code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # BFS
# <!-- - <span style= "orange">
#
# </span> -->
# - <span style="color:yellow">O(V+E)</span>
# +
class Node(object):
def __init__(self, name):
self.name = name
self.adjacentList = []
self.visited = False
self.predecssor = None
class BreadthFirstSearch(object):
def bfs(self, startNode):
queue = []
queue.append(startNode)
startNode.visited = True
while queue:
actualNode = queue.pop(0)
print("%s " % actualNode.name)
for n in actualNode.adjacentList:
if not n.visited:
n.visited = True
queue.append(n)
# +
node1 = Node("A")
node2 = Node("B")
node3 = Node("C")
node4 = Node("D")
node5 = Node("E")
node1.adjacentList.append(node2)
node1.adjacentList.append(node3)
node2.adjacentList.append(node4)
node4.adjacentList.append(node5)
bfs = BreadthFirstSearch()
bfs.bfs(node1)
# -
# # DFS
# - <span style="color:yellow">O(V+E)</span>
# +
class Node(object):
def __init__(self, name):
self.name = name
self.adjacentList = []
self.visited = False
self.predecssor = None
class DepthFirstSearch(object):
def dfs(self, node):
node.visited = True
print("%s " % node.name)
for n in node.adjacentList:
if not n.visited:
self.dfs(n)
# +
node1 = Node("A")
node2 = Node("B")
node3 = Node("C")
node4 = Node("D")
node5 = Node("E")
node1.adjacentList.append(node2)
node1.adjacentList.append(node3)
node2.adjacentList.append(node4)
node4.adjacentList.append(node5)
dfs = DepthFirstSearch()
dfs.dfs(node1)
# -
|
CODING/Graph.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import theano
import theano.tensor as T
import numpy as np
# +
X = theano.shared(np.zeros((4,10,20)).astype('float32'))
mask = theano.shared(np.zeros((4,10)).astype('float32'))
newX = T.tensor3('newX',dtype='float32')
newMask=T.matrix('newMask',dtype='float32')
resetX= theano.function([newX,newMask],None,updates=[(X,newX),(mask,newMask)])
statX = theano.function([],[X.mean(),X.max(),X.sum()])
# -
mnX,maX,smX = statX()
print mnX,maX,smX
nX = np.ones((12,4,32)).astype('float32')
nM = np.ones((17,3)).astype('float32')
resetX(newX=nX,newMask=nM)
mnX,maX,smX = statX()
print mnX,maX,smX
|
ResearchIpynb.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# +
# slow down a bit when hacking something together, e.g. I forgot to add a simple function call
# tuple unpacking is nice, but cannot be done in a nested list comprehension
# -
from dataclasses import dataclass
from math import gcd, ceil
import re
from collections import Counter, defaultdict, namedtuple, deque
import numpy as np
from matplotlib import pyplot as plt
import aoc
import networkx as nx
f=open('input.txt')
lines = [sorted(aoc.to_int(line.rstrip('\n').split('x'))) for line in f]
print(f'N lines: {len(lines)}, firstline looks like\n {lines[0][:100]}')
# part 1
count = 0
for line in lines:
l,w,h = line
count += (l*w*3+l*h*2+w*h*2)
count
# part
count = 0
for line in lines:
l,w,h = line
count+= l+l+w+w+l*w*h
count
|
advent_of_code_2015/day 2/solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Traffic Sign Classification with Keras
#
# Keras exists to make coding deep neural networks simpler. To demonstrate just how easy it is, you’re going to use Keras to build a convolutional neural network in a few dozen lines of code.
#
# You’ll be connecting the concepts from the previous lessons to the methods that Keras provides.
# ## Dataset
#
# The network you'll build with Keras is similar to the example in Keras’s GitHub repository that builds out a [convolutional neural network for MNIST](https://github.com/fchollet/keras/blob/master/examples/mnist_cnn.py).
#
# However, instead of using the [MNIST](http://yann.lecun.com/exdb/mnist/) dataset, you're going to use the [German Traffic Sign Recognition Benchmark](http://benchmark.ini.rub.de/?section=gtsrb&subsection=news) dataset that you've used previously.
#
# You can download pickle files with sanitized traffic sign data here:
# +
from urllib.request import urlretrieve
from os.path import isfile
from tqdm import tqdm
class DLProgress(tqdm):
last_block = 0
def hook(self, block_num=1, block_size=1, total_size=None):
self.total = total_size
self.update((block_num - self.last_block) * block_size)
self.last_block = block_num
if not isfile('train.p'):
with DLProgress(unit='B', unit_scale=True, miniters=1, desc='Train Dataset') as pbar:
urlretrieve(
'https://s3.amazonaws.com/udacity-sdc/datasets/german_traffic_sign_benchmark/train.p',
'train.p',
pbar.hook)
if not isfile('test.p'):
with DLProgress(unit='B', unit_scale=True, miniters=1, desc='Test Dataset') as pbar:
urlretrieve(
'https://s3.amazonaws.com/udacity-sdc/datasets/german_traffic_sign_benchmark/test.p',
'test.p',
pbar.hook)
print('Training and Test data downloaded.')
# -
# ## Overview
#
# Here are the steps you'll take to build the network:
#
# 1. Load the training data.
# 2. Preprocess the data.
# 3. Build a feedforward neural network to classify traffic signs.
# 4. Build a convolutional neural network to classify traffic signs.
# 5. Evaluate the final neural network on testing data.
#
# Keep an eye on the network’s accuracy over time. Once the accuracy reaches the 98% range, you can be confident that you’ve built and trained an effective model.
# +
import pickle
import numpy as np
import math
# Fix error with TF and Keras
import tensorflow as tf
tf.python.control_flow_ops = tf
print('Modules loaded.')
# -
# ## Load the Data
#
# Start by importing the data from the pickle file.
# +
with open('train.p', 'rb') as f:
data = pickle.load(f)
# TODO: Load the feature data to the variable X_train
X_train = data['features']
# TODO: Load the label data to the variable y_train
y_train = data['labels']
# -
# STOP: Do not change the tests below. Your implementation should pass these tests.
assert np.array_equal(X_train, data['features']), 'X_train not set to data[\'features\'].'
assert np.array_equal(y_train, data['labels']), 'y_train not set to data[\'labels\'].'
print('Tests passed.')
# ## Preprocess the Data
#
# 1. Shuffle the data
# 2. Normalize the features using Min-Max scaling between -0.5 and 0.5
# 3. One-Hot Encode the labels
#
# ### Shuffle the data
# Hint: You can use the [scikit-learn shuffle](http://scikit-learn.org/stable/modules/generated/sklearn.utils.shuffle.html) function to shuffle the data.
# TODO: Shuffle the data
from sklearn.utils import shuffle
X_train, y_train = shuffle(X_train, y_train)
# STOP: Do not change the tests below. Your implementation should pass these tests.
assert X_train.shape == data['features'].shape, 'X_train has changed shape. The shape shouldn\'t change when shuffling.'
assert y_train.shape == data['labels'].shape, 'y_train has changed shape. The shape shouldn\'t change when shuffling.'
assert not np.array_equal(X_train, data['features']), 'X_train not shuffled.'
assert not np.array_equal(y_train, data['labels']), 'y_train not shuffled.'
print('Tests passed.')
# ### Normalize the features
# Hint: You solved this in [TensorFlow lab](https://github.com/udacity/CarND-TensorFlow-Lab/blob/master/lab.ipynb) Problem 1.
# +
# TODO: Normalize the data features to the variable X_normalized
def normalize_grayscale(image_data):
a = -0.5
b = 0.5
grayscale_min = 0
grayscale_max = 255
return a + ( ( (image_data - grayscale_min)*(b - a) )/( grayscale_max - grayscale_min ) )
X_normalized = normalize_grayscale(X_train)
# -
# STOP: Do not change the tests below. Your implementation should pass these tests.
assert math.isclose(np.min(X_normalized), -0.5, abs_tol=1e-5) and math.isclose(np.max(X_normalized), 0.5, abs_tol=1e-5), 'The range of the training data is: {} to {}. It must be -0.5 to 0.5'.format(np.min(X_normalized), np.max(X_normalized))
print('Tests passed.')
# ### One-Hot Encode the labels
# Hint: You can use the [scikit-learn LabelBinarizer](http://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.LabelBinarizer.html) function to one-hot encode the labels.
# TODO: One Hot encode the labels to the variable y_one_hot
from sklearn.preprocessing import LabelBinarizer
label_binarizer = LabelBinarizer()
y_one_hot = label_binarizer.fit_transform(y_train)
# +
# STOP: Do not change the tests below. Your implementation should pass these tests.
import collections
assert y_one_hot.shape == (39209, 43), 'y_one_hot is not the correct shape. It\'s {}, it should be (39209, 43)'.format(y_one_hot.shape)
assert next((False for y in y_one_hot if collections.Counter(y) != {0: 42, 1: 1}), True), 'y_one_hot not one-hot encoded.'
print('Tests passed.')
# -
# ## Keras Sequential Model
# ```python
# from keras.models import Sequential
#
# # Create the Sequential model
# model = Sequential()
# ```
# The `keras.models.Sequential` class is a wrapper for the neural network model. Just like many of the class models in scikit-learn, it provides common functions like `fit()`, `evaluate()`, and `compile()`. We'll cover these functions as we get to them. Let's start looking at the layers of the model.
#
# ## Keras Layer
# A Keras layer is just like a neural network layer. It can be fully connected, max pool, activation, etc. You can add a layer to the model using the model's `add()` function. For example, a simple model would look like this:
# ```python
# from keras.models import Sequential
# from keras.layers.core import Dense, Activation, Flatten
#
# # Create the Sequential model
# model = Sequential()
#
# # 1st Layer - Add a flatten layer
# model.add(Flatten(input_shape=(32, 32, 3)))
#
# # 2nd Layer - Add a fully connected layer
# model.add(Dense(100))
#
# # 3rd Layer - Add a ReLU activation layer
# model.add(Activation('relu'))
#
# # 4th Layer - Add a fully connected layer
# model.add(Dense(60))
#
# # 5th Layer - Add a ReLU activation layer
# model.add(Activation('relu'))
# ```
# Keras will automatically infer the shape of all layers after the first layer. This means you only have to set the input dimensions for the first layer.
#
# The first layer from above, `model.add(Flatten(input_shape=(32, 32, 3)))`, sets the input dimension to (32, 32, 3) and output dimension to (3072=32\*32\*3). The second layer takes in the output of the first layer and sets the output dimenions to (100). This chain of passing output to the next layer continues until the last layer, which is the output of the model.
# ## Build a Multi-Layer Feedforward Network
#
# Build a multi-layer feedforward neural network to classify the traffic sign images.
#
# 1. Set the first layer to a `Flatten` layer with the `input_shape` set to (32, 32, 3)
# 2. Set the second layer to `Dense` layer width to 128 output.
# 3. Use a ReLU activation function after the second layer.
# 4. Set the output layer width to 43, since there are 43 classes in the dataset.
# 5. Use a softmax activation function after the output layer.
#
# To get started, review the Keras documentation about [models](https://keras.io/models/sequential/) and [layers](https://keras.io/layers/core/).
#
# The Keras example of a [Multi-Layer Perceptron](https://github.com/fchollet/keras/blob/master/examples/mnist_mlp.py) network is similar to what you need to do here. Use that as a guide, but keep in mind that there are a number of differences.
# +
from keras.models import Sequential
model = Sequential()
# TODO: Build a Multi-layer feedforward neural network with Keras here.
from keras.models import Sequential
from keras.layers.core import Dense, Activation, Flatten
model.add(Flatten(input_shape=(32, 32, 3)))
model.add(Dense(128))
model.add(Activation('relu'))
model.add(Dense(43))
model.add(Activation('softmax'))
# +
# STOP: Do not change the tests below. Your implementation should pass these tests.
from keras.layers.core import Dense, Activation, Flatten
from keras.activations import relu, softmax
def check_layers(layers, true_layers):
assert len(true_layers) != 0, 'No layers found'
for layer_i in range(len(layers)):
assert isinstance(true_layers[layer_i], layers[layer_i]), 'Layer {} is not a {} layer'.format(layer_i+1, layers[layer_i].__name__)
assert len(true_layers) == len(layers), '{} layers found, should be {} layers'.format(len(true_layers), len(layers))
check_layers([Flatten, Dense, Activation, Dense, Activation], model.layers)
assert model.layers[0].input_shape == (None, 32, 32, 3), 'First layer input shape is wrong, it should be (32, 32, 3)'
assert model.layers[1].output_shape == (None, 128), 'Second layer output is wrong, it should be (128)'
assert model.layers[2].activation == relu, 'Third layer not a relu activation layer'
assert model.layers[3].output_shape == (None, 43), 'Fourth layer output is wrong, it should be (43)'
assert model.layers[4].activation == softmax, 'Fifth layer not a softmax activation layer'
print('Tests passed.')
# -
# ## Training a Sequential Model
# You built a multi-layer neural network in Keras, now let's look at training a neural network.
# ```python
# from keras.models import Sequential
# from keras.layers.core import Dense, Activation
#
# model = Sequential()
# ...
#
# # Configures the learning process and metrics
# model.compile('sgd', 'mean_squared_error', ['accuracy'])
#
# # Train the model
# # History is a record of training loss and metrics
# history = model.fit(x_train_data, Y_train_data, batch_size=128, nb_epoch=2, validation_split=0.2)
#
# # Calculate test score
# test_score = model.evaluate(x_test_data, Y_test_data)
# ```
# The code above configures, trains, and tests the model. The line `model.compile('sgd', 'mean_squared_error', ['accuracy'])` configures the model's optimizer to `'sgd'`(stochastic gradient descent), the loss to `'mean_squared_error'`, and the metric to `'accuracy'`.
#
# You can find more optimizers [here](https://keras.io/optimizers/), loss functions [here](https://keras.io/objectives/#available-objectives), and more metrics [here](https://keras.io/metrics/#available-metrics).
#
# To train the model, use the `fit()` function as shown in `model.fit(x_train_data, Y_train_data, batch_size=128, nb_epoch=2, validation_split=0.2)`. The `validation_split` parameter will split a percentage of the training dataset to be used to validate the model. The model can be further tested with the test dataset using the `evaluation()` function as shown in the last line.
# ## Train the Network
#
# 1. Compile the network using adam optimizer and categorical_crossentropy loss function.
# 2. Train the network for ten epochs and validate with 20% of the training data.
# TODO: Compile and train the model here.
model.compile('adam', 'categorical_crossentropy', ['accuracy'])
history = model.fit(X_normalized, y_one_hot, nb_epoch=10, validation_split=0.2)
# +
# STOP: Do not change the tests below. Your implementation should pass these tests.
from keras.optimizers import Adam
assert model.loss == 'categorical_crossentropy', 'Not using categorical_crossentropy loss function'
assert isinstance(model.optimizer, Adam), 'Not using adam optimizer'
assert len(history.history['acc']) == 10, 'You\'re using {} epochs when you need to use 10 epochs.'.format(len(history.history['acc']))
assert history.history['acc'][-1] > 0.92, 'The training accuracy was: %.3f. It shoud be greater than 0.92' % history.history['acc'][-1]
assert history.history['val_acc'][-1] > 0.85, 'The validation accuracy is: %.3f. It shoud be greater than 0.85' % history.history['val_acc'][-1]
print('Tests passed.')
# -
# ## Convolutions
# 1. Re-construct the previous network
# 2. Add a [convolutional layer](https://keras.io/layers/convolutional/#convolution2d) with 32 filters, a 3x3 kernel, and valid padding before the flatten layer.
# 3. Add a ReLU activation after the convolutional layer.
#
# Hint 1: The Keras example of a [convolutional neural network](https://github.com/fchollet/keras/blob/master/examples/mnist_cnn.py) for MNIST would be a good example to review.
# +
# TODO: Re-construct the network and add a convolutional layer before the flatten layer.
from keras.models import Sequential
from keras.models import Sequential
from keras.layers.core import Dense, Activation, Flatten
from keras.layers.convolutional import Convolution2D
model = Sequential()
model.add(Convolution2D(32, 3, 3, input_shape=(32, 32, 3)))
model.add(Activation('relu'))
model.add(Flatten())
model.add(Dense(128))
model.add(Activation('relu'))
model.add(Dense(43))
model.add(Activation('softmax'))
# +
# STOP: Do not change the tests below. Your implementation should pass these tests.
from keras.layers.core import Dense, Activation, Flatten
from keras.layers.convolutional import Convolution2D
check_layers([Convolution2D, Activation, Flatten, Dense, Activation, Dense, Activation], model.layers)
assert model.layers[0].input_shape == (None, 32, 32, 3), 'First layer input shape is wrong, it should be (32, 32, 3)'
assert model.layers[0].nb_filter == 32, 'Wrong number of filters, it should be 32'
assert model.layers[0].nb_col == model.layers[0].nb_row == 3, 'Kernel size is wrong, it should be a 3x3'
assert model.layers[0].border_mode == 'valid', 'Wrong padding, it should be valid'
model.compile('adam', 'categorical_crossentropy', ['accuracy'])
history = model.fit(X_normalized, y_one_hot, batch_size=128, nb_epoch=2, validation_split=0.2)
assert(history.history['val_acc'][-1] > 0.91), "The validation accuracy is: %.3f. It should be greater than 0.91" % history.history['val_acc'][-1]
print('Tests passed.')
# -
# ## Pooling
# 1. Re-construct the network
# 2. Add a 2x2 [max pooling layer](https://keras.io/layers/pooling/#maxpooling2d) immediately following your convolutional layer.
# +
# TODO: Re-construct the network and add a pooling layer after the convolutional layer.
from keras.models import Sequential
from keras.models import Sequential
from keras.layers.core import Dense, Activation, Flatten
from keras.layers.convolutional import Convolution2D
from keras.layers.pooling import MaxPooling2D
model = Sequential()
model.add(Convolution2D(32, 3, 3, input_shape=(32, 32, 3)))
model.add(MaxPooling2D((2, 2)))
model.add(Activation('relu'))
model.add(Flatten())
model.add(Dense(128))
model.add(Activation('relu'))
model.add(Dense(43))
model.add(Activation('softmax'))
# +
# STOP: Do not change the tests below. Your implementation should pass these tests.
from keras.layers.core import Dense, Activation, Flatten
from keras.layers.convolutional import Convolution2D
from keras.layers.pooling import MaxPooling2D
check_layers([Convolution2D, MaxPooling2D, Activation, Flatten, Dense, Activation, Dense, Activation], model.layers)
assert model.layers[1].pool_size == (2, 2), 'Second layer must be a max pool layer with pool size of 2x2'
model.compile('adam', 'categorical_crossentropy', ['accuracy'])
history = model.fit(X_normalized, y_one_hot, batch_size=128, nb_epoch=2, validation_split=0.2)
assert(history.history['val_acc'][-1] > 0.91), "The validation accuracy is: %.3f. It should be greater than 0.91" % history.history['val_acc'][-1]
print('Tests passed.')
# -
# ## Dropout
# 1. Re-construct the network
# 2. Add a [dropout](https://keras.io/layers/core/#dropout) layer after the pooling layer. Set the dropout rate to 50%.
# +
# TODO: Re-construct the network and add dropout after the pooling layer.
from keras.models import Sequential
from keras.models import Sequential
from keras.layers.core import Dense, Activation, Flatten, Dropout
from keras.layers.convolutional import Convolution2D
from keras.layers.pooling import MaxPooling2D
model = Sequential()
model.add(Convolution2D(32, 3, 3, input_shape=(32, 32, 3)))
model.add(MaxPooling2D((2, 2)))
model.add(Dropout(0.5))
model.add(Activation('relu'))
model.add(Flatten())
model.add(Dense(128))
model.add(Activation('relu'))
model.add(Dense(43))
model.add(Activation('softmax'))
# +
# STOP: Do not change the tests below. Your implementation should pass these tests.
from keras.layers.core import Dense, Activation, Flatten, Dropout
from keras.layers.convolutional import Convolution2D
from keras.layers.pooling import MaxPooling2D
check_layers([Convolution2D, MaxPooling2D, Dropout, Activation, Flatten, Dense, Activation, Dense, Activation], model.layers)
assert model.layers[2].p == 0.5, 'Third layer should be a Dropout of 50%'
model.compile('adam', 'categorical_crossentropy', ['accuracy'])
history = model.fit(X_normalized, y_one_hot, batch_size=128, nb_epoch=2, validation_split=0.2)
assert(history.history['val_acc'][-1] > 0.91), "The validation accuracy is: %.3f. It should be greater than 0.91" % history.history['val_acc'][-1]
print('Tests passed.')
# -
# ## Optimization
# Congratulations! You've built a neural network with convolutions, pooling, dropout, and fully-connected layers, all in just a few lines of code.
#
# Have fun with the model and see how well you can do! Add more layers, or regularization, or different padding, or batches, or more training epochs.
#
# What is the best validation accuracy you can achieve?
# +
# TODO: Build a model
from keras.models import Sequential
from keras.models import Sequential
from keras.layers.core import Dense, Activation, Flatten, Dropout
from keras.layers.convolutional import Convolution2D
from keras.layers.pooling import MaxPooling2D
model = Sequential()
model.add(Convolution2D(32, 3, 3, input_shape=(32, 32, 3)))
model.add(MaxPooling2D((2, 2)))
model.add(Dropout(0.5))
model.add(Activation('relu'))
model.add(Flatten())
model.add(Dense(128))
model.add(Activation('relu'))
model.add(Dense(43))
model.add(Activation('softmax'))
# There is no right or wrong answer. This is for you to explore model creation.
# TODO: Compile and train the model
model.compile('adam', 'categorical_crossentropy', ['accuracy'])
history = model.fit(X_normalized, y_one_hot, nb_epoch=10, validation_split=0.2)
# -
# **Best Validation Accuracy:** (fill in here)
# ## Testing
# Once you've picked out your best model, it's time to test it.
#
# Load up the test data and use the [`evaluate()` method](https://keras.io/models/model/#evaluate) to see how well it does.
#
# Hint 1: The `evaluate()` method should return an array of numbers. Use the [`metrics_names`](https://keras.io/models/model/) property to get the labels.
# +
# TODO: Load test data
with open('test.p', 'rb') as f:
data_test = pickle.load(f)
X_test = data_test['features']
y_test = data_test['labels']
# TODO: Preprocess data & one-hot encode the labels
X_normalized_test = normalize_grayscale(X_test)
y_one_hot_test = label_binarizer.fit_transform(y_test)
# TODO: Evaluate model on test data
metrics = model.evaluate(X_normalized_test, y_one_hot_test)
for metric_i in range(len(model.metrics_names)):
metric_name = model.metrics_names[metric_i]
metric_value = metrics[metric_i]
print('{}: {}'.format(metric_name, metric_value))
# -
# **Test Accuracy:** (fill in here)
# ## Summary
# Keras is a great tool to use if you want to quickly build a neural network and evaluate performance.
|
Lesson 1 - Deep Learning and Computer Vision/CarND-KerasLab/traffic-sign-classification-with-keras-solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basics of NLP
# ## Topics covered:
# 1. Tokenization
# 2. Removing stop words
# 3. Normalization
# !pip install wordcloud
# +
import wordcloud
import nltk #natural language tool kit
import pandas as pd
import matplotlib.pyplot as plt
import io
import unicodedata
import numpy as np
import re
import string
# -
nltk.download('stopwords')
nltk.download('wordnet')
nltk.download('punkt')
nltk.download('averaged_perceptron_tagger')
# # Tokenization
# ## Tokenization is the process of tokenizing or splitting a string, text into a list of tokens. One can think of token as parts like a word is a token in a sentence, and a sentence is a token in a paragraph.
from nltk.tokenize import WhitespaceTokenizer, WordPunctTokenizer, TreebankWordTokenizer
sentence_1 = "ram, shyam and bob are good boys. hello world. bob's a nice boy."
# ## Sentence tokenization
# ### Delimiters = fullstops
# +
#tokens = sentence
sentence_tokens = nltk.sent_tokenize(sentence_1)
print("tokens = ",sentence_tokens)
print("number of tokens = ",len(sentence_tokens))
# -
# ## White space tokenization
# ### Delimiter = white space
# +
tk = WhitespaceTokenizer()
whitespace_tokens = tk.tokenize(sentence_1)
print("tokens = ",whitespace_tokens)
print("number of tokens = ",len(whitespace_tokens))
# -
# ## Word punctuation tokenizer
# ### Seperates punctuation from words
# +
tk = WordPunctTokenizer()
word_punctuation_tokens = tk.tokenize(sentence_1)
print("tokens = ",word_punctuation_tokens)
print("number of tokens = ",len(word_punctuation_tokens))
# -
# ## Tree bank Word Tokenizer
#
# +
tk = TreebankWordTokenizer()
tree_bank_tokens = tk.tokenize(sentence_1)
print("tokens = ",tree_bank_tokens)
print("number of tokens = ",len(tree_bank_tokens))
# -
# ------
# # Token normalization
# ## Text normalization is the process of transforming a text into a canonical form. That is, bringing a sentence to a predefined standard. it can be done in two ways,
#
# ### 1. Stemming
# ### 2. Lemmatization
# ----
# # Stop Words
#
# ## Before we normalisze, we need to get rid of the stop words, stop words are words that are common in any language like 'the', 'and' etc which dont add value to the analysis
# +
from nltk.corpus import stopwords
stop_words = set(stopwords.words('english'))
print("stop words count = ", len(stop_words))
print("stop words in english are : \n")
print(stop_words)
# -
# removing stop words
whitespace_tokens_op = [w for w in whitespace_tokens if not w in stop_words]
print("with tokens :: ",whitespace_tokens)
print("without tokens :: ",whitespace_tokens_op)
# # Stemming
# ## Stemming is the procress of removing or replacing the suffix of a word to get the root words. for example,
# ### wolf, wolves -> wolf
# ### talk, talks -> talk
# ### bob, bob's -> bob
from nltk.stem import PorterStemmer
|
Natural Language Procressing/1 Basics of NLP.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import sys
if "../" not in sys.path:
sys.path.append("../")
from lib.envs.blackjack import BlackjackEnv
env = BlackjackEnv()
# +
def print_observation(observation):
score, dealer_score, usable_ace = observation
print("Player Score: {} (Usable Ace: {}), Dealer Score: {}".format(
score, usable_ace, dealer_score))
def strategy(observation):
score, dealer_score, usable_ace = observation
# Stick (action 0) if the score is > 20, hit (action 1) otherwise
return 0 if score >= 20 else 1
for i_episode in range(20):
observation = env.reset()
for t in range(100):
print_observation(observation)
action = strategy(observation)
print("Taking action: {}".format( ["Stick", "Hit"][action]))
observation, reward, done, _ = env.step(action)
if done:
print_observation(observation)
print("Game end. Reward: {}\n".format(float(reward)))
break
# -
|
RL/chapter5MC/Blackjack Playground.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# %matplotlib notebook
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
import pyflow
from pyflow.potential import MurmanCole
from pyflow.potential import CircularAirfoil
# -
# # Full Potential Equation
#
# If the flow is irrotational, $\nabla \times \mathbf{u} = 0$, then the governing equations of the fluid dynamics can be determined by a potential function. Starting with the steady Euler equations,
#
# $$
# \begin{align}
# & \nabla \cdot (\rho \mathbf{u}) = 0 \\
# & \rho \mathbf{u} \cdot \nabla \mathbf{u} = -\nabla p
# \end{align}
# $$
#
# Using the above definition,
#
# $$
# \begin{align}
# & \mathbf{u} \cdot \nabla \rho + \rho \nabla \cdot \mathbf{u} = 0 \\
# & \rho \left( \mathbf{u} \cdot \nabla \right) \mathbf{u} = -a^2 \nabla \rho
# \end{align}
# $$
#
# Multiplying the momentum equation with the velocity vector,
#
# $$
# \mathbf{u} \cdot \left( \rho \left( \mathbf{u} \cdot \nabla \right) \mathbf{u}\right) = a^2 \mathbf{u} \cdot \nabla \rho
# $$
#
# and then using the mass conservation equation,
#
# $$
# \rho \left( \mathbf{u} \cdot \left( \mathbf{u} \cdot \nabla\right) \mathbf{u} \right) = a^2 \rho \nabla \cdot \mathbf{u}
# $$
#
# Dividing by the density, and the rearranging
#
# $$
# \nabla \cdot \mathbf{u} - \frac{ \mathbf{u} \cdot \left( \mathbf{u} \cdot \nabla\right) }{a^2} \mathbf{u} = 0
# $$
#
# Using the definition of the potential, $\mathbf{u} = \nabla \phi$, we get the following
#
# $$
# \nabla^2 \phi - \frac{ \nabla \phi \cdot \left( \nabla \phi \cdot \nabla\right) }{a^2} \nabla \phi = 0
# $$
#
# If $a$ is the speed of sound, then we can define a local Mach number vector as
#
# $$
# \mathbf{M} = \frac{\mathbf{u}}{a} = \frac{1}{a} \nabla \phi
# $$
#
#
# Using this definition,
#
# $$
# \nabla^2 \phi - \mathbf{M} \cdot \left( \mathbf{M} \cdot \nabla\right) \nabla \phi = 0
# $$
#
#
# $$
# \nabla \times \mathbf{u} = \nabla \times \left( \nabla \phi \right) = 0
# $$
#
# where in Cartesian coordinates, the velocity vector is given by the gradient of the potential, $\phi$,
#
# $$
# \mathbf{u} = \nabla \phi = \left[ \frac{\partial \phi}{\partial x} \, , \frac{\partial \phi}{\partial y} \, , \frac{\partial \phi}{\partial z} \right]
# $$
#
#
# ## Linearization of the potential equation
#
# In two-dimensions, the full potential equation is written as
#
# $$
# (a^2 - \phi_x^2)\phi_{xx} - 2 \phi_x \phi_y \phi_{xy} +
# (a^2 - \phi_y^2)\phi_{yy} = 0
# $$
#
# where
#
# $$
# \phi_x = \frac{\partial \phi}{\partial x} \,\, \textrm{and} \,\,
# \phi_{xx} = \frac{\partial^2 \phi}{\partial x^2}
# $$
#
# Let us assume the freestream flow is uniform and steady and the flow conditions are defined by $M_{\infty}$, $a_{\infty}$, $\rho_{\infty}$, etc. Now consider what happens to the flow when the gas moves over a slender object. If the disturbance or pertubation of the flow is small, then we can simplify our governing equations. We assume that velocity components are perturbed by $\hat{u}$ and $\hat{v}$, such that
#
# $$
# \begin{align}
# &u = u_{\infty} + \hat{u} \\
# &v = \hat{v}
# \end{align}
# $$
#
# where
#
# $$
# \frac{\hat{u}}{u_{\infty}} << 1
# $$
#
# Using the definition of the potential,
#
# $$
# \hat{u} = \frac{\partial \hat{\phi}}{\partial x} = \hat{\phi}_x
# \,\, \textrm{and} \,\, \hat{v} = \hat{\phi}_y
# $$
#
# which results in
#
# $$
# \begin{align}
# &u = u_{\infty} + \hat{\phi}_x \\
# &v = \hat{v}
# \end{align}
# $$
#
# or in terms of the potential $(\phi_{\infty} = u_{\infty}x)$,
#
# $$
# \begin{align}
# &\phi_x = u_{\infty} + \hat{\phi}_x \\
# &\phi_y = \hat{\phi}_y
# \end{align}
# $$
#
# The second derivatives of the potential are
#
# $$
# \begin{align}
# &\phi_{xx} = \hat{\phi}_{xx} \\
# &\phi_{yy} = \hat{\phi}_{yy} \\
# &\phi_{xy} = \hat{\phi}_{xy}
# \end{align}
# $$
#
# Using these expressions, we get
#
# $$
# \Big( a^2 - (u_{\infty} + \hat{\phi}_x)^2 \Big) \hat{\phi}_{xx} - 2 (u_{\infty} + \hat{\phi}_x) \hat{\phi}_y \hat{\phi}_{xy} +
# (a^2 - \hat{\phi}_y^2)\hat{\phi}_{yy} = 0
# $$
#
# Since we require that $\hat{u}$ to be much less than $u_{\infty}$, terms like
#
# $$
# \hat{\phi}_x^2, \,\,\, \hat{\phi}_y^2, \,\,\, \hat{\phi}_x\hat{\phi}_y
# $$
#
# are assumed to be negilible. Removing those terms from the above equation results in
#
# $$
# \Big( a^2 - (u_{\infty} + \hat{\phi}_x)^2 \Big) \hat{\phi}_{xx} +
# a^2 \hat{\phi}_{yy} = 0
# $$
#
# $$
# \Big( a^2 - u_{\infty}^2 - \hat{\phi}_x^2 - 2 u_{\infty}\hat{\phi}_x \Big)\hat{\phi}_{xx} +
# a^2 \hat{\phi}_{yy} = 0
# $$
#
# and assuming $\hat{\phi}_x^2 << 1$,
#
# $$
# \Big( a^2 - u_{\infty}^2 - 2 u_{\infty} \hat{\phi}_x \Big) \hat{\phi}_{xx} + a^2 \hat{\phi}_{yy} = 0
# $$
#
# With the assumption of isentropic flow, the energy equation can be written as
#
# $$
# \frac{u^2 + v^2}{2} + \frac{a^2}{\gamma - 1} = \frac{u^2_{\infty}}{2} + \frac{a^2_{\infty}}{\gamma - 1}
# $$
#
# which we can write as
#
# $$
# \frac{(u_{\infty} + \hat{u})^2 + \hat{v}^2}{2} + \frac{a^2}{\gamma - 1} = \frac{u^2_{\infty}}{2} + \frac{a^2_{\infty}}{\gamma - 1}
# $$
#
# and can be expanded as
#
# $$
# \frac{a^2}{\gamma - 1} = \frac{u^2_{\infty}}{2} - \frac{u^2_{\infty} + \hat{u}^2 + 2u_{\infty}\hat{u} + \hat{v}^2}{2} + \frac{a^2_{\infty}}{\gamma - 1}
# $$
#
# simplifying
#
# $$
# a^2 = a^2_{\infty} - \left({\gamma - 1} \right)\frac{\hat{u}^2 + 2u_{\infty}\hat{u} + \hat{v}^2}{2}
# $$
#
# Since $\hat{u}^2$ and $\hat{v}^2$ are much smaller than one, we can assume they are negilible,
#
# $$
# a^2 = a^2_{\infty} - (\gamma -1) u_{\infty} \hat{\phi}_x
# $$
#
# Using this expression for $a^2$, let us plug it into the linearized potential equation,
#
# $$
# \Big(a^2_{\infty} - (\gamma -1) u_{\infty} \hat{\phi}_x - u_{\infty}^2 - 2 u_{\infty} \hat{\phi}_x \Big) \hat{\phi}_{xx} +
# \Big(a^2_{\infty} - (\gamma -1) u_{\infty} \hat{\phi}_x\Big) \hat{\phi}_{yy} = 0
# $$
#
# Assume
#
# $$
# \hat{\phi}_x \hat{\phi}_{yy} << 1
# $$
#
# which results in
#
# $$
# \Big(a^2_{\infty} - (\gamma -1) u_{\infty} \hat{\phi}_x - u_{\infty}^2 - 2 u_{\infty} \hat{\phi}_x \Big) \hat{\phi}_{xx} +
# a^2_{\infty} \hat{\phi}_{yy} = 0
# $$
#
# Dividing by $a_{\infty}^2$ and simplifying results in the transonic small distrubance equation is
#
# $$
# \left[ 1 - M^2_{\infty} - \left(\gamma + 1\right) M^2_{\infty} \frac{\hat{\phi}_x}{u_{\infty}} \right] \hat{\phi}_{xx} + \hat{\phi}_{yy} = 0
# $$
#
# $$
# \rho \approx \rho_{\infty}\left(1 - M^2_{\infty}\left(\frac{u}{u_{\infty}} - 1\right)\right)
# $$
#
# $$
# p \approx p_{\infty}\left(1 - \gamma M^2_{\infty}\left(\frac{u}{u_{\infty}} - 1\right)\right)
# $$
#
# $$
# c_p = \frac{p - p_{\infty}}{\frac{1}{2}\rho_{\infty} u^2_{\infty}} \approx -2 \frac{\hat{u}}{u_{\infty}}
# $$
#
# As a result of the linearization, the transonic small distrubance equation describes the full around slender bodies, for larger distrubances, e.g., blunt-nosed airfoils, the full potential equations must be used. Both equations, however, assume the flow to be isentropic. The assumption of isentropic is that the flow is reversible, requiring only gradual changes in the thermodynamic quantities. As such the formation of a shock wave violates this assumption, since the flow is now irreversible.
#
# **Only weak shocks.** Strong shocks increase the entropy within the flow significantly, which would violate the assumption that the gas is isentropic. The isentropic equations are approximately valid to Mach numbers less than 1.3.
# # Model Elliptic Equation
#
# The second-order partial differential equation
#
# $$
# A \phi_{xx} + \phi_{yy} = 0
# $$
#
# where $A > 0$ is classified as an elliptic PDE. Elliptic equations have no characteristic curves. For both hyperbolic and parabolic equations, information ("perturbations") travels along characteristic curves. As a result, for elliptic equations, there is no mathematical mechanism present to support the concept of information propagation. In physics, elliptic equations often describe equilibrium states or potentials, e.g., gravity, electrical, etc.
#
#
# We also note that if $A < 0$, then the equation is hyperbolic. We should expect this, since in the transonic small perturbation equation, we have
#
# $$
# A =1 - M^2_{\infty} - \left(\gamma + 1\right) M^2_{\infty} \frac{\hat{\phi}_x}{u_{\infty}}
# $$
#
# which is less than one when the Mach number is greater than 1.
#
#
# ## Boundary conditions
#
# Since the PDE is elliptic, we must specify a boundary condition for the far-field boundary of the domain. We will apply two different types of boundary condition. A Dirchilet boundary condition,
#
# $$
# \phi = \theta_{bdry}
# $$
#
# where the value of $\phi$ along the boundary is specified directly and held constant throughout the simulation. The other type of boundary condition is a von Neumann boundary condition,
#
# $$
# \frac{\partial \phi}{\partial n}\bigg\rvert_{bdry} = 0
# $$
#
# where the gradient of $\phi$ normal to the boundary is specified. Notice that for the potential equation, specifying the gradient is conceptually equivalent to specifying the velocity. At the surface, the velocity vector should be tangential to the body, or stated another way perpendicular to the normal vector of the body. Let the function $f(x,y)$ represent the curve of the surface, then we can construct a boundary condition from the requirement that
#
# $$
# \mathbf{u} \cdot \nabla f = 0
# $$
#
# which for $u=u_{\infty} + \hat{u}$ and $v=\hat{v}$ is
#
# $$
# \Big(u_{\infty} + \hat{u}\Big) \frac{\partial f}{\partial x} + \hat{v}\frac{\partial f}{\partial y} = 0
# $$
#
# Since $u_{\infty} >> \hat{u}$, we can state
#
# $$
# \frac{\hat{v}}{u_{\infty}} \approx - \frac{\partial f / \partial x}{\partial f / \partial y} = -\frac{\textrm{d}y}{\textrm{d}x}
# $$
#
# Since the body must be "thin" in order for the perturbations to be small, then we can approximate the velocity perturbation using a Taylor series expansion about $y=0$,
#
# $$
# \hat{v}(x,y) = \hat{v}(x,0) + \left(\frac{\partial \hat{v}}{\partial y}\right)\bigg\rvert_{y=0}y + \dots
# $$
#
# Assuming the gradient of the $\hat{v}$ is small at the body,
#
# $$
# \frac{\partial \phi}{\partial y} = \hat{v}(x,y) = \hat{v}(x,0) = u_{\infty} \left( \frac{\textrm{d}y}{\textrm{d}x}\right)_{\textrm{body}}
# $$
#
# ## Simulation Domain
#
# The chord length is defined as $c$. We define the width and height of the domain as $L_x$ and $L_y$, respectively. Let us use, $L_x = 50c$ and $L_y = 50c$, where the airfoil is centered in domain along the $y=0$ boundary. Note that because we assume the airfoil is thin (small distrubance theory), then we do not need to use a body-fitted mesh.
#
# <img src="symmetric_arifoil.png" alt="symmetric_arifoil.png" width="500" height="600">
#
#
# Even though we do not need to use a body-fitted mesh, we can visualize the circular arc airfoil using the following code.
# +
mesh = CircularAirfoil(6, stretch_mesh=True, Nx=51, Ny=51)
xx, yy = mesh.plot_circular_arc()
fig = plt.figure(figsize=(9,6))
ax = fig.add_subplot(111)
line = ax.plot(xx, yy, c='k', ls='-', label="Symmetric Airfoil")
ax.legend(fontsize=12)
ax.set_xlabel(r'$x$', fontsize=12)
ax.set_ylabel(r'$y$', fontsize=12)
ax.set_ylim((0,0.1))
# -
# ### Two-dimensional Mesh
#
# We will use a $N_x \times N_y$ mesh where $N_x = N_y = 51$. In the $x$-direction, the mesh is uniformally spaced across the chord of the airfoil using 21 points and then exponentially stretched to the left and right far-field boundaries $50c$ away. In the $y$-direction, the mesh is exponentially stretched to the top far-field boundary, also $50c$ away.
#
# The mesh spacing in the x-direction across the airfoil is
#
# $$
# \Delta x = \frac{c}{N_{\textrm{chord}} - 1}
# $$
#
# where $N_{\textrm{chord}} = 21$. We will define the leading-edge of the airfoil as the origin of our simulation domain, thus we can state,
#
# $$
# \Delta x = \frac{c}{N_{\textrm{chord}} - 1} \qquad \textrm{for} \, 0 \leq x \leq c
# $$
#
# For $x < 0$ and $x > c$, we will exponentially stretch the mesh using the remaining points. Let us evenally distribute the remaining points to the left and right and define
#
# $$
# N_{\textrm{flow}} = \frac{1}{2} \big(N_x - N_{\textrm{chord}} \big)
# $$
#
# For $N_x = 51$ and $N_{\textrm{chord}}$, $N_{\textrm{flow}} = 15$. We will use the following function to define the mesh size
#
# $$
# x_i - x_0 = \big( L_{\textrm{flow}} \big) \frac{e^{\kappa_x \left(\frac{i-1}{N_{\textrm{flow}}\,-\,1}\right)} - 1}{e^{\kappa_x} - 1}
# $$
#
# However, the parameter $\kappa_x$ is unknown, since all we are given is $L_{\textrm{flow}}$, $N_{\textrm{flow}}$, and $x_0$. To find a value of $\kappa_x$, we will specify a value for the minimum spacing, $\Delta x_{\textrm{min}}$, and then solve for $\kappa_x$.
#
# For the $y$-direction, we use the same exponential stretching function,
#
# $$
# y_i - y_0 = \big( L_y \big) \frac{e^{\kappa_y \left(\frac{i-1}{N_y\,-\,1}\right)} - 1}{e^{\kappa_y} - 1}
# $$
#
# where once again the parameter $\kappa_y$ is unknown, since all we are given is $L_y$, $N_y$, and $y_0$. To find a value of $\kappa_y$, we will specify a value for the minimum spacing at the airfoil, $\Delta y_{\textrm{min}}$ and then solve for $\kappa_y$. We will use
#
# $$
# \Delta y_{\textrm{min}} = 0.01 \, t
# $$
#
# where $t$ is the thickness of the circular-arc airfoil.
#
# ### Solving for the stretching factor, $\kappa$
#
# Take the $x$-direction as an example. The same procedure described below applies the $y$-direction as well. The minimum
#
# $$
# \Delta x_{\textrm{min}} = x_1 - x_0 = \big( L_{\textrm{flow}} \big) \frac{e^{\kappa \left(\frac{1}{N_{\textrm{flow}}\,-\,1}\right)} - 1}{e^{\kappa} - 1}
# $$
#
# To use Newton's method, we can define the following function,
#
# $$
# f(\kappa) = \big( L_{\textrm{flow}} \big) \frac{e^{\kappa \left(\frac{1}{N_{\textrm{flow}}\,-\,1}\right)} - 1}{e^{\kappa} - 1} - \Delta x_{\textrm{min}}
# $$
#
# By inspection of the function above, if $f(\kappa)$ is zero, then the value of $\kappa$, corresponds to the desired value of $\Delta x_{\textrm{min}}$. Starting with some intial guess for $\kappa^n$, we can compute what our next value of $\kappa^{n+1}$ should be according to the following expansion,
#
# $$
# 0 = \frac{\partial f}{\partial \kappa} \big(\kappa^{n+1} - \kappa^n\big) + f(\kappa^n) = f^{\prime}(\kappa^{n}) \big[\kappa^{n+1} - \kappa^n\big] + f(\kappa^n)
# $$
#
# or
#
# $$
# \kappa^{n+1} = \kappa^n - \frac{f(\kappa^n)}{f^{\prime}( \kappa)}
# $$
# # Murman-Cole Method
#
# Use the Murman-Cole method to find a solution to the transonic small distrubance equation for flow past a thin airfoil. Consider two different freestream conditions
#
# - Case 1: Subsonic flow with $M_{\infty} = 0.5$
# - Case 2: Transonic flow with $M_{\infty} = 0.908$
#
# The thin airfoil in the above figure is symmetric and defined as a circular arc with a thickness of $t$, where
#
# $$
# t = \frac{\textrm{Thickness Percentage}}{100}
# $$
#
# Using geometry, the radius of curvature is defined as
#
# $$
# R = \frac{t c}{4} + \frac{1}{4} \frac{c^2}{t * c}
# $$
#
# Using $x = R \cos(\theta)$ and $y = R \sin(\theta)$, we can define the geometry of the airfoil. However, because the airfoil is thin and we are using the transonic, small distrubance equation, we do not need to use a body-fitted mesh. The only geometrical definition we need occurs at the airfoil surface boundary, where the flow tangency boundary condition requires,
#
# $$
# \frac{\partial \hat{\phi}}{\partial y} = V_{\infty} \left[ \frac{\textrm{d} y(x)}{\textrm{d} x}\right]_{\textrm{body}}
# $$
#
# From geometry, we know that for $0 \leq x < c/2$
#
# $$
# \frac{\textrm{d} y(x)}{\textrm{d} x}\bigg\rvert_{\textrm{body}} = \frac{t}{c}
# $$
#
# and $c/2 \leq x < c$
#
# $$
# \frac{\textrm{d} y(x)}{\textrm{d} x}\bigg\rvert_{\textrm{body}} = -\frac{t}{c}
# $$
#
# Assume that the pressure and density is normalized such that $V_{\infty}=1$ and $\rho_{\infty} = 1$ with $\gamma = 1.4$. Using $V_{\infty} = M_{\infty}a_{\infty}$,
#
# $$
# p_{\infty} = \frac{1}{\gamma} \rho_{\infty} a^2_{\infty} = \frac{1}{\gamma M_{\infty}^2}
# $$
#
# Since $\hat{\phi}$ is the distrubance of the potential, we can set it to any constant initially. Let us use, $\hat{\phi} = 1$. Remember we are interested in the derivatives of $\hat{\phi}$ with respect to $x$ and $y$. Take the coefficient of pressure, it is computed as
#
# $$
# c_p = \frac{p - p_{\infty}}{q_{\infty}}
# $$
#
# where the dynamic pressure is given by
#
# $$
# q_{\infty} = \frac{1}{2} \rho_{\infty} V^2_{\infty}
# $$
#
# and the pressure as
#
# $$
# p = p_{\infty}\left[1 + \frac{\gamma - 1}{2} M_{\infty}^2 \left(\frac{u^2 + v^2}{V_{\infty}^2} -1 \right)\right]^{\frac{\gamma}{\gamma - 1}}
# $$
#
# The solutions to the transonic, small distrubance equation give the value of the distrubance potential, $\hat{\phi}$, which is related to the velocity by
#
# $$
# \begin{align}
# u &= V_{\infty} + \hat{\phi}_x = V_{\infty} + \frac{\partial \hat{\phi}}{\partial x}\\
# v &= \hat{\phi}_y = \frac{\partial \hat{\phi}}{\partial y}
# \end{align}
# $$
# ## Subsonic Flow
mesh = CircularAirfoil(6, stretch_mesh=True, Nx=51, Ny=51)
flow = MurmanCole(mesh, M_inf=0.5)
flow.solve(print_residuals=100,max_residual=1.0e-5,max_iterations=2000)
# +
fig = plt.figure(figsize=(9,6))
ax = fig.add_subplot(111)
ax.plot(mesh.x, -flow.cp[:,2], marker='o')
ax.set_xlim((-0.1,1.1))
ax.set_xlabel(r'$x$', fontsize=12)
ax.set_ylabel(r'$-c_p$', fontsize=12)
# +
fig = plt.figure(figsize=(9,6))
ax = fig.add_subplot(111)
CS = plt.contour(flow.X, flow.Y, flow.M)#, levels=[0.8, 0.9, 1.0, 1.1, 1.2 ], colors=['y', 'b', 'r', 'g','k'])
cbar = fig.colorbar(CS)
cbar.ax.set_ylabel('Mach Number')
ax.minorticks_on()
ax.set_xlim((-0.25,1.25))
ax.set_ylim((0,1))
ax.set_xlabel(r'$x$', fontsize=12)
ax.set_ylabel(r'$y$', fontsize=12)
# +
fig = plt.figure(figsize=(9,6))
ax = fig.add_subplot(111)
CS = plt.pcolor(flow.X, flow.Y, flow.phi)
cbar = fig.colorbar(CS)
cbar.ax.set_ylabel('Potential')
ax.minorticks_on()
ax.set_xlim((-0.25,1.25))
ax.set_ylim((0,0.1))
ax.set_xlabel(r'$x$', fontsize=12)
ax.set_ylabel(r'$y$', fontsize=12)
# -
|
notebooks/MurmanColeMethod.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h2>Scipy</h2>
# This notebook shows the most important functions of Scipy, a Python library for scientific mathematical and statistical routines.
#
# Usually, Scipy is bundled with common Python distribution, such as Anaconda (https://www.continuum.io/anaconda). If it is not bundled with your distribution and you use pip, you can install Scipy with the command "pip install scipy". A free book which is including lectures about Scipy can be found under http://www.scipy-lectures.org (most of the examples shown in this notebook are derived from this book).
#
# After you installed Scipy on your device, you can import it into your Python scripts with the following commands:
from scipy import optimize
import numpy
# <h3>1. Optimization</h3>
# The following function shall be searched for global minima
def to_optimize(x):
return x**2 + 10*x
# The BFGS global minimum finder starts from a start point (here: 0)
# It may only find a local minimum, depending on the start point
optimize.fmin_bfgs(to_optimize, 0)
# The basinhopper algorithm is an alternative to BFGS
# It is said to be more reliable than BFGS
optimize.basinhopping(to_optimize, 0)
# In order to find a local minimum in an interval, an other method is used
optimize.fminbound(to_optimize, 0, 10)
# Find one of the roots of a function, probalby using an approximated Newton Method (the numeric parameter is an initial guess)
# In order to find more roots, other inital guesses have to be used
root = optimize.fsolve(to_optimize, 1)
root
# <h3>2. Fitting</h3>
# +
# Complex Curve Fitting
# We shall find parameters a and b which shall give the best fit to a number of points
# For polynomial fitting, you can also use NumPy
def to_fit(x, a, b):
return a*(x/2) + b*x
xpoints = numpy.linspace(-5, 5, num=10)
ypoints = numpy.linspace(5, 15, num=10)
initial_guess = [0, 0]
parameters, paramameters_covariance = optimize.curve_fit(to_fit, xpoints, ypoints, initial_guess)
parameters
# -
paramameters_covariance
# <h3>3. Numeric integration</h3>
# For symbolic integration, you can use SymPy.
from scipy.integrate import quad
# quad is a very general integrator
result, error = quad(numpy.cos, 0, 1)
result
from scipy.integrate import odeint
# This ordinary differential equation shall be solved (dy/dt=-y)...
def ode(y, t):
return -y
# ... which is done by using odeint
timepoints = numpy.linspace(0, 4, 5)
yvector, information = odeint(ode, 1, timepoints,
args=(), full_output=True)
yvector
information
# <h3>4. Statistics</h3>
from scipy import stats
# Fit data points to normal distribution
to_fit = numpy.random.normal(size=100)
loc, std = stats.norm.fit(to_fit)
loc
std
# Random numbers should be used using NumPy
numpy.random.rand(5)
# Many more statistical methods and tests are available. A table of available routines can be found under https://docs.scipy.org/doc/scipy/reference/stats.html.
# PSB 2017
|
python_module_tutorials/Scipy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:nlrc] *
# language: python
# name: conda-env-nlrc-py
# ---
# +
# Copyright (c) Microsoft Corporation. All rights reserved.
# Licensed under the MIT License.
# +
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = 'all' # default is ‘last_expr’
# %load_ext autoreload
# %autoreload 2
# -
import sys
sys.path.append('/Users/siyuyang/Source/repos/DevOps_AIforGood/nlrc.building-damage-assessment')
# +
import json
import os
from collections import defaultdict
import rasterio.features
import numpy as np
import shapely.geometry
from shapely.geometry import mapping, Polygon
from PIL import Image
from geospatial.visualization.raster_label_visualizer import RasterLabelVisualizer
# need to change the path in data/create_label_masks.py file where the label_map_file is loaded
# to an absolute path when running this notebook or add "../" to go one directory up (cwd is in the 'eval' folder)
from data.create_label_masks import get_feature_info, read_json
# -
# # Evaluate model output at the building level
#
# This notebook was used to develop `eval_building_level.py` and to create test data - use the script version instead when evaluating the model.
# +
viz = RasterLabelVisualizer(label_map='../constants/class_lists/xBD_label_map.json')
all_classes = set([1, 2, 3, 4, 5])
allowed_classes = set([1, 2, 3, 4]) # 5 is Unclassified, not used during evaluation
# -
# ### Sample tile
# A mask of the ground truth tile
# +
l = viz.plot_color_legend()
p = 'test_data/joplin-tornado_00000050_post_disaster_b1.png'
p_np = np.array(Image.open(p))
p_np.shape
np.unique(p_np)
im, buf = viz.show_label_raster(p_np, size=(10, 10))
im
# -
# ### Create a test tile from the sample with corner cases
#
# Manipulate the above tile to get one where one building blob contains pixels of multiple classes
# +
new_test_tile_np = np.array(Image.open(p))
new_test_tile_np.shape
new_test_tile_np[100:125, 700:725] = 2 # should remain classed as class 4
new_test_tile_np[80:85, 670:690] = 3
new_test_tile_np[300:320, 210:235] = 1 # should remain classed as class 4
new_test_tile_np[750:800, 880:945] = 1 # should be classed as class 1
new_test_tile_np[760:765, 890:900] = 2
# a hole of background class - to test the blob-forming logic
new_test_tile_np[20:30, 230:240] = 0
# a false positive
new_test_tile_np[800:900, 200:400] = 2
# a false negative - cover one building in black pixels
new_test_tile_np[400:600, 800:1000] = 0
im, buf = viz.show_label_raster(new_test_tile_np, size=(10, 10))
im
new_test_tile = Image.fromarray(new_test_tile_np)
new_test_tile.save('test_data/test_tile_0.png')
# -
# ### Label polygons
json_post = read_json('test_data/joplin-tornado_00000050_post_disaster.json')
polys = get_feature_info(json_post)
# +
label_polygons_and_class = [] # tuples of (shapely polygon, damage_class_num)
for uid, tup in polys.items():
poly, damage_class_num = tup # poly is a np.ndarray
polygon = Polygon(poly)
if damage_class_num in allowed_classes:
label_polygons_and_class.append((polygon, damage_class_num))
# -
len(label_polygons_and_class)
# ### Predicted polygons
#
# Test with a mask produced from the labels for training
mask_post = np.asarray(Image.open('test_data/test_tile_0.png'))
mask_post.shape
mask_post.dtype
# Method
#
# 1. Detect the connected components by all non-background classes to determine the predicted building blobs first (if we do this per class, a building with some pixels predicted to be in another class will result in more buildings than connected components)
# 2. The majority class for each building blob is assigned to be that building's predicted class.
# +
background_and_others_mask = np.where(mask_post > 0, 1, 0).astype(np.int16) # all non-background classes become 1
# rasterio.features.shapes:
# default is 4-connected for connectivity - see https://www.mathworks.com/help/images/pixel-connectivity.html
# specify the `mask` parameter, otherwise the background will be returned as a shape
connected_components = rasterio.features.shapes(background_and_others_mask, mask=mask_post > 0)
polygons = []
for component_geojson, pixel_val in connected_components:
# reference: https://shapely.readthedocs.io/en/stable/manual.html#python-geo-interface
shape = shapely.geometry.shape(component_geojson)
assert isinstance(shape, Polygon)
polygons.append(shape)
len(polygons) # 15: we took away one and added one (14), and one polygon is of the Unclassified class
# + jupyter={"outputs_hidden": true}
for p in polygons: # flipped when visualized this way
p
# +
polygons_by_class = []
for c in all_classes:
# default is 4-connected for connectivity
shapes = rasterio.features.shapes(mask_post, mask=mask_post == c)
for shape_geojson, pixel_val in shapes:
shape = shapely.geometry.shape(shape_geojson)
assert isinstance(shape, Polygon)
polygons_by_class.append((shape, int(pixel_val)))
len(polygons_by_class) # 20 shapes including the one of class Unclassified
# + jupyter={"outputs_hidden": true}
for p in polygons_by_class: # flipped when visualized this way
p[0]
# +
# we take the class of the shape with the maximum overlap with the building polygon to be the class of the building - majority vote
polygons_max_overlap = [0.0] * len(polygons) # indexed by polygon_i
polygons_max_overlap_class = [None] * len(polygons)
assert isinstance(polygons, list) # need the order constant
for polygon_i, polygon in enumerate(polygons):
for shape, shape_class in polygons_by_class:
intersection_area = polygon.intersection(shape).area
if intersection_area > polygons_max_overlap[polygon_i]:
polygons_max_overlap[polygon_i] = intersection_area
polygons_max_overlap_class[polygon_i] = shape_class
pred_polygons_and_class = [] # include all classes
for polygon_i, (max_overlap_area, clss) in enumerate(zip(polygons_max_overlap, polygons_max_overlap_class)):
pred_polygons_and_class.append(
(polygons[polygon_i], clss)
)
len(pred_polygons_and_class)
# -
for polygon, clss in pred_polygons_and_class:
print(f'area: {polygon.area}, class {clss}')
# ### Matching predicted and label polygons
#
# Method
# - For each predicted polygon, we find the maximum value of IoU it has with any ground truth polygon within the tile. This ground truth polygon is its "match".
# - Using the threshold IoU specified (typically and by default 0.5), if a prediction has overlap above the threshold AND the correct class, it is considered a true positive. All other predictions, no matter what their IOU is with any gt, are false positives.
# - Note that it is possible for one ground truth polygon to be the match for multiple predictions, especially if the IoU threshold is low, but each prediction only has one matching ground truth polygon.
# - For ground truth polygon not matched by any predictions, it is a false negative.
# - Given the TP, FP, and FN counts for each class, we can calculate the precision and recall for each tile *for each class*.
#
#
# - To plot a confusion table, we output two lists, one for the predictions and one for the ground truth polygons (because the set of polygons to confuse over are not the same...)
# 1. For the list of predictions, each item is associated with the ground truth class of the polygon that it matched, or a "false positive" attribute.
# 2. For the list of ground truth polygons, each is associated with the predicted class of the polygon it matched, or a "false negative" attribute.
# +
# DRAFT - see eval_building_level.py
def _evaluate_tile(pred_polygons_and_class: list,
label_polygons_and_class: list,
allowed_classes,
iou_threshold: float=0.5):
# the matched label polygon's IoU with the pred polygon, and the label polygon's index
pred_max_iou_w_label = [(0.0, None)] * len(pred_polygons_and_class)
for i_pred, (pred_poly, pred_class) in enumerate(pred_polygons_and_class):
# cannot skip pred_class if it's not in the allowed list, as the list above relies on their indices
for i_label, (label_poly, label_class) in enumerate(label_polygons_and_class):
intersection = pred_poly.intersection(label_poly)
union = pred_poly.union(label_poly) # they should not have zero area
iou = intersection.area / union.area
if iou > pred_max_iou_w_label[i_pred][0]:
pred_max_iou_w_label[i_pred] = (iou, i_label)
results = defaultdict(lambda: defaultdict(int)) # class: {tp, fp, fn} counts
i_label_polygons_matched = set()
list_preds = []
list_labels = []
for i_pred, (pred_poly, pred_class) in enumerate(pred_polygons_and_class):
if pred_class not in allowed_classes:
continue
max_iou, matched_i_label = pred_max_iou_w_label[i_pred]
if matched_i_label is not None:
i_label_polygons_matched.add(matched_i_label)
item = {
'pred': pred_class,
'label': label_polygons_and_class[matched_i_label][1] if matched_i_label is not None else None
}
list_preds.append(item)
list_labels.append(item)
if max_iou > iou_threshold and label_polygons_and_class[matched_i_label][1] == pred_class:
# true positive
results[pred_class]['tp'] += 1
else:
# false positive - all other predictions
results[pred_class]['fp'] += 1 # note that it is a FP for the prediction's class
# calculate the number of false negatives - how many label polygons are not matched by any predictions
for i_label, (label_poly, label_class) in enumerate(label_polygons_and_class):
if label_class not in allowed_classes:
continue
if i_label not in i_label_polygons_matched:
results[label_class]['fn'] += 1
list_labels.append({
'pred': None,
'label': label_class
})
return results, list_preds, list_labels
# -
results, list_preds, list_labels = _evaluate_tile(pred_polygons_and_class,
label_polygons_and_class,
allowed_classes)
for clss, res in results.items():
print(clss)
print(res)
print()
list_preds
list_labels
|
eval/eval_building_level.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# Método para resolver las energías y eigenfunciones de un sistema cuántico numéricamente
# Modelado Molecular 2
# By: <NAME>
import numpy as np
from sympy import *
from sympy import init_printing; init_printing(use_latex = 'mathjax')
import matplotlib.pyplot as plt
# Variables utilizadas
var('x l m hbar w k')
def Metodo_variaciones():
print('En este problema están definidas como variables la masa, el parámetro l (se define), el parámetro k(se optimiza) y x')
print('')
# La energía cinética está definida por: K=(-hbar**2)/(2*m)*diff(fx,x,2)
print('La energía cinética esta definida como: K=(-hbar**2)/(2*m)*diff(f(x),x,2)');print('')
# Declarar el potencial
V=sympify(input('Introduce la función de potencial: '));print('')
lim_inf_V=sympify(input('¿Cuál es el límite inferior de la función potencial? '))
lim_sup_V=sympify(input('¿Cuál es el límite superior de la función potencial? '));print('')
n = int(input('Introduce el número de funciones que vas a utilizar para resolver el problema: '));print('')
# Lista para ingresar las funciones
f=[]
# Matriz con integral de solapamiento
Sm=[]
# Matriz con integral de intercambio
Hm=[]
print('Ahora vamos definir las constantes del problema');print('')
mass=input('¿Cuánto es la masa de tu partícula? ')
large=input('Define el parámetro l: ');print('')
# Declarar funciones y límites de dichas funciones
lim_inf=[]
lim_sup=[]
for i in range(n):
f.append((input('Introduce la función %d: ' %(i+1))))
lim_inf.append(input('¿Cuál es el límite inferior de la función? '))
lim_sup.append(input('¿Cuál es el límite superior de la función? '));print('')
f=sympify(f)
lim_inf=sympify(lim_inf)
lim_sup=sympify(lim_sup)
# Para partícula en un pozo de potencial de 0 a l
# El siguiente ciclo for resuelve las integrales para formar las matrices Sij (Integrales de solapamiento)
# y Hij(integrale de intercambio)
# Aproximación de las energías
li=0
ls=0
for i in range(n):
for j in range(n):
integrandoT=(f[i])*((-hbar**2)/(2*m)*diff(f[j],x,2))
integrandoV=(f[i])*V*(f[j])
integrandoN=(f[i])*f[j]
# Definir los limites de integracion
# Límites inferiores
if lim_inf[i].subs({l:large})<=lim_inf[j].subs({l:large}):
li=lim_inf[j]
if li.subs({l:large})>=lim_inf_V.subs({l:large}):
liV=li
else:
liV=lim_inf_V
if lim_inf[i].subs({l:large})>=lim_inf[j].subs({l:large}):
li=lim_inf[i]
if li.subs({l:large})>=lim_inf_V.subs({l:large}):
liV=li
else:
liV=lim_inf_V
# Límites superiores
if lim_sup[i].subs({l:large})>=lim_sup[j].subs({l:large}):
ls=lim_sup[j]
if ls.subs({l:large})<=lim_sup_V.subs({l:large}):
lsV=ls
else:
lsV=lim_sup_V
if lim_sup[i].subs({l:large})<=lim_sup[j].subs({l:large}):
ls=lim_sup[i]
ls=lim_sup[j]
if ls.subs({l:large})<=lim_sup_V.subs({l:large}):
lsV=ls
else:
lsV=lim_sup_V
c=Integral(integrandoT,(x,li,ls))
e=Integral(integrandoV,(x,liV,lsV))
g=c+e
d=Integral(integrandoN,(x,li,ls))
g=g.doit()
Hm.append(g)
d=d.doit()
Sm.append(d)
Sm=np.reshape(Sm,(n,n))
Hm=np.reshape(Hm,(n,n))
# Matriz M: (Hij-Sij)*w
M=(Hm-Sm*w)
H=sympify(Matrix(M))
Hdet=H.det()
# Resolver el determinante para encontrar las energías
E=solve(Hdet,w)
# Ordenar energías
Eord=solve(Hdet,w)
energies=np.zeros(n)
for i in range (n):
energies[i]=E[i].subs({m: mass, l: large, hbar:1.0545718e-34})
energies_ord=sorted(energies)
for i in range(n):
for j in range(n):
if energies[i]==energies_ord[j]:
Eord[i]=E[j]
# Matriz de constantes para todas las eigenfunciones
c=zeros(n)
for i in range(n):
for j in range(n):
c[i,j]=Symbol('c %d %d' %(i+1,j+1))
# Solución a esas constantes
sol=[]
for i in range (n):
a=np.reshape(c[0+n*i:(n)+n*i],(n))
SE=Matrix(np.dot(M,a.transpose()))
SE=sympify((SE.subs({w:Eord[i]})))
sol.append(solve(SE,c[0+n*i:(n+1)+n*i]))
if n!= 1:
csol=zeros(n)
CTS,cts,Cdet=[],[],[]
for i in range (n):
for j in range(n):
csol[i,j]=(sol[i]).get(c[i,j])
if csol[i,j] is None:
csol[i,j]=c[i,j]
CTS.append(c[i,j]); cts.append(c[i,j]); Cdet.append(c[i,j])
# Impresión en pantalla de los resultados
print('<NAME>')
print(sympify(Matrix(Hm)));print('')
print('Matriz Sij')
print(sympify(Matrix(Sm)));print('')
print('Energías ordenadas')
print(Eord);print('')
# Normalizar las funciones de onda y graficar
graficar=input('Desea graficar las eigenfunciones calculadas: ');print('')
if graficar=="si":
if n>1:
fa=(np.reshape(f,(n)))
ef=csol*fa
for i in range(n):
integrando=ef[i]*ef[i]
integ=Integral(integrando,(x,lim_inf[i],lim_sup[i]))
integ=integ.doit()
cts[i]=solve(integ-1,Cdet[i])
if abs(cts[i][0])==cts[0][0]:
CTS[i]=cts[i][0]
else:
CTS[i]=cts[i][1]
ef=ef.subs({Cdet[i]:CTS[i]})
print('Constantes de cada una de las eigenfunciones (cada eigenfunción tiene una constante extra que se debe normalizar)')
print(csol);print('')
print('Para graficar se normalizaron las constantes mostradas anteriormente, cuyos resultados fueron:')
print(CTS);print('')
for i in range(n):
plot(ef[i].subs({l:1}),xlim=(0,1),ylim=(-2,2),title='Eigenfunción: %d' %(i+1))
# Falta automatizar los limites de las funciones para graficar y que te grafique la primer función
if n==1:
ct=Symbol('C22')
ef=ct*f[0]
integrando=(ef)*(ef)
integ=Integral(integrando,(x,lim_inf[0],lim_sup[0]))
integr=integ.doit()
cte=solve(integr-1,ct)
if cte[0].subs({l:large})>cte[1].subs({l:large}):
ctr=cte[0]
else:
ctr=cte[1]
ef=ef.subs({ct:ctr})
#print('Constantes de cada una de las eigenfunciones (cada eigenfunción tiene una constante extra que se debe normalizar)')
#print(csol);print('')
#print('Para graficar se normalizó las constante mostrada anteriormente, cuyo resultado fue:')
#print(CTS);print('')
plot(ef.subs({l:1}),xlim=(0,1),ylim=(-1,2))
return()
Metodo_variaciones()
|
Huckel_M0/Chema/Teorema_de_variaciones(1).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + language="javascript"
# IPython.OutputArea.prototype._should_scroll = function(lines) {
# return false;
# }
# +
# %matplotlib notebook
# Import modules
import numpy as np
import matplotlib.pyplot
from pyne import serpent
from pyne import nucname
from scipy.stats.stats import pearsonr
import itertools
vol_fuel_se = [18.0E+6]
vol_blank_se = [7.3E+6]
vol_storage = [7.3E+6*(50/63498.695312)]#[7.3E+6/1.9097e+04]
iso = 'u233'
#dep0 = serpent.parse_dep('/home/andrei2/Desktop/ornl/msfr/serpent/no_repr_depletion/msfr_depl.inp_dep.m', make_mats=False)
#dep1 = serpent.parse_dep('/home/andrei2/Desktop/ornl/msfr/serpent/gas_removal/msfr_gas_rem.inp_dep.m', make_mats=False)
#dep2 = serpent.parse_dep('/home/andrei2/Desktop/ornl/msfr/serpent/all_removal/msfr_gas_rem.inp_dep.m', make_mats=False)
#res0 = serpent.parse_res('/home/andrei2/Desktop/ornl/msfr/serpent/no_repr_depletion/msfr_depl.inp_res.m')
#res1 = serpent.parse_res('/home/andrei2/Desktop/ornl/msfr/serpent/gas_removal/msfr_gas_rem.inp_res.m')
#res2 = serpent.parse_res('/home/andrei2/Desktop/ornl/msfr/serpent/all_removal/msfr_gas_rem.inp_res.m')
#days = dep0['DAYS'] # Time array parsed from *_dep.m file
#n_se = dep0['NAMES'][0].split() # Names of isotopes parsed from *_dep.m file
#n_se1 = dep1['NAMES'][0].split() # Names of isotopes parsed from *_dep.m file
#n_se2 = dep2['NAMES'][0].split() # Names of isotopes parsed from *_dep.m file
#EOC = np.amax(days) # End of cycle (simulation time length)
#total_mass_list = dep0['TOT_MASS']
#adens_fuel_se = dep0['MAT_fuel_ADENS'] # atomic density for each isotope in material 'fuel'
#mdens_fuel_se = dep0['MAT_fuel_MDENS'] # mass density for each isotope in material 'fuel'
#mdens_fuel_se1 = dep1['MAT_fuel_MDENS'] # mass density for each isotope in material 'fuel'
#mdens_fuel_se2 = dep2['MAT_fuel_MDENS'] # mass density for each isotope in material 'fuel'
#vol_fuel_se = dep0['MAT_fuel_VOLUME'] # total volume of material 'fuel'
#adens_blank_se = dep0['MAT_blank_ADENS'] # atomic density for each isotope in material 'blank'
#mdens_blank_se = dep0['MAT_blank_MDENS'] # mass density for each isotope in material 'blank'
#mdens_blank_se1 = dep1['MAT_blank_MDENS'] # mass density for each isotope in material 'blank'
#mdens_blank_se2 = dep2['MAT_blank_MDENS'] # mass density for each isotope in material 'blank'
#vol_blank_se = dep0['MAT_blank_VOLUME'] # total volume of material 'blank'
# SCALE output
#filename_fuel = '/home/andrei2/Desktop/ornl/msfr/scale/no_reproc_depl/510efpd/quarter_cell2.000000000000000000.plt'
#filename_blank = '/home/andrei2/Desktop/ornl/msfr/scale/no_reproc_depl/510efpd/quarter_cell2.000000000000000001.plt'
#filename_fuel1 = '/home/andrei2/Desktop/ornl/msfr/scale/gas_removal/msfr_unit_vol_gases_removal.000000000000000000.plt'
#filename_blank1 = '/home/andrei2/Desktop/ornl/msfr/scale/gas_removal/msfr_unit_vol_gases_removal.000000000000000001.plt'
#filename_fuel2 = '/home/andrei2/Desktop/ornl/msfr/scale/gas_noble_removal/msfr_unit_vol_noble_removal.000000000000000000.plt'
#filename_blank2 = '/home/andrei2/Desktop/ornl/msfr/scale/gas_noble_removal/msfr_unit_vol_noble_removal.000000000000000001.plt'
#filename_fuel3 = '/home/andrei2/Desktop/ornl/msfr/scale/all_removal/msfr_unit_vol_noble_rare_removal.000000000000000000.plt'
#filename_blank3 = '/home/andrei2/Desktop/ornl/msfr/scale/all_removal/msfr_unit_vol_noble_rare_removal.000000000000000001.plt'
#filename_fuel4 = '/home/andrei2/Desktop/ornl/msfr/scale/th_feed_60yrs/msfr_all_removal_th_feed_60yrs.000000000000000000.plt'
#filename_blank4 = '/home/andrei2/Desktop/ornl/msfr/scale/th_feed_60yrs/msfr_all_removal_th_feed_60yrs.000000000000000001.plt'
filename_fuel4 = '/home/andrei2/Dropbox/study/internships/ornl/science/forAndrei/msfr6_withblank/msfr_all_removal_th_feed_60yrs_2.000000000000000000.plt'
filename_blank4 = '/home/andrei2/Dropbox/study/internships/ornl/science/forAndrei/msfr6_withblank/msfr_all_removal_th_feed_60yrs_2.000000000000000001.plt'
filename_u = '/home/andrei2/Dropbox/study/internships/ornl/science/forAndrei/msfr6_withblank/opus_print.000000000000000000.plt'
#k_file0 = '/home/andrei2/Desktop/ornl/msfr/scale/no_reproc_depl/510efpd/quarter_cell2.out'
#k_file1 = '/home/andrei2/Desktop/ornl/msfr/scale/gas_removal/msfr_unit_vol_gases_removal.out'
#k_file2 = '/home/andrei2/Desktop/ornl/msfr/scale/gas_noble_removal/msfr_unit_vol_noble_removal.out'
#k_file3 = '/home/andrei2/Desktop/ornl/msfr/scale/all_removal/msfr_unit_vol_noble_rare_removal.out'
k_file4 = '/home/andrei2/Dropbox/study/internships/ornl/science/forAndrei/msfr6_withblank/msfr_all_removal_th_feed_60yrs_2.out'
def read_scale_out (filename):
iso = []
adens = []
days_list = []
with open(filename,'r') as infile:
for line in itertools.islice(infile, 5, None): # Skip file header start=6, stop=None
p = line.split()
iso.append(str(p[0]))
adens.append(p[1:])
#iso.append(str(p[2]))
#adens.append(str(p[3]))
#u_en = 1e-6* np.flip (np.array (upp_enrg, dtype=float), 0 ) # Convert eV to MeV
#flux = np.flip ( np.array (flux_list, dtype=float), 0 )
#num_gr = len (u_en)
days_list.append (iso[0])
days_list = days_list + adens[0][:]
adens_arr = np.asarray(adens[1:][:], dtype=np.float32)
days = np.array (days_list, dtype=np.float32)
return iso[1:], days/365, adens_arr/1e+6
def read_scale_k (filename):
kinf = []
with open(filename) as openfile:
for line in openfile:
if line.startswith(' Infinite neutron multiplication'):
num = line.split(' ')[-1].strip()
kinf.append(float(num))
return kinf[1:]
#kinf0 = read_scale_k (k_file0)
#kinf1 = read_scale_k (k_file1)
#kinf2 = read_scale_k (k_file2)
#kinf3 = read_scale_k (k_file3)
kinf4 = read_scale_k (k_file4)
#n_sc, days_sc, mdens_fuel_sc = read_scale_out (filename_fuel)
#n_sc_blanket, days_sc, mdens_blank_sc = read_scale_out (filename_blank)
#n_sc1, days_sc1, mdens_fuel_sc1 = read_scale_out (filename_fuel1)
#n_sc_blanket1, days_sc1, mdens_blank_sc1 = read_scale_out (filename_blank1)
#n_sc2, days_sc2, mdens_fuel_sc2 = read_scale_out (filename_fuel2)
#n_sc_blanket2, days_sc2, mdens_blank_sc2 = read_scale_out (filename_blank2)
#n_sc3, days_sc3, mdens_fuel_sc3 = read_scale_out (filename_fuel3)
#n_sc_blanket3, days_sc3, mdens_blank_sc3 = read_scale_out (filename_blank3)
n_sc4, days_sc, mdens_fuel_sc4 = read_scale_out (filename_fuel4)
n_sc_blanket4, days_sc, mdens_blank_sc4 = read_scale_out (filename_blank4)
n_sc_u, days_sc_u, mdens_u = read_scale_out (filename_u)
#print (days_sc_u[:len(days_sc)])
#print (vol_blank_se[0]*mdens_u[n_sc_u.index(iso),(3*len(days_sc)):4*len(days_sc)])
tot_mass_sc = mdens_fuel_sc4[n_sc4.index(iso),]*vol_fuel_se[0]+mdens_blank_sc4[n_sc_blanket4.index(iso),]*vol_blank_se[0]+vol_storage[0]*mdens_u[n_sc_u.index(iso),]
#print (tot_mass_sc)
gain_rate = 1e-3*365*(tot_mass_sc[-1] - tot_mass_sc[0])/days_sc[-1]
# Initialize figure
fig_1 = matplotlib.pyplot.figure(1)
ax = fig_1.add_subplot(111)
ax.grid(True)
#ax.ticklabel_format (style='sci',scilimits=(0,0),axis='y')
#ax.set_ylim(0,0.00555)
#plot_title = 'Relative error in mass ' + str(100*abs(mdens_fuel_sc[n_sc.index(iso),-1]-mdens_fuel_se[n_se.index(iso.capitalize()),-1])/
# mdens_fuel_se[n_se.index(iso.capitalize()),-1] ) + ' %\n'
#for i in [n_se.index(iso.capitalize())]:
#ax.plot(days, mdens_fuel_se[i,:]*vol_fuel_se[0], '-',color='#ff8100', label=nucname.serpent(n_se[i])+' (no removals)')
#ax.plot(days, mdens_fuel_se[i,:]*vol_fuel_se[0]+mdens_blank_se[i,:]*vol_blank_se[0], '--',color='#ff8100',
# label=nucname.serpent(n_se[i])+'(w/ blanket, no removals)')
#ax.plot(days, mdens_blank_se[i,:]*vol_blank_se[0], '-',color='#ff8100', label=nucname.serpent(n_se[i])+'(Serpent, blanket)')
#for i in [n_se1.index(iso.capitalize())]:
# ax.plot(days, mdens_fuel_se1[i,:]*vol_fuel_se[0], '-',color='red',
# label=nucname.serpent(n_se1[i])+' Serpent(gases removal)')
#for i in [n_se2.index(iso.capitalize())]:
# ax.plot(days, mdens_fuel_se2[i,:]*vol_fuel_se[0], '--',color='green',
# label=nucname.serpent(n_se2[i])+' Serpent(volatile gases, noble metals, \nrare earths&discard)')
#for k in [iso]:
# ax.plot(days_sc, mdens_fuel_sc[n_sc.index(k)]*vol_fuel_se[0], '-',color='#ff8100', label=k+'(no removals)')
#for k in [iso]:
# ax.plot(days_sc, mdens_fuel_sc1[n_sc1.index(k)]*vol_fuel_se[0], '+--',color='blue', label=k+'(gases)')
#for k in [iso]:
# ax.plot(days_sc, mdens_fuel_sc2[n_sc2.index(iso)]*vol_fuel_se[0], '-',color='black', label=k+'(gases&noble)')
#for k in [iso]:
# ax.plot(days_sc, mdens_fuel_sc3[n_sc3.index(k)]*vol_fuel_se[0], '-',color='red', label=k+'(gases&noble\n&rare earths)')
for k in [iso]:
ax.plot(days_sc, mdens_fuel_sc4[n_sc4.index(k)]*vol_fuel_se[0], '-', label=k)
#for k in [iso]:
# ax.plot(days_sc, mdens_blank_sc4[n_sc_blanket4.index(k)]*vol_blank_se[0], '-', label='Blanket')
#for k in [iso]:
# ax.plot(days_sc, mdens_fuel_sc4[n_sc4.index(k)]*vol_fuel_se[0]+mdens_blank_sc4[n_sc_blanket4.index(k)]*vol_blank_se[0],
# '-', label='Total')
ax.legend(loc=0)
ax.set_ylabel('Mass [t]')
ax.set_xlabel('EFPY')
#ax.set_title('Mass balance in fuel salt for ' + str (iso)+'\n')
#ax.text(0.95, 0.5, 'Breeding gain '+"{0:.1f}".format(gain_rate)+ ' kg/year',
# horizontalalignment='right',verticalalignment='center', transform=ax.transAxes)
#ax.text(0.95, 0.45, 'Breeding gain coeff '+"{0:.4f}".format((tot_mass_sc[-1] - tot_mass_sc[0])/(tot_mass_sc[0])),
# horizontalalignment='right',verticalalignment='center', transform=ax.transAxes)
ax.set_xlim([0,np.amax(days_sc)])
#ax.set_ylim([0, np.amax(tot_mass_sc)])
fig_1.show()
fig_1.savefig(str(iso)+'_th_feed.png',bbox_inches='tight', dpi=900)
# Initialize figure
fig_2 = matplotlib.pyplot.figure(2)
ax = fig_2.add_subplot(111)
ax.grid(True)
#ax.ticklabel_format (style='sci',scilimits=(0,0),axis='y')
#for i in [n_se.index(iso.capitalize())]:
# ax.plot(days, mdens_blank_se[i,:]*vol_blank_se[0], '-',color='#ff8100', label=nucname.serpent(n_se[i])+' (no removals)')
#for i in [n_se1.index(iso.capitalize())]:
# ax.plot(days, mdens_blank_se1[i,:]*vol_blank_se[0], '-',color='red', label=nucname.serpent(n_se1[i])+' Serpent(gases removal)')
#for i in [n_se2.index(iso.capitalize())]:
# ax.plot(days, mdens_blank_se2[i,:]*vol_blank_se[0], '--',color='blue', label=nucname.serpent(n_se2[i])+' Serpent(volatile gases, noble metals, \nrare earths&discard)')
#for k in [iso]:
# ax.plot(days_sc, mdens_blank_sc[n_sc_blanket.index(k)]*vol_blank_se[0], '-',color='#ff8100', label=k+'(no removals)')
#for k in [iso]:
# ax.plot(days_sc, mdens_blank_sc1[n_sc_blanket1.index(k)]*vol_blank_se[0], '+--',color='blue', label=k+'(gases)')
#for k in [iso]:
# ax.plot(days_sc, mdens_blank_sc2[n_sc_blanket2.index(k)]*vol_blank_se[0], '-',color='black', label=k+'(gases&noble)')
#for k in [iso]:
# ax.plot(days_sc, mdens_blank_sc3[n_sc_blanket3.index(k)]*vol_blank_se[0], '-',color='red', label=k+'(gases&noble\n&rare earths)')
for k in [iso]:
ax.plot(days_sc, mdens_blank_sc4[n_sc_blanket4.index(k)]*vol_blank_se[0], '-',color='green', label=k+'(w/ Th-232 feed)')
ax.legend(loc=0)
ax.set_ylabel('Mass [t]')
ax.set_xlabel('EFPY')
ax.set_title('Mass balance in fertile salt for ' + str (iso))
ax.set_xlim([0,np.amax(days_sc)])
fig_2.show()
#fig_2.savefig(str(iso)+'_blanket_chemtr_rem.png',bbox_inches='tight', dpi=700)
# Initialize figure
fig_3 = matplotlib.pyplot.figure(3)
ax = fig_3.add_subplot(111)
ax.grid(True)
#ax.ticklabel_format (style='sci',scilimits=(0,0),axis='y')
#for k in [iso]:
# ax.plot(days_sc, mdens_fuel_sc[n_sc.index(k)]*vol_fuel_se[0]+mdens_blank_sc[n_sc_blanket.index(k)]*vol_blank_se[0], '-',
# color='#ff8100', label=k+'(no removals)')
#for k in [iso]:
# ax.plot(days_sc, mdens_fuel_sc1[n_sc1.index(k)]*vol_fuel_se[0]+mdens_blank_sc1[n_sc_blanket1.index(k)]*vol_blank_se[0], '+--',
# color='blue', label=k+'(gases)')
#for k in [iso]:
# ax.plot(days_sc, mdens_fuel_sc2[n_sc2.index(k)]*vol_fuel_se[0]+mdens_blank_sc2[n_sc_blanket2.index(k)]*vol_blank_se[0], '-',
# color='black', label=k+'(gases&noble)')
#for k in [iso]:
# ax.plot(days_sc, mdens_fuel_sc3[n_sc3.index(k)]*vol_fuel_se[0]+mdens_blank_sc3[n_sc_blanket3.index(k)]*vol_blank_se[0], '-',
# color='red', label=k+'(gases&noble\n&rare earths)')
for k in [iso]:
ax.plot(days_sc, vol_storage[0]*mdens_u[n_sc_u.index(iso)], '-',
color='green', label=k)
ax.legend(loc=0)
ax.set_ylabel('Mass [t]')
ax.set_xlabel('EFPY')
ax.set_title('U storage mass balance for ' + str (iso))
#ax.text(0.95, 0.5, 'Breeding gain '+"{0:.1f}".format(gain_rate)+ ' kg/year',
# horizontalalignment='right',verticalalignment='center', transform=ax.transAxes)
#ax.text(0.95, 0.45, 'Breeding gain coeff '+"{0:.4f}".format((tot_mass_sc[-1] - tot_mass_sc[0])/(tot_mass_sc[0])),
# horizontalalignment='right',verticalalignment='center', transform=ax.transAxes)
ax.set_xlim([0,np.amax(days_sc)])
fig_3.show()
#fig_3.savefig(str(iso)+'_total_chemtr_rem_n_feed.png',bbox_inches='tight', dpi=700)
# Initialize figure
fig_4 = matplotlib.pyplot.figure(4)
ax = fig_4.add_subplot(111)
ax.grid(True)
#ax.plot(days_sc, kinf0, '-',color='#ff8100', label='no removals')
#ax.plot(days_sc, kinf1, '-',color='blue', label='gases')
#ax.plot(days_sc, kinf2, '-',color='black', label='gases&noble')
#ax.plot(days_sc, kinf3, '-',color='red', label='gases&noble\n&rare earths')
ax.plot(days_sc, kinf4, '-',color='green', label='K$_{inf}$')
ax.legend(loc=0)
ax.set_ylabel('Infinite multiplication factor (k$_{\inf)}$)')
ax.set_xlabel('EFPY')
ax.set_title('Infinite muliplication factor')
ax.set_xlim([0,np.amax(days_sc)])
fig_4.show()
#fig_4.savefig('k_inf_Th_feed.png',bbox_inches='tight', dpi=700)
'''
print ("Correlation between mass of target isotope (Serpent-Unit vs SCALE-Unit) is "
+ str(pearsonr(mdens_fuel_sc[n_sc.index(iso)], mdens_fuel_se[n_se.index(iso.capitalize())])) )
print ('Relative error for fuel salt in target isotope mass after ' + str(days[-1]) + ' days: ' +
str(100*abs(mdens_fuel_sc[n_sc.index(iso),-1]-mdens_fuel_se[n_se.index(iso.capitalize()),-1])/
mdens_fuel_se[n_se.index(iso.capitalize()),-1] ) + ' %')
print ('Relative error in total target isotope mass after ' + str(days[-1]) + ' days: ' +
str (100*abs(tot_mass_se[-1]-tot_mass_sc[-1]) / tot_mass_se[-1]) + ' %' )
print ('Relative error in blanket in total target isotope mass after ' + str(days[-1]) + ' days: ' +
str (100*abs(mdens_blank_sc[n_sc_blanket.index(iso),-1]-mdens_blank_se[n_se.index(iso.capitalize()),-1]) / mdens_blank_se[n_se.index(iso.capitalize()),-1]) + ' %' )
'''
print ('\nFrom SCALE')
print ('Breeding gain ' + str (1e+3*(tot_mass_sc[-1] - tot_mass_sc[0])/days_sc[-1]) + ' kg/year' )
print ('Breeding gain coefficient ' + str ((tot_mass_sc[-1] - tot_mass_sc[0])/(tot_mass_sc[0] * days_sc[-1])) )
print ('Breeding gain in blanket ' + str (1e+3*(mdens_blank_sc4[n_sc_blanket4.index(iso),-1]*vol_blank_se[0] - mdens_blank_sc4[n_sc_blanket4.index(iso),0]*vol_blank_se[0])/days_sc[-1]) + ' kg/year' )
print ('\nDoubling time (net) ' + str( 2*tot_mass_sc[0]/ ((tot_mass_sc[-1] - tot_mass_sc[0])/days_sc[-1] )) )
print (tot_mass_sc[0])
# -
|
msfr/plots/MSFR_reprocessing_depletion_SCALE.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: py35-paddle1.2.0
# ---
# # 使用卷积神经网络进行图像分类
#
# **作者:** [PaddlePaddle](https://github.com/PaddlePaddle) <br>
# **日期:** 2022.5 <br>
# **摘要:** 本示例教程将会演示如何使用飞桨的卷积神经网络来完成图像分类任务。这是一个较为简单的示例,将会使用一个由三个卷积层组成的网络完成[cifar10](https://www.cs.toronto.edu/~kriz/cifar.html)数据集的图像分类任务。
# ## 一、环境配置
#
# 本教程基于PaddlePaddle 2.3.0 编写,如果你的环境不是本版本,请先参考官网[安装](https://www.paddlepaddle.org.cn/install/quick) PaddlePaddle 2.3.0 。
# +
import paddle
import paddle.nn.functional as F
from paddle.vision.transforms import ToTensor
import numpy as np
import matplotlib.pyplot as plt
print(paddle.__version__)
# -
# ## 二、加载数据集
#
# 本案例将会使用飞桨提供的API完成数据集的下载并为后续的训练任务准备好数据迭代器。cifar10数据集由60000张大小为32 * 32的彩色图片组成,其中有50000张图片组成了训练集,另外10000张图片组成了测试集。这些图片分为10个类别,将训练一个模型能够把图片进行正确的分类。
transform = ToTensor()
cifar10_train = paddle.vision.datasets.Cifar10(mode='train',
transform=transform)
cifar10_test = paddle.vision.datasets.Cifar10(mode='test',
transform=transform)
# ## 三、组建网络
# 接下来使用飞桨定义一个使用了三个二维卷积( ``Conv2D`` ) 且每次卷积之后使用 ``relu`` 激活函数,两个二维池化层( ``MaxPool2D`` ),和两个线性变换层组成的分类网络,来把一个(32, 32, 3)形状的图片通过卷积神经网络映射为10个输出,这对应着10个分类的类别。
class MyNet(paddle.nn.Layer):
def __init__(self, num_classes=1):
super(MyNet, self).__init__()
self.conv1 = paddle.nn.Conv2D(in_channels=3, out_channels=32, kernel_size=(3, 3))
self.pool1 = paddle.nn.MaxPool2D(kernel_size=2, stride=2)
self.conv2 = paddle.nn.Conv2D(in_channels=32, out_channels=64, kernel_size=(3,3))
self.pool2 = paddle.nn.MaxPool2D(kernel_size=2, stride=2)
self.conv3 = paddle.nn.Conv2D(in_channels=64, out_channels=64, kernel_size=(3,3))
self.flatten = paddle.nn.Flatten()
self.linear1 = paddle.nn.Linear(in_features=1024, out_features=64)
self.linear2 = paddle.nn.Linear(in_features=64, out_features=num_classes)
def forward(self, x):
x = self.conv1(x)
x = F.relu(x)
x = self.pool1(x)
x = self.conv2(x)
x = F.relu(x)
x = self.pool2(x)
x = self.conv3(x)
x = F.relu(x)
x = self.flatten(x)
x = self.linear1(x)
x = F.relu(x)
x = self.linear2(x)
return x
# ## 四、模型训练&预测
#
# 接下来,用一个循环来进行模型的训练,将会: <br>
# - 使用 ``paddle.optimizer.Adam`` 优化器来进行优化。
# - 使用 ``F.cross_entropy`` 来计算损失值。
# - 使用 ``paddle.io.DataLoader`` 来加载数据并组建batch。
epoch_num = 10
batch_size = 32
learning_rate = 0.001
# +
val_acc_history = []
val_loss_history = []
def train(model):
print('start training ... ')
# turn into training mode
model.train()
opt = paddle.optimizer.Adam(learning_rate=learning_rate,
parameters=model.parameters())
train_loader = paddle.io.DataLoader(cifar10_train,
shuffle=True,
batch_size=batch_size)
valid_loader = paddle.io.DataLoader(cifar10_test, batch_size=batch_size)
for epoch in range(epoch_num):
for batch_id, data in enumerate(train_loader()):
x_data = data[0]
y_data = paddle.to_tensor(data[1])
y_data = paddle.unsqueeze(y_data, 1)
logits = model(x_data)
loss = F.cross_entropy(logits, y_data)
if batch_id % 1000 == 0:
print("epoch: {}, batch_id: {}, loss is: {}".format(epoch, batch_id, loss.numpy()))
loss.backward()
opt.step()
opt.clear_grad()
# evaluate model after one epoch
model.eval()
accuracies = []
losses = []
for batch_id, data in enumerate(valid_loader()):
x_data = data[0]
y_data = paddle.to_tensor(data[1])
y_data = paddle.unsqueeze(y_data, 1)
logits = model(x_data)
loss = F.cross_entropy(logits, y_data)
acc = paddle.metric.accuracy(logits, y_data)
accuracies.append(acc.numpy())
losses.append(loss.numpy())
avg_acc, avg_loss = np.mean(accuracies), np.mean(losses)
print("[validation] accuracy/loss: {}/{}".format(avg_acc, avg_loss))
val_acc_history.append(avg_acc)
val_loss_history.append(avg_loss)
model.train()
model = MyNet(num_classes=10)
train(model)
# +
plt.plot(val_acc_history, label = 'validation accuracy')
plt.xlabel('Epoch')
plt.ylabel('Accuracy')
plt.ylim([0.5, 0.8])
plt.legend(loc='lower right')
# -
# ## The End
# 从上面的示例可以看到,在cifar10数据集上,使用简单的卷积神经网络,用飞桨可以达到70%以上的准确率。你也可以通过调整网络结构和参数,达到更好的效果。
|
docs/practices/cv/convnet_image_classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Facets
import pandas as pd
import altair as alt
import os
os.getcwd()
churn = pd.read_csv("../data/churn.csv")
alt.data_transformers.enable('json')
# ## Facets Examples
alt.Chart(churn).mark_bar().encode(
alt.X("Tenure:Q", bin=alt.BinParams(maxbins=20)),
alt.Y("count()"),
alt.Column("Churn"),
alt.Color("Churn")
).properties(
width=250,
height=250,
title="Facet Example"
)
alt.Chart(churn).mark_bar().encode(
alt.X("Tenure:Q", bin=alt.BinParams(maxbins=20)),
alt.Y("count()"),
alt.Column("PaymentMethod"),
).properties(
width=250,
height=250
)
alt.Chart(churn).mark_bar().encode(
alt.X("Tenure:Q", bin=alt.BinParams(maxbins=20)),
alt.Y("count()"),
alt.Column("Churn"),
alt.Color("Churn")
).properties(
width=250,
height=250,
title="Facet Example"
).resolve_scale(color="independent")
base = alt.Chart(churn).mark_bar().encode(
alt.X("Tenure:Q", bin=alt.BinParams(maxbins=20)),
alt.Y("count()")
).properties(
width=250,
height=250
)
base
base.facet(
column="PaymentMethod"
)
churn['TotalCharges'] = pd.to_numeric(churn.TotalCharges.str.replace(" ",""))
# ### Repeat
alt.Chart(churn).mark_bar().encode(
alt.X("Tenure:Q", bin=alt.BinParams(maxbins=20)),
alt.Y(alt.repeat("column"), aggregate='average',type="quantitative"),
color="Churn"
).properties(
width=180,
height=180
).repeat(
column=["MonthlyCharges", "TotalCharges"]
)
# ### Layers
alt.layer(
alt.Chart(churn).mark_bar().encode(
alt.X("PaymentMethod"),
alt.Y(alt.repeat('column'), aggregate="average", type="quantitative")
),
alt.Chart(churn).mark_rule(color="black").encode(
alt.Y(alt.repeat('column'), aggregate="average", type="quantitative")
)
).properties(
width=180,
height=180
).repeat(
column = ['MonthlyCharges', "TotalCharges"]
)
|
notebooks/03-Facets.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import pandas as pd
import numpy as np
# +
mydata = pd.read_csv('features.csv', delimiter=';', usecols=['is_featured', 'version', 'tags_number', 'score',
'is_best_time_to_launch', 'is_best_day_to_launch', 'is_weekend',
'discretized_positive_description_score',
'discretized_negative_description_score',
'text_description_length', 'sentence_length_in_the_description',
'bullet_points_explicit_features', 'emoji_in_description',
'tagline_length', 'emoji_in_tagline', 'are_there_video',
'are_there_tweetable_images', 'are_there_gif_images',
'number_of_gif', 'offers', 'promo_discount_codes',
'are_there_questions', 'hunter_has_twitter', 'hunter_has_website',
'hunter_followers', 'hunter_apps_made',
'hunter_follows_up_on_comments', 'hunter_is_maker',
'maker_has_twitter', 'maker_has_website', 'maker_followers',
'maker_follows_up_on_comments',
'discretized_maker_positive_comment_score',
'discretized_maker_negative_comment_score',
'discretized_others_positive_comment_score',
'discretized_others_negative_comment_score',
'topic'])
mydata = mydata.rename(columns = {'discretized_positive_description_score': 'positive_description_sentiment',
'discretized_negative_description_score': 'negative_description_sentiment',
'discretized_maker_positive_comment_score': 'maker_positive_comment',
'discretized_maker_negative_comment_score': 'maker_negative_comment',
'discretized_others_positive_comment_score': 'others_positive_comment',
'discretized_others_negative_comment_score': 'others_negative_comment'})
pd.set_option('display.max_columns', 38)
mydata.head()
# -
# # Standardizzazione dati dataset
# +
from sklearn.preprocessing import StandardScaler
# Create the Scaler object
scaler = StandardScaler()
# Fit your data on the scaler object
# mydata = scaler.fit_transform(mydata)
# mydata['version'] = scaler.fit_transform(mydata['version'].values.reshape(-1,1))
mydata[['version', 'tags_number', 'score', 'number_of_gif']] = scaler.fit_transform(mydata[['version', 'tags_number', 'score',
'number_of_gif']])
# -
mydata.to_csv('standardized_features.csv', sep=';', index=False)
mydata.head()
# # Matrice di correlazione tra le feature numeriche nel dataset
import seaborn as sn
import matplotlib.pyplot as plt
# %matplotlib inline
numerical_data = pd.read_csv('standardized_features.csv', delimiter=';',
usecols=['version', 'tags_number', 'score', 'number_of_gif'])
correlation_matrix = numerical_data.corr(method='pearson')
correlation_matrix
plt.figure(figsize=(10,7))
sn.heatmap(correlation_matrix, annot=True)
plt.savefig('correlation_matrix_of_features_initial_model.png')
# # Impostazione delle variabili di default per la regressione logistica
# Per impostare le variabili di default durante l'esecuzione della regressione logistica sono state create delle variabili `dummy`. In particolare:
# - per le variabili booleane è stato impostato come default il valore *No*
# - per la lunghezza della descrizione, delle frasi presenti nella descrizione e della tagline di ogni post è stato impostato come default il valore *Short*
# - per il numero di follower degli hunter, per il numero di follower dei maker e per il numero di applicazioni/prodotti fatti dall'hunter è stato impostato come default il valore *High*
# - per la variabile topic è stato impostato come default il valore *web development*
# +
mydata = pd.get_dummies(mydata, columns=['is_best_time_to_launch', 'is_best_day_to_launch', 'is_weekend',
'positive_description_sentiment', 'negative_description_sentiment',
'bullet_points_explicit_features', 'emoji_in_description', 'emoji_in_tagline',
'are_there_video', 'are_there_tweetable_images', 'are_there_gif_images', 'offers',
'promo_discount_codes', 'are_there_questions', 'hunter_has_twitter',
'hunter_has_website', 'hunter_follows_up_on_comments', 'hunter_is_maker',
'maker_has_twitter', 'maker_has_website', 'maker_follows_up_on_comments',
'maker_positive_comment', 'maker_negative_comment', 'others_positive_comment',
'others_negative_comment'], drop_first=True)
mydata = mydata.rename(columns = {'positive_description_sentiment_True': 'positive_description_sentiment',
'negative_description_sentiment_True': 'negative_description_sentiment',
'maker_positive_comment_True': 'maker_positive_comment',
'maker_negative_comment_True': 'maker_negative_comment',
'others_positive_comment_True': 'others_positive_comment',
'others_negative_comment_True': 'others_negative_comment'})
# +
mydata = pd.get_dummies(mydata, columns=['text_description_length', 'sentence_length_in_the_description', 'tagline_length',
'hunter_followers', 'hunter_apps_made', 'maker_followers'])
mydata = mydata.drop(['text_description_length_Short', 'sentence_length_in_the_description_Short', 'tagline_length_Short',
'hunter_followers_High', 'hunter_apps_made_High', 'maker_followers_High'], axis=1)
# -
mydata = pd.get_dummies(mydata, columns = ['topic'])
mydata = mydata.drop(['topic_web development'], axis=1)
pd.set_option('display.max_columns', 46)
mydata.head()
# # Implementazione del modello per la regressione logistica
# Per realizzare la regressione logistica è stato richiamato il metodo **from_formula** fornito dalla classe **GLM() (Generalized Linear Models)** presente nel package `statsmodels`. Questo metodo ha come parametri di input:
# * **formula** è una stringa rappresentante una formula che separa la variabile dipendente dalle variabili indipendenti
# * **data** rappresenta il dataset necessario per costruire il modello
# * **family** indica la distribuzione della variabile dipendente. In questo caso è Binomial che prende come parametro di default il link *logit* in quanto dobbiamo realizzare la regressione logistica.
#
# Successivamente è stato costruito il modello richiamando la funzione `fit`.
import statsmodels.api as sm
myformula = 'is_featured ~ version + tags_number + score + is_best_time_to_launch_Yes + is_best_day_to_launch_Yes + is_weekend_Yes + positive_description_sentiment + negative_description_sentiment + text_description_length_Medium + text_description_length_Long + sentence_length_in_the_description_Medium + sentence_length_in_the_description_Long + bullet_points_explicit_features_Yes + emoji_in_description_Yes + tagline_length_Medium + tagline_length_Long + emoji_in_tagline_Yes + are_there_video_Yes + are_there_tweetable_images_Yes + are_there_gif_images_Yes + number_of_gif + offers_Yes + promo_discount_codes_Yes + are_there_questions_Yes + hunter_has_twitter_Yes + hunter_has_website_Yes + hunter_followers_Low + hunter_followers_Medium + hunter_apps_made_Low + hunter_apps_made_Medium + hunter_follows_up_on_comments_Yes + hunter_is_maker_Yes + maker_has_twitter_Yes + maker_has_website_Yes + maker_followers_Low + maker_followers_Medium + maker_follows_up_on_comments_Yes + maker_positive_comment + maker_negative_comment + others_positive_comment + others_negative_comment + topic_community + topic_creativity'
model = sm.GLM.from_formula(formula=myformula, data=mydata, family=sm.families.Binomial())
results = model.fit()
# Richiamando la funzione `summary` è possibile osservare i risultati del modello costruito.
print(results.summary().tables[0])
print("Likelihood of specified model: {}".format(round(results.llf , 2)))
print("Likelihood of Intercept only model: {}".format(round(results.llnull, 2)))
print("Number of observations: {}".format(model.nobs))
# ### Goodness of fit (Nagelkerke' R-squared) del modello
# +
import math
power = 2 / model.nobs
cox_and_snell_r_squared = 1 - math.pow((results.llnull / results.llf), power)
denominator = 1 - math.pow((-results.llnull), power)
nagelkerke_r_squared = cox_and_snell_r_squared / denominator
print("Nagelkerke' r squared: {}".format(round(nagelkerke_r_squared, 3)))
# -
# ### Risultati del modello
# Note that the summary table is a list. The table at index 1 is the "core" table.
# Additionally, read_html puts dataframe in a list, so we want index 0
results_summary = results.summary()
results_as_html = results_summary.tables[1].as_html()
logistic = pd.read_html(results_as_html, header=0, index_col=0)[0]
# Convert html table into csv file
logistic.to_csv('logistic_regression_results.csv', sep=';', columns=logistic.columns)
temp = pd.read_csv('logistic_regression_results.csv', delimiter=';')
temp.rename(columns={temp.columns[0]: "Predictor"}, inplace = True)
temp.to_csv('logistic_regression_results.csv', sep=';', index=False)
# Successivamente è stato calcolato il valore di **Odds Ratio** per ogni coefficiente utilizzando la funzione *exp** fornita dal package `numpy` e che prende come parametro di input il coefficiente.
# +
# Calculate Odds Ratio
def calculate_odds_ratio(coefficient_column):
odds_ratio = []
for i in range(0, len(coefficient_column)):
if not odds_ratio:
odds_ratio.append('-')
else:
odds_ratio.append(round(np.exp(coefficient_column[i]), 3))
return odds_ratio
df = pd.read_csv('logistic_regression_results.csv', delimiter=';')
odds_ratio = calculate_odds_ratio(df['coef'])
print(odds_ratio)
# -
# Add Odds Ratio column to logistic regression summary table
df.insert(2, 'Odds Ratio', odds_ratio)
df.to_csv('logistic_regression_results.csv', sep=';', index=False)
# +
def make_bold(s):
if s['P>|z|'] < 0.05:
return ['font-weight: bold']*8
else:
return ['font-weight: normal']*8
logistic_regression_results = pd.read_csv('logistic_regression_results.csv', delimiter=';')
logistic_regression_results.style.apply(make_bold, axis=1)
# -
|
analysis/notebook/initial_model_logistic_regression/logistic_regression_notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Prediction of time required for quality inspection of mercedes cars
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
df=pd.read_csv("G:/DATA/Python/Mercedes/train.csv")
df.head()
df.shape
# for displaying all columns
pd.set_option("display.max_columns",380)
df.info()
# dropping ID as it is an unique identifier
df.drop(["ID"],inplace=True,axis=1)
df.head()
df.y.max()
# checking for correlation which is really tough since we have so many variables
df.corr()
# since there were few categorical variables we are converting them into numeric by using label encoder
from sklearn.preprocessing import LabelEncoder
le=LabelEncoder()
df[df.select_dtypes(include="object").columns] =df[df.select_dtypes(include="object").columns].apply(le.fit_transform)
# sampling
df_x= df.iloc[:,1:]
df_y=df.iloc[:,0]
from sklearn.model_selection import train_test_split
df_x_train, df_x_test, df_y_train, df_y_test= train_test_split(df_x,df_y, test_size=0.2)
# # using Linear Regression
from sklearn.linear_model import LinearRegression
reg=LinearRegression()
reg.fit(df_x_train,df_y_train)
reg.coef_
# +
# now we check R2 and adjR2
Rsquare= reg.score(df_x_train,df_y_train)
print("values of Rsquare is ",Rsquare)
N=df_x_train.shape[0]
K=df_x_train.shape[1]
Adjusted_Rsquare= 1-(1-Rsquare)*(N-1)/(N-K-1)
print("value of Adjusted_Rsquare is ", Adjusted_Rsquare)
# -
pred_linear= reg.predict(df_x_test)
pred_linear.max()
error_linear= df_y_test-pred_linear
from sklearn.metrics import mean_squared_error
Rmse= np.sqrt(mean_squared_error(df_y_test,pred_linear))
Rmse
#MAPE
# Here we can see that the mape value is very high also the RMSE value is really high
MAPE=np.mean(np.abs(error_linear/df_y_test)*100)
MAPE
# # Using Randomforest
from sklearn.ensemble import RandomForestRegressor
rf=RandomForestRegressor(n_estimators=300,max_depth=7)
rf.fit(df_x_train, df_y_train)
# +
R2=rf.score(df_x_train,df_y_train)
print("Rsquare is ",R2)
N=df_x_train.shape[0]
K=df_x_train.shape[1]
Adjusted_Rsquare= 1-(1-R2)*(N-1)/(N-K-1)
print("value of Adjusted_Rsquare is ", Adjusted_Rsquare)
# -
pred_rf=rf.predict(df_x_test)
error_rf=df_y_test-pred_rf
Rmse= np.sqrt(mean_squared_error(df_y_test,pred_rf))
Rmse
# Here we can see the mape value
MAPE=np.mean(np.abs(error_rf/df_y_test)*100)
MAPE
# accuracy using randomforest
accuracy = 100- MAPE
accuracy
# +
# XGBOOST REGRESSOR
# -
from xgboost import XGBRegressor
xg=XGBRegressor(booster="gbtree",learning_rate=0.15, max_depth=7,n_estimators=200)
xg.fit(df_x_train,df_y_train)
xg.feature_importances_
# +
r2=xg.score(df_x_train,df_y_train)
print("Rsquare is ",r2)
N=df_x_train.shape[0]
K=df_x_train.shape[1]
Adjusted_Rsquare= 1-(1-r2)*(N-1)/(N-K-1)
print("value of Adjusted_Rsquare is ", Adjusted_Rsquare)
# -
pred_xg= xg.predict(df_x_test)
pred_xg
error_xg=df_y_test-pred_xg
Rmse= np.sqrt(mean_squared_error(df_y_test,pred_xg))
Rmse
MAPE=np.mean(np.abs(error_xg/df_y_test)*100)
MAPE
accuracy= 100 - MAPE
accuracy
# the r2 adjusted R2 and the final accuracy is good for xgboost regressor
# +
# now using test data file that was seperatly given
# -
test=pd.read_csv("G:/DATA/Python/Mercedes/test.csv")
test2=pd.read_csv("G:/DATA/Python/Mercedes/test.csv")
test.head()
test.drop(["ID"],axis=1,inplace=True)
test[test.select_dtypes(include="object").columns] =test[test.select_dtypes(include="object").columns].apply(le.fit_transform)
pred_test= xg.predict(test)
test_solution=pd.DataFrame({"ID":test2.ID,"y":pred_test})
test_solution
# # PCA
# +
# PCA was applied just to check how dimensionality reduction affect the accuracy of model by using xgboost regressor
# -
from sklearn import decomposition
from sklearn.decomposition import PCA
pca=decomposition.PCA()
pca_dfx=pca.fit(df_x)
# variation explained by principal components
variation=pca.explained_variance_ratio_.cumsum() * 100
variation
# +
plt.plot(variation, marker=".")
