repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
OWASP/Nettacker | automation | 321 | This project isn't shown in https://owasp.org/projects | So its visibility can be very small.
I think you should talk with someone of OWASP website staff.
Thanks | closed | 2020-07-19T15:12:23Z | 2020-07-19T16:04:38Z | https://github.com/OWASP/Nettacker/issues/321 | [] | q2dg | 2 |
jeffknupp/sandman2 | rest-api | 37 | sqlalchemy_utils.PasswordType makes JSONEncoder's serialization barf | Here's my stacktrace:
```
File "/Library/Frameworks/Python.framework/Versions/3.4/lib/python3.4/site-packages/flask/json.py", line 83, in default
return _json.JSONEncoder.default(self, o)
File "/Library/Frameworks/Python.framework/Versions/3.4/lib/python3.4/json/encoder.py", line 173, in default
raise Type... | closed | 2016-07-31T02:53:17Z | 2016-08-05T20:28:43Z | https://github.com/jeffknupp/sandman2/issues/37 | [
"invalid",
"wontfix"
] | Datamance | 3 |
JoeanAmier/TikTokDownloader | api | 58 | 关于UserAgent 旁边的参数 | 我看代码里写了几条UserAgent的值,并且还有对应的二维数组。这些二维数组的值是怎么获取到的,或者可以加更多的UserAgent吗 | open | 2023-09-06T08:03:39Z | 2023-09-08T10:22:46Z | https://github.com/JoeanAmier/TikTokDownloader/issues/58 | [] | BaoStorm | 3 |
allenai/allennlp | nlp | 4,862 | save git status when run commands | Sometimes after changing many versions of the code, I'm confused about how I got this result. It would be nice if allennlp could log the current git status to `serialization_dir` when running `train` command.
Here is an example of a transformers record(`git_log.json`):
```
{
"repo_id": "<git.repo.base.Repo '/... | open | 2020-12-14T11:30:54Z | 2022-08-10T03:41:38Z | https://github.com/allenai/allennlp/issues/4862 | [
"Good First Issue",
"Contributions welcome",
"Feature request"
] | tshu-w | 12 |
nerfstudio-project/nerfstudio | computer-vision | 2,868 | How to find Nerf++ in Nerfstudio? | I have used Lenovo's computer system.How do I find the Nerf++ in the NerfStudio?Hope someone can help me. | closed | 2024-02-03T12:07:17Z | 2024-02-04T09:57:05Z | https://github.com/nerfstudio-project/nerfstudio/issues/2868 | [] | shehuirenwy | 1 |
OFA-Sys/Chinese-CLIP | nlp | 95 | Chinese-CLIP是如何修改context length的?How does Chinese-CLIP change the context length? | 我看到cn-clip是能够修改tokenizer的context length,但是我没有找到相关的代码是如何实现这个的。
在clip中,tokenizer的max context length为77,因为text-encoder在训练的时候就是如此。所以我想问一下,cn-clip是如何做到的呢?具体的代码又是在哪?
I see that cn-clip is able to modify the context length of the tokenizer, but I can't find the relevant code to implement this.
In clip, the tokenizer ... | closed | 2023-04-28T11:56:21Z | 2023-05-27T14:42:55Z | https://github.com/OFA-Sys/Chinese-CLIP/issues/95 | [] | DengXianqi | 2 |
OpenInterpreter/open-interpreter | python | 1,050 | Generated code is trimmed when using "-m gemini/gemini-pro" | ### Describe the bug
Please note that I'm using the `gemini/gemini-pro` implementation (which uses Google AI Studio / free) instead of the `gemini-pro` implementation (which uses Google Vertex AI / trial).
### Command ###
docker run --rm -it --name interpreter-instance openinterpreter interpreter -m gemini/ge... | open | 2024-03-02T04:04:59Z | 2024-10-31T08:34:20Z | https://github.com/OpenInterpreter/open-interpreter/issues/1050 | [
"Bug",
"More Information Required"
] | kripper | 8 |
microsoft/unilm | nlp | 755 | LayoutLM V2 error srcIndex < srcSelectDimSize | **Describe**
I am using LayoutLM V2 model. I am trying to finetune the the model by using my custom dataset. I got bellow error message.
Please tell me how to resolve the error.
you can download the code and dataset along with notebook
https://drive.google.com/file/d/1VdTvn580pGgVBlN03UX5alaFqSbc8Q5_/view?usp=sh... | open | 2022-06-10T10:45:35Z | 2023-02-22T08:50:39Z | https://github.com/microsoft/unilm/issues/755 | [] | koyelseba | 3 |
bendichter/brokenaxes | matplotlib | 49 | Hide the break | Want to hide the break between the axes and want to make it continuous. Any solution?
And also if is it possible to show only one number at the break for this continuous axis? | closed | 2020-05-19T10:24:02Z | 2020-06-03T04:17:17Z | https://github.com/bendichter/brokenaxes/issues/49 | [] | shivanshi13 | 3 |
man-group/arctic | pandas | 500 | import _compress results in no suitable image found | #### Arctic Version
```
1.59.0
```
#### Arctic Store
```
from . import _compress as clz4
```
#### Platform and version
macOSx 10.11.6
#### Description of problem and/or code sample that reproduces the issue
when I run a simple import arctic, I get a problem where it gets hung on
from . impo... | closed | 2018-02-06T06:33:37Z | 2018-08-25T15:41:31Z | https://github.com/man-group/arctic/issues/500 | [] | cavnerj | 2 |
davidteather/TikTok-Api | api | 519 | get_Video_By_Url issues | I use get_Video_By_Url to download video,it doesn' t work
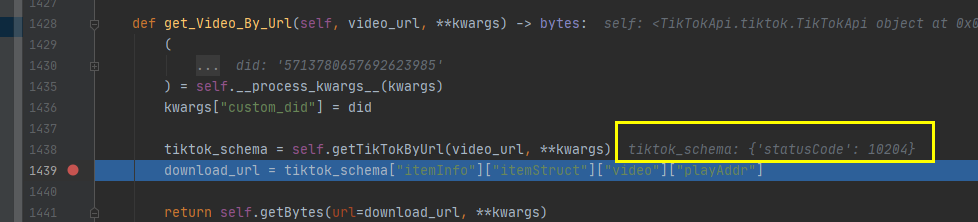
| closed | 2021-03-04T04:49:16Z | 2021-03-20T18:24:42Z | https://github.com/davidteather/TikTok-Api/issues/519 | [] | xyjw | 1 |
matplotlib/matplotlib | matplotlib | 29,008 | [Bug]: intersphinx on meson-python is broken | ### Bug summary
Since recently, sphinx-build error with ( e.g. https://app.circleci.com/pipelines/github/matplotlib/matplotlib/33363/workflows/c7423837-956c-4d75-85de-93e55fbdb8a5/jobs/85311):
> intersphinx inventory 'https://meson-python.readthedocs.io/en/stable/objects.inv' not readable due to ValueError: unkno... | closed | 2024-10-22T08:56:12Z | 2024-10-22T09:38:40Z | https://github.com/matplotlib/matplotlib/issues/29008 | [
"Documentation: build"
] | timhoffm | 1 |
Avaiga/taipy | automation | 2,293 | Have part or dialog centered to the element clicked | ### Description
Here, I have clicked on an icon and I have a dropdown menu of labels next to where I clicked:

Here, I have clicked on icon and I see a dialog/part showing up next to where I clicked:


request id:
f9bcb26c-a7a6-43c6-82a6-b284cc3889f3
| closed | 2025-01-17T10:01:57Z | 2025-01-17T16:13:01Z | https://github.com/yeongpin/cursor-free-vip/issues/35 | [] | geeklx | 0 |
jupyter-incubator/sparkmagic | jupyter | 516 | Release on Anaconda missing PySpark3 | The latest release of Anaconda on the Anaconda channel is missing PySpark3. The conda-forge channel appears to be fine.
