text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Interactive Visualization of Inferences (InVizIn)
```
import dash
import dash_table
import pandas as pd
import dash_core_components as dcc
import dash_html_components as html
def createTable(df):
table_header_style = {
"backgroundColor": "rgb(2,21,70)",
"color": "white",
"textAlign": "center",
}
tab = dash_table.DataTable(
id='table',
columns=[{"name": i, "id": i} for i in df.columns],
data=df.to_dict("rows"),
style_header=table_header_style,
style_table={'maxHeight': '40%',
'maxWidth': '50%'},
fixed_rows={ 'headers': True, 'data': 0 },
)
return tab
def createScatter(df):
dataTemplate = dict(
x=df[df.columns[0]],
y=df[df.columns[1]], #text=df.columns[0],
mode='markers',
opacity=0.7,
marker={
'size': 15,
'line': {'width': 0.5, 'color': 'white'}
},
name=df.columns[0])
dataDict = []
dataDict.append(dataTemplate)
scatterGraph = dcc.Graph(
id='scatter',
figure={
'data': dataDict,
'layout': dict(
xaxis={'title': df.columns[0]},
yaxis={'title': df.columns[1]},
hovermode='closest')
}
)
return scatterGraph
df = pd.read_csv('C:\\Users\\212613144\\Repository\\DARPA-ASKE-TA1-Ext\\Datasets\\Force_dataset.csv', header = [0,1])
colNames = []
for i in range(len(df.columns)):
colNames.append(df.columns[i][0])
df.columns = colNames
app = dash.Dash(__name__)
table = createTable(df)
graph = createScatter(df)
app.layout = html.Div(
className="container",
children=[
html.Div([
html.Label(
[
html.Div(["Mass"]),
dcc.Input(
id="mass-input",
placeholder="Enter a value...",
type="number",
value=1.0,
min=0.0,
max=10.0,
),
]
),
html.Label(
[
html.Div(["Acceleration"]),
dcc.Input(
id="acc-input",
placeholder="Enter a value...",
type="number",
value=0.5,
min=-10.0,
max=10.0,
),
]
),
]
),
html.Div(
className="row",
style={},
children=[
html.Div(className="table",
children=[table]
),
html.Div(className="graph",
children=[graph]
)
]
)
]
)
app.run_server(debug=False)
from plotly.subplots import make_subplots
import plotly.graph_objects as go
fig = make_subplots(rows=1, cols=2)
fig.add_trace(
go.Scatter(x=[1, 2, 3], y=[4, 5, 6]),
row=1, col=1
)
fig.add_trace(
go.Scatter(x=[20, 30, 40], y=[50, 60, 70]),
row=1, col=2
)
fig.update_layout(height=600, width=800, title_text="Subplots")
fig.show()
app = dash.Dash(__name__)
graph = createScatter(df)
app.layout = html.Div(children=[graph])
if __name__ == '__main__':
app.run_server(debug=False)
#imports needed for demonstration
#for communicating with services
import requests
#URL to interact with build
url_build = 'http://localhost:12345/darpa/aske/kchain/build'
#URL to interact with append service
url_append = 'http://localhost:12345/darpa/aske/kchain/append'
#URL to interact with evaluate service
url_evaluate = 'http://localhost:12345/darpa/aske/kchain/evaluate'
inputPacket = {
"inputVariables": [
{
"name": "mach",
"type": "float"
},
{
"name": "gamma",
"type": "float",
"value":"1.4"
}
],
"outputVariables": [
{
"name": "aflow",
"type": "float"
},
{
"name": "fac1",
"type": "float"
},
{
"name": "fac2",
"type": "float"
}
],
"equationModel" : """# Utility to get the corrected airflow per area given the Mach number
fac2 = (gamma + 1.0) / (2.0 * (gamma - 1.0))
fac1 = tf.math.pow((1.0 + 0.5 * (gamma - 1.0) * mach * mach), fac2)
number = 0.50161 * tf.math.sqrt(gamma) * mach / fac1
aflow = number
""",
"modelName" : "getAir"
}
r = requests.post(url_build, json=inputPacket)
r.json()
evalPacket = {
"inputVariables": [
{
"name": "mach",
"type": "float",
"value" : "0.9",
#"minValue": "0.0",
#"maxValue": "3.0"
},
{
"name": "gamma",
"type": "float",
"value" : "1.4",
#"minValue": "1.01",
#"maxValue": "2.0"
},
],
"outputVariables": [
{
"name": "aflow",
"type": "float"
},
{
"name": "fac1",
"type": "float"
},
{
"name": "fac2",
"type": "float"
}
],
"modelName" : "getAir"
}
r = requests.post(url_evaluate, json=evalPacket)
r.json()
r.status_code
from plotly.subplots import make_subplots
import plotly.colors as colors
import plotly.graph_objects as go
import pandas as pd
import numpy as np
import copy
def getOutputValue(evalPacket, url_evaluate, inputVal, index):
if inputVal is not None:
evalPacket['inputVariables'][index]['value'] = str(inputVal)
r = requests.post(url_evaluate, json=evalPacket)
rj = r.json()
assert r.status_code == 200
#print(rj)
outVals = []
for ii in range(len(rj['outputVariables'])):
outVals.append(float(rj['outputVariables'][ii]['value'][1:-1]))
return outVals
def getOutputDataframe(evalPacket, url_evaluate, X, index):
df = pd.DataFrame()
for inputVal in X:
outVals = getOutputValue(evalPacket, url_evaluate, inputVal, index)
dat = {}
dat[evalPacket['inputVariables'][index]['name']] = inputVal #X
for ix, outVal in enumerate(outVals):
dat[evalPacket['outputVariables'][ix]['name']] = outVal
df = df.append(dat, ignore_index=True)
return df
def getMinMax(pck, index):
frac = 0.1
if 'minValue' in pck['inputVariables'][index].keys():
MIN = float(pck['inputVariables'][index]['minValue'])
else:
MIN = float(pck['inputVariables'][index]['value'])*(1 - frac)
if 'maxValue' in pck['inputVariables'][index].keys():
MAX = float(pck['inputVariables'][index]['maxValue'])
else:
MAX = float(pck['inputVariables'][index]['value'])*(1 + frac)
return MIN, MAX
def createSensitivityGraphOAT(evalPacket):
pck = copy.deepcopy(evalPacket)
NUM = 20
refDat = {}
for ii, inputVariable in enumerate(pck['inputVariables']):
inVal = float(inputVariable['value'])
refDat[inputVariable['name']]=inVal
outVals = getOutputValue(evalPacket, url_evaluate, inputVal=None, index=None)
for ix, outVal in enumerate(outVals):
refDat[pck['outputVariables'][ix]['name']] = outVal
fig = make_subplots(rows=len(pck['outputVariables']), cols=len(pck['inputVariables']),
shared_xaxes=True, horizontal_spacing=0.075, vertical_spacing=0.05) #shared_yaxes= True, shared_xaxes=True,
for ii, inputVariable in enumerate(pck['inputVariables']):
#only one input is changed while others stay at base
MIN, MAX = getMinMax(pck, ii)
X = np.linspace(MIN, MAX, num=NUM)
df = getOutputDataframe(copy.deepcopy(pck), url_evaluate, X, index = ii)
for jj, outputVariable in enumerate(pck['outputVariables']):
fig.add_trace(go.Scatter(x = X, y=df[outputVariable['name']],
mode="lines",name = inputVariable['name'],
marker = {"size": 10}),
row=jj+1, col=ii+1)
fig.add_trace(go.Scatter(x = [refDat[inputVariable['name']]],
y = [refDat[outputVariable['name']]],
name = "Reference", mode="markers",
marker = {"symbol":"diamond-open","size": 10, "color": "black"}),
row=jj+1, col=ii+1)
if ii == 0:
fig.update_yaxes(title_text=outputVariable['name'], row=jj+1, col=ii+1, hoverformat=".2f")
else:
fig.update_yaxes(row=jj+1, col=ii+1, hoverformat=".2f")
if jj == len(pck['outputVariables'])-1:
fig.update_xaxes(title_text=inputVariable['name'], row=jj+1, col=ii+1, hoverformat=".2f")
else:
fig.update_xaxes(row=jj+1, col=ii+1, hoverformat=".2f")
fig.update_layout(title='Sensitivity: One-at-a-time Parametric Analysis', showlegend=False)
fig.show()
return fig
import dash
import dash_core_components as dcc
app = dash.Dash(
__name__, meta_tags=[{"name": "viewport", "content": "width=device-width"}]
)
fig = createSensitivityGraphOAT(evalPacket)
app.layout = dcc.Graph(figure=fig)
#app.run_server(debug=False, port=7779)
evalPacket = {
"inputVariables": [
{
"name": "u0d",
"type": "float",
"value": "100.0"
},
{
"name": "altd",
"type": "float",
"value": "10000.0"
}
],
"outputVariables": [
{
"name": "fsmach",
"type": "float"
},
{
"name": "a0",
"type": "float"
},
{
"name": "cpair",
"type": "float"
}
],
"modelName" : "getResponse"
}
r = requests.post(url_evaluate, json=evalPacket)
r.json()
```
# Local Sensitivity Analysis with Gradients
```
inputPacket = {
"inputVariables": [
{
"name": "u0d",
"type": "float"
},
{
"name": "altd",
"type": "float"
},
{
"name": "fsmach",
"type": "float"
}
],
"outputVariables": [
{
"name": "grad_fsmach_u0d",
"type": "float"
},
{
"name": "grad_fsmach_altd",
"type": "float"
},
{
"name": "sens_fsmach_u0d",
"type": "float"
},
{
"name": "sens_fsmach_altd",
"type": "float"
}
],
"equationModel" : """
grad_fsmach_u0d = tf.gradients(fsmach, u0d, stop_gradients = [u0d])[0]
grad_fsmach_altd = tf.gradients(fsmach, altd, stop_gradients = [altd])[0]
sens_fsmach_u0d = tf.math.abs(grad_fsmach_u0d)*u0d/fsmach
sens_fsmach_altd = tf.math.abs(grad_fsmach_altd)*altd/fsmach
""",
"modelName" : "grad_fsmach",
"targetModelName" : "getResponse"
}
#send request to build model
r = requests.post(url_append, json=inputPacket)
#see the response
r.json()
evalPacket = {
"inputVariables": [
{
"name": "u0d",
"type": "float",
"value": "100.0"
},
{
"name": "altd",
"type": "float",
"value": "10000.0"
}
],
"outputVariables": [
{
"name": "fsmach",
"type": "float"
},
{
"name": "grad_fsmach_u0d",
"type": "float"
},
{
"name": "grad_fsmach_altd",
"type": "float"
},
{
"name": "sens_fsmach_u0d",
"type": "float"
},
{
"name": "sens_fsmach_altd",
"type": "float"
}
],
"modelName" : "getResponse"
}
r = requests.post(url_evaluate, json=evalPacket)
r.json()
```
| github_jupyter |
<h1> Time series prediction using RNNs, with TensorFlow and Cloud ML Engine </h1>
This notebook illustrates:
<ol>
<li> Creating a Recurrent Neural Network in TensorFlow
<li> Creating a Custom Estimator in tf.estimator
<li> Training on Cloud ML Engine
</ol>
<p>
<h3> Simulate some time-series data </h3>
Essentially a set of sinusoids with random amplitudes and frequencies.
```
import os
PROJECT = 'cloud-training-demos' # REPLACE WITH YOUR PROJECT ID
BUCKET = 'cloud-training-demos-ml' # REPLACE WITH YOUR BUCKET NAME
REGION = 'us-central1' # REPLACE WITH YOUR BUCKET REGION e.g. us-central1
os.environ['TFVERSION'] = '1.8' # Tensorflow version
# for bash
os.environ['PROJECT'] = PROJECT
os.environ['BUCKET'] = BUCKET
os.environ['REGION'] = REGION
%%bash
gcloud config set project $PROJECT
gcloud config set compute/region $REGION
import tensorflow as tf
print(tf.__version__)
import numpy as np
import seaborn as sns
import pandas as pd
SEQ_LEN = 10
def create_time_series():
freq = (np.random.random() * 0.5) + 0.1 # 0.1 to 0.6
ampl = np.random.random() + 0.5 # 0.5 to 1.5
x = np.sin(np.arange(0, SEQ_LEN) * freq) * ampl
return x
for i in range(0, 5):
sns.tsplot( create_time_series() ); # 5 series
def to_csv(filename, N):
with open(filename, 'w') as ofp:
for lineno in range(0, N):
seq = create_time_series()
line = ",".join(map(str, seq))
ofp.write(line + '\n')
to_csv('train.csv', 1000) # 1000 sequences
to_csv('valid.csv', 50)
!head -5 train.csv valid.csv
```
<h2> RNN </h2>
For more info, see:
<ol>
<li> http://colah.github.io/posts/2015-08-Understanding-LSTMs/ for the theory
<li> https://www.tensorflow.org/tutorials/recurrent for explanations
<li> https://github.com/tensorflow/models/tree/master/tutorials/rnn/ptb for sample code
</ol>
Here, we are trying to predict from 9 values of a timeseries, the tenth value.
<p>
<h3> Imports </h3>
Several tensorflow packages and shutil
```
import tensorflow as tf
import shutil
import tensorflow.contrib.metrics as metrics
import tensorflow.contrib.rnn as rnn
```
<h3> Input Fn to read CSV </h3>
Our CSV file structure is quite simple -- a bunch of floating point numbers (note the type of DEFAULTS). We ask for the data to be read BATCH_SIZE sequences at a time. The Estimator API in tf.contrib.learn wants the features returned as a dict. We'll just call this timeseries column 'rawdata'.
<p>
Our CSV file sequences consist of 10 numbers. We'll assume that 9 of them are inputs and we need to predict the last one.
```
DEFAULTS = [[0.0] for x in range(0, SEQ_LEN)]
BATCH_SIZE = 20
TIMESERIES_COL = 'rawdata'
# In each sequence, column index 0 to N_INPUTS - 1 are features, and column index N_INPUTS to SEQ_LEN are labels
N_OUTPUTS = 1
N_INPUTS = SEQ_LEN - N_OUTPUTS
```
Reading data using the Estimator API in tf.estimator requires an input_fn. This input_fn needs to return a dict of features and the corresponding labels.
<p>
So, we read the CSV file. The Tensor format here will be a scalar -- entire line. We then decode the CSV. At this point, all_data will contain a list of scalar Tensors. There will be SEQ_LEN of these tensors.
<p>
We split this list of SEQ_LEN tensors into a list of N_INPUTS Tensors and a list of N_OUTPUTS Tensors. We stack them along the first dimension to then get a vector Tensor for each. We then put the inputs into a dict and call it features. The other is the ground truth, so labels.
```
# Read data and convert to needed format
def read_dataset(filename, mode, batch_size = 512):
def _input_fn():
# Provide the ability to decode a CSV
def decode_csv(line):
# all_data is a list of scalar tensors
all_data = tf.decode_csv(line, record_defaults = DEFAULTS)
inputs = all_data[:len(all_data) - N_OUTPUTS] # first N_INPUTS values
labels = all_data[len(all_data) - N_OUTPUTS:] # last N_OUTPUTS values
# Convert each list of rank R tensors to one rank R+1 tensor
inputs = tf.stack(inputs, axis = 0)
labels = tf.stack(labels, axis = 0)
# Convert input R+1 tensor into a feature dictionary of one R+1 tensor
features = {TIMESERIES_COL: inputs}
return features, labels
# Create list of files that match pattern
file_list = tf.gfile.Glob(filename)
# Create dataset from file list
dataset = tf.data.TextLineDataset(file_list).map(decode_csv)
if mode == tf.estimator.ModeKeys.TRAIN:
num_epochs = None # indefinitely
dataset = dataset.shuffle(buffer_size = 10 * batch_size)
else:
num_epochs = 1 # end-of-input after this
dataset = dataset.repeat(num_epochs).batch(batch_size)
iterator = dataset.make_one_shot_iterator()
batch_features, batch_labels = iterator.get_next()
return batch_features, batch_labels
return _input_fn
```
<h3> Define RNN </h3>
A recursive neural network consists of possibly stacked LSTM cells.
<p>
The RNN has one output per input, so it will have 8 output cells. We use only the last output cell, but rather use it directly, we do a matrix multiplication of that cell by a set of weights to get the actual predictions. This allows for a degree of scaling between inputs and predictions if necessary (we don't really need it in this problem).
<p>
Finally, to supply a model function to the Estimator API, you need to return a EstimatorSpec. The rest of the function creates the necessary objects.
