repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
holoviz/panel | matplotlib | 7,292 | "Debugging in VS Code" Documentation insufficient? | #### ALL software version info
```plaintext
panel==1.5.0
VS Code Version: 1.93.1 (Universal)
```
#### Description of expected behavior and the observed behavior
When adding a VS Code debugging configuration as suggested in the documentation, I expect to see Panel site variables in the debugging pane (ev... | closed | 2024-09-18T05:16:34Z | 2024-09-18T07:25:30Z | https://github.com/holoviz/panel/issues/7292 | [] | michaelweinold | 3 |
kubeflow/katib | scikit-learn | 1,744 | [Proposal] Support JSON format for `file-metrics-collector` | /kind feature
Describe the solution you'd like
[A clear and concise description of what you want to happen.]
## Motivation
Currently, it is difficult to parse JSON format files by `file-metrics-collector` using regexp filter since `file-metrics-collector` is designed to use TEXT format files.
I believe if `f... | closed | 2021-11-25T13:46:43Z | 2022-04-05T15:20:37Z | https://github.com/kubeflow/katib/issues/1744 | [
"kind/feature"
] | tenzen-y | 15 |
PrefectHQ/prefect | automation | 17,129 | `DeploymentScheduleUpdate.active` server api does not match the client api and cause an InternalError on the API | ### Bug summary
With the new prefect version 3.2.0, existing deployments cannot be updated since the client API for a `DeploymentScheduleUpdate` object does not match the `DeploymentScheduleUpdate` server API for the field `active`. This causes a prefect internal server failure when applying Deployments client side wh... | closed | 2025-02-13T17:37:26Z | 2025-02-13T17:52:07Z | https://github.com/PrefectHQ/prefect/issues/17129 | [
"bug"
] | marcm-ml | 1 |
neuml/txtai | nlp | 674 | Add support for dynamic vector dimensions | Add support for dynamic vector dimensions. This enables using models trained with [Matryoshka Representation Learning](https://arxiv.org/pdf/2205.13147.pdf). Support for this method was added with Sentence Transformers 2.4.
See [this blog post](https://huggingface.co/blog/matryoshka) for more. | closed | 2024-02-24T11:20:28Z | 2024-02-28T02:22:16Z | https://github.com/neuml/txtai/issues/674 | [] | davidmezzetti | 0 |
proplot-dev/proplot | matplotlib | 137 | Would you add the "readshapefile" method in proplot? | <!-- Thanks for helping us make proplot a better package! If this is a bug report, please use the template provided below. If this is a feature request, you can delete the template text (just try to be descriptive with your request). -->
### Description
[Description of the bug or feature.]
### Steps to reprodu... | closed | 2020-04-07T04:14:20Z | 2020-04-22T23:15:45Z | https://github.com/proplot-dev/proplot/issues/137 | [
"feature"
] | sfhua | 2 |
plotly/dash-table | plotly | 829 | Dash DataTable with RadioButton as cell content | I’m trying to create a table containing players, games and a set of configurations that can be toggled on/off.
The ideal would be to use a Dash DataTable with RadioButtons (similar as for DropDowns). Is this possible? Any ideas on how to achieve this? Screenshot 2020-09-16 at 23.27.41
![Screenshot 2020-09-16 at 23 ... | open | 2020-09-18T22:11:20Z | 2020-09-18T22:11:20Z | https://github.com/plotly/dash-table/issues/829 | [] | TomRoger | 0 |
opengeos/leafmap | plotly | 885 | Add support for selecting/editing multiple features | Selecting and highlight multiple features
```python
import os
import json
import requests
import copy
from ipyleaflet import Map, GeoJSON, LayersControl
def create_geojson_map(data, style, hover_style, highlight_style, center=(50.6252978589571, 0.34580993652344), zoom=4):
"""
Create a GeoJSON map w... | closed | 2024-09-06T03:23:44Z | 2024-09-11T03:01:50Z | https://github.com/opengeos/leafmap/issues/885 | [
"Feature Request"
] | giswqs | 3 |
paperless-ngx/paperless-ngx | django | 9,141 | [BUG] Concise description of the issue | ### Description
hello,
your script does not work. It says that "docker compose" does not exist. It is not calling "docker-compose", it is calling "docker compose". Tried different ways to get paperless up and running but was not able to get it done. Very frustrating.
Regards
Thomas
### Steps to reproduce
run the s... | closed | 2025-02-17T18:19:08Z | 2025-03-20T03:12:52Z | https://github.com/paperless-ngx/paperless-ngx/issues/9141 | [
"not a bug"
] | higgyforever | 8 |
xonsh/xonsh | data-science | 5,071 | Suppress subprocess traceback in case `XONSH_SHOW_TRACEBACK=False` and `$RAISE_SUBPROC_ERROR=True` | Source - https://github.com/xonsh/xonsh/discussions/4708
```xsh
echo @("""
$XONSH_SHOW_TRACEBACK = False
$RAISE_SUBPROC_ERROR = True
print('LINE 1')
cp nonexisting_file.txt other_name.txt
print("LINE 2")
""") > /tmp/1.xsh
xonsh /tmp/1.xsh
#
# CURRENT OUTPUT: full traceback
#
#
# EXPECTED OUTPUT - ... | closed | 2023-02-24T07:48:09Z | 2023-03-08T07:12:35Z | https://github.com/xonsh/xonsh/issues/5071 | [
"error",
"priority-medium"
] | anki-code | 6 |
pydantic/FastUI | pydantic | 337 | disable/enable/hide form elements dependent on input in another element | Hi,
Thanks for this great tool.
We often have forms where some questions do not make sense if a question was answered in a specific way before. For example, on a feedback form to an invitation, we ask if the invitee will attend the event (a required `bool` field), and if they attend, we ask how many guests they wil... | open | 2024-06-27T19:08:33Z | 2024-06-27T19:08:33Z | https://github.com/pydantic/FastUI/issues/337 | [] | PHvL | 0 |
vitalik/django-ninja | django | 1,180 | Static Method Not Reflecting Instance-Specific Argument in Dynamically Created ModelSchema Class | **Description:**
I encountered an issue with dynamically creating schema classes where a static method within the class does not reflect an instance-specific argument. Below is the code snippet illustrating the problem:
```Python
class get_some_schema(a=False):
class SomeSchema(ModelSchema):
class Me... | open | 2024-05-31T10:46:43Z | 2024-05-31T10:46:43Z | https://github.com/vitalik/django-ninja/issues/1180 | [] | Alex-Sichkar | 0 |
huggingface/datasets | numpy | 7,287 | Support for identifier-based automated split construction | ### Feature request
As far as I understand, automated construction of splits for hub datasets is currently based on either file names or directory structure ([as described here](https://huggingface.co/docs/datasets/en/repository_structure))
It would seem to be pretty useful to also allow splits to be based on ide... | open | 2024-11-10T07:45:19Z | 2024-11-19T14:37:02Z | https://github.com/huggingface/datasets/issues/7287 | [
"enhancement"
] | alex-hh | 3 |
ultralytics/yolov5 | machine-learning | 13,476 | mAP50计算与iou_thres的关系 | ### Search before asking
- [X] I have searched the YOLOv5 [issues](https://github.com/ultralytics/yolov5/issues) and [discussions](https://github.com/ultralytics/yolov5/discussions) and found no similar questions.
