'])
# You can also get the data as a pandas DataFrame by calling the `dataset.as_df` method:
dataset.as_df()
# <a id="gs-tutorial-1-save-artifcats-in-run-specific-paths"></a>
# Saving the Artifacts in Run-Specific Paths
# +
out = artifact_path
prep_data_run = data_prep_func.run(name='prep_data',
handler=prep_data,
inputs={'source_url': source_url},
local=True,
artifact_path=path.join(out, '{{run.uid}}'))
# -
# <a id="gs-tutorial-1-step-run-func-on-cluster"></a>
# ## Step 4: Running the Function on a Cluster
#
from mlrun.platforms import auto_mount
data_prep_func.apply(auto_mount())
prep_data_run = data_prep_func.run(name='prep_data',
handler='prep_data',
inputs={'source_url': source_url},
local=False)
print(prep_data_run.outputs)
# <a id="gs-tutorial-1-step-ui-jobs-view"></a>
# <a id="gs-tutorial-1-step-schedule-jobs"></a>
|
Data-Speaks-Laviosa1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Credit Risk Resampling Techniques
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
from pathlib import Path
from collections import Counter
# # Read the CSV and Perform Basic Data Cleaning
# +
columns = [
"loan_amnt", "int_rate", "installment", "home_ownership",
"annual_inc", "verification_status", "issue_d", "loan_status",
"pymnt_plan", "dti", "delinq_2yrs", "inq_last_6mths",
"open_acc", "pub_rec", "revol_bal", "total_acc",
"initial_list_status", "out_prncp", "out_prncp_inv", "total_pymnt",
"total_pymnt_inv", "total_rec_prncp", "total_rec_int", "total_rec_late_fee",
"recoveries", "collection_recovery_fee", "last_pymnt_amnt", "next_pymnt_d",
"collections_12_mths_ex_med", "policy_code", "application_type", "acc_now_delinq",
"tot_coll_amt", "tot_cur_bal", "open_acc_6m", "open_act_il",
"open_il_12m", "open_il_24m", "mths_since_rcnt_il", "total_bal_il",
"il_util", "open_rv_12m", "open_rv_24m", "max_bal_bc",
"all_util", "total_rev_hi_lim", "inq_fi", "total_cu_tl",
"inq_last_12m", "acc_open_past_24mths", "avg_cur_bal", "bc_open_to_buy",
"bc_util", "chargeoff_within_12_mths", "delinq_amnt", "mo_sin_old_il_acct",
"mo_sin_old_rev_tl_op", "mo_sin_rcnt_rev_tl_op", "mo_sin_rcnt_tl", "mort_acc",
"mths_since_recent_bc", "mths_since_recent_inq", "num_accts_ever_120_pd", "num_actv_bc_tl",
"num_actv_rev_tl", "num_bc_sats", "num_bc_tl", "num_il_tl",
"num_op_rev_tl", "num_rev_accts", "num_rev_tl_bal_gt_0",
"num_sats", "num_tl_120dpd_2m", "num_tl_30dpd", "num_tl_90g_dpd_24m",
"num_tl_op_past_12m", "pct_tl_nvr_dlq", "percent_bc_gt_75", "pub_rec_bankruptcies",
"tax_liens", "tot_hi_cred_lim", "total_bal_ex_mort", "total_bc_limit",
"total_il_high_credit_limit", "hardship_flag", "debt_settlement_flag"
]
target = ["loan_status"]
# +
# Load the data
file_path = Path('../Resources/LoanStats_2019Q1.csv.zip')
df = pd.read_csv(file_path, skiprows=1)[:-2]
df = df.loc[:, columns].copy()
# Drop the null columns where all values are null
df = df.dropna(axis='columns', how='all')
# Drop the null rows
df = df.dropna()
# Remove the `Issued` loan status
issued_mask = df['loan_status'] != 'Issued'
df = df.loc[issued_mask]
# convert interest rate to numerical
df['int_rate'] = df['int_rate'].str.replace('%', '')
df['int_rate'] = df['int_rate'].astype('float') / 100
# Convert the target column values to low_risk and high_risk based on their values
x = {'Current': 'low_risk'}
df = df.replace(x)
x = dict.fromkeys(['Late (31-120 days)', 'Late (16-30 days)', 'Default', 'In Grace Period'], 'high_risk')
df = df.replace(x)
df.reset_index(inplace=True, drop=True)
df.head()
# -
# # Split the Data into Training and Testing
# +
# Create our features
X = df.drop(columns=['loan_status'])
X = pd.get_dummies(X)
# Create our target
y = df[["loan_status"]]
# -
X.describe()
# Check the balance of our target values
y['loan_status'].value_counts()
# Create X_train, X_test, y_train, y_test
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=1, stratify=y)
# ## Data Pre-Processing
#
# Scale the training and testing data using the `StandardScaler` from `sklearn`. Remember that when scaling the data, you only scale the features data (`X_train` and `X_testing`).
# Create the StandardScaler instance
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
# Fit the Standard Scaler with the training data
# When fitting scaling functions, only train on the training dataset
X_scaler = scaler.fit(X_train)
# Scale the training and testing data
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
# # Oversampling
#
# In this section, you will compare two oversampling algorithms to determine which algorithm results in the best performance. You will oversample the data using the naive random oversampling algorithm and the SMOTE algorithm. For each algorithm, be sure to complete the folliowing steps:
#
# 1. View the count of the target classes using `Counter` from the collections library.
# 3. Use the resampled data to train a logistic regression model.
# 3. Calculate the balanced accuracy score from sklearn.metrics.
# 4. Print the confusion matrix from sklearn.metrics.
# 5. Generate a classication report using the `imbalanced_classification_report` from imbalanced-learn.
#
# Note: Use a random state of 1 for each sampling algorithm to ensure consistency between tests
# ### Naive Random Oversampling
# +
# Resample the training data with the RandomOversampler
from imblearn.over_sampling import RandomOverSampler
ros = RandomOverSampler(random_state=1)
X_resampled, y_resampled = ros.fit_resample(X_train, y_train)
Counter(y_resampled)
# +
# Train the Logistic Regression model using the resampled data
from sklearn.linear_model import LogisticRegression
nro_model = LogisticRegression(solver='lbfgs', random_state=1)
nro_model.fit(X_resampled, y_resampled)
# +
# Calculate the balanced accuracy score
from sklearn.metrics import balanced_accuracy_score
y_pred_nro = nro_model.predict(X_test)
balanced_accuracy_score(y_test, y_pred_nro)
# +
# Display the confusion matrix
from sklearn.metrics import confusion_matrix
confusion_matrix(y_test, y_pred_nro)
# +
# Print the imbalanced classification report
from imblearn.metrics import classification_report_imbalanced
print(classification_report_imbalanced(y_test, y_pred_nro))
# -
# ### SMOTE Oversampling
# +
# Resample the training data with SMOTE
from imblearn.over_sampling import SMOTE
X_resampled, y_resampled = SMOTE(random_state=1, sampling_strategy=1.0).fit_resample(X_train, y_train)
Counter(y_resampled)
# -
# Train the Logistic Regression model using the resampled data
so_model = LogisticRegression(solver='lbfgs', random_state=1)
so_model.fit(X_resampled, y_resampled)
# Calculated the balanced accuracy score
y_pred_so = so_model.predict(X_test)
balanced_accuracy_score(y_test, y_pred_so)
# Display the confusion matrix
confusion_matrix(y_test, y_pred_so)
# Print the imbalanced classification report
print(classification_report_imbalanced(y_test, y_pred_so))
# # Undersampling
#
# In this section, you will test an undersampling algorithms to determine which algorithm results in the best performance compared to the oversampling algorithms above. You will undersample the data using the Cluster Centroids algorithm and complete the folliowing steps:
#
# 1. View the count of the target classes using `Counter` from the collections library.
# 3. Use the resampled data to train a logistic regression model.
# 3. Calculate the balanced accuracy score from sklearn.metrics.
# 4. Print the confusion matrix from sklearn.metrics.
# 5. Generate a classication report using the `imbalanced_classification_report` from imbalanced-learn.
#
# Note: Use a random state of 1 for each sampling algorithm to ensure consistency between tests
# +
# Resample the data using the ClusterCentroids resampler
from imblearn.under_sampling import ClusterCentroids
cc = ClusterCentroids(random_state=1)
X_resampled, y_resampled = cc.fit_resample(X_train, y_train)
Counter(y_resampled)
# -
# Train the Logistic Regression model using the resampled data
us_model = LogisticRegression(solver='lbfgs', random_state=1)
us_model.fit(X_resampled, y_resampled)
# Calculated the balanced accuracy score
y_pred_us = us_model.predict(X_test)
balanced_accuracy_score(y_test, y_pred_us)
# Display the confusion matrix
y_pred_us = us_model.predict(X_test)
confusion_matrix(y_test, y_pred_us)
# Print the imbalanced classification report
print(classification_report_imbalanced(y_test, y_pred_us))
# # Combination (Over and Under) Sampling
#
# In this section, you will test a combination over- and under-sampling algorithm to determine if the algorithm results in the best performance compared to the other sampling algorithms above. You will resample the data using the SMOTEENN algorithm and complete the folliowing steps:
#
# 1. View the count of the target classes using `Counter` from the collections library.
# 3. Use the resampled data to train a logistic regression model.
# 3. Calculate the balanced accuracy score from sklearn.metrics.
# 4. Print the confusion matrix from sklearn.metrics.
# 5. Generate a classication report using the `imbalanced_classification_report` from imbalanced-learn.
#
# Note: Use a random state of 1 for each sampling algorithm to ensure consistency between tests
# +
# Resample the training data with SMOTEENN
from imblearn.combine import SMOTEENN
sm = SMOTEENN(random_state=1)
X_resampled, y_resampled = sm.fit_resample(X_train, y_train)
Counter(y_resampled)
# -
# Train the Logistic Regression model using the resampled data
comb_model = LogisticRegression(random_state=1)
comb_model.fit(X_resampled, y_resampled)
# Calculated the balanced accuracy score
y_pred_comb = comb_model.predict(X_test)
balanced_accuracy_score(y_test, y_pred_comb)
# Display the confusion matrix
confusion_matrix(y_test, y_pred_comb)
# Print the imbalanced classification report
print(classification_report_imbalanced(y_test, y_pred_comb))
|
Starter_Code/credit_risk_resampling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Challenge 1 - T-test
#
# In statistics, t-test is used to test if two data samples have a significant difference between their means. There are two types of t-test:
#
# * **Student's t-test** (a.k.a. independent or uncorrelated t-test). This type of t-test is to compare the samples of **two independent populations** (e.g. test scores of students in two different classes). `scipy` provides the [`ttest_ind`](https://docs.scipy.org/doc/scipy-0.15.1/reference/generated/scipy.stats.ttest_ind.html) method to conduct student's t-test.
#
# * **Paired t-test** (a.k.a. dependent or correlated t-test). This type of t-test is to compare the samples of **the same population** (e.g. scores of different tests of students in the same class). `scipy` provides the [`ttest_re`](https://docs.scipy.org/doc/scipy-0.15.1/reference/generated/scipy.stats.ttest_rel.html) method to conduct paired t-test.
#
# Both types of t-tests return a number which is called the **p-value**. If p-value is below 0.05, we can confidently declare the null-hypothesis is rejected and the difference is significant. If p-value is between 0.05 and 0.1, we may also declare the null-hypothesis is rejected but we are not highly confident. If p-value is above 0.1 we do not reject the null-hypothesis.
#
# Read more about the t-test in [this article](https://researchbasics.education.uconn.edu/t-test/) and [this Quora](https://www.quora.com/What-is-the-difference-between-a-paired-and-unpaired-t-test). Make sure you understand when to use which type of t-test.
# +
# Import libraries
import pandas as pd
from scipy import stats
# -
# #### Import dataset
#
# In this challenge we will work on the Pokemon dataset you have used last week. The goal is to test whether different groups of pokemon (e.g. Legendary vs Normal, Generation 1 vs 2, single-type vs dual-type) have different stats (e.g. HP, Attack, Defense, etc.).
# +
# Import dataset
pokemon = pd.read_csv('../../lab-df-calculation-and-transformation/your-code/Pokemon.csv')
pokemon.head()
# -
# #### First we want to define a function with which we can test the means of a feature set of two samples.
#
# In the next cell you'll see the annotations of the Python function that explains what this function does and its arguments and returned value. This type of annotation is called **docstring** which is a convention used among Python developers. The docstring convention allows developers to write consistent tech documentations for their codes so that others can read. It also allows some websites to automatically parse the docstrings and display user-friendly documentations.
#
# Follow the specifications of the docstring and complete the function.
def t_test_features(s1, s2, features=['HP', 'Attack', 'Defense', 'Sp. Atk', 'Sp. Def', 'Speed', 'Total']):
"""Test means of a feature set of two samples
Args:
s1 (dataframe): sample 1
s2 (dataframe): sample 2
features (list): an array of features to test
Returns:
dict: a dictionary of t-test scores for each feature where the feature name is the key and the p-value is the value
"""
# Your code here
results = {}
for feature in features:
results[feature] = stats.ttest_ind(s1[feature], s2[feature], equal_var = False).pvalue
return results
# #### Using the `t_test_features` function, conduct t-test for Lengendary vs non-Legendary pokemons.
#
# *Hint: your output should look like below:*
#
# ```
# {'HP': 1.0026911708035284e-13,
# 'Attack': 2.520372449236646e-16,
# 'Defense': 4.8269984949193316e-11,
# 'Sp. Atk': 1.5514614112239812e-21,
# 'Sp. Def': 2.2949327864052826e-15,
# 'Speed': 1.049016311882451e-18,
# 'Total': 9.357954335957446e-47}
# ```
# +
# Your code here
legendary = pokemon.loc[pokemon['Legendary'] == True]
normal = pokemon.loc[pokemon['Legendary'] == False]
t_test_features(legendary, normal)
# -
# #### From the test results above, what conclusion can you make? Do Legendary and non-Legendary pokemons have significantly different stats on each feature?
# +
# Your comment here
# -
# #### Next, conduct t-test for Generation 1 and Generation 2 pokemons.
# +
# Your code here
type_1 = pokemon.loc[pokemon['Generation'] == 1]
type_2 = pokemon.loc[pokemon['Generation'] == 2]
t_test_features(type_1, type_2)
# -
# #### What conclusions can you make?
# +
# Your comment here
# -
# #### Compare pokemons who have single type vs those having two types.
# +
# Your code here
single_type = pokemon.loc[pokemon['Type 2'].isnull()]
dual_type = pokemon.loc[pokemon['Type 2'].notnull()]
t_test_features(single_type, dual_type)
# -
# #### What conclusions can you make?
# +
# Your comment here
# -
# #### Now, we want to compare whether there are significant differences of `Attack` vs `Defense` and `Sp. Atk` vs `Sp. Def` of all pokemons. Please write your code below.
#
# *Hint: are you comparing different populations or the same population?*
# +
# Your code here
print(stats.ttest_rel(pokemon.Attack, pokemon.Defense))
print(stats.ttest_rel(pokemon['Sp. Atk'], pokemon['Sp. Def']))
# -
# #### What conclusions can you make?
# +
# Your comment here
|
module-2/lab-hypothesis-testing/your-code/challenge-1-solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from packages.nxt import *
brick = connectar(5)
say = Say(brick)
say.hello()
say.game_over()
say.good_job()
say.have_a_nice_day()
sound = Sound(brick)
sound.click()
sound.applause()
sound.arm()
sound.attention()
sound.error()
sound.startup()
desconnectar(brick)
|
Sound Test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
try:
import spnspecs
spnspecs.set_graph_specifications()
except:
spnspecs = None
figpth = '../Figures'
width = 6.8
dpi = 300
def no_smoothing(sat):
f = sat.copy()
f[sat < 0] = 0.
f[sat >= 0] = 1.
return f
def linear_smoothing(sat):
f = sat.copy()
f[sat < 0] = 0.
f[sat > 1] = 1.
return f
def cubic_smoothing(sat, c1=-1, c2=2):
f = c1 * sat**3 + c2 * sat**2
f[sat < 0] = 0
f[sat > 1] = 1
return f
def cubic_smoothing_dervsat(sat, c1=-1, c2=2):
dfdx = 3 * c1 * sat**2 + 2. * c2 * sat
dfdx[sat < 0] = 0
dfdx[sat > 1] = 0
return dfdx
def cubic_smoothingalt(x, xrange, c1=-1, c2=2):
sat = x / xrange
cof1 = c1 / (xrange)**3
cof2 = c2 / (xrange)**2
f = cof1 * x**3 + cof2 * x**2
f[sat < 0] = 0
f[sat > 1] = 1
return f
def cubic_smoothing_dervh(x, xrange, c1=-1, c2=2):
sat = x / xrange
cof1 = 3 * c1 / (xrange)**3
cof2 = 2 * c2 / (xrange)**2
dfdx = cof1 * x**2 + cof2 * x
dfdx[sat < 0] = 0
dfdx[sat > 1] = 0
return dfdx
def fd_derv(x, v):
isize = v.shape[0]
derv = np.ones((isize), dtype=v.dtype) * np.nan
for i in range(1, isize-2):
dv = v[i+1] - v[i-1]
dx = x[i+1] - x[i-1]
derv[i] = dv / dx
return derv
dtype = np.float
step = 0.001
s0, s1 = -0.1, 1.1
s = np.arange(s0, s1, step=step, dtype=dtype)
sp = s.copy()
sp[s < 0] = 0.
sp[s > 1] = 1.
s1, s.shape
plt.plot(s, sp*sp)
f0 = cubic_smoothing(s)
f1 = cubic_smoothing(s, c1=-2., c2=3.)
dseep = 1.
z0 = 0.
xdiff = dseep * s
h = xdiff + z0
dsdh = 1. / dseep
plt.plot(s, sp, lw=1.5, color='black', label='Linear')
plt.plot(s, f0, lw=0.75, color='blue', label='GWSEEP')
plt.plot(s, f1, lw=0.75, color='red', label='WELL')
if spnspecs is not None:
spnspecs.graph_legend(plt.gca())
#plt.xlim(0.9, 1.1)
#plt.ylim(0.9, 1.1)
f0p = cubic_smoothingalt(xdiff, dseep)
f1p = cubic_smoothingalt(xdiff, dseep, c1=-2., c2=3.)