```
LSTM_SIZE = 3 # number of hidden layers in each of the LSTM cells
# Create the inference model
def simple_rnn(features, labels, mode):
# 0. Reformat input shape to become a sequence
x = tf.split(features[TIMESERIES_COL], N_INPUTS, 1)
# 1. Configure the RNN
lstm_cell = rnn.BasicLSTMCell(LSTM_SIZE, forget_bias = 1.0)
outputs, _ = rnn.static_rnn(lstm_cell, x, dtype = tf.float32)
# Slice to keep only the last cell of the RNN
outputs = outputs[-1]
# Output is result of linear activation of last layer of RNN
weight = tf.get_variable("weight", initializer=tf.initializers.random_normal, shape=[LSTM_SIZE, N_OUTPUTS])
bias = tf.get_variable("bias", initializer=tf.initializers.random_normal, shape=[N_OUTPUTS])
predictions = tf.matmul(outputs, weight) + bias
# 2. Loss function, training/eval ops
if mode == tf.estimator.ModeKeys.TRAIN or mode == tf.estimator.ModeKeys.EVAL:
loss = tf.losses.mean_squared_error(labels, predictions)
train_op = tf.contrib.layers.optimize_loss(
loss = loss,
global_step = tf.train.get_global_step(),
learning_rate = 0.01,
optimizer = "SGD")
eval_metric_ops = {
"rmse": tf.metrics.root_mean_squared_error(labels, predictions)
}
else:
loss = None
train_op = None
eval_metric_ops = None
# 3. Create predictions
predictions_dict = {"predicted": predictions}
# 4. Create export outputs
export_outputs = {"predict_export_outputs": tf.estimator.export.PredictOutput(outputs = predictions)}
# 5. Return EstimatorSpec
return tf.estimator.EstimatorSpec(
mode = mode,
predictions = predictions_dict,
loss = loss,
train_op = train_op,
eval_metric_ops = eval_metric_ops,
export_outputs = export_outputs)
```
<h3> Estimator </h3>
Distributed training is launched off using an Estimator. The key line here is that we use tf.estimator.Estimator rather than, say tf.estimator.DNNRegressor. This allows us to provide a model_fn, which will be our RNN defined above. Note also that we specify a serving_input_fn -- this is how we parse the input data provided to us at prediction time.
```
# Create functions to read in respective datasets
def get_train():
return read_dataset(filename = 'train.csv', mode = tf.estimator.ModeKeys.TRAIN, batch_size = 512)
def get_valid():
return read_dataset(filename = 'valid.csv', mode = tf.estimator.ModeKeys.EVAL, batch_size = 512)
# Create serving input function
def serving_input_fn():
feature_placeholders = {
TIMESERIES_COL: tf.placeholder(tf.float32, [None, N_INPUTS])
}
features = {
key: tf.expand_dims(tensor, -1)
for key, tensor in feature_placeholders.items()
}
features[TIMESERIES_COL] = tf.squeeze(features[TIMESERIES_COL], axis = [2])
return tf.estimator.export.ServingInputReceiver(features, feature_placeholders)
# Create custom estimator's train and evaluate function
def train_and_evaluate(output_dir):
estimator = tf.estimator.Estimator(model_fn = simple_rnn,
model_dir = output_dir)
train_spec = tf.estimator.TrainSpec(input_fn = get_train(),
max_steps = 1000)
exporter = tf.estimator.LatestExporter('exporter', serving_input_fn)
eval_spec = tf.estimator.EvalSpec(input_fn = get_valid(),
steps = None,
exporters = exporter)
tf.estimator.train_and_evaluate(estimator, train_spec, eval_spec)
# Run the model
shutil.rmtree('outputdir', ignore_errors = True) # start fresh each time
train_and_evaluate('outputdir')
```
<h3> Standalone Python module </h3>
To train this on Cloud ML Engine, we take the code in this notebook and make a standalone Python module.
```
%%bash
# Run module as-is
echo $PWD
rm -rf outputdir
export PYTHONPATH=${PYTHONPATH}:${PWD}/simplernn
python -m trainer.task \
--train_data_paths="${PWD}/train.csv*" \
--eval_data_paths="${PWD}/valid.csv*" \
--output_dir=outputdir \
--job-dir=./tmp
```
Try out online prediction. This is how the REST API will work after you train on Cloud ML Engine
```
%writefile test.json
{"rawdata_input": [0,0.214,0.406,0.558,0.655,0.687,0.65,0.549,0.393]}
# local predict doesn't work with Python 3 yet.
# %%bash
# MODEL_DIR=$(ls ./outputdir/export/exporter/)
# gcloud ml-engine local predict --model-dir=./outputdir/export/exporter/$MODEL_DIR --json-instances=test.json
```
<h3> Cloud ML Engine </h3>
Now to train on Cloud ML Engine.
```
%%bash
# Run module on Cloud ML Engine
OUTDIR=gs://${BUCKET}/simplernn/model_trained
JOBNAME=simplernn_$(date -u +%y%m%d_%H%M%S)
gsutil -m rm -rf $OUTDIR
gcloud ml-engine jobs submit training $JOBNAME \
--region=$REGION \
--module-name=trainer.task \
--package-path=${PWD}/simplernn/trainer \
--job-dir=$OUTDIR \
--staging-bucket=gs://$BUCKET \
--scale-tier=BASIC \
--runtime-version=1.4 \
-- \
--train_data_paths="gs://${BUCKET}/train.csv*" \
--eval_data_paths="gs://${BUCKET}/valid.csv*" \
--output_dir=$OUTDIR
```
<h2> Variant: long sequence </h2>
To create short sequences from a very long sequence.
```
import tensorflow as tf
import numpy as np
def breakup(sess, x, lookback_len):
N = sess.run(tf.size(x))
windows = [tf.slice(x, [b], [lookback_len]) for b in range(0, N-lookback_len)]
windows = tf.stack(windows)
return windows
x = tf.constant(np.arange(1,11, dtype=np.float32))
with tf.Session() as sess:
print('input=', x.eval())
seqx = breakup(sess, x, 5)
print('output=', seqx.eval())
```
## Variant: Keras
You can also invoke a Keras model from within the Estimator framework by creating an estimator from the compiled Keras model:
```
def make_keras_estimator(output_dir):
from tensorflow import keras
model = keras.models.Sequential()
model.add(keras.layers.Dense(32, input_shape=(N_INPUTS,), name=TIMESERIES_INPUT_LAYER))
model.add(keras.layers.Activation('relu'))
model.add(keras.layers.Dense(1))
model.compile(loss = 'mean_squared_error',
optimizer = 'adam',
metrics = ['mae', 'mape']) # mean absolute [percentage] error
return keras.estimator.model_to_estimator(model)
%%bash
# Run module as-is
echo $PWD
rm -rf outputdir
export PYTHONPATH=${PYTHONPATH}:${PWD}/simplernn
python -m trainer.task \
--train_data_paths="${PWD}/train.csv*" \
--eval_data_paths="${PWD}/valid.csv*" \
--output_dir=${PWD}/outputdir \
--job-dir=./tmp --keras
```
Copyright 2017 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| github_jupyter |
```
# imports
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.datasets.samples_generator import make_regression
from scipy import stats
%matplotlib inline
```
# Basic Concepts
## What is "learning from data"?
> In general **Learning from Data** is a scientific discipline that is concerned with the design and development of algorithms that allow computers to infer (from data) a model that allows *compact representation* (unsupervised learning) and/or *good generalization* (supervised learning).
This is an important technology because it enables computational systems to adaptively improve their performance with experience accumulated from the observed data.
Most of these algorithms are based on the *iterative solution* of a mathematical problem that involves data and model. If there was an analytical solution to the problem, this should be the adopted one, but this is not the case for most of the cases.
So, the most common strategy for **learning from data** is based on solving a system of equations as a way to find a series of parameters of the model that minimizes a mathematical problem. This is called **optimization**.
The most important technique for solving optimization problems is **gradient descend**.
## Preliminary: Nelder-Mead method for function minimization.
The most simple thing we can try to minimize a function $f(x)$ would be to sample two points relatively near each other, and just repeatedly take a step down away from the largest value. This simple algorithm has a severe limitation: it can't get closer to the true minima than the step size.
The Nelder-Mead method dynamically adjusts the step size based off the loss of the new point. If the new point is better than any previously seen value, it **expands** the step size to accelerate towards the bottom. Likewise if the new point is worse it **contracts** the step size to converge around the minima. The usual settings are to half the step size when contracting and double the step size when expanding.
This method can be easily extended into higher dimensional examples, all that's required is taking one more point than there are dimensions. Then, the simplest approach is to replace the worst point with a point reflected through the centroid of the remaining n points. If this point is better than the best current point, then we can try stretching exponentially out along this line. On the other hand, if this new point isn't much better than the previous value, then we are stepping across a valley, so we shrink the step towards a better point.
> See "An Interactive Tutorial on Numerical Optimization": http://www.benfrederickson.com/numerical-optimization/
## Gradient descend (for *hackers*) for function minimization: 1-D
Let's suppose that we have a function $f: \mathbb{R} \rightarrow \mathbb{R}$. For example:
$$f(x) = x^2$$
Our objective is to find the argument $x$ that minimizes this function (for maximization, consider $-f(x)$). To this end, the critical concept is the **derivative**.
The derivative of $f$ of a variable $x$, $f'(x)$ or $\frac{\partial f}{\partial x}$, is a measure of the rate at which the value of the function changes with respect to the change of the variable.
It is defined as the following limit:
$$ f'(x) = \lim_{h \rightarrow 0} \frac{f(x + h) - f(x)}{h} $$
The derivative specifies how to scale a small change in the input in order to obtain the corresponding change in the output:
$$ f(x + h) \approx f(x) + h f'(x)$$
```
# numerical derivative at a point x
def f(x):
return x**2
def fin_dif(x,
f,
h = 0.00001):
'''
This method returns the derivative of f at x
by using the finite difference method
'''
return (f(x+h) - f(x))/h
x = 2.0
print("{:2.4f}".format(fin_dif(x,f)))
```
The limit as $h$ approaches zero, if it exists, should represent the **slope of the tangent line** to $(x, f(x))$.
For values that are not zero it is only an approximation.
> **NOTE**: It can be shown that the “centered difference formula" is better when computing numerical derivatives:
> $$ \lim_{h \rightarrow 0} \frac{f(x + h) - f(x - h)}{2h} $$
> The error in the "finite difference" approximation can be derived from Taylor's theorem and, assuming that $f$ is differentiable, is $O(h)$. In the case of “centered difference" the error is $O(h^2)$.
The derivative tells how to chage $x$ in order to make a small improvement in $f$.
Then, we can follow these steps to decrease the value of the function:
+ Start from a random $x$ value.
+ Compute the derivative $f'(x) = \lim_{h \rightarrow 0} \frac{f(x + h) - f(x - h)}{2h}$.
+ Walk a small step (possibly weighted by the derivative module) in the **opposite** direction of the derivative, because we know that $f(x - h \mbox{ sign}(f'(x)) < f(x)$ for small enough $h$.
The search for the minima ends when the derivative is zero because we have no more information about which direction to move. $x$ is a critical o stationary point if $f'(x)=0$.
+ A **minimum (maximum)** is a critical point where $f(x)$ is lower (higher) than at all neighboring points.
+ There is a third class of critical points: **saddle points**.
If $f$ is a **convex function**, this should be the minimum (maximum) of our functions. In other cases it could be a local minimum (maximum) or a saddle point.
```
x = np.linspace(-15,15,100)
y = x**2
fig, ax = plt.subplots(1, 1)
fig.set_facecolor('#EAEAF2')
plt.plot(x,y, 'r-')
plt.plot([0],[0],'o')
plt.ylim([-10,250])
plt.gcf().set_size_inches((10,3))
plt.grid(True)
ax.text(0,
20,
'Minimum',
ha='center',
color=sns.xkcd_rgb['pale red'],
)
plt.show
x = np.linspace(-15,15,100)
y = -x**2
fig, ax = plt.subplots(1, 1)
fig.set_facecolor('#EAEAF2')
plt.plot(x,y, 'r-')
plt.plot([0],[0],'o')
plt.ylim([-250,10])
plt.gcf().set_size_inches((10,3))
plt.grid(True)
ax.text(0,
-30,
'Maximum',
ha='center',
color=sns.xkcd_rgb['pale red'],
)
plt.show
x = np.linspace(-15,15,100)
y = x**3
fig, ax = plt.subplots(1, 1)
fig.set_facecolor('#EAEAF2')
plt.plot(x,y, 'r-')
plt.plot([0],[0],'o')
plt.ylim([-3000,3000])
plt.gcf().set_size_inches((10,3))
plt.grid(True)
ax.text(0,
400,
'Saddle Point',
ha='center',
color=sns.xkcd_rgb['pale red'],
)
plt.show
```
There are two problems with numerical derivatives:
+ It is approximate.
+ It is very slow to evaluate (two function evaluations: $f(x + h) , f(x - h)$ ).
Our knowledge from Calculus could help!
We know that we can get an **analytical expression** of the derivative for **some** functions.
For example, let's suppose we have a simple quadratic function, $f(x)=x^2−6x+5$, and we want to find the minimum of this function.
#### First approach
We can solve this analytically using Calculus, by finding the derivate $f'(x) = 2x-6$ and setting it to zero:
\begin{equation}
\begin{split}
2x-6 & = & 0 \\
2x & = & 6 \\
x & = & 3 \\
\end{split}
\end{equation}
```
x = np.linspace(-10,20,100)
y = x**2 - 6*x + 5
fig, ax = plt.subplots(1, 1)
fig.set_facecolor('#EAEAF2')
plt.plot(x,y, 'r-')
plt.plot([3],[3**2 - 6*3 + 5],'o')
plt.ylim([-10,250])
plt.gcf().set_size_inches((10,3))
plt.grid(True)
ax.text(3,
10,
'Min: x = 3',
ha='center',
color=sns.xkcd_rgb['pale red'],
)
plt.show
```
#### Second approach
To find the local minimum using **gradient descend**: you start at a random point, and move into the direction of steepest **descent** relative to the derivative:
+ Start from a random $x$ value.
+ Compute the derivative $f'(x)$ analitically.
+ Walk a small step in the **opposite** direction of the derivative.
In this example, let's suppose we start at $x=15$. The derivative at this point is $2×15−6=24$.
Because we're using gradient descent, we need to subtract the gradient from our $x$-coordinate: $f(x - f'(x))$. However, notice that $15−24$ gives us $−9$, clearly overshooting over target of $3$.
```
x = np.linspace(-10,20,100)
y = x**2 - 6*x + 5
start = 15
fig, ax = plt.subplots(1, 1)
fig.set_facecolor('#EAEAF2')
plt.plot(x,y, 'r-')
plt.plot([start],[start**2 - 6*start + 5],'o')
ax.text(start,
start**2 - 6*start + 35,
'Start',
ha='center',
color=sns.xkcd_rgb['blue'],
)
d = 2 * start - 6
end = start - d
plt.plot([end],[end**2 - 6*end + 5],'o')
plt.ylim([-10,250])
plt.gcf().set_size_inches((10,3))
plt.grid(True)
ax.text(end,
start**2 - 6*start + 35,
'End',
ha='center',
color=sns.xkcd_rgb['green'],
)
plt.show
```
To fix this, we multiply the gradient by a step size. This step size (often called **alpha**) has to be chosen carefully, as a value too small will result in a long computation time, while a value too large will not give you the right result (by overshooting) or even fail to converge.
In this example, we'll set the step size to 0.01, which means we'll subtract $24×0.01$ from $15$, which is $14.76$.
This is now our new temporary local minimum: We continue this method until we either don't see a change after we subtracted the derivative step size (or until we've completed a pre-set number of iterations).
```
old_min = 0
temp_min = 15
step_size = 0.01
precision = 0.0001
def f(x):
return x**2 - 6*x + 5
def f_derivative(x):
import math
return 2*x -6
mins = []
cost = []
while abs(temp_min - old_min) > precision:
old_min = temp_min
gradient = f_derivative(old_min)
move = gradient * step_size
temp_min = old_min - move
cost.append((3-temp_min)**2)
mins.append(temp_min)
# rounding the result to 2 digits because of the step size
print("Local minimum occurs at {:3.6f}.".format(round(temp_min,2)))
```
An important feature of gradient descent is that **there should be a visible improvement over time**: In this example, we simply plotted the squared distance from the local minima calculated by gradient descent and the true local minimum, ``cost``, against the iteration during which it was calculated. As we can see, the distance gets smaller over time, but barely changes in later iterations.
```
x = np.linspace(-10,20,100)
y = x**2 - 6*x + 5
x, y = (zip(*enumerate(cost)))
fig, ax = plt.subplots(1, 1)
fig.set_facecolor('#EAEAF2')
plt.plot(x,y, 'r-', alpha=0.7)
plt.ylim([-10,150])
plt.gcf().set_size_inches((10,3))
plt.grid(True)
plt.show
x = np.linspace(-10,20,100)
y = x**2 - 6*x + 5
fig, ax = plt.subplots(1, 1)
fig.set_facecolor('#EAEAF2')
plt.plot(x,y, 'r-')
plt.ylim([-10,250])
plt.gcf().set_size_inches((10,3))
plt.grid(True)
plt.plot(mins,cost,'o', alpha=0.3)
ax.text(start,
start**2 - 6*start + 25,
'Start',
ha='center',
color=sns.xkcd_rgb['blue'],
)
ax.text(mins[-1],
cost[-1]+20,
'End (%s steps)' % len(mins),
ha='center',
color=sns.xkcd_rgb['blue'],
)
plt.show
```
## From derivatives to gradient: $n$-dimensional function minimization.
Let's consider a $n$-dimensional function $f: \Re^n \rightarrow \Re$. For example:
$$f(\mathbf{x}) = \sum_{n} x_n^2$$
Our objective is to find the argument $\mathbf{x}$ that minimizes this function.