### Question
mAP50计算与iou_thres有什么关系,运行val.py文件时,改变iou_thres为0.5,精度上升了,召回率稍微下降了,这是为什么?

<ipython-input-50-1055f09e4b48> in <module>
----> 1 profile.to_widgets()
~\anaconda3\lib\site-packages\pandas_profiling\profile_report.py in to_widgets(self)
... | open | 2021-10-07T19:17:58Z | 2021-11-30T19:23:01Z | https://github.com/ydataai/ydata-profiling/issues/853 | [] | jijasmx | 7 |
huggingface/datasets | pytorch | 6,942 | Import sorting is disabled by flake8 noqa directive after switching to ruff linter | When we switched to `ruff` linter in PR:
- #5519
import sorting was disabled in all files containing the `# flake8: noqa` directive
- https://github.com/astral-sh/ruff/issues/11679
We should re-enable import sorting on those files. | closed | 2024-06-02T09:43:34Z | 2024-06-04T09:54:24Z | https://github.com/huggingface/datasets/issues/6942 | [
"maintenance"
] | albertvillanova | 0 |
indico/indico | sqlalchemy | 6,357 | Better UX when uploading files in the editing module | **Is your feature request related to a problem? Please describe.**
The dropzones for file uploads in the editing module are too small. At the same time it is not clear (at least to me, every time I try to upload a file) that the dropzone is actually just the area at the bottom with the dashed border and not the whol... | open | 2024-05-22T08:42:40Z | 2024-05-22T08:42:40Z | https://github.com/indico/indico/issues/6357 | [
"enhancement"
] | tomasr8 | 0 |
pyjanitor-devs/pyjanitor | pandas | 1,247 | Add `how='outer'` to `conditional_join` | closed | 2023-02-20T08:17:21Z | 2023-05-07T00:05:19Z | https://github.com/pyjanitor-devs/pyjanitor/issues/1247 | [] | samukweku | 0 | |
LAION-AI/Open-Assistant | python | 3,196 | Feature Request: Integration of Music Functionality in Open Assistant Project | As an active user of the Open Assistant project, I believe it would greatly enhance the user experience to include music functionality within the project. Music has become an integral part of our lives and can significantly contribute to a more enjoyable and immersive user interaction. Adding music capabilities would n... | closed | 2023-05-18T17:25:54Z | 2023-06-09T12:01:37Z | https://github.com/LAION-AI/Open-Assistant/issues/3196 | [
"feature",
"needs discussion"
] | CodeQueeninBuissness123 | 2 |
scikit-optimize/scikit-optimize | scikit-learn | 1,102 | Model is fed wrong values by `BayesSearchCV` | Hi there, this is my first time trying `scikit-optimize` and I have followed the [minimal example on your webpage](https://scikit-optimize.github.io/stable/auto_examples/sklearn-gridsearchcv-replacement.html), with the only modification of using `joblib` to distribute jobs to a dask cluster.
```python
from skopt im... | open | 2022-01-20T11:15:57Z | 2022-01-20T11:16:17Z | https://github.com/scikit-optimize/scikit-optimize/issues/1102 | [] | gcaria | 0 |
dunossauro/fastapi-do-zero | pydantic | 121 | `datetime.utcnow` está deprecado no python 3.12 | Como algumas pessoas estão fazendo o curso com a versão mais recente do python, essa chamada deve ser alterada para uma versão que seja compatível com 3.11 (o recomendado) com as versões mais recentes da linguagem.
Uma solução para manter a mesma funcionalidade, sem fazer a chamada de forma ingênua do datetime é cha... | closed | 2024-04-01T19:03:42Z | 2024-04-17T08:38:35Z | https://github.com/dunossauro/fastapi-do-zero/issues/121 | [] | dunossauro | 0 |
rougier/scientific-visualization-book | numpy | 47 | Minor typo on page 177 | This sentence "Filled contours with dropshadows is a nice effet that allows" has effect misspelled. | closed | 2022-02-24T17:23:09Z | 2022-08-08T14:48:28Z | https://github.com/rougier/scientific-visualization-book/issues/47 | [] | tjnd89 | 1 |
seleniumbase/SeleniumBase | pytest | 3,227 | driver.uc_gui_click_captcha() Broken | Since chrome version - Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/130.0.0.0 Safari/537.36.
**driver.uc_gui_click_captcha()** is broken. It is not clicking on the cloudflare turnstile captcha when accessing the website.
Tried below it was working earlier, but suddenly ... | closed | 2024-10-27T10:01:19Z | 2024-10-27T23:14:24Z | https://github.com/seleniumbase/SeleniumBase/issues/3227 | [
"can't reproduce",
"UC Mode / CDP Mode"
] | p-rk | 15 |
huggingface/transformers | pytorch | 36,224 | Incompatibility in flash_attention_2 + Llama + Transformers>=4.43 + Autocast to fp16 | ### System Info
setting: Inference or Training Llama with Automatic Mixed Precision (AMP) autocast from fp32 to fp16 + FlashAttention 2 (FA2).