plt.plot(s, f0, lw=0.75, color='blue', label='GWSEEP')
plt.plot(s, f0p, lw=0., color='blue', marker='o', mfc='none', ms=4, markevery=50)
plt.plot(s, f1, lw=0.75, color='red', label='WELL')
plt.plot(s, f1p, lw=0., color='red', marker='o', mfc='none', ms=4, markevery=50)
plt.plot(s, f0-f0p, lw=0.75, color='blue', label='GWSEEP')
plt.plot(s, f1-f1p, lw=0.75, color='red', label='WELL')
spderv = np.ones(s.shape, dtype=np.float)
spderv[s < 0] = 0.
spderv[s > 1] = 0.
f0derv = cubic_smoothing_dervsat(s)
f1derv = cubic_smoothing_dervsat(s, c1=-2., c2=3.)
f0derv2 = cubic_smoothing_dervh(xdiff, dseep)
f1derv2 = cubic_smoothing_dervh(xdiff, dseep, c1=-2., c2=3.)
plt.plot(s, f0derv, lw=4, color='blue', label='GWSEEP', alpha=0.5)
plt.plot(s, f0derv2/dsdh, lw=0.75, color='black', ls='-.') #marker='o', mfc='none', ms=4, markevery=50)
plt.plot(s, f1derv, lw=4, color='red', label='WELL', alpha=0.5)
plt.plot(s, f1derv2/dsdh, lw=0.75, color='black', ls='-.') #marker='o', mfc='none', ms=4, markevery=50)
plt.plot(s, fd_derv(h, f0), lw=0.75, color='blue', label='GWSEEP')
plt.plot(s, fd_derv(h, f1), lw=0.75, color='red', label='WELL')
plt.ylabel(r'$\frac{\partial F}{\partial h}$');
plt.plot(s, fd_derv(h, f0p), lw=0.75, color='blue', label='GWSEEP')
plt.plot(s, fd_derv(h, f1p), lw=0.75, color='red', label='WELL')
plt.ylabel(r'$\frac{\partial F}{\partial h}$');
# ### Plot data
# +
fig, axes = plt.subplots(nrows=1, ncols=2, tight_layout=True, figsize=(width, (1.4/s1) * width/2))
letters = ['A', 'B']
for idx, ax in enumerate(axes):
ax.set_xlim(s0, s1)
ax.set_ylim(s0, 1.4)
ax.set_xticks([0, 0.25, 0.5, 0.75, 1])
ax.set_xticklabels(['0', '0.25', '0.50', '0.75', '1.00'])
ax.set_yticks([0, 0.25, 0.5, 0.75, 1, 1.25])
ax.set_yticklabels(['0', '0.25', '0.50', '0.75', '1.00', '1.25'])
if spnspecs is not None:
spnspecs.remove_edge_ticks(ax)
spnspecs.heading(ax, letter=letters[idx])
ax = axes[0]
ax.axhline(0, lw=0.5, ls='-.', color='black')
ax.axhline(1, lw=0.5, ls='-.', color='black')
ax.axvline(0, lw=0.5, ls='-.', color='black')
ax.axvline(1, lw=0.5, ls='-.', color='black')
ax.plot(s, sp, lw=3.5, color='blue', label=r'$F_{DRN}$')
ax.plot(s, f0, lw=1.75, color='red', label=r'$F_{DRN}^*$')
ax.set_xlabel(r'$\frac{h - ZDRN}{DDRN}$, unitless')
ax.set_ylabel('Discharge scale factor, unitless')
ax = axes[1]
ax.axhline(0, lw=0.5, ls='-.', color='black')
ax.axhline(1, lw=0.5, ls='-.', color='black')
ax.axvline(0, lw=0.5, ls='-.', color='black')
ax.axvline(1, lw=0.5, ls='-.', color='black')
ax.plot([0, 1], [1, 1], lw=3.5, color='blue', label=r'$F_{DRN}$ (linear)')
ax.plot([s0, 0], [0, 0], lw=3.5, color='blue', label=None)
ax.plot([1, s1], [0, 0], lw=3.5, color='blue', label=None)
ax.plot(s[s<=1], f0derv[s<=1], lw=1.75, color='red', label=r'$F_{DRN}^*$ (cubic)')
ax.plot([1, s1], [0, 0], lw=1.75, color='red', label=None)
if spnspecs is not None:
spnspecs.graph_legend(ax, loc='lower right', bbox_to_anchor=(0.9,0.05))
ax.set_xlabel(r'$\frac{h - ZDRN}{DDRN}$, unitless')
ax.set_ylabel('Discharge scale factor derivative with respect to ' + r'$\frac{h - ZDRN}{DDRN}$, unitless')
fpth = os.path.join(figpth, 'DischargeScaleFactor.pdf')
fig.savefig(fpth, dpi=dpi)
# -
dc = np.arange(0, 1, 0.0001)
z = np.zeros(dc.shape, dtype=np.float)
slope = 0.2
dadd = 0.1
d = dadd
for idx in range(1, dc.shape[0]):
v = dc[idx]
if v > d:
slope *= -1.
d += dadd
z[idx] = z[idx - 1] + slope * (v - dc[idx - 1])
z -= z.max() / 2
zmin, zmax = z.min(), z.max()
ymin, ymax = -slope / 10, slope / 10
t = np.arange(s.shape[0]) / s.shape[0]
xdiffq = xdiff.copy()
xdiffq[xdiff < 0] = 0
q0 = no_smoothing(s) * xdiffq
ql = linear_smoothing(s) * xdiffq
qc = cubic_smoothing(s) * xdiffq
arrowprops=dict(arrowstyle='->',
connectionstyle="arc3")
# +
fig = plt.figure(tight_layout=True, figsize=(width, (2/3)*width))
gs = gridspec.GridSpec(2, 3)
axes = [fig.add_subplot(gs[0, :]), fig.add_subplot(gs[1, 0]),
fig.add_subplot(gs[1, 1]), fig.add_subplot(gs[1, 2])]
xlims = [(0., 1.), (0., 1.), (0., 1.), (0., 1.)]
ylims = [(ymax, ymin), (s0, s1), (0, s1), (0, 600)]
letters = ['A', 'B', 'C', 'D']
for idx, ax in enumerate(axes):
ax.set_xlim(xlims[idx])
ax.set_ylim(ylims[idx])
if idx > 0:
ax.set_xticks([0, 0.25, 0.5, 0.75, 1])
ax.set_xticklabels(['0', '0.25', '0.50', '0.75', '1.00'])
else:
ax.tick_params(axis='both', length=0, labelbottom=False, labelleft=False)
if spnspecs is not None:
spnspecs.remove_edge_ticks(ax)
spnspecs.heading(ax, letter=letters[idx])
ax = axes[0]
ax.fill_between(dc, ymax, 0, color='0.9')
# ax.plot(dc, z, lw=1, color='black')
ax.axhline(zmin, lw=0.5, ls='--', color='black')
ax.axhline(zmax, lw=.5, ls='--', color='black')
ax.axhline(0, lw=0.75, color='black')
if spnspecs is not None:
text = 'DDRN'
#spnspecs.add_text(ax, 'DDRN', x=1.01, y=0.5, ha='center', va='center',
# bold=False, rotation=-90)
spnspecs.add_annotation(ax, text=text, xy=(1.01, 0.77), xytext=(1.01, 0.5),
bold=False, rotation=-90, ha='center', va='center',
xycoords='axes fraction', textcoords='axes fraction',
arrowprops=arrowprops)
spnspecs.add_annotation(ax, text=text, xy=(1.01, 0.23), xytext=(1.01, 0.5),
bold=False, rotation=-90, ha='center', va='center',
xycoords='axes fraction', textcoords='axes fraction',
arrowprops=arrowprops)
text = 'Land surface elevation'
# spnspecs.add_annotation(ax, text=text, xy=(0.6, 0.48), xytext=(0.68, 0.9),
# bold=False, ha='center', va='center',
# xycoords='axes fraction', textcoords='axes fraction',
# arrowprops=arrowprops)
spnspecs.add_text(ax, text=text, x=0.5, y=0.5, bold=False, ha='center', va='bottom')
text = 'Cell top'
spnspecs.add_text(ax, text=text, x=0.01, y=0.5, bold=False, ha='left', va='bottom')
text = 'Cell bottom'
spnspecs.add_text(ax, text=text, x=0.01, y=0.01, bold=False, ha='left', va='bottom')
text = r'Land surface elevation + $\frac{DDRN}{2}$'
spnspecs.add_text(ax, text=text, x=0.5, y=zmax, transform=False, bold=False, ha='center')
text = r'Land surface elevation - $\frac{DDRN}{2}$'
spnspecs.add_text(ax, text=text, x=0.5, y=zmin, transform=False, bold=False,
ha='center', va='top')
text = r'$ZDRN + DDN$'
spnspecs.add_text(ax, text=text, x=0.99, y=zmax, transform=False, bold=False, ha='right')
text = r'$ZDRN$'
spnspecs.add_text(ax, text=text, x=0.99, y=zmin, transform=False, bold=False,
ha='right', va='top')
ax = axes[1]
ax.axhline(0, lw=0.5, ls='-.', color='black')
ax.axhline(1, lw=0.5, ls='-.', color='black')
ax.plot(t, s, lw=1.5, color='0.5', ls='--', label=r'$F_{DRN}$')
ax.set_xlabel('Fractional simulation time, unitless')
ax.set_ylabel(r'$\frac{h - ZDRN}{DDRN}$, unitless')
text = r'$h = ZDRN + DDN$'
spnspecs.add_text(ax, text=text, x=0.02, y=.98, transform=False, bold=False, ha='left', va='top')
text = r'$h = ZDRN$'
spnspecs.add_text(ax, text=text, x=0.99, y=0.01, transform=False, bold=False,
ha='right', va='bottom')
ax = axes[2]
ax.plot(t, q0, lw=0.75, color='black')
ax.plot(t, ql, lw=1.5, color='blue')
ax.plot(t, qc, lw=0.75, color='red', label=r'$F_{DRN}^*$')
ax.set_xlabel('Fractional simulation time, unitless')
ax.set_ylabel(r'Drain discharge rate, , L$^3$/T')
text = r'Area = 1 L$^2$' + '\n' + r'K$_v$ = 1 L/T'
if spnspecs is not None:
spnspecs.add_text(ax, text=text, x=0.1, y=0.93, italic=False, bold=False, ha='left', va='top')
ax = axes[3]
ax.plot(t, q0.cumsum(), lw=0.75, color='black', label=r'$F_{DRN}^0$ (original)')
ax.plot(t, ql.cumsum(), lw=1.5, color='blue', label=r'$F_{DRN}$ (linear)')
ax.plot(t, qc.cumsum(), lw=0.75, color='red', label=r'$F_{DRN}^*$ (cubic)')
ax.set_xlabel('Fractional simulation time, unitless')
ax.set_ylabel(r'Cumulative drain discharge, L$^3$')
if spnspecs is not None:
spnspecs.graph_legend(ax, loc='upper left', labelspacing=0.15)
fpth = os.path.join(figpth, 'DRNDischargeDifferences.pdf')
fig.savefig(fpth, dpi=dpi)
# -
q0.max(), ql.max(), qc.max()
q0.cumsum().max(), ql.cumsum().max(), qc.cumsum().max()
|
doc/SuppTechInfo/python/DRN-Discharge-Scaling.ipynb
|
// ---
// jupyter:
// jupytext:
// formats: ipynb,md:myst
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Scala
// language: scala
// name: scala
// ---
// # Introduction to Functional Programming
//
// (Content adapted from Critchlow & Eck)
//
// Functions are fundamental in computer programming,
// although not everything in programming that goes by the name of "function"
// is a function according to the mathematical definition.
//
// In computer programming, a function is a routine that is given
// some data as input and that will calculate and return an
// answer based on that data. For example, in the Java programming
// language, a function that calculates the cube of an integer
// could be written
// ```java
// int cube(int n) {
// return n * n * n;
// }
// ```
// In Java, _int_ is a data type. From the mathematical
// point of view, a data type is a set. The data type _int_
// is the set of all integers that can be represented as 32-bit
// binary numbers. Mathematically, then, $\textit{int}\subseteq\mathbb{Z}$.
// (You should get used to the fact that sets and functions can
// have names that consist of more than one character, since
// it's done all the time in computer programming.)
// The first line of the above function definition,
// "`int cube(int n)`", says that we are defining
// a function named _cube_ whose range is _int_
// and whose domain is _int_. In the usual notation for
// functions, we would express this as $\textit{cube}\colon \textit{int}\to\textit{int}$,
// or possibly as $\textit{cube}\in{\textit{int}}^{\textit{int}}$,
// where ${\textit{int}}^{\textit{int}}$ is the set of all
// functions that map the set _int_ to the set _int_.
//
// The first line of the function, `int cube(int n)`, is called
// the **signature** of the function (in some languages, such as C++, it
// is called the **prototype**). The signature specifies the
// name, the domain, and the range of the function and so carries
// exactly the same information as the notation "$f\colon A\to B$".
// The "$n$" in "`int cube(int n)`" is a name for
// an arbitrary element of the data type _int_. In computer
// jargon, $n$ is called a **parameter** of the function.
// The rest of the definition of _cube_ tells the computer
// to calculate the value of $\textit{cube}(n)$ for any $n\in\textit{int}$
// by multiplying $n\times n\times n$. The statement "`return n * n * n`"
// says that $n\times n\times n$ is the value that is computed, or "returned,"
// by the function. (The $*$ stands for multiplication.)
//
// Java has many data types in addition to _int_. There is
// a boolean data type named _boolean_. The values of type
// _boolean_ are _true_ and _false_. Mathematically,
// _boolean_ is a name for the set $\{\textit{true},\,\textit{false}\}$.
// The type _double_ consists of real numbers, which can
// include a decimal point. Of course, on a computer, it's not
// possible to represent the entire infinite set of real numbers,
// so _double_ represents some subset of the mathematical set
// of real numbers. There is also a data type whose values are
// strings of characters, such as "Hello world" or "xyz152QQZ".
// The name for this data type in Java is _String_. All these
// types, and many others, can be used in functions. For example,
// in Java, $m\,\%\,n$ is the remainder when the integer $m$ is
// divided by the integer $n$. We can define a function to test
// whether an integer is even as follows:
// ```java
// boolean even(int k) {
// if ( k % 2 == 1 )
// return false;
// else
// return true;
// }
// ```
// You don't need to worry about all the details here, but you should
// understand that the signature, `boolean even(int k)`,
// says that _even_ is a function from the set _int_
// to the set _boolean_. That is,
// $\textit{even}\colon\textit{int}\to\textit{boolean}$. Given
// an integer $N$, $\textit{even}(N)$ has the value _true_
// if $N$ is an even integer, and it has the value _false_
// if $N$ is an odd integer.
//
// A function can have more than one parameter. For example, we might
// define a function with signature `int index(String str, String sub)`.
// If $s$ and $t$ are strings, then $\textit{index}(s,t)$ would be the
// _int_ that is the value of the function at the ordered pair
// $(s,t)$. We see that the domain of _index_ is the cross product
// $\textit{String}\times\textit{String}$, and we can write
// $\textit{index}\colon \textit{String}\times\textit{String}\to\textit{int}$
// or, equivalently, $\textit{index}\in\textit{int}^{\textit{String}\times\textit{String}}$.
//
// ## Partial and Impure Functions
//
// Not every Java function is actually a function in the mathematical
// sense. In mathematics, a function must associate a single value in
// its range to each value in its domain. There are two things
// that can go wrong: The value of the function might not be defined
// for every element of the domain, and the function might associate
// several different values to the same element of the domain.
// Both of these things can happen with Java functions.
//
// In computer programming, it is very common for a "function" to be
// undefined for some values of its parameter. In mathematics,
// a **partial function** from a set $A$ to
// a set $B$ is defined to be a function from a subset of $A$ to $B$.
// A partial function from $A$ to $B$ can be undefined for some
// elements of $A$, but when it is defined for some $a\in A$,
// it associates just one element of $B$ to $a$. Many functions
// in computer programs are actually partial functions. (When
// dealing with partial functions, an ordinary function, which is
// defined for every element of its domain, is sometimes referred to
// as a **total function**. Note that—with the mind-boggling
// logic that is typical of mathematicians—a total function is
// a type of partial function, because a set is a subset of itself.)
//
// It's also very common for a "function" in a computer program
// to produce a variety of values for the same value of its parameter.
// This most frequently occurs because the function is **impure**—either
// it has a **side-effect** (such as changing the value of a global
// variable) beyond just computing a result, or it relies on a value that
// has been changed as a side-effect of some other part of the program.
// Another way to say that a function is impure is that it depends on a
// **hidden state**—some extra information affecting the result that
// is not provided by the function's direct inputs.
// A common example is the method in the _java.util.Random_ class with signature
// `int nextInt(int N)`, which returns a random integer between
// 0 and $N-1$. The value of _nextInt_(5) could be 0, 1, 2, 3, or 4.
// This is not the behavior of a mathematical function! Behind the scenes,
// this function is storing a variable called the "seed", which is used to
// generate a random value, plus a new value for the seed, each time the
// function is called. Each time the seed changes, the next value to be
// returned is likely to change.
//
// Even though many functions in computer programs are not really
// mathematical functions, I will continue to refer to them as
// functions in this section. Mathematicians will just have to stretch
// their definitions a bit to accommodate the realities of computer
// programming.
//
// ## Pure Functional Programming
//
// Unlike Java, a typical functional programming language such as Scala,
// Haskell, or ReasonML will actively discourage the use of side-effects
// in functions.[^io] The benefit of restricting the programmer to
// **pure** functions, that always return the same value for a given
// argument, is that it becomes possible to reason about the behavior
// **algebraically**, freely substituting the returned values in place
// of function calls without having to worry about whether some "hidden state"
// might have changed since the last time the function was called.
//
// For example, suppose that we have a function that computes the value of
// some polynomial, such as $f(x)=x^2+2x+1$. If we know that another function
// $g\colon\textit{String}\to\textit{int}$ is pure, then we can be sure
// that $f(g(s))$ is the same as $g(s)^2+2\cdot g(s)+1$, as well as
// $(g(s)+1)^2$. All of these are algebraically equivalent, as long as $g$
// is pure. This allows the programmer to reason more easily about the
// correctness of their program, and it also enables the computer to choose
// any equivalent expression for evaluation, ideally choosing the most
// efficient version.