# +
# Now we select the top 28 vaiarbles which have maximum variance retained from above graph
# -
x_pca= pca.transform(df_x)
# Now we create new x values using the selected principal components
x_pca=pd.DataFrame(x_pca)
x_pca
x_pca=x_pca.iloc[:,0:28]
x_pca
# now we do train test split
x_pca_train, x_pca_test, y_pca_train, y_pca_test= train_test_split(x_pca,df_y, test_size=0.2)
xg.fit(x_pca_train,y_pca_train)
pred_pca=xg.predict(x_pca_test)
error_pca=y_pca_test-pred_pca
Rmse= np.sqrt(mean_squared_error(y_pca_test,pred_pca))
Rmse
MAPE=np.mean(np.abs(error_pca/y_pca_test)*100)
MAPE
# +
# here we can clearly see that pca is actually not affecting the final accuracy to that extent in case of xgboost regressor
# +
# Now applying pca on the given test file
# -
test_pca_x=pca.fit(test)
test_pca=pca.transform(test)
test_pca= pd.DataFrame(test_pca)
test_pca
test_pca=test_pca.iloc[:,0:28]
pred_test_pca= xg.predict(test_pca)
test_solutionpca=pd.DataFrame({"ID":test2.ID,"y":pred_test_pca})
test_solutionpca
|
mercedes data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
crime=pd.read_csv('samplecrime.csv')
crime
type(crime)
crime.dtypes #to find datatypes of the crime dataset variables; if python doesn't recognise datatype it gives object as output
crime.head(10) #top 5 rows to be displayed: paramter if passed will give that many rows
crime['Category']
crime1=crime[['Category','DayOfWeek','PdDistrict']]
crime1
count_category=crime['Category'].value_counts() # value_counts() gives the count of offences of each category
count_category
type(count_category)
# converting Series into a 2D data structure
df=pd.DataFrame(count_category)
df
# the leftmost column should be a proper index, so we need to do the following:
DF1=df.reset_index()
DF1
# renaming the column names of the dataframe
DF1.columns=['Category','Frequency']
DF1
Sample=DF1.head();
Sample
# +
# create a bar chart: Data visualisation
plt.bar(Sample['Category'],Sample['Frequency'],color="Blue")
plt.xlabel("Cateory")
plt.ylabel("Frequency")
plt.xticks(rotation=45)
plt.title("The frequency of Category of Crime")
plt.show()
# -
|
CrimeCategoryFrequency.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import rosbag
import pymap3d as pm
from scipy.signal import savgol_filter
# %matplotlib inline
# +
def wrap_angle(angle):
return (angle + np.pi) % (2 * np.pi) - np.pi
def to_euler(x, y, z, w):
"""Return as xyz (roll pitch yaw) Euler angles."""
roll = np.arctan2(2 * (w * x + y * z), 1 - 2 * (x**2 + y**2))
pitch = np.arcsin(2 * (w * y - z * x))
yaw = np.arctan2(2 * (w * z + x * y), 1 - 2 * (y**2 + z**2))
return np.array([roll, pitch, yaw])
# -
# ls
bag = rosbag.Bag('waypoint_u turn_kiri_1.bag')
rpy = []
imu_t = []
for topic, msg, _ in bag.read_messages(topics=['/imu']):
qt = msg.orientation
rpy.append(to_euler(qt.x, qt.y, qt.z, qt.w))
imu_t.append(msg.header.stamp.to_sec())
imu_t = np.array(imu_t)
rpy = np.array(rpy)
# +
lat0, lon0, h0 = -6.8712, 107.5738, 768
gps_t = []
gps_pos = []
gps_cov = []
for topic, msg, _ in bag.read_messages(topics=['/fix']):
gps_t.append(msg.header.stamp.to_sec())
gps_pos.append(pm.geodetic2enu(msg.latitude, msg.longitude, msg.altitude, lat0, lon0, h0))
gps_cov.append(msg.position_covariance)
gps_t = np.array(gps_t)
gps_pos = np.array(gps_pos)
gps_cov = np.array(gps_cov).reshape(-1,3,3)
# Ambil yang bagus-bagus aja
gnss_pos = np.copy(gps_pos[:-1])
gnss_t = np.copy(gps_t[:-1])
gnss_cov = np.copy(gps_cov[:-1])
# -
gps_pos = (-1) * np.copy(gnss_pos[:,:2])
gps_t = np.copy(gnss_t)
# # PAKAI CLASS KF_gps
from kf_gps import KF_gps
# +
f = 100 #Hz
dt = 1/f #s
Tf = int(gps_t[-1] - gps_t[0] + 0.5)
var_gps_pos = 0.5 **2
var_gps_speed = 0.25 **2
var_gps_yaw = 0.1 **2
var_gps_w = 0.01 **2
Q = np.eye(8)
Q[:2,:2] = np.eye(2) * 3.**2
Q[2:4,2:4] = np.eye(2) * 3.**2
Q[4:6,4:6] = np.eye(2) * 0.1**2
Q[6,6] = 1.**2
Q[7,7] = 0.1**2
t = np.array([i*dt for i in range(Tf*f)]) + gps_t[0]
# +
x = np.zeros((f*Tf, 2))
x[0] = gps_pos[0]
v = np.zeros(x.shape)
v[0] = np.array([.0, 0.]) # Tebakan awal
a = np.zeros_like(v) # Tebakan awal
yaw = np.zeros(x.shape[0])
yaw[0] = -0.0
w = np.zeros_like(yaw)
w[0] = 0.05
P = np.zeros((x.shape[0], 8, 8))
P[0, 2:, 2:] = np.eye(6) * 1.
temp_pos_yaw = np.copy(gps_pos[0])
kf = KF_gps(var_gps_pos, var_gps_speed, var_gps_yaw, var_gps_w,
Q, x[0], v[0], a[0], yaw[0], w[0], P[0])
gps_idx = 1
# -
for i in range(1, x.shape[0]):
dt = t[i] - t[i-1]
x[i], v[i], a[i], yaw[i], w[i], P[i] = kf.predict(dt)
if (gps_idx != gps_t.shape[0]) and (gps_t[gps_idx] < t[i]):
dt_gps = gps_t[gps_idx] - gps_t[gps_idx - 1]
# Correct Position
x[i], v[i], a[i], yaw[i], w[i], P[i] = kf.correct_position(gps_pos[gps_idx])
# Correct Velocity
gps_vel = (gps_pos[gps_idx] - gps_pos[gps_idx-1]) / dt_gps
x[i], v[i], a[i], yaw[i], w[i], P[i] = kf.correct_velocity(gps_vel)
# Correct Yaw dan Omega
if np.linalg.norm(gps_vel) <= 1e-2: # If the car doesn't move
x[i], v[i], a[i], yaw[i], w[i], P[i] = kf.correct_w(0.0)
#print("doesn't move !")
else:
dpos = x[i] - temp_pos_yaw
gps_yaw = np.arctan2(dpos[1], dpos[0])
x[i], v[i], a[i], yaw[i], w[i], P[i] = kf.correct_yaw(gps_yaw)
temp_pos_yaw = np.copy(x[i])
gps_idx += 1
# +
width = 16
height = 9
plt.figure(figsize=(width, height))
plt.subplot(2,2,1)
plt.plot(x[:,0], x[:,1])
plt.subplot(2,2,2)
dgps = gps_pos[1:] - gps_pos[:-1]
v_gps = np.zeros((dgps.shape[0],2))
v_gps[:,0] = dgps[:,0] / (gps_t[1:] - gps_t[:-1])
v_gps[:,1] = dgps[:,1] / (gps_t[1:] - gps_t[:-1])
v_gps_norm = np.linalg.norm(v_gps, axis=-1)
plt.plot(gps_t[:-1], v_gps_norm)
plt.plot(t, np.linalg.norm(v,axis=-1))
plt.subplot(2,2,3)
diff = gps_pos[1:] - gps_pos[:-1]
plt.plot(gps_t[:-1], np.arctan2(diff[:,1], diff[:,0]))
plt.plot(t, yaw)
plt.subplot(2,2,4)
plt.plot(t, w)
# +
plt.figure(figsize=(8,4.5))
plt.plot(gps_t[:-1] - gps_t[0], v_gps_norm, label='gps murni')
plt.plot(t - t[0], np.linalg.norm(v,axis=-1), label='filtered')
plt.title("Kelajuan")
plt.xlabel("Waktu (s)")
plt.ylabel("Kelajuan (m/s)")
plt.legend()
plt.savefig('speed.png', dpi=500, transparent=True)
plt.show()
# +
plt.figure(figsize=(8,4.5))
plt.plot(gps_t[:-1] - gps_t[0], np.arctan2(diff[:,1], diff[:,0]), label='gps murni')
plt.plot(t - t[0], yaw, label='filtered')
plt.title("Sudut Yaw")
plt.xlabel("Waktu (s)")
plt.ylabel("Yaw (rad)")
plt.legend()
#plt.savefig('yaw.png', dpi=500, transparent=True)
plt.show()
# -
|
Archieved/kf_gps/test_kf_gps.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# Modelo de pronóstico para la pierna con datos de entrada TSM, CLa
# + slideshow={"slide_type": "-"}
from pandas import DataFrame
from pandas import concat
from pandas import read_csv
from datetime import datetime
from matplotlib import pyplot
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import MinMaxScaler
from keras.models import Sequential
from keras.layers.core import Dense, Activation, Dropout
from keras.layers.recurrent import LSTM
import lstm, time
import numpy as np
def series_to_supervised(data, n_in=1, n_out=1, dropnan=True):
"""
Frame a time series as a supervised learning dataset.
Arguments:
data: Sequence of observations as a list or NumPy array.
n_in: Number of lag observations as input (X).
n_out: Number of observations as output (y).
dropnan: Boolean whether or not to drop rows with NaN values.
Returns:
Pandas DataFrame of series framed for supervised learning.
"""
n_vars = 1 if type(data) is list else data.shape[1]
df = DataFrame(data)
cols, names = list(), list()
# input sequence (t-n, ... t-1)
for i in range(n_in, 0, -1):
cols.append(df.shift(i))
names += [('var%d(t-%d)' % (j+1, i)) for j in range(n_vars)]
# forecast sequence (t, t+1, ... t+n)
for i in range(0, n_out):
cols.append(df.shift(-i))
if i == 0:
names += [('var%d(t)' % (j+1)) for j in range(n_vars)]
else:
names += [('var%d(t+%d)' % (j+1, i)) for j in range(n_vars)]
# put it all together
agg = concat(cols, axis=1)
agg.columns = names
# drop rows with NaN values
if dropnan:
agg.dropna(inplace=True)
return agg
# -
dataset = read_csv('verdillo.csv', header=0, index_col=0)
values = dataset.values
encoder = LabelEncoder()
values[:,1] = encoder.fit_transform(values[:,1])
print(values)
# ensure all data is float
values = values.astype('float32')
# normalize features
scaler = MinMaxScaler(feature_range=(0, 1))
scaled = scaler.fit_transform(values)
reframed = series_to_supervised(scaled, 1, 1)
reframed.drop(reframed.columns[[4,5]], axis=1, inplace=True)
print(values.shape)
# +
# split into train and test sets
values = reframed.values
n_train_hours = 132-24
train = values[:n_train_hours, :]
test = values[n_train_hours:, :]
print(values.shape,train.shape,test.shape)
# split into input and outputs
train_X, train_y = train[:, :-1], train[:, -1]
test_X, test_y = test[:, :-1], test[:, -1]
# reshape input to be 3D [samples, timesteps, features]
train_X = train_X.reshape((train_X.shape[0], 1, train_X.shape[1]))
test_X = test_X.reshape((test_X.shape[0], 1, test_X.shape[1]))
# +
# design network
model = Sequential()
model.add(LSTM(50, input_shape=(train_X.shape[1], train_X.shape[2])))
model.add(Dropout(0.2))
model.add(Activation('tanh'))
model.add(Dense(1))
model.compile(loss='mae', optimizer='adam')
history = model.fit(train_X, train_y, epochs=35, batch_size=10,validation_data=(test_X, test_y),shuffle=False)
# -
pyplot.figure(figsize=(20,10))
pyplot.plot(history.history['loss'], label='train')
pyplot.plot(history.history['val_loss'], label='test')
pyplot.legend()
pyplot.show()
# +
# make a prediction
yhat = model.predict(test_X)
test_X = test_X.reshape((test_X.shape[0], test_X.shape[2]))
# -
print(test_X.shape)
print(yhat.shape)
# invert scaling for forecast
inv_yhat = np.concatenate((yhat, test_X[:, 1:]), axis=1)
inv_yhat = scaler.inverse_transform(inv_yhat)
inv_yhat = inv_yhat[1:,0]
# invert scaling for actual
test_y = test_y.reshape((len(test_y), 1))
inv_y = np.concatenate((test_y, test_X[:, 1:]), axis=1)
inv_y = scaler.inverse_transform(inv_y)
inv_y = inv_y[:22,0]
from sklearn.metrics import mean_squared_error
from scipy.stats.stats import pearsonr
# calculate RMSE
rmse = np.sqrt(mean_squared_error(inv_y, inv_yhat))
r=pearsonr(inv_y, inv_yhat)
print('Test RMSE: %.3f' % rmse)
print('Test R %.3f' %r[0])
pyplot.figure(figsize=(20,10))
pyplot.plot(inv_y, label='y')
pyplot.plot(inv_yhat, label='yhat')
pyplot.legend()
pyplot.show()
|
LSTM_Verdillo_Multi.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import csv
# Dependencies and Setup
import pandas as pd
import numpy as np
terror_data = pd.read_csv("terrorism.csv", encoding = "ISO-8859-1")
terror_data.head()
# -
terror_data_df = terror_data[['iyear', 'country_txt', 'nkill', 'latitude', 'longitude', 'attacktype1_txt',
'targtype1_txt', 'weaptype1_txt']].copy()
terror_data_df.head()
terror_data_df['nkill'] = terror_data_df['nkill'].fillna(0)
terror_data_df.head()
# +
terror_data_complete = terror_data_df.rename(columns={"iyear": "Year", "country_txt":
"Country", "nkill": "Deaths", "latitude": "Latitude", "longitude": "Longitide", "attacktype1_txt": "Attack",
"targtype1_txt": "Target", "weaptype1_txt": "Weapon" })
terror_data_complete.head()
# +
# Create country dictionary to store unique country names and their corresponding death totals
country_dict={}
# Initial counter to tally total deaths
tl_deaths = 0
for index, row in terror_data_complete.iterrows():
# print(row["Country"])
country = (row["Country"])
year = (row["Year"])
if year >= 1990 and year <= 1999:
if country not in country_dict.keys():
# the country just suffered its first terrorism attack
country_dict[row["Country"]] = row["Deaths"]
else:
# add to the death toll
country_dict[row["Country"]] = country_dict[row["Country"]] + row["Deaths"]
print(country_dict)
# -
# Convert dictionary to dataframe
terror_df = pd.DataFrame([country_dict])
terror_df.head()
total_terror_df = terror_df.T
total_terror_df.head()
total_terror_df.reset_index(inplace=True)
total_terror_df.head()
total_terror_df = total_terror_df.rename(columns={"index": "Country", 0: "Deaths"})
total_terror_df.head()
country_codes = pd.read_excel("country_abbrev.xls")
country_codes.head()
# +
# Combine the data into a single dataset
terrorism_complete = pd.merge(total_terror_df, country_codes, how="left", left_on=["Country"], right_on=["country_name"])
terrorism_complete.head()
# -
terrorism_complete=terrorism_complete.rename(columns={"country_code": "code", "Country": "total" })
terrorism_complete.head()
terrorism_complete.drop(columns=["country_name"],inplace=True)
terrorism_complete.head()
# Output results to new csv
terrorism_complete.to_csv("terror90.csv", index=False)
|
terrorism-90.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# PyTorch: nn
# -----------
#
# A fully-connected ReLU network with one hidden layer, trained to predict y from x
# by minimizing squared Euclidean distance.
#
# This implementation uses the nn package from PyTorch to build the network.
# PyTorch autograd makes it easy to define computational graphs and take gradients,
# but raw autograd can be a bit too low-level for defining complex neural networks;
# this is where the nn package can help. The nn package defines a set of Modules,
# which you can think of as a neural network layer that has produces output from
# input and may have some trainable weights.
#
#
# +
import torch
# N is batch size; D_in is input dimension;
# H is hidden dimension; D_out is output dimension.
N, D_in, H, D_out = 64, 1000, 100, 10
# Create random Tensors to hold inputs and outputs
x = torch.randn(N, D_in)
y = torch.randn(N, D_out)
# Use the nn package to define our model as a sequence of layers. nn.Sequential
# is a Module which contains other Modules, and applies them in sequence to
# produce its output. Each Linear Module computes output from input using a
# linear function, and holds internal Tensors for its weight and bias.
model = torch.nn.Sequential(
torch.nn.Linear(D_in, H),
torch.nn.ReLU(),
torch.nn.Linear(H, D_out),
)
# The nn package also contains definitions of popular loss functions; in this
# case we will use Mean Squared Error (MSE) as our loss function.
loss_fn = torch.nn.MSELoss(reduction='sum')
learning_rate = 1e-4
for t in range(500):
# Forward pass: compute predicted y by passing x to the model. Module objects
# override the __call__ operator so you can call them like functions. When
# doing so you pass a Tensor of input data to the Module and it produces
# a Tensor of output data.
y_pred = model(x)
# Compute and print loss. We pass Tensors containing the predicted and true
# values of y, and the loss function returns a Tensor containing the
# loss.
loss = loss_fn(y_pred, y)
print(t, loss.item())
# Zero the gradients before running the backward pass.
model.zero_grad()
# Backward pass: compute gradient of the loss with respect to all the learnable
# parameters of the model. Internally, the parameters of each Module are stored
# in Tensors with requires_grad=True, so this call will compute gradients for
# all learnable parameters in the model.
loss.backward()
# Update the weights using gradient descent. Each parameter is a Tensor, so
# we can access its gradients like we did before.
with torch.no_grad():
for param in model.parameters():
param -= learning_rate * param.grad
|
Deep_Learning/PyTorch/Neural_nets/two_layer_net_nn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Statistical concepts
#
# ## Holding period rates of return
#
# ### Holding period return of any financial instrument with our without cash payouts
#
# Consider a rate of return, such as that generated by a financial instrument over time. We call it rate of return $r$.
#
# When we talk about the return of a financial instrument, such as a stock or a bond, what we really mean is the holding-period return over a specific period during which we hold (own) the instrument.
#
# The *holding period return* would then be
#
# $$HPR=r=\frac{Final price-Starting price+Cash payout(if any)}{Starting price}
# $$
#
# #### Example
# $FinalPrice=100$
# $StartingPrice=90$
# $Cash payout=5$
#
# Then we get
#
# $HPR=\frac{100-90+5}{90}$
#
(100-90+5)/90
# ## How can we think about uncertainty?
#
# - There is considerable uncertainty in financial markets.
# - For example, we cannot be sure about a given share price one year from now.
# - We can organize our *beliefs* about possible *outcomes* by expressing them as *economic scenarios* with specific *probabilities*.
# - For example
#
|
Session002_Preliminaries.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import argparse
import pandas_datareader.data as web
from datetime import datetime
import os
import sys
from ax.plot.contour import plot_contour
from ax.plot.trace import optimization_trace_single_method
from ax.service.managed_loop import optimize
from ax.metrics.branin import branin
from ax.utils.measurement.synthetic_functions import hartmann6
from ax.utils.notebook.plotting import render, init_notebook_plotting
from ax import Experiment, save
from ax import load
# -
from relative_strength import *
spy = pd.DataFrame(web.DataReader("SPY", "av-daily-adjusted", start=datetime(2000, 2, 9), api_key=os.getenv('ALPHAVANTAGE_API_KEY')).close.map(np.log))
spy
thefunc = r_evaluator(spy, 'close')
best_parameters, values, experiment, model = optimize(
parameters = [
{
"name": "longdur",
"type": "range",
"bounds" : [1, 40],
"value_type" : "int",
},
{
"name": "shortdur",
"type": "range",
"bounds": [1, 40],
"value_type" : "int",
},
{
"name": "holdtime",
"type": "range",
"bounds": [1, 40],
"value_type" : "int",
},
{
"name": "thebound",
"type": "range",
"bounds": [-0.3, 0.3],
"value_type" : "float"
},
{
"name": "thesign",
"type": "choice",
"values": [-1, 1],
"value_type" : "float"
},
],
experiment_name="r_test",
objective_name="r_eval",
evaluation_function = thefunc,
minimize=True,
total_trials=1000,
)
best_parameters
values
# +
import numpy as np
from ax.plot.contour import plot_contour
from ax.plot.trace import optimization_trace_single_method
from ax.service.managed_loop import optimize
from ax.metrics.branin import branin
from ax.utils.measurement.synthetic_functions import hartmann6
from ax.utils.notebook.plotting import render, init_notebook_plotting
init_notebook_plotting()
# -
render(plot_contour(model=model, param_x='longdur', param_y='shortdur', metric_name='r_eval'))
theresults= doaverages(spy, 'close', best_parameters['longdur'],best_parameters['shortdur'], best_parameters['holdtime'])
df = theresults[0]
filtered = df[df.close_long - df.close_short * best_parameters['thesign'] > best_parameters['thebound'] * best_parameters['thesign']]
filtered
filtered.dropna(inplace=True)
filtered.describe()
filtered.close_gain.plot()
df.describe()
spy.describe()
spy.head()
spy.tail()
(5.71-4.92)/5000
|
shortnote.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .js
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernel_info:
// name: node_nteract
// kernelspec:
// display_name: Node.js (nteract)
// language: javascript
// name: node_nteract
// ---
// # Exploring Custom Revival with JSON.parse
//
// <svg xmlns="http://www.w3.org/2000/svg" version="1.1" viewBox="0 0 630 630" height="90">
// <g id="logo">
// <rect id="background" x="0" y="0" width="630" height="630" fill="#f7df1e" />
// <path id="j" d="m 165.65,526.47375 48.2125,-29.1775 C 223.16375,513.7875 231.625,527.74 251.92,527.74 c 19.45375,0 31.71875,-7.60975 31.71875,-37.21 l 0,-201.3 59.20375,0 0,202.1375 c 0,61.32 -35.94375,89.23125 -88.385,89.23125 -47.36125,0 -74.8525,-24.52875 -88.8075,-54.13" />
// <path id="s" d="m 375,520.13 48.20625,-27.91125 c 12.69,20.72375 29.1825,35.9475 58.36125,35.9475 24.53125,0 40.17375,-12.26475 40.17375,-29.18125 0,-20.29875 -16.06875,-27.48875 -43.135,-39.32625 l -14.7975,-6.3475 c -42.715,-18.18125 -71.05,-41.0175 -71.05,-89.2275 0,-44.40375 33.83125,-78.2375 86.695,-78.2375 37.6375,0 64.7025,13.11125 84.15375,47.36625 l -46.09625,29.60125 c -10.15,-18.1825 -21.1425,-25.37125 -38.0575,-25.37125 -17.33875,0 -28.335,10.995 -28.335,25.37125 0,17.7625 10.99625,24.9525 36.3675,35.94875 l 14.8,6.3425 c 50.325,21.56875 78.66,43.5575 78.66,93.03375 0,53.2875 -41.86625,82.465 -98.11,82.465 -54.97625,0 -90.5,-26.2175 -107.83625,-60.47375" />
// </g>
// </svg>
// + outputExpanded=false
var Immutable = require('immutable')
var _ = require('lodash')
// + [markdown] outputExpanded=false
// # Revival on Parse
//
// JSON.parse takes an extra argument called a reviver:
//
// ```
// JSON.parse(text[, reviver])
// ```
//
// The reviver accepts two parameters, `key` and `value` and returns the intended `value`. The key will either be a text key on Objects or numbers for when the value is in an Array.
//
// Let's walk through some sample code to check this out.
// + outputExpanded=false
// Classic JSON.parse
JSON.parse('{"a": 2, "b": { "name": "dave" }}')
// + outputExpanded=false
function reviver(key, value) {
if(key === 'name') {
return value + " senior";
}
return value
}
JSON.parse('{"a": 2, "b": { "name": "dave" }}', reviver)
// + [markdown] outputExpanded=false
// This means you can use this to change values based on a key, though you won't know the nested path of the overall JSON object.
//
// Since the string is (expected to be) JSON, there are only two types which are not immutable: `Array` and `Object`. You can use this to your advantage to create frozen or Immutable.js objects while parsing.
// + outputExpanded=false
JSON.parse('{"a": 2, "b": { "name": "dave" }}', (k, v) => Object.freeze(v))
// + outputExpanded=false
function immutableReviver(key, value) {
if (Array.isArray(value)) {
return Immutable.List(value);
}
if (typeof value === 'object') {
return Immutable.Map(value)
}
return value;
}
// + [markdown] outputExpanded=false
// Since it seemed handy enough, I put [`immutable-reviver`](https://github.com/rgbkrk/immutable-reviver) on npm. We'll just use the version written here for now though.
// + outputExpanded=false
revived = JSON.parse('{"a": 2, "b": { "name": "dave" }}', immutableReviver)
// + outputExpanded=false
revived.getIn(['b', 'name'])
// + [markdown] outputExpanded=false
// The reason I started looking into this was because I was trying to see if I could optimize loading of notebooks in nteract. We currently rely on a strategy that goes like:
//
// ```
// notebook = JSON.parse(rawNotebook)
// immutableNotebook = Immutable.fromJS(notebook)
//
// ourNotebook = immutableNotebook.map(...).map(...)... // A series of transformations to create our in-memory representation
// ```
//
// These transformations are mostly to turn notebook cells from this:
//
//
// ```
// {
// "metadata": {
// "collapsed": false,
// "outputExpanded": false
// },
// "cell_type": "markdown",
// "source": [
// "# Outputs you can update by name\n",
// "\n",
// "This notebook demonstrates the new name-based display functionality in the notebook. Previously, notebooks could only attach output to the cell that was currently being executed:\n",
// "\n"
// ]
// }
// ```
//
// into:
//
// ```
// {
// "metadata": {
// "collapsed": false,
// "outputExpanded": false
// },
// "cell_type": "markdown",
// "source": "# Outputs you can update by name\n\nThis notebook demonstrates the new name-based display functionality in the notebook. Previously, notebooks could only attach output to the cell that was currently being executed:\n\n"
// }
// ```
//
// This multi-line string format, introduced by Jupyter, is to accomodate diffing of notebooks in tools like git and GitHub. It's applied to source on cells as well as some output types.
// + [markdown] outputExpanded=false
// We can set up a reviver that handles all the keys that are most likely to have [multi-line strings](https://github.com/jupyter/nbformat/blob/62d6eb8803616d198eaa2024604d1fe923f2a7b3/nbformat/v4/nbformat.v4.schema.json#L386). We'll start with those that are media types that we know end up being encoded as an array of strings.
// + outputExpanded=false
var multilineStringMimetypes = new Set([
'application/javascript',
'text/html',
'text/markdown',
'text/latex',
'image/svg+xml',
'image/gif',
'image/png',
'image/jpeg',
'application/pdf',
'text/plain',
]);
function immutableNBReviver(key, value) {
if (Array.isArray(value)) {
if(multilineStringMimetypes.has(key)) {
return value.join('')
}
return Immutable.List(value);
}
if (typeof value === 'object') {
return Immutable.Map(value)
}
return value;
}
// + [markdown] outputExpanded=false
// We can also set up a "greedy" reviver that will also convert `source` and `text` fields. The primary problem with this though, because of how JSON.parse works is that we have no idea if it's a key in a cell where we expect, part of someone else's JSON payload, or in metadata.
// + outputExpanded=false
var specialKeys = new Set([
'application/javascript',
'text/html',
'text/markdown',
'text/latex',
'image/svg+xml',
'image/gif',
'image/png',
'image/jpeg',
'application/pdf',
'text/plain',
'source',
'text',
]);
function immutableGreedyReviver(key, value) {
if (Array.isArray(value)) {
if(specialKeys.has(key)) {
return value.join('')
}
return Immutable.List(value);
}
if (typeof value === 'object') {
return Immutable.Map(value)
}
return value;
}
// + [markdown] outputExpanded=false
// # Our runtime harnesses
//
// To evaluate the speed at which we can revive our objects, we'll set up a little testing harness.
// + outputExpanded=false
// Some logger that uses process.hrtime that I ripped off Stack Overflow, since we want to use timing in a way that we can't with console.time
[ a, o, ms, s, log ] = ( function * () {
yield * [
( process.hrtime )(),
process.hrtime,
ms => ( ( ms[ 0 ] * 1e9 + ms[ 1 ] ) / 1000000 ),
s => s / 1000,
() => {
const f = o( a ), msf = ms( f ), sf = s( msf );
return { a, o: f, ms: msf, s: sf };
}
];
} )();
// + outputExpanded=false
// Calculate the milliseconds it takes to run f
function measure(f) {
start = log()
f()
end = log()
return end.ms - start.ms
}
// measure the function run n times, return the mean
function runTrials(f, n=1000) {
values = []
for(var ii=0; ii < n; ii++) {
values.push(measure(f))
}
return values.reduce((a, b) => a + b, 0)/n
}
// + [markdown] outputExpanded=false
// With our harness all set up, we can run through all the notebooks we have locally to see how they perform with different revivers.
// -
notebooks = require('glob').sync('./*.ipynb')
// + outputExpanded=false
for(var notebookPath of notebooks) {
console.log("\n ----- ", path.basename(notebookPath))
raw = fs.readFileSync(notebookPath)
var tests = [
{ name: 'straight JSON.parse', f: () => { JSON.parse(raw) } },
{ name: 'Object.freeze', f: () => { JSON.parse(raw, (k, v) => Object.freeze(v)) } },
{ name: 'basic Immutable', f: () => { JSON.parse(raw, immutableReviver) } },
{ name: 'immutable notebook', f: () => { JSON.parse(raw, immutableNBReviver) } },
{ name: 'immutable greedy nb', f: () => { JSON.parse(raw, immutableGreedyReviver) } },
// { name: 'fromJS', f: () => { JSON.parse(raw, (k, v) => Immutable.fromJS(v)) } },
// { name: 'current commutable way', f: () => { commutable.fromJS(JSON.parse(raw)) } },
]
for(var test of tests) {
mean = runTrials(test.f, 100)
console.log(_.padEnd(test.name, 30), mean)
}
}
// + [markdown] outputExpanded=false
// # Evaluating revivers for notebook loading.
//
// Within nteract we are inevitably going to end up creating an immutable structure. These measurements only make sense in the context of running both the initial `JSON.parse` followed by the transformations. To give it a rough guess, I'll only compare a few I can evaluate.
// + outputExpanded=false
for(var notebookPath of notebooks) {
console.log("\n ----- ", path.basename(notebookPath))
raw = fs.readFileSync(notebookPath)
var tests = [
{ name: 'straight JSON.parse baseline', f: () => { JSON.parse(raw) } },
{ name: 'Object.freeze baseline', f: () => { JSON.parse(raw, (k,v) => Object.freeze(v)) } },
{ name: 'immutable greedy nb', f: () => { JSON.parse(raw, immutableGreedyReviver) } },
]
for(var test of tests) {
mean = runTrials(test.f, 100)
console.log(_.padEnd(test.name, 50), mean.toString().slice(0,10), 'ms')
}
}
// + [markdown] outputExpanded=false
// Since these are in milliseconds and the difference is not much, it seems like maybe this doesn't need to be optimized. In the case of the altair notebook, which has a pretty big JSON structure inside (and only one!), perhaps it would make sense if some of our structure is frozen objects (don't force vega payloads to be Immutable Maps).
//
// ```
// ----- altair.ipynb
// straight JSON.parse baseline 1.10996391 ms
// Object.freeze baseline 2.29745900 ms
// straight JSON.parse then commutable conversion 6.84918417 ms
// immutable greedy nb 5.85418076 ms
// ```
|
applications/desktop/example-notebooks/immutable-revival.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div style="width: 100%; overflow: hidden;">
# <div style="width: 150px; float: left;"> <img src="https://raw.githubusercontent.com/DataForScience/Networks/master/data/D4Sci_logo_ball.png" alt="Data For Science, Inc" align="left" border="0" width=150px> </div>
# <div style="float: left; margin-left: 10px;"> <h1>Causal Inference In Statistics - A Primer</h1>
# <h1>1.4 Graphs</h1>
# <p><NAME><br/>
# <a href="http://www.data4sci.com/">www.data4sci.com</a><br/>
# @bgoncalves, @data4sci</p></div>
# <div style="float: right; margin-right:10px;"> <p><a href="https://amzn.to/3gsFlkO" target=_blank><img src='data/causality.jpeg' width='100px'>
# <!--Amazon Affiliate Link--></a></p></div>
# </div>
# +
from collections import Counter
from pprint import pprint
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import networkx as nx
from networkx.drawing.nx_pydot import graphviz_layout
import watermark
# %load_ext watermark
# %matplotlib inline
# -
# We start by print out the versions of the libraries we're using for future reference
# %watermark -n -v -m -g -iv
# Load default figure style
plt.style.use('./d4sci.mplstyle')
colors = plt.rcParams['axes.prop_cycle'].by_key()['color']
# ## Graphs
# Graphs allow us to represent relationships (edges) between entities (nodes). We will use the [NetworkX](https://networkx.github.io/) Python package to handle all of our graphs for the rest of the course.
# We start by instanciating a new Graph object
G = nx.Graph()
# We can add nodes one by one or from a list of nodes. Any Python hashable object can be a node:
G.add_nodes_from(['X', 'Y', 'Z'])
# Similarly, we can add edges one by one of from a list. Edges are a pair of nodes with an optional dictionary contianing data about the edge:
G.add_edges_from([('X', 'Y', {'label':'A'}), ('Y', 'Z', {'label':'B'})])
# NetworkX has an extensive set of drawing functionality. We can simply draw the graph directly:
nx.draw(G)
# Or customize the plot by definining our own layout (node coordinates):
pos = {'X':(-1, 0), 'Y':(0, 0), 'Z':(1, 0)}
labels = {(node_i, node_j) : label for node_i, node_j, label in G.edges(data='label', default='')}
# And the way we want the graph to be plotted using Matplotlib.
fig, ax = plt.subplots(1, figsize=(4,1))
nx.draw(G, pos, ax=ax, node_color=colors[0])
nx.draw_networkx_labels(G, pos, ax=ax)
nx.draw_networkx_edge_labels(G, pos, labels, ax=ax)
fig.tight_layout()
# To create a directed graph, we simply use DiGraph()
G = nx.DiGraph()
G.add_nodes_from(['X', 'Y', 'Z'])
G.add_edges_from([('X', 'Y', {'label':'A'}), ('Y', 'Z', {'label':'B'})])
# And we can plot it in just the same way
# +
pos = {'X':(-1, 0), 'Y':(0, 0), 'Z':(1, 0)}
labels = {(node_i, node_j) : label for node_i, node_j, label in G.edges(data='label', default='')}
fig, ax = plt.subplots(1, figsize=(4,1))
nx.draw(G, pos, ax=ax, node_color=colors[0])
nx.draw_networkx_labels(G, pos, ax=ax)
nx.draw_networkx_edge_labels(G, pos, labels, ax=ax)
fig.tight_layout()
# -
# And if we add mode edges, the layout naturally needs to change as well
G = nx.DiGraph()
G.add_edges_from([
('U', 'X', {'label':'A'}),
('U', 'Y', {'label':'C'}),
('X', 'Y', {'label':'B'})])
# +
pos = {'U': (0, 1), 'X': (-1, 0), 'Y': (1, 0)}
labels = {(node_i, node_j) : label for node_i, node_j, label in G.edges(data='label', default='')}
fig, ax = plt.subplots(1, figsize=(3,2.2))
nx.draw(G, pos, ax=ax, node_color=colors[0])
nx.draw_networkx_labels(G, pos, ax=ax)
#nx.draw_networkx_edge_labels(G, pos, labels, ax=ax)
fig.tight_layout()
# -
# And yet another example
G = nx.DiGraph()
G.add_edges_from([
('X', 'Y'),
('X', 'W'),
('W', 'Y'),
('W', 'Z'),
('Y', 'T'),
('Y', 'Z'),
('Z', 'T')])
# +
pos = {'X': (-1, 0), 'Y': (0, 0), 'Z': (1, 0), 'W': (0, 1), 'T' : (0.5, -0.5)}
fig, ax = plt.subplots(1, figsize=(4,2.3))
nx.draw(G, pos, ax=ax, node_color=colors[0])
nx.draw_networkx_labels(G, pos, ax=ax)
fig.tight_layout()
# -
# NetworkX also gives us simple ways to directly ask questions of this DAG
# We can get the parents (predecessors) and ancestors of Z
print('Parents:', list(G.predecessors('Z')), 'Ancestors:', list(nx.ancestors(G, 'Z')))
# And the children (successors)and decendents of W:
print('Childrens:', list(G.successors('W')), 'Descendants:', list(nx.descendants(G, 'W')))
# We can easily find all the directed paths between X and T
directed_paths = {tuple(path) for path in nx.all_simple_paths(G, 'X', 'T')}
print(directed_paths)
# And write a helper function to plot them all out on top of our DAG:
def plot_path(G, pos, path, ax=None):
fig = None
if ax == None:
fig, ax = plt.subplots(1, figsize=(4,2.3))
edgelist = {(path[i], path[i+1]) for i in range(len(path)-1)}
edges = set(G.edges()) - set(edgelist)
nx.draw(G, pos, node_color=colors[0], ax=ax, edgelist=[])
nx.draw_networkx_labels(G, pos, ax=ax)
nx.draw_networkx_edges(G, pos,
edgelist=edgelist,
width=3, edge_color=colors[1], ax=ax)
nx.draw_networkx_edges(G, pos,
edgelist=edges,
width=1, ax=ax)
if fig is not None:
fig.tight_layout()
# And easily plot them all side by side
# +
fig, axs = plt.subplots(3, 2, figsize=(8, 6.9))
axs = np.array(axs).flatten()
for i, path in enumerate(directed_paths):
plot_path(G, pos, path, axs[i])
axs[-1].axis('off')
fig.tight_layout()
# -
# In some cases, we're also interested in all the paths, even if they break the direction of the edges. We can extract them all by converting our __DiGraph__ into a regular __Graph__ before calling __nx.all_simple_paths__
all_paths = {tuple(path) for path in nx.all_simple_paths(G.to_undirected(G), 'X', 'T')}
# And we see that we find two extra paths
print(all_paths-directed_paths)
# That we can also visualize
# +
fig, axs = plt.subplots(1, 2, figsize=(8, 2.3))
axs = np.array(axs).flatten()
new_paths = all_paths-directed_paths
for i, path in enumerate(new_paths):
plot_path(G, pos, path, axs[i])
axs[-1].axis('off')
fig.tight_layout()
# -
# <div style="width: 100%; overflow: hidden;">
# <img src="data/D4Sci_logo_full.png" alt="Data For Science, Inc" align="center" border="0" width=300px>
# </div>
|
1.4 - Graphs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div style="width: 100%; overflow: hidden;">
# <div style="width: 150px; float: left;"> <img src="https://raw.githubusercontent.com/DataForScience/Graphs4Sci/master/data/D4Sci_logo_ball.png" alt="Data For Science, Inc" align="left" border="0" width=150px> </div>
# <div style="float: left; margin-left: 10px;"><h1>Visualization for Science</h1>
# <h1>Timerseries Map</h1>
# <a href="http://www.data4sci.com/">www.data4sci.com</a><br/>
# @bgoncalves, @data4sci</p></div>
# </div>
# +
import pandas as pd
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
from matplotlib.collections import PatchCollection
# %matplotlib inline
# -
# Load default figure style
colors = [
'#51a7f9',
'#cf51f9',
'#70bf41',
'#f39019',
'#f9e351',
'#f9517b',
'#6d904f',
'#8b8b8b',
'#810f7c']
# ## Johns Hopkins University Dataset
# The JHU CoVID dataset has steadly become the unofficial standard dataset for the number of cases and deaths aroudn the world. You can download the continuously updated data from their [github repository](https://github.com/CSSEGISandData/COVID-19/):
url = 'https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/'
# The repository contains 4 time series files:
#
# - time_series_covid19_confirmed_global.csv
# - time_series_covid19_deaths_global.csv
# - time_series_covid19_confirmed_US.csv
# - time_series_covid19_deaths_US.csv
#
# correponding to confirmed cases and number of deaths globally by country and for the US by state.
#
# These curated files result from an aggregation and standardization of several dozen national health authority reports and have become the go-to
# ## US Data
#
#
# Now we take a detailed look at the situation within the US. The JHU github also has data for the US by state and county (admin2) level.
confirmed_US = pd.read_csv(url + 'time_series_covid19_confirmed_US.csv')
# As in the case of the country level data, there are several other columns that we don't need
confirmed_US.head()
# We remove the extraneous columns
del_columns = [
'UID',
'iso2',
'iso3',
'code3',
'FIPS',
'Country_Region',
'Lat',
'Long_',
'Combined_Key',
'Admin2'
]
# And aggregate at the state level.
confirmed_US.drop(columns=del_columns, inplace=True)
confirmed_US = confirmed_US.groupby('Province_State').sum().T
confirmed_US.index = pd.to_datetime(confirmed_US.index)
# ### Country Overview
# A simply way of getting a quick overview of the situation in all the states is to include some degree of geographical information to plot the time series of all the states side by side. We opt to use a block map where each state is represented by an equal size square block where we can plot the time series.
# We load the coordinates of each state from a csv file
states = pd.read_csv('https://raw.githubusercontent.com/DataForScience/Epidemiology101/master/data/states.csv', index_col=0)
# And create a convenient dictionary to convert from state abbreviations to full state names
state_dict = dict(states.reset_index()[['index', 'name']].values)
# And now we plot the time series for each state, normalized by the peak value, within each block. Further, we color the block based on the current value of the number of cases:
#
# - Green, below 25% of the peak value
# - Orange, between 25-75% of the peak value
# - Red, above 75% of the peak value
#
# as an indication of how well each state is doing.
confirmed_US = confirmed_US[:'2020']
# +
fig, ax = plt.subplots(1, figsize=(16, 22))
ax.set_aspect(1.)
ax.invert_yaxis()
patches = []
color_list = []
x = np.linspace(0., 0.75, confirmed_US.shape[0])
for state in states.index:
daily = confirmed_US[state_dict[state]].diff(1).rolling(7).mean()
timeline = (daily/daily.max()).fillna(0).values
color = colors[3]
if timeline[-1] > 0.75:
color = colors[5]
elif timeline[-1] < 0.25:
color = colors[2]
fancybox = mpatches.FancyBboxPatch([states.x[state]-0.5,
states.y[state]-0.5], 0.8, 0.8,
boxstyle=mpatches.BoxStyle("Round",
pad=0.06))
patches.append(fancybox)
color_list.append(color)
ax.text(y=states.y[state]-0.4, x=states.x[state]-0.4,
s=state, horizontalalignment='center',
verticalalignment='center', fontsize=15)
ax.plot(states.x[state]+x-0.5, states.y[state]-timeline/1.4+0.25,
lw=1, color=colors[1])
collection = PatchCollection(patches, facecolors=color_list, alpha=0.3)
ax.add_collection(collection)
ax.text(x=3.4,y=8, s='> 75% Maximum', color=colors[5], fontsize=15, ha='left')
ax.text(x=5, y=8, s='Otherwise', color=colors[3], fontsize=15, ha='left')
ax.text(x=6, y=8, s='< 25% Maximum', color=colors[2], fontsize=15, ha='left')
ax.set_title(confirmed_US.index[-1].strftime('%Y-%m-%d'), fontsize=32)
ax.axis('off')
fig.patch.set_facecolor('#FFFFFF')
fig.tight_layout()
# -
# <div style="width: 100%; overflow: hidden;">
# <img src="https://raw.githubusercontent.com/DataForScience/Graphs4Sci/master/data/D4Sci_logo_full.png" alt="Data For Science, Inc" align="center" border="0" width=300px>
# </div>
|
Timeseries Map.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
from brain import models
robot_info = requests.get('http://plfelt-mbp.local:8080/robots/marion').json()
models.create_end_to_end_model1(robot_info)
from keras.layers import concatenate
sensor_info = robot_info['sensors'][0]
mm=models.create_input_model_for(sensor_info)
ii=mm.get_input_at(0)
ss=ii.get_shape()
ss.as_list()
mm.name
|
server/volumes/notebooks/sandbox.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import warnings
warnings.filterwarnings('ignore')
warnings.filterwarnings("ignore", category=DeprecationWarning)
# -
# Importing the datasetm
dataset = pd.read_csv(r'C:\Users\IDEAPAD 320\Desktop\datasets\Churn_Modelling.csv')
X = dataset.iloc[:, 3:13].values
y = dataset.iloc[:, 13].values
dataset.head()
X
y
# +
from sklearn.preprocessing import LabelEncoder, OneHotEncoder
from sklearn.compose import ColumnTransformer
labelencoder_X_2 = LabelEncoder()
X[:, 2] = labelencoder_X_2.fit_transform(X[:, 2])
columnTransformer = ColumnTransformer([('encoder', OneHotEncoder(), [1])], remainder='passthrough')
X = np.array(columnTransformer.fit_transform(X), dtype = np.str)
X = X[:, 1:]
# -
# Splitting the dataset into the Training set and Test set
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 0)
# Feature Scaling
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
# Importing the Keras libraries and packages
import keras
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
# +
# Initialising the ANN
classifier = Sequential()
# Adding the input layer and the first hidden layer
classifier.add(Dense(output_dim = 6, init = 'uniform', activation = 'relu', input_dim = 11))
# Adding the second hidden layer
classifier.add(Dense(output_dim = 6, init = 'uniform', activation = 'relu'))
# Adding the output layer
classifier.add(Dense(output_dim = 1, init = 'uniform', activation = 'sigmoid'))
# Compiling the ANN
classifier.compile(optimizer = 'adam', loss = 'binary_crossentropy', metrics = ['accuracy'])
# -
# Fitting the ANN to the Training set
classifier.fit(X_train, y_train, batch_size = 10, nb_epoch = 100)
# +
# Predicting the Test set results
y_pred = classifier.predict(X_test)
y_pred = (y_pred > 0.5)
# -
# Making the Confusion Matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
cm
correct_prediction = cm[0][0]+cm[1][1]
wrong_prediction = cm[0][1]+cm[1][0]
total = correct_prediction + wrong_prediction
correct_prediction
wrong_prediction
accuracy = (correct_prediction/total)*100
accuracy
|
Deep Learning using Tensorflow Keras/Churn-Model-using-ANN/ANN - Churn Modelling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
import json
import random
from os import path
import pandas as pd
import numpy as np
from Bio import SeqIO
from Bio.Seq import Seq
from Bio.SeqRecord import SeqRecord
from Bio import AlignIO
from Bio.Align import MultipleSeqAlignment
from Bio.Align import AlignInfo
import matplotlib.pyplot as plt
import matplotlib.lines as mlines
import seaborn as sns
from scipy import stats
from collections import Counter
import matplotlib.ticker as ticker
def count_muts_by_site(cov, kind, window_size, lineage=None):
reference_file = '../'+str(cov)+'/config/'+str(cov)+'_spike_reference.gb'
for seq_record in SeqIO.parse(reference_file, "genbank"):
spike_len = len(seq_record)
for feature in seq_record.features:
if feature.type == 'CDS':
if feature.qualifiers['gene'] == ['s1']:
s1_nt_loc = [int(feature.location.start), int(feature.location.end)]
s1_aa_loc = [int(s1_nt_loc[0]/3 +1), int(s1_nt_loc[1]/3)]
if feature.qualifiers['gene'] == ['s2']:
s2_nt_loc = [int(feature.location.start), int(feature.location.end)]
s2_aa_loc = [int(s2_nt_loc[0]/3 +1), int(s2_nt_loc[1]/3)]
if kind == 'aa':
muts_file = '../'+str(cov)+'/results/aa_muts_'+str(cov)+'_spike.json'
mut_key = 'aa_muts'
elif kind == 'nt':
muts_file = '../'+str(cov)+'/results/nt_muts_'+str(cov)+'_spike.json'
mut_key = 'muts'
if lineage != None:
clades_file = '../'+str(cov)+'/results/clades_spike.json'
with open(clades_file) as clade_handle:
clade_dict = json.load(clade_handle)
#find nodes within specified lineage
lineage_nodes = []
for node, node_dict in clade_dict['nodes'].items():
if node_dict['clade_membership'] == lineage:
lineage_nodes.append(node)
with open(muts_file) as json_handle:
mut_dict = json.load(json_handle)
##store all muts in spike
muts = []
for node, node_dict in mut_dict['nodes'].items():
if lineage!= None:
if node in lineage_nodes:
if kind == 'aa':
if 'spike' in node_dict['aa_muts'].keys():
muts+=node_dict['aa_muts']['spike']
elif kind == 'nt':
muts+=node_dict['muts']
elif lineage == None:
if kind == 'aa':
if 'spike' in node_dict['aa_muts'].keys():
muts+=node_dict['aa_muts']['spike']
elif kind == 'nt':
muts+=node_dict['muts']
##tally all muts in spike
##by mut
muts_count = Counter(muts)
#by site
mut_sites = [int(x[1:-1]) for x in muts]
sites_count = Counter(mut_sites)
#add zeros for all unmutated sites
if kind == 'nt':
for pos in range(spike_len):
if pos not in sites_count.keys():
sites_count[pos] = 0
elif kind == 'aa':
for pos in range(int(spike_len/3)):
if pos not in sites_count.keys():
sites_count[pos] = 0
#count mutations in 20 nt window
if kind == 'aa':
s1_muts_per_window = count_muts_per_window(s1_aa_loc, sites_count, window_size)
s2_muts_per_window = count_muts_per_window(s2_aa_loc, sites_count, window_size)
s1_avg_muts = average_num_muts_per_site(s1_aa_loc, sites_count)
s2_avg_muts = average_num_muts_per_site(s2_aa_loc, sites_count)
spike_avg_muts = average_num_muts_per_site([s1_aa_loc[0], s2_aa_loc[1]], sites_count)
if kind == 'nt':
s1_muts_per_window = count_muts_per_window(s1_nt_loc, sites_count, window_size)
s2_muts_per_window = count_muts_per_window(s2_nt_loc, sites_count, window_size)
s1_avg_muts = average_num_muts_per_site(s1_nt_loc, sites_count)
s2_avg_muts = average_num_muts_per_site(s2_nt_loc, sites_count)
spike_avg_muts = average_num_muts_per_site([s1_nt_loc[0], s2_nt_loc[1]], sites_count)
return muts_count, sites_count, s1_muts_per_window, s2_muts_per_window, s1_avg_muts, s2_avg_muts, spike_avg_muts
# +
#sites that mutate the most
muts_count, sites_count, s1_muts_per_window, s2_muts_per_window, s1_avg_muts, s2_avg_muts, spike_avg_muts = count_muts_by_site('oc43', 'aa', 20, 'A')
for site, count in sites_count.items():
if count>5:
print(site)
# muts_count, sites_count, s1_muts_per_window, s2_muts_per_window = count_muts_by_site('oc43', 'aa', 'B')
# for site, count in sites_count.items():
# if count>2:
# print(site)
# +
#average number of muts per site in S1, S2
def average_num_muts_per_site(loc, sites_count):
total_muts = 0
for site, count in sites_count.items():
if site in range(loc[0], loc[1]):
total_muts+=count
region_len = len(range(loc[0], loc[1]))
# print(total_muts, region_len)
avg_muts_per_site = total_muts/region_len
return avg_muts_per_site
# -
def count_muts_per_window(loc, sites_count, window_size):
region_length = loc[1]-loc[0]
windows = []
for i in range(loc[0], loc[1], window_size):
if i+20>loc[1]:
window = range(i, loc[1])
else:
window = range(i,i + window_size)
windows.append(window)
window_counts = {}
for window in windows:
window_count = 0
for site, count in sites_count.items():
if site in window:
window_count+=count
window_counts[window[0]] = window_count
return window_counts
### Not all CoV Spikes are the same length, give each residue a relative position along spike? Or plot CoVs separately?
### Add zero counts for unmutated sites
#average number of mutations for S1 and S2 of each CoV
def average_muts_per_site_s1_s2(covs, kind, window_size):
to_plot = []
for cov in covs:
if cov == '229e':
muts_count, sites_count, s1_muts_per_window, s2_muts_per_window, s1_avg_muts, s2_avg_muts, spike_avg_muts = count_muts_by_site(cov, kind, window_size)
to_plot.append({'cov' : cov, 'cov_lineage' : cov, 'type_of_mut': kind,
's1_avg_muts': s1_avg_muts, 's2_avg_muts': s2_avg_muts, 'spike_avg_muts': spike_avg_muts})
else:
lineages = ['A', 'B']
for lineage in lineages:
muts_count, sites_count, s1_muts_per_window, s2_muts_per_window, s1_avg_muts, s2_avg_muts, spike_avg_muts = count_muts_by_site(cov, kind, window_size, lineage=lineage)
to_plot.append({'cov' : cov, 'cov_lineage' : cov+lineage, 'type_of_mut': kind,
's1_avg_muts': s1_avg_muts, 's2_avg_muts': s2_avg_muts, 'spike_avg_muts': spike_avg_muts})
to_plot_df = pd.DataFrame(to_plot)
print(to_plot_df)
def plot_muts_by_pos_separate_axes(covs, kind, no_hku1_lineages=False, window_size=20, filename=None):
#make dataframe of mutation counts at all sites for all CoVs
to_plot = []
#positions of S1 and S2 for each CoV
domain_positions = {}
#list of all cov/lineages to plot
cov_lineages = []
for cov in covs:
reference_file = '../'+str(cov)+'/config/'+str(cov)+'_spike_reference.gb'
for seq_record in SeqIO.parse(reference_file, "genbank"):
spike_len = len(seq_record)
for feature in seq_record.features:
if feature.type == 'CDS':
if feature.qualifiers['gene'][0] == 's1':
domain_positions[cov] = {'s1_start': feature.location.start,
's1_end': feature.location.end, 'spike_len': spike_len}
no_lineage_covs = ['229e', 'nl63']
if no_hku1_lineages==True:
no_lineage_covs.append('hku1')
if cov in no_lineage_covs:
if cov not in cov_lineages:
cov_lineages.append(str(cov))
muts_count, sites_count, s1_muts_per_window, s2_muts_per_window, s1_avg_muts, s2_avg_muts, spike_avg_muts = count_muts_by_site(cov, kind, window_size)
for site, count in sites_count.items():
to_plot.append({'cov' : cov, 'cov_lineage' : cov, 'position': int(site),
'count': int(count), 'type_of_mut': kind})
else:
lineages = ['A', 'B']
for lineage in lineages:
if cov+lineage not in cov_lineages:
cov_lineages.append(str(cov+lineage))
muts_count, sites_count, s1_muts_per_window, s2_muts_per_window, s1_avg_muts, s2_avg_muts, spike_avg_muts = count_muts_by_site(cov, kind, window_size, lineage=lineage)
for site, count in sites_count.items():
to_plot.append({'cov' : cov, 'cov_lineage' : cov+lineage, 'position': int(site),
'count': int(count), 'type_of_mut': kind})
to_plot_df = pd.DataFrame(to_plot)
color_map = {'oc43A': '#208288', 'oc43B':'#76C7BE', '229e': '#0B194C',
'nl63A': '#87C735', 'nl63B': '#009888', 'nl63': '#87C735',
'hku1A': '#2E74B3', 'hku1B': '#92B2DE', 'hku1': '#255191'}
fig, axes = plt.subplots(len(cov_lineages), 1, figsize=(14, 10), sharex=False, sharey=True)
for ax in range(len(axes)):
cov_lineage = cov_lineages[ax]
if cov_lineage in covs:
cov = cov_lineage
else:
cov = str(cov_lineage[:-1])
sns.scatterplot(x="position", y="count", hue='cov_lineage', linewidth=0.3,
palette=color_map, data=to_plot_df[to_plot_df['cov_lineage']==cov_lineage], ax=axes[ax])
if kind == 'aa':
axes[ax].set_ylabel("amino acid substitutions per site", fontsize=12)
axes[ax].set_xlabel("", fontsize=12)
for tick in axes[ax].xaxis.get_major_ticks():
tick.label.set_fontsize(14)
for tick in axes[ax].yaxis.get_major_ticks():
tick.label.set_fontsize(14)
elif kind == 'nt':
axes[ax].set(xlabel="", ylabel="nucleotide mutations per site")
trans = axes[ax].get_xaxis_transform()
if kind == 'nt':
axes[ax].annotate('S1', xy=(int(domain_positions[cov]['s1_start']), -.18),
fontsize=14, xycoords=trans, ha="center", va="top")
axes[ax].plot([int(domain_positions[cov]['s1_start']),int(domain_positions[cov]['s1_end'])],
[-.14,-.14], color="#878787", linewidth=10, transform=trans, clip_on=False)
elif kind == 'aa':
axes[ax].fill_between([0,(int(domain_positions[cov]['s1_end'])/3)], y1=[0,0],
y2=[13.5,13.5],
alpha=0.2, color="#a5a5a5")
axes[ax].fill_between([((int(domain_positions[cov]['s1_end'])/3)+3),(int(domain_positions[cov]['spike_len'])/3)], y1=[0,0],
y2=[13.5,13.5],
alpha=0.05, color="white")
axes[ax].annotate('S1', xy=((int(domain_positions[cov]['s1_start'])/3), 1.05),
fontsize=14, color= '#515151', xycoords=trans, ha="center", va="top")
axes[ax].annotate('S2', xy=((int(domain_positions[cov]['s1_end'])/3+3), 1.05),
fontsize=14, color="#878787", xycoords=trans, ha="center", va="top")
axes[ax].legend(bbox_to_anchor=(1, 1))
axes[ax].spines['right'].set_visible(False)
axes[ax].spines['top'].set_visible(False)
fig.tight_layout(pad=3.0)
if filename:
fig.savefig(filename, dpi=300)
#Dec 18: improve aestheics
plot_muts_by_pos_separate_axes(['oc43', '229e'], 'aa', filename = 'plots/fig2_dec18.png')
#Dec 18: improve aestheics
plot_muts_by_pos_separate_axes(['nl63', 'hku1'], 'aa', no_hku1_lineages=True, filename = 'plots/fig2supp_dec18.png')
#Dec 18: improve aestheics
plot_muts_by_pos_separate_axes(['hku1'], 'aa', filename = 'plots/fig2suppb_dec18.png')
|
antigenic_evolution/site_mutation_rank.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ArpitaChatterjee/Comedian-Routine-Analysis/blob/main/Topic_Modeling.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="PqLkVyXf__Pe"
# # Topic Modeling
# + colab={"base_uri": "https://localhost:8080/"} id="jD-kJ8PCAED4" outputId="e39c4f74-7f84-4cd2-c427-59503419f180"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="-8ifrKPN__Ph"
# ## Introduction
# + [markdown] id="1JquCT1N__Pi"
# The ultimate goal of topic modeling is to find various topics that are present in the corpus. Each document in the corpus will be made up of at least one topic, if not multiple topics.
#
# Here, I'll be using **Latent Dirichlet Allocation (LDA)**, which is one of many topic modeling techniques. It was specifically designed for text data.
#
# To use a topic modeling technique, i need (1) a document-term matrix and (2) the number of topics you would like the algorithm to pick up.
# + [markdown] id="Ov_ahXon__Pi"
# ## Topic Modeling - Attempt #1 (All Text)
# + colab={"base_uri": "https://localhost:8080/", "height": 475} id="jqABSzMX__Pj" outputId="9db398e6-25c1-41ea-9cb0-1082f06fb0d4"
# Let's read in our document-term matrix
import pandas as pd
import pickle
data = pd.read_pickle('/content/drive/MyDrive/Colab Notebooks/NLP/dtm_stop.pkl')
data
# + id="_J0Jki-r__Pj"
# Import the necessary modules for LDA with gensim
# Terminal / Anaconda Navigator: conda install -c conda-forge gensim
from gensim import matutils, models
import scipy.sparse
# import logging
# logging.basicConfig(format='%(asctime)s : %(levelname)s : %(message)s', level=logging.INFO)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="LBhbvv1Z__Pk" outputId="5866cde6-aa5f-4550-c3d0-7af8ef564679"
# One of the required inputs is a term-document matrix
tdm = data.transpose()
tdm.head()
# + id="TiLjon3h__Pk"
# We're going to put the term-document matrix into a new gensim format, from df --> sparse matrix --> gensim corpus
sparse_counts = scipy.sparse.csr_matrix(tdm)
corpus = matutils.Sparse2Corpus(sparse_counts)
# + colab={"base_uri": "https://localhost:8080/"} id="sHJXzoOn__Pl" outputId="ea75d403-379b-44af-a87f-5cad0ae22169"
# Gensim also requires dictionary of the all terms and their respective location in the term-document matrix
cv = pickle.load(open("/content/drive/MyDrive/Colab Notebooks/NLP/cv_stop.pkl", "rb"))
id2word = dict((v, k) for k, v in cv.vocabulary_.items())
# + [markdown] id="1SgeWj7I__Pl"
# Now that we have the corpus (term-document matrix) and id2word (dictionary of location: term), we need to specify two other parameters - the number of topics and the number of passes. start the number of topics at 2, see if the results make sense, and increase the number from there.
# + colab={"base_uri": "https://localhost:8080/"} id="0vOcwYR9__Pl" outputId="e6a1cdb1-4e88-4f69-a19f-b79af516b06f"
# Now that we have the corpus (term-document matrix) and id2word (dictionary of location: term),
# we need to specify two other parameters as well - the number of topics and the number of passes
lda = models.LdaModel(corpus=corpus, id2word=id2word, num_topics=2, passes=10)
lda.print_topics()
# + colab={"base_uri": "https://localhost:8080/"} id="KX32o3hi__Pm" outputId="c4e8cdb6-047b-49e1-ae9a-279728a7b06a"
# LDA for num_topics = 3
lda = models.LdaModel(corpus=corpus, id2word=id2word, num_topics=3, passes=10)
lda.print_topics()
# + id="xH2GfUVE__Pm"
# LDA for num_topics = 4
lda = models.LdaModel(corpus=corpus, id2word=id2word, num_topics=4, passes=10)
lda.print_topics()
# + [markdown] id="iXgNW0kM__Pm"
# These topics aren't looking too great. We've tried modifying our parameters. Let's try modifying our terms list as well.
# + [markdown] id="4JXw6nQm__Pn"
# ## Topic Modeling - Attempt #2 (Nouns Only)
# + [markdown] id="qO-Q2gDG__Pn"
# One popular trick is to look only at terms that are from one part of speech (only nouns, only adjectives, etc.). Check out the UPenn tag set: https://www.ling.upenn.edu/courses/Fall_2003/ling001/penn_treebank_pos.html.
# + id="FGeRMZqn__Pn"
# Let's create a function to pull out nouns from a string of text
from nltk import word_tokenize, pos_tag
def nouns(text):
'''Given a string of text, tokenize the text and pull out only the nouns.'''
is_noun = lambda pos: pos[:2] == 'NN'
tokenized = word_tokenize(text)
all_nouns = [word for (word, pos) in pos_tag(tokenized) if is_noun(pos)]
return ' '.join(all_nouns)
# + colab={"base_uri": "https://localhost:8080/", "height": 425} id="ps2keqoG__Pn" outputId="b2339b83-9129-4396-e4de-25e041c6f6b8"
# Read in the cleaned data, before the CountVectorizer step
data_clean = pd.read_pickle('/content/drive/MyDrive/Colab Notebooks/NLP/data_clean.pkl')
data_clean
# + colab={"base_uri": "https://localhost:8080/"} id="I5q4zIocClN7" outputId="6a583d68-a43c-4afb-8f6f-36222aa5a62f"
import nltk
nltk.download('punkt')
# + colab={"base_uri": "https://localhost:8080/"} id="tQAS_sz2Cxeg" outputId="345d9a39-77ae-4093-c9a2-82c9c635a9ea"
nltk.download('averaged_perceptron_tagger')
# + colab={"base_uri": "https://localhost:8080/", "height": 425} id="--METIGL__Pn" outputId="51bd840e-0fc4-41dc-c001-30eeb61c9dc7"
# Apply the nouns function to the transcripts to filter only on nouns
data_nouns = pd.DataFrame(data_clean.transcript.apply(nouns))
data_nouns
# + colab={"base_uri": "https://localhost:8080/", "height": 475} id="_iRyb2uq__Po" outputId="6a92d179-14d1-47ae-b6fe-c3c35a2fc02f"
# Create a new document-term matrix using only nouns
from sklearn.feature_extraction import text
from sklearn.feature_extraction.text import CountVectorizer
# Re-add the additional stop words since we are recreating the document-term matrix
add_stop_words = ['like', 'im', 'know', 'just', 'dont', 'thats', 'right', 'people',
'youre', 'got', 'gonna', 'time', 'think', 'yeah', 'said']
stop_words = text.ENGLISH_STOP_WORDS.union(add_stop_words)
# Recreate a document-term matrix with only nouns
cvn = CountVectorizer(stop_words=stop_words)
data_cvn = cvn.fit_transform(data_nouns.transcript)
data_dtmn = pd.DataFrame(data_cvn.toarray(), columns=cvn.get_feature_names())
data_dtmn.index = data_nouns.index
data_dtmn
# + id="ikjYjqQc__Po"
# Create the gensim corpus
corpusn = matutils.Sparse2Corpus(scipy.sparse.csr_matrix(data_dtmn.transpose()))
# Create the vocabulary dictionary
id2wordn = dict((v, k) for k, v in cvn.vocabulary_.items())
# + colab={"base_uri": "https://localhost:8080/"} id="EKU-za3C__Pp" outputId="1ec176dd-c32e-4826-c530-6dbd22db7100"
# Let's start with 2 topics
ldan = models.LdaModel(corpus=corpusn, num_topics=2, id2word=id2wordn, passes=10)
ldan.print_topics()
# + colab={"base_uri": "https://localhost:8080/"} id="avmbYuWv__Pp" outputId="fa640d0a-029d-4904-81a0-d04b88be9dcf"
# Let's try topics = 3
ldan = models.LdaModel(corpus=corpusn, num_topics=3, id2word=id2wordn, passes=10)
ldan.print_topics()
# + colab={"base_uri": "https://localhost:8080/"} id="jh2SNxcK__Pp" outputId="0056fa6a-e24e-427b-b084-95c6c93bd02e"
# Let's try 4 topics
ldan = models.LdaModel(corpus=corpusn, num_topics=4, id2word=id2wordn, passes=10)
ldan.print_topics()
# + [markdown] id="8TiFZcL3__Pp"
# ## Topic Modeling - Attempt #3 (Nouns and Adjectives)
# + id="CzxOkZZF__Pp"
# Let's create a function to pull out nouns from a string of text
def nouns_adj(text):
'''Given a string of text, tokenize the text and pull out only the nouns and adjectives.'''
is_noun_adj = lambda pos: pos[:2] == 'NN' or pos[:2] == 'JJ'
tokenized = word_tokenize(text)
nouns_adj = [word for (word, pos) in pos_tag(tokenized) if is_noun_adj(pos)]
return ' '.join(nouns_adj)
# + colab={"base_uri": "https://localhost:8080/", "height": 425} id="VYaDvDKk__Pq" outputId="868dd56f-97d5-46f3-8d60-9e2a7b47c094"
# Apply the nouns function to the transcripts to filter only on nouns
data_nouns_adj = pd.DataFrame(data_clean.transcript.apply(nouns_adj))
data_nouns_adj
# + colab={"base_uri": "https://localhost:8080/", "height": 475} id="22e_9vXn__Pq" outputId="fdc8fb69-5dde-4d45-bbcb-5ce4d5ab09de"
# Create a new document-term matrix using only nouns and adjectives, also remove common words with max_df
cvna = CountVectorizer(stop_words=stop_words, max_df=.8)
data_cvna = cvna.fit_transform(data_nouns_adj.transcript)
data_dtmna = pd.DataFrame(data_cvna.toarray(), columns=cvna.get_feature_names())
data_dtmna.index = data_nouns_adj.index
data_dtmna
# + id="WXfuRIx9__Pq"
# Create the gensim corpus
corpusna = matutils.Sparse2Corpus(scipy.sparse.csr_matrix(data_dtmna.transpose()))
# Create the vocabulary dictionary
id2wordna = dict((v, k) for k, v in cvna.vocabulary_.items())
# + colab={"base_uri": "https://localhost:8080/"} id="ZADdfguV__Pq" outputId="3c086f05-4c49-49c4-8aa9-0a44396dd6c9"
# Let's start with 2 topics
ldana = models.LdaModel(corpus=corpusna, num_topics=2, id2word=id2wordna, passes=10)
ldana.print_topics()
# + colab={"base_uri": "https://localhost:8080/"} id="pyN52JHC__Pr" outputId="8e1ea0d0-4889-4a2c-db49-bcc3a9fe1a05"
# Let's try 3 topics
ldana = models.LdaModel(corpus=corpusna, num_topics=3, id2word=id2wordna, passes=10)
ldana.print_topics()
# + colab={"base_uri": "https://localhost:8080/"} id="k1c7Pfnp__Pr" outputId="49082e21-3353-4462-b9d0-61d51d0b2e20"
# Let's try 4 topics
ldana = models.LdaModel(corpus=corpusna, num_topics=4, id2word=id2wordna, passes=10)
ldana.print_topics()
# + [markdown] id="sIojwUY4__Pr"
# ## Identify Topics in Each Document
# + [markdown] id="D4kfDL6K__Pr"
# Out of the 9 topic models we looked at, the nouns and adjectives, 4 topic one made the most sense. So let's pull that down here and run it through some more iterations to get more fine-tuned topics.
# + colab={"base_uri": "https://localhost:8080/"} id="HgeTlb_O__Pr" outputId="f61fad42-3c78-442d-ba44-65657ba8bee8"
# Our final LDA model (for now)
ldana = models.LdaModel(corpus=corpusna, num_topics=4, id2word=id2wordna, passes=80)
ldana.print_topics()
# + [markdown] id="x6nvsdkA__Ps"
# These four topics look pretty decent. I'll on these for now.
# * Topic 0: mom, parents
# * Topic 1: husband, wife
# * Topic 2: guns
# * Topic 3: profanity
# + colab={"base_uri": "https://localhost:8080/"} id="KOqP75Aw__Ps" outputId="e59ce30c-adbb-4229-bc39-b2bf6bca0909"
# Let's take a look at which topics each transcript contains
corpus_transformed = ldana[corpusna]
list(zip([a for [(a,b)] in corpus_transformed], data_dtmna.index))
# + [markdown] id="20yLNb7R__Pt"
# For a first pass of LDA, these kind of make sense to me, for now.
# * Topic 0: mom, parents [<NAME>, Louis, Ricky]
# * Topic 1: husband, wife [<NAME>]
# * Topic 2: guns [Bill, Bo, Jim]
# * Topic 3: profanity [<NAME>]
|
Topic_Modeling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Instructions
#
# You work as a Data Analyst for a finance company which is closely eyeing the Android market before it launches its new app into Google Play. You have been asked to present an analysis of Google Play apps so that the team gets a comprehensive overview of different categories of apps, their ratings, and other metrics.
#
# This will require you to use your data manipulation and data analysis skills.
#
# Your three questions are as follows:
#
# 1. **Read the `apps.csv` file and clean the `Installs` column to convert it into integer data type.** Save your answer as a DataFrame `apps`. Going forward, you will do all your analysis on the `apps` DataFrame.
#
# 2. **Find the number of apps in each category, the average price, and the average rating.** Save your answer as a DataFrame `app_category_info`. Your should rename the four columns as: `Category`, `Number of apps`, `Average price`, `Average rating`.
#
# 3. **Find the top 10 free `FINANCE` apps having the highest average sentiment score.** Save your answer as a DataFrame `top_10_user_feedback`. Your answer should have exactly 10 rows and two columns named: `App` and `Sentiment Score`, where the average `Sentiment Score` is sorted from **highest to lowest**.
|
the_android_app_market_on_google_play/project_instructions_unguided.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# __Вопросы для повторения:__
# * зачем нужны исключения?
#
# * какие конструкторы-деструкторы здесь будут вызваны?
#
# ```c++
# class M { ... };
#
# class D {
# public:
# D() { throw std::runtime_error("error"); }
# M m_;
# };
#
# D d;
# ```
#
# * откуда в `collect_team` может быть брошено исключение?
#
# ```c++
# const Dwarf& find_dwarf(int id) noexcept;
#
# std::vector<int> get_dwarfs_ids();
#
# auto collect_team()
# {
# const auto ids = get_dwarfs_ids();
#
# std::vector<Dwarf> dwarfs;
# for (int id : ids)
# dwarfs.push_back(find_dwarf(id));
#
# std::puts("dream team:");
# for (const auto drawf : dwarfs)
# std::cout << dwarf.name << '\n';
#
# return dwarfs;
# }
# ```
#
# * почему move-операции желательно делать `noexcept`?
# * исключения и деструкторы?
|
2019/sem1/lecture8_smart_pointers/repetition.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
star_wars = pd.read_csv("star_wars.csv", encoding="ISO-8859-1")
star_wars.head(3)
# get rows without nulls
star_wars = star_wars[star_wars["RespondentID"].notnull()].copy()
# reset index to 0...n
star_wars = star_wars.reset_index(drop=True)
star_wars.head(3)
star_wars["Have you seen any of the 6 films in the Star Wars franchise?"].value_counts()
# +
yes_no = {
"Yes": True,
"No": False,
}
star_wars["Have you seen any of the 6 films in the Star Wars franchise?"] = star_wars["Have you seen any of the 6 films in the Star Wars franchise?"].map(yes_no)
star_wars["Do you consider yourself to be a fan of the Star Wars film franchise?"] = star_wars["Do you consider yourself to be a fan of the Star Wars film franchise?"].map(yes_no)
# -
star_wars["Do you consider yourself to be a fan of the Star Wars film franchise?"].value_counts()
star_wars.columns[3:9]
star_wars["Unnamed: 8"].head()
col_film_map = {
"Which of the following Star Wars films have you seen? Please select all that apply.": "Star Wars: Episode I The Phantom Menace",
"Unnamed: 4": "Star Wars: Episode II Attack of the Clones",
"Unnamed: 5": "Star Wars: Episode III Revenge of the Sith",
"Unnamed: 6": "Star Wars: Episode IV A New Hope",
"Unnamed: 7": "Star Wars: Episode V The Empire Strikes Back",
"Unnamed: 8": "Star Wars: Episode VI Return of the Jedi"
}
for col in star_wars.columns[3:9]:
film_map = {
col_film_map[col]: True,
np.NaN: False
}
print(film_map)
star_wars[col] = star_wars[col].map(film_map)
star_wars.iloc[:,3:9].isnull().values.any()
rename_map = {
"Which of the following Star Wars films have you seen? Please select all that apply.": "seen_1",
"Unnamed: 4": "seen_2",
"Unnamed: 5": "seen_3",
"Unnamed: 6": "seen_4",
"Unnamed: 7": "seen_5",
"Unnamed: 8": "seen_6"
}
star_wars = star_wars.rename(columns=rename_map)
star_wars.iloc[:,3:9].head()
star_wars[star_wars.columns[9:15]] = star_wars[star_wars.columns[9:15]].astype(float)
star_wars.info()
rename_map = {
"Please rank the Star Wars films in order of preference with 1 being your favorite film in the franchise and 6 being your least favorite film.": "ranking_1",
"Unnamed: 10": "ranking_2",
"Unnamed: 11": "ranking_3",
"Unnamed: 12": "ranking_4",
"Unnamed: 13": "ranking_5",
"Unnamed: 14": "ranking_6"
}
star_wars = star_wars.rename(columns=rename_map)
star_wars.info()
rank_cols = ["ranking_{}".format(x) for x in range(1,7)]
star_wars[rank_cols].mean()
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
# +
fig = plt.figure(figsize=(10,6))
ax = fig.add_subplot(1,1,1)
width = 0.45 # the width of the bars
x = np.arange(6) # the x axis values, just a range from 1 to 6
ax.bar(x, star_wars[rank_cols].mean().tolist(), width, align='center')
# add some text for labels, title and axes ticks
ax.set_xlabel('Star Wars Episode')
ax.set_ylabel('Mean Ranking')
ax.set_title('Star Wars Film Rankings')
ax.set_xticks(x)
ax.set_xticklabels(x + 1)
plt.show()
# +
# how many people saw each movie?
seen_cols = ["seen_{}".format(x) for x in range(1,7)]
star_wars[seen_cols].sum()
# +
fig = plt.figure(figsize=(10,6))
ax = fig.add_subplot(1,1,1)
width = 0.45 # the width of the bars
x = np.arange(6) # the x axis values, just a range from 1 to 6
ax.bar(x, star_wars[seen_cols].sum().tolist(), width, align='center')
# add some text for labels, title and axes ticks
ax.set_xlabel('Star Wars Episode')
ax.set_ylabel('People')
ax.set_title('Star Wars Viewers')
ax.set_xticks(x)
ax.set_xticklabels(x + 1)
plt.show()
# -
ranks = star_wars[rank_cols].mean()
seens = star_wars[seen_cols].sum()
ranks.index = ["episode_{}".format(x) for x in range(1,7)]
ranks.index
seens.index = ["episode_{}".format(x) for x in range(1,7)]
seens.index
rank_seen = pd.DataFrame({"rank": ranks,
"seen": seens})
rank_seen
rank_seen.plot(kind="bar", figsize=(10,6), secondary_y="seen")
plt.show()
trekkies = star_wars[star_wars["Do you consider yourself to be a fan of the Star Trek franchise?"] == "Yes"]
non_trekkies = star_wars[star_wars["Do you consider yourself to be a fan of the Star Trek franchise?"] == "No"]
# +
ranks_trekkies = trekkies[rank_cols].mean()
ranks_trekkies.index = ["episode_{}".format(x) for x in range(1,7)]
seens_trekkies = trekkies[seen_cols].sum()
seens_trekkies.index = ["episode_{}".format(x) for x in range(1,7)]
# -
ranks_trekkies_vs_all = pd.DataFrame({"trekkies": ranks_trekkies, "all": ranks})
ranks_trekkies_vs_all
ranks_trekkies_vs_all.plot(kind="bar", figsize=(10,6), secondary_y="all")
plt.show()
# +
seen_trekkies_norm = seens_trekkies / seens_trekkies.sum()
seens_norm = seens / seens.sum()
print(seen_trekkies_norm.sum())
print(seens_norm.sum())
seens_trekkies_vs_all = pd.DataFrame({"trekkies": seen_trekkies_norm, "all": seens_norm})
seens_trekkies_vs_all
# -
seens_trekkies_vs_all.plot(kind="bar", figsize=(10,6), secondary_y="all")
plt.show()
fans = star_wars[star_wars["Do you consider yourself to be a fan of the Star Wars film franchise?"] == True]
non_fans = star_wars[star_wars["Do you consider yourself to be a fan of the Star Wars film franchise?"] == False]
# +
ranks_fans = fans[rank_cols].mean()
ranks_fans.index = ["episode_{}".format(x) for x in range(1,7)]
ranks_non_fans = non_fans[rank_cols].mean()
ranks_non_fans.index = ["episode_{}".format(x) for x in range(1,7)]
seens_fans = fans[seen_cols].sum()
seens_fans.index = ["episode_{}".format(x) for x in range(1,7)]
# -
ranks_fans_vs_all = pd.DataFrame({"fans": ranks_fans, "all": ranks})
ranks_fans_vs_all
ranks_fans_vs_all.plot(kind="bar", figsize=(10,6), secondary_y="all")
plt.show()
ranks_non_fans_vs_all = pd.DataFrame({"non_fans": ranks_non_fans, "all": ranks})
ranks_non_fans_vs_all
ranks_non_fans_vs_all.plot(kind="bar", figsize=(10,6), secondary_y="all")
plt.show()
|
star_wars/star_wars.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## This Notebook contains:
# - Reading the datset from Northeastern university
# - Modelling for classification models
# - Calculation of Error metrices
# - Summarizing Models at the end
#
# Required Python Packages
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.svm import SVC
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
from sklearn.utils import resample
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import KFold
import pickle
import matplotlib.pyplot as plt
import numpy as np
from sklearn.exceptions import DataConversionWarning
import warnings
warnings.filterwarnings(action='ignore', category=DataConversionWarning)
# Importing the dataset
dataset = pd.read_csv('../../data/all_records_northeastern.csv')
# Dropping the column, Unnamed as it is not necessary
dataset.drop(columns=['Unnamed: 0'],inplace=True)
dataset.head()
# Count of accept and reject in dataset
dataset.status.value_counts()
# As we see from above stats, our data is baised so we need to resample the data, in order to balance the dataset
# +
balanced_data=resample(dataset[dataset.status=='accept'],replace=True,n_samples=1000,random_state=123)
balanced_data=balanced_data.append(dataset[dataset.status=='reject'])
# -
balanced_data.status.value_counts()
encoded_dataset=balanced_data
encoded_dataset.head(2)
# As we see from above stats, our data is baised so we need to resample the data, in order to balance the dataset
X=encoded_dataset[['gre_score_quant','gre_score_verbal','test_score_toefl','undergraduation_score','work_ex','papers_published']].copy()
Y=encoded_dataset[['status']].copy()
# Split the dataset into test and train data
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.2,random_state=1)
# Training the model
def modeltraining(model,X_train,X_test,Y_train,Y_test):
sc = StandardScaler()
sc.fit(X_train)
X_train = sc.transform(X_train)
X_test = sc.transform(X_test)
model.fit(X_train,Y_train)
predicted_labels_test=model.predict(X_test)
predicted_labels_train=model.predict(X_train)
accuracy_test=accuracy_score(Y_test,predicted_labels_test)
accuracy_train=accuracy_score(Y_train,predicted_labels_train)
return model,predicted_labels_test,predicted_labels_train,accuracy_test,accuracy_train,sc
# Calling Support vector classifier
model=SVC(kernel='linear')
svclassifier,Y_Pred_Test,Y_Pred_Train,accuracyTest,accuracyTrain,sc=modeltraining(model,X_train,X_test,Y_train,Y_test)
# Accuracy of test data
accuracyTest
# Accuracy of train data
accuracyTrain
svclassifier
# Confusion matrix:
print(confusion_matrix(Y_test,Y_Pred_Test))
# Classification report:
print(classification_report(Y_test,Y_Pred_Test))
print(classification_report(Y_train,Y_Pred_Train))
# Hypertuning the parameters using grid search, using c, degree and class_weight
# +
kf = KFold(n_splits=5)
kf.get_n_splits(X)
param_grid = {"C":[1],
"degree":[3,4,5],
"class_weight":['balanced']
}
# run grid search
grid_search = GridSearchCV(svclassifier, param_grid, cv=5,return_train_score=True)
svclassifier,Y_Pred_Test,Y_Pred_Train,accuracyTest,accuracyTrain,sc=modeltraining(grid_search,X_train,X_test,Y_train,Y_test)
# -
# Accuracy of train data after grid search
accuracyTest
# Accuracy of test data after grid search
accuracyTrain
# Best estimator after grid search
grid_search.best_estimator_
y_test_list = Y_test['status'].tolist()
y_test_list_new = []
for i in y_test_list:
if i == 'accept':
y_test_list_new.append(0)
elif i == 'reject':
y_test_list_new.append(1)
y_pred_test_list = Y_Pred_Test.tolist()
y_test_list_new = []
for i in y_pred_test_list:
if i == 'accept':
y_pred_test_list_new.append(0)
elif i == 'reject':
y_pred_test_list_new.append(1)
len(y_test_list_new)
plt.figure(figsize=(40,10))
plt.bar(np.arange(0,100,1),y_pred_test_list_new[:100],y_test_list_new[:100])
#plt.plot(np.arange(0,100,1),y_pred_test_list_new[:100],color ='r')
# F1 score for Test and Train:
# <table style="width:100%">
# <tr>
# <th>Model</th>
# <th>Test Accuracy</th>
# <th>Train Accuracy</th>
# <th>Grid Search - Test Accuracy</th>
# <th>Grid Search - Test Accuracy</th>
# <th>Test F1 Score</th>
# <th>Train F1 Score</th>
# <th>Best Parameter</th>
# <th>Interpretability</th>
# <th>Reproducability</th>
# </tr>
# <tr>
# <th>Model-SupportVectorMachine</th>
# <th>0.6658</th>
# <th>0.6844</th>
# <th>Grid Search Test - 0.6686</th>
# <th>Grid Search Train - 0.6944</th>
# <th>0.66 </th>
# <th>0.65</th>
# <th>(C=1, cache_size=200, class_weight='balanced', coef0=0.0,
# decision_function_shape='ovr', degree=3, gamma='auto_deprecated',
# kernel='linear', max_iter=-1, probability=False, random_state=None,
# shrinking=True, tol=0.001, verbose=False)</th>
# <th>Non-Interpretable</th>
# <th>Non-reproducable</th>
# </tr>
#
# </table>
|
Final Project _ Graduate Admission Predictor/Code/Modelling/UniversitySupportVectorMachine.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # BioSentVec Tutorial
# This tutorial provides a fundemental introduction to our BioSentVec models. It illustrates (1) how to load the model, (2) an example function to preprocess sentences, (3) an example application that uses the model and (4) further resources for using the model more broadly.
# ## 1. Prerequisites
# Please download BioSentVec model and install all the related python libraries
# + tags=[]
import sent2vec
from nltk import word_tokenize
from nltk.corpus import stopwords
from string import punctuation
from scipy.spatial import distance
import pickle
from tqdm import tqdm
import numpy as np
# + tags=[]
import nltk
nltk.download('stopwords')
nltk.download('punkt')
# -
# ## 2. Load BioSentVec model
# Please specify the location of the BioSentVec model to model_path. It may take a while to load the model at the first time.
#
# Get the model here: https://github.com/ncbi-nlp/BioSentVec
# + tags=[]
model_path = '/home/thetaphipsi/Downloads/BioSentVec_PubMed_MIMICIII-bigram_d700.bin'
model = sent2vec.Sent2vecModel()
try:
model.load_model(model_path)
except Exception as e:
print(e)
print('model successfully loaded')
# -
# ## 3. Preprocess sentences
# There is no one-size-fits-all solution to preprocess sentences. We demonstrate a representative code example as below. This is also consistent with the preprocessing appaorach when we trained BioSentVec models.
stop_words = set(stopwords.words('english'))
def preprocess_sentence(text):
text = text.replace('/', ' / ')
text = text.replace('.-', ' .- ')
text = text.replace('.', ' . ')
text = text.replace('\'', ' \' ')
text = text.lower()
tokens = [token for token in word_tokenize(text) if token not in punctuation and token not in stop_words]
return ' '.join(tokens)
# An example of using the preprocess_sentence function:
sentence = preprocess_sentence('Breast cancers with HER2 amplification have a higher risk of CNS metastasis and poorer prognosis.')
print(sentence)
# ## 4. Retrieve a sentence vector
# Once a sentence is preprocessed, we can pass it to the BioSentVec model to retrieve a vector representation of the sentence.
sentence_vector = model.embed_sentence(sentence)
print(sentence_vector)
# Note that you can also use embed_sentences to retrieve vector representations of multiple sentences.
# The shape of the vector representation depends on the dimension parameter. In this case, we set the dimension to 700:
print(sentence_vector.shape)
# ## 5. Compute sentence similarity
# In this section, we demonstrate how to compute the sentence similarity between a sentence pair using the BioSentVec model. We firstly use the above code examples to get vector representations of sentences. Then we compute the cosine similarity between the pair.
# +
sentence_vector1 = model.embed_sentence(preprocess_sentence('Breast cancers with HER2 amplification have a higher risk of CNS metastasis and poorer prognosis.'))
sentence_vector2 = model.embed_sentence(preprocess_sentence('Breast cancers with HER2 amplification are more aggressive, have a higher risk of CNS metastasis, and poorer prognosis.'))
cosine_sim = 1 - distance.cosine(sentence_vector1, sentence_vector2)
print('cosine similarity:', cosine_sim)
# -
# Here is another example for a pair that is relatively less similar.
sentence_vector3 = model.embed_sentence(preprocess_sentence('Furthermore, increased CREB expression in breast tumors is associated with poor prognosis, shorter survival and higher risk of metastasis.'))
cosine_sim = 1 - distance.cosine(sentence_vector1, sentence_vector3)
print('cosine similarity:', cosine_sim)
# # more experiments with MIMIC-III data
text = "The patient is a 46 year old gentleman with past medical history including end stage HIV, placement on Hospital. Patient also has past medical history significant for polysubstance abuse including cocaine, hepatitis C and cirrhosis. Patient was transferred to from an outside hospital for recurrent v-fib arrest in the setting of having an ICD placed one week prior to admission. Patient was found down on the morning of admission by a neighbor. unresponsive, called EMS. When EMS came, patient remained unresponsive, but his vital signs were stable. He had a paced rhythm. He had normal blood pressure and normal oxygen saturation. Patient was then transferred to an outside hospital in . While there his ICD discharged at least two times for episodes of ventricular tachycardia. Rhythm strips from the outside hospital were not available at the time of admission. Per report patient was given amiodarone 150 mg IV three times and he was given magnesium sulfate 4 gm IV for magnesium level of 1.6. Med-Flight then transferred patient to as CCU beds were not available elsewhere. En route to the hospital patient experienced another two episodes of v-fib and his ICD discharged each time, converting patient back into paced rhythm. During the shocks patient was reported to be alert. He denied chest pain, shortness of breath, had stable blood pressure and stable oxygen saturation. On arrival to the CCU patient experienced two shocks from his ICD for torsades. Amiodarone was continued at 1 mg per minute. Patient went into v-fib arrest two times just after his admission. Again the ICD converted him back into a stable paced rhythm. Again his sats and blood pressure remained stable with these episodes.|HIV. Last CD4 count 438, last viral load less than 50 copies per ml from his outpatient infectious disease physician's notes in his medical record. These levels were from . HIV course is complicated by nonischemic cardiomyopathy and also complicated by candidal esophagitis. Hepatitis C and cirrhosis presumed secondary to IV drug abuse. Hep A positive as well. Nonischemic cardiomyopathy with EF of 20%. Patient had cardiac cath in that was negative for coronary artery disease. Patient is also status post ICD placement on Hospital. Patient is status post DDD pacer placed four years ago for complete heart block. Polysubstance abuse including crack cocaine, heroin and alcohol. Patient stated on admission that he had been using crack cocaine for the week prior to admission. Most recent use was three days prior to admission. Chronic renal insufficiency. Uncertain of baseline creatinine. Old hospital records from indicate that the renal failure was secondary to diuretic over-use. Peripheral neuropathy. B-12 deficiency and macrocytic anemia. Stasis dermatitis and eosinophilic folliculitis. Asthma.|Abacavir 300 mg b.i.d., saquinavir 400 mg b.i.d., Combivir tabs one b.i.d., Norvir|00 mg b.i.d., Protonix 40 q.day, trazodone 100 q.h.s., lisinopril 10 b.i.d., Lasix 80 b.i.d., methadone 250 mg q.d., Lexapro 10 mg q.d., digoxin 125 mcg q.d., Bactrim one double strength q.d., Zyrtec 10 q.d., sublingual nitroglycerin p.r.n., triamcinolone cream, albuterol inhaler p.r.n.|The patient denied any recent illness. He states that since the ICD had been placed, he has felt fatigued, but denies chest pain, shortness of breath. He does admit to recent crack cocaine use, most recently three days prior to admission.|The patient states that he was clean and sober for three years until this past week. He states that the stress of his illness and his mother's illness, who is bed bound status post CVA, had prompted this use. Patient lives alone and states that he has been very depressed at home and felt that his family avoids him because of his HIV status and feels that he cannot take care of himself appropriately. His sister and his niece do look in on him frequently, however. Per report of his sister and his niece, his house is disordered and they comment that patient is barely functional. Patient has a history of smoking, alcohol abuse and cocaine use as stated previously.|Vital signs on admission included temperature of 98.2, pulse 66, blood pressure 104 to 130 over|4 to 65, respiratory rate 14, sat 100% on 10 liters nasal cannula, weight 74 kg. In general, patient was alert and oriented times two, alert to person and place, but not time. He denied any distress. He appeared disheveled, but otherwise did not appear in any discomfort. HEENT pupils equal, round and reactive to light and accommodation. Oropharynx clear without lesions. Heart regular rate and rhythm, S1, S2, prominent S3, to holosystolic murmur at the left upper and left lower sternal borders, does not radiate to carotids, does not radiate to axillae. PMI laterally displaced. Right ventricular heave. JVP markedly elevated at 14 cm. Lungs bibasilar crackles, left greater than right. Abdominal exam soft, nondistended, nontender, hepatomegaly 3 cm beneath the costal margin. Extremities no cyanosis, clubbing or edema with good distal pulses. Neuro cranial nerves II-XII intact. Strength grossly normal. Sensation grossly normal.|From the outside hospital data included magnesium of 1.6. Tox screen included serum, ethanol, salicylates, acetaminophen and tricyclic levels which were all negative. Digoxin level was normal at 0.85, normal being|.9 to 2.0. CK 226, MB 3.3, troponin I 0.1. Hematocrit|0.8, white count 5.4, platelets 155. Chem-7 132, 4.0, 92,|6, 16, 1.5, 138. Calcium 8.2. Chest x-ray from the outside hospital showed cardiomegaly with clear lung fields. Head CT at the outside hospital, performed because patient had an unwitnessed fall, was read at the outside hospital as possible evolving left parietal infarct with no evidence of bleed. However, this head CT was read by our radiologists as no evidence of stroke or hemorrhage. EKG showed wide complex rhythm of left bundle morphology, rate 100 with paced beat preceded by a P and followed by a nonpaced ventricular beat, QTc 433. Strips from Med-Flight consistent with polymorphic VT of torsade type. Echo one week prior to admission from outside hospital showed moderate MR , left ventricular hypokinesis, EF 20%. Lab data from admission to hematocrit 32.8, white count 4.7, platelets 159. Chemistry sodium 133, potassium 4.8, chloride 96, bicarb 26, BUN 20, creatinine 1.2, glucose 148. Calcium 8.6, mag 2.6, phos 4.9, albumin 3.6. Serum tox was negative for benzo, negative for barbiturates. Urine tox was performed that was positive for cocaine and positive for methadone. UA was negative for UTI.|1. Cardiology. Rhythm. For patient's recurrent torsades he was initially continued on an amio drip at 1 mg per minute. As the QT was prolonged, patient was switched from amio to a lidocaine drip as lidocaine decreases the QT interval while amio increases the QT interval. Patient's pacer was interrogated on the night of admission and reprogrammed. On interrogation it was revealed that patient had 16 episodes of v-fib with heart rate between 200 to 230 beats per minute, requiring 17 joules and then 31 joules to convert patient back into a paced rhythm. His pacer was a DDD that was originally set for 50 to 120 beats per minute with paced AV at 180 msec. The mode was changed by the EP Fellow to DDD at|0 to 120 beats per minute with the thinking that increasing the rate of pacing would shorten the QT interval and hopefully prevent recurrent torsade. The pacer was also changed so that the VF zone first shock to patient at 31 joules rather than 17 joules as patient was requiring 31 joules of energy to convert him back to a paced rhythm. Patient's magnesium was checked frequently and magnesium after repletion was stable at 2.6. The patient was kept on a lidocaine drip from admission until 5:00 a.m. on the morning of . At this time patient began to experience visual hallucinations and became slightly agitated. Therefore, the lidocaine drip was decreased from 3 mg per minute to 1 mg per minute. However, as patient's agitation continued even with the decreased dose of lidocaine, the lidocaine drip was stopped at 7:00 a.m. Patient was switched to mexiletine 100 mg p.o. b.i.d. as this is also a class Ib antiarrhythmic. Patient tolerated this medication much better and his neurological status cleared. He no longer had visual hallucinations and he tolerated the mexiletine dose quite well. Patient was also started on labetalol 100 mg b.i.d. This beta blocker was chosen as patient's urine tox was positive for cocaine. He also tolerated this medication well. Patient had no repeat episodes of ventricular tachycardia or ventricular fibrillation during his hospital course.| 17:18 T: 12:39 JOB#: "
# +
sentence_vector1 = model.embed_sentence(preprocess_sentence(text[:100]))
sentence_vector2 = model.embed_sentence(preprocess_sentence(text[:500]))
cosine_sim = 1 - distance.cosine(sentence_vector1, sentence_vector2)
print('cosine similarity:', cosine_sim)
# +
import pickle
train_data = pickle.load(open('/Users/jplasser/Documents/AI Master/WS2021/MastersThesis/code.nosync/CNEP/src/data/mimic3/full_train_data_unique.pickle', 'rb'))
#val_data = pickle.load(open('/home/thetaphipsi/MasterAI/src/CNEP/src/data/mimic3/full_val_data_unique.pickle', 'rb'))
#test_data = pickle.load(open('/home/thetaphipsi/MasterAI/src/CNEP/src/data/mimic3/full_test_data_unique.pickle', 'rb'))
# +
import seaborn as sns
import numpy as np
import torch
def plot_similarity(labels, features, rotation, print_labels=True):
print(f"{features.shape=}")
corr = np.inner(features, features)
#print(corr)
labels = [m[:25] + '/' + str(len(m)) for m in labels]
sns.set(rc = {'figure.figsize':(20,12)})
sns.set(font_scale=1.2)
g = sns.heatmap(corr,
xticklabels=labels,
yticklabels=labels,
vmin=0,
vmax=1,
annot=print_labels, fmt='.1f',
cmap="YlOrRd")
g.set_xticklabels(labels, rotation=rotation)
g.set_title("Semantic Textual Similarity")
def run_and_plot(messages_, seq_len):
message_embeddings_ = torch.stack([torch.nn.functional.normalize(
torch.tensor(
model.embed_sentence(
preprocess_sentence(
m[:seq_len] + m[-seq_len:]
)
)
)
) for m in messages_])
message_embeddings_.shape
message_embeddings_ = message_embeddings_.reshape(len(messages_),-1)
plot_similarity(messages_, message_embeddings_, 90)
# +
import random
n = 20
idx = random.sample(list(np.arange(len(train_data['notes']))), n)
print(idx)
#messages = train_data['notes'][:-1:len(train_data['notes'])//10]
messages = train_data['notes'][idx]
run_and_plot(messages, 2000)
# -
text[:20], text[-20:]
np.arange(len(train_data['notes']))
# +
from tqdm import tqdm
embeds = []
seq_len = 2000
text1 = train_data['notes'][0]
max_cosine = 0.
best_idx = 0
sentence_vector1 = model.embed_sentence(preprocess_sentence(text1[:seq_len]))
sims = []
embeds.append(sentence_vector1)
for i in tqdm(range(len(train_data['notes'])-1)):
text2 = train_data['notes'][i+1]
sentence_vector2 = model.embed_sentence(preprocess_sentence(text2[:seq_len]))
cosine_sim = 1 - distance.cosine(sentence_vector1, sentence_vector2)
sims.append(cosine_sim)
embeds.append(sentence_vector2)
if cosine_sim > max_cosine:
max_cosine = cosine_sim
best_idx = i+1
print(f"cosine similarity {max_cosine} with index {best_idx}.")
# -
np.argmax(sims)+1, np.argmin(sims)+1
sns.histplot(sims, kde=True)
# +
text1 = train_data['notes'][0]
text2 = train_data['notes'][best_idx]
sentence_vector1 = embeds[0] #model.embed_sentence(preprocess_sentence(text1))
sentence_vector2 = embeds[best_idx] #model.embed_sentence(preprocess_sentence(text2))
cosine_sim = 1 - distance.cosine(sentence_vector1, sentence_vector2)
print(f"cosine similarity {cosine_sim} with index {best_idx}.")
# +
text1 = train_data['notes'][0]
text2 = train_data['notes'][best_idx]
sims_se = []
for i in tqdm(range(50,2000)):
sentence_vector1 = model.embed_sentence(preprocess_sentence(text1[:i+1]))
sentence_vector2 = model.embed_sentence(preprocess_sentence(text2[:i+1]))
cosine_sim = 1 - distance.cosine(sentence_vector1, sentence_vector2)
sims_se.append(cosine_sim)
#print(f"cosine similarity {cosine_sim} with index {best_idx}.")
# -
sims[1]
#sns.histplot(sims)
sns.scatterplot(x=sims_se, y=range(50,2000))
a = np.array(sims)
np.where(np.logical_and(a>=0.88, a<=1.))
# +
worst_idx = np.argmin(sims)+1
text1 = train_data['notes'][0]
text2 = train_data['notes'][worst_idx]
sentence_vector1 = embeds[0] #model.embed_sentence(preprocess_sentence(text1))
sentence_vector2 = embeds[worst_idx] #model.embed_sentence(preprocess_sentence(text2))
cosine_sim = 1 - distance.cosine(sentence_vector1, sentence_vector2)
print(f"cosine similarity {cosine_sim} with index {worst_idx}.")
# +
text1 = train_data['notes'][0]
text2 = train_data['notes'][worst_idx]
sims_se = []
for i in tqdm(range(50,seq_len)):
sentence_vector1 = model.embed_sentence(preprocess_sentence(text1[:i+1]))
sentence_vector2 = model.embed_sentence(preprocess_sentence(text2[:i+1]))
cosine_sim = 1 - distance.cosine(sentence_vector1, sentence_vector2)
sims_se.append(cosine_sim)
#print(f"cosine similarity {cosine_sim} with index {best_idx}.")
# -
sns.scatterplot(x=sims_se, y=range(50,seq_len))
a = np.array(sims_se)
np.where(np.logical_and(a>=0.21, a<=0.2169094979763031))
# # generate data sets with embeds included
# + tags=[]
seq_len = 2000 # original sequence len was 2000
#data_path = '/Users/jplasser/Documents/AI Master/WS2021/MastersThesis/code.nosync/CNEP/src/data/mimic3/'
data_path = '/home/thetaphipsi/MasterAI/src/CNEP/src/data/mimic3/'
datasets = ['train','val','test']
for dataset in datasets:
embeds = []
train_data = pickle.load(open(f'{data_path}full_{dataset}_data_unique.pickle', 'rb'))
for i in tqdm(range(len(train_data['notes']))):
sentence_vector = model.embed_sentence(preprocess_sentence(train_data['notes'][i])) #[:seq_len]))
embeds.append(sentence_vector.reshape(-1))
embeds = np.array(embeds)
print(train_data['inputs'].shape, embeds.shape)
train_data['embeds'] = embeds
pickle.dump(train_data, open(f'{data_path}full_{dataset}_data_unique_embed_fullsequence.pickle', 'wb'))
# -
# ## 6. More resources
# The above example demonstrates an unsupervised way to use the BioSentVec model. In addition, we summarize a few useful resources:
#
# #### (1) The Sent2vec homepage (https://github.com/epfml/sent2vec) has a few pre-trained sentence embeddings from general English copora.
# #### (2) You can also develop deep learning models to learn sentence similarity in a supervised manner.
# #### (3) You can also use the BioSentVec in other applications, such as multi-label classification.
# ## Reference
# When using some of our pre-trained models for your application, please cite the following paper:
#
# <NAME>, <NAME>, <NAME>. BioSentVec: creating sentence embeddings for biomedical texts. 2018. arXiv:1810.09302.
|
src/notebooks/BioSentVec_tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # maps.ipynb
#
# Display interactive maps of the most recent COVID-19 statistics with and without normalization by population.
#
# Inputs:
# * `data/us_counties_clean.csv`: The contents of `data/us_counties.csv` after data cleaning by `clean.ipynb`
# * `data/us_counties_clean_meta.json`: Column type metadata for reading `data/us_counties_clean.csv` with `pd.read_csv()`
# * [U.S. map in GeoJSON format, from Plotly](https://raw.githubusercontent.com/plotly/datasets/master/geojson-counties-fips.json)
# +
# Initialization boilerplate
import json
import pandas as pd
import numpy as np
from urllib.request import urlopen
from typing import *
import text_extensions_for_pandas as tp
# Local file of utility functions
import util
# +
# Read in the CSV file and apply the saved type information
csv_file = "data/us_counties_clean.csv"
meta_file = "data/us_counties_clean_meta.json"
# Read column type metadata
with open(meta_file) as f:
cases_meta = json.load(f)
# Pandas does not currently support parsing datetime64 from CSV files.
# As a workaround, read the "Date" column as objects and manually
# convert after.
cases_meta["Date"] = "object"
cases_vertical = pd.read_csv(csv_file, dtype=cases_meta, parse_dates=["Date"])
cases_vertical
# -
# Slice off the last element of each time series.
cases = cases_vertical[cases_vertical["Date"] == cases_vertical["Date"].max()].set_index("FIPS")
cases
# +
# Normalize the Confirmed and Deaths counts by population.
cases["confirmed_per_100"] = cases["Confirmed"] / cases["Population"] * 100
cases["deaths_per_100"] = cases["Deaths"] / cases["Population"] * 100
cases
# -
# Download a U.S. map in GEOJSON format
with urlopen('https://raw.githubusercontent.com/plotly/datasets/master/geojson-counties-fips.json') as response:
counties = json.load(response)
# +
# Common code to generate choropleth maps.
# NOTE: In order for this to work you need the JupyterLab extensions for Plotly:
# > jupyter labextension install jupyterlab-plotly
# (env.sh will run the above command for you)
import plotly.express as px
def draw_map(col_name, label_str):
# Each series may have NAs in different locations
valid_data = cases[~cases[col_name].isna()]
fig = px.choropleth(valid_data, geojson=counties,
locations=["{:05d}".format(f) for f in valid_data.index],
color=col_name,
# See https://plotly.com/python/builtin-colorscales/
color_continuous_scale="viridis",
# Top of scale == 95th percentile
range_color=(0, valid_data[col_name].quantile(0.95)),
scope="usa",
labels={col_name: label_str},
hover_name=valid_data["County"],
title=label_str)
fig.update_layout(margin={"r":0,"t":0,"l":0,"b":0})
fig.show()
# -
# Draw a map of number of confirmed cases by county.
# Yellow == 95th percentile
draw_map("Confirmed", "Confirmed Cases ")
# Draw a map of number of confirmed cases per 100 residents by county
# Yellow == 95th percentile
draw_map("confirmed_per_100", "Confirmed per 100")
# Draw a map of number of deaths by county
# Yellow == 95th percentile
draw_map("Deaths", "Total Deaths ")
# Draw a map of number of deaths per 100 residents by county
# Yellow == 95th percentile
draw_map("deaths_per_100", "Deaths per 100")
|
maps.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: flow3.7
# language: python
# name: flow3.7
# ---
import tensorflow as tf
import tensorflow_probability as tfp
from tensorflow.keras import Input
from tensorflow.keras.layers import Dense
import numpy as np
input_dim, action_dim, fcnet_hiddens = 24, 1, [32, 32, 32]
def build_neural_net_stochastic(input_dim, action_dim, fcnet_hiddens):
"""Build a keras model to output a stochastic policy.