https://anaconda.org/anaconda/sparkmagic/files | closed | 2019-03-12T22:47:01Z | 2019-06-27T14:38:08Z | https://github.com/jupyter-incubator/sparkmagic/issues/516 | [] | jaipreet-s | 2 |
ranaroussi/yfinance | pandas | 2,262 | Intraday data returned omits last daily datapoint | Running a demo using the below code and returning the expected range of data, but for each day the closing datapoint at 1600 is being omitted.
```
tick = 'vxf'
en = datetime.now()
st = en - timedelta(days = 59)
data = yf.Ticker(tick).history(interval='30m', start=st.strftime('%Y-%m-%d'), end=en.strftime('%Y-%m-%d')... | closed | 2025-02-12T20:26:48Z | 2025-02-12T22:11:18Z | https://github.com/ranaroussi/yfinance/issues/2262 | [] | cppt | 1 |
FactoryBoy/factory_boy | sqlalchemy | 352 | Custom provider declaration example in add_provider documentation | Example https://factoryboy.readthedocs.io/en/latest/reference.html#factory.Faker.add_provider showing how to create `SmileyProvider` would be nice.
BTW where are sources for the docs? https://github.com/FactoryBoy/factory_boy/blob/master/docs/reference.rst don't have following section. | open | 2017-03-09T12:17:37Z | 2017-11-01T00:12:32Z | https://github.com/FactoryBoy/factory_boy/issues/352 | [] | buoto | 2 |
JoeanAmier/TikTokDownloader | api | 380 | 建议加下载文件命名自定义功能 | 几个字段可以自定义设置
假设按照以下这些来定义
发布日期:YYYYMMDD
发布时间:hhmm
发布用户:user
作品标题:title
作评ID:id
这边举个例子,我个人是按这样命名保存的
YYYY.MM.DD_hhmm_user_title | open | 2025-01-16T16:55:55Z | 2025-01-17T01:05:27Z | https://github.com/JoeanAmier/TikTokDownloader/issues/380 | [] | yingfeng-i | 2 |
huggingface/datasets | numpy | 7,378 | Allow pushing config version to hub | ### Feature request
Currently, when datasets are created, they can be versioned by passing the `version` argument to `load_dataset(...)`. For example creating `outcomes.csv` on the command line
```
echo "id,value\n1,0\n2,0\n3,1\n4,1\n" > outcomes.csv
```
and creating it
```
import datasets
dataset = datasets.load_dat... | open | 2025-01-21T22:35:07Z | 2025-01-30T13:56:56Z | https://github.com/huggingface/datasets/issues/7378 | [
"enhancement"
] | momeara | 1 |
globaleaks/globaleaks-whistleblowing-software | sqlalchemy | 3,894 | Deduplicated notification emails | ### Proposal
Notification emails from errors should be deduplicated to prevent spamming. This should be according to content / stack trace, not timestamp. As in, if I get `FooException` 100 times in an hour, I want maybe the first email and then a summary ("100 cases in the last hour") to let me know it's repeating. A... | open | 2023-12-16T09:23:52Z | 2023-12-16T09:23:52Z | https://github.com/globaleaks/globaleaks-whistleblowing-software/issues/3894 | [
"T: Feature",
"Triage"
] | brassy-endomorph | 0 |
hbldh/bleak | asyncio | 586 | Using write without response causes exception on disconnect | * bleak version: 0.12.0
* Python version: 3.9.5
* Operating System: macOS 11.4
* BlueZ version (`bluetoothctl -v`) in case of Linux: N/A
### Description
It seems that calling `write_gatt_char` with `response` set to False results in an exception being thrown when `disconnect()` is called.
### What I Did
He... | closed | 2021-07-02T04:15:15Z | 2021-07-07T17:23:49Z | https://github.com/hbldh/bleak/issues/586 | [
"Backend: Core Bluetooth"
] | adamincera | 3 |
charlesq34/pointnet | tensorflow | 258 | Why the accuracy of train is high and the result of val is poor | Hello, I am a newbie in deep learning.
I would like to ask, I use the part_seg program to classify a large-scale urban point cloud data set (500m*500m). The training data is the data of the assigned categories in this data set, and the verification data is part of this data set.
I divide the input training data into ... | open | 2020-11-30T09:10:04Z | 2021-03-19T20:22:34Z | https://github.com/charlesq34/pointnet/issues/258 | [] | yasongguo | 1 |
keras-team/keras | tensorflow | 20,136 | Keras 3 doesn't map dictionary inputs by "key" | The code below runs in Tensorflow 2.11 (keras 2) but not in tf-nightly (Keras 3.4.1 ). I think Keras 3 doesn't map inputs by dict key
Epoch 1/10
Traceback (most recent call last):
File "/home/wangx286/rnn-base-caller/base_caller/scripts/example_metric.py", line 32, in <module>
model.fit({'before': x_train... | closed | 2024-08-19T19:33:43Z | 2024-08-21T04:08:16Z | https://github.com/keras-team/keras/issues/20136 | [
"type:Bug"
] | MeowTheCat | 2 |
DistrictDataLabs/yellowbrick | scikit-learn | 1,289 | Use classification visualizers directly from predictions, targets and logits? | Hi,
I work on classification problems and really like the design of the classification visualizers and their plots.
Nevertheless, I am a pytorch user. I usually store the model's output on test set as "prediction", "target", and "logits" (probability of each class).