//
// One of the most common ways that a functional language will encourage
// pure functions is to do away with, or at least severely restrict, the
// ability to update the value assigned to a variable. In pure Scala, there
// is no assignment statement. When a value is bound to a variable with a
// `val` statement, that variable will then remain bound to that value for
// as long as the variable exists. A variable will cease to exist when the
// block (such as a function body) containing it is finished:
{
val x = 42
println(x) // prints 42
}
// x no longer exists here
// A variable may be temporarily **shadowed** by another variable with the same
// name. This may look like an assignment of a changed value to a variable,
// but each use of the `val` statement will create a new named location in
// memory; if the shadowing variable goes away, the original one will become
// visible again with its correct value:
val x = 42
println(x) // prints 42
{
val x = 17 // shadows earlier definition of x
println(x) // prints 17
}
println(x) // prints 42 again
// Again, this behavior permits algebraic reasoning about the program. The above
// code is equivalent to
val x = 42
println(x)
{
val y = 17
println(y)
}
println(x)
// where we have uniformly renamed the inner variable _x_ as _y_ to make it clear
// that they are distinct variables. It is also equivalent to
println(42)
println(17)
println(42)
// where we have replaced each use of our identifiers with its value. The output is
// slightly different in this case, because we no longer have the top-level
// binding of 42 to _x_ that would have been available if we wrote additional lines
// at the bottom of the program, but it is equivalent if we just look at the printed
// results.
//
// ## First-Class Functions
//
// In most programming languages, functions are not first-class
// objects. That is, a function cannot
// be treated as a data value in the same way as a _String_
// or an _int_. However, recent versions of Java have taken a step in this
// direction. It is possible for a function to be a parameter
// to another function, as long as it is wrapped in a "function object".
// For example, consider the function signature
// ```java
// int sumten( Function<Integer, Integer> f )
// ```
// This is a signature for a function named _sumten_ whose
// parameter is a function object. The parameter is specified by the
// type "`Function<Integer, Integer>`". If _S_ and _T_ are types, then
// the type `Function<S, T>` represents functions from _S_ to _T_. Therefore,
// the parameter of _sumten_ is essentially a function from _int_ to _int_.[^int]
// The parameter name, $f$, stands for an arbitrary such function. Mathematically,
// $f\in \textit{int}^{\textit{int}}$, and so
// $\textit{sumten}\colon \textit{int}^{\textit{int}}\to\textit{int}$.
//
// My idea is that $\textit{sumten}(f)$ would compute
// $f(1)+f(2)+\cdots+f(10)$. A more useful function would
// be able to compute $f(a)+f(a+1)+\cdots+f(b)$ for any integers
// $a$ and $b$. This just means that $a$ and $b$ should be
// parameters to the function. The signature for the improved
// function would look like
// ```java
// int sum( Function<Integer, Integer> f, int a, int b )
// ```
// The parameters to _sum_ form an ordered triple in which
// the first coordinate is a function and the second and third
// coordinates are integers. So, we could write
// $$
// \textit{sum}\colon \textit{int}^{\textit{int}}
// \times\textit{int}\times\textit{int}\to\textit{int}
// $$
// It's interesting that computer programmers deal routinely
// with such complex objects.
//
// There are several ways of providing a function object as an
// argument in Java. If we want to pass the method _m_ of an object
// _x_, where the signature of _m_ is `int m( int i )`, then
// we can call our function as `sum(x::m, a, b)`. However, a more
// general technique is to use an **anonymous function**, also known
// as a **lambda**.[^lambda]
//
// ## Anonymous functions
//
// In Java, an expression such as `i -> { return i * i * i; }`
// creates a function object that takes an _int_ (this will be
// determined from the context) and returns another _int_. This
// particular function cubes its input. Thus, if we call our _sum_ function as
// follows:
// ```java
// sum(i -> { return i * i * i; }, 3, 5)
// ```
// the result will be $3^3 + 4^3 + 5^3$,
// which is 216.
//
// Many languages now support a similar syntax for creating anonymous
// function values, and offer some facility for working with functions
// as (mostly) first-class objects. For example, the same function
// is expressed in Scala as `(i: Int) => { i * i * i }`. Since one of the hallmarks of
// the functional languages is their ability to work with function
// values, you can imagine that they tend to provide the most
// thorough integration of functions with other kinds of values.
//
// Here are complete demos of the _sum_ example, first in Java and
// then in Scala:
//
// ```java
// import java.util.function.Function;
//
// public class SumDemo {
// private static int sum(Function<Integer, Integer> f, int a, int b) {
// int total = 0;
// for (int i = a; i <= b; i++) {
// total += f.apply(i);
// }
// return total;
// }
//
// public static void main(String[] args) {
// System.out.println(sum(i -> { return i * i * i; }, 3, 5));
// }
// }
// ```
// +
def sum(f: Int => Int, a: Int, b: Int): Int = {
if (a > b) {
0
} else {
f(a) + sum(f, a+1, b)
}
}
println(sum((i: Int) => { i * i * i }, 3, 5))
// -
// **REWRITE THIS SECTION**
//
// Note that we define the function _sum_ in ReasonML by binding
// a function value to the name _sum_, just as we can bind other
// types of values to identifiers. The keyword "rec" after the "let"
// indicates that the right-hand side expression is allowed to make
// use of the name that is currently being defined, so that we can
// make our function recursive. Without the "rec", any use of _sum_
// in the right-hand expression would refer to an older binding to
// that name.
//
// As a simpler function definition in ReasonML, not needing the "rec"
// keyword, here is our _cube_ function again:
// ```reason demo
// let cube = n => { n * n * n };
// ```
// This is just binding the anonymous function we have been using above
// to the name _cube_. Note that the function `n => { n * n * n }` is
// exactly the same as the function `i => { i * i * i }`, because the name
// of the parameter does not matter outside the function.
//
// An interesting fact about ReasonML is that the operators are also functions,
// bound to names made out of operator symbols instead of letters and digits. To
// refer to an operator as a function value, just put the operator in parentheses:
// `(+)`, `(*)`, `(==)`, …. Therefore, an expression such as `a + b * c`
// can also be written as `(+)(a, (*)(b, c))` (note that this takes into account
// the usual higher precedence of multiplication over addition).
// For example, if we wanted to define an exponentiation operator on _int_, and
// call it `***`, we could define it as follows:[^4]
//
// [^4]: The code here is based on the solution to an exercise in the [Recursion](../logic/recursion) section.]
//
// ```reason demo
// let rec (***) = (n, p) => {
// if (p == 0) {
// 1
// } else if (p mod 2 == 0) {
// (n * n) *** (p / 2)
// } else {
// n * (n *** (p - 1))
// }
// };
// ```
//
// It is even possible in functional languages for a function to return
// another function as its value. For example,
// ```reason demo
// let monomial = (a, n) => {
// x => { a * x *** n }
// };
// ```
// Here, `x *** n` is our exponentiation operator from above, which computes $x^n$, so for any
// integers $a$ and $n$, the value of $\textit{monomial}(a,n)$ is
// a function that computes $ax^n$. Thus,
// ```reason edit
// let f = monomial(2, 3);
// ```
// would define $f$ to be the function that satisfies $f(x)=2x^3$. This is
// now ready to be handed to our _sum_ function:
// ```reason edit
// print_int( sum( f, 3, 6 ) );
// ```
// would compute $2*3^3+2*4^3+2*5^3+2*6^3$. In fact, _monomial_
// can be used to create an unlimited number of new functions
// from scratch. It is even possible to write _monomial_(2, 3)(5)
// to indicate the result of applying the function _monomial_(2, 3)
// to the value 5. The value represented by _monomial_(2, 3)(5)
// is $2*5^3$, or 250. This is real functional programming and
// might give you some idea of its power.
//
// ## Exercises
//
// 1. For each of the following Java function signatures, translate the
// signature into a standard mathematical function specification, such
// as $\textit{func}\colon\textit{double}\to\textit{int}$.
// * `int strlen(String s)`
// * `double pythag(double x, double y)`
// * `int round(double x)`
// * `String sub(String s, int n, int m)`
// * `String unlikely(Function<String, Integer> f)`
// * `int h(Function<Integer, Integer> f, Function<Integer, Integer> g)`
//
// 2. Write a Java function signature for a function that
// belongs to each of the following sets.
//
// * $\textit{String}^{\textit{String}}$
// * $\textit{boolean}^{\textit{double}\times\textit{double}}$
// * $\textit{double}^{ \textit{int}^{\textit{int}} }$
//
// 3. It is possible to define new types in Java. For example, the
// definition
// ```java
// public class Point {
// public double x;
// public double y;
// }
// ```
// defines a new type named _Point_. A value of type _Point_
// contains two values of type _double_. What mathematical operation
// corresponds to the construction of this data type? Why?
//
// 4. Let _cube_, _sum_ and _monomial_
// be the ReasonML functions described in this section. What is the
// value of each of the following?
// * _sum_(_cube_, 2, 4)
// * _sum_(_monomial_(5, 2), 1, 3)
// * _monomial_(_cube_(2), 7)
// * _sum_($n\Rightarrow\{ 2*n \}$, 1, 5)
// * _cube_(_sum_(_monomial_(2, 3), 1, 2))
// }
//
// 5. Write a ReasonML function named _compose_
// that computes the composition of two functions. That
// is, $\textit{compose}(f, g)$ is $f\circ g$, where
// $f$ and $g$ are functions of one parameter. Recall that
// $f\circ g$ is the function defined by $(f\circ g)(x)=f(g(x))$.
//
// 6. Consider the following ReasonML function:
// ```reason edit
// let exercise = (a, b) => {
// let m = (x, y) => { x - y };
// let s = n => { n * n };
// let c = m(a, b);
// let d = m(a, c);
// let e = c + d;
// s(d) - s(e)
// };
// ```
// * What is the value of `exercise(4, 5)`?
// * What is the value of `exercise(12, 13)`?
// * Use algebraic substitution to evaluate `exercise(a, b)` in terms of the
// variables `a` and `b`.
//
// [^io]: The most common exception is for functions that send
// output to the console, such as the `print_int` function in ReasonML.
// Being able to track execution and easily display results is very
// useful, and printing a line on the console is a fairly benign
// side-effect—it won't cause the function to return different
// values for the same arguments. However, printing a result can still
// interfere with algebraic reasoning about a program, because interchanging
// such a function call with its value can affect whether and how many times
// the output is printed.
//
// [^int]: For our purposes we may ignore the distinction between _int_ and _Integer_ in Java.
//
// [^lambda]: The mathematician Alonzo Church introduced in
// the 1930's the use of the Greek letter lambda ($\lambda$) to indicate an
// otherwise unnamed function defined by a formula. That is, instead
// of writing "the function $f$ where $f(x) = \textit{some formula}$",
// he wrote "$\lambda x(\textit{some formula})$". When the first
// functional programming language, LISP (invented in the late 1950's),
// needed a way to create function values, <NAME> adopted Church's
// use of lambda, and the name has stuck.
|
book/fp/intro.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
class LetYourImaginationBeYourGuide:
pass
obj = LetYourImaginationBeYourGuide()
obj.this_class = "rules"
obj.this_class
import numpy as np
class MyDataset:
def __init__(self, size):
self.data = np.random.random(size)
self.size = size
def __add__(self, other):
obj = MyDataset(self.size)
obj.data = self.data + other.data
return obj
def plot(self, filename, ylabel = "", xlabel="x"):
plt.clf()
plt.plot(self.data)
plt.ylabel(ylabel)
plt.xlabel(xlabel)
plt.savefig(filename)
def __repr__(self):
return "Hi there!"
my_dataset = MyDataset(128)
my_dataset.plot("hi.png", xlabel = "my_x")
my_other_dataset = MyDataset(128)
my_dataset.data[0], my_other_dataset.data[0]
print(my_dataset)
my_dataset + my_other_dataset
a = MyDataset(128)
b = MyDataset(256)
c = a + b
d = b + a
class BigMistake:
def __init__(self, default_values = []):
self.default_values = default_values[:]
mistake1 = BigMistake()
mistake2 = BigMistake()
mistake1.
mistake2.something_else
mistake1.something_else[3] = 4
mistake2.something_else
mistake1.something_else is mistake2.something_else
mistake1.default_values.append(" SPOILER ALERT ")
mistake1.default_values
mistake2.default_values
|
week05/examples_lecture07_classes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bootstrap fit exploration
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
import os
import re
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
# catalogues to compare to
simard_df = pd.read_csv('lib/simard-catalog_fixed-n.csv', index_col=0)
simard_df_free = pd.read_csv('lib/simard-catalog_free-n.csv', index_col=0)
lackner_nb1 = pd.read_csv('lib/lackner_nb1.csv', index_col=0)
lackner_nb4 = pd.read_csv('lib/lackner_nb4.csv', index_col=0)
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
catalog_results = pd.concat((
simard_df[['__B_T_r', 'e__B_T_r']]
.rename(columns=dict(__B_T_r='Sbt', e__B_T_r='eSbt')),
simard_df_free[['__B_T_r', 'e__B_T_r']]
.rename(columns=dict(__B_T_r='Sbt_free', e__B_T_r='eSbt_free')),
lackner_nb1[['BULGE_TO_TOT_R', 'BULGE_TO_TOT_R_ERR']]
.rename(columns=dict(BULGE_TO_TOT_R='Lbt1', BULGE_TO_TOT_R_ERR='eLbt1')),
lackner_nb4[['BULGE_TO_TOT_R', 'BULGE_TO_TOT_R_ERR']]
.rename(columns=dict(BULGE_TO_TOT_R='Lbt4', BULGE_TO_TOT_R_ERR='eLbt4')),
), axis=1)
display_column_map = dict(
Sbt='Exponential + De Vaucoulers\n(Simard, 2011)',
Sbt_free='Exponential + Sersic\n(Simard, 2011)',
Lbt1='Exponential + Exponential\n(Lackner, 2012)',
Lbt4='Exponential + De Vaucoulers\n(Lackner, 2012)',
)
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
minima_info_nb1 = {
int(k.split('.')[0]): pd.read_csv(os.path.join('2comp_fits/minima', k), index_col=0)
for k in os.listdir('2comp_fits/minima')
if re.match(r'[0-9]+\.csv', k)
}
bt_nb1 = pd.DataFrame({
k: v.query('accepted == True and bulge_frac > 0')['bulge_frac']
for k, v in minima_info_nb1.items()
}).replace(0, np.nan)
best_bt_nb1 = pd.DataFrame({
k: v.query('accepted == True and bulge_frac > 0').sort_values(by='chisq')['bulge_frac']
for k, v in minima_info_nb1.items()
}).replace(0, np.nan).iloc[0].dropna()
closest_index_nb1 = np.abs(
bt_nb1 - catalog_results['Lbt1'].reindex_like(bt_nb1.T)
).idxmin().dropna()
closest_df_nb1 = pd.Series({
i: minima_info_nb1[i].loc[int(closest_index_nb1[i])]
for i in closest_index_nb1.index
}).apply(pd.Series).T
closest_bt_nb1 = closest_df_nb1.loc['bulge_frac']
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
plt.figure(figsize=(8, 8))
plt.scatter(
catalog_results['Lbt1'].reindex_like(closest_bt_nb1),
closest_bt_nb1,
marker='o', ec='C0', s=8**2, alpha=0.8,
label='Closest minimum to Lackner (2012) exp+exp model',
)
plt.scatter(
catalog_results['Lbt1'].reindex_like(best_bt_nb1),
best_bt_nb1,
marker='x', ec='C1', s=8**2,
label=r'Lowest $\chi_\nu^2$'
)
plt.plot([0, 1], [0, 1], c='k', alpha=0.2)
plt.ylim(0., 1)
plt.xlim(0., 1)
plt.legend();