Parameters
----------
input_dim : int
dimension of input layer
action_dim : int
action_space dimension
fcnet_hiddens : list
list containing size of each hidden layer (length of list is number of hidden layers)
Returns
-------
Keras model (untrained)
"""
input_layer = Input(shape=(input_dim, ))
curr_layer = input_layer
for i in range(len(fcnet_hiddens)):
size = fcnet_hiddens[i]
dense = Dense(size, activation="tanh")
curr_layer = dense(curr_layer)
out = Dense(2 * action_dim, activation=None)(curr_layer)
model = tf.keras.Model(inputs=input_layer, outputs=out, name="policy_network")
return model
# +
def get_loss(stochastic, variance_regularizer):
"""Get appropriate loss function for training.
Parameters
----------
stochastic : bool
determines if policy to be learned is deterministic or stochastic
variance_regularizer : float
regularization hyperparameter to penalize high variance policies
Returns
-------
Keras loss function to use for imitation learning.
"""
if stochastic:
return negative_log_likelihood_loss(variance_regularizer)
else:
return tf.keras.losses.mean_squared_error
def negative_log_likelihood_loss(variance_regularizer):
"""Negative log likelihood loss for learning stochastic policies.
Parameters
----------
variance_regularizer : float
regularization hyperparameter to penalize high variance policies
Returns
-------
Negative log likelihood loss function with variance regularization.
"""
def nll_loss(y, network_output):
assert network_output.shape[1] % 2 == 0, "Stochastic policies must output vectors of even length"
action_dim = network_output.shape[1] // 2
# first half of network_output is mean, second half is log_std
means, log_stds = network_output[:, :action_dim], network_output[:, action_dim:]
stds = tf.math.exp(log_stds)
variances = tf.math.square(stds)
# Multivariate Gaussian distribution
dist = tfp.distributions.MultivariateNormalDiag(loc=means, scale_diag=variances)
loss = dist.log_prob(y)
loss = tf.negative(loss)
loss = tf.reduce_mean(loss) + (variance_regularizer * tf.norm(variances))
return loss
return nll_loss
# -
def compile_network(model):
"""
Compiles Keras network with appropriate loss and optimizer
"""
loss = get_loss(True, 10)
model.compile(loss=loss, optimizer='adam')
model = build_neural_net_stochastic(input_dim, action_dim, fcnet_hiddens)
compile_network(model)
# model
action_batch = np.random.rand(600, 1)
action_batch = action_batch.reshape(action_batch.shape[0], action_dim)
observation_batch = np.random.rand(600, 24)
# ### Error 1
model.train_on_batch(observation_batch, action_batch, sample_weight=np.random.rand(600,))
# ### Error 2
model.train_on_batch(observation_batch, action_batch, sample_weight=np.random.rand(600,1))
|
Keras_sample_weight_1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Day 13: An Example of Markov Chain Monte Carlo Sampling
#
# We will be looking at a specific version of MCMC sampling called gibbs sampling for a 2D multivariate normal. This will be covered in detail in a few lectures, but the goal for now it get a sense of what it means to sample from a Markov Chain.
#
# # Some helper code
# +
##Imports
import numpy as np
import pandas as pd
import scipy.stats
np.set_printoptions(precision=3, suppress=True)
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style("whitegrid")
sns.set_context("notebook", font_scale=1.25)
np.random.seed( 12 )
# +
##Gibbs sampling helper functions
def draw_z0_given_z1(z1):
''' Sample a value of z[0] from its conditional given z[1]
Args
----
z1 : float
Value of random variable z[1]
random_state : numpy RandomState object
Random number generator
Returns
-------
z0_samp : float
Sample from the conditional p(z[0] | z[1])
'''
## First, use Bishop textbook formulas to compute the conditional mean/var
mean_01 = 0.4 * z1
var_01 = 0.68
# Draw a value u from the Standard Normal (zero mean, unit variance)
u_samp = np.random.randn()
# Transform this value into a sample from Normal(mean_01, var_01)
# Remember, if u ~ Normal(0, 1), a "standard" normal with mean 0 variance 1,
# then using transform: x <- T(u), with T(u) = \mu + \sigma * u
# we can say x ~ Normal(\mu, \sigma^2)
z0_samp = mean_01 + np.sqrt(var_01) * u_samp
return z0_samp
def draw_z1_given_z0(z0):
''' Sample a value of z[1] from its conditional given z[0]
Args
----
z0 : float
Value of random variable z[0]
random_state : numpy RandomState object
Random number generator
Returns
-------
z1_samp : float
Sample from the conditional p(z[1] | z[0])
'''
## First, use Bishop textbook formulas to compute conditional mean/var
mean_10 = 0.8 * z0
var_10 = 1.36
# Draw a value u from the Standard Normal (zero mean, unit variance)
u_samp = np.random.randn()
# Transform this value into a sample from Normal(mean_01, var_01)
# Remember, if u ~ Normal(0, 1), a "standard" normal with mean 0 variance 1,
# then using transform: x <- T(u), with T(u) = \mu + \sigma * u
# we can say x ~ Normal(\mu, \sigma^2)
z1_samp = mean_10 + np.sqrt(var_10) * u_samp
return z1_samp
# +
##Using gibbs sampling
def draw_sample_from_mcmc( prev_sample ):
new_sample = np.zeros( 2 )
new_sample[ 0 ] = draw_z0_given_z1(prev_sample[1])
new_sample[ 1 ] = draw_z1_given_z0(prev_sample[0])
return new_sample
##Using numpy's built in MVN
def draw_sample_from_mvn():
true_mu_2 = np.zeros(2)
true_cov_22 = np.asarray([[1.0, 0.8], [0.8, 2.0]])
true_sample_SD = np.random.multivariate_normal(true_mu_2, true_cov_22 )
return true_sample_SD
# -
def plot_samples_from_mcmc_and_numpy_mvn( z_samples_SD , true_samples_SD ):
true_density = []
for i in range( 1000 ):
true_density.append( draw_sample_from_mvn() )
true_density = np.array( true_density )
z_samples_SD = np.array( z_samples_SD )
true_samples_SD = np.array( true_samples_SD )
fig, ax_grid = plt.subplots(nrows=1, ncols=2, sharex=True, sharey=True, figsize=(10,4))
ax_grid[0].plot(z_samples_SD[0,0], z_samples_SD[0,1], 'rx' )
ax_grid[0].legend( 'Start state' )
ax_grid[0].plot(z_samples_SD[:,0], z_samples_SD[:,1], 'k' )
ax_grid[0].set_title('MCMC sampler')
ax_grid[0].set_aspect('equal', 'box');
ax_grid[0].set_xlabel('$z_0$');
ax_grid[0].set_ylabel('$z_1$');
ax_grid[1].plot(true_samples_SD[:,0], true_samples_SD[:,1], 'k.')
ax_grid[1].set_title('np.random.multivariate_normal')
ax_grid[1].set_aspect('equal', 'box');
#ax_grid[1].set_xlim([-6, 6]);
#ax_grid[1].set_ylim([-6, 6]);
ax_grid[1].set_xlabel('$z_0$');
ax_grid[1].set_ylabel('$z_1$');
# # Exercise: Compare results of your MCMC-sampler and Numpy's built-in sampler side-by-side
# ## Task 1: implement a for loop to draw S total samples from both an MCMC method for sampling, and numpy's built in sampler for an MVN and visualize the results.
#
# You have been given functions that draw 1 sample from each of these methods, and an example of how to do so. Be careful to pass in the correct input to the function that draws an mcmc sample! It should always be the previously drawn MCMC sample.
# +
S = 100
mcmc_sample = np.zeros( 2 ) ##Start state for mcmc
##Draw 1 mcmc sample
mcmc_sample = draw_sample_from_mcmc( mcmc_sample )
##Draw 1 sample from numpy's MVN
mvn_sample = draw_sample_from_mvn()
mcmc_samples = [ mcmc_sample ]
mvn_samples = [ mvn_sample ]
##TODO: sample S-1 more samples from each method and add them to the arrays of samples
plot_samples_from_mcmc_and_numpy_mvn( mcmc_samples , mvn_samples )
# -
# ## Task 2: Discuss the 2 methods. Do both sets of samples that look similar? Does MCMC work?
#
# Note: You can treat the samples drawn from the numpy function as ground truth.
# ## Task 3: Now draw only 5 samples and start the mcmc chain in a "bad" state at [-10, 10]. Then draw 5 samples again, but this time start the mcmc chain in a "better" state at [ 0, 0].
# +
S = 5
##Bad mcmc start state example
mcmc_sample = [ -10 , 10 ] ##Bad start state for mcmc
##Draw 1 mcmc sample
mcmc_sample = draw_sample_from_mcmc( mcmc_sample )
##Draw 1 sample from numpy's MVN
mvn_sample = draw_sample_from_mvn()
mcmc_samples = [ mcmc_sample ]
mvn_samples = [ mvn_sample ]
##TODO: sample S-1 more samples from each method and add them to the arrays of samples
##You can use the same for loop as above.
plot_samples_from_mcmc_and_numpy_mvn( mcmc_samples , mvn_samples )
############################################################################################################
##Good mcmc start state example
mcmc_sample = [ 0 , 0 ] ##Good start state for mcmc
##Draw 1 mcmc sample
mcmc_sample = draw_sample_from_mcmc( mcmc_sample )
##Draw 1 sample from numpy's MVN
mvn_sample = draw_sample_from_mvn()
mcmc_samples = [ mcmc_sample ]
mvn_samples = [ mvn_sample ]
##TODO: sample S-1 more samples from each method and add them to the arrays of samples
##You can use the same for loop as above.
plot_samples_from_mcmc_and_numpy_mvn( mcmc_samples , mvn_samples )
# -
# ## Task 4: Discuss--did the "bad" start state affect the MCMC sampler in a different way than the "good" start state? Is there any analogue for the numpy function?
#
# Note: you can use the numpy samples from task 1 as ground truth for what true samples from the distribution should look like.
#
# Note: Make sure to look at the x and y axes of the graph
|
in_class_exercises/day12MCMC-Example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/simonsanvil/ECG-classification-MLH/blob/master/notebooks/MLH_FINAL_PROJECT_QUIQUE.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="OTuf_KnD4A8H"
# IMPORTING REQUIRED LIBRARIES
from google.colab import drive
from collections import Counter
from shutil import copyfile
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import pickle
import zipfile
import os, sys
# + id="Fyn1Ronm4fgd" colab={"base_uri": "https://localhost:8080/"} outputId="8f21b7a6-e0e6-47a1-9933-13ece457072d"
# MOUNTING DRIVE
drive.mount('/content/drive')
drive_path = "drive/MyDrive/UNI/4º/MLH"
# + id="pFs6pIUj5GcA" colab={"base_uri": "https://localhost:8080/"} outputId="c76427a0-1b8c-454f-8f92-b995c468e6be"
# LOADING ORIGINAL DATA
"""
zipPath='./training2017.zip' #path of the 1st zip file
copyfile(drive_path+'/training2017.zip', zipPath) #Copy data file to our working folder
dataFolder='./data' #We extract files to the current folder
with zipfile.ZipFile(zipPath, 'r') as zip_ref:
zip_ref.extractall(dataFolder)
%ls data/training2017
"""
# + colab={"base_uri": "https://localhost:8080/"} id="ehS__G9v3cjK" outputId="71830e9a-3fa2-4c10-fe5e-43513b1a4335"
# LOADING THE PREPROCESSED DATA
with open(drive_path+'/challenge2017.pkl', 'rb') as fin:
res = pickle.load(fin)
all_data = res['data']
all_label = res['label']
print(Counter(all_label))
# + colab={"base_uri": "https://localhost:8080/"} id="f07koSWMX0In" outputId="46250104-0bed-4c07-9de4-6d4f87dcf83b"
# !git clone https://github.com/hsd1503/ENCASE
# + colab={"base_uri": "https://localhost:8080/"} id="1SF2vvY_e-CI" outputId="e9089e3e-b691-401a-8a04-7898e20e6d30"
# %cd ..
# !bash setup.sh
# + colab={"base_uri": "https://localhost:8080/", "height": 630} id="eyJJB0R4XQ7T" outputId="5e7f9085-be18-4d76-befe-3895f751dac4"
#with open('../'+drive_path+'/v2.5_xgb5_all_v2.pkl', 'rb') as f:
# %cd /content/ENCASE/code/
with open('../model/v2.5_xgb5_all_v2.pkl', 'rb') as f:
model = pickle.load(f)
model
# + id="Wr_-MzdBJG7O"
# OVERVIEW OF THE DIFFERENT CLASSES
for ax in axs.flat:
fig, axs = plt.subplots(2, 2)
ax[0, 0].plot(x, y)
ax[0, 0].set_title('Axis [0, 0]')
ax.set(xlabel='x-label', ylabel='y-label')
# Hide x labels and tick labels for top plots and y ticks for right plots.
ax.label_outer()
# + id="6zuk-W6YM5sf"
|
notebooks/quique-exploration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Recommender System with Python
# ## Overview
# we will be working with movielens dataset. The dataset consist of two .csv files, movies.csv and ratings.csv. If you want, you can directly downloaded these files from the provided link.
# ##### Let's import some libraries
import numpy as np
import pandas as pd
# ##### Let's read the datafiles
movies = pd.read_csv('movies.csv')
ratings = pd.read_csv('ratings.csv')
# ##### Let's check the head of movies and ratings
movies.head()
ratings.head()
# In the ratings dataset, we have userId, movieId, rating that the movie got from a specific user, and the timestamp at which the movie got rating from the user.
# If we look at both ratings and movies datasets, movieId column is a common column. Let's merge the dataframes on movieId and output the head() of the merged data.
df = pd.merge(ratings,movies, on = 'movieId')
df.head()
# ##### Let's do some pre-processing on data
# The first thing we can do is to create a dataframe that tells us the average rating for the movie and number of ratings the movie we got.
# To do this, we can use the groupby method.
# We need to groupby the movies on title and then grab the rating column to get its mean, so that we have the mean rating of each movie. We can also use sort_values(ascending = False) to re-arrange the entries from higher to lower mean rating and then check the head of dataframe, all in a single line code.
df.groupby('title')['rating'].mean().sort_values(ascending=False).head()
# There is an important situation to consider in the above groupby operation, while getting the mean rating.
# What if only 1 or 2 persons have watched a movie and gave a 5 stars rating to it.
# On the other hand, there could be a movie that got a good number of ratings between 1 and 5 stars. The average in such situation will be less then 5 but the number of watcher are higher.
# Once again, groupby on title and grab the ratings column. Instead of mean, we want count for this task. We can again use sort_values(ascending =False)
# to order the entries and check the head of our dataset.
df.groupby('title')['rating'].count().sort_values(ascending=False).head()
# It looks like the top 5 famous movies don't have the 5 star average rating.
# This is fine, you can imagine, if the movie, e.g. <NAME> got 341 ratings, all ratings must be 5 star to get average value of 5.
#
# Many of us may not know even the name of the movie with average 5 star rating, but, most of us might be familiar with the movies with most number of ratings, such as Start Wars in the list on number 5.
# ##### Let's create a new dataframe rating, in which we group the movies with their mean or average rating.
# we are going to use groupby on title and grab the rating columns to get its mean.
rating = pd.DataFrame(df.groupby('title')['rating'].mean())
rating.head()
# In the above dataframe, we got the average rating for each movie in rating column and name of the movie in title column. This does not make much sense for the movies who got several number of rating as compare to those who got very few.
# We need to look at the number of ratings as well.
# ##### Let's create another column n_ratings that tell us the number of ratings for each movie.
rating['n_ratings'] = pd.DataFrame(df.groupby('title')['rating'].count())
rating.head()
# We have a dataframe rating with three columns title, rating which is actually average of mean rating, and n_rating.
# ## Exploratory Data Analysis
# ##### Let's import some librairies again
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('white')
# %matplotlib inline
# To learn from our dataset, let's plot two histograms side-by-side, one for No. of ratings for movies, n_ratings column, and the other for average rating which is rating column in the dataframe.
# We can creates two subplots and unpacks the output array immediately.
# +
f , (ax1, ax2) = plt.subplots(nrows=1,ncols=2,figsize=(10,4))
ax1.set_title('No of ratings')
ax1.hist(rating['n_ratings'], bins = 30)
ax2.set_title('Rating')
ax2.hist(rating['rating'], bins = 30)
plt.show()
# -
# If we look at the "No. of Ratings" on the left, we learn that most of the movies have 0 or 1 number of ratings! So, either, people have not watched those movies and if they have watched, they did not rate them. The first argument somehow make sense, because, most of the time we prefer to watch only the famous movies.
# If we look at the "Rating, which is actually the mean rating" on the right, we may find that there are peaks at the whole numbers {1,2,3,4,5}. This is again a usual act, most of the time people rate the things with the whole numbers.
#
# Notice, there are some movies with average 5 star rating, they might be the outstanding movies or may got only few ratings. You can also point out some movies with really bad rating. However, most of the movies got average rating between 3 and 4.
# ##### Let's check the relationship between rating and no. of rating with a seaborn jointplot()
sns.jointplot(x='rating', y = 'n_ratings', data=rating)
plt.show()
# The joint plot is is making sense, more the number of rating a movie have, more average rating it gets.
# Good the movie is, more people will watch it and the movie will get more number of ratings or reviews. This is a normal act.
# We also see from the plot that the 1 or 2 stars movie have very few number of ratings.
# ## Recommender System for similar movie
# ##### Let's develop a simple recommender system that can recommend similar movies to the user.
df.head()
# Let's create a matrix that will have the userId on one axis (index) and the title on another axis (columns) whereas, rating as its value. In the way, each cell will consist of the rating that the user gave to a certain movie.
# We need userId, title and rating columns for such matrix. We are going to use pivot_table() method to get our required matrix.
# Our parameter for the pivot_table() will be:
# + active=""
# index = userId
# columns = title
# values = rating
# -
rating_mat = df.pivot_table(values='rating', index='userId', columns='title')
rating_mat.head()
# ##### Let's check the most rated movies once again from our rating dataframne.
rating.sort_values('n_ratings', ascending=False).head(10)
movies[movies['title']=='Forrest Gump (1994)']
movies[movies['title']=='Matrix, The (1999)']
FG_user_ratings = rating_mat['Forrest Gump (1994)']
Matrix_user_ratings = rating_mat['Matrix, The (1999)']
FG_user_ratings.head(), Matrix_user_ratings.head()
# we got the user ratings for the selected movies.
# Now, we want to know how these movies are correlated to the other movies in the dataframe.
# Let's see how the user rating of Forrest Gump (FG_user_ratings) is correlated with the user rating of all other movies in the rating_mat.
# We are going to compute the correlation of FG_user_ratings to the user rating or user behavior for all other movies and passing that to similar_to_FG and similar_to_Matrix.
similar_to_FG = rating_mat.corrwith(FG_user_ratings)
similar_to_Matrix =rating_mat.corrwith(Matrix_user_ratings)
# similar_to_FG.head()
# We need to clean the data for null value using dropna().
# First, We can create a dataframe instead of series so that it look little nicer, and then we will deal with NaN.
# In order to create a dataframe, we need to pass similar_to_FG and similar_to_matrix to pandas DataFrame(). We can set the column name as correlation.
corr_FG = pd.DataFrame(similar_to_FG, columns=['correlation'])
corr_FG.head()
# ##### Let's drop NaN and check the head
# corr_FG.dropna(inplace = True)
# corr_FG.head()
# ##### Let's do the same for Matrix
# corr_matrix = pd.DataFrame(similar_to_Matrix, columns=['correlation'])
# corr_matrix.dropna(inplace = True)
# corr_matrix.head()
# ##### Let's sort Matrix
corr_matrix.sort_values('correlation', ascending=False).head()
# We may not have ever heard about most of these movies but the correlation that we got is perfect. The results does not make much sense.
# We need to fix this and we know what the reason is. Most likely, these movies are watched only once by the same users who also watched Matrix and rated both with similar stars.
# To fix this, we can set a threshold value for the ratings.
# Let's re-plot a histogram for n_rating to see which could be a good threshold value for no of ratings.
plt.hist(rating['n_ratings'], bins = 50)
plt.show()
# We can see that the drop is significant after n_rating = 50.
# We can select 50 as a minimum no. of rating in order to be considered into our recommender system.
# ##### Let's sort the value again with the condition (n_rating >50) and also join the n_ratings column from rating dataframe to corr_matrix dataframe and apply the condition for n_rating > 50
corr_matrix = corr_matrix.join(rating['n_ratings'])
corr_matrix.head()
# ##### Let's sort the values in order from high to low
corr_matrix[corr_matrix['n_ratings']>50].sort_values('correlation', ascending = False).head()
# The results make much more sense! Matrix has the perfect correlation to itself. The Star Trek is the closely correlated with Matrix.
# ##### Let's do the same for <NAME>
corr_FG = corr_FG.join(rating['n_ratings'])
corr_FG[corr_FG['n_ratings']>50].sort_values(
'correlation', ascending = False).head()
|
Recommander_Systems_Project.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Q1 of Assignment # 3
# Make a calculator using Python with addition , subtraction , multiplication ,
# division and power.
# ---------------------------------------
operand_1 = float(input(" Enter first number "))
operand_2 = float(input("\n Enter second number "))
operator = input("\n Enter operation to be performed between val_1 and val_2\n \
+ for addition\n \
- for subtraction\n \
* for multiplication\n \
/ for division\n \
and \ ** for power")
if operator == "+" :
summation = operand_1 + operand_2
print(" \n the sum of two operands is :", summation)
if operator == "-" :
subtraction = operand_1 - operand_2
print(" \n the difference of two operands is :", subtraction)
if operator == "*" :
multiplication = operand_1 * operand_2
print(" \n the product of two operands is :", multiplication)
if operator == "/" :
division = operand_1 / operand_2
print(" \n the quotient of two operands is :", division)
if operator =="**" :
power = operand_1 ** operand_2
print(" \n the operand_2 as the power of operand_1 is :", power)
# +
# Q2 of Assignment # 3
# Write a program to check if there is any numeric value in list using
# for loop
# ---------------------------------------
mix_list = ["<NAME>", 88, "PIAIC", "Saylani", "Mass Training"]
for element_value in mix_list :
if element_value == mix_list[1] :
print("There is numeric value which is :", element_value)
# +
# Q3 of Assignment # 3
# Write a Python script to add a key to a dictionary
# ---------------------------------------
# new empty dictionary
new_dict = {}
# adding a key to new_dict
new_dict['firsr_key'] = '1000'
#printing new_dict
print(new_dict)
# +
# Q4 of Assignment # 3
# Write a Python program to sum all the numeric items in a dictionary
# ---------------------------------------
# new dictionary
new_dict = {'item_1': 1, 'item_2': 2, 'item_3': 3, 'item_4': 4, 'item_5': 5}
summation = 0
for item in new_dict :
# print(new_dict[item])
summation = summation + new_dict[item]
print(summation)
# +
# Q5 of Assignment # 3
# Write a program to identify duplicate values from list
# ---------------------------------------
listing = [90, 20, 40, 20, 20, 70, 40, 90, -20, 60, 40, -20, 70]
_size = len(listing)
repeated = []
for i in range(_size):
k = i + 1
for j in range(k, _size):
if listing[i] == listing[j] and listing[i] not in repeated:
repeated.append(listing[i])
print (repeated)
# +
# Q6 of Assignment # 3
# Write a Python script to check if a given key already exists in a dictionary
# ---------------------------------------
def checkKey(dict, key):
if key in dict.keys():
print("Present, ", end =" ")
print("value =", dict[key])
else:
print("Not present")
# Driver Code
dict = {'Jan': 1, 'Feb':2, 'Mar':3, 'April':4, 'May':500,
'Jun': 6, 'Jul': 7, 'Aug': 8, 'Sep': 9, 'Oct': 10, 'Nov': 11, 'Dec': 12}
key = 'Jan'
checkKey(dict, key)
key = 'Jul'
checkKey(dict, key)
# -
|
PY02303 , Muhammad Huzefa , Assignment # 3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
input_patterns=np.load('input_patterns.npy')
weights=np.load('weights.npy')
print(weights)
print(input_patterns.shape)
# +
def sigmoid(x):
return 1 / (1 + np.exp(-x))
def one_step(weights, pre_syn, post_syn):
dt=0.01
tau=10
new_post_syn=post_syn+dt/10*(-post_syn+np.dot(weights,sigmoid(pre_syn)))
return new_post_syn
# -
xs=[0]
for j in range(0,10000):
xs.append(one_step(weights,input_patterns[:,j],xs[-1]))
print(xs)
plt.plot(xs[5000:])
|
WorkInProgress3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# - - - -
# # Mechpy
# a mechanical engineer's toolbox
#
# To view this notebook, use the [nbviewer](http://nbviewer.jupyter.org/github/nagordon/mechpy/blob/master/mechpy.ipynb)
# - - - -
#
# - - - -
# ### Modules
#
#
# ## 1) [Statics](#Statics)
# * [Example 1: A simple supported beam with shear-bending plots](#Statics-Example-1)
# * [Example 2: Vector calculation method to calculate 3-D moments](#Statics-Example-2)
# * [Distributed Loads Calculations](#Distributed-Loads-Calculations)
#
# ## 2) [Materials](#Materials)
# * [composite mechanics](#Composite-Mechanics)
#
#
# ## 3) Kinematics
# * [double_pendulum](http://matplotlib.org/examples/animation/double_pendulum_animated.html)
#
#
# ## 4) Dynamics
# * [dynamics](#Dynamics)
# * [Python Vibration Codes](http://vibrationdata.com/software.htm)
# * [Dynamics Notes](#Dynamics-Vibrations-Notes)
#
# ## Appendix A: [Engineering Mathematics with Python](#Engineering-Mathematics-with-Python)
# [Differential Equations](#Differential-Equations)
# [Linear Algebra](#Linear-Algebra)
# [Signal Processing](#Signal-Processing)
# [Finite Element Method](#Finite-Element-Method)
# * [solids FEM example](#FEM-Example-1)
#
# [Units](#Units)
#
# - - - -
# - - - -
#
# ## References
# Hibbler - Statics
# Hibbler - Mechanics of Materials
#
#
# ## Python Initilaization with module imports
# +
# setup
import numpy as np
import sympy as sp
import scipy
from pprint import pprint
sp.init_printing(use_latex='mathjax')
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = (12, 8) # (width, height)
plt.rcParams['font.size'] = 14
plt.rcParams['legend.fontsize'] = 16
from matplotlib import patches
get_ipython().magic('matplotlib') # seperate window
get_ipython().magic('matplotlib inline') # inline plotting
# -
# - - - -
# # Statics
# [index](#Mechpy)
# - - - -
#
# The sum of the forces is zero
# $$
# \Sigma F_x =0 , \Sigma F_y =0 , \Sigma F_z =0
# $$
# The sum of the moments is zero
# $$
# \Sigma M_x =0 , \Sigma M_y =0 , \Sigma M_z =0
# $$
#
# Dot Product
#
# $$
# \vec{A} \bullet \vec{B} = ABcos\left(\theta\right)= A_xB_x + A_yB_y+A_zB_z
# $$
#
# Cross-Product
#
# $$
# \vec{C}=\vec{A} \times \vec{B} =
# \begin{vmatrix}
# \widehat{i} & \widehat{j} & \widehat{k}\\
# A_{x} & A_{y} & A_{z}\\
# B_{x} & B_{y} & B_{z}
# \end{vmatrix}
# $$
#
# Moment of Force
# $$
# M_0 = Fd \\
# \vec{M_0}=\vec{r}\times \vec{F} =
# \begin{vmatrix}
# \widehat{i} & \widehat{j} & \widehat{k}\\
# r_{x} & r_{y} & r_{z}\\
# F_{x} & F_{y} & F_{z}
# \end{vmatrix}
# $$
#
# Moment of Force about a Specified axis
# $$
# \vec{M_a}=\vec{u}\bullet\vec{r}\times \vec{F} =
# \begin{vmatrix}
# u_{x} & u_{y} & u_{z} \\
# r_{x} & r_{y} & r_{z} \\
# F_{x} & F_{y} & F_{z}
# \end{vmatrix}
# $$
#
# ### Statics-Example 1
# #### A simple supported beam with shear-bending plots
from mechpy.statics import simple_support
simple_support()
# ### Statics-Example 2
# ### Vector calculation method to calculate 3-D moments
# $
# \vec{M_{R_0}}=\Sigma \left( \vec{r} \times \vec{F} \right) = \vec{r_A}\times\vec{F_1} +\vec{r_A}\times\vec{F_2} + \vec{r_B}\times\vec{F_3}
# $
from mechpy.statics import moment_calc
moment_calc()
# ### Distributed Loads Calculations
#
# $
# F_R = \Sigma F=\int_L w(x) \,dx = \int_A dA \,dx
# $
#
# example, hibbler, pg 187
#
# $$
# F_R=\int_A dA \, = \int_{0}^{2} \, 60x^2 \, dx = 160 N
# $$
#
# $$
# \overline{x} = \frac{\int_A x \, dA}{\int_A dA} = \frac{\int_{0}^{2} x60x^2 \, dx}{\int_{0}^{2} \, 60x^2 \, dx} = \frac{240}{160}
# $$
x = sp.var('x')
w = 60*x**2# N/m
sp.plot(w, (x,0,2));
w.subs(x,2)
sp.Integral(w,(x,0,2))
sp.integrate(w)
sp.integrate(w,(x,0,2))
sp.Integral(x*w,(x,0,2))
sp.integrate(w*x)
sp.integrate(x*w,(x,0,2))
FR= float(sp.integrate(w,(x,0,2)))
xbar = float(sp.integrate(x*w,(x,0,2)))/FR
print('center of load of an exponential distributed load is %f' % xbar)
# +
#hibbler pg 346
import numpy as np
r = np.array([ 0, 3 , 5.25])
F = np.array([-13.5, 0 ,6.376])
M = -np.cross(r,F)
# moments
print('M_x = %f \nM_y = %f \nM_z = %f' % (M[0], M[1], M[2] ))
# -
# # Materials
# [index](#Mechpy)
#
# ## Stress and Strain
# Stress is a tensor that can be broken into
#
# $$
# \overline{\sigma}=\begin{bmatrix}
# \sigma_{xx} & \sigma_{xy} & \sigma_{xz}\\
# \sigma_{yx} & \sigma_{yy} & \sigma_{yz}\\
# \sigma_{zx} & \sigma_{zy} & \sigma_{zz}
# \end{bmatrix}
# $$
#
#
#
# ## Factors of safety
# In aerospace, typically 1.2 for civilian aircraft and 1.15 for military
#
# $$FS=\frac{\sigma_{yield}}{\sigma}-1$$
#
# ## Fastener Notes and Formulas
#
# Finding the centroid of a bolt with coordinates, $\overline{x},\overline{y}$
# $$ \overline{x}=\frac{\sum_{i=1}^{n_b}{A_i x_i} }{\sum_{i=1}^{n_b}{A_i} } \ \ \overline{y}=\frac{\sum_{i=1}^{n_b}{A_i y_i} }{\sum_{i=1}^{n_b}{A_i}}$$
#
# Joint/Polar Moment of Inertia, $r=$ distance from centroid to fastener
# $$J= \int{r^2dA}= \sum_{i=1}^{n_b}{A_k r_k^2}$$
#
# Bearing Stress on a bolt
# $$\sigma^i_{bearing}=\frac{V_{max}}{Dt}$$
#
# Shear Stress on each bolt i due to shear force
# $$\tau_f^i = \frac{P}{\sum_{i=1}^{n_b}{A_i} }$$
# Where $A_i=$ the area of ith bolt, $n_b=$number of bolts, and $P=$ shear force
#
# Shear Stress on each bolt i due to moment
# $$\tau_t^i = \frac{T r_i}{J} $$
#
# ### Modes of failure of fastened Joints
# 1. Tensile Plate Failure across the net section between rivets/bolts
# 2. Failure of rivets through shear
# 3. Compression failure between rivet and plate
# 4. Edge shear-out at rivet hole
# 5. Edge tearing at rivet hole
#
# #### 1.
#
# $$\sigma_t =\frac{F_s}{(b-nd)t}$$
#
# #### 2.
#
# #### 3.
#
# #### 4.
#
# #### 5.
#
#
#
# ## Adhesive Joints
#
# With members, or adherends, joined with adhesives, either the member will fail due to tensile loads or the adhesive will fail in shear.
#
# The simple solution to finding the stress of bonded surfaces is taking the average stress
# $$\tau_{avg}=\frac{P}{bL}$$, is not an accurate way to model maximum stress. A good rule of thumb based on the calculations below is
# $$\tau_{max}=2.08\tau_{avg}$$
#
# The maximum shearing stress of an adhesive layer, $\tau_{max}$, can be computed as
# $$\tau_{max}=K_s\tau_{avg}=K_s\left(\frac{P}{bL_L}\right)$$
# with $P$ as applied load, $b$ as the width ofthe adhesive layer, and $L_L$ as the length ofthe adhesive layer. The stress distribution factor, $K_s$, can be defined as $K_s=\frac{cL}{tanh(CL/2)}$ where $c=\sqrt{\frac{2G_a}{Et_mt_a}}$, where the shear modulus, $G_a=\frac{\tau}{\gamma}$, and $E$ as the modulus of elasticity.
#
#
# The max shearing stress, $\tau_{max}$ in a scarf joint can be found with
# $$\tau_{max}=K_s\tau_{avg}=K_s\left[ \frac{Pcos\theta}{\left(\frac{bt}{sin\theta} \right) } \right] = K_s\left( \frac{P}{bt} sin\theta cos\theta \right)$$
# where $t$ is the thickness of the adherend members and $\theta=tan^{-1}\frac{t}{L_s}$ is the scarf angle
#
# *Mechanical Design of Machine Elements and Machines by <NAME>., <NAME>., <NAME>. (2009)*
# %matplotlib inline
# +
## Bolted Joint Example
# fastener Location
from mechpy.design import fastened_joint
fx = [0,1,2,3,0,1,2,3]
fy = [0,0,0,0,1,1,1,1]
# Force magnitude(x,y)
P = [-300,-500]
# Force location
l = [2,1]
df = fastened_joint(fx, fy, P, l)
df.plot(kind='scatter', x='x', y='y');
#df.plot(style='o', x='x', y='y')
plt.plot(df.xbar[0],df.ybar[0],'*')
df
#ax = plt.gca()
#ax.arrow(l[0], l[1], Pnorm[0],Pnorm[1], head_width=0.05, head_length=0.1, fc='k', ec='k')
#x.arrow(xbar, ybar, Pnorm[0],0, head_width=0.05, head_length=0.1, fc='k', ec='k')
#ax.arrow(xbar, ybar, 0,Pnorm[1], head_width=0.05, head_length=0.1, fc='k', ec='k')
# -
# ## Composite Mechanics
# [index](#Mechpy)
from mechpy.math import T3rot, T6rot
from mechpy.composites import qbar_transformtion, composite_plate, vary_ply_direction_plot, laminate_gen
from mechpy.math import T2rot
T2rot(45)
T6rot(45,45,45)
# +
from IPython.html.widgets import *
plt.figure(figsize=(12,8))
x = [-1,1, 0,-1,]
y = [-1,-1,1,-1]
xy = np.array([x,y])
plt.xlim([-11.1,11.1])
plt.ylim([-11.1,11.1])
xyR = np.dot(T2rot(30),xy)
#plt.plot(xyR[0,:],xyR[1,:])
def rot2(th, xt,yt,zt):
xyR = np.dot(T2rot(th),xy*zt)
xyR[0,:]+=xt
xyR[1,:]+=yt
plt.plot(xyR[0,:],xyR[1,:])
plt.axis('square')
plt.xlim([-11.1,11.1])
plt.ylim([-11.1,11.1])
plt.show()
interact(rot2, th=(0,np.pi,np.pi/90), yt=(1,10,1), xt=(1,10,1), zt=(1,10,1));
# -
print(T6rot(45,45,45))
vary_ply_direction_plot()
qbar_transformtion()
help(laminate_gen)
laminate_gen()
composite_plate()
# +
from ipywidgets import IntSlider
IntSlider()
# +
# Principal Stresses
sx = 63.66
sy = 0
sz = 0
txy = 63.66
txz = 0
tyz = 0
S = np.matrix([[sx, txy, txz],
[txy, sy, tyz],
[txy, txz, sz]])
print(S)
# -
principal_stresses = np.linalg.eigvals(S)
print(principal_stresses)
import sympy as sp
from sympy.abc import tau, sigma
#s,s11,s22,s33,s12 = sp.var('s,s11,s22,s33,s12')
s,s11,s22,s33,s12,s13 = sp.symbols('sigma, sigma11,sigma22,sigma33,sigma12,sigma13')
s = sp.Matrix([[s11,s12,0],[s12,s22,0],[0,0,s33]])
s
s**2
s.eigenvals() # hmm looks familiar
s1 = s.subs(s11,2.2).subs(s22,3).subs(s33,sp.pi).subs(s12,7.3)
s1
# or
s2 = s.evalf(subs={s11:2.2, s22:3, s33:sp.pi, s12:7.3})
s2
s1.eigenvals()
s2.eigenvals()
s2.inv()
C = sp.symbols('C1:100')
C
from mechpy.math import ode1
ode1()
# ## Dynamics Vibrations Notes
# **Jul 1, 2015**
#
# ## Introduction
# <div id="sec:intro"></div>
#
# modal analysis is similar to frequency analysis. In frequency analysis a complex signal is resolved into a set of simple sine waves with individual frequency and amplitude and phase parameters. In modal analysis, a complex deflection pattern of a vibrating structure is resolved into a set of simple mode shapes with the same individual parameters.
#
#
# ## Structural Dynamics Background
# <div id="sec:stdybg"></div>
#
# Most systems are actually multiple degrees of freedom (MDOF) and have some non-linearity, but can be simplified with a superposition of SDOF linear systems
#
# Newtons law states that acceleration is a function of the applied force and the mass of the object, or
# $$
# [inertial forces] + [Dissipative forces] + [Restoring Forces] = [External Forces] \\
# m\ddot{x} + c\dot{x} + kx = f(t) \\
# \zeta<1 is\ underdamped \\
# $$
#
# some other dynamic characteristics are
# $$
# \omega = frequency \\
# \zeta = damping \\
# \{\phi\} = mode shape \\
# \omega^{2}_{n}=\frac{k}{m} = natural frequency \\
# \zeta = \frac{c}{\sqrt{2km}} \\
# H(\omega)=Frequency\ Response \\
# \phi(\omega)=Phase
# $$
#
# ## Damping Model
#
# Where there is energy dissipation, there is damping. The system can be broken into the system inputs/excitation, a system G(s), and the output response, in Laplace or space
#
# The transfer function is a math model defining the input/output relationship of a physical system. Another definition is the Laplace transform ( $\mathcal{L}$) of the output divided by the Laplace transform of the input.
#
# The frequency response function (FRF) is defined in a similar manner such that FRF is the fourier transform ($ \mathcal{F} $) of the input divided by the fourier transform of the output
#
# $$
# Transfer\ Function=\frac{Output}{Input} \\
# G(s) = \frac{Y(s)}{X(s)}
# $$
#
# ## Modal Testing
#
# These relationships can be further explained by the modal test process. The measurements taken during a test are frequency response function measurements. The parameter estimation routines are curve fits in the Laplace domain and result in transfer functions.
#
# Frequency Response Matrix
#
# $$
# \begin{bmatrix}
# H_{11} & H_{12} & \cdots & H_{1n} \\
# H_{21} & H_{22} & \cdots & H_{2n} \\
# \vdots & \vdots & \ddots & \vdots \\
# H_{n1} & H_{n2} & \cdots & H_{nn}
# \end{bmatrix}
# $$
#
# ## Random Notes
# <div id="sec:rand"></div>
#
# The signal-analysis approach is done by measuring vibration with accelerometers and determine the frequency spectrum. The other moethod is a system -analysis where a dual-channel FFT anlayzer is used to measure the ratio of the response to the input giving the frequency response function (FRF)
#
# a modal model allows the analysis of structural systems
#
# a mode shape is a deflection-pattern associated with a particular modal frequency or pole location. It is not tangible or easily observed. The actual displacement of the structure will be a sum of all the mode shapes. A harmonic exitation close to the modal frequency, 95% of the displacement may be due to the particular modeshape
#
# Modal Descriptions Assumes Linearity
# * Superposition of the component waves will result in the final wave. A swept sinosoid will give the same result as a broadband excitation
#
# * Homogeneity is when a measured FRF is independent of excitation level
#
# * Reciprocity implies that the FRF measured between any two DOFs is independent of which of them for excitation or response
#
# * small deflections - cannot predict buckling or catastrophic failure
#
# * casual - the structure will not vibrate before it is excited
#
# * stable - the vibrations will die out when the excitation is removd
#
# * time-invariant - the dynamic characteristics will not change during the measurments
#
# ## The Lumped-Parameter Model and Modal Theory
#
# [Physical Coordinates] = [Modal Matrix][Modal Coordinates]
#
# $$
# [x] = [\phi][q]
# $$
#
# ## Keywords and Notations
#
# $$
# m=mass \\
# k=stiffness \\
# c = damping coefficient \\
# c_c = critical damping coefficient \\
# $$
#
#
# ## Finite-Element-Method
# [index](#Mechpy)
# ### FEM-Example-1
from mechpy.fem import cst_fem
cst_fem(structure='9node')
# ## Linear Algebra with Python
# [index](#Mechpy)
#
# Python's numpy package allows python, a generic computing language to perform powerful mathematical calculations. Although python's math syntax is not as obvious as MATLAB's, the functionality is comparable. This document is designed to be an intro to that syntax
#
# Some references
#
# http://nbviewer.ipython.org/github/carljv/cython_testing/blob/master/cython_linalg.ipynb
#
# We can either use scipy, which includes numpy,
# http://docs.scipy.org/doc/
#
# or use numpy directly
# http://docs.scipy.org/doc/numpy/
#
# Since there are many ways to solve linear algebra problems, (eg Octave/Matlab, julia, scipy, numpy)
# I tend to prefer the most matlabesc approaches due to the ubiquity of Matlab and the simplicity of the syntax, which frankly, python suffers with.
#
# The major difference between arrays and matrices in python is that arrays are n-dimensions, where matrices are only up to 2-dimensions
# m
import numpy as np
from scipy import linalg
# Pythons list is a generic data storage object. it can be easily extended to a numpy array, which is specialized for numerical and scientific computation
np.zeros((5,3))
np.array([[1,2],[3,4]])
np.matrix(np.zeros((5,3)))
np.matrix([[1,2],[3,4]])
# Matrix multiplication can be achieved using the dot method
i = [[1,0,0],[0,1,0],[0,0,1]] # identiy matrix
a = [[4,3,1],[5,7,2],[2,2,2]]
np.dot(i,a)
#Or, matrix multiplication can be done if a matrix is explicitly defined
np.matrix(i)*np.matrix(a)
# Notice, when arrays are mutliplied, we get the dot product
np.array(i)*np.array(a)
# convert an array to a matrix
m = np.matrix(a)
m
m.T # transpose
m.I # inverse
m**2
np.array(a)**2
m
m[:,2]
m[2,:]
m[:2,:2]
m[1:,1:]
# ## Sympy Linear Algebra
# import sympy
import sympy as sp
#from sympy.mpmath import *
x = sp.Symbol('x') # x = var('x')
M = sp.Matrix([[2,x],[x,3]])
M
M.eigenvals()
M.eigenvects()
M.eigenvects()[1][0]
Mval = M.eigenvects()[1][0]
Mval.evalf(subs={x:3.14})
print(sp.latex(M))
# # copy and paste into markdown
#
# $ \left[\begin{matrix}2 & x\\x & 3\end{matrix}\right] $
#
# ## Signal Processing
# Page 174 Introduction for python for Science - <NAME>
import numpy as np
from scipy import fftpack
import matplotlib.pyplot as plt
width = 2.0
freq = 0.5
t = np.linspace(-10, 10, 101) # linearly space time array
g = np.exp(-np.abs(t)/width)*np.sin(2.0 * np.pi * freq * t)
dt = t[1]-t[0] # increment between times in time array
G = fftpack.fft(g) # FFT of g
f = fftpack.fftfreq(g.size, d=dt) # frequenies f[i] of g[i]
f = fftpack.fftshift(f) # shift frequencies from min to max
G = fftpack.fftshift(G) # shift G order to coorespond to f
fig = plt.figure(1, figsize=(8,6), frameon=False)
ax1 = fig.add_subplot(211)
ax1.plot(t, g)
ax1.set_xlabel('t')
ax1.set_ylabel('g(t)')
ax2 = fig.add_subplot(212)
ax2.plot(f, np.real(G), color='dodgerblue', label='real part')
ax2.plot(f, np.imag(G), color='coral', label='imaginary part')
ax2.legend()
ax2.set_xlabel('f')
ax2.set_ylabel('G(f)')
plt.show()
# # Engineering Mathematics with Python
# [index](#Mechpy)
from numpy import *
r_[1:11]
arange(1,11)
linspace(1,10,10)
# # Units
# [index](#Mechpy)
from mechunits import uc1
# uc1 uses sympy
uc1(1.0,'psi','kPa')
uc1(1.0,'newton','pound')
from mechunits import uc2
# uses pint
uc2(17.5,'lbf','newton')
uc2(300,'pascal','psi')
from mechunits import in_mm
in_mm()
# %load_ext version_information
# %version_information pydy, numpy, scipy, matplotlib
|
mechpy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="HHV8gH1Hr2Ll" colab_type="text"
# ## Define the Convolutional Neural Network
#
# After you've looked at the data you're working with and, in this case, know the shapes of the images and of the keypoints, you are ready to define a convolutional neural network that can *learn* from this data.
#
# In this notebook and in `models.py`, you will:
# 1. Define a CNN with images as input and keypoints as output
# 2. Construct the transformed FaceKeypointsDataset, just as before
# 3. Train the CNN on the training data, tracking loss
# 4. See how the trained model performs on test data
# 5. If necessary, modify the CNN structure and model hyperparameters, so that it performs *well* **\***
#
# **\*** What does *well* mean?
#
# "Well" means that the model's loss decreases during training **and**, when applied to test image data, the model produces keypoints that closely match the true keypoints of each face. And you'll see examples of this later in the notebook.
#
# ---
#
# + [markdown] id="oi9UjKCar2Lo" colab_type="text"
# ## CNN Architecture
#
# Recall that CNN's are defined by a few types of layers:
# * Convolutional layers
# * Maxpooling layers
# * Fully-connected layers
#
# You are required to use the above layers and encouraged to add multiple convolutional layers and things like dropout layers that may prevent overfitting. You are also encouraged to look at literature on keypoint detection, such as [this paper](https://arxiv.org/pdf/1710.00977.pdf), to help you determine the structure of your network.
#
#
# ### TODO: Define your model in the provided file `models.py` file
#
# This file is mostly empty but contains the expected name and some TODO's for creating your model.
#
# ---
# + [markdown] id="FUqQpu_7r2Lr" colab_type="text"
# ## PyTorch Neural Nets
#
# To define a neural network in PyTorch, you define the layers of a model in the function `__init__` and define the feedforward behavior of a network that employs those initialized layers in the function `forward`, which takes in an input image tensor, `x`. The structure of this Net class is shown below and left for you to fill in.
#
# Note: During training, PyTorch will be able to perform backpropagation by keeping track of the network's feedforward behavior and using autograd to calculate the update to the weights in the network.
#
# #### Define the Layers in ` __init__`
# As a reminder, a conv/pool layer may be defined like this (in `__init__`):
# ```
# # 1 input image channel (for grayscale images), 32 output channels/feature maps, 3x3 square convolution kernel
# self.conv1 = nn.Conv2d(1, 32, 3)
#
# # maxpool that uses a square window of kernel_size=2, stride=2
# self.pool = nn.MaxPool2d(2, 2)
# ```
#
# #### Refer to Layers in `forward`
# Then referred to in the `forward` function like this, in which the conv1 layer has a ReLu activation applied to it before maxpooling is applied:
# ```
# x = self.pool(F.relu(self.conv1(x)))
# ```
#
# Best practice is to place any layers whose weights will change during the training process in `__init__` and refer to them in the `forward` function; any layers or functions that always behave in the same way, such as a pre-defined activation function, should appear *only* in the `forward` function.
# + [markdown] id="TdKYKFtUr2Lu" colab_type="text"
# #### Why models.py
#
# You are tasked with defining the network in the `models.py` file so that any models you define can be saved and loaded by name in different notebooks in this project directory. For example, by defining a CNN class called `Net` in `models.py`, you can then create that same architecture in this and other notebooks by simply importing the class and instantiating a model:
# ```
# from models import Net
# net = Net()
# ```
# + id="hbyuASaReYiO" colab_type="code" outputId="b9efcbe9-cc0f-447a-f7b7-8e69c2a4a66c" executionInfo={"status": "ok", "timestamp": 1567441771439, "user_tz": -480, "elapsed": 1118, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07513880639336204123"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
from google.colab import drive
drive.mount("/content/drive")
# + id="LbU-Lzfcelo4" colab_type="code" outputId="11cb7559-13cf-4942-d7de-3d7367950d17" executionInfo={"status": "ok", "timestamp": 1567441771441, "user_tz": -480, "elapsed": 1101, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07513880639336204123"}} colab={"base_uri": "https://localhost:8080/", "height": 34}
# %cd drive/"My Drive"/CVND/CVND_projects/P1_Facial_Keypoints
# + [markdown] id="2EVNawhPz8Oy" colab_type="text"
#
# + id="JrGveU-Jr2Lw" colab_type="code" colab={}
# import the usual resources
import matplotlib.pyplot as plt
import numpy as np
# watch for any changes in model.py, if it changes, re-load it automatically
# %load_ext autoreload
# %autoreload 2
# + id="_wM_KmCir2L3" colab_type="code" outputId="bbff39bc-b900-45ca-e8ab-ca4f6dedcac8" executionInfo={"status": "ok", "timestamp": 1567441780048, "user_tz": -480, "elapsed": 9649, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07513880639336204123"}} colab={"base_uri": "https://localhost:8080/", "height": 391}
## TODO: Define the Net in models.py
import torch
import torch.nn as nn
import torch.nn.functional as F
## TODO: Once you've define the network, you can instantiate it
# one example conv layer has been provided for you
from models import NaimishNet
net = NaimishNet()
print(net)
# + [markdown] id="8aTIt-6sr2L9" colab_type="text"
# ## Transform the dataset
#
# To prepare for training, create a transformed dataset of images and keypoints.
#
# ### TODO: Define a data transform
#
# In PyTorch, a convolutional neural network expects a torch image of a consistent size as input. For efficient training, and so your model's loss does not blow up during training, it is also suggested that you normalize the input images and keypoints. The necessary transforms have been defined in `data_load.py` and you **do not** need to modify these; take a look at this file (you'll see the same transforms that were defined and applied in Notebook 1).
#
# To define the data transform below, use a [composition](http://pytorch.org/tutorials/beginner/data_loading_tutorial.html#compose-transforms) of:
# 1. Rescaling and/or cropping the data, such that you are left with a square image (the suggested size is 224x224px)
# 2. Normalizing the images and keypoints; turning each RGB image into a grayscale image with a color range of [0, 1] and transforming the given keypoints into a range of [-1, 1]
# 3. Turning these images and keypoints into Tensors
#
# These transformations have been defined in `data_load.py`, but it's up to you to call them and create a `data_transform` below. **This transform will be applied to the training data and, later, the test data**. It will change how you go about displaying these images and keypoints, but these steps are essential for efficient training.
#
# As a note, should you want to perform data augmentation (which is optional in this project), and randomly rotate or shift these images, a square image size will be useful; rotating a 224x224 image by 90 degrees will result in the same shape of output.
# + id="CmQcyz9Rr2L-" colab_type="code" colab={}
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
# the dataset we created in Notebook 1 is copied in the helper file `data_load.py`
from data_load import FacialKeypointsDataset
# the transforms we defined in Notebook 1 are in the helper file `data_load.py`
from data_load import Rescale, RandomCrop, Normalize, ToTensor
## TODO: define the data_transform using transforms.Compose([all tx's, . , .])
# order matters! i.e. rescaling should come before a smaller crop
data_transform = transforms.Compose([Rescale(250),
RandomCrop(224),
Normalize(),
ToTensor()])
# testing that you've defined a transform
assert(data_transform is not None), 'Define a data_transform'
# + id="E-83XyHwr2MD" colab_type="code" outputId="89986b62-4689-4fb8-ffe5-606c5f94fa6d" executionInfo={"status": "ok", "timestamp": 1567441808007, "user_tz": -480, "elapsed": 37591, "user": {"displayName": "nazrul ismail", "photoUrl": "", "userId": "07513880639336204123"}} colab={"base_uri": "https://localhost:8080/", "height": 102}
# create the transformed dataset
transformed_dataset = FacialKeypointsDataset(csv_file='data/training_frames_keypoints.csv',
root_dir='data/training/',
transform=data_transform)
print('Number of images: ', len(transformed_dataset))
# iterate through the transformed dataset and print some stats about the first few samples
for i in range(4):
sample = transformed_dataset[i]
print(i, sample['image'].size(), sample['keypoints'].size())
# + [markdown] id="7f7xK-vYr2MJ" colab_type="text"
# ## Batching and loading data
#
# Next, having defined the transformed dataset, we can use PyTorch's DataLoader class to load the training data in batches of whatever size as well as to shuffle the data for training the model. You can read more about the parameters of the DataLoader, in [this documentation](http://pytorch.org/docs/master/data.html).
#
# #### Batch size
# Decide on a good batch size for training your model. Try both small and large batch sizes and note how the loss decreases as the model trains.
#
# **Note for Windows users**: Please change the `num_workers` to 0 or you may face some issues with your DataLoader failing.
# + id="UH2WOb6kr2MK" colab_type="code" colab={}
# load training data in batches
batch_size = 64
train_loader = DataLoader(transformed_dataset,
batch_size=batch_size,
shuffle=True,
num_workers=4)
# + [markdown] id="TcvaDZHor2MO" colab_type="text"
# ## Before training
#
# Take a look at how this model performs before it trains. You should see that the keypoints it predicts start off in one spot and don't match the keypoints on a face at all! It's interesting to visualize this behavior so that you can compare it to the model after training and see how the model has improved.
#
# #### Load in the test dataset
#
# The test dataset is one that this model has *not* seen before, meaning it has not trained with these images. We'll load in this test data and before and after training, see how your model performs on this set!
#
# To visualize this test data, we have to go through some un-transformation steps to turn our images into python images from tensors and to turn our keypoints back into a recognizable range.
# + id="BHRSI-Afr2MQ" colab_type="code" colab={}
# load in the test data, using the dataset class
# AND apply the data_transform you defined above
# create the test dataset
test_dataset = FacialKeypointsDataset(csv_file='data/test_frames_keypoints.csv',
root_dir='data/test/',
transform=data_transform)
# + id="YnwS9xI3r2MT" colab_type="code" colab={}
# load test data in batches
batch_size = 64
test_loader = DataLoader(test_dataset,
batch_size=batch_size,
shuffle=True,
num_workers=4)
# + [markdown] id="TJqYCpe7r2MY" colab_type="text"
# ## Apply the model on a test sample
#
# To test the model on a test sample of data, you have to follow these steps:
# 1. Extract the image and ground truth keypoints from a sample
# 2. Make sure the image is a FloatTensor, which the model expects.
# 3. Forward pass the image through the net to get the predicted, output keypoints.
#
# This function test how the network performs on the first batch of test data. It returns the images, the transformed images, the predicted keypoints (produced by the model), and the ground truth keypoints.
# + id="FyUF_u8lr2MZ" colab_type="code" colab={}
# test the model on a batch of test images
def net_sample_output():
# iterate through the test dataset
for i, sample in enumerate(test_loader):
# get sample data: images and ground truth keypoints
images = sample['image']
key_pts = sample['keypoints']
print(key_pts)
# convert images to FloatTensors
images = images.type(torch.FloatTensor)
# forward pass to get net output
output_pts = net(images)
# reshape to batch_size x 68 x 2 pts
output_pts = output_pts.view(output_pts.size()[0], 68, -1)
# break after first image is tested
if i == 0:
return images, output_pts, key_pts
# + [markdown] id="n5SCv2Rwr2Mc" colab_type="text"
# #### Debugging tips
#
# If you get a size or dimension error here, make sure that your network outputs the expected number of keypoints! Or if you get a Tensor type error, look into changing the above code that casts the data into float types: `images = images.type(torch.FloatTensor)`.
# + id="ZrAx0J9kr2Md" colab_type="code" colab={}
# call the above function
# returns: test images, test predicted keypoints, test ground truth keypoints
test_images, test_outputs, gt_pts = net_sample_output()
# print out the dimensions of the data to see if they make sense
print(test_images.data.size())
print(test_outputs.data.size())
print(gt_pts.size())
# + [markdown] id="Dj2dUKOgr2Mg" colab_type="text"
# ## Visualize the predicted keypoints
#
# Once we've had the model produce some predicted output keypoints, we can visualize these points in a way that's similar to how we've displayed this data before, only this time, we have to "un-transform" the image/keypoint data to display it.
#
# Note that I've defined a *new* function, `show_all_keypoints` that displays a grayscale image, its predicted keypoints and its ground truth keypoints (if provided).
# + [markdown] id="R0sHZNAopp9i" colab_type="text"
#
# + id="YUOig_5Rr2Mg" colab_type="code" colab={}
def show_all_keypoints(image, predicted_key_pts, gt_pts=None):
"""Show image with predicted keypoints"""
# image is grayscale
plt.imshow(image, cmap='gray')
plt.scatter(predicted_key_pts[:, 0], predicted_key_pts[:, 1], s=20, marker='.', c='m')
# plot ground truth points as green pts
if gt_pts is not None:
plt.scatter(gt_pts[:, 0], gt_pts[:, 1], s=20, marker='.', c='g')
# + [markdown] id="z9_2lHavr2Mj" colab_type="text"
# #### Un-transformation
#
# Next, you'll see a helper function. `visualize_output` that takes in a batch of images, predicted keypoints, and ground truth keypoints and displays a set of those images and their true/predicted keypoints.
#
# This function's main role is to take batches of image and keypoint data (the input and output of your CNN), and transform them into numpy images and un-normalized keypoints (x, y) for normal display. The un-transformation process turns keypoints and images into numpy arrays from Tensors *and* it undoes the keypoint normalization done in the Normalize() transform; it's assumed that you applied these transformations when you loaded your test data.
# + id="dXeWqAYCr2Mj" colab_type="code" outputId="b5a8c2cb-596a-4c7e-9f22-e7d79e03b093" executionInfo={"status": "ok", "timestamp": 1567441885695, "user_tz": -480, "elapsed": 99251, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07513880639336204123"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
# visualize the output
# by default this shows a batch of 10 images
def visualize_output(test_images, test_outputs, gt_pts=None, batch_size=10):
for i in range(batch_size):
plt.figure(figsize=(30,50))
ax = plt.subplot(1, batch_size, i+1)
# un-transform the image data
image = test_images[i].data # get the image from it's wrapper
image = image.numpy() # convert to numpy array from a Tensor
image = np.transpose(image, (1, 2, 0)) # transpose to go from torch to numpy image
# un-transform the predicted key_pts data
predicted_key_pts = test_outputs[i].data
predicted_key_pts = predicted_key_pts.numpy()
# undo normalization of keypoints
predicted_key_pts = predicted_key_pts*50 + 100
# plot ground truth points for comparison, if they exist
ground_truth_pts = None
if gt_pts is not None:
ground_truth_pts = gt_pts[i]
ground_truth_pts = ground_truth_pts* 50 + 100
# call show_all_keypoints
show_all_keypoints(np.squeeze(image), predicted_key_pts, ground_truth_pts)
plt.axis('off')
plt.show()
# call it
visualize_output(test_images, test_outputs, gt_pts)
#Output prior to training the network
# + [markdown] id="1pli_hGzr2Ml" colab_type="text"
# ## Training
#
# #### Loss function
# Training a network to predict keypoints is different than training a network to predict a class; instead of outputting a distribution of classes and using cross entropy loss, you may want to choose a loss function that is suited for regression, which directly compares a predicted value and target value. Read about the various kinds of loss functions (like MSE or L1/SmoothL1 loss) in [this documentation](http://pytorch.org/docs/master/_modules/torch/nn/modules/loss.html).
#
# ### TODO: Define the loss and optimization
#
# Next, you'll define how the model will train by deciding on the loss function and optimizer.
#
# ---
# + id="fkDJlx1qr2Ml" colab_type="code" colab={}
## TODO: Define the loss and optimization
import torch.optim as optim
#criterion = nn.MSELoss()
criterion = nn.SmoothL1Loss()
power = .95
n_epochs = 100
init_lr = 3e-3
def poly_lr_decay(epoch):
print(epoch)
decay = (1 - (epoch/n_epochs)) ** power
new_lr = init_lr*decay
print(new_lr)
return new_lr
def poly_lr_scheduler(optimizer, init_lr, epoch, lr_decay_iter=1,
n_epoch=100, power=0.9):
"""Polynomial decay of learning rate
:param init_lr is base learning rate
:param iter is a current iteration
:param lr_decay_iter how frequently decay occurs, default is 1
:param max_iter is number of maximum iterations
:param power is a polymomial power
"""
if epoch % lr_decay_iter or epoch > n_epoch:
return optimizer
lr = init_lr*(1 - epoch/float(n_epoch))**power
print(f"Current Learning rate: {lr}")
for param_group in optimizer.param_groups:
param_group['lr'] = lr
return lr
optimizer = optim.RMSprop(net.parameters(), lr=init_lr, momentum=0.9)
#optimizer = optim.SGD(net.parameters(), lr=init_lr, momentum=.9)
#optimizer = optim.Adam(net.parameters(), lr=1e-3, weight_decay=1e-5)
# + [markdown] id="ZK4F0zfar2Mn" colab_type="text"
# ## Training and Initial Observation
#
# Now, you'll train on your batched training data from `train_loader` for a number of epochs.
#
# To quickly observe how your model is training and decide on whether or not you should modify it's structure or hyperparameters, you're encouraged to start off with just one or two epochs at first. As you train, note how your the model's loss behaves over time: does it decrease quickly at first and then slow down? Does it take a while to decrease in the first place? What happens if you change the batch size of your training data or modify your loss function? etc.
#
# Use these initial observations to make changes to your model and decide on the best architecture before you train for many epochs and create a final model.
# + id="37PRCPLHr2Mo" colab_type="code" colab={}
def train_net(n_epochs):
print_every = 10
# prepare the net for training
net.train()
net.cuda()
for epoch in range(n_epochs): # loop over the dataset multiple times
running_loss = 0.0
poly_lr_scheduler(optimizer, init_lr, epoch, lr_decay_iter=1, n_epoch=n_epochs, power=power)
# train on batches of data, assumes you already have train_loader
for batch_i, data in enumerate(train_loader):
# get the input images and their corresponding labels
images = data['image'].cuda()
key_pts = data['keypoints'].cuda()
# flatten pts
key_pts = key_pts.view(key_pts.size(0), -1)
# convert variables to floats for regression loss
key_pts = key_pts.type(torch.cuda.FloatTensor)
images = images.type(torch.cuda.FloatTensor)
# forward pass to get outputs
output_pts = net(images)
# calculate the loss between predicted and target keypoints
loss = criterion(output_pts, key_pts)
# zero the parameter (weight) gradients
optimizer.zero_grad()
# backward pass to calculate the weight gradients
loss.backward()
# update the weights
optimizer.step()
# print loss statistics
# to convert loss into a scalar and add it to the running_loss, use .item()
running_loss += loss.item()
if batch_i % 10 == 0: # print every 10 batches
print('Epoch: {}, Batch: {}, Avg. Loss: {}'.format(epoch + 1, batch_i+1, running_loss/print_every))
running_loss = 0.0
print('Finished Training')
# + id="jBL6icSar2Mq" colab_type="code" outputId="5254c136-4935-4542-b8a3-03af56a6fd22" executionInfo={"status": "ok", "timestamp": 1567444307286, "user_tz": -480, "elapsed": 2498451, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07513880639336204123"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
train_net(n_epochs)
# + [markdown] id="kRQGwP1sr2Mt" colab_type="text"
# ## Test data
#
# See how your model performs on previously unseen, test data. We've already loaded and transformed this data, similar to the training data. Next, run your trained model on these images to see what kind of keypoints are produced. You should be able to see if your model is fitting each new face it sees, if the points are distributed randomly, or if the points have actually overfitted the training data and do not generalize.
# + id="p7IaOZA0r2Mt" colab_type="code" colab={}
net.cpu()
net.eval()
# get a sample of test data again
test_images, test_outputs, gt_pts = net_sample_output()
print(test_images.data.size())
print(test_outputs.data.size())
print(gt_pts.size())
# + id="0LFY0o9Gr2Mv" colab_type="code" outputId="35f3a4c6-dbcc-4985-f84c-584ec7eb536a" executionInfo={"status": "ok", "timestamp": 1567444367765, "user_tz": -480, "elapsed": 13211, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07513880639336204123"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
## TODO: visualize your test output
# you can use the same function as before, by un-commenting the line below:
visualize_output(test_images, test_outputs, gt_pts, batch_size=10)
# + [markdown] id="E6jebywyr2Mx" colab_type="text"
# Once you've found a good model (or two), save your model so you can load it and use it later!
# + id="yhKguzSer2My" colab_type="code" colab={}
net.cpu()
## TODO: change the name to something uniqe for each new model
model_dir = 'saved_models/'
model_name = 'NaimishNet-100epochs_w_lr_decay_2.pt'
config = {'epoch': 62,
'model_state_dict': net.state_dict(),
'loss': criterion,
'optimizer': optimizer
}
# after training, save your model parameters in the dir 'saved_models'
torch.save(config, model_dir+model_name)
#net.load_state_dict(torch.load(model_dir+model_name))
# + id="BUJK7sudRXWw" colab_type="code" colab={}
# !ls saved_models/
# + [markdown] id="3FRSRnNvfG8H" colab_type="text"
#
# + [markdown] id="78drXGgyr2M0" colab_type="text"
# After you've trained a well-performing model, answer the following questions so that we have some insight into your training and architecture selection process. Answering all questions is required to pass this project.
# + [markdown] id="KGBiZQpTr2M1" colab_type="text"
# ### Question 1: What optimization and loss functions did you choose and why?
#
# + [markdown] id="v0LOS78Yr2M2" colab_type="text"
# **Answer**:
#
# 1. **Optimization**: RMSprop (stands for Root mean square prop), Proposed by <NAME>
# **Best performance:** RMSprop (lr = 3e-3 wih momentum set to 0.9)
#
# <ul>
#
# <li>I have experimented with Adam which was originally used in the paper, SGD and RMSprop. I noticed a significant slow convergence rate and unstable using Adam SGD provided that the hyperparameter was carefully selected.</li>
#
# <li>RMSprop had given me much success with lowest loss rate with the fastest convergence rate among the three. This is because RMSprop updates the gradient by divinding by its squares (element-wise) thus giving us the effect of smoothing the cost function or as we call it the <i> exponentially moving average</i>.
# </li>
#
# </ul>
#
# **Update rule for RMSprop**
# <ul>
#
# <li>$s \delta w = ({\delta w * \beta}) + (({1-\beta}) *\delta w^2)$ </li>
# <li>$s \delta b = ({\delta b * \beta}) + (({1-\beta}) *\delta b^2)$</li>
# <li>$w = w - \alpha * \frac{\delta w}{\sqrt{s \delta w}}$</li>
# <li>$b = b - \alpha * \frac{\delta b}{\sqrt{s \delta b}}$</li>
# </ul>
# Where w, b represents weights and bias respectively
#
#
# 2. **Loss function:** SmoothL1Loss
# **Experimented:** Huber Loss/Smooth Mean Absolute Error (SmoothL1Loss) and Mean Square Error (MSELoss/L2Error)
#
#
# The we are trying to learn a mapping for $x \rightarrow (x,y)$ where this are the coordinates for each of the facial keypoints that we are predicting. A regression loss is to be used in this matter. As the dataset consisted of points that were annotated and no outliers were present loss functioon Smooth MAE is used.
#
#
#
#
# + [markdown] id="zX0qmGWHr2M2" colab_type="text"
# ### Question 2: What kind of network architecture did you start with and how did it change as you tried different architectures? Did you decide to add more convolutional layers or any layers to avoid overfitting the data?
# + [markdown] id="F5BGbd6Xr2M3" colab_type="text"
# **Answer**:
#
# I initially started implemented NaimishNet which is based on <b><i>Facial Key Points Detection using Deep Convolutional Neural Network (2017)</li></b></i>, however having the input the network must be of size 96px which could lead to loss of finer details in extraction features from small images, I had tweak the hyperparameter of the architecture to work with images of size 224x224px.
#
# To counter overfitting, I have added several Regularization techniques such as Dropouts and batch normalizations into the architecture.
#
#
#
# + [markdown] id="4m80Wv6Dr2M3" colab_type="text"
# ### Question 3: How did you decide on the number of epochs and batch_size to train your model?
# + [markdown] id="GvUnXrQqr2M4" colab_type="text"
# **Answer**:
#
# 1. I have trained the network initially with a small value of 30 and gradually increase to 100 and monitor the loss in the test set to make sure that it is reducing monotonically.
# 2. total number of batch size I had determined is the maximum no. of images that my GPU could handle per Epoch.
#
#
# + [markdown] id="Z8HRRmxOr2M4" colab_type="text"
# ## Feature Visualization
#
# Sometimes, neural networks are thought of as a black box, given some input, they learn to produce some output. CNN's are actually learning to recognize a variety of spatial patterns and you can visualize what each convolutional layer has been trained to recognize by looking at the weights that make up each convolutional kernel and applying those one at a time to a sample image. This technique is called feature visualization and it's useful for understanding the inner workings of a CNN.
# + [markdown] id="W2YkkS-8r2M5" colab_type="text"
# In the cell below, you can see how to extract a single filter (by index) from your first convolutional layer. The filter should appear as a grayscale grid.
# + id="hcYDG5GTr2M5" colab_type="code" outputId="1c3c1860-c2f4-4ad0-b904-855df1e93b1f" executionInfo={"status": "ok", "timestamp": 1567440894872, "user_tz": -480, "elapsed": 2015280, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07513880639336204123"}} colab={"base_uri": "https://localhost:8080/", "height": 388}
net.cpu()
# Get the weights in the first conv layer, "conv1"
# if necessary, change this to reflect the name of your first conv layer
weights1 = net.conv1.weight.data
w = weights1.numpy()
print(w.shape)
filter_index = 31
print(w[filter_index][0])
print(w[filter_index][0].shape)
# display the filter weights
plt.imshow(w[filter_index][0], cmap='gray')
# + [markdown] id="IKwdAv7ar2M6" colab_type="text"
# ## Feature maps
#
# Each CNN has at least one convolutional layer that is composed of stacked filters (also known as convolutional kernels). As a CNN trains, it learns what weights to include in it's convolutional kernels and when these kernels are applied to some input image, they produce a set of **feature maps**. So, feature maps are just sets of filtered images; they are the images produced by applying a convolutional kernel to an input image. These maps show us the features that the different layers of the neural network learn to extract. For example, you might imagine a convolutional kernel that detects the vertical edges of a face or another one that detects the corners of eyes. You can see what kind of features each of these kernels detects by applying them to an image. One such example is shown below; from the way it brings out the lines in an the image, you might characterize this as an edge detection filter.
#
# <img src='https://drive.google.com/uc?id=1qZIKoowtryL-a0kA8Q33du_6pna2Lxiy' width=50% height=50%/>
#
#
# Next, choose a test image and filter it with one of the convolutional kernels in your trained CNN; look at the filtered output to get an idea what that particular kernel detects.
#
# ### TODO: Filter an image to see the effect of a convolutional kernel
# ---
# + id="5Gnpc-r3r2M7" colab_type="code" colab={}
##TODO: load in and display any image from the transformed test dataset
## TODO: Using cv's filter2D function,
## apply a specific set of filter weights (like the one displayed above) to the test image
sample = next(iter(test_loader))
# + id="Z2jVKDF4g7I1" colab_type="code" colab={}
idx = 10
images = sample['image']
key_pts = sample['keypoints']
# + id="tzXDKwm_hW1i" colab_type="code" colab={}
test_image = images[idx].numpy()
test_image = np.transpose(test_image, (1, 2, 0))
test_image = np.squeeze(test_image)
# + id="aOPEp3rirC1x" colab_type="code" outputId="5a71f124-7732-4196-c9e9-b877b28316c3" executionInfo={"status": "ok", "timestamp": 1567440898147, "user_tz": -480, "elapsed": 2018507, "user": {"displayName": "nazrul ismail", "photoUrl": "", "userId": "07513880639336204123"}} colab={"base_uri": "https://localhost:8080/", "height": 286}
img = np.squeeze(test_images[idx].data.numpy())
plt.imshow(img, cmap="gray")
# + id="-q7amFZ_rLiM" colab_type="code" outputId="f2dc3f5b-397a-4bd0-8598-94cad93f5241" executionInfo={"status": "ok", "timestamp": 1567440898148, "user_tz": -480, "elapsed": 2018497, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07513880639336204123"}} colab={"base_uri": "https://localhost:8080/", "height": 286}
import cv2
filtered_img = cv2.filter2D(img, -1, w[filter_index][0])
plt.imshow(filtered_img, cmap="gray")
# + [markdown] id="N5CEhYtUr2M8" colab_type="text"
# ### Question 4: Choose one filter from your trained CNN and apply it to a test image; what purpose do you think it plays? What kind of feature do you think it detects?
#
# + [markdown] id="x_Ms22Ver2M9" colab_type="text"
# **Answer**: Conv1 seems to learn to blur images
# + [markdown] id="mAEmvYKbr2M9" colab_type="text"
# ---
# ## Moving on!
#
# Now that you've defined and trained your model (and saved the best model), you are ready to move on to the last notebook, which combines a face detector with your saved model to create a facial keypoint detection system that can predict the keypoints on *any* face in an image!
|
P1_Facial_Keypoints/2. Define the Network Architecture.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# TreeNode{val: 1, left: None, right: None} None
#
# TreeNode{val: 3, left: TreeNode{val: 1, left: None, right: None},
# right: TreeNode{val: 4, left: TreeNode{val: 2, left: None, right: None}, right: None}}
# TreeNode{val: 4, left: TreeNode{val: 2, left: None, right: None}, right: None}
#
#
#
# TreeNode{val: 2, left: None, right: None} None
# TreeNode{val: 4, left: TreeNode{val: 2, left: None, right: None}, right: None} None
# TreeNode{val: 3, left: TreeNode{val: 1, left: None, right: None}, right: TreeNode{val: 4, left: TreeNode{val: 2, left: None, right: None}, right: None}}
#
# TreeNode{val: 4, left: TreeNode{val: 2, left: None, right: None}, right: None}
# TreeNode{val: 2, left: None, right: None} None
# TreeNode{val: 4, left: TreeNode{val: 2, left: None, right: None}, right: None} None
#
# +
class TreeNode:
def __init__(self, x):
self.val = x
self.left = None
self.right = None
class Solution:
def recoverTree(self, root: TreeNode) -> None:
def dfs(node):
nonlocal fir, sec, prev
if not node:
return
dfs(node.left)
if node.val >= prev.val:
prev = node
else:
if not fir:
fir = pre
sec = node
pre = node
else:
sec = node
dfs(node.right)
fir, sec = None, None
prev = TreeNode(-float('inf'))
dfs(root)
fir.val, sec.val = sec.val, fir.val
# -
# +
# Definition for a binary tree node.
class TreeNode:
def __init__(self, x):
self.val = x
self.left = None
self.right = None
class Solution:
def recoverTree(self, root: TreeNode) -> None:
"""
Do not return anything, modify root in-place instead.
"""
firstNode = None
secondNode = None
pre = TreeNode(float("-inf"))
stack = []
p = root
while p or stack:
while p:
stack.append(p)
p = p.left
p = stack.pop()
if not firstNode and pre.val > p.val:
firstNode = pre
if firstNode and pre.val > p.val:
#print(firstNode.val,pre.val, p.val)
secondNode = p
pre = p
p = p.right
firstNode.val, secondNode.val = secondNode.val, firstNode.val
# +
# Definition for a binary tree node.
class TreeNode:
def __init__(self, x):
self.val = x
self.left = None
self.right = None
class Solution:
def recoverTree(self, root: TreeNode) -> None:
self.firstNode = None
self.secondNode = None
self.preNode = TreeNode(float("-inf"))
def in_order(root):
if not root:
return
in_order(root.left)
if self.firstNode == None and self.preNode.val >= root.val:
self.firstNode = self.preNode
if self.firstNode and self.preNode.val >= root.val:
self.secondNode = root
self.preNode = root
in_order(root.right)
in_order(root)
self.firstNode.val, self.secondNode.val = self.secondNode.val, self.firstNode.val
# -
-33 321 55 71 146 231 -13 399
-inf 146 -1 -1
-inf 71 -1 -1
-inf 55 -1 -1
-inf 321 -1 -1
-inf -33 -1 -1
321 -13 -1 146
321 231 -1 146
321 399 -1 -13
pre node fir sec [node >= pre 如果 node.val < pre: 异常点]
-inf -33 # #
-33 321 # #
321 55 # #
321 71 321 55
321 146 321 71
321 231 321 146
321 -13 321 231
321 399 321 -13
# +
321 71 55
321 146 71
321 231 146
321 -13 231
|
Tree/1224/99. Recover Binary Search Tree.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ZrF1N_UoWCek"
# ### Nueral Machine Translation `(NMT)` with Transformers for language understanding.
#
# This notebook is a follow up to the tensorflow [tutorial](https://www.tensorflow.org/text/tutorials/transformer) on NMT. We are going to create a model that will translate Portuguese to English using [this dataset](https://www.tensorflow.org/datasets/catalog/ted_hrlr_translate#ted_hrlr_translatept_to_en)
#
#
# ### Introduction
# The cored idea behind Transformers model is `self-attention` —the ability to attend to different positions of the input sequence to compute a representation of that sequence. Transformer creates stacks of self-attention layers and is explained below in the sections Scaled dot product attention and Multi-head attention.
#
# A transformer model handles variable-sized input using stacks of self-attention layers instead of RNNs or CNNs. This general architecture has a number of advantages:
# * It makes no assumptions about the temporal/spatial relationships across the data.
# * Distant items can affect each other's output without passing through many RNN-steps, or convolution layers (see [Scene Memory Transformer](https://arxiv.org/pdf/1903.03878.pdf) for example).
# * It can learn long-range dependencies. This is a challenge in many sequence tasks.
#
#
# The downsides of this architecture are:
# * For a time-series, the output for a time-step is calculated from the entire history instead of only the inputs and current hidden-state. This may be less efficient.
# * If the input does have a temporal/spatial relationship, like text, some positional encoding must be added or the model will effectively see a bag of words.
#
# ### Setup
#
# + colab={"base_uri": "https://localhost:8080/"} id="344J2ke9VtKK" outputId="208cce20-cd44-4360-db99-f3a25ad1d2bf"
# !pip install tensorflow_datasets
# !pip install -U tensorflow-text
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="1XvtAV7uV82J" outputId="139bcc64-7d05-4267-ae78-16b621460a3e"
import tensorflow as tf
from tensorflow import keras
import tensorflow_datasets as tfds
import tensorflow_text as text
import string, sys, os, time, collections, pathlib, re
import numpy as np
import matplotlib.pyplot as plt
tf.__version__
# + [markdown] id="F4gjvn_lYF88"
# ### Downloading the dataset using `tfds`.
#
# We are going to load the Portuguese-English translation dataset from the TED Talks Open Translation Project.
#
# This dataset contains approximately 50000 training examples, 1100 validation examples, and 2000 test examples.
# + id="avXXjt3OV8zJ"
examples, metadata = tfds.load('ted_hrlr_translate/pt_to_en', with_info=True,
as_supervised=True)
train_examples, val_examples = examples['train'], examples['validation']
# + colab={"base_uri": "https://localhost:8080/"} id="7U7C19-4eQka" outputId="11ff7187-3720-431c-d682-94714de52c1f"
train_examples
# + [markdown] id="sx8Oe8FPYxQY"
# The ``tf.data.Dataset`` object returned by TensorFlow datasets yields pairs of text examples:
# + id="xtbjz0qrZxY4"
from prettytable import PrettyTable
def tabulate(column_names, data):
table = PrettyTable(column_names)
table.align[column_names[0]] = 'l'
table.align[column_names[1]] = 'l'
table._max_width = {column_names[0] :50, column_names[1]: 50}
for row in data:
table.add_row(row)
print(table)
# + colab={"base_uri": "https://localhost:8080/"} id="cXSU13e1V8v7" outputId="85229893-fd5c-4721-c1d0-15411c616b4a"
pairs = []
for pt_examples, en_examples in train_examples.batch(3).take(1):
for pt, en in zip(pt_example.numpy(), en_example.numpy()):
pairs.append((pt.decode('utf-8'), en.decode('utf-8')))
pairs.append(("-"*50, "-"*50 ))
tabulate(("pt", "eng"), pairs)
# + [markdown] id="7HmS-Buwaje5"
# ### Text Tokenization and detokenization
# We can't train a model directly on text. The text needs to be converted to some numeric representation first. Typically, we convert the text to sequences of token IDs, which are used as indices into an embedding.
#
# One popular implementation is demonstrated in the [Subword tokenizertutorial](https://www.tensorflow.org/text/guide/subwords_tokenizer) builds subword tokenizers ([`text.BertTokenizer`](https://www.tensorflow.org/text/api_docs/python/text/BertTokenizer)) optimized for this dataset and exports them in a ``saved_model``.
#
# Download and unzip and import the ``saved_model:``
#
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="q9oZ6nOQV8s1" outputId="0b980ab5-f1b5-4abd-a8b1-0a7a687c4e45"
model_name = "ted_hrlr_translate_pt_en_converter"
keras.utils.get_file(
f"{model_name}.zip",
f"https://storage.googleapis.com/download.tensorflow.org/models/{model_name}.zip",
cache_dir='.', cache_subdir='', extract=True
)
# + id="b1fJCMUKV8qY"
tokenizers = tf.saved_model.load(model_name)
# + [markdown] id="ONEkqxsqbnWx"
# The ``tf.saved_model`` contains two text tokenizers, one for English and one for Portuguese. Both have the same methods:
# + colab={"base_uri": "https://localhost:8080/"} id="KcrBwYpSV8nS" outputId="370e5d90-2b6b-48ab-b5fc-6728d1171975"
[item for item in dir(tokenizers.en) if not item.startswith("_")]
# + colab={"base_uri": "https://localhost:8080/"} id="DoOfv8LRV8hy" outputId="1753ef85-f1c0-4c83-f381-4768abba735e"
tokenizers.en.vocab
# + [markdown] id="1O_8v4yecDnQ"
# The ``tokenize`` method converts a batch of strings to a padded-batch of token IDs. This method splits punctuation, lowercases and unicode-normalizes the input before tokenizing. That standardization is not visible here because the input data is already standardized.
# + colab={"base_uri": "https://localhost:8080/"} id="76hVySSVcDe6" outputId="489317d4-eda6-49f0-9334-f34f1a4693dc"
for en in en_examples.numpy():
print(en.decode('utf-8'))
# + colab={"base_uri": "https://localhost:8080/"} id="60t_KDvAcDcP" outputId="1a7a1454-b8b3-4f0c-f8b4-e7966f1352c3"
encoded = tokenizers.en.tokenize(en_examples)
for row in encoded.to_list():
print(row)
# + [markdown] id="l1tz6HaachUY"
# The ``detokenize`` method attempts to convert these token IDs back to human readable text:
# + colab={"base_uri": "https://localhost:8080/"} id="hYzbu2IjcDZI" outputId="5d5b6fc3-c354-4d98-d904-3ab44aafa502"
round_trip = tokenizers.en.detokenize(encoded)
for line in round_trip.numpy():
print(line.decode('utf-8'))
# + [markdown] id="uEs70Sifcstx"
# The lower level ``lookup`` method converts from token-IDs to token text:
# + colab={"base_uri": "https://localhost:8080/"} id="Sl-rrmpncDVr" outputId="57dd9cbf-fa75-41e3-d1be-51d378f957fb"
tokens = tokenizers.en.lookup(encoded)
tokens
# + [markdown] id="3QtB5gt9c4jZ"
# Here you can see the "subword" aspect of the tokenizers. The word "searchability" is decomposed into "search ##ability" and the word "serendipity" into "s ##ere ##nd ##ip ##ity"
#
# ### Input pipeline.
# To build an input pipeline suitable for training we'll apply some transformations to the dataset.
#
# This function will be used to encode the batches of raw text:
# + id="HNyAOW5icDSx"
def tokenize_pairs(pt, en):
pt = tokenizers.pt.tokenize(pt)
# Convert from ragged to dense, padding with zeros.
pt = pt.to_tensor()
en = tokenizers.en.tokenize(en)
# Convert from ragged to dense, padding with zeros.
en = en.to_tensor()
return pt, en
# + [markdown] id="RBH9OeRVdc2o"
# Here's a simple input pipeline that processes, shuffles and batches the data:
# + id="45eqid5GcDPp"
BUFFER_SIZE = 20000
BATCH_SIZE = 64
# + id="hkuMh4IdcDGi"
def make_batches(ds):
return (
ds
.cache()
.shuffle(BUFFER_SIZE)
.batch(BATCH_SIZE)
.map(tokenize_pairs, num_parallel_calls=tf.data.AUTOTUNE)
.prefetch(tf.data.AUTOTUNE)
)
train_batches = make_batches(train_examples)
val_batches = make_batches(val_examples)
# + [markdown] id="tdovhB9hhD9z"
# ### Positional encoding.
#
# Attention layers see their input as a set of vectors, with no sequential order. This model also doesn't contain any recurrent or convolutional layers. Because of this a "positional encoding" is added to give the model some information about the relative position of the tokens in the sentence.
#
# The positional encoding vector is added to the embedding vector. Embeddings represent a token in a d-dimensional space where tokens with similar meaning will be closer to each other. But the embeddings do not encode the relative position of tokens in a sentence. So after adding the positional encoding, tokens will be closer to each other based on the similarity of their meaning and their position in the sentence, in the d-dimensional space.
#
# The formula for calculating the positional encoding is as follows:
#
#
# $$\Large{PE_{(pos, 2i)} = \sin(pos / 10000^{2i / d_{model}})} $$
# $$\Large{PE_{(pos, 2i+1)} = \cos(pos / 10000^{2i / d_{model}})} $$
#
#
# + id="GIifuSgKV8fI"
def get_angles(pos, i, d_model):
angle_rates = 1 / np.power(10000, (2 * (i//2)) / np.float32(d_model))
return pos * angle_rates
# + id="iUfor47hV8b6"
def positional_encoding(position, d_model):
angle_rads = get_angles(np.arange(position)[:, np.newaxis],
np.arange(d_model)[np.newaxis, :],
d_model)
# apply sin to even indices in the array; 2i
angle_rads[:, 0::2] = np.sin(angle_rads[:, 0::2])
# apply cos to odd indices in the array; 2i+1
angle_rads[:, 1::2] = np.cos(angle_rads[:, 1::2])
pos_encoding = angle_rads[np.newaxis, ...]
return tf.cast(pos_encoding, dtype=tf.float32)
# + colab={"base_uri": "https://localhost:8080/", "height": 300} id="34xOPQ_Hibra" outputId="ab9d3e03-6f61-4389-a972-ad9d9e40aea8"
n, d = 2048, 512
pos_encoding = positional_encoding(n, d)
print(pos_encoding.shape)
pos_encoding = pos_encoding[0]
# Juggle the dimensions for the plot
pos_encoding = tf.reshape(pos_encoding, (n, d//2, 2))
pos_encoding = tf.transpose(pos_encoding, (2, 1, 0))
pos_encoding = tf.reshape(pos_encoding, (d, n))
plt.pcolormesh(pos_encoding, cmap='RdBu')
plt.ylabel('Depth')
plt.xlabel('Position')
plt.colorbar()
plt.show()
# + [markdown] id="SCyuwNlSig3R"
# ### Masking
# Mask all the pad tokens in the batch of sequence. It ensures that the model does not treat padding as the input. The mask indicates where pad value ``0`` is present: it outputs a ``1`` at those locations, and a ``0`` otherwise.
# + id="mKKlpiKmidW0"
def create_padding_mask(seq):
seq = tf.cast(tf.math.equal(seq, 0), tf.float32)
# add extra dimensions to add the padding
# to the attention logits.
return seq[:, tf.newaxis, tf.newaxis, :] # (batch_size, 1, 1, seq_len)
# + colab={"base_uri": "https://localhost:8080/"} id="r6AaP7vqi5ay" outputId="78597594-5b86-49cf-bb76-c0d6d672ad12"
x = tf.constant([[7, 6, 0, 0, 1], [1, 2, 3, 0, 0], [0, 0, 0, 4, 5]])
create_padding_mask(x)
# + [markdown] id="n3ROQsFgi-Sr"
# The look-ahead mask is used to mask the future tokens in a sequence. In other words, the mask indicates which entries should not be used.
#
# This means that to predict the third token, only the first and second token will be used. Similarly to predict the fourth token, only the first, second and the third tokens will be used and so on.
# + id="cj4DlJq5i5Wo"
def create_look_ahead_mask(size):
mask = 1 - tf.linalg.band_part(tf.ones((size, size)), -1, 0)
return mask # (seq_len, seq_len)
# + colab={"base_uri": "https://localhost:8080/"} id="VcSPwLMKi5UV" outputId="69abdd9b-2b8c-47ff-a897-c44a32640b26"
x = tf.random.uniform((1, 3))
temp = create_look_ahead_mask(x.shape[1])
temp
# + [markdown] id="mJSyDUJ9mHzp"
# ### Scaled dot product attention
#
# 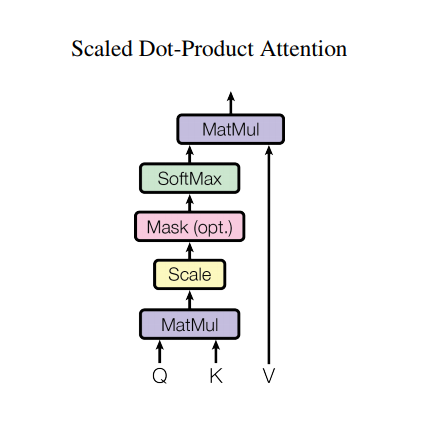
#
# The attention function used by the transformer takes three inputs: Q (query), K (key), V (value). The equation used to calculate the attention weights is:
#
# $$\Large{Attention(Q, K, V) = softmax_k\left(\frac{QK^T}{\sqrt{d_k}}\right) V} $$
#
# The dot-product attention is scaled by a factor of square root of the depth. This is done because for large values of depth, the dot product grows large in magnitude pushing the softmax function where it has small gradients resulting in a very hard softmax.
#
# For example, consider that `Q` and `K` have a mean of 0 and variance of 1. Their matrix multiplication will have a mean of 0 and variance of `dk`. So the *square root of `dk`* is used for scaling, so you get a consistent variance regardless of the value of `dk`. If the variance is too low the output may be too flat to optimize effectively. If the variance is too high the softmax may saturate at initialization making it difficult to learn.
#
# The mask is multiplied with -1e9 (close to negative infinity). This is done because the mask is summed with the scaled matrix multiplication of Q and K and is applied immediately before a softmax. The goal is to zero out these cells, and large negative inputs to softmax are near zero in the output.
#
#
# + id="ls1Y_BfKi5QS"
def scaled_dot_product_attention(q, k, v, mask):
"""Calculate the attention weights.
q, k, v must have matching leading dimensions.
k, v must have matching penultimate dimension, i.e.: seq_len_k = seq_len_v.
The mask has different shapes depending on its type(padding or look ahead)
but it must be broadcastable for addition.
Args:
q: query shape == (..., seq_len_q, depth)
k: key shape == (..., seq_len_k, depth)
v: value shape == (..., seq_len_v, depth_v)
mask: Float tensor with shape broadcastable
to (..., seq_len_q, seq_len_k). Defaults to None.
Returns:
output, attention_weights
"""
matmul_qk = tf.matmul(q, k, transpose_b=True) # (..., seq_len_q, seq_len_k)
# scale matmul_qk
dk = tf.cast(tf.shape(k)[-1], tf.float32)
scaled_attention_logits = matmul_qk / tf.math.sqrt(dk)
# add the mask to the scaled tensor.
if mask is not None:
scaled_attention_logits += (mask * -1e9)
# softmax is normalized on the last axis (seq_len_k) so that the scores
# add up to 1.
attention_weights = tf.nn.softmax(scaled_attention_logits, axis=-1) # (..., seq_len_q, seq_len_k)
output = tf.matmul(attention_weights, v) # (..., seq_len_q, depth_v)
return output, attention_weights
# + [markdown] id="EgwC1781nITx"
# As the softmax normalization is done on ``K``, its values decide the amount of importance given to Q.
#
# The output represents the multiplication of the attention weights and the V (value) vector. This ensures that the tokens you want to focus on are kept as-is and the irrelevant tokens are flushed out.
# + id="C0jdbhVbi5Ni"
def print_out(q, k, v):
temp_out, temp_attn = scaled_dot_product_attention(
q, k, v, None)
print('Attention weights are:')
print(temp_attn)
print('Output is:')
print(temp_out)
# + colab={"base_uri": "https://localhost:8080/"} id="vnNCmzzai5Lb" outputId="7c10c47a-dfea-402d-9118-9504b789128b"
np.set_printoptions(suppress=True)
temp_k = tf.constant([[10, 0, 0],
[0, 10, 0],
[0, 0, 10],
[0, 0, 10]], dtype=tf.float32) # (4, 3)
temp_v = tf.constant([[1, 0],
[10, 0],
[100, 5],
[1000, 6]], dtype=tf.float32) # (4, 2)
# This `query` aligns with the second `key`,
# so the second `value` is returned.
temp_q = tf.constant([[0, 10, 0]], dtype=tf.float32) # (1, 3)
print_out(temp_q, temp_k, temp_v)
# + colab={"base_uri": "https://localhost:8080/"} id="0pPIcBCni5Hx" outputId="b7746418-2d48-48d7-874e-d73911f3fa0d"
# This query aligns with a repeated key (third and fourth),
# so all associated values get averaged.
temp_q = tf.constant([[0, 0, 10]], dtype=tf.float32) # (1, 3)
print_out(temp_q, temp_k, temp_v)
# + colab={"base_uri": "https://localhost:8080/"} id="gJlsPLk8i5E1" outputId="5aa62f82-da8d-413b-fe90-b47d827454f3"
# This query aligns equally with the first and second key,
# so their values get averaged.
temp_q = tf.constant([[10, 10, 0]], dtype=tf.float32) # (1, 3)
print_out(temp_q, temp_k, temp_v)
# + [markdown] id="rJKBOwifnXTY"
# Pass all the queries together.
# + colab={"base_uri": "https://localhost:8080/"} id="mXdnG2mVnUnJ" outputId="74bd6d7b-a2b4-4dba-ad86-c91a4f9d599c"
temp_q = tf.constant([[0, 0, 10],
[0, 10, 0],
[10, 10, 0]], dtype=tf.float32) # (3, 3)
print_out(temp_q, temp_k, temp_v)
# + [markdown] id="PK8Dupv3naKx"
# ### Multi-head attention
#
# <img src="https://www.tensorflow.org/images/tutorials/transformer/multi_head_attention.png" width="500" alt="multi-head attention">
#
#
# Multi-head attention consists of four parts:
# * Linear layers.
# * Scaled dot-product attention.
# * Final linear layer.
#
#
# Each multi-head attention block gets three inputs; Q (query), K (key), V (value). These are put through linear (Dense) layers before the multi-head attention function.
#
# In the diagram above `(K,Q,V)` are passed through sepearte linear (`Dense`) layers for each attention head. For simplicity/efficiency the code below implements this using a single dense layer with `num_heads` times as many outputs. The output is rearranged to a shape of `(batch, num_heads, ...)` before applying the attention function.
#
# The `scaled_dot_product_attention` function defined above is applied in a single call, broadcasted for efficiency. An appropriate mask must be used in the attention step. The attention output for each head is then concatenated (using `tf.transpose`, and `tf.reshape`) and put through a final `Dense` layer.
#
# Instead of one single attention head, Q, K, and V are split into multiple heads because it allows the model to jointly attend to information from different representation subspaces at different positions. After the split each head has a reduced dimensionality, so the total computation cost is the same as a single head attention with full dimensionality.
# + id="XULAkG8CnY8A"
class MultiHeadAttention(tf.keras.layers.Layer):
def __init__(self, d_model, num_heads):
super(MultiHeadAttention, self).__init__()
self.num_heads = num_heads
self.d_model = d_model
assert d_model % self.num_heads == 0
self.depth = d_model // self.num_heads
self.wq = tf.keras.layers.Dense(d_model)
self.wk = tf.keras.layers.Dense(d_model)
self.wv = tf.keras.layers.Dense(d_model)
self.dense = tf.keras.layers.Dense(d_model)
def split_heads(self, x, batch_size):
"""Split the last dimension into (num_heads, depth).
Transpose the result such that the shape is (batch_size, num_heads, seq_len, depth)
"""
x = tf.reshape(x, (batch_size, -1, self.num_heads, self.depth))
return tf.transpose(x, perm=[0, 2, 1, 3])
def call(self, v, k, q, mask):
batch_size = tf.shape(q)[0]
q = self.wq(q) # (batch_size, seq_len, d_model)
k = self.wk(k) # (batch_size, seq_len, d_model)
v = self.wv(v) # (batch_size, seq_len, d_model)
q = self.split_heads(q, batch_size) # (batch_size, num_heads, seq_len_q, depth)
k = self.split_heads(k, batch_size) # (batch_size, num_heads, seq_len_k, depth)
v = self.split_heads(v, batch_size) # (batch_size, num_heads, seq_len_v, depth)
# scaled_attention.shape == (batch_size, num_heads, seq_len_q, depth)
# attention_weights.shape == (batch_size, num_heads, seq_len_q, seq_len_k)
scaled_attention, attention_weights = scaled_dot_product_attention(
q, k, v, mask)
scaled_attention = tf.transpose(scaled_attention, perm=[0, 2, 1, 3]) # (batch_size, seq_len_q, num_heads, depth)
concat_attention = tf.reshape(scaled_attention,
(batch_size, -1, self.d_model)) # (batch_size, seq_len_q, d_model)
output = self.dense(concat_attention) # (batch_size, seq_len_q, d_model)
return output, attention_weights
# + [markdown] id="3OVsS1LsnyFq"
# Create a `MultiHeadAttention` layer to try out. At each location in the sequence, `y`, the `MultiHeadAttention` runs all 8 attention heads across all other locations in the sequence, returning a new vector of the same length at each location.
# + colab={"base_uri": "https://localhost:8080/"} id="xGVKwW0qnvh3" outputId="5bccefd5-c30b-4328-fd07-39348ee6d87d"
temp_mha = MultiHeadAttention(d_model=512, num_heads=8)
y = tf.random.uniform((1, 60, 512)) # (batch_size, encoder_sequence, d_model)
out, attn = temp_mha(y, k=y, q=y, mask=None)
out.shape, attn.shape
# + [markdown] id="j04TBTNPnx1Q"
# ## Point wise feed forward network"
# Point wise feed forward network consists of two fully-connected layers with a ReLU activation in between.
# + id="IvHVfAshnxzm"
def point_wise_feed_forward_network(d_model, dff):
return tf.keras.Sequential([
tf.keras.layers.Dense(dff, activation='relu'), # (batch_size, seq_len, dff)
tf.keras.layers.Dense(d_model) # (batch_size, seq_len, d_model)
])
# + colab={"base_uri": "https://localhost:8080/"} id="mcbp0iMgnxvR" outputId="<PASSWORD>"
sample_ffn = point_wise_feed_forward_network(512, 2048)
sample_ffn(tf.random.uniform((64, 50, 512))).shape
# + [markdown] id="CKkoytyEoBRO"
# ###Encoder Decoder Model
# 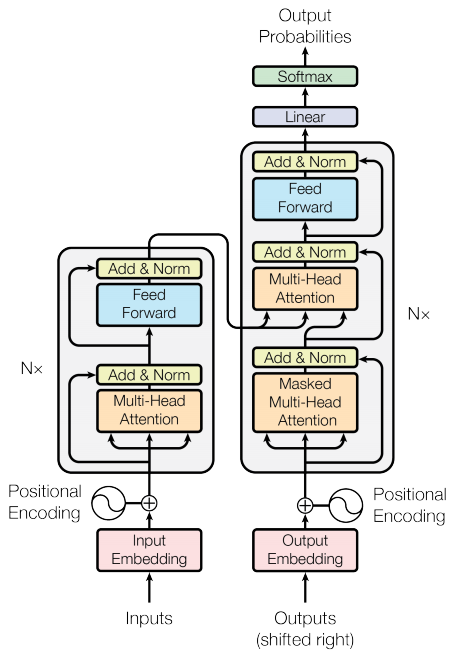
#
# The transformer model follows the same general pattern as a standard [sequence to sequence with attention model](https://www.tensorflow.org/text/tutorials/nmt_with_attention.ipynb).
#
# * The input sentence is passed through `N` encoder layers that generates an output for each token in the sequence.
# * The decoder attends to the encoder's output and its own input (self-attention) to predict the next word.
#
# ### Encoder Layer
# Each encoder layer consists of sublayers:
#
# 1. Multi-head attention (with padding mask)
# 2. Point wise feed forward networks.
#
# Each of these sublayers has a residual connection around it followed by a layer normalization. Residual connections help in avoiding the vanishing gradient problem in deep networks.
#
# The output of each sublayer is `LayerNorm(x + Sublayer(x))`. The normalization is done on the `d_model` (last) axis. There are N encoder layers in the transformer.
# + id="aTItri-boEcf"
class EncoderLayer(keras.layers.Layer):
def __init__(self, d_model, num_heads, dff, rate=0.1):
super(EncoderLayer, self).__init__()
self.mha = MultiHeadAttention(d_model, num_heads)
self.ffn = point_wise_feed_forward_network(d_model, dff)
self.layernorm1 = keras.layers.LayerNormalization(epsilon=1e-6)
self.layernorm2 = keras.layers.LayerNormalization(epsilon=1e-6)
self.dropout1 = keras.layers.Dropout(rate)
self.dropout2 = tf.keras.layers.Dropout(rate)
def call(self, x, training, mask):
attn_output, _ = self.mha(x, x, x, mask) # (batch_size, input_seq_len, d_model)
attn_output = self.dropout1(attn_output, training=training)
out1 = self.layernorm1(x + attn_output) # (batch_size, input_seq_len, d_model)
ffn_output = self.ffn(out1) # (batch_size, input_seq_len, d_model)
ffn_output = self.dropout2(ffn_output, training=training)
out2 = self.layernorm2(out1 + ffn_output) # (batch_size, input_seq_len, d_model)
return out2
# + colab={"base_uri": "https://localhost:8080/"} id="bSEEklBFooUM" outputId="a3e37f40-d229-4900-f7cb-c58383542a49"
sample_encoder_layer = EncoderLayer(512, 8, 2048)
sample_encoder_layer_output = sample_encoder_layer(
tf.random.uniform((64, 43, 512)), False, None)
sample_encoder_layer_output.shape # (batch_size, input_seq_len, d_model)
# + [markdown] id="iPTxuEUToBJB"
# ### Decoder layer
#
# Each decoder layer consists of sublayers:
#
# 1. Masked multi-head attention (with look ahead mask and padding mask)
# 2. Multi-head attention (with padding mask). V (value) and K (key) receive the *encoder output* as inputs. Q (query) receives the *output from the masked multi-head attention sublayer.*
# 3. Point wise feed forward networks
#
# Each of these sublayers has a residual connection around it followed by a layer normalization. The output of each sublayer is `LayerNorm(x + Sublayer(x))`. The normalization is done on the `d_model` (last) axis.
#
# There are N decoder layers in the transformer.
#
# As Q receives the output from decoder's first attention block, and K receives the encoder output, the attention weights represent the importance given to the decoder's input based on the encoder's output. In other words, the decoder predicts the next token by looking at the encoder output and self-attending to its own output. See the demonstration above in the scaled dot product attention section.
# + id="9HhMBgOTnxsU"
class DecoderLayer(tf.keras.layers.Layer):
def __init__(self, d_model, num_heads, dff, rate=0.1):
super(DecoderLayer, self).__init__()
self.mha1 = MultiHeadAttention(d_model, num_heads)
self.mha2 = MultiHeadAttention(d_model, num_heads)
self.ffn = point_wise_feed_forward_network(d_model, dff)
self.layernorm1 = tf.keras.layers.LayerNormalization(epsilon=1e-6)
self.layernorm2 = tf.keras.layers.LayerNormalization(epsilon=1e-6)
self.layernorm3 = tf.keras.layers.LayerNormalization(epsilon=1e-6)
self.dropout1 = tf.keras.layers.Dropout(rate)
self.dropout2 = tf.keras.layers.Dropout(rate)
self.dropout3 = tf.keras.layers.Dropout(rate)
def call(self, x, enc_output, training,
look_ahead_mask, padding_mask):
# enc_output.shape == (batch_size, input_seq_len, d_model)
attn1, attn_weights_block1 = self.mha1(x, x, x, look_ahead_mask) # (batch_size, target_seq_len, d_model)
attn1 = self.dropout1(attn1, training=training)
out1 = self.layernorm1(attn1 + x)
attn2, attn_weights_block2 = self.mha2(
enc_output, enc_output, out1, padding_mask) # (batch_size, target_seq_len, d_model)
attn2 = self.dropout2(attn2, training=training)
out2 = self.layernorm2(attn2 + out1) # (batch_size, target_seq_len, d_model)
ffn_output = self.ffn(out2) # (batch_size, target_seq_len, d_model)
ffn_output = self.dropout3(ffn_output, training=training)
out3 = self.layernorm3(ffn_output + out2) # (batch_size, target_seq_len, d_model)
return out3, attn_weights_block1, attn_weights_block2
# + colab={"base_uri": "https://localhost:8080/"} id="_WDfr8JBoy7h" outputId="f8246a59-da7b-446a-e9f0-d732e9650f79"
sample_decoder_layer = DecoderLayer(512, 8, 2048)
sample_decoder_layer_output, _, _ = sample_decoder_layer(
tf.random.uniform((64, 50, 512)), sample_encoder_layer_output,
False, None, None)
sample_decoder_layer_output.shape # (batch_size, target_seq_len, d_model)
# + [markdown] id="Qu_xjuBto3sD"
# ### Encoder
#
# The `Encoder` consists of:
# 1. Input Embedding
# 2. Positional Encoding
# 3. N encoder layers
#
# The input is put through an embedding which is summed with the positional encoding. The output of this summation is the input to the encoder layers. The output of the encoder is the input to the decoder.
# + id="sIVzxBeVo0tY"
class Encoder(tf.keras.layers.Layer):
def __init__(self, num_layers, d_model, num_heads, dff, input_vocab_size,
maximum_position_encoding, rate=0.1):
super(Encoder, self).__init__()
self.d_model = d_model
self.num_layers = num_layers
self.embedding = tf.keras.layers.Embedding(input_vocab_size, d_model)
self.pos_encoding = positional_encoding(maximum_position_encoding,
self.d_model)
self.enc_layers = [EncoderLayer(d_model, num_heads, dff, rate)
for _ in range(num_layers)]
self.dropout = tf.keras.layers.Dropout(rate)
def call(self, x, training, mask):
seq_len = tf.shape(x)[1]
# adding embedding and position encoding.
x = self.embedding(x) # (batch_size, input_seq_len, d_model)
x *= tf.math.sqrt(tf.cast(self.d_model, tf.float32))
x += self.pos_encoding[:, :seq_len, :]
x = self.dropout(x, training=training)
for i in range(self.num_layers):
x = self.enc_layers[i](x, training, mask)
return x # (batch_size, input_seq_len, d_model)
# + colab={"base_uri": "https://localhost:8080/"} id="leBGPlkoo_yt" outputId="4983814f-2c61-4700-f825-70a2cea72683"
sample_encoder = Encoder(num_layers=2, d_model=512, num_heads=8,
dff=2048, input_vocab_size=8500,
maximum_position_encoding=10000)
temp_input = tf.random.uniform((64, 62), dtype=tf.int64, minval=0, maxval=200)
sample_encoder_output = sample_encoder(temp_input, training=False, mask=None)
print(sample_encoder_output.shape) # (batch_size, input_seq_len, d_model)
# + [markdown] id="dAus4nBHpDcb"
# The `Decoder` consists of:
# 1. Output Embedding
# 2. Positional Encoding
# 3. N decoder layers
#
# The target is put through an embedding which is summed with the positional encoding. The output of this summation is the input to the decoder layers. The output of the decoder is the input to the final linear layer.
