repo_name stringlengths 9 75 | topic stringclasses 30
values | issue_number int64 1 203k | title stringlengths 1 976 | body stringlengths 0 254k | state stringclasses 2
values | created_at stringlengths 20 20 | updated_at stringlengths 20 20 | url stringlengths 38 105 | labels listlengths 0 9 | user_login stringlengths 1 39 | comments_count int64 0 452 |
|---|---|---|---|---|---|---|---|---|---|---|---|
unytics/bigfunctions | data-visualization | 160 | [improve]: `sleep`: fully in BigQuery SQL (to save cloud run / python costs) | ### Check your idea has not already been reported
- [X] I could not find the idea in [existing issues](https://github.com/unytics/bigfunctions/issues?q=is%3Aissue+is%3Aopen+label%3Abug-bigfunction)
### Edit `function_name` and the short idea description in title above
- [X] I wrote the correct function name and a short idea description in the title above
### Tell us everything
current [sleep](https://github.com/unytics/bigfunctions/blob/main/bigfunctions/sleep.yaml) BigFunctions uses cloud run & python ... therefore generating cost
Here is a all BigQuery SQL solution found on stackoverflow : [is-there-a-wait-method-for-google-bigquery-sql](https://stackoverflow.com/questions/66097351/is-there-a-wait-method-for-google-bigquery-sql)
```sql
-- procedure in order to wait X seconds (made only with full BigQuery SQL without any cloudrun)
CREATE OR REPLACE PROCEDURE `bigfunctions.eu.sleep(seconds INT64)
BEGIN
DECLARE i INT64 DEFAULT 0;
DECLARE seconds_to_iterations_ratio INT64 DEFAULT 75;
DECLARE num_iterations INT64 DEFAULT seconds * seconds_to_iterations_ratio;
WHILE i < num_iterations DO
SET i = i + 1;
END WHILE;
END;
```
Shall I do a MR & swap existing sleep function to this SQL logic ? | closed | 2024-07-26T06:55:13Z | 2024-09-19T15:21:31Z | https://github.com/unytics/bigfunctions/issues/160 | [
"improve-bigfunction"
] | AntoineGiraud | 3 |
google-research/bert | tensorflow | 1,278 | OP_REQUIRES failed at save_restore_v2_ops.cc:184 : Not found: Key output_bias not found in checkpoint | I have pre-trained a model based on the bert-large model using my dataset and get the checkpoint successfully,
however ,when I use the checkpoint to do predict task I get the **Restoring from checkpoint failed. **error
i just run
****
`python run_classifier.py
--task_name=XNLI
\
--do_train=true
\
--do_eval=true
\
--data_dir=./GLUE_DIR/XNLI
\
--vocab_file=./wwm_uncased_L-24_H-1024_A-16/vocab.txt
\
--bert_config_file=./wwm_uncased_L-24_H-1024_A-16/bert_config.json
\
--init_checkpoint=./tmp/pretraining_output/model.ckpt-20
\
--max_seq_length=64
\
--train_batch_size=8
\
--learning_rate=2e-5
\
--num_train_epochs=5.0
\
--output_dir=./tmp/pretraining_output
`
2021-11-24 20:28:22.908571: W tensorflow/core/framework/op_kernel.cc:1651] OP_REQUIRES failed at save_restore_v2_ops.cc:184 : Not found: Key output_bias not found in checkpoint
ERROR:tensorflow:Error recorded from training_loop: Restoring from checkpoint failed. This is most likely due to a Variable name or other graph key that is missing from the checkpoint. Please ensure that you have not altered the graph expected based on the checkpoint. Original error:
Key output_bias not found in checkpoint
[[node save/RestoreV2 (defined at /anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/framework/ops.py:1748) ]]
Original stack trace for 'save/RestoreV2':
File "/xuweixinxi/run_classifier.py", line 1007, in <module>
tf.app.run()
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/platform/app.py", line 40, in run
_run(main=main, argv=argv, flags_parser=_parse_flags_tolerate_undef)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/absl/app.py", line 303, in run
_run_main(main, args)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/absl/app.py", line 251, in _run_main
sys.exit(main(argv))
File "/xuweixinxi/run_classifier.py", line 906, in main
estimator.train(input_fn=train_input_fn, max_steps=num_train_steps)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_estimator/python/estimator/tpu/tpu_estimator.py", line 3030, in train
saving_listeners=saving_listeners)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_estimator/python/estimator/estimator.py", line 370, in train
loss = self._train_model(input_fn, hooks, saving_listeners)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_estimator/python/estimator/estimator.py", line 1161, in _train_model
return self._train_model_default(input_fn, hooks, saving_listeners)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_estimator/python/estimator/estimator.py", line 1195, in _train_model_default
saving_listeners)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_estimator/python/estimator/estimator.py", line 1490, in _train_with_estimator_spec
log_step_count_steps=log_step_count_steps) as mon_sess:
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/training/monitored_session.py", line 584, in MonitoredTrainingSession
stop_grace_period_secs=stop_grace_period_secs)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/training/monitored_session.py", line 1014, in __init__
stop_grace_period_secs=stop_grace_period_secs)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/training/monitored_session.py", line 725, in __init__
self._sess = _RecoverableSession(self._coordinated_creator)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/training/monitored_session.py", line 1207, in __init__
_WrappedSession.__init__(self, self._create_session())
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/training/monitored_session.py", line 1212, in _create_session
return self._sess_creator.create_session()
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/training/monitored_session.py", line 878, in create_session
self.tf_sess = self._session_creator.create_session()
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/training/monitored_session.py", line 638, in create_session
self._scaffold.finalize()
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/training/monitored_session.py", line 237, in finalize
self._saver.build()
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/training/saver.py", line 840, in build
self._build(self._filename, build_save=True, build_restore=True)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/training/saver.py", line 878, in _build
build_restore=build_restore)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/training/saver.py", line 502, in _build_internal
restore_sequentially, reshape)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/training/saver.py", line 381, in _AddShardedRestoreOps
name="restore_shard"))
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/training/saver.py", line 328, in _AddRestoreOps
restore_sequentially)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/training/saver.py", line 575, in bulk_restore
return io_ops.restore_v2(filename_tensor, names, slices, dtypes)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/ops/gen_io_ops.py", line 1696, in restore_v2
name=name)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/framework/op_def_library.py", line 794, in _apply_op_helper
op_def=op_def)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/util/deprecation.py", line 507, in new_func
return func(*args, **kwargs)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/framework/ops.py", line 3357, in create_op
attrs, op_def, compute_device)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/framework/ops.py", line 3426, in _create_op_internal
op_def=op_def)
File "/anaconda3/envs/tf_ruan/lib/python3.7/site-packages/tensorflow_core/python/framework/ops.py", line 1748, in __init__
self._traceback = tf_stack.extract_stack()
| open | 2021-11-24T12:47:32Z | 2021-11-24T12:47:32Z | https://github.com/google-research/bert/issues/1278 | [] | Faker0918 | 0 |
frol/flask-restplus-server-example | rest-api | 17 | _invalid_response (potential) misusage | [Here](https://github.com/frol/flask-restplus-server-example/blob/master/app/extensions/auth/oauth2.py#L115), you're trying to override `_invalid_response` in `OAuthProvider` object.
Do you really need this trick? AFAIU, the proper way to do this would be to register your custom function using [`invalid_response`](https://github.com/lepture/flask-oauthlib/blob/e1bb478b2015239dd12f1d3ee13fb2627a5a3407/flask_oauthlib/provider/oauth2.py#L211).
Example using it as a decorator (as seen [here](https://github.com/lepture/flask-oauthlib/blob/8d4faabf17f2fa99ab5f927d3f94fbb1641ab914/tests/oauth2/server.py#L313)):
``` py
@oauth.invalid_response
def require_oauth_invalid(req):
api.abort(code=http_exceptions.Unauthorized.code)
```
| closed | 2016-08-29T10:09:59Z | 2016-08-29T10:59:19Z | https://github.com/frol/flask-restplus-server-example/issues/17 | [] | lafrech | 1 |
JaidedAI/EasyOCR | deep-learning | 986 | Fine-Tune on Korean handwritten dataset | Dear @JaidedTeam
I would like to fine-tune the EASY OCR in the handwritten Korean language, I am assuming that the pre-trained model is already trained in Korean and English vocabulary and I will enhance the Korean handwritten accuracy on EASY OCR. How do I achieve it? I know how to train custom models but due to the large size of English datasets, I don't want to train in Korean and English from scratch. I have already 10 M KOREAN handwritten images.
Regards,
Khawar | open | 2023-04-10T06:31:13Z | 2024-01-10T00:52:55Z | https://github.com/JaidedAI/EasyOCR/issues/986 | [] | khawar-islam | 6 |
dunossauro/fastapi-do-zero | sqlalchemy | 231 | fastapi_madr | https://github.com/Romariolima1998/fastapi_MADR | closed | 2024-08-22T16:55:11Z | 2024-08-27T17:45:31Z | https://github.com/dunossauro/fastapi-do-zero/issues/231 | [] | Romariolima1998 | 1 |
holoviz/colorcet | plotly | 45 | Add glasbey_tableau10? | **Is your feature request related to a problem? Please describe.**
I sometimes want to plot lots of colors but I want them to look nice
**Describe the solution you'd like**
Similar to #11, could you add a Glasbey color list that starts with the (new) Tableau 10 series?
```
4e79a7
f28e2c
e15759
76b7b2
59a14f
edc949
af7aa1
ff9da7
9c755f
bab0ab
```
| closed | 2020-02-18T05:06:31Z | 2020-02-18T22:37:11Z | https://github.com/holoviz/colorcet/issues/45 | [] | endolith | 3 |
akfamily/akshare | data-science | 5,182 | AKShare 接口问题报告index_value_hist_funddb无法使用 |
1. AKShare 已升级至1.14.74
index_value_hist_funddb和index_value_name_funddb 均无法使用

| closed | 2024-09-13T08:50:23Z | 2024-09-13T08:57:09Z | https://github.com/akfamily/akshare/issues/5182 | [
"bug"
] | sourgelp | 1 |
sammchardy/python-binance | api | 1,190 | BLVT info websockets endpoints | Hi,
it seems that BLVT info websockets (https://binance-docs.github.io/apidocs/spot/en/#websocket-blvt-info-streams) are not supported. Are there any plans to implement this functionality?
Thanks. | open | 2022-05-22T13:56:12Z | 2022-05-22T13:57:32Z | https://github.com/sammchardy/python-binance/issues/1190 | [] | jacek-jablonski | 0 |
microsoft/nni | pytorch | 4,803 | Error: PolicyBasedRL | **Describe the issue**:
I tried running the the following models space with PolicyBasedRL and I will also put in the experiment configuration:
#BASELINE NAS USING v2.7
from nni.retiarii.serializer import model_wrapper
import torch.nn.functional as F
import nni.retiarii.nn.pytorch as nn
class Block1(nn.Module):
def __init__(self, layer_size):
super().__init__()
self.conv1 = nn.Conv2d(3, layer_size*2, 3, stride=1,padding=1)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(layer_size*2, layer_size*8, 3, stride=1, padding=1)
def forward(self, x):
x = F.relu(self.conv1(x))
x = self.pool(x)
x = F.relu(self.conv2(x))
x = self.pool(x)
return x
class Block2(nn.Module):
def __init__(self, layer_size):
super().__init__()
self.conv1 = nn.Conv2d(3, layer_size, 3, stride=1,padding=1)
self.conv2 = nn.Conv2d(layer_size, layer_size*2, 3, stride=1,padding=1)
self.pool = nn.MaxPool2d(2, 2)
self.conv3 = nn.Conv2d(layer_size*2, layer_size*8, 3, stride=1,padding=1)
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.relu(self.conv2(x))
x = self.pool(x)
x = F.relu(self.conv3(x))
x = self.pool(x)
return x
class Block3(nn.Module):
def __init__(self, layer_size):
super().__init__()
self.conv1 = nn.Conv2d(3, layer_size, 3, stride=1,padding=1)
self.conv2 = nn.Conv2d(layer_size, layer_size*2, 3, stride=1,padding=1)
self.pool = nn.MaxPool2d(2, 2)
self.conv3 = nn.Conv2d(layer_size*2, layer_size*4, 3, stride=1,padding=1)
self.conv4 = nn.Conv2d(layer_size*4, layer_size*8, 3, stride=1, padding=1)
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.relu(self.conv2(x))
x = self.pool(x)
x = F.relu(self.conv3(x))
x = F.relu(self.conv4(x))
x = self.pool(x)
return x
@model_wrapper
class Net(nn.Module):
def __init__(self):
super().__init__()
rand_var = nn.ValueChoice([32,64])
self.conv1 = nn.LayerChoice([Block1(rand_var),Block2(rand_var),Block3(rand_var)])
self.conv2 = nn.Conv2d(rand_var*8,rand_var*16 , 3, stride=1, padding=1)
self.fc1 = nn.Linear(rand_var*16*8*8, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
def forward(self, x):
x = self.conv1(x)
x = F.relu(self.conv2(x))
x = x.reshape(x.shape[0],-1)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
model = Net()
from nni.retiarii.experiment.pytorch import RetiariiExeConfig, RetiariiExperiment
exp = RetiariiExperiment(model, trainer, [], RL_strategy)
exp_config = RetiariiExeConfig('local')
exp_config.experiment_name = '5%_RL_10_epochs_64_batch'
exp_config.trial_concurrency = 2
exp_config.max_trial_number = 100
#exp_config.trial_gpu_number = 2
exp_config.max_experiment_duration = '660m'
exp_config.execution_engine = 'base'
exp_config.training_service.use_active_gpu = False
--> This led to the following error:
[2022-04-24 23:49:22] ERROR (nni.runtime.msg_dispatcher_base/Thread-5) 3
Traceback (most recent call last):
File "/Users/sh/opt/anaconda3/lib/python3.7/site-packages/nni/runtime/msg_dispatcher_base.py", line 88, in command_queue_worker
self.process_command(command, data)
File "/Users/sh/opt/anaconda3/lib/python3.7/site-packages/nni/runtime/msg_dispatcher_base.py", line 147, in process_command
command_handlers[command](data)
File "/Users/sh/opt/anaconda3/lib/python3.7/site-packages/nni/retiarii/integration.py", line 170, in handle_report_metric_data
self._process_value(data['value']))
File "/Users/sh/opt/anaconda3/lib/python3.7/site-packages/nni/retiarii/execution/base.py", line 111, in _intermediate_metric_callback
model = self._running_models[trial_id]
KeyError: 3
What does this error mean/ why does it occur/ how can I fix it?
Thanks for your help!
**Environment**:
- NNI version:
- Training service (local|remote|pai|aml|etc):
- Client OS:
- Server OS (for remote mode only):
- Python version:
- PyTorch/TensorFlow version:
- Is conda/virtualenv/venv used?:
- Is running in Docker?:
**Configuration**:
- Experiment config (remember to remove secrets!):
- Search space:
**Log message**:
- nnimanager.log:
- dispatcher.log:
- nnictl stdout and stderr:
<!--
Where can you find the log files:
LOG: https://github.com/microsoft/nni/blob/master/docs/en_US/Tutorial/HowToDebug.md#experiment-root-director
STDOUT/STDERR: https://github.com/microsoft/nni/blob/master/docs/en_US/Tutorial/Nnictl.md#nnictl%20log%20stdout
-->
**How to reproduce it?**: | open | 2022-04-25T09:36:27Z | 2023-04-25T13:28:22Z | https://github.com/microsoft/nni/issues/4803 | [] | NotSure2732 | 2 |
hankcs/HanLP | nlp | 1,697 | crf模型在处理emoji时报错 | <!--
提问请上论坛,不要发这里!
提问请上论坛,不要发这里!
提问请上论坛,不要发这里!
以下必填,否则恕不受理。
-->
**问题描述:**
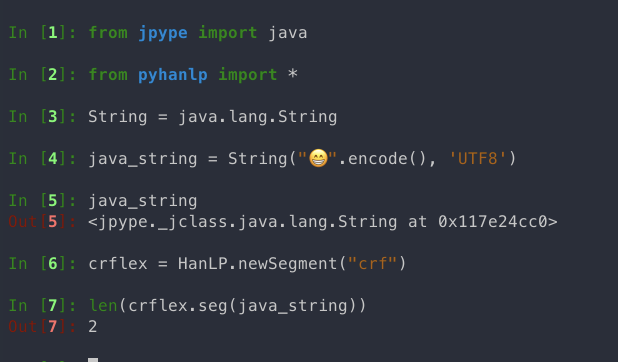
crf模型在处理emoji时报错(wordbasedsegmenter不会报错), 样例代码如下:
**复现代码**
```
from jpype import java
from pyhanlp import *
text = '😱😱😱你好,欢迎在😱Pytho😱😱n中😱调用HanLP的API 😱😱😱😱😱😱😱😱😱😱'
String = java.lang.String
java_string = String(text.encode(), 'UTF8')
string_from_java = str(java_string).encode('utf-16', errors='surrogatepass').decode('utf-16')
print('原文:' + string_from_java)
print('HanLP分词结果')
crflex = HanLP.newSegment("crf")
for term in crflex.seg(java_string):
print(str(term.word).encode('utf-16', errors='surrogatepass').decode('utf-16'))
```
**报错信息**
UnicodeDecodeError: 'utf-8' codec can't decode byte 0xed in position 0: invalid continuation byte
**System information**
系统: MAC Big Sur, I7.
python 3.6.8
HanLP Jar version: 1.8.2
pyhanlp version: 0.1.79
**其他信息**
crf模型在输入八字节的单emoji时, 会拆成两个4字节的字返回
* [x] I've completed this form and searched the web for solutions.