It looks to me that classification report, c... | closed | 2022-11-29T20:57:18Z | 2022-11-29T21:48:00Z | https://github.com/DistrictDataLabs/yellowbrick/issues/1289 | [] | 2533245542 | 1 |
ultrafunkamsterdam/undetected-chromedriver | automation | 916 | detected at mastersportal.com | ### Link: `https://www.mastersportal.com/studies/294909/environmental-economics-and-management.html?ref=search_card`
### Code:
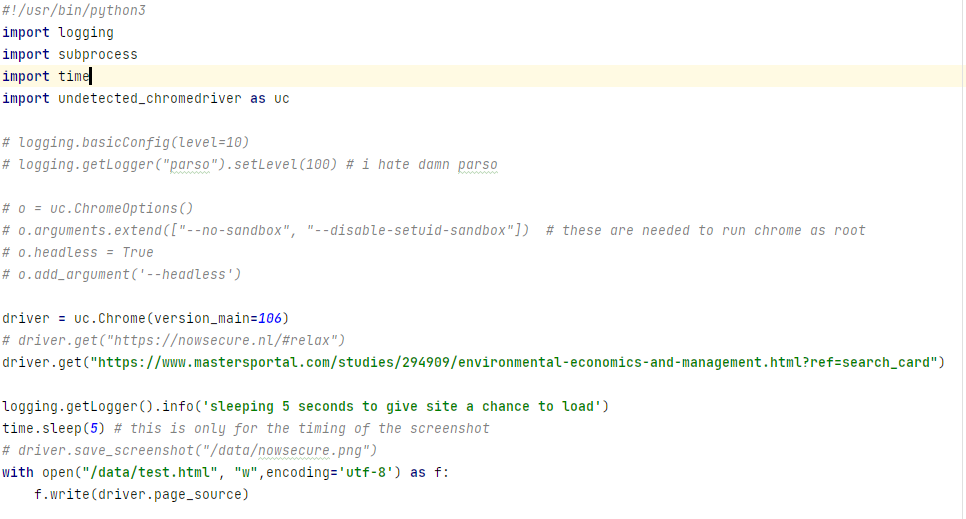
### Result:
 KeyError: 'pass_ticket' | itchat 1.3.5 版本。
这个在login.py 里面提示,`self.loginInfo['pass_ticket'])
KeyError: 'pass_ticket'`,是啥情况啊 | closed | 2017-06-26T13:51:02Z | 2018-02-28T04:13:17Z | https://github.com/littlecodersh/ItChat/issues/427 | [
"question"
] | lucasjinreal | 8 |
vitalik/django-ninja | pydantic | 1,222 | Resolve method is not being called in a nested schema | Hey everyone, I am trying to output a nested schema in a response but the resolve method of the nested schema is not being called and therefore, the following error is being raised:
The main schema for the response (shortened for brevity):
```
class DetailedAlbumOut(Schema):
id: int
artists: List[... | closed | 2024-07-07T20:29:41Z | 2024-07-11T04:35:19Z | https://github.com/vitalik/django-ninja/issues/1222 | [] | millejon | 4 |
indico/indico | flask | 6,018 | Unschedule contribution icon change | Currently, the unschedule contribution icon is a trash can. It is confusing for users, who think the action will be deleting the contribution altogether.
There is no "clock with cross" icon in the icomoon collection, but maybe just a cross would do? Or a composition of icons?
| open | 2023-11-01T14:47:02Z | 2023-11-01T14:47:19Z | https://github.com/indico/indico/issues/6018 | [
"enhancement",
"new-timetable"
] | javfg | 0 |
amdegroot/ssd.pytorch | computer-vision | 343 | line 83, in __call__\n label_idx = self.class_to_ind[name]\nKeyError: | open | 2019-05-08T08:47:24Z | 2022-09-09T09:08:08Z | https://github.com/amdegroot/ssd.pytorch/issues/343 | [] | sxyxf66 | 11 | |
aidlearning/AidLearning-FrameWork | jupyter | 50 | How to reorganize keyboard layout? | I wish there would be a-left-arrow button,how could I do this?
thanks a lot | closed | 2019-09-12T07:00:04Z | 2019-09-14T04:16:20Z | https://github.com/aidlearning/AidLearning-FrameWork/issues/50 | [] | dobefore | 1 |
deepinsight/insightface | pytorch | 2,714 | error installing on windows 11 |
C:\Users\jeffr\AppData\Local\Temp\pip-install-fw8krnga\insightface_ac45d088a86942fe933a54455be8f4c2\insightface\thirdparty\face3d\mesh\cython\mesh_core.h(4): fatal error C1083: Cannot open include file: 'stdio.h': No such file or directory
error: command 'C:\Program Files (x86)\Microsoft Visual Studio\2022\BuildTool... | open | 2025-01-03T11:55:56Z | 2025-03-10T03:06:35Z | https://github.com/deepinsight/insightface/issues/2714 | [] | J-Ai-57 | 1 |
donnemartin/data-science-ipython-notebooks | machine-learning | 64 | alexnet.ipynb contains incomplete architecture of alexnet(2 cnn layers missing) | Alexnet implementation in tensorflow has incomplete architecture where 2 convolution neural layers are missing. This issue is in reference to the python notebook mentioned below.
https://github.com/donnemartin/data-science-ipython-notebooks/blob/master/deep-learning/tensor-flow-examples/notebooks/3_neural_networks/a... | open | 2019-04-09T18:08:10Z | 2020-09-27T16:39:19Z | https://github.com/donnemartin/data-science-ipython-notebooks/issues/64 | [
"needs-review"
] | harshitsaini | 5 |
mwaskom/seaborn | matplotlib | 3,352 | Categorical scatter plots on symlog-scaled axis | Hi,
On the current dev version (eb2b5a2) and matplotlib 3.7.1, consider the following code that uses `stripplot` to draw two points on unscaled and symlog-scaled yaxis:
```python
import seaborn as sns
import matplotlib.pyplot as plt
x = [0.1,2]
y = [0.1,5]
fig, axs = plt.subplots(ncols=2)
sns.stripplot(x=x,... | closed | 2023-05-01T09:18:52Z | 2023-08-20T22:08:34Z | https://github.com/mwaskom/seaborn/issues/3352 | [
"bug",
"mod:categorical"
] | MaozGelbart | 0 |
python-security/pyt | flask | 36 | Add readthedocs | If you look at https://github.com/trailofbits/manticore/blob/master/README.md you can see a nice link at the top to the docs. I'll write the docs once the layout is there, please see
https://www.slideshare.net/mobile/JohnCosta/how-to-readthedocs
(So the [easy] issues are good for new people who want to start contr... | closed | 2017-04-25T17:55:22Z | 2017-07-13T00:47:01Z | https://github.com/python-security/pyt/issues/36 | [
"enhancement",
"easy"
] | KevinHock | 9 |
peerchemist/finta | pandas | 27 | possible error in calculation | may be you can double check..
but i think this line https://github.com/peerchemist/finta/blob/master/finta/finta.py#L798
should be
ohlc["down_move"] = -ohlc["low"].diff() | closed | 2019-04-28T17:04:30Z | 2019-05-05T11:36:51Z | https://github.com/peerchemist/finta/issues/27 | [] | livinter | 2 |
davidsandberg/facenet | computer-vision | 757 | when i train a classifier on own images,the accuracy = 0 | open | 2018-05-24T08:01:28Z | 2018-07-18T13:54:49Z | https://github.com/davidsandberg/facenet/issues/757 | [] | chankillo | 1 | |
benbusby/whoogle-search | flask | 559 | [QUESTION] How to use social media alternatives using url parameter | The official instance, by default, doesn't use social media alternatives.