# + id="QItIS7TrpCKA"
class Decoder(tf.keras.layers.Layer):
def __init__(self, num_layers, d_model, num_heads, dff, target_vocab_size,
maximum_position_encoding, rate=0.1):
super(Decoder, self).__init__()
self.d_model = d_model
self.num_layers = num_layers
self.embedding = tf.keras.layers.Embedding(target_vocab_size, d_model)
self.pos_encoding = positional_encoding(maximum_position_encoding, d_model)
self.dec_layers = [DecoderLayer(d_model, num_heads, dff, rate)
for _ in range(num_layers)]
self.dropout = tf.keras.layers.Dropout(rate)
def call(self, x, enc_output, training,
look_ahead_mask, padding_mask):
seq_len = tf.shape(x)[1]
attention_weights = {}
x = self.embedding(x) # (batch_size, target_seq_len, d_model)
x *= tf.math.sqrt(tf.cast(self.d_model, tf.float32))
x += self.pos_encoding[:, :seq_len, :]
x = self.dropout(x, training=training)
for i in range(self.num_layers):
x, block1, block2 = self.dec_layers[i](x, enc_output, training,
look_ahead_mask, padding_mask)
attention_weights[f'decoder_layer{i+1}_block1'] = block1
attention_weights[f'decoder_layer{i+1}_block2'] = block2
# x.shape == (batch_size, target_seq_len, d_model)
return x, attention_weights
# + colab={"base_uri": "https://localhost:8080/"} id="Szhfec_npHqW" outputId="8f81528b-9a0a-4443-c9e2-1907e6ade385"
sample_decoder = Decoder(num_layers=2, d_model=512, num_heads=8,
dff=2048, target_vocab_size=8000,
maximum_position_encoding=5000)
temp_input = tf.random.uniform((64, 26), dtype=tf.int64, minval=0, maxval=200)
output, attn = sample_decoder(temp_input,
enc_output=sample_encoder_output,
training=False,
look_ahead_mask=None,
padding_mask=None)
output.shape, attn['decoder_layer2_block2'].shape
# + [markdown] id="_1mRPXEgpKGJ"
# ### Creating the Transformer
#
# Transformer consists of the encoder, decoder and a final linear layer. The output of the decoder is the input to the linear layer and its output is returned.
# + id="UNZDHsmUpJbE"
class Transformer(tf.keras.Model):
def __init__(self, num_layers, d_model, num_heads, dff, input_vocab_size,
target_vocab_size, pe_input, pe_target, rate=0.1):
super().__init__()
self.encoder = Encoder(num_layers, d_model, num_heads, dff,
input_vocab_size, pe_input, rate)
self.decoder = Decoder(num_layers, d_model, num_heads, dff,
target_vocab_size, pe_target, rate)
self.final_layer = tf.keras.layers.Dense(target_vocab_size)
def call(self, inputs, training):
# Keras models prefer if you pass all your inputs in the first argument
inp, tar = inputs
enc_padding_mask, look_ahead_mask, dec_padding_mask = self.create_masks(inp, tar)
enc_output = self.encoder(inp, training, enc_padding_mask) # (batch_size, inp_seq_len, d_model)
# dec_output.shape == (batch_size, tar_seq_len, d_model)
dec_output, attention_weights = self.decoder(
tar, enc_output, training, look_ahead_mask, dec_padding_mask)
final_output = self.final_layer(dec_output) # (batch_size, tar_seq_len, target_vocab_size)
return final_output, attention_weights
def create_masks(self, inp, tar):
# Encoder padding mask
enc_padding_mask = create_padding_mask(inp)
# Used in the 2nd attention block in the decoder.
# This padding mask is used to mask the encoder outputs.
dec_padding_mask = create_padding_mask(inp)
# Used in the 1st attention block in the decoder.
# It is used to pad and mask future tokens in the input received by
# the decoder.
look_ahead_mask = create_look_ahead_mask(tf.shape(tar)[1])
dec_target_padding_mask = create_padding_mask(tar)
look_ahead_mask = tf.maximum(dec_target_padding_mask, look_ahead_mask)
return enc_padding_mask, look_ahead_mask, dec_padding_mask
# + colab={"base_uri": "https://localhost:8080/"} id="mcuAXpv3pRBa" outputId="e8089a0b-a910-4510-a221-0d329cc7a672"
sample_transformer = Transformer(
num_layers=2, d_model=512, num_heads=8, dff=2048,
input_vocab_size=8500, target_vocab_size=8000,
pe_input=10000, pe_target=6000)
temp_input = tf.random.uniform((64, 38), dtype=tf.int64, minval=0, maxval=200)
temp_target = tf.random.uniform((64, 36), dtype=tf.int64, minval=0, maxval=200)
fn_out, _ = sample_transformer([temp_input, temp_target], training=False)
fn_out.shape # (batch_size, tar_seq_len, target_vocab_size)
# + [markdown] id="spSueiFopVUU"
# ### Hyper params
#
# To keep this example small and relatively fast, the values for `num_layers, d_model, dff` have been reduced.
#
# The base model described in the [paper](https://arxiv.org/abs/1706.03762) used: `num_layers=6, d_model=512, dff=2048`.
# + id="wLe0nNq5pSok"
num_layers = 4
d_model = 128
dff = 512
num_heads = 8
dropout_rate = 0.1
# + [markdown] id="7eyIKP5QpccZ"
# ### Optimizer
# Use the Adam optimizer with a custom learning rate scheduler according to the formula in the [paper](https://arxiv.org/abs/1706.03762).
#
# $$\Large{lrate = d_{model}^{-0.5} * \min(step{\_}num^{-0.5}, step{\_}num \cdot warmup{\_}steps^{-1.5})}$$
# + id="UiVVh8ChpadS"
class CustomSchedule(keras.optimizers.schedules.LearningRateSchedule):
def __init__(self, d_model, warmup_steps=4000):
super(CustomSchedule, self).__init__()
self.d_model = d_model
self.d_model = tf.cast(self.d_model, tf.float32)
self.warmup_steps = warmup_steps
def __call__(self, step):
arg1 = tf.math.rsqrt(step)
arg2 = step * (self.warmup_steps ** -1.5)
return tf.math.rsqrt(self.d_model) * tf.math.minimum(arg1, arg2)
# + id="3CQKn8evpjiT"
learning_rate = CustomSchedule(d_model)
optimizer = tf.keras.optimizers.Adam(learning_rate, beta_1=0.9, beta_2=0.98,
epsilon=1e-9)
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="nuuA9sXEpm5d" outputId="b9835191-b8da-4355-cb49-497ebdad5eaa"
temp_learning_rate_schedule = CustomSchedule(d_model)
plt.plot(temp_learning_rate_schedule(tf.range(40000, dtype=tf.float32)))
plt.ylabel("Learning Rate")
plt.xlabel("Train Step")
# + [markdown] id="r5zdfaTMprZj"
# ### Loss and metrics
#
# Since the target sequences are padded, it is important to apply a padding mask when calculating the loss.
# + id="ajigzXSzpol0"
loss_object = tf.keras.losses.SparseCategoricalCrossentropy(
from_logits=True, reduction='none')
# + id="864DPLhtpvbg"
def loss_function(real, pred):
mask = tf.math.logical_not(tf.math.equal(real, 0))
loss_ = loss_object(real, pred)
mask = tf.cast(mask, dtype=loss_.dtype)
loss_ *= mask
return tf.reduce_sum(loss_)/tf.reduce_sum(mask)
def accuracy_function(real, pred):
accuracies = tf.equal(real, tf.argmax(pred, axis=2))
mask = tf.math.logical_not(tf.math.equal(real, 0))
accuracies = tf.math.logical_and(mask, accuracies)
accuracies = tf.cast(accuracies, dtype=tf.float32)
mask = tf.cast(mask, dtype=tf.float32)
return tf.reduce_sum(accuracies)/tf.reduce_sum(mask)
# + id="3Uy1jHHDpxJd"
train_loss = tf.keras.metrics.Mean(name='train_loss')
train_accuracy = tf.keras.metrics.Mean(name='train_accuracy')
# + [markdown] id="fNUsxSIxpzy7"
# ### Trainibg and checkpointing
# + id="ZGCO9KvNpzRW"
transformer = Transformer(
num_layers=num_layers,
d_model=d_model,
num_heads=num_heads,
dff=dff,
input_vocab_size=tokenizers.pt.get_vocab_size().numpy(),
target_vocab_size=tokenizers.en.get_vocab_size().numpy(),
pe_input=1000,
pe_target=1000,
rate=dropout_rate)
# + [markdown] id="9Ge1qUaUp61N"
# Create the checkpoint path and the checkpoint manager. This will be used to save checkpoints every `n` epochs.
# + id="kGEPgsQTp4fM"
checkpoint_path = "./checkpoints/train"
ckpt = tf.train.Checkpoint(transformer=transformer,
optimizer=optimizer)
ckpt_manager = tf.train.CheckpointManager(ckpt, checkpoint_path, max_to_keep=5)
# if a checkpoint exists, restore the latest checkpoint.
if ckpt_manager.latest_checkpoint:
ckpt.restore(ckpt_manager.latest_checkpoint)
print('Latest checkpoint restored!!')
# + [markdown] id="-Pv9L4kjp_wW"
# The target is divided into tar_inp and tar_real. tar_inp is passed as an input to the decoder. `tar_real` is that same input shifted by 1: At each location in `tar_input`, `tar_real` contains the next token that should be predicted.
#
# For example, `sentence` = "SOS A lion in the jungle is sleeping EOS"
#
# `tar_inp` = "SOS A lion in the jungle is sleeping"
#
# `tar_real` = "A lion in the jungle is sleeping EOS"
#
# The transformer is an auto-regressive model: it makes predictions one part at a time, and uses its output so far to decide what to do next.
#
# During training this example uses teacher-forcing (like in the [text generation tutorial](https://www.tensorflow.org/text/tutorials/text_generation)). Teacher forcing is passing the true output to the next time step regardless of what the model predicts at the current time step.
#
# As the transformer predicts each token, *self-attention* allows it to look at the previous tokens in the input sequence to better predict the next token.
#
# To prevent the model from peeking at the expected output the model uses a look-ahead mask.
# + id="Q82O0O-1p9rZ"
EPOCHS = 20
# + id="qPHyq5oIqCdB"
# The @tf.function trace-compiles train_step into a TF graph for faster
# execution. The function specializes to the precise shape of the argument
# tensors. To avoid re-tracing due to the variable sequence lengths or variable
# batch sizes (the last batch is smaller), use input_signature to specify
# more generic shapes.
train_step_signature = [
tf.TensorSpec(shape=(None, None), dtype=tf.int64),
tf.TensorSpec(shape=(None, None), dtype=tf.int64),
]
@tf.function(input_signature=train_step_signature)
def train_step(inp, tar):
tar_inp = tar[:, :-1]
tar_real = tar[:, 1:]
with tf.GradientTape() as tape:
predictions, _ = transformer([inp, tar_inp],
training = True)
loss = loss_function(tar_real, predictions)
gradients = tape.gradient(loss, transformer.trainable_variables)
optimizer.apply_gradients(zip(gradients, transformer.trainable_variables))
train_loss(loss)
train_accuracy(accuracy_function(tar_real, predictions))
# + colab={"base_uri": "https://localhost:8080/"} id="BwcUd73IqHQs" outputId="04b8e4e8-1324-4cb6-b6ee-656413ce2dd3"
for epoch in range(EPOCHS):
start = time.time()
train_loss.reset_states()
train_accuracy.reset_states()
# inp -> portuguese, tar -> english
for (batch, (inp, tar)) in enumerate(train_batches):
train_step(inp, tar)
if batch % 50 == 0:
print(f'Epoch {epoch + 1} Batch {batch} Loss {train_loss.result():.4f} Accuracy {train_accuracy.result():.4f}')
if (epoch + 1) % 5 == 0:
ckpt_save_path = ckpt_manager.save()
print(f'Saving checkpoint for epoch {epoch+1} at {ckpt_save_path}')
print(f'Epoch {epoch + 1} Loss {train_loss.result():.4f} Accuracy {train_accuracy.result():.4f}')
print(f'Time taken for 1 epoch: {time.time() - start:.2f} secs\n')
# + [markdown] id="fk1gsZbrqM6S"
# ### Mode Inference
#
# The following steps are used for inference:
#
# * Encode the input sentence using the Portuguese tokenizer (`tokenizers.pt`). This is the encoder input.
# * The decoder input is initialized to the `[START]` token.
# * Calculate the padding masks and the look ahead masks.
# * The `decoder` then outputs the predictions by looking at the `encoder output` and its own output (self-attention).
# * Concatenate the predicted token to the decoder input and pass it to the decoder.
# * In this approach, the decoder predicts the next token based on the previous tokens it predicted.
#
#
# + [markdown] id="FLos50dUqK8v"
# Note: The model is optimized for _efficient training_ and makes a next-token prediction for each token in the output simultaneously. This is redundant during inference, and only the last prediction is used. This model can be made more efficient for inference if you only calculate the last prediction when running in inference mode (`training=False`).
# + id="fpp0b5vUqahr"
class Translator(tf.Module):
def __init__(self, tokenizers, transformer):
self.tokenizers = tokenizers
self.transformer = transformer
def __call__(self, sentence, max_length=20):
# input sentence is portuguese, hence adding the start and end token
assert isinstance(sentence, tf.Tensor)
if len(sentence.shape) == 0:
sentence = sentence[tf.newaxis]
sentence = self.tokenizers.pt.tokenize(sentence).to_tensor()
encoder_input = sentence
# as the target is english, the first token to the transformer should be the
# english start token.
start_end = self.tokenizers.en.tokenize([''])[0]
start = start_end[0][tf.newaxis]
end = start_end[1][tf.newaxis]
# `tf.TensorArray` is required here (instead of a python list) so that the
# dynamic-loop can be traced by `tf.function`.
output_array = tf.TensorArray(dtype=tf.int64, size=0, dynamic_size=True)
output_array = output_array.write(0, start)
for i in tf.range(max_length):
output = tf.transpose(output_array.stack())
predictions, _ = self.transformer([encoder_input, output], training=False)
# select the last token from the seq_len dimension
predictions = predictions[:, -1:, :] # (batch_size, 1, vocab_size)
predicted_id = tf.argmax(predictions, axis=-1)
# concatentate the predicted_id to the output which is given to the decoder
# as its input.
output_array = output_array.write(i+1, predicted_id[0])
if predicted_id == end:
break
output = tf.transpose(output_array.stack())
# output.shape (1, tokens)
text = tokenizers.en.detokenize(output)[0] # shape: ()
tokens = tokenizers.en.lookup(output)[0]
# `tf.function` prevents us from using the attention_weights that were
# calculated on the last iteration of the loop. So recalculate them outside
# the loop.
_, attention_weights = self.transformer([encoder_input, output[:,:-1]], training=False)
return text, tokens, attention_weights
# + id="qOPEHE9pqab4"
translator = Translator(tokenizers, transformer)
# + id="iISHWREAqaWC"
def print_translation(sentence, tokens, ground_truth):
print(f'{"Input:":15s}: {sentence}')
print(f'{"Prediction":15s}: {tokens.numpy().decode("utf-8")}')
print(f'{"Ground truth":15s}: {ground_truth}')
# + colab={"base_uri": "https://localhost:8080/"} id="PBO1ie7uqaTH" outputId="0f7bfb6d-cab4-4141-9baa-ab42241b0a9d"
sentence = "este é um problema que temos que resolver."
ground_truth = "this is a problem we have to solve ."
translated_text, translated_tokens, attention_weights = translator(
tf.constant(sentence))
print_translation(sentence, translated_text, ground_truth)
# + colab={"base_uri": "https://localhost:8080/"} id="oghQLLdUqjpr" outputId="393a1d16-959d-496b-d906-d7219f4e0cd1"
sentence = "os meus vizinhos ouviram sobre esta ideia."
ground_truth = "and my neighboring homes heard about this idea ."
translated_text, translated_tokens, attention_weights = translator(
tf.constant(sentence))
print_translation(sentence, translated_text, ground_truth)
# + colab={"base_uri": "https://localhost:8080/"} id="_S3SyyoIqk5j" outputId="fc55b457-e7e5-493f-ec18-bf1522072df3"
sentence = "vou então muito rapidamente partilhar convosco algumas histórias de algumas coisas mágicas que aconteceram."
ground_truth = "so i \'ll just share with you some stories very quickly of some magical things that have happened ."
translated_text, translated_tokens, attention_weights = translator(
tf.constant(sentence))
print_translation(sentence, translated_text, ground_truth)
# + [markdown] id="7LpD-BcLqm4I"
# ## Attention plots
# The `Translator` class returns a dictionary of attention maps you can use to visualize the internal working of the model:
# + colab={"base_uri": "https://localhost:8080/"} id="1eEbDCC6qnki" outputId="169c7b47-3586-4436-a77e-cd7046330dee"
sentence = "este é o primeiro livro que eu fiz."
ground_truth = "this is the first book i've ever done."
translated_text, translated_tokens, attention_weights = translator(
tf.constant(sentence))
print_translation(sentence, translated_text, ground_truth)
# + id="GncETAxSqtTi"
def plot_attention_head(in_tokens, translated_tokens, attention):
# The plot is of the attention when a token was generated.
# The model didn't generate `<START>` in the output. Skip it.
translated_tokens = translated_tokens[1:]
ax = plt.gca()
ax.matshow(attention)
ax.set_xticks(range(len(in_tokens)))
ax.set_yticks(range(len(translated_tokens)))
labels = [label.decode('utf-8') for label in in_tokens.numpy()]
ax.set_xticklabels(
labels, rotation=90)
labels = [label.decode('utf-8') for label in translated_tokens.numpy()]
ax.set_yticklabels(labels)
# + colab={"base_uri": "https://localhost:8080/"} id="u15CW7zKqvDM" outputId="e906523d-2b4d-463d-d163-d0d236c88cc2"
head = 0
# shape: (batch=1, num_heads, seq_len_q, seq_len_k)
attention_heads = tf.squeeze(
attention_weights['decoder_layer4_block2'], 0)
attention = attention_heads[head]
attention.shape
# + colab={"base_uri": "https://localhost:8080/"} id="dCLebdXdqwPk" outputId="2b87cc12-bb64-45a9-e726-b9438a7e97bd"
in_tokens = tf.convert_to_tensor([sentence])
in_tokens = tokenizers.pt.tokenize(in_tokens).to_tensor()
in_tokens = tokenizers.pt.lookup(in_tokens)[0]
in_tokens
# + colab={"base_uri": "https://localhost:8080/"} id="LTRivI3Kqxqk" outputId="1e1ba7dd-8510-4277-9da6-cae4e54b9589"
translated_tokens
# + colab={"base_uri": "https://localhost:8080/", "height": 301} id="Ayx_SzzPqy_J" outputId="ebeca5c0-c243-42c1-e173-e8d2824fb9fd"
plot_attention_head(in_tokens, translated_tokens, attention)
# + id="gTrwsmwRq05P"
def plot_attention_weights(sentence, translated_tokens, attention_heads):
in_tokens = tf.convert_to_tensor([sentence])
in_tokens = tokenizers.pt.tokenize(in_tokens).to_tensor()
in_tokens = tokenizers.pt.lookup(in_tokens)[0]
in_tokens
fig = plt.figure(figsize=(16, 8))
for h, head in enumerate(attention_heads):
ax = fig.add_subplot(2, 4, h+1)
plot_attention_head(in_tokens, translated_tokens, head)
ax.set_xlabel(f'Head {h+1}')
plt.tight_layout()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 568} id="lg9s1QiFq2cU" outputId="acef9e92-013b-4f90-9c3f-cc67d9503335"
plot_attention_weights(sentence, translated_tokens,
attention_weights['decoder_layer4_block2'][0])
# + [markdown] id="IqZsajmaq4sQ"
# The model does okay on unfamiliar words. Neither "triceratops" or "encyclopedia" are in the input dataset and the model almost learns to transliterate them, even without a shared vocabulary:
# + colab={"base_uri": "https://localhost:8080/", "height": 615} id="POxTxt_lq5DA" outputId="081d6aad-dac7-475a-f4f4-4fdb735c3f64"
sentence = "Eu li sobre triceratops na enciclopédia."
ground_truth = "I read about triceratops in the encyclopedia."
translated_text, translated_tokens, attention_weights = translator(
tf.constant(sentence))
print_translation(sentence, translated_text, ground_truth)
plot_attention_weights(sentence, translated_tokens,
attention_weights['decoder_layer4_block2'][0])
# + [markdown] id="7NswXgybq-h5"
# ### Conclusion
#
# All thanks to [this](https://colab.research.google.com/github/tensorflow/text/blob/master/docs/tutorials/transformer.ipynb#scrollTo=9cxysY7uh3jg) tensorflow tutorial, next we are going to load and create our own dataset insead of using the ``tfds`` dataset and perform the same task.
# + id="aD4hFzXdrQPf"
|
14_NLP/01_Neural_Machine_Translation/10_NMT_Transformers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 그래프
#
# ## 그래프의 정의
# * 정점의 모음과 간선의 모음의 결합
# * 정점의 집합을 V, 간선의 집합을 E, 그리고 그래프를 G라고 했을 때, G=(V,E)이다.
# * 인접(adjacent), 이웃 관계: 간선으로 연결된 두 정점
# * 경로(path): 간선을 통해 이웃 관계가 된 정점들이 그래프 내에서 길(경로)를 이룸
# * 사이클(cycle): 경로 중 정점 하나를 두 번 이상 거치는 경로
# * 방향성 그래프(directed graph): 간선이 방향성을 가지는 그래프
# * 무방향성 그래프(undirected graph): 간선이 방샹성을 가지지 않는 그래프
# * 연결성(connectivity): 무방향성 그래프 내의 두 정점 사이에 경로가 존재하는 경우 "두 정점이 연결되어 있다"라고 한다.
# ## 그래프의 표현
# * 정점: 어떤 자료구조로도 쉽게 표현 가능
# * 결국 그래프의 표현은 간선의 표현 문제로 귀결
# - 인접 행렬(adjacency matrix)
# - 인접 리스트(adjacency list)
#
# | | 장점 | 단점 |
# | --- | --- | --- |
# | 인접 행렬 | 인접 여부를 빠르게 파악 | 메모리소비가 큼 |
# | 인접 리스트 | 메모리 소비가 적음. 정점, 간선의 삽입이 빠름 | 인접 여부를 파악하기 위해 순차 탐색이 필요 |
|
python/graph/note.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # <strong>Road networks and robustness to flooding on US Atlantic and Gulf barrier islands</strong>
# ## <strong>- Statistics -</strong>
# ### This notebook generates the stats included in the manuscript
# +
### Packages
import os
import geopandas as gpd
import pandas as pd
import networkx as nx
import numpy as np
# -
path=''
os.chdir(path)
# +
### Area and shoreline length of the US Atlantic and Gulf barrier islands
# Calculate area (in km2) and shoreline length (in km) for all barriers
barriers= gpd.read_file('./Data/Exceedance/US_barriers.shp')
barriers= barriers.to_crs('esri:102003')
barriers["area"] = barriers['geometry'].area/ 10**6
barriers["length"]= barriers['geometry'].length/10**3
# Filter those that belong to the 72 networks with more than 100 nodes (sampled) and keep the remaining in another df (unsampled)
table = pd.read_csv("./Results/Results_AllBarriers.csv", sep=",", header=0)
sampled_barriers= list(table.Barrier) # >100 nodes
sampled = barriers.query('name in @sampled_barriers')
unsampled = barriers.query('name not in @sampled_barriers') #
number_sampled= len(sampled_barriers) # number of barrier with more than 100 nodes
# Stats
area_sampled_sum=sampled['area'].sum()
area_total_sum=barriers['area'].sum()
area_sampled_mean=sampled['area'].mean()
area_unsampled_mean=unsampled['area'].mean()
area_sampled_100=len(sampled.loc[sampled['area']<100])
area_sampled_100_perc=len(sampled.loc[sampled['area']<100])/number_sampled*100
area_sampled_25=len(sampled.loc[sampled['area']<25])
area_sampled_25_perc=len(sampled.loc[sampled['area']<25])/number_sampled*100
length_sampled_sum=sampled['length'].sum()
length_total_sum=barriers['length'].sum()
# +
### Street length
rootdir = './Results/Statistics'
extensions = ('.csv')
barrier_names=[]
length_street=[]
# Loop through files with statistics and open each csv to retrieve street length
for subdir, dirs, files in os.walk(rootdir):
for file in files:
ext = os.path.splitext(file)[-1].lower()
if ext in extensions:
file_path = os.path.join(subdir, file)
barrier = file.replace(".csv","")
barrier = barrier.replace("_geo","")
barrier_names.append(barrier)
table = pd.read_csv(file_path, sep=",", header=0)
table.rename(columns={ table.columns[0]: "stats", table.columns[1]:"values"}, inplace = True)
length=table.loc[table['stats'] == 'street_length_total', 'values'].iloc[0]/10**3
length_street.append(length)
# Create new dataframe with results and filter those that have more than 100 nodes
df = list(zip(barrier_names, length_street))
df = pd.DataFrame(df, columns=['Barrier','Street_length'])
table = pd.read_csv("./Results/Results_AllBarriers.csv", sep=",", header=0)
sampled_barriers= list(table.Barrier)
sampled = df.query('Barrier in @sampled_barriers')
# Stats
street_length_min=sampled['Street_length'].min()
street_length_max=sampled['Street_length'].max()
street_length_mean=sampled['Street_length'].mean()
street_length_200=len(sampled.loc[sampled['Street_length']>200])
street_length_200_perc=len(sampled.loc[sampled['Street_length']>200])/number_sampled*100
# +
### Number of nodes
rootdir = './Data/Roads'
extensions = ('.graphml')
barrier_names=[]
nodes=[]
# Loop through files and open barrier graphml to retrieve number of nodes in each drivable road network
for subdir, dirs, files in os.walk(rootdir):
for file in files:
ext = os.path.splitext(file)[-1].lower()
if ext in extensions:
file_path = os.path.join(subdir, file)
barrier = file.replace(".graphml","")
barrier_names.append(barrier)
G = nx.read_graphml(file_path)
N= len(G.nodes(data=True))
nodes.append(N)
df = list(zip(barrier_names, nodes))
df= pd.DataFrame(df, columns=['Barrier','Nodes'])
number_drivable=len(df)
# Filter those that have more than 100 nodes (sampled)
table = pd.read_csv("./Results/Results_AllBarriers.csv", sep=",", header=0)
sampled_barriers= list(table.Barrier)
sampled = df.query('Barrier in @sampled_barriers')
# Stats
nodes_min=sampled['Nodes'].min()
nodes_max=sampled['Nodes'].max()
nodes_mean=sampled['Nodes'].mean()
nodes_1000=len(sampled.loc[sampled['Nodes']>1000])
nodes_1000_perc=len(sampled.loc[sampled['Nodes']>1000])/number_sampled*100
# +
### Nodes elevation
rootdir = './Data/Roads'
extensions = ('.graphml')
elevations=[]
barrier_names=[]
# Loop through files and open barrier graphml to retrieve the elevation of each node in each drivable network
for subdir, dirs, files in os.walk(rootdir):
for file in files:
ext = os.path.splitext(file)[-1].lower()
if ext in extensions:
file_path = os.path.join(subdir, file)
barrier = file.replace(".graphml","")
barrier_names.append(barrier)
G = nx.read_graphml(file_path)
N= len(G.nodes(data=True))
# select only those that have more than 100 nodes
if N>100:
df=pd.DataFrame.from_dict(dict(G.nodes(data=True)), orient='index')
Elev= pd.to_numeric(df.Elevations)
Elev=list(Elev.values)
elevations.append(Elev)
merged_list = []
for l in elevations:
merged_list += l
elevations=np.array(merged_list)
# Stats
elevations_mean= elevations.mean()
elevations_1btw3=len(elevations[(elevations>1)&(elevations<3)])/len(elevations)*100
elevations_1=len(elevations[(elevations<1)])/len(elevations)*100
elevations_5=len(elevations[(elevations>5)])/len(elevations)*100
elevations_10=len(elevations[(elevations>10)])/len(elevations)*100
# +
### Critical elevation
table = pd.read_csv("./Results/Results_AllBarriers.csv", sep=",", header=0)
df = table[['Barrier','Critical_elevation']]
# Stats
z_5=len(df.loc[df['Critical_elevation']<5])
z_5_perc=len(df.loc[df['Critical_elevation']<5])/72*100
z_25=len(df.loc[df['Critical_elevation']<2.5])
z_25_perc=len(df.loc[df['Critical_elevation']<2.5])/72*100
z_15=len(df.loc[df['Critical_elevation']<1.5])
z_15_perc=len(df.loc[df['Critical_elevation']<1.5])/72*100
z_1=len(df.loc[df['Critical_elevation']<1])
z_1_perc=len(df.loc[df['Critical_elevation']<1])/72*100
# +
### Critical exceedance
table = pd.read_csv("./Results/Results_AllBarriers.csv", sep=",", header=0)
df = table[['Barrier','Critical_exceedance']]
# Stats
e_100=len(df.loc[df['Critical_exceedance']>0.01])
e_100_perc=len(df.loc[df['Critical_exceedance']>0.01])/72*100
e_10=len(df.loc[df['Critical_exceedance']>0.1])
e_10_perc=len(df.loc[df['Critical_exceedance']>0.1])/72*100
# +
### Critical elevation and critical exceedance
table = pd.read_csv("./Results/Results_AllBarriers.csv", sep=",", header=0)
df = table[['Barrier','Critical_exceedance','Critical_elevation']]
# Stats
e_10=df.loc[df['Critical_exceedance']<10]
e_10_meanz=e_10['Critical_elevation'].mean()
# +
### Robustness
table = pd.read_csv("./Results/Results_AllBarriers.csv", sep=",", header=0)
df = table[['Barrier','Robustness']]
# Stats
R_03btw05= len(df.loc[(df['Robustness']>0.3)&(df['Robustness']<0.4)])
R_03btw05_perc= len(df.loc[(df['Robustness']>0.3)&(df['Robustness']<0.4)])/72*100
R_04= len(df.loc[df['Robustness']>0.4])
R_04_perc= len(df.loc[df['Robustness']>0.4])/72*100
R_045= len(df.loc[df['Robustness']>0.45])
R_045_perc= len(df.loc[df['Robustness']>0.45])/72*100
R_03= len(df.loc[df['Robustness']<0.3])
R_03_perc= len(df.loc[df['Robustness']<0.3])/72*100
R_02= len(df.loc[df['Robustness']<0.2])
R_02_perc= len(df.loc[df['Robustness']<0.2])/72*100
R_max=df.loc[df['Robustness']==df['Robustness'].max()]
R_min=df.loc[df['Robustness']==df['Robustness'].min()]
|
Notebooks/Statistics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="cDpd_7I-1Sd2" colab_type="text"
# # Cross-Entropy Method
#
# ---
#
# In this notebook, we will train the Cross-Entropy Method with OpenAI Gym's MountainCarContinuous environment.
# + [markdown] id="-xVNrbEH1Sd3" colab_type="text"
# ### 1. Import the Necessary Packages
# + id="X-YwbaFf1Sd5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1020} outputId="4c62e1a8-c54d-4b07-f7d6-c540503371c1"
# !pip install gym torch
import gym
import math
import numpy as np
from collections import deque
import matplotlib.pyplot as plt
# %matplotlib inline
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
# + [markdown] id="R4AFxdJb1Sd9" colab_type="text"
# ### 2. Instantiate the Environment and Agent
# + id="wImHCiHu1Sd-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="fc9d72f8-e834-411d-c435-38b6e35242ad"
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
env = gym.make('MountainCarContinuous-v0')
env.seed(101)
np.random.seed(101)
print('observation space:', env.observation_space)
print('action space:', env.action_space)
print(' - low:', env.action_space.low)
print(' - high:', env.action_space.high)
class Agent(nn.Module):
def __init__(self, env, h_size=16):
super(Agent, self).__init__()
self.env = env
# state, hidden layer, action sizes
self.s_size = env.observation_space.shape[0]
self.h_size = h_size
self.a_size = env.action_space.shape[0]
# define layers
self.fc1 = nn.Linear(self.s_size, self.h_size)
self.fc2 = nn.Linear(self.h_size, self.a_size)
def set_weights(self, weights):
s_size = self.s_size
h_size = self.h_size
a_size = self.a_size
# separate the weights for each layer
fc1_end = (s_size*h_size)+h_size
fc1_W = torch.from_numpy(weights[:s_size*h_size].reshape(s_size, h_size))
fc1_b = torch.from_numpy(weights[s_size*h_size:fc1_end])
fc2_W = torch.from_numpy(weights[fc1_end:fc1_end+(h_size*a_size)].reshape(h_size, a_size))
fc2_b = torch.from_numpy(weights[fc1_end+(h_size*a_size):])
# set the weights for each layer
self.fc1.weight.data.copy_(fc1_W.view_as(self.fc1.weight.data))
self.fc1.bias.data.copy_(fc1_b.view_as(self.fc1.bias.data))
self.fc2.weight.data.copy_(fc2_W.view_as(self.fc2.weight.data))
self.fc2.bias.data.copy_(fc2_b.view_as(self.fc2.bias.data))
def get_weights_dim(self):
return (self.s_size+1)*self.h_size + (self.h_size+1)*self.a_size
def forward(self, x):
x = F.relu(self.fc1(x))
x = F.tanh(self.fc2(x))
return x.cpu().data
def evaluate(self, weights, gamma=1.0, max_t=5000):
self.set_weights(weights)
episode_return = 0.0
state = self.env.reset()
for t in range(max_t):
state = torch.from_numpy(state).float().to(device)
action = self.forward(state)
state, reward, done, _ = self.env.step(action)
episode_return += reward * math.pow(gamma, t)
if done:
break
return episode_return
agent = Agent(env).to(device)
# + [markdown] id="HsGAdxji1SeE" colab_type="text"
# ### 3. Train the Agent with the Cross-Entropy Method
#
# Run the code cell below to train the agent from scratch. Alternatively, you can skip to the next code cell to load the pre-trained weights from file.
# + id="-UIf4Bip1SeE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 602} outputId="c410c22a-d889-415c-ce3c-2f4dc4752c99"
def cem(n_iterations=500, max_t=1000, gamma=1.0, print_every=10, pop_size=50, elite_frac=0.2, sigma=0.5):
"""PyTorch implementation of the cross-entropy method.
Params
======
n_iterations (int): maximum number of training iterations
max_t (int): maximum number of timesteps per episode
gamma (float): discount rate
print_every (int): how often to print average score (over last 100 episodes)
pop_size (int): size of population at each iteration
elite_frac (float): percentage of top performers to use in update
sigma (float): standard deviation of additive noise
"""
n_elite=int(pop_size*elite_frac)
scores_deque = deque(maxlen=100)
scores = []
best_weight = sigma*np.random.randn(agent.get_weights_dim())
for i_iteration in range(1, n_iterations+1):
weights_pop = [best_weight + (sigma*np.random.randn(agent.get_weights_dim())) for i in range(pop_size)]
rewards = np.array([agent.evaluate(weights, gamma, max_t) for weights in weights_pop])
elite_idxs = rewards.argsort()[-n_elite:]
elite_weights = [weights_pop[i] for i in elite_idxs]
best_weight = np.array(elite_weights).mean(axis=0)
reward = agent.evaluate(best_weight, gamma=1.0)
scores_deque.append(reward)
scores.append(reward)
torch.save(agent.state_dict(), 'checkpoint.pth')
if i_iteration % print_every == 0:
print('Episode {}\tAverage Score: {:.2f}'.format(i_iteration, np.mean(scores_deque)))
if np.mean(scores_deque)>=90.0:
print('\nEnvironment solved in {:d} iterations!\tAverage Score: {:.2f}'.format(i_iteration-100, np.mean(scores_deque)))
break
return scores
scores = cem()
# plot the scores
fig = plt.figure()
ax = fig.add_subplot(111)
plt.plot(np.arange(1, len(scores)+1), scores)
plt.ylabel('Score')
plt.xlabel('Episode #')
plt.show()
# + [markdown] id="O_BipjVz1SeI" colab_type="text"
# ### 4. Watch a Smart Agent!
#
# In the next code cell, you will load the trained weights from file to watch a smart agent!
# + id="586X51tH1SeJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="576c2b16-c59e-41d6-935c-205dc5ffc38a"
# load the weights from file
agent.load_state_dict(torch.load('checkpoint.pth'))
state = env.reset()
while True:
state = torch.from_numpy(state).float().to(device)
with torch.no_grad():
action = agent(state)
# env.render()
next_state, reward, done, _ = env.step(action)
state = next_state
if done:
break
env.close()
|
cross-entropy/CEM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <p><font size="6"><b> Case study: air quality data of European monitoring stations (AirBase)</b></font></p><br>
#
# > *DS Data manipulation, analysis and visualisation in Python*
# > *December, 2018*
#
# > *© 2016, <NAME> and <NAME> (<mailto:<EMAIL>>, <mailto:<EMAIL>>). Licensed under [CC BY 4.0 Creative Commons](http://creativecommons.org/licenses/by/4.0/)*
#
# ---
# + run_control={"frozen": false, "read_only": false} slideshow={"slide_type": "-"}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# -
# We processed some raw data files of the AirBase air quality data. The data contains hourly concentrations of nitrogen dioxide (NO2) for 4 different measurement stations:
#
# - FR04037 (PARIS 13eme): urban background site at Square de Choisy
# - FR04012 (Paris, Place Victor Basch): urban traffic site at Rue d'Alesia
# - BETR802: urban traffic site in Antwerp, Belgium
# - BETN029: rural background site in Houtem, Belgium
#
# See http://www.eea.europa.eu/themes/air/interactive/no2
# + [markdown] slideshow={"slide_type": "slide"}
# # Importing and quick exploration
# + [markdown] slideshow={"slide_type": "subslide"}
# A set of pre-processed data is available in `../data/airbase_data.csv`):
# + run_control={"frozen": false, "read_only": false}
alldata = pd.read_csv('data/airbase_data.csv', index_col=0, parse_dates=True)
# -
# We only use the data from 1999 onwards:
data = alldata['1999':].copy()
# Some first exploration with the *typical* functions:
# + run_control={"frozen": false, "read_only": false}
data.head() # tail()
# + run_control={"frozen": false, "read_only": false}
data.info()
# + run_control={"frozen": false, "read_only": false}
data.describe(percentiles=[0.1, 0.5, 0.9])
# -
data.plot(figsize=(12,6))
# <div class="alert alert-warning">
# <b>ATTENTION!</b>: <br><br>
#
# When just using `.plot()` without further notice (selection, aggregation,...)
# <ul>
# <li>Risk of running into troubles by overloading your computer processing (certainly with looooong time series)</li>
# <li>Not always the most informative/interpretable visualisation</li>
# </ul>
# </div>
# **Plot only a subset**
# Why not just using the `head`/`tail` possibilities?
data.tail(500).plot(figsize=(12,6))
# **Summary figures**
# Use summary statistics...
data.plot(kind='box', ylim=[0,250])
# Also with seaborn plots function, just start with some subsets as first impression...
#
# As we already have seen previously, the plotting library [seaborn](http://seaborn.pydata.org/generated/seaborn.heatmap.html) provides some high-level plotting functions on top of matplotlib (check the [docs](http://seaborn.pydata.org/examples/index.html)!). One of those functions is `pairplot`, which we can use here to quickly visualize the concentrations at the different stations and their relation:
import seaborn as sns
sns.pairplot(data.tail(5000).dropna())
# # Is this a tidy dataset ?
data.head()
# In principle this is not a tidy dataset. The variable that was measured is the NO2 concentration, and is divided in 4 columns. Of course those measurements were made at different stations, so one could interpret it as separate variables. But in any case, such format does not always work well with libraries like `seaborn` which expects a pure tidy format.
#
#
# Reason to not use a tidy dataset here:
#
# * smaller memory use
# * timeseries functionality like resample works better
# * pandas plotting already does what we want when having different columns for *some* types of plots (eg line plots of the timeseries)
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>Create a tidy version of this dataset <code>data_tidy</code>, ensuring the result has new columns 'station' and 'no2'.</li>
# <li>Check how many missing values are contained in the 'no2' column.</li>
# <li>Drop the rows with missing values in that column.</li>
# </ul>
# </div>
# + clear_cell=true
data_tidy = data.reset_index().melt(id_vars=["datetime"], var_name='station', value_name='no2')
data_tidy.head()
# + clear_cell=true
data_tidy['no2'].isnull().sum()
# + clear_cell=true
data_tidy = data_tidy.dropna()
# -
# In the following exercises we will mostly do our analysis on `data`and often use pandas plotting, but once we produced some kind of summary dataframe as the result of an analysis, then it becomes more interesting to convert that result to a tidy format to be able to use the more advanced plotting functionality of seaborn.
# # Exercises
# <div class="alert alert-warning">
#
# <b>REMINDER</b>: <br><br>
#
# Take a look at the [Timeseries notebook](pandas_04_time_series_data.ipynb) when you require more info about:
#
# <ul>
# <li><code>resample</code></li>
# <li>string indexing of DateTimeIndex</li>
# </ul><br>
#
# Take a look at the [matplotlib](visualization_01_matplotlib.ipynb) and [seaborn](visualization_02_seaborn.ipynb) notebooks when you require more info about the plot requirements.
#
# </div>
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>Plot the monthly mean and median concentration of the 'FR04037' station for the years 2009 - 2013 in a single figure/ax</li>
# </ul>
# </div>
# + clear_cell=true
fig, ax = plt.subplots()
data.loc['2009':, 'FR04037'].resample('M').mean().plot(ax=ax, label='mean')
data.loc['2009':, 'FR04037'].resample('M').median().plot(ax=ax, label='median')
ax.legend(ncol=2)
ax.set_title("FR04037");
# + clear_cell=true
data.loc['2009':, 'FR04037'].resample('M').agg(['mean', 'median']).plot()
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>
#
# <ul>
# <li>Make a violin plot for January 2011 until August 2011 (check out the documentation to improve the plotting settings)</li>
# <li>Change the y-label to 'NO$_2$ concentration (µg/m³)'</li>
# </ul><br>
#
# NOTE:
#
# In this case, we can use seaborn both with the data not in a long format but when having different columns for which you want to make violin plots, as with the tidy data.
# </div>
# + clear_cell=true
# with wide dataframe
fig, ax = plt.subplots()
sns.violinplot(data=data['2011-01': '2011-08'], palette="GnBu_d", ax=ax)
ax.set_ylabel("NO$_2$ concentration (µg/m³)")
# + clear_cell=true
# with tidy dataframe
data_tidy_subset = data_tidy[(data_tidy['datetime'] >= "2011-01") & (data_tidy['datetime'] < "2011-09")]
fig, ax = plt.subplots()
sns.violinplot(data=data_tidy_subset, x="station", y="no2", palette="GnBu_d", ax=ax)
ax.set_ylabel("NO$_2$ concentration (µg/m³)")
# + clear_cell=true
# with figure-level function
sns.catplot(data=data_tidy_subset, x="station", y="no2", kind="violin", palette="GnBu_d")
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>
#
# <ul>
# <li>Make a bar plot with pandas of the mean of each of the stations in the year 2012 (check the documentation of Pandas plot to adapt the rotation of the labels) and make sure all bars have the same color.</li>
# <li>Using the matplotlib objects, change the y-label to 'NO$_2$ concentration (µg/m³)</li>
# <li>Add a 'darkorange' horizontal line on the ax for the y-value 40 µg/m³ (command for horizontal line from matplotlib: <code>axhline</code>).</li>
# <li><a href="visualization_01_matplotlib.ipynb">Place the text</a> 'Yearly limit is 40 µg/m³' just above the 'darkorange' line.</li>
# </ul>
#
# </div>
# + clear_cell=true
fig, ax = plt.subplots()
data['2012'].mean().plot(kind='bar', ax=ax, rot=0, color='C0')
ax.set_ylabel("NO$_2$ concentration (µg/m³)")
ax.axhline(y=40., color='darkorange')
ax.text(0.01, 0.48, 'Yearly limit is 40 µg/m³',
horizontalalignment='left', fontsize=13,
transform=ax.transAxes, color='darkorange');
# -
# <div class="alert alert-success">
#
# <b>EXERCISE:</b> Did the air quality improve over time?
#
# <ul>
# <li>For the data from 1999 till the end, plot the yearly averages</li>
# <li>For the same period, add the overall mean (all stations together) as an additional line to the graph, use a thicker black line (<code>linewidth=4</code> and <code>linestyle='--'</code>)</li>
# <li>[OPTIONAL] Add a legend above the ax for all lines</li>
#
#
# </ul>
# </div>
#
# + clear_cell=true
fig, ax = plt.subplots()
data['1999':].resample('A').mean().plot(ax=ax)
data['1999':].mean(axis=1).resample('A').mean().plot(color='k',
linestyle='--',
linewidth=4,
ax=ax,
label='Overall mean')
ax.legend(loc='center', ncol=3,
bbox_to_anchor=(0.5, 1.06))
ax.set_ylabel("NO$_2$ concentration (µg/m³)");
# -
# <div class="alert alert-info">
#
# <b>REMEMBER</b>: <br><br>
#
# `resample` is a special version of a`groupby` operation. For example, taking annual means with `data.resample('A').mean()` is equivalent to `data.groupby(data.index.year).mean()` (but the result of `resample` still has a DatetimeIndex).<br><br>
#
# Checking the index of the resulting DataFrame when using **groupby** instead of resample: You'll notice that the Index lost the DateTime capabilities:
#
# <code>
# > data.groupby(data.index.year).mean().index
# </code>
#
# Results in:
#
# <code>
# Int64Index([1990, 1991, 1992, 1993, 1994, 1995, 1996, 1997, 1998, 1999, 2000,
# 2001, 2002, 2003, 2004, 2005, 2006, 2007, 2008, 2009, 2010, 2011,
# 2012],
# dtype='int64')$
# </code>
#
# <br>
#
# When using **resample**, we keep the DateTime capabilities:
#
# <code>
# > data.resample('A').mean().index
# </code>
#
# Results in:
#
# <code>
# DatetimeIndex(['1999-12-31', '2000-12-31', '2001-12-31', '2002-12-31',
# '2003-12-31', '2004-12-31', '2005-12-31', '2006-12-31',
# '2007-12-31', '2008-12-31', '2009-12-31', '2010-12-31',
# '2011-12-31', '2012-12-31'],
# dtype='datetime64[ns]', freq='A-DEC')
# </code>
# <br>
#
# But, `groupby` is more flexible and can also do resamples that do not result in a new continuous time series, e.g. by grouping by the hour of the day to get the diurnal cycle.
# </div>
# <div class="alert alert-success">
#
# <b>EXERCISE</b>
#
# <ul>
# <li>How does the <i>typical yearly profile</i> (typical averages for the different months over the years) look like for the different stations? (add a 'month' column as a first step)</li>
#
# </ul>
# </div>
# + clear_cell=true
# add a column to the dataframe that indicates the month (integer value of 1 to 12):
data['month'] = data.index.month
# now, we can calculate the mean of each month over the different years:
data.groupby('month').mean()
# plot the typical monthly profile of the different stations:
data.groupby('month').mean().plot()
# -
data = data.drop("month", axis=1)
# Note: Technically, we could reshape the result of the groupby operation to a tidy format (we no longer have a real time series), but since we already have the things we want to plot as lines in different columns, doing `.plot` already does what we want.
# <div class="alert alert-success">
#
# <b>EXERCISE</b>
#
# <ul>
# <li>Plot the weekly 95% percentiles of the concentration in 'BETR801' and 'BETN029' for 2011</li>
#
# </ul>
# </div>
# + clear_cell=true
# Groupby wise
df2011 = data['2011']
df2011.groupby(df2011.index.week)[['BETN029', 'BETR801']].quantile(0.95).plot()
# + clear_cell=true
# Resample wise
# Note the different x-axis labels
df2011[['BETN029', 'BETR801']].resample('W').quantile(0.75).plot()
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>
#
# <ul>
# <li>Plot the typical diurnal profile (typical hourly averages) for the different stations taking into account the whole time period.</li>
#
# </ul>
# </div>
# + clear_cell=true
data.groupby(data.index.hour).mean().plot()
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b> <br><br>
#
# What is the difference in the typical diurnal profile between week and weekend days? (and visualise it)<br><br>
#
# Start with only visualizing the different in diurnal profile for the BETR801 station. In a next step, make the same plot for each station.<br><br>
#
# **Hints:**
#
# <ul>
# <li>Add a column 'weekend' defining if a value of the index is in the weekend (i.e. weekdays 5 and 6) or not</li>
# <li>Add a column 'hour' with the hour of the day for each row.</li>
# <li>You can groupby on multiple items at the same time.</li>
#
# </ul>
# </div>
# + clear_cell=true
data['weekend'] = data.index.weekday.isin([5, 6])
data['weekend'] = data['weekend'].replace({True: 'weekend', False: 'weekday'})
data['hour'] = data.index.hour
# + clear_cell=true
data_weekend = data.groupby(['weekend', 'hour']).mean()
data_weekend.head()
# + clear_cell=true
# using unstack and pandas plotting
data_weekend_BETR801 = data_weekend['BETR801'].unstack(level=0)
data_weekend_BETR801.plot()
# + clear_cell=true
# using a tidy dataset and seaborn
data_weekend_BETR801_tidy = data_weekend['BETR801'].reset_index()
sns.lineplot(data=data_weekend_BETR801_tidy, x="hour", y="BETR801", hue="weekend")
# + clear_cell=true
# tidy dataset that still includes all stations
data_weekend_tidy = pd.melt(data_weekend.reset_index(), id_vars=['weekend', 'hour'],
var_name='station', value_name='no2')
data_weekend_tidy.head()
# + clear_cell=true
# when still having multiple factors, it becomes useful to convert to tidy dataset and use seaborn
sns.relplot(data=data_weekend_tidy, x="hour", y="no2", kind="line",
hue="weekend", col="station", col_wrap=2)
# -
data = data.drop(['hour', 'weekend'], axis=1)
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:<br><br>
#
# <ul>
# <li>Calculate the correlation between the different stations (check in the documentation, google "pandas correlation" or use the magic function <code>%psearch</code>)</li>
#
# </ul>
# </div>
#
# + clear_cell=true
data[['BETR801', 'BETN029', 'FR04037', 'FR04012']].corr()
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:<br><br>
#
# Count the number of exceedances of hourly values above the European limit 200 µg/m3 for each year and station after 2005. Make a barplot of the counts. Add an horizontal line indicating the maximum number of exceedances (which is 18) allowed per year?<br><br>
#
# **Hints:**
#
# <ul>
# <li>Create a new DataFrame, called <code>exceedances</code>, (with boolean values) indicating if the threshold is exceeded or not</li>
# <li>Remember that the sum of True values can be used to count elements</li>
# <li>Adding a horizontal line can be done with the matplotlib function <code>ax.axhline</code></li>
#
#
# </ul>
# </div>
# + clear_cell=true
exceedances = data > 200
# + clear_cell=true
# group by year and count exceedances (sum of boolean)
exceedances = exceedances.groupby(exceedances.index.year).sum()
# + clear_cell=true
# Make a barplot of the yearly number of exceedances
ax = exceedances.loc[2005:].plot(kind='bar')
ax.axhline(18, color='k', linestyle='--')
# + [markdown] slideshow={"slide_type": "slide"}
# # More advanced exercises...
# -
data = alldata['1999':].copy()
# <div class="alert alert-success">
#
# <b>EXERCISE</b>: Perform the following actions for the station `'FR04012'` only:
#
# <ul>
# <li>Remove the rows containing <code>NaN</code> or zero values</li>
# <li>Sort the values of the rows according to the air quality values (low to high values)</li>
# <li>Rescale the values to the range [0-1] and store result as <code>FR_scaled</code> (Hint: check <a href="https://en.wikipedia.org/wiki/Feature_scaling#Rescaling">wikipedia</a>)</li>
# <li>Use pandas to plot these values sorted, not taking into account the dates</li>
# <li>Add the station name 'FR04012' as y-label</li>
# <li>[OPTIONAL] Add a vertical line to the plot where the line (hence, the values of variable FR_scaled) reach the value <code>0.3</code>. You will need the documentation of <code>np.searchsorted</code> and matplotlib's <code>axvline</code></li>
# </ul>
# </div>
# + clear_cell=true
FR_station = data['FR04012'] # select the specific data series
FR_station = FR_station[(FR_station.notnull()) & (FR_station != 0.0)] # exclude the Nan and zero values
# + clear_cell=true
FR_sorted = FR_station.sort_values(ascending=True)
FR_scaled = (FR_sorted - FR_sorted.min())/(FR_sorted.max() - FR_sorted.min())
# + clear_cell=true run_control={"frozen": false, "read_only": false}
fig, axfr = plt.subplots()
FR_scaled.plot(use_index=False, ax = axfr) #alternative version: FR_scaled.reset_index(drop=True).plot(use_index=False)
axfr.set_ylabel('FR04012')
# optional addition, just in case you need this
axfr.axvline(x=FR_scaled.searchsorted(0.3), color='0.6', linestyle='--', linewidth=3)
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>Create a Figure with two subplots (axes), for which both ax<b>i</b>s are shared</li>
# <li>In the left subplot, plot the histogram (30 bins) of station 'BETN029', only for the year 2009</li>
# <li>In the right subplot, plot the histogram (30 bins) of station 'BETR801', only for the year 2009</li>
# <li>Add the title representing the station name on each of the subplots, you do not want to have a legend</li>
# </ul>
# </div>
# + clear_cell=true
# Mixing an matching matplotlib and Pandas
fig, (ax1, ax2) = plt.subplots(1, 2,
sharex=True,
sharey=True)
data.loc['2009', ['BETN029', 'BETR801']].plot(kind='hist', subplots=True,
bins=30, legend=False,
ax=(ax1, ax2))
ax1.set_title('BETN029')
ax2.set_title('BETR801')
# Remark: the width of the bins is calculated over the x data range for both plots together
# + clear_cell=true
# A more step by step approach (equally valid)
fig, (ax1, ax2) = plt.subplots(1, 2, sharey=True, sharex=True)
data.loc['2009', 'BETN029'].plot(kind='hist', bins=30, ax=ax1)
ax1.set_title('BETN029')
data.loc['2009', 'BETR801'].plot(kind='hist', bins=30, ax=ax2)
ax2.set_title('BETR801')
# Remark: the width of the bins is calculated over the x data range for each plot individually
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>
#
# <ul>
# <li>Make a selection of the original dataset of the data in January 2009, call the resulting variable <code>subset</code></li>
# <li>Add a new column, called 'weekday', to the variable <code>subset</code> which defines for each data point the day of the week</li>
# <li>From the <code>subset</code> DataFrame, select only Monday (= day 0) and Sunday (=day 6) and remove the others (so, keep this as variable <code>subset</code>)</li>
# <li>Change the values of the weekday column in <code>subset</code> according to the following mapping: <code>{0:"Monday", 6:"Sunday"}</code></li>
# <li>With seaborn, make a scatter plot of the measurements at 'BETN029' vs 'FR04037', with the color variation based on the weekday. Add a linear regression to this plot.</li>
# </ul><br>
#
# **Note**: If you run into the **SettingWithCopyWarning** and do not know what to do, recheck [pandas_03b_indexing](pandas_03b_indexing.ipynb)
#
# </div>
# + clear_cell=true
subset = data['2009-01'].copy()
subset["weekday"] = subset.index.weekday
subset = subset[subset['weekday'].isin([0, 6])]
# + clear_cell=true
subset["weekday"] = subset["weekday"].replace(to_replace={0:"Monday", 6:"Sunday"})
# + clear_cell=true
sns.set_style("whitegrid")
# + clear_cell=true
sns.lmplot(
data=subset, x="BETN029", y="FR04037", hue="weekday"
)
# -
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>The maximum daily, 8 hour mean, should be below 100 µg/m³. What is the number of exceedances of this limit for each year/station?</li><br>
# </ul>
#
# **Tip:**<br>
#
# Have a look at the `rolling` method to perform moving window operations.<br><br>
#
# **Note:**<br>
# This is not an actual limit for NO$_2$, but a nice exercise to introduce the `rolling` method. Other pollutans, such as 0$_3$ have actually such kind of limit values based on 8-hour means.
#
# </div>
# + clear_cell=true run_control={"frozen": false, "read_only": false}
exceedances = data.rolling(8).mean().resample('D').max() > 100
# + clear_cell=true run_control={"frozen": false, "read_only": false}
exceedances = exceedances.groupby(exceedances.index.year).sum()
ax = exceedances.plot(kind='bar')
# + [markdown] slideshow={"slide_type": "subslide"}
# <div class="alert alert-success">
#
# <b>EXERCISE</b>:
#
# <ul>
# <li>Visualize the typical week profile for station 'BETR801' as boxplots (where the values in one boxplot are the <i>daily means</i> for the different <i>weeks</i> for a certain weekday).</li><br>
# </ul>
#
#
# **Tip:**<br>
#
# The boxplot method of a DataFrame expects the data for the different boxes in different columns. For this, you can either use `pivot_table` or a combination of `groupby` and `unstack`
#
#
# </div>
#
# -
# Calculating daily means and add weekday information:
# + clear_cell=true
data_daily = data.resample('D').mean()
# + clear_cell=true run_control={"frozen": false, "read_only": false} slideshow={"slide_type": "fragment"}
# add a weekday column
data_daily['weekday'] = data_daily.index.weekday
data_daily.head()
# -
# Plotting with seaborn:
# + clear_cell=true
# seaborn
sns.boxplot(data=data_daily["2012":], x='weekday', y='BETR801', color="grey")
# -
# Reshaping and plotting with pandas:
# + clear_cell=true run_control={"frozen": false, "read_only": false} slideshow={"slide_type": "subslide"}
# when using pandas to plot, the different boxplots should be different columns
# therefore, pivot table so that the weekdays are the different columns
data_daily['week'] = data_daily.index.week
data_pivoted = data_daily['2012'].pivot_table(columns='weekday', index='week', values='BETR801')
data_pivoted.head()
data_pivoted.boxplot();
# + clear_cell=true run_control={"frozen": false, "read_only": false}
# An alternative method using `groupby` and `unstack`
data_daily['2012'].groupby(['weekday', 'week'])['BETR801'].mean().unstack(level=0).boxplot();
# -
|
_solved/case4_air_quality.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Machine Learning Notebooks
#
# *Welcome to the Machine Learning Notebooks!*
#
# [Prerequisites](#Prerequisites) (see below)
#
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/ageron/handson-ml3/blob/main/index.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# </td>
# <td>
# <a target="_blank" href="https://homl.info/kaggle/"><img src="https://kaggle.com/static/images/open-in-kaggle.svg" /></a>
# </td>
# </table>
# ## Notebooks
# 1. [The Machine Learning landscape](01_the_machine_learning_landscape.ipynb)
# 2. [End-to-end Machine Learning project](02_end_to_end_machine_learning_project.ipynb)
# 3. [Classification](03_classification.ipynb)
# 4. [Training Models](04_training_linear_models.ipynb)
# 5. [Support Vector Machines](05_support_vector_machines.ipynb)
# 6. [Decision Trees](06_decision_trees.ipynb)
# 7. [Ensemble Learning and Random Forests](07_ensemble_learning_and_random_forests.ipynb)
# 8. [Dimensionality Reduction](08_dimensionality_reduction.ipynb)
# 9. [Unsupervised Learning Techniques](09_unsupervised_learning.ipynb)
# 10. [Artificial Neural Nets with Keras](10_neural_nets_with_keras.ipynb)
# 11. [Training Deep Neural Networks](11_training_deep_neural_networks.ipynb)
# 12. [Custom Models and Training with TensorFlow](12_custom_models_and_training_with_tensorflow.ipynb)
# 13. [Loading and Preprocessing Data](13_loading_and_preprocessing_data.ipynb)
# 14. [Deep Computer Vision Using Convolutional Neural Networks](14_deep_computer_vision_with_cnns.ipynb)
# 15. [Processing Sequences Using RNNs and CNNs](15_processing_sequences_using_rnns_and_cnns.ipynb)
# 16. [Natural Language Processing with RNNs and Attention](16_nlp_with_rnns_and_attention.ipynb)
# 17. [Representation Learning and Generative Learning Using Autencoders and GANs](17_autoencoders_and_gans.ipynb)
# 18. [Reinforcement Learning](18_reinforcement_learning.ipynb)
# 19. [Training and Deploying TensorFlow Models at Scale](19_training_and_deploying_at_scale.ipynb)
# ## Scientific Python tutorials
# * [NumPy](tools_numpy.ipynb)
# * [Matplotlib](tools_matplotlib.ipynb)
# * [Pandas](tools_pandas.ipynb)
# ## Math Tutorials
# * [Linear Algebra](math_linear_algebra.ipynb)
# * [Differential Calculus](math_differential_calculus.ipynb)
# ## Extra Material
# * [Auto-differentiation](extra_autodiff.ipynb)
# ## Misc.
# * [Equations](book_equations.pdf) (list of equations in the book)
# ## Prerequisites
# ### To understand
# * **Python** – you don't need to be an expert python programmer, but you do need to know the basics. If you don't, the official [Python tutorial](https://docs.python.org/3/tutorial/) is a good place to start.
# * **Scientific Python** – We will be using a few popular python libraries, in particular NumPy, matplotlib and pandas. If you are not familiar with these libraries, you should probably start by going through the tutorials in the Tools section (especially NumPy).
# * **Math** – We will also use some notions of Linear Algebra, Calculus, Statistics and Probability theory. You should be able to follow along if you learned these in the past as it won't be very advanced, but if you don't know about these topics or you need a refresher then go through the appropriate introduction in the Math section.
# ### To run the examples
# * **Jupyter** – These notebooks are based on Jupyter. You can run these notebooks in just one click using a hosted platform such as Binder, Deepnote or Colaboratory (no installation required), or you can just view them using Jupyter.org's viewer, or you can install everything on your machine, as you prefer. Check out the [home page](https://github.com/ageron/handson-ml3/) for more details.
|
index.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Example 1: Sandstone Model
# + code_folding=[]
# Importing
import theano.tensor as T
import sys, os
sys.path.append("../GeMpy")
# Importing GeMpy modules
import GeMpy_core
import Visualization
# Reloading (only for development purposes)
import importlib
importlib.reload(GeMpy_core)
importlib.reload(Visualization)
# Usuful packages
import numpy as np
import pandas as pn
import matplotlib.pyplot as plt
# This was to choose the gpu
os.environ['CUDA_LAUNCH_BLOCKING'] = '1'
# Default options of printin
np.set_printoptions(precision = 6, linewidth= 130, suppress = True)
# %matplotlib inline
# #%matplotlib notebook
# -
# First we make a GeMpy instance with most of the parameters default (except range that is given by the project). Then we also fix the extension and the resolution of the domain we want to interpolate. Finally we compile the function, only needed once every time we open the project (the guys of theano they are working on letting loading compiled files, even though in our case it is not a big deal).
#
# *General note. So far the reescaling factor is calculated for all series at the same time. GeoModeller does it individually for every potential field. I have to look better what this parameter exactly means*
# +
# Setting extend, grid and compile
# Setting the extent
sandstone = GeMpy_core.GeMpy()
# Create Data class with raw data
sandstone.import_data( [696000,747000,6863000,6950000,-20000, 2000],[ 40, 40, 80],
path_f = os.pardir+"/input_data/a_Foliations.csv",
path_i = os.pardir+"/input_data/a_Points.csv")
# -
# All input data is stored in pandas dataframes under, ```self.Data.Interances``` and ```self.Data.Foliations```:
sandstone.Data.Foliations.head()
# In case of disconformities, we can define which formation belong to which series using a dictionary. Until python 3.6 is important to specify the order of the series otherwise is random
sandstone.Data.set_series({"EarlyGranite_Series":sandstone.Data.formations[-1],
"BIF_Series":(sandstone.Data.formations[0], sandstone.Data.formations[1]),
"SimpleMafic_Series":sandstone.Data.formations[2]},
order = ["EarlyGranite_Series",
"BIF_Series",
"SimpleMafic_Series"])
# Now in the data frame we should have the series column too
sandstone.Data.Foliations.head()
# Next step is the creating of a grid. So far only regular. By default it takes the extent and the resolution given in the `import_data` method.
# Create a class Grid so far just regular grid
sandstone.create_grid()
sandstone.Grid.grid
# ## Plotting raw data
# The object Plot is created automatically as we call the methods above. This object contains some methods to plot the data and the results.
#
# It is possible to plot a 2D projection of the data in a specific direction using the following method. Also is possible to choose the series you want to plot. Additionally all the key arguments of seaborn lmplot can be used.
sandstone.Plot.plot_data(series = sandstone.Data.series.columns.values[1])
# ## Class Interpolator
# This class will take the data from the class Data and calculate potential fields and block. We can pass as key arguments all the variables of the interpolation. I recommend not to touch them if you do not know what are you doing. The default values should be good enough. Also the first time we execute the method, we will compile the theano function so it can take a bit of time.
# + code_folding=[]
sandstone.set_interpolator()
# -
# Now we could visualize the individual potential fields as follow:
# ### Early granite
sandstone.Plot.plot_potential_field(10, n_pf=0)
# ### BIF Series
sandstone.Plot.plot_potential_field(13, n_pf=1, cmap = "magma", plot_data = True,
verbose = 5 )
# ### SImple mafic
sandstone.Plot.plot_potential_field(10, n_pf=2)
# ## Optimizing the export of lithologies
#
# But usually the final result we want to get is the final block. The method `compute_block_model` will compute the block model, updating the attribute `block`. This attribute is a theano shared function that can return a 3D array (raveled) using the method `get_value()`.
# +
# Reset the block
sandstone.Interpolator.block.set_value(np.zeros_like(sandstone.Grid.grid[:,0]))
# Compute the block
sandstone.Interpolator.compute_block_model([0,1,2], verbose = 0)
# -
sandstone.Interpolator.block.get_value(), np.unique(sandstone.Interpolator.block.get_value())
# And again after computing the model in the Plot object we can use the method `plot_block_section` to see a 2D section of the model
sandstone.Plot.plot_block_section(13, interpolation = 'nearest', direction='y')
plt.savefig("sandstone_example.png")
# ## Export to vtk. (*Under development*)
# +
"""Export model to VTK
Export the geology blocks to VTK for visualisation of the entire 3-D model in an
external VTK viewer, e.g. Paraview.
..Note:: Requires pyevtk, available for free on: https://github.com/firedrakeproject/firedrake/tree/master/python/evtk
**Optional keywords**:
- *vtk_filename* = string : filename of VTK file (default: output_name)
- *data* = np.array : data array to export to VKT (default: entire block model)
"""
vtk_filename = "noddyFunct2"
extent_x = 10
extent_y = 10
extent_z = 10
delx = 0.2
dely = 0.2
delz = 0.2
from pyevtk.hl import gridToVTK
# Coordinates
x = np.arange(0, extent_x + 0.1*delx, delx, dtype='float64')
y = np.arange(0, extent_y + 0.1*dely, dely, dtype='float64')
z = np.arange(0, extent_z + 0.1*delz, delz, dtype='float64')
# self.block = np.swapaxes(self.block, 0, 2)
gridToVTK(vtk_filename, x, y, z, cellData = {"geology" : sol})
# -
# ## Performance Analysis
# One of the advantages of theano is the posibility to create a full profile of the function. This has to be included in at the time of the creation of the function. At the moment it should be active (the downside is larger compilation time and I think also a bit in the computation so be careful if you need a fast call)
# ### CPU
# The following profile is with a 2 core laptop. Nothing spectacular.
# +
# %%timeit
# Reset the block
sandstone.Interpolator.block.set_value(np.zeros_like(sandstone.Grid.grid[:,0]))
# Compute the block
sandstone.Interpolator.compute_block_model([0,1,2], verbose = 0)
# -
# Looking at the profile we can see that most of time is in pow operation (exponential). This probably is that the extent is huge and we are doing it with too much precision. I am working on it
esandstone.Interpolator._interpolate.profile.summary()
# ### GPU
# +
# %%timeit
# Reset the block
sandstone.block.set_value(np.zeros_like(sandstone.grid[:,0]))
# Compute the block
sandstone.compute_block_model([0,1,2], verbose = 0)
# -
sandstone.block_export.profile.summary()
|
Prototype Notebook/.ipynb_checkpoints/Sandstone Testing new structure-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Synapse PySpark
# name: synapse_pyspark
# ---
# # Getting started with Azure Cosmos DB's API for MongoDB and Synapse Link
#
# In this sample we will execute the following tasks:
#
# 1. Insert a dataset using the traditional MongoDB client.
# 1. Execute aggregation queries against the Analytical Store from the transactional data we inserted.
# 1. Insert another dataset, but this time using the MongoSpark connector.
# 1. Execute aggregation queries again, consolidating both datasets.
#
# ## Pre-requisites
# 1. Have you created a MongoDB API account in Azure Cosmos DB? If not, go to [Create an account for Azure Cosmos DB's API for MongoDB](https://docs.microsoft.com/azure/cosmos-db/create-cosmosdb-resources-portal#create-an-azure-cosmos-db-account). Be sure to create an account using MongoDB as the API option.
# 1. For your Cosmos DB account, have you enabled Synapse Link? If not, go to [Enable Synapse Link for Azure Cosmos DB accounts](https://docs.microsoft.com/azure/cosmos-db/configure-synapse-link#enable-synapse-link).
# 1. Have you created a Synapse Workspace? If not, go to [Create a Synapse Workspace](https://docs.microsoft.com/azure/synapse-analytics/quickstart-create-workspace).
#
# ## Create a Cosmos DB collection with Synapse Link
# 1. Create a database named `test`.
# 1. Create a collection named `htap` with a partition key called `pk`.
# - Make sure you set the `Analytical store` option to `On` when you create your collection.
#
# ## Connect your collection to Synapse
# 1. Go to your Synapse Analytics workspace.
# 1. Create a `Linked Data` connection for your MongoDB API account.
# 1. Under the `Data` blade, select the + (plus) sign.
# 1. Select the `Connect to external data` option.
# 1. Now select the `Azure Cosmos DB (MongoDB API)` option.
# 1. Enter all the information regarding your specific Azure Cosmos DB account either by using the dropdowns or by entering the connection string. Take note of the name you assigned to your `Linked Data` connection.
# - Alternatively, you can also use the connection parameters from your account overview.
# 1. Test the connection by looking for your database accounts in the `Data` blade, and under the `Linked` tab.
# - There should be a list that contains all accounts and collections.
# - Collections that have an `Analytical Store` enabled will have a distinctive icon.
# ### Let's get the environment ready
#
# This environment allows you to install and use any python libraries that you want to run. For this sample, you need to add the following libraries to your Spark pool:
#
# ```
# pymongo==2.8.1
# bson
# randint
# aenum==2.1.2
# backports-abc==0.5
# bson==0.5.10
# ```
#
# Learn how to import libraries into your Spark pools in [this article](https://docs.microsoft.com/en-us/azure/synapse-analytics/spark/apache-spark-azure-portal-add-libraries). We recommend creating a new pool for this.
#
# You can execute the following command to make sure all the libraries are installed correctly:
# + outputCollapsed=true
import pip #needed to use the pip functions
for i in pip.get_installed_distributions(local_only=True):
print(i)
# The output might be long... you can collapse it by clicking on the 'Collapse output' option on the upper left corner of the output cell.
# -
# ### Write your database account specific secrets here!
#
# We won't tell anybody.
#
DATABASE_ACCOUNT_NAME = 'lbosq-mongo-synapse'
DATABASE_ACCOUNT_READWRITE_KEY = '<KEY>
# ## Let's initialize the MongoDB client
#
# You are only going to need the following parameters from your account overview:
# - Connection string.
# - Primary or secondary ready/write key.
#
# Remember that we named our database `test` and our collection `htap`.
#
# The code snippet below shows how to initialize the `MongoClient` object.
# +
from pymongo import MongoClient
from bson import ObjectId # For ObjectId to work
client = MongoClient("mongodb://{account}.mongo.cosmos.azure.com:10255/?ssl=true&replicaSet=globaldb".format(account = DATABASE_ACCOUNT_NAME)) # Your own database account endpoint.
db = client.test # Select the database
db.authenticate(name=DATABASE_ACCOUNT_NAME,password=DATABASE_ACCOUNT_READWRITE_KEY) # Use your database account name and any of your read/write keys.
# -
# ## Inserting data with the MongoClient driver
#
# The following sample will generate 500 items based on random data. Each item will contain the following fields:
# - Item, string
# - Price, float
# - Rating, integer
# - Timestamp, [epoch integer](http://unixtimestamp.50x.eu/about.php)
#
# This cell depends on the cell above to create an instance of the connection to the Cosmos DB MongoDB API account.
#
# This data will be inserted into the MongoDB store of your database. This emulates the transactional data that an application would generate.
# +
from random import randint
import time
orders = db["htap"]
items = ['Pizza','Sandwich','Soup', 'Salad', 'Tacos']
prices = [2.99, 3.49, 5.49, 12.99, 54.49]
for x in range(1, 501):
order = {
'item' : items[randint(0, (len(items)-1))],
'price' : prices[randint(0, (len(prices)-1))],
'rating' : randint(1, 5),
'timestamp' : time.time()
}
result=orders.insert(order)
print('finished creating 500 orders')
# -
# ## Read data from the Analytical Store.
#
# Now that we have inserted some transactional data, let's read it from the Analytical Store.
#
# The data will be automatically transformed into the columnar format, which will make it fast and easy to execute large aggregation queries.
#
# +
# Load the Analytical Store data into a dataframe
# Make sure to run the cell with the secrets to get the DATABASE_ACCOUNT_NAME and the DATABASE_ACCOUNT_READWRITE_KEY variables.
df = spark.read.format("cosmos.olap")\
.option("spark.cosmos.accountEndpoint", "https://{account}.documents.azure.com:443/".format(account = DATABASE_ACCOUNT_NAME))\
.option("spark.cosmos.accountKey", DATABASE_ACCOUNT_READWRITE_KEY)\
.option("spark.cosmos.database", "test")\
.option("spark.cosmos.container", "htap")\
.load()
# Let's find out all the revenue from Pizza orders
df.groupBy(df.item.string).sum().show()
# df[df.item.string == 'Pizza'].show(10)
# df.select(df['item'] == Struct).show(10)
# df.select("timestamp.float64").show(10)
#df.select("timestamp.string", when(df.timestamp.string != null)).show(10)
# + [markdown] nteract={"transient": {"deleting": false}}
#
# ## A quick note about the MongoDB schema in Analytical Store
#
# For MongoDB accounts we make use of a **Full Fidelity Schema**. This is a representation of property names extended with their data types to provide an accurate representation of their values and avoid ambiguity.
#
# This is why, when we called the fields above, we used their datatype as a suffix. Like in the example below:
#
# ```
# df.filter((df.item.string == "Pizza")).show(10)
# ```
#
# Notice how we specified the `string` type after the name of the property. Here is a map of all potential properties and their suffix representations in the Analytical Store:
#
# | Original Data Type | Suffix | Example |
# |---------------|----------------|--------|
# | Double | ".float64" | `24.99` |
# | Array | ".array" | `["a", "b"]` |
# | Binary | ".binary" | `0` |
# | Boolean | ".bool" | `True` |
# | Int32 | ".int32" | `123` |
# | Int64 | ".int64" | `255486129307` |
# | Null | ".null" | `null` |
# | String | ".string" | `"ABC"` |
# | Timestamp | ".timestamp" | `Timestamp(0, 0)` |
# | DateTime | ".date" | `ISODate("2020-08-21T07:43:07.375Z")` |
# | ObjectId | ".objectId" | `ObjectId("5f3f7b59330ec25c132623a2")` |
# | Document | ".object" | `{"a": "a"}` |
#
# These types are inferred from the data that is inserted in the transactional store. You can see the schema by executing the following command:
# ```
# df.schema
# ```
# -
# ## Let's insert more orders!
#
# This time we will use slightly different data. Each item will contain the following fields:
# - Item, string
# - Price, float
# - Rating, integer
# - Timestamp, [ISO String format](https://en.wikipedia.org/wiki/ISO_8601)
#
# Notice how the `Timestamp` field is now in a string format. This will help us understand how the different data fields can be read based on their data type.
# +
from random import randint
from time import strftime
orders = db["htap"]
items = ['Pizza','Sandwich','Soup', 'Salad', 'Tacos']
prices = [2.99, 3.49, 5.49, 12.99, 54.49]
for x in range(1, 501):
order = {
'item' : items[randint(0, (len(items)-1))],
'price' : prices[randint(0, (len(prices)-1))],
'rating' : randint(1, 5),
'timestamp' : strftime("%Y-%m-%d %H:%M:%S")
}
result=orders.insert(order)
print('finished creating 500 orders')
# -
# ## Let's read that data again!
#
# This time, we will be reading the ISO string dates separately by specifying the `timestamp.string` parameter.
#
# +
# Load the Analytical Store data into a dataframe
# Make sure to run the cell with the secrets to get the DATABASE_ACCOUNT_NAME and the DATABASE_ACCOUNT_READWRITE_KEY variables.
df = spark.read.format("cosmos.olap")\
.option("spark.cosmos.accountEndpoint", "https://{account}.documents.azure.com:443/".format(account = DATABASE_ACCOUNT_NAME))\
.option("spark.cosmos.accountKey", DATABASE_ACCOUNT_READWRITE_KEY)\
.option("spark.cosmos.database", "test")\
.option("spark.cosmos.container", "htap")\
.load()
# Let's find out all the revenue from Pizza orders
df.filter( (df.timestamp.string != "")).show(10)
|
Notebooks/PySpark/Synapse Link for Cosmos DB samples/MongoDB/spark-notebooks/pyspark/01-CosmosDBSynapseMongoDB.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ESPEI
#
# ### Extensible Self-optimizating Phase Equilibria Infrastructure
#
# Documentation for internal and external APIs can be found at https://espei.org
#
# Solutions to this notebook can be found at https://github.com/materialsgenomefoundation/2021-workshop-material
#
# ## Markov Chain Monte Carlo (MCMC)
#
# ### Running MCMC
#
# The most minimal MCMC settings file that could be used for ESPEI only requires setting the source of the database (i.e. the database from parameter selection) and the number of iterations.
#
# ```yaml
# system:
# phase_models: phases.json
# datasets: input-data/run
# output:
# output_db: mcmc.tdb
# verbosity: 2
# mcmc:
# iterations: 100
# input_db: dft-aicc_penalty.tdb
# ```
#
# By default, ESPEI will run in parallel using the `dask` package. If you try to run this locally, you may need to do an [extra step to configure dask](https://espei.org/en/latest/installation.html#configuration).
#
# However, since we are using limited and shared cloud resources, we will make some compromises in terms of accuracy and storage. The settings we'll use for running MCMC simulations are as follows (saved as `mcmc_settings.yaml`):
#
# ```yaml
# system:
# phase_models: Cr-Ni_phases.json
# datasets: input-data
# tags:
# dft:
# excluded_model_contributions: ['idmix', 'mag']
# weight: 0.1
# nomag:
# excluded_model_contributions: ['mag']
# estimated-entropy:
# excluded_model_contributions: ['idmix', 'mag']
# weight: 0.1
# output:
# output_db: mcmc.tdb
# verbosity: 2
# tracefile: null # don't write a trace file
# probfile: null # don't write a lnprob file
# logfile: null # write output to console
# mcmc:
# iterations: 3
# save_interval: 1
# scheduler: null # no parallelization
# input_db: mcmc-start.tdb # same as our generated-aicc.tdb
# chains_per_parameter: 2
# data_weights:
# ZPF: 40.0
# SM: 0.1
# ```
#
# All MCMC options are explained in [ESPEI's YAML settings file documentation](https://espei.org/en/latest/writing_input.html#mcmc). Compared to parameter generation, the options are more extensive and worth being familiar with.
import yaml
from espei import run_espei
from pycalphad import Database, binplot, equilibrium, variables as v
with open('mcmc_settings.yaml') as fp:
mcmc_settings = yaml.safe_load(fp)
# Now we'll run the MCMC simulation for just two iterations. The outputs are the database which has the most optimal parameters of all samples and an `emcee.EnsembleSampler` object that contains the trace (contains samples of the parameters for every chain and iteration) and the log-likelihood.
dbf_mcmc, sampler = run_espei(mcmc_settings)
# The key output from the sampler are the trace (`emcee` calls this the "chain") and the log-probability (lnprob). The trace has the shape `(number of chains, number of iterations, number of parameters)`. The log-probability has the shape `(number of chains, number of iterations)`.
#
#
# +
trace = sampler.chain
lnprob = sampler.lnprobability
print(f"Trace shape: {trace.shape}")
print(f"Log-probability shape: {lnprob.shape}")
# -
# With the MCMC simulation complete, we can see what the phase diagram looks like.
# +
from espei.plot import dataplot
from espei.datasets import recursive_glob, load_datasets
# load our JSON datasets into an in-memory database
datasets = load_datasets(recursive_glob('input-data', '*.json'))
# -
dbf_start = Database('mcmc-start.tdb')
comps = ['CR', 'NI']
phases = ['FCC_A1', 'BCC_A2', 'LIQUID']
conds = {v.N: 1.0, v.P: 101325, v.T: (300, 2300, 20), v.X('NI'): (0, 1, 0.02)}
ax = binplot(dbf_start, comps, phases, conds)
dataplot(comps, phases, conds, datasets, ax=ax)
comps = ['CR', 'NI']
phases = ['FCC_A1', 'BCC_A2', 'LIQUID']
conds = {v.N: 1.0, v.P: 101325, v.T: (300, 2300, 20), v.X('NI'): (0, 1, 0.02)}
ax = binplot(dbf_mcmc, comps, phases, conds)
dataplot(comps, phases, conds, datasets, ax=ax)
# After just three MCMC iterations through 20 chains, the phase diagram shows a small improvement.
# ### Simulated MCMC analysis
#
# Typically, a fully converged MCMC simulation with enough samples to do uncertainty quantification will require a few hundred to a few thousand calculations.
#
#
# Since performing an MCMC simulation for a significant period of time is not possible in this workshop, an existing pre-computed trace and log-probability are loaded that took 1000 iterations of sampling 40 chains for a total of 40,000 samples in parameter space. These 40,000 samples took 3.5 hours to run across 6 cores on a 2015 MacBook Pro (2.2 GHz Intel i7).
#
import numpy as np
from espei.analysis import truncate_arrays
# +
trace = np.load('Cr-Ni-trace.npy')
lnprob = np.load('Cr-Ni-lnprob.npy')
trace, lnprob = truncate_arrays(trace, lnprob)
print(trace.shape)
print(lnprob.shape)
# -
# #### Visualizing convergence of the simulation
# +
import matplotlib.pyplot as plt
plt.plot(lnprob.T)
plt.title('Log-probability convergence\n(1 line = 1 chain)')
plt.xlabel('Iterations')
plt.ylabel('Log-probability')
plt.figure()
plt.plot(lnprob.T)
plt.title('Zoomed Log-probability convergence\n(1 line = 1 chain)')
plt.xlabel('Iterations')
plt.ylabel('Log-probability')
plt.ylim(-4000, -2000)
# -
# #### Visualizing change in a particular parameter
# +
# index of parameter of interest within the chain
# could be looped to produce figures for all parameters
parameter_idx = 5
num_chains = trace.shape[0]
ax = plt.figure().gca()
ax.set_xlabel('Iterations')
ax.set_ylabel('Parameter value')
ax.plot(trace[..., parameter_idx].T)
ax.set_title('Parameter Convergence')
# -
# #### Finding the optimal set of parameters and plotting the phase diagram
#
# An MCMC simulation has many samples, but we are still likely interested in getting the set of parameters that's the best point estimate of the data.
#
# ESPEI provides an `optimal_parameters` function that will extract the parameter set with the highest log-probability, which can be used to update the symbols that we fit in the database ($ \mathrm{VV0001} $, ...)..
# +
from espei.utils import database_symbols_to_fit, optimal_parameters
import copy
# make an in-memory copy of the database because updating the
# symbols with the optimal solutions will erase the old ones
dbf_opt = copy.deepcopy(dbf_mcmc)
# Find the optimal parameters and replace the values in the symbols dictionary
opt_params = dict(zip(database_symbols_to_fit(dbf_opt), optimal_parameters(trace, lnprob)))
dbf_opt.symbols.update(opt_params)
# +
# plot the phase diagram
ax = binplot(dbf_opt, comps, phases, conds)
dataplot(comps, phases, conds, datasets, ax=ax)
|
ESPEI/ESPEI MCMC.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import warnings
warnings.filterwarnings('ignore')
import os
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from tqdm import tqdm, tqdm_notebook, tqdm_pandas
tqdm.pandas(tqdm_notebook)
from sklearn.model_selection import StratifiedKFold, train_test_split, GroupKFold
import lightgbm as lgb
from sklearn.metrics import roc_auc_score
from sklearn import preprocessing
# +
train_transaction = pd.read_csv('../input/train_transaction.csv')
train_identity = pd.read_csv('../input/train_identity.csv')
test_transaction = pd.read_csv('../input/test_transaction.csv')
test_identity = pd.read_csv('../input/test_identity.csv')
# -
train_transaction = train_transaction.merge(train_identity, on=['TransactionID'],how='left')
test_transaction = test_transaction.merge(test_identity, on=['TransactionID'],how='left')
all_data = pd.concat([train_transaction,test_transaction])
id_feature = [ c for c in all_data.columns if c.find('id_') !=-1]
v_feature = [ c for c in all_data.columns if c.find('V') !=-1]
card_feature = [ c for c in all_data.columns if c.find('card') !=-1]
C_feature = [ c for c in all_data.columns if c.find('C') !=-1 and c != 'ProductCD']
D_feature = [ c for c in all_data.columns if c.find('D') !=-1 and c not in ['ProductCD','TransactionID','TransactionDT','DeviceType','DeviceInfo']]
M_feature = [ c for c in all_data.columns if c.find('M') !=-1]
# +
# 청구자와 수납자의 mail domain이 같은지?
all_data['email_check'] = np.where(all_data['P_emaildomain']==all_data['R_emaildomain'],1,0)
# 모두 null인지
all_data['email_check_nan_all'] = np.where((all_data['P_emaildomain'].isna())&(all_data['R_emaildomain'].isna()),1,0)
# 하나만 null인지
all_data['email_check_nan_any'] = np.where((all_data['P_emaildomain'].isna())|(all_data['R_emaildomain'].isna()),1,0)
# 제공안된 email fillna
all_data['P_emaildomain'] = all_data['P_emaildomain'].fillna('email_not_provided')
all_data['R_emaildomain'] = all_data['R_emaildomain'].fillna('email_not_provided')
all_data['P_emaildomain_Country'] = all_data['P_emaildomain'].apply(lambda x: x.split('.')[-1])
all_data['P_emaildomain_Country'] = all_data['P_emaildomain_Country'].apply(lambda x: np.nan if x in ['email_not_provided', 'com', 'net', 'gmail', 'edu'] else x)
all_data['R_emaildomain_Country'] = all_data['R_emaildomain'].apply(lambda x: x.split('.')[-1])
all_data['R_emaildomain_Country'] = all_data['R_emaildomain_Country'].apply(lambda x: np.nan if x in ['email_not_provided', 'com', 'net', 'gmail', 'edu'] else x)
all_data['P_emaildomain_prefix'] = all_data['P_emaildomain'].apply(lambda x: x.split('.')[0])
all_data['R_emaildomain_prefix'] = all_data['R_emaildomain'].apply(lambda x: x.split('.')[0])
# +
M_feature_col = M_feature.copy()
M_feature_col.remove('M4')
all_data['M_not_null'] = all_data[M_feature_col].notnull().sum(axis=1)
for m in M_feature_col:
all_data[m] = all_data[m].map({'T':1,'F':0})
all_data['M_sum'] = all_data[M_feature_col].sum(axis=1)
all_data['collect_agree_ratio'] = all_data['M_sum'] / all_data['M_not_null']
# +
all_data['DeviceInfo'] = all_data['DeviceInfo'].fillna('unknown_device').str.lower()
all_data['DeviceInfo_c'] = all_data['DeviceInfo']
device_match_dict = {
'sm':'samsung',
'samsung':'samsung',
'huawei':'huawei',
'moto':'moto',
'rv':'rv:',
'trident':'trident',
'lg':'lg',
'htc':'htc',
'blade':'blade',
'windows':'windows',
'lenovo':'lenovo',
'linux':'linux',
'f3':'f3',
'f5':'f5',
'ios':'apple',
'mac':'apple'
}
for dev_type_key, dev_type_value in device_match_dict.items():
print(dev_type_key)
all_data['DeviceInfo_c'] = all_data['DeviceInfo_c'].apply(lambda x: dev_type_value if x.find(dev_type_key)!=-1 else x)
all_data['DeviceInfo_c'] = all_data['DeviceInfo_c'].apply(lambda x: 'other_d_type' if x not in device_match_dict.values() else x)
# +
all_data['id_30'] = all_data['id_30'].fillna('unknown').str.lower()
id_30_c_dict={'win':'windows','mac':'mac','ios':'ios','and':'android','linux':'linux'}
all_data['id_30_c'] = all_data['id_30']
for k, v in id_30_c_dict.items():
all_data['id_30_c'] = all_data['id_30_c'].apply(lambda x: v if x.find(k)!=-1 else x)
all_data['id_30_v'] = all_data['id_30'].apply(lambda x: ''.join([i for i in x if i.isdigit()]))
all_data.loc[all_data['id_30']=='unknown','id_30_c'] = np.nan
# +
all_data['id_31'] = all_data['id_31'].fillna('unknown').str.lower()
all_data['id_31_c'] = all_data['id_31']
br_list = ['safari','ie','chrome','edge','firefox','samsung','opera','google','android']
for b in br_list:
all_data['id_31_c'] = all_data['id_31_c'].apply(lambda x: b if x.find(b)!=-1 else x)
all_data.loc[all_data['id_31']=='unknown','id_31_c'] = np.nan
all_data.loc[(~all_data['id_31_c'].isin(br_list+['other']))&(all_data['id_31_c'].notnull()),'id_31_c'] = 'other'
all_data['is_mobile_browser'] = all_data['id_31'].apply(lambda x: 1 if x.find('mobile')!=-1 else 0)
all_data['id_31_v'] = all_data['id_31'].apply(lambda x: ''.join([i for i in x if i.isdigit()]))
# +
START_DATE = '2017-11-30'
import datetime
startdate = datetime.datetime.strptime(START_DATE, '%Y-%m-%d')
all_data['TransactionDT'] = all_data['TransactionDT'].apply(lambda x: (startdate + datetime.timedelta(seconds = x)))
all_data['dow'] = all_data['TransactionDT'].dt.dayofweek
all_data['day'] = all_data['TransactionDT'].dt.day
all_data['Transaction_hour'] = all_data['TransactionDT'].dt.hour
# -
# encoding 해도 좋은 feature, category수가 많고, 개수에 따라 분포가 변함
category_encoding = ['card1','card2','card3','card5',
'C1','C2','C3','C4','C5','C6','C7','C8','C9','C10','C11','C12','C13','C14',
'D1','D2','D3','D4','D5','D6','D7','D8','D9',
'addr1','addr2',
'dist1','dist2',
'P_emaildomain', 'R_emaildomain',
'id_01','id_02','id_03','id_04','id_05','id_06','id_07','id_08','id_09','id_10',
'id_11','id_13','id_14','id_17','id_18','id_19','id_20','id_21','id_22','id_24',
'id_25','id_26','id_30','id_31','id_32','id_33',
'DeviceInfo','DeviceInfo_c','id_30_c','id_30_v','id_31_v',
]
for col in tqdm_notebook(category_encoding):
fq_encode = all_data[col].value_counts().to_dict()
all_data[col+'_fq_enc'] = all_data[col].map(fq_encode)
def fast_groupby(data, col1, col2, agg_type):
new_col_name = f'{col1}_{col2}_{agg_type}'
temp = data.groupby([col1])[col2].agg([agg_type]).reset_index().rename(columns={agg_type:new_col_name})
temp.index = list(temp[col1])
temp = temp[new_col_name].to_dict()
data[new_col_name] = data[col1].map(temp)
return data
# +
########################### TransactionAmt
# Let's add some kind of client uID based on cardID ad addr columns
# The value will be very specific for each client so we need to remove it
# from final feature. But we can use it for aggregations.
all_data['uid'] = all_data['card1'].astype(str)+'_'+all_data['card2'].astype(str)
all_data['uid2'] = all_data['uid'].astype(str)+'_'+all_data['card3'].astype(str)+'_'+all_data['card4'].astype(str)
all_data['uid3'] = all_data['uid2'].astype(str)+'_'+all_data['addr1'].astype(str)+'_'+all_data['addr2'].astype(str)
# For our model current TransactionAmt is a noise
# https://www.kaggle.com/kyakovlev/ieee-check-noise
# (even if features importances are telling contrariwise)
# There are many unique values and model doesn't generalize well
# Lets do some aggregations
i_cols = ['card1','card2','card3','card5','uid','uid2','uid3']
for col in tqdm_notebook(i_cols):
for agg_type in ['mean','std']:
all_data = fast_groupby(all_data, col, 'TransactionAmt', agg_type )
# -
all_data['ymd'] = all_data['TransactionDT'].dt.year*10000+all_data['TransactionDT'].dt.month*100 + all_data['TransactionDT'].dt.day
all_data = fast_groupby(all_data,'ymd','TransactionDT','count')
all_data = fast_groupby(all_data,'ymd','TransactionAmt','sum')
uid3_product_cd_agg = all_data.groupby(['uid3','ProductCD'])['dow'].agg({'uid2_dayofweek_mean':'mean','uid2_dayofweek_std':'std'}).reset_index()
all_data = all_data.merge(uid3_product_cd_agg, how='left', on = ['uid3','ProductCD'])
all_data['TransactionAmt_decimal_count'] = ((all_data['TransactionAmt'] - all_data['TransactionAmt'].astype(int))).astype(str).apply(lambda x: len(x.split('.')[1]))
all_data['TransactionAmt_decimal'] = ((all_data['TransactionAmt'] - all_data['TransactionAmt'].astype(int)) * 1000).astype(int)
# +
def account_start_date(val):
if np.isnan(val) :
return np.NaN
else:
days= int( str(val).split('.')[0])
return pd.Timedelta( str(days) +' days')
for i in ['D1', 'D2', 'D4', 'D8','D10', 'D15']:
all_data['account_start_day'] = all_data[i].apply(account_start_date)
# account_make_date 컴퓨터가 인식할 수 있도록 수치형으로 바꿔 줌.
all_data['account_make_date'] = (all_data['TransactionDT'] - all_data['account_start_day']).dt.date
all_data['account_make_date_{}'.format(i)] = (10000 * pd.to_datetime(all_data['account_make_date']).dt.year) + (100 * pd.to_datetime(all_data['account_make_date']).dt.month) + (1 * pd.to_datetime(all_data['account_make_date']).dt.day)
del all_data['account_make_date']
del all_data['account_start_day']
# -
card1_d1_pcd_mean = all_data.groupby(['card1','account_make_date_D1','ProductCD'])['TransactionAmt'].agg({'card1_make_date_D1_productCD_Amt_mean':'mean','card1_make_date_D1_productCD_Amt_std':'std'}).reset_index()
all_data = all_data.merge(card1_d1_pcd_mean, how='left', on = ['card1','account_make_date_D1','ProductCD'])
# +
from sklearn.decomposition import LatentDirichletAllocation as LDA
from sklearn.feature_extraction.text import CountVectorizer
import gc
def add_lda_feature(all_data, col1, col2):
n_comp = 5
print("add_lda_feature", col1, col2, n_comp)
temp = all_data[[col1,col2]]
col1col2_dict = {}
def col1col2(row):
nonlocal col1, col2
if np.isnan(row[col1]):
return
if np.isnan(row[col2]):
return
col1col2_dict.setdefault(row[col1], []).append(str(row[col2]))
temp.apply(lambda row:col1col2(row) , axis=1)
col1_keys = list(col1col2_dict.keys())
col1col2_dict_as_sentence = [' '.join(col1col2_dict[c]) for c in col1_keys]
_as_matrix = CountVectorizer().fit_transform(col1col2_dict_as_sentence)
topics_of_col1 = LDA(n_components=n_comp, n_jobs=-1,random_state=0).fit_transform(_as_matrix)
col1_frame = pd.DataFrame(col1col2_dict.keys(),columns=[col1])
topics_of_col1 = pd.DataFrame(topics_of_col1, columns=[f'{col1}_{col2}_LDA_{i}' for i in range(n_comp)])
col1_frame = pd.concat([col1_frame, topics_of_col1], axis=1)
col1_frame.index = list(col1_frame[col1])
new_col_name=[f'{col1}_{col2}_LDA_{i}' for i in range(n_comp)]
for c in new_col_name:
print(c)
temp_df = col1_frame[c].to_dict()
all_data[c] = all_data[col1].map(temp_df)
#all_data = all_data.merge(col1_frame, on=[col1], how='left')
del col1col2_dict
del topics_of_col1
del _as_matrix
del col1_frame
#del topics_of_col1
gc.collect()
return all_data, new_col_name
# +
from sklearn.decomposition import NMF
def add_nmf_feature(all_data, col1, col2):
n_comp = 5
print("add_nmf_feature", col1, col2, n_comp)
temp = all_data[[col1,col2]]
col1col2_dict = {}
def col1col2(row):
nonlocal col1, col2
if pd.isna(row[col1]):
return
if pd.isna(row[col2]):
return
col1col2_dict.setdefault(row[col1], []).append(str(row[col2]))
temp.apply(lambda row:col1col2(row) , axis=1)
col1_keys = list(col1col2_dict.keys())
col1col2_dict_as_sentence = [' '.join(col1col2_dict[c]) for c in col1_keys]
_as_matrix = CountVectorizer().fit_transform(col1col2_dict_as_sentence)
topics_of_col1 = LDA(n_components=n_comp, n_jobs=-1,random_state=0).fit_transform(_as_matrix)
col1_frame = pd.DataFrame(col1col2_dict.keys(),columns=[col1])
topics_of_col1 = pd.DataFrame(topics_of_col1, columns=[f'{col1}_{col2}_NMF_{i}' for i in range(n_comp)])
col1_frame = pd.concat([col1_frame, topics_of_col1], axis=1)
col1_frame.index = list(col1_frame[col1])
new_col_name=[f'{col1}_{col2}_NMF_{i}' for i in range(n_comp)]
for c in new_col_name:
print(c)
temp_df = col1_frame[c].to_dict()
all_data[c] = all_data[col1].map(temp_df)
#all_data = all_data.merge(col1_frame, on=[col1], how='left')
del col1col2_dict
del topics_of_col1
del _as_matrix
del col1_frame
#del topics_of_col1
gc.collect()
return all_data, new_col_name
# -
# (0.9404788466588169, 0.9353552350432497, 0.9117537267548267, 0.9589567433316727)
all_data,_ = add_lda_feature(all_data, 'card1','addr1')
all_data,_ = add_lda_feature(all_data, 'card1','id_20')
all_data,_ = add_nmf_feature(all_data, 'card4','DeviceInfo_c')
def category_combine_feature(all_data, c1, c2):
new_col_list = []
new_col = f'{c1}_{c2}_combine'
all_data[new_col] = all_data[c1].astype(str) + '_' + all_data[c2].astype(str)
all_data[new_col] = pd.factorize(all_data[new_col], sort=True)[0]
new_col_freq = new_col + '_fq_enc_combine'
fq_encode = all_data[new_col].value_counts().to_dict()
all_data[new_col_freq] = all_data[new_col].map(fq_encode)
new_col_list.append(new_col)
new_col_list.append(new_col_freq)
if new_col not in ['card1_card2_combine']:
for agg_type in ['mean','std']:
new_col_name = new_col+'_TransactionAmt_'+agg_type
#temp_df['TransactionAmt'] = temp_df['TransactionAmt'].astype(int)
temp_df = all_data.groupby([new_col])['TransactionAmt'].agg([agg_type]).reset_index().rename(
columns={agg_type: new_col_name})
temp_df.index = list(temp_df[new_col])
temp_df = temp_df[new_col_name].to_dict()
all_data[new_col_name] = all_data[new_col].map(temp_df)
new_col_list.append(new_col_name)
print(new_col_list)
return all_data, new_col_list
# (0.9765178882384341, 0.9750723088222574, 0.9691847487013969, 0.9809598689431178)
# +
for a in ['addr1']:
for i in ['id_32','id_14','id_19']:
new_col = f'{a}_{i}_combine'
if new_col in all_data.columns:
continue
print(new_col)
all_data, new_col_list = category_combine_feature(all_data,a,i)
for c in ['card1']:
for i in ['card2','addr1']:
new_col = f'{c}_{i}_combine'
if new_col in all_data.columns:
continue
print(new_col)
all_data, new_col_list = category_combine_feature(all_data,c,i)
#all_data, new_col_list = category_combine_feature(all_data,'id_17','id_33')
# -
all_data_temp = pd.concat([train_transaction,test_transaction])
all_data_temp['uid'] = all_data_temp['card1'].astype(str)+'_'+all_data_temp['card2'].astype(str)
all_data_temp['uid2'] = all_data_temp['uid'].astype(str)+'_'+all_data_temp['card3'].astype(str)+'_'+all_data_temp['card4'].astype(str)
all_data_temp['uid3'] = all_data_temp['uid2'].astype(str)+'_'+all_data_temp['addr1'].astype(str)+'_'+all_data_temp['addr2'].astype(str)
all_data_temp['uid3_next_click'] = all_data_temp['TransactionDT'] - all_data_temp.groupby(['uid3'])['TransactionDT'].shift(-1)
all_data_temp = all_data_temp.reset_index(drop=True)
all_data['uid3_next_click'] = all_data_temp['uid3_next_click']
card1_pcd_mean = all_data.groupby(['card1','account_make_date_D2','ProductCD'])['TransactionAmt'].agg({'card1_make_date_D2_productCD_Amt_mean':'mean','card1_make_date_D2_productCD_Amt_std':'std'}).reset_index()
all_data = all_data.merge(card1_pcd_mean, how='left', on = ['card1','account_make_date_D2','ProductCD'])
card1_pcd_mean = all_data.groupby(['card1','addr1','account_make_date_D1','ProductCD'])['TransactionAmt'].agg({'card1_addr1_make_date_D1_ProductCD_Amt_mean':'mean','card1_addr1_make_date_D1_ProductCD_Amt_std':'std'}).reset_index()
all_data = all_data.merge(card1_pcd_mean, how='left', on = ['card1','addr1','ProductCD','account_make_date_D1'])
card1_pcd_mean = all_data.groupby(['card1','id_20','account_make_date_D1','ProductCD'])['TransactionAmt'].agg({'card1_id_20_make_date_D1_ProductCD_Amt_mean':'mean','card1_id_20_make_date_D1_ProductCD_Amt_std':'std'}).reset_index()
all_data = all_data.merge(card1_pcd_mean, how='left', on = ['card1','id_20','ProductCD','account_make_date_D1'])
for p in all_data['ProductCD'].unique():
all_data.loc[all_data['ProductCD']==p, 'ProductCD_Amt_expandmean'] = all_data.loc[all_data['ProductCD']==p, 'TransactionAmt'].cumsum()/np.arange(1,all_data.loc[all_data['ProductCD']==p, 'TransactionAmt'].shape[0]+1)
from sklearn.preprocessing import StandardScaler,MinMaxScaler
for p in all_data['ProductCD'].unique():
mm = MinMaxScaler()
all_data.loc[all_data['ProductCD']==p, 'TransactionAmt'] = mm.fit_transform(all_data.loc[all_data['ProductCD']==p, 'TransactionAmt'].values.reshape(-1,1))
# +
card1_pcd_mean = all_data.groupby(['card1','account_make_date_D1','ProductCD'])['dist1'].agg({'card1_make_date_D1_ProductCD_dist1_mean':'mean','card1_make_date_D1_ProductCD_dist1_std':'std'}).reset_index()
all_data = all_data.merge(card1_pcd_mean, how='left', on = ['card1','ProductCD','account_make_date_D1'])
card1_pcd_mean = all_data.groupby(['card1','addr1','account_make_date_D1','ProductCD'])['dist1'].agg({'card1_addr1_make_date_D1_ProductCD_dist1_mean':'mean','card1_addr1_make_date_D1_ProductCD_dist1_std':'std'}).reset_index()
all_data = all_data.merge(card1_pcd_mean, how='left', on = ['card1','addr1','ProductCD','account_make_date_D1'])
card1_pcd_mean = all_data.groupby(['card1','card2','account_make_date_D1','ProductCD'])['dist1'].agg({'card1_card2_make_date_D1_ProductCD_dist1_mean':'mean','card1_card2_make_date_D1_ProductCD_dist1_std':'std'}).reset_index()
all_data = all_data.merge(card1_pcd_mean, how='left', on = ['card1','card2','ProductCD','account_make_date_D1'])
# +
card1_pcd_mean = all_data.groupby(['id_02','ProductCD'])['TransactionAmt'].agg({'id_02_ProductCD_amt_mean':'mean'}).reset_index()
all_data = all_data.merge(card1_pcd_mean, how='left', on = ['id_02','ProductCD'])
# -
card1_d1_pcd_mean = all_data.groupby(['card1','account_make_date_D1','ProductCD'])['Transaction_hour'].agg({'card1_make_date_D1_productCD_hour_mean':'mean'}).reset_index()
all_data = all_data.merge(card1_d1_pcd_mean, how='left', on = ['card1','account_make_date_D1','ProductCD'])
all_data['card1_maked1_pcd'] = all_data['card1'].astype(str) + all_data['account_make_date_D1'].astype(str) + all_data['ProductCD'].astype(str)
all_data['card1_maked1_pcd'] = pd.factorize(all_data['card1_maked1_pcd'])[0]
new_col_freq = 'card1_maked1_pcd' + '_fq_enc_combine'
fq_encode = all_data['card1_maked1_pcd'].value_counts().to_dict()
all_data[new_col_freq] = all_data['card1_maked1_pcd'].map(fq_encode)
for col in ['account_make_date_D1']:
new_col_freq = col + '_fq_enc_combine'
fq_encode = all_data[col].value_counts().to_dict()
all_data[new_col_freq] = all_data[col].map(fq_encode)
# +
all_data['d1_d2'] = all_data['account_make_date_D1'].astype(str) + '_' + all_data['account_make_date_D2'].astype(str) + '_' + all_data['ProductCD'].astype(str)
all_data['same_d1_d2'] = 0
all_data.loc[all_data['account_make_date_D1'] == all_data['account_make_date_D2'], 'same_d1_d2'] = 1
fq_encode = all_data['d1_d2'].value_counts().to_dict()
all_data['d1_d2_fq_enc'] = all_data['d1_d2'].map(fq_encode)
# +
object_col = []
for col in all_data.columns:
if all_data[col].dtypes == 'object':
object_col.append(col)
for col in object_col:
all_data[col] = pd.factorize(all_data[col], sort=True)[0]
# +
x_train = all_data.loc[all_data['isFraud'].notnull()]#.sample(frac=0.3,random_state=0).reset_index(drop=True)
#x_train = x_train.loc[x_train['TransactionDT']>2500000].reset_index(drop=True) # like lb hist dist
y_train = x_train.isFraud
x_test = all_data.loc[all_data['isFraud'].isnull()].reset_index(drop=True)
# -
def mean_encoding(train, test, col):
temp_dict = train.groupby([col])['isFraud'].agg(['mean']).reset_index().rename(
columns={'mean': col+'_target_mean'})
temp_dict.index = temp_dict[col].values
temp_dict = temp_dict[col+'_target_mean'].to_dict()
train[col+'_target_mean'] = train[col].map(temp_dict)
test[col+'_target_mean'] = test[col].map(temp_dict)
return train, test
x_train, x_test = mean_encoding(x_train, x_test, 'ProductCD')
x_train, x_test = mean_encoding(x_train, x_test, 'M4')
param = {
#'bagging_freq': 5,
#'bagging_fraction': 0.8,
'boost_from_average':'true',
'boost': 'gbdt',
'feature_fraction': 0.7,
'bagging_fraction': 0.7,
'learning_rate': 0.01,
'subsample_freq':1,
'max_bin':255,
'max_depth': -1,
'metric':'auc',
'num_leaves': 256,
'num_threads': 32,
'tree_learner': 'serial',
'objective': 'binary',
#'scale_pos_weight':97,
'verbosity': 1,
'seed':42
#'reg_lambda': 0.3,
}
def get_train_column(train, must_delete_col):
delete_col = must_delete_col.copy()
unique_col = []
for c in train.columns:
if train[c].fillna(-9999).nunique()<2:
unique_col.append(c)
delete_col = delete_col + unique_col
train_columns = [c for c in train.columns if c not in delete_col]
return train_columns, delete_col
# +
category_feature_dict = dict()
delete_feature_dict = dict()
w_type = [x_train['ProductCD'].value_counts().index[0]]
other_type = list(x_train['ProductCD'].value_counts().index[1:].values)
category_feature_dict[str(w_type)] = ['addr1','addr2','P_emaildomain']
category_feature_dict[str(other_type)] = ['addr1','addr2','P_emaildomain','R_emaildomain','id_13','id_15', 'id_33', 'id_37', 'id_38','id_30_v','id_31_v']
#import pickle
#delete_v_feature = []
#with open('noise_v_cols.pickle', 'rb') as file:
# delete_v_feature = pickle.load(file)
# print(delete_v_feature)
#'same_d1_d2','d1_d2','d1_d2_fq_enc'
delete_feautre_list = ['DT_M','card1_addr1_maked1_pcd','card1_addr1_maked1_pcd_fq_enc_combine','C13_over_cf_sum','TransactionID','isFraud','TransactionDT','bank_type','uid','uid2','uid3','ymd','ym','is_D_max','addr1_null']
delete_feature_dict[str(w_type)] = delete_feautre_list +['P_emaildomain_Country','card1_TransactionAmt_std']#+ delete_v_feature #+ M_feature
delete_feature_dict[str(other_type)] = delete_feautre_list
# +
#must_del_col_list = ['DT_M','card1_addr1_maked1_pcd','card1_addr1_maked1_pcd_fq_enc_combine','C13_over_cf_sum','TransactionID','isFraud','TransactionDT','bank_type','uid','uid2','uid3','ymd','ym','is_D_max','addr1_null'] + delete_v_feature #+ M_feature
debug = False
oof_train = np.zeros(len(x_train))
oof_test = np.zeros(len(x_test))
NFOLD =5
SEED = 42
w_type = [x_train['ProductCD'].value_counts().index[0]]
other_type = list(x_train['ProductCD'].value_counts().index[1:].values)
product_type = list()
product_type.append(w_type)
product_type.append(other_type)
feature_importance_df = pd.DataFrame()
cv_list ={}
for p_type in product_type:
x_train_type = x_train.loc[x_train['ProductCD'].isin(p_type)]
y_train_type = x_train_type['isFraud']
x_test_type = x_test.loc[x_test['ProductCD'].isin(p_type)]
category_feature = category_feature_dict[str(p_type)].copy()
must_del_col_list = delete_feature_dict[str(p_type)].copy()
#param = org_param.copy()#lgbm_param_dict[str(p_type)]
#must_del_col_list_temp = must_del_col_list.copy()
#must_del_col_list_temp = must_del_col_list_temp + cat_combinefeature_dict[str(p_type)]
train_columns, _ = get_train_column(x_train_type, must_del_col_list)
print(x_train_type.shape)
print(len(train_columns), train_columns)
kfold = StratifiedKFold(n_splits=NFOLD, shuffle=True, random_state=SEED)
#split_groups = x_train_type['DT_M']
#kfold = GroupKFold(n_splits=6)
oof_train_type = np.zeros((x_train_type.shape[0],))
oof_test_type = np.zeros((x_test_type.shape[0],))
feature_importance_type = pd.DataFrame()
type_score = 0
for i, (train_index, cross_index) in enumerate(kfold.split(x_train_type,y_train_type),1):
print(f"TYPE {p_type} - {i} FOLD Start")
x_tr = x_train_type.iloc[train_index][train_columns]
x_cr = x_train_type.iloc[cross_index][train_columns]
y_tr = y_train_type.iloc[train_index]
y_cr = y_train_type.iloc[cross_index]
dtrain = lgb.Dataset(x_tr, label=y_tr, silent=True)
dcross = lgb.Dataset(x_cr, label=y_cr, silent=True)
clf = lgb.train(param, train_set=dtrain, num_boost_round=15000, valid_sets=[dtrain, dcross],
early_stopping_rounds=100, verbose_eval=500, categorical_feature=category_feature)
oof_train_type[cross_index] = clf.predict(x_cr)
oof_test_type += clf.predict(x_test_type[train_columns])/NFOLD
feature_importance = pd.DataFrame()
feature_importance["Feature"] = x_tr.columns
feature_importance["Importance"] = clf.feature_importance()
feature_importance["FOLD"] = i
feature_importance_type = pd.concat([feature_importance_type, feature_importance])
cv_score = roc_auc_score(y_cr, oof_train_type[cross_index])
type_score += cv_score / NFOLD
print(f"{i} FOLD Score: ", cv_score)
if debug is True:
break
print("Total CV: ", type_score)
print("Total CV2: ", roc_auc_score(y_train_type, oof_train_type))
cv_list[str(p_type)] = roc_auc_score(y_train_type, oof_train_type)
oof_train[x_train_type.index] = oof_train_type
oof_test[x_test_type.index] = oof_test_type
feature_importance_type["TYPE"] = str(p_type)
feature_importance_df = pd.concat([feature_importance_df, feature_importance_type])
# -
pd.DataFrame(oof_train,columns=[f'train_seed_{SEED}']).to_csv(f'../oof/goss_oof_train_seed_{SEED}.csv',index=False)
pd.DataFrame(oof_test,columns=[f'test_seed_{SEED}']).to_csv(f'../oof/goss_oof_test_seed_{SEED}.csv',index=False)
#card1_addr1 addr1_id_32 card1 card2 addr1_id_19 addr1_id_14
print(roc_auc_score(y_train, oof_train))
print(np.mean([c for c in cv_list.values()]))
print(cv_list)
#card1_addr1 addr1_id_32 card1 card2 addr1_id_19 addr1_id_14
print(roc_auc_score(y_train, oof_train))
print(np.mean([c for c in cv_list.values()]))
print(cv_list)
#card1_addr1 addr1_id_32 card1 card2 addr1_id_19 addr1_id_14
print(roc_auc_score(y_train, oof_train))
print(np.mean([c for c in cv_list.values()]))
print(cv_list)
# +
fi = feature_importance_df.groupby(['Feature'])['Importance'].mean().reset_index()
fi = fi.sort_values('Importance',ascending=False)
fi.to_csv('../importance/goss.csv',index=False)
fi
# -
sub = pd.read_csv('../input/sample_submission.csv')
sub['isFraud'] = oof_test
sub.to_csv('../output/goss.csv',index=False)
# #### Seed Sampling
# +
#must_del_col_list = ['DT_M','card1_addr1_maked1_pcd','card1_addr1_maked1_pcd_fq_enc_combine','C13_over_cf_sum','TransactionID','isFraud','TransactionDT','bank_type','uid','uid2','uid3','ymd','ym','is_D_max','addr1_null'] + delete_v_feature #+ M_feature
debug = False
oof_train = np.zeros(len(x_train))
oof_test = np.zeros(len(x_test))
NFOLD =5
SEED = 42
w_type = [x_train['ProductCD'].value_counts().index[0]]
other_type = list(x_train['ProductCD'].value_counts().index[1:].values)
product_type = list()
product_type.append(w_type)
product_type.append(other_type)
for seed in range(10):
print(seed)
param['seed'] = seed
feature_importance_df = pd.DataFrame()
cv_list ={}
for p_type in product_type:
x_train_type = x_train.loc[x_train['ProductCD'].isin(p_type)]
y_train_type = x_train_type['isFraud']
x_test_type = x_test.loc[x_test['ProductCD'].isin(p_type)]
category_feature = category_feature_dict[str(p_type)].copy()
must_del_col_list = delete_feature_dict[str(p_type)].copy()
#param = org_param.copy()#lgbm_param_dict[str(p_type)]
#must_del_col_list_temp = must_del_col_list.copy()
#must_del_col_list_temp = must_del_col_list_temp + cat_combinefeature_dict[str(p_type)]
train_columns, _ = get_train_column(x_train_type, must_del_col_list)
print(x_train_type.shape)
print(len(train_columns), train_columns)
kfold = StratifiedKFold(n_splits=NFOLD, shuffle=True, random_state=SEED)
#split_groups = x_train_type['DT_M']
#kfold = GroupKFold(n_splits=6)
oof_train_type = np.zeros((x_train_type.shape[0],))
oof_test_type = np.zeros((x_test_type.shape[0],))
feature_importance_type = pd.DataFrame()
type_score = 0
for i, (train_index, cross_index) in enumerate(kfold.split(x_train_type,y_train_type),1):
print(f"TYPE {p_type} - {i} FOLD Start")
x_tr = x_train_type.iloc[train_index][train_columns]
x_cr = x_train_type.iloc[cross_index][train_columns]
y_tr = y_train_type.iloc[train_index]
y_cr = y_train_type.iloc[cross_index]
dtrain = lgb.Dataset(x_tr, label=y_tr, silent=True)
dcross = lgb.Dataset(x_cr, label=y_cr, silent=True)
clf = lgb.train(param, train_set=dtrain, num_boost_round=15000, valid_sets=[dtrain, dcross],
early_stopping_rounds=100, verbose_eval=500, categorical_feature=category_feature)
oof_train_type[cross_index] = clf.predict(x_cr)
oof_test_type += clf.predict(x_test_type[train_columns])/NFOLD
feature_importance = pd.DataFrame()
feature_importance["Feature"] = x_tr.columns
feature_importance["Importance"] = clf.feature_importance()
feature_importance["FOLD"] = i
feature_importance_type = pd.concat([feature_importance_type, feature_importance])
cv_score = roc_auc_score(y_cr, oof_train_type[cross_index])
type_score += cv_score / NFOLD
print(f"{i} FOLD Score: ", cv_score)
if debug is True:
break
print("Total CV: ", type_score)
print("Total CV2: ", roc_auc_score(y_train_type, oof_train_type))
cv_list[str(p_type)] = roc_auc_score(y_train_type, oof_train_type)
oof_train[x_train_type.index] = oof_train_type
oof_test[x_test_type.index] = oof_test_type
feature_importance_type["TYPE"] = str(p_type)
feature_importance_df = pd.concat([feature_importance_df, feature_importance_type])
pd.DataFrame(oof_train,columns=[f'train_seed_{seed}']).to_csv(f'../oof/goss_oof_train_seed_{seed}.csv',index=False)
pd.DataFrame(oof_test,columns=[f'test_seed_{seed}']).to_csv(f'../oof/goss_oof_test_seed_{seed}.csv',index=False)
# -
# Total CV2: 0.974203181381347, Total CV2: 0.9831092711580688
|
IEEE-CIS Fraud Detection/code/Model/Model2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Object Oriented Programming
#
# For this lesson we will construct our knowledge of OOP in Python by building on the following topics:
#
# * Objects
# * Using the *class* keyword
# * Creating class attributes
# * Creating methods in a class
# * Learning about Inheritance
# * Learning about Polymorphism
# * Learning about Special Methods for classes
# ## Objects
# In Python, *everything is an object*. Remember from previous lectures we can use type() to check the type of object something is:
print(type(1))
print(type([]))
print(type(()))
print(type({}))
# So we know all these things are objects, so how can we create our own Object types? That is where the <code>class</code> keyword comes in.