<!-- 发表前先搜索,此处一定要勾选! -->
<!-- 发表前先搜索,此处一定要勾选! -->
<!-- 发表前先搜索,此处一定要勾选! --> | closed | 2021-11-10T03:44:24Z | 2021-11-10T15:47:31Z | https://github.com/hankcs/HanLP/issues/1697 | [
"question"
] | Gooklim | 4 |
sinaptik-ai/pandas-ai | data-science | 747 | Translate all docs to pt-br | ### 🚀 The feature
Translate all docs to pt-br to expand the use of PandasAI. Each doc will be a task inside this issue.
### Motivation, pitch
I'm a lusophone (Portuguese speaker) and want to facilitate pandas AI use for non-English speakers.
### Alternatives
As alternatives, there could be translations for other languages, too.
### Additional context
I'm evaluating using the [Transifex](https://explore.transifex.com/) tool to control the translation better and get it most updated.
- [ ] README.md
- [ ] docs/API/helpers.md
- [ ] docs/API/llms.md
- [ ] docs/API/pandasai.md
- [ ] docs/API/prompts.md
- [ ] docs/LLMs/langchain.md
- [ ] docs/LLMs/llms.md
- [ ] docs/building_docs.md
- [ ] docs/cache.md
- [ ] docs/callbacks.md
- [ ] docs/connectors.md
- [ ] docs/CONTRIBUTING.md
- [ ] docs/custom-head.md
- [ ] docs/custom-instructions.md
- [ ] docs/custom-prompts.md
- [ ] docs/custom-response.md
- [ ] docs/custom-whitelisted-dependencies.md
- [ ] docs/determinism.md
- [ ] docs/examples.md
- [ ] docs/getting-started.md
- [ ] docs/index.md
- [ ] docs/license.md
- [ ] docs/middlewares.md
- [ ] docs/release-notes.md
- [ ] docs/save-dataframes.md
- [ ] docs/shortcuts.md
- [ ] docs/skills.md | closed | 2023-11-12T16:23:22Z | 2024-06-01T00:20:17Z | https://github.com/sinaptik-ai/pandas-ai/issues/747 | [] | HenriqueAJNB | 4 |
numpy/numpy | numpy | 27,588 | DISCUSS: About issue of masked array | While reading the code and addressing some bugs related to `numpy/ma`, I encountered a few questions:
1. **Filling Values in Masked Arrays**:
Do we actually care about the exact fill value in masked arrays, given that they are masked by other values? If not, I believe I can resolve [bug 27580](https://github.com/numpy/numpy/issues/27580) by simply removing the check for `inf`.
2. **Default Fill Value for Masked Arrays**:
Currently, the default fill value for masked arrays is defined by [`default_filler`](https://github.com/numpy/numpy/blob/bdc8d4e03181deac5280166aec4188318050570d/numpy/ma/core.py#L167C1-L183C63) for Python data types. However, Python doesn’t have unsigned integer types, so for `np.uint` arrays, the default fill value is stored as `np.int64(999999)`. This causes issues in operations like `copyto(..., casting='samekind')`, as seen in [bug 27580](https://github.com/numpy/numpy/issues/27580) and [bug 27269](https://github.com/numpy/numpy/issues/27269). Should we consider using NumPy data types for the default fill value to ensure that the fill value matches the data type of the array (e.g., using a fill value that corresponds to integers or unsigned integers as appropriate)?
3. **Large Default Fill Values**:
Some default fill values seem quite large, such as `999999` for `np.int8` and `1.e20` for `np.float16`. What would be an appropriate default fill value for masked arrays, particularly for small data types like `int8` and `float16`? ([bug25677](https://github.com/numpy/numpy/issues/25677))
4. **Reviewing `copyto` in Masked Arrays**:
Should we perform a comprehensive review of `copyto` functionality for masked arrays? It seems likely that similar bugs could exist due to the same root cause.
5. **Testing for Small Data Types**:
Should we extend the test suite to include small data types (e.g., `int8` and `float16`) to ensure that functions handle these cases correctly?
6. **Checking Method Consistency**
Should we check the consistency of method between (no-masked) `masked array` and `ndarray`? There is some difference between methods and behaviors of (no-masked) `masked array` and `ndarray`, for example, see [bug27258](https://github.com/numpy/numpy/issues/27258).
7. **Making Standard Clear**
Some methods' standard is not clear. For example, should we auto mask the invalid result? In some function (such as `sqrt` , `std`) it does, but in other function (such as `median`, `mean`). Something more worse is that in the document some function don't mention it but auto change the mask (`sqrt` [`std`](https://numpy.org/devdocs/reference/generated/numpy.ma.masked_array.std.html#numpy.ma.masked_array.std)) , and others do mention it but not change ([`mean`](https://numpy.org/devdocs/reference/generated/numpy.ma.mean.html#numpy.ma.mean)).
And something more worse is that, some important methods don't have clear explanation both in document and doc string, some of them are really important. For example, `__array_wrap__` , most of the callings to ufunc call it, and I think it might be the cause of the [bug25635](https://github.com/numpy/numpy/issues/25635).
Since I'm not sure where to place these questions, I’ve marked this as a discussion for now.
| open | 2024-10-18T13:51:16Z | 2025-02-01T22:23:01Z | https://github.com/numpy/numpy/issues/27588 | [
"15 - Discussion",
"component: numpy.ma"
] | fengluoqiuwu | 10 |
matterport/Mask_RCNN | tensorflow | 3,044 | GPU memory is occupied but cannot proceed | GPU is fully occupied, as shown in the figure below,:
<img width="547" alt="dd652e615e6ff64f59abfe21fd586b3" src="https://github.com/matterport/Mask_RCNN/assets/110774617/c7e1fc94-cfe8-44c0-bdcb-d7e31632f83b">
But training cannot proceed and is stuck at 1/1:

The code can proceed on cpu, but it's too slow,how can I solve this problem | open | 2024-07-05T14:41:28Z | 2024-08-17T10:36:55Z | https://github.com/matterport/Mask_RCNN/issues/3044 | [] | r2race | 1 |
OpenInterpreter/open-interpreter | python | 926 | Content size exceeds 20 MB | ### Describe the bug
I have two monitors. One 4k and one samsung G9 (5120x1440).
I'm getting:
```
convert_to_openai_messages.py", line 138, in convert_to_openai_messages
assert content_size_mb < 20, "Content size exceeds 20 MB"
AssertionError: Content size exceeds 20 MB
```
I assume this is because my screen image to too big?
### Reproduce
Probably only repos on large screens...
### Expected behavior
I not sure I expect anything different, perhaps a message telling my how big the image is. Or in multi screen set ups an option to only screen shot the active screen?
### Screenshots
_No response_
### Open Interpreter version
Open Interpreter 0.2.0 New Computer
### Python version
Python 3.10.12
### Operating System name and version
PopOS
### Additional context
_No response_ | closed | 2024-01-15T20:00:07Z | 2024-10-31T17:01:41Z | https://github.com/OpenInterpreter/open-interpreter/issues/926 | [
"Bug"
] | commandodev | 1 |
chaoss/augur | data-visualization | 2,593 | Contributors metric API | The canonical definition is here: https://chaoss.community/?p=3464 | open | 2023-11-30T17:34:29Z | 2023-11-30T18:21:03Z | https://github.com/chaoss/augur/issues/2593 | [
"API",
"first-timers-only"
] | sgoggins | 0 |
huggingface/datasets | pandas | 7,159 | JSON lines with missing struct fields raise TypeError: Couldn't cast array | JSON lines with missing struct fields raise TypeError: Couldn't cast array of type.
See example: https://huggingface.co/datasets/wikimedia/structured-wikipedia/discussions/5
One would expect that the struct missing fields are added with null values. | closed | 2024-09-23T07:57:58Z | 2024-10-21T08:07:07Z | https://github.com/huggingface/datasets/issues/7159 | [
"bug"
] | albertvillanova | 1 |
mwaskom/seaborn | pandas | 2,811 | kdeplot log normalization | Is it possible to apply a log normalization to a bivariate density plot? I only see the `log_scale` parameter which can apply a log scale to either one or both of the variables, but I want to scale the density values. With Matplotlib.pyplot.hist2d there is a norm parameter to which I can pass `norm=matplotlib.colors.LogNorm()` to apply a log normalization. Is this functionality available for kdeplot? | closed | 2022-05-16T15:03:54Z | 2022-05-17T16:19:05Z | https://github.com/mwaskom/seaborn/issues/2811 | [] | witherscp | 2 |
PablocFonseca/streamlit-aggrid | streamlit | 48 | CVE's in npm package immer | The package-lock.json contains the following transitive npm dependency:
"immer": {
"version": "8.0.1",
"resolved": "https://registry.npmjs.org/immer/-/immer-8.0.1.tgz",
"integrity": "sha512-aqXhGP7//Gui2+UrEtvxZxSquQVXTpZ7KDxfCcKAF3Vysvw0CViVaW9RZ1j1xlIYqaaaipBoqdqeibkc18PNvA=="
},
This package contains the CVE's below. Could you please explicitly bump the immer package to "^9.06" ([as in the streamlit main package](https://github.com/streamlit/streamlit/blob/1.2.0/frontend/package.json))?
[https://nvd.nist.gov/vuln/detail/CVE-2021-23436](https://nvd.nist.gov/vuln/detail/CVE-2021-23436 )
[https://nvd.nist.gov/vuln/detail/CVE-2021-3757](https://nvd.nist.gov/vuln/detail/CVE-2021-3757)
| closed | 2021-11-29T09:03:48Z | 2022-01-24T01:59:43Z | https://github.com/PablocFonseca/streamlit-aggrid/issues/48 | [] | vtslab | 2 |
ets-labs/python-dependency-injector | asyncio | 93 | Think about binding providers to catalogs that they are in | closed | 2015-09-29T19:57:32Z | 2015-10-16T22:09:29Z | https://github.com/ets-labs/python-dependency-injector/issues/93 | [
"research"
] | rmk135 | 1 | |
sammchardy/python-binance | api | 1,211 | API: "futures_coin_recent_trades" doesn't work? | It looks USDT-M Futures API works, while COIN-M Futures API doesn't work.
For example, the following for USDT-M Futures works!
recent_trades = client.futures_recent_trades(symbol='BTCUSDT')
While the following for COIN-M Futures doesn't work.
recent_trades = client.futures_coin_recent_trades(symbol='BTCUSD')
https://github.com/sammchardy/python-binance/blob/59e3c8045fa13ca0ba038d4a2e0481212b3b665f/binance/client.py#L6205
Error message as below: APIError(code=-1121): Invalid symbol.
Is this a bug? Thank you!
```
---------------------------------------------------------------------------
BinanceAPIException Traceback (most recent call last)
~\AppData\Local\Temp/ipykernel_13620/2129933085.py in <module>
11 """
12
---> 13 recent_trades = client.futures_coin_recent_trades(symbol='BTCUSD')
D:\ProgramData\Anaconda3\lib\site-packages\binance\client.py in futures_coin_recent_trades(self, **params)
6209
6210 """
-> 6211 return self._request_futures_coin_api("get", "trades", data=params)
6212
6213 def futures_coin_historical_trades(self, **params):
D:\ProgramData\Anaconda3\lib\site-packages\binance\client.py in _request_futures_coin_api(self, method, path, signed, version, **kwargs)
347 uri = self._create_futures_coin_api_url(path, version=version)
348
--> 349 return self._request(method, uri, signed, True, **kwargs)
350
351 def _request_futures_coin_data_api(self, method, path, signed=False, version=1, **kwargs) -> Dict:
D:\ProgramData\Anaconda3\lib\site-packages\binance\client.py in _request(self, method, uri, signed, force_params, **kwargs)
313
314 self.response = getattr(self.session, method)(uri, **kwargs)
--> 315 return self._handle_response(self.response)
316
317 @staticmethod
D:\ProgramData\Anaconda3\lib\site-packages\binance\client.py in _handle_response(response)
322 """
323 if not (200 <= response.status_code < 300):
--> 324 raise BinanceAPIException(response, response.status_code, response.text)
325 try:
326 return response.json()
BinanceAPIException: APIError(code=-1121): Invalid symbol.
``` | closed | 2022-07-03T00:54:42Z | 2022-07-03T17:38:13Z | https://github.com/sammchardy/python-binance/issues/1211 | [] | yiluzhou | 4 |
scikit-optimize/scikit-optimize | scikit-learn | 771 | [Feature request] Score confidence parameter. | Hello, I was wondering if there could be a way to assign confidence values to the scores.
Suppose I have a probabilistic algorithm that after `n` iterations assures me to have found the optimum with 70% probability and after `m` 90%. I may want to iterate with the bayesian optimizer over the number of iterations, but the prior update within the bayesian optimizer would require to include the score confidence.
How could I achieve this?
(P.S. If it makes any sense.) | open | 2019-06-06T07:27:42Z | 2020-03-06T11:43:30Z | https://github.com/scikit-optimize/scikit-optimize/issues/771 | [
"Enhancement"
] | LucaCappelletti94 | 5 |
deepspeedai/DeepSpeed | deep-learning | 6,752 | Some Demos on How to config to offload tensors to nvme device | Dear authors:
I am wondering if I could get some demos on config which is used to train a Language Model with ZeRO-Infinity?
It confused me a lot that how to config the "offload_param" and "offload_optimizer"
Thanks! | open | 2024-11-15T08:23:57Z | 2024-12-06T22:14:10Z | https://github.com/deepspeedai/DeepSpeed/issues/6752 | [] | niebowen666 | 3 |
pallets-eco/flask-sqlalchemy | flask | 755 | Documentation PDF | Hi,
is it possible to get the documentation as pdf? I saw some closed issues about this, but the solutions given there do not work anymore. It would be nice to have a link to a pdf version on this site: http://flask.pocoo.org/docs/1.0/
| closed | 2019-06-23T13:54:19Z | 2020-12-05T20:21:47Z | https://github.com/pallets-eco/flask-sqlalchemy/issues/755 | [] | sorenwacker | 1 |
Josh-XT/AGiXT | automation | 1,125 | Documentation | ### Description

Documentation shows this. Never saw this.
### Steps to Reproduce the Bug
./AGiXT.ps1
### Expected Behavior
Documentation matches experience. Also, below you need an option for Python 3.12. Also, is latest stable considered latest?
### Operating System
- [ ] Linux
- [X] Microsoft Windows
- [ ] Apple MacOS
- [ ] Android
- [ ] iOS
- [ ] Other
### Python Version
- [ ] Python <= 3.9
- [ ] Python 3.10
- [ ] Python 3.11
### Environment Type - Connection
- [X] Local - You run AGiXT in your home network
- [ ] Remote - You access AGiXT through the internet
### Runtime environment
- [ ] Using docker compose
- [ ] Using local
- [ ] Custom setup (please describe above!)
### Acknowledgements
- [X] I have searched the existing issues to make sure this bug has not been reported yet.
- [X] I am using the latest version of AGiXT.
- [X] I have provided enough information for the maintainers to reproduce and diagnose the issue. | closed | 2024-03-01T22:31:14Z | 2024-03-06T11:22:56Z | https://github.com/Josh-XT/AGiXT/issues/1125 | [
"type | report | bug",
"needs triage"
] | oldgithubman | 4 |
piccolo-orm/piccolo | fastapi | 522 | Python 3.10: DeprecationWarning: There is no current event loop | ```
piccolo/utils/sync.py:20: DeprecationWarning: There is no current event loop
loop = asyncio.get_event_loop()
```
If my understanding of that function is correct we want to use [asyncio.get_running_loop](https://docs.python.org/3.10/library/asyncio-eventloop.html#asyncio.get_running_loop) instead to get the currently running event loop, which will then raise a RuntimeError is one hasn't been created, this was added in python3.7 | closed | 2022-05-22T14:11:40Z | 2022-05-24T17:12:12Z | https://github.com/piccolo-orm/piccolo/issues/522 | [
"enhancement"
] | Drapersniper | 1 |
art049/odmantic | pydantic | 400 | Embedded models have to be defined before any normal models that reference them | # Bug
No matter how you type hint a field in the base model that references an embedded one (including future annotations), defining indexes for fields that are in the embedded model, or trying to query/sort by fields that are in the embedded model throws an error when the embedded model is defined after the base model, but not when it is defined before.
### Current Behavior
```python
from odmantic import AIOEngine, EmbeddedModel, Index, Model
from odmantic.query import asc
class Bar(Model):
bar: "Foo"
model_config = {
"indexes": lambda: [Index(asc(Bar.bar.foo))]
}
class Foo(EmbeddedModel):
foo: int
engine = AIOEngine()
await engine.configure_database([Bar]) # throws error
await engine.find(Bar, sort=asc(Bar.bar.foo)) # also throws error
```
The error that gets thrown both times is: `AttributeError: operator foo not allowed for ODMField fields`
However, when you switch the model definitions around like this:
```python
...
class Foo(EmbeddedModel):
foo: int
class Bar(Model):
bar: "Foo"
model_config = {
"indexes": lambda: [Index(asc(Bar.bar.foo))]
}
...
```
it behaves as expected (see "Expected behavior").
### Expected behavior
No errors are thrown and the engine methods work as intended.
If it's intentional or expected behavior, it should be clear in the documentation.
### Environment
- ODMantic version: 1.0.0
- MongoDB version: ...
- Pydantic version: 2.5.3
- Pydantic-core version: 2.14.6
- Python version: 3.12.0
**Additional context**
It doesn't matter if you use `foo: int = Field(...)` or just have it as `foo: int`. The behavior is unchanged.