The **gradient** of $f$ is the vector whose components are the $n$ partial derivatives of $f$. It is thus a vector-valued function.
The gradient points in the direction of the greatest rate of **increase** of the function.
$$\nabla {f} = (\frac{\partial f}{\partial x_1}, \dots, \frac{\partial f}{\partial x_n})$$
```
def f(x):
return sum(x_i**2 for x_i in x)
def fin_dif_partial_centered(x,
f,
i,
h=1e-6):
'''
This method returns the partial derivative of the i-th
component of f at x
by using the centered finite difference method
'''
w1 = [x_j + (h if j==i else 0) for j, x_j in enumerate(x)]
w2 = [x_j - (h if j==i else 0) for j, x_j in enumerate(x)]
return (f(w1) - f(w2))/(2*h)
def fin_dif_partial_old(x,
f,
i,
h=1e-6):
'''
This method returns the partial derivative of the i-th
component of f at x
by using the (non-centered) finite difference method
'''
w1 = [x_j + (h if j==i else 0) for j, x_j in enumerate(x)]
return (f(w1) - f(x))/h
def gradient_centered(x,
f,
h=1e-6):
'''
This method returns the gradient vector of f at x
by using the centered finite difference method
'''
return[round(fin_dif_partial_centered(x,f,i,h), 10) for i,_ in enumerate(x)]
def gradient_old(x,
f,
h=1e-6):
'''
This method returns the the gradient vector of f at x
by using the (non-centered)ç finite difference method
'''
return[round(fin_dif_partial_old(x,f,i,h), 10) for i,_ in enumerate(x)]
x = [1.0,1.0,1.0]
print('{:.6f}'.format(f(x)), gradient_centered(x,f))
print('{:.6f}'.format(f(x)), gradient_old(x,f))
```
The function we have evaluated, $f({\mathbf x}) = x_1^2+x_2^2+x_3^2$, is $3$ at $(1,1,1)$ and the gradient vector at this point is $(2,2,2)$.
Then, we can follow this steps to maximize (or minimize) the function:
+ Start from a random $\mathbf{x}$ vector.
+ Compute the gradient vector.
+ Walk a small step in the opposite direction of the gradient vector.
> It is important to be aware that gradient computation is very expensive: if $\mathbf{x}$ has dimension $n$, we have to evaluate $f$ at $2*n$ points.
### How to use the gradient.
$f(x) = \sum_i x_i^2$, takes its mimimum value when all $x$ are 0.
Let's check it for $n=3$:
```
def euc_dist(v1,v2):
import numpy as np
import math
v = np.array(v1)-np.array(v2)
return math.sqrt(sum(v_i ** 2 for v_i in v))
```
Let's start by choosing a random vector and then walking a step in the opposite direction of the gradient vector. We will stop when the difference between the new solution and the old solution is less than a tolerance value.
```
# choosing a random vector
import random
import numpy as np
x = [random.randint(-10,10) for i in range(3)]
x
def step(x,
grad,
alpha):
'''
This function makes a step in the opposite direction of
the gradient vector
in order to compute a new value for the target function.
'''
return [x_i - alpha * grad_i for x_i, grad_i in zip(x,grad)]
tol = 1e-15
alpha = 0.01
while True:
grad = gradient_centered(x,f)
next_x = step(x,grad,alpha)
if euc_dist(next_x,x) < tol:
break
x = next_x
print([round(i,10) for i in x])
```
### Alpha
The step size, **alpha**, is a slippy concept: if it is too small we will slowly converge to the solution, if it is too large we can diverge from the solution.
There are several policies to follow when selecting the step size:
+ Constant size steps. In this case, the size step determines the precision of the solution.
+ Decreasing step sizes.
+ At each step, select the optimal step.
The last policy is good, but too expensive. In this case we would consider a fixed set of values:
```
step_size = [100, 10, 1, 0.1, 0.01, 0.001, 0.0001, 0.00001]
```
## Learning from data
In general, we have:
+ A dataset $(\mathbf{x},y)$ of $n$ examples.
+ A target function $f_\mathbf{w}$, that we want to minimize, representing the **discrepancy between our data and the model** we want to fit. The model is represented by a set of parameters $\mathbf{w}$.
+ The gradient of the target function, $g_f$.
In the most common case $f$ represents the errors from a data representation model $M$. To fit the model is to find the optimal parameters $\mathbf{w}$ that minimize the following expression:
$$ f_\mathbf{w} = \frac{1}{n} \sum_{i} (y_i - M(\mathbf{x}_i,\mathbf{w}))^2 $$
For example, $(\mathbf{x},y)$ can represent:
+ $\mathbf{x}$: the behavior of a "Candy Crush" player; $y$: monthly payments.
+ $\mathbf{x}$: sensor data about your car engine; $y$: probability of engine error.
+ $\mathbf{x}$: finantial data of a bank customer; $y$: customer rating.
> If $y$ is a real value, it is called a *regression* problem.
> If $y$ is binary/categorical, it is called a *classification* problem.
Let's suppose that our model is a one-dimensional linear model $M(\mathbf{x},\mathbf{w}) = w \cdot x $.
### Batch gradient descend
We can implement **gradient descend** in the following way (*batch gradient descend*):
```
import numpy as np
import random
# f = 2x
x = np.arange(10)
y = np.array([2*i for i in x])
# f_target = 1/n Sum (y - wx)**2
def target_f(x,y,w):
return np.sum((y - x * w)**2.0) / x.size
# gradient_f = 2/n Sum 2wx**2 - 2xy
def gradient_f(x,y,w):
return 2 * np.sum(2*w*(x**2) - 2*x*y) / x.size
def step(w,grad,alpha):
return w - alpha * grad
def BGD_multi_step(target_f,
gradient_f,
x,
y,
toler = 1e-6):
'''
Batch gradient descend by using a multi-step approach
'''
alphas = [100, 10, 1, 0.1, 0.001, 0.00001]
w = random.random()
val = target_f(x,y,w)
i = 0
while True:
i += 1
gradient = gradient_f(x,y,w)
next_ws = [step(w, gradient, alpha) for alpha in alphas]
next_vals = [target_f(x,y,w) for w in next_ws]
min_val = min(next_vals)
next_w = next_ws[next_vals.index(min_val)]
next_val = target_f(x,y,next_w)
if (abs(val - next_val) < toler):
return w
else:
w, val = next_w, next_val
print('{:.6f}'.format(BGD_multi_step(target_f, gradient_f, x, y)))
%%timeit
BGD_multi_step(target_f, gradient_f, x, y)
def BGD(target_f,
gradient_f,
x,
y,
toler = 1e-6,
alpha=0.01):
'''
Batch gradient descend by using a given step
'''
w = random.random()
val = target_f(x,y,w)
i = 0
while True:
i += 1
gradient = gradient_f(x,y,w)
next_w = step(w, gradient, alpha)
next_val = target_f(x,y,next_w)
if (abs(val - next_val) < toler):
return w
else:
w, val = next_w, next_val
print('{:.6f}'.format(BGD(target_f, gradient_f, x, y)))
%%timeit
BGD(target_f, gradient_f, x, y)
```
### Stochastic Gradient Descend
The last function evals the whole dataset $(\mathbf{x}_i,y_i)$ at every step.
If the dataset is large, this strategy is too costly. In this case we will use a strategy called **SGD** (*Stochastic Gradient Descend*).
When learning from data, the cost function is additive: it is computed by adding sample reconstruction errors.
Then, we can compute the estimate the gradient (and move towards the minimum) by using only **one data sample** (or a small data sample).
Thus, we will find the minimum by iterating this gradient estimation over the dataset.
A full iteration over the dataset is called **epoch**. During an epoch, data must be used in a random order.
If we apply this method we have some theoretical guarantees to find a good minimum:
+ SGD essentially uses the inaccurate gradient per iteration. Since there is no free food, what is the cost by using approximate gradient? The answer is that the convergence rate is slower than the gradient descent algorithm.
+ The convergence of SGD has been analyzed using the theories of convex minimization and of stochastic approximation: it converges almost surely to a global minimum when the objective function is convex or pseudoconvex, and otherwise converges almost surely to a local minimum.
```
import numpy as np
x = np.arange(10)
y = np.array([2*i for i in x])
data = zip(x,y)
for (x_i,y_i) in data:
print('{:3d} {:3d}'.format(x_i,y_i))
print()
def in_random_order(data):
'''
Random data generator
'''
import random
indexes = [i for i,_ in enumerate(data)]
random.shuffle(indexes)
for i in indexes:
yield data[i]
for (x_i,y_i) in in_random_order(data):
print('{:3d} {:3d}'.format(x_i,y_i))
import numpy as np
import random
def SGD(target_f,
gradient_f,
x,
y,
toler = 1e-6,
epochs=100,
alpha_0=0.01):
'''
Stochastic gradient descend with automatic step adaptation (by
reducing the step to its 95% when there are iterations with
no increase)
'''
data = list(zip(x,y))
w = random.random()
alpha = alpha_0
min_w, min_val = float('inf'), float('inf')
epoch = 0
iteration_no_increase = 0
while epoch < epochs and iteration_no_increase < 100:
val = target_f(x, y, w)
if min_val - val > toler:
min_w, min_val = w, val
alpha = alpha_0
iteration_no_increase = 0
else:
iteration_no_increase += 1
alpha *= 0.95
for x_i, y_i in in_random_order(data):
gradient_i = gradient_f(x_i, y_i, w)
w = w - (alpha * gradient_i)
epoch += 1
return min_w
print('w: {:.6f}'.format(SGD(target_f, gradient_f, x, y)))
```
## Exercise: Stochastic Gradient Descent and Linear Regression
The linear regression model assumes a linear relationship between data:
$$ y_i = w_1 x_i + w_0 $$
Let's generate a more realistic dataset (with noise), where $w_1 = 2$ and $w_0 = 0$.
```
%reset
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.datasets.samples_generator import make_regression
from scipy import stats
import random
%matplotlib inline
# x: input data
# y: noisy output data
x = np.random.uniform(0,1,20)
# f = 2x + 0
def f(x): return 2*x + 0
noise_variance =0.1
noise = np.random.randn(x.shape[0])*noise_variance
y = f(x) + noise
fig, ax = plt.subplots(1, 1)
fig.set_facecolor('#EAEAF2')
plt.xlabel('$x$', fontsize=15)
plt.ylabel('$f(x)$', fontsize=15)
plt.plot(x, y, 'o', label='y')
plt.plot([0, 1], [f(0), f(1)], 'b-', label='f(x)')
plt.ylim([0,2])
plt.gcf().set_size_inches((10,3))
plt.grid(True)
plt.show
```
Complete the following code in order to:
+ Compute the value of $w$ by using a estimator based on minimizing the squared error.
+ Get from SGD function a list, `target_value`, representing the value of the target function at each iteration.
```
# Write your target function as f_target 1/n Sum (y - wx)**2
def target_f(x,y,w):
# your code here
return
# Write your gradient function
def gradient_f(x,y,w):
# your code here
return
def in_random_order(data):
'''
Random data generator
'''
import random
indexes = [i for i,_ in enumerate(data)]
random.shuffle(indexes)
for i in indexes:
yield data[i]
# Modify the SGD function to return a 'target_value' vector
def SGD(target_f,
gradient_f,
x,
y,
toler = 1e-6,
epochs=100,
alpha_0=0.01):
# Insert your code among the following lines
data = zip(x,y)
w = random.random()
alpha = alpha_0
min_w, min_val = float('inf'), float('inf')
iteration_no_increase = 0
epoch = 0
while epoch < epochs and iteration_no_increase < 100:
val = target_f(x, y, w)
if min_val - val > toler:
min_w, min_val = w, val
alpha = alpha_0
iteration_no_increase = 0
else:
iteration_no_increase += 1
alpha *= 0.95
for x_i, y_i in in_random_order(data):
gradient_i = gradient_f(x_i, y_i, w)
w = w - (alpha * gradient_i)
epoch += 1
return min_w
# Print the value of the solution
w, target_value = SGD(target_f, gradient_f, x, y)
print('w: {:.6f}'.format(w))
# Visualize the solution regression line
fig, ax = plt.subplots(1, 1)
fig.set_facecolor('#EAEAF2')
plt.plot(x, y, 'o', label='t')
plt.plot([0, 1], [f(0), f(1)], 'b-', label='f(x)', alpha=0.5)
plt.plot([0, 1], [0*w, 1*w], 'r-', label='fitted line', alpha=0.5, linestyle='--')
plt.xlabel('input x')
plt.ylabel('target t')
plt.title('input vs. target')
plt.ylim([0,2])
plt.gcf().set_size_inches((10,3))
plt.grid(True)
plt.show
# Visualize the evolution of the target function value during iterations.
fig, ax = plt.subplots(1, 1)
fig.set_facecolor('#EAEAF2')
plt.plot(np.arange(target_value.size), target_value, 'o', alpha = 0.2)
plt.xlabel('Iteration')
plt.ylabel('Cost')
plt.grid()
plt.gcf().set_size_inches((10,3))
plt.grid(True)
plt.show()
```
## Mini-batch Gradient Descent
In code, general batch gradient descent looks something like this:
```python
nb_epochs = 100
for i in range(nb_epochs):
grad = evaluate_gradient(target_f, data, w)
w = w - learning_rate * grad
```
For a pre-defined number of epochs, we first compute the gradient vector of the target function for the whole dataset w.r.t. our parameter vector.
**Stochastic gradient descent** (SGD) in contrast performs a parameter update for each training example and label:
```python
nb_epochs = 100
for i in range(nb_epochs):
np.random.shuffle(data)
for sample in data:
grad = evaluate_gradient(target_f, sample, w)
w = w - learning_rate * grad
```
Mini-batch gradient descent finally takes the best of both worlds and performs an update for every mini-batch of $n$ training examples:
```python
nb_epochs = 100
for i in range(nb_epochs):
np.random.shuffle(data)
for batch in get_batches(data, batch_size=50):
grad = evaluate_gradient(target_f, batch, w)
w = w - learning_rate * grad
```
Minibatch SGD has the advantage that it works with a slightly less noisy estimate of the gradient. However, as the minibatch size increases, the number of updates done per computation done decreases (eventually it becomes very inefficient, like batch gradient descent).
There is an optimal trade-off (in terms of computational efficiency) that may vary depending on the data distribution and the particulars of the class of function considered, as well as how computations are implemented.
## Loss Funtions
Loss functions $L(y, f(\mathbf{x})) = \frac{1}{n} \sum_i \ell(y_i, f(\mathbf{x_i}))$ represent the price paid for inaccuracy of predictions in classification/regression problems.
In classification this function is often the **zero-one loss**, that is, $ \ell(y_i, f(\mathbf{x_i}))$ is zero when $y_i = f(\mathbf{x}_i)$ and one otherwise.
This function is discontinuous with flat regions and is thus extremely hard to optimize using gradient-based methods. For this reason it is usual to consider a proxy to the loss called a *surrogate loss function*. For computational reasons this is usually convex function. Here we have some examples:
### Square / Euclidean Loss
In regression problems, the most common loss function is the square loss function:
$$ L(y, f(\mathbf{x})) = \frac{1}{n} \sum_i (y_i - f(\mathbf{x}_i))^2 $$
The square loss function can be re-written and utilized for classification:
$$ L(y, f(\mathbf{x})) = \frac{1}{n} \sum_i (1 - y_i f(\mathbf{x}_i))^2 $$
### Hinge / Margin Loss (i.e. Suport Vector Machines)
The hinge loss function is defined as:
$$ L(y, f(\mathbf{x})) = \frac{1}{n} \sum_i \mbox{max}(0, 1 - y_i f(\mathbf{x}_i)) $$
The hinge loss provides a relatively tight, convex upper bound on the 0–1 Loss.
<img src="https://raw.githubusercontent.com/DataScienceUB/DeepLearningfromScratch2018/master/images/loss_functions.png">
### Logistic Loss (Logistic Regression)
This function displays a similar convergence rate to the hinge loss function, and since it is continuous, simple gradient descent methods can be utilized.
$$ L(y, f(\mathbf{x})) = \frac{1}{n} \sum_i log(1 + exp(-y_i f(\mathbf{x}_i))) $$
### Sigmoid Cross-Entropy Loss (Softmax classifier)
Cross-Entropy is a loss function that is very used for training **multiclass problems**. We'll focus on models that assume that classes are mutually exclusive.
In this case, our labels have this form $\mathbf{y}_i =(1.0,0.0,0.0)$. If our model predicts a different distribution, say $ f(\mathbf{x}_i)=(0.4,0.1,0.5)$, then we'd like to nudge the parameters so that $f(\mathbf{x}_i)$ gets closer to $\mathbf{y}_i$.
C.Shannon showed that if you want to send a series of messages composed of symbols from an alphabet with distribution $y$ ($y_j$ is the probability of the $j$-th symbol), then to use the smallest number of bits on average, you should assign $\log(\frac{1}{y_j})$ bits to the $j$-th symbol.