I observed that in newer versions of the Transformers library (>=4.43), training (and inference) fails with the error `RuntimeError: FlashAttention only supports fp16 and bf16... | open | 2025-02-17T08:14:24Z | 2025-03-20T08:03:39Z | https://github.com/huggingface/transformers/issues/36224 | [
"bug"
] | poedator | 2 |
deepset-ai/haystack | pytorch | 8,026 | Sentence Window retrieval documentation | Add documentation for the new `SentenceWindowRetrieval` abstraction: https://github.com/deepset-ai/haystack/blob/0411cd938a8b1d4a7153b0c269c6cd11d7da2efd/haystack/components/retrievers/sentence_window_retrieval.py#L13
A suggestion to add under the advanced RAG techniques alongside Hyde. | closed | 2024-07-15T14:26:25Z | 2024-07-17T11:55:47Z | https://github.com/deepset-ai/haystack/issues/8026 | [
"type:documentation",
"P1"
] | mrm1001 | 2 |
quantumlib/Cirq | api | 6,329 | `synchronize_terminal_measurements()` misorders measurements with the same key | **Description of the issue**
(not 100% sure this isn't intended behavior)
when two measurements have the same key, reordering them results in a logically different circuit. Transformers like `align_left()` and `align_right()` correctly account for this by preventing the reordering of measurements with the same ke... | open | 2023-10-25T18:26:53Z | 2025-03-22T00:30:02Z | https://github.com/quantumlib/Cirq/issues/6329 | [
"kind/bug-report",
"triage/needs-feasibility",
"triage/needs-more-evidence"
] | richrines1 | 13 |
encode/databases | sqlalchemy | 495 | The documentation does not describe the return values for the database.execute (LAST_INSERT_ID) | ```
async with database.transaction() as transaction:
query = insert(Session).values({"foo": "bar"})
last_insert_id = await database.execute(query)
(uid, ) = await database.fetch_one("SELECT LAST_INSERT_ID() as id")
assert uid == last_inserted_id # True
```... | open | 2022-06-02T16:05:12Z | 2022-06-02T16:05:12Z | https://github.com/encode/databases/issues/495 | [] | AntonGsv | 0 |
MaartenGr/BERTopic | nlp | 1,574 | Modeling taking a long time after progress bar complete | I was surprised that my model was taking over 8 hours to run, so I set 'verbose = True' to monitor the progress. I was surprised to see that the progress bar completed within an hour, but the cell was left running for over several hours after (and still is). I've added a screenshot of my code and output for reference. ... | open | 2023-10-11T17:40:14Z | 2023-10-12T13:48:57Z | https://github.com/MaartenGr/BERTopic/issues/1574 | [] | vlawlor | 2 |
statsmodels/statsmodels | data-science | 8,764 | I did this code and this error occurs after running it | ```
import pandas as pd
import numpy as np
import statsmodels.api as sm
from statsmodels.tsa.arima.model import ARIMA
from sklearn.neural_network import MLPRegressor
from sklearn.model_selection import TimeSeriesSplit
from sklearn.base import BaseEstimator, RegressorMixin
import optuna
# Load your dataset fr... | open | 2023-04-02T01:00:27Z | 2023-04-02T01:09:10Z | https://github.com/statsmodels/statsmodels/issues/8764 | [] | akeller1992 | 0 |
Yorko/mlcourse.ai | numpy | 640 | Mistake in assignment 8 | There is no true formula for updating weights using gradient descent:

Right formula is:
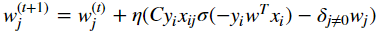
... | closed | 2019-10-30T22:11:54Z | 2019-11-04T14:09:57Z | https://github.com/Yorko/mlcourse.ai/issues/640 | [
"enhancement"
] | Ecclesiast | 1 |
KaiyangZhou/deep-person-reid | computer-vision | 252 | Pytorch to ONNX | Hellow,I‘m interested in your work, and I want to make a demo with your resnet50 model, but when I followed by the introduction, there will be a misktake. Could you please tell me the pretrained model's detail or if it is different with the nomal pytorch model?
Thank you very much!!!
my code as following:
... | closed | 2019-11-03T07:28:40Z | 2019-11-04T06:07:58Z | https://github.com/KaiyangZhou/deep-person-reid/issues/252 | [] | zhuyu-cs | 0 |
WZMIAOMIAO/deep-learning-for-image-processing | deep-learning | 226 | 大神 这边的tensorflow2内容有木有tensorflow1的对应的 ?比如rest 迁移 在tensorflow1的版本? 这差异太大啦 | **System information**
* Have I written custom code:
* OS Platform(e.g., window10 or Linux Ubuntu 16.04):
* Python version:
* Deep learning framework and version(e.g., Tensorflow2.1 or Pytorch1.3):
* Use GPU or not:
* CUDA/cuDNN version(if you use GPU):
* The network you trained(e.g., Resnet34 network):
**Des... | closed | 2021-04-15T02:53:02Z | 2021-04-16T10:10:13Z | https://github.com/WZMIAOMIAO/deep-learning-for-image-processing/issues/226 | [] | nanfangyuan | 1 |
home-assistant/core | asyncio | 141,117 | Device Location not working |

Hey there!
I have set up my companion app with Home Assistant, which is running on a Raspberry Pi 5. However, I'm experiencing issues with the "device tracker," as it always shows "away" even when I am at home.
I have checked t... | open | 2025-03-22T13:58:18Z | 2025-03-22T23:20:45Z | https://github.com/home-assistant/core/issues/141117 | [] | tommdq | 1 |
dunossauro/fastapi-do-zero | pydantic | 193 | FastAPI_Du_Zero |
Link do projeto | Seu @ no git | Comentário (opcional)
-- | -- | --
[FastAPI_Du_Zero](https://github.com/rodten23/FastAPI_Du_Zero) | [@rodten23](https://github.com/rodten23) | Implementação do material do curso sem alterações. Muito Obrigado, Dunossauro!
| closed | 2024-07-07T00:09:55Z | 2024-07-10T01:39:32Z | https://github.com/dunossauro/fastapi-do-zero/issues/193 | [] | rodten23 | 1 |
ultralytics/yolov5 | deep-learning | 12,846 | YOLOv5 interface - predict problem | ### Search before asking
- [X] I have searched the YOLOv5 [issues](https://github.com/ultralytics/yolov5/issues) and found no similar bug report.
### YOLOv5 Component
Other
### Bug
TypeError Traceback (most recent call last)
[<ipython-input-21-e91aa8dd130a>](https://loc... | closed | 2024-03-24T15:13:59Z | 2024-10-20T19:42:10Z | https://github.com/ultralytics/yolov5/issues/12846 | [
"bug",
"Stale"
] | paulikoe | 3 |
httpie/cli | api | 1,255 | Warn the user when there is no incoming data after a certain time passed on stdin | E.g
```
$ cat | http POST pie.dev/post 861ms
> no stdin data read in 10.0s (perhaps you want to --ignore-stdin)
> https://httpie.io/docs/cli/best-practices
``` | closed | 2021-12-30T15:06:40Z | 2022-01-12T14:07:34Z | https://github.com/httpie/cli/issues/1255 | [
"enhancement",
"new"
] | isidentical | 0 |
public-apis/public-apis | api | 4,158 | 1 | open | 2025-02-25T02:06:30Z | 2025-02-25T02:06:30Z | https://github.com/public-apis/public-apis/issues/4158 | [] | 2629728088 | 0 | |
CorentinJ/Real-Time-Voice-Cloning | python | 903 | Synthesizer fine-tuning ruined the output | We were trying to fine-tune the existing model on some of our own data. We were faced with this error.

We went into the synthesizer/train.py file and commented out the line 192(which ... | closed | 2021-11-23T17:24:05Z | 2021-11-24T04:23:43Z | https://github.com/CorentinJ/Real-Time-Voice-Cloning/issues/903 | [] | Fzr2k | 2 |
huggingface/transformers | tensorflow | 36,537 | Bug when computing positional IDs from embeddings | ### System Info
https://github.com/huggingface/transformers/blob/c0f8d055ce7a218e041e20a06946bf0baa8a7d6a/src/transformers/models/esm/modeling_esm.py#L243
I think it should be `1, sequence_length + 1, dtype=torch.long, device=inputs_embeds.device`
I don't see how the index of my padding token in the alphabet has anyt... | open | 2025-03-04T14:01:31Z | 2025-03-04T14:27:26Z | https://github.com/huggingface/transformers/issues/36537 | [
"bug"
] | SabrinaRichter | 1 |
vitalik/django-ninja | django | 947 | Management command to output schema json | We're starting to use the openapi client generators. It's not too hard to just `curl` the json file in local dev, but for CI it would be slightly annoying to have to do.