It doesn't matter if you use `bar: "Foo"`, `bar: Foo`, or `bar: Foo # (with future annotations)`. The behavior is unchanged.
This is also the case for reference fields.
| open | 2024-01-05T07:37:52Z | 2024-01-17T06:43:02Z | https://github.com/art049/odmantic/issues/400 | [
"bug"
] | duhby | 4 |
voila-dashboards/voila | jupyter | 1,255 | Missing list of configs available | All through the documentation there is text saying to change different options in config.json. Is there a list of all config options? I'm worried I might just not know about some that I would like to use.
| open | 2022-11-09T23:18:26Z | 2022-11-09T23:18:26Z | https://github.com/voila-dashboards/voila/issues/1255 | [
"documentation"
] | pmarcellino | 0 |
wkentaro/labelme | computer-vision | 1,206 | labelme | ### Provide environment information
(labelme) C:\Users\s\Desktop\example>python labelme2voc.py images target --labels labels.txt
Creating dataset: target
class_names: ('_background_', 'SAT', 'VAT', 'Muscle', 'Bone', 'Gas', 'Intestine')
Saved class_names: target\class_names.txt
Generating dataset from: images\A004.json
Traceback (most recent call last):
File "labelme2voc.py", line 106, in <module>
main()
File "labelme2voc.py", line 95, in main
viz = imgviz.label2rgb(
TypeError: label2rgb() got an unexpected keyword argument 'img'
### What OS are you using?
python3.6 labelme3.16
### Describe the Bug
My file is labelme2voc.py. I can't find img in label2rgb(). And all my try are wrong.
### Expected Behavior
_No response_
### To Reproduce
_No response_ | closed | 2022-10-29T15:42:04Z | 2023-07-10T07:29:20Z | https://github.com/wkentaro/labelme/issues/1206 | [
"status: wip-by-author"
] | she-666 | 7 |
CorentinJ/Real-Time-Voice-Cloning | tensorflow | 1,083 | PortAudio library not found | open | 2022-06-08T10:05:35Z | 2022-11-23T19:58:33Z | https://github.com/CorentinJ/Real-Time-Voice-Cloning/issues/1083 | [] | bonad | 5 | |
keras-team/keras | deep-learning | 20,518 | Need to explicitly specify `x` and `y` instead of using a generator when the model has two inputs. | tf.__version__: '2.17.0'
tf.keras.__version__: '3.5.0'
for a image caption model, one input is vector representation of images, another is caption of encoder.
```python
def batch_generator(batch_size, tokens_train, transfer_values_train, is_random_caption=False):
# ...
x_data = \
{
'transfer_values_input': transfer_values, #transfer_values
'decoder_input': decoder_input_data # decoder_input_data
}
y_data = \
{
'decoder_output': decoder_output_data
}
yield (x_data, y_data)
```
then train on this is not work, it mistake the "decoder_input" or "decoder_output" as the "transfer_values_input" for the model, even with right parameters name in the model.
```python
decoder_model.fit(x=flick_generator,
steps_per_epoch=flick_steps_per_epoch,
epochs=20,
callbacks=flick_callbacks)
```
only explicitly specify `x` and `y` will work.
```python
from tqdm import tqdm
for epoch in tqdm(range(2)):
for step in range(flick_steps_per_epoch):
x_data, y_data = next(flick_generator)
decoder_model.fit(
x=x_data,
y=y_data,
batch_size=len(x_data[0]),
verbose=0,
)
print(f"Epoch {epoch+1} completed.")
``` | closed | 2024-11-19T20:20:59Z | 2024-12-25T02:01:11Z | https://github.com/keras-team/keras/issues/20518 | [
"stat:awaiting response from contributor",
"stale"
] | danica-zhu | 3 |
joeyespo/grip | flask | 25 | Unexpected paragraphs in lists items are messing their margin up | When using lists (any type of lists), once you introduce a paragraph inside one of the list items (by any mean: explicitly making a new paragraph or introducing a blockquote which will wrap the rest of the list item's text in a paragraph), all the following list items will be wrapped in paragraphs. You'll see it clearly in the attached screenshot.
Also, these paragraphs have a `margin-top` property that makes the lists look dirty once there are paragraphs inside. In the screenshot, from the `Item 3`, every Item have a `margin-top: 15px` applied, while the previous elements don't.

If you'd like to test on your side, here's the source:
\* Item #1
\* Item #2
With a newline inside.
\* Item #3
With another paragraph inside.
\* Item #4
\* Item #5
Can also be reproduced in numbered lists and with blockquotes.
1. Item #6
2. Item #7
3. Item #8
> With blockquote inside.
4. Item #9
While I'm here, thanks a lot for Grip. It's great tool and I'm using it daily to render MD, making my doc reading sessions much more comfy !
| closed | 2013-09-06T13:05:31Z | 2014-08-02T18:47:55Z | https://github.com/joeyespo/grip/issues/25 | [
"not-a-bug"
] | qn7o | 4 |
graphql-python/graphene-sqlalchemy | sqlalchemy | 286 | Use with SqlAlchemy classical mapping | I'm trying to use graphene-sqlalchemy in a project where classical mapping is used (https://docs.sqlalchemy.org/en/13/orm/mapping_styles.html#classical-mappings). I can see in some reports here (https://github.com/graphql-python/graphene-sqlalchemy/issues/130#issuecomment-386870986) that setting Base.query is a needed step to get things working, and there is no declararive base in classical mapping. Is there a way to make this work without having to change the entire code to use declarative mapping?
When I try to make a query I get `An error occurred while resolving field Equipment.id` `graphql.error.located_error.GraphQLLocatedError: 'Equipment' object has no attribute '__mapper__'` | closed | 2020-09-09T06:12:57Z | 2023-02-25T00:49:17Z | https://github.com/graphql-python/graphene-sqlalchemy/issues/286 | [] | eduardomezencio | 2 |
Yorko/mlcourse.ai | matplotlib | 659 | Issue related to Lasso and Ridge regression notebook file - mlcourse.ai/jupyter_english/topic06_features_regression/lesson6_lasso_ridge.ipynb / | While plotting Ridge coefficient vs weights, the alphas used are different from Ridge_alphas. And while fitting the the model and calculating the ridge_cv.alpha_ we're using ridge_alphas. So,in below code its taking alpha values from alpha defined for **Lasso**. if we plot using ridge alphas plot is quite different. Please suggest if this is correct plot.
n_alphas=200
ridge_alphas=np.logspace(-2,6,n_alphas)
coefs = []
for a in alphas: # alphas = np.linspace(0.1,10,200) it's from Lasso
model.set_params(alpha=a)
model.fit(X, y)
coefs.append(model.coef_)
| closed | 2020-03-24T11:08:45Z | 2020-03-24T11:21:44Z | https://github.com/Yorko/mlcourse.ai/issues/659 | [
"minor_fix"
] | sonuksh | 1 |
scikit-learn/scikit-learn | data-science | 30,904 | PowerTransformer overflow warnings | ### Describe the bug
I'm running into overflow warnings using PowerTransformer in some not-very-extreme scenarios. I've been able to find at least one boundary of the problem, where a vector of `[[1]] * 354 + [[0]] * 1` works fine, while `[[1]] * 355 + [[0]] * 1` throws up ("overflow encountered in multiply"). Also, an additional warning starts happening at `[[1]] * 359 + [[0]] * 1` ("overflow encountered in reduce").
Admittedly, I haven't looked into the underlying math of Yeo-Johnson, so an overflow might make sense in that light. (If that's the case, though, perhaps this is an opportunity for a clearer warning?)
### Steps/Code to Reproduce
```python
import sys
from sklearn.preprocessing import PowerTransformer
for n in range(350, 360):
print(f"[[1]] * {n}, [[0]] * 1", file=sys.stderr)
_ = PowerTransformer().fit_transform([[1]] * n + [[0]] * 1)
print(file=sys.stderr)
```
### Expected Results
```
[[1]] * 350, [[0]] * 1
[[1]] * 351, [[0]] * 1
[[1]] * 352, [[0]] * 1
[[1]] * 353, [[0]] * 1
[[1]] * 354, [[0]] * 1
[[1]] * 355, [[0]] * 1
[[1]] * 356, [[0]] * 1
[[1]] * 357, [[0]] * 1
[[1]] * 358, [[0]] * 1
[[1]] * 359, [[0]] * 1
```
### Actual Results
```
[[1]] * 350, [[0]] * 1
[[1]] * 351, [[0]] * 1
[[1]] * 352, [[0]] * 1
[[1]] * 353, [[0]] * 1
[[1]] * 354, [[0]] * 1
[[1]] * 355, [[0]] * 1
/Users/*****/lib/python3.11/site-packages/numpy/_core/_methods.py:194: RuntimeWarning: overflow encountered in multiply
x = um.multiply(x, x, out=x)
[[1]] * 356, [[0]] * 1
/Users/*****/lib/python3.11/site-packages/numpy/_core/_methods.py:194: RuntimeWarning: overflow encountered in multiply
x = um.multiply(x, x, out=x)
[[1]] * 357, [[0]] * 1
/Users/*****/lib/python3.11/site-packages/numpy/_core/_methods.py:194: RuntimeWarning: overflow encountered in multiply
x = um.multiply(x, x, out=x)
[[1]] * 358, [[0]] * 1
/Users/*****/lib/python3.11/site-packages/numpy/_core/_methods.py:194: RuntimeWarning: overflow encountered in multiply
x = um.multiply(x, x, out=x)
[[1]] * 359, [[0]] * 1
/Users/*****/lib/python3.11/site-packages/numpy/_core/_methods.py:194: RuntimeWarning: overflow encountered in multiply
x = um.multiply(x, x, out=x)
/Users/*****/lib/python3.11/site-packages/numpy/_core/_methods.py:205: RuntimeWarning: overflow encountered in reduce
ret = umr_sum(x, axis, dtype, out, keepdims=keepdims, where=where)
```
### Versions
```shell
System:
python: 3.11.9 (main, May 16 2024, 15:17:37) [Clang 14.0.3 (clang-1403.0.22.14.1)]
executable: /Users/*****/.pyenv/versions/3.11.9/envs/disposable/bin/python
machine: macOS-15.2-arm64-arm-64bit
Python dependencies:
sklearn: 1.6.1
pip: 24.0
setuptools: 65.5.0
numpy: 2.2.3
scipy: 1.15.2
Cython: None
pandas: None
matplotlib: None
joblib: 1.4.2
threadpoolctl: 3.5.0
Built with OpenMP: True
threadpoolctl info:
user_api: openmp
internal_api: openmp
num_threads: 8
prefix: libomp
filepath: /Users/*****/.pyenv/versions/3.11.9/envs/disposable/lib/python3.11/site-packages/sklearn/.dylibs/libomp.dylib
version: None
``` | open | 2025-02-26T01:45:05Z | 2025-02-28T11:39:21Z | https://github.com/scikit-learn/scikit-learn/issues/30904 | [] | rcgale | 1 |
NullArray/AutoSploit | automation | 993 | Ekultek, you are correct. | Kek | closed | 2019-04-19T16:46:38Z | 2019-04-19T16:58:00Z | https://github.com/NullArray/AutoSploit/issues/993 | [] | AutosploitReporter | 0 |
ijl/orjson | numpy | 5 | Publish a new version with `default` support? | I build and test the code from master(https://github.com/ijl/orjson/commit/15757e064586d5c6cf60528cea6ada13bf302b15) on macOS, It works fine, but I want to deploy it to Ubuntu server. | closed | 2019-01-14T08:47:39Z | 2019-01-28T14:51:03Z | https://github.com/ijl/orjson/issues/5 | [] | phyng | 3 |
supabase/supabase-py | flask | 127 | how to use .range() using supabase-py | please tell me how to use .range() using supabase-py | closed | 2022-01-18T02:35:55Z | 2023-10-07T12:57:15Z | https://github.com/supabase/supabase-py/issues/127 | [] | alif-arrizqy | 1 |
cvat-ai/cvat | computer-vision | 8,756 | issues labels |
Good afternoon, in the previous version of CVAT, I could change the numbers assigned to the labels for quick label switching, making annotation faster. How do I change this in the new version? Thanks in advance, best regards

| closed | 2024-11-28T13:51:13Z | 2024-11-29T06:36:51Z | https://github.com/cvat-ai/cvat/issues/8756 | [
"duplicate"
] | lolo-05 | 1 |
dropbox/PyHive | sqlalchemy | 286 | TTransportException: Could not start SASL: b'Error in sasl_client_start (-4) SASL(-4): no mechanism available: Unable to find a callback: 2' | Hi ,
I am trying connect Hive server 2 using PyHive using the below code
conn = hive.Connection(host='host', port=port, username='id', password='paswd',**auth='LDAP'**,database = 'default') and it generate error below.
File "C:\Users\userabc\AppData\Local\Continuum\Anaconda3\lib\site-packages\thrift_sasl\__init__.py", line 79, in open
message=("Could not start SASL: %s" % self.sasl.getError()))
TTransportException: Could not start SASL: b'Error in sasl_client_start (-4) SASL(-4): no mechanism available: Unable to find a callback: 2'.
Could you please let me know how to fix this issue.
| open | 2019-05-20T07:31:30Z | 2021-09-18T03:47:27Z | https://github.com/dropbox/PyHive/issues/286 | [] | DrYoonus | 8 |
biolab/orange3 | numpy | 6,814 | Orange3 Widget Connection Error | <!--
Thanks for taking the time to report a bug!
If you're raising an issue about an add-on (i.e., installed via Options > Add-ons), raise an issue in the relevant add-on's issue tracker instead. See: https://github.com/biolab?q=orange3
To fix the bug, we need to be able to reproduce it. Please answer the following questions to the best of your ability.
-->
**What's wrong?**
<!-- Be specific, clear, and concise. Include screenshots if relevant. -->
<!-- If you're getting an error message, copy it, and enclose it with three backticks (```). -->
I'm trying to create a classification model for dogs and cats, and I've connected it to the Test and Score widget to see which of the three models performs the best. However, the Test and Score and Confusion Matirx widgets are not connecting to each other. When I looked closely, I noticed that the percentages were rising and then stopping as the data was being passed to the models after embedding. Maybe this is why the Test and Score widget doesn't show any information about the models, but when I connected the models to the Predictions widget, the models performed well. What could be the reason for this disconnect?

The logs below were pulled just in case - could this be the reason?
C:\Program Files\Orange\lib\site-packages\orangewidget\gui.py:2068: UserWarning: decorate OWNNLearner.apply with @gui.deferred and then explicitly call apply.now or apply.deferred.
warnings.warn(
C:\Program Files\Orange\lib\site-packages\orangewidget\gui.py:2068: UserWarning: decorate OWKNNLearner.apply with @gui.deferred and then explicitly call apply.now or apply.deferred.
warnings.warn(
C:\Program Files\Orange\lib\site-packages\orangewidget\gui.py:2068: UserWarning: decorate OWLogisticRegression.apply with @gui.deferred and then explicitly call apply.now or apply.deferred.
warnings.warn(
[archive.zip](https://github.com/user-attachments/files/15506429/archive.zip)
**How can we reproduce the problem?**
<!-- Upload a zip with the .ows file and data. -->
[archive.zip](https://github.com/user-attachments/files/15506429/archive.zip)
<!-- Describe the steps (open this widget, click there, then add this...) -->
Import the train folder into the top import widget and the test folder into the bottom widget.
**What's your environment?**
<!-- To find your Orange version, see "Help → About → Version" or `Orange.version.full_version` in code -->
- Operating system: windows 11
- Orange version: 3.36.2
- How you installed Orange: Official website
| closed | 2024-05-30T22:31:33Z | 2024-06-06T15:50:53Z | https://github.com/biolab/orange3/issues/6814 | [
"bug report"
] | JISEOP-P | 1 |
autokey/autokey | automation | 974 | Unable to save changes when abbreviation is defined | ### AutoKey is a Xorg application and will not function in a Wayland session. Do you use Xorg (X11) or Wayland?
Xorg
### Has this issue already been reported?
- [X] I have searched through the existing issues.
### Is this a question rather than an issue?
- [X] This is not a question.
### What type of issue is this?
Bug
### Choose one or more terms that describe this issue:
- [ ] autokey triggers
- [X] autokey-gtk
- [ ] autokey-qt
- [ ] beta
- [X] bug
- [ ] critical
- [ ] development
- [ ] documentation
- [ ] enhancement
- [ ] installation/configuration
- [ ] phrase expansion
- [ ] scripting
- [ ] technical debt
- [ ] user interface
### Other terms that describe this issue if not provided above:
_No response_
### Which Linux distribution did you use?
Distributor ID: Arch
Description: Arch Linux
### Which AutoKey GUI did you use?
GTK
### Which AutoKey version did you use?
0.96.0
### How did you install AutoKey?
[AUR](https://aur.archlinux.org/pkgbase/autokey)
### Can you briefly describe the issue?
Using the GTK-GUI, when trying to edit a phrase that as at least one abbreviation set I'm unable to save.
Additionally, when having the Abbreviations dialog open and at least one abbreviation set, clicking on “OK” does nothing.
### Can the issue be reproduced?
Always
### What are the steps to reproduce the issue?
1. Open AutoKey GTK-GUI.
2. Select one already set phrase that already has an abbreviation.
3. Edit it.
4. Click on “Save”.
### What should have happened?
Changes should have been saved.
### What actually happened?
The GUI remains responsive but I'm now unable to navigate to other phrases or exit it by clicking “Quit” in the tray's context menu.