The optimal number of bits is known as **entropy**:
$$ H(\mathbf{y}) = \sum_j y_j \log\frac{1}{y_j} = - \sum_j y_j \log y_j$$
**Cross entropy** is the number of bits we'll need if we encode symbols by using a wrong distribution $\hat y$:
$$ H(y, \hat y) = - \sum_j y_j \log \hat y_j $$
In our case, the real distribution is $\mathbf{y}$ and the "wrong" one is $f(\mathbf{x}_i)$. So, minimizing **cross entropy** with respect our model parameters will result in the model that best approximates our labels if considered as a probabilistic distribution.
Cross entropy is used in combination with **Softmax** classifier. In order to classify $\mathbf{x}_i$ we could take the index corresponding to the max value of $f(\mathbf{x}_i)$, but Softmax gives a slightly more intuitive output (normalized class probabilities) and also has a probabilistic interpretation:
$$ P(\mathbf{y}_i = j \mid \mathbf{x_i}) = - log \left( \frac{e^{f_j(\mathbf{x_i})}}{\sum_k e^{f_k(\mathbf{x_i})} } \right) $$
where $f_k$ is a linear classifier.
## Advanced gradient descend
### Momentum
SGD has trouble navigating ravines, i.e. areas where the surface curves much more steeply in one dimension than in another, which are common around local optima. In these scenarios, SGD oscillates across the slopes of the ravine while only making hesitant progress along the bottom towards the local optimum.
<img src="https://github.com/DataScienceUB/DeepLearningfromScratch2018/blob/master/images/ridge2.png?raw=true">
Momentum is a method that helps accelerate SGD in the relevant direction and dampens oscillations. It does this by adding a fraction of the update vector of the past time step to the current update vector:
$$ v_t = m v_{t-1} + \alpha \nabla_w f $$
$$ w = w - v_t $$
The momentum $m$ is commonly set to $0.9$.
### Nesterov
However, a ball that rolls down a hill, blindly following the slope, is highly unsatisfactory. We'd like to have a smarter ball, a ball that has a notion of where it is going so that it knows to slow down before the hill slopes up again.
Nesterov accelerated gradient (NAG) is a way to give our momentum term this kind of prescience. We know that we will use our momentum term $m v_{t-1}$ to move the parameters $w$. Computing
$w - m v_{t-1}$ thus gives us an approximation of the next position of the parameters (the gradient is missing for the full update), a rough idea where our parameters are going to be. We can now effectively look ahead by calculating the gradient not w.r.t. to our current parameters $w$ but w.r.t. the approximate future position of our parameters:
$$ w_{new} = w - m v_{t-1} $$
$$ v_t = m v_{t-1} + \alpha \nabla_{w_{new}} f $$
$$ w = w - v_t $$
### Adagrad
All previous approaches manipulated the learning rate globally and equally for all parameters. Tuning the learning rates is an expensive process, so much work has gone into devising methods that can adaptively tune the learning rates, and even do so per parameter.
Adagrad is an algorithm for gradient-based optimization that does just this: It adapts the learning rate to the parameters, performing larger updates for infrequent and smaller updates for frequent parameters.
$$ c = c + (\nabla_w f)^2 $$
$$ w = w - \frac{\alpha}{\sqrt{c}} $$
### RMProp
RMSProp update adjusts the Adagrad method in a very simple way in an attempt to reduce its aggressive, monotonically decreasing learning rate. In particular, it uses a moving average of squared gradients instead, giving:
$$ c = \beta c + (1 - \beta)(\nabla_w f)^2 $$
$$ w = w - \frac{\alpha}{\sqrt{c}} $$
where $\beta$ is a decay rate that controls the size of the moving average.
<img src="https://github.com/DataScienceUB/DeepLearningfromScratch2018/blob/master/images/g1.gif?raw=true">
(Image credit: Alec Radford)
<img src="https://github.com/DataScienceUB/DeepLearningfromScratch2018/blob/master/images/g2.gif?raw=true">
(Image credit: Alec Radford)
```
%reset
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.datasets.samples_generator import make_regression
from scipy import stats
import random
%matplotlib inline
# the function that I'm going to plot
def f(x,y):
return x**2 + 5*y**2
x = np.arange(-3.0,3.0,0.1)
y = np.arange(-3.0,3.0,0.1)
X,Y = np.meshgrid(x, y, indexing='ij') # grid of point
Z = f(X, Y) # evaluation of the function on the grid
plt.pcolor(X, Y, Z, cmap=plt.cm.gist_earth)
plt.axis([x.min(), x.max(), y.min(), y.max()])
plt.gca().set_aspect('equal', adjustable='box')
plt.gcf().set_size_inches((6,6))
plt.show()
def target_f(x):
return x[0]**2.0 + 5*x[1]**2.0
def part_f(x,
f,
i,
h=1e-6):
w1 = [x_j + (h if j==i else 0) for j, x_j in enumerate(x)]
w2 = [x_j - (h if j==i else 0) for j, x_j in enumerate(x)]
return (f(w1) - f(w2))/(2*h)
def gradient_f(x,
f,
h=1e-6):
return np.array([round(part_f(x,f,i,h), 10) for i,_ in enumerate(x)])
def SGD(target_f,
gradient_f,
x,
alpha_0=0.01,
toler = 0.000001):
alpha = alpha_0
min_val = float('inf')
steps = 0
iteration_no_increase = 0
trace = []
while iteration_no_increase < 100:
val = target_f(x)
if min_val - val > toler:
min_val = val
alpha = alpha_0
iteration_no_increase = 0
else:
alpha *= 0.95
iteration_no_increase += 1
trace.append(x)
gradient_i = gradient_f(x, target_f)
x = x - (alpha * gradient_i)
steps += 1
return x, val, steps, trace
x = np.array([2,-2])
x, val, steps, trace = SGD(target_f, gradient_f, x)
print(x)
print('Val: {:.6f}, steps: {:.0f}'.format(val, steps))
def SGD_M(target_f,
gradient_f,
x,
alpha_0=0.01,
toler = 0.000001,
m = 0.9):
alpha = alpha_0
min_val = float('inf')
steps = 0
iteration_no_increase = 0
v = 0.0
trace = []
while iteration_no_increase < 100:
val = target_f(x)
if min_val - val > toler:
min_val = val
alpha = alpha_0
iteration_no_increase = 0
else:
alpha *= 0.95
iteration_no_increase += 1
trace.append(x)
gradient_i = gradient_f(x, target_f)
v = m * v + (alpha * gradient_i)
x = x - v
steps += 1
return x, val, steps, trace
x = np.array([2,-2])
x, val, steps, trace2 = SGD_M(target_f, gradient_f, x)
print('\n',x)
print('Val: {:.6f}, steps: {:.0f}'.format(val, steps))
x2 = np.array(range(len(trace)))
x3 = np.array(range(len(trace2)))
plt.xlim([0,len(trace)])
plt.gcf().set_size_inches((10,3))
plt.plot(x3, trace2)
plt.plot(x2, trace, '-')
```
| github_jupyter |
# The Gibbs phenomenon
The Fourier series of a periodic function with fine jumps presents oscillations close to the discontinuities.
For example, the square wave defined by the odd-extension of $ f(x)=1 $ for $ 0<x<1 $ (namely let $ f(x)=-1 $ for $ -1<x<0 $, and repeat periodically the pattern) is
$$
f(x)=\frac4\pi \sum_{n\in\mathbb{Z_+}} \frac1{2n+1} \sin((2n+1)\pi x)
$$
In this notebook we will explore the Gibbs phenomenon plotting truncated sums of the above series.
```
# preliminary inclusions
import numpy as np
from matplotlib import pyplot as plt
from math import floor
from scipy import linspace
class PartialSum:
def __init__(self, f, n, l=1):
self.n = n
self.l = l
# an index is already shifted: a_1 == an[0], a_k == an[k-1]
self.an = [f(n) for n in range(1,self.n+1)]
def __call__(self, x):
return sum(self.an[n] * np.sin((n+1)*np.pi/self.l*x) for n in range(0,self.n))
def plot_fn(an, ran, steps, l=1, xlim=None, ylim=None, pts=200, orig=None):
x0, xl = ran
x = linspace(x0, xl, pts)
f = PartialSum(an, steps, l)
if xlim or ylim:
plt.figure(figsize=(14,4))
ax1 = plt.subplot(121)
ax1.plot(x, f(x))
ax1.set_title("Truncated to $a_{{{}}}$".format(steps))
if orig:
ax1.plot(x, orig(x), 'r', linewidth=1)
ax2 = plt.subplot(122)
if xlim:
ax2.set_xlim(xlim)
if ylim:
ax2.set_ylim(ylim)
if orig:
ax2.plot(x, orig(x), 'r', linewidth=1)
ax2.plot(x, f(x))
else:
plt.plot(x, f(x))
if orig:
plt.plot(x, orig(x), 'r', linewidth=1)
plt.show()
def an_const_one(n):
"Coefficient a_n = 4/(n*pi) if n is odd and 0 oterwise for the odd extension of f=1 in (0,1)"
return 0.0 if n % 2 == 0 else 4. / (np.pi * n)
orig = lambda x: np.array([1 if floor(x)%2==0 == 0 else -1 for x in x])
plot_fn(an_const_one, (-1,1), 10, orig=orig)
for steps in (20, 30, 50, 100, 1000):
plot_fn(an_const_one, (-1,1), steps, xlim=(0,0.08), ylim=(0.7, 1.3), pts=min(steps*20, 5000), orig=orig)
```
The following is the series for the odd extension of $f(x) = 1-x$ on $(0,1)$:
$$
f(x)=\frac2\pi \sum_{n\in\mathbb{Z_+}} \frac1{n} \sin(n\pi x)
$$
```
def an_one_minus_x(n):
"Coefficient a_n = 2/(n*pi)"
return 2.0/(n*np.pi)
orig = lambda x: np.array([1-x+floor(x) if floor(x)%2==0 == 0 else -x-1-floor(-x) for x in x])
plot_fn(an_one_minus_x, (-1.5,1.5), 10, orig=orig)
for steps in (20, 30, 50, 100, 1000):
plot_fn(an_one_minus_x, (-1.5,1.5), steps, xlim=(-0.1,0.1), ylim=(0.7, 1.3), pts=min(steps*20, 5000), orig=orig)
```
Once `sagemath` will be easily installable, all this will become immensely easier. Indeed one can replicate the first of our plot in `sage` [practically trivially](http://doc.sagemath.org/html/en/reference/functions/sage/functions/piecewise.html#sage.functions.piecewise.PiecewiseFunction.EvaluationMethods.fourier_series_partial_sum):
```python
f2(x) = 1; f1(x) = -1
f = piecewise([[(-1,0),f1],[(0,1),f2]])
S = f.fourier_series_partial_sum(10)
g = plot(S, x, -1, 1, color='blue')
# We did not do this in our plots, but it is
# nice to see how easy it is to do it in sage:
saw(x) = x - 2 * floor((x + 1) / 2)
g += plot(saw(x) / abs(saw(x)), x, -1, 1, color='red')
g
```
Finally, the series for the odd extension of $f(x) = x(\pi - x)$ on $(0,\pi)$:
$$
f(x)=\frac8\pi \sum_{n\in\mathbb{Z_+}} \frac1{(2n+1)^3} \sin((2n+1) x)
$$
where we expect the convergence to be nice and fast.
```
def an_pix_minus_xsq(n):
"Coefficient a_n = 8/(n^3*pi) if n is odd and 0 oterwise for the odd extension of f=1 in (0,1)"
return 0.0 if n % 2 == 0 else 8.0/(n**3*np.pi)
def orig(x):
def f(x):
x = x/np.pi
one_xscaled = 1-x+floor(x) if floor(x)%2==0 else -x-1-floor(-x)
pi_x = np.pi*one_xscaled
x = x-floor(x) if floor(x)%2==0 else -x-floor(-x)
x = np.pi*x
return x*pi_x
return np.array([f(x) for x in x])
plot_fn(an_pix_minus_xsq, (-np.pi,np.pi), 1, l=np.pi, orig=orig)
for steps in (2, 3, 5, 10, 20):
plot_fn(an_pix_minus_xsq, (-np.pi,np.pi), steps, xlim=(-0.1,0.1), ylim=(0.1, -0.1), pts=min(steps*20, 5000), l=np.pi, orig=orig)
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torch.backends.cudnn as cudnn
import torchvision
import torchvision.transforms as transforms
import numpy as np
import os
from matplotlib import pyplot as plt
%matplotlib inline
from torch.utils.data import Dataset, DataLoader
# from create_foreground_background import fg_bg_data,fg_data
# from mini_inception import inception_net
# from resnet import ResNet18,ResNet34,ResNet50
device = 'cuda' if torch.cuda.is_available() else 'cpu'
print(device)
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'plane', 'car', 'bird'}
background_classes = {'cat', 'deer', 'dog', 'frog','horse','ship', 'truck'}
# print(type(foreground_classes))
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(5000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]])
j+=1
else:
image_list.append(foreground_data[fg_idx])
label = foreground_label[fg_idx] # minus 7 because our fore ground classes are 7,8,9 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
desired_num = 30000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(desired_num):
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
batch = 250
msd = MosaicDataset(mosaic_list_of_images, mosaic_label , fore_idx)
train_loader = DataLoader( msd,batch_size= batch ,shuffle=True)
test_images =[] #list of mosaic images, each mosaic image is saved as laist of 9 images
fore_idx_test =[] #list of indexes at which foreground image is present in a mosaic image
test_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(10000):
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx_test.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
test_images.append(image_list)
test_label.append(label)
test_data = MosaicDataset(test_images,test_label,fore_idx_test)
test_loader = DataLoader( test_data,batch_size= batch ,shuffle=False)
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torch.backends.cudnn as cudnn
import torchvision
import torchvision.transforms as transforms
import numpy as np
device = 'cuda' if torch.cuda.is_available() else 'cpu'
class BasicBlock(nn.Module):
expansion = 1
def __init__(self, in_planes, planes, stride=1):
super(BasicBlock, self).__init__()
self.conv1 = nn.Conv2d(
in_planes, planes, kernel_size=3, stride=stride, padding=1, bias=False)
self.bn1 = nn.BatchNorm2d(planes)
self.conv2 = nn.Conv2d(planes, planes, kernel_size=3,
stride=1, padding=1, bias=False)
self.bn2 = nn.BatchNorm2d(planes)
self.shortcut = nn.Sequential()
if stride != 1 or in_planes != self.expansion*planes:
self.shortcut = nn.Sequential(
nn.Conv2d(in_planes, self.expansion*planes,
kernel_size=1, stride=stride, bias=False),
nn.BatchNorm2d(self.expansion*planes)
)
def forward(self, x):
out = F.relu(self.bn1(self.conv1(x)))
out = self.bn2(self.conv2(out))
out += self.shortcut(x)
out = F.relu(out)
return out
class Bottleneck(nn.Module):
expansion = 4
def __init__(self, in_planes, planes, stride=1):
super(Bottleneck, self).__init__()
self.conv1 = nn.Conv2d(in_planes, planes, kernel_size=1, bias=False)
self.bn1 = nn.BatchNorm2d(planes)
self.conv2 = nn.Conv2d(planes, planes, kernel_size=3,
stride=stride, padding=1, bias=False)
self.bn2 = nn.BatchNorm2d(planes)
self.conv3 = nn.Conv2d(planes, self.expansion *
planes, kernel_size=1, bias=False)
self.bn3 = nn.BatchNorm2d(self.expansion*planes)
self.shortcut = nn.Sequential()
if stride != 1 or in_planes != self.expansion*planes:
self.shortcut = nn.Sequential(
nn.Conv2d(in_planes, self.expansion*planes,
kernel_size=1, stride=stride, bias=False),
nn.BatchNorm2d(self.expansion*planes)
)
def forward(self, x):
out = F.relu(self.bn1(self.conv1(x)))
out = F.relu(self.bn2(self.conv2(out)))
out = self.bn3(self.conv3(out))
out += self.shortcut(x)
out = F.relu(out)
return out
class ResNet(nn.Module):
def __init__(self, block, num_blocks, num_classes=10):
super(ResNet, self).__init__()
self.in_planes = 64
self.conv1 = nn.Conv2d(27, 64, kernel_size=3,
stride=1, padding=1, bias=False)
self.bn1 = nn.BatchNorm2d(64)
self.layer1 = self._make_layer(block, 64, num_blocks[0], stride=1)
self.layer2 = self._make_layer(block, 128, num_blocks[1], stride=2)
self.layer3 = self._make_layer(block, 256, num_blocks[2], stride=2)
self.layer4 = self._make_layer(block, 512, num_blocks[3], stride=2)
self.linear = nn.Linear(512*block.expansion, num_classes)
def _make_layer(self, block, planes, num_blocks, stride):
strides = [stride] + [1]*(num_blocks-1)
layers = []
for stride in strides:
layers.append(block(self.in_planes, planes, stride))
self.in_planes = planes * block.expansion
return nn.Sequential(*layers)
def forward(self, x):
out = F.relu(self.bn1(self.conv1(x)))
out = self.layer1(out)
out = self.layer2(out)
out = self.layer3(out)
out = self.layer4(out)
out = F.avg_pool2d(out, 4)
out = out.view(out.size(0), -1)
out = self.linear(out)
return out
def ResNet18(num_classes):
return ResNet(BasicBlock, [2, 2, 2, 2],num_classes=num_classes)
# def ResNet34(num_classes):
# return ResNet(BasicBlock, [3,4,6,3],num_classes=num_classes)
# def ResNet50(num_classes):
# return ResNet(Bottleneck, [3,4,6,3],num_classes=num_classes)
net = ResNet18(9*3)
net = net.to("cuda").double()
if device == 'cuda':
net = torch.nn.DataParallel(net)
cudnn.benchmark = True
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(net.parameters(), lr=0.01,
momentum=0.9, weight_decay=5e-4)
# Training
def train(epoch):
print('\nEpoch: %d' % epoch)
net.train()
train_loss = 0
correct = 0
total = 0
for batch_idx, data in enumerate(train_loader):
inputs , labels , fore_idx = data
inputs = inputs.double()
# zero the parameter gradients
# print(inputs.shape)
inputs = torch.reshape(inputs,(batch,9*3,32,32))
# print(inputs.shape)
inputs , labels , fore_idx = inputs.to("cuda") , labels.to("cuda"), fore_idx.to("cuda")
optimizer.zero_grad()
outputs = net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
train_loss += loss.item()
_, predicted = outputs.max(1)
total += labels.size(0)
correct += predicted.eq(labels).sum().item()
print(batch_idx, len(trainloader), 'Loss: %.3f | Acc: %.3f%% (%d/%d)'% (train_loss/(batch_idx+1), 100.*correct/total, correct, total))
return train_loss/(batch_idx+1)
# Commented out IPython magic to ensure Python compatibility.
def test(epoch):
global best_acc
net.eval()
test_loss = 0
correct = 0
total = 0
with torch.no_grad():
for batch_idx, data in enumerate(test_loader):
inputs , labels , fore_idx = data
inputs = inputs.double()
# zero the parameter gradients
# print(inputs.shape)
inputs = torch.reshape(inputs,(batch,9*3,32,32))
# print(inputs.shape)
inputs , labels , fore_idx = inputs.to("cuda") , labels.to("cuda"), fore_idx.to("cuda")
outputs = net(inputs)
loss = criterion(outputs, labels)
test_loss += loss.item()
_, predicted = outputs.max(1)
total += labels.size(0)
correct += predicted.eq(labels).sum().item()
print(batch_idx, len(testloader), 'Loss: %.3f | Acc: %.3f%% (%d/%d)' % (test_loss/(batch_idx+1), 100.*correct/total, correct, total))
