repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
pyro-ppl/numpyro | numpy | 1,155 | Improved Error Message for Incorrect Sample Shape | I have a jax array of D parameters for D independent Bernoulli distributions. I wanted to draw N samples from each of the D Bernoulli distributions, but when I tried the following code, I received the error `unsupported operand type(s) for +: 'int' and 'tuple'`.
```
with numpyro.plate('data', num_obs):... | closed | 2021-09-13T03:42:06Z | 2021-09-24T14:18:34Z | https://github.com/pyro-ppl/numpyro/issues/1155 | [
"documentation"
] | RylanSchaeffer | 2 |
Morizeyao/GPT2-Chinese | nlp | 38 | argparse中参数gradient_accumulation类型错误 | **错误原因**
`train.py` 脚本中第 57 行 `parser` 设置参数 `gradient_accumulation` 的类型为 `str`
> parser.add_argument('--gradient_accumulation', default=1, type=str, required=False, help='梯度积累')
会导致第 139 进行除运算时抛出类型错误,不能对 str 做除法
> total_steps = int(full_len / stride * epochs / batch_size / gradient_accumulation)
**更正建议**
... | closed | 2019-08-27T09:44:47Z | 2019-08-28T06:36:53Z | https://github.com/Morizeyao/GPT2-Chinese/issues/38 | [] | xinfeng1i | 3 |
ultralytics/ultralytics | python | 19,261 | Assign tracker ID with unique ID | ### Search before asking
- [x] I have searched the Ultralytics YOLO [issues](https://github.com/ultralytics/ultralytics/issues) and [discussions](https://github.com/orgs/ultralytics/discussions) and found no similar questions.
### Question
Hello. Thanks for create such rich Ultralytics library.
I want to know Is i... | open | 2025-02-15T16:37:33Z | 2025-02-15T22:51:55Z | https://github.com/ultralytics/ultralytics/issues/19261 | [
"question",
"track"
] | erfansafaie | 4 |
ExpDev07/coronavirus-tracker-api | fastapi | 168 | Recovered number always zero | As I said, the 'recovered' field is always 0, in all countries. | closed | 2020-03-24T16:49:54Z | 2020-03-24T19:10:22Z | https://github.com/ExpDev07/coronavirus-tracker-api/issues/168 | [
"duplicate"
] | tonjo | 1 |
paperless-ngx/paperless-ngx | machine-learning | 8,828 | Docker image / Many duplicate files under the language assets directories | ### Description
I felt on this by accident while looking in the v2..14.x docker image.
Nothing important nor urgent, but it could help to reclaim easily 100M+ on the docker image.
Under both `/usr/src/paperless/src/documents/static/frontend` there are multiples files for each language that are repeated, while they ar... | closed | 2025-01-20T11:44:21Z | 2025-02-20T03:07:47Z | https://github.com/paperless-ngx/paperless-ngx/issues/8828 | [
"not a bug"
] | Daryes | 2 |
davidteather/TikTok-Api | api | 130 | [FEATURE_REQUEST] - Does the current release support http proxies? | **Is your feature request related to a problem? Please describe.**
A clear and concise description of what the problem is. Ex. I'm always frustrated when [...]
**Describe the solution you'd like**
A clear and concise description of what you want to happen.
**Describe alternatives you've considered**
A clear an... | closed | 2020-06-07T21:24:35Z | 2020-06-10T21:52:42Z | https://github.com/davidteather/TikTok-Api/issues/130 | [
"feature_request"
] | dj2ball | 3 |
hankcs/HanLP | nlp | 1,850 | " unpack (expected 4, got 3)" from HanLP(['XXXXX']) 运行错误 | <!--
感谢找出bug,请认真填写下表:
-->
**Describe the bug**
在本地运行SDP语义依存分析模型时候出现了bug
**Code to reproduce the issue**
```python
import hanlp
import torch
xx=hanlp.pretrained.sdp.SEMEVAL15_PSD_BIAFFINE_EN
HanLP = hanlp.load(xx,devices=torch.device('cpu'))
HanLP(['abc def ghk'])
```
**Describe the current behav... | closed | 2023-10-14T08:18:18Z | 2023-10-15T05:37:51Z | https://github.com/hankcs/HanLP/issues/1850 | [
"invalid"
] | 1558359609 | 1 |
neuml/txtai | nlp | 131 | Multi-labels classification ? | This tutorial seems to be a single-label classification case.
https://github.com/neuml/txtai/blob/master/examples/07_Apply_labels_with_zero_shot_classification.ipynb
Is it possible to have multilabel classification? | closed | 2021-10-28T22:16:44Z | 2021-11-02T15:46:29Z | https://github.com/neuml/txtai/issues/131 | [] | PetricaR | 3 |
biolab/orange3 | numpy | 6,009 | Allow vertical minimization of Paint Data window | Paint Data widget prevents me to make its window really small. I can minimize it any way I want to horizontally, but vertically it resists.
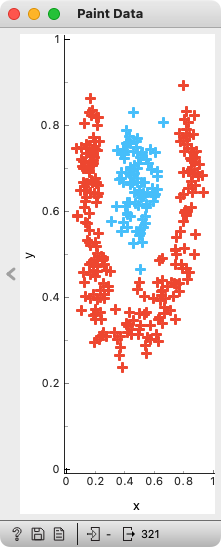
For instance, I can make Scatter Plot window much smaller, both ... | closed | 2022-06-08T19:53:16Z | 2022-06-10T13:41:11Z | https://github.com/biolab/orange3/issues/6009 | [] | BlazZupan | 0 |
nonebot/nonebot2 | fastapi | 3,038 | Plugin: 涩图插件 | ### PyPI 项目名
nonebot-plugin-picsetu
### 插件 import 包名
nonebot_plugin_picsetu
### 标签
[]
### 插件配置项
_No response_ | closed | 2024-10-19T13:48:44Z | 2024-10-20T02:19:51Z | https://github.com/nonebot/nonebot2/issues/3038 | [
"Plugin"
] | zhongwen-4 | 2 |
OpenInterpreter/open-interpreter | python | 1,571 | 是否可以使用 Gitee AI 的模型 API ? | ### Is your feature request related to a problem? Please describe.