### Do you have screenshots?
_No response_
### Can you provide the output of the AutoKey command?
```bash
Traceback (most recent call last):
File "/usr/lib/python3.12/site-packages/autokey/gtkui/configwindow.py", line 979, in on_save
if self.__getCurrentPage().validate():
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/usr/lib/python3.12/site-packages/autokey/gtkui/configwindow.py", line 717, in validate
errors += self.settingsWidget.validate()
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/usr/lib/python3.12/site-packages/autokey/gtkui/configwindow.py", line 201, in validate
abbreviations, modifiers, key, filterExpression = self.get_item_details()
^^^^^^^^^^^^^^^^^^^^^^^
File "/usr/lib/python3.12/site-packages/autokey/gtkui/configwindow.py", line 221, in get_item_details
abbreviations = self.abbrDialog.get_abbrs()
^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/usr/lib/python3.12/site-packages/autokey/gtkui/dialogs.py", line 336, in get_abbrs
text = model.get_value(i.next().iter, 0)
^^^^^^
AttributeError: 'TreeModelRowIter' object has no attribute 'next'
```
### Anything else?
_No response_
<br/>
<hr/>
<details><summary>This repo is using Opire - what does it mean? 👇</summary><br/>💵 Everyone can add rewards for this issue commenting <code>/reward 100</code> (replace <code>100</code> with the amount).<br/>🕵️♂️ If someone starts working on this issue to earn the rewards, they can comment <code>/try</code> to let everyone know!<br/>🙌 And when they open the PR, they can comment <code>/claim #974</code> either in the PR description or in a PR's comment.<br/><br/>🪙 Also, everyone can tip any user commenting <code>/tip 20 @muekoeff</code> (replace <code>20</code> with the amount, and <code>@muekoeff</code> with the user to tip).<br/><br/>📖 If you want to learn more, check out our <a href="https://docs.opire.dev">documentation</a>.</details> | open | 2024-11-18T21:33:22Z | 2024-12-27T16:03:42Z | https://github.com/autokey/autokey/issues/974 | [
"0.96.1",
"environment"
] | muekoeff | 14 |
ivy-llc/ivy | pytorch | 27,925 | Missing `assert` leading to useless comparison | In the following test function, these 3 lines are missing `assert`
https://github.com/unifyai/ivy/blob/f057407029669edeaf1fc49b90883978e4f0c3aa/ivy_tests/test_ivy/test_frontends/test_jax/test_numpy/test_mathematical_functions.py#L1106-L1108
Reference:
Comment under that PR (https://github.com/unifyai/ivy/pull/27003#discussion_r1365411148)/) | closed | 2024-01-16T05:10:07Z | 2024-01-16T10:33:51Z | https://github.com/ivy-llc/ivy/issues/27925 | [] | Sai-Suraj-27 | 0 |
nerfstudio-project/nerfstudio | computer-vision | 3,384 | No scene option in viewer | When i use the viewer, i find the GUI don't have "scene"option. There's only control, render and export? How can i solve this?

| open | 2024-08-25T09:20:56Z | 2024-08-27T20:59:44Z | https://github.com/nerfstudio-project/nerfstudio/issues/3384 | [] | hanyangyu1021 | 2 |
tensorpack/tensorpack | tensorflow | 861 | segmentation fault while reproducing 'R50-C4' | Thanks for great work.
I used 2 GPU with horovod.
While training example 'R50-C4' segmentation fault occurs.
Also fastrcnn-losses show 'nan' (not sure its relevant to crashing issues)
An issue :
- Unexpected Problems / Potential Bugs
1. What you did:
+ If you're using examples:
+ What's the command you run:
> CUDA_VISIBLE_DEVICES=0,1 mpirun -np 2 --output-filename R50_pretrained_log python3 ./train.py --logdir=./train_log/Fasternn_R50_ImageNet_pretrained --config MODE_MASK=False FRCNN.BATCH_PER_IM=64 PREPROC.SHORT_EDGE_SIZE=600 PREPROC.MAX_SIZE=1024 TRAIN
.LR_SCHEDULE=[100000,230000,280000] TRAIN.NUM_GPUS=2
+ Have you made any changes to code? Paste them if any:
On "config.py"
> _C.TRAINER = 'horovod'
> _C.DATA.BASEDIR = '/home/cvhsh/proj_knowledge_trans_tspack/COCO'
> _C.BACKBONE.WEIGHTS = '/home/cvhsh/proj_knowledge_trans_tspack/COCO/ImageNet-ResNet50.npz' # /path/to/weights.npz
2. What you observed, including but not limited to the __entire__ logs.
+ Better to paste what you observed instead of describing them.
> [0813 19:28:17 @base.py:237] Start Epoch 52 ...
>
> 100%|##########|500/500[01:47<00:00, 4.65it/s][0813 19:30:04 @base.py:247] Epoch 52 (global_step 26000) finished, time:1 minute 47 seconds.
> [0813 19:30:04 @graph.py:73] Running Op horovod_broadcast/group_deps ...
> 100%|##########|500/500[01:47<00:00, 4.64it/s][0813 19:30:04 @base.py:247] Epoch 52 (global_step 26000) finished, time:1 minute 47 seconds.
> [0813 19:30:04 @graph.py:73] Running Op horovod_broadcast/group_deps ...
> [0813 19:30:05 @monitor.py:435] PeakMemory(MB)/gpu:0: 4393.2
> [0813 19:30:05 @base.py:237] Start Epoch 53 ...
> [0813 19:30:05 @misc.py:99] Estimated Time Left: 2 days 17 hours 40 minutes 46 seconds
>
> 0%| |0/500[00:00<?,?it/s][0813 19:30:05 @monitor.py:435] PeakMemory(MB)/gpu:0: 4418.7
> [0813 19:30:05 @monitor.py:435] QueueInput/queue_size: 50
> [0813 19:30:05 @monitor.py:435] fastrcnn_losses/box_loss: nan
> [0813 19:30:05 @monitor.py:435] fastrcnn_losses/label_loss: nan
> [0813 19:30:05 @monitor.py:435] fastrcnn_losses/label_metrics/accuracy: 0.76824
> [0813 19:30:05 @monitor.py:435] fastrcnn_losses/label_metrics/false_negative: 1
> [0813 19:30:05 @monitor.py:435] fastrcnn_losses/label_metrics/fg_accuracy: 2.246e-37
> [0813 19:30:05 @monitor.py:435] learning_rate: 0.01
> [0813 19:30:05 @monitor.py:435] rpn_losses/box_loss: 0.056564
> [0813 19:30:05 @monitor.py:435] rpn_losses/label_loss: 0.24104
> [0813 19:30:05 @monitor.py:435] rpn_losses/label_metrics/precision_th0.1: 0.38716
> [0813 19:30:05 @monitor.py:435] rpn_losses/label_metrics/precision_th0.2: 0.4666
> [0813 19:30:05 @monitor.py:435] rpn_losses/label_metrics/precision_th0.5: 0.54889
> [0813 19:30:05 @monitor.py:435] rpn_losses/label_metrics/recall_th0.1: 0.79813
> [0813 19:30:05 @monitor.py:435] rpn_losses/label_metrics/recall_th0.2: 0.71973
> [0813 19:30:05 @monitor.py:435] rpn_losses/label_metrics/recall_th0.5: 0.3714
> [0813 19:30:05 @monitor.py:435] rpn_losses/num_pos_anchor: 40.375
> [0813 19:30:05 @monitor.py:435] rpn_losses/num_valid_anchor: 256
> [0813 19:30:05 @monitor.py:435] sample_fast_rcnn_targets/num_bg: 49.167
> [0813 19:30:05 @monitor.py:435] sample_fast_rcnn_targets/num_fg: 14.833
> [0813 19:30:05 @monitor.py:435] sample_fast_rcnn_targets/proposal_metrics/best_iou_per_gt: nan
> [0813 19:30:05 @monitor.py:435] sample_fast_rcnn_targets/proposal_metrics/recall_iou0.3: 0.63788
> [0813 19:30:05 @monitor.py:435] sample_fast_rcnn_targets/proposal_metrics/recall_iou0.5: 0.53133
> [0813 19:30:05 @monitor.py:435] total_cost: nan
> [0813 19:30:05 @monitor.py:435] wd_cost: nan
> [0813 19:30:05 @base.py:237] Start Epoch 53 ...
>
> 0%| |0/500[00:00<?,?it/s][cvhsh-labdevice:25111] *** Process received signal ***
> [cvhsh-labdevice:25111] Signal: Segmentation fault (11)
> [cvhsh-labdevice:25111] Signal code: Address not mapped (1)
> [cvhsh-labdevice:25111] Failing at address: (nil)
> [cvhsh-labdevice:25111] [ 0] /lib/x86_64-linux-gnu/libpthread.so.0(+0x11390)[0x7f7ddd406390]
> [cvhsh-labdevice:25111] [ 1] /usr/local/lib/python3.5/dist-packages/horovod/common/mpi_lib.cpython-35m-x86_64-linux-gnu.so(+0x22047)[0x7f7d2cf2f047]
> [cvhsh-labdevice:25111] [ 2] /usr/local/lib/python3.5/dist-packages/horovod/common/mpi_lib.cpython-35m-x86_64-linux-gnu.so(+0x2409b)[0x7f7d2cf3109b]
> [cvhsh-labdevice:25111] [ 3] /usr/lib/x86_64-linux-gnu/libstdc++.so.6(+0xb8c80)[0x7f7dd5c0fc80]
> [cvhsh-labdevice:25111] [ 4] /lib/x86_64-linux-gnu/libpthread.so.0(+0x76ba)[0x7f7ddd3fc6ba]
> [cvhsh-labdevice:25111] [ 5] /lib/x86_64-linux-gnu/libc.so.6(clone+0x6d)[0x7f7ddd13241d]
> [cvhsh-labdevice:25111] *** End of error message ***
> \-------------------------------------------------------
> Primary job terminated normally, but 1 process returned
> a non-zero exit code. Per user-direction, the job has been aborted.
> \-------------------------------------------------------
> \--------------------------------------------------------------------------
> mpirun noticed that process rank 0 with PID 0 on node cvhsh-labdevice exited on signal 11 (Segmentation fault).
> \--------------------------------------------------------------------------
3. What you expected, if not obvious.
4. Your environment:
+ Python version.
: 3.5.2
+ TF version: `python -c 'import tensorflow as tf; print(tf.GIT_VERSION, tf.VERSION)'`.
: 1.9.0
+ Tensorpack version: `python -c 'import tensorpack; print(tensorpack.__version__)'`.
: 0.8.6
+ Hardware information, if relevant.
: GPU PASCAL TITAN Xp 12G
+ others
: ubuntu 16.04, nvidia drivier 396.37, CUDA 9.0
: horovod(0.13.11), mpirun (Open MPI) 3.1.1, opencv-contrib-python (3.4.2.17) | closed | 2018-08-13T11:06:49Z | 2018-08-22T18:18:11Z | https://github.com/tensorpack/tensorpack/issues/861 | [
"examples"
] | sehyun03 | 4 |
httpie/cli | rest-api | 971 | Basic authentication: unexpected extra leading space in generated base64 token | Hi,
I was giving `httpie` a try, in order to replace `curl` especially because of the session management.
And I might miss something, but I believe the Basic authentication token computed has an unexpected extra leading space.
**Steps to reproduce**
I run:
```
$ https --offline -a 'a:b' domain.tld/auth | grep Authorization
```
**Expected result**
- I get:
```
Authorization: Basic YTpi
```
- Which, when decoded, gives:
```
$ echo \"$(printf "YTpi"|base64 -d)\"
"a:b"
```
**Actual result**
- I get:
```
Authorization: Basic wqBhOmI=
```
- Which, when decoded, gives an unexpected extra leading space:
```
$ echo \"$(printf "wqBhOmI="|base64 -d)\"
" a:b"
```
**Extra info**
- I checked the output given by `curl`, which gives the expected result:
```
$ curl --verbose -u 'a:b' https://httpbin.org/auth 2>&1 | grep Authorization
> Authorization: Basic YTpi
```
- Debug output
```
$ https --debug --offline -a 'a:b' domain.tld/auth
HTTPie 2.2.0
Requests 2.24.0
Pygments 2.6.1
Python 3.8.5 (default, Jul 21 2020, 10:48:26)
[Clang 11.0.3 (clang-1103.0.32.62)]
/usr/local/Cellar/httpie/2.2.0/libexec/bin/python3.8
Darwin 19.6.0
<Environment {'colors': 256,
'config': {'default_options': []},
'config_dir': PosixPath('/Users/eroubion/.config/httpie'),
'is_windows': False,
'log_error': <function Environment.log_error at 0x1057450d0>,
'program_name': 'https',
'stderr': <_io.TextIOWrapper name='<stderr>' mode='w' encoding='utf-8'>,
'stderr_isatty': True,
'stdin': <_io.TextIOWrapper name='<stdin>' mode='r' encoding='utf-8'>,
'stdin_encoding': 'utf-8',
'stdin_isatty': True,
'stdout': <_io.TextIOWrapper name='<stdout>' mode='w' encoding='utf-8'>,
'stdout_encoding': 'utf-8',
'stdout_isatty': True}>
>>> requests.request(**{'auth': <httpie.plugins.builtin.HTTPBasicAuth object at 0x105814520>,
'data': RequestJSONDataDict(),
'files': RequestFilesDict(),
'headers': {'User-Agent': b'HTTPie/2.2.0'},
'method': 'get',
'params': RequestQueryParamsDict(),
'url': 'https://domain.tld/auth'})
GET /auth HTTP/1.1
Accept: */*
Accept-Encoding: gzip, deflate
Authorization: Basic wqBhOmI=
Connection: keep-alive
Host: domain.tld
User-Agent: HTTPie/2.2.0
```
Of course, the header could be better parsed server-side: actually it just does not authenticate the request. Maybe it could be smarter (?), but seems weird to me.
As I said, I may miss something here, first experience with `httpie` here , and it looks "wow": thanks! | closed | 2020-09-30T16:40:18Z | 2020-10-01T10:28:11Z | https://github.com/httpie/cli/issues/971 | [] | maanuair | 3 |
strawberry-graphql/strawberry | fastapi | 3,305 | Expore schema directives in schema introspection API | Add schema directives in schema introspection API
<!--- This template is entirely optional and can be removed, but is here to help both you and us. -->
<!--- Anything on lines wrapped in comments like these will not show up in the final text. -->
## Feature Request Type
- [ ] Core functionality
- [x] Alteration (enhancement/optimization) of existing feature(s)
- [ ] New behavior
## Description
Right now strawberry doesn\`t expose schema directives usage to schema, only declaration. So you can look whatever such schema directive defined on server, but you can\`t look where it is used. | open | 2023-12-22T16:27:57Z | 2025-03-20T15:56:32Z | https://github.com/strawberry-graphql/strawberry/issues/3305 | [] | Voldemat | 1 |
vanna-ai/vanna | data-visualization | 338 | Support additional js chart libraries (like highcharts / d3 js ) in addition to plotly. | **Support customization of charting libraries for front end.**
Exploring on using vann-flask/API or built in VannaFlaskApp(vn) but this seems to be tightly coupled with Plotly as visualization library.
Trying to extend on the example given below link but the js assets are compressed and minified.
Can I know from where can I find the uncompressed src.js or jsx files or the information regarding building those js files for the javascript code shared in the below
https://github.com/vanna-ai/vanna-flask
and assest.py in src/vanna/flask folder.
**Describe the solution you'd like**
Make the front end samples decoupled with plotting libraries.
| closed | 2024-04-04T15:46:39Z | 2024-04-08T07:43:50Z | https://github.com/vanna-ai/vanna/issues/338 | [] | rrimmana | 2 |
davidsandberg/facenet | tensorflow | 513 | do you plan to release finetuning code on own datasets? | closed | 2017-11-02T07:11:19Z | 2018-04-01T21:33:50Z | https://github.com/davidsandberg/facenet/issues/513 | [] | ronyuzhang | 1 | |
graphql-python/graphene | graphql | 867 | How to make a required argument? | Apologies for writing a question in the bugtracker but I simply can not figure out how to create a required custom argument for a query. Can anyone show me how it's done? | closed | 2018-11-23T23:24:01Z | 2018-11-23T23:28:41Z | https://github.com/graphql-python/graphene/issues/867 | [] | jmn | 0 |
graphql-python/graphene | graphql | 1,065 | Support for GEOS fields-- PointField, LineString etc... | Can we add support for fields offered by ``django.contrib.gis.geos``-- [GEOS](https://docs.djangoproject.com/en/2.2/ref/contrib/gis/geos/). ``django-graphql-geojson`` package comes closest to solving the problem, however, the particular implementation has a major drawback that it converts the whole *model* into *GeoJSON*, which causes problems with external interfaces which are defined on *non-geometric* fields. (May also break internal ones.)
Converting *GEOS* fields into *GeoJSON* should more than sufficient for most use cases because it provides you with pre-built interfaces and filters on non-geometric types.
How would one go about adding support for serialization of these fields? Having gone through the codebase-- adding them to ``converter.py`` coupled with ``tests.py`` should be sufficient or am I missing something?