Since I am using Cookie AutoDelete extension, if I change config, it won't persist.
So is it possible to do this via url parameter? | closed | 2021-11-28T04:51:45Z | 2021-12-17T15:48:31Z | https://github.com/benbusby/whoogle-search/issues/559 | [
"question"
] | specter78 | 5 |
Kanaries/pygwalker | pandas | 654 | Make Streamlit Bike Sharing app contained within Pygwalker universe | I had two issues when trying to convert the Streamlit Bike Sharing app to a panel app.
- The [gw_config.json](https://github.com/Kanaries/pygwalker-in-streamlit/blob/main/gw_config.json) cannot be used by `GraphicWalker` React directly. I need to unwrap it by taking the inner `configuration` when using with `Graphic... | open | 2024-11-06T04:18:36Z | 2024-11-09T04:06:50Z | https://github.com/Kanaries/pygwalker/issues/654 | [] | MarcSkovMadsen | 2 |
openapi-generators/openapi-python-client | fastapi | 545 | Delete request with body | **Is your feature request related to a problem? Please describe.**
In our API we handle a lot of items and came to a point where we want to delete a lot of this items at the same time. Our first approach was to call a DELETE on every single ID. This works, but it is very slow.
Then we added a new delete functionality... | closed | 2021-12-16T09:53:02Z | 2022-01-19T15:28:43Z | https://github.com/openapi-generators/openapi-python-client/issues/545 | [
"✨ enhancement"
] | MalteBecker | 4 |
python-restx/flask-restx | flask | 480 | How do I document the required header in and endpoint, when I'm passing a model to expect() instead of a regparse? | Regparser is not good at documented nested models, so I have switched to passing a model into the `@api.expect()` decorator
`@api.expect(models.input_model_request_generation, validate=True)`
However that does not document my headers.
For headers, the documentation says the following
 in the docs. When I load the playground and issue the query `subscription { counter }`, instead of a working counter, an "unsupported operand" error is raised:
``... | closed | 2020-02-14T05:28:55Z | 2021-01-01T16:24:09Z | https://github.com/mirumee/ariadne/issues/320 | [
"bug",
"roadmap"
] | command-tab | 8 |
ageitgey/face_recognition | python | 993 | How can I add Percentage rate? | * face_recognition version: 1.2.3
* Python version: 3.6.8
* Operating System: windows 10
### Description
Hi Adam, Firstly thanks for great library, I already 21k faces(one of mine) encoded and insert to sql , I am testing three different photos of mine. Results are below.
First photo : 40 different person (o... | closed | 2019-12-03T15:22:07Z | 2021-05-27T01:38:43Z | https://github.com/ageitgey/face_recognition/issues/993 | [] | HAKANMAZI | 4 |
JaidedAI/EasyOCR | machine-learning | 412 | Missing information while extracting text from similar images | I have similar set of images from which I am trying to extract. On some images it is working good but on certain it misses necessary information. The images have texts written on them in German.
In the first image the information "Verkauft" could not be extracted, while in the next image it was extracted. I had such... | closed | 2021-04-01T11:30:21Z | 2021-04-06T17:53:16Z | https://github.com/JaidedAI/EasyOCR/issues/412 | [] | RishikMani | 2 |
modin-project/modin | data-science | 7,315 | Avoid unnecessary length checks in `df.squeeze` | It is possible that when `axis=1` in squeeze we still check `len(self.index)`, which is never necessary when `axis=1`. Link to code here: https://github.com/modin-project/modin/blob/eac3c77baf456c7bd7e1e5fde81790a4ed3ebb27/modin/pandas/dataframe.py#L2074-L2084
This is an easy fix, also see https://github.com/snowfla... | closed | 2024-06-14T15:48:36Z | 2024-09-20T18:46:25Z | https://github.com/modin-project/modin/issues/7315 | [] | sfc-gh-dpetersohn | 0 |
scikit-optimize/scikit-optimize | scikit-learn | 1,130 | BayesSearchCV returns different results when n_points is changed | Hello,
I'm using the 'unofficial' version of BayesSearchCV with multimetrics. In order to improve parallel processing and speed up run times with my new machine, I increased the n_points parameter from the default 1.
However, for every different value of n_points I used, I got different sets of results all else b... | open | 2022-10-14T02:16:47Z | 2022-10-14T12:17:53Z | https://github.com/scikit-optimize/scikit-optimize/issues/1130 | [] | RNarayan73 | 0 |
autokey/autokey | automation | 635 | Support for multiple languages (l10n) | ## Classification:
UI/Usability
## Reproducibility:
Always
## AutoKey version:
Not relevant
## Used GUI:
Gtk
## Installed via:
Package manager
## Linux distribution:
Not relevant
## Summary:
Currently GUI speaks only English. It would be great if support for other languages is added.
## Step... | open | 2021-12-01T15:25:58Z | 2023-12-10T06:36:59Z | https://github.com/autokey/autokey/issues/635 | [
"enhancement",
"help-wanted",
"user interface"
] | jose1711 | 4 |
CorentinJ/Real-Time-Voice-Cloning | pytorch | 633 | Installation solutions for people with multiple python versions? | I've been doing a lot of troubleshooting trying to get this to work. though I _believe_ im on python 3.8 right now, ive been installing everything using **pip** instead of **pip3**. thus installing tensorflow version 1.15 didnt work, instead i installed the newest tensorflow. installing everything else worked fine.
... | closed | 2021-01-20T00:15:48Z | 2021-01-20T17:31:28Z | https://github.com/CorentinJ/Real-Time-Voice-Cloning/issues/633 | [] | Woolton | 2 |
coqui-ai/TTS | python | 3,299 | [Bug] CUDA crash when running xttx inference in Fastapi for streaming endpoint. | ### Describe the bug
I am using code at: https://github.com/hengjiUSTC/xtts-streaming-server/blob/main/server/main.py Building a Fastapi server for streaming TTS service. Got following error
```
Traceback (most recent call last):
File "/home/ubuntu/xtts-streaming-server/server/venv/lib/python3.10/site-packages/... | closed | 2023-11-24T10:40:24Z | 2024-09-12T11:14:08Z | https://github.com/coqui-ai/TTS/issues/3299 | [
"bug"
] | hengjiUSTC | 8 |
jupyterlab/jupyter-ai | jupyter | 385 | Allow REQUESTS_CA_BUNDLE | Re: https://github.com/jupyterlab/jupyter-ai/issues/321#issuecomment-1714127620
### Problem
* using different OpenAI base url for chat UI I am getting connection failed or timeout error
* my OpenAI base url starts with https
### Proposed Solution
* allow adding https connection option with certificate
#... | closed | 2023-09-12T07:34:32Z | 2024-06-26T15:58:08Z | https://github.com/jupyterlab/jupyter-ai/issues/385 | [
"enhancement",
"scope:chat-ux"
] | sqlreport | 2 |
nerfstudio-project/nerfstudio | computer-vision | 3,078 | 如何评估和渲染结果? | **Is your feature request related to a problem? Please describe.**
A clear and concise description of what the problem is. Ex. I'm always frustrated when [...]