# -
# What if we restrict to models Lackner & Gunn identified as Exp + Exp models?
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
lackner_nb1['TYPE'].value_counts()
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
best_bt_nb1_masked = best_bt_nb1.where(lackner_nb1['TYPE'] == 'nb1').dropna()
closest_bt_nb1_masked = closest_bt_nb1.where(lackner_nb1['TYPE'] == 'nb1').dropna()
plt.figure(figsize=(8, 8))
plt.scatter(
catalog_results['Lbt1'].reindex_like(closest_bt_nb1_masked),
closest_bt_nb1_masked,
marker='o', ec='C0', s=8**2, alpha=0.8,
label='Closest minimum to Lackner (2012) exp+exp model',
)
plt.scatter(
catalog_results['Lbt1'].reindex_like(best_bt_nb1_masked),
best_bt_nb1_masked,
marker='x', ec='C1', s=8**2,
label=r'Lowest $\chi_\nu^2$'
)
plt.plot([0, 1], [0, 1], c='k', alpha=0.2)
plt.ylim(0., 1)
plt.xlim(0., 1)
plt.legend();
# -
# Better, but this is to be expected, as Lackner & Gunn are more likely to have found the global minimum in the cases where they select exp+exp as the best model
# ### Equivalent for exp+deV models
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
minima_info_nb4 = {
int(k.split('.')[0]): pd.read_csv(os.path.join('2comp_fits_nb4/minima', k), index_col=0)
for k in os.listdir('2comp_fits_nb4/minima')
if re.match(r'[0-9]+\.csv', k)
}
bt_nb4 = pd.DataFrame({
k: v.query('accepted == True and bulge_frac > 0')['bulge_frac']
for k, v in minima_info_nb4.items()
}).replace(0, np.nan)
best_bt_nb4 = pd.DataFrame({
k: v.query('accepted == True and bulge_frac > 0').sort_values(by='chisq')['bulge_frac']
for k, v in minima_info_nb4.items()
}).replace(0, np.nan).iloc[0].dropna()
closest_index_nb4 = np.abs(
bt_nb4 - catalog_results['Lbt4'].reindex_like(bt_nb4.T)
).idxmin().dropna()
closest_df_nb4 = pd.Series({
i: minima_info_nb4[i].loc[int(closest_index_nb4[i])]
for i in closest_index_nb4.index
}).apply(pd.Series).T
closest_bt_nb4 = closest_df_nb4.loc['bulge_frac']
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
plt.figure(figsize=(8, 8))
plt.scatter(
catalog_results['Lbt4'].reindex_like(closest_bt_nb4),
closest_bt_nb4,
marker='o', ec='C0', s=8**2, alpha=0.8,
label='Closest minimum to Lackner (2012) exp+deV model'
)
plt.scatter(
catalog_results['Lbt4'].reindex_like(best_bt_nb4),
best_bt_nb4,
marker='x', ec='C1', s=8**2,
label=r'Lowest $\chi_\nu^2$'
)
plt.plot([0, 1], [0, 1], c='k', alpha=0.2)
plt.ylim(0., 1)
plt.xlim(0., 1)
plt.legend();
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
best_bt_nb4_masked = best_bt_nb4.where(lackner_nb1['TYPE'] == 'nb4').dropna()
closest_bt_nb4_masked = closest_bt_nb4.where(lackner_nb1['TYPE'] == 'nb4').dropna()
plt.figure(figsize=(8, 8))
plt.scatter(
catalog_results['Lbt4'].reindex_like(closest_bt_nb4_masked),
closest_bt_nb4_masked,
marker='o', ec='C0', s=8**2, alpha=0.8,
label='Closest minimum to Lackner (2012) exp+deV model'
)
plt.scatter(
catalog_results['Lbt4'].reindex_like(best_bt_nb4_masked),
best_bt_nb4_masked,
marker='x', ec='C1', s=8**2,
label=r'Lowest $\chi_\nu^2$'
)
plt.plot([0, 1], [0, 1], c='k', alpha=0.2)
plt.ylim(0., 1)
plt.xlim(0., 1)
plt.legend();
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
closest_index_simard = np.abs(
bt_nb4 - catalog_results['Sbt'].reindex_like(bt_nb4.T)
).idxmin().dropna()
closest_df_simard = pd.Series({
i: minima_info_nb4[i].loc[int(closest_index_simard[i])]
for i in closest_index_simard.index
}).apply(pd.Series).T
closest_bt_simard = closest_df_simard.loc['bulge_frac']
plt.figure(figsize=(8, 8))
plt.scatter(
catalog_results['Lbt4'].reindex_like(closest_bt_simard),
closest_bt_simard,
marker='o', ec='C0', s=8**2, alpha=0.8,
label='Closest minimum to Simard (2011) exp+deV model'
)
plt.scatter(
catalog_results['Lbt4'].reindex_like(best_bt_nb4),
best_bt_nb4,
marker='x', ec='C1', s=8**2,
label=r'Lowest $\chi_\nu^2$'
)
plt.plot([0, 1], [0, 1], c='k', alpha=0.2)
plt.ylim(0., 1)
plt.xlim(0., 1)
plt.legend();
# + inputHidden=false jupyter={"outputs_hidden": false} outputHidden=false
|
2comp_results.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: dog-project
# language: python
# name: dog-project
# ---
# # Artificial Intelligence Nanodegree
#
# ## Convolutional Neural Networks
#
# ## Project: Write an Algorithm for a Dog Identification App
#
# ---
#
# In this notebook, some template code has already been provided for you, and you will need to implement additional functionality to successfully complete this project. You will not need to modify the included code beyond what is requested. Sections that begin with **'(IMPLEMENTATION)'** in the header indicate that the following block of code will require additional functionality which you must provide. Instructions will be provided for each section, and the specifics of the implementation are marked in the code block with a 'TODO' statement. Please be sure to read the instructions carefully!
#
# > **Note**: Once you have completed all of the code implementations, you need to finalize your work by exporting the iPython Notebook as an HTML document. Before exporting the notebook to html, all of the code cells need to have been run so that reviewers can see the final implementation and output. You can then export the notebook by using the menu above and navigating to \n",
# "**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
#
# In addition to implementing code, there will be questions that you must answer which relate to the project and your implementation. Each section where you will answer a question is preceded by a **'Question X'** header. Carefully read each question and provide thorough answers in the following text boxes that begin with **'Answer:'**. Your project submission will be evaluated based on your answers to each of the questions and the implementation you provide.
#
# >**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. Markdown cells can be edited by double-clicking the cell to enter edit mode.
#
# The rubric contains _optional_ "Stand Out Suggestions" for enhancing the project beyond the minimum requirements. If you decide to pursue the "Stand Out Suggestions", you should include the code in this IPython notebook.
#
#
#
# ---
# ### Why We're Here
#
# In this notebook, you will make the first steps towards developing an algorithm that could be used as part of a mobile or web app. At the end of this project, your code will accept any user-supplied image as input. If a dog is detected in the image, it will provide an estimate of the dog's breed. If a human is detected, it will provide an estimate of the dog breed that is most resembling. The image below displays potential sample output of your finished project (... but we expect that each student's algorithm will behave differently!).
#
# 
#
# In this real-world setting, you will need to piece together a series of models to perform different tasks; for instance, the algorithm that detects humans in an image will be different from the CNN that infers dog breed. There are many points of possible failure, and no perfect algorithm exists. Your imperfect solution will nonetheless create a fun user experience!
#
# ### The Road Ahead
#
# We break the notebook into separate steps. Feel free to use the links below to navigate the notebook.
#
# * [Step 0](#step0): Import Datasets
# * [Step 1](#step1): Detect Humans
# * [Step 2](#step2): Detect Dogs
# * [Step 3](#step3): Create a CNN to Classify Dog Breeds (from Scratch)
# * [Step 4](#step4): Use a CNN to Classify Dog Breeds (using Transfer Learning)
# * [Step 5](#step5): Create a CNN to Classify Dog Breeds (using Transfer Learning)
# * [Step 6](#step6): Write your Algorithm
# * [Step 7](#step7): Test Your Algorithm
#
# ---
# <a id='step0'></a>
# ## Step 0: Import Datasets
#
# ### Import Dog Dataset
#
# In the code cell below, we import a dataset of dog images. We populate a few variables through the use of the `load_files` function from the scikit-learn library:
# - `train_files`, `valid_files`, `test_files` - numpy arrays containing file paths to images
# - `train_targets`, `valid_targets`, `test_targets` - numpy arrays containing onehot-encoded classification labels
# - `dog_names` - list of string-valued dog breed names for translating labels
# +
from sklearn.datasets import load_files
from keras.utils import np_utils
import numpy as np
from glob import glob
# define function to load train, test, and validation datasets
def load_dataset(path):
data = load_files(path)
dog_files = np.array(data['filenames'])
dog_targets = np_utils.to_categorical(np.array(data['target']), 133)
return dog_files, dog_targets
# load train, test, and validation datasets
train_files, train_targets = load_dataset('dogImages/train')
valid_files, valid_targets = load_dataset('dogImages/valid')
test_files, test_targets = load_dataset('dogImages/test')
# load list of dog names
dog_names = [item[20:-1] for item in sorted(glob("dogImages/train/*/"))]
# print statistics about the dataset
print('There are %d total dog categories.' % len(dog_names))
print('There are %s total dog images.\n' % len(np.hstack([train_files, valid_files, test_files])))
print('There are %d training dog images.' % len(train_files))
print('There are %d validation dog images.' % len(valid_files))
print('There are %d test dog images.'% len(test_files))
# -
# ### Import Human Dataset
#
# In the code cell below, we import a dataset of human images, where the file paths are stored in the numpy array `human_files`.
# +
import random
random.seed(8675309)
# load filenames in shuffled human dataset
human_files = np.array(glob("lfw/*/*"))
random.shuffle(human_files)
# print statistics about the dataset
print('There are %d total human images.' % len(human_files))
# -
# ---
# <a id='step1'></a>
# ## Step 1: Detect Humans
#
# We use OpenCV's implementation of [Haar feature-based cascade classifiers](http://docs.opencv.org/trunk/d7/d8b/tutorial_py_face_detection.html) to detect human faces in images. OpenCV provides many pre-trained face detectors, stored as XML files on [github](https://github.com/opencv/opencv/tree/master/data/haarcascades). We have downloaded one of these detectors and stored it in the `haarcascades` directory.
#
# In the next code cell, we demonstrate how to use this detector to find human faces in a sample image.
# +
import cv2
import matplotlib.pyplot as plt
# %matplotlib inline
# extract pre-trained face detector
face_cascade = cv2.CascadeClassifier('haarcascades/haarcascade_frontalface_alt.xml')
# load color (BGR) image
img = cv2.imread(human_files[3])
# convert BGR image to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
# find faces in image
faces = face_cascade.detectMultiScale(gray)
# print number of faces detected in the image
print('Number of faces detected:', len(faces))
# get bounding box for each detected face
for (x,y,w,h) in faces:
# add bounding box to color image
cv2.rectangle(img,(x,y),(x+w,y+h),(255,0,0),2)
# convert BGR image to RGB for plotting
cv_rgb = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
# display the image, along with bounding box
plt.imshow(cv_rgb)
plt.show()
# -
# Before using any of the face detectors, it is standard procedure to convert the images to grayscale. The `detectMultiScale` function executes the classifier stored in `face_cascade` and takes the grayscale image as a parameter.
#
# In the above code, `faces` is a numpy array of detected faces, where each row corresponds to a detected face. Each detected face is a 1D array with four entries that specifies the bounding box of the detected face. The first two entries in the array (extracted in the above code as `x` and `y`) specify the horizontal and vertical positions of the top left corner of the bounding box. The last two entries in the array (extracted here as `w` and `h`) specify the width and height of the box.
#
# ### Write a Human Face Detector
#
# We can use this procedure to write a function that returns `True` if a human face is detected in an image and `False` otherwise. This function, aptly named `face_detector`, takes a string-valued file path to an image as input and appears in the code block below.
# returns "True" if face is detected in image stored at img_path
def face_detector(img_path):
img = cv2.imread(img_path)
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
faces = face_cascade.detectMultiScale(gray)
return len(faces) > 0
# ### (IMPLEMENTATION) Assess the Human Face Detector
#
# __Question 1:__ Use the code cell below to test the performance of the `face_detector` function.
# - What percentage of the first 100 images in `human_files` have a detected human face?
# - What percentage of the first 100 images in `dog_files` have a detected human face?
#
# Ideally, we would like 100% of human images with a detected face and 0% of dog images with a detected face. You will see that our algorithm falls short of this goal, but still gives acceptable performance. We extract the file paths for the first 100 images from each of the datasets and store them in the numpy arrays `human_files_short` and `dog_files_short`.
#
# __Answer:__ 97% of the first 100 human images have a detected human face, and 11% of the first 100 dog images have a detected human face.
# +
human_files_short = human_files[:100]
dog_files_short = train_files[:100]
# Do NOT modify the code above this line.
## TODO: Test the performance of the face_detector algorithm
## on the images in human_files_short and dog_files_short.
human_as_human = len(list(x for x in map(face_detector, human_files_short) if x))/100
dog_as_human = len(list(x for x in map(face_detector, dog_files_short) if x))/100
print(human_as_human, dog_as_human)
# -
# __Question 2:__ This algorithmic choice necessitates that we communicate to the user that we accept human images only when they provide a clear view of a face (otherwise, we risk having unneccessarily frustrated users!). In your opinion, is this a reasonable expectation to pose on the user? If not, can you think of a way to detect humans in images that does not necessitate an image with a clearly presented face?
#
# __Answer:__ Given that the function is only meant to detect whether a human face is present, I think this is not a reasonable expectation to pose on the user. We can further train the detector with blurred face images such that it could recognise other (more abstract) features, for example the shape of the head.
#
# We suggest the face detector from OpenCV as a potential way to detect human images in your algorithm, but you are free to explore other approaches, especially approaches that make use of deep learning :). Please use the code cell below to design and test your own face detection algorithm. If you decide to pursue this _optional_ task, report performance on each of the datasets.
# +
## (Optional) TODO: Report the performance of another
## face detection algorithm on the LFW dataset
### Feel free to use as many code cells as needed.
# -
# ---
# <a id='step2'></a>
# ## Step 2: Detect Dogs
#
# In this section, we use a pre-trained [ResNet-50](http://ethereon.github.io/netscope/#/gist/db945b393d40bfa26006) model to detect dogs in images. Our first line of code downloads the ResNet-50 model, along with weights that have been trained on [ImageNet](http://www.image-net.org/), a very large, very popular dataset used for image classification and other vision tasks. ImageNet contains over 10 million URLs, each linking to an image containing an object from one of [1000 categories](https://gist.github.com/yrevar/942d3a0ac09ec9e5eb3a). Given an image, this pre-trained ResNet-50 model returns a prediction (derived from the available categories in ImageNet) for the object that is contained in the image.
# +
from keras.applications.resnet50 import ResNet50
# define ResNet50 model
ResNet50_model = ResNet50(weights='imagenet')
# -
# ### Pre-process the Data
#
# When using TensorFlow as backend, Keras CNNs require a 4D array (which we'll also refer to as a 4D tensor) as input, with shape
#
# $$
# (\text{nb_samples}, \text{rows}, \text{columns}, \text{channels}),
# $$
#
# where `nb_samples` corresponds to the total number of images (or samples), and `rows`, `columns`, and `channels` correspond to the number of rows, columns, and channels for each image, respectively.
#
# The `path_to_tensor` function below takes a string-valued file path to a color image as input and returns a 4D tensor suitable for supplying to a Keras CNN. The function first loads the image and resizes it to a square image that is $224 \times 224$ pixels. Next, the image is converted to an array, which is then resized to a 4D tensor. In this case, since we are working with color images, each image has three channels. Likewise, since we are processing a single image (or sample), the returned tensor will always have shape
#
# $$
# (1, 224, 224, 3).
# $$
#
# The `paths_to_tensor` function takes a numpy array of string-valued image paths as input and returns a 4D tensor with shape
#
# $$
# (\text{nb_samples}, 224, 224, 3).
# $$
#
# Here, `nb_samples` is the number of samples, or number of images, in the supplied array of image paths. It is best to think of `nb_samples` as the number of 3D tensors (where each 3D tensor corresponds to a different image) in your dataset!
# +
from keras.preprocessing import image
from tqdm import tqdm
def path_to_tensor(img_path):
# loads RGB image as PIL.Image.Image type
img = image.load_img(img_path, target_size=(224, 224))
# convert PIL.Image.Image type to 3D tensor with shape (224, 224, 3)
x = image.img_to_array(img)
# convert 3D tensor to 4D tensor with shape (1, 224, 224, 3) and return 4D tensor
return np.expand_dims(x, axis=0)
def paths_to_tensor(img_paths):
list_of_tensors = [path_to_tensor(img_path) for img_path in tqdm(img_paths)]
return np.vstack(list_of_tensors)
# -
# ### Making Predictions with ResNet-50
#
# Getting the 4D tensor ready for ResNet-50, and for any other pre-trained model in Keras, requires some additional processing. First, the RGB image is converted to BGR by reordering the channels. All pre-trained models have the additional normalization step that the mean pixel (expressed in RGB as $[103.939, 116.779, 123.68]$ and calculated from all pixels in all images in ImageNet) must be subtracted from every pixel in each image. This is implemented in the imported function `preprocess_input`. If you're curious, you can check the code for `preprocess_input` [here](https://github.com/fchollet/keras/blob/master/keras/applications/imagenet_utils.py).
#
# Now that we have a way to format our image for supplying to ResNet-50, we are now ready to use the model to extract the predictions. This is accomplished with the `predict` method, which returns an array whose $i$-th entry is the model's predicted probability that the image belongs to the $i$-th ImageNet category. This is implemented in the `ResNet50_predict_labels` function below.
#
# By taking the argmax of the predicted probability vector, we obtain an integer corresponding to the model's predicted object class, which we can identify with an object category through the use of this [dictionary](https://gist.github.com/yrevar/942d3a0ac09ec9e5eb3a).
# +
from keras.applications.resnet50 import preprocess_input, decode_predictions
def ResNet50_predict_labels(img_path):
# returns prediction vector for image located at img_path
img = preprocess_input(path_to_tensor(img_path))
return np.argmax(ResNet50_model.predict(img))
# -
# ### Write a Dog Detector
#
# While looking at the [dictionary](https://gist.github.com/yrevar/942d3a0ac09ec9e5eb3a), you will notice that the categories corresponding to dogs appear in an uninterrupted sequence and correspond to dictionary keys 151-268, inclusive, to include all categories from `'Chihuahua'` to `'Mexican hairless'`. Thus, in order to check to see if an image is predicted to contain a dog by the pre-trained ResNet-50 model, we need only check if the `ResNet50_predict_labels` function above returns a value between 151 and 268 (inclusive).
#
# We use these ideas to complete the `dog_detector` function below, which returns `True` if a dog is detected in an image (and `False` if not).
### returns "True" if a dog is detected in the image stored at img_path
def dog_detector(img_path):
prediction = ResNet50_predict_labels(img_path)
return ((prediction <= 268) & (prediction >= 151))
# ### (IMPLEMENTATION) Assess the Dog Detector
#
# __Question 3:__ Use the code cell below to test the performance of your `dog_detector` function.
# - What percentage of the images in `human_files_short` have a detected dog?
# - What percentage of the images in `dog_files_short` have a detected dog?
#
# __Answer:__ A dog is detected in 2% of the human_files_short images and 100% of the dog_files_short images respectively.
### TODO: Test the performance of the dog_detector function
### on the images in human_files_short and dog_files_short.
human_as_dog = sum(1 for x in map(dog_detector, human_files_short) if x)/100
dog_as_dog = sum(1 for x in map(dog_detector, dog_files_short) if x)/100
print(human_as_dog, dog_as_dog)
# ---
# <a id='step3'></a>
# ## Step 3: Create a CNN to Classify Dog Breeds (from Scratch)
#
# Now that we have functions for detecting humans and dogs in images, we need a way to predict breed from images. In this step, you will create a CNN that classifies dog breeds. You must create your CNN _from scratch_ (so, you can't use transfer learning _yet_!), and you must attain a test accuracy of at least 1%. In Step 5 of this notebook, you will have the opportunity to use transfer learning to create a CNN that attains greatly improved accuracy.
#
# Be careful with adding too many trainable layers! More parameters means longer training, which means you are more likely to need a GPU to accelerate the training process. Thankfully, Keras provides a handy estimate of the time that each epoch is likely to take; you can extrapolate this estimate to figure out how long it will take for your algorithm to train.
#
# We mention that the task of assigning breed to dogs from images is considered exceptionally challenging. To see why, consider that *even a human* would have great difficulty in distinguishing between a Brittany and a Welsh Springer Spaniel.