**Edit** Wrong Repository. Sorry.
| closed | 2019-09-05T16:46:43Z | 2019-09-05T17:01:27Z | https://github.com/graphql-python/graphene/issues/1065 | [] | EverWinter23 | 0 |
matplotlib/matplotlib | data-science | 29,779 | [Bug]: get_default_filename removes '0' from file name instead of '\0' from window title | ### Bug summary
removed_chars = r'<>:"/\|?*\0 ' in get_default_filename has r before string so escape char '\' is ignored in the string, so it just replace regular zero character '0' with '_'
just replace removed_chars = r'<>:"/\|?*\0 ' with removed_chars = '<>:"/\\|?*\0 '
https://github.com/matplotlib/matplotlib/blob/c887ecbc753763ff4232041cc84c9c6a44d20fd4/lib/matplotlib/backend_bases.py#L2223
### Code for reproduction
```Python
# create a plot and set title to sth with 0 in it
fig.canvas.set_window_title('120')
# then click on save button
```
### Actual outcome
It shows save dialog with 12_.png file name
### Expected outcome
It should show save dialog with 120.png file name
### Additional information
_No response_
### Operating system
_No response_
### Matplotlib Version
All
### Matplotlib Backend
All
### Python version
All
### Jupyter version
All
### Installation
None | closed | 2025-03-19T01:51:05Z | 2025-03-20T19:30:34Z | https://github.com/matplotlib/matplotlib/issues/29779 | [] | htalaco | 0 |
piskvorky/gensim | machine-learning | 3,028 | AttributeError: 'Word2Vec' object has no attribute 'most_similar' |
#### Problem description
When I was trying to use a trained word2vec model to find the similar word, it showed that 'Word2Vec' object has no attribute 'most_similar'.
I haven't seen that what are changed of the 'most_similar' attribute from gensim 4.0. When I was using the gensim in Earlier versions, most_similar() can be used as:
model_hasTrain=word2vec.Word2Vec.load(saveBinPath)
y=model_hasTrain.most_similar('price',topn=100)
I don't know that are most_similar() removed or changed? Or do I need to reinstall the gensim? Thank you for solving my problem. Thanks very much.
#### Versions
Please provide the output of:
```python
Windows-2012Server-6.2.9200-SP0
Python 3.8.3 (default, Jul 2 2020, 17:30:36) [MSC v.1916 64 bit (AMD64)]
Bits 64
NumPy 1.18.5
SciPy 1.5.0
gensim 4.0.0beta
FAST_VERSION 1
```
| closed | 2021-01-19T08:44:24Z | 2021-01-19T21:41:21Z | https://github.com/piskvorky/gensim/issues/3028 | [
"bug",
"documentation"
] | freedomSummer1964 | 5 |
horovod/horovod | deep-learning | 3,772 | Support newer version of pytorch_lightning | **Is your feature request related to a problem? Please describe.**
Do we have plans to support newer version of pytorch_lightning with horovod? Such as v1.8.0? There're lots of new features coming in and horovod is stil depending on v1.5.x
**Describe the solution you'd like**
Update package version and fix incompatible stuff.
**Describe alternatives you've considered**
No applicable.
**Additional context**
I can help contribute :)
| open | 2022-11-17T06:25:13Z | 2022-11-17T17:55:25Z | https://github.com/horovod/horovod/issues/3772 | [
"enhancement",
"contribution welcome"
] | serena-ruan | 2 |
hzwer/ECCV2022-RIFE | computer-vision | 341 | Reproducing 3.6 HD model | Hello! I am trying to reproduce the 3.6 model linked in the Readme.md
However, the training procedure in RIFE_HDv3.py file shipped with the pretrained model appears to be broken
`if training:
self.optimG.zero_grad()
loss_G = **loss_cons** + loss_smooth * 0.1
loss_G.backward()
self.optimG.step()
else:
flow_teacher = flow[2]
return merged[2], {
'mask': mask,
'flow': flow[2][:, :2],
'loss_l1': loss_l1,
'loss_cons': **loss_cons**,
'loss_smooth': loss_smooth,
}`
The loss_cons variable is not assigned anywhere, also L1 loss is calculated but not used
What loss combination should i use to reproduce v3.6?
Also what file should i use for training, RIFE_HDv3.py shipped with the model or RIFE.py presented in the repo?
Also, was that model also trained just on Vimeo triplet dataset? | closed | 2023-10-23T00:13:02Z | 2023-10-27T01:37:08Z | https://github.com/hzwer/ECCV2022-RIFE/issues/341 | [] | Lum1nus | 5 |
deepspeedai/DeepSpeed | machine-learning | 6,945 | DeepSpeed Installation Fails During Docker Build (NVML Initialization Issue) | Hello,
I encountered an issue while building a Docker image for deep learning model training, specifically when attempting to install DeepSpeed.
**Issue**
When building the Docker image, the DeepSpeed installation fails with a warning that NVML initialization is not possible.
However, if I create a container from the same image and install DeepSpeed inside the container, the installation works without any issues.
**Environment**
Base Image: `nvcr.io/nvidia/pytorch:23.01-py3`
DeepSpeed Version: `0.16.2`
**Build Log**
[docker_build.log](https://github.com/user-attachments/files/18396187/docker_build.log)
**Additional Context**
The problem does not occur with the newer base image `nvcr.io/nvidia/pytorch:24.05-py3`.
Thank you. | closed | 2025-01-13T11:39:39Z | 2025-01-24T00:32:39Z | https://github.com/deepspeedai/DeepSpeed/issues/6945 | [] | asdfry | 8 |
davidsandberg/facenet | computer-vision | 415 | The FLOPS and param on Inception-ResNet-v1? | @davidsandberg
Thans for you share your FaceNet work.
Now I consider hardwar . In your FaceNet paper, the FLOPS is 1.6B on NN2. I wants to know the FLOPS and param on Inception-ResNet-v1. Do you have the information about FLOPS and param on Inception-ResNet-v1?
thank you very much! | open | 2017-08-10T08:42:30Z | 2019-01-23T03:47:28Z | https://github.com/davidsandberg/facenet/issues/415 | [] | lsx66 | 2 |
scikit-multilearn/scikit-multilearn | scikit-learn | 224 | MlKNN:TypeError: __init__() takes 1 positional argument but 2 were given | When mlknn is used, X is passed in_ train,Y_ When running a train, it always reports an error, saying that one more parameter has been passed. You can obviously pass two parameters. I don't know where the error is | closed | 2021-10-26T02:06:07Z | 2024-03-01T05:02:28Z | https://github.com/scikit-multilearn/scikit-multilearn/issues/224 | [] | Foreverkobe1314 | 13 |
littlecodersh/ItChat | api | 812 | 博客 | 图片都不能看的 | closed | 2019-04-10T14:23:51Z | 2019-05-22T02:14:53Z | https://github.com/littlecodersh/ItChat/issues/812 | [] | juzj | 0 |
flasgger/flasgger | api | 364 | DispatcherMiddleware has moved | Werkzeug==1.0.0
Please change:
from werkzeug.wsgi import DispatcherMiddleware
to:
from werkzeug.middleware.dispatcher import DispatcherMiddleware | open | 2020-02-10T21:02:50Z | 2020-02-10T21:02:50Z | https://github.com/flasgger/flasgger/issues/364 | [] | fenchu | 0 |
xlwings/xlwings | automation | 1,797 | OS Error: -1743 (User has declined permission) when trying to use xlwings | Hi,
I have made a program with a GUI (PyQt5) that edits excel sheets with Python, Openpyxl, and Xlwings. I have used PyInstaller to make this into an app.
Now I have the .app format and an exec. When I run the .exec by double-clicking from Finder the program works fine. When I run the .app from the terminal with this command:
`./dist/main_window.app/Contents/MacOS/main_window`
The program works fine.
However, when I try and double click the .app the GUI launches and the GUI works perfectly, but when I hit the button that uses openpyxl and xlwings the app crashes.
I tried catching the exception and got this:
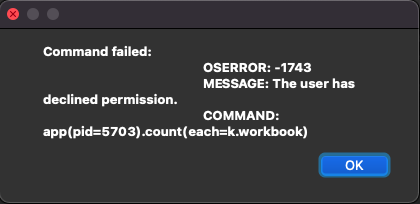
I have checked my System Preferences -> Security & Privacy and have given the .app file Full Disk Access and Developer Tools Access.
This is how the structure of my code is:
A button in the GUI calls this function (this is where I use OpenPyxl):
```
def creating_planner(year, uni_name, bin_day, bin_week, add_school):
wb = Workbook()
jan = wb.create_sheet("JANUARY")
feb = wb.create_sheet("FEBRUARY")...
... desktop = os.path.join(os.path.join(os.path.expanduser('~')), 'Desktop')
xlsx_name = str(year) + " Planner.xlsx"
wb.save(desktop + "/" + xlsx_name)
adding_colours_PH(year)
```
The code works well up to here, it saves the excel sheet on the Desktop. After this step, I use a function that opens this excel workbook in XlWings, the code is below:
```
def adding_colours_PH(year):
info, reason = publicHolidays(year)
desktop = os.path.join(os.path.join(os.path.expanduser('~')), 'Desktop')
xlsx_name = str(year) + " Planner.xlsx"
wb = xw.Book(desktop + "/" + xlsx_name)...
```
I believe this is where the error message comes up.
Could you please help me fix this error? | closed | 2022-01-18T14:06:33Z | 2022-01-19T03:11:54Z | https://github.com/xlwings/xlwings/issues/1797 | [] | febinjose24 | 2 |
collerek/ormar | fastapi | 419 | Creating records directly in tests in a FastAPI appliction | **Describe the bug**
Suppose I have an endpoint like this:
```python
@router.get("/", response_model=List[Item])
async def get_items(keys: List[str] = Query([])) -> List[Item]:
return await Item.objects.filter(key__in=keys).all()
```
I'd like to test this with something like:
```python
@pytest.mark.asyncio
async def test_success(self, client: TestClient) -> None:
item_1 = await Item.objects.create(key="1")
item_2 = await Item.objects.create(key="2")
url = router.url_path_for("get_items")
response = client.get(
f"{url}?keys=1&keys=2&keys=3",
)
data = response.json()
assert len(data) == 2
assert data[0]["id"] == item_1.id
assert data[1]["id"] == item_2.id
```
However, running this i get errors of this nature:
```
asyncpg.exceptions._base.InterfaceError: cannot perform operation: another operation is in progress
```
I'm using the following fixtures:
```python
@pytest.fixture(autouse=True, scope="class")
def create_test_database() -> Generator[None, None, None]:
"""Create a test database before tests and tear down afterwards."""
engine = sqlalchemy.create_engine(settings.sqlalchemy_database_uri)
metadata.create_all(engine)
yield
metadata.drop_all(engine)
@pytest.fixture(scope="module")
def client() -> Generator[TestClient, None, None]:
"""Initialize a `TestClient` to be used in tests.
Yields:
the test client
"""
with TestClient(app) as test_client:
yield test_client
```
Is there a way around this? I notice in your tests you create dummy routes for the test client app to create objects in the tests, is that the only way to create these factory style records in tests?
**To Reproduce**
Steps to reproduce the behavior:
This is a little tricky given that it relies on a postgres database but I can give a full example if needs be
**Expected behavior**
I should be able to create a record in test
**Screenshots**
If applicable, add screenshots to help explain your problem.
**Versions (please complete the following information):**
- Database backend used (mysql/sqlite/postgress): postgres:13
- Python version: 3.9.0
- `ormar` version: 0.10.22
- `pydantic` version: 1.8.2
- if applicable `fastapi` version: 0.70.0
| closed | 2021-11-09T17:04:06Z | 2021-11-09T17:21:41Z | https://github.com/collerek/ormar/issues/419 | [
"bug"
] | dbatten5 | 2 |
encode/databases | asyncio | 594 | Handle errors in transaction.commit() (for sqlite) | ## Context
For `sqlite` backend, when inside a transaction, errors are raised at the execution of the `COMMIT` statement instead of the SQL that causes the error, for example
```sql
PRAGMA foreign_keys = ON;
BEGIN TRANSACTION;
INSERT INTO "follow" ("user_id", "target_id") VALUES (0, 0);
COMMIT;
```
It was line 3 causing the `SQLITE_CONSTRAINT_FOREIGNKEY` error, but the error is raised during the `COMMIT` execution;
## Problem
in `databases/core.py` line 463:
```python
class Transaction:
async def commit(self) -> None:
async with self._connection._transaction_lock:
assert self._connection._transaction_stack[-1] is self
self._connection._transaction_stack.pop() # this line
assert self._transaction is not None
await self._transaction.commit()
await self._connection.__aexit__()
self._transaction = None
```
the `_transaction_stack.pop()` is called before the `_transaction.commit()` is called, so if `_transaction.commit()` raised an error, the current transaction is lost in the connection and cannot be rolled back, thus leading the database to be locked;
## Fix
To prevent this problem, I came up with a workaround:
1. to catch errors in `commit()` and call `rollback()` manually
2. move the `_transaction_stack.pop()` after the `commit()` statement, so the transaction will be poped after committed successfully
namely:
```python
from databases.core import Transaction
class _Transaction(Transaction):
async def commit(self) -> None:
async with self._connection._transaction_lock:
assert self._connection._transaction_stack[-1] is self
assert self._transaction is not None
await self._transaction.commit()
# POP after committing successfully
self._connection._transaction_stack.pop()
await self._connection.__aexit__()
self._transaction = None
async def __aexit__(self, exc_type, exc_value, traceback):
"""
Called when exiting `async with database.transaction()`
"""
if exc_type is not None or self._force_rollback:
await self.rollback()
else:
try:
await self.commit()
except Exception as e:
await self.rollback()
raise e
```
I am following the contributing guide here to submit an issue to discuss this modification before I make a pull request, and hope this can be merged to the repo soon
| open | 2024-10-06T08:18:24Z | 2024-10-06T08:18:24Z | https://github.com/encode/databases/issues/594 | [] | voidZXL | 0 |
kizniche/Mycodo | automation | 1,393 | DS18B20 Issue | ### Describe the problem/bug
DS18B20 Temperature sensor stopped working overnight and got stuck at 0C. Config page for w1thermsensor shows two "00-00000000." device ID
Tried a config page for OW-Shell and device ID shows 2 different unique IDs
The input with ow-shell library provides erratic measurements ranging from 0C to 96C jumping all around.
I bought a new sensor (both from adafruit) and same behavior happens with brand new sensor. Sensors are installed with 4.7k pullup resistor between signal and VCC.
### Versions:
- Mycodo Version: 8.15.13
- Raspberry Pi Version: 4b
- Raspbian OS Version: *[e.g. Buster Lite]*
### Screenshots


### Additional context
Logs show:
2024-08-23 23:40:33,446 - ERROR - mycodo.inputs.ds18s20_f2982e8f - Error 101: Device not set up. See https://kizniche.github.io/Mycodo/Error-Codes#error-101 for more info.
2024-08-23 23:40:36,663 - ERROR - mycodo.inputs.ds18s20_f2982e8f - Initialization errored 3 times; giving up. Maybe the following traceback can help diagnose the issue.
Traceback (most recent call last):
File "/var/mycodo-root/mycodo/abstract_base_controller.py", line 57, in try_initialize
self.initialize()
File "/home/mfittipaldi/Mycodo/mycodo/inputs/ds18s20.py", line 98, in initialize
self.sensor = W1ThermSensor(
^^^^^^^^^^^^^^
File "/var/mycodo-root/env/lib/python3.11/site-packages/w1thermsensor/core.py", line 154, in __init__
raise NoSensorFoundError(
w1thermsensor.errors.NoSensorFoundError:
Could not find sensor of type DS18S20 with id 2e0000000000
Please check cabling and check your /boot/config.txt for
dtoverlay=w1-gpio
2024-08-23 23:40:36,664 - DEBUG - mycodo.controllers.controller_input_f2982e8f - get_measurement() found
2024-08-23 23:40:36,665 - DEBUG - mycodo.controllers.controller_input_f2982e8f - listener() not found
2024-08-23 23:40:36,665 - INFO - mycodo.controllers.controller_input_f2982e8f - Activated in 10172.7 ms
2024-08-23 23:40:36,665 - ERROR - mycodo.inputs.ds18s20_f2982e8f - Error 101: Device not set up. See https://kizniche.github.io/Mycodo/Error-Codes#error-101 for more info.
2024-08-23 23:40:41,669 - ERROR - mycodo.inputs.ds18s20_f2982e8f - Error 101: Device not set up. See https://kizniche.github.io/Mycodo/Error-Codes#error-101 for more info.
2024-08-23 23:40:48,471 - ERROR - mycodo.inputs.ds18s20_f2982e8f - Error 101: Device not set up. See https://kizniche.github.io/Mycodo/Error-Codes#error-101 for more info.
2024-08-23 23:40:56,683 - ERROR - mycodo.inputs.ds18s20_f2982e8f - Error 101: Device not set up. See https://kizniche.github.io/Mycodo/Error-Codes#error-101 for more info.
2024-08-23 23:40:56,683 - ERROR - mycodo.controllers.controller_input_f2982e8f - StopIteration raised 3 times. Possibly could not read input. Ensure it's connected properly and detected.
2024-08-23 23:41:03,487 - ERROR - mycodo.inputs.ds18s20_f2982e8f - Error 101: Device not set up. See https://kizniche.github.io/Mycodo/Error-Codes#error-101 for more info.