# ## class
# +
# Create a new object type called Sample
class Sample:
pass
# Instance of Sample
x = Sample()
print(type(x))
# -
# By convention we give classes a name that starts with a capital letter. x **instantiates** the Sample class.
#
# Inside of the class we currently just have pass. But we can define class attributes and methods.
#
# ## Instance Attributes
# The syntax for creating an attribute is:
#
# self.attribute = something
#
# There is a special method called:
#
# __init__()
#
# This method is used to initialize the attributes of an object. For example:
# +
class Dog:
def __init__(self,breed='sdd'):
self.breed = breed
sam = Dog(breed='Lab')
frank = Dog(breed='Huskie')
# -
# The special method
#
# __init__()
# is called automatically right after the object has been created:
#
# def __init__(self, breed):
# Each attribute in a class definition begins with a reference to the instance object. It is by convention named self. The breed is the argument. The value is passed during the class instantiation.
#
# self.breed = breed
# Now we have created two instances of the Dog class. With two breed types, we can then access these attributes like this:
sam.breed
frank.breed
# ### Class Attributes
# These Class Object Attributes are the same for any instance of the class. For example, we could create the attribute *species* for the Dog class. Dogs, regardless of their breed, name, or other attributes, will always be mammals. We apply this logic in the following manner:
class Dog:
# Class Object Attribute
species = 'mammal'
def __init__(self,breed,name):
self.breed = breed
self.name = name
sam = Dog('Lab','Sam')
# Note that the Class Object Attribute is defined outside of any methods in the class. Also by convention, we place them first before the init.
sam.species
# ### Python Names
# Name (also called identifier) is simply a name given to objects. Name is a way to access the underlying object.For example, when we do the assignment a = 2, here 2 is an object stored in memory and a is the name we associate it with. We can get the address (in RAM) of some object through the built-in function, id().
# Please visit https://www.programiz.com/python-programming/namespace
a = 2
print('id(2) =', id(2))
print('id(a) =', id(a))
# +
a = 2
print('id(a) =', id(a))
a = a+1
print('id(a) =', id(a))
print('id(3) =', id(3))
b = 2
print('id(2) =', id(b))
print('id(2) =', id(2))
# -
# Initially, an object 2 is created and the name a is associated with it, when we do a = a+1, a new object 3 is created and now a associates with this object.
#
# Note that id(a) and id(3) have same values. Furthermore, when we do b = 2, the new name b gets associated with the previous object 2.
#
# ### Python Namespace
#
# #### namespace is a collection of names. In Python, you can imagine a namespace as a mapping of every name, you have defined, to corresponding objects. They’re usually implemented as Python dictionaries, although this is abstracted away ( __dict__ ).
#
# #### Different namespaces can co-exist at a given time but are completely isolated.
#
# #### A namespace containing all the built-in names is created when we start the Python interpreter and exists as long we don't exit.
#
# #### This is the reason that built-in functions like id(), print() etc. are always available to us from any part of the program. Each module creates its own global namespace.
#
# #### These different namespaces are isolated. Hence, the same name that may exist in different modules do not collide.
#
# #### Modules can have various functions and classes. A local namespace is created when a function is called, which has all the names defined in it. Similar, is the case with class. Following diagram may help to clarify this concept.
#
# Nested Namespaces in Python Programming
#
# 
# ### Python Class attribute vs. Instance attribute: What’s the Difference?
# A Python class attribute is an attribute of the class (circular, I know), rather than an attribute of an instance of a class.
class MyClass(object):
class_var = 1
def __init__(self, i_var):
self.i_var = i_var
# Note that all instances of the class have access to class_var, and that it can also be accessed as a property of the class itself:
# +
foo = MyClass(2)
bar = MyClass(3)
foo.class_var, foo.i_var
## 1, 2
bar.class_var, bar.i_var
## 1, 3
MyClass.class_var ## <— This is key
## 1
# -
# For Java or C++ programmers, the class attribute is similar—but not identical—to the static member.
# ### Python Class Attributes: An Overly Thorough Guide
# - For details see: https://www.toptal.com/python/python-class-attributes-an-overly-thorough-guide
# #### Private variable in classes
# In Python, you don't write to other classes' instance or class variables. Python drops that pretence of security and encourages programmers to be responsible. If you want to emulate private variables for some reason, you can always use the __ prefix from PEP 8. Python mangles the names of variables like __foo so that they're not easily visible to code outside the class that contains them (although you can get around it if you're determined enough, just like you can get around Java's protections if you work at it).
#
# In Python, mangling is used for "private" class members which are designated as such by giving them a name with two leading underscores and no more than one trailing underscore. For example, __thing will be mangled, as will ___thing and __thing_, but __thing__ and __thing___ will not. Python's runtime does not restrict access to such members, the mangling only prevents name collisions if a derived class defines a member with the same name.
#
# On encountering name mangled attributes, Python transforms these names by prepending a single underscore and the name of the enclosing class
# +
class Test(object):
def __mangled_name(self):
pass
def normal_name(self):
pass
t = Test()
f = [attr for attr in dir(t) if 'name' in attr]
#['_Test__mangled_name', 'normal_name']
# -
# ### Descriptive: Naming Styles according to PEP 8
#
# The following special forms using leading or trailing underscores are recognized (these can generally be combined with any case convention):
#
#
#
# 1. _single_leading_underscore: weak "internal use" indicator. E.g. from M
# import * does not import objects whose name starts with an underscore.
# 2. single_trailing_underscore_: used by convention to avoid conflicts with Python keyword, e.g. Tkinter.Toplevel(master, class_='ClassName')
# 3. \__double_leading_underscore: when naming a class attribute, invokes name mangling (inside class FooBar, \_\_boo becomes_FooBar__boo).
# 4. \_\_double_leading_and_trailing_underscore__: "magic" objects or attributes that live in user-controlled namespaces. E.g. \_\_init__, \_\_import__ or \_\_file__. Never invent such names; only use them as documented.
#
class A:
def __init__(self):
self.__var = 123
def printVar(self):
print self.__var
# ## Methods
#
# Methods are functions defined inside the body of a class. They are used to perform operations with the attributes of our objects.
# +
class Circle:
pi = 3.14
# Circle gets instantiated with a radius (default is 1)
def __init__(self, radius=1):
self.radius = radius
self.area = radius * radius * Circle.pi
# Method for resetting Radius
def setRadius(self, new_radius):
self.radius = new_radius
self.area = new_radius * new_radius * self.pi
# Method for getting Circumference
def getCircumference(self):
return self.radius * self.pi * 2
c = Circle()
print('Radius is: ',c.radius)
print('Area is: ',c.area)
print('Circumference is: ',c.getCircumference())
# -
# In the \__init__ method above, in order to calculate the area attribute, we had to call Circle.pi. This is because the object does not yet have its own .pi attribute, so we call the Class Object Attribute pi instead.<br>
# In the setRadius method, however, we'll be working with an existing Circle object that does have its own pi attribute. Here we can use either Circle.pi or self.pi.<br><br>
# Now let's change the radius and see how that affects our Circle object:
# +
c.setRadius(2)
print('Radius is: ',c.radius)
print('Area is: ',c.area)
print('Circumference is: ',c.getCircumference())
# -
# Notice how we used self. notation to reference attributes of the class within the method calls. Review how the code above works and try creating your own method.
#
# ## Inheritance
#
# Inheritance is a way to form new classes using classes that have already been defined. The newly formed classes are called derived classes, the classes that we derive from are called base classes. Important benefits of inheritance are code reuse and reduction of complexity of a program. The derived classes (descendants) override or extend the functionality of base classes (ancestors).
# +
class Animal:
def __init__(self):
print("Animal created")
def whoAmI(self):
print("Animal")
def eat(self):
print("Eating")
class Dog(Animal):
def __init__(self):
Animal.__init__(self)
print("Dog created")
def whoAmI(self):
print("Dog")
def bark(self):
print("Woof!")
# -
d = Dog()
d.whoAmI()
d.eat()
d.bark()
# In this example, we have two classes: Animal and Dog. The Animal is the base class, the Dog is the derived class.
#
# The derived class inherits the functionality of the base class.
#
# * It is shown by the eat() method.
#
# The derived class modifies existing behavior of the base class.
#
# * shown by the whoAmI() method.
#
# Finally, the derived class extends the functionality of the base class, by defining a new bark() method.
# ## Polymorphism
# In Python, *polymorphism* refers to the way in which different object classes can share the same method name, and those methods can be called from the same place even though a variety of different objects might be passed in.
# +
class Dog:
def __init__(self, name):
self.name = name
def speak(self):
return self.name+' says Woof!'
class Cat:
def __init__(self, name):
self.name = name
def speak(self):
return self.name+' says Meow!'
niko = Dog('Niko')
felix = Cat('Felix')
print(niko.speak())
print(felix.speak())
# -
# Here we have a Dog class and a Cat class, and each has a `.speak()` method. When called, each object's `.speak()` method returns a result unique to the object.
#
# There a few different ways to demonstrate polymorphism. First, with a for loop:
for pet in [niko,felix]:
print(pet.speak())
# Another is with functions:
# +
def pet_speak(pet):
print(pet.speak())
pet_speak(niko)
pet_speak(felix)
# + [markdown] slideshow={"slide_type": "slide"}
# In both cases we were able to pass in different object types, and we obtained object-specific results from the same mechanism.
#
# A more common practice is to use abstract classes and inheritance. An abstract class is one that never expects to be instantiated. For example, we will never have an Animal object, only Dog and Cat objects, although Dogs and Cats are derived from Animals:
# +
class Animal:
def __init__(self, name): # Constructor of the class
self.name = name
def speak(self): # Abstract method, defined by convention only
raise NotImplementedError("Subclass must implement abstract method")
class Dog(Animal):
def speak(self):
return self.name+' says Woof!'
class Cat(Animal):
def speak(self):
return self.name+' says Meow!'
fido = Dog('Fido')
isis = Cat('Isis')
print(fido.speak())
print(isis.speak())
# -
# Real life examples of polymorphism include:
# * opening different file types - different tools are needed to display Word, pdf and Excel files
# * adding different objects - the `+` operator performs arithmetic and concatenation
# ## Special Methods
# Finally let's go over special methods. Classes in Python can implement certain operations with special method names. These methods are not actually called directly but by Python specific language syntax. For example let's create a Book class:
class Book:
def __init__(self, title, author, pages):
print("A book is created")
self.title = title
self.author = author
self.pages = pages
def __str__(self):
return "Title: %s, author: %s, pages: %s" %(self.title, self.author, self.pages)
def __len__(self):
return self.pages
def __del__(self):
print("A book is destroyed")
# +
book = Book("Python Rocks!", "<NAME>", 159)
#Special Methods
print(book)
print(len(book))
del book
# -
# The __init__(), __str__(), __len__() and __del__() methods
# These special methods are defined by their use of underscores. They allow us to use Python specific functions on objects created through our class.
|
1.Chapter-Python/2-Python_Basis/courses/04-Object Oriented Programming/01-Object Oriented Programming.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div align="center"><h1>Model Preparation Notebook</h1></div>
# This notebook contains the sequential steps to create the model from the raw dataset.
#
# In this notebook the following steps will be
#
# 1. Importing the required libraries.
# 2. Downloading the data
# 3. Loading the data
# 4. Data Visualization
# 5. Preprocessing & EDA
# 6. Prepare trainable data
# 7. Create Model
# 8. Load it into .pkl file
#
#
# Follow the steps.
# # 1. Importing Libraries :
from google_drive_downloader import GoogleDriveDownloader as gdd
import pandas as pd
# # 2. Downloading the data :
# +
"""
Data Link : https://drive.google.com/file/d/1whUKZ-BB4-VKEanDRVteti5mawjnux5o/view?usp=sharing
We'll take the id section and download the dataset
"""
gdd.download_file_from_google_drive(file_id='1whUKZ-BB4-VKEanDRVteti5mawjnux5o',
dest_path='assets/spotify.csv')
# -
# # 3. Loading the Data :
# +
df = pd.read_csv('assets/spotify.csv')
df.head()
# -
# # 4. Data Visualization:
df.columns
# ### The most important task is to preprocess the data and get the intuition of each features so that we can perform the preprocessing.
#
#
# AcousticNess : Ratio of lyrics and music
#
# Danceability : Level of ease to dance with the song
#
# duration : total time
#
# Energy : Loudness
#
# Explicit : Child prohibited
#
# instrumentalness : instrumental
#
# Key : keys of music
#
# Liveness : Fresh songs
#
# Loudness : Volume
#
# popularity : Likes fom people
#
# release date : date of publish
#
# speechiness : Level of present of lyrics in music
#
# tempo : speed of the song
#
# valence : positivity in music
df.info()
df.describe()
# # 5. Preprocessing & EDA:
# # 6. Prepare trainable data:
# # 7. Create Model :
# # 8. Load it into `.pkl` file :
|
Rhythm-Finder Data Processing and Model Preparation Notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re, pandas as pd, numpy as np
df = pd.read_csv("trantxt/qna.csv", sep="|", header=None, names=["txt","analyst_num", "q_num", "q", "a"])
from pandasql import *
# +
# match = lambda q: sqldf(q, globals())
# q = """
# select distinct txt, analyst_num, max(q_num)
# from df group by txt, analyst_num
# """
# q_num = match(q)
# +
# q_num.to_csv('~/q_num.csv', header=True, sep="|")
# -
df_sub = df.head(100)
df_sub["q_string"] = df_sub["q"].str.replace(r"\W","").str.upper()
df_sub["a_string"] = df_sub["a"].str.replace(r"\W","").str.upper()
future = re.compile(r"ANTICIPAT(?:ED?|ING|ION)|EXPECT(?:ATION|ED)?|FORECAST(?:ED)?|GUIDANCE|OUTLOOK|(?:GOINGTO|WILL|SHOULD)BE|PROJECT(?:ED|ION)|REVISED|TRAJECTORY")
df_sub["future"] = np.where(df_sub["q_string"].str.contains(future, regex=True)|df_sub["a_string"].str.contains(future, regex=True), 1, 0)
df_sub = df_sub.drop(columns=["q_num","q", "q_string", "a", "a_string"])
|
qna_measures.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Imports and server setup
# + deletable=true editable=true
# parsing & plotting, later
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from collections import defaultdict
# %matplotlib inline
# + deletable=true editable=true
# for workflow management
import json
import os
from google.cloud import storage
import cromwell_manager as cwm
with open(os.path.expanduser('~/.ssh/mint_cromwell_config.json')) as f:
cromwell_server = cwm.Cromwell(**json.load(f))
storage_client = storage.Client(project='broad-dsde-mint-dev')
# -
# ## Define and run the un-optimized workflows
# + deletable=true editable=true
wdl = '../count/count.wdl'
inputs = 'pbmc8k_inputs.json'
options = 'https://raw.githubusercontent.com/ambrosejcarr/cromwell-manager/master/src/accessories/long_call_caching.json'
# + deletable=true editable=true
full_monitored = cwm.Workflow.from_submission(wdl=wdl, inputs_json=inputs, options_json=options, cromwell_server=cromwell_server, storage_client=storage_client)
full_monitored_from_get_trimmed_aligned = cwm.Workflow.from_submission(wdl=wdl, inputs_json=inputs, options_json=options, cromwell_server=cromwell_server, storage_client=storage_client)
# + deletable=true editable=true
full_monitored = cwm.Workflow('52b9df91-088d-44b0-b3a8-5c5a6b542268', cromwell_server=cromwell_server, storage_client=storage_client)
full_monitored_from_get_trimmed_aligned = cwm.Workflow('d89225c5-b804-4b13-a8fb-98a71557c800', cromwell_server=cromwell_server, storage_client=storage_client)
# -
# ## Parse the data; identify and save cost information
# + deletable=true editable=true
# here's what I did last time. This time I'll need to:
# 1. merge the two tasks -- shouldn't be too bad, the second is a superset of the first. Account for possibility of None's (also make sure to commit that change! if it works)
# 2. find the average of each task over all the shards when possible, otherwise use the max
max_memory = defaultdict(list)
available_memory = defaultdict(list)
max_disk = defaultdict(list)
available_disk = defaultdict(list)
def extract_info(task):
"""awful little side-effecting function to fill dictionaries"""
ru = task.resource_utilization
if ru is None: # allow for None
return
max_memory[ru.task_name].append(ru.max_memory)
max_disk[ru.task_name].append(ru.max_disk)
available_memory[ru.task_name].append(ru.total_memory)
available_disk[ru.task_name].append(ru.total_disk)
for task_name, calledtask1 in full_monitored_from_get_trimmed_aligned.tasks.items():
# get the other run's tasks
if task_name in full_monitored.tasks:
calledtask2 = full_monitored.tasks[task_name]
for shard in calledtask2._shards:
extract_info(shard)
for shard in calledtask1._shards:
extract_info(shard)
mean_max_memory = {key: np.mean(val) for key, val in max_memory.items()}
max_max_memory = {key: np.max(val) for key, val in max_memory.items()}
std_max_memory = {key: np.std(val) for key, val in max_memory.items()}
mean_max_disk = {key: np.mean(val) for key, val in max_disk.items()}
max_max_disk = {key: np.max(val) for key, val in max_disk.items()}
std_max_disk = {key: np.std(val) for key, val in max_disk.items()}
available_memory = {key: np.mean(val) for key, val in available_memory.items()}
available_disk = {key: np.mean(val) for key, val in available_disk.items()}
parsed_data = pd.DataFrame({
'mean_max_memory': mean_max_memory,
'max_max_memory': max_max_memory,
'std_max_memory': std_max_memory,
'available_memory': available_memory,
'mean_max_disk': mean_max_disk,
'max_max_disk': max_max_disk,
'std_max_disk': std_max_disk,
'available_disk': available_disk
})
# + deletable=true editable=true
parsed_data # memory utilization
# + deletable=true editable=true
# save the data to file
parsed_data = pd.read_csv('pbmc8k_mem_diskusage_data.csv', index_col=0)
parsed_data.to_csv('pbmc8k_mem_diskusage_data.csv')
# -
# ## Plot utilization vs requested resources
# + deletable=true editable=true
def barplot(dataframe, column, label, ax, *args, **kwargs):
ax.bar(left=np.arange(dataframe.shape[0]), height=dataframe[column], *args, **kwargs)
ax.set_xticks(np.arange(dataframe.shape[0]));
ax.set_xticklabels(list(i.replace('singlesample.Ss2RunSingleSample.', '') for i in dataframe.index), rotation=90)
ax.set_xlim((-0.6, dataframe.shape[0] - 0.4));
ax.set_ylabel(label)
# + deletable=true editable=true
f, ax = plt.subplots(figsize=(12, 5))
barplot(dataframe=parsed_data, column='available_memory', label='memory (MB)', zorder=0, ax=ax, facecolor='b')
barplot(dataframe=parsed_data, column='mean_max_memory', label='memory (MB)', zorder=1, ax=ax, facecolor='r',
yerr=parsed_data['std_max_memory'])
plt.scatter(np.arange(parsed_data.shape[0]), parsed_data['max_max_memory'], c='r')
plt.title('memory usage (red) vs. allocation (blue) by task');
# + deletable=true editable=true
f, ax = plt.subplots(figsize=(12, 5))
barplot(dataframe=parsed_data, column='available_disk', label='disk (KB)', zorder=0, ax=ax, facecolor='b')
barplot(dataframe=parsed_data, column='mean_max_disk', label='disk (KB)', zorder=1, ax=ax, facecolor='r',
yerr=parsed_data['std_max_disk'])
plt.scatter(np.arange(parsed_data.shape[0]), parsed_data['max_max_disk'], c='r')
plt.title('disk usage (red) vs. allocation (blue) by task');
# + [markdown] deletable=true editable=true
# ## Run the resource-optimized count on test data.
# -
# I'm running with call caching, and docker was doing weird things, so the cost won't be accurate.
# + deletable=true editable=true
wdl = '/Users/carra1/projects/skylab/10x/count/count.wdl'
pbmc8k_inputs = 'pbmc8k_inputs.json'
inputs_json = '/Users/carra1/projects/skylab/10x/count/example_count_input.json'
options = '/Users/carra1/projects/cromwell-manager/src/accessories/options.json'
monitor = {"monitoring_script": "gs://broad-dsde-mint-dev-teststorage/10x/benchmark/scripts/monitor.sh"}
# + deletable=true editable=true
test_monitored_small = cwm.Workflow.from_submission(wdl=wdl, inputs_json=inputs_json, cromwell_server=cromwell_server, storage_client=storage_client)
# + deletable=true editable=true
test_monitored_small.status
# + deletable=true editable=true
os.environ['wdltool'] = os.path.expanduser('~/google_drive/software/wdltool-0.14.jar')
test_monitored = cwm.Workflow.validate(
wdl=wdl,
inputs_json=pbmc8k_inputs,
options_json={"monitoring_script": "gs://broad-dsde-mint-dev-teststorage/10x/benchmark/scripts/monitor_long.sh"},
cromwell_server=cromwell_server,
storage_client=storage_client)
# + deletable=true editable=true
test_monitored = cwm.Workflow.from_submission(
wdl=wdl,
inputs_json=pbmc8k_inputs,
options_json={"monitoring_script": "gs://broad-dsde-mint-dev-teststorage/10x/benchmark/scripts/monitor_long.sh"},
cromwell_server=cromwell_server,
storage_client=storage_client)
# + deletable=true editable=true
test_monitored.status
# -
# Carry out the cost analysis even though docker took 100x longer than it should have.
# +
import requests
script_data = requests.get('https://raw.githubusercontent.com/ambrosejcarr/cromwell-manager/master/src/accessories/calculate_workflow_cost.py')
with open('get_cost.py', 'wb') as f:
f.write(script_data.content)
md = test_monitored.metadata
with open('run_metadata.json', 'w') as f:
json.dump(md, f)
# -
# !python2.7 get_cost.py -m run_metadata.json --only_total
|
benchmarking/10x_count/cost/scale_pbmc8k_with_monitoring.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import matplotlib.pyplot as plt
import pandas as pd
plt.style.use('seaborn')
URL='https://data.seattle.gov/api/views/mdbt-9ykn/rows.csv?accessType=DOWNLOAD'
def get_freemont_data(filename='Fremont.csv',url=URL,force_download=False):
if force_download or not os.path.exists(filename):
urlretrieve(url,filename)
data=pd.read_csv('Fremont.csv',index_col='Date',parse_dates=True)
data.columns=['West','East']
data['Total']=data['West']+data['East']
return data
# -
data=get_freemont_data()
data.head()
# %matplotlib inline
data.resample('W').sum().plot();
# +
ax=data.resample('D').sum().rolling(365).sum().plot();
ax.set_ylim(0,None);
# -
data.groupby(data.index.time).mean()
pivoted=data.pivot_table('Total',index=data.index.time,columns=data.index.date)
pivoted.iloc[:5,:5]
|
Freemont_bikeshare.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This is a demonstration code
from main import __main__ as main
parser = main(JSON_FOLDER=cfg.figmap)
# Demonstration
parser.css_sify({ '#btn1':{ 'background': 'red', 'border-radius': '1rem' } })
parser.__attr_list__()
|
demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CS 533 Project: TVCA - General and Bike traffic correlation
# by Ravishankar and <NAME>
#
# ## Purpose and Context
# The purpose of this notebook is to understand how does the bike traffic changes with the general traffic.
#
# ## Setup
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from simpledbf import Dbf5
import glob
bike_count_overall = pd.read_excel(r'clean-data/Gen_traf_analysis_data/traffic_master_count.xlsx')
bike_count_overall.head()
data = bike_count_overall.replace(0.0, np.nan)
df1 = data.sort_values('count', ascending=False)
df = df1.head(10)
df
achd = pd.read_csv('clean-data/Gen_traf_analysis_data/out.csv')
achd.head()
merge_df = pd.merge(achd,df,how='inner',left_on='Stname',right_on='Street_name')
merge_df
split1 = merge_df[['Stname','Traffic_count']]
split2_2 = merge_df[['Street_name','Total']]
split2 = split2_2.rename(columns={'Street_name' : 'Stname', 'Total' : 'Traffic_count'})
split2
split1['category']='general traffic'
split2['category']='Bike traffic'
split1
final_df=pd.concat([split1,split2])
final_df
# +
sns.set(rc={'figure.figsize':(13.7,13.27)})
w = sns.scatterplot(x='Stname',y='Traffic_count',hue='category',data =final_df)
#w.savefig(r'C:\Users\Admini\Desktop\TVCA2\TVCA\orig-data\fig1.png')
# -
# **Inferences:**
#
# - For Bannock street, the decrease in general traffic positively resulted in an increase in bike traffic.
# - The same trend is not seen for Fort or Latah street.Therefore we can conclude general traffic doesn't influence bike traffic.
# - Latah street doesn't have enough data points at this point of time compared to Bannock.
# - There are some hidden factors that influence bike traffic.For example, speed limit of a street, Directions.
#
# **Directions:**
#
# Let us see, how does the bike traffic change when directions are taken in to account.
#
bike_dir = pd.read_csv('clean-data/Gen_traf_analysis_data/directions.csv')
bike_dir.head()
bike_dir_sum = bike_dir.groupby('Location')['Total: Location 1','Total: Location 2'].sum().reset_index()
bike_dir_sum.head()
split_1 = bike_dir_sum[['Location','Total: Location 1']]
split_2 = bike_dir_sum[['Location','Total: Location 2']]
df1 = split_1.rename(columns={'Total: Location 1' : 'bike_traffic'})
df2 = split_2.rename(columns={'Total: Location 2' : 'bike_traffic'})
# +
#final_bike_dir.info()
# -
df1['category']='N-S_direction'
df2['category']='E-W_direction'
df1.head()
final_bike_dir=pd.concat([df1,df2])
dir_final_busy = pd.read_csv('clean-data/Gen_traf_analysis_data/dir_final_busy.csv')
# +
sns.set(rc={'figure.figsize':(16.7,13.27)})
sns.scatterplot(x="Location",y="bike_traffic",hue="category",data=dir_final_busy)
# -
# **Inference:**
#
# - The skewed bike traffic at 08/Bannock and 08/River suggests that *the streets are more important to bikers than intersections.*
# - The other intersection have almost equally distributed bike traffic in both the directions.
|
04- General_traffic correlation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <NAME>, <NAME>
# October 11 , 2016
# ## Demonstration of ComplexPlaneNP
# We import our module and set the attributes for the plane window using the constructor. With the addition of a `__repr__` method, we are able to view our plane via `print`. The use of pandas includes labels corresponding the the x-axis (row 0) and the y-axis (column 0).
import cplane_np as cp
testPlane = cp.ComplexPlaneNP(-5,5,11,-5,5,11)
print(testPlane)
# The method `set_f()` transforms the complexplane by applting the function `f` to every point. Let us transform the plane by squaring each point with the function myFunction, then print the result.
myFunction = lambda x:x**2
testPlane.set_f(myFunction)
print(testPlane)
# The `zoom` function changes the viewing window of the plane, either by changing the minimum and maximum of the axes, by changing the number of points per axis, or both. Then we `print` the plane to view it.
testPlane.zoom(-1,1,4,0,3,5)
print(testPlane)
# ## Julia
#
# create a function `julia(c, max=100)` that takes a complex parameter c and an optional positive integer max
# The function takes one complex parameter z as an input, and returns a positive integer n.
# The returned integer n should count how many times the complex parameter z can be transformed
# as $z = z^2$ + c before the resulting magnitude $|z|$ exceeds 2.
# If the number max is reached before the magnitude of z exceeds 2, the function should return the number 0.
# If the number z already has a magnitude larger than 2, the function should return 1.
# ### Testing the Julia function
# To test the julia function, we used a few simple cases, where |z| exceeds 2 after a small number of transformation. This can be done for several z values at once by applying the function generated by julia(0) to our complex plane.
someFunction = cp.julia(0)
testPlane.set_f(someFunction)
testPlane.zoom(-1,1,3,-1,1,3)
print(testPlane)
# As expected, the value for 0 (which will never have a magnitude greater than 2 with zero being added to it) is 0, since the magnitude does not exceed 2 even after 100 (the default max) tries. Our other values, for small real and imaginary parts of z, are also as we expected. This is also how we created our test functions: with values that were easy to work with, but that also covered each output case (0, 1, or $n>1$).
# ## Difference between CW5 and CW6
#
# The biggest difference is out introduction of panda and numpy, rather than vanilla python. This allowed us to simplify our algorithm for generating the plane by eliminating loops completely. The numpy function `linspace` generated the points for each axis, and the numpy function `meshgrid` matched each real component with every imaginary component, creating a 2D grid of complex numbers. The added vectorizing functionality also meant that we no longer needed two of our attributes, xtick and ytick. The `linspace` method has an argument (number of points in the interval) that accomplishes everything those attributes had done. The other change we made between CW5 and CW6 was the reorganization of our class. In response to peer critiques, we moved the creation of the plane to the `refresh()` method, which is then called in the `__init__`, `zoom`, and `set_f` methods. Lastly, we added the `__repr__` method so `print` could be called on our class objects, which is particularly useful in both testing and this demonstration.
|
cw06-cplane-np.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import functions as func
import re
import pandas as pd
import warnings
warnings.filterwarnings("ignore")
#Um diretorio que contém todos os datasets
path = 'data'
df_dict = func.readCSV_DATA(path)
train_df = df_dict['train.csv']
test_df = df_dict['test.csv']
# +
#question_title
#question_body
#answer
#question_title, question_body
#answer
# +
question_lst = []
answer_lst = []
for column in train_df.columns[11:]:
match = re.match('([a-z]*)_', column)
if match.group(1) == 'question':
question_lst.append(column)
else:
answer_lst.append(column)
# -
train_df = func.prepareData(train_df)
train_df
|
clusteringAnswer .ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mancinimassimiliano/DeepLearningLab/blob/master/Lab2/solution/convolutional_neural_networks_solution.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="kaht-FPA1Jvq" colab_type="text"
# # Introduction
#
# ## Lab2: Train a Convolutional Neural Network (CNN).
#
# In this Lab session we will learn how to train a CNN from scratch for classifying MNIST digits.
# + id="UvxtTYHlVfRK" colab_type="code" colab={}
import torch
import torchvision
from torchvision import transforms as T
import torch.nn.functional as F
# + [markdown] id="HYCvhGxKWyN7" colab_type="text"
# ### Define LeNet
#
# 
#
# Here we are going to define our first CNN which is **LeNet** in this case. To construct a LeNet we will be using some convolutional layers followed by some fully-connected layers. The convolutional layers can be simply defined using `torch.nn.Conv2d` module of `torch.nn` package. Details can be found [here](https://pytorch.org/docs/stable/nn.html#conv2d). Moreover, we will use pooling operation to reduce the size of convolutional feature maps. For this case we are going to use `torch.nn.functional.max_pool2d`. Details about maxpooling can be found [here](https://pytorch.org/docs/stable/nn.html#max-pool2d)
#
# Differently from our previous Lab, we will use a Rectified Linear Units (ReLU) as activation function with the help of `torch.nn.functional.relu`, replacing `torch.nn.Sigmoid`. Details about ReLU can be found [here](https://pytorch.org/docs/stable/nn.html#id26).
# + id="dMC_LDYdWkI7" colab_type="code" colab={}
class LeNet(torch.nn.Module):
def __init__(self):
super(LeNet, self).__init__()
# input channel = 1, output channels = 6, kernel size = 5
# input image size = (28, 28), image output size = (24, 24)
self.conv1 = torch.nn.Conv2d(in_channels=1, out_channels=6, kernel_size=(5, 5))
# input channel = 6, output channels = 16, kernel size = 5
# input image size = (12, 12), output image size = (8, 8)
self.conv2 = torch.nn.Conv2d(in_channels=6, out_channels=16, kernel_size=(5, 5))
# input dim = 4 * 4 * 16 ( H x W x C), output dim = 120
self.fc3 = torch.nn.Linear(in_features=4 * 4 * 16, out_features=120)
# input dim = 120, output dim = 84
self.fc4 = torch.nn.Linear(in_features=120, out_features=84)
# input dim = 84, output dim = 10
self.fc5 = torch.nn.Linear(in_features=84, out_features=10)
def forward(self, x):
x = self.conv1(x)
x = F.relu(x)
# Max Pooling with kernel size = 2
# output size = (12, 12)
x = F.max_pool2d(x, kernel_size=2)
x = self.conv2(x)
x = F.relu(x)
# Max Pooling with kernel size = 2
# output size = (4, 4)
x = F.max_pool2d(x, kernel_size=2)
# flatten the feature maps into a long vector
x = x.view(x.shape[0], -1)
x = self.fc3(x)
x = F.relu(x)
x = self.fc4(x)
x = F.relu(x)
x = self.fc5(x)
return x
# + [markdown] id="gChf6TvWonrV" colab_type="text"
# ### Define cost function
# + id="6j5UrBH3oek8" colab_type="code" colab={}
def get_cost_function():
cost_function = torch.nn.CrossEntropyLoss()
return cost_function
# + [markdown] id="U2TjXeVdorV9" colab_type="text"
# ### Define the optimizer
# + id="hBZN-WPboulR" colab_type="code" colab={}
def get_optimizer(net, lr, wd, momentum):
optimizer = torch.optim.SGD(net.parameters(), lr=lr, weight_decay=wd, momentum=momentum)
return optimizer
# + [markdown] id="wTkfrV64oxIL" colab_type="text"
# ### Train and test functions
# + id="t-sE5vFio0lf" colab_type="code" colab={}
def test(net, data_loader, cost_function, device='cuda:0'):
samples = 0.
cumulative_loss = 0.
cumulative_accuracy = 0.
net.eval() # Strictly needed if network contains layers which has different behaviours between train and test
with torch.no_grad():
for batch_idx, (inputs, targets) in enumerate(data_loader):
# Load data into GPU
inputs = inputs.to(device)
targets = targets.to(device)
# Forward pass
outputs = net(inputs)
# Apply the loss
loss = cost_function(outputs, targets)
# Better print something
samples+=inputs.shape[0]
cumulative_loss += loss.item() # Note: the .item() is needed to extract scalars from tensors
_, predicted = outputs.max(1)
cumulative_accuracy += predicted.eq(targets).sum().item()
return cumulative_loss/samples, cumulative_accuracy/samples*100
def train(net,data_loader,optimizer,cost_function, device='cuda:0'):
samples = 0.
cumulative_loss = 0.
cumulative_accuracy = 0.
net.train() # Strictly needed if network contains layers which has different behaviours between train and test
for batch_idx, (inputs, targets) in enumerate(data_loader):
# Load data into GPU
inputs = inputs.to(device)
targets = targets.to(device)
# Forward pass
outputs = net(inputs)
# Apply the loss
loss = cost_function(outputs,targets)
# Reset the optimizer
# Backward pass
loss.backward()
# Update parameters
optimizer.step()
optimizer.zero_grad()
# Better print something, no?
samples+=inputs.shape[0]
cumulative_loss += loss.item()
_, predicted = outputs.max(1)
cumulative_accuracy += predicted.eq(targets).sum().item()
return cumulative_loss/samples, cumulative_accuracy/samples*100
# + [markdown] id="T6IT0Lsgo8AM" colab_type="text"
# ### Define the function that fetches a data loader that is then used during iterative training.
#
# We will learn a new thing in this function as how to Normalize the inputs given to the network.
#
# ***Why Normalization is needed***?
#
# To have nice and stable training of the network it is recommended to normalize the network inputs between \[-1, 1\].
#
# ***How it can be done***?
#
# This can be simply done using `torchvision.transforms.Normalize()` transform. Details can be found [here](https://pytorch.org/docs/stable/torchvision/transforms.html#torchvision.transforms.Normalize).
# + id="qDxpo6uVo_8k" colab_type="code" colab={}
def get_data(batch_size, test_batch_size=256):
# Prepare data transformations and then combine them sequentially
transform = list()
transform.append(T.ToTensor()) # converts Numpy to Pytorch Tensor
transform.append(T.Normalize(mean=[0.5], std=[0.5])) # Normalizes the Tensors between [-1, 1]
transform = T.Compose(transform) # Composes the above transformations into one.
# Load data
full_training_data = torchvision.datasets.MNIST('./data', train=True, transform=transform, download=True)
test_data = torchvision.datasets.MNIST('./data', train=False, transform=transform, download=True)
# Create train and validation splits
num_samples = len(full_training_data)
training_samples = int(num_samples*0.5+1)
validation_samples = num_samples - training_samples
training_data, validation_data = torch.utils.data.random_split(full_training_data, [training_samples, validation_samples])
# Initialize dataloaders
train_loader = torch.utils.data.DataLoader(training_data, batch_size, shuffle=True)
val_loader = torch.utils.data.DataLoader(validation_data, test_batch_size, shuffle=False)
test_loader = torch.utils.data.DataLoader(test_data, test_batch_size, shuffle=False)
return train_loader, val_loader, test_loader
# + [markdown] id="OHcB8f0AsY4n" colab_type="text"
# ### Wrapping everything up
#
# Finally, we need a main function which initializes everything + the needed hyperparameters and loops over multiple epochs (printing the results).
# + id="ip_R-hruse0Q" colab_type="code" colab={}
'''
Input arguments
batch_size: Size of a mini-batch
device: GPU where you want to train your network
weight_decay: Weight decay co-efficient for regularization of weights
momentum: Momentum for SGD optimizer
epochs: Number of epochs for training the network
'''
def main(batch_size=128,
device='cuda:0',
learning_rate=0.01,
weight_decay=0.000001,
momentum=0.9,
epochs=25):
train_loader, val_loader, test_loader = get_data(batch_size)
net = LeNet().to(device)
optimizer = get_optimizer(net, learning_rate, weight_decay, momentum)
cost_function = get_cost_function()
print('Before training:')
train_loss, train_accuracy = test(net, train_loader, cost_function)
val_loss, val_accuracy = test(net, val_loader, cost_function)
test_loss, test_accuracy = test(net, test_loader, cost_function)
print('\t Training loss {:.5f}, Training accuracy {:.2f}'.format(train_loss, train_accuracy))
print('\t Validation loss {:.5f}, Validation accuracy {:.2f}'.format(val_loss, val_accuracy))
print('\t Test loss {:.5f}, Test accuracy {:.2f}'.format(test_loss, test_accuracy))
print('-----------------------------------------------------')
for e in range(epochs):
train_loss, train_accuracy = train(net, train_loader, optimizer, cost_function)
val_loss, val_accuracy = test(net, val_loader, cost_function)
print('Epoch: {:d}'.format(e+1))
print('\t Training loss {:.5f}, Training accuracy {:.2f}'.format(train_loss, train_accuracy))
print('\t Validation loss {:.5f}, Validation accuracy {:.2f}'.format(val_loss, val_accuracy))
print('-----------------------------------------------------')
print('After training:')
train_loss, train_accuracy = test(net, train_loader, cost_function)
val_loss, val_accuracy = test(net, val_loader, cost_function)
test_loss, test_accuracy = test(net, test_loader, cost_function)
print('\t Training loss {:.5f}, Training accuracy {:.2f}'.format(train_loss, train_accuracy))
print('\t Validation loss {:.5f}, Validation accuracy {:.2f}'.format(val_loss, val_accuracy))
print('\t Test loss {:.5f}, Test accuracy {:.2f}'.format(test_loss, test_accuracy))
print('-----------------------------------------------------')
# + [markdown] id="ltdCMiB3t18h" colab_type="text"
# Lets train!
# + id="6d-z20H4tziL" colab_type="code" outputId="d0590fdc-52b5-4612-f5d1-c26bbece67cc" colab={"base_uri": "https://localhost:8080/", "height": 2699}
main()
|
Lab2/solution/convolutional_neural_networks_solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/shailymishra/Paper-Presentation-Summary-Implementation/blob/main/RegretNet_in_pytorch.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="AZGkMNO5XFZn" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="43c3637f-426e-49e2-9cf4-8c0ccb7f7308"
## Imports
import torch
import os
import numpy as np
import pandas as pd
import pandas.util.testing as tm
from tqdm import tqdm
import seaborn as sns
from pylab import rcParams
import matplotlib.pyplot as plt
from matplotlib import rc
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, classification_report
from torch import nn, optim
import torch.nn.functional as F
# + id="OqNlvKVyX7Dg"
## Config
import os
import os.path as osp
import numpy as np
# `pip install easydict` if you don't have it
from easydict import EasyDict as edict
__C = edict()
cfg = __C
# Output-dir to write log-files and save model
__C.dir_name = os.path.join("experiments", "additive_1x2_uniform")
# Auction params
__C.num_agents = 1
__C.num_items = 2
__C.distribution_type = "uniform"
__C.agent_type = "additive"
# Save data for restore.
__C.save_data = False
# Neural Net parameters
__C.net = edict()
# initialization g - glorot, h - he + u - uniform, n - normal [gu, gn, hu, hn]
__C.net.init = "gu"
# activations ["tanh", "sigmoid", "relu"]
# num_a_layers, num_p_layers - total number of hidden_layers + output_layer, [a - alloc, p - pay]
# num_p_hidden_units, num_p_hidden_units - number of hidden units, [a - alloc, p - pay]
__C.net.num_a_layers = 3
__C.net.num_a_activation = ["tanh", "tanh", "softmax"]
__C.net.num_a_units = [__C.num_items*__C.num_agents, 100,100,__C.num_items*__C.num_agents]
__C.net.num_p_layers = 3
__C.net.num_p_activation = ["tanh", "tanh", "sigmoid"]
__C.net.num_p_units = [__C.num_items*__C.num_agents ,100,100, __C.num_agents]
# Train paramters
__C.train = edict()
# Random seed
__C.train.seed = 42
# Iter from which training begins. If restore_iter = 0 for default. restore_iter > 0 for starting
# training form restore_iter [needs saved model]
__C.train.restore_iter = 0
# max iters to train
# __C.train.max_iter = 400000 ##CHANGED
__C.train.max_iter = 100
# Learning rate of network param updates
__C.train.learning_rate = 1e-3
# Regularization
__C.train.wd = None
""" Train-data params """
# Choose between fixed and online. If online, set adv_reuse to False
__C.train.data = "fixed"
# Number of batches
# __C.train.num_batches = 5000 ##CHANGED
__C.train.num_batches = 1
# Train batch size
# __C.train.batch_size = 128 ##CHANGED
__C.train.batch_size = 5
""" Train-misreport params """
# Cache-misreports after misreport optimization
__C.train.adv_reuse = True
# Number of misreport initialization for training
__C.train.num_misreports = 1
# Number of steps for misreport computation
__C.train.gd_iter = 25
# Learning rate of misreport computation
__C.train.gd_lr = 0.1
""" Lagrange Optimization params """
# Initial update rate
__C.train.update_rate = 1.0
# Initial Lagrange weights
__C.train.w_rgt_init_val = 5.0
# Lagrange update frequency
# __C.train.update_frequency = 100 ##CHANGED
__C.train.update_frequency = 10
# Value by which update rate is incremented
__C.train.up_op_add = 50.0
# Frequency at which update rate is incremented
__C.train.up_op_frequency = 10000
""" train summary and save params"""
# Number of models to store on disk
__C.train.max_to_keep = 25
# Frequency at which models are saved-
__C.train.save_iter = 20000
# Train stats print frequency
# __C.train.print_iter = 1000 ##changed
__C.train.print_iter = 10
""" Validation params """
__C.val = edict()
# Number of steps for misreport computation
# __C.val.gd_iter = 2000 ##changed
__C.val.gd_iter = 20
# Learning rate for misreport computation
__C.val.gd_lr = 0.1
# Number of validation batches
__C.val.num_batches = 20
# Frequency at which validation is performed
# __C.val.print_iter = 10000 ##changed
__C.val.print_iter = 10
# Validation data frequency
__C.val.data = "fixed"
""" Test params """
# Test set
__C.test = edict()
# Test Seed
__C.test.seed = 100
# Model to be evaluated
__C.test.restore_iter = 400000
# Number of misreports
__C.test.num_misreports = 1000
# Number of steps for misreport computation
__C.test.gd_iter = 2000
# Learning rate for misreport computation
__C.test.gd_lr = 0.1
# Test data
__C.test.data = "online"
# Number of test batches
__C.test.num_batches = 100
# Test batch size
__C.test.batch_size = 100
# Save Ouput
__C.test.save_output = False
# Fixed Val params
__C.val.batch_size = __C.train.batch_size
__C.val.num_misreports = __C.train.num_misreports
# Compute number of samples
__C.train.num_instances = __C.train.num_batches * __C.train.batch_size
__C.val.num_instances = __C.val.num_batches * __C.val.batch_size
__C.test.num_instances = __C.test.num_batches * __C.test.batch_size
# + id="HOpfDfJnXvY_"
## Create Data
## data n_samples x n_agents x n_items
## n_samples x 1 x 2
def generate_random_X(shape):
return np.random.rand(*shape)
def generate_random_ADV(shape):
return np.random.rand(*shape)
def preprocessdata(data):
return torch.squeeze(data,1)
# + id="WfxLMceXXx5Z"
## Create Net
class RegretNet(nn.Module):
def __init__(self, config):
super(RegretNet, self).__init__()
self.num_items = config.num_items
self.num_agents = config.num_agents
self.num_a_units = config.net.num_a_units
self.num_a_layers = config.net.num_a_layers
self.num_a_activation = config.net.num_a_activation
self.num_p_units = config.net.num_p_units
self.num_p_layers = config.net.num_p_layers
self.num_p_activation = config.net.num_p_activation
self.num_misreports = config.train.num_misreports
# self.u_shape = [self.num_agents, config.train.num_misreports, config.train.batch_size, self.num_agents]
self.relu = nn.ReLU()
self.update_rate = config.train.update_rate
self.w_rgt_init_val = config.train.w_rgt_init_val
self.w_rgt = np.ones(self.num_agents).astype(np.float32) * self.w_rgt_init_val
self.w_rgt = torch.tensor(self.w_rgt)
print(' w_rgt ', self.w_rgt)
self.activation = {'sigmoid': nn.Sigmoid() , 'tanh' : nn.Tanh() , 'softmax' : nn.Softmax() }
## Layers
self.allocationNetwork = nn.ModuleList([ nn.Linear( self.num_a_units[i] , self.num_a_units[i+1] ) for i in range(self.num_a_layers)])
self.paymentNetwork = nn.ModuleList([ nn.Linear( self.num_p_units[i] , self.num_p_units[i+1] ) for i in range(self.num_p_layers)])
for i in range(self.num_a_layers) :
torch.nn.init.xavier_uniform(self.allocationNetwork[i].weight)
self.allocationNetwork[i].bias.data.fill_(0.00)
for i in range(self.num_p_layers) :
torch.nn.init.xavier_uniform(self.paymentNetwork[i].weight)
self.paymentNetwork[i].bias.data.fill_(0.00)
def forward(self, x):
allocation = x
payment = x
for i in range(self.num_a_layers):
allocation = self.allocationNetwork[i](allocation)
allocation = self.activation[self.num_a_activation[i]](allocation)
for i in range(self.num_p_layers):
payment = self.paymentNetwork[i](payment)
payment = self.activation[self.num_p_activation[i]](payment)
n_samples = allocation.shape[0]
allocXval = torch.reshape(allocation * x, (n_samples, self.num_agents ,self.num_items ))
payment = payment * torch.sum( allocXval, dim=2) ## summing for each agent, over all items
return allocation , payment
def compute_rev(self, payment):
return torch.mean(torch.sum(payment, dim=1))
def compute_utility(self, x, allocation, payment):
n_samples = x.shape[0]
# x = x.reshape(n_samples, self.num_agents , self.num_items)
allocXval = torch.reshape(allocation * x, (n_samples, self.num_agents , self.num_items ))
utility = torch.sum( allocXval, dim=2) - payment
return utility
def compute_regret(self,x, misreports, utility_true):
n_samples = misreports.shape[1]
misreports_allocation = []
misreports_payments = []
for i in range(self.num_misreports):
a , p = net.forward(preprocessdata(misreports[i]))
misreports_allocation.append(a)
misreports_payments.append(p)
misreports_allocation = torch.stack(misreports_allocation)
misreports_payments = torch.stack(misreports_payments)
misreports_utility = [ net.compute_utility(x, misreports_allocation[i], misreports_payments[i] ) for i in range(self.num_misreports)]
misreports_utility = torch.stack(misreports_utility)
difference = self.relu(misreports_utility - utility_true)
maxdifference , indices= torch.max(difference , dim=0)
regret = torch.mean(maxdifference, dim=1)
return regret , misreports_utility
def loss_function(self, allocation, payment , train_data , train_misreports_data ):
revenue = self.compute_rev(payment)
utility = self.compute_utility(train_data, allocation, payment)
regret , misreports_utility = self.compute_regret(train_data, train_misreports_data , utility)
print(' revenue is ', revenue)
rgt_mean = torch.mean(regret)
irp_mean = torch.mean(self.relu(-utility))
rgt_penalty = self.update_rate * torch.sum(torch.square(regret)) / 2.0
lag_loss = torch.sum(self.w_rgt * regret)
loss_1 = -revenue + rgt_penalty + lag_loss
loss_2 = -torch.sum(misreports_utility)
loss_3 = -lag_loss
# reg_losses = tf.get_collection('reg_losses')
# if len(reg_losses) > 0:
# reg_loss_mean = tf.reduce_mean(reg_losses)
# loss_1 = loss_1 + reg_loss_mean
self.metrics = [revenue, rgt_mean, rgt_penalty, lag_loss, loss_1, torch.mean(self.w_rgt), self.update_rate]
self.metric_names = ["Revenue", "Regret", "Reg_Loss", "Lag_Loss", "Net_Loss", "w_rgt_mean", "update_rate"]
return loss_1 , loss_2 , loss_3
# + id="TTnqTxwoXy7f" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="ee4fac21-b385-4fb9-c5e0-da1c85fcfded"
## Create train , val , test
np.random.seed(cfg.train.seed)
net = RegretNet(cfg)
train_data_shape = [cfg.train.num_instances , cfg.num_agents, cfg.num_items]
train_adv_shape = [cfg.train.num_misreports, cfg.train.num_instances, cfg.num_agents, cfg.num_items]
train_data = generate_random_X(train_data_shape)
train_data = torch.from_numpy(train_data).float()
train_data = preprocessdata(train_data)
train_misreports_data = generate_random_ADV(train_adv_shape)
train_misreports_data = torch.from_numpy(train_misreports_data).float()
# temp = [train_misreports_data]
temp = [torch.tensor(train_misreports_data,requires_grad=True )]
optimizer_1 = optim.Adam(net.parameters(), cfg.train.learning_rate)
optimizer_2 = optim.Adam(temp, lr = cfg.train.gd_lr )
allocation , payment = net(train_data)
loss1, loss2,loss3 = net.loss_function(allocation, payment , train_data, train_misreports_data)
print(' Update for Misreports')
for _ in range(cfg.train.gd_iter):
print(_)
optimizer_2.zero_grad()
loss1, loss2,loss3 = net.loss_function(allocation, payment , train_data, temp[0])
loss2.backward()
optimizer_2.step()
temp[0].data.clamp_(0,1)
print()
print(temp)
print()
print('___________________________________________________')
for i in range(len(net.metric_names)):
print( net.metric_names[i] , ' : ' , net.metrics[i] )
print('___________________________________________________')
# optimizer_1.zero_grad()
# loss1.backward()
# optimizer_1.step()
# allocation , payment = net(train_data)
# optimizer.zero_grad()
# output = model(input)
# loss = loss_fn(output, target)
# loss.backward()
# optimizer.step()
# loss1, loss2,loss3 = net.loss_function(allocation, payment , train_data, train_misreports_data)
# optimizer_1.zero_grad()
# loss1.backward()
# r = optimizer_1.step()
# print(' loss 1 ', loss1 , ' loss 2 ', loss2 , ' loss 3 ', loss3)
# print('________________________')
# print(train_misreports_data)
# print('________________________')
# Optimizer
# print('___________________________________________________')
# for i in range(len(net.metric_names)):
# print( net.metric_names[i] , ' : ' , net.metrics[i] )
# print('___________________________________________________')
# revenue = net.compute_rev(payment)
# utility = net.compute_utility(train_data, allocation, payment)
# regret = net.compute_regret(train_misreports_data , utility)
### update loss and check if that is also fine
### then for on batches
### then save model , and
# + id="LfMG8ECDcngm"
n_samples = 2
n_items = 2
n_agents = 3
allocation = torch.tensor([[0.4 , 0.2 , 0.5 , 0.6, 0.7,0.4 ], [0.1 , 0.3 , 0.7 , 0.2,0.1,0.2 ] ])
valuation = torch.tensor([[0.1 , 0.2 , 0.5 , 0.3 , 0.8,0.6], [0.54 , 0.2 , 0.7 , 0.8, 0.8,0.9 ] ])
payment = torch.tensor([[0.13 , 0.33, 0.6], [0.76, 0.65, 0.6]])
allocXval = torch.reshape(allocation * valuation, (n_samples, n_agents , n_items ))
# payment = payment * torch.sum( allocXval, dim=2)
utility = torch.sum( allocXval, dim=2) - payment
print('________________________________________________________________')
print(utility)
print('________________________________________________________________')
v_misreports = torch.tensor([ [[0.3 , 0.24 , 0.25 , 0.38 , 0.38,0.16], [0.5 , 0.12 , 0.47 , 0.28, 0.81,0.5 ] ],
[[0.5 , 0.25 , 0.25 , 0.73 , 0.877,0.61], [0.4 , 0.22 , 0.37 , 0.38, 0.5,0.3 ] ]
])
a_misreports = torch.tensor([ [[0.3 , 0.24 , 0.25 , 0.38 , 0.38,0.16], [0.5 , 0.12 , 0.47 , 0.28, 0.81,0.5 ] ],
[[0.5 , 0.25 , 0.25 , 0.73 , 0.877,0.61], [0.4 , 0.22 , 0.37 , 0.38, 0.5,0.3 ] ]
])
p_misreports = torch.tensor([ [[0.7 , 0.7 , 0.7 ], [0.5 , 0.12 , 0.47 , ] ],
[[0.5 , 0.25 , 0.25], [0.4 , 0.22 , 0.37 ] ]
])
n_misreports = 2
allocXvalu_misreports = torch.reshape(a_misreports * v_misreports, (n_misreports, n_samples, n_agents , n_items ))
u_misreports = torch.sum( allocXvalu_misreports, dim=3) - p_misreports
print('________________________________________________________________')
print(u_misreports)
print('________________________________________________________________')
relu = nn.ReLU()
difference = relu(u_misreports - utility)
print('________________________________________________________________')
print(difference)
print('________________________________________________________________')
maxdiff , indices= torch.max(difference , dim=0)
print('________________________________________________________________')
print(maxdiff)
print('________________________________________________________________')
regret = torch.mean(maxdiff, dim=1)
print('________________________________________________________________')
print(regret)
print('________________________________________________________________')
# utility2 = [ net.compute_utility(v_misreports[i], a_misreports[i], p_misreports[i] ) for i in range(n_misreports)]
# print('________________________________________________________________')
# utility2 = torch.stack(utility2)
# print(utility2)
# print('________________________________________________________________')
# print(misreports.shape)
# print('payment')
# print(payment)
# print('__________________________________________')
# print(' Revenue ', torch.mean(torch.sum(payment, dim=1)))
# print('__________________________________________')
# utility = torch.sum( allocXval, dim=2) - payment
# print('__________________________________________')
# print(utility)
# print('__________________________________________')
# print('allocation')
# print(allocation)
# print('__________________________________________')
# print('valuation')
# print(valuation)
# print('__________________________________________')
# print('__________________________________________')
# print(torch.sum(allocXval, dim=2))
# print('__________________________________________')
# + id="8TjGgt0Aeoko"
# + id="nYrNpA2ADLdv"
|
Paper Implementations/Optimal Auctions RegretNet/RegretNet_in_pytorch.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df=pd.read_csv("common.csv")
df.head()
g=df.groupby(['file_id'])
remove_index_list=[]
for name,sub in g:
num_classes=sub['class'].value_counts().shape[0]
if num_classes!=2:
remove_index_list.extend(sub.index.values)
print(remove_index_list)
print(df.shape)
df2=df.drop(index=remove_index_list)
print(df2.shape)
# +
def find_conjunction_images_from_df(df,id_list,object_pattern):
"""Used to remove unqualified entries in an AND search."""
remove_index_list=[]
grouped_df=df.groupby(['file_id'])
for each_file_id,grouped_set in grouped_df:
if object_pattern=='class':
num_unique=grouped_set['class'].value_counts().shape[0]
else:
num_unique=grouped_set['object'].value_counts().shape[0]
if num_unique!=len(id_list):
remove_index_list.extend(grouped_set.index.values)
df=df.drop(index=remove_index_list)
return df
|
semlog_mongo/example/Find common.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from PIL import Image
# adjust pixel of image to avoid out of bound [0,255]
def adjust_pixel(pixel):
if pixel < 0: return 0
elif pixel > 255: return 255
return pixel
def changeBrightness():
img = Image.open("lena.png") # import image
pixels = img.load() # load pixel of image
img_convert = Image.new(img.mode, img.size) # mode, size, color(default: 0)
pixels_convert = img_convert.load() # load pixel of new image
for i in range(img_convert.size[0]):
for j in range(img_convert.size[1]):
R, G, B = pixels[i, j] # get R,G,B value
# take R,G,B of old image plus for a value brightness and adjust new value to avoid out of bound
# increase value 50 to get image with more brighter, this values usually in range [-255,255]
_R, _G, _B = adjust_pixel(R + 50), adjust_pixel(G + 50), adjust_pixel(B + 50)
pixels_convert[i, j] = (_R, _G, _B, 255) # update new pixel of image
img_convert.save("lena_brighter.png") # save the output image
changeBrightness()
# -
img1 = mpimg.imread("lena.png")
plt.imshow(img1)
plt.title("Original lena image", fontsize=18, fontweight='bold')
img2 = mpimg.imread("lena_brighter.png")
plt.imshow(img2)
plt.title("Lena image with more brighter", fontsize=18, fontweight='bold')
# +
def changeContrast():
img = Image.open("lena.png") # import image
pixels = img.load() # load pixel of image
img_convert = Image.new(img.mode, img.size) # mode, size, color(default: 0)
pixels_convert = img_convert.load() # load pixel of new image
for i in range(img_convert.size[0]):
for j in range(img_convert.size[1]):
contrast_value = 100 # value to change contrast, it's usually in range [-255,255]
# formula factor: F = (259(C + 255)) / (255(259 - C))
factor = (259 * (contrast_value + 255)) / (255 * (259 - contrast_value))
R, G, B = pixels[i, j] # get R,G,B value
# formula adjust R,G,B: F(R - 128) + 128
_R = adjust_pixel(factor * (R - 128) + 128)
_G = adjust_pixel(factor * (G - 128) + 128)
_B = adjust_pixel(factor * (B - 128) + 128)
pixels_convert[i, j] = (int(_R), int(_G), int(_B), 255) # update new pixel of image, this must be int not float
img_convert.save("lena_more_contrast.png") # save the output image
changeContrast()
# -
img3 = mpimg.imread("lena_more_contrast.png")
plt.imshow(img3)
plt.title("Lena with more contrast", fontsize=18, fontweight='bold')
# +
def changeGrey():
img = Image.open("lena.png") # import image
pixels = img.load() # load pixel of image
img_convert = Image.new(img.mode, img.size) # mode, size, color(default: 0)
pixels_convert = img_convert.load() # load pixel of new image
for i in range(img_convert.size[0]):
for j in range(img_convert.size[1]):
R, G, B = pixels[i, j] # get R,G,B value
# grey image: R,G,B have same value, so get average of 3 sub-pixels and create a new pixel
# we also have another formula for better grayscale: x = (0.299*R + 0.587*G + 0.114*B)
sub_pixel_avg = int(round((R + G + B) / 3))
pixels_convert[i, j] = (sub_pixel_avg, sub_pixel_avg,sub_pixel_avg, 0) # update new pixel of image
img_convert.save("lena_grey.png") # save the output image
changeGrey()
# +
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
img4 = mpimg.imread("lena_grey.png")
plt.imshow(img4)
plt.title("Lena grayscale", fontsize=18, fontweight='bold')
# +
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
img5 = mpimg.imread('lena.png') # import image
# example our matrix of image is: [[1,2,3], [4,5,6], [7,8,9]]
# fliplr will make matrix become: [[7,8,9], [4,5,6], [1,2,3]]
img6 = np.fliplr(img5)
plt.imshow(img6)
plt.title("Mirror of lena", fontsize=18, fontweight='bold')
# +
from PIL import Image
# import image and convert to RGBA to make sure image got an alpha channel
theme = Image.open("red_square.png").convert("RGBA")
overlay = Image.open("yellow_star.png").convert("RGBA")
# blend the image
img13 = Image.blend(theme, overlay, 0.5) # alpha = 0.5
img13.save("flag_vn_blend.png", "PNG")
# -
img15 = mpimg.imread("red_square.png")
plt.imshow(img15)
plt.title("Red square", fontsize=18, fontweight='bold')
img16 = mpimg.imread("yellow_star.png")
plt.imshow(img16)
plt.title("Yellow star", fontsize=18, fontweight='bold')
# +
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
def convertToGrayscale(rgb):
# follow the formula: Y' = 0.2989*R + 0.5870*G + 0.1140*B
return np.dot(rgb[...,:3], [0.2989, 0.5870, 0.1140])
img14 = mpimg.imread('flag_vn_blend.png')
grayscale = convertToGrayscale(img14)
plt.imshow(gray, cmap=plt.get_cmap('gray'), vmin=0, vmax=1)
plt.title("Image after overlaying", fontsize=18, fontweight='bold')
plt.show()
# +
from PIL import Image, ImageDraw
img15 = Image.open("lena.png") # import image
pixels = img15.load() # load pixel of image
# box blur kernel 3x3: 1/9 * [[1,1,1],[1,1,1],[1,1,1]]
box_blur = [[1/9, 1/9, 1/9],
[1/9, 1/9, 1/9],
[1/9, 1/9, 1/9]]
centerKernel = len(box_blur) // 2 # get the center of kernel, centerKernel = 3//2 = 1
img16 = Image.new("RGB", img15.size) # output image
# calculate convolution
# for each pixel in image row (image row in input image), i = [1,218]
# img15.width = img15.height = 220 (default resolution of image)
for i in range(centerKernel, img15.width - centerKernel):
# for each pixel in image column (image column in input image), j = [1,218]
for j in range(centerKernel, img15.height - centerKernel):
accumulator = [0, 0, 0] # set accumulator to zero
# for each element in kernel row (kernel row in kernel), [0,0,0,1,1,1,2,2,2,...]
for element1 in range(len(box_blur)):
# for each element in kernel column (kernel column in kernel), [0,1,2,0,1,2,0,1,2,...]
for element2 in range(len(box_blur)):
if centerKernel < 0: centerKernel = 0
# if element position corresponding to pixel position
pixel_position = pixels[i + element1 - centerKernel, j + element2 - centerKernel]
# multiply element value corresponding to pixel value and add result to accumulator
accumulator[0] += pixel_position[0] * box_blur[element1][element2]
accumulator[1] += pixel_position[1] * box_blur[element1][element2]
accumulator[2] += pixel_position[2] * box_blur[element1][element2]
# set output image pixel to accumulator
draw = ImageDraw.Draw(img16)
draw.point((i, j), (int(accumulator[0]), int(accumulator[1]), int(accumulator[2])))
img16.save("lena_blur.png") # save the image
# +
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
img17 = mpimg.imread("lena_blur.png")
plt.imshow(img17)
plt.title("Lena blur", fontsize=18, fontweight='bold')
# -
|
Lab02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] Collapsed="false"
# # Twins 数据集上因果推断
#
# DoWhy example on Twins dataset
# + [markdown] Collapsed="false"
# Here we study the twins dataset as studied by <a href="https://arxiv.org/pdf/1705.08821.pdf" target="_blank">Louizos et al</a>. We focus on twins which are the same sex and weigh less than 2kgs. The treatment t = 1 is being born the heavier twin and the outcome is mortality of each of the twins in their first year of life.The confounding variable taken is 'gestat10', the number of gestational weeks prior to birth, as it is highly correlated with the outcome. The results using the methods below are in coherence with those obtained in the paper.
# + Collapsed="false"
import os, sys
sys.path.append(os.path.abspath("../../../"))
import pandas as pd
import numpy as np
import dowhy
from dowhy import CausalModel
from dowhy import causal_estimators
# + [markdown] Collapsed="false"
# **<font size="4">Load the Data</font>**
# + [markdown] Collapsed="false"
# The data loading process involves combining the covariates, treatment and outcome, and resolving the pair property in the data. Since there are entries for both the twins, their mortalities can be treated as two potential outcomes. The treatment is given in terms of weights of the twins.Therefore, to get a binary treatment, each child's information is added in a separate row instead of both's information being condensed in a single row as in the original data source.
# + Collapsed="false"
#The covariates data has 46 features
x = pd.read_csv("https://raw.githubusercontent.com/AMLab-Amsterdam/CEVAE/master/datasets/TWINS/twin_pairs_X_3years_samesex.csv")
#The outcome data contains mortality of the lighter and heavier twin
y = pd.read_csv("https://raw.githubusercontent.com/AMLab-Amsterdam/CEVAE/master/datasets/TWINS/twin_pairs_Y_3years_samesex.csv")
#The treatment data contains weight in grams of both the twins
t = pd.read_csv("https://raw.githubusercontent.com/AMLab-Amsterdam/CEVAE/master/datasets/TWINS/twin_pairs_T_3years_samesex.csv")
# + Collapsed="false"
#_0 denotes features specific to the lighter twin and _1 denotes features specific to the heavier twin
lighter_columns = ['pldel', 'birattnd', 'brstate', 'stoccfipb', 'mager8',
'ormoth', 'mrace', 'meduc6', 'dmar', 'mplbir', 'mpre5', 'adequacy',
'orfath', 'frace', 'birmon', 'gestat10', 'csex', 'anemia', 'cardiac',
'lung', 'diabetes', 'herpes', 'hydra', 'hemo', 'chyper', 'phyper',
'eclamp', 'incervix', 'pre4000', 'preterm', 'renal', 'rh', 'uterine',
'othermr', 'tobacco', 'alcohol', 'cigar6', 'drink5', 'crace',
'data_year', 'nprevistq', 'dfageq', 'feduc6', 'infant_id_0',
'dlivord_min', 'dtotord_min', 'bord_0',
'brstate_reg', 'stoccfipb_reg', 'mplbir_reg']
heavier_columns = [ 'pldel', 'birattnd', 'brstate', 'stoccfipb', 'mager8',
'ormoth', 'mrace', 'meduc6', 'dmar', 'mplbir', 'mpre5', 'adequacy',
'orfath', 'frace', 'birmon', 'gestat10', 'csex', 'anemia', 'cardiac',
'lung', 'diabetes', 'herpes', 'hydra', 'hemo', 'chyper', 'phyper',
'eclamp', 'incervix', 'pre4000', 'preterm', 'renal', 'rh', 'uterine',
'othermr', 'tobacco', 'alcohol', 'cigar6', 'drink5', 'crace',
'data_year', 'nprevistq', 'dfageq', 'feduc6',
'infant_id_1', 'dlivord_min', 'dtotord_min', 'bord_1',
'brstate_reg', 'stoccfipb_reg', 'mplbir_reg']
# + Collapsed="false"
#Since data has pair property,processing the data to get separate row for each twin so that each child can be treated as an instance
data = []
for i in range(len(t.values)):
#select only if both <=2kg
if t.iloc[i].values[1]>=2000 or t.iloc[i].values[2]>=2000:
continue
this_instance_lighter = list(x.iloc[i][lighter_columns].values)
this_instance_heavier = list(x.iloc[i][heavier_columns].values)
#adding weight
this_instance_lighter.append(t.iloc[i].values[1])
this_instance_heavier.append(t.iloc[i].values[2])
#adding treatment, is_heavier
this_instance_lighter.append(0)
this_instance_heavier.append(1)
#adding the outcome
this_instance_lighter.append(y.iloc[i].values[1])
this_instance_heavier.append(y.iloc[i].values[2])
data.append(this_instance_lighter)
data.append(this_instance_heavier)
# + Collapsed="false"
cols = [ 'pldel', 'birattnd', 'brstate', 'stoccfipb', 'mager8',
'ormoth', 'mrace', 'meduc6', 'dmar', 'mplbir', 'mpre5', 'adequacy',
'orfath', 'frace', 'birmon', 'gestat10', 'csex', 'anemia', 'cardiac',
'lung', 'diabetes', 'herpes', 'hydra', 'hemo', 'chyper', 'phyper',
'eclamp', 'incervix', 'pre4000', 'preterm', 'renal', 'rh', 'uterine',
'othermr', 'tobacco', 'alcohol', 'cigar6', 'drink5', 'crace',
'data_year', 'nprevistq', 'dfageq', 'feduc6',
'infant_id', 'dlivord_min', 'dtotord_min', 'bord',
'brstate_reg', 'stoccfipb_reg', 'mplbir_reg','wt','treatment','outcome']
df = pd.DataFrame(columns=cols,data=data)
df.head()
# + Collapsed="false"
df = df.astype({"treatment":'bool'}, copy=False) #explicitly assigning treatment column as boolean
df.fillna(value=df.mean(),inplace=True) #filling the missing values
df.fillna(value=df.mode().loc[0],inplace=True)
data_1 = df[df["treatment"]==1]
data_0 = df[df["treatment"]==0]
print(np.mean(data_1["outcome"]))
print(np.mean(data_0["outcome"]))
print("ATE", np.mean(data_1["outcome"])- np.mean(data_0["outcome"]))
# + [markdown] Collapsed="false"
# **<font size="4">1. Model</font>**
# + Collapsed="false"
#The causal model has "treatment = is_heavier", "outcome = mortality" and "gestat10 = gestational weeks before birth"
model=CausalModel(
data = df,
treatment='treatment',
outcome='outcome',
common_causes='gestat10'
)
# + [markdown] Collapsed="false"
# **<font size="4">2. Identify</font>**
# + Collapsed="false"
identified_estimand = model.identify_effect()
# + [markdown] Collapsed="false"
# **<font size="4">3. Estimate Using Various Methods</font>**
# + [markdown] Collapsed="false"
# **<font size="3">3.1 Using Linear Regression</font>**
# + Collapsed="false"
estimate = model.estimate_effect(identified_estimand,
method_name="backdoor.linear_regression", test_significance=True
)
print(estimate)
print("ATE", np.mean(data_1["outcome"])- np.mean(data_0["outcome"]))
print("Causal Estimate is " + str(estimate.value))
# + [markdown] Collapsed="false"
# **<font size="3">3.2 Using Propensity Score Matching</font>**
# + Collapsed="false"
estimate = model.estimate_effect(identified_estimand,
method_name="backdoor.propensity_score_matching"
)
print("Causal Estimate is " + str(estimate.value))
print("ATE", np.mean(data_1["outcome"])- np.mean(data_0["outcome"]))
# + [markdown] Collapsed="false"
# **<font size="4">4. Refute</font>**
# + [markdown] Collapsed="false"
# **<font size="3">4.1 Adding a random cause</font>**
# + Collapsed="false"
refute_results=model.refute_estimate(identified_estimand, estimate,
method_name="random_common_cause")
print(refute_results)
# + [markdown] Collapsed="false"
# **<font size="3">4.2 Using a placebo treatment</font>**
# + Collapsed="false"
res_placebo=model.refute_estimate(identified_estimand, estimate,
method_name="placebo_treatment_refuter", placebo_type="permute")
print(res_placebo)
# + [markdown] Collapsed="false"
# **<font size="3">4.3 Using a data subset refuter</font>**
# + Collapsed="false"
res_subset=model.refute_estimate(identified_estimand, estimate,
method_name="data_subset_refuter", subset_fraction=0.9)
print(res_subset)
# + Collapsed="false"
|
docs/source/example_notebooks/dowhy_twins_example .ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Image Classification
# In this project, you'll classify images from the [CIFAR-10 dataset](https://www.cs.toronto.edu/~kriz/cifar.html). The dataset consists of airplanes, dogs, cats, and other objects. You'll preprocess the images, then train a convolutional neural network on all the samples. The images need to be normalized and the labels need to be one-hot encoded. You'll get to apply what you learned and build a convolutional, max pooling, dropout, and fully connected layers. At the end, you'll get to see your neural network's predictions on the sample images.
# ## Get the Data
# Run the following cell to download the [CIFAR-10 dataset for python](https://www.cs.toronto.edu/~kriz/cifar-10-python.tar.gz).
# + deletable=true editable=true
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
from urllib.request import urlretrieve
from os.path import isfile, isdir
from tqdm import tqdm
import problem_unittests as tests
import tarfile
cifar10_dataset_folder_path = 'cifar-10-batches-py'
# Use Floyd's cifar-10 dataset if present
floyd_cifar10_location = '/input/cifar-10/python.tar.gz'
if isfile(floyd_cifar10_location):
tar_gz_path = floyd_cifar10_location
else:
tar_gz_path = 'cifar-10-python.tar.gz'
class DLProgress(tqdm):
last_block = 0
def hook(self, block_num=1, block_size=1, total_size=None):
self.total = total_size
self.update((block_num - self.last_block) * block_size)
self.last_block = block_num
if not isfile(tar_gz_path):
with DLProgress(unit='B', unit_scale=True, miniters=1, desc='CIFAR-10 Dataset') as pbar:
urlretrieve(
'https://www.cs.toronto.edu/~kriz/cifar-10-python.tar.gz',
tar_gz_path,
pbar.hook)
if not isdir(cifar10_dataset_folder_path):
with tarfile.open(tar_gz_path) as tar:
tar.extractall()
tar.close()
tests.test_folder_path(cifar10_dataset_folder_path)
# + [markdown] deletable=true editable=true
# ## Explore the Data
# The dataset is broken into batches to prevent your machine from running out of memory. The CIFAR-10 dataset consists of 5 batches, named `data_batch_1`, `data_batch_2`, etc.. Each batch contains the labels and images that are one of the following:
# * airplane
# * automobile
# * bird
# * cat
# * deer
# * dog
# * frog
# * horse
# * ship
# * truck
#
# Understanding a dataset is part of making predictions on the data. Play around with the code cell below by changing the `batch_id` and `sample_id`. The `batch_id` is the id for a batch (1-5). The `sample_id` is the id for a image and label pair in the batch.
#
# Ask yourself "What are all possible labels?", "What is the range of values for the image data?", "Are the labels in order or random?". Answers to questions like these will help you preprocess the data and end up with better predictions.
# + deletable=true editable=true
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import helper
import numpy as np
# Explore the dataset
batch_id = 1
sample_id = 5
helper.display_stats(cifar10_dataset_folder_path, batch_id, sample_id)
# + [markdown] deletable=true editable=true
# ## Implement Preprocess Functions
# ### Normalize
# In the cell below, implement the `normalize` function to take in image data, `x`, and return it as a normalized Numpy array. The values should be in the range of 0 to 1, inclusive. The return object should be the same shape as `x`.
# + deletable=true editable=true
def normalize(x):
"""
Normalize a list of sample image data in the range of 0 to 1
: x: List of image data. The image shape is (32, 32, 3)
: return: Numpy array of normalize data
"""
mini = np.min(x)
maxi = np.max(x)
return (x-mini)/(maxi-mini)
# TODO: Implement Function
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_normalize(normalize)
# + [markdown] deletable=true editable=true
# ### One-hot encode
# Just like the previous code cell, you'll be implementing a function for preprocessing. This time, you'll implement the `one_hot_encode` function. The input, `x`, are a list of labels. Implement the function to return the list of labels as One-Hot encoded Numpy array. The possible values for labels are 0 to 9. The one-hot encoding function should return the same encoding for each value between each call to `one_hot_encode`. Make sure to save the map of encodings outside the function.
#
# Hint: Don't reinvent the wheel.
# + deletable=true editable=true
def one_hot_encode(x):
"""
One hot encode a list of sample labels. Return a one-hot encoded vector for each label.
: x: List of sample Labels
: return: Numpy array of one-hot encoded labels
"""
# TODO: Implement Function
values = x
n_values = np.max(values) + 1
return np.eye(n_values)[values]
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_one_hot_encode(one_hot_encode)
# + [markdown] deletable=true editable=true
# ### Randomize Data
# As you saw from exploring the data above, the order of the samples are randomized. It doesn't hurt to randomize it again, but you don't need to for this dataset.
# + [markdown] deletable=true editable=true
# ## Preprocess all the data and save it
# Running the code cell below will preprocess all the CIFAR-10 data and save it to file. The code below also uses 10% of the training data for validation.
# + deletable=true editable=true
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# Preprocess Training, Validation, and Testing Data
helper.preprocess_and_save_data(cifar10_dataset_folder_path, normalize, one_hot_encode)
# + [markdown] deletable=true editable=true
# # Check Point
# This is your first checkpoint. If you ever decide to come back to this notebook or have to restart the notebook, you can start from here. The preprocessed data has been saved to disk.
# + deletable=true editable=true
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
import pickle
import problem_unittests as tests
import helper
# Load the Preprocessed Validation data
valid_features, valid_labels = pickle.load(open('preprocess_validation.p', mode='rb'))
# + [markdown] deletable=true editable=true
# ## Build the network
# For the neural network, you'll build each layer into a function. Most of the code you've seen has been outside of functions. To test your code more thoroughly, we require that you put each layer in a function. This allows us to give you better feedback and test for simple mistakes using our unittests before you submit your project.
#
# >**Note:** If you're finding it hard to dedicate enough time for this course each week, we've provided a small shortcut to this part of the project. In the next couple of problems, you'll have the option to use classes from the [TensorFlow Layers](https://www.tensorflow.org/api_docs/python/tf/layers) or [TensorFlow Layers (contrib)](https://www.tensorflow.org/api_guides/python/contrib.layers) packages to build each layer, except the layers you build in the "Convolutional and Max Pooling Layer" section. TF Layers is similar to Keras's and TFLearn's abstraction to layers, so it's easy to pickup.
#
# >However, if you would like to get the most out of this course, try to solve all the problems _without_ using anything from the TF Layers packages. You **can** still use classes from other packages that happen to have the same name as ones you find in TF Layers! For example, instead of using the TF Layers version of the `conv2d` class, [tf.layers.conv2d](https://www.tensorflow.org/api_docs/python/tf/layers/conv2d), you would want to use the TF Neural Network version of `conv2d`, [tf.nn.conv2d](https://www.tensorflow.org/api_docs/python/tf/nn/conv2d).
#
# Let's begin!
#
# ### Input
# The neural network needs to read the image data, one-hot encoded labels, and dropout keep probability. Implement the following functions
# * Implement `neural_net_image_input`
# * Return a [TF Placeholder](https://www.tensorflow.org/api_docs/python/tf/placeholder)
# * Set the shape using `image_shape` with batch size set to `None`.
# * Name the TensorFlow placeholder "x" using the TensorFlow `name` parameter in the [TF Placeholder](https://www.tensorflow.org/api_docs/python/tf/placeholder).
# * Implement `neural_net_label_input`
# * Return a [TF Placeholder](https://www.tensorflow.org/api_docs/python/tf/placeholder)
# * Set the shape using `n_classes` with batch size set to `None`.
# * Name the TensorFlow placeholder "y" using the TensorFlow `name` parameter in the [TF Placeholder](https://www.tensorflow.org/api_docs/python/tf/placeholder).
# * Implement `neural_net_keep_prob_input`
# * Return a [TF Placeholder](https://www.tensorflow.org/api_docs/python/tf/placeholder) for dropout keep probability.
# * Name the TensorFlow placeholder "keep_prob" using the TensorFlow `name` parameter in the [TF Placeholder](https://www.tensorflow.org/api_docs/python/tf/placeholder).
#
# These names will be used at the end of the project to load your saved model.
#
# Note: `None` for shapes in TensorFlow allow for a dynamic size.
# + deletable=true editable=true
import tensorflow as tf
def neural_net_image_input(image_shape):
"""
Return a Tensor for a batch of image input
: image_shape: Shape of the images
: return: Tensor for image input.
"""
# TODO: Implement Function
image_height, image_width, color_channels = image_shape
return tf.placeholder(tf.float32, shape=[None, image_height, image_width, color_channels], name='x')
def neural_net_label_input(n_classes):
"""
Return a Tensor for a batch of label input
: n_classes: Number of classes
: return: Tensor for label input.
"""
return tf.placeholder(tf.float32, [None, n_classes], name='y')
def neural_net_keep_prob_input():
"""
Return a Tensor for keep probability
: return: Tensor for keep probability.
"""
# TODO: Implement Function
return tf.placeholder(tf.float32, name='keep_prob')
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tf.reset_default_graph()
tests.test_nn_image_inputs(neural_net_image_input)
tests.test_nn_label_inputs(neural_net_label_input)
tests.test_nn_keep_prob_inputs(neural_net_keep_prob_input)
# + [markdown] deletable=true editable=true
# ### Convolution and Max Pooling Layer
# Convolution layers have a lot of success with images. For this code cell, you should implement the function `conv2d_maxpool` to apply convolution then max pooling:
# * Create the weight and bias using `conv_ksize`, `conv_num_outputs` and the shape of `x_tensor`.
# * Apply a convolution to `x_tensor` using weight and `conv_strides`.
# * We recommend you use same padding, but you're welcome to use any padding.
# * Add bias
# * Add a nonlinear activation to the convolution.
# * Apply Max Pooling using `pool_ksize` and `pool_strides`.
# * We recommend you use same padding, but you're welcome to use any padding.
#
# **Note:** You **can't** use [TensorFlow Layers](https://www.tensorflow.org/api_docs/python/tf/layers) or [TensorFlow Layers (contrib)](https://www.tensorflow.org/api_guides/python/contrib.layers) for **this** layer, but you can still use TensorFlow's [Neural Network](https://www.tensorflow.org/api_docs/python/tf/nn) package. You may still use the shortcut option for all the **other** layers.
# + deletable=true editable=true
def conv2d_maxpool(x_tensor, conv_num_outputs, conv_ksize, conv_strides, pool_ksize, pool_strides):
"""
Apply convolution then max pooling to x_tensor
:param x_tensor: TensorFlow Tensor
:param conv_num_outputs: Number of outputs for the convolutional layer
:param conv_ksize: kernal size 2-D Tuple for the convolutional layer
:param conv_strides: Stride 2-D Tuple for convolution
:param pool_ksize: kernal size 2-D Tuple for pool
:param pool_strides: Stride 2-D Tuple for pool
: return: A tensor that represents convolution and max pooling of x_tensor
"""
# TODO: Implement Function
#print(type(conv_ksize[0]))
#print(type(conv_ksize[1]))
#print(type(int(x_tensor.get_shape()[3])))
#print(type(conv_num_outputs))
#conv_ksize[0], conv_ksize[1], int(x_tensor.get_shape()[3]), conv_num_outputs
weight = tf.Variable(tf.truncated_normal([conv_ksize[0], conv_ksize[1], int(x_tensor.get_shape()[3]), conv_num_outputs], stddev=0.1))
bias = tf.Variable(tf.zeros(conv_num_outputs))
conv_layer = tf.nn.conv2d(x_tensor, weight, strides=[1] + list(conv_strides) + [1], padding='SAME')
conv_layer = tf.nn.bias_add(conv_layer, bias)
conv_layer = tf.nn.relu(conv_layer)
conv_layer = tf.nn.max_pool(conv_layer, ksize=[1] + list(pool_ksize) + [1], strides=[1] + list(pool_strides) + [1], padding='SAME')
#print(conv_layer)
return conv_layer
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_con_pool(conv2d_maxpool)
# + [markdown] deletable=true editable=true
# ### Flatten Layer
# Implement the `flatten` function to change the dimension of `x_tensor` from a 4-D tensor to a 2-D tensor. The output should be the shape (*Batch Size*, *Flattened Image Size*). Shortcut option: you can use classes from the [TensorFlow Layers](https://www.tensorflow.org/api_docs/python/tf/layers) or [TensorFlow Layers (contrib)](https://www.tensorflow.org/api_guides/python/contrib.layers) packages for this layer. For more of a challenge, only use other TensorFlow packages.
# + deletable=true editable=true
def flatten(x_tensor):
"""
Flatten x_tensor to (Batch Size, Flattened Image Size)
: x_tensor: A tensor of size (Batch Size, ...), where ... are the image dimensions.
: return: A tensor of size (Batch Size, Flattened Image Size).
"""
# TODO: Implement Function
return tf.contrib.layers.flatten(x_tensor)
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_flatten(flatten)
# + [markdown] deletable=true editable=true
# ### Fully-Connected Layer
# Implement the `fully_conn` function to apply a fully connected layer to `x_tensor` with the shape (*Batch Size*, *num_outputs*). Shortcut option: you can use classes from the [TensorFlow Layers](https://www.tensorflow.org/api_docs/python/tf/layers) or [TensorFlow Layers (contrib)](https://www.tensorflow.org/api_guides/python/contrib.layers) packages for this layer. For more of a challenge, only use other TensorFlow packages.
# + deletable=true editable=true
def fully_conn(x_tensor, num_outputs):
"""
Apply a fully connected layer to x_tensor using weight and bias
: x_tensor: A 2-D tensor where the first dimension is batch size.
: num_outputs: The number of output that the new tensor should be.
: return: A 2-D tensor where the second dimension is num_outputs.
"""
# TODO: Implement Function
return tf.contrib.layers.fully_connected(x_tensor, num_outputs, activation_fn=None)
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_fully_conn(fully_conn)
# + [markdown] deletable=true editable=true
# ### Output Layer
# Implement the `output` function to apply a fully connected layer to `x_tensor` with the shape (*Batch Size*, *num_outputs*). Shortcut option: you can use classes from the [TensorFlow Layers](https://www.tensorflow.org/api_docs/python/tf/layers) or [TensorFlow Layers (contrib)](https://www.tensorflow.org/api_guides/python/contrib.layers) packages for this layer. For more of a challenge, only use other TensorFlow packages.
#
# **Note:** Activation, softmax, or cross entropy should **not** be applied to this.
# + deletable=true editable=true
def output(x_tensor, num_outputs):
"""
Apply a output layer to x_tensor using weight and bias
: x_tensor: A 2-D tensor where the first dimension is batch size.
: num_outputs: The number of output that the new tensor should be.
: return: A 2-D tensor where the second dimension is num_outputs.
"""
# TODO: Implement Function
return tf.contrib.layers.fully_connected(x_tensor, num_outputs, activation_fn=tf.nn.relu)
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_output(output)
# + [markdown] deletable=true editable=true
# ### Create Convolutional Model
# Implement the function `conv_net` to create a convolutional neural network model. The function takes in a batch of images, `x`, and outputs logits. Use the layers you created above to create this model:
#
# * Apply 1, 2, or 3 Convolution and Max Pool layers
# * Apply a Flatten Layer
# * Apply 1, 2, or 3 Fully Connected Layers
# * Apply an Output Layer
# * Return the output
# * Apply [TensorFlow's Dropout](https://www.tensorflow.org/api_docs/python/tf/nn/dropout) to one or more layers in the model using `keep_prob`.
# + deletable=true editable=true
def conv_net(x, keep_prob):
"""
Create a convolutional neural network model
: x: Placeholder tensor that holds image data.
: keep_prob: Placeholder tensor that hold dropout keep probability.
: return: Tensor that represents logits
"""
# TODO: Apply 1, 2, or 3 Convolution and Max Pool layers
# Play around with different number of outputs, kernel size and stride
# Function Definition from Above:
# conv2d_maxpool(x_tensor, conv_num_outputs, conv_ksize, conv_strides, pool_ksize, pool_strides)
c1s2 = conv2d_maxpool(x, 6, (5,5), (1,1), (2, 2), (2, 2))
c3s4 = conv2d_maxpool(c1s2, 16, (5,5), (1,1), (2, 2), (2, 2))
# TODO: Apply a Flatten Layer
# Function Definition from Above:
# flatten(x_tensor)
c5 = flatten(c3s4)
c5 = tf.nn.dropout(c5, keep_prob)
# TODO: Apply 1, 2, or 3 Fully Connected Layers
# Play around with different number of outputs
# Function Definition from Above:
# fully_conn(x_tensor, num_outputs)
f6 = fully_conn(c5, 84)
# TODO: Apply an Output Layer
# Set this to the number of classes
# Function Definition from Above:
# output(x_tensor, num_outputs)
f7 = output(f6, 10)
# TODO: return output
return f7
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
##############################
## Build the Neural Network ##
##############################
# Remove previous weights, bias, inputs, etc..
tf.reset_default_graph()
# Inputs
x = neural_net_image_input((32, 32, 3))
y = neural_net_label_input(10)
keep_prob = neural_net_keep_prob_input()
# Model
logits = conv_net(x, keep_prob)
# Name logits Tensor, so that is can be loaded from disk after training
logits = tf.identity(logits, name='logits')
# Loss and Optimizer
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=logits, labels=y))
optimizer = tf.train.AdamOptimizer().minimize(cost)
# Accuracy
correct_pred = tf.equal(tf.argmax(logits, 1), tf.argmax(y, 1))
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32), name='accuracy')
tests.test_conv_net(conv_net)
# + [markdown] deletable=true editable=true
# ## Train the Neural Network
# ### Single Optimization
# Implement the function `train_neural_network` to do a single optimization. The optimization should use `optimizer` to optimize in `session` with a `feed_dict` of the following:
# * `x` for image input
# * `y` for labels
# * `keep_prob` for keep probability for dropout
#
# This function will be called for each batch, so `tf.global_variables_initializer()` has already been called.
#
# Note: Nothing needs to be returned. This function is only optimizing the neural network.
# + deletable=true editable=true
def train_neural_network(session, optimizer, keep_probability, feature_batch, label_batch):
"""
Optimize the session on a batch of images and labels
: session: Current TensorFlow session
: optimizer: TensorFlow optimizer function
: keep_probability: keep probability
: feature_batch: Batch of Numpy image data
: label_batch: Batch of Numpy label data
"""
# TODO: Implement Function
session.run(optimizer, feed_dict={x: feature_batch, y: label_batch, keep_prob: keep_probability})
"""
DON'T MODIFY ANYTHING IN THIS CELL THAT IS BELOW THIS LINE
"""
tests.test_train_nn(train_neural_network)
# + [markdown] deletable=true editable=true
# ### Show Stats
# Implement the function `print_stats` to print loss and validation accuracy. Use the global variables `valid_features` and `valid_labels` to calculate validation accuracy. Use a keep probability of `1.0` to calculate the loss and validation accuracy.
# + deletable=true editable=true
def print_stats(session, feature_batch, label_batch, cost, accuracy):
"""
Print information about loss and validation accuracy
: session: Current TensorFlow session
: feature_batch: Batch of Numpy image data
: label_batch: Batch of Numpy label data
: cost: TensorFlow cost function
: accuracy: TensorFlow accuracy function
"""
# TODO: Implement Function
loss = session.run(cost, feed_dict={x: feature_batch, y: label_batch,keep_prob: 1.})
valid_acc = session.run(accuracy, feed_dict={x: feature_batch, y: label_batch, keep_prob: 1.})
print('loss is {}'.format(loss))
print('validation_accuracy is {}'.format(valid_acc))
# + [markdown] deletable=true editable=true
# ### Hyperparameters
# Tune the following parameters:
# * Set `epochs` to the number of iterations until the network stops learning or start overfitting
# * Set `batch_size` to the highest number that your machine has memory for. Most people set them to common sizes of memory:
# * 64
# * 128
# * 256
# * ...
# * Set `keep_probability` to the probability of keeping a node using dropout
# + deletable=true editable=true
# TODO: Tune Parameters
epochs = 10
batch_size = 128
keep_probability = 0.75
# + [markdown] deletable=true editable=true
# ### Train on a Single CIFAR-10 Batch
# Instead of training the neural network on all the CIFAR-10 batches of data, let's use a single batch. This should save time while you iterate on the model to get a better accuracy. Once the final validation accuracy is 50% or greater, run the model on all the data in the next section.
# + deletable=true editable=true
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
print('Checking the Training on a Single Batch...')
with tf.Session() as sess:
# Initializing the variables
sess.run(tf.initialize_all_variables())
# Training cycle
for epoch in range(epochs):
batch_i = 1
for batch_features, batch_labels in helper.load_preprocess_training_batch(batch_i, batch_size):
train_neural_network(sess, optimizer, keep_probability, batch_features, batch_labels)
print('Epoch {:>2}, CIFAR-10 Batch {}: '.format(epoch + 1, batch_i), end='')
print_stats(sess, batch_features, batch_labels, cost, accuracy)
# + [markdown] deletable=true editable=true
# ### Fully Train the Model
# Now that you got a good accuracy with a single CIFAR-10 batch, try it with all five batches.
# + deletable=true editable=true
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
save_model_path = './image_classification'
print('Training...')
with tf.Session() as sess:
# Initializing the variables
sess.run(tf.initialize_all_variables())
# Training cycle
for epoch in range(epochs):
# Loop over all batches
n_batches = 5
for batch_i in range(1, n_batches + 1):
for batch_features, batch_labels in helper.load_preprocess_training_batch(batch_i, batch_size):
train_neural_network(sess, optimizer, keep_probability, batch_features, batch_labels)
print('Epoch {:>2}, CIFAR-10 Batch {}: '.format(epoch + 1, batch_i), end='')
print_stats(sess, batch_features, batch_labels, cost, accuracy)
# Save Model
saver = tf.train.Saver()
save_path = saver.save(sess, save_model_path)
# + [markdown] deletable=true editable=true
# # Checkpoint
# The model has been saved to disk.
# ## Test Model
# Test your model against the test dataset. This will be your final accuracy. You should have an accuracy greater than 50%. If you don't, keep tweaking the model architecture and parameters.
# + deletable=true editable=true
"""
DON'T MODIFY ANYTHING IN THIS CELL
"""
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import tensorflow as tf
import pickle
import helper
import random
# Set batch size if not already set
try:
if batch_size:
pass
except NameError:
batch_size = 64
save_model_path = './image_classification'
n_samples = 4
top_n_predictions = 3
def test_model():
"""
Test the saved model against the test dataset
"""
test_features, test_labels = pickle.load(open('preprocess_test.p', mode='rb'))
loaded_graph = tf.Graph()
with tf.Session(graph=loaded_graph) as sess:
# Load model
loader = tf.train.import_meta_graph(save_model_path + '.meta')
loader.restore(sess, save_model_path)
# Get Tensors from loaded model
loaded_x = loaded_graph.get_tensor_by_name('x:0')
loaded_y = loaded_graph.get_tensor_by_name('y:0')
loaded_keep_prob = loaded_graph.get_tensor_by_name('keep_prob:0')
loaded_logits = loaded_graph.get_tensor_by_name('logits:0')
loaded_acc = loaded_graph.get_tensor_by_name('accuracy:0')
# Get accuracy in batches for memory limitations
test_batch_acc_total = 0
test_batch_count = 0
for test_feature_batch, test_label_batch in helper.batch_features_labels(test_features, test_labels, batch_size):
test_batch_acc_total += sess.run(
loaded_acc,
feed_dict={loaded_x: test_feature_batch, loaded_y: test_label_batch, loaded_keep_prob: 1.0})
test_batch_count += 1
print('Testing Accuracy: {}\n'.format(test_batch_acc_total/test_batch_count))
# Print Random Samples
random_test_features, random_test_labels = tuple(zip(*random.sample(list(zip(test_features, test_labels)), n_samples)))
random_test_predictions = sess.run(
tf.nn.top_k(tf.nn.softmax(loaded_logits), top_n_predictions),
feed_dict={loaded_x: random_test_features, loaded_y: random_test_labels, loaded_keep_prob: 1.0})
helper.display_image_predictions(random_test_features, random_test_labels, random_test_predictions)
test_model()
# + [markdown] deletable=true editable=true
# ## Why 50-80% Accuracy?
# You might be wondering why you can't get an accuracy any higher. First things first, 50% isn't bad for a simple CNN. Pure guessing would get you 10% accuracy. However, you might notice people are getting scores [well above 80%](http://rodrigob.github.io/are_we_there_yet/build/classification_datasets_results.html#43494641522d3130). That's because we haven't taught you all there is to know about neural networks. We still need to cover a few more techniques.
# ## Submitting This Project
# When submitting this project, make sure to run all the cells before saving the notebook. Save the notebook file as "dlnd_image_classification.ipynb" and save it as a HTML file under "File" -> "Download as". Include the "helper.py" and "problem_unittests.py" files in your submission.
|
Udacity-DL/image_classification_2/dlnd_image_classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Furniture Sales Time Series Model
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# -
#loading the data
df= pd.read_excel('Furniture-Sales.xls',sheet_name='Orders',index_col='Row ID')
df.shape
df.columns
df.head()
df.dtypes
# ## Data Preprocessing
# Taking the order date and sales in our data as we only need that to predict future sales as per time series model.
data= df.drop(['Order ID','Ship Date', 'Ship Mode', 'Customer ID',
'Customer Name', 'Segment', 'Country', 'City', 'State', 'Postal Code',
'Region', 'Product ID', 'Category', 'Sub-Category', 'Product Name',
'Quantity', 'Discount', 'Profit'], axis=1)
data.head()
# Now as we see in data we have more than one sales for a particular date
# so we will group the data by date and will do the sum of its sales
data = data.groupby('Order Date', sort=True)['Sales'].sum().reset_index()
data.set_index('Order Date', inplace=True)
# So if you want you can slice it now with day, date or year
data['2014-01']
# Now as we know mainly sales are done with Monthly prediction so we will change the freq of daily to monthly with avg monthly sales and will take first of every month.
data = data.resample('MS').mean()
data['2017']
# ## Seasonality Check
# Visulaizing the data
data.plot(figsize=(15,5))
plt.show()
# Now if we will see in graph, Per year sales is less in starting month and max in end month with low sales in few mid month. Which represent the seasonality trend both
# We have seasonal decompose class whcih returns all the following plot data
from statsmodels.tsa.seasonal import seasonal_decompose
decom= seasonal_decompose(data, model= 'additive')
# Seasonal
decom.seasonal.plot(figsize= (15,5))
# Trend
decom.trend.plot(figsize= (15,5))
# We can see we have very less increasing trend in our data, starting from july 2015
#Residuals
decom.resid.plot(figsize=(15,5))
#original data plot
decom.observed.plot(figsize=(15,5))
# Now as we have Trend we will try to use lag to check trend.
diff= data.diff(periods=1)
seasonal_decompose(diff.dropna(),model='additive').trend.plot(figsize=(15,5))
# Now we can see with One lag , we can see increasing trend is gone, which we can use for model.
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
plot_acf(data)
# Acf plot also suggest with no linear trend
# now getting training and testing data set
train= data.iloc[0:36,:]
test= data.iloc[36:48,:]
print(train.shape, test.shape)
import warnings
warnings.filterwarnings('ignore')
import itertools as it
p=d=q=range(0,2)
pdq= list(it.product(p,d,q))
seasonal_pdq=[(x[0], x[1], x[2], 12) for x in list(it.product(p, d, q))]
print('Examples of parameter combinations for Seasonal ARIMA...')
print('SARIMAX: {} x {}'.format(pdq[1], seasonal_pdq[1]))
print('SARIMAX: {} x {}'.format(pdq[1], seasonal_pdq[2]))
print('SARIMAX: {} x {}'.format(pdq[2], seasonal_pdq[3]))
print('SARIMAX: {} x {}'.format(pdq[2], seasonal_pdq[4]))
from statsmodels.tsa.api import SARIMAX
pair= {}
for param in pdq:
for param_seasonal in seasonal_pdq:
try:
mod = SARIMAX(data,order=param,
seasonal_order=param_seasonal,
enforce_stationarity=False,
enforce_invertibility=False)
results = mod.fit()
pair[results.aic]=(param,param_seasonal)
# print('ARIMA{}x{}12 - AIC:{}'.format(param, param_seasonal, results.aic))
except:
continue
best_order,best_s_order= pair[min(pair.keys())]
print(best_order,best_s_order)
final_model= SARIMAX(data,order=best_order,seasonal_order=best_s_order).fit()
final_model.summary()
final_model.plot_diagnostics(figsize=(16, 8))
plt.show()
# our model diagnostics suggests that the model residuals are near normally distributed.
yp= final_model.predict(start='2017-01-01', end='2017-12-01')
plt.figure(figsize=(15,5))
plt.plot(train,'blue',label='Train')
plt.plot(test,'red',label='Test')
plt.plot(yp,'green',label='Pred')
plt.legend(loc='best')
plt.show()
# +
from sklearn.metrics import mean_squared_error, r2_score
print('rmse:{}'.format(np.sqrt(mean_squared_error(test,yp))))
print('R2 Score:{}'.format(r2_score(test,yp)))
# -
plt.figure(figsize=(15,5))
plt.plot(data,'blue',label='Train')
plt.plot(final_model.forecast(steps=100),'green')
plt.show()
# So now with R2 of .65 and above we cn consider it as a good model. Which captured trend and seasonality both for future also.