这里是 Gitee AI 平台,隶属于开源中国项目。我们目前正在积极拓展合作伙伴。据我们了解,您的这款产品可以集成到我们的 Serverless API 应用中,并为您提供详细的集成配置指南。我们认为,通过合作,我们可以提升高端并共同提升用户的AI使用体验。
这是对应的活动文档:https:
[//ai.gitee.com/docs/openapi/serverless](https://ai.gitee.com/docs/openapi/serverless)
也可以用openai兼... | open | 2024-12-20T09:19:15Z | 2024-12-20T09:19:51Z | https://github.com/OpenInterpreter/open-interpreter/issues/1571 | [] | pittosporum1 | 0 |
polakowo/vectorbt | data-visualization | 98 | Getting the timestamp of a sell order | Is there a simple method to retrieve the Sell order date, currently, i'm doing this
import yfinance as yf
import numpy as np
import pandas as pd
import vectorbt as vbt
price = yf.Ticker('BTC-USD').history(period='max')['Close']
size = pd.Series.vbt.empty_like(price, 0.)
size.iloc[0] = np.inf # go all in
po... | closed | 2021-02-08T15:58:13Z | 2021-02-08T16:51:29Z | https://github.com/polakowo/vectorbt/issues/98 | [] | ben1628 | 2 |
iMerica/dj-rest-auth | rest-api | 489 | PasswordResetConfirm view does not include UID and Token values in the corresponding fields in the browsable API | After the email is sent, the PasswordResetConfirmView view of the `dj-rest-auth` package, the `uidb64` and `token` values are not passed automatically to the Token and UID fields in the Browsable API as I see for example in the demo example.
I'm using Django version 4.1.4, and my `urls.py` code is as follows:
``... | open | 2023-02-22T16:19:00Z | 2024-01-02T12:46:27Z | https://github.com/iMerica/dj-rest-auth/issues/489 | [] | rochdikhalid | 1 |
peerchemist/finta | pandas | 72 | Relative Strength Index (RSI) calculation | First of all, many thanks for this nice python library. I am using the version 0.4.4.
I computed the RSI on data fetched using `ccxt`, and compared it with the RSI shown on cryptowat.ch.
Unfortunately, the RSI calculated by finta did not match the RSI calculated by cryptowat.ch.
After investigation, I realized t... | closed | 2020-07-12T06:21:04Z | 2020-07-23T11:40:46Z | https://github.com/peerchemist/finta/issues/72 | [] | charlyisidore | 1 |
scikit-tda/kepler-mapper | data-visualization | 205 | Mismatch in the total number of samples | **Describe the bug**
I am having trouble with the number of samples in the output of `mapper`.
My starting point is a `769x769` distance matrix, `D`.
```
In [107]: D.shape
Out[107]: (769, 769)
```
I then run,
```
import kmapper as km
mapper = km.KeplerMapper(verbose=2)
lens = mapper.fit_transform(D, d... | closed | 2020-12-16T17:44:30Z | 2020-12-16T19:32:39Z | https://github.com/scikit-tda/kepler-mapper/issues/205 | [] | andreacortis | 2 |
widgetti/solara | fastapi | 563 | Functions run very slow when they are called under use_thread | **Problem**
A function runs very slow when it is run under `solara.use_thread`.
**How much slow:**
On my Mac, it is about 20 times. On Hugging Face (which uses Linux), 26 times slower.
**How to reproduce:**
I deployed the [code](https://huggingface.co/spaces/hkayabilisim/test_solara_use_thread/blob/main/test_... | closed | 2024-03-17T19:55:39Z | 2024-03-27T05:59:38Z | https://github.com/widgetti/solara/issues/563 | [] | hkayabilisim | 3 |
coqui-ai/TTS | deep-learning | 3,075 | [Feature request] xTTS server configuration example | <!-- Welcome to the 🐸TTS project!
We are excited to see your interest, and appreciate your support! --->
**🚀 Feature Description**
Not sure xTTS is supported in the server but in case it is could you please provide model and server config example?
**Solution**
model configutation example for multi speaker... | closed | 2023-10-17T14:43:49Z | 2023-11-28T11:06:05Z | https://github.com/coqui-ai/TTS/issues/3075 | [
"feature request"
] | darkzbaron | 1 |
SALib/SALib | numpy | 56 | Broken links on the io webpages | Links to readme and contributing are broken.
Not sure whether these are supposed to link to the github repository, or copies of these files on the web-page.
| closed | 2015-06-15T13:59:57Z | 2015-06-17T14:50:00Z | https://github.com/SALib/SALib/issues/56 | [] | willu47 | 1 |
deezer/spleeter | tensorflow | 923 | It's not working i don't know what to do | There's eroor
Progress idle
Starting processing of all songs
Processing: Users/user/Desktop/###mp3
2025-01-05 12:11:40.960984. F tensorflow/stream_executor/cuda/cuda_driver.cc:351]
Check falled:CUDA_SUCCESS==cuDevicePrimaryCtxGetState(device/former_primary_context_flags &former primary context_is_active) (... | open | 2025-01-08T05:19:58Z | 2025-01-08T05:28:12Z | https://github.com/deezer/spleeter/issues/923 | [
"question"
] | nohur7 | 0 |
xinntao/Real-ESRGAN | pytorch | 412 | bad tool trash | your tool does not work
just create a video from bad frames
creating a video out of phase with the audio | open | 2022-08-19T01:57:19Z | 2022-08-30T18:23:55Z | https://github.com/xinntao/Real-ESRGAN/issues/412 | [] | marana22 | 6 |
HIT-SCIR/ltp | nlp | 72 | 无用代码清理 | 以下列表中的文件LTP已经不再使用
```
__util/conversion_utf.h
__util/decode_gbk.h
__util/EncodeUtil.cpp
__util/EncodeUtil.h
__util/gbk_u16.h
__util/IniReader.cpp
__util/IniReader.h
__util/Logger.cpp
__util/Logger.h
__util/md5.cpp
__util/md5.h
__util/SBC2DBC.cpp
__util/SBC2DBC.h
__util/TextProcess.cpp
__util/TextProcess.h
__util/Timer... | closed | 2014-09-17T07:15:32Z | 2014-09-17T07:26:45Z | https://github.com/HIT-SCIR/ltp/issues/72 | [
"enhancement"
] | Oneplus | 0 |
aleju/imgaug | deep-learning | 638 | AttributeError: 'Array' object has no attribute 'deepcopy' (version 0.40) - python 3.7.6 | ~\AppData\Local\Continuum\anaconda3\envs\singan\lib\site-packages\imgaug\augmentables\utils.py in copy_augmentables(augmentables)
17 result.append(np.copy(augmentable))
18 else:
---> 19 result.append(augmentable.deepcopy())
20 return result
21
AttributeEr... | open | 2020-03-12T04:28:26Z | 2020-04-13T06:03:27Z | https://github.com/aleju/imgaug/issues/638 | [] | bluetyson | 4 |
microsoft/UFO | automation | 116 | Why the page is always stoping in "Round 1, Step 1, HostAgent: Analyzing the user intent and decomposing the request..." | I find the page is always stopping at "Round 1, Step 1, HostAgent: Analyzing the user intent and decomposing the request..."" when I use Gemini, please help me how to fix it.