#
# Brittany | Welsh Springer Spaniel
# - | -
# <img src="images/Brittany_02625.jpg" width="100"> | <img src="images/Welsh_springer_spaniel_08203.jpg" width="200">
#
# It is not difficult to find other dog breed pairs with minimal inter-class variation (for instance, Curly-Coated Retrievers and American Water Spaniels).
#
# Curly-Coated Retriever | American Water Spaniel
# - | -
# <img src="images/Curly-coated_retriever_03896.jpg" width="200"> | <img src="images/American_water_spaniel_00648.jpg" width="200">
#
#
# Likewise, recall that labradors come in yellow, chocolate, and black. Your vision-based algorithm will have to conquer this high intra-class variation to determine how to classify all of these different shades as the same breed.
#
# Yellow Labrador | Chocolate Labrador | Black Labrador
# - | -
# <img src="images/Labrador_retriever_06457.jpg" width="150"> | <img src="images/Labrador_retriever_06455.jpg" width="240"> | <img src="images/Labrador_retriever_06449.jpg" width="220">
#
# We also mention that random chance presents an exceptionally low bar: setting aside the fact that the classes are slightly imabalanced, a random guess will provide a correct answer roughly 1 in 133 times, which corresponds to an accuracy of less than 1%.
#
# Remember that the practice is far ahead of the theory in deep learning. Experiment with many different architectures, and trust your intuition. And, of course, have fun!
#
# ### Pre-process the Data
#
# We rescale the images by dividing every pixel in every image by 255.
# +
from PIL import ImageFile
ImageFile.LOAD_TRUNCATED_IMAGES = True
# pre-process the data for Keras
train_tensors = paths_to_tensor(train_files).astype('float32')/255
valid_tensors = paths_to_tensor(valid_files).astype('float32')/255
test_tensors = paths_to_tensor(test_files).astype('float32')/255
# -
# ### (IMPLEMENTATION) Model Architecture
#
# Create a CNN to classify dog breed. At the end of your code cell block, summarize the layers of your model by executing the line:
#
# model.summary()
#
# We have imported some Python modules to get you started, but feel free to import as many modules as you need. If you end up getting stuck, here's a hint that specifies a model that trains relatively fast on CPU and attains >1% test accuracy in 5 epochs:
#
# 
#
# __Question 4:__ Outline the steps you took to get to your final CNN architecture and your reasoning at each step. If you chose to use the hinted architecture above, describe why you think that CNN architecture should work well for the image classification task.
#
# __Answer:__ I started with an architecture with the one hinted above because convolutional neural networks are generally good at image classification, the convolutional layers can each specialise on recognising specific features in the images. In addition, the layers in my CNN has more filters and my CNN contains an extra layer to increase the number of trainable parameters, but at the same time have added dropout layers an an attempt to reduce overfitting.
# +
from keras.layers import Conv2D, MaxPooling2D, GlobalAveragePooling2D
from keras.layers import Dropout, Flatten, Dense
from keras.models import Sequential
model = Sequential()
### TODO: Define your architecture.
model.add(Conv2D(filters=32, kernel_size=2, padding='same', activation='relu',
input_shape=(224, 224, 3)))
model.add(MaxPooling2D(pool_size=2))
model.add(Conv2D(filters=64, kernel_size=2, padding='same', activation='relu'))
model.add(MaxPooling2D(pool_size=2))
model.add(Conv2D(filters=64, kernel_size=2, padding='same', activation='relu'))
model.add(MaxPooling2D(pool_size=2))
model.add(Conv2D(filters=32, kernel_size=2, padding='same', activation='relu'))
model.add(GlobalAveragePooling2D())
model.add(Dropout(0.4))
model.add(Dense(133, activation='softmax'))
model.summary()
# -
# ### Compile the Model
model.compile(optimizer='rmsprop', loss='categorical_crossentropy', metrics=['accuracy'])
# ### (IMPLEMENTATION) Train the Model
#
# Train your model in the code cell below. Use model checkpointing to save the model that attains the best validation loss.
#
# You are welcome to [augment the training data](https://blog.keras.io/building-powerful-image-classification-models-using-very-little-data.html), but this is not a requirement.
# +
from keras.callbacks import ModelCheckpoint
### TODO: specify the number of epochs that you would like to use to train the model.
epochs = 5
### Do NOT modify the code below this line.
checkpointer = ModelCheckpoint(filepath='saved_models/weights.best.from_scratch.hdf5',
verbose=1, save_best_only=True)
model.fit(train_tensors, train_targets,
validation_data=(valid_tensors, valid_targets),
epochs=epochs, batch_size=50, callbacks=[checkpointer], verbose=1)
# -
# ### Load the Model with the Best Validation Loss
model.load_weights('saved_models/weights.best.from_scratch.hdf5')
# ### Test the Model
#
# Try out your model on the test dataset of dog images. Ensure that your test accuracy is greater than 1%.
# +
# get index of predicted dog breed for each image in test set
dog_breed_predictions = [np.argmax(model.predict(np.expand_dims(tensor, axis=0))) for tensor in test_tensors]
# report test accuracy
test_accuracy = 100*np.sum(np.array(dog_breed_predictions)==np.argmax(test_targets, axis=1))/len(dog_breed_predictions)
print('Test accuracy: %.4f%%' % test_accuracy)
# -
# ---
# <a id='step4'></a>
# ## Step 4: Use a CNN to Classify Dog Breeds
#
# To reduce training time without sacrificing accuracy, we show you how to train a CNN using transfer learning. In the following step, you will get a chance to use transfer learning to train your own CNN.
#
# ### Obtain Bottleneck Features
bottleneck_features = np.load('bottleneck_features/DogVGG16Data.npz')
train_VGG16 = bottleneck_features['train']
valid_VGG16 = bottleneck_features['valid']
test_VGG16 = bottleneck_features['test']
# ### Model Architecture
#
# The model uses the the pre-trained VGG-16 model as a fixed feature extractor, where the last convolutional output of VGG-16 is fed as input to our model. We only add a global average pooling layer and a fully connected layer, where the latter contains one node for each dog category and is equipped with a softmax.
# +
VGG16_model = Sequential()
VGG16_model.add(GlobalAveragePooling2D(input_shape=train_VGG16.shape[1:]))
VGG16_model.add(Dense(133, activation='softmax'))
VGG16_model.summary()
# -
# ### Compile the Model
VGG16_model.compile(loss='categorical_crossentropy', optimizer='rmsprop', metrics=['accuracy'])
# ### Train the Model
# +
checkpointer = ModelCheckpoint(filepath='saved_models/weights.best.VGG16.hdf5',
verbose=1, save_best_only=True)
VGG16_model.fit(train_VGG16, train_targets,
validation_data=(valid_VGG16, valid_targets),
epochs=20, batch_size=20, callbacks=[checkpointer], verbose=1)
# -
# ### Load the Model with the Best Validation Loss
VGG16_model.load_weights('saved_models/weights.best.VGG16.hdf5')
# ### Test the Model
#
# Now, we can use the CNN to test how well it identifies breed within our test dataset of dog images. We print the test accuracy below.
# +
# get index of predicted dog breed for each image in test set
VGG16_predictions = [np.argmax(VGG16_model.predict(np.expand_dims(feature, axis=0))) for feature in test_VGG16]
# report test accuracy
test_accuracy = 100*np.sum(np.array(VGG16_predictions)==np.argmax(test_targets, axis=1))/len(VGG16_predictions)
print('Test accuracy: %.4f%%' % test_accuracy)
# -
# ### Predict Dog Breed with the Model
# +
from extract_bottleneck_features import *
def VGG16_predict_breed(img_path):
# extract bottleneck features
bottleneck_feature = extract_VGG16(path_to_tensor(img_path))
# obtain predicted vector
predicted_vector = VGG16_model.predict(bottleneck_feature)
# return dog breed that is predicted by the model
return dog_names[np.argmax(predicted_vector)]
# -
# ---
# <a id='step5'></a>
# ## Step 5: Create a CNN to Classify Dog Breeds (using Transfer Learning)
#
# You will now use transfer learning to create a CNN that can identify dog breed from images. Your CNN must attain at least 60% accuracy on the test set.
#
# In Step 4, we used transfer learning to create a CNN using VGG-16 bottleneck features. In this section, you must use the bottleneck features from a different pre-trained model. To make things easier for you, we have pre-computed the features for all of the networks that are currently available in Keras:
# - [VGG-19](https://s3-us-west-1.amazonaws.com/udacity-aind/dog-project/DogVGG19Data.npz) bottleneck features
# - [ResNet-50](https://s3-us-west-1.amazonaws.com/udacity-aind/dog-project/DogResnet50Data.npz) bottleneck features
# - [Inception](https://s3-us-west-1.amazonaws.com/udacity-aind/dog-project/DogInceptionV3Data.npz) bottleneck features
# - [Xception](https://s3-us-west-1.amazonaws.com/udacity-aind/dog-project/DogXceptionData.npz) bottleneck features
#
# The files are encoded as such:
#
# Dog{network}Data.npz
#
# where `{network}`, in the above filename, can be one of `VGG19`, `Resnet50`, `InceptionV3`, or `Xception`. Pick one of the above architectures, download the corresponding bottleneck features, and store the downloaded file in the `bottleneck_features/` folder in the repository.
#
# ### (IMPLEMENTATION) Obtain Bottleneck Features
#
# In the code block below, extract the bottleneck features corresponding to the train, test, and validation sets by running the following:
#
# bottleneck_features = np.load('bottleneck_features/Dog{network}Data.npz')
# train_{network} = bottleneck_features['train']
# valid_{network} = bottleneck_features['valid']
# test_{network} = bottleneck_features['test']
### TODO: Obtain bottleneck features from another pre-trained CNN.
bottleneck_features = np.load('bottleneck_features/DogXceptionData.npz')
train_Xception = bottleneck_features['train']
valid_Xception = bottleneck_features['valid']
test_Xception = bottleneck_features['test']
# ### (IMPLEMENTATION) Model Architecture
#
# Create a CNN to classify dog breed. At the end of your code cell block, summarize the layers of your model by executing the line:
#
# <your model's name>.summary()
#
# __Question 5:__ Outline the steps you took to get to your final CNN architecture and your reasoning at each step. Describe why you think the architecture is suitable for the current problem.
#
# __Answer:__ We started with the Xception model and the output is being fed to a global average pooling layer, which is further connected to a dense layer. The Xception model is pretrained with data from ImageNet, it is highly likely that the first few convolution layer is capable of identifying simple features (e.g. lines, shapes) within an image, which should be relevant to our task. I believe one dense layer would be sufficient provided that our dog images are possibly similar to subsets from ImageNet. The dense layer has 133 nodes which correspond to the number of dog categories of interest.
# +
### TODO: Define your architecture.
Xception_model = Sequential()
Xception_model.add(GlobalAveragePooling2D(input_shape=train_Xception.shape[1:]))
Xception_model.add(Dense(133, activation='softmax'))
Xception_model.summary()
# -
# ### (IMPLEMENTATION) Compile the Model
### TODO: Compile the model.
Xception_model.compile(loss='categorical_crossentropy', optimizer='rmsprop', metrics=['accuracy'])
# ### (IMPLEMENTATION) Train the Model
#
# Train your model in the code cell below. Use model checkpointing to save the model that attains the best validation loss.
#
# You are welcome to [augment the training data](https://blog.keras.io/building-powerful-image-classification-models-using-very-little-data.html), but this is not a requirement.
# +
### TODO: Train the model.
checkpointer = ModelCheckpoint(filepath='saved_models/weights.best.Xception.hdf5',
verbose=1, save_best_only=True)
Xception_model.fit(train_Xception, train_targets,
validation_data=(valid_Xception, valid_targets),
epochs=20, batch_size=20, callbacks=[checkpointer], verbose=1)
# -
# ### (IMPLEMENTATION) Load the Model with the Best Validation Loss
### TODO: Load the model weights with the best validation loss.
Xception_model.load_weights('saved_models/weights.best.Xception.hdf5')
# ### (IMPLEMENTATION) Test the Model
#
# Try out your model on the test dataset of dog images. Ensure that your test accuracy is greater than 60%.
# +
### TODO: Calculate classification accuracy on the test dataset.
Xception_predictions = [np.argmax(Xception_model.predict(np.expand_dims(feature, axis=0))) for feature in test_Xception]
# report test accuracy
test_accuracy = 100*np.sum(np.array(Xception_predictions)==np.argmax(test_targets, axis=1))/len(Xception_predictions)
print('Test accuracy: %.4f%%' % test_accuracy)
# -
# ### (IMPLEMENTATION) Predict Dog Breed with the Model
#
# Write a function that takes an image path as input and returns the dog breed (`Affenpinscher`, `Afghan_hound`, etc) that is predicted by your model.
#
# Similar to the analogous function in Step 5, your function should have three steps:
# 1. Extract the bottleneck features corresponding to the chosen CNN model.
# 2. Supply the bottleneck features as input to the model to return the predicted vector. Note that the argmax of this prediction vector gives the index of the predicted dog breed.
# 3. Use the `dog_names` array defined in Step 0 of this notebook to return the corresponding breed.
#
# The functions to extract the bottleneck features can be found in `extract_bottleneck_features.py`, and they have been imported in an earlier code cell. To obtain the bottleneck features corresponding to your chosen CNN architecture, you need to use the function
#
# extract_{network}
#
# where `{network}`, in the above filename, should be one of `VGG19`, `Resnet50`, `InceptionV3`, or `Xception`.
### TODO: Write a function that takes a path to an image as input
### and returns the dog breed that is predicted by the model.
def Xception_predict_breed(img_path):
bottleneck_feature = extract_Xception(path_to_tensor(img_path))
predicted_vector = Xception_model.predict(bottleneck_feature)
return dog_names[np.argmax(predicted_vector)]
# ---
# <a id='step6'></a>
# ## Step 6: Write your Algorithm
#
# Write an algorithm that accepts a file path to an image and first determines whether the image contains a human, dog, or neither. Then,
# - if a __dog__ is detected in the image, return the predicted breed.
# - if a __human__ is detected in the image, return the resembling dog breed.
# - if __neither__ is detected in the image, provide output that indicates an error.
#
# You are welcome to write your own functions for detecting humans and dogs in images, but feel free to use the `face_detector` and `dog_detector` functions developed above. You are __required__ to use your CNN from Step 5 to predict dog breed.
#
# Some sample output for our algorithm is provided below, but feel free to design your own user experience!
#
# 
#
#
# ### (IMPLEMENTATION) Write your Algorithm
# +
### TODO: Write your algorithm.
### Feel free to use as many code cells as needed.
from PIL import Image # I commented this out so my notebook doesn't require image to run
from matplotlib.pyplot import imshow, show
def resembling_dog_breed(img_path):
#image = Image.open(img_path)
#imshow(np.asarray(image))
#show()
if (dog_detector(img_path)):
breed = Xception_predict_breed(img_path)
print('Detected a dog which looks like a', breed)
elif (face_detector(img_path)):
breed = Xception_predict_breed(img_path)
print('Detected a human who resembles a ', breed)
else:
print('Unable to detect a human nor dog in the input image.')
# -
# ---
# <a id='step7'></a>
# ## Step 7: Test Your Algorithm
#
# In this section, you will take your new algorithm for a spin! What kind of dog does the algorithm think that __you__ look like? If you have a dog, does it predict your dog's breed accurately? If you have a cat, does it mistakenly think that your cat is a dog?
#
# ### (IMPLEMENTATION) Test Your Algorithm on Sample Images!
#
# Test your algorithm at least six images on your computer. Feel free to use any images you like. Use at least two human and two dog images.
#
# __Question 6:__ Is the output better than you expected :) ? Or worse :( ? Provide at least three possible points of improvement for your algorithm.
#
# __Answer:__ The output is actually better than I expected. It misclassified my dog as a human but is able to identify the correct breed (golden retriever). It is also able to tell when the input is neither a human nor dog.
# - I could have augmented the training data such that the dog breed prediction may have a higher accuracy.
# - Evaluate both the probability of the image containing a human or dog to improve the chances to correctly identifying the subject within the input image (e.g. there could be cases where both detectors return true).
# - Train the network with images containing a human with other animals.
## TODO: Execute your algorithm from Step 6 on
## at least 6 images on your computer.
## Feel free to use as many code cells as needed.
img_paths = [
'images/nova_bb.jpg', # golden retriever
'images/nova.jpg', # golden retriever
'images/alex.jpg', # poddle
'images/lamb.jpg', # toy
'images/corgi.jpg', # corgi
'images/peanut.jpg', # cat
'images/lya.jpg', # human
'images/kty.jpg' # human
]
for path in img_paths:
resembling_dog_breed(path)
Xception_predict_breed('images/peanut.jpg')
|
dog_app.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + language="javascript"
#
# IPython.OutputArea.prototype._should_scroll = function(lines) {
# return false;
# }
# -
# %aiida
import ipywidgets as ipw
from IPython.display import clear_output
from aiida.cmdline.utils.ascii_vis import format_call_graph
import urllib.parse as urlparse
from aiidalab_widgets_base import ProcessFollowerWidget, ProgressBarWidget, ProcessReportWidget
from aiidalab_widgets_base import ProcessInputsWidget, ProcessOutputsWidget, ProcessCallStackWidget, RunningCalcJobOutputWidget
url = urlparse.urlsplit(jupyter_notebook_url)
url_dict = urlparse.parse_qs(url.query)
if 'id' in url_dict:
pk = int(url_dict['id'][0])
process = load_node(pk)
else:
process = None
# ## Process inputs.
display(ProcessInputsWidget(process))
# ## Process outputs.
display(ProcessOutputsWidget(process))
follower = ProcessFollowerWidget(
process,
followers=[ProgressBarWidget(), ProcessReportWidget(), ProcessCallStackWidget(), RunningCalcJobOutputWidget()], path_to_root='../',
update_interval=2)
display(follower)
follower.follow(detach=True)
|
process.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .fs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: F#
// language: fsharp
// name: ifsharp
// ---
// Surprisingly there are only three numbers that can be written as the sum of fourth powers of their digits:
//
// 1634 = 1^4 + 6^4 + 3^4 + 4^4
// 8208 = 8^4 + 2^4 + 0^4 + 8^4
// 9474 = 9^4 + 4^4 + 7^4 + 4^4
// As 1 = 1^4 is not a sum it is not included.
//
// The sum of these numbers is `1634 + 8208 + 9474 = 19316`.
//
// Find the sum of all the numbers that can be written as the sum of fifth powers of their digits.