#
from pmdarima.arima import auto_arima
final_model= auto_arima(data, start_p=0, start_q=0,d=1,
max_p=3,max_q=3, m=12,
start_P=0,D=1,start_Q=0 ,seasonal=True,
max_P=3,max_Q=3, trace=True,
error_action='ignore',
suppress_warnings=True,
stepwise=True)
model= final_model.fit(data)
forecast = model.predict(n_periods=len(test))
forecast = pd.DataFrame(forecast,index = test.index, columns=['Prediction'])
print('rmse:{}'.format(np.sqrt(mean_squared_error(test['Sales'],forecast['Prediction']))))
print('R2 Score:{}'.format(r2_score(test['Sales'],forecast['Prediction'])))
plt.figure(figsize=(15,5))
plt.plot(train['Sales'],'blue',label='Train')
plt.plot(test['Sales'],'red',label='Test')
plt.plot(forecast['Prediction'],'green',label='Pred')
plt.legend(loc='best')
plt.show()
# Auto_Arima is not good at all as per R2 Score
|
furniture-tsa.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import rcParams
rcParams.update({'figure.autolayout': True})
import seaborn as sns
import json
import pandas as pd
from IPython.core.debugger import Tracer
# -
def make_dataframe(filename, trial_id, stride = 1, stop_point=500):
with open(filename) as jfile:
dat = json.load(jfile)
for i in range(len(dat)):
dat[i] = [trial_id] + dat[i]
stop_point = min(stop_point, len(dat)-5)
dat = dat[:stop_point:stride]
df = pd.DataFrame(dat, columns=['trial_id','wall_time', 'step', 'value'])
return df
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'error_test_cond_cifar_bs900.pdf'
# bs900_drop
file_names_ccnf_bs900 = ['./figures_neurips/result_data_published/run-900-ccnf-drop-1_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-900-ccnf-drop-2_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-900-ccnf-drop-3_error_validation-tag-error.json',]
keys_ccnf_bs900 = ['run1','run2','run3']
id_ccnf_bs900 = 'CCNF (baseline)'
stop_point_ccnf_bs900 = 500
# bs900_mc
file_names_info_bs900 = ['./figures_neurips/result_data_published/run-900-infocnf-1_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-900-infocnf-2_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-900-infocnf-3_error_validation-tag-error.json',]
keys_info_bs900 = ['run1','run2','run3']
id_info_bs900 = 'InfoCNF'
stop_point_info_bs900 = 500
# bs900_mc
file_names_infotune_bs900 = ['./figures_neurips/result_data_published/run-900-infotunecnf-1_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-900-infotunecnf-2_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-900-infotunecnf-3_error_validation-tag-error.json',]
keys_infotune_bs900 = ['run1','run2','run3']
id_infotune_bs900 = 'InfoCNF_tuned'
stop_point_infotune_bs900 = 500
# bs900_mc_drop
file_names_gate_bs900 = ['./figures_neurips/result_data_published/run-900-gatecnf-2_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-900-gatecnf-3_error_validation-tag-error.json',]
keys_gate_bs900 = ['run1','run2','run3']
id_gate_bs900 = 'Gated InfoCNF'
stop_point_gate_bs900 = 500
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None):
df = []
df_min = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
stop_indx = min(500, len(df[-1]['value']))
df_min.append(min(df[-1]['value'][:stop_indx]))
print(np.argmin(df[-1]['value'][:stop_indx]))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color)
return sns_plot, df_min
# sns_plot, df_min_ccnf = plot_line(file_names_ccnf_bs900, keys_ccnf_bs900, id_ccnf_bs900, stride=10, stop_point=stop_point_ccnf_bs900, color='red')
# sns_plot, df_min_info = plot_line(file_names_info_bs900, keys_info_bs900, id_info_bs900, stride=10, stop_point=stop_point_info_bs900, color='green')
sns_plot, df_min_gate = plot_line(file_names_gate_bs900, keys_gate_bs900, id_gate_bs900, stride=1, stop_point=stop_point_gate_bs900, color='blue')
#sns_plot, df_min_infotune = plot_line(file_names_infotune_bs900, keys_infotune_bs900, id_infotune_bs900, stride=10, stop_point=stop_point_infotune_bs900, color='orange')
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('Test error', fontsize = 16)
sns_plot.set_xlim(0,380)
sns_plot.set_ylim(0.2,0.6)
sns_plot.legend().set_visible(False)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'nll_test_cond_cifar_bs900.pdf'
# bs900_drop
file_names_ccnf_bs900 = ['./figures_neurips/result_data_published/run-900-ccnf-drop-1_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-900-ccnf-drop-2_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-900-ccnf-drop-3_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_ccnf_bs900 = ['run1','run2','run3']
id_ccnf_bs900 = 'CCNF (baseline)'
stop_point_ccnf_bs900 = 500
# bs900_mc
file_names_info_bs900 = ['./figures_neurips/result_data_published/run-900-infocnf-1_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-900-infocnf-2_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-900-infocnf-3_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_info_bs900 = ['run1','run2','run3']
id_info_bs900 = 'InfoCNF'
stop_point_info_bs900 = 500
file_names_infotune_bs900 = ['./figures_neurips/result_data_published/run-900-infotunecnf-1_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-900-infotunecnf-2_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-900-infotunecnf-3_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_infotune_bs900 = ['run1','run2','run3']
id_infotune_bs900 = 'InfoCNF_tuned'
stop_point_infotune_bs900 = 500
# bs900_mc_drop
file_names_gate_bs900 = ['./figures_neurips/result_data_published/run-900-gatecnf-2_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-900-gatecnf-3_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_gate_bs900 = ['run1','run2','run3']
id_gate_bs900 = 'Gated InfoCNF'
stop_point_gate_bs900 = 500
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None):
df = []
df_min = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
stop_indx = min(500, len(df[-1]['value']))
df_min.append(min(df[-1]['value'][:stop_indx]))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color)
return sns_plot, df_min
sns_plot, df_min_ccnf = plot_line(file_names_ccnf_bs900, keys_ccnf_bs900, id_ccnf_bs900, stride=10, stop_point=stop_point_ccnf_bs900, color='red')
sns_plot, df_min_info = plot_line(file_names_info_bs900, keys_info_bs900, id_info_bs900, stride=10, stop_point=stop_point_info_bs900, color='green')
sns_plot, df_min_gate = plot_line(file_names_gate_bs900, keys_gate_bs900, id_gate_bs900, stride=10, stop_point=stop_point_gate_bs900, color='blue')
# sns_plot, df_min_infotune = plot_line(file_names_infotune_bs900, keys_infotune_bs900, id_infotune_bs900, stride=10, stop_point=stop_point_infotune_bs900, color='orange')
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('NLL (bits/dim)', fontsize = 16)
sns_plot.set_xlim(0,380)
sns_plot.set_ylim(3.5,4.4)
sns_plot.legend().set_visible(False)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'nfe_cond_cifar_bs900.pdf'
# bs900_drop
file_names_ccnf_bs900 = ['./figures_neurips/result_data_published/run-900-ccnf-drop-1_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-900-ccnf-drop-2_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-900-ccnf-drop-3_nfe_train_epoch-tag-nfe.json',]
keys_ccnf_bs900 = ['run1','run2','run3']
id_ccnf_bs900 = 'CCNF (baseline)'
stop_point_ccnf_bs900 = 500
# bs900_mc
file_names_info_bs900 = ['./figures_neurips/result_data_published/run-900-infocnf-1_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-900-infocnf-2_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-900-infocnf-3_nfe_train_epoch-tag-nfe.json',]
keys_info_bs900 = ['run1','run2','run3']
id_info_bs900 = 'InfoCNF'
stop_point_info_bs900 = 500
# bs900_mc
file_names_infotune_bs900 = ['./figures_neurips/result_data_published/run-900-infotunecnf-1_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-900-infotunecnf-2_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-900-infotunecnf-3_nfe_train_epoch-tag-nfe.json',]
keys_infotune_bs900 = ['run1','run2','run3']
id_infotune_bs900 = 'InfoCNF_tuned'
stop_point_infotune_bs900 = 500
# bs900_mc_drop
file_names_gate_bs900 = ['./figures_neurips/result_data_published/run-900-gatecnf-2_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-900-gatecnf-3_nfe_train_epoch-tag-nfe.json',]
keys_gate_bs900 = ['run1','run2','run3']
id_gate_bs900 = 'Gated InfoCNF'
stop_point_gate_bs900 = 500
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None):
df = []
df_mean = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
stop_indx = min(500, len(df[-1]['value']))
df_mean.append(np.mean(df[-1]['value'][:stop_indx]))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color)
return sns_plot, df_mean
sns_plot, df_mean_ccnf = plot_line(file_names_ccnf_bs900, keys_ccnf_bs900, id_ccnf_bs900, stride=10, stop_point=stop_point_ccnf_bs900, color='red')
sns_plot, df_mean_info = plot_line(file_names_info_bs900, keys_info_bs900, id_info_bs900, stride=10, stop_point=stop_point_info_bs900, color='green')
sns_plot, df_mean_gate = plot_line(file_names_gate_bs900, keys_gate_bs900, id_gate_bs900, stride=10, stop_point=stop_point_gate_bs900, color='blue')
# sns_plot, df_mean_infotune = plot_line(file_names_infotune_bs900, keys_infotune_bs900, id_infotune_bs900, stride=10, stop_point=stop_point_infotune_bs900, color='orange')
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('# Function Evaluations', fontsize = 16)
sns_plot.set_xlim(0,380)
sns_plot.legend().set_visible(False)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'nfe_cond_cifar_bs900.pdf'
# bs900_drop
file_names_ccnf_bs900 = ['./figures_neurips/result_data_published/run-900-ccnf-drop-1_epoch_time_validation-tag-epoch_time.json',
'./figures_neurips/result_data_published/run-900-ccnf-drop-2_epoch_time_validation-tag-epoch_time.json',
'./figures_neurips/result_data_published/run-900-ccnf-drop-3_epoch_time_validation-tag-epoch_time.json',]
keys_ccnf_bs900 = ['run1','run2','run3']
id_ccnf_bs900 = 'CCNF (baseline)'
stop_point_ccnf_bs900 = 380
# bs900_mc
file_names_info_bs900 = ['./figures_neurips/result_data_published/run-900-infocnf-1_epoch_time_validation-tag-epoch_time.json',
'./figures_neurips/result_data_published/run-900-infocnf-2_epoch_time_validation-tag-epoch_time.json',
'./figures_neurips/result_data_published/run-900-infocnf-3_epoch_time_validation-tag-epoch_time.json',]
keys_info_bs900 = ['run1','run2','run3']
id_info_bs900 = 'InfoCNF'
stop_point_info_bs900 = 380
# bs900_mc
file_names_infotune_bs900 = ['./figures_neurips/result_data_published/run-900-infotunecnf-1_epoch_time_validation-tag-epoch_time.json',
'./figures_neurips/result_data_published/run-900-infotunecnf-2_epoch_time_validation-tag-epoch_time.json',
'./figures_neurips/result_data_published/run-900-infotunecnf-3_epoch_time_validation-tag-epoch_time.json',]
keys_infotune_bs900 = ['run1','run2','run3']
id_infotune_bs900 = 'InfoCNF_tuned'
stop_point_infotune_bs900 = 380
# bs900_mc_drop
file_names_gate_bs900 = ['./figures_neurips/result_data_published/run-900-gatecnf-1_epoch_time_validation-tag-epoch_time.json',
'./figures_neurips/result_data_published/run-900-gatecnf-2_epoch_time_validation-tag-epoch_time.json',
'./figures_neurips/result_data_published/run-900-gatecnf-3_epoch_time_validation-tag-epoch_time.json',]
keys_gate_bs900 = ['run1','run2','run3']
id_gate_bs900 = 'Gated InfoCNF'
stop_point_gate_bs900 = 380
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None):
df = []
df_mean = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
stop_indx = min(500, len(df[-1]['value']))
df_mean.append(np.mean(df[-1]['value'][:stop_indx]))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color)
return sns_plot, df_mean
sns_plot, df_mean_ccnf = plot_line(file_names_ccnf_bs900, keys_ccnf_bs900, id_ccnf_bs900, stride=10, stop_point=stop_point_ccnf_bs900, color='red')
sns_plot, df_mean_info = plot_line(file_names_info_bs900, keys_info_bs900, id_info_bs900, stride=10, stop_point=stop_point_info_bs900, color='green')
sns_plot, df_mean_gate = plot_line(file_names_gate_bs900, keys_gate_bs900, id_gate_bs900, stride=10, stop_point=stop_point_gate_bs900, color='blue')
# sns_plot, df_mean_infotune = plot_line(file_names_infotune_bs900, keys_infotune_bs900, id_infotune_bs900, stride=10, stop_point=stop_point_infotune_bs900, color='orange')
print(df_mean_ccnf)
print(df_mean_info)
print(df_mean_gate)
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('# Function Evaluations', fontsize = 16)
sns_plot.set_xlim(0,380)
sns_plot.legend().set_visible(False)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'error_test_cond_cifar_bs8k.pdf'
# bs8k_drop
file_names_ccnf_bs8k = ['./figures_neurips/result_data_published/run-8k-ccnf-drop-1_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-8k-ccnf-drop-2_error_validation-tag-error.json',
]
keys_ccnf_bs8k = ['run1','run2']
id_ccnf_bs8k = 'CCNF (baseline)'
stop_point_ccnf_bs8k = 500
# bs8k_mc
file_names_info_bs8k = ['./figures_neurips/result_data_published/run-8k-infocnf-drop-1_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-8k-infocnf-drop-2_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-8k-infocnf-drop-3_error_validation-tag-error.json',]
keys_info_bs8k = ['run1','run2','run3']
id_info_bs8k = 'InfoCNF'
stop_point_info_bs8k = 500
# b8k_mc_drop
file_names_gate_std15_bs8k = ['./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-1_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-2_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-3_error_validation-tag-error.json',]
keys_gate_std15_bs8k = ['run1','run2','run3']
id_gate_std15_bs8k = 'Gated InfoCNF'
stop_point_std15_gate_bs8k = 500
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None):
df = []
df_min = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
stop_indx = min(500, len(df[-1]['value']))
df_min.append(min(df[-1]['value'][:stop_indx]))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color)
return sns_plot, df_min
sns_plot, _ = plot_line(file_names_ccnf_bs8k, keys_ccnf_bs8k, id_ccnf_bs8k, stride=20, stop_point=stop_point_ccnf_bs8k, color='red')
sns_plot, _ = plot_line(file_names_info_bs8k, keys_info_bs8k, id_info_bs8k, stride=10, stop_point=stop_point_info_bs8k, color='green')
sns_plot, _ = plot_line(file_names_gate_std15_bs8k, keys_gate_std15_bs8k, id_gate_std15_bs8k, stride=10, stop_point=stop_point_std15_gate_bs8k, color='blue')
# sns_plot, df_min_ccnf = plot_line(file_names_ccnf_bs8k, keys_ccnf_bs8k, id_ccnf_bs8k, stride=1, stop_point=stop_point_ccnf_bs8k, color='red')
# sns_plot, df_min_info = plot_line(file_names_info_bs8k, keys_info_bs8k, id_info_bs8k, stride=1, stop_point=stop_point_info_bs8k, color='green')
# sns_plot, df_min_gate = plot_line(file_names_gate_std15_bs8k, keys_gate_std15_bs8k, id_gate_std15_bs8k, stride=1, stop_point=stop_point_std15_gate_bs8k, color='blue')
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('Test error', fontsize = 16)
sns_plot.set_xlim(0,380)
sns_plot.set_ylim(0.2,0.6)
sns_plot.legend().set_visible(False)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'nll_test_cond_cifar_bs8k.pdf'
# bs8k_drop
file_names_ccnf_bs8k = ['./figures_neurips/result_data_published/run-8k-ccnf-drop-1_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-8k-ccnf-drop-2_bits_per_dim_validation-tag-bits_per_dim.json',
]
keys_ccnf_bs8k = ['run1','run2']
id_ccnf_bs8k = 'CCNF (baseline)'
stop_point_ccnf_bs8k = 500
# bs8k_mc
file_names_info_bs8k = ['./figures_neurips/result_data_published/run-8k-infocnf-drop-1_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-8k-infocnf-drop-2_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-8k-infocnf-drop-3_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_info_bs8k = ['run1','run2','run3']
id_info_bs8k = 'InfoCNF'
stop_point_info_bs8k = 500
# b8k_mc_drop
file_names_gate_std15_bs8k = ['./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-1_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-2_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-3_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_gate_std15_bs8k = ['run1','run2','run3']
id_gate_std15_bs8k = 'Gated InfoCNF'
stop_point_std15_gate_bs8k = 500
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None):
df = []
df_min = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
stop_indx = min(500, len(df[-1]['value']))
df_min.append(min(df[-1]['value'][:stop_indx]))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color)
return sns_plot, df_min
sns_plot, _ = plot_line(file_names_ccnf_bs8k, keys_ccnf_bs8k, id_ccnf_bs8k, stride=20, stop_point=stop_point_ccnf_bs8k, color='red')
sns_plot, _ = plot_line(file_names_info_bs8k, keys_info_bs8k, id_info_bs8k, stride=10, stop_point=stop_point_info_bs8k, color='green')
sns_plot, _ = plot_line(file_names_gate_std15_bs8k, keys_gate_std15_bs8k, id_gate_std15_bs8k, stride=10, stop_point=stop_point_std15_gate_bs8k, color='blue')
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('NLL (bits/dim)', fontsize = 16)
sns_plot.set_xlim(0,380)
sns_plot.set_ylim(3.6,4.5)
sns_plot.legend().set_visible(False)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'nfe_cond_cifar_bs8k.pdf'
# bs8k_drop
file_names_ccnf_bs8k = ['./figures_neurips/result_data_published/run-8k-ccnf-drop-1_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-8k-ccnf-drop-2_nfe_train_epoch-tag-nfe.json',
]
keys_ccnf_bs8k = ['run1','run2']
id_ccnf_bs8k = 'CCNF (baseline)'
stop_point_ccnf_bs8k = 500
# bs8k_mc
file_names_info_bs8k = ['./figures_neurips/result_data_published/run-8k-infocnf-drop-1_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-8k-infocnf-drop-2_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-8k-infocnf-drop-3_nfe_train_epoch-tag-nfe.json',]
keys_info_bs8k = ['run1','run2','run3']
id_info_bs8k = 'InfoCNF'
stop_point_info_bs8k = 500
# b8k_mc_drop
file_names_gate_std15_bs8k = ['./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-1_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-2_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-3_nfe_train_epoch-tag-nfe.json',]
keys_gate_std15_bs8k = ['run1','run2','run3']
id_gate_std15_bs8k = 'Gated InfoCNF'
stop_point_std15_gate_bs8k = 500
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None):
df = []
df_mean = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
stop_indx = min(500, len(df[-1]['value']))
df_mean.append(np.mean(df[-1]['value'][:stop_indx]))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color)
return sns_plot, df_mean
sns_plot, _ = plot_line(file_names_ccnf_bs8k, keys_ccnf_bs8k, id_ccnf_bs8k, stride=20, stop_point=stop_point_ccnf_bs8k, color='red')
sns_plot, _ = plot_line(file_names_info_bs8k, keys_info_bs8k, id_info_bs8k, stride=20, stop_point=stop_point_info_bs8k, color='green')
sns_plot, _ = plot_line(file_names_gate_std15_bs8k, keys_gate_std15_bs8k, id_gate_std15_bs8k, stride=20, stop_point=stop_point_std15_gate_bs8k, color='blue')
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('# Function Evaluations', fontsize = 16)
sns_plot.set_xlim(0,380)
sns_plot.legend().set_visible(False)
# sns_plot.set_ylim(0.27,0.6)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'nll_test_uncond_cifar_bs900.pdf'
# bs900_drop
file_names_cnf_bs900 = ['./figures_neurips/result_data_published/run-900_run2_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-900_run3_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_cnf_bs900 = ['run2','run3']
id_cnf_bs900 = 'CNF (baseline)'
stop_point_cnf_bs900 = 500
# bs900_mc_drop
file_names_gate_bs900 = ['./figures_neurips/result_data_published/run-900_rl_std_6_run2_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-900_rl_std_6_run3_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_gate_bs900 = ['run2','run3']
id_gate_bs900 = 'Gated CNF'
stop_point_gate_bs900 = 500
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None):
df = []
df_min = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
stop_indx = min(500, len(df[-1]['value']))
df_min.append(min(df[-1]['value'][:stop_indx]))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color)
return sns_plot, df_min
sns_plot, df_min_ccnf = plot_line(file_names_cnf_bs900, keys_cnf_bs900, id_cnf_bs900, stride=10, stop_point=stop_point_cnf_bs900, color='red')
sns_plot, df_min_gate = plot_line(file_names_gate_bs900, keys_gate_bs900, id_gate_bs900, stride=10, stop_point=stop_point_gate_bs900, color='blue')
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('NLL (bits/dim)', fontsize = 16)
sns_plot.set_xlim(0,380)
sns_plot.set_ylim(3.35,3.85)
sns_plot.legend().set_visible(False)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'nfe_uncond_cifar_bs900.pdf'
# bs900_drop
file_names_cnf_bs900 = ['./figures_neurips/result_data_published/run-900_run2_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-900_run3_nfe_train_epoch-tag-nfe.json',]
keys_cnf_bs900 = ['run2','run3']
id_cnf_bs900 = 'CNF (baseline)'
stop_point_cnf_bs900 = 500
# bs900_mc_drop
file_names_gate_bs900 = ['./figures_neurips/result_data_published/run-900_rl_std_6_run2_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-900_rl_std_6_run3_nfe_train_epoch-tag-nfe.json',]
keys_gate_bs900 = ['run2','run3']
id_gate_bs900 = 'Gated CNF'
stop_point_gate_bs900 = 500
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None):
df = []
df_mean = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
stop_indx = min(500, len(df[-1]['value']))
df_mean.append(np.mean(df[-1]['value'][:stop_indx]))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color)
return sns_plot, df_mean
sns_plot, df_mean_ccnf = plot_line(file_names_cnf_bs900, keys_cnf_bs900, id_cnf_bs900, stride=10, stop_point=stop_point_cnf_bs900, color='red')
sns_plot, df_mean_gate = plot_line(file_names_gate_bs900, keys_gate_bs900, id_gate_bs900, stride=10, stop_point=stop_point_gate_bs900, color='blue')
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('# Function Evaluations', fontsize = 16)
sns_plot.set_xlim(0,380)
sns_plot.legend().set_visible(False)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'error_test_large_batches_methods.pdf'
# bs8k_drop
file_names_ccnf_bs8k = ['./figures_neurips/result_data_published/run-8k-ccnf-drop-1_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-8k-ccnf-drop-2_error_validation-tag-error.json',
]
keys_ccnf_bs8k = ['run1','run2']
id_ccnf_bs8k = 'CCNF'
stop_point_ccnf_bs8k = 500
# bs8k_mc
file_names_info_bs8k = ['./figures_neurips/result_data_published/run-8k-infocnf-drop-1_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-8k-infocnf-drop-2_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-8k-infocnf-drop-3_error_validation-tag-error.json',]
keys_info_bs8k = ['run1','run2','run3']
id_info_bs8k = 'InfoCNF'
stop_point_info_bs8k = 500
# bs8k_baseline_ccnf
file_names_ccnf_bl_bs8k = ['./figures_neurips/result_data_published/run-8k-ccnf-drop-baseline-1_error_validation-tag-error.json',]
keys_ccnf_bl_bs8k = ['run1',]
id_ccnf_bl_bs8k = 'CCNF without large lr \n and adjusted tolerance'
stop_point_ccnf_bl_bs8k = 500
# bs8k_baseline_ccnf
file_names_info_bl_bs8k = ['./figures_neurips/result_data_published/run-8k-infocnf-drop-baseline-1_error_validation-tag-error.json',]
keys_info_bl_bs8k = ['run1',]
id_info_bl_bs8k = 'InfoCNF without large lr \n and adjusted tolerance'
stop_point_info_bl_bs8k = 500
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None):
df = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color)
return sns_plot
sns_plot = plot_line(file_names_ccnf_bs8k, keys_ccnf_bs8k, id_ccnf_bs8k, stride=20, stop_point=stop_point_ccnf_bs8k, color='red')
sns_plot = plot_line(file_names_info_bs8k, keys_info_bs8k, id_info_bs8k, stride=10, stop_point=stop_point_info_bs8k, color='green')
sns_plot = plot_line(file_names_ccnf_bl_bs8k, keys_ccnf_bl_bs8k, id_ccnf_bl_bs8k, stride=10, stop_point=stop_point_ccnf_bl_bs8k, color='magenta')
sns_plot = plot_line(file_names_info_bl_bs8k, keys_info_bl_bs8k, id_info_bl_bs8k, stride=10, stop_point=stop_point_info_bl_bs8k, color='purple')
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('Test error', fontsize = 16)
sns_plot.set_xlim(0,380)
sns_plot.set_ylim(0.3,0.8)
sns_plot.legend().set_visible(False)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'nll_test_large_batches_methods.pdf'
# bs8k_drop
file_names_ccnf_bs8k = ['./figures_neurips/result_data_published/run-8k-ccnf-drop-1_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-8k-ccnf-drop-2_bits_per_dim_validation-tag-bits_per_dim.json',
]
keys_ccnf_bs8k = ['run1','run2']
id_ccnf_bs8k = 'CCNF'
stop_point_ccnf_bs8k = 500
# bs8k_mc
file_names_info_bs8k = ['./figures_neurips/result_data_published/run-8k-infocnf-drop-1_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-8k-infocnf-drop-2_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-8k-infocnf-drop-3_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_info_bs8k = ['run1','run2','run3']
id_info_bs8k = 'InfoCNF'
stop_point_info_bs8k = 500
# bs8k_baseline_ccnf
file_names_ccnf_bl_bs8k = ['./figures_neurips/result_data_published/run-8k-ccnf-drop-baseline-1_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_ccnf_bl_bs8k = ['run1',]
id_ccnf_bl_bs8k = 'CCNF without large lr \n and adjusted tolerance'
stop_point_ccnf_bl_bs8k = 500
# bs8k_baseline_ccnf
file_names_info_bl_bs8k = ['./figures_neurips/result_data_published/run-8k-infocnf-drop-baseline-1_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_info_bl_bs8k = ['run1',]
id_info_bl_bs8k = 'InfoCNF without large lr \n and adjusted tolerance'
stop_point_info_bl_bs8k = 500
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None):
df = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color)
return sns_plot
sns_plot = plot_line(file_names_ccnf_bs8k, keys_ccnf_bs8k, id_ccnf_bs8k, stride=20, stop_point=stop_point_ccnf_bs8k, color='red')
sns_plot = plot_line(file_names_info_bs8k, keys_info_bs8k, id_info_bs8k, stride=10, stop_point=stop_point_info_bs8k, color='green')
sns_plot = plot_line(file_names_ccnf_bl_bs8k, keys_ccnf_bl_bs8k, id_ccnf_bl_bs8k, stride=10, stop_point=stop_point_ccnf_bl_bs8k, color='magenta')
sns_plot = plot_line(file_names_info_bl_bs8k, keys_info_bl_bs8k, id_info_bl_bs8k, stride=10, stop_point=stop_point_info_bl_bs8k, color='purple')
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('NLL (bits/dim)', fontsize = 16)
sns_plot.set_xlim(0,380)
sns_plot.set_ylim(3.6,5.0)
sns_plot.legend().set_visible(False)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'nfe_large_batches_methods.pdf'
# bs8k_drop
file_names_ccnf_bs8k = ['./figures_neurips/result_data_published/run-8k-ccnf-drop-1_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-8k-ccnf-drop-2_nfe_train_epoch-tag-nfe.json',
]
keys_ccnf_bs8k = ['run1','run2']
id_ccnf_bs8k = 'CCNF'
stop_point_ccnf_bs8k = 500
# bs8k_mc
file_names_info_bs8k = ['./figures_neurips/result_data_published/run-8k-infocnf-drop-1_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-8k-infocnf-drop-2_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-8k-infocnf-drop-3_nfe_train_epoch-tag-nfe.json',]
keys_info_bs8k = ['run1','run2','run3']
id_info_bs8k = 'InfoCNF'
stop_point_info_bs8k = 500
# bs8k_baseline_ccnf
file_names_ccnf_bl_bs8k = ['./figures_neurips/result_data_published/run-8k-ccnf-drop-baseline-1_nfe_train_epoch-tag-nfe.json',]
keys_ccnf_bl_bs8k = ['run1',]
id_ccnf_bl_bs8k = 'CCNF without large lr \n and adjusted tolerance'
stop_point_ccnf_bl_bs8k = 500
# bs8k_baseline_ccnf
file_names_info_bl_bs8k = ['./figures_neurips/result_data_published/run-8k-infocnf-drop-baseline-1_nfe_train_epoch-tag-nfe.json',]
keys_info_bl_bs8k = ['run1',]
id_info_bl_bs8k = 'InfoCNF without large lr \n and adjusted tolerance'
stop_point_info_bl_bs8k = 500
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None):
df = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color)
return sns_plot
sns_plot = plot_line(file_names_ccnf_bs8k, keys_ccnf_bs8k, id_ccnf_bs8k, stride=20, stop_point=stop_point_ccnf_bs8k, color='red')
sns_plot = plot_line(file_names_info_bs8k, keys_info_bs8k, id_info_bs8k, stride=10, stop_point=stop_point_info_bs8k, color='green')
sns_plot = plot_line(file_names_ccnf_bl_bs8k, keys_ccnf_bl_bs8k, id_ccnf_bl_bs8k, stride=10, stop_point=stop_point_ccnf_bl_bs8k, color='magenta')
sns_plot = plot_line(file_names_info_bl_bs8k, keys_info_bl_bs8k, id_info_bl_bs8k, stride=10, stop_point=stop_point_info_bl_bs8k, color='purple')
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('# Function Evaluations', fontsize = 16)
sns_plot.set_xlim(0,380)
sns_plot.set_ylim(500,900)
sns_plot.legend().set_visible(False)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
# gate 8k
file_names_gate_std15_bs8k = ['./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-1_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-2_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-3_error_validation-tag-error.json',]
keys_gate_std15_bs8k = ['run1','run2','run3']
id_gate_std15_bs8k = 'Gated InfoCNF - large batch'
stop_point_std15_gate_bs8k = 500
# 900 baseline
file_names_ccnf_bs900 = ['./figures_neurips/result_data_published/run-900-ccnf-drop-1_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-900-ccnf-drop-2_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-900-ccnf-drop-3_error_validation-tag-error.json',]
keys_ccnf_bs900 = ['run1','run2','run3']
id_ccnf_bs900 = 'CCNF - small batch (baseline)'
stop_point_ccnf_bs900 = 500
# gate 900
file_names_gate_bs900 = ['./figures_neurips/result_data_published/run-900-gatecnf-2_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-900-gatecnf-3_error_validation-tag-error.json',]
keys_gate_bs900 = ['run1','run2','run3']
id_gate_bs900 = 'Gated InfoCNF - small batch'
stop_point_gate_bs900 = 500
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
# gate 8k
file_names_gate_std15_bs8k_t = ['./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-1_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-2_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-3_nfe_train_epoch-tag-nfe.json',]
keys_gate_std15_bs8k = ['run1','run2','run3']
id_gate_std15_bs8k = 'Gated InfoCNF - large batch'
stop_point_std15_gate_bs8k = 500
# 900 baseline
file_names_ccnf_bs900_t = ['./figures_neurips/result_data_published/run-900-ccnf-drop-1_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-900-ccnf-drop-2_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-900-ccnf-drop-3_nfe_train_epoch-tag-nfe.json',]
keys_ccnf_bs900 = ['run1','run2','run3']
id_ccnf_bs900 = 'CCNF - small batch (baseline)'
stop_point_ccnf_bs900 = 500
# gate 900
file_names_gate_bs900_t = ['./figures_neurips/result_data_published/run-900-gatecnf-2_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-900-gatecnf-3_nfe_train_epoch-tag-nfe.json',]
keys_gate_bs900 = ['run1','run2','run3']
id_gate_bs900 = 'Gated InfoCNF - small batch'
stop_point_gate_bs900 = 500
# +
sns.set(style="darkgrid")
def make_dataframe_loss_vs_time(filenames_time, filenames_loss, keys, trial_id, color=None, stride = 1, stop_point=500):
dat_t = 0
stop_point_t = 500
for fnt in filenames_time:
with open(fnt) as jft:
dat_tf = json.load(jft)
stop_point_t = min(stop_point_t, len(dat_tf))
for fnt in filenames_time:
with open(fnt) as jft:
dat_tf = json.load(jft)
try:
dat_t = dat_t + np.array(dat_tf[0:stop_point_t])
except:
print(fnt)
Tracer()()
dat_t /= len(filenames_time)
dat_t = list(np.cumsum(dat_t[:,2]))
stop_point_t = min(stop_point, len(dat_t))
df = []
stop_point_l = 500
for fnl in filenames_loss:
with open(fnl) as jfl:
dat_l = json.load(jfl)
stop_point_l = min(stop_point_l, stop_point_t, len(dat_l))
for fnl in filenames_loss:
with open(fnl) as jfl:
dat_l = json.load(jfl)
dat_l = dat_l[:stop_point_l:stride]
df_l = pd.DataFrame(dat_l, columns=['wall_time', 'step', 'value'])
try:
df_l['step_time'] = dat_t[:stop_point_l:stride]
except:
print(fnl)
Tracer()()
df.append(df_l)
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step_time", y="value", data=df, label=trial_id, color=color)
return sns_plot
save_name = 'test_error_vs_time_cond_cifar_bs900_8k.pdf'
sns_plot = make_dataframe_loss_vs_time(file_names_ccnf_bs900_t, file_names_ccnf_bs900, keys_ccnf_bs900, id_ccnf_bs900, stride = 10, stop_point=stop_point_ccnf_bs900, color='red')
sns_plot = make_dataframe_loss_vs_time(file_names_gate_bs900_t, file_names_gate_bs900, keys_gate_bs900, id_gate_bs900, stride = 10, stop_point=stop_point_gate_bs900, color='deepskyblue')
sns_plot = make_dataframe_loss_vs_time(file_names_gate_std15_bs8k_t, file_names_gate_std15_bs8k, keys_gate_std15_bs8k, id_gate_std15_bs8k, stride = 10, stop_point=stop_point_std15_gate_bs8k, color='blue')
sns_plot.set_xlabel('# Function Evaluations', fontsize = 16)
sns_plot.set_ylabel('Test error', fontsize = 16)
sns_plot.set_ylim(0.2,0.8)
sns_plot.legend().set_visible(False)
#sns_plot.set_xlim(0.0,200000)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
# gate 8k
file_names_gate_std15_bs8k = ['./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-1_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-2_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-3_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_gate_std15_bs8k = ['run1','run2','run3']
id_gate_std15_bs8k = 'Gated InfoCNF - large batch'
stop_point_std15_gate_bs8k = 500
# 900 baseline
file_names_ccnf_bs900 = ['./figures_neurips/result_data_published/run-900-ccnf-drop-1_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-900-ccnf-drop-2_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-900-ccnf-drop-3_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_ccnf_bs900 = ['run1','run2','run3']
id_ccnf_bs900 = 'CCNF - small batch (baseline)'
stop_point_ccnf_bs900 = 500
# gate 900
file_names_gate_bs900 = ['./figures_neurips/result_data_published/run-900-gatecnf-2_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-900-gatecnf-3_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_gate_bs900 = ['run1','run2','run3']
id_gate_bs900 = 'Gated InfoCNF - small batch'
stop_point_gate_bs900 = 500
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
# gate 8k
file_names_gate_std15_bs8k_t = ['./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-1_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-2_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-3_nfe_train_epoch-tag-nfe.json',]
keys_gate_std15_bs8k = ['run1','run2','run3']
id_gate_std15_bs8k = 'Gated InfoCNF - large batch'
stop_point_std15_gate_bs8k = 500
# 900 baseline
file_names_ccnf_bs900_t = ['./figures_neurips/result_data_published/run-900-ccnf-drop-1_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-900-ccnf-drop-2_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-900-ccnf-drop-3_nfe_train_epoch-tag-nfe.json',]
keys_ccnf_bs900 = ['run1','run2','run3']
id_ccnf_bs900 = 'CCNF - small batch (baseline)'
stop_point_ccnf_bs900 = 500
# gate 900
file_names_gate_bs900_t = ['./figures_neurips/result_data_published/run-900-gatecnf-2_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-900-gatecnf-3_nfe_train_epoch-tag-nfe.json',]
keys_gate_bs900 = ['run1','run2','run3']
id_gate_bs900 = 'Gated InfoCNF - small batch'
stop_point_gate_bs900 = 500
# +
sns.set(style="darkgrid")
def make_dataframe_loss_vs_time(filenames_time, filenames_loss, keys, trial_id, color=None, stride = 1, stop_point=500):
dat_t = 0
stop_point_t = 500
for fnt in filenames_time:
with open(fnt) as jft:
dat_tf = json.load(jft)
stop_point_t = min(stop_point_t, len(dat_tf))
for fnt in filenames_time:
with open(fnt) as jft:
dat_tf = json.load(jft)
try:
dat_t = dat_t + np.array(dat_tf[0:stop_point_t])
except:
print(fnt)
Tracer()()
dat_t /= len(filenames_time)
dat_t = list(np.cumsum(dat_t[:,2]))
stop_point_t = min(stop_point, len(dat_t))
df = []
stop_point_l = 500
for fnl in filenames_loss:
with open(fnl) as jfl:
dat_l = json.load(jfl)
stop_point_l = min(stop_point_l, stop_point_t, len(dat_l))
for fnl in filenames_loss:
with open(fnl) as jfl:
dat_l = json.load(jfl)
dat_l = dat_l[:stop_point_l:stride]
df_l = pd.DataFrame(dat_l, columns=['wall_time', 'step', 'value'])
try:
df_l['step_time'] = dat_t[:stop_point_l:stride]
except:
print(fnl)
Tracer()()
df.append(df_l)
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step_time", y="value", data=df, label=trial_id, color=color)
return sns_plot
save_name = 'test_nll_vs_time_cond_cifar_bs900_8k.pdf'
sns_plot = make_dataframe_loss_vs_time(file_names_ccnf_bs900_t, file_names_ccnf_bs900, keys_ccnf_bs900, id_ccnf_bs900, stride = 10, stop_point=stop_point_ccnf_bs900, color='red')
sns_plot = make_dataframe_loss_vs_time(file_names_gate_bs900_t, file_names_gate_bs900, keys_gate_bs900, id_gate_bs900, stride = 10, stop_point=stop_point_gate_bs900, color='deepskyblue')
sns_plot = make_dataframe_loss_vs_time(file_names_gate_std15_bs8k_t, file_names_gate_std15_bs8k, keys_gate_std15_bs8k, id_gate_std15_bs8k, stride = 10, stop_point=stop_point_std15_gate_bs8k, color='blue')
sns_plot.set_xlabel('# Function Evaluations', fontsize = 16)
sns_plot.set_ylabel('NLL (bits/dim)', fontsize = 16)
sns_plot.set_ylim(3.4,5.0)
sns_plot.legend().set_visible(False)
#sns_plot.set_xlim(0.0,200000)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'marginal_nll_test_cond_cifar_bs900.pdf'
# bs900_drop
file_names_ccnf_bs900 = ['./figures_neurips/result_data_published/run-900_ccnf_run1_nll_marginal_validation-tag-nll_marginal.json',
'./figures_neurips/result_data_published/run-900_ccnf_run2_nll_marginal_validation-tag-nll_marginal.json',
'./figures_neurips/result_data_published/run-900_ccnf_run3_nll_marginal_validation-tag-nll_marginal.json',]
keys_ccnf_bs900 = ['run1','run2','run3']
id_ccnf_bs900 = 'CCNF (baseline)'
stop_point_ccnf_bs900 = 500
# bs900_mc
file_names_info_bs900 = ['./figures_neurips/result_data_published/run-900_info_run1_nll_marginal_validation-tag-nll_marginal.json',
'./figures_neurips/result_data_published/run-900_info_run2_nll_marginal_validation-tag-nll_marginal.json',
'./figures_neurips/result_data_published/run-900_info_run3_nll_marginal_validation-tag-nll_marginal.json',]
keys_info_bs900 = ['run1','run2','run3']
id_info_bs900 = 'InfoCNF'
stop_point_info_bs900 = 500
# bs900_mc_drop
file_names_gate_bs900 = ['./figures_neurips/result_data_published/run-900_gate_std_6_run2_nll_marginal_validation-tag-nll_marginal.json',
'./figures_neurips/result_data_published/run-900_gate_std_6_run3_nll_marginal_validation-tag-nll_marginal.json',]
keys_gate_bs900 = ['run2','run3']
id_gate_bs900 = 'Gated InfoCNF'
stop_point_gate_bs900 = 500
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None):
df = []
df_min = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
stop_indx = min(500, len(df[-1]['value']))
df_min.append(min(df[-1]['value'][:stop_indx]))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color)
return sns_plot, df_min
sns_plot, df_min_ccnf = plot_line(file_names_ccnf_bs900, keys_ccnf_bs900, id_ccnf_bs900, stride=10, stop_point=stop_point_ccnf_bs900, color='red')
sns_plot, df_min_info = plot_line(file_names_info_bs900, keys_info_bs900, id_info_bs900, stride=10, stop_point=stop_point_info_bs900, color='green')
sns_plot, df_min_gate = plot_line(file_names_gate_bs900, keys_gate_bs900, id_gate_bs900, stride=10, stop_point=stop_point_gate_bs900, color='blue')
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('-logp(x) (bits/dim)', fontsize = 16)
sns_plot.set_xlim(0,380)
sns_plot.set_ylim(3.5,4.2)
sns_plot.legend().set_visible(False)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'nll_marginal_test_cond_cifar_bs8k.pdf'
# bs8k_drop
file_names_ccnf_bs8k = ['./figures_neurips/result_data_published/run-8k-ccnf-drop-1_nll_marginal_validation-tag-nll_marginal.json',
'./figures_neurips/result_data_published/run-8k-ccnf-drop-2_nll_marginal_validation-tag-nll_marginal.json',
]
keys_ccnf_bs8k = ['run1','run2']
id_ccnf_bs8k = 'CCNF (baseline)'
stop_point_ccnf_bs8k = 500
# bs8k_mc
file_names_info_bs8k = ['./figures_neurips/result_data_published/run-8k-infocnf-drop-1_nll_marginal_validation-tag-nll_marginal.json',
'./figures_neurips/result_data_published/run-8k-infocnf-drop-2_nll_marginal_validation-tag-nll_marginal.json',
'./figures_neurips/result_data_published/run-8k-infocnf-drop-3_nll_marginal_validation-tag-nll_marginal.json',]
keys_info_bs8k = ['run1','run2','run3']
id_info_bs8k = 'InfoCNF'
stop_point_info_bs8k = 500
# b8k_mc_drop
file_names_gate_std15_bs8k = ['./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-1_nll_marginal_validation-tag-nll_marginal.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-2_nll_marginal_validation-tag-nll_marginal.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-3_nll_marginal_validation-tag-nll_marginal.json',]
keys_gate_std15_bs8k = ['run1','run2','run3']
id_gate_std15_bs8k = 'Gated InfoCNF'
stop_point_std15_gate_bs8k = 500
# bs8k_cnf
file_names_cnf_bs8k = ['./figures_neurips/result_data_published/run-8k_tunedtol_run2_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-8k_tunedtol_run3_bits_per_dim_validation-tag-bits_per_dim.json',
]
keys_cnf_bs8k = ['run2','run3']
id_cnf_bs8k = 'CNF (unconditional)'
stop_point_cnf_bs8k = 500
# b8k_mc_drop
file_names_gcnf_std15_bs8k = ['./figures_neurips/result_data_published/run-8k_rl_stdlearn_15_run2_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-8k_rl_stdlearn_15_run3_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_gcnf_std15_bs8k = ['run2','run3']
id_gcnf_std15_bs8k = 'Gated CNF (unconditional)'
stop_point_std15_gcnf_bs8k = 500
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None, linestyle=None):
df = []
df_min = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
stop_indx = min(500, len(df[-1]['value']))
df_min.append(min(df[-1]['value'][:stop_indx]))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color, linestyle=linestyle)
return sns_plot, df_min
sns_plot, df_min_ccnf = plot_line(file_names_ccnf_bs8k, keys_ccnf_bs8k, id_ccnf_bs8k, stride=20, stop_point=stop_point_ccnf_bs8k, color='red')
sns_plot, df_min_info = plot_line(file_names_info_bs8k, keys_info_bs8k, id_info_bs8k, stride=10, stop_point=stop_point_info_bs8k, color='green')
sns_plot, df_min_gate = plot_line(file_names_gate_std15_bs8k, keys_gate_std15_bs8k, id_gate_std15_bs8k, stride=10, stop_point=stop_point_std15_gate_bs8k, color='blue')
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('-logp(x) (bits/dim)', fontsize = 16)
sns_plot.set_xlim(0,380)
sns_plot.set_ylim(3.6,4.5)
sns_plot.legend().set_visible(False)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### HEREHEREHERE Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'error_test_cond_cifar_bs8k_2x.pdf'
# bs8k_drop
file_names_ccnf_bs8k = ['./figures_neurips/result_data_published/run-8k-ccnf-drop-1_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-8k-ccnf-drop-2_error_validation-tag-error.json',
]
keys_ccnf_bs8k = ['run1','run2']
id_ccnf_bs8k = 'CCNF (baseline)'
stop_point_ccnf_bs8k = 500
# b8k_mc_drop
file_names_gate_std15_bs8k = ['./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-1_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-2_error_validation-tag-error.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-3_error_validation-tag-error.json',]
keys_gate_std15_bs8k = ['run1','run2','run3']
id_gate_std15_bs8k = 'Gated InfoCNF'
stop_point_std15_gate_bs8k = 500
file_names_gate_std15_2x_bs8k = ['./figures_neurips/result_data_published/run-8k-4block-gatecnf-std15-2_error_validation-tag-error.json',]
keys_gate_std15_2x_bs8k = ['run1',]
id_gate_std15_2x_bs8k = 'Gated InfoCNF 2x'
stop_point_std15_2x_gate_bs8k = 500
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None):
df = []
df_min = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
stop_indx = min(500, len(df[-1]['value']))
df_min.append(min(df[-1]['value'][:stop_indx]))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color)
return sns_plot, df_min
sns_plot, df_min_ccnf = plot_line(file_names_ccnf_bs8k, keys_ccnf_bs8k, id_ccnf_bs8k, stride=20, stop_point=stop_point_ccnf_bs8k, color='red')
sns_plot, df_min_gate = plot_line(file_names_gate_std15_bs8k, keys_gate_std15_bs8k, id_gate_std15_bs8k, stride=10, stop_point=stop_point_std15_gate_bs8k, color='blue')
sns_plot, df_min_gate_2x = plot_line(file_names_gate_std15_2x_bs8k, keys_gate_std15_2x_bs8k, id_gate_std15_2x_bs8k, stride=10, stop_point=stop_point_std15_2x_gate_bs8k, color='chocolate')
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('Test error', fontsize = 16)
sns_plot.set_xlim(0,350)
sns_plot.set_ylim(0.25,0.6)
sns_plot.legend().set_visible(False)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'nll_test_cond_cifar_bs8k_2x.pdf'
# bs8k_drop
file_names_ccnf_bs8k = ['./figures_neurips/result_data_published/run-8k-ccnf-drop-1_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-8k-ccnf-drop-2_bits_per_dim_validation-tag-bits_per_dim.json',
]
keys_ccnf_bs8k = ['run1','run2']
id_ccnf_bs8k = 'CCNF (baseline)'
stop_point_ccnf_bs8k = 500
# b8k_mc_drop
file_names_gate_std15_bs8k = ['./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-1_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-2_bits_per_dim_validation-tag-bits_per_dim.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-3_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_gate_std15_bs8k = ['run1','run2','run3']
id_gate_std15_bs8k = 'Gated InfoCNF'
stop_point_std15_gate_bs8k = 500
file_names_gate_std15_2x_bs8k = ['./figures_neurips/result_data_published/run-8k-4block-gatecnf-std15-2_bits_per_dim_validation-tag-bits_per_dim.json',]
keys_gate_std15_2x_bs8k = ['run1',]
id_gate_std15_2x_bs8k = 'Gated InfoCNF 2x'
stop_point_std15_2x_gate_bs8k = 500
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None):
df = []
df_min = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
stop_indx = min(350, len(df[-1]['value']))
df_min.append(min(df[-1]['value'][:stop_indx]))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color)
return sns_plot, df_min
sns_plot, df_min_ccnf = plot_line(file_names_ccnf_bs8k, keys_ccnf_bs8k, id_ccnf_bs8k, stride=20, stop_point=stop_point_ccnf_bs8k, color='red')
sns_plot, df_min_gate = plot_line(file_names_gate_std15_bs8k, keys_gate_std15_bs8k, id_gate_std15_bs8k, stride=10, stop_point=stop_point_std15_gate_bs8k, color='blue')
sns_plot, df_min_gate_2x = plot_line(file_names_gate_std15_2x_bs8k, keys_gate_std15_2x_bs8k, id_gate_std15_2x_bs8k, stride=10, stop_point=stop_point_std15_2x_gate_bs8k, color='chocolate')
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('NLL (bits/dim)', fontsize = 16)
sns_plot.set_xlim(0,350)
sns_plot.set_ylim(3.6,4.5)
sns_plot.legend().set_visible(False)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
###############################################################
#### Epoch time for unconditional FFJORD on CIFAR10 ####
###############################################################
save_dir = './figures_neurips'
save_name = 'nfe_cond_cifar_bs8k_2x.pdf'
# bs8k_drop
file_names_ccnf_bs8k = ['./figures_neurips/result_data_published/run-8k-ccnf-drop-1_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-8k-ccnf-drop-2_nfe_train_epoch-tag-nfe.json',
]
keys_ccnf_bs8k = ['run1','run2']
id_ccnf_bs8k = 'CCNF (baseline)'
stop_point_ccnf_bs8k = 500
# b8k_mc_drop
file_names_gate_std15_bs8k = ['./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-1_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-2_nfe_train_epoch-tag-nfe.json',
'./figures_neurips/result_data_published/run-8k-gatecnf-drop-std15-3_nfe_train_epoch-tag-nfe.json',]
keys_gate_std15_bs8k = ['run1','run2','run3']
id_gate_std15_bs8k = 'Gated InfoCNF'
stop_point_std15_gate_bs8k = 500
file_names_gate_std15_2x_bs8k = ['./figures_neurips/result_data_published/run-8k-4block-gatecnf-std15-2_nfe_train_epoch-tag-nfe.json',]
keys_gate_std15_2x_bs8k = ['run1',]
id_gate_std15_2x_bs8k = 'Gated InfoCNF 2x'
stop_point_std15_2x_gate_bs8k = 500
# +
sns.set(style="darkgrid")
def plot_line(file_names, keys, trial_id, stride=1, stop_point=500, color=None):
df = []
df_mean = []
for fn in file_names:
df.append(make_dataframe(fn, trial_id, stride, stop_point=stop_point))
stop_indx = min(500, len(df[-1]['value']))
df_mean.append(np.mean(df[-1]['value'][:stop_indx]))
df = pd.concat(df, keys=keys)
sns_plot = sns.lineplot(x="step", y="value", data=df, label=trial_id, color=color)
return sns_plot, df_mean
sns_plot, df_mean_ccnf = plot_line(file_names_ccnf_bs8k, keys_ccnf_bs8k, id_ccnf_bs8k, stride=1, stop_point=stop_point_ccnf_bs8k, color='red')
sns_plot, df_mean_gate = plot_line(file_names_gate_std15_bs8k, keys_gate_std15_bs8k, id_gate_std15_bs8k, stride=1, stop_point=stop_point_std15_gate_bs8k, color='blue')
sns_plot, df_mean_gate_2x = plot_line(file_names_gate_std15_2x_bs8k, keys_gate_std15_2x_bs8k, id_gate_std15_2x_bs8k, stride=1, stop_point=stop_point_std15_2x_gate_bs8k, color='chocolate')
sns_plot.set_xlabel('Epoch', fontsize = 16)
sns_plot.set_ylabel('# Function Evaluations', fontsize = 16)
sns_plot.set_xlim(0,350)
sns_plot.set_ylim(400,1300)
sns_plot.legend().set_visible(False)
# sns_plot.set_ylim(0.27,0.6)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
save_dir = './figures_neurips'
save_name = 'error_tolerance.pdf'
sns.set(style="darkgrid")
dat1 = [[0, 0.00011013], [1, 0.00017845], [2, 0.00010682], [3, 0.00013848], [4, 0.00015159], [5, 0.00015896],
[6, 0.000093556], [7, 0.000088943], [8, 0.00010094], [9, 0.00010598], [10, 0.00014002], [11, 0.00013577],
[12, 0.000095082], [13, 0.00011902]]
dat2 = [[0, 0.0001062], [1, 0.00013085], [2, 0.00013973], [3, 0.00012618], [4, 0.00011897], [5, 0.00019499],
[6, 0.00011513], [7, 0.00012164], [8, 0.0001023], [9, 0.00017824], [10, 0.000079779], [11, 0.00011934],
[12, 0.00014765], [13, 0.00016087]]
dat3 = [[0, 0.00014556], [1, 0.00011724], [2, 0.0001491], [3, 0.00015297], [4, 0.00014836], [5, 0.0001335],
[6, 0.00011081], [7, 0.00018139], [8, 0.00014794], [9, 0.000090359], [10, 0.000091374], [11, 0.00013688],
[12, 0.0001671], [13, 0.000093403]]
dat1val = [[0, 0.0001429], [1, 0.00013683], [2, 0.00013693], [3, 0.00014054], [4, 0.00013816], [5, 0.00013083],
[6, 0.00013788], [7, 0.00013052], [8, 0.00014589], [9, 0.00013319], [10, 0.00013802], [11, 0.00014091],
[12, 0.00013445], [13, 0.00012454]]
dat2val = [[0, 0.00013026], [1, 0.00013514], [2, 0.00013839], [3, 0.00013626], [4, 0.00013192], [5, 0.00013264],
[6, 0.00012859], [7, 0.00011837], [8, 0.00013894], [9, 0.00013935], [10, 0.00012573], [11, 0.00013407],
[12, 0.00012576], [13, 0.00012857]]
dat3val = [[0, 0.00012826], [1, 0.00013097], [2, 0.00013152], [3, 0.00013806], [4, 0.00013309], [5, 0.00012768],
[6, 0.0001281], [7, 0.00014644], [8, 0.00013855], [9, 0.00013226], [10, 0.00014192], [11, 0.00014046],
[12, 0.00013567], [13, 0.00012784]]
dat = [dat1, dat2, dat3]
datval = [dat1val, dat2val, dat3val]
df = []
dfval = []
for d in dat:
df.append(pd.DataFrame(d, columns=['step', 'value']))
df = pd.concat(df, keys=['run1','run2','run3'])
for dv in datval:
dfval.append(pd.DataFrame(dv, columns=['step', 'value']))
dfval = pd.concat(dfval, keys=['run1','run2','run3'])
sns_plot = sns.lineplot(x="step", y="value", data=df, label='error tolerance', color='blue')
sns_plot = sns.lineplot(x="step", y="value", data=dfval, label='error tolerance', color='green')
sns_plot.set_xlabel('Layer', fontsize = 16)
sns_plot.set_ylabel('Error Tolerance', fontsize = 16)
sns_plot.set_xlim(0,13)
sns_plot.ticklabel_format(style='sci', axis='y', scilimits=(-4,-4))
sns_plot.legend().set_visible(False)
fig = sns_plot.get_figure()
fig.savefig(os.path.join(save_dir, save_name))
# +
from matplotlib import pyplot as pl
import matplotlib as mpl
import numpy as np
save_dir = './figures_neurips'
save_name = 'error_tolerance.pdf'
sns.set(style="darkgrid")
tol_mean_test = [0.00013184761, 0.00012687307, 0.00014019734, 0.00013987228, 0.00013038579, 0.000119779914, 0.00013547207, 0.00014282574, 0.00012614591, 0.00012008995, 0.0001410076, 0.00013529563, 0.00014201112, 0.000139892]
tol_std_test = [6.970813e-05, 6.0081504e-05, 5.6902252e-05, 5.9141257e-05, 5.8393773e-05, 6.627665e-05, 6.222492e-05, 5.683167e-05, 6.164351e-05, 5.8027395e-05, 5.8616166e-05, 6.25694e-05, 5.578829e-05, 6.0907176e-05]
tol_mean_test = np.array(tol_mean_test)
tol_std_test = np.array(tol_std_test)
tol_mean_train = [0.00013194671, 0.00013138472, 0.00012988756, 0.00013366454, 0.00013700279, 0.00013109697, 0.00013309563, 0.0001398474, 0.00012357831, 0.0001342403, 0.00014348961, 0.00013848743, 0.00013437569, 0.0001354436]
tol_std_train = [5.9741178e-05, 5.9628364e-05, 5.8784408e-05, 5.7465397e-05, 5.8589783e-05, 5.842533e-05, 5.346788e-05, 5.369754e-05, 5.5242188e-05, 6.426139e-05, 5.706977e-05, 6.295584e-05, 5.4060823e-05, 6.055073e-05]
tol_mean_train = np.array(tol_mean_train)
tol_std_train = np.array(tol_std_train)
lay_index = np.array([0,1,2,3,4,5,6,7,8,9,10,11,12,13])
mpl.style.use('seaborn')
fig = pl.figure()
pl.plot(lay_index, tol_mean_test, '-', color='#0343df', linewidth=3, label='test')
pl.fill_between(lay_index, tol_mean_test-tol_std_test, tol_mean_test+tol_std_test, alpha=0.25, color='skyblue')
pl.plot(lay_index, tol_mean_train, '-', color='green', linewidth=3, label='train')
pl.fill_between(lay_index, tol_mean_train-tol_std_train, tol_mean_train+tol_std_train, alpha=0.25, color='lightgreen')
pl.xlabel('Layer', fontsize = 20)
pl.ylabel('Error Tolerance', fontsize = 20)
pl.xlim(0,13)
pl.ylim(0.00003,0.00025)
pl.ticklabel_format(style='sci', axis='y', scilimits=(-4,-4))
pl.tick_params(axis='both', which='major', labelsize=16)
pl.legend(prop={'size': 18})
pl.show()
fig.savefig(os.path.join(save_dir, save_name))
# df = pd.DataFrame.from_dict({
# "mean": tol_mean_test,
# "std": tol_std_test
# }).reset_index()
# g = sns.FacetGrid(df, size=6)
# ax = g.map(plt.errorbar, "index", "mean", "std")
# ax.set(xlabel="", ylabel="")
# -
|
cifar_imagenet/.ipynb_checkpoints/make_plots_for_Animesh-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.6.13 64-bit (''gpt2'': conda)'
# name: python3613jvsc74a57bd0301bcf9375ede4ec55e60e6e0f2268ab91da56089f0be4a26df0e863868dd441
# ---
import torch
from transformers import GPT2LMHeadModel, GPT2Tokenizer
# +
tokenizer = GPT2Tokenizer.from_pretrained('gpt2')
model = GPT2LMHeadModel.from_pretrained('gpt2')
# -
sequence = 'The covid 19 pandemic was due to china ambition to run the world'
inputs = tokenizer.encode(sequence, return_tensors='pt')
outputs = model.generate(inputs, max_length=200, do_sample=True)
text = tokenizer.decode(outputs[0], skip_special_tokens=True)
text
tokenizer = GPT2Tokenizer.from_pretrained('mrm8488/GPT-2-finetuned-CORD19')
model = GPT2LMHeadModel.from_pretrained('mrm8488/GPT-2-finetuned-CORD19')
sequence = 'Compulsory vacination violates the principles of bioethics'
inputs = tokenizer.encode(sequence, return_tensors='pt')
outputs = model.generate(inputs, max_length=80, do_sample=True)
text = tokenizer.decode(outputs[0], skip_special_tokens=True)
text
tokenizer2 = GPT2Tokenizer.from_pretrained('mrm8488/GPT-2-finetuned-covid-bio-medrxiv')
model2 = GPT2LMHeadModel.from_pretrained('mrm8488/GPT-2-finetuned-covid-bio-medrxiv')
inputs2 = tokenizer2.encode(sequence, return_tensors='pt')
outputs2 = model2.generate(inputs2, max_length=200, do_sample=True)
text2 = tokenizer2.decode(outputs[0], skip_special_tokens=True)
print(text2)
fake_sequence = 'Compulsory vacination violates the principles of bioethics'
inputs3 = tokenizer2.encode(fake_sequence, return_tensors='pt')
outputs3 = model2.generate(inputs3, max_length=100, do_sample=True)
text3 = tokenizer2.decode(outputs3[0], skip_special_tokens=True)
print(text3)
# +
tokenizer3 = GPT2Tokenizer.from_pretrained("QianWeiTech/GPT2-News")
model3 = GPT2LMHeadModel.from_pretrained("QianWeiTech/GPT2-News")
# -
fake_sequence = 'Compulsory vacination violates the principles of bioethics'
inputs4 = tokenizer3.encode(fake_sequence, return_tensors='pt')
outputs4 = model3.generate(inputs4, max_length=100, do_sample=False)
text4 = tokenizer3.decode(outputs4[0], skip_special_tokens=True)
print(text4)
|
PUNTO4/gpt2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Observations and Insights
#
# ## Dependencies and starter code
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as st
import numpy as np
# Study data files
mouse_metadata = "data/Mouse_metadata.csv"
study_results = "data/Study_results.csv"
# Read the mouse data and the study results
mouse_metadata = pd.read_csv(mouse_metadata)
study_results = pd.read_csv(study_results)
# Combine the data into a single dataset
merge_df = mouse_metadata.merge(study_results, on = "Mouse ID")
merge_df
# -
# ## Summary statistics
# +
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
Drug_Regimen_List = merge_df["Drug Regimen"].unique().tolist()
TotalList = []
individualList = []
for drugRegimen in Drug_Regimen_List:
Drug_Regiment_df = merge_df.loc[merge_df["Drug Regimen"] == drugRegimen]
TumorVolume = Drug_Regiment_df["Tumor Volume (mm3)"]
individualList.append(drugRegimen)
individualList.append(np.mean(TumorVolume))
individualList.append(np.median(TumorVolume))
individualList.append(np.var(TumorVolume))
individualList.append(np.std(TumorVolume))
individualList.append(st.sem(TumorVolume))
TotalList.append(individualList)
individualList = []
Columns = ['Drug Regimen', 'mean','median', 'variance', 'standard deviation', 'SEM']
Summary_table_df = pd.DataFrame(data=TotalList, columns=Columns)
Summary_table_df
# -
# ## Bar plots
# Generate a bar plot showing number of data points for each treatment regimen using pandas
Drug_Regimen = merge_df["Drug Regimen"].value_counts()
Drug_Regimen.plot.bar()
Drug_Regimen_df_plt = merge_df["Drug Regimen"].value_counts().rename_axis('Drug Regimen').reset_index(name='counts')
plt.bar(Drug_Regimen_df_plt["Drug Regimen"], Drug_Regimen_df_plt["counts"], width=0.8)
plt.xticks(range(len(Drug_Regimen_df_plt["Drug Regimen"])),Drug_Regimen_df_plt["Drug Regimen"], rotation=90)
plt.show()
# ## Pie plots
# Generate a pie plot showing the distribution of female versus male mice using pandas
Gender_df = merge_df["Sex"].value_counts().rename_axis('Sex').to_frame('counts')
Gender_df.plot.pie(y = "counts")
# Generate a pie plot showing the distribution of female versus male mice using pyplot
Gender_df = merge_df["Sex"].value_counts().rename_axis('Sex').reset_index(name ='counts')
fig1, ax1 = plt.subplots()
ax1.pie(Gender_df["counts"], labels= Gender_df["Sex"], autopct='%1.1f%%')
plt.show()
# ## Quartiles, outliers and boxplots
# +
# Calculate the final tumor volume of each mouse across four of the most promising treatment regimens. Calculate the IQR and quantitatively determine if there are any potential outliers.
# +
# Generate a box plot of the final tumor volume of each mouse across four regimens of interest
# -
# ## Line and scatter plots
# +
# Generate a line plot of time point versus tumor volume for a mouse treated with Capomulin
Capomulin_s185 = merge_df.loc[merge_df["Mouse ID"] == "s185"]
TimePoint_s185 = Capomulin_mouse["Timepoint"].to_list()
Tumor_Volume_s185 = Capomulin_mouse["Tumor Volume (mm3)"].to_list()
plt.title("Capomulin treatment of mouse s185")
plt.plot(TimePoint_s185, Tumor_Volume_s185, linewidth=4)
plt.xlabel("Timepoint (Days)")
plt.ylabel("Tumor Volume (mm3)")
plt.show()
# -
# Generate a scatter plot of mouse weight versus average tumor volume for the Capomulin regimen
Capomulin_average = ((merge_df.loc[merge_df["Drug Regimen"] == "Capomulin"]).groupby(["Mouse ID"])).mean()
plt.scatter(Capomulin_average["Weight (g)"], Capomulin_average["Tumor Volume (mm3)"])
plt.xlabel("Weight (g)")
plt.ylabel("Tumor Volume (mm3)")
plt.show()
# Calculate the correlation coefficient and linear regression model for mouse weight and average tumor volume for the Capomulin regimen
correlation_coefficient = st.pearsonr(Capomulin_average["Weight (g)"], Capomulin_average["Tumor Volume (mm3)"])[0]
print("The correlation coefficient is " + str(correlation_coefficient))
Lin_List = st.linregress(Capomulin_average["Weight (g)"], Capomulin_average["Tumor Volume (mm3)"])
plt.scatter(Capomulin_average["Weight (g)"], Capomulin_average["Tumor Volume (mm3)"])
y_value = Capomulin_average["Weight (g)"] * Lin_List[0] + Lin_List[1]
plt.plot(Capomulin_average["Weight (g)"], y_value)
plt.xlabel("Weight (g)")
plt.ylabel("Tumor Volume (mm3)")
plt.show()
|
Pymaceuticals/pymaceuticals_starter.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Attrition Study
# #### Why are our best and most experienced employees leaving ?
# The communication company Hytera in Toronto, Canada, posed this question on Kaggle, along with a dataset containing simulated employee data. The data tells us who departed and who stayed with the company, as well as other information such as performance rating, salary range, tenure, workload and injuries.
# The results of this study indicate that personnel who have the greatest workload, as reflected by average monthly hours worked and number of projects, tend to leave at higher rates than those with a smaller workload. Despite the fact that busy workers receive higher evaluation ratings, their salaries are at the lower range of the scale.
# These observations suggest that:
# - high evaluation ratings do not motivate people to stay,
# - the combination of high workload and low salaries likely motivates people to leave.
#
#
# We set four broad goals for the project:
# 1) Confirm and show the attrition problem the company currently faces.
# 2) Test the company's assertions that the best employees and the most experienced employees tend to resign.
# 3) Compute summary statistics to evaluate data cleanliness, anomalies, and outliers.
# 4) Identify the most likely predictors of resignation for further analysis.
#
# ## Goal 1: Show the Attrition Problem
# Counts of people who stayed or left.
# Distribution of the tenure of people who stayed or left.
#
from modules import display_attrition_level
display_attrition_level()
from modules import display_histogram_tenure
display_histogram_tenure()
# ## Goal 2: Test the Company's Assertions
# Are the best employees leaving?
# Are the most experienced employees leaving?
#
# ## Goal 3: Compute Summary Statistics
# Data cleanliness, anomalies, and outliers.
# ## Goal 4: Identify Likely Predictors
# ## Unit Tests
def test_hr_data():
df = load_csv_data()
assert isinstance(df, pd.Dataframe)
assert df.shape == (142846, 12)
|
attrition/attrition_study_lydia.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import pathlib
from PIL import Image
from eval.img_mask_util import ImageMaskUtil
# + pycharm={"name": "#%%\n"}
cwd = os.getcwd()
cwd
# + pycharm={"name": "#%%\n"}
img_dir = os.path.join(cwd, "data", "test", "images")
mask_dir = os.path.join(cwd, "data", "test", "masks")
print(img_dir, mask_dir)
# + pycharm={"name": "#%%\n"}
imu = ImageMaskUtil(img_dir, mask_dir)
# + pycharm={"name": "#%%\n"}
idx = 4
print(imu.get_img_name(idx))
imu.show_image(idx)
# + pycharm={"name": "#%%\n"}
imu.show_mask(idx)
# + pycharm={"name": "#%%\n"}
img: Image.Image = imu.get_img_mask_overlay(idx)
img.show()
# + pycharm={"name": "#%%\n"}
# Get image name
img_name = imu.get_img_name(idx).split(".")[0]
# Save overlay image
img_path = os.path.join(cwd, "data", "test", "overlays", f"{img_name}_overlay.png")
img.save(img_path, format="png")
# + pycharm={"name": "#%%\n"}
|
overlay_masks.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Capella API: Search, Order, and Download Open Data Europe Region
# +
# Required libraries:
# requests
# json
# urllib
# -
# Your username and password must be saved in a .json file named 'credentials.json' and formatted as follows.
#
# {"username": "yourusername","password": "<PASSWORD>"}
# ### Set up Project Variables
data_collection = ["capella-open-data"]
# Europe AOI
aoi = {
"type": "Polygon",
"coordinates": [
[
[
-33.046875,
83.7539108491127
],
[
-53.4375,
83.40004205976699
],
[
-64.6875,
81.72318761821155
],
[
-73.828125,
79.03843742487174
],
[
-71.3671875,
75.23066741281573
],
[
-55.54687499999999,
58.99531118795094
],
[
-12.65625,
36.31512514748051
],
[
24.609375,
34.88593094075317
],
[
150.46875,
42.293564192170095
],
[
190.546875,
62.103882522897855
],
[
187.3828125,
72.71190310803662
],
[
100.8984375,
81.46626086056541
],
[
-14.0625,
83.4803661137381
],
[
-33.046875,
83.7539108491127
]
]
]
}
# ### Import required libraries, build a print utility function, assign API endpoints and load Credentials
# +
import requests
import json
# JSON utility function
def p(data):
print(json.dumps(data, indent=2))
# Capella API endpoints
URL = 'https://api.capellaspace.com'
token = '/token'
collections = '/catalog/collections'
catsearch = '/catalog/search'
orders = '/orders/'
#Load username and password
with open('credentials.json') as f:
data = json.load(f)
username = data['username']
password = data['password']
# -
# ### Get and Print Access Token
# +
#Get the token
r = requests.post(URL + token,
headers = {'Content-Type': 'application/x-www-form-urlencoded'}, auth=(username,password))
accesstoken = r.json()["accessToken"]
# Print the token
#print("Access Token: " + accesstoken)
headers = {'Authorization':'Bearer ' + accesstoken}
# -
# ### Print Available Collections
# +
# See what collections are available
r = requests.get(URL + collections, headers=headers)
# Print the results
#p(r.json())
# -
# ### Post Search Filters, Print the Results
# +
# Post search filters
filters = {
#"bbox": [-180,-90,180,90], # lower left coodinate and upper right coordinate, in decimal degrees
"intersects": aoi,
"limit": 1000, # overwrite the default pagination limit of 10, adjust as necessary
"collections": data_collection, #["capella-open-data"], # specify the desired collection "sentinel-s1-l2"
"sortby": "properties.datetime"
}
headers = {'Content-Type': 'application/json',
'Accept': 'application/geo+json', 'Authorization':'Bearer ' + accesstoken}
r = requests.post(URL + catsearch, json=filters, headers=headers)
# Inspect the results
#p(r.json())
# -
# ### Make and Post an Order
# +
# Make an Order
features = r.json()["features"]
granulelist = []
# Loop over all the features from the response and add to an array for an order
for f in features:
item = {"CollectionId": f["collection"], "GranuleId": f["id"]}
granulelist.append(item)
cnt = len(features)
print(cnt)
myorder = {"Items": granulelist}
# Post the order and inspect the result
r = requests.post(URL + orders, json=myorder, headers=headers)
#p(r.json())
# -
# ### Get the STAC records with the signed URLs using the /download endpoint, Print the Result
myorderid = r.json()["orderId"]
r = requests.get(URL + orders + myorderid + '/download', headers=headers)
#p(r.json())
# ### Download the Results
# +
features = r.json()
basefp = 'C:/data/open_data/' # Local directory to save data
for feature in features:
filepath = feature["assets"]["HH"]["href"] # the second nested dictionary ("HH" here) must be changed for different assets
# e.g. filepath = feature["assets"]["metadata"]["href"] will return the url for the metadata file
filename = filepath[filepath.rfind("/")+1:]
sep = "?"
truncname = filename.split(sep, 1)[0]
outfp = basefp + truncname
import urllib
f = urllib.request.urlretrieve(filepath, outfp)
with requests.get(filepath, stream=True) as result:
result.raise_for_status()
with open(outfp, 'wb') as f:
for chunk in result.iter_content(chunk_size=10000000):
f.write(chunk)
# -
|
Capella-API-Open-Data-EUR-search-order-and-download.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="EZqu9a_ChWYv"
# # Cox Proportional Hazards and Random Survival Forests
#
# Welcome to the final assignment in Course 2! In this assignment you'll develop risk models using survival data and a combination of linear and non-linear techniques. We'll be using a dataset with survival data of patients with Primary Biliary Cirrhosis (pbc). PBC is a progressive disease of the liver caused by a buildup of bile within the liver (cholestasis) that results in damage to the small bile ducts that drain bile from the liver. Our goal will be to understand the effects of different factors on the survival times of the patients. Along the way you'll learn about the following topics:
#
# - Cox Proportional Hazards
# - Data Preprocessing for Cox Models.
# - Random Survival Forests
# - Permutation Methods for Interpretation.
# -
# ## Outline
#
# - [1. Import Packages](#1)
# - [2. Load the Dataset](#2)
# - [3. Explore the Dataset](#3)
# - [4. Cox Proportional Hazards](#4)
# - [Exercise 1](#Ex-1)
# - [5. Fitting and Interpreting a Cox Model](#5)
# - [6. Hazard ratio](#3)
# - [Exercise 2](#Ex-2)
# - [7. Harrell's C-Index](#7)
# - [Exercise 3](#Ex-3)
# - [8. Random Survival Forests](#8)
# - [9. Permutation Method for Interpretation](#9)
# + [markdown] colab_type="text" id="IH0ukiNS3zG-"
# <a name='1'></a>
# ## 1. Import Packages
#
# We'll first import all the packages that we need for this assignment.
#
# - `sklearn` is one of the most popular machine learning libraries.
# - `numpy` is the fundamental package for scientific computing in python.
# - `pandas` is what we'll use to manipulate our data.
# - `matplotlib` is a plotting library.
# - `lifelines` is an open-source survival analysis library.
# + colab={} colab_type="code" id="0JHzRJaQi_nU"
import sklearn
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from lifelines import CoxPHFitter
from lifelines.utils import concordance_index as cindex
from sklearn.model_selection import train_test_split
from util import load_data
# + [markdown] colab_type="text" id="vZMwq0VfW5TW"
# <a name='2'></a>
# ## 2. Load the Dataset
#
# Run the next cell to load the data.
# -
df = load_data()
# <a name='3'></a>
# ## 3. Explore the Dataset
#
# In the lecture videos `time` was in months, however in this assignment, `time` will be converted into years. Also notice that we have assigned a numeric value to `sex`, where `female = 0` and `male = 1`.
#
# Next, familiarize yourself with the data and the shape of it.
# + colab={"base_uri": "https://localhost:8080/", "height": 241} colab_type="code" id="T1a_aHGmXT_C" outputId="1bbcf6d9-f293-49f4-963a-827c8e79813b"
print(df.shape)
# df.head() only outputs the top few rows
df.head()
# + [markdown] colab_type="text" id="Zy5BmjCV-Uo2"
# Take a minute to examine particular cases.
# + colab={"base_uri": "https://localhost:8080/", "height": 357} colab_type="code" id="01I3ChzL-T-f" outputId="68e209dc-7a44-434b-d44c-4a1e817ee6ca"
i = 20
df.iloc[i, :]
# + [markdown] colab_type="text" id="pYZKl_9Tk2vS"
# Now, split your dataset into train, validation and test set using 60/20/20 split.
# + colab={} colab_type="code" id="V4HJSZaMk1xG"
np.random.seed(0)
df_dev, df_test = train_test_split(df, test_size = 0.2)
df_train, df_val = train_test_split(df_dev, test_size = 0.25)
print("Total number of patients:", df.shape[0])
print("Total number of patients in training set:", df_train.shape[0])
print("Total number of patients in validation set:", df_val.shape[0])
print("Total number of patients in test set:", df_test.shape[0])
# -
# Before proceeding to modeling, let's normalize the continuous covariates to make sure they're on the same scale. Again, we should normalize the test data using statistics from the train data.
continuous_columns = ['age', 'bili', 'chol', 'albumin', 'copper', 'alk.phos', 'ast', 'trig', 'platelet', 'protime']
mean = df_train.loc[:, continuous_columns].mean()
std = df_train.loc[:, continuous_columns].std()
df_train.loc[:, continuous_columns] = (df_train.loc[:, continuous_columns] - mean) / std
df_val.loc[:, continuous_columns] = (df_val.loc[:, continuous_columns] - mean) / std
df_test.loc[:, continuous_columns] = (df_test.loc[:, continuous_columns] - mean) / std
# Let's check the summary statistics on our training dataset to make sure it's standardized.
df_train.loc[:, continuous_columns].describe()
# + [markdown] colab_type="text" id="BX3woHz-jit1"
# <a name='4'></a>
# ## 4. Cox Proportional Hazards
#
# Our goal is to build a risk score using the survival data that we have. We'll begin by fitting a Cox Proportional Hazards model to your data.
#
# Recall that the Cox Proportional Hazards model describes the hazard for an individual $i$ at time $t$ as
#
# $$
# \lambda(t, x) = \lambda_0(t)e^{\theta^T X_i}
# $$
#
# The $\lambda_0$ term is a baseline hazard and incorporates the risk over time, and the other term incorporates the risk due to the individual's covariates. After fitting the model, we can rank individuals using the person-dependent risk term $e^{\theta^T X_i}$.
#
# Categorical variables cannot be used in a regression model as they are. In order to use them, conversion to a series of variables is required.
#
# Since our data has a mix of categorical (`stage`) and continuous (`wblc`) variables, before we proceed further we need to do some data engineering. To tackle the issue at hand we'll be using the `Dummy Coding` technique. In order to use Cox Proportional Hazards, we will have to turn the categorical data into one hot features so that we can fit our Cox model. Luckily, Pandas has a built-in function called `get_dummies` that will make it easier for us to implement our function. It turns categorical features into multiple binary features.
#
# <img src="1-hot-encode.png" style="padding-top: 5px;width: 60%;left: 0px;margin-left: 150px;margin-right: 0px;">
#
#
#
# -
# <a name='Ex-1'></a>
# ### Exercise 1
# In the cell below, implement the `to_one_hot(...)` function.
# <details>
# <summary>
# <font size="3" color="darkgreen"><b>Hints</b></font>
# </summary>
# <p>
# <ul>
# <li>Remember to drop the first dummy for each each category to avoid convergence issues when fitting the proportional hazards model.</li>
# <li> Check out the <a href="https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.get_dummies.html" > get_dummies() </a> documentation. </li>
# <li>Use <code>dtype=np.float64</code>.</li>
# </ul>
# </p>
# + colab={} colab_type="code" id="VMzvx0xF_C3I"
# UNQ_C1 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def to_one_hot(dataframe, columns):
'''
Convert columns in dataframe to one-hot encoding.
Args:
dataframe (dataframe): pandas dataframe containing covariates
columns (list of strings): list categorical column names to one hot encode
Returns:
one_hot_df (dataframe): dataframe with categorical columns encoded
as binary variables
'''
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
one_hot_df = pd.get_dummies(dataframe, columns = columns, dtype = np.float64, drop_first = True)
### END CODE HERE ###
return one_hot_df
# + [markdown] colab_type="text" id="rM2tIzvG_ifc"
# Now we'll use the function you coded to transform the training, validation, and test sets.
# + colab={} colab_type="code" id="SGZfLeup_fUL"
# List of categorical columns
to_encode = ['edema', 'stage']
one_hot_train = to_one_hot(df_train, to_encode)
one_hot_val = to_one_hot(df_val, to_encode)
one_hot_test = to_one_hot(df_test, to_encode)
print(one_hot_val.columns.tolist())
print(f"There are {len(one_hot_val.columns)} columns")
# -
# #### Expected output
# ```Python
# ['time', 'status', 'trt', 'age', 'sex', 'ascites', 'hepato', 'spiders', 'bili', 'chol', 'albumin', 'copper', 'alk.phos', 'ast', 'trig', 'platelet', 'protime', 'edema_0.5', 'edema_1.0', 'stage_2.0', 'stage_3.0', 'stage_4.0']
# There are 22 columns
# ```
# ### Look for new features
# Now, let's take a peek at one of the transformed data sets. Do you notice any new features?
# + colab={"base_uri": "https://localhost:8080/", "height": 241} colab_type="code" id="w8EG8A9gXcpu" outputId="384d9ade-2c96-4979-d3b7-da2b8e50f2e0"
print(one_hot_train.shape)
one_hot_train.head()
# + [markdown] colab_type="text" id="hNxuymLwyjqM"
# <a name='5'></a>
# ## 5. Fitting and Interpreting a Cox Model
# + [markdown] colab_type="text" id="ygiFcUKcAFQk"
# Run the following cell to fit your Cox Proportional Hazards model using the `lifelines` package.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="dDCS7p3xjbXB" outputId="41b12f82-8b35-43e1-d2a9-05258ac50b20"
cph = CoxPHFitter()
cph.fit(one_hot_train, duration_col = 'time', event_col = 'status', step_size=0.1)
# + [markdown] colab_type="text" id="5MUITR0QANDH"
# You can use `cph.print_summary()` to view the coefficients associated with each covariate as well as confidence intervals.
# + colab={"base_uri": "https://localhost:8080/", "height": 903} colab_type="code" id="fH5AZs8vjcEv" outputId="5429f7d5-5669-431f-a014-cf609c90997f"
cph.print_summary()
# -
# **Question:**
#
# - According to the model, was treatment `trt` beneficial?
# - What was its associated hazard ratio?
# - Note that the hazard ratio is how much an incremental increase in the feature variable changes the hazard.
# <details>
# <summary>
# <font size="3" color="darkgreen"><b>Check your answer!</b></font>
# </summary>
# <p>
# <ul>
# <ul>
# <li>You should see that the treatment (trt) was beneficial because it has a negative impact on the hazard (the coefficient is negative, and exp(coef) is less than 1).</li>
# <li>The associated hazard ratio is ~0.8, because this is the exp(coef) of treatment.</li>
# </ul>
# </p>
# We can compare the predicted survival curves for treatment variables. Run the next cell to plot survival curves using the `plot_covariate_groups()` function.
# - The y-axis is th survival rate
# - The x-axis is time
# + colab={"base_uri": "https://localhost:8080/", "height": 282} colab_type="code" id="Uxl0icyBS4Dr" outputId="5fa08369-e89e-424f-f9f0-60cf7a1cfbcd"
cph.plot_covariate_groups('trt', values=[0, 1]);
# -
# Notice how the group without treatment has a lower survival rate at all times (the x-axis is time) compared to the treatment group.
# <a name='6'></a>
# ## 6. Hazard Ratio
#
# Recall from the lecture videos that the Hazard Ratio between two patients was the likelihood of one patient (e.g smoker) being more at risk than the other (e.g non-smoker).
# $$
# \frac{\lambda_{smoker}(t)}{\lambda_{nonsmoker}(t)} = e^{\theta (X_{smoker} - X_{nonsmoker})^T}
# $$
#
# Where
#
# $$
# \lambda_{smoker}(t) = \lambda_0(t)e^{\theta X_{smoker}^T}
# $$
# and
# $$
# \lambda_{nonsmoker}(t) = \lambda_0(t)e^{\theta X_{nonsmoker}^T} \\
# $$
# <a name='Ex-2'></a>
# ### Exercise 2
# In the cell below, write a function to compute the hazard ratio between two individuals given the cox model's coefficients.
# <details>
# <summary>
# <font size="3" color="darkgreen"><b>Hints</b></font>
# </summary>
# <p>
# <ul>
# <li>use numpy.dot</li>
# <li>use nump.exp</li>
# </ul>
# </p>
#
# + colab={} colab_type="code" id="WbBmxbeDA3k1"
# UNQ_C2 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def hazard_ratio(case_1, case_2, cox_params):
'''
Return the hazard ratio of case_1 : case_2 using
the coefficients of the cox model.
Args:
case_1 (np.array): (1 x d) array of covariates
case_2 (np.array): (1 x d) array of covariates
model (np.array): (1 x d) array of cox model coefficients
Returns:
hazard_ratio (float): hazard ratio of case_1 : case_2
'''
### START CODE HERE (REPLACE INSTANCES OF 'None' with your code) ###
hr = np.exp(np.dot(cox_params,(case_1 - case_2).T))
### END CODE HERE ###
return hr
# + [markdown] colab_type="text" id="zbDQUxE6CcA3"
# Now, evaluate it on the following pair of indivduals: `i = 1` and `j = 5`
# + colab={} colab_type="code" id="7flsvTRXCgqO"
i = 1
case_1 = one_hot_train.iloc[i, :].drop(['time', 'status'])
j = 5
case_2 = one_hot_train.iloc[j, :].drop(['time', 'status'])
print(hazard_ratio(case_1.values, case_2.values, cph.params_.values))
# -
# #### Expected Output:
# ```CPP
# 15.029017732492221
# ```
# **Question:**
#
# Is `case_1` or `case_2` at greater risk?
# <details>
# <summary>
# <font size="3" color="darkgreen"><b>Check your answer!</b></font>
# </summary>
# <p>
# <ul>
# <ul>
# Important! The following answer only applies if you picked i = 1 and j = 5
# <li>You should see that `case_1` is at higher risk.</li>
# <li>The hazard ratio of case 1 / case 2 is greater than 1, so case 1 had a higher hazard relative to case 2</li>
# </ul>
# </p>
# Inspect different pairs, and see if you can figure out which patient is more at risk.
# + colab={"base_uri": "https://localhost:8080/", "height": 374} colab_type="code" id="g2PZ3sGvCs0K" outputId="59336868-d421-4645-d88e-76a8a8cffc9f"
i = 4
case_1 = one_hot_train.iloc[i, :].drop(['time', 'status'])
j = 7
case_2 = one_hot_train.iloc[j, :].drop(['time', 'status'])
print("Case 1\n\n", case_1, "\n")
print("Case 2\n\n", case_2, "\n")
print("Hazard Ratio:", hazard_ratio(case_1.values, case_2.values, cph.params_.values))
# -
# <details>
# <summary>
# <font size="3" color="darkgreen"><b>Check your answer!</b></font>
# </summary>
# <p>
# <ul>
# <ul>
# Important! The following answer only applies if you picked i = 4 and j = 7
# <li>You should see that `case_2` is at higher risk.</li>
# <li>The hazard ratio of case 1 / case 2 is less than 1, so case 2 had a higher hazard relative to case 1</li>
# </ul>
# </p>
# + [markdown] colab_type="text" id="KUa6r-KOyySp"
# <a name='7'></a>
# ## 7. Harrell's C-index
# + [markdown] colab_type="text" id="woQAtSmRXrgr"
# To evaluate how good our model is performing, we will write our own version of the C-index. Similar to the week 1 case, C-index in the survival context is the probability that, given a randomly selected pair of individuals, the one who died sooner has a higher risk score.
#
# However, we need to take into account censoring. Imagine a pair of patients, $A$ and $B$.
#
# #### Scenario 1
# - A was censored at time $t_A$
# - B died at $t_B$
# - $t_A < t_B$.
#
# Because of censoring, we can't say whether $A$ or $B$ should have a higher risk score.
#
# #### Scenario 2
# Now imagine that $t_A > t_B$.
#
# - A was censored at time $t_A$
# - B died at $t_B$
# - $t_A > t_B$
#
# Now we can definitively say that $B$ should have a higher risk score than $A$, since we know for a fact that $A$ lived longer.
#
# Therefore, when we compute our C-index
# - We should only consider pairs where at most one person is censored
# - If they are censored, then their censored time should occur *after* the other person's time of death.
#
# The metric we get if we use this rule is called **Harrel's C-index**.
#
# Note that in this case, being censored at time $t$ means that the true death time was some time AFTER time $t$ and not at $t$.
# - Therefore if $t_A = t_B$ and A was censored:
# - Then $A$ actually lived longer than $B$.
# - This will effect how you deal with ties in the exercise below!
#
#
# -
# <a name='Ex-3'></a>
# ### Exercise 3
# Fill in the function below to compute Harrel's C-index.
# <details>
# <summary>
# <font size="3" color="darkgreen"><b>Hints</b></font>
# </summary>
# <p>
# <ul>
# <li>If you get a division by zero error, consider checking how you count when a pair is permissible (in the case where one patient is censored and the other is not censored).</li>
# </ul>
# </p>
# UNQ_C3 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
def harrell_c(y_true, scores, event):
'''
Compute Harrel C-index given true event/censoring times,
model output, and event indicators.
Args:
y_true (array): array of true event times
scores (array): model risk scores
event (array): indicator, 1 if event occurred at that index, 0 for censorship
Returns:
result (float): C-index metric
'''
n = len(y_true)
assert (len(scores) == n and len(event) == n)
concordant = 0.0
permissible = 0.0
ties = 0.0
result = 0.0
### START CODE HERE (REPLACE INSTANCES OF 'None' and 'pass' with your code) ###
# use double for loop to go through cases
for i in range(n):
# set lower bound on j to avoid double counting
for j in range(i+1, n):
# check if at most one is censored
if event[i]==1 or event[j]==1:
# check if neither are censored
if event[i]==1 and event[j]==1:
permissible+=1
# check if scores are tied
if scores[i]==scores[j]:
ties = ties + 1
# check for concordant
elif scores[i]>scores[j] and y_true[i]<y_true[j]:
concordant = concordant + 1
elif scores[i]<scores[j] and y_true[j]<y_true[i]:
concordant = concordant + 1
# check if one is censored
elif event[i]==0 or event[j]==0:
# get censored index
censored = j
uncensored = i
if event[i] == 0:
censored = i
uncensored = j
# check if permissible
# Note: in this case, we are assuming that censored at a time
# means that you did NOT die at that time. That is, if you
# live until time 30 and have event = 0, then you lived THROUGH
# time 30.
if y_true[uncensored] <= y_true[censored]:
permissible +=1
# check if scores are tied
if scores[uncensored]==scores[censored]:
# update ties
ties = ties + 1
# check if scores are concordant
if scores[uncensored]>scores[censored]:
concordant = concordant + 1
# set result to c-index computed from number of concordant pairs,
# number of ties, and number of permissible pairs (REPLACE 0 with your code)
result = (concordant + (0.5 * ties)) / permissible
### END CODE HERE ###
return result
# You can test your function on the following test cases:
# +
y_true = [30, 12, 84, 9]
# Case 1
event = [1, 1, 1, 1]
scores = [0.5, 0.9, 0.1, 1.0]
print("Case 1")
print("Expected: 1.0, Output: {}".format(harrell_c(y_true, scores, event)))
# Case 2
scores = [0.9, 0.5, 1.0, 0.1]
print("\nCase 2")
print("Expected: 0.0, Output: {}".format(harrell_c(y_true, scores, event)))
# Case 3
event = [1, 0, 1, 1]
scores = [0.5, 0.9, 0.1, 1.0]
print("\nCase 3")
print("Expected: 1.0, Output: {}".format(harrell_c(y_true, scores, event)))
# Case 4
y_true = [30, 30, 20, 20]
event = [1, 0, 1, 0]
scores = [10, 5, 15, 20]
print("\nCase 4")
print("Expected: 0.75, Output: {}".format(harrell_c(y_true, scores, event)))
# Case 5
y_true = list(reversed([30, 30, 30, 20, 20]))
event = [0, 1, 0, 1, 0]
scores = list(reversed([15, 10, 5, 15, 20]))
print("\nCase 5")
print("Expected: 0.583, Output: {}".format(harrell_c(y_true, scores, event)))
# Case 6
y_true = [10,10]
event = [0,1]
scores = [4,5]
print("\nCase 6")
print(f"Expected: 1.0 , Output:{harrell_c(y_true, scores, event):.4f}")
# + [markdown] colab_type="text" id="CtQVe4pAn8ic"
# Now use the Harrell's C-index function to evaluate the cox model on our data sets.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="8nzHc_Qbn7dM" outputId="bc2f960d-16e5-46b2-a41f-695892c311c7"
# Train
scores = cph.predict_partial_hazard(one_hot_train)
cox_train_scores = harrell_c(one_hot_train['time'].values, scores.values, one_hot_train['status'].values)
# Validation
scores = cph.predict_partial_hazard(one_hot_val)
cox_val_scores = harrell_c(one_hot_val['time'].values, scores.values, one_hot_val['status'].values)
# Test
scores = cph.predict_partial_hazard(one_hot_test)
cox_test_scores = harrell_c(one_hot_test['time'].values, scores.values, one_hot_test['status'].values)
print("Train:", cox_train_scores)
print("Val:", cox_val_scores)
print("Test:", cox_test_scores)
# -
# What do these values tell us ?
# + [markdown] colab_type="text" id="AuNjR_wNkpWz"
# <a name='8'></a>
# ## 8. Random Survival Forests
#
# This performed well, but you have a hunch you can squeeze out better performance by using a machine learning approach. You decide to use a Random Survival Forest. To do this, you can use the `RandomForestSRC` package in R. To call R function from Python, we'll use the `r2py` package. Run the following cell to import the necessary requirements.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 530} colab_type="code" id="ZgSy-Dj6kquK" outputId="4aa5d2fa-30f4-4328-ae29-a2ff05223e22"
# %load_ext rpy2.ipython
# %R require(ggplot2)
from rpy2.robjects.packages import importr
# import R's "base" package
base = importr('base')
# import R's "utils" package
utils = importr('utils')
# import rpy2's package module
import rpy2.robjects.packages as rpackages
forest = rpackages.importr('randomForestSRC', lib_loc='R')
from rpy2 import robjects as ro
R = ro.r
from rpy2.robjects import pandas2ri
pandas2ri.activate()
# + [markdown] colab_type="text" id="LXBgqQBfuMA5"
# Instead of encoding our categories as binary features, we can use the original dataframe since trees deal well with raw categorical data (can you think why this might be?).
#
# Run the code cell below to build your forest.
# + colab={} colab_type="code" id="B-pio4o4mdVJ"
model = forest.rfsrc(ro.Formula('Surv(time, status) ~ .'), data=df_train, ntree=300, nodedepth=5, seed=-1)
# + colab={"base_uri": "https://localhost:8080/", "height": 289} colab_type="code" id="zZfcUvJ3nL04" outputId="27d00bd8-ea33-4c1b-f5d7-ca2c73ead721"
print(model)
# + [markdown] colab_type="text" id="9Mwzm55H-QKV"
# Finally, let's evaluate on our validation and test sets, and compare it with our Cox model.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="vfl4LbGfpbKp" outputId="13f8b560-e171-41e9-f6dc-cf0468a3f786"
result = R.predict(model, newdata=df_val)
scores = np.array(result.rx('predicted')[0])
print("Cox Model Validation Score:", cox_val_scores)
print("Survival Forest Validation Score:", harrell_c(df_val['time'].values, scores, df_val['status'].values))
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="uhqSQJhrplSG" outputId="752c266e-0234-45c5-d53f-554e2ff17a5a"
result = R.predict(model, newdata=df_test)
scores = np.array(result.rx('predicted')[0])
print("Cox Model Test Score:", cox_test_scores)
print("Survival Forest Validation Score:", harrell_c(df_test['time'].values, scores, df_test['status'].values))
# + [markdown] colab_type="text" id="Gp_SgUXreAWn"
# Your random forest model should be outperforming the Cox model slightly. Let's dig deeper to see how they differ.
# + [markdown] colab_type="text" id="ZtPMPaSli8GB"
# <a name='9'></a>
# ## 9. Permutation Method for Interpretation
#
# We'll dig a bit deeper into interpretation methods for forests a bit later, but for now just know that random surival forests come with their own built in variable importance feature. The method is referred to as VIMP, and for the purpose of this section you should just know that higher absolute value of the VIMP means that the variable generally has a larger effect on the model outcome.
#
# Run the next cell to compute and plot VIMP for the random survival forest.
# + colab={"base_uri": "https://localhost:8080/", "height": 281} colab_type="code" id="u7M4_N_d-YJu" outputId="7e1830cb-4b67-444f-8ba5-d49d3ff2f172"
vimps = np.array(forest.vimp(model).rx('importance')[0])
y = np.arange(len(vimps))
plt.barh(y, np.abs(vimps))
plt.yticks(y, df_train.drop(['time', 'status'], axis=1).columns)
plt.title("VIMP (absolute value)")
plt.show()
# + [markdown] colab_type="text" id="2YGhK2xwjkiA"
# ### Question:
#
# How does the variable importance compare to that of the Cox model? Which variable is important in both models? Which variable is important in the random survival forest but not in the Cox model? You should see that `edema` is important in both the random survival forest and the Cox model. You should also see that `bili` is important in the random survival forest but not the Cox model .
# -
# # Congratulations!
#
# You've finished the last assignment in course 2! Take a minute to look back at the analysis you've done over the last four assignments. You've done a great job!
|
AI for Medical Prognosis/Week4/C2M4_Assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Imports
# +
from tensorflow import keras
from keras.utils import np_utils
from keras.constraints import maxnorm
from keras.models import model_from_json
from io import BytesIO
from PIL import Image
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import MultinomialNB
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis
from sklearn.naive_bayes import GaussianNB
from sklearn.svm import SVC
from sklearn.linear_model import SGDClassifier
from sklearn.ensemble import VotingClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import Pipeline
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.model_selection import cross_validate, KFold
from sklearn.metrics import recall_score, make_scorer
from sklearn.ensemble import RandomForestClassifier
from sklearn.base import TransformerMixin, BaseEstimator
from sklearn.preprocessing import Normalizer
from sklearn.preprocessing import FunctionTransformer
from joblib import dump, load
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import warnings
import sys
import time
import pymongo
import copy
import os
warnings.filterwarnings('ignore')
print(tf.__version__)
# -
# # Helper Functions
# +
# Helper functions
def cprint(text):
sys.stdout.write("\r" + text)
sys.stdout.flush()
def split_list(a_list, amount):
return a_list[amount:], a_list[:amount]
def plot_image(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array[i], true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'blue'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
def plot_value_array(i, predictions_array, true_label):
predictions_array, true_label = predictions_array[i], true_label[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
thisplot = plt.bar(range(10), predictions_array, color="#777777")
plt.ylim([0, 1])
predicted_label = np.argmax(predictions_array)
thisplot[predicted_label].set_color('red')
thisplot[true_label].set_color('blue')
def save_model(model, fileName):
dump(model, 'models/%s.joblib' % fileName)
def load_model(fileName):
try:
if not os.path.exists('models'):
os.makedirs('models')
return load('models/%s.joblib' % fileName)
except FileNotFoundError:
return None
def merge_probabilities(prob1, prob2):
temp = prob1 + prob2
return [x / 2 for x in temp]
# -
# # Retrieving data
# +
def get_ml_data():
cbs_db_ties = pymongo.MongoClient(
"mongodb://cbs_user:cbs_pwd@<EMAIL>:27017/test")["test"]
data = cbs_db_ties["data"]
nrofrecords = cbs_db_ties.command("collstats", "data")['count']
posts = data.aggregate([{ '$sample': { 'size': nrofrecords } }])
post = posts.next()
img = Image.open(BytesIO(post['image']))
imgs = np.array([np.asarray(img)])
texts = np.array([post['bericht tekst']])
lbls= np.array(post['label'])
for index, post in enumerate(posts):
newPost = copy.deepcopy(post)
del newPost['_id']
cprint("Getting data " + str(round((index / nrofrecords) * 100)) + "% completed")
try:
img = Image.open(BytesIO(newPost['image']))
imgs = np.append(imgs, np.array([np.asarray(img)]), axis=0)
texts = np.append(texts, newPost['bericht tekst'])
lbls = np.append(lbls, newPost['label'])
except Exception as ex:
print(ex)
return (imgs,texts,lbls)
img_data, text_data, labels = get_ml_data()
# +
np_img = np.array(img_data)
np_text = np.array(text_data)
np_lbls = np.array(labels)
np.save('img.npy', np_img)
np.save('text.npy', np_text)
np.save('lbls.npy', np_lbls)
# -
img_data = np.load('img.npy')
text_data = np.load('text.npy')
labels = np.load('lbls.npy')
# # Prepare data
# this step is neccesary for the image handling of the models
# +
# labels for end results
class_names = ['sport','vakantie','werk','IT','politiek','school','events','veiligheid','lifestyle','social media']
num_classes = 10 #
# Splitting data
train_imgs, test_imgs = split_list(img_data, round(len(img_data)/10))
train_text, test_text = split_list(text_data, round(len(text_data)/10))
train_lbls, test_lbls = split_list(labels, round(len(labels)/10))
train_img_data = train_imgs
test_img_data = test_imgs
train_labels = train_lbls
test_labels = test_lbls
# Normalizing image data
train_lbls = np_utils.to_categorical(train_lbls, num_classes)
test_lbls = np_utils.to_categorical(test_lbls, num_classes)
train_imgs = train_imgs.astype('float32')
test_imgs = test_imgs.astype('float32')
train_imgs /= 255
test_imgs /= 255
# -
# # Deep learning
# ## Building model
# +
# Setup the model
deep_model = keras.Sequential([
# feature extraction layers
keras.layers.Conv2D(32, kernel_size=(3,3), padding="same", input_shape=train_imgs.shape[1:], activation=tf.nn.relu),
keras.layers.Dropout(0.2),
keras.layers.Conv2D(32, kernel_size=(3,3), padding="same", activation=tf.nn.relu),
keras.layers.MaxPool2D(pool_size=(2,2)),
keras.layers.Conv2D(64, kernel_size=(3,3), padding="same", activation=tf.nn.relu),
keras.layers.Dropout(0.2),
keras.layers.Conv2D(64, kernel_size=(3,3), padding="same", activation=tf.nn.relu),
keras.layers.MaxPool2D(pool_size=(2,2)),
keras.layers.Conv2D(128, kernel_size=(3,3), padding="same", activation=tf.nn.relu),
keras.layers.Dropout(0.2),
keras.layers.Conv2D(128, kernel_size=(3,3), padding="same", activation=tf.nn.relu),
keras.layers.MaxPool2D(pool_size=(2,2)),
# hidden layers
keras.layers.Flatten(),
keras.layers.Dense(1024, activation=tf.nn.relu, kernel_constraint=maxnorm(3)),
keras.layers.Dropout(0.2),
keras.layers.Dense(10, activation=tf.nn.softmax)
])
# compile the model
deep_model.compile(optimizer=keras.optimizers.Adam(),
loss='categorical_crossentropy',
metrics=['accuracy'])
# -
# ## Train model
# train the model
deep_model.fit(train_imgs, train_lbls, epochs=20)
# ## Save model
# +
# Save the weights
deep_model.save_weights('deep_model_weights.h5')
# Save the model architecture
with open('deep_model.json', 'w') as f:
f.write(deep_model.to_json())
# -
# ## Load model
# +
# Model reconstruction from JSON file
with open('insta_architecture_24k.json', 'r') as f:
deep_model = model_from_json(f.read())
# Load weights into the new model
deep_model.load_weights('insta_weights_24k.h5')
# -
# # Machine learning
#
# ## setting up models
def refreshModel():
models = []
models.append((
'LogisticRegression',
LogisticRegression(
max_iter=1000,
class_weight='balanced'
),
[{'clf__C': [1]}],
None
))
models.append((
'LogisticRegression (Skewed)',
LogisticRegression(
max_iter=1000
),
[{'clf__C': [10]}],
None
))
models.append((
'SVM',
SVC(class_weight='balanced', probability=True),
[{'clf__C': [1000]}],
None
))
models.append((
'SVM (Skewed)',
SVC(probability=True),
[{'clf__C': [1000]}],
None
))
models.append((
'SGDClassifier',
SGDClassifier(loss='log', max_iter=1000, class_weight='balanced'),
[{
'vect__ngram_range': [(1, 2)],
'tfidf__use_idf': (True, False),
'clf__alpha': (1e-2, 1e-3)
}],
None
))
models.append((
'SGDClassifier (Skewed)',
SGDClassifier(max_iter=1000, loss='log'),
[{
'vect__ngram_range': [(1, 2)],
'tfidf__use_idf': (True, False),
'clf__alpha': (1e-2, 1e-3)
}],
None
))
return models
# +
models = refreshModel()
classifiers = []
seed = 7;
print(train_text.shape)
print(train_labels.shape)
for name, model, param, dense in models:
classifiers.append((name, model))
start = time.time()
text_model = Pipeline([
('vect', CountVectorizer(stop_words='english')),
('normalizer', Normalizer()),
('tfidf', TfidfTransformer()),
('clf', VotingClassifier(estimators=classifiers, voting='soft')),
])
text_model.fit(train_text, train_labels)
dump(text_model, 'text_model.joblib')
print('time: %s \n' % (time.time() - start))
# -
text_model = load_model("text_model")
# # Combining the results
# +
use_data = img_data.astype('float32')
use_data /= 255
text_proba = text_model.predict_proba(text_data)
img_proba = deep_model.predict(use_data)
# -
final_proba = merge_probabilities(text_proba, img_proba)
predicted_lbls = np.array([np.argmax(x) for x in final_proba])
plot_value_array(0, final_proba, labels)
acc = sum(1 for x,y in zip(labels, predicted_lbls) if x == y) / float(len(labels))
print("Overall accuray: " + str(acc))
offset = 10
num_rows = 5
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(2*2*num_cols, 2*num_rows))
for i in range(num_images):
plt.subplot(num_rows, 2*num_cols, 2*i+1)
plot_image(i, final_proba, labels, img_data)
plt.subplot(num_rows, 2*num_cols, 2*i+2)
plot_value_array(i, final_proba, labels)
# ## Accuracy per model
# +
text_model_pred = np.array([np.argmax(x) for x in img_proba])
deep_model_pred = np.array([np.argmax(x) for x in img_proba])
text_acc = sum(1 for x,y in zip(labels, text_model_pred) if x == y) / float(len(labels))
deep_acc = sum(1 for x,y in zip(labels, deep_model_pred) if x == y) / float(len(labels))
print("Text accuray: " + str(text_acc))
print("Image accuray: " + str(deep_acc))
# -
# ## Graph Text model
num_rows = 5
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(2*2*num_cols, 2*num_rows))
for i in range(num_images):
plt.subplot(num_rows, 2*num_cols, 2*i+1)
plot_image(i, text_proba, labels, img_data)
plt.subplot(num_rows, 2*num_cols, 2*i+2)
plot_value_array(i, text_proba, labels)
# ## Graph Image model
num_rows = 5
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(2*2*num_cols, 2*num_rows))
for i in range(num_images):
plt.subplot(num_rows, 2*num_cols, 2*i+1)
plot_image(i, img_proba, labels, img_data)
plt.subplot(num_rows, 2*num_cols, 2*i+2)
plot_value_array(i, img_proba, labels)
# # Do predictions on whole dataset
# +
def get_predict_data():
cbs_db_ties = pymongo.MongoClient(
"mongodb://cbs_user:cbs_<EMAIL>:27017/test")["test"]
data = cbs_db_ties["instagram"]
nrofrecords = cbs_db_ties.command("collstats", "instagram")['count']
print(nrofrecords)
posts = data.aggregate([{ '$sample': { 'size': nrofrecords } }], allowDiskUse=True)
post = posts.next()
img = Image.open(BytesIO(post['image']))
imgs = np.array([np.asarray(img)])
texts = np.array([post['bericht tekst']])
for index, post in enumerate(posts):
newPost = copy.deepcopy(post)
del newPost['_id']
cprint("Getting data " + str(round((index / nrofrecords) * 100)) + "% completed")
try:
img = Image.open(BytesIO(newPost['image']))
imgs = np.append(imgs, np.array([np.asarray(img)]), axis=0)
texts = np.append(texts, newPost['bericht tekst'])
except Exception as ex:
print(ex)
return (imgs,texts)
img_data, text_data = get_predict_data()
# +
use_data = img_data.astype('float32')
use_data /= 255
print(len(use_data))
# -
text_proba = text_model.predict_proba(text_data)
img_proba = deep_model.predict(use_data)
final_proba = merge_probabilities(text_proba, img_proba)
predicted_lbls = np.array([np.argmax(x) for x in final_proba])
# +
class_names = ['sport','vakantie','werk','IT','politiek','school','events','veiligheid','lifestyle','social media']
data = {
'vakantie': 0,
'lifestyle': 0,
'events': 0,
'sport': 0,
'werk': 0,
'school': 0,
'social media': 0,
'politiek': 0,
'IT': 0,
'veiligheid': 0
}
for label in predicted_lbls:
data[class_names[label]] = data[class_names[label]] + 1
print(data)
# -
plt.figure(figsize=(15,15))
plt.bar(data.keys(), data.values())
|
machine learning/Deep Learning/Combine the machine learning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: MRIenv
# language: python
# name: mrienv
# ---
from nilearn import datasets, plotting, image
ds = datasets.fetch_development_fmri(n_subjects=1)
ds['description']
ds['func'][0]
# +
mean_func = image.mean_img(ds['func'][0])
for smoothing in range(0, 25, 5):
smoothed_img = image.smooth_img(mean_func, smoothing)
plotting.plot_epi(
smoothed_img,
title="Smoothing %imm" % smoothing)
# -
plotting.plot_glass_brain("/data/perlman/moochie/analysis/CARE/MRI_data_clean/sub-50802/ses-0/anat/sub-50802_ses-0_run-1_T1w.nii.gz")
|
scripts/MRI/nilearn-practice.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="-Jv7Y4hXwt0j"
# # Assignment 4: Question duplicates
#
# Welcome to the fourth assignment of course 3. In this assignment you will explore Siamese networks applied to natural language processing. You will further explore the fundamentals of Trax and you will be able to implement a more complicated structure using it. By completing this assignment, you will learn how to implement models with different architectures.