| open | 2024-08-05T08:58:18Z | 2024-08-05T09:15:40Z | https://github.com/microsoft/UFO/issues/116 | [] | lovegit2021 | 3 |
explosion/spaCy | nlp | 13,658 | Spacy installation on python 3.13 fails | <!-- NOTE: For questions or install related issues, please open a Discussion instead. -->
## How to reproduce the behaviour
<!-- Include a code example or the steps that led to the problem. Please try to be as specific as possible. -->
`pip3.13 install spacy`
C:\Users\talta\AppData\Local\Programs\Python\Python3... | open | 2024-10-12T07:42:28Z | 2025-03-24T13:11:56Z | https://github.com/explosion/spaCy/issues/13658 | [] | RubTalha | 35 |
piskvorky/gensim | data-science | 2,986 | Faster evaluation metrics (baked into the library?) | Before getting into the issue, I'd like to thank you all for maintaining this library! It's been great so far, and I really appreciate the thorough documentation.
---
#### Problem description
I'm trying to train Word2Vec embedding vectors on my own dataset. Things have been going well so far, but as I've start... | closed | 2020-10-19T21:53:32Z | 2021-04-19T15:49:40Z | https://github.com/piskvorky/gensim/issues/2986 | [] | dataframing | 5 |
keras-team/keras | machine-learning | 20,297 | Webpage not rendering correctly | Please see the attached image. As you can see, the documentation for the `Attention` layer is not rendering correctly.

| closed | 2024-09-27T11:44:15Z | 2024-09-28T01:01:30Z | https://github.com/keras-team/keras/issues/20297 | [
"type:Bug"
] | dkgaraujo | 2 |
labmlai/annotated_deep_learning_paper_implementations | machine-learning | 189 | can not run ViT(vision transformer) experiment file (failed to connect to https://api.labml.ai/api/vl/track?run%20wuid-87829.c05191leeae2db06088ee9ee4&labml%20version=0.4.162) | When I try to run experiment.py, it gives an error message.
"failed to connect: https://api.labml.ai/api/vl/track?run%20wuid-87829.c05191leeae2db06088ee9ee4&labml%20version=0.4.162"
I also can't visit this site.
[https://api.labml.ai/api/vl/track?run%20wuid-87829.c05191leeae2db06088ee9ee4&labml%20version=0.4.162](ur... | closed | 2023-06-07T10:05:26Z | 2023-07-03T03:26:36Z | https://github.com/labmlai/annotated_deep_learning_paper_implementations/issues/189 | [] | HiFei4869 | 2 |
tflearn/tflearn | tensorflow | 623 | Warning! ***HDF5 library version mismatched error*** | when i run the example lstm_generate_cityname.py on my pc (win10 + anaconda python3.5 + tensorflow1.0 ), it will show this . | closed | 2017-02-23T02:41:12Z | 2017-02-23T04:07:06Z | https://github.com/tflearn/tflearn/issues/623 | [] | chenggui53 | 1 |
ResidentMario/geoplot | matplotlib | 30 | Assigning projections to plt.subplots()-generated axes generates rotated images | First off, this is a great library, thanks for making it!
I'm running into some non-intuitive behavior when using projections with `plt.subplots()`-generated axes. Briefly, passing a `projection` argument to both `plt.subplots` (via `subplot_kw=`) and `gplt.polyplot` generates images which are rotated by ~30 degrees... | closed | 2017-04-27T15:38:54Z | 2019-07-05T14:53:12Z | https://github.com/ResidentMario/geoplot/issues/30 | [] | hinnefe2 | 3 |
twopirllc/pandas-ta | pandas | 379 | How to get eri data in dataframe columns? | I don't understand how I get the eri values in a dataframe.
Here is a small example to explain myself better.
```python
import pandas as pd
import pandas_ta as pta
import numpy as np
df_size = 10
data = np.random.random_integers(10000, 30000, size=df_size)
df = pd.DataFrame(data, columns=['volume'])
data... | closed | 2021-08-26T13:19:18Z | 2021-08-26T19:06:43Z | https://github.com/twopirllc/pandas-ta/issues/379 | [
"info"
] | BillGatesIII | 2 |
huggingface/transformers | deep-learning | 36,022 | Transformers are untraceable with FX after 4.38 | ### System Info
- `transformers` version: 4.48.0
- Platform: Linux-6.8.0-1020-gcp-x86_64-with-glibc2.39
- Python version: 3.12.3
- Huggingface_hub version: 0.27.1
- Safetensors version: 0.4.5
- Accelerate version: not installed
- Accelerate config: not found
- PyTorch version (GPU?): 2.6.0a0+df5bbc09d1.nv24.12 (True)
... | closed | 2025-02-03T20:43:11Z | 2025-03-14T11:29:58Z | https://github.com/huggingface/transformers/issues/36022 | [
"bug"
] | Li357 | 7 |
ultralytics/ultralytics | machine-learning | 19,235 | Val mode - No json predictions & plots in return | ### Search before asking
- [x] I have searched the Ultralytics YOLO [issues](https://github.com/ultralytics/ultralytics/issues) and [discussions](https://github.com/orgs/ultralytics/discussions) and found no similar questions.
### Question
Hello,
When I run a Yolo val mode on custom data , I get my confusion matri... | closed | 2025-02-13T16:54:00Z | 2025-02-14T15:44:07Z | https://github.com/ultralytics/ultralytics/issues/19235 | [
"question",
"detect"
] | adriengoleb | 2 |
2noise/ChatTTS | python | 243 | hf的几个pt分别是做什么用的? | <img width="655" alt="image" src="https://github.com/2noise/ChatTTS/assets/3038472/84351770-176f-4fc9-9fc3-040a541ad031">
| closed | 2024-06-04T03:52:17Z | 2024-07-20T04:01:27Z | https://github.com/2noise/ChatTTS/issues/243 | [
"stale"
] | zh794390558 | 1 |
nolar/kopf | asyncio | 1,019 | Client-side throttling | ### Problem
I want to be able to use kopf to watch resources and update `last-handled-configuration` annotations for a large number of resources. (We are "migrating" from `kubectl apply --patch` to using kopf as our k8s "reconciler" and so for all previously managed k8s resources we need to patch those resources w/ ... | open | 2023-03-28T14:20:18Z | 2023-08-09T07:12:13Z | https://github.com/nolar/kopf/issues/1019 | [
"enhancement"
] | mecampbellsoup | 2 |
PokeAPI/pokeapi | api | 704 | Pokemon locations for Galar and Hisui | Galar and Hisui locations are missing. When might these are added? Thank you! | closed | 2022-03-22T19:11:49Z | 2022-03-28T17:52:00Z | https://github.com/PokeAPI/pokeapi/issues/704 | [] | 684efs3 | 1 |
xlwings/xlwings | automation | 2,422 | xlwings Server: respect the sync/async definition of custom functions | Currently, all custom functions run on an async server endpoint, even if they are defined as a sync function, which will block the event loop.