**Describe the solution you'd like**
A clear and concise description of what you want to happen.
**Describe alternatives you've considered**
A clear an... | open | 2024-04-16T03:17:07Z | 2024-04-26T06:29:50Z | https://github.com/nerfstudio-project/nerfstudio/issues/3078 | [] | Fanjunyi55 | 2 |
unit8co/darts | data-science | 2,119 | Cannot get optuna gridSearch to work. TypeError: Unknown type of parameter:series, got:TimeSeries | I am following your guides on optuna and Ray tune. With ray tune i keep getting time out error and dont know why , but i will start asking about optuna. I want to use lightgbm (as i understand it , any model in darts , i should be able to use). Will ask about optuna since i did manage to get it to work some time ago wi... | closed | 2023-12-13T10:23:49Z | 2023-12-14T08:50:49Z | https://github.com/unit8co/darts/issues/2119 | [
"triage"
] | Allena101 | 1 |
home-assistant/core | asyncio | 140,373 | VMB7IN state is 0 | ### The problem
The measurement state for a VMB7IN entity stays at 0.0
<img width="1428" alt="Image" src="https://github.com/user-attachments/assets/dda46a7b-b729-4460-be21-413c7b01f7b2" />
(the counter is still working)
<img width="1422" alt="Image" src="https://github.com/user-attachments/assets/4eba6215-d2cf-475... | closed | 2025-03-11T13:05:05Z | 2025-03-17T07:31:56Z | https://github.com/home-assistant/core/issues/140373 | [
"integration: velbus"
] | CasperBE | 8 |
coqui-ai/TTS | deep-learning | 2,997 | [Bug]Training using multiple GPU's | ### Describe the bug
RuntimeError: [!] 2 active GPUs. Define the target GPU by `CUDA_VISIBLE_DEVICES`. For multi-gpu training use `TTS/bin/distribute.py`.
But i cannot find distribute.py in that location also distribute.py is in TTS/utils/distribute.py
I am trying use multiple GPU for training on custom data, b... | closed | 2023-09-25T21:56:46Z | 2024-09-03T01:13:48Z | https://github.com/coqui-ai/TTS/issues/2997 | [
"bug"
] | 18Raksha | 5 |
strawberry-graphql/strawberry | asyncio | 3,154 | Make HTTP request data available when logging errors | <!--- Provide a general summary of the changes you want in the title above. -->
When logging errors, I am not aware of a method to add the IP address and similar info to the logged data. Specifically, I'm looking to set the base properties of [GCP HTTP request log entries](https://cloud.google.com/logging/docs/refer... | open | 2023-10-17T09:01:41Z | 2025-03-20T15:56:26Z | https://github.com/strawberry-graphql/strawberry/issues/3154 | [] | moritz89 | 0 |
django-import-export/django-import-export | django | 1,026 | How can we import a csv file with ANSI encoding | I'm trying to import a file that has ANSI encoding, Accented characters like "ç" etc.
The import shows an error as mentioned below.
> Imported file has a wrong encoding: 'utf-8' codec can't decode byte 0xf3 in position 18710: invalid continuation byte
Changing it to utf-8 breaks the characters into "Blockers" | closed | 2019-11-06T08:50:47Z | 2020-05-28T07:25:50Z | https://github.com/django-import-export/django-import-export/issues/1026 | [
"stale"
] | farhankn | 3 |
holoviz/panel | jupyter | 6,946 | Tabulator selectable is broken | panel==1.4.4
I believe Tabulator js has changed how `selectable` works and Panel needs to adapt. It will change even for from 5.5 (current panel version) to 6.2 (latest js version).
```python
import panel as pn
import pandas as pd
import numpy as np
pn.extension("tabulator")
sel_df = pd.DataFrame(np.random... | closed | 2024-06-28T10:14:17Z | 2024-07-28T20:17:42Z | https://github.com/holoviz/panel/issues/6946 | [
"component: tabulator"
] | MarcSkovMadsen | 2 |
aiortc/aiortc | asyncio | 331 | Error using MediaRecorder creating HLS segments | Hi, this is a great library.
I am attempting to use the MediaRecorder to create hls segments, but ffmpeg encounteres the following error during transmuxing:
`Application provided invalid, non monotonically increasing dts to muxer in stream 1: 2217000 >= 2217000
`
I am using the MediaRecorder as in the server ... | closed | 2020-04-06T19:51:06Z | 2022-03-11T17:56:00Z | https://github.com/aiortc/aiortc/issues/331 | [] | tlaz4 | 16 |
pallets/quart | asyncio | 99 | LifespanFailure Quart 11.3 | I am getting the following error in my app in the latest version:
```python
File "app.py", line 118, in <module>
app.run(host='0.0.0.0', port=port)
File "C:\xxxxxx\Anaconda3\envs\api\lib\site-packages\quart\app.py", line 1615, in run
loop.run_until_complete(task)
File "C:\xxxxxx\Anaconda3\envs\api\l... | closed | 2020-02-26T21:12:00Z | 2022-07-05T01:59:06Z | https://github.com/pallets/quart/issues/99 | [] | slyduda | 6 |
unionai-oss/pandera | pandas | 932 | @pa.check_types won't validate in MyPy using type hints that trigger the method | #### Question about pandera
The actual code I am using and testing:
```python
import typing
import pandera as pa
from pandera.typing import DataFrame, Index, Series
from pandera.typing.common import DataFrameBase
class EntitySchema(pa.SchemaModel):
"""EntitySchema - base class for nodes and edges.... | open | 2022-09-01T08:32:38Z | 2022-09-01T08:32:38Z | https://github.com/unionai-oss/pandera/issues/932 | [
"question"
] | rjurney | 0 |
OFA-Sys/Chinese-CLIP | nlp | 371 | 关于调用cn_clip进行特征提取报错格式错误的问题 | Text的特征提取代码如下:
import cn_clip.clip as clip
from cn_clip.clip import load_from_name, available_models
class TextCLIPModel(nn.Module):
def __init__(self, config, device):
super().__init__()
self.device = device
self.model, self.preprocess = self._load_model(config)
def _load_mo... | open | 2024-12-02T09:01:24Z | 2024-12-02T09:01:24Z | https://github.com/OFA-Sys/Chinese-CLIP/issues/371 | [] | Seing-yu | 0 |
httpie/cli | python | 1,252 | Choco installed packages conflict with the user's own site-packages | See [this](https://discord.com/channels/725351238698270761/799982808122523648/924860935074635777) thread on our discord server for details. We should try to be more isolated for package installations on windows. | open | 2021-12-27T09:26:24Z | 2021-12-28T10:39:07Z | https://github.com/httpie/cli/issues/1252 | [
"windows",
"packaging",
"low-priority"
] | isidentical | 0 |
AirtestProject/Airtest | automation | 949 | airtest多次run case以后手机变得卡顿 | (请尽量按照下面提示内容填写,有助于我们快速定位和解决问题,感谢配合。否则直接关闭。)
**(重要!问题分类)**
* 测试开发环境AirtestIDE使用问题 -> https://github.com/AirtestProject/AirtestIDE/issues
* 控件识别、树状结构、poco库报错 -> https://github.com/AirtestProject/Poco/issues
* 图像识别、设备控制相关问题 -> 按下面的步骤
**描述问题bug**
使用"python3 -m airtest run {case} --device Android:///”启动airtest ru... | closed | 2021-08-04T01:42:36Z | 2021-10-14T02:29:12Z | https://github.com/AirtestProject/Airtest/issues/949 | [] | ZhangOscar | 3 |
marshmallow-code/flask-smorest | rest-api | 99 | Deserialization at point of request handling | Hi!