It would be nice if there was a clean `python manage.py ninja-schema` that would output the json schema without having to start up gunicorn or the ... | closed | 2023-11-22T02:17:09Z | 2023-11-22T18:45:01Z | https://github.com/vitalik/django-ninja/issues/947 | [
"documentation"
] | shughes-uk | 3 |
jessevig/bertviz | nlp | 58 | no attribute 'bias' while loading a finetuned BERT from TF | Hi!
I have a finetuned BERT model trained in Tensorflow and I would like to visualize its attentions.
I tried to load the model in different ways, but I always get the same error. Any suggestion at how to solve this no attribute bias error?
```
model = BertModel.from_pretrained(model_path, from_tf=True, output_... | closed | 2020-10-29T14:47:17Z | 2021-01-31T14:41:48Z | https://github.com/jessevig/bertviz/issues/58 | [] | GorkaUrbizu | 2 |
glumpy/glumpy | numpy | 285 | Installing from pypi on python 3.8+ | The update to handle the depreciated function `time.clock` in python 3.8 which was pushed on feb 23, 2020 is not on pypi resulting in an unusable installation for python 3.8+ users.
The work around fix is simply to go into the app/clock.py and change line 164 which reads:
`_default_time_function = time.clock`
and ... | closed | 2021-05-06T15:03:52Z | 2021-05-17T19:23:49Z | https://github.com/glumpy/glumpy/issues/285 | [] | merny93 | 2 |
mwaskom/seaborn | data-visualization | 3,192 | TypeError: ufunc 'isfinite' not supported with numpy 1.24.0 | This is the code that I ran
```python
import matplotlib.pyplot as plt
import seaborn as sns
fmri = sns.load_dataset("fmri")
fmri.info()
sns.set(style="darkgrid")
sns.lineplot(data=fmri, x="timepoint", y="signal", hue="region", style="event")
plt.show()
```
This is the error I got
```python
❯ python3... | closed | 2022-12-19T13:57:57Z | 2024-02-28T20:42:27Z | https://github.com/mwaskom/seaborn/issues/3192 | [
"upstream"
] | Rizwan-Hasan | 3 |
open-mmlab/mmdetection | pytorch | 11,556 | 在使用resume文件继续训练时卡在了advance dataloader步骤中 | 你好,最近我在MMdetection训练时服务器意外重启,所以想用resume继续训练遇到了这个情况,请问这要怎么解决
 | open | 2024-03-16T06:56:38Z | 2024-05-30T05:37:33Z | https://github.com/open-mmlab/mmdetection/issues/11556 | [] | Chengnotwang | 13 |
miguelgrinberg/python-socketio | asyncio | 1,292 | All connections stop processing events at the same time | Hi guys!
I'm recently facing an issue that all my connections turn into a strange state **almost at the same time**
So basically, I have a customised "telemetry" packet a little bit under 1k size sending every 5 seconds. After running them for couple hours, all of such "telemetry" events starting timeout. I double ... | closed | 2023-12-31T06:55:37Z | 2023-12-31T11:48:25Z | https://github.com/miguelgrinberg/python-socketio/issues/1292 | [] | morland96 | 0 |
lanpa/tensorboardX | numpy | 229 | AssertionError: %30 : Dynamic = onnx::Shape(%8) has empty scope name in add_graph() | I'm writing a PyTorch LSTM model and want to use tensorboardX to visualize the graph.
But when I call ``writer.add_graph(rnn, r)``, I got error:
```bat
C:\Python36\python.exe I:/github_repos/botrainer/train.py
Traceback (most recent call last):
File "I:/github_repos/botrainer/train.py", line 238, in <module>... | closed | 2018-09-19T08:15:04Z | 2018-12-29T17:58:33Z | https://github.com/lanpa/tensorboardX/issues/229 | [
"seems fixed"
] | hsluoyz | 1 |
google/seq2seq | tensorflow | 213 | where to get the English-Chinese data? | hello, I want to do the translation for English-Chinese or Chinese-English, but where to get the English-Chinese data link? can you give some advises?
thank you in advance! | open | 2017-05-09T12:08:58Z | 2017-07-01T15:18:29Z | https://github.com/google/seq2seq/issues/213 | [] | PapaMadeleine2022 | 1 |
deepfakes/faceswap | machine-learning | 786 | issue converting with filter | Hi, I'm trying to run convert with a face filter (to process/not process certain person) and it shows the following error.
(It runs no problem if I don't include the face filter)
face_filter = dict(detector=self.args.detector.replace("-", "_").lower(),
AttributeError: 'Namespace' object has no attribute 'detect... | closed | 2019-07-09T09:17:27Z | 2019-07-10T09:54:11Z | https://github.com/deepfakes/faceswap/issues/786 | [] | khcy82dyc | 1 |
mwaskom/seaborn | data-visualization | 3,439 | so.plot not working in version 0.12.2 | The following code does to display the intended graph, just a blank white space.
```
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
import seaborn.objects as so
dataLi = pd.read_csv("lithium_data.csv")
dataLi
(
so.Plot(dataL... | closed | 2023-08-16T02:17:21Z | 2023-08-16T16:19:59Z | https://github.com/mwaskom/seaborn/issues/3439 | [] | RSelvaratnam | 1 |
MaartenGr/BERTopic | nlp | 1,054 | Length of weights not compatible with specified axis. | umap_model = UMAP(n_components=10, n_neighbors=50, min_dist=0.0)
print('umap done')
hdbscan_model = HDBSCAN(min_samples=15, gen_min_span_tree=True,prediction_data=True)
print('hdbscan done')
ctfidf_model = ClassTfidfTransformer()
representation_model = MaximalMarginalRelevance(diversity=0.6)
vectorizer_model = ... | closed | 2023-03-01T15:54:19Z | 2023-05-23T09:35:15Z | https://github.com/MaartenGr/BERTopic/issues/1054 | [] | josepius-clemson | 4 |
gunthercox/ChatterBot | machine-learning | 2,213 | can't get the same response if I type same input twice | I'm trying to develop a chatbot for my project, the problem is I cannot get the same response with the same input. Can anyone help to solve this issue?
it looks like that:
[https://ibb.co/19b5zxP](url)
here is the code below:
from chatterbot import ChatBot
from chatterbot.trainers import... | open | 2021-11-01T07:46:59Z | 2021-12-01T17:26:09Z | https://github.com/gunthercox/ChatterBot/issues/2213 | [] | samsam3215 | 1 |
huggingface/datasets | nlp | 7,449 | Cannot load data with different schemas from different parquet files | ### Describe the bug
Cannot load samples with optional fields from different files. The schema cannot be correctly derived.
### Steps to reproduce the bug
When I place two samples with an optional field `some_extra_field` within a single parquet file, it can be loaded via `load_dataset`.