Instead, sync functions should behave like a sync fastapi endpoint, possibly via `run_in_threadpool`: https://github.com/tiangolo/fastapi/discussions/10768
Might need to be so... | closed | 2024-03-25T10:31:38Z | 2024-07-19T09:10:56Z | https://github.com/xlwings/xlwings/issues/2422 | [
"bug",
"Server"
] | fzumstein | 1 |
biolab/orange3 | scikit-learn | 6,480 | Forward selection in feature suggestion | I really like the feature suggestion in _Linear projection_ and _Radviz_ but finding a combination of as little as 4 features among 100+ attributes takes forever. I tried to use _Rank_ to reduce their number but often cannot go so low that it would be worth the wait (also independently measuring feature importance does... | closed | 2023-06-17T14:03:00Z | 2023-06-23T10:36:51Z | https://github.com/biolab/orange3/issues/6480 | [] | processo | 1 |
neuml/txtai | nlp | 678 | Move vector model caching from Embeddings to Vectors | Currently, `Vectors` instances are cached in an `Embeddings` when a models cache is set. This works well in most cases but it would be better if only the actual underlying model was cached and not the configuration parameters.
For example, with the new `dimensionality` parameter, it would be nice to have that be va... | closed | 2024-02-28T01:30:25Z | 2024-02-28T02:22:17Z | https://github.com/neuml/txtai/issues/678 | [] | davidmezzetti | 0 |
rthalley/dnspython | asyncio | 587 | Setting edns options in a query | Hi, I am trying to set EDNS options on a DNS query that should be equivalent to setting `+ednsopt=100:test_data` in `dig` options.
I tried the following:
```python
request = dns.message.make_query(domain, 'A')
opt = dns.edns.GenericOption(100, b'test_data')
request.use_edns(edns=True, options=[opt])... | closed | 2020-09-24T03:16:09Z | 2020-09-25T22:45:09Z | https://github.com/rthalley/dnspython/issues/587 | [
"Cannot Reproduce"
] | ChamaraG | 6 |
jupyter/nbgrader | jupyter | 927 | `nbgrader feedback` flag to add new feedback file instead of overwriting | Sometimes, I have a back and forth exchanging an assignment with students as they're resolving various issues. In these cases, it would be really helpful to have a flag for `nbgrader feedback` to generate a new feedback file while keeping the previous one (with the original version of the code and my comments) intact. ... | open | 2018-02-07T11:10:10Z | 2022-06-23T10:21:07Z | https://github.com/jupyter/nbgrader/issues/927 | [
"enhancement"
] | dlukes | 1 |
fastapi/sqlmodel | fastapi | 31 | How to deal with Postgres Enum columns? | ### First Check
- [X] I added a very descriptive title to this issue.
- [X] I used the GitHub search to find a similar issue and didn't find it.
- [X] I searched the SQLModel documentation, with the integrated search.
- [X] I already searched in Google "How to X in SQLModel" and didn't find any information.
- [X] I al... | closed | 2021-08-26T09:15:44Z | 2022-09-10T00:13:19Z | https://github.com/fastapi/sqlmodel/issues/31 | [
"question",
"answered"
] | gregsifr | 7 |
errbotio/errbot | automation | 986 | PySide not Working with Python 3.5 | In order to let us help you better, please fill out the following fields as best you can:
### I am...

* [ ] Reporting a bug
* [ ] Suggesting a new feature
* [ ] Requesti... | closed | 2017-04-04T21:06:50Z | 2021-07-23T05:22:27Z | https://github.com/errbotio/errbot/issues/986 | [
"type: bug",
"backend: Common",
"backend: GUI"
] | gyleodhis | 3 |
serengil/deepface | machine-learning | 828 | How to change max_threshold_to_verify | Currently, the default `max_threshold_to_verify` of `DeepFace.verify()` is 0.4, how do I change that value? | closed | 2023-08-20T07:24:10Z | 2023-08-20T07:29:13Z | https://github.com/serengil/deepface/issues/828 | [
"question"
] | brownfox2k6 | 2 |
junyanz/pytorch-CycleGAN-and-pix2pix | computer-vision | 1,575 | Issues with running repository on custom dataset | Hi, I am training the cycle Gan on a custom dataset that contains images of people wearing the jewellery and no jewellery. The images i train A and train B are of the same people with and without the jewellery. The fakes generated during the training, has people wearing the jewellery slowly. But when I use any of the t... | open | 2023-05-18T11:31:52Z | 2023-07-11T11:41:40Z | https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/issues/1575 | [] | vandana-sreenivasan3 | 1 |
kubeflow/katib | scikit-learn | 2,422 | SDK is broken when installed by `git+https` | ### What happened?
When I run the following code with `kubeflow-katib` installed by: `pip install git+https://github.com/kubeflow/katib.git@master#subdirectory=sdk/python/v1beta1`
```Python
import kubeflow.katib as katib
# Step 1. Create an objective function with push-based metrics collection.
def objective... | closed | 2024-09-05T06:30:06Z | 2024-09-05T16:09:17Z | https://github.com/kubeflow/katib/issues/2422 | [
"help wanted",
"good first issue",
"kind/bug",
"area/sdk",
"lifecycle/needs-triage"
] | Electronic-Waste | 4 |
WZMIAOMIAO/deep-learning-for-image-processing | pytorch | 642 | 关于HRNet中目标检测的json文件加载之后会报错的问题 | 环境配置是严格按照readme里面的,用GT是没有问题的,我想大概不是环境的事
请问一下大佬,当我把GT信息换成所提供的目标检测的json文件之后,无论是train.py还是validation.py在运行的过程中都会出现报错。
下面我描述一下validation.py中我遇到的情况
其中我注意到,validation的代码中,您好像忘了将args.person_det参数加进去了,因此我首先修改了
val_dataset = CocoKeypoint(data_root, "val", transforms=data_transform["val"], det_json_path=args.person_det... | closed | 2022-09-19T09:20:23Z | 2022-09-20T15:40:29Z | https://github.com/WZMIAOMIAO/deep-learning-for-image-processing/issues/642 | [] | Caicaizi-cdy | 4 |
feature-engine/feature_engine | scikit-learn | 449 | add inverse_transform method and functionality to BoxCoxTransformer | We can use the inv_boxcox functionality from scipy.special: https://docs.scipy.org/doc/scipy-1.8.0/html-scipyorg/reference/generated/scipy.special.inv_boxcox.html | closed | 2022-05-10T08:51:52Z | 2022-06-12T10:28:28Z | https://github.com/feature-engine/feature_engine/issues/449 | [
"good first issue",
"enhancement",
"easy"
] | solegalli | 3 |
python-restx/flask-restx | api | 176 | flask_restx is not compatible with gunicorn | I am trying to use gunicorn to run the flask API that I just developed. It worked well when I use flask_restplus, but it had No module named 'flask_restx' when using flask_restx even I have flask_restx installed in my virtual environment. I think it might because gunicorn does not support flask_restx. Anyone had this i... | closed | 2020-07-20T12:46:09Z | 2022-01-09T23:02:41Z | https://github.com/python-restx/flask-restx/issues/176 | [
"bug"
] | Serena-Xu | 1 |
abhiTronix/vidgear | dash | 159 | processed frames web streaming | Hey,
First of all thank for great repo.