#
# ## Outline
#
# - [Overview](#0)
# - [Part 1: Importing the Data](#1)
# - [1.1 Loading in the data](#1.1)
# - [1.2 Converting a question to a tensor](#1.2)
# - [1.3 Understanding the iterator](#1.3)
# - [Exercise 01](#ex01)
# - [Part 2: Defining the Siamese model](#2)
# - [2.1 Understanding Siamese Network](#2.1)
# - [Exercise 02](#ex02)
# - [2.2 Hard Negative Mining](#2.2)
# - [Exercise 03](#ex03)
# - [Part 3: Training](#3)
# - [3.1 Training the model](#3.1)
# - [Exercise 04](#ex04)
# - [Part 4: Evaluation](#4)
# - [4.1 Evaluating your siamese network](#4.1)
# - [4.2 Classify](#4.2)
# - [Exercise 05](#ex05)
# - [Part 5: Testing with your own questions](#5)
# - [Exercise 06](#ex06)
# - [On Siamese networks](#6)
#
# <a name='0'></a>
# ### Overview
# In this assignment, concretely you will:
#
# - Learn about Siamese networks
# - Understand how the triplet loss works
# - Understand how to evaluate accuracy
# - Use cosine similarity between the model's outputted vectors
# - Use the data generator to get batches of questions
# - Predict using your own model
#
# By now, you are familiar with trax and know how to make use of classes to define your model. We will start this homework by asking you to preprocess the data the same way you did in the previous assignments. After processing the data you will build a classifier that will allow you to identify whether to questions are the same or not.
# <img src = "meme.png" style="width:550px;height:300px;"/>
#
#
# You will process the data first and then pad in a similar way you have done in the previous assignment. Your model will take in the two question embeddings, run them through an LSTM, and then compare the outputs of the two sub networks using cosine similarity. Before taking a deep dive into the model, start by importing the data set.
#
# + [markdown] colab_type="text" id="4sF9Hqzgwt0l"
# <a name='1'></a>
# # Part 1: Importing the Data
# <a name='1.1'></a>
# ### 1.1 Loading in the data
#
# You will be using the Quora question answer dataset to build a model that could identify similar questions. This is a useful task because you don't want to have several versions of the same question posted. Several times when teaching I end up responding to similar questions on piazza, or on other community forums. This data set has been labeled for you. Run the cell below to import some of the packages you will be using.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="zdACgs491cs2" outputId="b31042ef-845b-46b8-c783-185e96b135f7"
import os
import nltk
import trax
from trax import layers as tl
from trax.supervised import training
from trax.fastmath import numpy as fastnp
import numpy as np
import pandas as pd
import random as rnd
# set random seeds
trax.supervised.trainer_lib.init_random_number_generators(34)
rnd.seed(34)
# + [markdown] colab_type="text" id="3GYhQRMspitx"
# **Notice that for this assignment Trax's numpy is referred to as `fastnp`, while regular numpy is referred to as `np`.**
#
# You will now load in the data set. We have done some preprocessing for you. If you have taken the deeplearning specialization, this is a slightly different training method than the one you have seen there. If you have not, then don't worry about it, we will explain everything.
# + colab={"base_uri": "https://localhost:8080/", "height": 528} colab_type="code" id="sXWBVGWnpity" outputId="afa90d4d-fed7-43b8-bcba-48c95d600ad5"
data = pd.read_csv("questions.csv")
N=len(data)
print('Number of question pairs: ', N)
data.head()
# + [markdown] colab_type="text" id="gkSQTu7Ypit0"
# We first split the data into a train and test set. The test set will be used later to evaluate our model.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="z00A7vEMpit1" outputId="c12ae7e8-a959-4f56-aa29-6ad34abc1c81"
N_train = 300000
N_test = 10*1024
data_train = data[:N_train]
data_test = data[N_train:N_train+N_test]
print("Train set:", len(data_train), "Test set:", len(data_test))
del(data) # remove to free memory
# + [markdown] colab_type="text" id="FbqIRRyEpit4"
# As explained in the lectures, we select only the question pairs that are duplicate to train the model. <br>
# We build two batches as input for the Siamese network and we assume that question $q1_i$ (question $i$ in the first batch) is a duplicate of $q2_i$ (question $i$ in the second batch), but all other questions in the second batch are not duplicates of $q1_i$.
# The test set uses the original pairs of questions and the status describing if the questions are duplicates.
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="Xi_TwXxxpit4" outputId="f146046f-9c0d-4d8a-ecf8-8d6a4a5371f7"
td_index = (data_train['is_duplicate'] == 1).to_numpy()
td_index = [i for i, x in enumerate(td_index) if x]
print('number of duplicate questions: ', len(td_index))
print('indexes of first ten duplicate questions:', td_index[:10])
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="3I9oXSsKpit7" outputId="6f6bd3a1-219f-4fb3-a524-450c38bf44ba"
print(data_train['question1'][5]) # Example of question duplicates (first one in data)
print(data_train['question2'][5])
print('is_duplicate: ', data_train['is_duplicate'][5])
# + colab={} colab_type="code" id="XHpZO58Dss_v"
Q1_train_words = np.array(data_train['question1'][td_index])
Q2_train_words = np.array(data_train['question2'][td_index])
Q1_test_words = np.array(data_test['question1'])
Q2_test_words = np.array(data_test['question2'])
y_test = np.array(data_test['is_duplicate'])
# + [markdown] colab_type="text" id="P5vBkxunpiuB"
# Above, you have seen that you only took the duplicated questions for training our model. <br>You did so on purpose, because the data generator will produce batches $([q1_1, q1_2, q1_3, ...]$, $[q2_1, q2_2,q2_3, ...])$ where $q1_i$ and $q2_k$ are duplicate if and only if $i = k$.
#
# <br>Let's print to see what your data looks like.
# + colab={"base_uri": "https://localhost:8080/", "height": 170} colab_type="code" id="joyrS1XEpLWn" outputId="3257cde7-3164-40d9-910e-fa91eae917a0"
print('TRAINING QUESTIONS:\n')
print('Question 1: ', Q1_train_words[0])
print('Question 2: ', Q2_train_words[0], '\n')
print('Question 1: ', Q1_train_words[5])
print('Question 2: ', Q2_train_words[5], '\n')
print('TESTING QUESTIONS:\n')
print('Question 1: ', Q1_test_words[0])
print('Question 2: ', Q2_test_words[0], '\n')
print('is_duplicate =', y_test[0], '\n')
# + [markdown] colab_type="text" id="WC_BZU3XpiuF"
# You will now encode each word of the selected duplicate pairs with an index. <br> Given a question, you can then just encode it as a list of numbers.
#
# First you tokenize the questions using `nltk.word_tokenize`. <br>
# You need a python default dictionary which later, during inference, assigns the values $0$ to all Out Of Vocabulary (OOV) words.<br>
# Then you encode each word of the selected duplicate pairs with an index. Given a question, you can then just encode it as a list of numbers.
# + colab={} colab_type="code" id="QbCoIgLQpiuF"
#create arrays
Q1_train = np.empty_like(Q1_train_words)
Q2_train = np.empty_like(Q2_train_words)
Q1_test = np.empty_like(Q1_test_words)
Q2_test = np.empty_like(Q2_test_words)
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="m9ZmfpGWpiuI" outputId="d2995c9a-92b4-4892-d34b-c77b94b27134"
# Building the vocabulary with the train set (this might take a minute)
from collections import defaultdict
vocab = defaultdict(lambda: 0)
vocab['<PAD>'] = 1
for idx in range(len(Q1_train_words)):
Q1_train[idx] = nltk.word_tokenize(Q1_train_words[idx])
Q2_train[idx] = nltk.word_tokenize(Q2_train_words[idx])
q = Q1_train[idx] + Q2_train[idx]
for word in q:
if word not in vocab:
vocab[word] = len(vocab) + 1
print('The length of the vocabulary is: ', len(vocab))
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="TTMRF8eZpiuK" outputId="f81d4dc1-7cf9-4476-a454-467b54fe4dc4"
print(vocab['<PAD>'])
print(vocab['Astrology'])
print(vocab['Astronomy']) #not in vocabulary, returns 0
print(Q1_train[0])
print(Q2_train[0])
# + colab={} colab_type="code" id="5sDs36m81g6f"
for idx in range(len(Q1_test_words)):
Q1_test[idx] = nltk.word_tokenize(Q1_test_words[idx])
Q2_test[idx] = nltk.word_tokenize(Q2_test_words[idx])
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="3QgGE9KlpiuP" outputId="19c3cf93-cf0d-4f8f-da99-e75481f16599"
print('Train set has reduced to: ', len(Q1_train) )
print('Test set length: ', len(Q1_test) )
# + [markdown] colab_type="text" id="BDcxEmX31y3d"
# <a name='1.2'></a>
# ### 1.2 Converting a question to a tensor
#
# You will now convert every question to a tensor, or an array of numbers, using your vocabulary built above.
# + colab={} colab_type="code" id="zOhNa-sapiuS"
# Converting questions to array of integers
for i in range(len(Q1_train)):
Q1_train[i] = [vocab[word] for word in Q1_train[i]]
Q2_train[i] = [vocab[word] for word in Q2_train[i]]
for i in range(len(Q1_test)):
Q1_test[i] = [vocab[word] for word in Q1_test[i]]
Q2_test[i] = [vocab[word] for word in Q2_test[i]]
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="Dpawm38dpiuU" outputId="ef1aa65b-c89b-46f9-a9cf-f73748f1ee56"
print('first question in the train set:\n')
print(Q1_train_words[0], '\n')
print('encoded version:')
print(Q1_train[0],'\n')
print('first question in the test set:\n')
print(Q1_test_words[0], '\n')
print('encoded version:')
print(Q1_test[0])
# + [markdown] colab_type="text" id="SuggGPaQpiuY"
# You will now split your train set into a training/validation set so that you can use it to train and evaluate your Siamese model.
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="BmhrWPtgpiuY" outputId="7272fb74-79e6-499a-ce95-d11b9edcd64a"
# Splitting the data
cut_off = int(len(Q1_train)*.8)
train_Q1, train_Q2 = Q1_train[:cut_off], Q2_train[:cut_off]
val_Q1, val_Q2 = Q1_train[cut_off: ], Q2_train[cut_off:]
print('Number of duplicate questions: ', len(Q1_train))
print("The length of the training set is: ", len(train_Q1))
print("The length of the validation set is: ", len(val_Q1))
# + [markdown] colab_type="text" id="iFOR19cX2TQs"
# <a name='1.3'></a>
# ### 1.3 Understanding the iterator
#
# Most of the time in Natural Language Processing, and AI in general we use batches when training our data sets. If you were to use stochastic gradient descent with one example at a time, it will take you forever to build a model. In this example, we show you how you can build a data generator that takes in $Q1$ and $Q2$ and returns a batch of size `batch_size` in the following format $([q1_1, q1_2, q1_3, ...]$, $[q2_1, q2_2,q2_3, ...])$. The tuple consists of two arrays and each array has `batch_size` questions. Again, $q1_i$ and $q2_i$ are duplicates, but they are not duplicates with any other elements in the batch.
#
# <br>
#
# The command ```next(data_generator)```returns the next batch. This iterator returns the data in a format that you could directly use in your model when computing the feed-forward of your algorithm. This iterator returns a pair of arrays of questions.
#
# <a name='ex01'></a>
# ### Exercise 01
#
# **Instructions:**
# Implement the data generator below. Here are some things you will need.
#
# - While true loop.
# - if `index >= len_Q1`, set the `idx` to $0$.
# - The generator should return shuffled batches of data. To achieve this without modifying the actual question lists, a list containing the indexes of the questions is created. This list can be shuffled and used to get random batches everytime the index is reset.
# - Append elements of $Q1$ and $Q2$ to `input1` and `input2` respectively.
# - if `len(input1) == batch_size`, determine `max_len` as the longest question in `input1` and `input2`. Ceil `max_len` to a power of $2$ (for computation purposes) using the following command: `max_len = 2**int(np.ceil(np.log2(max_len)))`.
# - Pad every question by `vocab['<PAD>']` until you get the length `max_len`.
# - Use yield to return `input1, input2`.
# - Don't forget to reset `input1, input2` to empty arrays at the end (data generator resumes from where it last left).
# + colab={} colab_type="code" id="ibchgos48MtA"
# UNQ_C1 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: data_generator
def data_generator(Q1, Q2, batch_size, pad=1, shuffle=True):
"""Generator function that yields batches of data
Args:
Q1 (list): List of transformed (to tensor) questions.
Q2 (list): List of transformed (to tensor) questions.
batch_size (int): Number of elements per batch.
pad (int, optional): Pad character from the vocab. Defaults to 1.
shuffle (bool, optional): If the batches should be randomnized or not. Defaults to True.
Yields:
tuple: Of the form (input1, input2) with types (numpy.ndarray, numpy.ndarray)
NOTE: input1: inputs to your model [q1a, q2a, q3a, ...] i.e. (q1a,q1b) are duplicates
input2: targets to your model [q1b, q2b,q3b, ...] i.e. (q1a,q2i) i!=a are not duplicates
"""
input1 = []
input2 = []
idx = 0
len_q = len(Q1)
question_indexes = [*range(len_q)]
if shuffle:
rnd.shuffle(question_indexes)
### START CODE HERE (Replace instances of 'None' with your code) ###
while True:
if idx >= len_q:
# if idx is greater than or equal to len_q, set idx accordingly
# (Hint: look at the instructions above)
idx = 0
# shuffle to get random batches if shuffle is set to True
if shuffle:
rnd.shuffle(question_indexes)
# get questions at the `question_indexes[idx]` position in Q1 and Q2
q1 = Q1[question_indexes[idx]]
q2 = Q2[question_indexes[idx]]
# increment idx by 1
idx += 1
# append q1
input1.append(q1)
# append q2
input2.append(q2)
if len(input1) == batch_size:
# determine max_len as the longest question in input1 & input 2
# Hint: use the `max` function.
# take max of input1 & input2 and then max out of the two of them.
max_len = max([max([ len(i) for i in input1]), max([len(i) for i in input2])])
# print('max lenth value is ',max_len)
# pad to power-of-2 (Hint: look at the instructions above)
max_len = 2**int(np.ceil(np.log2(max_len)))
b1 = []
b2 = []
for q1, q2 in zip(input1, input2):
# add [pad] to q1 until it reaches max_len
q1 = q1[:max_len] + [pad]*(max_len - len(q1))
# add [pad] to q2 until it reaches max_len
q2 = q2[:max_len] + [pad]*(max_len - len(q2))
# append q1
b1.append(q1)
# append q2
b2.append(q2)
# use b1 and b2
yield np.array(b1), np.array(b2)
### END CODE HERE ###
# reset the batches
input1, input2 = [], [] # reset the batches
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="ZFZeBPnW8Mlb" outputId="7a31cd19-55dc-4b97-f288-6c59c6a34b53"
batch_size = 2
res1, res2 = next(data_generator(train_Q1, train_Q2, batch_size))
print("First questions : ",'\n', res1, '\n')
print("Second questions : ",'\n', res2)
# + [markdown] colab_type="text" id="tWJ1L9m2piui"
# **Note**: The following expected output is valid only if you run the above test cell **_once_** (first time). The output will change on each execution.
#
# If you think your implementation is correct and it is not matching the output, make sure to restart the kernel and run all the cells from the top again.
#
# **Expected Output:**
# ```CPP
# First questions :
# [[ 30 87 78 134 2132 1981 28 78 594 21 1 1 1 1
# 1 1]
# [ 30 55 78 3541 1460 28 56 253 21 1 1 1 1 1
# 1 1]]
#
# Second questions :
# [[ 30 156 78 134 2132 9508 21 1 1 1 1 1 1 1
# 1 1]
# [ 30 156 78 3541 1460 131 56 253 21 1 1 1 1 1
# 1 1]]
# ```
# Now that you have your generator, you can just call it and it will return tensors which correspond to your questions in the Quora data set.<br>Now you can go ahead and start building your neural network.
#
#
# + [markdown] colab_type="text" id="KmZRBoaMwt0w"
# <a name='2'></a>
# # Part 2: Defining the Siamese model
#
# <a name='2.1'></a>
#
# ### 2.1 Understanding Siamese Network
# A Siamese network is a neural network which uses the same weights while working in tandem on two different input vectors to compute comparable output vectors.The Siamese network you are about to implement looks like this:
#
# <img src = "siamese.png" style="width:600px;height:300px;"/>
#
# You get the question embedding, run it through an LSTM layer, normalize $v_1$ and $v_2$, and finally use a triplet loss (explained below) to get the corresponding cosine similarity for each pair of questions. As usual, you will start by importing the data set. The triplet loss makes use of a baseline (anchor) input that is compared to a positive (truthy) input and a negative (falsy) input. The distance from the baseline (anchor) input to the positive (truthy) input is minimized, and the distance from the baseline (anchor) input to the negative (falsy) input is maximized. In math equations, you are trying to maximize the following.
#
# $$\mathcal{L}(A, P, N)=\max \left(\|\mathrm{f}(A)-\mathrm{f}(P)\|^{2}-\|\mathrm{f}(A)-\mathrm{f}(N)\|^{2}+\alpha, 0\right)$$
#
# $A$ is the anchor input, for example $q1_1$, $P$ the duplicate input, for example, $q2_1$, and $N$ the negative input (the non duplicate question), for example $q2_2$.<br>
# $\alpha$ is a margin; you can think about it as a safety net, or by how much you want to push the duplicates from the non duplicates.
# <br>
#
# <a name='ex02'></a>
# ### Exercise 02
#
# **Instructions:** Implement the `Siamese` function below. You should be using all the objects explained below.
#
# To implement this model, you will be using `trax`. Concretely, you will be using the following functions.
#
#
# - `tl.Serial`: Combinator that applies layers serially (by function composition) allows you set up the overall structure of the feedforward. [docs](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.combinators.Serial) / [source code](https://github.com/google/trax/blob/1372b903bb66b0daccee19fd0b1fdf44f659330b/trax/layers/combinators.py#L26)
# - You can pass in the layers as arguments to `Serial`, separated by commas.
# - For example: `tl.Serial(tl.Embeddings(...), tl.Mean(...), tl.Dense(...), tl.LogSoftmax(...))`
#
#
# - `tl.Embedding`: Maps discrete tokens to vectors. It will have shape (vocabulary length X dimension of output vectors). The dimension of output vectors (also called d_feature) is the number of elements in the word embedding. [docs](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.core.Embedding) / [source code](https://github.com/google/trax/blob/1372b903bb66b0daccee19fd0b1fdf44f659330b/trax/layers/core.py#L113)
# - `tl.Embedding(vocab_size, d_feature)`.
# - `vocab_size` is the number of unique words in the given vocabulary.
# - `d_feature` is the number of elements in the word embedding (some choices for a word embedding size range from 150 to 300, for example).
#
#
# - `tl.LSTM` The LSTM layer. It leverages another Trax layer called [`LSTMCell`](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.rnn.LSTMCell). The number of units should be specified and should match the number of elements in the word embedding. [docs](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.rnn.LSTM) / [source code](https://github.com/google/trax/blob/1372b903bb66b0daccee19fd0b1fdf44f659330b/trax/layers/rnn.py#L87)
# - `tl.LSTM(n_units)` Builds an LSTM layer of n_units.
#
#
# - `tl.Mean`: Computes the mean across a desired axis. Mean uses one tensor axis to form groups of values and replaces each group with the mean value of that group. [docs](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.core.Mean) / [source code](https://github.com/google/trax/blob/1372b903bb66b0daccee19fd0b1fdf44f659330b/trax/layers/core.py#L276)
# - `tl.Mean(axis=1)` mean over columns.
#
#
# - `tl.Fn` Layer with no weights that applies the function f, which should be specified using a lambda syntax. [docs](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.base.Fn) / [source doce](https://github.com/google/trax/blob/70f5364dcaf6ec11aabbd918e5f5e4b0f5bfb995/trax/layers/base.py#L576)
# - $x$ -> This is used for cosine similarity.
# - `tl.Fn('Normalize', lambda x: normalize(x))` Returns a layer with no weights that applies the function `f`
#
#
# - `tl.parallel`: It is a combinator layer (like `Serial`) that applies a list of layers in parallel to its inputs. [docs](https://trax-ml.readthedocs.io/en/latest/trax.layers.html#trax.layers.combinators.Parallel) / [source code](https://github.com/google/trax/blob/37aba571a89a8ad86be76a569d0ec4a46bdd8642/trax/layers/combinators.py#L152)
#
# + colab={} colab_type="code" id="hww76f8_wt0x"
# UNQ_C2 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: Siamese
def Siamese(vocab_size=len(vocab), d_model=128, mode='train'):
"""Returns a Siamese model.
Args:
vocab_size (int, optional): Length of the vocabulary. Defaults to len(vocab).
d_model (int, optional): Depth of the model. Defaults to 128.
mode (str, optional): 'train', 'eval' or 'predict', predict mode is for fast inference. Defaults to 'train'.
Returns:
trax.layers.combinators.Parallel: A Siamese model.
"""
def normalize(x): # normalizes the vectors to have L2 norm 1
return x / fastnp.sqrt(fastnp.sum(x * x, axis=-1, keepdims=True))
### START CODE HERE (Replace instances of 'None' with your code) ###
q_processor = tl.Serial( # Processor will run on Q1 and Q2.
tl.Embedding(vocab_size,d_model), # Embedding layer
tl.LSTM(d_model), # LSTM layer
tl.Mean(axis=1), # Mean over columns
tl.Fn('Normalize', lambda x: normalize(x)) # Apply normalize function
) # Returns one vector of shape [batch_size, d_model].
### END CODE HERE ###
# Run on Q1 and Q2 in parallel.
model = tl.Parallel(q_processor, q_processor)
return model
# + [markdown] colab_type="text" id="es2gfwZypiul"
# Setup the Siamese network model
# + colab={"base_uri": "https://localhost:8080/", "height": 255} colab_type="code" id="kvQ_jf52-JAn" outputId="d409460d-2ffb-4ae6-8745-ddcfa1d892ad"
# check your model
model = Siamese()
print(model)
# + [markdown] colab_type="text" id="LMK9zqhHpiuo"
# **Expected output:**
#
# ```CPP
# Parallel_in2_out2[
# Serial[
# Embedding_41699_128
# LSTM_128
# Mean
# Normalize
# ]
# Serial[
# Embedding_41699_128
# LSTM_128
# Mean
# Normalize
# ]
# ]
# ```
# + [markdown] colab_type="text" id="KVo1Gvripiuo"
# <a name='2.2'></a>
#
# ### 2.2 Hard Negative Mining
#
#
# You will now implement the `TripletLoss`.<br>
# As explained in the lecture, loss is composed of two terms. One term utilizes the mean of all the non duplicates, the second utilizes the *closest negative*. Our loss expression is then:
#
# \begin{align}
# \mathcal{Loss_1(A,P,N)} &=\max \left( -cos(A,P) + mean_{neg} +\alpha, 0\right) \\
# \mathcal{Loss_2(A,P,N)} &=\max \left( -cos(A,P) + closest_{neg} +\alpha, 0\right) \\
# \mathcal{Loss(A,P,N)} &= mean(Loss_1 + Loss_2) \\
# \end{align}
#
#
# Further, two sets of instructions are provided. The first set provides a brief description of the task. If that set proves insufficient, a more detailed set can be displayed.
#
# <a name='ex03'></a>
# ### Exercise 03
#
# **Instructions (Brief):** Here is a list of things you should do: <br>
#
# - As this will be run inside trax, use `fastnp.xyz` when using any `xyz` numpy function
# - Use `fastnp.dot` to calculate the similarity matrix $v_1v_2^T$ of dimension `batch_size` x `batch_size`
# - Take the score of the duplicates on the diagonal `fastnp.diagonal`
# - Use the `trax` functions `fastnp.eye` and `fastnp.maximum` for the identity matrix and the maximum.
# + [markdown] colab_type="text" id="GWsX-Wz3piup"
# <details>
# <summary>
# <font size="3" color="darkgreen"><b>More Detailed Instructions </b></font>
# </summary>
# We'll describe the algorithm using a detailed example. Below, V1, V2 are the output of the normalization blocks in our model. Here we will use a batch_size of 4 and a d_model of 3. As explained in lecture, the inputs, Q1, Q2 are arranged so that corresponding inputs are duplicates while non-corresponding entries are not. The outputs will have the same pattern.
# <img src = "C3_W4_triploss1.png" style="width:1021px;height:229px;"/>
# This testcase arranges the outputs, v1,v2, to highlight different scenarios. Here, the first two outputs V1[0], V2[0] match exactly - so the model is generating the same vector for Q1[0] and Q2[0] inputs. The second outputs differ, circled in orange, we set, V2[1] is set to match V2[**2**], simulating a model which is generating very poor results. V1[3] and V2[3] match exactly again while V1[4] and V2[4] are set to be exactly wrong - 180 degrees from each other, circled in blue.
#
# The first step is to compute the cosine similarity matrix or `score` in the code. As explained in lecture, this is $$V_1 V_2^T$$ This is generated with `fastnp.dot`.
# <img src = "C3_W4_triploss2.png" style="width:959px;height:236px;"/>
# The clever arrangement of inputs creates the data needed for positive *and* negative examples without having to run all pair-wise combinations. Because Q1[n] is a duplicate of only Q2[n], other combinations are explicitly created negative examples or *Hard Negative* examples. The matrix multiplication efficiently produces the cosine similarity of all positive/negative combinations as shown above on the left side of the diagram. 'Positive' are the results of duplicate examples and 'negative' are the results of explicitly created negative examples. The results for our test case are as expected, V1[0]V2[0] match producing '1' while our other 'positive' cases (in green) don't match well, as was arranged. The V2[2] was set to match V1[3] producing a poor match at `score[2,2]` and an undesired 'negative' case of a '1' shown in grey.
#
# With the similarity matrix (`score`) we can begin to implement the loss equations. First, we can extract $$cos(A,P)$$ by utilizing `fastnp.diagonal`. The goal is to grab all the green entries in the diagram above. This is `positive` in the code.
#
# Next, we will create the *closest_negative*. This is the nonduplicate entry in V2 that is closest (has largest cosine similarity) to an entry in V1. Each row, n, of `score` represents all comparisons of the results of Q1[n] vs Q2[x] within a batch. A specific example in our testcase is row `score[2,:]`. It has the cosine similarity of V1[2] and V2[x]. The *closest_negative*, as was arranged, is V2[2] which has a score of 1. This is the maximum value of the 'negative' entries (blue entries in the diagram).
#
# To implement this, we need to pick the maximum entry on a row of `score`, ignoring the 'positive'/green entries. To avoid selecting the 'positive'/green entries, we can make them larger negative numbers. Multiply `fastnp.eye(batch_size)` with 2.0 and subtract it out of `scores`. The result is `negative_without_positive`. Now we can use `fastnp.max`, row by row (axis=1), to select the maximum which is `closest_negative`.
#
# Next, we'll create *mean_negative*. As the name suggests, this is the mean of all the 'negative'/blue values in `score` on a row by row basis. We can use `fastnp.eye(batch_size)` and a constant, this time to create a mask with zeros on the diagonal. Element-wise multiply this with `score` to get just the 'negative values. This is `negative_zero_on_duplicate` in the code. Compute the mean by using `fastnp.sum` on `negative_zero_on_duplicate` for `axis=1` and divide it by `(batch_size - 1)` . This is `mean_negative`.
#
# Now, we can compute loss using the two equations above and `fastnp.maximum`. This will form `triplet_loss1` and `triplet_loss2`.
#
# `triple_loss` is the `fastnp.mean` of the sum of the two individual losses.
#
# Once you have this code matching the expected results, you can clip out the section between ### START CODE HERE and ### END CODE HERE it out and insert it into TripletLoss below.
#
#
# <\details>
# + colab={} colab_type="code" id="oJM8EQiopiuv"
# UNQ_C3 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: TripletLossFn
def TripletLossFn(v1, v2, margin=0.25):
"""Custom Loss function.
Args:
v1 (numpy.ndarray): Array with dimension (batch_size, model_dimension) associated to Q1.
v2 (numpy.ndarray): Array with dimension (batch_size, model_dimension) associated to Q2.
margin (float, optional): Desired margin. Defaults to 0.25.
Returns:
jax.interpreters.xla.DeviceArray: Triplet Loss.
"""
### START CODE HERE (Replace instances of 'None' with your code) ###
# use fastnp to take the dot product of the two batches (don't forget to transpose the second argument)
scores = fastnp.dot(v1, v2.T) # pairwise cosine sim
# print("scores:", scores)
# calculate new batch size
batch_size = len(scores)
# use fastnp to grab all postive `diagonal` entries in `scores`
positive = fastnp.diagonal(scores) # the positive ones (duplicates)
# print("positive ", positive)
# multiply `fastnp.eye(batch_size)` with 2.0 and subtract it out of `scores`
negative_without_positive = scores - fastnp.eye(batch_size) * 2.0
# print("negative_without_positive ", negative_without_positive)
# take the row by row `max` of `negative_without_positive`.
# Hint: negative_without_positive.max(axis = [?])
closest_negative = negative_without_positive.max(axis=1)
# print("closest_negative ", closest_negative)
# subtract `fastnp.eye(batch_size)` out of 1.0 and do element-wise multiplication with `scores`
negative_zero_on_duplicate = (1.0-fastnp.eye(batch_size)) * scores
# print("negative_zero_on_duplicate ", negative_zero_on_duplicate)
# use `fastnp.sum` on `negative_zero_on_duplicate` for `axis=1` and divide it by `(batch_size - 1)`
mean_negative = fastnp.sum(negative_zero_on_duplicate,axis=1)/(batch_size - 1)
# print("mean_negative ", mean_negative)
# compute `fastnp.maximum` among 0.0 and `A`
# A = subtract `positive` from `margin` and add `closest_negative`
triplet_loss1 = fastnp.maximum(margin - positive + closest_negative,0.0)
# print("triplet_loss1 ", triplet_loss1)
# compute `fastnp.maximum` among 0.0 and `B`
# B = subtract `positive` from `margin` and add `mean_negative`
triplet_loss2 = fastnp.maximum(margin - positive + mean_negative,0.0)
# print("triplet_loss2 ", triplet_loss2)
# add the two losses together and take the `fastnp.mean` of it
triplet_loss = fastnp.mean(triplet_loss1 + triplet_loss2)
# print("triplet_loss ", triplet_loss)
### END CODE HERE ###
return triplet_loss
# -
v1 = np.array([[0.26726124, 0.53452248, 0.80178373],[0.5178918 , 0.57543534, 0.63297887]])
v2 = np.array([[ 0.26726124, 0.53452248, 0.80178373],[-0.5178918 , -0.57543534, -0.63297887]])
TripletLossFn(v2,v1)
print("Triplet Loss:", TripletLossFn(v2,v1))
# **Expected Output:**
# ```CPP
# Triplet Loss: 0.5
# ```
# + [markdown] colab_type="text" id="r974ozuHYAom"
# To make a layer out of a function with no trainable variables, use `tl.Fn`.
# -
from functools import partial
def TripletLoss(margin=0.25):
triplet_loss_fn = partial(TripletLossFn, margin=margin)
return tl.Fn('TripletLoss', triplet_loss_fn)
# + [markdown] colab_type="text" id="lsvjaCQ6wt02"
# <a name='3'></a>
#
# # Part 3: Training
#
# Now you are going to train your model. As usual, you have to define the cost function and the optimizer. You also have to feed in the built model. Before, going into the training, we will use a special data set up. We will define the inputs using the data generator we built above. The lambda function acts as a seed to remember the last batch that was given. Run the cell below to get the question pairs inputs.
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="iPk7gh-nzCBg" outputId="a2e8525d-f89a-4d9d-c0d6-bd7406f0246a"
batch_size = 256
train_generator = data_generator(train_Q1, train_Q2, batch_size, vocab['<PAD>'])
val_generator = data_generator(val_Q1, val_Q2, batch_size, vocab['<PAD>'])
print('train_Q1.shape ', train_Q1.shape)
print('val_Q1.shape ', val_Q1.shape)
# + [markdown] colab_type="text" id="IgFMfH5awt07"
# <a name='3.1'></a>
#
# ### 3.1 Training the model
#
# You will now write a function that takes in your model and trains it. To train your model you have to decide how many times you want to iterate over the entire data set; each iteration is defined as an `epoch`. For each epoch, you have to go over all the data, using your training iterator.
#
# <a name='ex04'></a>
# ### Exercise 04
#
# **Instructions:** Implement the `train_model` below to train the neural network above. Here is a list of things you should do, as already shown in lecture 7:
#
# - Create `TrainTask` and `EvalTask`
# - Create the training loop `trax.supervised.training.Loop`
# - Pass in the following depending on the context (train_task or eval_task):
# - `labeled_data=generator`
# - `metrics=[TripletLoss()]`,
# - `loss_layer=TripletLoss()`
# - `optimizer=trax.optimizers.Adam` with learning rate of 0.01
# - `lr_schedule=lr_schedule`,
# - `output_dir=output_dir`
#
#
# You will be using your triplet loss function with Adam optimizer. Please read the [trax](https://trax-ml.readthedocs.io/en/latest/trax.optimizers.html?highlight=adam#trax.optimizers.adam.Adam) documentation to get a full understanding.
#
# This function should return a `training.Loop` object. To read more about this check the [docs](https://trax-ml.readthedocs.io/en/latest/trax.supervised.html?highlight=loop#trax.supervised.training.Loop).
# + colab={} colab_type="code" id="_kbtfz4T_m7x"
lr_schedule = trax.lr.warmup_and_rsqrt_decay(400, 0.01)
# UNQ_C4 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: train_model
def train_model(Siamese, TripletLoss, lr_schedule, train_generator=train_generator, val_generator=val_generator, output_dir='model/'):
"""Training the Siamese Model
Args:
Siamese (function): Function that returns the Siamese model.
TripletLoss (function): Function that defines the TripletLoss loss function.
lr_schedule (function): Trax multifactor schedule function.
train_generator (generator, optional): Training generator. Defaults to train_generator.
val_generator (generator, optional): Validation generator. Defaults to val_generator.
output_dir (str, optional): Path to save model to. Defaults to 'model/'.
Returns:
trax.supervised.training.Loop: Training loop for the model.
"""
output_dir = os.path.expanduser(output_dir)
### START CODE HERE (Replace instances of 'None' with your code) ###
train_task = training.TrainTask(
labeled_data=train_generator, # Use generator (train)
loss_layer=TripletLoss(), # Use triplet loss. Don't forget to instantiate this object
optimizer=trax.optimizers.Adam(learning_rate=0.01), # Don't forget to add the learning rate parameter
lr_schedule=lr_schedule, # Use Trax multifactor schedule function
)
eval_task = training.EvalTask(
labeled_data=val_generator, # Use generator (val)
metrics=[TripletLoss()], # Use triplet loss. Don't forget to instantiate this object
)
### END CODE HERE ###
training_loop = training.Loop(Siamese(),
train_task,
eval_task=eval_task,
output_dir=output_dir)
return training_loop
# + colab={"base_uri": "https://localhost:8080/", "height": 391} colab_type="code" id="-3KXjmBo_6Xa" outputId="9d57f731-1534-4218-e744-783359d5cd19"
train_steps = 5
training_loop = train_model(Siamese, TripletLoss, lr_schedule)
training_loop.run(train_steps)
# -
# The model was only trained for 5 steps due to the constraints of this environment. For the rest of the assignment you will be using a pretrained model but now you should understand how the training can be done using Trax.
# + [markdown] colab_type="text" id="abKPe7d4wt1C"
# <a name='4'></a>
#
# # Part 4: Evaluation
#
# <a name='4.1'></a>
#
# ### 4.1 Evaluating your siamese network
#
# In this section you will learn how to evaluate a Siamese network. You will first start by loading a pretrained model and then you will use it to predict.
# + colab={} colab_type="code" id="3OtmlEuOwt1D"
# Loading in the saved model
model = Siamese()
model.init_from_file('model.pkl.gz')
# + [markdown] colab_type="text" id="QDi4MBiKpivF"
# <a name='4.2'></a>
# ### 4.2 Classify
# To determine the accuracy of the model, we will utilize the test set that was configured earlier. While in training we used only positive examples, the test data, Q1_test, Q2_test and y_test, is setup as pairs of questions, some of which are duplicates some are not.
# This routine will run all the test question pairs through the model, compute the cosine simlarity of each pair, threshold it and compare the result to y_test - the correct response from the data set. The results are accumulated to produce an accuracy.
#
#
# <a name='ex05'></a>
# ### Exercise 05
#
# **Instructions**
# - Loop through the incoming data in batch_size chunks
# - Use the data generator to load q1, q2 a batch at a time. **Don't forget to set shuffle=False!**
# - copy a batch_size chunk of y into y_test
# - compute v1, v2 using the model
# - for each element of the batch
# - compute the cos similarity of each pair of entries, v1[j],v2[j]
# - determine if d > threshold
# - increment accuracy if that result matches the expected results (y_test[j])
# - compute the final accuracy and return
#
# Due to some limitations of this environment, running classify multiple times may result in the kernel failing. If that happens *Restart Kernal & clear output* and then run from the top. During development, consider using a smaller set of data to reduce the number of calls to model().
# + colab={} colab_type="code" id="K-h6ZH507fUm"
# UNQ_C5 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: classify
def classify(test_Q1, test_Q2, y, threshold, model, vocab, data_generator=data_generator, batch_size=64):
"""Function to test the accuracy of the model.
Args:
test_Q1 (numpy.ndarray): Array of Q1 questions.
test_Q2 (numpy.ndarray): Array of Q2 questions.
y (numpy.ndarray): Array of actual target.
threshold (float): Desired threshold.
model (trax.layers.combinators.Parallel): The Siamese model.
vocab (collections.defaultdict): The vocabulary used.
data_generator (function): Data generator function. Defaults to data_generator.
batch_size (int, optional): Size of the batches. Defaults to 64.
Returns:
float: Accuracy of the model.
"""
accuracy = 0
### START CODE HERE (Replace instances of 'None' with your code) ###
for i in range(0, len(test_Q1), batch_size):
# Call the data generator (built in Ex 01) with shuffle=False using next()
# use batch size chuncks of questions as Q1 & Q2 arguments of the data generator. e.g x[i:i + batch_size]
# Hint: use `vocab['<PAD>']` for the `pad` argument of the data generator
q1, q2 = next(data_generator(test_Q1[i:i+batch_size], test_Q2[i:i+batch_size], batch_size=batch_size,pad=vocab['<PAD>'], shuffle=False))
# use batch size chuncks of actual output targets (same syntax as example above)
y_test = y[i:i+batch_size]
# Call the model
v1, v2 = model((q1,q2))
for j in range(batch_size):
# take dot product to compute cos similarity of each pair of entries, v1[j], v2[j]
# don't forget to transpose the second argument
d = fastnp.dot(v1[j], v2[j].T)
# is d greater than the threshold?
res = d>threshold
# increment accurancy if y_test is equal `res`
accuracy += (y_test[j] == res)
# compute accuracy using accuracy and total length of test questions
accuracy = accuracy / len(test_Q1)
### END CODE HERE ###
return accuracy
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="yeQjHxkfpivH" outputId="103b8449-896f-403d-f011-583df70afdae"
# this takes around 1 minute
accuracy = classify(Q1_test,Q2_test, y_test, 0.7, model, vocab, batch_size = 512)
print("Accuracy", accuracy)
# + [markdown] colab_type="text" id="CsokYZwhpivJ"
# **Expected Result**
# Accuracy ~0.69
# + [markdown] colab_type="text" id="4-STC44Ywt1I"
# <a name='5'></a>
#
# # Part 5: Testing with your own questions
#
# In this section you will test the model with your own questions. You will write a function `predict` which takes two questions as input and returns $1$ or $0$ depending on whether the question pair is a duplicate or not.
#
# But first, we build a reverse vocabulary that allows to map encoded questions back to words:
# + [markdown] colab_type="text" id="21h3Y0FNpivK"
# Write a function `predict`that takes in two questions, the model, and the vocabulary and returns whether the questions are duplicates ($1$) or not duplicates ($0$) given a similarity threshold.
#
# <a name='ex06'></a>
# ### Exercise 06
#
#
# **Instructions:**
# - Tokenize your question using `nltk.word_tokenize`
# - Create Q1,Q2 by encoding your questions as a list of numbers using vocab
# - pad Q1,Q2 with next(data_generator([Q1], [Q2],1,vocab['<PAD>']))
# - use model() to create v1, v2
# - compute the cosine similarity (dot product) of v1, v2
# - compute res by comparing d to the threshold
#
# + colab={} colab_type="code" id="kg0wQ8qhpivL"
# UNQ_C6 (UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: predict
def predict(question1, question2, threshold, model, vocab, data_generator=data_generator, verbose=False):
"""Function for predicting if two questions are duplicates.
Args:
question1 (str): First question.
question2 (str): Second question.
threshold (float): Desired threshold.
model (trax.layers.combinators.Parallel): The Siamese model.
vocab (collections.defaultdict): The vocabulary used.
data_generator (function): Data generator function. Defaults to data_generator.
verbose (bool, optional): If the results should be printed out. Defaults to False.
Returns:
bool: True if the questions are duplicates, False otherwise.
"""
### START CODE HERE (Replace instances of 'None' with your code) ###
# use `nltk` word tokenize function to tokenize
q1 = nltk.word_tokenize(question1) # tokenize
q2 = nltk.word_tokenize(question2) # tokenize
Q1, Q2 = [], []
for word in q1: # encode q1
# increment by checking the 'word' index in `vocab`
Q1 += [vocab[word]]
for word in q2: # encode q2
# increment by checking the 'word' index in `vocab`
Q2 += [vocab[word]]
# Call the data generator (built in Ex 01) using next()
# pass [Q1] & [Q2] as Q1 & Q2 arguments of the data generator. Set batch size as 1
# Hint: use `vocab['<PAD>']` for the `pad` argument of the data generator
Q1, Q2 = next(data_generator([Q1], [Q2], batch_size=1,pad=vocab['<PAD>'], shuffle=False))
# Call the model
v1, v2 = model((Q1,Q2))
# take dot product to compute cos similarity of each pair of entries, v1, v2
# don't forget to transpose the second argument
d = fastnp.dot(v1[0], v2[0].T)
# is d greater than the threshold?
res = d>threshold
### END CODE HERE ###
if(verbose):
print("Q1 = ", Q1, "\nQ2 = ", Q2)
print("d = ", d)
print("res = ", res)
return res
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="Raojyhw3z7HE" outputId="b0907aaf-63c0-448d-99b0-012359381a97"
# Feel free to try with your own questions
question1 = "When will I see you?"
question2 = "When can I see you again?"
# 1 means it is duplicated, 0 otherwise
predict(question1 , question2, 0.7, model, vocab, verbose = True)
# + [markdown] colab_type="text" id="7OEKCa_hpivP"
# ##### Expected Output
# If input is:
# ```CPP
# question1 = "When will I see you?"
# question2 = "When can I see you again?"
# ```
#
# Output is (d may vary a bit):
# ```CPP
# Q1 = [[585 76 4 46 53 21 1 1]]
# Q2 = [[ 585 33 4 46 53 7280 21 1]]
# d = 0.88113236
# res = True
# True
# ```
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="DZccIQ_lpivQ" outputId="3ed0af7e-5d44-4eb3-cebe-d6f74abe3e41"
# Feel free to try with your own questions
question1 = "Do they enjoy eating the dessert?"
question2 = "Do they like hiking in the desert?"
# 1 means it is duplicated, 0 otherwise
predict(question1 , question2, 0.7, model, vocab, verbose=True)
# + [markdown] colab_type="text" id="lWrt-yCMpivS"
# ##### Expected output
#
# If input is:
# ```CPP
# question1 = "Do they enjoy eating the dessert?"
# question2 = "Do they like hiking in the desert?"
# ```
#
# Output (d may vary a bit):
#
# ```CPP
# Q1 = [[ 443 1145 3159 1169 78 29017 21 1]]
# Q2 = [[ 443 1145 60 15302 28 78 7431 21]]
# d = 0.477536
# res = False
# False
# ```
# + [markdown] colab_type="text" id="NAfV3l5Zwt1L"
# You can see that the Siamese network is capable of catching complicated structures. Concretely it can identify question duplicates although the questions do not have many words in common.
#
# + [markdown] colab_type="text" id="FsE8tdTLwt1M"
# <a name='6'></a>
#
# ### <span style="color:blue"> On Siamese networks </span>
#
# Siamese networks are important and useful. Many times there are several questions that are already asked in quora, or other platforms and you can use Siamese networks to avoid question duplicates.
#
# Congratulations, you have now built a powerful system that can recognize question duplicates. In the next course we will use transformers for machine translation, summarization, question answering, and chatbots.
#
|
3 - NLP with Sequence Models/Week 4/C3_W4_Assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Amazon Lookout for Metrics에 대한 기록 데이터 백테스팅
#
# Amazon Lookout for Metrics는 기록 정보에 대한 백테스팅을 지원하며 본 노트북에서는 방금까지 준비한 동일 데이터 세트에서 해당 기능을 시연할 것입니다. 백테스팅 작업이 완료되면 기록 데이터의 마지막 30%에 대해 Amazon Lookout for Metrics가 감지한 모든 이상치를 살펴볼 수 있습니다. 데이터 스트리밍을 신규로 시작했을 때 Amazon Lookout for Metrics에서 보게될 결과를 여기서 미리 파악해볼 수 있습니다. **참고: 실시간 데이터를 활용하려면 탐지기를 신규 생성해야 합니다. 백테스팅은 탐색 용도입니다.**
#
# 이 노트북은 `1.PrereqSetupData.ipynb`의 사전 요구 사항을 이미 완료했다고 가정합니다. 그렇지 않은 경우 돌아가서 완료 먼저 하세요.
#
# ## 초기 단계
#
# 먼저 이전 노트북에서 만든 변수를 복원한 다음 필요한 라이브러리를 임포트합니다.
# %store -r
# 이전 노트북과 마찬가지로 SDK를 통해 AWS와 연결합니다.
import boto3
import utility
# **아래의 셀에서 오류가 발생하는 경우:** 이는 완전히 정상이며, Boto3가 단순히 최신 버전이 아님을 의미합니다. 해당 문제를 해결하려면 오류가 발생한 셀 아래의 셀로 이동하여 SageMaker 내부의 Boto3 최신 버전을 업데이트하도록 하세요. SageMaker 노트북을 사용하지 않는다면 Python 환경에 대한 지침을 따르세요.
#
# 업그레이드 셀을 실행한 후 상단 메뉴에서 `Kernel`을 클릭한 다음 `Restart Kernel`을 클릭하세요. 완료되면 본 노트북의 맨 윗부분부터 다시 시작하세요.
#
# **필요한 경우가 아니면 업그레이드 셀을 실행하지 마세요**
L4M = boto3.client("lookoutmetrics", region_name="us-west-2")
# 이것은 선택 사항입니다. 필요하지 않은 경우 실행하지 마세요.
# !pip install --upgrade boto3
# ## 탐지기 생성하기
#
# 이제 기본적인 외부 리소스가 준비되었으므로 `Detector`부터 생성하면서 Amazon Lookout for Metrics를 시작해봅시다.
#
# ### 탐지기
#
# 이상치를 탐지하기 위해 Amazon Lookout for Metrics는 기계 학습 모델을 원천 데이터로 훈련하여 구축합니다. `Detector`라고 하는 이 모델은 데이터와 사용 사례에 가장 적합한 기계 학습 알고리즘으로 자동 훈련됩니다. 훈련을 위해 기록 데이터를 제공하거나 (데이터가 있는 경우) 아니면 실시간 데이터로 시작해서 Amazon Lookout for Metrics는 데이터를 입수하는 동시에 모델을 훈련시킵니다. 본 `Backtesting` 예제에서는 기록 데이터만 제공합니다.
#
# 이 예제에서는 기록 데이터의 S3 위치를 지정하지만 `Continous` 탐지기를 생성하는 경우 Amazon Lookout for Metrics가 지속적으로 신규 데이터를 모니터링할 수 있게 Amazon S3 위치를 지정해줘야 합니다. `Detector`를 생성하면서 `탐지 도메인`과 `극단값 탐지 빈도`도 지정합니다.
#
# 탐지기를 얼마나 자주 활성화해서 신규 데이터를 찾고, 분석을 실행하고, 흥미로운 결과를 알려줄지 `극단값 탐지 빈도`로 정의합니다.
# +
project = "initial-lookoutmetrics-backtesting-test"
frequency = "PT1H" # 'P1D', 'PT1H', 'PT10M'과 'PT5M' 중 하나, 이것은 1시간마다 반복함을 의미합니다
# +
response = L4M.create_anomaly_detector(
AnomalyDetectorName = project + "-detector",
AnomalyDetectorDescription = "My Detector",
AnomalyDetectorConfig = {
"AnomalyDetectorFrequency": "PT1H",
},
)
anomaly_detector_arn = response["AnomalyDetectorArn"]
# print(anomaly_detector_arn)
# -
# ## 메트릭 정의하기
#
# ### 측정값과 차원
#
# `Measures`는 고객이 극단값을 탐지하고자하는 변수 또는 핵심 성과 지표이고, `Dimensions`은 측정값에 대한 범주 정보를 나타내는 메타 데이터입니다.
#
# 이 전자 상거래 예제에서 조회 수와 수익은 측정값이고 플랫폼과 시장은 차원입니다. 고객은 모든 플랫폼과 시장, 이 두 조합에 대해 조회 수 또는 수익의 이상치가 있는지 데이터를 모니터링할 수 있습니다. 데이터 집합 당 최대 5개의 측정값과 5개의 차원을 지정할 수 있습니다.
#
# ### 메트릭
#
# 탐지기를 생성하고 측정값과 차원을 매핑한 후 Amazon Lookout for Metrics는 이러한 측정값과 차원의 각 조합을 분석합니다. 위의 예제에서 시장에는 7개의 고유한 값 (us, jp, de 등) 이 있고 플랫폼에는 3개의 고유한 값 (모바일 웹, 모바일 앱, PC 웹) 이 있으므로 총 21개의 고유한 조합이 있습니다. 각 측정값과 차원의 고유한 조합은 (예: us / 모바일 앱 / 수익) `메트릭` 시계열을 정의합니다. 이 경우 21가지의 차원과 2가지의 측정값, 즉 총 42개의 `메트릭` 시계열이 존재합니다.
#
# Amazon Lookout for Metrics는 가장 세분화된 수준에서 이상치를 탐지하므로 데이터의 예기치 않은 동작을 정확히 찾아낼 수 있습니다.
#
# ### 데이터셋
#
# 측정값, 차원과 메트릭은 원천 데이터의 Amazon S3 위치, 해당 Amazon S3 위치에 대한 읽기와 쓰기 권한이 모두 있는 IAM 역할, 원천 데이터 위치에서 데이터 수집 속도 (업로드 빈도와 데이터 수집 지연의 정도)를 포함하는 `데이터셋`에 매핑됩니다.
# 이제 S3의 백테스트 데이터와 연결된 탐지기의 메트릭 집합을 만듭니다.
#
# 먼저 아래 셀은 S3에 대한 백테스팅 경로를 생성한 다음 인자를 API로 전달됩니다.
s3_path_backtest = "s3://" + s3_bucket + "/ecommerce/backtest/"
# print(s3_path_backtest)
# +
params = {
"AnomalyDetectorArn": anomaly_detector_arn,
"MetricSetName": project + "-metric-set-1",
"MetricList": [
{
"MetricName": "views",
"AggregationFunction": "SUM",
},
{
"MetricName": "revenue",
"AggregationFunction": "SUM",
},
],
"DimensionList": ["platform", "marketplace"],
"TimestampColumn" : {
"ColumnName": "timestamp",
"ColumnFormat": "yyyy-MM-dd HH:mm:ss",
},
# "Delay": 120, # 현재 시간으로부터 아래의 탐지 빈도에 따라 최신 데이터를 읽기 전 탐지기가 대기하는 시간 (초)
"MetricSetFrequency": frequency,
"MetricSource": {
"S3SourceConfig": {
"RoleArn": role_arn,
"HistoricalDataPathList": [
s3_path_backtest,
],
# "TemplatedPathList": [
# s3_path_format,
# ],
"FileFormatDescriptor": {
"CsvFormatDescriptor": {
"FileCompression": "NONE",
"Charset": "UTF-8",
"ContainsHeader": True,
"Delimiter": ",",
# "HeaderList" : [
# "platform",
# "marketplace",
# "timestamp",
# "views",
# "revenue"
# ],
"QuoteSymbol" : '"'
},
}
}
},
}
# print(params)
# -
# 아래 셀은 해당 인자를 가져와서 `MetricSet`을 생성합니다. 이제 다음 단계에서 활성화시킬 준비가 됐습니다.
# +
response = L4M.create_metric_set(**params)
metric_set_arn = response["MetricSetArn"]
# print(metric_set_arn)
# -
# ## 탐지기 활성화하고 백테스팅 실행하기
#
# 이제 `MetricSet`이 지정되었으므로 백테스트 탐지기를 활성화해서 백테스팅을 시작할 준비가 되었습니다. 백테스팅 과정은 약 25분 정도 소요되므로 간식을 먹으며 휴식을 취하고 못읽은 신문 기사를 마저 읽으세요. `BACK_TEST_ACTIVE`라고 표시되면 서비스의 모델 훈련이 완료된 것이며 이제 따로 빼어 둔 기간에 대해 평가를 시작합니다.
L4M.back_test_anomaly_detector(AnomalyDetectorArn=anomaly_detector_arn)
# 참고: 셀은 먼저 활성화 중이라고 출력한 다음 백테스팅 작업이 활성화되었음을 알려줍니다. 이것은 단지 추론 프로세스를 실행하고 있음을 의미하며 이상치를 찾기 전에 셀이 완전히 완료될 때까지 더 많은 시간을 기다려야 합니다.
_ = utility.wait_anomaly_detector(L4M, anomaly_detector_arn)
# ## 결과를 검증하기
#
# 백테스팅이 완료된 다음 콘솔을 통해 기록 이상치를 시각적으로 확인하거나 아래 명령을 실행하여 결과를 검증할 수 있습니다. 그러나 콘솔에서 탐색을 시작하는 편이 좋습니다. 콘솔은 나중에 온라인 모드에서 경고를 보고 이해할 수 있는 도구가 되므로 이런 방식으로 프로세스에 익숙해질 수 있습니다.
# +
anomaly_groups = []
next_token = None
first_response = None
while True:
params = {
"AnomalyDetectorArn": anomaly_detector_arn,
"SensitivityThreshold": 50,
"MaxResults": 100,
}
if next_token:
params["NextToken"] = next_token
response = L4M.list_anomaly_group_summaries(**params)
if first_response is None:
first_response = response
anomaly_groups += response["AnomalyGroupSummaryList"]
if "NextToken" in response:
next_token = response["NextToken"]
continue
break
# print(first_response)
# +
import pandas as pd
anomaly_groups = pd.DataFrame(anomaly_groups)
anomaly_groups.head()
# -
# 특정 이상치 그룹을 더 깊이 살펴보려면 관심있는 이상치 그룹을 단순 선택하고 해당 시계열를 세분화 분석해봅니다. 여기에서는 해당 List의 첫 번째 이상치 그룹을 사용했습니다.
# ## 리소스 정리하기
#
# 백테스팅을 완료하면 생성한 리소스 정리를 시작할 수 있습니다. 정리하기 전 Amazon Lookout for Metrics 콘솔의 "이상치" 페이지를 방문하여 탐지된 이상치를 시각적으로 확인해보세요.
#
# 본 작업을 진행하면 생성한 모든 리소스가 지워지므로 전체 삭제하려는 것이 확실할 때에만 실행하세요.
#
# **참고: 아래에서 역할을 삭제했다면 연속 탐지기를 구축하기 전에 다시 만들어야합니다.**
# +
answer = input("Delete resources? (y/n)")
if answer == "y":
delete_resources = True
else:
delete_resources = False
if delete_resources:
role_name = "L4MTestRole"
L4M.delete_anomaly_detector(AnomalyDetectorArn=anomaly_detector_arn)
utility.wait_delete_anomaly_detector(L4M, anomaly_detector_arn)
utility.delete_iam_role(role_name)
else:
print("Not deteleting resources.")
|
getting_started/2.BacktestingWithHistoricalData.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Creates Tensorflow Graphs for spark-nlp DL Annotators and Models
#
# +
import numpy as np
import os
import tensorflow as tf
import string
import random
import math
import sys
import shutil
from ner_model import NerModel
from dataset_encoder import DatasetEncoder
from ner_model_saver import NerModelSaver
from pathlib import Path
# -
# ## SETTINGS
# +
use_contrib = False if os.name == 'nt' else True
name_prefix = 'blstm-noncontrib' if not use_contrib else 'blstm'
# -
def create_graph(ntags, embeddings_dim, nchars, lstm_size = 128):
if sys.version_info[0] != 3 or sys.version_info[1] >= 7:
print('Python 3.7 or above not supported by tensorflow')
return
if tf.__version__ != '1.12.0':
print('Spark NLP is compiled with Tensorflo 1.12.0. Please use such version.')
return
tf.reset_default_graph()
model_name = name_prefix+'_{}_{}_{}_{}'.format(ntags, embeddings_dim, lstm_size, nchars)
with tf.Session() as session:
ner = NerModel(session=None, use_contrib=use_contrib)
ner.add_cnn_char_repr(nchars, 25, 30)
ner.add_bilstm_char_repr(nchars, 25, 30)
ner.add_pretrained_word_embeddings(embeddings_dim)
ner.add_context_repr(ntags, lstm_size, 3)
ner.add_inference_layer(True)
ner.add_training_op(5)
ner.init_variables()
saver = tf.train.Saver()
file_name = model_name + '.pb'
tf.train.write_graph(ner.session.graph, './', file_name, False)
ner.close()
session.close()
# ### Attributes info
# - 1st attribute: max number of tags (Must be at least equal to the number of unique labels, including O if IOB)
# - 2nd attribute: embeddings dimension
# - 3rd attribute: max number of characters processed (Must be at least the largest possible amount of characters)
# - 4th attribute: LSTM Size (128)
create_graph(80, 200, 125)
# create_graph(10, 200, 100)
# create_graph(10, 300, 100)
# create_graph(10, 768, 100)
# create_graph(10, 1024, 100)
# create_graph(25, 300, 100)
|
python/tensorflow/ner/create_models.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Self-Driving Car Engineer Nanodegree
#
#
# ## Project: Build a Traffic Sign Recognition Classifier
#
# In this project, deep neural networks and convolutional neural networks were used to classify traffic signs. I have trained and validated the model so it can classify traffic sign images using the [German Traffic Sign Dataset](http://benchmark.ini.rub.de/?section=gtsrb&subsection=dataset). After the model is trained, I then tried out my model on images of German traffic signs which I downloaded from the web.
# ---
# ## Step 0: Load The Data
# +
# Import pickle library to serialize and de-serialize the data
import pickle
# Path to training, testing and validation data
training_filepath = '../data/train.p'
validation_filepath='../data/valid.p'
testing_filepath = '../data/test.p'
# Load the training, testing and validation data
with open(training_filepath, mode='rb') as f:
train = pickle.load(f)
with open(validation_filepath, mode='rb') as f:
valid = pickle.load(f)
with open(testing_filepath, mode='rb') as f:
test = pickle.load(f)
# Load features (input data) and lables (output data) seperately in an array for training, testing and validation data
X_train, y_train = train['features'], train['labels']
X_valid, y_valid = valid['features'], valid['labels']
X_test, y_test = test['features'], test['labels']
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape, X_valid.shape, y_valid.shape)
# -
# ---
#
# ## Step 1: Dataset Summary & Exploration
#
# The pickled data is a dictionary with 4 key/value pairs:
#
# - `'features'` is a 4D array containing raw pixel data of the traffic sign images, (num examples, width, height, channels).
# - `'labels'` is a 1D array containing the label/class id of the traffic sign. The file `signnames.csv` contains id -> name mappings for each id.
# - `'sizes'` is a list containing tuples, (width, height) representing the original width and height the image.
# - `'coords'` is a list containing tuples, (x1, y1, x2, y2) representing coordinates of a bounding box around the sign in the image. **THESE COORDINATES ASSUME THE ORIGINAL IMAGE. THE PICKLED DATA CONTAINS RESIZED VERSIONS (32 by 32) OF THESE IMAGES**
# ### Basic Summary of the Data Set Using Python, Numpy and/or Pandas
# +
# Import libraries
import pandas as pd
import numpy as np
# Number of training examples
train_num = len(X_train)
# Number of validation examples
validation_num = len(X_valid)
# Number of testing examples.
test_num = len(X_test)
# The shape of an traffic sign image
image_shape = X_train[0].shape
# The number of unique classes/labels there are in the dataset.
classes_num = len(pd.Series(y_train).unique())
print("Size of the training set =", train_num)
print("Size of the validation set =", validation_num)
print("Size of the testing set =", test_num)
print("The shape of a traffic sign image =", image_shape)
print("The number of unique classes/labels in the data set is =", classes_num)
# -
# ### Include an exploratory visualization of the dataset
# Visualize the German Traffic Signs Dataset using the pickled file(s).
#
# The [Matplotlib](http://matplotlib.org/) [examples](http://matplotlib.org/examples/index.html) and [gallery](http://matplotlib.org/gallery.html) pages are a great resource for doing visualizations in Python.
# +
### Data exploration visualization code goes here.
import matplotlib.pyplot as plt
import random
# Visualizations will be shown in the notebook.
# %matplotlib inline
# Randomly select the image from dataset and disply the image
index = random.randint(0, len(X_train))
image=X_train[index]
print(y_train[index])
plt.figure()
plt.imshow(image)
# +
# Plot the histogram of traffic sign signals for training data
plt.hist(y_train, bins=classes_num)
plt.title('Histogram of Traffic Sign Signals (Training set)') # Display the title
plt.xlabel('Traffic Signs') # Display the x-axis
plt.ylabel('Total Count') # Display the y-axis
plt.show() # Plot the histogram
# Plot the histogram of traffic sign signals for test data
plt.hist(y_test, bins=classes_num)
plt.title('Histogram of Traffic Sign Signals (Test set)') # Display the title
plt.xlabel('Traffic Signs') # Display the x-axis
plt.ylabel('Total Count') # Display the y-axis
plt.show() # Plot the histogram
# Plot the histogram of traffic sign signals for validation data
plt.hist(y_valid, bins=classes_num)
plt.title('Histogram of Traffic Sign Signals (Validation set)') # Display the title
plt.xlabel('Traffic Signs') # Display the x-axis
plt.ylabel('Total Count') # Display the y-axis
plt.show() # Plot the histogram
# -
# #### Number of Images in each category
# +
label = pd.read_csv('signnames.csv')
num_img_category = np.zeros(classes_num)
for i in range(0, len(y_train)):
num_img_category[y_train[i]] += 1
# print the total count of images in each category
for i in range(0, classes_num):
print("{:2d}".format(i), "\t Count := " + str(num_img_category[i]),
"\t Traffic Sign := " + label.loc[i, 'SignName'],
)
# -
# ### Pre-process the Data Set (normalization, grayscale, etc.)
# Initial step followed in data pre-processing is to shuffle the data. It is very important to shuffle the training data otherwise ordering of data might have huge effect on how the network trends (Neural Network training).
# +
# Import library to shuffle the data
from sklearn.utils import shuffle
X_train, y_train = shuffle(X_train, y_train) # Shuffle training data
X_valid, y_valid = shuffle(X_valid, y_valid) # Shuffle validation data
X_test, y_test = shuffle(X_test, y_test) # Shuffle test data
# -
# The image data should be normalized so that the data has mean zero and equal variance. For image data, `(pixel - pixel_mean)/ (max_pixel-min_pixel)` is used to normalize the data. After normalization pixel values will be in the range -1 to +1.
# Other method used in normalizing the data is `(pixel - 128)/ 128`.
# +
import numpy as np
#Nomralisation
X_train = (X_train-X_train.mean())/(np.max(X_train)-np.min(X_train)) # Apply Normalization to training data
X_valid = (X_valid-X_valid.mean())/(np.max(X_valid)-np.min(X_valid)) # Apply Normalization to validation data
X_test = (X_test-X_test.mean())/(np.max(X_test)-np.min(X_test)) # Apply Normalization to test data
print("Min pixel value in normalized train dataset", np.amin(X_train[0]))
print("Max pixel value in normalized train dataset", np.amax(X_train[0]))
# -
# ### Model Architecture
# LeNet Architecture is used to build the Traffic Sign Classifier model. It includes two Convolutional Layers and three Fully-Connected layers.
#
# EPOCH and BATCH SIZE values affects the training speed and model accuracy.
# EPOCH variable is used to tell the TensorFlow how many times to run our training data through the network. More number of EPOCHS results in better model training but it takes longer time to train the network.
#
# BATCH_SIZE variable is used to tell the TensorFlow how many training images to run through the network at a time. If the BATCH_SIZE is larger, the model gets trained faster but our processor may have a memory limit on how large a batch it can run.
#
# #### Dimesionality
# The number of neurons of each layer in our CNN can be calculated by using below formula,
#
# output_height = [(input_height - filter_height + 2 * padding) / vertical_stride] + 1
#
# output_width = [(input_width - filter_width + 2 * padding) / vertical_stride] + 1
#
# output_depth = number of filters
# +
# Import libraries
import tensorflow as tf # Import tensorflow library
from tensorflow.contrib.layers import flatten # Flattens the input layer
EPOCHS = 50 # Number of epochs
BATCH_SIZE = 128 # Batch size
learning_rate = 0.001 # Learning rate tells the TensorFlow how quickly to update the network weights
# Hyperparameters
# Both these parameters relate to how we initialze our weights.
mu = 0
sigma = 0.1
# Define LeNet architecture
def LeNet(x):
# Layer 1: Convolutional. Input = 32x32x3. Output = 28x28x6.
# Use 5X5 filter with an input depth of 3 and an output depth of 6.
conv1_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 3, 6), mean = mu, stddev = sigma))
# Initialize the bias
conv1_b = tf.Variable(tf.zeros(6))
# Use conv2D function to convolve the filter over images and add bias at the end.
# y = x*W+b
conv1 = tf.nn.conv2d(x, conv1_W, strides=[1, 1, 1, 1], padding='VALID') + conv1_b
#output_height = [(32-5+2*0)/1]+1 = 28.
#output_width = [(32-5+2*0)/1]+1 = 28.
#output_depth = filter_depth = 6
# Activation 1.
conv1 = tf.nn.relu(conv1) # Activate the output of the convolutional layer using ReLU activation function
# Pooling. Input = 28x28x6. Output = 14x14x6.
# Pool the output using 2x2 kernel with a 2x2 stride
conv1 = tf.nn.max_pool(conv1, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
#output_height = 28/2 = 14.
#output_width = 28/2 = 14.
#output_depth = 6
# Layer 2: Convolutional. Input = 14x14x6. Output = 10x10x16.
conv2_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 6, 16), mean = 0, stddev = 0.1))
conv2_b = tf.Variable(tf.zeros(16))
conv2 = tf.nn.conv2d(conv1, conv2_W, strides=[1, 1, 1, 1], padding='VALID') + conv2_b
#output_height = [(14-5+2*0)/1]+1 = 10.
#output_width = [(14-5+2*0)/1]+1 = 10.
#output_depth = filter_depth = 16
# Activation 2.
conv2 = tf.nn.relu(conv2) # Activate the output of the convolutional layer using ReLU activation function
# Pooling. Input = 10x10x16. Output = 5x5x16.
# Pool the output using 2x2 kernel with a 2x2 stride
conv2 = tf.nn.max_pool(conv2, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
#output_height = 10/2 = 5.
#output_width = 10/2 = 5.
#output_depth = 16
# Flatten the output into a vector. Input = 5x5x16. Output=5x5x16=400.
flattened = flatten(conv2)
# Layer 3: Fully Connected. Input = 400. Output = 120.
fc1_W = tf.Variable(tf.truncated_normal(shape=(400, 120), mean = mu, stddev = sigma))
fc1_b = tf.Variable(tf.zeros(120))
fc1 = tf.matmul(flattened, fc1_W) + fc1_b
# Fully connected layer activation 1.
fc1 = tf.nn.relu(fc1)
# Layer 4: Fully Connected. Input = 120. Output = 84.
fc2_W = tf.Variable(tf.truncated_normal(shape=(120, 84), mean = mu, stddev = sigma))
fc2_b = tf.Variable(tf.zeros(84))
fc2 = tf.matmul(fc1, fc2_W) + fc2_b
# Full connected layer activation 2.
fc2 = tf.nn.relu(fc2)
# Layer 5: Fully Connected. Input = 84. Output = 43.
# Set the output with a width equal to number of classes in our label set. These outrputs are called as logits.
fc3_W = tf.Variable(tf.truncated_normal(shape=(84, 43), mean = mu, stddev = sigma))
fc3_b = tf.Variable(tf.zeros(43))
logits = tf.matmul(fc2, fc3_W) + fc3_b
return logits
# -
# ### Features and Labels
# +
# x is a placeholder that will store our input batches
# batch_size=None allows the placeholder to later accept the batch of any size
# set the image dimensions to 32x32x3
# In this case, our labels come through sparse variables, which means they are integers. They are not one-hot encoded yet.
x = tf.placeholder(tf.float32, (None, 32, 32, 3))
# y is a placeholder that will store our output batches (y stores the labels)
y = tf.placeholder(tf.int32, (None))
# One-hot encode the labels
one_hot_y = tf.one_hot(y, 43) # Number of outputs = 43
# -
# ### Training Pipeline
# Create a training pipeline that uses the model to classify traffic sign images.
# +
# Pass the input data to the LeNet function to calculate our logits
logits = LeNet(x)
# 'tf.nn.softmax_cross_entropy_with_logits' function is used to compare the logits to the ground truth training labels and
# calculate the cross entropy. Cross Entropy is just a measure of how different the logits are from the ground truth training labels.
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(labels=one_hot_y, logits=logits)
# 'tf.reduce_mean' function averages the cross entropy from all of the training images
loss_operation = tf.reduce_mean(cross_entropy)
# Adam Optimizer uses the Adam algorithm to minimize the loss function using learning rate.
optimizer = tf.train.AdamOptimizer(learning_rate = learning_rate)
# Run the minimize function on the optimizer which uses back-propagation to update the network and minimize our training loss.
training_operation = optimizer.minimize(loss_operation)
# -
# ### Model Evaluation
# Evaluate how well the model the loss and accuracy of the model for a given dataset.
#
# A validation set can be used to assess how well the model is performing. A low accuracy on the training and validation
# sets imply underfitting. A high accuracy on the training set but low accuracy on the validation set implies overfitting.
# +
# Measure whether given prediction is correct by comparing logit prediction to the one-hot encoded ground truth label.
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1))
# Calculte the model's overall accuracy by averaging the individual prediction accuracies.
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
saver = tf.train.Saver()
def evaluate(X_data, y_data):
num_examples = len(X_data)
total_accuracy = 0
total_loss = 0
sess = tf.get_default_session()
# Break the training data into batches and train the model on each batch.
for offset in range(0, num_examples, BATCH_SIZE):
batch_x, batch_y = X_data[offset:offset+BATCH_SIZE], y_data[offset:offset+BATCH_SIZE] # batch the dataset
loss, accuracy = sess.run([loss_operation, accuracy_operation], feed_dict={x: batch_x, y: batch_y}) # run through evaluation pipeline
total_accuracy += (accuracy * len(batch_x)) # Calculate the avarage accuracy of each batch to calculate the total accuracy of the model
total_loss += (loss * len(batch_x))
return total_accuracy/num_examples
# -
# ### Train the Model
# Run the training data through the training pipeline to train the model.
#
# Before each epoch shuffle the training set to ensure that our training is not biased by the order of the images.
#
# Break the training data into batches and train the model on each batch.
#
# At the end of each epoch, we evaluate the model on our validation data.
#
# Once we have completly trained the model, save it. We can load it later or modify it or evaluate our model on test datset.
# Create TensorFlow session and initialize variables
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
num_examples = len(X_train)
print("Training...")
print()
# Train over 'N' number of epochs
for i in range(EPOCHS):
# Before each epoch shuffle the training set.
X_train, y_train = shuffle(X_train, y_train)
# Break the training data into batches and train the model on each batch.
for offset in range(0, num_examples, BATCH_SIZE):
end = offset + BATCH_SIZE
batch_x, batch_y = X_train[offset:end], y_train[offset:end]
sess.run(training_operation, feed_dict={x: batch_x, y: batch_y})
# At the end of each epoch, we evaluate the model on our validation data
validation_accuracy = evaluate(X_valid, y_valid)
print("EPOCH {} ...".format(i+1))
print("Validation Accuracy = {:.3f}".format(validation_accuracy))
print()
# Save the model after the completion of training.
saver.save(sess, './lenet')
print("Model saved")
# As we train the model, we see that validation accuracy starts off really high and stays there. This is the result of the powerful Convolutional Network Acrchitecture LeNet and because of the choice hyperparameters.
# ### Evalute the Model
# Evalate the performance of the model on test set. This has to be done only once after the completion of training. Otherwise, we would be using the test dataset to choose the best model and then the test dataset would not provide a good estimate of how well the model would do in the real world.
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
test_accuracy = evaluate(X_test, y_test)
print("Test Accuracy = {:.3f}".format(test_accuracy))
# ---
#
# ## Step 3: Test a Model on New Images
#
# To give more insight into how the model is working, five pictures of German traffic signs are downloded from the web and the model is used to predict the traffic sign type.
#
# We can find `signnames.csv` useful as it contains mappings from the class id (integer) to the actual sign name.
# ### Load the Images
# +
### Import the libraries
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import cv2
# List of test images
x_test_images = ['test_images/1.png', 'test_images/2.png', 'test_images/3.png', 'test_images/4.png', 'test_images/5.png']
y_new_test = np.array([11,12,1,25,38])
imgs = []
for i in range(0,len(x_test_images)):
img = mpimg.imread(x_test_images[i])
imgs.append(cv2.cvtColor(cv2.imread(x_test_images[i]), cv2.COLOR_BGR2RGB))
plt.imshow(img)
plt.figure()
# Create an array of images
X_new_test = np.asarray(imgs)
# Normalize the test data
X_new_test = (X_new_test-X_new_test.mean())/(np.max(X_new_test)-np.min(X_new_test))
# -
# ### Predict the Sign Type for Each Image
# +
### Run the predictions here and use the model to output the prediction for each image.
# set the batch size to 64
batch_size = 64
result = None
# Read the sign id and name from csv file
sign_names = pd.read_csv('signnames.csv', names=['id', 'name'], header=0)
# Function to evaluate the prediction
def evaluate_prediction(X_new_test, batch_size):
sess = tf.get_default_session()
steps_per_epoch = len(X_new_test) // batch_size + (len(X_new_test)%batch_size > 0)
predictions = np.zeros((len(X_new_test), classes_num))
for step in range(steps_per_epoch):
batch_x = X_new_test[step*batch_size:(step+1)*batch_size]
batch_y = np.zeros((len(batch_x), classes_num))
prediction = sess.run(tf.nn.softmax(logits), feed_dict={x: batch_x})
predictions[step*batch_size:(step+1)*batch_size] = prediction
return predictions
# Create tensorflow session
with tf.Session() as sess:
# Restore the model which was saved earlier
saver.restore(sess, tf.train.latest_checkpoint('.'))
prediction = evaluate_prediction(X_new_test, batch_size)
result = sess.run(tf.nn.top_k(tf.constant(prediction),k=5))
values, indices = result
for each in indices:
print('{} -- {}'.format(each[0], sign_names.name[each[0]]))
# -
# ### Analyze Performance
# +
### Calculate the accuracy for these 5 new images.
### For example, if the model predicted 1 out of 5 signs correctly, it's 20% accurate on these new images.
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
accuracy = sess.run(accuracy_operation, feed_dict={
x: X_new_test,
y: y_new_test,
})
print('Accuracy: {:.3f}'.format(accuracy))
# -
# ### Output Top 5 Softmax Probabilities For Each Image Found on the Web
# For each of the new images, print out the model's softmax probabilities to show the **certainty** of the model's predictions (limit the output to the top 5 probabilities for each image). [`tf.nn.top_k`](https://www.tensorflow.org/versions/r0.12/api_docs/python/nn.html#top_k) could prove helpful here.
#
# The example below demonstrates how tf.nn.top_k can be used to find the top k predictions for each image.
#
# `tf.nn.top_k` will return the values and indices (class ids) of the top k predictions. So if k=3, for each sign, it'll return the 3 largest probabilities (out of a possible 43) and the correspoding class ids.
#
# Take this numpy array as an example. The values in the array represent predictions. The array contains softmax probabilities for five candidate images with six possible classes. `tf.nn.top_k` is used to choose the three classes with the highest probability:
#
# ```
# # (5, 6) array
# a = np.array([[ 0.24879643, 0.07032244, 0.12641572, 0.34763842, 0.07893497,
# 0.12789202],
# [ 0.28086119, 0.27569815, 0.08594638, 0.0178669 , 0.18063401,
# 0.15899337],
# [ 0.26076848, 0.23664738, 0.08020603, 0.07001922, 0.1134371 ,
# 0.23892179],
# [ 0.11943333, 0.29198961, 0.02605103, 0.26234032, 0.1351348 ,
# 0.16505091],
# [ 0.09561176, 0.34396535, 0.0643941 , 0.16240774, 0.24206137,
# 0.09155967]])
# ```
#
# Running it through `sess.run(tf.nn.top_k(tf.constant(a), k=3))` produces:
#
# ```
# TopKV2(values=array([[ 0.34763842, 0.24879643, 0.12789202],
# [ 0.28086119, 0.27569815, 0.18063401],
# [ 0.26076848, 0.23892179, 0.23664738],
# [ 0.29198961, 0.26234032, 0.16505091],
# [ 0.34396535, 0.24206137, 0.16240774]]), indices=array([[3, 0, 5],
# [0, 1, 4],
# [0, 5, 1],
# [1, 3, 5],
# [1, 4, 3]], dtype=int32))
# ```
#
# Looking just at the first row we get `[ 0.34763842, 0.24879643, 0.12789202]`, you can confirm these are the 3 largest probabilities in `a`. You'll also notice `[3, 0, 5]` are the corresponding indices.
### Print out the top five softmax probabilities for the predictions on the German traffic sign images found on the web.
for i, img in enumerate(X_new_test):
correct = label.loc[y_new_test[i], 'SignName']
print('True Label is --> {:>5}:{:<30}'.format(y_new_test[i], correct))
print("")
for j, k in zip(indices[i], values[i]):
name = label.loc[j, 'SignName']
print('{:>5}: {:<30} {:>3.3f}%'.format(j, name, k*100.0))
print("##############################################")
print("##############################################")
# ### Project Writeup
#
# Once you have completed the code implementation, document your results in a project writeup using this [template](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/writeup_template.md) as a guide. The writeup can be in a markdown or pdf file.
# > **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to \n",
# "**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
# ---
#
# ## Step 4 (Optional): Visualize the Neural Network's State with Test Images
#
# This Section is not required to complete but acts as an additional excersise for understaning the output of a neural network's weights. While neural networks can be a great learning device they are often referred to as a black box. We can understand what the weights of a neural network look like better by plotting their feature maps. After successfully training your neural network you can see what it's feature maps look like by plotting the output of the network's weight layers in response to a test stimuli image. From these plotted feature maps, it's possible to see what characteristics of an image the network finds interesting. For a sign, maybe the inner network feature maps react with high activation to the sign's boundary outline or to the contrast in the sign's painted symbol.
#
# Provided for you below is the function code that allows you to get the visualization output of any tensorflow weight layer you want. The inputs to the function should be a stimuli image, one used during training or a new one you provided, and then the tensorflow variable name that represents the layer's state during the training process, for instance if you wanted to see what the [LeNet lab's](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) feature maps looked like for it's second convolutional layer you could enter conv2 as the tf_activation variable.
#
# For an example of what feature map outputs look like, check out NVIDIA's results in their paper [End-to-End Deep Learning for Self-Driving Cars](https://devblogs.nvidia.com/parallelforall/deep-learning-self-driving-cars/) in the section Visualization of internal CNN State. NVIDIA was able to show that their network's inner weights had high activations to road boundary lines by comparing feature maps from an image with a clear path to one without. Try experimenting with a similar test to show that your trained network's weights are looking for interesting features, whether it's looking at differences in feature maps from images with or without a sign, or even what feature maps look like in a trained network vs a completely untrained one on the same sign image.
#
# <figure>
# <img src="visualize_cnn.png" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> Your output should look something like this (above)</p>
# </figcaption>
# </figure>
# <p></p>
#
# +
### Visualize your network's feature maps here.
### Feel free to use as many code cells as needed.
# image_input: the test image being fed into the network to produce the feature maps
# tf_activation: should be a tf variable name used during your training procedure that represents the calculated state of a specific weight layer
# activation_min/max: can be used to view the activation contrast in more detail, by default matplot sets min and max to the actual min and max values of the output
# plt_num: used to plot out multiple different weight feature map sets on the same block, just extend the plt number for each new feature map entry
def outputFeatureMap(image_input, tf_activation, activation_min=-1, activation_max=-1 ,plt_num=1):
# Here make sure to preprocess your image_input in a way your network expects
# with size, normalization, ect if needed
# image_input =
# Note: x should be the same name as your network's tensorflow data placeholder variable
# If you get an error tf_activation is not defined it may be having trouble accessing the variable from inside a function
activation = tf_activation.eval(session=sess,feed_dict={x : image_input})
featuremaps = activation.shape[3]
plt.figure(plt_num, figsize=(15,15))
for featuremap in range(featuremaps):
plt.subplot(6,8, featuremap+1) # sets the number of feature maps to show on each row and column
plt.title('FeatureMap ' + str(featuremap)) # displays the feature map number
if activation_min != -1 & activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin =activation_min, vmax=activation_max, cmap="gray")
elif activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmax=activation_max, cmap="gray")
elif activation_min !=-1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin=activation_min, cmap="gray")
else:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", cmap="gray")
|
Traffic_Sign_Classifier.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Novo Capstone
# language: python
# name: novo
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
grid_search_data = pd.read_csv('first_grid_search_results.csv', index_col=0)
# -
grid_search_data.sort_values('means', ascending=False)
print('Pearson correlation of each hyperparameter \nwith accuracy '
'from 10-fold cv:')
grid_search_data.corr()['means'][:4]
# Without any further analysis, it seems there is a slight preference for small filter_size and large pool_size.
# We should also vary pool_stride.
grouped = grid_search_data.groupby('filter_size')
grouped.mean()['means'].plot(yerr=grouped.mean()['stddev'], marker='o', ms=7, mfc='k')
grouped = grid_search_data.groupby('hidden_dims')
grouped.mean()['means'].plot(yerr=grouped.mean()['stddev'], marker='o', ms=7, mfc='k')
grouped = grid_search_data.groupby('num_filters')
grouped.mean()['means'].plot(yerr=grouped.mean()['stddev'], marker='o', ms=7, mfc='k')
grouped = grid_search_data.groupby('pool_size')
grouped.mean()['means'].plot(yerr=grouped.mean()['stddev'], marker='o', ms=7, mfc='k')
# # correlation between sets of 2?
grouped = grid_search_data.groupby(['pool_size', 'filter_size'])
grouped.mean()['means'].plot(yerr=grouped.mean()['stddev'], marker='o', ms=7, mfc='k')
grouped = grid_search_data.groupby(['pool_size', 'num_filters'])
grouped.mean()['means'].plot(yerr=grouped.mean()['stddev'], marker='o', ms=7, mfc='k')
grouped = grid_search_data.groupby(['filter_size', 'num_filters'])
grouped.mean()['means'].plot(yerr=grouped.mean()['stddev'], marker='o', ms=7, mfc='k')
grouped = grid_search_data.groupby(['pool_size', 'filter_size', 'num_filters'])
grouped.mean()['means'].plot(yerr=grouped.mean()['stddev'], marker='o', ms=7, mfc='k')
# +
# current results indicate GlobalMaxPooling1D with small filter size will improve results
# -
# # second attempt, switching to max pooling and multiple fitler sizes
grid_search_two = pd.read_csv('second_grid_search_results.csv', index_col=0)
grid_search_two.sort_values('means', ascending=False)
print('Pearson correlation of each hyperparameter \nwith accuracy '
'from 10-fold cv:')
grid_search_two.corr()['means']
grouped = grid_search_two.groupby('num_filters')
grouped.mean()['means'].plot(yerr=grouped.mean()['stddev'], marker='o', ms=7, mfc='k')
grouped = grid_search_two.groupby('filter_sizes')
grouped.mean()['means'].plot(yerr=grouped.mean()['stddev'], marker='o', ms=7, mfc='k')
grouped = grid_search_two.groupby(['filter_sizes', 'num_filters'])
grouped.mean()['means'].plot(yerr=grouped.mean()['stddev'], marker='o', ms=7, mfc='k')
|
archive/josh/grid_search_synthesis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
num = int(input("Enter a Numbver to reverse:"))
reverse = 0
while num != 0:
digit = num % 10
reverse = reverse * 10 + digit
num //= 10
print('Reverse Number: '+str(reverse))
# -
num = int(input("Enter a Numbver to reverse:"))
print(str(num)[::-1])
num = (input("Enter a Number to reverse:"))
if num[::-1] == num:
print("Palindrome Number")
else:
print("Not a Palindrome")
# +
n = int(input("Enter the number"))
count = 0
sum = 0
original = n
temp=n
# its use for calculating number of digits in the number
while(n>0):
count=count+1
n=n//10
print("No of Digits in the number is :",count)
while(original > 0):
digit = original % 10
sum += digit ** count
original //= 10
if temp == sum:
print(temp,"is an armstrong number")
else:
print(temp,"is not an armstrong number")
# -
num = int(input("Enter a number : "))
n1,n2 = 0,1
for i in range(2,num):
n3 = n1+n2
n1 = n2
n2 = n3
print(n3)
num = int(input("Enter the number : "))
fact = 1
for i in range(1,num+1):
fact = fact*i
print("Factorial is : ",fact)
num = int(input("Enter the number : "))
num1 = int(input("Enter the number : "))
temp = 1
for i in range(0,num1):
temp = temp * num
print("Power of the number is",temp)
# +
num=int(input("enter a number"))
factors=[]
for i in range(1,num+1):
if num%i==0:
factors.append(i)
print ("Factors of {} = {}".format(num,factors))
# -
sum = 0
n = int(input ("Enter a number : "))
temp = n
while (n):
k = 1
fact = 1
r = n % 10
while(k <= r):
fact = fact * k
k = k+1
sum = sum + fact
n = n// 10
if(sum == temp):
print(str(temp)+"is a strong number")
else:
print(str(temp)+" is not a strong number")
# +
digit = 0
length = 0
square = 0
num = input("Enter a number: ")
length = len(num)
num = int(num)
square = num ** 2
digit = square % 10 ** length
if digit == num:
print(num,"IS automorphic number")
else:
print("Not Automorphic")
|
python2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Noisy Duelling Double Deep Q Learning - A simple ambulance dispatch point allocation model
#
# ## Reinforcement learning introduction
#
# ### RL involves:
# * Trial and error search
# * Receiving and maximising reward (often delayed)
# * Linking state -> action -> reward
# * Must be able to sense something of their environment
# * Involves uncertainty in sensing and linking action to reward
# * Learning -> improved choice of actions over time
# * All models find a way to balance best predicted action vs. exploration
#
# ### Elements of RL
# * *Environment*: all observable and unobservable information relevant to us
# * *Observation*: sensing the environment
# * *State*: the perceived (or perceivable) environment
# * *Agent*: senses environment, decides on action, receives and monitors rewards
# * *Action*: may be discrete (e.g. turn left) or continuous (accelerator pedal)
# * *Policy* (how to link state to action; often based on probabilities)
# * *Reward signal*: aim is to accumulate maximum reward over time
# * *Value function* of a state: prediction of likely/possible long-term reward
# * *Q*: prediction of likely/possible long-term reward of an *action*
# * *Advantage*: The difference in Q between actions in a given state (sums to zero for all actions)
# * *Model* (optional): a simulation of the environment
#
# ### Types of model
#
# * *Model-based*: have model of environment (e.g. a board game)
# * *Model-free*: used when environment not fully known
# * *Policy-based*: identify best policy directly
# * *Value-based*: estimate value of a decision
# * *Off-policy*: can learn from historic data from other agent
# * *On-policy*: requires active learning from current decisions
#
#
# ## Duelling Deep Q Networks for Reinforcement Learning
#
# Q = The expected future rewards discounted over time. This is what we are trying to maximise.
#
# The aim is to teach a network to take the current state observations and recommend the action with greatest Q.
#
# Duelling is very similar to Double DQN, except that the policy net splits into two. One component reduces to a single value, which will model the state *value*. The other component models the *advantage*, the difference in Q between different actions (the mean value is subtracted from all values, so that the advtantage always sums to zero). These are aggregated to produce Q for each action.
#
# <img src="./images/duelling_dqn.png" width="500"/>
#
# Q is learned through the Bellman equation, where the Q of any state and action is the immediate reward achieved + the discounted maximum Q value (the best action taken) of next best action, where gamma is the discount rate.
#
# $$Q(s,a)=r + \gamma.maxQ(s',a')$$
#
# ## Key DQN components
#
# <img src="./images/dqn_components.png" width="700"/>
#
#
# ## General method for Q learning:
#
# Overall aim is to create a neural network that predicts Q. Improvement comes from improved accuracy in predicting 'current' understood Q, and in revealing more about Q as knowledge is gained (some rewards only discovered after time).
#
# <img src="./images/dqn_process.png" width="600|"/>
#
# Target networks are used to stabilise models, and are only updated at intervals. Changes to Q values may lead to changes in closely related states (i.e. states close to the one we are in at the time) and as the network tries to correct for errors it can become unstable and suddenly lose signficiant performance. Target networks (e.g. to assess Q) are updated only infrequently (or gradually), so do not have this instability problem.
#
# ## Training networks
#
# Double DQN contains two networks. This ammendment, from simple DQN, is to decouple training of Q for current state and target Q derived from next state which are closely correlated when comparing input features.
#
# The *policy network* is used to select action (action with best predicted Q) when playing the game.
#
# When training, the predicted best *action* (best predicted Q) is taken from the *policy network*, but the *policy network* is updated using the predicted Q value of the next state from the *target network* (which is updated from the policy network less frequently). So, when training, the action is selected using Q values from the *policy network*, but the the *policy network* is updated to better predict the Q value of that action from the *target network*. The *policy network* is copied across to the *target network* every *n* steps (e.g. 1000).
#
# <img src="./images/dqn_training.png" width="700|"/>
#
# ## Noisy layers
# Noisy layers are an alternative to epsilon-greedy exploration (here, we leave the epsilon-greedy code in the model, but set it to reduce to zero immediately after the period of fully random action choice).
#
# For every weight in the layer we have a random value that we draw from the normal distribution. This random value is used to add noise to the output. The parameters for the extent of noise for each weight, sigma, are stored within the layer and get trained as part of the standard back-propogation.
#
# A modification to normal nosiy layers is to use layers with ‘factorized gaussian noise’. This reduces the number of random numbers to be sampled (so is less computationally expensive). There are two random vectors, one with the size of the input, and the other with the size of the output. A random matrix is created by calculating the outer product of the two vectors.
#
# ## References
#
# Double DQN:
# <NAME>, <NAME>, <NAME>. (2015) Deep Reinforcement Learning with Double Q-learning. arXiv:150906461 http://arxiv.org/abs/1509.06461
#
# Duelling DDQN:
# <NAME>, <NAME>, <NAME>, et al. (2016) Dueling Network Architectures for Deep Reinforcement Learning. arXiv:151106581 http://arxiv.org/abs/1511.06581
#
# Noisy networks:
# Fortunato M, Azar MG, Piot B, et al. (2019) Noisy Networks for Exploration. arXiv:170610295 http://arxiv.org/abs/1706.10295
#
# Code for the nosiy layers comes from:
#
# Lapan, M. (2020). Deep Reinforcement Learning Hands-On: Apply modern RL methods to practical problems of chatbots, robotics, discrete optimization, web automation, and more, 2nd Edition. Packt Publishing.
#
# ## Code structure
#
# <img src="./images/dqn_program_structure.png" width="700|"/>
# +
################################################################################
# 1 Import packages #
################################################################################
from amboworld.environment import Env
import math
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import random
import torch
import torch.nn as nn
import torch.optim as optim
from torch.nn import functional as F
# Use a double ended queue (deque) for memory
# When memory is full, this will replace the oldest value with the new one
from collections import deque
# Supress all warnings (e.g. deprecation warnings) for regular use
import warnings
warnings.filterwarnings("ignore")
# +
################################################################################
# 2 Define model parameters #
################################################################################
# Set whether to display on screen (slows model)
DISPLAY_ON_SCREEN = False
# Discount rate of future rewards
GAMMA = 0.99
# Learing rate for neural network
LEARNING_RATE = 0.003
# Maximum number of game steps (state, action, reward, next state) to keep
MEMORY_SIZE = 10000000
# Sample batch size for policy network update
BATCH_SIZE = 5
# Number of game steps to play before starting training (all random actions)
REPLAY_START_SIZE = 50000
# Number of steps between policy -> target network update
SYNC_TARGET_STEPS = 1000
# Exploration rate (epsilon) is probability of choosing a random action
EXPLORATION_MAX = 1.0
EXPLORATION_MIN = 0.0
# Reduction in epsilon with each game step
EXPLORATION_DECAY = 0.0
# Training episodes
TRAINING_EPISODES = 50
# Save results
RESULTS_NAME = 'noisy_d3qn'
# SIM PARAMETERS
RANDOM_SEED = 42
SIM_DURATION = 5000
NUMBER_AMBULANCES = 3
NUMBER_INCIDENT_POINTS = 1
INCIDENT_RADIUS = 2
NUMBER_DISPTACH_POINTS = 25
AMBOWORLD_SIZE = 50
INCIDENT_INTERVAL = 60
EPOCHS = 2
AMBO_SPEED = 60
AMBO_FREE_FROM_HOSPITAL = False
# +
################################################################################
# 3 Define DQN (Duelling Deep Q Network) class #
# (Used for both policy and target nets) #
################################################################################
"""
Code for nosiy layers comes from:
<NAME>. (2020). Deep Reinforcement Learning Hands-On: Apply modern RL methods
to practical problems of chatbots, robotics, discrete optimization,
web automation, and more, 2nd Edition. Packt Publishing.
"""
class NoisyLinear(nn.Linear):
"""
Noisy layer for network.
For every weight in the layer we have a random value that we draw from the
normal distribution.Paraemters for the noise, sigma, are stored within the
layer and get trained as part of the standard back-propogation.
'register_buffer' is used to create tensors in the network that are not
updated during back-propogation. They are used to create normal
distributions to add noise (multiplied by sigma which is a paramater in the
network).
"""
def __init__(self, in_features, out_features,
sigma_init=0.017, bias=True):
super(NoisyLinear, self).__init__(
in_features, out_features, bias=bias)
w = torch.full((out_features, in_features), sigma_init)
self.sigma_weight = nn.Parameter(w)
z = torch.zeros(out_features, in_features)
self.register_buffer("epsilon_weight", z)
if bias:
w = torch.full((out_features,), sigma_init)
self.sigma_bias = nn.Parameter(w)
z = torch.zeros(out_features)
self.register_buffer("epsilon_bias", z)
self.reset_parameters()
def reset_parameters(self):
std = math.sqrt(3 / self.in_features)
self.weight.data.uniform_(-std, std)
self.bias.data.uniform_(-std, std)
def forward(self, input):
self.epsilon_weight.normal_()
bias = self.bias
if bias is not None:
self.epsilon_bias.normal_()
bias = bias + self.sigma_bias * \
self.epsilon_bias.data
v = self.sigma_weight * self.epsilon_weight.data + self.weight
return F.linear(input, v, bias)
class NoisyFactorizedLinear(nn.Linear):
"""
NoisyNet layer with factorized gaussian noise. This reduces the number of
random numbers to be sampled (so less computationally expensive). There are
two random vectors. One with the size of the input, and the other with the
size of the output. A random matrix is create by calculating the outer
product of the two vectors.
'register_buffer' is used to create tensors in the network that are not
updated during back-propogation. They are used to create normal
distributions to add noise (multiplied by sigma which is a paramater in the
network).
"""
def __init__(self, in_features, out_features,
sigma_zero=0.4, bias=True):
super(NoisyFactorizedLinear, self).__init__(
in_features, out_features, bias=bias)
sigma_init = sigma_zero / math.sqrt(in_features)
w = torch.full((out_features, in_features), sigma_init)
self.sigma_weight = nn.Parameter(w)
z1 = torch.zeros(1, in_features)
self.register_buffer("epsilon_input", z1)
z2 = torch.zeros(out_features, 1)
self.register_buffer("epsilon_output", z2)
if bias:
w = torch.full((out_features,), sigma_init)
self.sigma_bias = nn.Parameter(w)
def forward(self, input):
self.epsilon_input.normal_()
self.epsilon_output.normal_()
func = lambda x: torch.sign(x) * torch.sqrt(torch.abs(x))
eps_in = func(self.epsilon_input.data)
eps_out = func(self.epsilon_output.data)
bias = self.bias
if bias is not None:
bias = bias + self.sigma_bias * eps_out.t()
noise_v = torch.mul(eps_in, eps_out)
v = self.weight + self.sigma_weight * noise_v
return F.linear(input, v, bias)
class DQN(nn.Module):
"""Deep Q Network. Udes for both policy (action) and target (Q) networks."""
def __init__(self, observation_space, action_space):
"""Constructor method. Set up neural nets."""
# nerurones per hidden layer = 2 * max of observations or actions
neurons_per_layer = 2 * max(observation_space, action_space)
# Set starting exploration rate
self.exploration_rate = EXPLORATION_MAX
# Set up action space (choice of possible actions)
self.action_space = action_space
# First layerswill be common to both Advantage and value
super(DQN, self).__init__()
self.feature = nn.Sequential(
nn.Linear(observation_space, neurons_per_layer),
nn.ReLU()
)
# Advantage has same number of outputs as the action space
self.advantage = nn.Sequential(
NoisyFactorizedLinear(neurons_per_layer, neurons_per_layer),
nn.ReLU(),
NoisyFactorizedLinear(neurons_per_layer, action_space)
)
# State value has only one output (one value per state)
self.value = nn.Sequential(
nn.Linear(neurons_per_layer, neurons_per_layer),
nn.ReLU(),
nn.Linear(neurons_per_layer, 1)
)
def act(self, state):
"""Act either randomly or by redicting action that gives max Q"""
# Act randomly if random number < exploration rate
if np.random.rand() < self.exploration_rate:
action = random.randrange(self.action_space)
else:
# Otherwise get predicted Q values of actions
q_values = self.forward(torch.FloatTensor(state))
# Get index of action with best Q
action = np.argmax(q_values.detach().numpy()[0])
return action
def forward(self, x):
x = self.feature(x)
advantage = self.advantage(x)
value = self.value(x)
action_q = value + advantage - advantage.mean()
return action_q
# +
################################################################################
# 4 Define policy net training function #
################################################################################
def optimize(policy_net, target_net, memory):
"""
Update model by sampling from memory.
Uses policy network to predict best action (best Q).
Uses target network to provide target of Q for the selected next action.
"""
# Do not try to train model if memory is less than reqired batch size
if len(memory) < BATCH_SIZE:
return
# Reduce exploration rate (exploration rate is stored in policy net)
policy_net.exploration_rate *= EXPLORATION_DECAY
policy_net.exploration_rate = max(EXPLORATION_MIN,
policy_net.exploration_rate)
# Sample a random batch from memory
batch = random.sample(memory, BATCH_SIZE)
for state, action, reward, state_next, terminal in batch:
state_action_values = policy_net(torch.FloatTensor(state))
# Get target Q for policy net update
if not terminal:
# For non-terminal actions get Q from policy net
expected_state_action_values = policy_net(torch.FloatTensor(state))
# Detach next state values from gradients to prevent updates
expected_state_action_values = expected_state_action_values.detach()
# Get next state action with best Q from the policy net (double DQN)
policy_next_state_values = policy_net(torch.FloatTensor(state_next))
policy_next_state_values = policy_next_state_values.detach()
best_action = np.argmax(policy_next_state_values[0].numpy())
# Get target net next state
next_state_action_values = target_net(torch.FloatTensor(state_next))
# Use detach again to prevent target net gradients being updated
next_state_action_values = next_state_action_values.detach()
best_next_q = next_state_action_values[0][best_action].numpy()
updated_q = reward + (GAMMA * best_next_q)
expected_state_action_values[0][action] = updated_q
else:
# For termal actions Q = reward (-1)
expected_state_action_values = policy_net(torch.FloatTensor(state))
# Detach values from gradients to prevent gradient update
expected_state_action_values = expected_state_action_values.detach()
# Set Q for all actions to reward (-1)
expected_state_action_values[0] = reward
# Set network to training mode
policy_net.train()
# Reset net gradients
policy_net.optimizer.zero_grad()
# calculate loss
loss_v = nn.MSELoss()(state_action_values, expected_state_action_values)
# Backpropogate loss
loss_v.backward()
# Update network gradients
policy_net.optimizer.step()
return
# +
################################################################################
# 5 Define memory class #
################################################################################
class Memory():
"""
Replay memory used to train model.
Limited length memory (using deque, double ended queue from collections).
- When memory full deque replaces oldest data with newest.
Holds, state, action, reward, next state, and episode done.
"""
def __init__(self):
"""Constructor method to initialise replay memory"""
self.memory = deque(maxlen=MEMORY_SIZE)
def remember(self, state, action, reward, next_state, done):
"""state/action/reward/next_state/done"""
self.memory.append((state, action, reward, next_state, done))
# +
################################################################################
# 6 Define results plotting function #
################################################################################
def plot_results(run, exploration, score, mean_call_to_arrival,
mean_assignment_to_arrival):
"""Plot and report results at end of run"""
# Set up chart (ax1 and ax2 share x-axis to combine two plots on one graph)
fig = plt.figure(figsize=(6,6))
ax1 = fig.add_subplot(111)
ax2 = ax1.twinx()
# Plot results
lns1 = ax1.plot(
run, exploration, label='exploration', color='g', linestyle=':')
lns2 = ax2.plot(run, mean_call_to_arrival,
label='call to arrival', color='r')
lns3 = ax2.plot(run, mean_assignment_to_arrival,
label='assignment to arrival', color='b', linestyle='--')
# Get combined legend
lns = lns1 + lns2 + lns3
labs = [l.get_label() for l in lns]
ax1.legend(lns, labs, loc='upper center', bbox_to_anchor=(0.5, -0.1), ncol=3)
# Set axes
ax1.set_xlabel('run')
ax1.set_ylabel('exploration')
ax2.set_ylabel('Response time')
filename = 'output/' + RESULTS_NAME +'.png'
plt.savefig(filename, dpi=300)
plt.show()
# +
################################################################################
# 7 Main program #
################################################################################
def qambo():
"""Main program loop"""
############################################################################
# 8 Set up environment #
############################################################################
# Set up game environemnt
sim = Env(
random_seed = RANDOM_SEED,
duration_incidents = SIM_DURATION,
number_ambulances = NUMBER_AMBULANCES,
number_incident_points = NUMBER_INCIDENT_POINTS,
incident_interval = INCIDENT_INTERVAL,
number_epochs = EPOCHS,
number_dispatch_points = NUMBER_DISPTACH_POINTS,
incident_range = INCIDENT_RADIUS,
max_size = AMBOWORLD_SIZE,
ambo_kph = AMBO_SPEED,
ambo_free_from_hospital = AMBO_FREE_FROM_HOSPITAL
)
# Get number of observations returned for state
observation_space = sim.observation_size
# Get number of actions possible
action_space = sim.action_number
############################################################################
# 9 Set up policy and target nets #
############################################################################
# Set up policy and target neural nets (and keep best net performance)
policy_net = DQN(observation_space, action_space)
target_net = DQN(observation_space, action_space)
best_net = DQN(observation_space, action_space)
# Set loss function and optimizer
policy_net.optimizer = optim.Adam(
params=policy_net.parameters(), lr=LEARNING_RATE)
# Copy weights from policy_net to target
target_net.load_state_dict(policy_net.state_dict())
# Set target net to eval rather than training mode
# We do not train target net - ot is copied from policy net at intervals
target_net.eval()
############################################################################
# 10 Set up memory #
############################################################################
# Set up memomry
memory = Memory()
############################################################################
# 11 Set up + start training loop #
############################################################################
# Set up run counter and learning loop
run = 0
all_steps = 0
continue_learning = True
best_reward = -np.inf
# Set up list for results
results_run = []
results_exploration = []
results_score = []
results_mean_call_to_arrival = []
results_mean_assignment_to_arrival = []
# Continue repeating games (episodes) until target complete
while continue_learning:
########################################################################
# 12 Play episode #
########################################################################
# Increment run (episode) counter
run += 1
########################################################################
# 13 Reset game #
########################################################################
# Reset game environment and get first state observations
state = sim.reset()
# Reset total reward and rewards list
total_reward = 0
rewards = []
# Reshape state into 2D array with state obsverations as first 'row'
state = np.reshape(state, [1, observation_space])
# Continue loop until episode complete
while True:
####################################################################
# 14 Game episode loop #
####################################################################
####################################################################
# 15 Get action #
####################################################################
# Get action to take (se eval mode to avoid dropout layers)
policy_net.eval()
action = policy_net.act(state)
####################################################################
# 16 Play action (get S', R, T) #
####################################################################
# Act
state_next, reward, terminal, info = sim.step(action)
total_reward += reward
# Update trackers
rewards.append(reward)
# Reshape state into 2D array with state observations as first 'row'
state_next = np.reshape(state_next, [1, observation_space])
# Update display if needed
if DISPLAY_ON_SCREEN:
sim.render()
####################################################################
# 17 Add S/A/R/S/T to memory #
####################################################################
# Record state, action, reward, new state & terminal
memory.remember(state, action, reward, state_next, terminal)
# Update state
state = state_next
####################################################################
# 18 Check for end of episode #
####################################################################
# Actions to take if end of game episode
if terminal:
# Get exploration rate
exploration = policy_net.exploration_rate
# Clear print row content
clear_row = '\r' + ' ' * 79 + '\r'
print(clear_row, end='')
print(f'Run: {run}, ', end='')
print(f'Exploration: {exploration: .3f}, ', end='')
average_reward = np.mean(rewards)
print(f'Average reward: {average_reward:4.1f}, ', end='')
mean_assignment_to_arrival = np.mean(info['assignment_to_arrival'])
print(f'Mean assignment to arrival: {mean_assignment_to_arrival:4.1f}, ', end='')
mean_call_to_arrival = np.mean(info['call_to_arrival'])
print(f'Mean call to arrival: {mean_call_to_arrival:4.1f}, ', end='')
demand_met = info['fraction_demand_met']
print(f'Demand met {demand_met:0.3f}')
# Add to results lists
results_run.append(run)
results_exploration.append(exploration)
results_score.append(total_reward)
results_mean_call_to_arrival.append(mean_call_to_arrival)
results_mean_assignment_to_arrival.append(mean_assignment_to_arrival)
# Save model if best reward
total_reward = np.sum(rewards)
if total_reward > best_reward:
best_reward = total_reward
# Copy weights to best net
best_net.load_state_dict(policy_net.state_dict())
################################################################
# 18b Check for end of learning #
################################################################
if run == TRAINING_EPISODES:
continue_learning = False
# End episode loop
break
####################################################################
# 19 Update policy net #
####################################################################
# Avoid training model if memory is not of sufficient length
if len(memory.memory) > REPLAY_START_SIZE:
# Update policy net
optimize(policy_net, target_net, memory.memory)
################################################################
# 20 Update target net periodically #
################################################################
# Use load_state_dict method to copy weights from policy net
if all_steps % SYNC_TARGET_STEPS == 0:
target_net.load_state_dict(policy_net.state_dict())
############################################################################
# 21 Learning complete - plot and save results #
############################################################################
# Target reached. Plot results
plot_results(results_run, results_exploration, results_score,
results_mean_call_to_arrival, results_mean_assignment_to_arrival)
# SAVE RESULTS
run_details = pd.DataFrame()
run_details['run'] = results_run
run_details['exploration '] = results_exploration
run_details['mean_call_to_arrival'] = results_mean_call_to_arrival
run_details['mean_assignment_to_arrival'] = results_mean_assignment_to_arrival
filename = 'output/' + RESULTS_NAME + '.csv'
run_details.to_csv(filename, index=False)
############################################################################
# Test best model #
############################################################################
print()
print('Test Model')
print('----------')
best_net.exploration_rate = 0
best_net.eval()
# Set up results dictionary
results = dict()
results['call_to_arrival'] = []
results['assign_to_arrival'] = []
results['demand_met'] = []
# Replicate model runs
for run in range(30):
# Reset game environment and get first state observations
state = sim.reset()
state = np.reshape(state, [1, observation_space])
# Continue loop until episode complete
while True:
# Get action to take (se eval mode to avoid dropout layers)
best_net.eval()
action = best_net.act(state)
# Act
state_next, reward, terminal, info = sim.step(action)
# Reshape state into 2D array with state observations as first 'row'
state_next = np.reshape(state_next, [1, observation_space])
# Update state
state = state_next
if terminal:
print(f'Run: {run}, ', end='')
mean_assignment_to_arrival = np.mean(info['assignment_to_arrival'])
print(f'Mean assignment to arrival: {mean_assignment_to_arrival:4.1f}, ', end='')
mean_call_to_arrival = np.mean(info['call_to_arrival'])
print(f'Mean call to arrival: {mean_call_to_arrival:4.1f}, ', end='')
demand_met = info['fraction_demand_met']
print(f'Demand met: {demand_met:0.3f}')
# Add to results
results['call_to_arrival'].append(mean_call_to_arrival)
results['assign_to_arrival'].append(mean_assignment_to_arrival)
results['demand_met'].append(demand_met)
# End episode loop
break
results = pd.DataFrame(results)
filename = './output/results_' + RESULTS_NAME +'.csv'
results.to_csv(filename, index=False)
print()
print(results.describe())
return run_details
# +
######################## MODEL ENTRY POINT #####################################
# Run model and return last run results
last_run = qambo()
|
experiments/1_incident_points_3_ambo/03_qambo_noisy_3dqn.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [LEGALST-123] Lab 18: Regular Expressions
#
# This lab will cover the basics of regular expression: finding, extracting and manipulating pieces of text based on specific patterns within strings.