2024-08-23 23:41:03,487 - ERROR - mycodo.controllers.controller_input_f2982e8f - StopIteration raised 3 times. Possibly could not read input. Ensure it's connected properly and detected. | open | 2024-08-24T03:15:25Z | 2024-09-20T19:59:44Z | https://github.com/kizniche/Mycodo/issues/1393 | [] | mfittipaldi90 | 27 |
miguelgrinberg/microblog | flask | 41 | Chapter 5: User Logins - app/models.py - can't find generate_password_hash ... | In app/models.py:
The import:
```from werkzeug import generate_password_hash, check_password_hash```
should be:
```from werkzeug.security import generate_password_hash, check_password_hash```
In order to prepare for V1.0 of werkzeug, the automatic redirect was removed.
| closed | 2017-12-11T17:36:07Z | 2017-12-12T02:24:28Z | https://github.com/miguelgrinberg/microblog/issues/41 | [
"question"
] | dastagg | 7 |
ultralytics/ultralytics | deep-learning | 19,120 | -- | closed | 2025-02-07T07:12:44Z | 2025-02-10T14:00:12Z | https://github.com/ultralytics/ultralytics/issues/19120 | [
"documentation",
"detect"
] | sabira-mcw | 2 | |
PeterL1n/BackgroundMattingV2 | computer-vision | 61 | Add an option to use a still image as target background | In example, a `--image-target-bgr` to use with a PNG image.
Or create a `--target-bgr` option that allows for both video and image. | open | 2021-02-12T10:21:48Z | 2021-02-12T10:21:48Z | https://github.com/PeterL1n/BackgroundMattingV2/issues/61 | [] | Peque | 0 |
pyro-ppl/numpyro | numpy | 1,831 | saving render_model() output to the desired file path | When calling `numpyro.render_model(model, filename=my_path)` I was expecting the generated file to be saved in `my_path`. But it does not.
Reading the source code, it seems that it deliberately use `filename.stem` instead of user provided file path. Is this the intended behavior? Unable to save the file to the path user requested make the filename option useless.
```python
def render_model(
model,
model_args=None,
model_kwargs=None,
filename=None,
render_distributions=False,
render_params=False,
):
"""
Wrap all functions needed to automatically render a model.
.. warning:: This utility does not support the
:func:`~numpyro.contrib.control_flow.scan` primitive.
If you want to render a time-series model, you can try
to rewrite the code using Python for loop.
:param model: Model to render.
:param model_args: Positional arguments to pass to the model.
:param model_kwargs: Keyword arguments to pass to the model.
:param str filename: File to save rendered model in.
:param bool render_distributions: Whether to include RV distribution annotations in the plot.
:param bool render_params: Whether to show params in the plot.
"""
relations = get_model_relations(
model,
model_args=model_args,
model_kwargs=model_kwargs,
)
graph_spec = generate_graph_specification(relations, render_params=render_params)
graph = render_graph(graph_spec, render_distributions=render_distributions)
if filename is not None:
filename = Path(filename)
graph.render(
filename.stem, view=False, cleanup=True, format=filename.suffix[1:]
) # remove leading period from suffix
return graph
``` | closed | 2024-07-08T19:23:31Z | 2024-09-16T23:47:48Z | https://github.com/pyro-ppl/numpyro/issues/1831 | [
"bug",
"help wanted"
] | znwang25 | 5 |
onnx/onnx | scikit-learn | 5,807 | Function infrastructure support for multiple target opset versions | # Bug Report
### Describe the bug
See [this issue](https://github.com/microsoft/onnxruntime/issues/16438) for more context and background.
(i) `SetContextDependentFunctionBodyBuilder` has a default value for the opset. But [the default behavior](https://github.com/onnx/onnx/blob/8a980683df9acbcb82dc3385fc7eb8cce4ed840f/onnx/defs/schema.cc#L719), which is to use the op's own opset-version, works only for ops in ONNX standard domain. For an op in a custon domain opset version 1, using ONNX opset version 1 as the default value doesn't make sense. So, we should either eliminate the use of the default-value, or use it only for ONNX standard domain, throwing an error/warning for other domains.
(ii) We should do something similar for [context-independent functions](https://github.com/onnx/onnx/blob/8a980683df9acbcb82dc3385fc7eb8cce4ed840f/onnx/defs/schema.cc#L814)
(iii) Yet another fix required in ONNX is that [BuildContextDependentFunction](https://github.com/onnx/onnx/blob/e9871305d4421ac9836e5fd2be2dc89bc0bb9dcf/onnx/defs/schema.cc#L748) invokes validation, but returns true even if validation fails.
### Notes
<!-- Any additional information -->
| closed | 2023-12-14T23:52:03Z | 2025-01-26T06:42:45Z | https://github.com/onnx/onnx/issues/5807 | [
"bug",
"stale",
"contributions welcome"
] | gramalingam | 0 |
plotly/dash | data-science | 2,813 | How does dash combine with flask jwt? | My previous project used flask jwt for authentication. After switching to dash, how can I support jwt? | closed | 2024-03-25T08:31:51Z | 2024-04-02T18:01:37Z | https://github.com/plotly/dash/issues/2813 | [] | jaxonister | 1 |
flairNLP/fundus | web-scraping | 344 | [Bug]: Fundus not installing on Google Colab | ### Describe the bug
I tried running the tutorial code in a fresh [colab environment](https://colab.research.google.com ), but when running
```
pip install fundus
```
the installation fails with the output
```
google-colab 1.0.0 requires requests==2.31.0, but you have requests 2.28.2 which is incompatible.
yfinance 0.2.36 requires requests>=2.31, but you have requests 2.28.2 which is incompatible.
```
### How to reproduce
```python
pip install fundus
from fundus import PublisherCollection, Crawler
# initialize the crawler for news publishers based in the US
crawler = Crawler(PublisherCollection.us)
# crawl 2 articles and print
for article in crawler.crawl(max_articles=2):
print(article)
```
### Expected behavior.
It installs correctly and I can run all tutorials on Google Colab.
### Logs and Stack traces
_No response_
### Screenshots
_No response_
### Additional Context
_No response_
### Environment
```markdown
Python 3.10.12, CPU-only, Google Colab
```
| closed | 2024-02-01T10:30:58Z | 2024-04-18T10:28:25Z | https://github.com/flairNLP/fundus/issues/344 | [
"bug"
] | alanakbik | 5 |
modin-project/modin | data-science | 6,641 | BUG: passing in large data object to DataFrame apply() function resulting in SegFault | ### Modin version checks
- [X] I have checked that this issue has not already been reported.
- [X] I have confirmed this bug exists on the latest released version of Modin.
- [ ] I have confirmed this bug exists on the main branch of Modin. (In order to do this you can follow [this guide](https://modin.readthedocs.io/en/stable/getting_started/installation.html#installing-from-the-github-master-branch).)
### Reproducible Example
```python
import modin.pandas as pd
import numpy as np
n_features = 20000
n_samples = 20000
features = [f'f{i+1}' for i in range(n_features)]
samples = [f's{i+1}' for i in range(n_samples)]
df = pd.DataFrame(np.random.rand(n_samples, n_features), columns=features, index=samples).round(2)
# mat_df = pd.DataFrame(np.random.rand(n_features, n_features), columns=features, index=features).round(2)
# res_df = df.dot(mat_df)
# this can be simplified to:
mat_arr = np.random.rand(n_features, n_features)
res_df = df.dot(mat_arr)
```
### Issue Description
Getting `Segmentation fault (core dumped)` when doing matrix multiplication on 20K x 20K matrices.
I have two servers, each with 16 threads, >50GB available RAM, >300GB available storage. Code tested on both py@3.8+modin@0.23.4 and py@3.10+modin@0.24.1, both showed same result.
The code is trying to do a dot() on two 20K x 20K matrices, each matrix is roughly 3GB (showing on object store memory in one of the nodes), the seg fault happens when calling `df.dot(mat_df)`. However, the code will work on small matrices like 500 x 500.
There's not much to show on the terminal, basically it just says `UserWarning: Ray Client is attempting to retrieve a 3.00 GiB object over the network, which may be slow. Consider serializing the object to a file and using S3 or rsync instead.` and the program gets killed.
I have tried to increase my object store memory and also specify a plasma directory, and still getting seg fault, the problem seems to happen after the computation has finished on remote node and being transferred by to client, I saw an increase to 12GB on the object store memory , and around 3GB increase on the client's main memory (when the warning message shows up), right when the main memory increased to 3GB, the program gets killed.
### Expected Behavior
should return the transformed df.
### Error Logs
<details>
```python-traceback
Replace this line with the error backtrace (if applicable).
```
</details>
### Installed Versions
<details>
UserWarning: Setuptools is replacing distutils.
INSTALLED VERSIONS
------------------
commit : 4c01f6464a1fb7a201a4e748c7420e5cb41d16ce
python : 3.10.13.final.0
python-bits : 64
OS : Linux
OS-release : 5.15.0-84-generic
Version : #93~20.04.1-Ubuntu SMP Wed Sep 6 16:15:40 UTC 2023
machine : x86_64
processor : x86_64
byteorder : little
LC_ALL : None
LANG : en_US.UTF-8
LOCALE : en_US.UTF-8
Modin dependencies
------------------
modin : 0.24.1
ray : 2.7.1
dask : 2023.9.3
distributed : 2023.9.3
hdk : None
pandas dependencies
-------------------
pandas : 2.1.1
numpy : 1.26.0
pytz : 2023.3.post1
dateutil : 2.8.2
setuptools : 65.5.0
pip : 23.0.1
Cython : None
pytest : None
hypothesis : None
sphinx : None
blosc : None
feather : None
xlsxwriter : None
lxml.etree : None
html5lib : None
pymysql : None
psycopg2 : None
jinja2 : 3.1.2
IPython : 8.16.1
pandas_datareader : None
bs4 : 4.12.2
bottleneck : None
dataframe-api-compat: None
fastparquet : None
fsspec : 2023.9.2
gcsfs : None
matplotlib : None
numba : None
numexpr : None
odfpy : None
openpyxl : None
pandas_gbq : None
pyarrow : 13.0.0
pyreadstat : None
pyxlsb : None
s3fs : None
scipy : None
sqlalchemy : None
tables : None
tabulate : None
xarray : None
xlrd : None
zstandard : None
tzdata : 2023.3
qtpy : None
pyqt5 : None
</details>
| open | 2023-10-10T21:58:13Z | 2023-10-16T16:04:28Z | https://github.com/modin-project/modin/issues/6641 | [
"bug 🦗",
"Blocked ❌",
"Ray ⚡",
"P2",
"External"
] | SiRumCz | 23 |
plotly/dash | plotly | 2,563 | provide option to set HTML tag attributes | **Is your feature request related to a problem? Please describe.**
I'm part of a small team embedding a Plotly Dash in a larger website that must be accessible. We recently added [Pa11y CI](https://github.com/pa11y/pa11y-ci) accessibility tests to our continuous integration and encountered the error:
```
<html> element must have a lang attribute
(https://dequeuniversity.com/rules/axe/4.7/html-has-lang?application=axeAPI)
(html)
<html><head><style>input:invalid { ...</html>
```
**Describe the solution you'd like**
Ideally, there'd be an option (perhaps a kwarg) to add attributes to the `html` tag, perhaps from a dict? e.g. maybe
```python
app = Dash('example', html_tags={'"lang": "en"})
```
**Describe alternatives you've considered**
Searching the community forum led me to a workaround by passing the `index_string` kwarg, so I now have something like
```python
from dash.dash import _default_index
index_string = _default_index.replace("<html>", "<html lang='en'>")
app = Dash('example', index_string=index_string)
```
which satisfies `pa11y-ci`.
| open | 2023-06-13T12:08:59Z | 2024-08-13T19:34:17Z | https://github.com/plotly/dash/issues/2563 | [
"feature",
"P3"
] | warrickball | 0 |
plotly/dash | data-visualization | 2,372 | [BUG] Background callback returns 502 | Thank you so much for helping improve the quality of Dash!
We do our best to catch bugs during the release process, but we rely on your help to find the ones that slip through.
**Describe your context**
I have some high CPU/RAM efficient workflow that handling by celery and in the same time I have background callback that triggering this workflow and partially doing some CPU-related calculations too.
- replace the result of `pip list | grep dash` below
```
│ dash │ 2.7.0 │ │
│ dash-dv2 │ 1.0.95 │ │
│ fef-dash │ 4.4.4 │ │
│ dash-table │ 5.0.0 │ │
│ dash-core-components │ 2.0.0 │ │
│ dash-html-components │ 2.0.0 │ │
```
- if frontend related, tell us your Browser, Version and OS
- OS: macOS
- Browser Chrome
- Version 108
**Describe the bug**
When I'm starting only one this loading in parallel – all is ok, when more than three (including) – I'm receiving 502 error:
The issue reproducing regularly, not one-time issue and not fixing by restarting containers (we're using k8s: 2 pods of dash app, 2 Redis instances, 2 scalable pods of nginx-ingress).
```
HTTP/1.1 502 Bad Gateway
Date: Thu, 29 Dec 2022 03:15:18 GMT
Content-Type: text/html
Content-Length: 552
Connection: keep-alive
Content-Security-Policy: <HIDDEN>
Referrer-Policy: no-referrer
Strict-Transport-Security: max-age=31536000; includeSubDomains; preload
X-Content-Type-Options: nosniff
X-Frame-Options: SAMEORIGIN
X-XSS-Protection: 1; mode=block
```
**Expected behavior**
Not receiving 502 or with described output.
**Screenshots**
<img width="666" alt="image" src="https://user-images.githubusercontent.com/82237255/209899363-09cf4ec7-6d24-491c-9266-434c9402533e.png">
| closed | 2022-12-29T03:29:51Z | 2024-07-25T13:06:03Z | https://github.com/plotly/dash/issues/2372 | [] | ArtsiomAntropau | 2 |
joke2k/django-environ | django | 60 | Environment variables starting with $ cause a recursion error | This SECRET_KEY declaration causes a maximum recursion depth exceeded error. (Note: I never used this secret key value anywhere since I was just starting the project)
```
SECRET_KEY = env("DJANGO_SECRET_KEY", default='$%ify^q7jg*o7(5me*g%+ae-7_1iy)gey*#eo%3c##-=1d=6mb')
```
Results in:
```
...snip...
File "/Users/chromakey/.virtualenvs/xxxx/lib/python3.4/site-packages/environ/environ.py", line 260, in get_value
value = self.get_value(value, cast=cast, default=default)
File "/Users/chromakey/.virtualenvs/xxxx/lib/python3.4/site-packages/environ/environ.py", line 260, in get_value
value = self.get_value(value, cast=cast, default=default)
File "/Users/chromakey/.virtualenvs/xxxx/lib/python3.4/site-packages/environ/environ.py", line 249, in get_value
value = os.environ[var]
File "/Users/chromakey/.virtualenvs/xxxx/bin/../lib/python3.4/os.py", line 630, in __getitem__
value = self._data[self.encodekey(key)]
RuntimeError: maximum recursion depth exceeded
```
This however, causes no error:
```
SECRET_KEY = env("DJANGO_SECRET_KEY", default='%ify^q7jg*o7(5me*g%+ae-7_1iy)gey*#eo%3c##-=1d=6mb')
```
Obviously not a blocker for me on anything, but thought I'd let you know. Thanks!
| closed | 2015-12-23T18:32:08Z | 2021-12-04T03:40:43Z | https://github.com/joke2k/django-environ/issues/60 | [
"duplicate"
] | chromakey | 8 |
errbotio/errbot | automation | 1,149 | Can't signin to Hipchat with SSO enabled | In order to let us help you better, please fill out the following fields as best you can:
### I am...
* [ ] Reporting a bug
* [ ] Suggesting a new feature
* [x] Requesting help with running my bot
* [ ] Requesting help writing plugins
* [ ] Here about something else
### I am running...
* Errbot version: 5.1.3
* OS version: Linux
* Python version: 3.7
* Using a virtual environment: no
### Issue description
We recently changed our Hipchat login to use SSO, which is preventing the bot from logging in. From what I can tell, it's because Hipchat requires a username, password, and a token. Is there a reason the hipchat login can't use just the token for logging in?
### Steps to reproduce
Enable SSO integration with hipchat, attempt to login.
| closed | 2018-01-03T19:27:12Z | 2018-04-19T22:42:35Z | https://github.com/errbotio/errbot/issues/1149 | [] | ImDevinC | 1 |
xonsh/xonsh | data-science | 5,644 | $XONSH_SUBPROC_OUTPUT_FORMAT affects downstream apps | ## Current Behavior
<!---
For general xonsh issues, please try to replicate the failure using `xonsh --no-rc --no-env`.
Short, reproducible code snippets are highly appreciated.
You can use `$XONSH_SHOW_TRACEBACK=1`, `$XONSH_TRACE_SUBPROC=2`, or `$XONSH_DEBUG=1`
to collect more information about the failure.
-->
Setting `$XONSH_SUBPROC_OUTPUT_FORMAT = 'list_lines'` breaks things when using starship.
It's unclear what the canonical way to use starship is.