I am connecting to an IP camera via VidGear, grab frame by frame, processed them on my local computer(e.g object detection) and want to stream that frames in real-time to AWS EC2 computer in order that my clients(react's app) will can see the processed frame's(e.g frame with bo... | closed | 2020-09-09T09:24:31Z | 2020-09-10T15:38:05Z | https://github.com/abhiTronix/vidgear/issues/159 | [
"QUESTION :question:",
"SOLVED :checkered_flag:"
] | idanmosh | 5 |
RomelTorres/alpha_vantage | pandas | 109 | Forex and pandas | Hi Romel,
Congrats for the module, it is really helpful and user friendly.
I would like to point out that the info in the README about
> Foreign Exchange (FX)
needs to be fixed a little bit because it is written that
> The foreign exchange is just metadata, thus only available as json format (using th... | closed | 2019-01-13T15:45:47Z | 2019-02-11T21:57:59Z | https://github.com/RomelTorres/alpha_vantage/issues/109 | [] | NTavou | 4 |
ultralytics/ultralytics | python | 19,438 | After train and export the model with ultralytics, can it be deployed lightweightly? | ### Search before asking
- [x] I have searched the Ultralytics [issues](https://github.com/ultralytics/ultralytics/issues) and found no similar feature requests.
### Description
First of all, this repo is very convenient for training and exporting models, because it takes into account many frameworks on the market,... | open | 2025-02-26T06:34:32Z | 2025-02-27T16:33:30Z | https://github.com/ultralytics/ultralytics/issues/19438 | [
"enhancement",
"dependencies"
] | qianyue76 | 5 |
plotly/dash-table | dash | 836 | When some rows have dropdown other rows are not editable | If I use dropdowns in a dash table with Editable = True, I cannot edit values in the other rows. This has been reported by several people, but I have not found a fix yet.
Some people say that it used to work, but not any longer. Any ideas? | open | 2020-10-06T19:30:30Z | 2023-11-22T02:56:12Z | https://github.com/plotly/dash-table/issues/836 | [] | Bastituta | 6 |
scrapy/scrapy | web-scraping | 6,378 | Edit Contributing.rst document to specify how to propose documentation suggestions | There are multiple types of contributions that the community can suggest including bug reports, feature requests, code improvements, security vulnerability reports, and documentation changes.
For the Scrapy.py project it was difficult to discern what process to follow to make a documentation improvement suggestion.... | closed | 2024-05-26T15:43:40Z | 2024-07-10T07:37:32Z | https://github.com/scrapy/scrapy/issues/6378 | [] | jtoallen | 11 |
tensorflow/tensor2tensor | machine-learning | 1,142 | Can one register callback functions for certain events (e.g. after eval)? | In `T2TModel` there is
```python
# Replace the two methods below in order to add custom SessionRunHooks to
# the training procedure.
@staticmethod
def train_hooks():
return []
@staticmethod
def eval_hooks():
return []
```
However, the problem is that these hooks are already living in a session.
... | closed | 2018-10-15T08:46:36Z | 2018-10-23T11:00:08Z | https://github.com/tensorflow/tensor2tensor/issues/1142 | [] | stefan-falk | 1 |
sczhou/CodeFormer | pytorch | 71 | How long does it take for traning stage one? | Hi, I am try to ReIP. the training code. I am wondering how long does it take for training stage one.
Best | open | 2022-11-24T11:05:23Z | 2022-11-24T11:05:23Z | https://github.com/sczhou/CodeFormer/issues/71 | [] | henanjun | 0 |
serengil/deepface | machine-learning | 774 | how to analyze age and gender in real time ? | how to analyze age and gender in real time ? | closed | 2023-06-10T19:36:32Z | 2023-06-11T10:01:11Z | https://github.com/serengil/deepface/issues/774 | [
"question"
] | Rasantis | 1 |
tensorpack/tensorpack | tensorflow | 1,066 | [FasterRCNN] Understanding clip_boxes in generate_fpn_proposals | I'm working with the Mask/Faster RCNN code and I'm confused about why the ROI proposals are being generated on a per-FPN-level basis, but the call to `generate_rpn_proposals` for each level passes in the same `image_shape2d`.
In the RPN code, you call `generate_fpn_proposals`. https://github.com/tensorpack/tensorpac... | closed | 2019-01-30T00:35:44Z | 2019-01-30T22:08:15Z | https://github.com/tensorpack/tensorpack/issues/1066 | [
"examples"
] | armandmcqueen | 4 |
python-visualization/folium | data-visualization | 1,687 | Click twice to exit full screen | **Describe the bug**
When I enter full screen mode, I need to click twice to exit full screen mode
| closed | 2022-12-30T06:35:32Z | 2023-10-17T08:33:17Z | https://github.com/python-visualization/folium/issues/1687 | [] | Winky678 | 4 |
jupyter-book/jupyter-book | jupyter | 1,810 | Cell error traceback incorrectly lexed as IPython | ### Describe the bug
I'm using Jupyter Book to produce a book with executable OCaml cells, including demoing some OCaml code that deliberately does not compile. I tag those cells with raises-exception of course. Some (not all) OCaml compiler error messages cause Sphinx to produce a lexer warning, Could not lex literal... | open | 2022-08-17T01:11:24Z | 2023-12-29T15:17:33Z | https://github.com/jupyter-book/jupyter-book/issues/1810 | [
"bug"
] | clarksmr | 2 |
numba/numba | numpy | 9,722 | optional type in `if x is not None` branch is still optional | <!--
Thanks for opening an issue! To help the Numba team handle your information
efficiently, please first ensure that there is no other issue present that
already describes the issue you have
(search at https://github.com/numba/numba/issues?&q=is%3Aissue).
-->
## Reporting a bug
<!--
Before submittin... | open | 2024-09-12T03:04:13Z | 2024-11-26T05:51:18Z | https://github.com/numba/numba/issues/9722 | [
"bug - typing"
] | auderson | 5 |
CorentinJ/Real-Time-Voice-Cloning | pytorch | 598 | Someone is selling this software!? | Hi,
Just found this: https://realtimevoicecloning.com
Is he selling your work as his own? | closed | 2020-11-19T02:56:36Z | 2020-12-05T08:32:15Z | https://github.com/CorentinJ/Real-Time-Voice-Cloning/issues/598 | [] | bostjan39 | 5 |
JoeanAmier/XHS-Downloader | api | 118 | 小红书下载主页视频时经常卡住不动 | 问题如下:
使用小红书下载功能批量下载主页内容时经常卡住进度不动,要么一就绪,要么就下载几十条的时候一直不动,下不了,重新停止后再开始下载又能下,过一会又会中断,如前面一样,希望能优化下,还有新版本下载时会出现签名服务器断联等问题,使用环境 windows 11。 | open | 2024-07-16T02:17:33Z | 2024-07-16T02:17:33Z | https://github.com/JoeanAmier/XHS-Downloader/issues/118 | [] | mizi6654 | 0 |
aimhubio/aim | data-visualization | 3,175 | AIM Client Process Termination Leaves Run in Active State | ## 🐛 Bug
When the AIM client process is killed, the corresponding run remains in the "In Progress" state indefinitely. I expect the run to transition to the "Finished" state upon client termination.