First off, let me say that this library is the closest thing to what I've been looking for as an API framework in flask. Awesome job pulling in the best practices of API framework tools! I will try to put a few hours a week to helping in any way I can.
My question. As shown in your documentation, even thoug... | closed | 2019-09-18T15:01:13Z | 2019-09-20T15:13:47Z | https://github.com/marshmallow-code/flask-smorest/issues/99 | [
"question"
] | georgesequeira | 2 |
Lightning-AI/pytorch-lightning | data-science | 20,045 | Skip certain step during training | ### Bug description
I want to ignore some batch step during training, how can I write the code? Any suggestions would be appreciated.Thanks in advance.
The chatGPT answer below:
```
def on_train_batch_start(self, batch, batch_idx, dataloader_idx):
# Define steps to skip
steps_to_skip = {19... | closed | 2024-07-04T13:13:00Z | 2024-07-14T10:32:56Z | https://github.com/Lightning-AI/pytorch-lightning/issues/20045 | [
"question",
"ver: 2.2.x"
] | real-junjiezhang | 3 |
home-assistant/core | python | 141,110 | Can't connect after reboot | ### The problem
After update HA Core to 2025.3.4 my Tado can't no more connect !

### What version of Home Assistant Core has the issue?
2025.3.4
### What was the last working version of Home Assistant Core?
2025.3.3
### Wha... | closed | 2025-03-22T12:29:08Z | 2025-03-23T01:03:09Z | https://github.com/home-assistant/core/issues/141110 | [] | beckynet | 14 |
newpanjing/simpleui | django | 3 | 再提个建议啊 | 作者你好:
在使用了你这个插件后,在首页左下角会有你项目的git地址,这个怎么去掉呢?虽然我很支持你,但如果不去掉这个地址,实在是无法应用到项目中,也不利于贵项目的推广呀。
如下:
Simpleui
项目主页:https://www.88cto.com/project/simpleui/
Github:https://github.com/newpanjing/simpleui | closed | 2018-12-13T14:47:35Z | 2018-12-21T03:46:20Z | https://github.com/newpanjing/simpleui/issues/3 | [] | wthahaha | 4 |
cupy/cupy | numpy | 8,779 | `cupy.ravel` behaves differently with `numpy.ravel` | ### Description
As claimed in [NumPy doc](https://numpy.org/doc/stable/reference/generated/numpy.ravel.html):
> When order is ‘K’, it will preserve orderings that are neither ‘C’ nor ‘F’, but won’t reverse axes:
```py
>>> a = np.arange(12).reshape(2,3,2).swapaxes(1,2); a
array([[[ 0, 2, 4],
[ 1, 3, 5]],
... | closed | 2024-12-02T00:52:15Z | 2025-02-07T00:14:39Z | https://github.com/cupy/cupy/issues/8779 | [
"issue-checked"
] | AnonymousPlayer2000 | 2 |
Kludex/mangum | fastapi | 119 | Store the 'requestContext' in WebSocket message events | Currently just store the initial connection event data, should add a key to the scope for updating the message request context. | closed | 2020-05-21T08:27:16Z | 2020-06-28T01:52:35Z | https://github.com/Kludex/mangum/issues/119 | [
"improvement",
"websockets"
] | jordaneremieff | 0 |
Sanster/IOPaint | pytorch | 356 | [BUG] | **Model**
Which model are you using?
**Describe the bug**
A clear and concise description of what the bug is.
**Screenshots**
If applicable, add screenshots to help explain your problem.
**System Info**
Software version used
- Platform: Windows-10-10.0.22000-SP0
- Python version: 3.11.3
- torch: 2.0.1
... | closed | 2023-08-04T05:19:58Z | 2023-08-30T03:27:50Z | https://github.com/Sanster/IOPaint/issues/356 | [] | szcelp | 0 |
fastapi-users/fastapi-users | fastapi | 630 | Use relative `tokenUrl` parameter for JWTAuthentication (and docs) | Currently the [JWTAuthentication docs page](https://frankie567.github.io/fastapi-users/configuration/authentication/jwt/) doesn't document the `tokenUrl` parameter, although it does document all its other parameters.
When/if this gets added, it would be worth mentioning that the `tokenUrl` must be relative (i.e. no ... | closed | 2021-05-12T19:38:00Z | 2021-05-20T09:47:24Z | https://github.com/fastapi-users/fastapi-users/issues/630 | [
"documentation",
"enhancement"
] | eddsalkield | 4 |
Anjok07/ultimatevocalremovergui | pytorch | 1,749 | salom | Last Error Received:
Process: VR Architecture
If this error persists, please contact the developers with the error details.
Raw Error Details:
RuntimeError: "Error(s) in loading state_dict for CascadedNet:
Missing key(s) in state_dict: "stg1_low_band_net.0.enc1.conv.0.weight", "stg1_low_band_net.0.enc1.conv.1.weig... | open | 2025-02-24T05:32:17Z | 2025-02-25T21:49:05Z | https://github.com/Anjok07/ultimatevocalremovergui/issues/1749 | [] | Jasurbek1987 | 1 |
widgetti/solara | jupyter | 831 | Can't use tooltips for children of `ToggleButtonsSingle` | This is somewhat related to #683
If I try to use tooltips for each Button inside a ToggleButtonsSingle component, the selected value is set to the tooltip value, not the one from the button.
## Correct behavior (but no tooltips)
```python
import solara
map_type = solara.Reactive("stack")
@solara.compon... | open | 2024-10-24T11:42:17Z | 2024-11-22T10:28:03Z | https://github.com/widgetti/solara/issues/831 | [
"bug"
] | lopezvoliver | 0 |
jupyter-book/jupyter-book | jupyter | 2,153 | Fix analytics config remapping | ### Describe the bug
The upstream `pydata-sphinx-theme` understands configuration sections for analytics information, namely `html.analytics`. These changes post-date the Jupyter Book config & documentation, so we need to update it to match.