If I follow starship's [docs](https://starship.rs/guide/#step-2-set-up-your-shell-to-use-starship) and add `execx($(starship init xonsh))` to `~/.xonshrc`, then try to set `$XONSH_SUBPROC_OUTPUT_FORMAT = 'list_lines'`, xonsh crashes hard:
<details>
```xsh
> $XONSH_SUBPROC_OUTPUT_FORMAT = 'list_lines'
ERROR:root:expected str, got list
Traceback (most recent call last):
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/xonsh/prompt/base.py", line 93, in __call__
toks = self._format_prompt(template=template, **kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/xonsh/shells/ptk_shell/formatter.py", line 46, in _format_prompt
toks = super()._format_prompt(
^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/xonsh/prompt/base.py", line 109, in _format_prompt
for literal, field, spec, conv in xt.FORMATTER.parse(tmpl):
^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/local/Library/Frameworks/Python.framework/Versions/3.12/lib/python3.12/string.py", line 288, in parse
return _string.formatter_parser(format_string)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
TypeError: expected str, got list
xonsh: For full traceback set: $XONSH_SHOW_TRACEBACK = True
TypeError: expected str, got list
Failed to format prompt `<function starship_prompt at 0x1062198a0>`-> <class 'TypeError'>:expected str, got list
ERROR:root:expected str, got list
Traceback (most recent call last):
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/xonsh/prompt/base.py", line 93, in __call__
toks = self._format_prompt(template=template, **kwargs)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/xonsh/shells/ptk_shell/formatter.py", line 46, in _format_prompt
toks = super()._format_prompt(
^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/xonsh/prompt/base.py", line 109, in _format_prompt
for literal, field, spec, conv in xt.FORMATTER.parse(tmpl):
^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/local/Library/Frameworks/Python.framework/Versions/3.12/lib/python3.12/string.py", line 288, in parse
return _string.formatter_parser(format_string)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
TypeError: expected str, got list
xonsh: For full traceback set: $XONSH_SHOW_TRACEBACK = True
TypeError: expected str, got list
Failed to format prompt `<function starship_rprompt at 0x106219940>`-> <class 'TypeError'>:expected str, got list
Traceback (most recent call last):
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/xonsh/main.py", line 512, in main
sys.exit(main_xonsh(args))
^^^^^^^^^^^^^^^^
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/xonsh/main.py", line 558, in main_xonsh
shell.shell.cmdloop()
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/xonsh/shells/ptk_shell/__init__.py", line 415, in cmdloop
line = self.singleline(auto_suggest=auto_suggest)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/xonsh/shells/ptk_shell/__init__.py", line 383, in singleline
line = self.prompter.prompt(**prompt_args)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/prompt_toolkit/shortcuts/prompt.py", line 1035, in prompt
return self.app.run(
^^^^^^^^^^^^^
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/prompt_toolkit/application/application.py", line 1002, in run
return asyncio.run(coro)
^^^^^^^^^^^^^^^^^
File "/opt/local/Library/Frameworks/Python.framework/Versions/3.12/lib/python3.12/asyncio/runners.py", line 194, in run
return runner.run(main)
^^^^^^^^^^^^^^^^
File "/opt/local/Library/Frameworks/Python.framework/Versions/3.12/lib/python3.12/asyncio/runners.py", line 118, in run
return self._loop.run_until_complete(task)
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/opt/local/Library/Frameworks/Python.framework/Versions/3.12/lib/python3.12/asyncio/base_events.py", line 687, in run_until_complete
return future.result()
^^^^^^^^^^^^^^^
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/prompt_toolkit/application/application.py", line 886, in run_async
return await _run_async(f)
^^^^^^^^^^^^^^^^^^^
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/prompt_toolkit/application/application.py", line 739, in _run_async
self._redraw()
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/prompt_toolkit/application/application.py", line 543, in _redraw
self.context.copy().run(run_in_context)
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/prompt_toolkit/application/application.py", line 526, in run_in_context
self.renderer.render(self, self.layout)
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/prompt_toolkit/renderer.py", line 684, in render
screen.draw_all_floats()
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/prompt_toolkit/layout/screen.py", line 262, in draw_all_floats
functions[0][1]()
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/prompt_toolkit/layout/containers.py", line 904, in _draw_float
width = fl.content.preferred_width(write_position.width).preferred
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/prompt_toolkit/layout/containers.py", line 1579, in preferred_width
return self._merge_dimensions(
^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/prompt_toolkit/layout/containers.py", line 1634, in _merge_dimensions
preferred = get_preferred()
^^^^^^^^^^^^^^^
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/prompt_toolkit/layout/containers.py", line 1569, in preferred_content_width
preferred_width = self.content.preferred_width(
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/prompt_toolkit/layout/controls.py", line 347, in preferred_width
text = fragment_list_to_text(self._get_formatted_text_cached())
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
File "/Users/wryfi/.local/pipx/venvs/xonsh/lib/python3.12/site-packages/prompt_toolkit/formatted_text/utils.py", line 73, in fragment_list_to_text
return "".join(item[1] for item in fragments if ZeroWidthEscape not in item[0])
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
TypeError: sequence item 0: expected str instance, list found
Xonsh encountered an issue during launch.
Please report to https://github.com/xonsh/xonsh/issues
Failback to /bin/zsh
```
</details>
If I use [xontrib-prompt-starship](https://github.com/anki-code/xontrib-prompt-starship), my prompt gets corrupted:
<details>
```xsh
> xontrib load prompt_starship
> $XONSH_SUBPROC_OUTPUT_FORMAT = 'list_lines'
['', '\x1b[38;2;214;93;14m\ue0b6\x1b[48;2;214;93;14;38;2;251;241;199m wryfi@beebo \x1b[48;2;215;153;33;38;2;214;93;14m\ue0b0\x1b[38;2;251;241;199m \uf015 \x1b[4
8;2;250;189;47;38;2;215;153;33m\ue0b0\x1b[38;2;251;241;199m xsh \x1b[48;2;104;157;106;38;2;250;189;47m\ue0b0\x1b[48;2;69;133;136;38;2;104;157;106m\ue0b0\x1b[48;2
;102;92;84;38;2;69;133;136m\ue0b0\x1b[48;2;60;56;54;38;2;102;92;84m\ue0b0\x1b[38;2;251;241;199m \uf43a 16:43 \x1b[0m\x1b[38;2;60;56;54m\ue0b4 \x1b[0m', '\x1b[1;3
8;2;104;157;106m\uf101\x1b[0m ']
```
</details>
Either way something seems off, though it is possible I'm doing something wrong as a xonsh newb.
## Expected Behavior
I expect starship to keep working and my prompt to not get corrupted when setting `$XONSH_SUBPROC_OUTPUT_FORMAT = 'list_lines'`.
## xonfig
When activating starship via `~/.xonshrc`:
<details>
```xsh
+-----------------------------+-----------------------+
| xonsh | 0.18.2 |
| Python | 3.12.4 |
| PLY | 3.11 |
| have readline | True |
| prompt toolkit | 3.0.47 |
| shell type | prompt_toolkit |
| history backend | json |
| pygments | 2.18.0 |
| on posix | True |
| on linux | False |
| on darwin | True |
| on windows | False |
| on cygwin | False |
| on msys2 | False |
| is superuser | False |
| default encoding | utf-8 |
| xonsh encoding | utf-8 |
| encoding errors | surrogateescape |
| xontrib | [] |
| RC file 1 | /Users/wryfi/.xonshrc |
| UPDATE_OS_ENVIRON | False |
| XONSH_CAPTURE_ALWAYS | False |
| XONSH_SUBPROC_OUTPUT_FORMAT | stream_lines |
| THREAD_SUBPROCS | True |
| XONSH_CACHE_SCRIPTS | True |
+-----------------------------+-----------------------+
```
</details>
When using xontrib-prompt-starship:
<details>
```xsh
+-----------------------------+-----------------------+
| xonsh | 0.18.2 |
| Python | 3.12.4 |
| PLY | 3.11 |
| have readline | True |
| prompt toolkit | 3.0.47 |
| shell type | prompt_toolkit |
| history backend | json |
| pygments | 2.18.0 |
| on posix | True |
| on linux | False |
| on darwin | True |
| on windows | False |
| on cygwin | False |
| on msys2 | False |
| is superuser | False |
| default encoding | utf-8 |
| xonsh encoding | utf-8 |
| encoding errors | surrogateescape |
| xontrib 1 | prompt_starship |
| RC file 1 | /Users/wryfi/.xonshrc |
| UPDATE_OS_ENVIRON | False |
| XONSH_CAPTURE_ALWAYS | False |
| XONSH_SUBPROC_OUTPUT_FORMAT | list_lines |
| THREAD_SUBPROCS | True |
| XONSH_CACHE_SCRIPTS | True |
+-----------------------------+-----------------------+
```
</details>
## For community
⬇️ **Please click the 👍 reaction instead of leaving a `+1` or 👍 comment**
| closed | 2024-08-06T00:10:37Z | 2024-08-06T11:20:52Z | https://github.com/xonsh/xonsh/issues/5644 | [
"stdout",
"subproc",
"downstream"
] | wryfi | 1 |
streamlit/streamlit | data-visualization | 10,656 | Button gets full width if it has a `help`/tooltip | ### Checklist
- [x] I have searched the [existing issues](https://github.com/streamlit/streamlit/issues) for similar issues.
- [x] I added a very descriptive title to this issue.
- [x] I have provided sufficient information below to help reproduce this issue.
### Summary
A button gets full width if it has a `help`/tooltip defined.
### Reproducible Code Example
[](https://issues.streamlitapp.com/?issue=gh-10656)
```Python
import streamlit as st
st.button("A") # Has expected width.
st.button("B", help="tooltip") # Has unexpected 100% width.
```
### Steps To Reproduce
Try the above code example.
### Expected Behavior
Button B should have `width: auto`, just like button A.
### Current Behavior
Button B has `width: 100%`.
<img width="750" alt="Image" src="https://github.com/user-attachments/assets/f3b750b0-461e-4c61-9e8d-b58d2b934c2c" />
### Is this a regression?
- [x] Yes, this used to work in a previous version.
### Debug info
- Streamlit version: 1.43.0
- Python version: 3.11.9
- Operating System: MacOS
- Browser: Chrome
### Additional Information
This is a regression from 1.43.0. It used to work as expected in 1.42.2. | closed | 2025-03-05T21:54:28Z | 2025-03-07T21:20:44Z | https://github.com/streamlit/streamlit/issues/10656 | [
"type:bug",
"type:regression",
"status:confirmed",
"priority:P1"
] | JosephMarinier | 4 |
sczhou/CodeFormer | pytorch | 124 | `--face_upsample` does nothing. | Looking through `inference_codeformer.py`, it seems that the `--face_upsample` flag is disabled for some reason:
https://github.com/sczhou/CodeFormer/blob/4c93a0ac8e309cb186165eba098fb8a379f9e349/inference_codeformer.py#L125-L132
It looks like this happened in [this commit](https://github.com/sczhou/CodeFormer/commit/396b5e1d92022ed776ed3d3174e26fd330ed0f74) enabling CPU support for realesrgan.
Is there a reason this should be disabled or was it just commented out during debugging and not uncommented afterwards? | closed | 2023-01-28T11:39:43Z | 2023-01-30T06:43:07Z | https://github.com/sczhou/CodeFormer/issues/124 | [] | bolshoytoster | 1 |
Yorko/mlcourse.ai | plotly | 375 | Assignment 1: Question 6 answer should be 1 of Olympics dataset. Because of men double medal. | closed | 2018-10-16T08:52:49Z | 2018-10-16T11:32:38Z | https://github.com/Yorko/mlcourse.ai/issues/375 | [
"invalid"
] | HammadZahidAli | 2 | |
newpanjing/simpleui | django | 359 | 对某一列进行排序后,点击下一页会排序会失效 | **bug描述**
对某一列进行排序后,点击下一页会排序会失效
**环境**
* Python Version:3.8
* Django Version:3.2
* SimpleUI Version:2021.4.1
| closed | 2021-04-10T01:54:23Z | 2021-05-11T09:12:50Z | https://github.com/newpanjing/simpleui/issues/359 | [
"bug"
] | gojuukaze | 0 |
lepture/authlib | flask | 341 | update_token with password grant | I'm trying to use [update_token](https://docs.authlib.org/en/latest/client/oauth2.html#refresh-auto-update-token). I've got this code:
```python
def update_token(token, refresh_token=None, access_token=None):
token_var = client.fetch_token(token_url, username=username, password=password)
print('in update_token')
# update old token
client.token = token_var
client = OAuth2Session(client_id, client_secret, scope=scope, update_token=update_token)
token_var = client.fetch_token(token_url, username=username, password=password)
client.token = token_var
for i in range(1, 900):
env = client.get('<MY SECURED ENDPOINT>')
time.sleep(1)
if i % 30 == 0:
print(str(i)+"s")
print(env)
```
If I print the token I see that it has 'expires_in' value of 300. But as we get near to that 'update_token' never gets called. I'm on 0.15.3 and wondering if it's something related to using the password_grant. More fully:
```python
<Response [200]>
180s
<Response [200]>
210s
<Response [200]>
Traceback (most recent call last):
File "password-grant.py", line 28, in <module>
env = client.get('<MY SECURED ENDPOINT>')
File "/usr/lib/python3.9/site-packages/requests/sessions.py", line 555, in get
return self.request('GET', url, **kwargs)
File "/home/ryan/.local/lib/python3.9/site-packages/authlib/integrations/requests_client/oauth2_session.py", line 112, in request
return super(OAuth2Session, self).request(
File "/usr/lib/python3.9/site-packages/requests/sessions.py", line 528, in request
prep = self.prepare_request(req)
File "/usr/lib/python3.9/site-packages/requests/sessions.py", line 456, in prepare_request
p.prepare(
File "/usr/lib/python3.9/site-packages/requests/models.py", line 320, in prepare
self.prepare_auth(auth, url)
File "/usr/lib/python3.9/site-packages/requests/models.py", line 551, in prepare_auth
r = auth(self)
File "/home/ryan/.local/lib/python3.9/site-packages/authlib/integrations/requests_client/oauth2_session.py", line 34, in __call__
self.ensure_active_token()
File "/home/ryan/.local/lib/python3.9/site-packages/authlib/integrations/requests_client/oauth2_session.py", line 31, in ensure_active_token
raise InvalidTokenError()
authlib.integrations.base_client.errors.InvalidTokenError: token_invalid:
```
| open | 2021-05-04T17:43:14Z | 2025-02-21T09:01:40Z | https://github.com/lepture/authlib/issues/341 | [
"client"
] | ryandawsonuk | 1 |
pydantic/FastUI | pydantic | 218 | Unable to pass a BaseModel into component from subfolder (Pydantic V1 issue) | I was attempting to create a simple pydantic object as below
```
class User(BaseModel):
name: str = Field(title='Name', max_lengh=200)
```
But I face some issues:
### Scenario 1 - Having the Pydantic Base Model define on the same page as entry point (i.e. `main.py`)
In this scenario, the BaseModel is implemented on the same file as the entry point file as below:
```
component = [
# Blah, other components
c.ModelForm(model=User),
# Blah....
]
```
This works fine and it shows the form as expected on UI.
Here's the problem:
### Scenario 2 - The Pydantic object is imported from somewhere else
```
from model.blah import User
...
component = [
# Blah, other components
c.ModelForm(model=User),
# Blah....
]
```
When I run this, I receive an exception of
```
Input should be a subclass of BaseModel [type=is_subclass_of, input_value=<class 'model.blah.User'>, input_type=ModelMetaclass]
For further information visit https://errors.pydantic.dev/2.6/v/is_subclass_of
```
Not entirely sure what happened here, will continue to investigate but will raise this problem I bumped into. | closed | 2024-02-26T13:57:51Z | 2024-02-26T14:26:54Z | https://github.com/pydantic/FastUI/issues/218 | [] | edwardmfho | 2 |
ageitgey/face_recognition | machine-learning | 1,290 | OpenCL | Hello everyone, how do I run the code on the embedded GPU? I understand correctly that I should use OpenCL? | open | 2021-03-05T12:10:56Z | 2021-04-22T12:59:54Z | https://github.com/ageitgey/face_recognition/issues/1290 | [] | RarDay | 2 |
docarray/docarray | pydantic | 1,780 | Release Note | # Release Note
This release contains 3 bug fixes and 4 documentation improvements, including 1 breaking change.
## 💥 Breaking Changes
### Changes to the return type of `DocList.to_json()` and `DocVec.to_json()`
In order to make the `to_json` method consistent across different classes, we changed its return type in `DocList` and `DocVec` to `str`.
This means that, if you use this method in your application, make sure to update your codebase to expect `str` instead of `bytes`.
## 🐞 Bug Fixes
### Make DocList.to_json() and DocVec.to_json() return str instead of bytes (#1769)
This release changes the return type of the methods `DocList.to_json()` and `DocVec.to_json()` in order to be consistent with `BaseDoc .to_json()` and other pydantic models. After this release, these methods will return `str ` type data instead of `bytes`.
💥 Since the return type is changed, this is considered a breaking change.
### Casting in reduce before appending (#1758)
This release introduces type casting internally in the `reduce `helper function, casting its inputs before appending them to the final result. This will make it possible to reduce documents whose schemas are compatible but not exactly the same.
### Skip doc attributes in `__annotations__` but not in `__fields__` (#1777)
This release fixes an issue in the create_pure_python_type_model helper function. Starting with this release, only attributes in the class `__fields__` will be considered during type creation.
The previous behavior broke applications when users introduced a ClassVar in an input class:
```python
class MyDoc(BaseDoc):
endpoint: ClassVar[str] = "my_endpoint"
input_test: str = ""
```
```text
field_info = model.__fields__[field_name].field_info
KeyError: 'endpoint'
```
Kudos to @NarekA for raising the issue and contributing a fix in the Jina project, which was ported in DocArray.
## 📗 Documentation Improvements
- Explain how to set Document config (#1773)
- Add workaround for torch compile (#1754)
- Add note about pickling dynamically created Doc class (#1763)
- Improve the docstring of `filter_docs` (#1762)
## 🤟 Contributors
We would like to thank all contributors to this release:
- Sami Jaghouar (@samsja )
- Johannes Messner (@JohannesMessner )
- AlaeddineAbdessalem (@alaeddine-13 )
- Joan Fontanals (@JoanFM ) | closed | 2023-09-07T07:09:57Z | 2023-09-07T13:50:39Z | https://github.com/docarray/docarray/issues/1780 | [] | JoanFM | 5 |
graphql-python/graphene-django | django | 1,503 | SerializerMutation return types are incompatible with the relay updater api | **Is your feature request related to a problem? Please describe.**
In relay, mutations use a store updater that has the ability to update automatically the store, without the need to write an updater function, if the mutation object return type is provided in a shape undersood by relay. The current implementation of the SerializerMutation is incompatible with this api.