```python
import pandas as ... | closed | 2025-03-13T08:14:49Z | 2025-03-17T07:27:48Z | https://github.com/huggingface/datasets/issues/7449 | [] | li-plus | 2 |
pytest-dev/pytest-django | pytest | 542 | Catching Django-specific warnings in pytest | I would like to turn the `RemovedInNextVersionWarning` warnings to errors, as described in https://docs.pytest.org/en/latest/warnings.html
Unfortunately, when I add to pytest.ini the following lines:
```
filterwarnings =
error::RemovedInDjango20Warning
```
I end up with (it is looking strictly for Pyth... | closed | 2017-11-21T16:28:45Z | 2019-02-03T23:21:00Z | https://github.com/pytest-dev/pytest-django/issues/542 | [
"bitesize",
"documentation",
"question"
] | MRigal | 3 |
JoeanAmier/TikTokDownloader | api | 397 | 请教如何去掉保存路径中的uid | 请教作者和各位前辈
想让视频文件保存路径不包含uid,直接放在root作为根目录, 然后直接存各自mark值对应的子文件夹里
比如
"root": "/Users/admin/Desktop/抖音下载",
{
"mark": "项目1",
"url": "https://www.douyin.com/user/MS4wsjABAAAAjhjh9m4Y2GglyvbImVZlMjhjhzYPJ4?from_tab_name=main&vid=7463746000604171572",
"tab": "post",
"earliest": "2025/1/23",
... | closed | 2025-01-27T13:04:43Z | 2025-01-27T14:29:14Z | https://github.com/JoeanAmier/TikTokDownloader/issues/397 | [] | 9ihbd2DZSMjtsf7vecXjz | 2 |
flasgger/flasgger | flask | 249 | Flasgger 0.9.1 does not reflect changes without restarting app | I'm using latest flasgger==0.9.1 and create documentation in Yaml files using `swag_from` decorator. Whenever I change something in Yaml file (major or very minor change, whatever) I need to restart Python app in order to get changes reflected on Flasgger UI. I think it's quite inconvenient - I see it worked well in fl... | closed | 2018-10-04T11:46:25Z | 2019-01-16T04:41:19Z | https://github.com/flasgger/flasgger/issues/249 | [
"0.9.2"
] | ab-gissupport | 9 |
google-research/bert | nlp | 959 | How to interprete results | I am a bit confused.
After the training im left with the following files in the output folder:
train.tf_record
model.ckpt-x.meta
model.ckpt-x.index
model.ckpt-x.data-00000of-x
graph.pbtxt
checkpoints.txt
events.out.tfevents.1575989900.MO-HSK-M-TEC057
i know i have to use the model.ckpt to load the model wh... | closed | 2019-12-10T16:05:48Z | 2020-06-06T16:47:56Z | https://github.com/google-research/bert/issues/959 | [] | raff7 | 1 |
huggingface/text-generation-inference | nlp | 2,832 | use pip install TGI3.0 | ### Feature request
I want to use
pip install tgi
but there are only 2.4.0
And I see the release already has 3.0.1.
Do I just have to wait to use it via pip install tgi 3.0+?
### Motivation
Easier to use tgi 3.0+
### Your contribution
https://pypi.org/project/tgi/ | open | 2024-12-12T13:53:46Z | 2024-12-14T10:09:25Z | https://github.com/huggingface/text-generation-inference/issues/2832 | [] | xiezhipeng-git | 3 |
flasgger/flasgger | rest-api | 285 | __init__() missing 2 required positional arguments: 'schema_name_resolver' and 'spec' | Hello, I'm getting the error below. Am I missing anything?
```
../../../env36/lib64/python3.6/site-packages/flask_base/app.py:1: in <module>
from flasgger import Swagger, LazyString, LazyJSONEncoder
../../../env36/lib64/python3.6/site-packages/flasgger/__init__.py:8: in <module>
from .base import Swagger... | open | 2019-02-10T15:14:05Z | 2019-07-25T05:52:22Z | https://github.com/flasgger/flasgger/issues/285 | [] | wobeng | 12 |
hzwer/ECCV2022-RIFE | computer-vision | 110 | Nothing is generating. | Hello, i'm trying to use RIFE using the d emo.mp4 and others video.
The outputs show that no FPS is generated and no videos are present when we finalize the command:
- python3 inference_video.py --exp=2 --video=sample.mp4 --montage --skip
Fri Feb 19 14:55:27 2021
+----------------------------------------... | closed | 2021-02-19T15:00:46Z | 2021-03-15T15:12:51Z | https://github.com/hzwer/ECCV2022-RIFE/issues/110 | [] | ktakanopy | 7 |
graphistry/pygraphistry | pandas | 385 | [FEA] anonymize graph | **Is your feature request related to a problem? Please describe.**
When sharing graphs with others, especially via going from private server / private account -> public hub, such as for publicizing or debugging, it'd help to have a way to quickly anonymize a graph
Sample use cases to make fast:
* show topolog... | open | 2022-07-29T21:01:18Z | 2022-09-09T20:08:45Z | https://github.com/graphistry/pygraphistry/issues/385 | [
"enhancement",
"help wanted",
"good-first-issue"
] | lmeyerov | 3 |
scrapy/scrapy | python | 6,361 | Remove top-level reactor imports from CrawlerProces/CrawlerRunner examples | There are several code examples on https://docs.scrapy.org/en/latest/topics/practices.html that have a top-level `from twisted.internet import reactor`, which is problematic (breaks when the settings specify a non-default reactor) and needs to be fixed. | closed | 2024-05-14T09:52:47Z | 2024-05-27T10:36:35Z | https://github.com/scrapy/scrapy/issues/6361 | [
"bug",
"good first issue",
"docs"
] | wRAR | 5 |
HIT-SCIR/ltp | nlp | 338 | 如何在./tools/train/ 中编译生成./otcws文件 | <!-- 中文模板起始:If you speak English, please remove the Chinese templates -->
在提问之前,请确认以下几点:
- [x] 由于您的问题可能与前任问题重复,在提交issue前,请您确认您已经搜索过之前的问题
## 问题*类型*
<!-- 例如:构建失败、内存错误、异常终止等 -->
## 出错*场景*
<!-- 例如:分析句子“xxx”时出错,运行4小时后出错,能否复现 -->
## 代码片段
## 如何复现这一错误
<!-- Please be specific as possible. Use dashes (-) or nu... | closed | 2019-05-07T09:29:55Z | 2020-06-25T11:19:51Z | https://github.com/HIT-SCIR/ltp/issues/338 | [] | Fireboyar | 2 |
youfou/wxpy | api | 169 | 所有的方法,能否放回json格式的数据 | 所有类似的API能够返回json格式数据?
bot.friends().search('', sex=2, city='深圳', province='广东') | open | 2017-08-29T02:32:45Z | 2017-08-29T02:57:52Z | https://github.com/youfou/wxpy/issues/169 | [] | leeyisoft | 1 |
apify/crawlee-python | automation | 1,034 | How to pass arbitrary data to context | Hello, I am looking for a way to inject or pass arbitrary data to the context object. Similar to what is possible in fastapi via `app.state` or via dependencies:
```python
from fastapi import FastAPI
app = FastAPI()
app.state.custom_context = {"a": "b"}
# In separate file
router = Router()
@router.get("/items/{i... | closed | 2025-02-27T20:25:16Z | 2025-03-03T14:15:51Z | https://github.com/apify/crawlee-python/issues/1034 | [
"t-tooling"
] | cirezd | 3 |
gradio-app/gradio | data-visualization | 10,673 | No mouse wheel zoom in `gr.Plot()` after updating to Gradio 5 | ### Describe the bug
Hey there 🙂
I recently upgraded from Gradio 4.44.1 to 5.18.0 (using Python 3.11.11).