### Reproduce the bug
NA
### List your environment
_No response_ | closed | 2024-05-28T10:37:00Z | 2024-05-28T12:24:32Z | https://github.com/jupyter-book/jupyter-book/issues/2153 | [
"bug"
] | agoose77 | 0 |
replicate/cog | tensorflow | 1,801 | Don't hold event lock while processing iterator models | https://github.com/replicate/cog/pull/1773/files#r1676200859 | closed | 2024-07-12T17:06:35Z | 2024-07-18T13:02:41Z | https://github.com/replicate/cog/issues/1801 | [] | nickstenning | 2 |
roboflow/supervision | machine-learning | 1,670 | Problem with minimum matching threshold parameter of ByteTracker | ### Search before asking
- [X] I have searched the Supervision [issues](https://github.com/roboflow/supervision/issues) and found no similar bug report.
### Bug
Hi folks. Amazing project, but I'm getting a peculiar behaviour in ByteTracker.
My assumption for the `minimum_matching_threshold` parameter of ... | open | 2024-11-15T01:54:36Z | 2025-01-09T15:57:27Z | https://github.com/roboflow/supervision/issues/1670 | [
"bug"
] | rsnk96 | 1 |
OFA-Sys/Chinese-CLIP | computer-vision | 236 | loss 为0 | 您好,我训练自己的数据,loss 为 0 可能是什么原因,日志如下:
2023-12-14,08:40:01 | INFO | Rank 0 | Global Steps: 240/270 | Train Epoch: 3 [60/90 (67%)] | Loss: 0.000000 | Image2Text Acc: 100.00 | Text2Image Acc: 100.00 | Data Time: 0.042s | Batch Time: 0.170s | LR: 0.000004 | logit_scale: 2.659 | Global Batch Size: 1 | open | 2023-12-14T08:47:31Z | 2023-12-20T03:19:45Z | https://github.com/OFA-Sys/Chinese-CLIP/issues/236 | [] | wwangxinhao | 1 |
zappa/Zappa | django | 812 | [Migrated] Streaming data with Flask's stream_with_context function does not behave as expected | Originally from: https://github.com/Miserlou/Zappa/issues/1980 by [ArmanMaesumi](https://github.com/ArmanMaesumi)
<!--- Provide a general summary of the issue in the Title above -->
## Context
I am trying to use Flask's stream_with_context function to stream a large file (100mb-500mb) while it is being created.
... | closed | 2021-02-20T12:51:57Z | 2022-08-18T02:01:26Z | https://github.com/zappa/Zappa/issues/812 | [] | jneves | 1 |
pyg-team/pytorch_geometric | pytorch | 9,222 | pyproject.toml doesn't list as dependencies modules imported at runtime: dgl, torch_sparse | ### 🐛 Describe the bug
dgl, torch_sparse are imported, but not mentioned in pyproject.toml
### Versions
HEAD | closed | 2024-04-21T03:06:15Z | 2024-04-22T15:43:44Z | https://github.com/pyg-team/pytorch_geometric/issues/9222 | [
"bug"
] | yurivict | 2 |
airtai/faststream | asyncio | 1,297 | Docs: dealing with different schema registries | It would be good to add documentation with examples how to deal with different schema registries. Again there are many registries and coupling router with a particular registry isn't a good idea, unless there will be a some Abstract class first, so later community can add implementation | open | 2024-03-11T10:23:41Z | 2024-08-21T19:09:52Z | https://github.com/airtai/faststream/issues/1297 | [
"documentation",
"Confluent"
] | davorrunje | 0 |
aimhubio/aim | tensorflow | 2,501 | Flag / option to auto-commit or store diff patch | ## 🚀 Feature
Flag or option on run instantiation (or maybe some config file somewhere) to auto-commit when a new run is started so that commits stored on Aim are synced with the git repo.
### Motivation
Often, commits on Aim are not in sync with the git repo state because uncommitted changes are not incorpora... | open | 2023-01-25T19:13:09Z | 2023-02-01T18:47:04Z | https://github.com/aimhubio/aim/issues/2501 | [
"type / enhancement",
"area / SDK-storage"
] | rodrigo-castellon | 1 |
litestar-org/litestar | pydantic | 3,893 | Ehancement: CLI - Better error message for invalid `--app` string | ### Description
A condition is missing for the case that `app_path` does not contain a colon.
```
Using Litestar app from env: 'invalid'
Traceback (most recent call last):
File "/home/henry/miniconda3/envs/facefusion/bin/litestar", line 8, in <module>
sys.exit(run_cli())
File "/home/henry/miniconda3/... | closed | 2024-12-07T13:27:44Z | 2025-03-20T15:55:03Z | https://github.com/litestar-org/litestar/issues/3893 | [
"Enhancement"
] | henryruhs | 3 |
facebookresearch/fairseq | pytorch | 5,510 | i have tried your hokkien demo before,it works well.but recently i found it not work .what's wrong | ## ❓ Questions and Help
### Before asking:
1. search the issues.
2. search the docs.
<!-- If you still can't find what you need: -->
#### What is your question?
#### Code
<!-- Please paste a code snippet if your question requires it! -->
#### What have you tried?
#### What's your environment?
... | open | 2024-06-21T09:42:28Z | 2024-06-21T09:42:28Z | https://github.com/facebookresearch/fairseq/issues/5510 | [
"question",
"needs triage"
] | Jackylee2032 | 0 |
gradio-app/gradio | data-science | 10,335 | How to present mathematical formulas? | Firstly, **I Tried gr.Markdown**. It doesn't work
Then, **I tried gr.Markdown and js** like this:
`<script type="text/javascript" async
src="https://cdnjs.cloudflare.com/ajax/libs/mathjax/2.7.7/MathJax.js?config=TeX-MML-AM_CHTML'"
</script>`
`out = gr.HTML(label="Answer", value=mathjax_script + "<div>This is... | closed | 2025-01-11T11:17:32Z | 2025-01-12T15:40:11Z | https://github.com/gradio-app/gradio/issues/10335 | [] | MrJs133 | 1 |
pyqtgraph/pyqtgraph | numpy | 2,417 | Precision issues in opengl renderer when zoomed in with high values | ### Short description
The gpu's ability to deal with interpolating numbers can be somewhat limited, it's not just the values you are sending down because the GPU needs to be able to interpolate between the points to draw the line which can require a bunch of additional precision.
The paintGL code winds up with 2 issu... | open | 2022-09-15T18:42:27Z | 2024-06-16T05:44:25Z | https://github.com/pyqtgraph/pyqtgraph/issues/2417 | [
"openGL"
] | gedalia | 8 |
globaleaks/globaleaks-whistleblowing-software | sqlalchemy | 3,550 | Select specific file types | ### Proposal
Hello,
is there a possibility that only certain file types can be sent with a report?