//
//
// +
let raiseTo pow n =
double(n) ** double(pow)
let sumPowerDigits pow n =
string(n).ToCharArray()
|> Array.map (string >> int >> (raiseTo pow))
|> Array.sum
|> int
[2..1000000] // assume less than 1 million...
|> List.map (fun n -> (n, sumPowerDigits 5 n))
|> List.filter (fun (a,b) -> a = b)
|> List.map fst
|> List.sum
// -
|
Problem 030 - Digit fifth powers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="89B27-TGiDNB"
# ## Imports
# + colab_type="code" id="U6rgexPXmY37" outputId="526059ec-83ae-4c68-b14d-22171c551d50" colab={"base_uri": "https://localhost:8080/", "height": 51}
import os, sys, math
import numpy as np
from matplotlib import pyplot as plt
if 'google.colab' in sys.modules: # Colab-only Tensorflow version selector
# %tensorflow_version 2.x
import tensorflow as tf
print("Tensorflow version " + tf.__version__)
# + cellView="form" colab_type="code" id="MPkvHdAYNt9J" colab={}
#@title "display utilities [RUN ME]"
def display_9_images_from_dataset(dataset):
plt.figure(figsize=(13,13))
subplot=331
for i, (image, label) in enumerate(dataset):
plt.subplot(subplot)
plt.axis('off')
plt.imshow(image.numpy().astype(np.uint8))
plt.title(label.numpy().decode("utf-8"), fontsize=16)
subplot += 1
if i==8:
break
plt.tight_layout()
plt.subplots_adjust(wspace=0.1, hspace=0.1)
plt.show()
# + [markdown] colab_type="text" id="w9S3uKC_iXY5"
# ## Configuration
# + colab_type="code" id="d8K6hL_kiWve" colab={}
GCS_PATTERN = 'gs://flowers-public/*/*.jpg'
CLASSES = ['daisy', 'dandelion', 'roses', 'sunflowers', 'tulips'] # flower labels (folder names in the data)
# + [markdown] colab_type="text" id="kvPXiovhi3ZZ"
# ## Read images and labels [WORK REQUIRED]
# 1. Use `fileset=`[`tf.data.Dataset.list_files`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#list_files) to scan the data folder
# 1. Iterate through the dataset of filenames: `for filename in fileset:...` .
# * Does it work ? Yes, but if you print the filename you get Tensors containing strings.
# * To display the string only, you can use filename.numpy(). This works on any Tensorflow tensor.
# * tip: to limit the size of the dataset for display, you can use [`Dataset.take()`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#take). Like this: `for data in dataset.take(10): ....`
# 1. Use [`tf.data.Dataset.map`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#map) to decode the JPEG files. You will find useful TF code snippets below.
# * Iterate on the image dataset. You can use .numpy().shape to only see the data sizes.
# * Are all images of the same size ?
# 1. Now create a training dataset: you have images but you also need labels:
# * the labels (flower names) are the directory names. You will find useful TF code snippets below for parsing them.
# * If you do "`return image, label`" in the decoding function, you will have a Dataset of pairs (image, label).
# 1. You can see the flowers and their labels with the `display_9_images_from_dataset` function. It expects the Dataset to have `(image, label)` elements.
# + colab_type="code" id="nwsZ8X59mu24" colab={}
nb_images = len(tf.io.gfile.glob(GCS_PATTERN))
print("Pattern matches {} images.".format(nb_images))
#
# YOUR CODE GOES HERE
#
#display_9_images_from_dataset(dataset)
# + [markdown] colab_type="text" id="ZX6Vg0YZwRCP"
# ## Useful code snippets
# + [markdown] colab_type="text" id="FszNm593wnky"
# ### Decode a JPEG in Tensorflow
# + colab_type="code" id="pmPz2WM2wTbS" colab={}
def decode_jpeg(filename):
bits = tf.io.read_file(filename)
image = tf.image.decode_jpeg(bits)
return image
# + [markdown] colab_type="text" id="Dax9B6W7wuxt"
# ### Decode a JPEG and extract folder name in TF
# + colab_type="code" id="kPhQl3BlxB7D" colab={}
def decode_jpeg_and_label(filename):
bits = tf.io.read_file(filename)
image = tf.image.decode_jpeg(bits)
# parse flower name from containing directory
label = tf.strings.split(tf.expand_dims(filename, axis=-1), sep='/')
label = label.values[-2]
return image, label
# + [markdown] id="6UuMEBez45s5" colab_type="text"
# ## License
#
#
#
# ---
#
#
# author: <NAME><br>
# twitter: @martin_gorner
#
#
# ---
#
#
# Copyright 2020 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
#
# ---
#
#
# This is not an official Google product but sample code provided for an educational purpose
#
|
courses/fast-and-lean-data-science/02_Dataset_playground.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python_3.6
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import mglearn
from IPython.display import display
# %matplotlib inline
# -
# # Generalization, Overfitting, and Underfitting
#
# ## Generalization, when the model is able to predict new data from traning data
# Overfitting, when the model contains more than the needed features to be considered on the predition phase, so this prevent us from get correct predittions with new data
# Underfitting, when we don't use enough features to predict correct the data
#
# The sweet spot is the best place for new data prediction
from IPython.display import Image
Image(filename="images/mode_complexity.png")
X, y = mglearn.datasets.make_forge()
# plot dataset
mglearn.discrete_scatter(X[:, 0], X[:, 1], y)
plt.legend(["Class 0", "Class 1"], loc=4)
plt.xlabel("First feature")
plt.ylabel("Second feature")
print("X.shape: {}".format(X.shape))
plt.show()
# this dataset consists of 26 data points, with 2 features
X, y = mglearn.datasets.make_wave(n_samples=40)
plt.plot(X, y, 'o')
plt.ylim(-3, 3)
plt.xlabel("Feature")
plt.ylabel("Target")
# Let's use the previous data set with the cancer dataset so we can do some analysis
from sklearn.datasets import load_breast_cancer
cancer = load_breast_cancer()
print("Cancer.keys(): \n{}".format(cancer.keys()))
print("Shape of cancer data: {}".format(cancer.data.shape))
print("Sample counts per class:\n{}".format({n: v
for n, v in zip(cancer.target_names, np.bincount(cancer.target))}))
print("Feture names:\n{}".format(cancer.feature_names))
# Dataset for Boston, the task is to predict the median value of home in sveral Boston neighborhoods in the 70's
from sklearn.datasets import load_boston
boston = load_boston()
print("Boston Data Shape: {}".format(boston.data.shape))
# We will actually expand this dataset by not only considering these 13 measurements as input features, but also looking at all products (also called interactions) between features. In other words, we will not only consider crime rate and highway accessibility as features, but also the product of crime rate and highway accessibility. Including derived feature like these is called feature engineering
X, y = mglearn.datasets.load_extended_boston()
print("X.shape: {}".format(X.shape))
|
Source/Chapter02/01 supervisedMLAlgorithms.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Do
# - run all subjects
# - subsample trials because e.g., S08 and S09 don't have the same number of trials
# - take that code, and plug in other classifiers
# - use on more subjects, each subject one by one
# - concatenate all the electrodes together across subjects, and see what you find
#
# ### Done
# - j-remi classifier over time
# - try S09 as well
# ### Import packages
# +
# general packages
import numpy as np
import matplotlib.pyplot as plt
# sklearn models
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LogisticRegression
# mne
import mne
import pickle
# edit here
# sample dataset
from mne.datasets import sample
from mne.decoding import (SlidingEstimator, GeneralizingEstimator,
cross_val_multiscore, LinearModel, get_coef)
# -
# # Load preprocessed data
# ## How can I load the pickled data or otherwise the dictionary?
with open('../data/decision-making/data/data_preproc/X.pickle', 'rb') as handle:
X = pickle.load(handle)
with open('../data/decision-making/data/data_preproc/y.pickle', 'rb') as handle:
y = pickle.load(handle)
X[0].shape
y[0].shape
# +
# # Save
# dictionary = {'hello':'world'}
# np.save('my_file.npy', dictionary)
# Load
# read_dictionary = np.load('../data/decision-making/data/data_preproc/X.npy')
# print(read_dictionary['hello'])
# -
read_dictionary.shape
favorite_color = pickle.load( open( '../data/decision-making/data/data_preproc/X.pickle', "rb" ) )
import pickle
with open('../data/decision-making/data/data_preproc/X.pickle', 'rb') as handle:
X = pickle.load(handle)
X[9].shape
target = '../data/decision-making/data/data_preproc/X.pickle'
if os.path.getsize(target) > 0:
with open(target, "rb") as f:
unpickler = pickle.Unpickler(f)
# if file is not empty scores will be equal
# to the value unpickled
scores = unpickler.load()
with open('../data/decision-making/data/data_preproc/X.pickle', 'wb') as handle:
pickle.load('../data/decision-making/data/data_preproc/X.pickle')
X = pickle.load('../data/decision-making/data/data_preproc/X.pickle')
X1 = np.load('../data/decision-making/data/data_preproc/X1.npy')
X8 = np.load('../data/decision-making/data/data_preproc/X8.npy')
# X9 = np.load('../data/decision-making/data/data_preproc/X9.npy')
X1.shape
# +
# X8.shape, X9.shape
# n_epochs (200) x n_channels (10) x n_times (3001)
# -
y1 = np.load('../data/decision-making/data/data_preproc/y1.npy')
y8 = np.load('../data/decision-making/data/data_preproc/y8.npy')
y9 = np.load('../data/decision-making/data/data_preproc/y9.npy')
y1.shape
y8.shape, y9.shape
# n_epochs (200)
# ## Try decoding exercise:
# https://martinos.org/mne/stable/auto_tutorials/plot_sensors_decoding.html#sphx-glr-auto-tutorials-plot-sensors-decoding-py
import warnings
warnings.filterwarnings("ignore")
# ## X8 and X9: decoding over time
X8.shape, y8.shape, # n_channels, n_times
X9.shape, y9.shape, # n_channels, n_times
np.unique(y9)
# +
# Might need to convert "gamble" to "1", and "safebet" to 0
# -
# ### subject 1
# +
# properties of model
clf = make_pipeline(StandardScaler(), LogisticRegression())
time_decod = SlidingEstimator(clf, n_jobs=1, scoring='roc_auc')
# notice subject here
scores = cross_val_multiscore(time_decod, X1, y1, cv=5, n_jobs=1)
# Mean scores across cross-validation splits
scores = np.mean(scores, axis=0)
# -
# Plot
fig, ax = plt.subplots()
# ax.plot(epochs.times, scores, label='score')
ax.plot(scores, label='score')
ax.axhline(.5, color='k', linestyle='--', label='chance')
ax.set_xlabel('Times')
ax.set_ylabel('AUC') # Area Under the Curve
ax.legend()
ax.axvline(1000, color='k', linestyle='-')
ax.set_title('Logistic Regression')
plt.show()
# ### X8
# +
# properties of model
clf = make_pipeline(StandardScaler(), LogisticRegression())
time_decod = SlidingEstimator(clf, n_jobs=1, scoring='roc_auc')
# replacing X and y with X8 and y8
scores = cross_val_multiscore(time_decod, X8, y8, cv=5, n_jobs=1)
# Mean scores across cross-validation splits
scores = np.mean(scores, axis=0)
# -
# Plot
fig, ax = plt.subplots()
# ax.plot(epochs.times, scores, label='score')
ax.plot(scores, label='score')
ax.axhline(.5, color='k', linestyle='--', label='chance')
ax.set_xlabel('Times')
ax.set_ylabel('AUC') # Area Under the Curve
ax.legend()
ax.axvline(1000, color='k', linestyle='-')
ax.set_title('Logistic Regression')
plt.show()
# ## X9
# +
# properties of model
clf = make_pipeline(StandardScaler(), LogisticRegression())
time_decod = SlidingEstimator(clf, n_jobs=1, scoring='roc_auc')
# replacing X and y with X9 and y9
scores = cross_val_multiscore(time_decod, X9, y9, cv=5, n_jobs=1)
# Mean scores across cross-validation splits
scores = np.mean(scores, axis=0)
# -
# Plot
fig, ax = plt.subplots()
# ax.plot(epochs.times, scores, label='score')
ax.plot(scores, label='score')
ax.axhline(.5, color='k', linestyle='--', label='chance')
ax.set_xlabel('Times')
ax.set_ylabel('AUC') # Area Under the Curve
ax.legend()
ax.axvline(1000, color='k', linestyle='-')
ax.set_title('Logistic Regression')
plt.show()
|
Model History/model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: popeye
# language: python
# name: popeye
# ---
import plaidml.keras
plaidml.keras.install_backend()
import os
os.environ["KERAS_BACKEND"] = "plaidml.keras.backend"
# +
# Importing useful libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.preprocessing import MinMaxScaler
from keras.models import Sequential
from keras.layers import Dense, LSTM, Dropout, GRU, Bidirectional, Conv1D, Flatten, MaxPooling1D
from keras.optimizers import SGD
import math
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
from keras import optimizers
import time
# -
# ### Data Processing
df = pd.read_csv('../data/num_data.csv')
dataset = df
dataset.shape
# + _uuid="b288a8e2caf6196daec9cd2bc4ca78fe50345845"
# Useful functions
def plot_predictions(test, predicted):
plt.figure(figsize=(30, 15));
plt.plot(test, color='red', alpha=0.5, label='Actual PM2.5 Concentration',)
plt.plot(predicted, color='blue', alpha=0.5, label='Predicted PM2.5 Concentation')
plt.title('PM2.5 Concentration Prediction')
plt.xlabel('Time')
plt.ylabel('PM2.5 Concentration')
plt.legend()
plt.show()
def return_rmse(test,predicted):
rmse = math.sqrt(mean_squared_error(test, predicted))
return rmse
# -
# +
data_size = dataset.shape[0]
train_size=int(data_size * 0.6)
test_size = 100
valid_size = data_size - train_size - test_size
test_next_day = [12, 24, 48]
# + _uuid="fb4c9db6d8a5bcf20ffad41747cfa5b6215ba220"
training_set = dataset[:train_size].values
valid_set = dataset[train_size:train_size+valid_size].values
test_set = dataset[data_size-test_size:].values
# -
y = dataset.iloc[:,4].values
y = y.reshape(-1,1)
n_feature = training_set.shape[1]
y.shape
# + _uuid="bcc9c36165fc07d258bd5ea87874d2da17fa4a4d"
# Scaling the dataset
sc = MinMaxScaler(feature_range=(0,1))
training_set_scaled = sc.fit_transform(training_set)
valid_set_scaled = sc.fit_transform(valid_set)
test_set_scaled = sc.fit_transform(test_set)
sc_y = MinMaxScaler(feature_range=(0,1))
y_scaled = sc_y.fit_transform(y)
# -
# split a multivariate sequence into samples
def split_sequences(sequences, n_steps_in, n_steps_out):
X_, y_ = list(), list()
for i in range(len(sequences)):
# find the end of this pattern
end_ix = i + n_steps_in
out_end_ix = end_ix + n_steps_out-1
# check if we are beyond the dataset
if out_end_ix > len(sequences):
break
# gather input and output parts of the pattern
seq_x, seq_y = sequences[i:end_ix, :], sequences[end_ix-1:out_end_ix, 0]
X_.append(seq_x)
y_.append(seq_y)
return np.array(X_), np.array(y_)
n_steps_in = 24
n_steps_out = 24
X_train, y_train = split_sequences(training_set_scaled, n_steps_in, n_steps_out)
X_valid, y_valid = split_sequences(valid_set_scaled, n_steps_in, n_steps_out)
X_test, y_test = split_sequences(test_set_scaled, n_steps_in, n_steps_out)
# +
GRU_LSTM_reg = Sequential()
GRU_LSTM_reg.add(GRU(units=50, return_sequences=True, input_shape=(X_train.shape[1],n_feature), activation='tanh'))
GRU_LSTM_reg.add(LSTM(units=50, activation='tanh'))
GRU_LSTM_reg.add(Dense(units=n_steps_out))
# Compiling the RNNs
adam = optimizers.Adam(lr=0.01)
GRU_LSTM_reg.compile(optimizer=adam,loss='mean_squared_error')
# +
RnnModelDict = {'GRU_LSTM': GRU_LSTM_reg}
X_test_24 = X_test[:24]
y_test_24 = y_test[:24]
rmse_df = pd.DataFrame(columns=['Model', 'train_rmse', 'valid_rmse', 'train_time'])
# RnnModelDict = {'LSTM_GRU': LSTM_GRU_reg}
# +
for model in RnnModelDict:
regressor = RnnModelDict[model]
print('training start for', model)
start = time.process_time()
regressor.fit(X_train,y_train,epochs=50,batch_size=1024)
train_time = round(time.process_time() - start, 2)
print('results for training set')
y_train_pred = regressor.predict(X_train)