I noticed that I can no longer zoom using the mouse wheel on a Plotly-generated map inside `gr.Plot()`. The zoom-in/out buttons provided by Plotly only work occasionally.
This issue occurs both locally and on H... | closed | 2025-02-25T13:47:48Z | 2025-02-25T14:00:27Z | https://github.com/gradio-app/gradio/issues/10673 | [
"bug"
] | marinasie | 1 |
sinaptik-ai/pandas-ai | data-visualization | 1,175 | Agent Method parameter save_charts acting like open_charts, and opens saved charts automatically | ### System Info
Python 3.11.7
PandasAI 2.0.37
Pandas 1.5.3
### 🐛 Describe the bug
``` python
def test():
os.environ["PANDASAI_API_KEY"] = os.environ.get("PANDASAI_API_KEY")
llm = OpenAI(api_token="os.environ.get("OPENAI_API_KEY")")
df = pd.DataFrame({
"country": ["United States", ... | closed | 2024-05-23T17:44:29Z | 2024-08-29T16:05:48Z | https://github.com/sinaptik-ai/pandas-ai/issues/1175 | [
"bug"
] | Emad-Eldin-G | 7 |
graphql-python/graphene-sqlalchemy | graphql | 140 | geoalchemy2 support | ## Help! geoalchemy2 support!
```
class CoordinateMixin():
location = db.Column('location', Geography(geometry_type='POINT' ,srid=4326, spatial_index=True, dimension=2), doc='gps coordinate')
```
Exception: Don't know how to convert the SQLAlchemy field user.location (<class 'sqlalchemy.sql.schema.Colu... | open | 2018-07-04T12:40:44Z | 2022-01-13T16:26:50Z | https://github.com/graphql-python/graphene-sqlalchemy/issues/140 | [] | wahello | 6 |
huggingface/datasets | nlp | 6,760 | Load codeparrot/apps raising UnicodeDecodeError in datasets-2.18.0 | ### Describe the bug
This happens with datasets-2.18.0; I downgraded the version to 2.14.6 fixing this temporarily.
```
Traceback (most recent call last):
File "/home/xxx/miniconda3/envs/py310/lib/python3.10/site-packages/datasets/load.py", line 2556, in load_dataset
builder_instance = load_dataset_builder... | open | 2024-03-28T03:44:26Z | 2024-06-19T07:06:40Z | https://github.com/huggingface/datasets/issues/6760 | [] | yucc-leon | 4 |
graphql-python/graphene-django | django | 995 | ModuleNotFoundError: No module named 'graphene_django' | **Note: for support questions, please use stackoverflow**. This repository's issues are reserved for feature requests and bug reports.
* **What is the current behavior?**
```
Watching for file changes with StatReloader
Exception in thread django-main-thread:
Traceback (most recent call last):
File "/usr/local... | closed | 2020-06-26T16:12:29Z | 2020-06-26T17:50:20Z | https://github.com/graphql-python/graphene-django/issues/995 | [
"🐛bug"
] | raymondfx | 1 |
junyanz/pytorch-CycleGAN-and-pix2pix | computer-vision | 1,599 | Cycle_A loss suddenly becomes very high | Hi, @junyanz @taesungp
does anyone encounter issues like the cycle_A loss suddenly becomes very high after a random iteration? my loss log shows that in epoch 78, the discrimintor loss suddenly decrease to close to zero and my generator loss goes from 0.2ish to almost 1 and 2. The cycle_A loss also looks like being... | open | 2023-09-23T22:15:30Z | 2023-10-31T17:06:13Z | https://github.com/junyanz/pytorch-CycleGAN-and-pix2pix/issues/1599 | [] | JunCS1 | 1 |
CorentinJ/Real-Time-Voice-Cloning | python | 302 | License should be GPL | First off, this is a really cool project. Thanks for sharing it and leaving it up!
I noticed, however, that PyQt5 uses the GPL. Since this is a copyleft license, your code must also be GPL. Otherwise, we're violating the terms of the license.
See also:
https://www.riverbankcomputing.com/commercial/license-faq
h... | closed | 2020-03-22T16:22:28Z | 2020-05-02T23:47:36Z | https://github.com/CorentinJ/Real-Time-Voice-Cloning/issues/302 | [] | rustygentile | 2 |
0b01001001/spectree | pydantic | 235 | Can the email validator dependency be optional? | `EmailStr` is used for the contact info, but the field is optional. It's a waste to install these additional dependencies if I don't plan on supplying a value for this field, and don't plan on using `EmailStr` for my own API's models. Or maybe I do want to supply an email in the contact info, but don't value validation... | closed | 2022-07-11T22:08:39Z | 2022-07-14T02:51:40Z | https://github.com/0b01001001/spectree/issues/235 | [] | MarkKoz | 6 |
3b1b/manim | python | 1,228 | There are no scenes inside that module |
[09/15/20 04:18:10] ERROR __main__.py:83
There are no scenes inside that module
| closed | 2020-09-15T01:22:48Z | 2021-02-16T10:35:03Z | https://github.com/3b1b/manim/issues/1228 | [] | zhangj563 | 5 |
openapi-generators/openapi-python-client | rest-api | 351 | Support non-file fields in Multipart requests | **Is your feature request related to a problem? Please describe.**
Given this schema for a POST request:
```
{
"requestBody": {
"content": {
"multipart/form-data": {
"schema": {
"type": "object",
"properties": {
"file": {
"type": "string",
... | closed | 2021-03-19T17:08:08Z | 2021-03-22T20:25:52Z | https://github.com/openapi-generators/openapi-python-client/issues/351 | [
"✨ enhancement"
] | csymeonides-mf | 1 |
piskvorky/gensim | nlp | 2,889 | Document the X2Vec refactoring in the change log. | Document the X2Vec refactoring in the change log.
What changed from the user's perspective?
_Originally posted by @mpenkov in https://github.com/RaRe-Technologies/gensim/pull/2698/review_comment/create_ | closed | 2020-07-19T13:14:09Z | 2020-09-28T12:23:19Z | https://github.com/piskvorky/gensim/issues/2889 | [] | mpenkov | 2 |
AutoViML/AutoViz | scikit-learn | 65 | Read CSV file with different encodings | Hi. I'm trying to use the library with a CSV file that uses "ISO-8859-1" encoding, and the log says:
`pandas ascii encoder does not work for this file. Continuing...`
` pandas utf-8 encoder does not work for this file. Continuing...`
` pandas iso-8859-1 encoder does not work for this file. Continuing...`
After ... | closed | 2022-03-29T08:19:39Z | 2022-04-08T19:00:53Z | https://github.com/AutoViML/AutoViz/issues/65 | [] | gaspar-avit | 1 |
roboflow/supervision | pytorch | 943 | `Detections.empty()` invalidates detections, causes crashes when `Detections.merge()` is called. | ### Search before asking
- [ ] I have searched the Supervision [issues](https://github.com/roboflow/supervision/issues) and found no similar bug report.