# Save checkpoint.
# acc = (100.0)*correct/total
# if acc > best_acc:
# print('Saving..')
# state = {
# 'net': net.state_dict(),
# 'acc': acc,
# 'epoch': epoch,
# }
# if not os.path.isdir('checkpoint'):
# os.mkdir('checkpoint')
# torch.save(state, './checkpoint/ckpt.pth')
# best_acc = acc
return test_loss/(batch_idx+1)
best_acc = 0
start_epoch =0
tr_loss = []
ts_loss = []
for epoch in range(start_epoch, start_epoch+50):
tr_loss.append(train(epoch))
ts_loss.append(test(epoch))
# model = ResNet34(3)
# #model = inception_net(3,3)
# model = model.to(device)
# if device == 'cuda':
# model = torch.nn.DataParallel(model)
# cudnn.benchmark = True
# checkpoint = torch.load('./checkpoint/ckpt.pth')
# model.load_state_dict(checkpoint['net'])
# epoch = checkpoint['epoch']
# test_acc = checkpoint['acc']
plt.plot(tr_loss,label='training_loss')
plt.plot(ts_loss,label = 'test_loss')
plt.xlabel("epochs")
plt.ylabel("cross_entropy loss")
plt.legend()
# test_acc
```
| github_jupyter |
<img align="right" src="images/ninologo.png" width="150"/>
<img align="right" src="images/tf-small.png" width="125"/>
<img align="right" src="images/dans.png" width="150"/>
# Quads
When simple signs get stacked we get composite signs.
Here we call them *quads*.
There are several ways to compose quads from sub-quads: there is always
an *operator* involved.
And a composition can again be subjected to an other composition.
And again ...
```
%load_ext autoreload
%autoreload 2
from tf.app import use
A = use("uruk:clone", checkout="clone", hoist=globals())
# A = use('uruk', hoist=globals())
```
We need our example tablet (again).
It is particularly relevant to this chapter in our tutorial:
it contains the most deeply nested quad in the whole corpus.
```
pNum = "P005381"
query = """
tablet catalogId=P005381
"""
results = A.search(query)
A.lineart(results[0][0], width=200)
A.show(results, withNodes=True)
```
The components of quads are either sub-quads or signs.
Sub-quads are also quads in TF, and they are always a composition.
Whenever a member of a sub-quad is no longer a composition, it is a *sign*.
Let's try to unravel the structure of the biggest quad in this tablet.
## Find the quad
First we need to get the node of this quad. Above we have seen the source code of the tablet in which
it occurs, from that we can pick the node of the case it is in:
```
case = A.nodeFromCase(("P005381", "obverse:2", "1"))
print(A.getSource(case))
A.pretty(case, withNodes=True)
```
We can easily read off the node number of this big quad.
But we can also do it programmatically.
In order to identify our super-quad, we list all quad nodes that are part of this case.
For every quad we list the node numbers of the signs contained in it.
In order to know what signs are contained in any given node, we use the feature `oslots`.
Like the feature `otype`, this is a standard feature that is always available in a TF dataset.
Unlike `otype`, `oslots` is an *edge* feature: there is an edge between every node and every slot contained in it.
Whereas you use `F` to do stuff with node features, you use `E` to do business with edge features.
And whereas you use `F.feature.v(node)` to get the feature value of a node, you use
`E.oslots.s(node)` to get the nodes for which there is an `oslots` edge from `node` to it.
```
for node in L.d(case, otype="quad"):
print(f"{node:>6} {E.oslots.s(node)}")
```
We see what the biggest quad is.
We could have been a bit more friendly to our selves by showing the actual graphemes in the quads.
```
for node in L.d(case, otype="quad"):
print(f'{node:>6} {" ".join(F.grapheme.v(s) for s in E.oslots.s(node))}')
```
So let us get the node of the biggest quad.
```
bigQuad = sorted(
(quad for quad in L.d(case, otype="quad")), key=lambda q: -len(E.oslots.s(q))
)[0]
bigQuad
```
Lo and behold, it is precisely the big quad.
This is what we are talking about:
```
A.lineart(bigQuad)
```
## Quad structure
Now we are going to retrieve its components by following *edges*.
When we converted the data to Text-Fabric, we have made
*edges* from quad nodes to the nodes of their component quads and signs.
We also have made edges between sibling quads and signs.
We can distinguish between kinds of edges by means of edge features.
The edges that go down in a structure have a feature `sub`.
In order to follow the `sub` edges from a node, you use
`E.sub.f(node)`.
This will give you a list of nodes that can be reached *from* `node` by following
a `sub` edge.
Edges can be traveled in the opposite direction as well:
`E.sub.t(node)`.
This will give you the nodes from which there is a `sub` edge *to* `node`.
```
E.sub.f(bigQuad)
```
or, more friendly:
```
for node in E.sub.f(bigQuad):
print(f'{node:>6} {" ".join(F.grapheme.v(s) for s in E.oslots.s(node))}')
```
Let us unravel the whole structure by means of a function:
```
def unravelQuad(quad):
if F.otype.v(quad) == "sign":
return F.grapheme.v(quad)
subQuads = E.sub.f(quad)
unraveledSubQuads = [unravelQuad(subQuad) for subQuad in subQuads]
return f'<{", ".join(unraveledSubQuads)}>'
unravelQuad(bigQuad)
```
## Operators
Where have the operators gone?
They are present as a feature `op` of edges between sibling quads and signs.
```
for child in E.sub.f(bigQuad):
for (right, op) in E.op.f(child):
print(child, op, right)
```
Note, that whereas `E.sub.f` yields a list of nodes,
`E.op.f` yields a list of pairs (node, op-value),
because the `op` edges carry a value.
The best way to know this, is to consult the
[Feature Doc](https://github.com/Nino-cunei/uruk/blob/master/docs/transcription.md).
This link as always present below the cell where you called `Cunei` for the first time.
Can we try to adapt the unravel function above to get the operators?
Yes:
```
def unravelQuad(quad):
if F.otype.v(quad) == "sign":
return F.grapheme.v(quad)
subQuads = E.sub.f(quad)
result = "<"
for sq in subQuads:
for (rq, operator) in E.op.f(sq):
leftRep = unravelQuad(sq)
rightRep = unravelQuad(rq)
result += f"{leftRep} {operator} {rightRep}"
result += ">"
return result
unravelQuad(bigQuad)
```
This technique is employed fully in the function `A.atfFromQuad()`:
```
print(A.atfFromQuad(bigQuad))
```
We have tested the function `A.atfFromQuad()` on all quads in the corpus, an it regenerates the exact ATF transliterations for them, except for two cases where the ATF has unnecessary brackets. See [checks](http://nbviewer.jupyter.org/github/Nino-cunei/uruk/blob/master/programs/checks.ipynb#Quads).
# Next
[jumps](jumps.ipynb)
*Leap to the next level ...*
All chapters:
[start](start.ipynb)
[imagery](imagery.ipynb)
[steps](steps.ipynb)
[search](search.ipynb)
[calc](calc.ipynb)
[signs](signs.ipynb)
**quads**
[jumps](jumps.ipynb)
[cases](cases.ipynb)
---
CC-BY Dirk Roorda
| github_jupyter |
```
import numpy as np
import pandas as pd
import types
import sys
from collections import defaultdict
from pprint import pformat
import math
from pyqstrat.evaluator import compute_return_metrics, display_return_metrics, plot_return_metrics
from pyqstrat.account import Account
from pyqstrat.pq_utils import *
from pyqstrat.pq_types import ContractGroup
from pyqstrat.plot import TimeSeries, trade_sets_by_reason_code, Subplot, Plot
def _get_time_series_list(timestamps, names, values, properties):
ts_list = []
for name in names:
line_type, color = None, None
if properties is not None and name in properties:
if 'line_type' in properties[name]: line_type = properties[name]['line_type']
if 'color' in properties[name]: color = properties[name]['color']
y = getattr(values, name)
if not len(y): continue
if y.dtype.type in [np.str_, np.object_, np.datetime64]: continue
ts = TimeSeries(name, timestamps, y, line_type = line_type, color = color)
ts_list.append(ts)
return ts_list
class Strategy:
def __init__(self, timestamps, contract_groups, price_function, starting_equity = 1.0e6, pnl_calc_time = 15 * 60 + 1, trade_lag = 1,
run_final_calc = True, strategy_context = None):
'''
Args:
timestamps (np.array of np.datetime64): The "heartbeat" of the strategy. We will evaluate trading rules and
simulate the market at these times.
price_function: A function that returns the price of a contract at a given timestamp
contract_groups (list of :obj:`ContractGroup`): The contract groups we will potentially trade.
starting_equity (float, optional): Starting equity in Strategy currency. Default 1.e6
pnl_calc_time (int, optional): Time of day used to calculate PNL. Default 15 * 60 (3 pm)
trade_lag (int, optional): Number of bars you want between the order and the trade. For example, if you think it will take
5 seconds to place your order in the market, and your bar size is 1 second, set this to 5. Set this to 0 if you
want to execute your trade at the same time as you place the order, for example, if you have daily bars. Default 1.
run_final_calc (bool, optional): If set, calculates unrealized pnl and net pnl as well as realized pnl when strategy is done.
If you don't need unrealized pnl, turn this off for faster run time. Default True
strategy_context (:obj:`types.SimpleNamespace`, optional): A storage class where you can store key / value pairs
relevant to this strategy. For example, you may have a pre-computed table of correlations that you use in the
indicator or trade rule functions.
If not set, the __init__ function will create an empty member strategy_context object that you can access.
'''
self.name = None
self.timestamps = timestamps
assert(len(contract_groups) and isinstance(contract_groups[0], ContractGroup))
self.contract_groups = contract_groups
if strategy_context is None: strategy_context = types.SimpleNamespace()
self.strategy_context = strategy_context
self.account = Account(contract_groups, timestamps, price_function, strategy_context, starting_equity, pnl_calc_time)
assert trade_lag >= 0, f'trade_lag cannot be negative: {trade_lag}'
self.trade_lag = trade_lag
self.run_final_calc = run_final_calc
self.indicators = {}
self.signals = {}
self.signal_values = defaultdict(types.SimpleNamespace)
self.rule_names = []
self.rules = {}
self.position_filters = {}
self.rule_signals = {}
self.market_sims = []
self._trades = []
self._orders = []
self._open_orders = defaultdict(list)
self.indicator_deps = {}
self.indicator_cgroups = {}
self.indicator_values = defaultdict(types.SimpleNamespace)
self.signal_indicator_deps = {}
self.signal_deps = {}
self.signal_cgroups = {}
self.trades_iter = [[] for x in range(len(timestamps))] # For debugging, we don't really need this as a member variable
def add_indicator(self, name, indicator, contract_groups = None, depends_on = None):
'''
Args:
name: Name of the indicator
indicator: A function that takes strategy timestamps and other indicators and returns a numpy array
containing indicator values. The return array must have the same length as the timestamps object.
Can also be a numpy array or a pandas Series in which case we just store the values.
contract_groups (list of :obj:`ContractGroup`, optional): Contract groups that this indicator applies to.
If not set, it applies to all contract groups. Default None.
depends_on (list of str, optional): Names of other indicators that we need to compute this indicator.
Default None.
'''
self.indicators[name] = indicator
self.indicator_deps[name] = [] if depends_on is None else depends_on
if contract_groups is None: contract_groups = self.contract_groups
if isinstance(indicator, np.ndarray) or isinstance(indicator, pd.Series):
indicator_values = series_to_array(indicator)
for contract_group in contract_groups:
setattr(self.indicator_values[contract_group], name, indicator_values)
self.indicator_cgroups[name] = contract_groups
def add_signal(self, name, signal_function, contract_groups = None, depends_on_indicators = None, depends_on_signals = None):
'''
Args:
name (str): Name of the signal
signal_function (function): A function that takes timestamps and a dictionary of indicator value arrays and
returns a numpy array
containing signal values. The return array must have the same length as the input timestamps
contract_groups (list of :obj:`ContractGroup`, optional): Contract groups that this signal applies to.
If not set, it applies to all contract groups. Default None.
depends_on_indicators (list of str, optional): Names of indicators that we need to compute this signal. Default None.
depends_on_signals (list of str, optional): Names of other signals that we need to compute this signal. Default None.
'''
self.signals[name] = signal_function
self.signal_indicator_deps[name] = [] if depends_on_indicators is None else depends_on_indicators
self.signal_deps[name] = [] if depends_on_signals is None else depends_on_signals
if contract_groups is None: contract_groups = self.contract_groups
self.signal_cgroups[name] = contract_groups
def add_rule(self, name, rule_function, signal_name, sig_true_values = None, position_filter = None):
'''Add a trading rule. Trading rules are guaranteed to run in the order in which you add them. For example, if you set trade_lag to 0,
and want to exit positions and re-enter new ones in the same bar, make sure you add the exit rule before you add the entry rule to the
strategy.
Args:
name (str): Name of the trading rule
rule_function (function): A trading rule function that returns a list of Orders
signal_name (str): The strategy will call the trading rule function when the signal with this name matches sig_true_values
sig_true_values (numpy array, optional): If the signal value at a bar is equal to one of these values,
the Strategy will call the trading rule function. Default [TRUE]
position_filter (str, optional): Can be "zero", "nonzero" or None. Zero rules are only triggered when
the corresponding contract positions are 0
Nonzero rules are only triggered when the corresponding contract positions are non-zero.
If not set, we don't look at position before triggering the rule.
Default None
'''
if sig_true_values is None: sig_true_values = [True]
if name in self.rule_names:
raise Exception(f'Rule {name} already exists')
# Rules should be run in order
self.rule_names.append(name)
self.rule_signals[name] = (signal_name, sig_true_values)
self.rules[name] = rule_function
if position_filter is not None:
assert(position_filter in ['zero', 'nonzero'])
self.position_filters[name] = position_filter
def add_market_sim(self, market_sim_function):
'''Add a market simulator. A market simulator takes a list of Orders as input and returns a list of Trade objects.
Args:
market_sim_function (function): A function that takes a list of Orders and Indicators as input
and returns a list of Trade objects
'''
self.market_sims.append(market_sim_function)
def run_indicators(self, indicator_names = None, contract_groups = None, clear_all = False):
'''Calculate values of the indicators specified and store them.
Args:
indicator_names (list of str, optional): List of indicator names. If None (default) run all indicators
contract_groups (list of :obj:`ContractGroup`, optional): Contract group to run this indicator for.
If None (default), we run it for all contract groups.
clear_all (bool, optional): If set, clears all indicator values before running. Default False.