### To reproduce
1. Create a Python script `test-aim.py` with the following content:
```
import time
from ai... | open | 2024-06-25T07:28:48Z | 2025-01-10T14:59:06Z | https://github.com/aimhubio/aim/issues/3175 | [
"type / bug",
"help wanted"
] | zhiyxu | 8 |
httpie/cli | api | 975 | Support some way to specify an header that shouldn't be persisted to the session | Although not the most common, some services sometimes have HTTP headers that should be sent only for some specific requests, but not all of them, and as such shouldn't be persisted in the session. httpie currently persists all headers except for the ones starting with `Content-` or `If-`. That means such headers will a... | open | 2020-10-19T18:04:48Z | 2020-12-21T15:33:44Z | https://github.com/httpie/cli/issues/975 | [
"enhancement"
] | segevfiner | 0 |
jazzband/django-oauth-toolkit | django | 1,140 | How to skip oauth2_views.AuthorizationView's LoginRequiredMixin | I want to use Authorization code grant type, but when I invoke /o/authorize/, it redirects to admin's login.
How can I skip admin login?
| closed | 2022-04-06T13:23:36Z | 2023-10-04T14:42:33Z | https://github.com/jazzband/django-oauth-toolkit/issues/1140 | [
"question"
] | luohaoGit | 3 |
freqtrade/freqtrade | python | 10,558 | Freqtrade no longer working with Mexc due to recent API changes | OS Ubuntu 22.04.4
CCXT 4.3.79
Freqtrade docker-2024.8-dev-4ca6e617
Freqtrade has not been working with Mexc for a number of days now due to changes to their API
[<https://mexcdevelop.github.io/apidocs/spot_v3_en/#change-log>](url)
The change on 16/08/2024 is what seems to have broken it. | closed | 2024-08-19T12:00:33Z | 2024-08-21T06:38:21Z | https://github.com/freqtrade/freqtrade/issues/10558 | [
"Wont fix / Not a bug",
"CCXT",
"unsupported exchange"
] | rmtucker | 3 |
graphistry/pygraphistry | pandas | 522 | [BUG] chain has excess edges | **Describe the bug**
In the new chain tutorial, we get excess edges during:
```
g2.chain([
n({'community_infomap': 2}),
e_undirected(),
n({'community_infomap': 2})
]).plot()
```
While the nodes are right, the edges have excess, and the backend will materialize a bunch of nodes we don't actual... | closed | 2023-12-04T06:51:48Z | 2023-12-23T00:21:37Z | https://github.com/graphistry/pygraphistry/issues/522 | [
"bug",
"p3"
] | lmeyerov | 1 |
influxdata/influxdb-client-python | jupyter | 119 | Creating buckets using BucketsApi | On using BucketsApi from influxdb_client, if the org_id is not passed, the api updates the org_id with the org field from influxdb_client (which is a name). Ref:
https://github.com/influxdata/influxdb-client-python/blob/91ffed11c270c295a93b3e7d0b94a69b4657a917/influxdb_client/client/bucket_api.py#L39
A possible op... | closed | 2020-06-29T03:13:29Z | 2020-07-20T05:21:04Z | https://github.com/influxdata/influxdb-client-python/issues/119 | [
"question"
] | MajorCarrot | 3 |
jowilf/starlette-admin | sqlalchemy | 267 | Bug: | **Describe the bug**
I try to deploy my FastAPI project where I use starlette_admin as an admin panel. My host is based on a HTTPS secured domain, but starlette admin uses static files and scripts through HTTP and it cannot load on my website.
**To Reproduce**
Try to access admin panel on a HTTPS secured domain
... | closed | 2023-08-24T19:47:42Z | 2023-08-24T19:58:25Z | https://github.com/jowilf/starlette-admin/issues/267 | [
"bug"
] | xorwise | 3 |
521xueweihan/HelloGitHub | python | 2,630 | 【开源自荐】TestAgent:国内首个测试行业大模型工具 | ## 推荐项目
<!-- 这里是 HelloGitHub 月刊推荐项目的入口,欢迎自荐和推荐开源项目,唯一要求:请按照下面的提示介绍项目。-->
<!-- 点击上方 “Preview” 立刻查看提交的内容 -->
<!--仅收录 GitHub 上的开源项目,请填写 GitHub 的项目地址-->
- 项目地址:https://github.com/codefuse-ai/Test-Agent
<!--请从中选择(C、C#、C++、CSS、Go、Java、JS、Kotlin、Objective-C、PHP、Python、Ruby、Rust、Swift、其它、书籍、机器学习)-->
- 类别:机器学习... | open | 2023-10-25T11:40:58Z | 2023-11-26T03:20:27Z | https://github.com/521xueweihan/HelloGitHub/issues/2630 | [
"机器学习"
] | hailianzhl | 0 |
zihangdai/xlnet | tensorflow | 255 | Docker support | #Feature Request
Docker support is needed. | open | 2020-01-01T14:53:09Z | 2020-01-01T14:53:09Z | https://github.com/zihangdai/xlnet/issues/255 | [] | sanjibnarzary | 0 |
AUTOMATIC1111/stable-diffusion-webui | deep-learning | 15,398 | [Bug]: inpaint zoom is broken on firefox | ### Checklist
- [X] The issue exists after disabling all extensions
- [X] The issue exists on a clean installation of webui
- [X] The issue is caused by an extension, but I believe it is caused by a bug in the webui
- [X] The issue exists in the current version of the webui
- [X] The issue has not been reported before... | open | 2024-03-27T22:37:07Z | 2024-03-31T15:21:20Z | https://github.com/AUTOMATIC1111/stable-diffusion-webui/issues/15398 | [
"bug-report"
] | BismutoGanymedes | 2 |
Netflix/metaflow | data-science | 1,812 | Update extras_require for tracing dependencies? | From the [2.10.4 release notes](https://github.com/Netflix/metaflow/releases/tag/2.10.4):
> Some additional dependencies are required for the tracing functionality in the execution environment. These can be installed in the base Docker image, or supplied through a conda environment. The relevant packages are
> `op... | open | 2024-04-25T00:44:58Z | 2024-04-25T00:44:58Z | https://github.com/Netflix/metaflow/issues/1812 | [] | chriselion | 0 |
davidsandberg/facenet | computer-vision | 1,252 | Question : Anybody has luck converting to coreml ? | Anybody has luck converting to coreml ?