This is the underlying reason for #928. I believe it is important enough to deserve a separate issue.
### Bug
`Detectons.empty()` creates a very specific t... | open | 2024-02-25T09:02:56Z | 2024-05-20T07:30:42Z | https://github.com/roboflow/supervision/issues/943 | [
"bug"
] | LinasKo | 3 |
wkentaro/labelme | deep-learning | 709 | how to label instance segmentation dataset when multiple objects of the same class in one picture | when run labelme_json_to_dataset xxx.json,no info.yaml generated.
I use the latest version.
| closed | 2020-07-01T09:11:43Z | 2020-09-07T10:21:04Z | https://github.com/wkentaro/labelme/issues/709 | [] | deep-practice | 9 |
frappe/frappe | rest-api | 31,849 | A method to disable Prepared Report permanently |
**Is your feature request related to a problem? Please describe.**
We have a script report that is used by all our users multiple times a day. There are defined limits on the amount of data that can be processed by this report and it should always execute in a fraction of a second.
Unfortunately once every few weeks ... | open | 2025-03-21T11:14:27Z | 2025-03-24T05:57:54Z | https://github.com/frappe/frappe/issues/31849 | [
"feature-request"
] | gscfogrady | 1 |
sherlock-project/sherlock | python | 2,400 | [SPAM] | [SPAM] | closed | 2025-01-29T01:36:40Z | 2025-01-29T02:04:35Z | https://github.com/sherlock-project/sherlock/issues/2400 | [
"spam"
] | Jhonatanbb | 0 |
recommenders-team/recommenders | deep-learning | 1,577 | [BUG] Misleading Example provided in LibffmConverter | The example provided in **LibffmConverter** is wrong
https://github.com/microsoft/recommenders/blob/27709229cdc4aa7d39ab715789f093a2d21d2661/recommenders/datasets/pandas_df_utils.py#L134-L141
The expected output for last column should be:
```python
field4
4:6:1
4:7:1
4:8:1
4:9:1
4:10:1
``` | closed | 2021-12-10T10:29:35Z | 2021-12-17T10:20:49Z | https://github.com/recommenders-team/recommenders/issues/1577 | [] | tim5go | 1 |
vimalloc/flask-jwt-extended | flask | 442 | access token isn't refreshing | Here is my code:
```python
from flask import Flask, url_for, redirect, jsonify, make_response, flash
from flask_mail import Mail
from flask_migrate import Migrate
from flask_restful import Api
from flask_sqlalchemy import SQLAlchemy
from config import config
from flask_login import LoginManager, logout_user, ... | closed | 2021-07-29T21:01:50Z | 2021-07-31T11:25:46Z | https://github.com/vimalloc/flask-jwt-extended/issues/442 | [] | V01D0 | 1 |
OFA-Sys/Chinese-CLIP | nlp | 250 | 中文和英文同时匹配图片时中文得分非常低 | 当对一张图片与中文和英文的文本分别相似度计算时, 中文文本的得分远小于英文的得分.
```
import torch
from PIL import Image
import requests
import cn_clip.clip as clip
from cn_clip.clip import load_from_name, available_models
print("Available models:", available_models())
# Available models: ['ViT-B-16', 'ViT-L-14', 'ViT-L-14-336', 'ViT-H-14', 'RN50... | open | 2024-01-22T11:07:26Z | 2024-01-22T11:07:26Z | https://github.com/OFA-Sys/Chinese-CLIP/issues/250 | [] | HiddenMarkovModel | 0 |
sigmavirus24/github3.py | rest-api | 609 | Support for source import API | [Support Import API](https://developer.github.com/v3/migration/source_imports/)
- [ ] Start an import
- [ ] Get import progress
- [ ] Update existing import
- [ ] Get commit authors
- [ ] Map a commit author
- [ ] Set Git LFS preference
- [ ] Get large files
- [ ] Cancel an import
I'll begin with "starting the import"... | open | 2016-05-07T20:09:12Z | 2016-11-15T19:39:34Z | https://github.com/sigmavirus24/github3.py/issues/609 | [] | itsmemattchung | 0 |
waditu/tushare | pandas | 1,004 | index_weight接口取到的指数成分和权重有不少没更新 | 比如,今天(20190411)取到的中小板399005和创业板399006权重还是去年20181228的,000807的权重还是20181130,这种情况还不少 | closed | 2019-04-11T06:17:04Z | 2019-04-16T15:17:48Z | https://github.com/waditu/tushare/issues/1004 | [] | deepfuzzy | 2 |
huggingface/datasets | pandas | 7,359 | There are multiple 'mteb/arguana' configurations in the cache: default, corpus, queries with HF_HUB_OFFLINE=1 | ### Describe the bug
Hey folks,
I am trying to run this code -
```python
from datasets import load_dataset, get_dataset_config_names
ds = load_dataset("mteb/arguana")
```
with HF_HUB_OFFLINE=1
But I get the following error -
```python
Using the latest cached version of the dataset since mteb/arguana... | open | 2025-01-06T17:42:49Z | 2025-01-06T17:43:31Z | https://github.com/huggingface/datasets/issues/7359 | [] | Bhavya6187 | 1 |
cvat-ai/cvat | computer-vision | 8,514 | Track has the incorrect number of interpolated frames equals to the total number of video frames | ### Actions before raising this issue
- [X] I searched the existing issues and did not find anything similar.
- [X] I read/searched [the docs](https://docs.cvat.ai/docs/)
### Steps to Reproduce
I am not having this error a few days ago, but I don't know why it started happening recently.
Basically, if a vi... | closed | 2024-10-05T22:07:19Z | 2024-10-06T08:05:53Z | https://github.com/cvat-ai/cvat/issues/8514 | [
"bug"
] | Microos | 1 |
flavors/django-graphql-jwt | graphql | 51 | Raise custom exception with decorators | It would be nice if the decorators would support raising a custom exception (see line in decorators). I would like to implement a `@verification_required` annotation that would extend the `@user_passes_test` function to include checking the custom verification field on the `User` model. However, it currently raises the... | closed | 2018-11-14T20:09:25Z | 2018-11-28T16:06:53Z | https://github.com/flavors/django-graphql-jwt/issues/51 | [] | kendallroth | 2 |
python-restx/flask-restx | api | 602 | Unit Test "EmailTest.test_invalid_values_check" is failing | ### Summary
The unit test case "EmailTest.test_invalid_values_check" fails because the `not-found .fr` was registered a few weeks ago and doesn't throw a `ValueError` exception anymore. This affects the CI process. ([Tests #2026](https://github.com/python-restx/flask-restx/actions/runs/8105255279))
For test cases, ... | closed | 2024-04-15T16:01:33Z | 2024-07-24T14:27:37Z | https://github.com/python-restx/flask-restx/issues/602 | [
"bug"
] | StellaContrail | 1 |
jupyter-book/jupyter-book | jupyter | 1,502 | MAINT: Remove mathjax_config migration to mathjax3_config due to sphinx>4 changes | ### Description / Summary
`jupyter-book/config.py` contains a `todo` item to remove some code when we release `jupyter-book>=0.14`
The purpose of this code is to check if someone using `sphinx>=4` has specified `mathjax_config` in:
```yaml
sphinx:
config:
mathjax_config:
```
Sphinx has made this con... | open | 2021-10-13T06:40:42Z | 2021-10-13T06:42:59Z | https://github.com/jupyter-book/jupyter-book/issues/1502 | [
"deprecate"
] | mmcky | 0 |
plotly/dash-table | dash | 634 | user friendly option for table to size to parent container | At this time it is a little tricky to get the table to vertically size to align with neighbouring components. This is exhibited while using Design Kit and including the table in a row, block, or card component.