'''
if indicator_names is None: indicator_names = self.indicators.keys()
if contract_groups is None: contract_groups = self.contract_groups
if clear_all: self.indicator_values = defaultdict(types.SimpleNamespace)
ind_names = []
for ind_name, cgroup_list in self.indicator_cgroups.items():
if len(set(contract_groups).intersection(cgroup_list)): ind_names.append(ind_name)
indicator_names = list(set(ind_names).intersection(indicator_names))
for cgroup in contract_groups:
cgroup_ind_namespace = self.indicator_values[cgroup]
for indicator_name in indicator_names:
# First run all parents
parent_names = self.indicator_deps[indicator_name]
for parent_name in parent_names:
if cgroup in self.indicator_values and hasattr(cgroup_ind_namespace, parent_name): continue
self.run_indicators([parent_name], [cgroup])
# Now run the actual indicator
if cgroup in self.indicator_values and hasattr(cgroup_ind_namespace, indicator_name): continue
indicator_function = self.indicators[indicator_name]
parent_values = types.SimpleNamespace()
for parent_name in parent_names:
setattr(parent_values, parent_name, getattr(cgroup_ind_namespace, parent_name))
if isinstance(indicator_function, np.ndarray) or isinstance(indicator_function, pd.Series):
indicator_values = indicator_function
else:
indicator_values = indicator_function(cgroup, self.timestamps, parent_values, self.strategy_context)
setattr(cgroup_ind_namespace, indicator_name, series_to_array(indicator_values))
def run_signals(self, signal_names = None, contract_groups = None, clear_all = False):
'''Calculate values of the signals specified and store them.
Args:
signal_names (list of str, optional): List of signal names. If None (default) run all signals
contract_groups (list of :obj:`ContractGroup`, optional): Contract groups to run this signal for.
If None (default), we run it for all contract groups.
clear_all (bool, optional): If set, clears all signal values before running. Default False.
'''
if signal_names is None: signal_names = self.signals.keys()
if contract_groups is None: contract_groups = self.contract_groups
if clear_all: self.signal_values = defaultdict(types.SimpleNamespace)
sig_names = []
for sig_name, cgroup_list in self.signal_cgroups.items():
if len(set(contract_groups).intersection(cgroup_list)):
sig_names.append(sig_name)
signal_names = list(set(sig_names).intersection(signal_names))
for cgroup in contract_groups:
for signal_name in signal_names:
if cgroup not in self.signal_cgroups[signal_name]: continue
# First run all parent signals
parent_names = self.signal_deps[signal_name]
for parent_name in parent_names:
if cgroup in self.signal_values and hasattr(self.signal_values[cgroup], parent_name): continue
self.run_signals([parent_name], [cgroup])
# Now run the actual signal
if cgroup in self.signal_values and hasattr(self.signal_values[cgroup], signal_name): continue
signal_function = self.signals[signal_name]
parent_values = types.SimpleNamespace()
for parent_name in parent_names:
sig_vals = getattr(self.signal_values[cgroup], parent_name)
setattr(parent_values, parent_name, sig_vals)
# Get indicators needed for this signal
indicator_values = types.SimpleNamespace()
for indicator_name in self.signal_indicator_deps[signal_name]:
setattr(indicator_values, indicator_name, getattr(self.indicator_values[cgroup], indicator_name))
setattr(self.signal_values[cgroup], signal_name, series_to_array(
signal_function(cgroup, self.timestamps, indicator_values, parent_values, self.strategy_context)))
def _generate_order_iterations(self, rule_names = None, contract_groups = None, start_date = None, end_date = None):
'''
>>> class MockStrat:
... def __init__(self):
... self.timestamps = timestamps
... self.account = self
... self.rules = {'rule_a' : rule_a, 'rule_b' : rule_b}
... self.market_sims = {ibm : market_sim_ibm, aapl : market_sim_aapl}
... self.rule_signals = {'rule_a' : ('sig_a', [1]), 'rule_b' : ('sig_b', [1, -1])}
... self.signal_values = {ibm : types.SimpleNamespace(sig_a = np.array([0., 1., 1.]),
... sig_b = np.array([0., 0., 0.]) ),
... aapl : types.SimpleNamespace(sig_a = np.array([0., 0., 0.]),
... sig_b = np.array([0., -1., -1])
... )}
... self.signal_cgroups = {'sig_a' : [ibm, aapl], 'sig_b' : [ibm, aapl]}
... self.indicator_values = {ibm : types.SimpleNamespace(), aapl : types.SimpleNamespace()}
>>>
>>> def market_sim_aapl(): pass
>>> def market_sim_ibm(): pass
>>> def rule_a(): pass
>>> def rule_b(): pass
>>> timestamps = np.array(['2018-01-01', '2018-01-02', '2018-01-03'], dtype = 'M8[D]')
>>> rule_names = ['rule_a', 'rule_b']
>>> ContractGroup.clear()
>>> ibm = ContractGroup.create('IBM')
>>> aapl = ContractGroup.create('AAPL')
>>> contract_groups = [ibm, aapl]
>>> start_date = np.datetime64('2018-01-01')
>>> end_date = np.datetime64('2018-02-05')
>>> strategy = MockStrat()
>>> Strategy._generate_order_iterations(strategy, rule_names, contract_groups, start_date, end_date)
>>> orders_iter = strategy.orders_iter
>>> assert(len(orders_iter[0]) == 0)
>>> assert(len(orders_iter[1]) == 2)
>>> assert(orders_iter[1][0][1] == ibm)
>>> assert(orders_iter[1][1][1] == aapl)
>>> assert(len(orders_iter[2]) == 0)
'''
start_date, end_date = str2date(start_date), str2date(end_date)
if rule_names is None: rule_names = self.rule_names
if contract_groups is None: contract_groups = self.contract_groups
num_timestamps = len(self.timestamps)
# List of lists, i -> list of orders
orders_iter = [[] for x in range(num_timestamps)]
for rule_name in rule_names:
rule_function = self.rules[rule_name]
for cgroup in contract_groups:
signal_name, sig_true_values = self.rule_signals[rule_name]
if cgroup not in self.signal_cgroups[signal_name]:
# We don't need to call this rule for this contract group
continue
sig_values = getattr(self.signal_values[cgroup], signal_name)
timestamps = self.timestamps
null_value = False if sig_values.dtype == np.dtype('bool') else np.nan
if start_date: sig_values[0:np.searchsorted(timestamps, start_date)] = null_value
if end_date: sig_values[np.searchsorted(timestamps, end_date):] = null_value
indices = np.nonzero(np.isin(sig_values[:num_timestamps], sig_true_values))[0]
# Don't run rules on last index since we cannot fill any orders
if len(indices) and indices[-1] == len(sig_values) -1: indices = indices[:-1]
indicator_values = self.indicator_values[cgroup]
iteration_params = {'indicator_values' : indicator_values, 'signal_values' : sig_values, 'rule_name' : rule_name}
for idx in indices: orders_iter[idx].append((rule_function, cgroup, iteration_params))
self.orders_iter = orders_iter
def run_rules(self, rule_names = None, contract_groups = None, start_date = None, end_date = None):
'''Run trading rules.
Args:
rule_names: List of rule names. If None (default) run all rules
contract_groups (list of :obj:`ContractGroup`, optional): Contract groups to run this rule for.
If None (default), we run it for all contract groups.
start_date: Run rules starting from this date. Default None
end_date: Don't run rules after this date. Default None
'''
start_date, end_date = str2date(start_date), str2date(end_date)
self._generate_order_iterations(rule_names, contract_groups, start_date, end_date)
# Now we know which rules, contract groups need to be applied for each iteration, go through each iteration and apply them
# in the same order they were added to the strategy
for i in range(len(self.orders_iter)):
self._run_iteration(i)
if self.run_final_calc:
self.account.calc(self.timestamps[-1])
def _run_iteration(self, i):
self._sim_market(i)
# Treat all orders as IOC, i.e. if the order was not executed, then its cancelled.
self._open_orders[i] = []
rules = self.orders_iter[i]
for (rule_function, contract_group, params) in rules:
orders = self._get_orders(i, rule_function, contract_group, params)
self._orders += orders
self._open_orders[i + self.trade_lag] += orders
# If the lag is 0, then run rules one by one, and after each rule, run market sim to generate trades and update
# positions. For example, if we have a rule to exit a position and enter a new one, we should make sure
# positions are updated after the first rule before running the second rule. If the lag is not 0,
# run all rules and collect the orders, we don't need to run market sim after each rule
if self.trade_lag == 0: self._sim_market(i)
# If we failed to fully execute any orders in this iteration, add them to the next iteration so we get another chance to execute
open_orders = self._open_orders.get(i)
if open_orders is not None and len(open_orders):
self._open_orders[i + 1] += open_orders
def run(self):
self.run_indicators()
self.run_signals()
self.run_rules()
def _get_orders(self, idx, rule_function, contract_group, params):
try:
indicator_values, signal_values, rule_name = (params['indicator_values'], params['signal_values'], params['rule_name'])
position_filter = self.position_filters[rule_name]
if position_filter is not None:
curr_pos = self.account.position(contract_group, self.timestamps[idx])
if position_filter == 'zero' and not math.isclose(curr_pos, 0): return []
if position_filter == 'nonzero' and math.isclose(curr_pos, 0): return []
orders = rule_function(contract_group, idx, self.timestamps, indicator_values, signal_values, self.account,
self.strategy_context)
except Exception as e:
raise type(e)(f'Exception: {str(e)} at rule: {type(rule_function)} contract_group: {contract_group} index: {idx}'
).with_traceback(sys.exc_info()[2])
return orders
def _sim_market(self, i):
'''
Go through all open orders and run market simulators to generate a list of trades and return any orders that were not filled.
'''
open_orders = self._open_orders.get(i)
if open_orders is None or len(open_orders) == 0: return [], []
# If there is more than one order for a contract, throw away any but the last one.
#seen = set()
#seen_add = seen.add
#open_orders = list(reversed([order for order in reversed(orders) if not (order.contract in seen or seen_add(order.contract))]))
for market_sim_function in self.market_sims:
try:
trades = market_sim_function(open_orders, i, self.timestamps, self.indicator_values, self.signal_values, self.strategy_context)
if len(trades): self.account.add_trades(trades)
self._trades += trades
except Exception as e:
raise type(e)(f'Exception: {str(e)} at index: {i} function: {market_sim_function}').with_traceback(sys.exc_info()[2])
self._open_orders[i] = [order for order in open_orders if order.status != 'filled']
def df_data(self, contract_groups = None, add_pnl = True, start_date = None, end_date = None):
'''
Add indicators and signals to end of market data and return as a pandas dataframe.
Args:
contract_groups (list of :obj:`ContractGroup`, optional): list of contract groups to include. All if set to None (default)
add_pnl: If True (default), include P&L columns in dataframe
start_date: string or numpy datetime64. Default None
end_date: string or numpy datetime64: Default None
'''
start_date, end_date = str2date(start_date), str2date(end_date)
if contract_groups is None: contract_groups = self.contract_groups
timestamps = self.timestamps
if start_date: timestamps = timestamps[timestamps >= start_date]
if end_date: timestamps = timestamps[timestamps <= end_date]
dfs = []
for contract_group in contract_groups:
df = pd.DataFrame({'timestamp' : self.timestamps})
if add_pnl:
df_pnl = self.df_pnl(contract_group)
indicator_values = self.indicator_values[contract_group]
for k in sorted(indicator_values.__dict__):
name = k
# Avoid name collisions
if name in df.columns: name = name + '.ind'
df.insert(len(df.columns), name, getattr(indicator_values, k))
signal_values = self.signal_values[contract_group]
for k in sorted(signal_values.__dict__):
name = k
if name in df.columns: name = name + '.sig'
df.insert(len(df.columns), name, getattr(signal_values, k))
if add_pnl: df = pd.merge(df, df_pnl, on = ['timestamp'], how = 'left')
# Add counter column for debugging
df.insert(len(df.columns), 'i', np.arange(len(df)))
dfs.append(df)
return pd.concat(dfs)
def trades(self, contract_group = None, start_date = None, end_date = None):
'''Returns a list of trades with the given contract group and with trade date between (and including) start date
and end date if they are specified.
If contract_group is None trades for all contract_groups are returned'''
start_date, end_date = str2date(start_date), str2date(end_date)
return self.account.trades(contract_group, start_date, end_date)
def df_trades(self, contract_group = None, start_date = None, end_date = None):
'''Returns a dataframe with data from trades with the given contract group and with trade date between (and including)
start date and end date
if they are specified. If contract_group is None trades for all contract_groups are returned'''
start_date, end_date = str2date(start_date), str2date(end_date)
return self.account.df_trades(contract_group, start_date, end_date)
def orders(self, contract_group = None, start_date = None, end_date = None):
'''Returns a list of orders with the given contract group and with order date between (and including) start date and
end date if they are specified.
If contract_group is None orders for all contract_groups are returned'''
orders = []
start_date, end_date = str2date(start_date), str2date(end_date)
if contract_group is None:
orders += [order for order in self._orders if (
start_date is None or order.date >= start_date) and (end_date is None or order.date <= end_date)]
else:
for contract in contract_group.contracts:
orders += [order for order in self._orders if (contract is None or order.contract == contract) and (
start_date is None or order.date >= start_date) and (end_date is None or order.date <= end_date)]
return orders
def df_orders(self, contract_group = None, start_date = None, end_date = None):
'''Returns a dataframe with data from orders with the given contract group and with order date between (and including)
start date and end date
if they are specified. If contract_group is None orders for all contract_groups are returned'''
start_date, end_date = str2date(start_date), str2date(end_date)
orders = self.orders(contract_group, start_date, end_date)
order_records = [(order.contract.symbol, type(order).__name__, order.timestamp, order.qty,
order.reason_code,
(str(order.properties.__dict__) if order.properties.__dict__ else ''),
(str(order.contract.properties.__dict__) if order.contract.properties.__dict__ else '')) for order in orders]
df_orders = pd.DataFrame.from_records(order_records,
columns = ['symbol', 'type', 'timestamp', 'qty', 'reason_code', 'order_props', 'contract_props'])
return df_orders
def df_pnl(self, contract_group = None):
'''Returns a dataframe with P&L columns. If contract group is set to None (default), sums up P&L across all contract groups'''
return self.account.df_account_pnl(contract_group)
def df_returns(self, contract_group = None, sampling_frequency = 'D'):
'''Return a dataframe of returns and equity indexed by date.
Args:
contract_group (:obj:`ContractGroup`, optional) : The contract group to get returns for.
If set to None (default), we return the sum of PNL for all contract groups
sampling_frequency: Downsampling frequency. Default is None. See pandas frequency strings for possible values
'''
pnl = self.df_pnl(contract_group)[['timestamp', 'net_pnl', 'equity']]
pnl.equity = pnl.equity.ffill()
pnl = pnl.set_index('timestamp').resample(sampling_frequency).last().reset_index()
pnl = pnl.dropna(subset = ['equity'])
pnl['ret'] = pnl.equity.pct_change()
return pnl
def plot(self,
contract_groups = None,
primary_indicators = None,
primary_indicators_dual_axis = None,
secondary_indicators = None,
secondary_indicators_dual_axis = None,
indicator_properties = None,
signals = None,
signal_properties = None,
pnl_columns = None,
title = None,
figsize = (20, 15),
date_range = None,
date_format = None,
sampling_frequency = None,
trade_marker_properties = None,
hspace = 0.15):
'''Plot indicators, signals, trades, position, pnl
Args:
contract_groups (list of :obj:`ContractGroup`, optional): Contract groups to plot or None (default) for all
contract groups.
primary indicators (list of str, optional): List of indicators to plot in the main indicator section.
Default None (plot everything)
primary indicators (list of str, optional): List of indicators to plot in the secondary indicator section.
Default None (don't plot anything)
indicator_properties (dict of str : dict, optional): If set, we use the line color, line type indicated
for the given indicators
signals (list of str, optional): Signals to plot. Default None (plot everything).
plot_equity (bool, optional): If set, we plot the equity curve. Default is True
title (list of str, optional): Title of plot. Default None
figsize (tuple of int): Figure size. Default (20, 15)
date_range (tuple of str or np.datetime64, optional): Used to restrict the date range of the graph.
Default None
date_format (str, optional): Date format for tick labels on x axis. If set to None (default),
will be selected based on date range. See matplotlib date format strings
sampling_frequency (str, optional): Downsampling frequency. The graph may get too busy if you have too many bars
of data, in which case you may want to downsample before plotting. See pandas frequency strings for
possible values. Default None.
trade_marker_properties (dict of str : tuple, optional): A dictionary of
order reason code -> marker shape, marker size, marker color for plotting trades with different reason codes.