#
# *Estimated Time: 45 minutes*
#
# ### Table of Contents
#
# [The Data](#section data)<br>
#
# [Overview](#section context)<br>
#
# 0- [Matching with Regular Expressions](#section 0)<br>
#
# 1 - [Introduction to Essential RegEx](#section 1)<br>
#
# 1 - [Special Characters](#subsection 1)
#
# 2 - [Quantifiers](#subsection 2)
#
# 3 - [Sets](#subsection 3)
#
# 4 - [Special Sequences](#subsection 4)
#
# 5 - [Groups and Logical OR](#subsection 4)
#
# 2- [Python RegEx Methods](#section 2)<br>
#
# 3 - [Valuation Extraction](#section 3)<br>
# ## The Data <a id='data'></a>
#
# You will again be working with the Old Bailey data set to practice matching and manipulating pieces of the textual data.
#
#
# ## Overview <a id='data'></a>
# Regular Expressions operations ("RegEx") are a very flexible version of the text search function that you find in most text processing software. In those regular search functions, you press `ctrl+F` (or `command+F`) and type in the search phrase you are looking for e.g. "Congress". If your software finds an exact match for your search phrase ("Congress"), it jumps to its position in the text and you can take it from there.
#
# Thinking a bit more abstractly about this, "Congress" is nothing else than a very specific search. In it, we ask the search function to report the position where it finds a capital "C" followed seven lower case letters ("o", "n", "g", "r", "e","s","s"), all in a specific order. Depending on your text, it may have been sufficient to let your search function look for all words starting with the captial letter "C", or for those words starting with "C" and ending with "ess". This kind of flexibility is exactly what RegEx provides.
#
# RegEx is more flexible than the customary search function as it does not restrict you to spell out the literal word, number or phrase you are looking for. Rather, in RegEx you can describe the necessary characteristics for a match. You can enter these characteristics based on rules and special characters that make RegEx what it is.
#
# Regular expressions are useful in a variety of applications, and can be used in different programs and programming languages. We will start by learning the general components of regular expressions, using a simple online tool, Regex101. Then at the end of the workshop, we'll learn how to use regular expressions to conduct devaluation exploration on the Old Bailey dataset - we will look at how often plaintiffs had the amount they were charged with stealing reduced when they were sentenced by matching valuations in the text such as 'value 8s 6p'.
#
# __IT IS IMPORTANT to have an experimental mindset as you go through today's practice problems.__ Practice and curiosity are the keys to success! Each indiviual character expression may output a simple pattern, but you will need to explore different combinations to match more and more complicated sets of strings. Feel free to go beyond what the questions ask and test different expressions as you work through this notebook.
#
#
# __Dependencies__: Run the cell below. We will go over what this python library does in the Python Methods section of this lab.
import re
# ---
#
# ## Introduction to Essential RegEx<a id='section 1'></a>
# ### 0. Matching with Regular Expressions <a id='subsection 0'></a>
#
#
# Before we dive into the different character expressions and their meanings, let's explore what it looks like to match some basic expressions. Open up [Regex101](https://regex101.com/r/Una9U7/4), an online Python regular expression editor. This editor will allow us to input any test string and practice using regular expressions while receiving verification and tips in real-time. There should already be an excerpt from the Old Bailey Set (edited, for the sake of practice problems) in the `Test String` box.
#
# You can think of the `Regular Expression` field like the familiar `ctrl+F` search box.
# Try typing in the following, one at a time, to the `Regular Expression` field:
# ~~~ {.input}
# 1. lowercase letter: d
# 2. uppercase letter: D
# 3. the word: lady
# 4. the word: Lady
# 5. the word: our
# 6. the word: Our
# 7. a single space
# 8. a single period
# ~~~
#
# __Question 1:__ What do you notice?
#
# __Your Answer:__
# *Write your Answer Here:*
# Note that:
# 1. RegEx is case sensitive: it matches _exactly_ what you tell it to match.
# 2. RegEx looks for the exact order of each character you input into the expression. In the entire text, it found 'our' in 'Hon`our`able' and 'F`our`score'. However, nowhere in the text was there the exact sequence of letters O-u-r starting with a capital 'O', so 'Our' doesn't match anything.
# 3. The space character ` ` highlights all the single spaces in the text.
# 4. the period character `.` matches all the characters in the text, not just the periods... why?
#
# This last question takes us now to what is called __special characters__.
#
# ---
# ### 1. Special Characters <a id='subsection 1'></a>
#
# Strings are composed of characters, and we are writing patterns to match specific sequences of characters.
# Various characters have special meaning in regular expressions. When we use these characters in an expression,
# we aren't matching the identical character, we're using the character as a placeholder for some other character(s)
# or part(s) of a string.
#
#
# ~~~ {.input}
#
# . any single character except newline character
# ^ start of string
# $ end of entire string
# \n new line
# \r carriage return
# \t tab
#
# ~~~
#
# Note: if you want to actually match a character that happens to be a special character, you have to escape it with a backslash
# `\`.
#
# __Question 2:__ Try typing the following special characters into the `Regular Expression` field on the same Regex101 site. What happens
# when you type:
#
# 1. `Samuel` vs. `^Samuel` vs. `Samuel$`?
#
# 2. `.` vs. `\.`
#
# 3. `the` vs. `th.` vs. `th..` ?
#
# __Your Answer:__
# *Write your Answer Here*:
#
# 1.
#
# 2.
#
# 3.
# SOLUTION:
# ~~~ {.input}
# 1.
# `Samuel` will match all instances of the pattern `Samuel` in the text
# `^Samuel` will match only the instances of `Samuel` at the beginning of the text
# `Samuel$` will match only the instance of `Samuel` at the end of the text
#
# 2.
# `.` matches all individual characters in the text
# `\.` matches all periods in the text
#
# 3.
# `the` matches all instances of the pattern `the` in the text
# `th.` matches all instances of patterns starting with `th` and ending in any character
# `th..` matches all instances of patterns starting with `th` any two characters follwing it
# ~~~
# ---
# ### 2. Quantifiers<a id='subsection 2'></a>
#
# Some special characters refer to optional characters, to a specific number of characters, or to an open-ended
# number of characters matching the preceding pattern.
#
# ~~~ {.input}
# * 0 or more of the preceding character/expression
# + 1 or more of the preceding character/expression
# ? 0 or 1 of the preceding character/expression
# {n} n copies of the preceding character/expression
# {n,m} n to m copies of the preceding character/expression
# ~~~
#
#
# __Question 3:__ For this question, click [here](https://regex101.com/r/ssAUXx/1) to open another Regex101 page.
#
# What do the expressions `of`, `of*`, `of+`, `of?`, `of{1}`, `of{1,2}` match? Remember that the quantifier only applies to the character *immediately* preceding it. For example, the `*` in `of*` applies only to the `f`, so the expression looks for a pattern starting with __exactly one__ `o` and __0 or more__ `f`'s.
#
# __Your Answer:__
# *Write your answer here:*
# SOLUTION:
#
# ~~~ {.input}
#
# - `of`: matches all instances of the pattern `of` in the text
# - `of+`: matches all intances of a pattern starting with exactly one `o` and 1 or more `f`'s
# - `of?`: matches all instances of a pattern starting with exactly one `o` and at most one `f` after it.
# - `of{1}`: matches all instances of a pattern starting with exactly one `o` and exactly one `f`
# - `of{1,2}`: matches all instances of a pattern starting with exactly one `o` and 1 OR 2 `f`'s after it.
#
# ~~~
# ---
# ### 3. Sets<a id='subsection 3'></a>
#
# A set by itself is merely a __collection__ of characters the computer may choose from to match a __single__ character in a pattern. We can define these sets of characters using `square brackets []`.
#
# Within a set of square brackets, you may list characters individually, e.g. `[aeiou]`, or in a range, e.g. `[A-Z]` (note that all regular expressions are case sensitive).
#
#
# You can also create a complement set by excluding certain characters, using `^` as the first character
# in the set. The set `[^A-Za-z]` will match any character except a letter. All other special characters loose
# their special meaning inside a set, so the set `[.?]` will look for a literal period or question mark.
#
# The set will match only one character contained within that set, so to find sequences of multiple characters from
# the same set, use a quantifier like `+` or a specific number or number range `{n,m}`.
#
# ~~~ {.input}
# [0-9] any numeric character
# [a-z] any lowercase alphabetic character
# [A-Z] any uppercase alphabetic character
# [aeiou] any vowel (i.e. any character within the brackets)
# [0-9a-z] to combine sets, list them one after another
# [^...] exclude specific characters
# ~~~
#
# __Question 4:__ Let's switch back to the excerpt from the Old Bailey data set (link [here](https://regex101.com/r/Una9U7/2) for convenience). Can you write a regular expression that matches __all consonants__ in the text string?
#
# __Your Answer:__
# +
# YOUR EXPRESSION HERE
# -
#SOLUTION:
[^aeiou]
# ---
#
# ### 4. Special sequences<a id='subsection 4'></a>
#
# If we want to define a set of all 26 characters of the alphabet, we would have to write an extremely long expression inside a square bracket. Fortunately, there are several special characters that denote special sequences. These begin with a `\` followed by a letter.
#
# Note that the uppercase version is usually the complement of the lowercase version.
#
# ~~~ {.input}
# \d Any digit
# \D Any non-digit character
# \w Any alphanumeric character [0-9a-zA-Z_]
# \W Any non-alphanumeric character
# \s Any whitespace (space, tab, new line)
# \S Any non-whitespace character
# \b Matches the beginning or end of a word (does not consume a character)
# \B Matches only when the position is not the beginning or end of a word (does not consume a character)
# ~~~
#
# __Question 5:__ Write a regular expression that matches all numbers (without punctuation marks or spaces) in the Old Bailey excerpt. Make sure you are matching whole numbers (i.e. `250`) as opposed to individual digits within the number (i.e. `2`, `5`, `0`).
#
# __Your Answer:__
# +
# YOUR EXPRESSION HERE
# -
#SOLUTION
\d+
# __Question 6:__ Write a regular expression that matches all patterns with __at least__ 2 and __at most__ 3 digit and/or white space characters in the Old Bailey excerpt.
#
# __Your Answer:__
# +
#YOUR EXPRESSION HERE
# -
#Solution
[\d\s]{2,3}
# ---
#
# ### 5. Groups and Logical OR<a id='subsection 5'></a>
#
# Parentheses are used to designate groups of characters, to aid in logical conditions, and to be able to retrieve the
# contents of certain groups separately.
#
# The pipe character `|` serves as a logical OR operator, to match the expression before or after the pipe. Group parentheses
# can be used to indicate which elements of the expression are being operated on by the `|`.
#
# ~~~ {.input}
# | Logical OR opeator
# (...) Matches whatever regular expression is inside the parentheses, and notes the start and end of a group
# (this|that) Matches the expression "this" or the expression "that"
# ~~~
#
# __Question 7:__ Write an expression that matches groups of `Samuel` or `Prisoner` in the Old Bailey excerpt.
#
# __Your Answer:__
# +
# YOUR EXPRESSION HERE
# -
#SOLUTION
(Samuel|Prisoner)
# ---
# ## Python RegEx Methods <a id='section 2'></a>
#
# So how do we actually use RegEx for analysis in Python?
#
# Python has a RegEx library called `re` that contains various methods so we can manipulate text using RegEx. The following are some useful Python Methods we may use for text analysis:
#
#
# - ``.findall(pattern, string)``: Checks whether your pattern appears somewhere inside your text (including the start). If so, it returns all phrases that matched your pattern, but not their position.
# - ``.sub(pattern, repl, string)``: Return the string obtained by replacing the leftmost non-overlapping occurrences of pattern in string by the replacement repl.
# - ``.split(pattern, string)``: Split string by the occurrences of pattern. If capturing parentheses are used in pattern, then the text of all groups in the pattern are also returned as part of the resulting list.
#
#
# We will only be using the `.findall()` method for the purposes of today's lab, so don't worry if the functionality of each method isn't clear right now. If you are curious about all the module content within the `re` library, take a look at the [documentation for `re`](https://docs.python.org/2/library/re.html) on your own time!
# ---
#
# ## Extracting Valuation from Old Bailey <a id='section 3'></a>
#
# Let's apply our new RegEx knowledge to extract all valuation information from the text!
#
# The next cell simply assigns a long string containing three separate theft cases to a variable called `old_bailey`. Within the text are valuations which indicate the worth of the items stolen. We will use this string, what we can observe about the format of valuation notes in the text, and what we just learned about regular expressions to __find all instances of valuations in the text__.
#
# Valuations will look something like: `val. 4 s. 6 d.`
#
# *Note:* British Currency before 1971 was divided into pounds (`l`), shillings (`s`), and pennies (`d`) - that's what the letters after the values represent. We want to make sure to keep the values and units together when extracting valuations.
#
# __STEP 1__: We will first write expression(s) that will match the valuations.
# Take a moment to look for a pattern you notice across the valuations:
old_bailey = """"<NAME>, of the Parish of St. James Westminster, was indicted for feloniously Stealing 58 Diamonds set in
Silver gilt, value 250 l. the Goods of the Honourable Catherine Lady Herbert, on the 28th of July last. It appeared that the
Jewels were put up in a Closet, which was lockt, and the Prisoner being a Coachman in the House, took his opportunity to take
them; the Lady, when missing them, offered a Reward of Fourscore Pounds to any that could give any notice of it; upon enquiry,
the Lady heard that a Diamond was sold on London-Bridge, and they described the Prisoner who sold it, and pursuing him, found
the Prisoner at East-Ham, with all his Goods bundled up ready to be gone, and in his Trunk found all the Diamonds but one, which
was found upon him in the Role of his Stocking, when searcht before the Justice. He denied the Fact, saying, He found them upon
a great Heap of Rubbish, but could not prove it; and that being but a weak Excuse, the Jury found him guilty.
<NAME>, was
indicted for stealing eleven crown pieces, twenty four half crowns, one Spanish piece, val. 4 s. 6 d. one silk purse, and
4 s. 6 d. in silver, the goods of Ann Kempster, in the dwelling house of <NAME>. December 17. Acquitted. He was a second
time indicted for stealing one pair of stockings, val. 6 d. the goods of <NAME> .
GEORGE MORGAN was indicted for that he, about the hour of ten in the night of the 10th of December , being in the dwelling-house
of <NAME> , feloniously did steal two hundred and three copper halfpence, five china bowls, value 30s. a tea-caddie,
value 5s. a pound of green tea, value 8s. four glass rummers, value 2s. and a wooden drawer, called a till, value 6d. the
property of the said George, and that he having committed the said felony about the hour of twelve at night, burglariously
did break the dwelling-house of the said George to get out of the same."""
# You might notice that there are multiple ways in which valuations are noted. It can take the form:
#
# ~~~ {.input}
# value 30s.
# val. 6 d.
# 4 s. 6 d.
# ~~~
#
# ...and so on.
#
# Fortunately, we only care about the values and the associaed units, so the ommission or abbreviation of the word `value` can be ignored - we only care about:
#
# ~~~ {.input}
# 30s.
# 6 d.
# 4 s. 6 d.
# ~~~
#
# Unfortunately, we can see that the format is still not consistent. The first one has no space between the number and unit, but the second and third do. The first and second have a single number and unit, but the third has two of each.
#
# How might you write an expression that would account for the variations in how valuations are written? Can you write a single regular expression that would match all the different forms of valuations exactly? Or do we need to have a few different expressions to account for these differnces, look for each pattern individually, and combine them somehow in the end?
#
# Real data is messy. When manipulating real data, you will inevitably encounter inconsistencies and you will need to ask yourself questions such as the above. You will have to figure out how to clean and/or work with the mess.
#
# With that in mind, click [here](https://regex101.com/r/2lal6d/1) to open up a new Regex101 with `old_bailey` already in the Test String. We will compose a regular expression, in three parts, that will account for all forms of valuations in the string above.
#
# __PART 1: Write an expression__ that matches __all__ valuations of the form `30s.` AND `6 d.`, but does not match _anything else_ (e.g. your expression should not match any dates). Try not to look at the hints on your first attempt! Save this expression __as a string__ in `exp1`.
#
#
# _Hint1:_ Notice the structure of valuations. It begins with a number, then an _optional_ space, then a single letter followed by a period.
#
# _Hint2:_ What _quantifier_ allows you to choose _0 or more of the previous character_?
#
# _Hint3:_ If you are still stuck, look back to the practice problems and see that we've explored/written expressions to match all components of this expression! It's just a matter of putting it together.
#
#Your Expression Here
exp1 =
#SOLUTION
exp1 = '\d+ ?[a-z]\.'
# __PART 2:__ For the third case we found above, there are multiple values and units in the valuation. What can you add to what you came up with above so that we have another expression that matches this specific case? Save this expression as a string in `exp2`.
#Your Expression Here
exp2 = ...
#SOLUTION
exp2 = '\d+ [a-z]\. \d+ [a-z]\.'
# __PART 3:__ Now that you have expressions that account for the different valuation formats, combine it into one long expression that looks for (_hint_) one expression __OR__ the other. Set this expression to `final`. Be careful about the order in which you ask the computer to look for patterns (i.e. should it look for the shorter expression first, or the longer expression first?). Save this final expression as a string in `final`.
#Your Expression Here
final =
#SOLUTION
final = '\d+ [a-z]\. \d+ [a-z]\.|\d+ ?[a-z]\.'
# __STEP 2:__ Now that you have the right regular expression that would match our valuations, how would you use it to _extract_ all instances of valuations from the text saved in `old_bailey`?
#
# Remember, you need to input your regular expression as a __string__ into the method.
# +
#Your Expression Here
# -
#SOLUTION
re.findall(final, old_bailey)
# __Congratulations!!__ You've successfully extracted all valuations from our sample. When you are extracting valuations from a larger text for your devaluation exploration, keep in mind all the possible variations in valuation that may not have been covered by our example above. You now have all the skills necessary to tweak the expression to account for such minor variations -- Good Luck!
# ---
#
# ## Bibliography
# - The Python Standard Library. (2018, February). Regular Expression Operations. https://docs.python.org/2/library/re.html
# - <NAME>. (2006, June). British Currency Before 2971. http://www.victorianweb.org/economics/currency.html
# - The Proceedings of the Old Bailey. https://www.oldbaileyonline.org/
#
# ---
#
# Notebook Developed by: <NAME>
#
# Data Science Modules: http://data.berkeley.edu/education/modules
|
labs/18_Regular Expressions/18_Regular_Expressions_solutions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
"""
===================================================
Faces recognition example using eigenfaces and SVMs
===================================================
The dataset used in this example is a preprocessed excerpt of the
"Labeled Faces in the Wild", aka LFW_:
http://vis-www.cs.umass.edu/lfw/lfw-funneled.tgz (233MB)
.. _LFW: http://vis-www.cs.umass.edu/lfw/
original source: http://scikit-learn.org/stable/auto_examples/applications/face_recognition.html
"""
print __doc__
from time import time
import logging
import pylab as pl
import numpy as np
from sklearn.cross_validation import train_test_split
from sklearn.datasets import fetch_lfw_people
from sklearn.grid_search import GridSearchCV
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.decomposition import RandomizedPCA
from sklearn.svm import SVC
from PIL import Image
# Display progress logs on stdout
logging.basicConfig(level=logging.INFO, format='%(asctime)s %(message)s')
###############################################################################
# Download the data, if not already on disk and load it as numpy arrays
lfw_people = fetch_lfw_people(min_faces_per_person=70, resize=0.4)
# introspect the images arrays to find the shapes (for plotting)
n_samples, h, w = lfw_people.images.shape
np.random.seed(42)
# for machine learning we use the data directly (as relative pixel
# position info is ignored by this model)
X = lfw_people.data
n_features = X.shape[1]
# the label to predict is the id of the person
y = lfw_people.target
target_names = lfw_people.target_names
n_classes = target_names.shape[0]
print "Total dataset size:"
print "n_samples: %d" % n_samples
print "n_features: %d" % n_features
print "n_classes: %d" % n_classes
###############################################################################
# Split into a training and testing set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=42)
###############################################################################
# Compute a PCA (eigenfaces) on the face dataset (treated as unlabeled
# dataset): unsupervised feature extraction / dimensionality reduction
n_components = 150
print "Extracting the top %d eigenfaces from %d faces" % (n_components, X_train.shape[0])
t0 = time()
pca = RandomizedPCA(n_components=n_components, whiten=True).fit(X_train)
print "done in %0.3fs" % (time() - t0)
eigenfaces = pca.components_.reshape((n_components, h, w))
print "Projecting the input data on the eigenfaces orthonormal basis"
t0 = time()
X_train_pca = pca.transform(X_train)
X_test_pca = pca.transform(X_test)
print "done in %0.3fs" % (time() - t0)
###############################################################################
# Train a SVM classification model
print "Fitting the classifier to the training set"
t0 = time()
param_grid = {
'C': [1e3, 5e3, 1e4, 5e4, 1e5],
'gamma': [0.0001, 0.0005, 0.001, 0.005, 0.01, 0.1],
}
# for sklearn version 0.16 or prior, the class_weight parameter value is 'auto'
clf = GridSearchCV(SVC(kernel='rbf', class_weight='balanced'), param_grid)
clf = clf.fit(X_train_pca, y_train)
print "done in %0.3fs" % (time() - t0)
print "Best estimator found by grid search:"
print clf.best_estimator_
###############################################################################
# Quantitative evaluation of the model quality on the test set
print "Predicting the people names on the testing set"
t0 = time()
y_pred = clf.predict(X_test_pca)
print "done in %0.3fs" % (time() - t0)
print classification_report(y_test, y_pred, target_names=target_names)
print confusion_matrix(y_test, y_pred, labels=range(n_classes))
###############################################################################
# Qualitative evaluation of the predictions using matplotlib
def plot_gallery(images, titles, h, w, n_row=3, n_col=4):
"""Helper function to plot a gallery of portraits"""
pl.figure(figsize=(1.8 * n_col, 2.4 * n_row))
pl.subplots_adjust(bottom=0, left=.01, right=.99, top=.90, hspace=.35)
for i in range(n_row * n_col):
pl.subplot(n_row, n_col, i + 1)
pl.imshow(images[i].reshape((h, w)), cmap=pl.cm.gray)
pl.title(titles[i], size=12)
pl.xticks(())
pl.yticks(())
# plot the result of the prediction on a portion of the test set
def title(y_pred, y_test, target_names, i):
pred_name = target_names[y_pred[i]].rsplit(' ', 1)[-1]
true_name = target_names[y_test[i]].rsplit(' ', 1)[-1]
return 'predicted: %s\ntrue: %s' % (pred_name, true_name)
prediction_titles = [title(y_pred, y_test, target_names, i)
for i in range(y_pred.shape[0])]
plot_gallery(X_test, prediction_titles, h, w)
# plot the gallery of the most significative eigenfaces
eigenface_titles = ["eigenface %d" % i for i in range(eigenfaces.shape[0])]
plot_gallery(eigenfaces, eigenface_titles, h, w)
pl.show()
# -
|
examples/pca/eigenfaces.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py362
# language: python
# name: py362
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 292} colab_type="code" id="36p8DSfO100T" outputId="e06ff593-1f9b-4c0e-adf5-d260df5023ab"
# !pip install networkx
# !pip install rdflib
# !pip install numpy
# !pip install sparqlwrapper
# + colab={} colab_type="code" id="1VvYZ1WNUfib"
import rdflib
import numpy as np
from collections import Counter
from SPARQLWrapper import SPARQLWrapper, JSON
import networkx as nx
import requests
# -
def query_wiki_article_title(query):
params = {
'action':"query",
'list':"search",
'srsearch': query,
'format':"json"
}
resp = requests.get("https://en.wikipedia.org/w/api.php", params)
if resp.status_code != 200:
return None
results = resp.json()
if len(results):
return results['query']['search'][0]['title'].replace(" ", "_")
# + colab={} colab_type="code" id="wT1sBX8dUfif"
def get_link_set(article_link):
q ="""PREFIX p: <http://www.wikidata.org/prop/>
SELECT DISTINCT ?thing ?relation
WHERE { ?thing ?relation <"""+ article_link +"""> . }
LIMIT 10"""
sparql = SPARQLWrapper("http://dbpedia.org/sparql")
sparql.setReturnFormat(JSON)
sparql.setQuery(q) # the previous query as a literal string
json = sparql.query().convert()
links = json["results"]["bindings"]
return set([obj["thing"]["value"] for obj in links])
# -
def get_link_set_outlinks(article_link):
q ="""PREFIX dbr: <http://dbpedia.org/resource/>
PREFIX owl: <http://www.w3.org/2002/07/owl#>
SELECT DISTINCT ?p ?o
WHERE {
<""" + article_link + """> ?p ?o .
?o a owl:Thing .
}
LIMIT 10"""
sparql = SPARQLWrapper("http://dbpedia.org/sparql")
sparql.setReturnFormat(JSON)
sparql.setQuery(q) # the previous query as a literal string
json = sparql.query().convert()
links = json["results"]["bindings"]
return set([obj["o"]["value"] for obj in links])
def create_graph(raw_titles, get_link_set_fn):
G = nx.Graph()
for cand in raw_titles:
formatted_cand = query_wiki_article_title(cand)
candlink = "http://dbpedia.org/resource/" + formatted_cand
G.add_node(candlink)
linkSet = get_link_set_fn(candlink)
for link in linkSet:
G.add_node(link)
# Explore 1x time
deg2linkSet = get_link_set_fn(link)
for link2 in deg2linkSet:
G.add_node(link2)
G.add_edge(link, link2)
# Create edges to links
G.add_edge(candlink, link)
return G
def generate_graph_pickles(output_candidates_fname="output_candidates"):
with open(output_candidates_fname) as f:
contents = f.readlines()
contents = [x.strip() for x in contents]
for i, titles in enumerate(contents):
print ("creating topic " + str(i) + " pickle file")
G = create_graph(titles.split(), get_link_set)
fname = "Topics/topic" +str(i)+ "G"
nx.write_gml(G, fname)
f.close()
def extract_ranked_cands(seed_cands, top_links):
"""
Extract the generated candidates' rank out from the raw order after
DBpedia exploration.
:param seed_cands: list of generated candidates
:param top_links: list of (link, score) tuples
"""
cand_pool = set(seed_cands)
print (cand_pool)
top_cands = []
for raw_link, _ in top_links:
last_sep = raw_link.rfind("/")
raw_title = raw_link[last_sep + 1:]
title = raw_title.lower()
if title in cand_pool:
top_cands.append(title)
return top_cands
def get_best_label(label_list,num):
fname = "Topics/topic" +str(num)+ "G"
G = nx.read_gml(fname)
Gc = max(nx.connected_component_subgraphs(G), key=len)
centrality_measure = nx.betweenness_centrality(Gc)
top_links = sorted(centrality_measure.items(), key=lambda x: x[1], reverse=True)
cands_ranks = extract_ranked_cands(label_list, top_links)
return cands_ranks
get_best_label(d[0],0)
mygraph = nx.read_gml("Topics/topic0G")
Gc = max(nx.connected_component_subgraphs(mygraph), key=len)
clo_gen = nx.betweenness_centrality(Gc)
sorted(clo_gen.items(), key=lambda x: x[1], reverse=True)[:20]
|
model_run/DBpedia_experiment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Uniview module for IceCube event 170922A
#
# *<NAME>, 2018*
#
# ### Imports and function definitions
# +
#This directory contains all the data needed for the module. It should be in the same directory as the notebook
dataFolder = "data"
import sys, os, shutil, errno, string, urllib
sys.path.append(( os.path.abspath( os.path.join(os.path.realpath("__file__"), os.pardir, os.pardir) )))
import uvmodlib.v1 as uvmod
# +
# # Run this if you want to build in 3.0 rather than 2.0
# from config import Settings
# Settings.uvcustommodulelocation='C:\\Users\\msubbarao\\SCISS\\Uniview Theater 3.0\\Custom Modules'
# -
# ### USES Conf Template
Template = """mesh
{
data ICgeom ./modules/$folderName/IceCubeGeometry.raw
data ICgeomTop ./modules/$folderName/IceCubeGeometryTop.raw
data ICevent ./modules/$folderName/ic170922.tsort.raw # cat ic170922.raw | sort -n -k 4 > ic170922.tsort.raw
data quad ./modules/$folderName/quad.3ds
dataHints ICgeom disablePostprocessing
dataHints ICgeomTop disablePostprocessing
dataHints ICevent disablePostprocessing
cullRadius $cr
glslVersion 330
propertyCollection
{
__objectName__
{
vec1f geomRadius 3.5 | public | desc "geometry point size " | widget slider | range 0 20
vec1f eventRadius 15 | public | desc "event scale point size " | widget slider | range 0 20
vec1f tfac 10 | public | desc "time width " | widget slider | range 0. 20
vec1f tmax 5000 | public | desc "max time color " | widget slider | range 0. 11800 #for color bar
vec4f lineColor 0.0 0.25 0 0.5 | public
bool buildUp false | public | desc "keep showing the observations after t"
vec1f eventTime -1000 | public | desc "event time "
vec1f transitionLength 10 | public | desc "transition length in seconds"
bool jump true | public | desc "jump to time without transition"
}
}
############# to hold the time information
renderTexture
{
name stateTexture
width 1
height 1
numTextures 1
isPingPong true
isPersistent true
isFramePersistent true
internalTextureFormat GL_RGB32F
magnify GL_NEAREST
minify GL_NEAREST
}
############# set Transition State
pass
{
useDataObject quad
renderTarget
{
name stateTexture
enableColorClear false
}
shader
{
type defaultMeshShader
{
vertexShader ./modules/$folderName/pass0.vs
fragmentShader ./modules/$folderName/state.fs
textureFBO stateTexture stateTexture
stateManagerVar __objectName__.transitionLength transitionLength
stateManagerVar __objectName__.jump jump
stateManagerVar __objectName__.eventTime eventTime
parameter2f timeRange -1000 11800
}
}
}
############# Detector geometry, points
pass
{
useDataObject ICgeom
shader
{
type defaultMeshShader
{
geometryShader ./modules/$folderName/ICgeom.gs
vertexShader ./modules/$folderName/ICgeom.vs
fragmentShader ./modules/$folderName/ICgeom.fs
stateManagerVar __objectName__.geomRadius geomRadius
glState
{
UV_CULL_FACE_ENABLE false
UV_BLEND_ENABLE true
UV_DEPTH_ENABLE false
UV_WRITE_MASK_DEPTH true
UV_BLEND_FUNC GL_SRC_ALPHA GL_ONE_MINUS_SRC_ALPHA
}
}
}
}
############# Detector geometry, lines
pass
{
useDataObject ICgeomTop
shader
{
type defaultMeshShader
{
geometryShader ./modules/$folderName/ICgeomLine.gs
vertexShader ./modules/$folderName/ICgeom.vs
fragmentShader ./modules/$folderName/ICgeomLine.fs
stateManagerVar __objectName__.lineColor lineColor
glState
{
UV_CULL_FACE_ENABLE false
#UV_CULL_MODE GL_FRONT
UV_BLEND_ENABLE true
UV_BLEND_FUNC GL_SRC_ALPHA GL_ONE_MINUS_SRC_ALPHA
UV_LINE_WIDTH 2
}
}
}
}
############# Event geometry
pass
{
useDataObject ICevent
shader
{
type defaultMeshShader
{
geometryShader ./modules/$folderName/ICevent.gs
vertexShader ./modules/$folderName/ICevent.vs
fragmentShader ./modules/$folderName/ICevent.fs
textureFBO stateTexture stateTexture
texture cmap ./modules/$folderName/cmap.png
{
wrapModeS GL_CLAMP_TO_EDGE
wrapModeR GL_CLAMP_TO_EDGE
colorspace linear
}
#stateManagerVar __objectName__.eventTime eventTime
stateManagerVar __objectName__.eventRadius eventRadius
stateManagerVar __objectName__.tfac tfac
stateManagerVar __objectName__.tmax tmax
stateManagerVar __objectName__.buildUp buildUp
glState
{
UV_CULL_FACE_ENABLE false
UV_BLEND_ENABLE true
UV_DEPTH_ENABLE false
UV_WRITE_MASK_DEPTH true
UV_BLEND_FUNC GL_SRC_ALPHA GL_ONE_MINUS_SRC_ALPHA
}
}
}
}
}"""
# ### IceCube class
class IceCube():
def __init__(self, object):
self.object = object
uvmod.Utility.ensurerelativepathexsists("ICgeom.gs",dataFolder)
uvmod.Utility.ensurerelativepathexsists("ICgeom.vs",dataFolder)
uvmod.Utility.ensurerelativepathexsists("ICgeom.fs",dataFolder)
uvmod.Utility.ensurerelativepathexsists("ICgeomLine.gs",dataFolder)
uvmod.Utility.ensurerelativepathexsists("ICgeomLine.fs",dataFolder)
uvmod.Utility.ensurerelativepathexsists("ICevent.gs",dataFolder)
uvmod.Utility.ensurerelativepathexsists("ICevent.vs",dataFolder)
uvmod.Utility.ensurerelativepathexsists("ICevent.fs",dataFolder)
self.cr = 1000
self.Scale = 1
def generatemod(self):
self.object.setgeometry(self.object.name+"Mesh.usesconf")
return self.object.generatemod()
def generatefiles(self, absOutDir, relOutDir):
fileName = self.object.name+"Mesh.usesconf"
s = string.Template(Template)
f = open(absOutDir+"\\"+fileName, 'w')
if f:
f.write(s.substitute(folderName = relOutDir,
cr = self.cr,
Scale = self.Scale
))
f.close()
uvmod.Utility.copyfoldercontents(os.getcwd()+"\\"+dataFolder, absOutDir)
# ### Object Instantiation
# +
model = IceCube(uvmod.OrbitalObject())
generator = uvmod.Generator()
scene = uvmod.Scene()
scene.setname("IceCube")
scene.setparent("Earth")
scene.setentrydist(100000)
scene.setunit(1)
scene.setsurfacepositionerrotation(0,0.,-90)
scene.setpositionfile(uvmod.PositionFileTypes.Surface("Earth", 0.0, -89., -0.2))
modinfo = uvmod.ModuleInformation()
# -
# ### Specify Settings and generate the module
# +
model.object.setcameraradius(10)
model.object.setcoord(scene.name)
model.object.setname("IceCube")
model.object.setguiname("/KavliLecture/Larson/IceCube")
model.object.settargetradius(20)
model.object.showatstartup(False)
model.cr = 10000
modinfo.setname("IceCube")
modinfo.setauthor("<NAME><sup>1</sup> and <NAME><sup>2</sup><br />(1)<NAME>,<br />(2)Northwestern University")
modinfo.cleardependencies()
modinfo.setdesc("Uniview module for IceCube event 170922A")
#modinfo.setthumbnail("data/R0010133.JPG")
modinfo.setversion("1.0")
generator.generate("IceCube",[scene],[model],modinfo)
uvmod.Utility.senduvcommand(model.object.name+".reload")
# -
# ## Helper Functions for modifing code
# *Reload Module and Shaders in Uniview*
uvmod.Utility.senduvcommand(model.object.name+".reload; system.reloadallshaders")
# *Copy modified Shader files and reload*
from config import Settings
uvmod.Utility.copyfoldercontents(os.getcwd()+"\\"+dataFolder, Settings.uvcustommodulelocation+'\\'+model.object.name)
uvmod.Utility.senduvcommand(model.object.name+".reload")
# ### Create colormap texture
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import cm
# %matplotlib inline
# +
gradient = np.linspace(0, 1, 256)
gradient = np.vstack((gradient, gradient))
def plot_cmap(colormap):
fig=plt.imshow(gradient, aspect=1, cmap=colormap)
plt.axis('off')
fig.axes.get_xaxis().set_visible(False)
fig.axes.get_yaxis().set_visible(False)
plt.savefig("data/cmap.png", bbox_inches='tight',pad_inches=0)
plot_cmap('viridis')
# -
|
generate.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from sentence_transformers import SentenceTransformer
from sklearn.metrics.pairwise import cosine_similarity
# # Read the posts and comments csv files
subreddit = "computerscience"
scrape_order = 'hot'
df_posts = pd.read_csv(f"../data/raw/{subreddit}_{scrape_order}_posts.csv")
df_posts.head()
df_comments = pd.read_csv(f"../data/raw/{subreddit}_{scrape_order}_comments.csv")
df_comments.head()
df = df_posts.merge(df_comments, left_on='post_id', right_on='post_id', how='left')
df.head()
# Check the number of posts with no comments
print("There are {} posts with no comments".format(df.comment.isna().sum()))
#print("There are {} posts with no content after filtering".format(len(df[df['comment'].str.len() == 0])))
# # Import the bert model
model = SentenceTransformer('bert-base-nli-mean-tokens')
# # Traverse through each post
similarities = np.array([])
for post_id in df['post_id'].unique():
temp = df[df['post_id'] == post_id]
if pd.isnull(temp.iloc[0]['comment']):
similarities = np.append(similarities, 0)
continue
title = temp.iloc[0]['title']
body = temp.iloc[0]['body']
topic = title
# topic = title + body
comments = temp['comment'].to_list()
comments.append(topic)
sentence_embeddings = model.encode(comments)
similarity = cosine_similarity([sentence_embeddings[-1]], sentence_embeddings[:-1])
similarities = np.append(similarities, similarity.flatten())
similarities
df['similarity'] = similarities
df
df.to_csv("../data/results/relevance_output.csv", index=False)
# Notes: this takes a long time to run. Suggestion from another paper: use faiss.
# Also perhaps compare the comments with something else other than title
# # Sample code below to generate embeddings and cosine similarity
sentence_embeddings = model.encode(df_posts['title'].to_list())
sentence_embeddings.shape
sentence_embeddings
cosine_similarity(
[sentence_embeddings[0]],
sentence_embeddings[1:]
)
|
notebooks/Relevance - Andrew 2-9-2022.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] toc="true"
# # Table of Contents
# <p><div class="lev1 toc-item"><a href="#Load-lightcone" data-toc-modified-id="Load-lightcone-1"><span class="toc-item-num">1 </span>Load lightcone</a></div><div class="lev2 toc-item"><a href="#Add-missing-column-for-the-simulations" data-toc-modified-id="Add-missing-column-for-the-simulations-11"><span class="toc-item-num">1.1 </span>Add missing column for the simulations</a></div><div class="lev1 toc-item"><a href="#Set-galaxy-parameters" data-toc-modified-id="Set-galaxy-parameters-2"><span class="toc-item-num">2 </span>Set galaxy parameters</a></div><div class="lev1 toc-item"><a href="#Set-survey-properties-and-header" data-toc-modified-id="Set-survey-properties-and-header-3"><span class="toc-item-num">3 </span>Set survey properties and header</a></div><div class="lev1 toc-item"><a href="#Generate-HI-data-cube" data-toc-modified-id="Generate-HI-data-cube-4"><span class="toc-item-num">4 </span>Generate HI data cube</a></div><div class="lev2 toc-item"><a href="#Generate-catalogue-of-spectra" data-toc-modified-id="Generate-catalogue-of-spectra-41"><span class="toc-item-num">4.1 </span>Generate catalogue of spectra</a></div><div class="lev3 toc-item"><a href="#Grid-spectra-to-data-cube" data-toc-modified-id="Grid-spectra-to-data-cube-411"><span class="toc-item-num">4.1.1 </span>Grid spectra to data cube</a></div>
# -
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
# +
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
import numpy as np
from astropy import units as u
from astropy.io import fits
from pyhis import surveys, galaxies, core, helpers
from pyhiframe.pyhiframe import HIConverter
# -
from matplotlib import rcParams
rcParams['figure.figsize'] = (12, 6)
# # Load lightcone
# Load the lightcone, which should be a table format such as `pandas`. Right now, the required columns are R.A., Dec., redshift, and HI mass.
# lightcone = pd.read_csv('../data/processed/lightcones/BolshoiP_Bethermin_Mstar_z_0_1.csv')
lightcone = pd.read_csv('../simulations/new_api_test/lightcone.csv')
lightcone = lightcone[lightcone['z'] > 0.5]
lightcone.head()
fig, (ax1, ax2) = plt.subplots(nrows=2)
sns.distplot(np.log10(lightcone['mhalo']), kde=False, ax=ax1)
sns.distplot(lightcone['z'], kde=False, ax=ax2);
# ## Add missing column for the simulations
# On top of the current column, we also require HI masses, radial velocities, distances, and HI fluxes.
lightcone = helpers.extend_lightcone(lightcone)
lightcone.head()
lightcone.to_csv('../simulations/new_api_test/lightcone.csv', index=False)
# +
fig, (ax1, ax2) = plt.subplots(nrows=2)
ax1.plot(lightcone['distance'], lightcone['z'])
ax1.set_xlabel('Distance [Mpc]')
ax1.set_ylabel('Redshift')
sns.distplot(np.log10(lightcone['flux']), kde=False, ax=ax2);
# -
# # Set galaxy parameters
# The galaxy shape parameters according to which HI is painted in the halos.
n_samples = int(1e5)
galaxy = galaxies.Stewart2014(n_samples)
fig, (ax1, ax2) = plt.subplots(nrows=2)
sns.distplot(galaxy.v_turb, ax=ax1)
sns.distplot(galaxy.solid_rot, ax=ax2)
# # Set survey properties and header
survey = surveys.GbtType()
# surveyproperties, header = surveys.SKA_type()
vars(survey).keys()
survey.header
# # Generate HI data cube
# ## Generate catalogue of spectra
# +
sample = lightcone.sample(n_samples)
spectra = core.generate_spectra(sample, survey, galaxy)
# -
spectra.shape
_ = [pl.plot(survey.nu_grid, spec, c='k', alpha=0.3) for spec in spectra[:120]]
pl.gca().invert_xaxis();
# ### Grid spectra to data cube
cube = core.generate_cube(spectra, survey.header, sample)
pl.plot(np.nanmean(cube, (1,2)))
fits.writeto('../simulations/new_api_test/datacube.fits', cube, survey.header, overwrite=True)
|
notebooks/explore_package.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#import json
import numpy as np
import pandas as pd
dir_name = "userAction_Recommendation/"
file_name1 = "boston.csv"
boston_info = pd.read_csv(dir_name + file_name1)
#print(boston_info.head())
file_name2 = "columbus.csv"
columbus_info = pd.read_csv(dir_name + file_name2)
file_name3 = "review_boston.csv"
review_bos = pd.read_csv(dir_name + file_name3)
file_name4 = "review_columbus.csv"
review_col = pd.read_csv(dir_name + file_name4)
#data cleaning
print(len(review_bos))
review_bos = review_bos.drop(review_bos[review_bos.user_id == '#NAME?'].index)
review_bos = review_bos.drop(review_bos[review_bos.user_id == '#VALUE!'].index)
review_bos = review_bos.drop(review_bos[review_bos.business_id == '#NAME?'].index)
review_bos = review_bos.drop(review_bos[review_bos.business_id == '#VALUE!'].index)
review_bos = review_bos.drop(review_bos[review_bos.review_id == '#NAME?'].index)
review_bos = review_bos.drop(review_bos[review_bos.review_id == '#VALUE!'].index)
print(len(review_bos))
#review_bos = review_bos.dropna(subset = ['labels'])
from sklearn.cluster import DBSCAN
X = np.array((boston_info.latitude,boston_info.longitude))
X = X.transpose()
#print(X.shape)
clustering = DBSCAN(eps = 0.004, min_samples = 2).fit(X)
print(clustering.labels_)
import matplotlib.pyplot as plt
import matplotlib.colors as colors
import matplotlib.cm as cm
plt.style.use('seaborn-whitegrid')
x_points = X[:, 0]
y_points = X[:, 1]
labels = clustering.labels_
colormap = cm.viridis
color_labels = [colors.rgb2hex(colormap(i)) for i in np.linspace(0, 0.9, max(labels)+1)]
color_list = [color_labels[j] for j in labels]
#print(color_list)
plt.scatter(x_points,y_points,c = color_list, s=5, alpha=0.3)
plt.legend()
plt.show()
labels = clustering.labels_
print(max(labels))
print(review_bos.labels)
#print(review_bos[0:1])
boston_info['labels'] = labels
print(len(boston_info))
print(boston_info.head())
boston_info = boston_info.dropna(subset = ['labels'])
print(len(boston_info))
#每一个用户ID生成一个linkedlist,每个linkedlist就是一串商家的聚类名按照日期排列
class Node:
def __init__(self, dataval = None, date = None):
self.dataval = dataval
self.date = date
#next指向下一个node
self.next = None
class LinkedList:
def __init(self):
self.head = None
def add(self, NewNode):
if self.head is None:
self.head = NewNode
return
last_node = self.head
while(last_node.next):
last_node = last_node.next
last_node.next = NewNode
def insert(self, NewNode):
if self.head is None:
self.head = NewNode
return
pre_node = self.head
while(pre_node.next):
cur_node = pre_node.next
if cur_node.date > NewNode.date:
#insert node
NewNode.next = cur_node
pre_node.next = NewNode
return
pre_node = pre_node.next
pre_node.next = NewNode
def printlist(self):
printval = self.head
print('->',end = "")
while printval is not None:
print(printval.dataval, ' -> ',end = "")
printval = printval.next
print('None')
def get_list(self):
list = []
node = self.head
while node is not None:
list.append(node.dataval)
node = node.next
self.list = list
def get_freq(self):
self.get_list(self)
dict = {}
for value, count in enumerate(self.list):
dict[value] = count
self.dict = dict
import math
users = {}
Labels = np.array([boston_info.business_id, boston_info.labels])
Labels = Labels.transpose()
user_label = []
dict_label = dict(zip(Labels[:,0],Labels[:,1]))
#insert label column in review_bos
count = 0
for i in range(len(review_bos)):
review = review_bos.iloc[i]
#print(review.business_id)
if(review.business_id == "nan"):
user_label.append(0)
count += 1
elif(review.business_id not in dict_label.keys()):
user_label.append(0)
count += 1
else:
#print(dict_label[review.business_id])
user_label.append(dict_label[review.business_id])
print(count)
review_bos.labels = user_label
#create linkedlist
for i in range(len(review_bos)):
review = review_bos.iloc[i]
#print(review['user_id'])
user_id = review['user_id']
#user_id = review.user_id.astype('str').astype('string')
#user_id = review['user_id'].astype(str)
#user_id = str(user_id)
#print(user_id)
if user_id not in users:
#user_id = review.user_id
node = Node(review.labels, review.date)
users[user_id] = LinkedList()
users[user_id].head = node
else:
#user_id = review.user_id
node = Node(review.labels, review.date)
users[user_id].insert(node)
print(len(users))
#print前10个user的linked list
j = 0
for i in users:
if j < 10:
users[i].printlist()
j+=1
#make networkx
import networkx as nx
import pylab
count = 0
for i in users:
if count < 10:
G = nx.DiGraph()
cur_node = users[i].head
if cur_node.next is None:
G.add_node(cur_node.dataval)
else:
add_list = []
weights = []
dict_node = {}
dict_node[cur_node.dataval] = 1
while cur_node.next is not None:
edge = [(cur_node.dataval, cur_node.next.dataval)]
if edge in add_list:
p = add_list.index(edge)
weights[p] += 1
else:
add_list.append(edge)
weights.append(1)
for i in range(len(weights)):
G.add_edges_from(add_list[1][:], weight = weights[i])
pos = nx.spring_layout(G)
nx.draw(G, pos)
pylab.show()
count += 1
else:
break
#test
G = nx.DiGraph()
i = "99RsBrARhhx60UnAC4yDoA"
cur_node = users[i].head
users[i].printlist()
if cur_node.next is None:
G.add_node(cur_node.dataval)
else:
add_list = []
weights = []
dict_node = {}
dict_node[cur_node.dataval] = 1
while cur_node.next is not None:
edge = [(cur_node.dataval, cur_node.next.dataval)]
if edge in add_list:
p = add_list.index(edge)
weights[p] += 1
else:
add_list.append(edge)
weights.append(1)
cur_node = cur_node.next
for i in range(len(weights)):
G.add_edges_from(add_list[1][:], weight = weights[i])
pos = nx.spring_layout(G)
nx.draw(G, pos)
pylab.show()
|
dataset/Part2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Miscellaneous
# Collection of useful functions and tricks to get the most out of brainrender
#
# ### Set up
# +
# We begin by adding the current path to sys.path to make sure that the imports work correctly
import sys
sys.path.append('../')
import os
# Set up VTKPLOTTER to work in Jupyter notebooks
from vtkplotter import *
# Import variables
from brainrender import * # <- these can be changed to personalize the look of your renders
# Import brainrender classes and useful functions
from brainrender.scene import Scene
# -
# ## Camera
#
# To specify the orientation of the camera in a `brainrender` scene, pass the name of the camera when creating an instance of `Scene`.
#
# The default cameras supported in `brainrender` are: `three_quarters` (default), `sagittal`, `coronal` and `top`.
# But you can also pass a dictionary instead of the camera name, just make sure that the dictionary has the correct camera parameters (check `brainrender.camera.py`).
# +
scene = Scene(jupyter=True, camera='sagittal')
scene.add_brain_regions(['TH'], use_default_colors=True)
# ... render
# -
# ### Edit actors
# You can change the look of actors very easily. Check `brainrender.Utils.actors_funcs.py` for more options.
# +
scene = Scene(jupyter=True)
scene.add_brain_regions(['MOs', 'MOp'], colors='red')
scene.edit_actors([scene.actors['regions']['MOs']], wireframe=True)
# ... render
# -
# ### Useful functions
#
# `Scene.get_n_random_points_in_region` let's you generate N random points in a brain region. It can be usefull when working with tractography data. For large brain regions projections to different locations within the region might differ, this function lets you sample the target region more copmletely.
#
# `Scene.edit_neurons` Can be used to edit neuron actors (e.g. mirror them across hemispheres). To get the neurons actors just use `Scene.actors['neurons']`.
|
Examples/notebooks/Misc.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + jupyter={"source_hidden": true} slideshow={"slide_type": "skip"} tags=[]
import numpy as np
import matplotlib.pyplot as plt
from IPython.display import display, HTML, IFrame
from ipywidgets import interact,fixed,FloatSlider
# import pandas as pd
from mpl_toolkits import mplot3d
from mpl_toolkits.mplot3d import axes3d
from numpy.linalg import norm
from numpy import cos,sin,tan,arctan,exp,log,pi,sqrt,array,arange,linspace,meshgrid
from ipykernel.pylab.backend_inline import flush_figures
from scipy.integrate import quad, dblquad, tplquad
# %matplotlib inline
plt.rcParams.update({
"figure.figsize": (6,6),
"text.usetex": True,
"font.family": "serif",
})
# Uncomment the one that corresponds to your Jupyter theme
plt.style.use('default')
# plt.style.use('dark_background')
# plt.style.use('fivethirtyeight')
# plt.style.use('Solarize_Light2')
# + [markdown] slideshow={"slide_type": "notes"}
# $\newcommand{\RR}{\mathbb{R}}$
# $\newcommand{\bv}[1]{\begin{bmatrix} #1 \end{bmatrix}}$
# $\renewcommand{\vec}{\mathbf}$
#
# + [markdown] slideshow={"slide_type": "fragment"}
# ### Example
#
# Consider the vector field $\vec F(x,y,z) = x\,\vec i$. Consider the surfaces
#
# - $\Omega_1$: piece of the plane $x=0$ for $0 \leq y,z \leq 1$
# - $\Omega_2$: piece of the surface $x= 3y(1-y)$ for $0 \leq y,z \leq 1$
#
# oriented in the positive $x$-direction. Through which surface is the flux of $\vec F$ greater?
# + hide_input=true jupyter={"source_hidden": true} slideshow={"slide_type": "fragment"} tags=[]
@interact
def _(angle = (-96,6,6),caps=False):
fig = plt.figure(figsize=(8,8))
ax = fig.add_subplot(projection='3d')
u = np.linspace(0,1,20)
v = np.linspace(0,1,10)
U,V = np.meshgrid(u,v)
ax.plot_surface(0*U*(1-U),U,V,alpha=.5)
ax.plot_wireframe(0*U*(1-U),U,V,alpha=1,color='k',rcount=10,ccount=10)
ax.plot_surface(3*U*(1-U),U,V,alpha=.5)
ax.plot_wireframe(3*U*(1-U),U,V,alpha=1,color='k',rcount=10,ccount=10)
if caps:
ax.plot_surface(3*V*U*(1-U),U,np.ones_like(V),alpha=.5)
ax.plot_wireframe(3*V*U*(1-U),U,np.ones_like(V),alpha=1,color='k',rcount=10,ccount=10)
ax.plot_surface(3*V*U*(1-U),U,np.zeros_like(V),alpha=.5)
ax.plot_wireframe(3*V*U*(1-U),U,np.zeros_like(V),alpha=1,color='k',rcount=10,ccount=10)
ax.view_init(30,angle)
u = v = np.linspace(0,1,7)
# v = np.linspace(0,2*pi,10)
U,V,W = np.meshgrid(u,v,u)
ax.quiver(U,V,W,U,0*U,0*U,length=.2);
flush_figures();
# + [markdown] jp-MarkdownHeadingCollapsed=true tags=[]
# #### Solution
# + [markdown] slideshow={"slide_type": "fragment"}
# Let $\Omega_1$ and $\Omega_2$ be as before (with a negative sign indicating an opposite orientation) and $T_1$ and $T_0$ be the enclosing, upward-oriented "caps" to the solid at $z=1$ and $z=0$, respectively.
#
# $$ \iint_{\Omega_2} \vec F\cdot d\vec S + \iint_{-\Omega_1} \vec F\cdot d\vec S + \iint_{T_1} \vec F\cdot d\vec S + \iint_{-T_0} \vec F\cdot d\vec S = \iiint_E \nabla\cdot \vec F\,dV $$
# + [markdown] slideshow={"slide_type": "fragment"}
# $$ \iint_{\Omega_2} \vec F\cdot d\vec S - \iint_{\Omega_1} \vec F\cdot d\vec S = \iiint_E dV $$
#
# $$ \iint_{\Omega_2} \vec F\cdot d\vec S = \int_0^1 \int_0^1 3y(1-y)\,dy\,dz = 1/2.$$
# -
# <p style="padding-bottom:40%;"> </p>
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Revisit Cone
#
# Find the flux of the curl $\nabla\times \vec F$ of the vector field $2y\,\vec i + 3z\,\vec j +x\,\vec k$ through the piece of the cone $z=\sqrt{x^2 + y^2}$ below $z=2$, oriented upward.
#
#
# + hide_input=false jupyter={"source_hidden": true} slideshow={"slide_type": "fragment"} tags=[]
@interact(angle=(-6,96,6))
def _(angle = 24,vangle=(0,90,6),fade=(0,1,.04)):
fig = plt.figure(figsize=(8,8))
ax = fig.add_subplot(projection='3d')
u = np.linspace(0,2,100)
v = np.linspace(0,2*pi,100)
U,V = np.meshgrid(u,v)
ax.view_init(vangle,angle)
ax.plot_surface(U*cos(V),U*sin(V),U,alpha=.5)
ax.plot_wireframe(U*cos(V),U*sin(V),U,alpha=1,color='k',rcount=10,ccount=10)
# ax.plot_surface(U*cos(V),U*sin(V),2*np.ones_like(U),alpha=.5)
# ax.plot_wireframe(U*cos(V),U*sin(V),2*np.ones_like(U),alpha=1,color='k',rcount=10,ccount=10)
u = v = np.linspace(-2,2,8)
# v = np.linspace(0,2*pi,10)
U,V,W = np.meshgrid(u,v,u)
ax.quiver(U,(V),W,2*V,3*W,U,length=.1,alpha=1-fade);
ax.quiver(U,(V),W,0*U - 3,0*W - 1,0*U - 2,length=.2,lw=2,color='r',alpha=fade);
flush_figures();
# + [markdown] jp-MarkdownHeadingCollapsed=true tags=[]
# #### Solution
# -
# We compute $$\nabla\times \vec F = \begin{vmatrix} \vec i & \vec j & \vec k \\
# \partial_x & \partial_y & \partial_z \\
# 2y & 3z & x \\
# \end{vmatrix} = -3\,\vec i - \vec j -2 \vec k$$
# <p style="padding-bottom:40%;"> </p>
# ## Exercise
#
# Compute the line integral $\oint_{\partial D} \vec F \cdot d\vec r$ around the ccw (from above) boundary of the surface $D$ given.
#
# $$ \vec F(x,y,z) = xyz \,\vec i + y \,\vec j + z \,\vec k$$
#
# where $D$ is the portion of the surface $z=x^2$ above $x^2 + y^2 \leq a^2$ in the first octant.
# + hide_input=false jupyter={"source_hidden": true} slideshow={"slide_type": "fragment"} tags=[]
@interact(angle=(-96,6,6))
def _(angle = -24,vangle=(0,90,6)):
fig = plt.figure(figsize=(8,8))
ax = fig.add_subplot(projection='3d')
u = np.linspace(0,2,40)
v = np.linspace(0,pi/2,40)
U,V = np.meshgrid(u,v)
ax.view_init(vangle,angle)
ax.plot_surface(U*cos(V),U*sin(V),U**2*cos(V)**2,alpha=.5)
ax.plot_wireframe(U*cos(V),U*sin(V),U**2*cos(V)**2,alpha=1,color='k',rcount=10,ccount=10)
# ax.plot_surface(U*cos(V),U*sin(V),2*np.ones_like(U),alpha=.5)
# ax.plot_wireframe(U*cos(V),U*sin(V),2*np.ones_like(U),alpha=1,color='k',rcount=10,ccount=10)
u = v = np.linspace(0,2,8)
# v = np.linspace(0,2*pi,10)
U,V,W = np.meshgrid(u,v,2*u)
ax.quiver(U,V,W,U*W*V,V,W,length=.05);
# ax.quiver(U,V,W,0*U - 3,0*W - 1,0*U - 2,length=.2,lw=2,color='r');
flush_figures();
# + [markdown] jp-MarkdownHeadingCollapsed=true slideshow={"slide_type": "fragment"} tags=[]
# #### Solution
# + [markdown] slideshow={"slide_type": "fragment"}
# Compute the curl.
#
# $$\nabla \times \vec F = xy\, \vec j - xz \,\vec k $$
#
# Parametrize the surface.
#
# $$ \vec r(u,v) = u \cos v \,\vec i + u \sin v \,\vec j + u^2 \cos^2 v \,\vec k$$
#
# for $0 \leq u \leq a$ and $0 \leq v \leq \pi/2$.
#
# $$ \vec r_u \times \vec r_v = \begin{vmatrix} \vec i & \vec j & \vec k \\
# \cos v & \sin v & 2u \cos^2 v \\
# -u\sin v & u \cos v & -2u^2 \sin v\cos v \\
# \end{vmatrix} = - 2u^2 \cos v \vec i + u \vec k$$
#
# which is consistent with an upward orientation.
#
# Finally, we apply Stokes'.
#
# $$\oint_{\partial \Omega} \vec F\cdot d\vec r = \iint_\Omega \nabla \times \vec F \cdot d\vec S $$
#
# Since the vectors only share a $\vec k$ component, the integrand simplifies.
#
# $$ = \int_0^{\pi/2} \int_0^a -u^4\cos ^3 v \, du\, dv =-\frac{2a^5}{15}$$
# -
# **Challenge** Parametrize the boundary of this surface and compute the line integral directly to verify this number.
# A little analysis makes this easier. $y\,\vec j + z\,\vec k$ is a conservative vector field, so its integral arround a closed loop will be 0. $xyz\,\vec i$ is $\vec 0$ on the coordinate planes, so we need only evaluate the curved edge.
#
# $$\vec r(t) = a \cos t\, \vec i + a \sin t\, \vec j + a^2 \cos^2 t\,\vec k$$
#
# with $0 \leq t \leq \pi/2$.
#
# $$\oint_{\partial \Omega} \vec F\cdot d\vec r = \int_0^{\pi/2} a^5 \cos^3 t \sin t (-\sin t)\,dt $$
# with $w = \sin t, dw = \cos t\,dt$, we get $$-\int_0^1 a^5 (1 - w^2)w^2\,dw -a^5(\frac 13 - \frac15) = - \frac{2a^5}{15}$$
|
exercises/L23-Exercises-Solutions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.5.0
# language: julia
# name: julia-0.5
# ---
# # Sums of squares certicates
# We examime whether $f(x,z) = 2 x^4 + 2 x^3 z - x^2 z^2 + 5 z^4$ can be written as a sum of squares.
using Polyopt
x, z = variables(["x", "z"]);
f = 2*x^4 + 2*x^3*z - x^2*z^2 + 5*z^4;
# The degree of $f$ is 4, so we need a second order relaxation,
prob = momentprob(2, f);
# We solve the problem,
X, Z, t, y, solsta = solve_mosek(prob);
# Since $t$ is zero, we have a sum-of-squares certificate
t
X[1]
# To verify the sum-of-squares certificate, we compute a monomial vector
v = monomials(2, [x, z])
u = chol(X[1])*v
# or discarding monomial terms with small coeffients,
u = [truncate(ui, 1e-8) for ui=u ]
# and the sum-of-squares term can be evaluated to
truncate(f - dot(u,u), 1e-8)
|
notebooks/SOS1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import pandas as pd
import gnucash_pandas
import numpy as np
def plot_df(df, **kwargs):
return df.loc[df.index > '2015-01-01'].plot(figsize=(16,8), grid=True, **kwargs)
# -
gnucash_file = input("Path to GnuCash file to load ")
df = gnucash_pandas.splits_dataframe(gnucash_file)
expenses = gnucash_pandas.daily(df, 'Expense')
expenses
monthly_expenses = expenses.rolling(31).sum()
monthly_expenses
# # Monthly Expenses
#
# This plot displays rolling 32-day expenses by category.
plt = plot_df(monthly_expenses)
plt.legend(loc='lower center', ncol=4, bbox_to_anchor=(0.5,-0.6))
plt
# # Total Monthly Expenses
total_monthly_expenses = monthly_expenses.sum(axis=1) # axis=1 means sum columns
plot_df(total_monthly_expenses)
income = -gnucash_pandas.daily(df, 'Income')
income
monthly_income = income.rolling(31).sum()
monthly_income
# # Monthly Income
plt = plot_df(monthly_income)
plt.legend(loc='lower center', ncol=4, bbox_to_anchor=(0.5,-0.3))
plt
# # Total Monthly Income
total_monthly_income = monthly_income.sum(axis=1) # axis=1 means sum columns
plot_df(total_monthly_income)
# # Percentage of income spent
#
# This is the proportion of monthly income spent on monthly expenses.
pct_expenses = total_monthly_expenses/total_monthly_income
plot_df(pct_expenses, ylim=(0,2), yticks=np.linspace(0, 2, num=9))
taxes = [
"CA Private Disability Employee",
"CA State Income Tax",
"Employee Medicare",
"Federal Income Tax",
"Social Security Employee Tax",
"Taxes",
]
tax_free_expenses = monthly_expenses.drop(taxes, axis=1)
tax_free_expenses
# # Monthly after-tax expenses
plt = plot_df(tax_free_expenses)
plt.legend(loc='lower center', ncol=4, bbox_to_anchor=(0.5,-0.55))
plt
# # Total after-tax expenses
total_tax_free_expenses = tax_free_expenses.sum(axis=1) # axis=1 means sum columns
plot_df(total_tax_free_expenses)
# # Total after-tax income
total_tax_expenses = total_monthly_expenses - total_tax_free_expenses
total_after_tax_income = total_monthly_income - total_tax_expenses
plot_df(total_after_tax_income)
# # Percentage of after-tax income spent
after_tax_pct_expenses = total_tax_free_expenses/total_after_tax_income
plot_df(after_tax_pct_expenses, ylim=(0,2), yticks=np.linspace(0, 2, num=9))
# # Food Expenses
food_expenses = monthly_expenses[["Dining", "Groceries"]].copy()
food_expenses["Total"] = food_expenses.sum(axis=1)
plot_df(food_expenses)
|
Monthly expenses.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Decoupling Simulation Accuracy from Mesh Quality
# Simple python notebook to regenrate the teaser image.
#
# The data and scripts to regenerate the other figures can be found
# [here](https://github.com/polyfem/Decoupling-Simulation-Accuracy-from-Mesh-Quality).
#
# Note, we used [polyfem](https://polyfem.github.io) [C++ version](https://github.com/polyfem/polyfem/) for all figures, with direct solver Pardiso, this notebook uses the default iterative algebraic multigrid Hypre.
# Note that this can be run direcly with binder!
# [](https://mybinder.org/v2/gh/polyfem/Decoupling-Simulation-Accuracy-from-Mesh-Quality.git/master?filepath=Decoupling-Simulation-Accuracy-from-Mesh-Quality.ipynb)
#
# Importing libraries
# +
import numpy as np
import polyfempy as pf
import json
#plotting:
import meshplot as mp
import plotly.offline as plotly
import plotly.graph_objs as go
#Necessary for the notebook
plotly.init_notebook_mode(connected=True)
# -
# Note that this notebook is designed to be run in the root folder.
# If you want to reproduce the results of the teaser you can find the images [here](https://github.com/polyfem/Decoupling-Simulation-Accuracy-from-Mesh-Quality/tree/master/fig1-teaser/meshes).
# Setup problem, the exact definition of the function can be found [here](https://github.com/polyfem/polyfem/blob/d598e1764512da28d62ecbbbd1a832407ffb42de/src/problem/MiscProblem.cpp#L23)
# +
settings = pf.Settings()
#Teaser uses laplacian
settings.set_pde(pf.PDEs.Laplacian)
problem = pf.Problem()
#Use quartic function f = (2*y-0.9)^4+0.1 for bounary conditions
#f''=\Deta f = 48*(2*y-.9)^2 for rhs
problem.add_dirichlet_value(1, "(2*y-0.9)^4+0.1")
problem.exact = "(2*y-0.9)^4+0.1"
problem.exact_grad = ["0", "8*(2*y-.9)^3"]
problem.rhs = "48*(2*y-.9)^2"
settings.problem = problem
# -
# First we solve using standard $P_1$ elements and our method which we enable with `settings.set_advanced_option("use_p_ref", True)`.
#
# **Note** this will take some time, the last mesh quality is really low!
# +
n_meshes = 8
total_solutions = {}
total_errors = {}
for use_pref in [False, True]:
settings.set_advanced_option("use_p_ref", use_pref)
solver = pf.Solver()
solver.set_log_level(6)
solver.settings(settings)
solutions = []
errors = []
#iterating over input files, named from 0 to 8
for i in range(n_meshes+1):
solver.load_mesh_from_path("fig1-teaser/meshes/large_angle_strip_{}.obj".format(i), normalize_mesh=True)
#we dont distinguis bc contitions, all sidesets to 1
solver.set_boundary_side_set_from_bary(lambda v: 1)
solver.solve()
solver.compute_errors()
#getting and storing solution
v, f, sol = solver.get_sampled_solution()
pts = np.append(v, sol, 1)
solutions.append({"pts": pts, "f": f})
#getting error:
log = solver.get_log()
log = json.loads(log)
errors.append(log['err_l2'])
#store these list in the total one
total_solutions["ours" if use_pref else "p1"] = solutions
total_errors["ours" if use_pref else "p1"] = errors
print("Done!")
# -
# **Plz wait until done!**
# ## Plot 3D results
# For nice plot we append the 2 meshses, ours on the left
def get_v_f_c(sols, i):
#append points and offset x by 1.1
pts_ours = np.array(total_solutions["ours"][i]["pts"])
pts_p1 = np.array(total_solutions["p1"][i]["pts"])
pts_p1[:, 0] += 1.1
v = np.append(pts_ours, pts_p1, 0)
#colors
c = np.array(v[:, 2])
#append faces
f_ours = np.array(total_solutions["ours"][i]['f'])
f_p1 = np.array(total_solutions["p1"][i]['f'])
f_p1 += np.max(f_ours)+1
f = np.append(f_ours, f_p1, 0)
return v, f, c
# Our results are on the left!
#
# You can rotate the view with the mouse!
# +
v, f, c = get_v_f_c(total_solutions, 0)
p = mp.plot(v, f, c, return_plot=True)
@mp.interact(mesh=[('mesh_{}'.format(i), i) for i in range(n_meshes+1)])
def ff(mesh):
v, f, c = get_v_f_c(total_solutions, mesh)
mp.plot(v, f, c, plot=p)
p
# -
# ## Error plor
# +
colors = {'ours': 'rgb(85, 239, 196)', 'p1': 'rgb(255, 201, 26)'}
marker_shape = 'circle'
marker_size = 6
def get_plot_trace(errors, method):
x = []
y = []
for i in range(len(errors[method])):
x.append(i)
y.append(errors[method][i])
x, y = zip(*sorted(zip(x, y))[:])
trace = go.Scatter(x=x, y=y,
mode='lines+markers',
name=method,
line=dict(color=colors[method]),
marker=dict(symbol=marker_shape, size=marker_size)
)
return trace
# +
plot_data = [get_plot_trace(total_errors, "ours"), get_plot_trace(total_errors, "p1")]
layout = go.Layout(
legend=dict(x=0.1, y=0.98),
xaxis=dict(
title="Mesh",
nticks=5,
),
yaxis=dict(
title="Error L2",
range=[0.0, 0.3]
),
hovermode='closest')
fig = go.Figure(data=plot_data, layout=layout)
plotly.iplot(fig)
# -
|
Decoupling-Simulation-Accuracy-from-Mesh-Quality.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R 3.6
# language: R
# name: ir36
# ---
# # Descriptive Statistics
#
# `R` has plenty of functions to describe data quantitatively and visually. Before we continue, let's set the seed since we are sampling.
set.seed(37)
# ## Summarization
# ### Summarization for data structures
#
# The `summary` function may be used for `vectors`, `factors`, `matrices` and `data frames`.
# +
x <- rnorm(1000, mean=10, sd=2)
print(summary(x))
# +
x <- factor(sample(c('water', 'soda', 'tea', 'coffee'), 1000, replace=TRUE))
print(summary(x))
# +
A <- matrix(rnorm(1000), ncol=2)
print(summary(A))
# +
df <- data.frame(
V1 <- rnorm(500),
V2 <- rnorm(500)
)
print(summary(df))
# -
# ### Scalar value summarization
#
# The following functions may be used to produce a scalar value summarization for `vectors`.
#
# * `min` returns the smallest number
# * `max` returns the largest number
# * `length` returns the length of the vector (number of elements)
# * `mean` returns the average of the elements in the vector
# * `sd` returns the standard deviation of the elements in the vector
# * `var` returns the variance of the elements in the vector
# * `mad` returns the mean absolute deviation of the elements in the vector
# +
library(purrr)
x <- rnorm(1000, mean=10, sd=2)
xMin <- min(x)
xMax <- max(x)
xLen <- length(x)
xSum <- sum(x)
xMean <- mean(x)
xMed <- median(x)
xSd <- sd(x)
xVar <- var(x)
xMad <- mad(x)
v1 <- c('min', 'max', 'length', 'sum', 'mean', 'median', 'standard deviation', 'variance', 'mean absolute deviation')
v2 <- c(xMin, xMax, xLen, xSum, xMean, xMed, xSd, xVar, xMax)
for (item in map2(v1, v2, function(x, y) paste(x, y))) {
print(item)
}
# -
# Some of these functions may also be applied to data frames.
# +
df <- data.frame(
V1 <- rnorm(500),
V2 <- rnorm(500)
)
xMin <- min(df)
xMax <- max(df)
xLen <- length(df)
xSum <- sum(df)
v1 <- c('min', 'max', 'length', 'sum')
v2 <- c(xMin, xMax, xLen, xSum)
for (item in map2(v1, v2, function(x, y) paste(x, y))) {
print(item)
}
# -
# ### Summarization with multiple results
#
# The `quantile` and `fivenum` functions will return summary with multiple results.
# +
x <- rnorm(100, mean=15, sd=3)
xQuant <- quantile(x)
print(xQuant)
# -
xFive <- fivenum(x)
print(xFive)
# ### Row and column summarization
#
# There are row and column summarization functions for matrices and data frames.
# +
df <- data.frame(
V1 <- c(1, 2, 3, 4, 5),
V2 <- c(6, 7, 8, 9, 10)
)
print(rowMeans(df))
print(rowSums(df))
print(colMeans(df))
print(colSums(df))
# -
# You may also use the `apply` method to produce summaries. The second parameter is either `1` or `2` for rows or columns, respectively.
print(apply(df, 1, sd))
print(apply(df, 2, sd))
# ## Cummulative
#
# You may apply the following functions to find the cummulative value
#
# * `cumsum` computes the cummulative sum
# * `cummax` computes the cummulative max
# * `cummin` computes the commulative minimum
# * `cummax` computes the commulative maximum
# +
x <- seq(1, 5)
print(cumsum(x))
# +
x <- seq(1, 5)
print(cummax(x))
# +
x <- seq(1, 5)
print(cummin(x))
# +
x <- seq(1, 5)
print(cumprod(x))
# -
# ## Tables
#
# The `table` command builds a contingency table. When `table` is used on a vector of numbers, a sort frequency distribution appears.
x <- c(5, 4, 4, 3, 3, 3, 2, 2, 2, 2, 1, 1, 1, 1, 1)
y <- table(x)
print(y)
# The `table` function may also be used on a vector of characters.
x <- c('hi', 'hi', 'hi', 'bye', 'bye')
y <- table(x)
print(y)
# Here's the `table` command used on a matrix.
s <- sample(seq(1:10), 100, replace=TRUE)
m <- matrix(s, ncol=5)
print(m)
y <- table(m)
print(y)
# You may subset a matrix and then apply `table`.
y <- table(m[,1], m[,2], dnn=c('V1', 'V2'))
print(y)
# The following shows using `table` on a table with numeric and character data.
# +
df <- data.frame(
x <- c(7, 7, 7, 7, 7, 8, 8, 8, 8),
y <- c('w', 'w', 'w', 'l', 'l', 'l', 'l', 'l', 'w')
)
y <- table(df, dnn=c('x', 'y'))
print(y)
# -
# The following shows using `table` on a table with numeric data.
# +
df <- data.frame(
x <- c(7, 7, 7, 7, 8, 8, 8, 8),
y <- c(1, 1, 2, 3, 1, 2, 3, 4)
)
y <- table(df, dnn=c('x', 'y'))
print(y)
# -
# Use the `with` command to avoid using `dnn` with the `table` function.
with(df, table(x, y))
# ## Stem
x <- sample(seq(1:10), 100, replace=TRUE)
s <- stem(x)
print(s)
# Compare the `stem` to the `table` function.
t <- table(x)
print(t)
# You may control the `scaling` of the `stem` function.
# ## Histogram
# +
x <- sample(seq(1:10), 100, replace=TRUE)
options(repr.plot.width=4, repr.plot.height=4)
hist(x, col='gray75')
# -
# ## Density
x <- sample(seq(1:10), 100, replace=TRUE)
d <- density(x, bw='nrd0', kernel='gaussian', na.rm=FALSE)
print(d)
# +
options(repr.plot.width=4, repr.plot.height=4)
plot(d, main='Density')
# +
options(repr.plot.width=4, repr.plot.height=4)
hist(x, freq=F, col='gray85')
lines(density(x), lty=2)
lines(density(x, k='rectangular'))
|
sphinx/r-intro/source/descriptive.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Vincent-Emma/DS-Unit-1-Sprint-2-Data-Wrangling-and-Storytelling/blob/master/Vince_Emma_LS_DS_123_Make_Explanatory_Visualizations.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="-8-trVo__vRE"
# _Lambda School Data Science_
#
# # Make Explanatory Visualizations
#
# ### Objectives
#
# - identify misleading visualizations and how to fix them
# - use Seaborn to visualize distributions and relationships with continuous and discrete variables
# - add emphasis and annotations to transform visualizations from exploratory to explanatory
# - remove clutter from visualizations
#
# ### Links
#
# - [How to Spot Visualization Lies](https://flowingdata.com/2017/02/09/how-to-spot-visualization-lies/)
# - [Visual Vocabulary - Vega Edition](http://ft.com/vocabulary)
# - [Choosing a Python Visualization Tool flowchart](http://pbpython.com/python-vis-flowchart.html)
# - [Searborn example gallery](http://seaborn.pydata.org/examples/index.html) & [tutorial](http://seaborn.pydata.org/tutorial.html)
# - [Strong Titles Are The Biggest Bang for Your Buck](http://stephanieevergreen.com/strong-titles/)
# - [Remove to improve (the data-ink ratio)](https://www.darkhorseanalytics.com/blog/data-looks-better-naked)
# - [How to Generate FiveThirtyEight Graphs in Python](https://www.dataquest.io/blog/making-538-plots/)
# + [markdown] id="s-24T844-8qv" colab_type="text"
# # Avoid Misleading Visualizations
#
# Did you find/discuss any interesting misleading visualizations in your Walkie Talkie?
# + [markdown] id="Qzxt9ntsNjs0" colab_type="text"
# ## What makes a visualization misleading?
#
# [5 Ways Writers Use Misleading Graphs To Manipulate You](https://venngage.com/blog/misleading-graphs/)
# + [markdown] id="q7_DUiENNvxk" colab_type="text"
# ## Two y-axes
#
# <img src="https://kieranhealy.org/files/misc/two-y-by-four-sm.jpg" width="800">
#
# Other Examples:
# - [Spurious Correlations](https://tylervigen.com/spurious-correlations)
# - <https://blog.datawrapper.de/dualaxis/>
# - <https://kieranhealy.org/blog/archives/2016/01/16/two-y-axes/>
# - <http://www.storytellingwithdata.com/blog/2016/2/1/be-gone-dual-y-axis>
# + [markdown] id="oIijNBDMNv2k" colab_type="text"
# ## Y-axis doesn't start at zero.
#
# <img src="https://i.pinimg.com/originals/22/53/a9/2253a944f54bb61f1983bc076ff33cdd.jpg" width="600">
# + [markdown] id="ISB2p8vZNv6r" colab_type="text"
# ## Pie Charts are bad
#
# <img src="https://i1.wp.com/flowingdata.com/wp-content/uploads/2009/11/Fox-News-pie-chart.png?fit=620%2C465&ssl=1" width="600">
# + [markdown] id="67CsAzu1NwBJ" colab_type="text"
# ## Pie charts that omit data are extra bad
#
# - A guy makes a misleading chart that goes viral
#
# What does this chart imply at first glance? You don't want your user to have to do a lot of work in order to be able to interpret you graph correctly. You want that first-glance conclusions to be the correct ones.
#
# <img src="https://pbs.twimg.com/media/DiaiTLHWsAYAEEX?format=jpg&name=medium" width='600'>
#
# <https://twitter.com/michaelbatnick/status/1019680856837849090?lang=en>
#
# - It gets picked up by overworked journalists (assuming incompetency before malice)
#
# <https://www.marketwatch.com/story/this-1-chart-puts-mega-techs-trillions-of-market-value-into-eye-popping-perspective-2018-07-18>
#
# - Even after the chart's implications have been refuted, it's hard a bad (although compelling) visualization from being passed around.
#
# <https://www.linkedin.com/pulse/good-bad-pie-charts-karthik-shashidhar/>
#
# **["yea I understand a pie chart was probably not the best choice to present this data."](https://twitter.com/michaelbatnick/status/1037036440494985216)**
# + [markdown] id="FYXmlToEOOTC" colab_type="text"
# ## Pie Charts that compare unrelated things are next-level extra bad
#
# <img src="http://www.painting-with-numbers.com/download/document/186/170403+Legalizing+Marijuana+Graph.jpg" width="600">
#
# + [markdown] id="IwtMQpY_QFUw" colab_type="text"
# ## Be careful about how you use volume to represent quantities:
#
# radius vs diameter vs volume
#
# <img src="https://static1.squarespace.com/static/5bfc8dbab40b9d7dd9054f41/t/5c32d86e0ebbe80a25873249/1546836082961/5474039-25383714-thumbnail.jpg?format=1500w" width="600">
# + [markdown] id="tTuAWjSBRsc7" colab_type="text"
# ## Don't cherrypick timelines or specific subsets of your data:
#
# <img src="https://wattsupwiththat.com/wp-content/uploads/2019/02/Figure-1-1.png" width="600">
#
# Look how specifically the writer has selected what years to show in the legend on the right side.
#
# <https://wattsupwiththat.com/2019/02/24/strong-arctic-sea-ice-growth-this-year/>
#
# Try the tool that was used to make the graphic for yourself
#
# <http://nsidc.org/arcticseaicenews/charctic-interactive-sea-ice-graph/>
#
# + [markdown] id="Xs13S7p4Srme" colab_type="text"
# ## Use Relative units rather than Absolute Units
#
# <img src="https://imgs.xkcd.com/comics/heatmap_2x.png" width="600">
# + [markdown] id="CIMt5OiuTlrr" colab_type="text"
# ## Avoid 3D graphs unless having the extra dimension is effective
#
# Usually you can Split 3D graphs into multiple 2D graphs
#
# 3D graphs that are interactive can be very cool. (See Plotly and Bokeh)
#
# <img src="https://thumbor.forbes.com/thumbor/1280x868/https%3A%2F%2Fblogs-images.forbes.com%2Fthumbnails%2Fblog_1855%2Fpt_1855_811_o.jpg%3Ft%3D1339592470" width="600">
# + [markdown] id="GATMu9IqUlIj" colab_type="text"
# ## Don't go against typical conventions
#
# <img src="http://www.callingbullshit.org/twittercards/tools_misleading_axes.png" width="600">
# + [markdown] id="g6bKgZ0m_ynS" colab_type="text"
# # Tips for choosing an appropriate visualization:
# + [markdown] id="WtBsVnO4VHiJ" colab_type="text"
# ## Use Appropriate "Visual Vocabulary"
#
# [Visual Vocabulary - Vega Edition](http://ft.com/vocabulary)
# + [markdown] id="H_QM9FHqVT7T" colab_type="text"
# ## What are the properties of your data?
# - Is your primary variable of interest continuous or discrete?
# - Is in wide or long (tidy) format?
# - Does your visualization involve multiple variables?
# - How many dimensions do you need to include on your plot?
#
# Can you express the main idea of your visualization in a single sentence?
#
# How hard does your visualization make the user work in order to draw the intended conclusion?
# + [markdown] id="5EqXxnJeB89_" colab_type="text"
# ## Which Visualization tool is most appropriate?
#
# [Choosing a Python Visualization Tool flowchart](http://pbpython.com/python-vis-flowchart.html)
# + [markdown] id="4mDuzLeNn23m" colab_type="text"
# ## Anatomy of a Matplotlib Plot
# + id="h-aIS1Vdn2RR" colab_type="code" outputId="7acbb0e3-2133-44e9-b684-8d0b5f9a4411" colab={"base_uri": "https://localhost:8080/", "height": 555}
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import AutoMinorLocator, MultipleLocator, FuncFormatter
np.random.seed(19680801)
X = np.linspace(0.5, 3.5, 100)
Y1 = 3+np.cos(X)
Y2 = 1+np.cos(1+X/0.75)/2
Y3 = np.random.uniform(Y1, Y2, len(X))
fig = plt.figure(figsize=(8, 8))
ax = fig.add_subplot(1, 1, 1, aspect=1)
def minor_tick(x, pos):
if not x % 1.0:
return ""
return "%.2f" % x
ax.xaxis.set_major_locator(MultipleLocator(1.000))
ax.xaxis.set_minor_locator(AutoMinorLocator(4))
ax.yaxis.set_major_locator(MultipleLocator(1.000))
ax.yaxis.set_minor_locator(AutoMinorLocator(4))
ax.xaxis.set_minor_formatter(FuncFormatter(minor_tick))
ax.set_xlim(0, 4)
ax.set_ylim(0, 4)
ax.tick_params(which='major', width=1.0)
ax.tick_params(which='major', length=10)
ax.tick_params(which='minor', width=1.0, labelsize=10)
ax.tick_params(which='minor', length=5, labelsize=10, labelcolor='0.25')
ax.grid(linestyle="--", linewidth=0.5, color='.25', zorder=-10)
ax.plot(X, Y1, c=(0.25, 0.25, 1.00), lw=2, label="Blue signal", zorder=10)
ax.plot(X, Y2, c=(1.00, 0.25, 0.25), lw=2, label="Red signal")
ax.plot(X, Y3, linewidth=0,
marker='o', markerfacecolor='w', markeredgecolor='k')
ax.set_title("Anatomy of a figure", fontsize=20, verticalalignment='bottom')
ax.set_xlabel("X axis label")
ax.set_ylabel("Y axis label")
ax.legend()
def circle(x, y, radius=0.15):
from matplotlib.patches import Circle
from matplotlib.patheffects import withStroke
circle = Circle((x, y), radius, clip_on=False, zorder=10, linewidth=1,
edgecolor='black', facecolor=(0, 0, 0, .0125),
path_effects=[withStroke(linewidth=5, foreground='w')])
ax.add_artist(circle)
def text(x, y, text):
ax.text(x, y, text, backgroundcolor="white",
ha='center', va='top', weight='bold', color='blue')
# Minor tick
circle(0.50, -0.10)
text(0.50, -0.32, "Minor tick label")
# Major tick
circle(-0.03, 4.00)
text(0.03, 3.80, "Major tick")
# Minor tick
circle(0.00, 3.50)
text(0.00, 3.30, "Minor tick")
# Major tick label
circle(-0.15, 3.00)
text(-0.15, 2.80, "Major tick label")
# X Label
circle(1.80, -0.27)
text(1.80, -0.45, "X axis label")
# Y Label
circle(-0.27, 1.80)
text(-0.27, 1.6, "Y axis label")
# Title
circle(1.60, 4.13)
text(1.60, 3.93, "Title")
# Blue plot
circle(1.75, 2.80)
text(1.75, 2.60, "Line\n(line plot)")
# Red plot
circle(1.20, 0.60)
text(1.20, 0.40, "Line\n(line plot)")
# Scatter plot
circle(3.20, 1.75)
text(3.20, 1.55, "Markers\n(scatter plot)")
# Grid
circle(3.00, 3.00)
text(3.00, 2.80, "Grid")
# Legend
circle(3.70, 3.80)
text(3.70, 3.60, "Legend")
# Axes
circle(0.5, 0.5)
text(0.5, 0.3, "Axes")
# Figure
circle(-0.3, 0.65)
text(-0.3, 0.45, "Figure")
color = 'blue'
ax.annotate('Spines', xy=(4.0, 0.35), xytext=(3.3, 0.5),
weight='bold', color=color,
arrowprops=dict(arrowstyle='->',
connectionstyle="arc3",
color=color))
ax.annotate('', xy=(3.15, 0.0), xytext=(3.45, 0.45),
weight='bold', color=color,
arrowprops=dict(arrowstyle='->',
connectionstyle="arc3",
color=color))
ax.text(4.0, -0.4, "Made with http://matplotlib.org",
fontsize=10, ha="right", color='.5')
plt.show()
# + [markdown] id="ORUwQD6F-VYg" colab_type="text"
# Today we will reproduce this [example by FiveThirtyEight:](https://fivethirtyeight.com/features/al-gores-new-movie-exposes-the-big-flaw-in-online-movie-ratings/)
#
#
# + id="MxehIOC4yUh_" colab_type="code" outputId="071204db-e6bf-47a3-cf53-ba0c43c24bb5" colab={"base_uri": "https://localhost:8080/", "height": 493}
##simple web scraper
from requests import get
url = 'https://www.imdb.com/title/tt6105098/ratings?ref_=tt_ov_rt'
response = get(url)
print(response.text[:500])
# + id="PNJ5u7aCzAhe" colab_type="code" outputId="e525ca4e-a796-46f1-be16-2e7961ad9edd" colab={"base_uri": "https://localhost:8080/", "height": 34}
from bs4 import BeautifulSoup
html_soup = BeautifulSoup(response.text, 'html.parser')
type(html_soup)
# + id="yZs106Mazfkk" colab_type="code" outputId="a38a83f3-0605-407e-91fa-9c1345d9a474" colab={"base_uri": "https://localhost:8080/", "height": 255}
vote_container = html_soup.find_all('div', class_ = 'leftAligned')
vote_container
# vote_container[1:11]
# + id="aIpz8WFw0RXC" colab_type="code" outputId="e1a8b031-87e6-487e-fb27-55d7af4501d6" colab={"base_uri": "https://localhost:8080/", "height": 34}
votes = [containers.text for containers in vote_container][1:11]
votes[1:11]
# + id="iNye77eX0rCT" colab_type="code" outputId="4fcd7a27-f4cf-4b53-9030-808a9f450245" colab={"base_uri": "https://localhost:8080/", "height": 163}
list(map(int, votes))
# + id="HEQhy4h81Mhk" colab_type="code" outputId="52a84a67-606b-4d0d-ac72-6db8ffc1c3e3" colab={"base_uri": "https://localhost:8080/", "height": 54}
urls = ['https://www.imdb.com/title/tt6105098/ratings?ref_=tt_ov_rt', # Lion King (2019)
'https://www.imdb.com/title/tt0323073/ratings?ref_=tt_ov_rt', # Lion King (1994)
'https://www.imdb.com/title/tt6139732/ratings?ref_=tt_ov_rt', # Aladdin (2019)
'https://www.imdb.com/title/tt0103639/ratings?ref_=tt_ov_rt'] # Aladdin (1992) w/ <NAME>
votes_list = []
for url in urls:
# Get raw HTML response
response = get(url)
# Convert to BS Object
html_soup = BeautifulSoup(response.text, 'html.parser')
# Find vote containers and extract star ratings
vote_containers = html_soup.find_all('div', class_ = 'leftAligned')
votes = [containers.text for containers in vote_containers][1:11]
# Append to initial list
votes_list.append(votes)
print(votes_list)
# + id="6MY4-jVe1Rv8" colab_type="code" outputId="c54fd0df-1eba-46d1-b9b9-c4454a8eeb29" colab={"base_uri": "https://localhost:8080/", "height": 359}
import pandas as pd
movies = ['The Lion King (2019)', 'The Lion King (1994)', 'Aladdin (2019)', 'Aladdin (1992)']
df = pd.DataFrame(votes_list)
df = df.T
df.columns = movies
df = df.apply(lambda x: x.str.replace(',',''))
df['Star Rating'] = range(1,11)[::-1]
df
# + id="ag7TrBva2Wjt" colab_type="code" outputId="38d97339-e376-44ef-c606-122119d1a0fa" colab={"base_uri": "https://localhost:8080/", "height": 1000}
df_tidy = df.melt(id_vars='Star Rating')
df_tidy = df_tidy.rename(columns={'variable' : 'Movie', 'value': 'Number of Votes'})
df_tidy['Number of Votes'] = pd.to_numeric(df_tidy['Number of Votes'])
df_tidy
# + id="K2QDG3nD4H1s" colab_type="code" outputId="a72ccbb8-a408-4a2c-854a-ee5e0a8d12d3" colab={"base_uri": "https://localhost:8080/", "height": 204}
df_tidy['Vote Percent'] = df_tidy.groupby('Movie')['Number of Votes'].apply(lambda x: x / x.sum() *100)
df_tidy.head()
# + id="isAtZRww3Js7" colab_type="code" outputId="28f5a3f3-e532-43ae-bfbc-c5e2519dc161" colab={"base_uri": "https://localhost:8080/", "height": 197}
import seaborn as sns
sns.catplot(x='Star Rating', y='Vote Percent', col='Movie', col_wrap=2, height=6, kind='bar', data=df_tidy)
# + [markdown] id="5_na7Oy3NGKA" colab_type="text"
# # Making Explanatory Visualizations with Seaborn
# + colab_type="code" id="ya_w5WORGs-n" outputId="0dbf77af-aa69-4d25-cdb7-bf0e7f8058ad" colab={"base_uri": "https://localhost:8080/", "height": 355}
from IPython.display import display, Image
url = 'https://fivethirtyeight.com/wp-content/uploads/2017/09/mehtahickey-inconvenient-0830-1.png'
example = Image(url=url, width=400)
display(example)
# + [markdown] colab_type="text" id="HP4DALiRG3sC"
# Using this data: https://github.com/fivethirtyeight/data/tree/master/inconvenient-sequel
# + [markdown] colab_type="text" id="HioPkYtUG03B"
# Links
# - [Strong Titles Are The Biggest Bang for Your Buck](http://stephanieevergreen.com/strong-titles/)
# - [Remove to improve (the data-ink ratio)](https://www.darkhorseanalytics.com/blog/data-looks-better-naked)
# - [How to Generate FiveThirtyEight Graphs in Python](https://www.dataquest.io/blog/making-538-plots/)
# + [markdown] colab_type="text" id="0w_iMnQ6-VoQ"
# ## Make prototypes
#
# This helps us understand the problem
# + colab_type="code" id="5uz0eEaEN-GO" outputId="38105367-c228-4b3a-f58a-56e7d4e2935c" colab={"base_uri": "https://localhost:8080/", "height": 285}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
plt.style.use('fivethirtyeight')
fake = pd.Series([38, 3, 2, 1, 2, 4, 6, 5, 5, 33],
index=range(1,11))
fake.plot.bar(color='C1', width=0.9);
# + colab_type="code" id="KZ0VLOV8OyRr" outputId="7bba6b4d-5ca3-49ab-cb0f-7857b8a8932f" colab={"base_uri": "https://localhost:8080/", "height": 289}
fake2 = pd.Series(
[1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
2, 2, 2,
3, 3, 3,
4, 4,
5, 5, 5,
6, 6, 6, 6,
7, 7, 7, 7, 7,
8, 8, 8, 8,
9, 9, 9, 9,
10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10])
fake2.value_counts().sort_index().plot.bar(color='C1', width=0.9);
# + [markdown] colab_type="text" id="mZb3UZWO-q05"
# ## Annotate with text
# + colab_type="code" id="f6U1vswr_uWp" outputId="a61a09c2-3ef8-4d87-b0e6-9121a42aaba0" colab={"base_uri": "https://localhost:8080/", "height": 387}
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
fake.plot.bar(color='C3', width=0.9);
plt.text(x=-1.5, y=50,fontsize=16, fontweight='bold',
s = "'An Inconvenient Sequel: Truth to Power' is divisive")
# plt.title("'An Inconvenient Sequel: Truth to Power' is divisive")
plt.yticks([0, 10, 20, 30, 40])
plt.xlabel('Rating', fontsize=10, fontweight='bold')
plt.ylabel('Percent of Total Votes', fontsize=10, fontweight='bold')
# + [markdown] colab_type="text" id="x8jRZkpB_MJ6"
# ## Reproduce with real data
# + colab_type="code" id="3SOHJckDUPI8" outputId="44b0cc7b-780f-4553-a502-eef9fbe229e7" colab={"base_uri": "https://localhost:8080/", "height": 326}
df = pd.read_csv('https://raw.githubusercontent.com/fivethirtyeight/data/master/inconvenient-sequel/ratings.csv')
print(df.shape)
df.head()
# + colab_type="code" id="cDltXxhC_yG-" outputId="f2ad9654-3414-4ac7-f070-0bc74fa9b8b2" colab={"base_uri": "https://localhost:8080/", "height": 357}
df.category.value_counts()
# + id="jVNKbNBX_Goj" colab_type="code" outputId="d4d23a26-6fd9-4981-91ed-0cfa538aae02" colab={"base_uri": "https://localhost:8080/", "height": 493}
df.dtypes
# + id="YmA70f4-_VDl" colab_type="code" outputId="63bbb77b-e1e2-4ddd-847e-018e19ef4ca8" colab={"base_uri": "https://localhost:8080/", "height": 136}
df['timestamp'] = pd.to_datetime(df['timestamp'])
df.timestamp.describe()
# + id="dMBjKVcJ_yGo" colab_type="code" outputId="7a549309-a492-437f-bf1a-9eea06add097" colab={"base_uri": "https://localhost:8080/", "height": 34}
df_imdb = df[df.category == 'IMDb users']
df_imdb.shape
# + id="Cs5DqPoKACdh" colab_type="code" outputId="9c9bead5-0b6a-42dd-aedf-86dec1847685" colab={"base_uri": "https://localhost:8080/", "height": 117}
final = df_imdb.tail(1)
final
# + id="sycvD4uxAUEl" colab_type="code" outputId="163a9450-e81b-405a-ce9b-e642485d3bb6" colab={"base_uri": "https://localhost:8080/", "height": 187}
columns = ['{}_pct'.format(i) for i in range(1,11)]
columns = [f'{i}_pct' for i in range(1,11)]
columns
# + id="AMTPXc5DBC0z" colab_type="code" outputId="029b5145-9f79-4472-df01-0e2d3d1b5c2b" colab={"base_uri": "https://localhost:8080/", "height": 359}
data = final[columns]
data= data.T
data.index = range(1,11)
data
# + id="mx3U9D_8AQYW" colab_type="code" outputId="36201b4f-755f-4fb0-c4ac-8b488f752454" colab={"base_uri": "https://localhost:8080/", "height": 68}
# !free -m
# + id="ivu_Sh5R--Of" colab_type="code" outputId="c852ed28-cd32-45ed-f974-79932b8eb701" colab={"base_uri": "https://localhost:8080/", "height": 384}
plt.style.use('fivethirtyeight')
data.plot.bar(color='C3', width=0.9, legend=False);
plt.text(x=-1.5, y=50,fontsize=16, fontweight='bold',
s = "'An Inconvenient Sequel: Truth to Power' is divisive")
# plt.title("'An Inconvenient Sequel: Truth to Power' is divisive")
plt.yticks([0, 10, 20, 30, 40])
plt.xlabel('Rating', fontsize=10, fontweight='bold')
plt.ylabel('Percent of Total Votes', fontsize=10, fontweight='bold')
# + [markdown] colab_type="text" id="NMEswXWh9mqw"
# # ASSIGNMENT
#
# Replicate the lesson code. I recommend that you [do not copy-paste](https://docs.google.com/document/d/1ubOw9B3Hfip27hF2ZFnW3a3z9xAgrUDRReOEo-FHCVs/edit).
#
# # STRETCH OPTIONS
#
# #### 1) Reproduce another example from [FiveThityEight's shared data repository](https://data.fivethirtyeight.com/).
#
# #### 2) Reproduce one of the following using a library other than Seaborn or Matplotlib.
#
# For example:
# - [thanksgiving-2015](https://fivethirtyeight.com/features/heres-what-your-part-of-america-eats-on-thanksgiving/) (try the [`altair`](https://altair-viz.github.io/gallery/index.html#maps) library)
# - [candy-power-ranking](https://fivethirtyeight.com/features/the-ultimate-halloween-candy-power-ranking/) (try the [`statsmodels`](https://www.statsmodels.org/stable/index.html) library)
# - or another example of your choice!
#
# #### 3) Make more charts!
#
# Choose a chart you want to make, from [Visual Vocabulary - Vega Edition](http://ft.com/vocabulary).
#
# Find the chart in an example gallery of a Python data visualization library:
# - [Seaborn](http://seaborn.pydata.org/examples/index.html)
# - [Altair](https://altair-viz.github.io/gallery/index.html)
# - [Matplotlib](https://matplotlib.org/gallery.html)
# - [Pandas](https://pandas.pydata.org/pandas-docs/stable/visualization.html)
#
# Reproduce the chart. [Optionally, try the "<NAME>."](https://docs.google.com/document/d/1ubOw9B3Hfip27hF2ZFnW3a3z9xAgrUDRReOEo-FHCVs/edit) If you want, experiment and make changes.
#
# Take notes. Consider sharing your work with your cohort!
#
#
#
#
#
#
#
#
# + [markdown] id="rtmXNOc2M6zv" colab_type="text"
# ###Assignment
# + id="j2fG3BRZGHY3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 544} outputId="1c8075bb-0910-4cfa-fc66-e695e6c813e7"
from requests import get
#importing
url = 'https://www.imdb.com/title/tt0121765/ratings?ref_=tt_ov_rt'
respoonse = get(url)
print(response.text[:600])
# + id="3wKTCm03oEsS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d7f04ac1-3960-40c5-acbb-61b844f14356"
from bs4 import BeautifulSoup
#scraping one movie
html_soup = BeautifulSoup(response.text, 'html.parser')
type(html_soup)
# + id="AcZAXDhoDu-1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="45e0dc4d-9e3e-447c-9657-520737acdd8d"
vote_container = html_soup.find_all('div', class_ ='leftAligned')
vote_container[1:11]
# + id="wjjrfJ4kGps9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="c1416651-0ddf-4c38-dbd8-fd1db4ee1f57"
votes = [containers.text for containers in vote_container][1:11]
votes
# + id="HM_LY24AGpyg" colab_type="code" colab={}
#scraping 3 movies
# + id="m4odVeVjxXdt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="0834d2e3-8fab-4423-953c-9f4bcba1ca6f"
urls = ['https://www.imdb.com/title/tt0121765/ratings?ref_=tt_ov_rt', #Star Wars: Episode II - Attack of the Clones (2002)
'https://www.imdb.com/title/tt0080684/ratings?ref_=tt_ov_rt', #Star Wars: Episode V - The Empire Strikes Back (1980)
'https://www.imdb.com/title/tt2527336/ratings?ref_=tt_ov_rt'] #Star Wars: The Last Jedi (2017)
votes_list = []
for url in urls:
response = get(url)
html_soup = BeautifulSoup(response.text, 'html.parser')
vote_containers = html_soup.find_all('div', class_ = 'leftAligned')
votes = [containers.text for containers in vote_containers][1:11]
votes_list.append(votes)
print(votes_list)
# + id="RoCkxKH0EMfE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="bbac9e8a-e8cc-45a5-f933-e206617063c3"
import pandas as pd
#creating a dataset out of scraped movies
movies = ['Attack of the Clones (2002)', 'The Empire Strikes Back (1980)', 'The Last Jedi (2017)']
df = pd.DataFrame(votes_list)
df = df.T
df.columns = movies
df = df.apply(lambda x: x.str.replace( ',',''))
df['Star Rating'] = range(1,11)[::-1]
df
# + id="y-X6q4S6FD4X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 979} outputId="e9e2a643-6d20-4d72-e9de-43b49c27d634"
#Needs to be tidied up
df_tidy = df.melt(id_vars='Star Rating')
df_tidy = df_tidy.rename(columns={'variable': 'Movie', 'value': 'Number of Votes'})
df_tidy['Number of Votes'] = pd.to_numeric(df_tidy['Number of Votes'])
df_tidy
# + id="1pK6q2zpH8iS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="02c4a43c-8d7f-4fd9-b6c3-16aa7c205918"
#creating another column
df_tidy['Vote Percent'] = df_tidy.groupby('Movie')['Number of Votes'].apply(lambda x: x / x.sum() * 100)
df_tidy.head()
# + id="IJeACAb9H_1s" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="551b7437-5384-4cc4-ed20-79990cd6d73f"
df_tidy.info()
# + id="ZvpcZwzMIJ8o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 533} outputId="b2bced00-95c2-4691-8498-1223aa649684"
# simple plotting
import seaborn as sns
sns.catplot(x='Star Rating', y='Vote Percent', col='Movie', data=df_tidy, col_wrap=4, height=7, kind='bar', palette='muted', legend=False);
# + id="aEaWoy_FJF0p" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 544} outputId="5adfc219-a330-4305-b324-dd8d94e0da5b"
url = ('https://www.imdb.com/title/tt4154796/ratings?ref_=tt_ov_rt')
response = get(url)
print(response.text[:600])
# + id="MzKuz49ZMmQd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="e78f9849-26ab-4d14-a0a8-89806c5b30ce"
html_soup = BeautifulSoup(response.text, 'html.parser')
vote_container = html_soup.find_all('div', class_ ='leftAligned')
vote = [containers.text for containers in vote_container][1:11]
vote
# + id="tbXBmeolQ6OD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="9f6e5f94-a095-40f0-abe0-94515950e145"
import pandas as pd
df = pd.DataFrame(vote)
df = df.apply(lambda x: x.str.replace(',',''))
df['Star Rating'] = range(1,11)[::-1]
df
# + id="4sj0LbhsaWob" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="52cf5b95-8bbb-400c-e1f7-fee6d2446f6d"
df_tidy = df.melt(id_vars='Star Rating')
df_tidy = df_tidy.rename(columns={'variable': 'Movie', 'value': 'Number of Votes'})
df_tidy['Number of Votes'] = pd.to_numeric(df_tidy['Number of Votes'])
df_tidy
# + id="wcjdDFSWaHAY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="5e9e4dae-4e1a-45c5-f724-eb058eeca71a"
df['Vote Percent'] = df_tidy.groupby('Movie')['Number of Votes'].apply(lambda x: x / x.sum() * 100)
df.head()
# + id="c9-9wCKfLr0I" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 355} outputId="a237431c-5467-42d9-ffc2-76a30583f48d"
from IPython.display import display, Image
url = 'https://fivethirtyeight.com/wp-content/uploads/2017/09/mehtahickey-inconvenient-0830-1.png'
example = Image(url=url, width=400)
display(example)
# + id="RrFvE9_TO6jh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 291} outputId="3a1a7ace-4943-4c0d-bb8d-39bc75dd82b6"
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('fivethirtyeight')
df.plot.bar(color='C3', width=0.9, legend=False);
plt.text(x=1.8, y=11,fontsize=16, fontweight='bold',
s = "Avengers: Endgame")
# plt.yticks([10, 9, 8, 7, 6, 5, 4, 3, 2, 1])
plt.xlabel('Rating', fontsize=10, fontweight='bold')
plt.ylabel('Percent of Total Votes', fontsize=10, fontweight='bold');
# + id="u4zp38vtYjxk" colab_type="code" colab={}
|
module3-make-explanatory-visualizations/Vince_Emma_LS_DS_123_Make_Explanatory_Visualizations.ipynb
|