Thanks | open | 2024-06-18T02:38:59Z | 2024-06-18T02:38:59Z | https://github.com/davidsandberg/facenet/issues/1252 | [] | x4080 | 0 |
wkentaro/labelme | computer-vision | 421 | [Suggestion] Add support for import existing annotation | Hello, i really appreciate your work, the software really helps me a lot.
There's some suggestions:
1. Add support for **import existing annotation**.
In some cases annotations are firstly generated by beginners or primary algorthms, then inspected or corrected by the experienced. I work for medical image processi... | closed | 2019-06-20T02:46:21Z | 2020-01-27T01:34:40Z | https://github.com/wkentaro/labelme/issues/421 | [] | coffeehat | 3 |
microsoft/nni | data-science | 5,740 | dispather comand:globals.args.pythonInterpreter, '-m', 'nni', '--exp_params', 占用内存太多 | **Describe the issue**:

**Environment**:
- NNI version: 2.9
- Training service (local|remote|pai|aml|etc): local
- Client OS:
- Server OS (for remote mode only):
- Python version:
- PyTorch/TensorFlow version:... | closed | 2024-01-25T03:42:39Z | 2024-01-26T01:51:23Z | https://github.com/microsoft/nni/issues/5740 | [] | yjjinjie | 1 |
huggingface/datasets | deep-learning | 7,458 | Loading the `laion/filtered-wit` dataset in streaming mode fails on v3.4.0 | ### Describe the bug
Loading https://huggingface.co/datasets/laion/filtered-wit in streaming mode fails after update to `datasets==3.4.0`. The dataset loads fine on v3.3.2.
### Steps to reproduce the bug
Steps to reproduce:
```
pip install datastes==3.4.0
python -c "from datasets import load_dataset; load_dataset('l... | closed | 2025-03-17T14:54:02Z | 2025-03-17T16:02:04Z | https://github.com/huggingface/datasets/issues/7458 | [] | nikita-savelyevv | 1 |
elliotgao2/gain | asyncio | 19 | Add homepage. | Add homepage. | closed | 2017-06-09T01:22:40Z | 2017-06-12T01:47:51Z | https://github.com/elliotgao2/gain/issues/19 | [] | elliotgao2 | 1 |
allure-framework/allure-python | pytest | 743 | Fix historyId to be dependent on dynamic allure parameters | #### I'm submitting a ...
- [x] bug report
- [ ] feature request
- [ ] support request => Please do not submit support request here, see note at the top of this template.
#### What is the current behavior?
Parameters added via the `allure.dynamic.parameter` inside a test body don't affect allure history o... | closed | 2023-04-19T10:04:13Z | 2023-04-26T09:02:03Z | https://github.com/allure-framework/allure-python/issues/743 | [
"bug",
"theme:pytest",
"contribute"
] | delatrie | 1 |
FlareSolverr/FlareSolverr | api | 224 | close suddently windows 10 | **Please use the search bar** at the top of the page and make sure you are not creating an already submitted issue.
Check closed issues as well, because your issue may have already been fixed.
### How to enable debug and html traces
[Follow the instructions from this wiki page](https://github.com/FlareSolverr/Fl... | closed | 2021-11-19T18:08:50Z | 2021-12-12T16:13:04Z | https://github.com/FlareSolverr/FlareSolverr/issues/224 | [
"more information needed"
] | 3xploiton3 | 10 |
NullArray/AutoSploit | automation | 901 | Divided by zero exception277 | Error: Attempted to divide by zero.277 | closed | 2019-04-19T16:03:08Z | 2019-04-19T16:37:01Z | https://github.com/NullArray/AutoSploit/issues/901 | [] | AutosploitReporter | 0 |
deeppavlov/DeepPavlov | tensorflow | 927 | Using ranking module with pretreined bert for prediction | How can I use pretrained on ubuntu dataset bert (representation based) model from ranking module in the following scenario: I have list of possible responses (in file) and I would like to load pretrained model in python and get ordering of possible responses for given context? My goal is to define function rank(possibl... | closed | 2019-07-15T09:27:38Z | 2020-05-13T09:44:28Z | https://github.com/deeppavlov/DeepPavlov/issues/927 | [] | norbertryc | 1 |
microsoft/qlib | deep-learning | 1,713 | how to convert 3-seconds market data to qlib bin? | I have my own 3-seconds market data in local mysql and want to conver these 3-sec data to qlib bin,
How do I convert?
I used to convert csv (3 seconds data) to qlib bin by using dump_bin.py, the result of bin files like this:
: finished with status 'error' | ### Checklist
- [ ] The issue exists after disabling all extensions
- [ ] The issue exists on a clean installation of webui
- [ ] The issue is caused by an extension, but I believe it is caused by a bug in the webui
- [ ] The issue exists in the current version of the webui
- [ ] The issue has not been reported before... | open | 2024-04-01T14:08:08Z | 2024-04-01T14:08:08Z | https://github.com/AUTOMATIC1111/stable-diffusion-webui/issues/15426 | [
"bug-report"
] | zutim | 0 |
python-gino/gino | asyncio | 15 | Delegate more asyncpg API | In #9 only `fetch` and `cursor` are delegated, we may still need to delegate `fetchrow` and `execute`. | closed | 2017-07-23T14:14:37Z | 2017-07-24T11:44:07Z | https://github.com/python-gino/gino/issues/15 | [
"help wanted",
"task"
] | fantix | 0 |
ultralytics/yolov5 | machine-learning | 12,768 | How to load models without pip install? | ### Search before asking
- [X] I have searched the YOLOv5 [issues](https://github.com/ultralytics/yolov5/issues) and [discussions](https://github.com/ultralytics/yolov5/discussions) and found no similar questions.