Default is None in which case the dictionary from the ReasonCode class is used
hspace (float, optional): Height (vertical) space between subplots. Default is 0.15
'''
date_range = strtup2date(date_range)
if contract_groups is None: contract_groups = self.contract_groups
if isinstance(contract_groups, ContractGroup): contract_groups = [contract_groups]
if pnl_columns is None: pnl_columns = ['equity']
for contract_group in contract_groups:
primary_indicator_names = [ind_name for ind_name in self.indicator_values[contract_group].__dict__ \
if hasattr(self.indicator_values[contract_group], ind_name)]
if primary_indicators:
primary_indicator_names = list(set(primary_indicator_names).intersection(primary_indicators))
secondary_indicator_names = []
if secondary_indicators:
secondary_indicator_names = secondary_indicators
signal_names = [sig_name for sig_name in self.signals.keys() if hasattr(self.signal_values[contract_group], sig_name)]
if signals:
signal_names = list(set(signal_names).intersection(signals))
primary_indicator_list = _get_time_series_list(self.timestamps, primary_indicator_names,
self.indicator_values[contract_group], indicator_properties)
secondary_indicator_list = _get_time_series_list(self.timestamps, secondary_indicator_names,
self.indicator_values[contract_group], indicator_properties)
signal_list = _get_time_series_list(self.timestamps, signal_names, self.signal_values[contract_group], signal_properties)
df_pnl_ = self.df_pnl(contract_group)
pnl_list = [TimeSeries(pnl_column, timestamps = df_pnl_.timestamp.values, values = df_pnl_[pnl_column].values
) for pnl_column in pnl_columns]
trades = [trade for trade in self._trades if trade.order.contract.contract_group == contract_group]
if trade_marker_properties:
trade_sets = trade_sets_by_reason_code(trades, trade_marker_properties, remove_missing_properties = True)
else:
trade_sets = trade_sets_by_reason_code(trades)
primary_indicator_subplot = Subplot(primary_indicator_list + trade_sets,
secondary_y = primary_indicators_dual_axis,
height_ratio = 0.5, ylabel = 'Primary Indicators')
if len(secondary_indicator_list):
secondary_indicator_subplot = Subplot(secondary_indicator_list,
secondary_y = secondary_indicators_dual_axis,
height_ratio = 0.5, ylabel = 'Secondary Indicators')
signal_subplot = Subplot(signal_list, ylabel = 'Signals', height_ratio = 0.167)
pnl_subplot = Subplot(pnl_list, ylabel = 'Equity', height_ratio = 0.167, log_y = True, y_tick_format = '${x:,.0f}')
position = df_pnl_.position.values
pos_subplot = Subplot([TimeSeries('position', timestamps = df_pnl_.timestamp, values = position,
plot_type = 'filled_line')], ylabel = 'Position', height_ratio = 0.167)
title_full = title
if len(contract_groups) > 1:
if title is None: title = ''
title_full = f'{title} {contract_group.name}'
plot_list = []
if len(primary_indicator_list): plot_list.append(primary_indicator_subplot)
if len(secondary_indicator_list): plot_list.append(secondary_indicator_subplot)
if len(signal_list) : plot_list.append(signal_subplot)
if len(position): plot_list.append(pos_subplot)
if len(pnl_list): plot_list.append(pnl_subplot)
if not len(plot_list): return
plot = Plot(plot_list, figsize = figsize, date_range = date_range, date_format = date_format,
sampling_frequency = sampling_frequency,
title = title_full, hspace = hspace)
plot.draw()
def evaluate_returns(self, contract_group = None, plot = True, display_summary = True, float_precision = 4, return_metrics = False):
'''Returns a dictionary of common return metrics.
Args:
contract_group (:obj:`ContractGroup`, optional): Contract group to evaluate or None (default) for all contract groups
plot (bool): If set to True, display plots of equity, drawdowns and returns. Default False
float_precision (float, optional): Number of significant figures to show in returns. Default 4
return_metrics (bool, optional): If set, we return the computed metrics as a dictionary
'''
returns = self.df_returns(contract_group)
ev = compute_return_metrics(returns.timestamp.values, returns.ret.values, self.account.starting_equity)
if display_summary:
display_return_metrics(ev.metrics(), float_precision = float_precision)
if plot: plot_return_metrics(ev.metrics())
if return_metrics:
return ev.metrics()
def plot_returns(self, contract_group = None):
'''Display plots of equity, drawdowns and returns for the given contract group or for all contract groups if contract_group
is None (default)'''
if contract_group is None:
returns = self.df_returns()
else:
returns = self.df_returns(contract_group)
ev = compute_return_metrics(returns.timestamp.values, returns.ret.values, self.account.starting_equity)
fig, ax = plot_return_metrics(ev.metrics())
return fig, ax
def __repr__(self):
return f'{pformat(self.indicators)} {pformat(self.rules)} {pformat(self.account)}'
def test_strategy():
#if __name__ == "__main__":
import math
import datetime
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import scipy.stats
import os
from types import SimpleNamespace
from pyqstrat.pq_types import Contract, ContractGroup, Trade
from pyqstrat.portfolio import Portfolio
from pyqstrat.orders import MarketOrder
try:
# If we are running from unit tests
ko_file_path = os.path.dirname(os.path.realpath(__file__)) + '/notebooks/support/coke_15_min_prices.csv.gz'
pep_file_path = os.path.dirname(os.path.realpath(__file__)) + '/notebooks/support/pepsi_15_min_prices.csv.gz'
except:
ko_file_path = '../notebooks/support/coke_15_min_prices.csv.gz'
pep_file_path = '../notebooks/support/pepsi_15_min_prices.csv.gz'
ko_prices = pd.read_csv(ko_file_path)
pep_prices = pd.read_csv(pep_file_path)
ko_prices['timestamp'] = pd.to_datetime(ko_prices.date)
pep_prices['timestamp'] = pd.to_datetime(pep_prices.date)
end_time = '2019-01-30 12:00'
ko_prices = ko_prices.query(f'timestamp <= "{end_time}"')
pep_prices = pep_prices.query(f'timestamp <= "{end_time}"')
timestamps = ko_prices.timestamp.values
ratio = ko_prices.c / pep_prices.c
def zscore_indicator(contract_group, timestamps, indicators, strategy_context): # simple moving average
ratio = indicators.ratio
r = pd.Series(ratio).rolling(window = 130)
mean = r.mean()
std = r.std(ddof = 0)
zscore = (ratio - mean) / std
zscore = np.nan_to_num(zscore)
return zscore
def pair_strategy_signal(contract_group, timestamps, indicators, parent_signals, strategy_context):
# We don't need any indicators since the zscore is already part of the market data
zscore = indicators.zscore
signal = np.where(zscore > 1, 2, 0)
signal = np.where(zscore < -1, -2, signal)
signal = np.where((zscore > 0.5) & (zscore < 1), 1, signal)
signal = np.where((zscore < -0.5) & (zscore > -1), -1, signal)
if contract_group.name == 'PEP': signal = -1. * signal
return signal
def pair_entry_rule(contract_group, i, timestamps, indicators, signal, account, strategy_context):
timestamp = timestamps[i]
assert(math.isclose(account.position(contract_group, timestamp), 0))
signal_value = signal[i]
risk_percent = 0.1
orders = []
symbol = contract_group.name
contract = contract_group.get_contract(symbol)
if contract is None: contract = Contract.create(symbol, contract_group = contract_group)
# if we don't already have a position, check if we should enter a trade
#if math.isclose(curr_pos, 0):
curr_equity = account.equity(timestamp)
order_qty = np.round(curr_equity * risk_percent / indicators.c[i] * np.sign(signal_value))
trigger_price = indicators.c[i]
print(f'order_qty: {order_qty} curr_equity: {curr_equity} timestamp: {timestamp}' + \
f' risk_percent: {risk_percent} indicator: {indicators.c[i]} signal_value: {signal_value}')
reason_code = ReasonCode.ENTER_LONG if order_qty > 0 else ReasonCode.ENTER_SHORT
orders.append(MarketOrder(contract, timestamp, order_qty, reason_code = reason_code))
return orders
def pair_exit_rule(contract_group, i, timestamps, indicators, signal, account, strategy_context):
timestamp = timestamps[i]
curr_pos = account.position(contract_group, timestamp)
assert(not math.isclose(curr_pos, 0))
signal_value = signal[i]
orders = []
symbol = contract_group.name
contract = contract_group.get_contract(symbol)
if contract is None: contract = Contract.create(symbol, contract_group = contract_group)
if (curr_pos > 0 and signal_value == -1) or (curr_pos < 0 and signal_value == 1):
order_qty = -curr_pos
reason_code = ReasonCode.EXIT_LONG if order_qty < 0 else ReasonCode.EXIT_SHORT
orders.append(MarketOrder(contract, timestamp, order_qty, reason_code = reason_code))
return orders
def market_simulator(orders, i, timestamps, indicators, signals, strategy_context):
trades = []
timestamp = timestamps[i]
for order in orders:
trade_price = np.nan
cgroup = order.contract.contract_group
ind = indicators[cgroup]
o, h, l, c = ind.o[i], ind.h[i], ind.l[i], ind.c[i]
assert isinstance(order, MarketOrder), f'Unexpected order type: {order}'
trade_price = 0.5 * (o + h) if order.qty > 0 else 0.5 * (o + l)
if np.isnan(trade_price): continue
trade = Trade(order.contract, timestamp, order.qty, trade_price, order = order, commission = 0, fee = 0)
order.status = 'filled'
print(f'trade: {trade}')
trades.append(trade)
return trades
def get_price(contract, timestamps, i, strategy_context):
if contract.symbol == 'KO':
return strategy_context.ko_price[i]
elif contract.symbol == 'PEP':
return strategy_context.pep_price[i]
raise Exception(f'Unknown contract: {contract}')
Contract.clear()
ContractGroup.clear()
ko_contract_group = ContractGroup.create('KO')
pep_contract_group = ContractGroup.create('PEP')
strategy_context = SimpleNamespace(ko_price = ko_prices.c.values, pep_price = pep_prices.c.values)
strategy = Strategy(timestamps, [ko_contract_group, pep_contract_group], get_price, strategy_context = strategy_context)
for tup in [(ko_contract_group, ko_prices), (pep_contract_group, pep_prices)]:
for column in ['o', 'h', 'l', 'c']:
strategy.add_indicator(column, tup[1][column], contract_groups = [tup[0]])
strategy.add_indicator('ratio', ratio)
strategy.add_indicator('zscore', zscore_indicator, depends_on = ['ratio'])
strategy.add_signal('pair_strategy_signal', pair_strategy_signal, depends_on_indicators = ['zscore'])
# ask pqstrat to call our trading rule when the signal has one of the values [-2, -1, 1, 2]
strategy.add_rule('pair_entry_rule', pair_entry_rule,
signal_name = 'pair_strategy_signal', sig_true_values = [-2, 2], position_filter = 'zero')
strategy.add_rule('pair_exit_rule', pair_exit_rule,
signal_name = 'pair_strategy_signal', sig_true_values = [-1, 1], position_filter = 'nonzero')
strategy.add_market_sim(market_simulator)
strategy.run_indicators()
strategy.run_signals()
strategy.run_rules()
metrics = strategy.evaluate_returns(plot = False, display_summary = False)
assert(round(metrics['gmean'], 6) == -0.062874) #-0.062878)
assert(round(metrics['sharpe'], 4) == -7.2709)
assert(round(metrics['mdd_pct'], 6) == -0.002841)
if __name__ == "__main__":
strategy = test_strategy()
import doctest
doctest.testmod(optionflags = doctest.NORMALIZE_WHITESPACE)
```
| github_jupyter |
```
#########
# Demo pour la composition du corpus
########
# you can export your mongodb collection data for ES indexing step
import pymongo
myclient = pymongo.MongoClient("mongodb://localhost:27017/")
mydb = myclient["inventaire_medo"]
mycol = mydb["agriculture_demo"]
mydoc = mycol.find() # to retriev all data
my_corpus = list(mydoc)
my_corpus = pd.DataFrame(my_corpus,copy =True)
# pd.DataFrame.from_dict(my_corpus)
# pd.DataFrame.from_records(my_corpus)
my_corpus
columns = ['name','text','title', 'SNE', 'TNE', 'pertinence']
new_corpus = pd.DataFrame(my_corpus, columns=columns)
new_corpus['SNE'][0]['ent0']
new_corpus['SNE'].apply(pd.Series)
corpus_data = pd.concat([new_corpus.drop(['SNE', 'TNE'], axis=1), new_corpus['SNE'].apply(pd.Series), new_corpus['TNE'].apply(pd.Series)], axis=1)
corpus_data.head()
# LOad Dataviz pkgs
import seaborn as sns
corpus_data['date'].value_counts()
# Ploting
sns.countplot(x = 'ent0', data = corpus_data, orient= 'h' )
fig, ax = plt.subplots(2,1)
ax = ax.flatten()
tab = pd.crosstab(corpus_data['ent0'])
for i,cat in enumerate(tab.index):
tab.loc[cat].plot.pie(ax=ax[i],startangle=90)
ax[i].set_ylabel('')
ax[i].set_title(cat, fontweight='bold')
corpus_data['ent0'].isna().sum()
len(corpus_data)
len(corpus_data)
corpus_data['ent0'].value_counts()[:2]
corpus_data[corpus_data['ent0'].notnull()].head(2)
# nms.name.notnull()
#### Spatiality
import pandas as pd
import plotly.graph_objects as go
df = pd.DataFrame({'node_names': ['Corpus', 'With_SNE', 'WithOut_SNE'],
'node_parent': ["", "Corpus", "Corpus"],
'node_labels': ['Agri Corpus<br>Data Spatiality<br>','With_SNE', 'WithOut_SNE'],
#'node_counts': [len(corpus), len(corpus_with_extend), len(corpus_without_extend)]
'node_counts': [len(corpus_data), corpus_data['ent0'].isna().sum(), len(corpus_data)- corpus_data['ent0'].isna().sum()]
}
)
colors = []
for p in df["node_labels"]:
if p in ["", 'Agri Corpus<br>Data Spatiality<br>']:
colors.append("white")
elif p in ['With_SNE']:
colors.append("green")
elif p in ["WithOut_SNE"]:
colors.append("blue")
fig=go.Figure(
data=go.Sunburst(
ids=df["node_names"],
labels=df["node_labels"],
parents=df["node_parent"],
marker=dict(colors=colors_),
values=df["node_counts"],
branchvalues="total",
texttemplate = ('%{label}',
'%{label}<br>%{percentParent:.1%}',
'%{label}<br>%{percentParent:.1%}',
'%{label}<br>%{percentParent:.1%}',
'%{label}<br>%{percentParent:.1%}',
'%{label}<br>%{percentParent:.1%}',
'%{label}<br>%{percentParent:.1%}',
'%{label}<br>%{percentParent:.1%}',
'%{label}<br>%{percentParent:.1%}'),
),
)
#fig.update_layout(margin = dict(t=0, l=0, r=0, b=0))
#fig.update_layout(template='none') #dict(font=dict(color='Blue'))
fig.show()
corpus_data['date'].isna().sum()
# df2 = pd.to_datetime(df['col1'])
corpus_data['date'] = pd.to_datetime(corpus_data['date'])
# corpus_data['date']
corpus_data_tne = corpus_data[corpus_data['date'].notnull()]
# nms.name.notnull()
corpus_data_tne.head(2)
range_1 = corpus_data_tne[corpus_data_tne['date']>='2019-01-01']
# range_1
mask = (corpus_data_tne['date'] > '2015-01-01') & (corpus_data_tne['date'] <= '2019-01-01')
range_2 = corpus_data_tne[mask]
# range_2
range_3 = corpus_data_tne[corpus_data_tne['date']<='2015-01-01']
# range_3
def sne_color(df):
colors = []
for p in df["node_labels"]:
if p in ["", 'Agri Corpus<br>Data Spatiality<br>']:
colors.append("white")
elif p in ['With_SNE']:
colors.append("green")
elif p in ["WithOut_SNE"]:
colors.append("blue")
return colors
def tne_color(df):
colors = []
for p in df["node_labels"]:
if p in ["", 'Agri Corpus<br>Data Temporality<br>']:
colors.append("white")
elif p in ["<1 an"]:
colors.append("blue")
elif p in ["1 à 5 ans"]:
colors.append("brown")
else:
colors.append("red")
return colors
def drw_pie(df, colors):
# colors = tne_color(df)
fig=go.Figure(
data=go.Sunburst(
ids=df["node_names"],
labels=df["node_labels"],
parents=df["node_parent"],
marker=dict(colors=colors),
values=df["node_counts"],
branchvalues="total",
texttemplate = ('%{label}',
'%{label}<br>%{percentParent:.1%}',
'%{label}<br>%{percentParent:.1%}',
'%{label}<br>%{percentParent:.1%}',
'%{label}<br>%{percentParent:.1%}',
'%{label}<br>%{percentParent:.1%}',
'%{label}<br>%{percentParent:.1%}',
'%{label}<br>%{percentParent:.1%}',
'%{label}<br>%{percentParent:.1%}'),),)
fig.show()
#### Spatiality
import pandas as pd
import plotly.graph_objects as go
SNE_NODE = {'node_names': ['Corpus', 'With_SNE', 'WithOut_SNE'],
'node_parent': ["", "Corpus", "Corpus"],
'node_labels': ['Agri Corpus<br>Data Spatiality<br>','With_SNE', 'WithOut_SNE'],
#'node_counts': [len(corpus), len(corpus_with_extend), len(corpus_without_extend)]
'node_counts': [len(corpus_data), corpus_data['ent0'].isna().sum(), len(corpus_data)- corpus_data['ent0'].isna().sum()]
}
TNE_NODE = {'node_names': ['Corpus',"WithOut_TNE",'With_TNE', "<1 an", "1 à 5 ans","> 5 ans"],
'node_parent': ["", "Corpus", "Corpus", "With_TNE",'With_TNE','With_TNE'],
'node_labels': ['Agri Corpus<br>Data Temporality<br>',"WithOut_TNE",'With_TNE',"<1 an", "1 à 5 ans","> 5 ans"],
#'node_counts': [len(corpus), len(corpus_with_extend), len(corpus_without_extend)]
'node_counts': [len(corpus_data),len(corpus_data)-len(corpus_data_tne),len(corpus_data_tne), len(range_1), len(range_2),len(range_3)]
}
df = pd.DataFrame(TNE_NODE)
# colors = sne_color(df)
colors = tne_color(df)
drw_pie(df,colors)
```
| github_jupyter |
# Calculate DMD Mode Clusters (gDMD version)
***
This notebook should be run after calculating the DMD modes in the notebook `1_Calculate_DMD`. It hierarchically clusters DMD modes, assigns them into flat clusters, and plots the corresponding output modes.