In particular, reading the latest [useMutation](https://relay.dev/docs/api-reference/use-mutation/) relay hook docs ([additional explanation here](https://relay.dev/docs/tutorial/mutations-updates/#how-relay-automatically-processes-mutation-responses)), I saw ` If updater is not provided, by default, Relay will know to automatically update the fields on the records referenced in the mutation response;`. The mutation default return type automatically generated (MyMutationPayload)is not an object recognized by relay for an automatic update of the store.
**Describe the solution you'd like**
Provided that the `SerializerMutation` already implements the `clientMutationId` api,
the cleanest would be simply to change the in `SerializerMutation` the mutation return type by doing the following : first providing a registry to the `__init_subclass_with__meta__` and reading from it in the `_meta` initialization, like this is done in the `DjangoObjectType`. The registry, is then used to fetch the Node corresponding to the `model_class` also available in the _meta defintion to create the return type. Something like :
```python
if not _meta:
_meta = RelaySerializerMutationOptions(cls)
_meta.lookup_field = lookup_field
_meta.model_operations = model_operations
_meta.serializer_class = serializer_class
_meta.model_class = model_class
if registry:
_meta.registry = registry
node_type = registry.get_type_for_model(model_class)
_meta.node_type = node_type
new_output_fields = OrderedDict()
new_output_fields["instance"] = graphene.Field(
node_type, description="The created/updated instance."
)
_meta.fields = yank_fields_from_attrs(new_output_fields, _as=Field)
```
Also, this would require the following change in `perform_mutate` method.
```python
@classmethod
def perform_mutate(cls, serializer, info):
obj = serializer.save()
kwargs = {}
for f, field in serializer.fields.items():
if not field.write_only:
if isinstance(field, serializers.SerializerMethodField):
kwargs[f] = field.to_representation(obj)
else:
kwargs[f] = field.get_attribute(obj)
if cls._meta.registry:
node_type = cls._meta.node_type
kwargs["id"] = to_global_id(node_type.__name__, obj.pk)
instance = node_type(**kwargs)
# without it it breaks (NodeType has no attribute pk. ).
instance.pk = obj.pk
return cls(errors=None, instance=instance)
return cls(errors=None, **kwargs)
```
The code above is working.
This solution however, implies a breaking change since we would not be spreading anymore the model instance alongside the clientMutationId, but in a separate relay node (`instance` in the snippet above).
In the two snippets, the `if registry:` tests are for rapid prototyping only, I would imagine an implementation where this is the default behaviour.
**Describe alternatives you've considered**
Other solutions would be less elegant, one would be to create a children class of SerializerMutation that implements this behaviour, this would not have the downside of being a breaking change rather than an opt in. However, it does also require passing the register to the SerializerMutation, which also needs to be changed to receive it as a param, but does not uses it itself.
Another one I considered early was to simply change the ID returned in the MutationPayload to a globalId. I managed to do the change, and I got an interesting result with useMutation. Instead of the store of relay not updating, it would update to a blank item - even if the graphql mutation payload received contained the data. This means that the automatic store updater mentioned in the introduction does indeed require a globalId as mentionned in the introduction, but also the __typename to be matching.
**Additional context**
Thank you for reading this ! I would enjoy reading your thoughts if you have any. Implementing this changewould ensure greater compatibility with the relay api with the downside of being a breaking change. I would be happy to provide a PR for consideration if the change appears relevant. | open | 2024-03-05T02:26:08Z | 2024-04-10T08:58:16Z | https://github.com/graphql-python/graphene-django/issues/1503 | [
"✨enhancement"
] | advl | 0 |
python-restx/flask-restx | api | 264 | No validation error thrown when strict mode for validating model fields set to true | When I pass a spurious parameter not defined in my model where the strict parameter is set to true and the method is decorated with api.expect(model, validate=True) no validation error is thrown. Is my implementation incorrect or is this a bug?
I have a simple Users resource with a put method to update all user information except user password. According to Flask-Restx docs when a model has set the strict and validation params to true, a validation error will be thrown if an unspecified param is provided in the request. However, this doesn't seem to be working for me.
Api instantiation:
```python
from flask import Blueprint
from flask_restx import Api
api_bp = Blueprint(
"api", __name__, url_prefix="/api/v0.1"
) # Used to add url prefix to all routes
authorizations = {"Bearer": {"type": "apiKey", "in": "header", "name": "Authorization"}}
api = Api(
api_bp,
version="0.1",
title="Asteroid Hotels API",
description="Welcome to the Asteroid Hotels api documentation site!",
doc="/docs",
authorizations=authorizations,
)
```
Model definition:
```python
from flask_restx import Namespace, Resource, fields, marshal
users_ns = Namespace("users")
user = users_ns.model(
"user",
{
"user_name": fields.String(example="some_user", required=True),
"email": fields.String(example="some.user@email", required=True),
"is_admin": fields.Boolean(example="False"),
"is_deactivated": fields.Boolean(example="False"),
"created_date": fields.DateTime(example="2020-12-01T01:59:39.297904"),
"last_modified_date": fields.DateTime(example="2020-12-01T01:59:39.297904"),
"uri": fields.Url("api.user"),
},
strict=True,
)
user_post = users_ns.inherit(
"user_post", user, {"password": fields.String(required=True)}
) # Used for when
```
Resource and method definition:
```python
from api.models import Users
class User(Resource):
@users_ns.marshal_with(user)
@users_ns.expect(user, validate=True)
def put(self, id):
"""
Update a specified user.
"""
user = Users.query.get_or_404(id)
body = request.get_json()
user.update(body)
return user
```
Failing Test:
```python
def test_update_user_invalid_password_param(self, client, db):
""" User endpoint should return 400 when user attempts to pass password param to update. """
data = {
"user_name": "some_user",
"email": "some.user@email.com",
"password": "newpassword",
}
response = client.put(url_for("api.user", id=1), json=data)
assert response.status_code == 400
```
The response.status_code here is 200 because no validation error is thrown for the unspecified password param passed in the body of the request.
@maho am I using the strict param improperly? Am I misunderstanding the behavior of strict?
UPDATED: I've added the test for strict model param from Flask-RestX repo (can be found [here](https://github.com/python-restx/flask-restx/blob/master/tests/test_namespace.py)) for more context on expected behavior:
```python
def test_api_payload_strict_verification(self, app, client):
api = restx.Api(app, validate=True)
ns = restx.Namespace("apples")
api.add_namespace(ns)
fields = ns.model(
"Person",
{
"name": restx.fields.String(required=True),
"age": restx.fields.Integer,
"birthdate": restx.fields.DateTime,
},
strict=True,
)
@ns.route("/validation/")
class Payload(restx.Resource):
payload = None
@ns.expect(fields)
def post(self):
Payload.payload = ns.payload
return {}
data = {
"name": "John Doe",
"agge": 15, # typo
}
resp = client.post_json("/apples/validation/", data, status=400)
assert re.match("Additional properties are not allowed \(u*'agge' was unexpected\)", resp["errors"][""])
```
### **Environment**
- Python 3.7.5
Flask==1.1.2
Flask-restx==0.2.0
| closed | 2020-12-17T17:47:24Z | 2020-12-18T15:55:44Z | https://github.com/python-restx/flask-restx/issues/264 | [
"bug"
] | clpatterson | 2 |
plotly/dash | data-science | 3,151 | add dash-pivottable to core components | We should move https://github.com/plotly/dash-pivottable into this repository as a component. | open | 2025-02-06T19:01:19Z | 2025-02-06T19:01:19Z | https://github.com/plotly/dash/issues/3151 | [
"feature",
"P2"
] | gvwilson | 0 |
drivendataorg/cookiecutter-data-science | data-science | 139 | Persisting large models | Are large models (say, between 600MB to 2GB) packaged with the code, or persisted somewhere else?
If they are persisted somewhere else, does that invalidate the `src/models/predict_model.py` method? | closed | 2018-08-21T19:02:15Z | 2019-01-29T17:52:57Z | https://github.com/drivendataorg/cookiecutter-data-science/issues/139 | [] | jtascensao | 1 |
chainer/chainer | numpy | 8,393 | Docs build is warned by Sphinx>=2 | Sphinx (`2.2.1` or `master`) produces the following two kinds of warnings in my environment.
### `duplicate object description`
I think cross refs to such object is ambiguous.
### `autosummary: stub file not found`
There are
- `chainer.dataset.Converter` base class and
- `chainer.dataset.converter` decorator.
Therefore the filesystem has to allow to store `chainer.dataset.Converter.html` and `chainer.dataset.converter.html` to build the docs.
BTW, the warning message is buggy: `.rst` is repeated in `/path/to/chainer/docs/source/reference/datasets.rst.rst:69: WARNING: autosummary: stub file not found 'chainer.dataset.converter'. Check your autosummary_generate setting.`
## Code to reproduce
```
cd docs
make clean
make html
```
### Error messages, stack traces, or logs
```
/path/to/chainer/docs/source/chainermn/model_parallel/index.rst:1: WARNING: duplicate object description of chainermn, other instance in chainermn/installation/index, use :noindex: for one of them
/path/to/chainer/docs/source/chainermn/reference/index.rst:1: WARNING: duplicate object description of chainermn, other instance in chainermn/model_parallel/index, use :noindex: for one of them
/path/to/chainer/docs/source/chainermn/tutorial/index.rst:1: WARNING: duplicate object description of chainermn, other instance in chainermn/reference/index, use :noindex: for one of them
/path/to/chainer/docs/source/chainerx/reference/backprop.rst:4: WARNING: duplicate object description of chainerx, other instance in chainerx/reference/backend_and_device, use :noindex: for one of them
/path/to/chainer/docs/source/chainerx/reference/index.rst:5: WARNING: duplicate object description of chainerx, other instance in chainerx/reference/backprop, use :noindex: for one of them
/path/to/chainer/docs/source/chainerx/reference/ndarray.rst:4: WARNING: duplicate object description of chainerx, other instance in chainerx/reference/index, use :noindex: for one of them
/path/to/chainer/docs/source/chainerx/reference/routines.rst:6: WARNING: duplicate object description of chainerx, other instance in chainerx/reference/ndarray, use :noindex: for one of them
/path/to/chainer/docs/source/reference/datasets.rst.rst:69: WARNING: autosummary: stub file not found 'chainer.dataset.converter'. Check your autosummary_generate setting.
/path/to/chainer/docs/source/reference/links.rst:215: WARNING: duplicate object description of chainer, other instance in reference/index, use :noindex: for one of them
/path/to/chainer/docs/source/reference/serializers.rst:37: WARNING: duplicate object description of chainer, other instance in reference/links, use :noindex: for one of them
```
| closed | 2019-11-08T06:26:27Z | 2020-06-12T07:30:09Z | https://github.com/chainer/chainer/issues/8393 | [
"cat:document",
"stale",
"prio:high"
] | toslunar | 4 |
httpie/cli | api | 752 | Does not handle HTTP informational responses correctly | httpie incorrectly treats informational responses (102-199) as a final response, printing the response headers and exiting.
```
$ http :5000/102
HTTP/1.1 102 Processing
$
```
Compare to what actually happens:
```
$ nc 127.0.0.1 5000
GET /102 HTTP/1.1
HTTP/1.1 102 Processing
HTTP/1.1 200 OK
…
```
httpie should print the zero or more informational responses (if response headers are enabled) before printing the final response and exiting.
| open | 2019-02-12T20:01:44Z | 2021-12-28T12:42:09Z | https://github.com/httpie/cli/issues/752 | [
"bug"
] | daguej | 0 |
absent1706/sqlalchemy-mixins | sqlalchemy | 116 | Exploring the Repository Pattern | I have been experimenting with the repository pattern using these mixins.
Here is the gist https://gist.github.com/aurthurm/0a654760174edc3f27f7ef80b905da72 | closed | 2024-09-01T16:16:59Z | 2024-09-09T14:13:48Z | https://github.com/absent1706/sqlalchemy-mixins/issues/116 | [] | aurthurm | 1 |
SYSTRAN/faster-whisper | deep-learning | 1,165 | how not to get the Smooth text | 
| open | 2024-11-22T09:21:10Z | 2024-11-22T09:21:10Z | https://github.com/SYSTRAN/faster-whisper/issues/1165 | [] | LRY1994 | 0 |
retentioneering/retentioneering-tools | data-visualization | 12 | Bugs with plot_graph() using Safari browser | Nodes labels misalignment. Arrows misplacements | closed | 2020-01-31T13:23:29Z | 2020-10-05T23:28:44Z | https://github.com/retentioneering/retentioneering-tools/issues/12 | [] | tokedo | 1 |
scikit-image/scikit-image | computer-vision | 7,157 | Gallery backlinks are underlined now since pydata-sphinx-theme 0.14 | I just noticed that in or dev docs the [backlinks in gallery examples are currently underlined](https://scikit-image.org/docs/dev/auto_examples/numpy_operations/plot_structuring_elements.html#sphx-glr-auto-examples-numpy-operations-plot-structuring-elements-py). I think that's due to the update to PyData theme 0.14 (was there before https://github.com/scikit-image/scikit-image/pull/7155 got merged). Do we want to override that in our CSS? I don't particularly mind it as it makes the links more visible... | closed | 2023-09-26T19:09:22Z | 2023-10-19T07:53:34Z | https://github.com/scikit-image/scikit-image/issues/7157 | [
":page_facing_up: type: Documentation"
] | lagru | 6 |
AirtestProject/Airtest | automation | 1,073 | Flutter App - how to distinguish icon name or key from the image widget | Airtest IDE: v1.2.14
Android phone: Pixel 4a
Android OS: 12
<img width="624" alt="Screenshot 2022-08-08 at 5 25 14 PM" src="https://user-images.githubusercontent.com/95615371/183386197-6bb1a33a-2577-4395-88a3-f63e9863559c.png">
| open | 2022-08-08T09:29:20Z | 2022-08-08T09:29:20Z | https://github.com/AirtestProject/Airtest/issues/1073 | [] | andychan06 | 0 |
ydataai/ydata-profiling | pandas | 1,639 | stationarity test | ### Missing functionality
It would be nice if the parameters for adfuller could be configured.
### Proposed feature
If you have very large data sets, then using the standard AIC takes too much time and too much RAM.
It would be nice to chose BIC or maxlags.
e.g.
result = adfuller(series.dropna(), autolag='BIC')
or
result = adfuller(series.dropna(), maxlag=30)
### Alternatives considered
_No response_
### Additional context
_No response_ | closed | 2024-08-08T14:12:29Z | 2024-10-28T17:41:37Z | https://github.com/ydataai/ydata-profiling/issues/1639 | [
"feature request 💬"
] | Blackandwhite23 | 1 |
lanpa/tensorboardX | numpy | 17 | Not compatible with tensorflow 1.3? | After I installed tensorflow 1.3 ,tensorboard can be used in terminal
But after I installed tensorboard-pytorch, it can not work:
```
Traceback (most recent call last):
File "/usr/local/bin/tensorboard", line 7, in <module>
from tensorboard.main import main
ImportError: No module named 'tensorboard.main'
``` | closed | 2017-08-22T14:18:57Z | 2021-12-04T08:50:30Z | https://github.com/lanpa/tensorboardX/issues/17 | [] | nsknsl | 13 |
microsoft/nni | tensorflow | 5,642 | documentation regarding environment variable set by nni | **What would you like to be added**:
documentation for environment variables used by nni.
**Why is this needed**:
I was trying to save pytorch model checkpoint while using `nnictl experiment create`. But in the trail script, I couldn't find a proper way to access path to current trial directory. I did some digging into the source code and found out some environment variables were saved in non-public api deep in the library.
**Without this feature, how does current nni work**:
User has to dig into nni source code and find where and what environment variable is set under the hood.
**Components that may involve changes**:
nni documentation and potentially nni api.
**Brief description of your proposal if any**:
add a documentation so that users know what environment variable is set. perhaps add additional helper function to allow easy access to values of environment variables. | open | 2023-07-19T20:00:59Z | 2023-08-11T06:56:08Z | https://github.com/microsoft/nni/issues/5642 | [] | busFred | 0 |
PrefectHQ/prefect | automation | 16,791 | Worker Logs UI not tailing nicely | ### Bug summary
When on the worker logs if you try to scroll down to the most recent logs the view constantly scrolls up a little bit, so that you can't easily sit and read the logs.
See attached video for demonstration.
https://github.com/user-attachments/assets/5b40ad8e-20c7-4ce1-8e48-db5126d0ef77
### Version info
```Text
Prefect workers using 3.1.16
```
### Additional context
_No response_ | open | 2025-01-21T04:30:43Z | 2025-02-10T21:48:59Z | https://github.com/PrefectHQ/prefect/issues/16791 | [] | jfloodnet | 1 |
dmlc/gluon-cv | computer-vision | 1,301 | Typos on the tutorials page of Object Tracking. | 01. Single object tracking with pre-trained SiamRPN models
> You need to prepare two things to start a tracking demo, the video and its first frame object coordinates. The coordinates show the region of interest where to track, and in the format of (center_x, center_y, weight, height).
I think this is correct.
> (corner_x, corner_y, width, height)
Thanks. | closed | 2020-05-12T03:59:25Z | 2020-06-08T05:02:03Z | https://github.com/dmlc/gluon-cv/issues/1301 | [] | dai-ichiro | 3 |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.