### Question
I need to execute this model on a different device as a pytorch model where I can'... | closed | 2024-02-27T02:47:48Z | 2024-04-08T02:16:25Z | https://github.com/ultralytics/yolov5/issues/12768 | [
"question",
"Stale"
] | useruser2023 | 4 |
JaidedAI/EasyOCR | deep-learning | 854 | easyOCR still uses GPU although not asked | Hi,
When I create the `reader` object with `gpu=False`, it doesnt directly use the GPU device, but upon calling `readtext`, it captures the first available GPU device apparently uses it according to `nvidia-smi`. I verified that its not `detect()` that triggers the GPU use so its probably the recognizer pipeline. I ... | open | 2022-09-15T13:42:32Z | 2022-09-29T13:31:02Z | https://github.com/JaidedAI/EasyOCR/issues/854 | [] | ozancaglayan | 4 |
abhiTronix/vidgear | dash | 254 | Release of Vidgear v0.2.3 | <!--- Add a brief but descriptive title for your issue above -->
# Release of Vidgear v0.2.3
## Question
<!--- Provide your question description here -->
When will this new version containing some hotfixes be released? I need version 0.2.3 to complete what I am working on currently. I can always help with the r... | closed | 2021-10-19T16:26:48Z | 2021-10-27T04:00:06Z | https://github.com/abhiTronix/vidgear/issues/254 | [
"QUESTION :question:",
"SOLVED :checkered_flag:",
"NEW RELEASE :fire:"
] | Vboivin | 3 |
Miksus/rocketry | pydantic | 31 | why do we need red engine? | What its advantages over Airflow and other mature schedule framework are? | closed | 2022-07-04T12:30:43Z | 2022-07-04T19:55:24Z | https://github.com/Miksus/rocketry/issues/31 | [] | lidh15 | 1 |
ivy-llc/ivy | numpy | 28,607 | Fix Frontend Failing Test: tensorflow - operators.jax.lax.real | closed | 2024-03-14T22:09:57Z | 2024-03-16T12:27:15Z | https://github.com/ivy-llc/ivy/issues/28607 | [
"Sub Task"
] | samthakur587 | 0 | |
google/trax | numpy | 848 | PyTorch backend | I notice that so far there are JAX, TensorFlow, and NumPy backends. Are there any plans to have a PyTorch backend in the future? | open | 2020-07-17T14:43:40Z | 2020-10-14T07:15:40Z | https://github.com/google/trax/issues/848 | [
"enhancement"
] | briankosw | 2 |
jupyterhub/zero-to-jupyterhub-k8s | jupyter | 3,623 | Ingress with a subpath has a hard-coded trailing slash | <!-- Thank you for contributing. These HTML comments will not render in the issue, but you can delete them once you've read them if you prefer! -->
### Bug description
I am trying to deploy JH to a subpath, e.g. `<mydomain>/jupyter`. I set `hub.baseUrl=/jupyter` and enable ingress, but the `Ingress` object is not cor... | open | 2025-02-14T16:28:51Z | 2025-02-14T16:28:51Z | https://github.com/jupyterhub/zero-to-jupyterhub-k8s/issues/3623 | [
"bug"
] | johnflavin | 0 |
marimo-team/marimo | data-science | 3,828 | Programs containing marimo cannot be frozen anymore | ### Describe the bug
When freezing a program containing marimo, it crashes during startup/module loading.
I tried cx-freeze and pyinstaller, and it worked on version 0.10.6 and breaks starting with 0.10.7.
To freeze it, I'm using fastapi and uvicorn, and it worked beautifully to deploy. (side note: marimo is great!)
... | closed | 2025-02-18T09:47:23Z | 2025-02-18T18:29:23Z | https://github.com/marimo-team/marimo/issues/3828 | [
"bug"
] | ABChristian | 3 |
dynaconf/dynaconf | django | 403 | [bug] Cannot get secrets from vault | **Describe the bug**
When trying to retrieve secrets from hashicorp vault without setting os envs the settings only contain the default dynaconf settings.
**To Reproduce**
Steps to reproduce the behavior:
1. Run local vault server as described in the documentation
2. Store some secrets in vault
3.Run this... | closed | 2020-09-02T11:17:43Z | 2020-10-23T18:57:28Z | https://github.com/dynaconf/dynaconf/issues/403 | [
"hacktoberfest",
"Docs"
] | clanzett | 4 |
open-mmlab/mmdetection | pytorch | 11,572 | the same image, the same model, after deployed with c++, sometimes has result, sometimes has nothings,why? | the same image, the same model, when deploy with c++, sometimes has result, sometimes has nothings,why?
the error is: "异常 dets.size() == 0".
code is below:
#include "mmdeploy/detector.hpp"
#include "opencv2/imgcodecs/imgcodecs.hpp"
#include "opencv2/core/utility.hpp"
#include "utilsisualize.h"
#include
... | open | 2024-03-20T05:18:04Z | 2024-03-20T05:18:04Z | https://github.com/open-mmlab/mmdetection/issues/11572 | [] | happybear1015 | 0 |
streamlit/streamlit | streamlit | 10,514 | toml.decoder.TomlDecodeError: Key name found without value. Reached end of line. | ### Summary
Hello, I created Google OpenID Connect client and tried to implement it on streamlit but the following error occured
```
Traceback (most recent call last):
File "<frozen runpy>", line 198, in _run_module_as_main
File "<frozen runpy>", line 88, in _run_code
File "/home/ugroon/.local/lib/python3.12/si... | closed | 2025-02-19T11:56:06Z | 2025-02-25T18:24:57Z | https://github.com/streamlit/streamlit/issues/10514 | [] | ug0x01 | 3 |
httpie/cli | rest-api | 1,126 | Session file with cookie cannot be parsed | **Checklist**
- [Y] I've searched for similar issues.
- [Y] I'm using the the latest version of HTTPie.
- httpie version, 2.4.0
- python version, 3.9
I prepared the **my-session-cookie.json** file, the request parameters are placed in **query.json**, and the expected response results are placed in the **resu... | closed | 2021-08-12T18:24:04Z | 2021-08-17T14:05:30Z | https://github.com/httpie/cli/issues/1126 | [
"bug",
"new"
] | caofanCPU | 6 |
ymcui/Chinese-LLaMA-Alpaca | nlp | 502 | Chinese Plus Alpaca 继续精调疑问 | 我想继续训练Chinese Plus Alpaca
--model_name_or_path: 合并Chinese-LLaMA-Plus-LoRA后的Chinese-LLaMA模型
问题1: --peft_path: 这里是使用Chinese-Alpaca 还是 Chinese-Plus-Alpaca 的权重目录?
问题2: 继续精调是只用提供新的数据的 json,微调后就能得到原Chinese-Plus-Alpaca + 新数据合并的效果吗?
问题3: 训练完后得到的 New-Alpaca-Lora 合并时是使用 Chinese-LLaMA + Chinese-LLaMA-Plus-LoRA + Chinese-Al... | closed | 2023-06-03T04:17:11Z | 2023-06-05T05:13:28Z | https://github.com/ymcui/Chinese-LLaMA-Alpaca/issues/502 | [] | Damonproto | 3 |
plotly/dash-bio | dash | 421 | AlignmentChart Component Improvements | The AlignmentChart component currently lacks a few features which are relevant to ease of use in creating app layouts and in specifying which data is displayed.
* One issue is the lack of dynamic auto-resizing to fit a container based on the size of the display. Although the `height` and `width` of the chart can be... | closed | 2019-09-25T15:50:44Z | 2021-10-27T01:44:00Z | https://github.com/plotly/dash-bio/issues/421 | [
"nice-to-have"
] | HammadTheOne | 2 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.