For example: in order to get it to size within a given row
<img width="1278" alt="Screen Shot 2019-10-30 a... | open | 2019-10-31T00:22:37Z | 2019-10-31T00:22:37Z | https://github.com/plotly/dash-table/issues/634 | [] | cldougl | 0 |
tensorflow/tensor2tensor | machine-learning | 1,278 | Documentation for batch_size contradicts behaviour on TPU for variable length sequences | ### Description
The focus of this issue the interaction between the hparams batch_size and max_length.
The [comments on the batch_size hparam](https://github.com/tensorflow/tensor2tensor/blob/master/tensor2tensor/layers/common_hparams.py#L31) state the following:
```
# If the problem consists of variable-len... | open | 2018-12-05T16:27:34Z | 2018-12-05T16:27:34Z | https://github.com/tensorflow/tensor2tensor/issues/1278 | [] | etragas-fathom | 0 |
tensorflow/tensor2tensor | machine-learning | 1,586 | How to get perplexity scores for Language model training? | ### Description
In language model training I am using transformer model and geting logs for loss value around 3.5 after 3 lac steps.what is perplexity score equivalent to it? Is it e^(loss)?
| open | 2019-05-27T09:01:21Z | 2019-05-28T04:29:41Z | https://github.com/tensorflow/tensor2tensor/issues/1586 | [] | ashu5644 | 0 |
codertimo/BERT-pytorch | nlp | 49 | Question about the loss of Masked LM | Thank you very much for this great contribution.
I found the loss of masked LM didn't decrease when it reaches the value around 7. However, in the official tensorflow implementation, the loss of MLM decreases to 1 easily. I think something went wrong in your implementation.
In additional, I found the code can not ... | open | 2018-12-07T12:07:29Z | 2023-06-14T09:36:40Z | https://github.com/codertimo/BERT-pytorch/issues/49 | [
"good first issue"
] | zhezhaoa | 5 |
InstaPy/InstaPy | automation | 5,963 | Allow commenting on specific posts that already contain comment from user | <!-- Did you know that we have a Discord channel ? Join us: https://discord.gg/FDETsht -->
<!-- Is this a Feature Request ? Please, check out our Wiki first https://github.com/timgrossmann/InstaPy/wiki -->
## Expected Behavior
Comment a specific post that already contain a comment of the user.
## Current Behavior... | closed | 2020-12-16T00:12:39Z | 2020-12-17T19:57:23Z | https://github.com/InstaPy/InstaPy/issues/5963 | [] | sokratis1988 | 2 |
plotly/dash | data-visualization | 2,294 | replace `libraryTarget` in webpack config | Please use [output.library.type](https://webpack.js.org/configuration/output/#outputlibrarytype) instead of [output.libraryTarget](https://webpack.js.org/configuration/output/#outputlibrarytarget) as they might drop support for `output.libraryTarget` in the future.
https://github.com/plotly/dash/blob/a9eb3434023880c... | closed | 2022-10-29T14:22:36Z | 2023-05-15T19:30:16Z | https://github.com/plotly/dash/issues/2294 | [] | archmoj | 0 |
scikit-learn/scikit-learn | machine-learning | 30,811 | Are there any pitfalls by combining `n_jobs` and `random_state`? |
### Discussed in https://github.com/scikit-learn/scikit-learn/discussions/30809
<div type='discussions-op-text'>
<sup>Originally posted by **adosar** February 11, 2025</sup>
In [Controlling randomness](https://scikit-learn.org/stable/common_pitfalls.html#common-pitfalls-and-recommended-practices), the guide is discu... | closed | 2025-02-11T15:52:47Z | 2025-02-20T10:39:55Z | https://github.com/scikit-learn/scikit-learn/issues/30811 | [
"Needs Triage"
] | adosar | 4 |
amidaware/tacticalrmm | django | 1,596 | Feature request: Network Load Check - like CPU Load/Memory Check | Please add an Check for Network Load, like that ones for CPU and Memory.
Ive tried it with an task that runs every second and outputs the Averange Bandwith of the Last 10 Seconds to an Custom Field, its OK but it doesnt have an Diagram like the CPU and Memory.
| open | 2023-08-11T15:02:07Z | 2023-08-16T12:58:08Z | https://github.com/amidaware/tacticalrmm/issues/1596 | [
"enhancement"
] | maieredv-manuel | 1 |
python-restx/flask-restx | flask | 540 | Validate Custom Field in @api.expect() | I want to make a custom field when making an API model and validate that it is either a string or boolean or a list
I have tried
```
class CustomField(fields.Raw):
__schema_type__ = ["String", "Boolean", "List"]
__schema_example__ = "string or boolean or list"
def format(self, value):
... | closed | 2023-05-04T03:55:57Z | 2023-05-09T01:12:12Z | https://github.com/python-restx/flask-restx/issues/540 | [
"question"
] | p-g-p-t | 1 |
ageitgey/face_recognition | python | 902 | opencv读取图像报错:352 环境为win10+pycharm+python3.7+opencv4.1.0 ,希望得到解决 | import cv2 as cv
import numpy as np
print("-------hello python---------")
scr = cv.imread(r"C://Users//ASUS//Pictures//Saved Pictures//d.jpg")
cv.namedWindow("input image", cv.WINDOW_AUTOSIZE)
cv.imshow("input image", scr)
cv.waitKey(0)
cv.destroyAllWindows()
报错:cv2.error: OpenCV(4.1.0) C:\projects\op... | open | 2019-08-09T06:33:34Z | 2020-05-24T09:46:57Z | https://github.com/ageitgey/face_recognition/issues/902 | [] | RehobothRoselle | 7 |
indico/indico | sqlalchemy | 6,714 | Replace `Y` with `y` when formatting dates w/ babel | The `Y` pattern is an [ISO week date](https://babel.pocoo.org/en/latest/dates.html) which can give unexpected results when the date is
in the last/first week of the year. We should use `y` instead. Luckily, there's not that many instances where we use it. | closed | 2025-01-27T11:43:30Z | 2025-01-28T16:00:07Z | https://github.com/indico/indico/issues/6714 | [
"bug",
"help wanted"
] | tomasr8 | 0 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.