# plot_predictions(y_train,y_train_pred)
train_rmse = return_rmse(y_train,y_train_pred)
print('results for valid set')
y_valid_pred = regressor.predict(X_valid)
# plot_predictions(y_valid,y_valid_pred)
valid_rmse = return_rmse(y_valid,y_valid_pred)
# print('results for test set - 24 hours')
# y_test_pred24 = regressor.predict(X_test_24)
# plot_predictions(y_test_24,y_test_pred24)
# test24_rmse = return_rmse(y_test_24,y_test_pred24)
one_df = pd.DataFrame([[model, train_rmse, valid_rmse, train_time]],
columns=['Model', 'train_rmse', 'valid_rmse', 'train_time'])
rmse_df = pd.concat([rmse_df, one_df])
# save the rmse results
# rmse_df.to_csv('../rmse_24h_plus_time.csv')
# -
regressor
# +
history = regressor.fit(X_train, y_train, epochs=50, batch_size=1024, validation_data=(X_valid, y_valid),
verbose=2, shuffle=False)
# plot history
plt.figure(figsize=(30, 15))
plt.plot(history.history['loss'], label='Training')
plt.plot(history.history['val_loss'], label='Validation')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
# -
600 - 554
# +
# Transform back and plot
y_train_origin = y[:train_size-46]
y_valid_origin = y[train_size:train_size+valid_size]
y_train_pred = regressor.predict(X_train)
y_train_pred_origin = sc_y.inverse_transform(y_train_pred)
y_valid_pred = regressor.predict(X_valid)
y_valid_pred_origin = sc_y.inverse_transform(y_valid_pred)
_y_train_pred_origin = y_train_pred_origin[:, 0:1]
_y_valid_pred_origin = y_valid_pred_origin[:, 0:1]
# -
plt.plot(y_train_pred_origin)
plot_predictions(y_valid_pred[:,0:1], y_valid_pred[:,0:1])
|
notebooks/.ipynb_checkpoints/DFS_2LSTM_pred_plots-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
import numpy as np
import pandas as pd
from pathlib import Path
# ### Getting the data
path = Path('data')
path.mkdir(parents=True, exist_ok=True)
path
# +
# #!wget http://files.fast.ai/part2/lesson14/rossmann.tgz -P {path}
# +
# #!tar -xvzf {path}/rossmann.tgz -C {path}
# #!rm {path}/rossmann.tgz
# +
# #!rm {path}/sample_submission.csv
# -
# ### Feature engineering
table_paths = sorted(list(path.glob('*.csv')))
table_paths
tables = [pd.read_csv(table_path, low_memory=False) for table_path in table_paths]
googletrend_df, state_names_df, store_df, store_states_df, test_df, train_df, weather_df = tables
# #### Train and test data
train_df.shape, test_df.shape
# +
train_df['Date'] = pd.to_datetime(train_df.Date)
train_df.Date.min(), train_df.Date.max()
# +
test_df['Date'] = pd.to_datetime(test_df.Date)
test_df.Date.min(), test_df.Date.max()
# -
train_df.columns.difference(test_df.columns)
test_df.drop('Id', axis=1, inplace=True)
test_df['Customers'] = np.nan
test_df['Sales'] = np.nan
# +
# Merge train and test to apply preprocessors on both of them at time
df = train_df.append(test_df, sort=False, ignore_index=True, verify_integrity=True)
df.Date.min(), df.Date.max()
# -
# 0 and 1 to boolean
df['Open'] = df['Open'] != 0
df['Promo'] = df['Promo'] != 0
df['StateHoliday'] = df['StateHoliday'] != '0' # object
df['SchoolHoliday'] = df['SchoolHoliday'] != 0
# +
import re
def add_datepart(df):
# Decompose date column into various parts for the purpose of constructing categoricals
date_col = df['Date']
targ_pre = re.sub('[Dd]ate$', '', 'Date')
attr = ['Year', 'Month', 'Week', 'Day', 'Dayofweek', 'Dayofyear',
'Is_month_end', 'Is_month_start', 'Is_quarter_end',
'Is_quarter_start', 'Is_year_end', 'Is_year_start']
for n in attr:
df[targ_pre + n] = getattr(date_col.dt, n.lower())
df[targ_pre + 'Elapsed'] = date_col.astype(np.int64) // 10 ** 9
add_datepart(df)
# -
df.drop('DayOfWeek', axis=1, inplace=True)
df.query('Date==20141231').iloc[0]
base_columns = df.columns
# #### Store data
def left_outer_join(left, right, left_on, right_on=None, suffixes=('', '_y')):
# Left outer join which makes unmatched values on the right side zero
# The suffixes argument describes the naming convention for duplicate fields
if right_on is None: right_on = left_on
return left.merge(right, how='left', left_on=left_on, right_on=right_on, suffixes=suffixes)
# +
# Populate stores with state name
store_df = left_outer_join(store_df, store_states_df, "Store")
store_df.State.isnull().sum()
# -
store_df['Promo2'] = store_df['Promo2'] != 0
# +
# Merge sales data with store information
df = left_outer_join(df, store_df, "Store")
df.StoreType.isnull().sum()
# -
df[df.columns.difference(base_columns)].iloc[0]
base_columns = df.columns
# #### Google Trends
store_states_df['State'].unique()
# +
# Extract date and state for merging purposes
googletrend_df['Date'] = googletrend_df.week.str.split(' - ', expand=True)[0]
googletrend_df['State'] = googletrend_df.file.str.split('_', expand=True)[2]
# Match the rest of the data
googletrend_df.loc[googletrend_df.State=='NI', "State"] = 'HB,NI'
# -
googletrend_df['Date'] = pd.to_datetime(googletrend_df['Date'])
add_datepart(googletrend_df)
# +
# Merge with google trends across states
df = left_outer_join(df, googletrend_df, ["State", "Year", "Week"])
df.trend.isnull().sum()
# +
# Merge with google trends across Germany
trend_de_df = googletrend_df[googletrend_df.file == 'Rossmann_DE']
df = left_outer_join(df, trend_de_df, ["Year", "Week"], suffixes=('', '_DE'))
df.trend_DE.isnull().sum()
# -
# Remove redundant columns
for c in df.columns:
if c in ['file', 'week'] or c.endswith('_y') or (c.endswith('_DE') and c != 'trend_DE'):
if c in df.columns: df.drop(c, inplace=True, axis=1)
df[df.columns.difference(base_columns)].iloc[0]
base_columns = df.columns
# #### Weather data
# +
weather_df = left_outer_join(weather_df, state_names_df, "file", "StateName")
weather_df.StateName.isnull().sum()
# -
weather_df['Date'] = pd.to_datetime(weather_df.Date)
# +
# Merge with weather data
df = left_outer_join(df, weather_df, ["State", "Date"])
df.Mean_TemperatureC.isnull().sum()
# -
df.drop(['file', 'StateName'], axis=1, inplace=True)
df[df.columns.difference(base_columns)].iloc[0]
base_columns = df.columns
# #### Promo and competition
# Pick an arbitrary signal value that doesn't otherwise appear in the data
df['Promo2SinceYear'] = df.Promo2SinceYear.fillna(1900).astype(np.int32)
df['Promo2SinceWeek'] = df.Promo2SinceWeek.fillna(1).astype(np.int32)
# +
from isoweek import Week
df["Promo2Since"] = pd.to_datetime(df.apply(lambda x: Week(x.Promo2SinceYear, x.Promo2SinceWeek).monday(), axis=1))
df["Promo2Days"] = df.Date.subtract(df["Promo2Since"]).dt.days
df.loc[df.Promo2Days<0, "Promo2Days"] = 0
df.loc[df.Promo2SinceYear<1990, "Promo2Days"] = 0
df["Promo2Weeks"] = df["Promo2Days"] // 7
df.loc[df.Promo2Weeks<0, "Promo2Weeks"] = 0
df.loc[df.Promo2Weeks>25, "Promo2Weeks"] = 25 # limit the number of categories
# -
df['CompetitionOpenSinceYear'] = df.CompetitionOpenSinceYear.fillna(1900).astype(np.int32)
df['CompetitionOpenSinceMonth'] = df.CompetitionOpenSinceMonth.fillna(1).astype(np.int32)
# +
# Populate some temporal columns related to competition
df["CompetitionOpenSince"] = pd.to_datetime(dict(year=df.CompetitionOpenSinceYear,
month=df.CompetitionOpenSinceMonth, day=15))
df["CompetitionDaysOpen"] = df.Date.subtract(df.CompetitionOpenSince).dt.days
df.loc[df.CompetitionOpenSinceYear<1990, "CompetitionDaysOpen"] = 0
df.loc[df.CompetitionDaysOpen<0, "CompetitionDaysOpen"] = 0
df["CompetitionMonthsOpen"] = df["CompetitionDaysOpen"] // 30
df.loc[df.CompetitionMonthsOpen<0, "CompetitionMonthsOpen"] = 0
df.loc[df.CompetitionMonthsOpen>24, "CompetitionMonthsOpen"] = 24
# -
df[df.columns.difference(base_columns)].iloc[0]
base_columns = df.columns
# #### Duration between events
def set_elapsed(df, fld, pre):
# Given a particular boolean field fld to monitor,
# this function will start tracking time since the last occurrence of that field
day1 = np.timedelta64(1, 'D')
last_date = np.datetime64()
last_store = 0
res = []
for s, v, d in zip(df.Store.values, df[fld].values, df.Date.values):
if s != last_store:
last_date = np.datetime64()
last_store = s
if v: last_date = d
res.append(abs((d-last_date).astype('timedelta64[D]') / day1))
df[pre+fld] = res
# +
# Create a temporary dataframe with relevant data
on_columns = ["Date", "Store", "Open", "Promo", "StateHoliday", "SchoolHoliday"]
elapsed_df = df[on_columns].copy()
# +
# Get durations of events
elapsed_df['Closed'] = ~elapsed_df['Open']
event_flds = ['Closed', 'SchoolHoliday', 'StateHoliday', 'Promo']
for fld in event_flds:
elapsed_df.sort_values(['Store', 'Date'], inplace=True)
set_elapsed(elapsed_df, fld, 'After')
elapsed_df.sort_values(['Store', 'Date'], ascending=[True, False], inplace=True)
set_elapsed(elapsed_df, fld, 'Before')
# -
elapsed_df.set_index("Date", inplace=True)
# Replace NaNs with 0
for suffix in event_flds:
for prefix in ['Before', 'After']:
column = prefix + suffix
elapsed_df[column] = elapsed_df[column].fillna(0).astype(int)
# Calculate the number of events in the previous 7 days
bwd_df = elapsed_df[['Store']+event_flds].sort_index().groupby("Store").rolling(7, min_periods=1).sum()
# _ next 7 days
fwd_df = elapsed_df[['Store']+event_flds].sort_index(ascending=False).groupby("Store").rolling(7, min_periods=1).sum()
# Drop index
bwd_df.drop('Store', axis=1, inplace=True)
bwd_df.reset_index(inplace=True)
fwd_df.drop('Store', axis=1, inplace=True)
fwd_df.reset_index(inplace=True)
bwd_df.columns
fwd_df.columns
elapsed_df.columns
elapsed_df = left_outer_join(elapsed_df, bwd_df, ['Date', 'Store'], suffixes=['', '_bw'])
elapsed_df = left_outer_join(elapsed_df, fwd_df, ['Date', 'Store'], suffixes=['', '_fw'])
elapsed_df.sort_values(by=['Store', 'Date']).head(10).T
# Drop not needed fields before join
elapsed_df.drop(['Open'] + event_flds, axis=1, inplace=True)
df = left_outer_join(df, elapsed_df, ['Store', 'Date'])
df[df.columns.difference(base_columns)].iloc[0]
# #### Closed stores and zero sales stores
(~df.Open).sum()
(df['Sales'] == 0).sum()
# Closed stores won't be counted into the forecasts
df = df[~((~df.Open) | (df['Sales'] == 0))]
df.drop('Open', axis=1, inplace=True)
# #### Save data
# +
df.sort_values(by=['Date', 'Store'], inplace=True)
df.Date.iloc[0], df.Date.iloc[-1]
# +
df.reset_index(inplace=True, drop=True)
df.index
# -
# Get columns with missing values
df[df.columns[df.isna().any()]].isnull().sum()
df[['SchoolHoliday_bw', 'StateHoliday_bw', 'Promo_bw', 'Closed_bw',
'SchoolHoliday_fw', 'StateHoliday_fw', 'Promo_fw', 'Closed_fw']] = \
df[['SchoolHoliday_bw', 'StateHoliday_bw', 'Promo_bw', 'Closed_bw',
'SchoolHoliday_fw', 'StateHoliday_fw', 'Promo_fw', 'Closed_fw']].astype('int64')
# +
train_df = df.loc[~df.Sales.isnull()]
train_df.shape
# -
train_df[['Sales', 'Customers']] = train_df[['Sales', 'Customers']].astype('int64')
train_df.describe(include=[np.number]).loc[['count', 'min', 'max']].T.astype(np.int64)
train_df.describe(exclude=[np.number]).T
# +
test_df = df.loc[df.Sales.isnull()]
test_df.drop('Sales', axis=1, inplace=True)
test_df.drop('Customers', axis=1, inplace=True)
test_df.shape
# -
# Save final tables
train_df.to_pickle(path/'train_df')
test_df.to_pickle(path/'test_df')
|
rossmann-sales-prediction/data-preparation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from sn_random_numbers import sn_random_numbers
from simulation_class import simulation_class
class geometric_brownian_motion(simulation_class):
def __init__(self,name,mar_env,corr=False):
super(geometric_brownian_motion,self).__init__(name,mar_env,corr)
def update(self,initial_value=None,volatility=None,final_date=None):
if initial_value is not None:
self.initial_value=initial_value
if volatility is not None:
self.volatility=volatility
if final_date is not None:
self.final_date=final_date
self.instrument_values=None
def generate_path(self,fixed_seed=False,day_count=365):
if self.time_grid is None:
self.generate_time_grid()
M=len(self.time_grid)
I=self.paths
paths=np.zeros((M,I))
paths[0]=self.initial_value
if not self.correlated:
rand=sn_random_numbers((1,M,I),fixed_seed=fixed_seed)
else:
rand=self.random_numbers
short_rate=self.discount_curve.short_rate
for t in range(1,len(self.time_grid)):
if not self.correlated:
ran=rand[t]
else:
ran=np.dot(self.cholesky_matrix,rand[:,t,:])
ran=ran[self.rn_set]
dt=(self.time_grid[t]-self.time_grid[t-1]).days/day_count
paths[t]=paths[t-1]*np.exp((short_rate-0.5*self.volatility**2)*dt+
self.volatility*np.sqrt(dt)*ran)
self.instrument_values=paths
|
geometric_brownian_motion.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
data = pd.read_csv("../countries.csv")
data.head()
# +
# Calculates GDP per Country
data['gdpTotal'] = data.population * data.gdpPerCapita
# Get the top 10 countries per GDP
country_gdp = data.sort_values('gdpTotal', ascending=False)
top_10_gdp = country_gdp[country_gdp.year == 2007].head(10)
# Sumarize their names for usage in the for loop
top_10_gdp = set(top_10_gdp.country)
# +
# Sort the data by Year
data.sort_values('year', ascending=True)
legend = []
for country in set(top_10_gdp):
country_data = data[data.country == country]
plt.plot(country_data.year, country_data.gdpTotal / country_data.gdpTotal.iloc[0], label=country)
legend.append(country)
# Change the figure (graph) zise
fig = plt.gcf()
fig.set_size_inches(20, 12)
plt.legend(legend)
plt.show()
# -
|
pluralsight/data_visualization_YK_Sugi/Module 9 - Solving Real World Problem/What's the Fastest Growing Country in GDP.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.014331, "end_time": "2022-04-03T05:38:04.951607", "exception": false, "start_time": "2022-04-03T05:38:04.937276", "status": "completed"} tags=[]
# # PTN Template
# This notebook serves as a template for single dataset PTN experiments
# It can be run on its own by setting STANDALONE to True (do a find for "STANDALONE" to see where)
# But it is intended to be executed as part of a *papermill.py script. See any of the
# experimentes with a papermill script to get started with that workflow.
# + papermill={"duration": 0.991614, "end_time": "2022-04-03T05:38:05.952320", "exception": false, "start_time": "2022-04-03T05:38:04.960706", "status": "completed"} tags=[]
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import os, json, sys, time, random
import numpy as np
import torch
from torch.optim import Adam
from easydict import EasyDict
import matplotlib.pyplot as plt
from steves_models.steves_ptn import Steves_Prototypical_Network
from steves_utils.lazy_iterable_wrapper import Lazy_Iterable_Wrapper
from steves_utils.iterable_aggregator import Iterable_Aggregator
from steves_utils.ptn_train_eval_test_jig import PTN_Train_Eval_Test_Jig
from steves_utils.torch_sequential_builder import build_sequential
from steves_utils.torch_utils import get_dataset_metrics, ptn_confusion_by_domain_over_dataloader
from steves_utils.utils_v2 import (per_domain_accuracy_from_confusion, get_datasets_base_path)
from steves_utils.PTN.utils import independent_accuracy_assesment
from steves_utils.stratified_dataset.episodic_accessor import Episodic_Accessor_Factory
from steves_utils.ptn_do_report import (
get_loss_curve,
get_results_table,
get_parameters_table,
get_domain_accuracies,
)
from steves_utils.transforms import get_chained_transform
# + [markdown] papermill={"duration": 0.009136, "end_time": "2022-04-03T05:38:05.970628", "exception": false, "start_time": "2022-04-03T05:38:05.961492", "status": "completed"} tags=[]
# # Required Parameters
# These are allowed parameters, not defaults
# Each of these values need to be present in the injected parameters (the notebook will raise an exception if they are not present)