### Motivation and context
Certain file types may contain malicious code | closed | 2023-07-24T06:48:41Z | 2023-07-28T05:26:30Z | https://github.com/globaleaks/globaleaks-whistleblowing-software/issues/3550 | [
"T: Feature"
] | JimpoTEDY | 1 |
pydantic/pydantic-settings | pydantic | 494 | `BaseSettings.__init_subclass__()` takes no keyword arguments | When creating a new BaseSettings class, I get an error that states the `__init_subclass__` function takes no keyword arguments. I've taken the following example directly from the [Pydantic Settings Docs](https://docs.pydantic.dev/latest/concepts/pydantic_settings/#cli-kebab-case-for-arguments)
```python
import sys
... | closed | 2024-12-06T18:17:43Z | 2024-12-09T14:24:18Z | https://github.com/pydantic/pydantic-settings/issues/494 | [
"unconfirmed"
] | JoshLoecker | 2 |
paperless-ngx/paperless-ngx | django | 8,513 | [BUG] File descriptors gone wild | ### Description
I have a weird issue where paperless is using hundreds of thousands of file handles and browing out the server. On a completely idle startup, it's using ~150,000 handles. When using it, it rapidly goes up and breaks when the system runs out of file handles.
We are not using notify, but polling on... | closed | 2024-12-18T09:43:53Z | 2024-12-18T14:50:19Z | https://github.com/paperless-ngx/paperless-ngx/issues/8513 | [
"dependencies",
"not a bug"
] | PhantomPhoton | 1 |
nolar/kopf | asyncio | 1,070 | on.create() handler keeps getting fired every time object is modified | ### Long story short
I've implemented a firewall operator that assigns externalIPs to LoadBalancer services. The problem it that the on.create() handler keeps getting fired not only upon service creation, but also upon every modification of the service.
Ive tested this by creating a simple Watch stream in python ... | open | 2023-10-19T14:48:50Z | 2023-10-26T07:21:34Z | https://github.com/nolar/kopf/issues/1070 | [
"bug"
] | michal0000000 | 1 |
pyg-team/pytorch_geometric | pytorch | 8,994 | Possible overwriting scenario with Jinja | ### 🐛 Describe the bug
I am getting the following error, not always but from time to time:
```
File "/usr/local/lib/python3.8/dist-packages/torch_geometric/nn/conv/cg_conv.py", line 57, in __init__
super().__init__(aggr=aggr, **kwargs)
File "/usr/local/lib/python3.8/dist-packages/torch_geometric/nn/conv... | closed | 2024-02-29T15:31:31Z | 2024-03-01T18:24:03Z | https://github.com/pyg-team/pytorch_geometric/issues/8994 | [
"bug"
] | jychoi-hpc | 3 |
browser-use/browser-use | python | 92 | How to stop Python script after agent is done? | It requires to press "Enter" to stop, but in a Docker environment it's not that handy and triggering `exit()` seems to be a workaround.
Is it happening on Playwright level or inside browser-use? Anyone knows how to stop it? | open | 2024-12-06T08:54:10Z | 2024-12-06T14:01:55Z | https://github.com/browser-use/browser-use/issues/92 | [] | n-sviridenko | 1 |
fastapi/sqlmodel | pydantic | 37 | FastAPI and Pydantic - Relationships Not Working | ### First Check
- [X] I added a very descriptive title to this issue.
- [X] I used the GitHub search to find a similar issue and didn't find it.
- [X] I searched the SQLModel documentation, with the integrated search.
- [X] I already searched in Google "How to X in SQLModel" and didn't find any information.
- [X... | closed | 2021-08-26T22:40:52Z | 2024-08-22T16:54:39Z | https://github.com/fastapi/sqlmodel/issues/37 | [
"question"
] | Chunkford | 24 |
litestar-org/polyfactory | pydantic | 27 | `OrmarModelFactory.get_field_value` TypeError("object of type 'bool' has no len()") | Hi @Goldziher!
First of all, thanks for this superb library, I just started integrating it into my project and it seems very promising. I stumbled upon a problem, though, and I think it might be a problem with the library itself.
The `OrmarModelFactory` overrides the `get_field_value` method to handle choices fie... | closed | 2022-02-15T21:59:28Z | 2022-02-18T08:20:33Z | https://github.com/litestar-org/polyfactory/issues/27 | [] | mciszczon | 1 |
oegedijk/explainerdashboard | dash | 140 | Tabs are freezing | I have dataset which has 100k row and 50 column. I use classification use case.
--
Feature Importances
Classification Stats
--
tabs are opened in ~1sec.
---
Individual Predictions
What if...
Feature Dependence
Decision Trees
---
This tabs are opend in 30 second... What is the problem? I think they are pr... | closed | 2021-08-10T11:41:13Z | 2021-12-23T15:27:31Z | https://github.com/oegedijk/explainerdashboard/issues/140 | [] | nailcankara | 2 |
Kanaries/pygwalker | matplotlib | 240 | Running PyGWalker in a Hugging Face space | It would be amazing to test PyGWalker in a [Hugging Face space](https://huggingface.co/spaces), particularly on datasets hosted on the Hub.
| open | 2023-09-25T12:47:33Z | 2023-10-25T14:18:57Z | https://github.com/Kanaries/pygwalker/issues/240 | [
"enhancement"
] | severo | 15 |
christabor/flask_jsondash | flask | 38 | Only load js files if the chart for it is enabled in that view. | closed | 2016-08-26T21:45:02Z | 2016-09-09T22:23:53Z | https://github.com/christabor/flask_jsondash/issues/38 | [
"enhancement",
"performance"
] | christabor | 0 | |
HIT-SCIR/ltp | nlp | 428 | requirement.txt 下面 transformers 的版本必须是是3.2 有什么特殊原因么? | transformers==3.2.0 跟transformers 最新版有冲突。 安装是会自动降级transformers。 有什么特殊原因必须固定在3.2 版本么?由于版本原因会导致transformers 里面一些example 无法运行 | closed | 2020-10-29T00:03:25Z | 2020-11-02T06:23:11Z | https://github.com/HIT-SCIR/ltp/issues/428 | [] | johnsonice | 1 |
apachecn/ailearning | python | 499 | LSTM深入浅出的好文这篇 blog 链接已挂 | 第2部分-深度学习基础-深度学习必学下第4个链接 _LSTM深入浅出的好文: https://blog.csdn.net/roslei/article/details/61912618_ ,此链接不可用。 | closed | 2019-04-22T02:09:03Z | 2019-04-26T02:06:29Z | https://github.com/apachecn/ailearning/issues/499 | [] | Sunjk21 | 1 |
albumentations-team/albumentations | deep-learning | 2,439 | [Feature request] Add apply_to_images to ToGray | open | 2025-03-11T01:19:11Z | 2025-03-11T01:19:17Z | https://github.com/albumentations-team/albumentations/issues/2439 | [
"enhancement",
"good first issue"
] | ternaus | 0 | |
junyanz/pytorch-CycleGAN-and-pix2pix | computer-vision | 1,646 | Training Parameters or Architecture Settings Recommendations | closed | 2024-04-22T12:21:35Z | 2024-05-03T07:36:13Z | https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/issues/1646 | [] | selimceylan | 0 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.