#
# Papermill uses the cell tag "parameters" to inject the real parameters below this cell.
# Enable tags to see what I mean
# + papermill={"duration": 0.02099, "end_time": "2022-04-03T05:38:06.000735", "exception": false, "start_time": "2022-04-03T05:38:05.979745", "status": "completed"} tags=[]
required_parameters = {
"experiment_name",
"lr",
"device",
"seed",
"dataset_seed",
"labels_source",
"labels_target",
"domains_source",
"domains_target",
"num_examples_per_domain_per_label_source",
"num_examples_per_domain_per_label_target",
"n_shot",
"n_way",
"n_query",
"train_k_factor",
"val_k_factor",
"test_k_factor",
"n_epoch",
"patience",
"criteria_for_best",
"x_transforms_source",
"x_transforms_target",
"episode_transforms_source",
"episode_transforms_target",
"pickle_name",
"x_net",
"NUM_LOGS_PER_EPOCH",
"BEST_MODEL_PATH",
"torch_default_dtype"
}
# + papermill={"duration": 0.02597, "end_time": "2022-04-03T05:38:06.036020", "exception": false, "start_time": "2022-04-03T05:38:06.010050", "status": "completed"} tags=["parameters"]
standalone_parameters = {}
standalone_parameters["experiment_name"] = "STANDALONE PTN"
standalone_parameters["lr"] = 0.0001
standalone_parameters["device"] = "cuda"
standalone_parameters["seed"] = 1337
standalone_parameters["dataset_seed"] = 1337
standalone_parameters["num_examples_per_domain_per_label_source"]=100
standalone_parameters["num_examples_per_domain_per_label_target"]=100
standalone_parameters["n_shot"] = 3
standalone_parameters["n_query"] = 2
standalone_parameters["train_k_factor"] = 1
standalone_parameters["val_k_factor"] = 2
standalone_parameters["test_k_factor"] = 2
standalone_parameters["n_epoch"] = 100
standalone_parameters["patience"] = 10
standalone_parameters["criteria_for_best"] = "target_accuracy"
standalone_parameters["x_transforms_source"] = ["unit_power"]
standalone_parameters["x_transforms_target"] = ["unit_power"]
standalone_parameters["episode_transforms_source"] = []
standalone_parameters["episode_transforms_target"] = []
standalone_parameters["torch_default_dtype"] = "torch.float32"
standalone_parameters["x_net"] = [
{"class": "nnReshape", "kargs": {"shape":[-1, 1, 2, 256]}},
{"class": "Conv2d", "kargs": { "in_channels":1, "out_channels":256, "kernel_size":(1,7), "bias":False, "padding":(0,3), },},
{"class": "ReLU", "kargs": {"inplace": True}},
{"class": "BatchNorm2d", "kargs": {"num_features":256}},
{"class": "Conv2d", "kargs": { "in_channels":256, "out_channels":80, "kernel_size":(2,7), "bias":True, "padding":(0,3), },},
{"class": "ReLU", "kargs": {"inplace": True}},
{"class": "BatchNorm2d", "kargs": {"num_features":80}},
{"class": "Flatten", "kargs": {}},
{"class": "Linear", "kargs": {"in_features": 80*256, "out_features": 256}}, # 80 units per IQ pair
{"class": "ReLU", "kargs": {"inplace": True}},
{"class": "BatchNorm1d", "kargs": {"num_features":256}},
{"class": "Linear", "kargs": {"in_features": 256, "out_features": 256}},
]
# Parameters relevant to results
# These parameters will basically never need to change
standalone_parameters["NUM_LOGS_PER_EPOCH"] = 10
standalone_parameters["BEST_MODEL_PATH"] = "./best_model.pth"
# uncomment for CORES dataset
from steves_utils.CORES.utils import (
ALL_NODES,
ALL_NODES_MINIMUM_1000_EXAMPLES,
ALL_DAYS
)
standalone_parameters["labels_source"] = ALL_NODES
standalone_parameters["labels_target"] = ALL_NODES
standalone_parameters["domains_source"] = [1]
standalone_parameters["domains_target"] = [2,3,4,5]
standalone_parameters["pickle_name"] = "cores.stratified_ds.2022A.pkl"
# Uncomment these for ORACLE dataset
# from steves_utils.ORACLE.utils_v2 import (
# ALL_DISTANCES_FEET,
# ALL_RUNS,
# ALL_SERIAL_NUMBERS,
# )
# standalone_parameters["labels_source"] = ALL_SERIAL_NUMBERS
# standalone_parameters["labels_target"] = ALL_SERIAL_NUMBERS
# standalone_parameters["domains_source"] = [8,20, 38,50]
# standalone_parameters["domains_target"] = [14, 26, 32, 44, 56]
# standalone_parameters["pickle_name"] = "oracle.frame_indexed.stratified_ds.2022A.pkl"
# standalone_parameters["num_examples_per_domain_per_label_source"]=1000
# standalone_parameters["num_examples_per_domain_per_label_target"]=1000
# Uncomment these for Metahan dataset
# standalone_parameters["labels_source"] = list(range(19))
# standalone_parameters["labels_target"] = list(range(19))
# standalone_parameters["domains_source"] = [0]
# standalone_parameters["domains_target"] = [1]
# standalone_parameters["pickle_name"] = "metehan.stratified_ds.2022A.pkl"
# standalone_parameters["n_way"] = len(standalone_parameters["labels_source"])
# standalone_parameters["num_examples_per_domain_per_label_source"]=200
# standalone_parameters["num_examples_per_domain_per_label_target"]=100
standalone_parameters["n_way"] = len(standalone_parameters["labels_source"])
# + papermill={"duration": 0.025308, "end_time": "2022-04-03T05:38:06.070699", "exception": false, "start_time": "2022-04-03T05:38:06.045391", "status": "completed"} tags=["injected-parameters"]
# Parameters
parameters = {
"experiment_name": "baseline_ptn_32bit_oracle.run1.framed",
"lr": 0.001,
"device": "cuda",
"seed": 1337,
"dataset_seed": 1337,
"labels_source": [
"3123D52",
"3123D65",
"3123D79",
"3123D80",
"3123D54",
"3123D70",
"3123D7B",
"3123D89",
"3123D58",
"3123D76",
"3123D7D",
"3123EFE",
"3123D64",
"3123D78",
"3123D7E",
"3124E4A",
],
"labels_target": [
"3123D52",
"3123D65",
"3123D79",
"3123D80",
"3123D54",
"3123D70",
"3123D7B",
"3123D89",
"3123D58",
"3123D76",
"3123D7D",
"3123EFE",
"3123D64",
"3123D78",
"3123D7E",
"3124E4A",
],
"x_transforms_source": [],
"x_transforms_target": [],
"episode_transforms_source": [],
"episode_transforms_target": [],
"num_examples_per_domain_per_label_source": 1000,
"num_examples_per_domain_per_label_target": 1000,
"n_shot": 3,
"n_way": 16,
"n_query": 2,
"train_k_factor": 1,
"val_k_factor": 2,
"test_k_factor": 2,
"torch_default_dtype": "torch.float32",
"n_epoch": 50,
"patience": 3,
"criteria_for_best": "target_loss",
"x_net": [
{"class": "nnReshape", "kargs": {"shape": [-1, 1, 2, 256]}},
{
"class": "Conv2d",
"kargs": {
"in_channels": 1,
"out_channels": 256,
"kernel_size": [1, 7],
"bias": False,
"padding": [0, 3],
},
},
{"class": "ReLU", "kargs": {"inplace": True}},
{"class": "BatchNorm2d", "kargs": {"num_features": 256}},
{
"class": "Conv2d",
"kargs": {
"in_channels": 256,
"out_channels": 80,
"kernel_size": [2, 7],
"bias": True,
"padding": [0, 3],
},
},
{"class": "ReLU", "kargs": {"inplace": True}},
{"class": "BatchNorm2d", "kargs": {"num_features": 80}},
{"class": "Flatten", "kargs": {}},
{"class": "Linear", "kargs": {"in_features": 20480, "out_features": 256}},
{"class": "ReLU", "kargs": {"inplace": True}},
{"class": "BatchNorm1d", "kargs": {"num_features": 256}},
{"class": "Linear", "kargs": {"in_features": 256, "out_features": 256}},
],
"NUM_LOGS_PER_EPOCH": 10,
"BEST_MODEL_PATH": "./best_model.pth",
"pickle_name": "oracle.Run1_framed_2000Examples_stratified_ds.2022A.pkl",
"domains_source": [8, 32, 50],
"domains_target": [14, 20, 26, 38, 44],
}
# + papermill={"duration": 0.021425, "end_time": "2022-04-03T05:38:06.101630", "exception": false, "start_time": "2022-04-03T05:38:06.080205", "status": "completed"} tags=[]
# Set this to True if you want to run this template directly
STANDALONE = False
if STANDALONE:
print("parameters not injected, running with standalone_parameters")
parameters = standalone_parameters
if not 'parameters' in locals() and not 'parameters' in globals():
raise Exception("Parameter injection failed")
#Use an easy dict for all the parameters
p = EasyDict(parameters)
supplied_keys = set(p.keys())
if supplied_keys != required_parameters:
print("Parameters are incorrect")
if len(supplied_keys - required_parameters)>0: print("Shouldn't have:", str(supplied_keys - required_parameters))
if len(required_parameters - supplied_keys)>0: print("Need to have:", str(required_parameters - supplied_keys))
raise RuntimeError("Parameters are incorrect")
# + papermill={"duration": 0.021623, "end_time": "2022-04-03T05:38:06.132588", "exception": false, "start_time": "2022-04-03T05:38:06.110965", "status": "completed"} tags=[]
###################################
# Set the RNGs and make it all deterministic
###################################
np.random.seed(p.seed)
random.seed(p.seed)
torch.manual_seed(p.seed)
torch.use_deterministic_algorithms(True)
# + papermill={"duration": 0.020805, "end_time": "2022-04-03T05:38:06.162860", "exception": false, "start_time": "2022-04-03T05:38:06.142055", "status": "completed"} tags=[]
###########################################
# The stratified datasets honor this
###########################################
torch.set_default_dtype(eval(p.torch_default_dtype))
# + papermill={"duration": 0.046428, "end_time": "2022-04-03T05:38:06.218809", "exception": false, "start_time": "2022-04-03T05:38:06.172381", "status": "completed"} tags=[]
###################################
# Build the network(s)
# Note: It's critical to do this AFTER setting the RNG
# (This is due to the randomized initial weights)
###################################
x_net = build_sequential(p.x_net)
# + papermill={"duration": 0.022505, "end_time": "2022-04-03T05:38:06.250970", "exception": false, "start_time": "2022-04-03T05:38:06.228465", "status": "completed"} tags=[]
start_time_secs = time.time()
# + papermill={"duration": 1.019244, "end_time": "2022-04-03T05:38:07.279901", "exception": false, "start_time": "2022-04-03T05:38:06.260657", "status": "completed"} tags=[]
###################################
# Build the dataset
###################################
if p.x_transforms_source == []: x_transform_source = None
else: x_transform_source = get_chained_transform(p.x_transforms_source)
if p.x_transforms_target == []: x_transform_target = None
else: x_transform_target = get_chained_transform(p.x_transforms_target)
if p.episode_transforms_source == []: episode_transform_source = None
else: raise Exception("episode_transform_source not implemented")
if p.episode_transforms_target == []: episode_transform_target = None
else: raise Exception("episode_transform_target not implemented")
eaf_source = Episodic_Accessor_Factory(
labels=p.labels_source,
domains=p.domains_source,
num_examples_per_domain_per_label=p.num_examples_per_domain_per_label_source,
iterator_seed=p.seed,
dataset_seed=p.dataset_seed,
n_shot=p.n_shot,
n_way=p.n_way,
n_query=p.n_query,
train_val_test_k_factors=(p.train_k_factor,p.val_k_factor,p.test_k_factor),
pickle_path=os.path.join(get_datasets_base_path(), p.pickle_name),
x_transform_func=x_transform_source,
example_transform_func=episode_transform_source,
)
train_original_source, val_original_source, test_original_source = eaf_source.get_train(), eaf_source.get_val(), eaf_source.get_test()
eaf_target = Episodic_Accessor_Factory(
labels=p.labels_target,
domains=p.domains_target,
num_examples_per_domain_per_label=p.num_examples_per_domain_per_label_target,
iterator_seed=p.seed,
dataset_seed=p.dataset_seed,
n_shot=p.n_shot,
n_way=p.n_way,
n_query=p.n_query,
train_val_test_k_factors=(p.train_k_factor,p.val_k_factor,p.test_k_factor),
pickle_path=os.path.join(get_datasets_base_path(), p.pickle_name),
x_transform_func=x_transform_target,
example_transform_func=episode_transform_target,
)
train_original_target, val_original_target, test_original_target = eaf_target.get_train(), eaf_target.get_val(), eaf_target.get_test()
transform_lambda = lambda ex: ex[1] # Original is (<domain>, <episode>) so we strip down to episode only
train_processed_source = Lazy_Iterable_Wrapper(train_original_source, transform_lambda)
val_processed_source = Lazy_Iterable_Wrapper(val_original_source, transform_lambda)
test_processed_source = Lazy_Iterable_Wrapper(test_original_source, transform_lambda)
train_processed_target = Lazy_Iterable_Wrapper(train_original_target, transform_lambda)
val_processed_target = Lazy_Iterable_Wrapper(val_original_target, transform_lambda)
test_processed_target = Lazy_Iterable_Wrapper(test_original_target, transform_lambda)
datasets = EasyDict({
"source": {
"original": {"train":train_original_source, "val":val_original_source, "test":test_original_source},
"processed": {"train":train_processed_source, "val":val_processed_source, "test":test_processed_source}
},
"target": {
"original": {"train":train_original_target, "val":val_original_target, "test":test_original_target},
"processed": {"train":train_processed_target, "val":val_processed_target, "test":test_processed_target}
},
})
# + papermill={"duration": 1.592898, "end_time": "2022-04-03T05:38:08.887532", "exception": false, "start_time": "2022-04-03T05:38:07.294634", "status": "completed"} tags=[]
# Some quick unit tests on the data
from steves_utils.transforms import get_average_power, get_average_magnitude
q_x, q_y, s_x, s_y, truth = next(iter(train_processed_source))
assert q_x.dtype == eval(p.torch_default_dtype)
assert s_x.dtype == eval(p.torch_default_dtype)
print("Visually inspect these to see if they line up with expected values given the transforms")
print('x_transforms_source', p.x_transforms_source)
print('x_transforms_target', p.x_transforms_target)
print("Average magnitude, source:", get_average_magnitude(q_x[0].numpy()))
print("Average power, source:", get_average_power(q_x[0].numpy()))
q_x, q_y, s_x, s_y, truth = next(iter(train_processed_target))
print("Average magnitude, target:", get_average_magnitude(q_x[0].numpy()))
print("Average power, target:", get_average_power(q_x[0].numpy()))
# + papermill={"duration": 0.074449, "end_time": "2022-04-03T05:38:08.979945", "exception": false, "start_time": "2022-04-03T05:38:08.905496", "status": "completed"} tags=[]
###################################
# Build the model
###################################
model = Steves_Prototypical_Network(x_net, device=p.device, x_shape=(2,256))
optimizer = Adam(params=model.parameters(), lr=p.lr)
# + papermill={"duration": 414.502485, "end_time": "2022-04-03T05:45:03.497862", "exception": false, "start_time": "2022-04-03T05:38:08.995377", "status": "completed"} tags=[]
###################################
# train
###################################
jig = PTN_Train_Eval_Test_Jig(model, p.BEST_MODEL_PATH, p.device)
jig.train(
train_iterable=datasets.source.processed.train,
source_val_iterable=datasets.source.processed.val,
target_val_iterable=datasets.target.processed.val,
num_epochs=p.n_epoch,
num_logs_per_epoch=p.NUM_LOGS_PER_EPOCH,
patience=p.patience,
optimizer=optimizer,
criteria_for_best=p.criteria_for_best,
)
# + papermill={"duration": 0.088325, "end_time": "2022-04-03T05:45:03.658272", "exception": false, "start_time": "2022-04-03T05:45:03.569947", "status": "completed"} tags=[]
total_experiment_time_secs = time.time() - start_time_secs
# + papermill={"duration": 19.148263, "end_time": "2022-04-03T05:45:22.878724", "exception": false, "start_time": "2022-04-03T05:45:03.730461", "status": "completed"} tags=[]
###################################
# Evaluate the model
###################################
source_test_label_accuracy, source_test_label_loss = jig.test(datasets.source.processed.test)
target_test_label_accuracy, target_test_label_loss = jig.test(datasets.target.processed.test)
source_val_label_accuracy, source_val_label_loss = jig.test(datasets.source.processed.val)
target_val_label_accuracy, target_val_label_loss = jig.test(datasets.target.processed.val)
history = jig.get_history()
total_epochs_trained = len(history["epoch_indices"])
val_dl = Iterable_Aggregator((datasets.source.original.val,datasets.target.original.val))
confusion = ptn_confusion_by_domain_over_dataloader(model, p.device, val_dl)
per_domain_accuracy = per_domain_accuracy_from_confusion(confusion)
# Add a key to per_domain_accuracy for if it was a source domain
for domain, accuracy in per_domain_accuracy.items():
per_domain_accuracy[domain] = {
"accuracy": accuracy,
"source?": domain in p.domains_source
}
# Do an independent accuracy assesment JUST TO BE SURE!
# _source_test_label_accuracy = independent_accuracy_assesment(model, datasets.source.processed.test, p.device)
# _target_test_label_accuracy = independent_accuracy_assesment(model, datasets.target.processed.test, p.device)
# _source_val_label_accuracy = independent_accuracy_assesment(model, datasets.source.processed.val, p.device)
# _target_val_label_accuracy = independent_accuracy_assesment(model, datasets.target.processed.val, p.device)
# assert(_source_test_label_accuracy == source_test_label_accuracy)
# assert(_target_test_label_accuracy == target_test_label_accuracy)
# assert(_source_val_label_accuracy == source_val_label_accuracy)
# assert(_target_val_label_accuracy == target_val_label_accuracy)
experiment = {
"experiment_name": p.experiment_name,
"parameters": dict(p),
"results": {
"source_test_label_accuracy": source_test_label_accuracy,
"source_test_label_loss": source_test_label_loss,
"target_test_label_accuracy": target_test_label_accuracy,
"target_test_label_loss": target_test_label_loss,
"source_val_label_accuracy": source_val_label_accuracy,
"source_val_label_loss": source_val_label_loss,
"target_val_label_accuracy": target_val_label_accuracy,
"target_val_label_loss": target_val_label_loss,
"total_epochs_trained": total_epochs_trained,
"total_experiment_time_secs": total_experiment_time_secs,
"confusion": confusion,
"per_domain_accuracy": per_domain_accuracy,
},
"history": history,
"dataset_metrics": get_dataset_metrics(datasets, "ptn"),
}
# + papermill={"duration": 0.199919, "end_time": "2022-04-03T05:45:23.151787", "exception": false, "start_time": "2022-04-03T05:45:22.951868", "status": "completed"} tags=[]
ax = get_loss_curve(experiment)
plt.show()
# + papermill={"duration": 0.21362, "end_time": "2022-04-03T05:45:23.442059", "exception": false, "start_time": "2022-04-03T05:45:23.228439", "status": "completed"} tags=[]
get_results_table(experiment)
# + papermill={"duration": 0.191786, "end_time": "2022-04-03T05:45:23.708795", "exception": false, "start_time": "2022-04-03T05:45:23.517009", "status": "completed"} tags=[]
get_domain_accuracies(experiment)
# + papermill={"duration": 0.089974, "end_time": "2022-04-03T05:45:23.875158", "exception": false, "start_time": "2022-04-03T05:45:23.785184", "status": "completed"} tags=[]
print("Source Test Label Accuracy:", experiment["results"]["source_test_label_accuracy"], "Target Test Label Accuracy:", experiment["results"]["target_test_label_accuracy"])
print("Source Val Label Accuracy:", experiment["results"]["source_val_label_accuracy"], "Target Val Label Accuracy:", experiment["results"]["target_val_label_accuracy"])
# + papermill={"duration": 0.088849, "end_time": "2022-04-03T05:45:24.039588", "exception": false, "start_time": "2022-04-03T05:45:23.950739", "status": "completed"} tags=["experiment_json"]
json.dumps(experiment)
|
experiments/baseline_ptn_32bit/oracle.run1.framed/trials/1/trial.ipynb
|