Note that there are two versions of this notebook: a **gDMD** and **sDMD** version. These notebooks are nearly identical except that their default parameters have been chosen to be more appropriate for each case.
```
%load_ext watermark
%watermark -a "James Kunert-Graf" -d -v -m -p os,h5py,tqdm,numpy,matplotlib,scipy -g -u
import os
import h5py
from tqdm import tqdm_notebook as tqdm
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import scipy.cluster.hierarchy as sch
import scipy.spatial.distance as ssd
from hcp_dmd_utils import flat2mat,mat2flat
%matplotlib inline
```
## Parameters and Filepaths
***
The cell below contains parameters for the execution of the algorithm, along with variables specifying which data to analyze
**Parameters**
- *mthresh*: threshold on spatial continuity measure which filters out discontinuous modes before clustering
- *zthresh*: threshold on
- *min_clust_size*: clusters containing fewer than this number of modes are not kept for visualization
- *fclust_thresh*: threshold on flat clustering using `scipy.cluster.hierarchy.fcluster`
**Filepath Information**
The set of runs to analyze is identified using the same labels as were used in `1_Calculate_DMD`, which should be run first as this notebook relies on its saved output.
- *runlabel*: run label to analyze (e.g. REST1, REST2)
- *scandir*: scan direction to analyze (LR/RL)
- *nmodes*: choice of *nmodes* parameter used in DMD calculation (number of modes calculated in each window)
- *nframes*: choice of *nframes* parameter used in DMD calculation (window length, in frames)
- *ulist*: list of individuals to include in the clustering. For example:
- **ulist=u120** is full gDMD as in the paper, clustering the modes from all 120 individuals
- **ulist=u120[:10]** would run a smaller gDMD consisting of the first ten individuals
- **ulist=[u120[0]]** would run sDMD on the first individual (clustering only modes from the single individual)
```
##CLUSTERING PARAMETERS:
mthresh=25
zthresh=2.0
min_clust_size=400
fclust_thresh=0.955
## SCAN TO CLUSTER
runlabel='REST2'
scandir='RL'
nmodes=8
nframes=32
# Choose individuals to include in clustering:
u120=np.genfromtxt('u120.txt').astype(int).astype(str)
ulist=u120
```
### Reference RSNs
***
We use reference RSNs from Gordon et al. ("Generation and evaluation of a cortical area parcellation from resting-state correlations", *Cerebral Cortex*, 26(1):288–303, 2016.). We load them here and convert them into a vector form where we can easily compare them against our average modes.
```
#load RSN masks and labels
with h5py.File('RSN.h5','r') as hf:
RSNs=np.array(hf['RSNs'])
RSNlabels=np.array(hf['RSNlabels'])
rsn=[]
for k in range(RSNs.shape[2]):
r=RSNs[:,:,k]
rsn.append(mat2flat(r))
rsn=np.vstack(rsn)
rsnNo=np.arange(RSNs.shape[2])
rsnNo=np.delete(rsnNo,[0,7])
rsn=rsn[rsnNo,:]
rlabs=RSNlabels[rsnNo]
```
## Calculate Clusters for DMD
**Load DMD modes**:
Load and collect DMD mode information for the scan and set of individuals identified above.
```
try:
os.mkdir('./DMD_Clusters/')
except:
pass
try:
os.mkdir('./Figures/')
except:
pass
freq=[]
power=[]
F=[]
ux=[]
jx=[]
jno=[]
#LOAD WINDOWED DMD RESULTS
rx=[]
runNo=-1
for uid,u in enumerate(tqdm(ulist)):
#print '{:}/{:}'.format(uid+1,len(ulist))
fdir='./DMD_results/nmodes{:}_nframes{:}_{:}_{:}/'.format(nmodes,nframes,runlabel,scandir)
try:
with h5py.File(fdir+'Modes_{:}.h5'.format(u),'r') as hf:
F.append(np.array(hf['F']))
with h5py.File(fdir+'{:}.h5'.format(u),'r') as hf:
freq.append(np.array(hf['freq']))
power.append(np.array(hf['power']))
ux.append(np.array(hf['ux']))
jx.append(np.array(hf['jx']))
jno.append(np.array(hf['jno']))
runNo+=1
rx.append(runNo*np.ones(np.array(hf['jx']).shape))
except:
pass
freq=np.hstack(freq)
power=np.hstack(power)
F=np.vstack(F)
ux=np.hstack(ux);
jx=np.hstack(jx)
rx=np.hstack(rx)
jno=np.hstack(jno)
winNo=np.unique(np.array(['{:03.0f}{:03.0f}{:01.0f}'.format(ux[k],jx[k],rx[k]) for k in range(len(ux))]),return_inverse=True)[1]
```
**Pre-clustering mode filtration**:
Filter out modes which do not meet a certain standard of spatial continuity (threshold set above by parameter 'mthresh'). Then, filter out modes for which no pixel has a z-score above the threshold zthresh.
We also calculate the 'overlap' array, which is the dot product between the DMD modes and the RSN masks. This gives a measure of how much of each DMD mode's power is concentrated inside of a particular reference RSN (roughly indicating how strongly a mode resembles a given RSN).
```
#CALCULATE/THRESHOLD ON SPATIAL CONTINUITY
Mflat=(F-np.mean(F,1)[:,None])/F.std(1)[:,None]
mpro=[]
for f in Mflat:
m=flat2mat(f>2.0).astype(float)
#mpro: product of masks with masks which have been shifted diagonally
mpro.append(np.sum(np.sum(m[1:,1:]*m[:-1,:-1],0),0))
mpro=np.hstack(mpro)
mwhere=mpro>mthresh
#convert modes to zscores and threshold to create masks to cluster
Fz=F[mwhere,:]
Fz=(Fz-np.mean(Fz,1)[:,None])/np.std(Fz,1)[:,None]
Fn=(Fz>zthresh).astype(float)
#calculate overlap between mode masks and rsn masks
overlap=rsn.dot(Mflat.T)
```
**Clustering**:
Hierarchically cluster the modes using `scipy.cluster.hierarchy`. Heuristically, hierarchically clustering upon the average correlation works well, though this could be adjusted below.
Flat clusters are then formed based upon the "fclust_thresh" parameter defined above.
```
#cluster modes based upon average correlation between groups
method='average';metric='correlation';
Z=sch.linkage(Fn, method, metric)
ix=sch.dendrogram(Z,no_plot=True)['leaves']
#form flat clusters
cgroups=sch.fcluster(Z,fclust_thresh,'distance')[ix]
cgroups=cgroups.astype(float)
#count the number of modes belonging to each cluster, and remove clusters with less than 'min_clust_size' members
cun,count=np.unique(cgroups,return_counts=True)
for k,c in enumerate(cun):
if count[k]<min_clust_size:
cgroups[cgroups==c]=-1
cgroups=np.unique(cgroups,return_inverse=True)[1].astype(float)
cgroups[cgroups==0]=np.nan
cgroups+=-1
#cif =
cif=np.isfinite(cgroups)
cuni=np.unique(cgroups[cif])
#get identified clusters in terms of original indices
#so that they can be easily reloaded in the future...
outfile='./DMD_Clusters/{:}_{:}_{:}m_{:}f_{:}clusters2.h5'.format(runlabel,scandir,nmodes,nframes,len(cuni))
indices=np.arange(len(jx))[mwhere][ix]
cluster_indices=[]
for c in cuni:
cix=indices[cgroups==c]
cluster_indices.append(cix)
print '{:} clusters'.format(len(cuni))
#sort the RSN overlap array by hierarchical clustering order
Ou=overlap[:,mwhere][:,ix][:,cif]
Ou=(Ou-Ou.mean(1)[:,None])/Ou.std(1)[:,None]
import skimage.transform as skit
#'scale_factor' is a factor to scale down plotted image size -- use for many thousands of modes
scale_factor=10
Fsub=skit.downscale_local_mean(Fn[ix][cif],(scale_factor,1))
C=ssd.cdist(Fsub,Fsub,metric=metric)
C=np.abs(1-C);
np.fill_diagonal(C,0)
```
### Figure
***
Plot a figure showing the hierarchically clustered correlation matrix, the overlap of each mode with canonical RSNs, the automatically identified flat clusters, and the average DMD mode within each cluster:
```
plt.figure(figsize=(16,9))
a1=plt.subplot2grid((6,1),(0,0))
plt.imshow(C,vmin=C.mean(),vmax=C.mean()+2.0*C.std())
yl=plt.ylim()
plt.ylim(yl)
plt.xticks([])
plt.ylabel('DMD Mode')
a2=plt.subplot2grid((6,1),(1,0))
vmin=Ou.mean();vmax=Ou.mean()+2.0*Ou.std()
plt.imshow(skit.downscale_local_mean(Ou,(1,scale_factor)),aspect='auto',vmin=vmin,vmax=vmax)
plt.yticks(np.arange(len(rlabs)),rlabs,rotation=0)
a2.tick_params(pad=1,length=2)
xl=plt.xlim()
plt.xlim(xl)
plt.xlabel('DMD Mode')
a4=plt.subplot2grid((6,1),(2,0))
cbig=np.ones(cgroups.shape)*np.nan
ck=-1
for c in cuni:
if np.sum(cgroups==c)>=min_clust_size:
ck+=1
cbig[cgroups==c]=ck
plt.imshow(cbig[cif][:,None]%12,cmap='Paired',aspect='auto',vmin=0,vmax=12)
plt.xticks([]);plt.yticks([])
a5=plt.subplot2grid((6,1),(3,0),frameon=False)
lplot=np.where(cbig[cif][1:]!=cbig[cif][:-1])[0].astype(float)/(C.shape[0]*scale_factor)
cut=0.95
for lp in lplot:
plt.plot([0,cut],[lp]*2,':',c=[1,1,1],lw=1.1)
plt.plot([cut,1],[lp]*2,':',c=[0,0,0],lw=1.1)
plt.xlim([0,1])
plt.ylim([0,1])
plt.xticks([])
plt.yticks([])
a5.invert_yaxis()
a6=plt.subplot2grid((6,1),(4,0),frameon=False)
lplot=np.where(cbig[cif][1:]!=cbig[cif][:-1])[0].astype(float)/(C.shape[0]*scale_factor)
cut=0.95
for lp in lplot:
plt.plot([lp]*2,[0,1],':',c=[1,1,1],lw=1.1)
plt.xlim([0,1])
plt.ylim([0,1])
plt.xticks([])
plt.yticks([])
x1=0.0578
y1=0.3
y2=0.08
w1=0.364
a1.set_position([0.00,y1,0.48,0.65])
a2.set_position([x1,y2,w1,0.21])
a4.set_position([0.426,y1,0.025*0.8,0.65])
a5.set_position([x1,y1,w1+0.025,0.65])
a6.set_position([x1,y2,w1,0.87])
glist=np.unique(cgroups[np.isnan(cgroups)==False]).astype(int)
#%
cf=[]
for gid in glist:
cf.append(np.mean(np.absolute(freq[mwhere][ix][cgroups==gid])))
cf=np.array(cf)
glist_big=[]
for gid in glist:
if np.sum(cgroups==gid)>=min_clust_size:
glist_big.append(gid)
glist_big=np.array(glist_big)
DMN=[]
for gk,gid in enumerate(glist_big):
a6=plt.axes([0.45+0.275*(gk%2),0.714-0.15*np.floor(gk/2),0.26,0.35])
colors=mpl.cm.Paired(range(12))
a6.set_facecolor(colors[gk%12])
dmn=np.mean(F[mwhere][ix][cgroups==gid],0)
vmin=dmn.mean()+0.0*dmn.std();vmax=dmn.mean()+2.0*dmn.std()
DMN.append(dmn)
D=flat2mat(dmn)
D[D==0]=np.nan
D=np.concatenate((D[:40],D[40:]),1)
cm=mpl.cm.viridis
cm.set_bad([1,1,1,0])
plt.imshow(D,vmin=vmin,vmax=vmax,cmap=cm)
plt.xticks([]);plt.yticks([])
cf=freq[mwhere][ix][cgroups==gid]
lum=np.sum(np.array([0.299,0.587,0.114])*colors[gid%12][:3])
if lum>0.65:
tcolor=[0,0,0]
else:
tcolor=[1,1,1]
plt.text(159,40,'{:0.2f} +/- {:0.2f} Hz'.format(cf.mean(),cf.std()),color=tcolor,ha='right',va='bottom')
DMN=np.vstack(DMN)
plt.savefig('./Figures/fig2b_{:}_{:}.svg'.format(runlabel,scandir))
```
### Output
***
Save the indices of the modes which constitute each of the plotted clusters, and the average mode within each cluster.
```
with h5py.File(outfile,'w') as hf:
dt = h5py.special_dtype(vlen=np.dtype('int32'))
hf.create_dataset('clusters',data=np.array(cluster_indices),dtype=dt)
hf.create_dataset('modes',data=DMN)
```
| github_jupyter |
# MW-M31 Post-Merger Remnant: masses and densities
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Setup" data-toc-modified-id="Setup-1"><span class="toc-item-num">1 </span>Setup</a></span></li><li><span><a href="#Shape" data-toc-modified-id="Shape-2"><span class="toc-item-num">2 </span>Shape</a></span></li><li><span><a href="#Mass-profiles" data-toc-modified-id="Mass-profiles-3"><span class="toc-item-num">3 </span>Mass profiles</a></span><ul class="toc-item"><li><span><a href="#Hernquist-fits" data-toc-modified-id="Hernquist-fits-3.1"><span class="toc-item-num">3.1 </span>Hernquist fits</a></span></li><li><span><a href="#Sersic-fits" data-toc-modified-id="Sersic-fits-3.2"><span class="toc-item-num">3.2 </span>Sersic fits</a></span></li></ul></li><li><span><a href="#Densities" data-toc-modified-id="Densities-4"><span class="toc-item-num">4 </span>Densities</a></span></li><li><span><a href="#Virial-radius/mass" data-toc-modified-id="Virial-radius/mass-5"><span class="toc-item-num">5 </span>Virial radius/mass</a></span></li></ul></div>
## Setup
```
# import modules
import numpy as np
import numpy.linalg as la
from numpy.linalg import norm, eigh
from scipy.optimize import curve_fit
import pandas as pd
import astropy.units as u
from astropy.constants import G
from astropy.cosmology import Planck15, z_at_value
# import plotting modules
import matplotlib
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
import mpl_scatter_density
from matplotlib import rcParams
from matplotlib.patches import Ellipse
#
# Make the norm object to define the image stretch
from astropy.visualization import LogStretch
from astropy.visualization.mpl_normalize import ImageNormalize
im_norm = ImageNormalize(vmin=0., vmax=1000, stretch=LogStretch())
%matplotlib inline
# my modules
from galaxy.galaxy import Galaxy
from galaxy.centerofmass import CenterOfMass
from galaxy.massprofile import MassProfile
from galaxy.plots import Plots
from galaxy.timecourse import TimeCourse
from galaxy.utilities import find_nearest, rotation_matrix_to_vector, z_rotation_matrix
from galaxy.remnant import Remnant
tc = TimeCourse()
p = Plots()
# Just luminous particles: ptype defaults to (2,3)
remnant = Remnant(usesql=True)
remnant.data, remnant.data.shape
def get_counts(remnant):
df = pd.DataFrame()
df['gal'] = remnant.data['galname']
df['type'] = remnant.data['type']
# create some better column names
types = {1: 'Halo', 2: 'Disk', 3: 'Bulge'}
df['typename'] = df['type'].map(types)
# get pandas to do most of the work
df_piv = pd.pivot_table(df, values='type',
index='gal', columns='typename',
aggfunc='count', fill_value=0, margins=True)
return df_piv
counts = get_counts(remnant) / 1000
counts
print(counts.to_latex())
```
## Shape
Get remnant-centered coordinates: just translation, no rotation at this stage
```
com = CenterOfMass(remnant, ptype=None)
xyz, vxyz = com.center_com()
xyz.shape
```
Define some methods to calculate principal axes of the (presumed) ellipsoid:
```
def get_where(gal, ptype):
if gal == 'MW':
gal = 'MW ' # annoying!
return np.where((remnant.data['galname']==gal) & (remnant.data['type']==ptype))
def sub_ellipsoid_axes(subset, r_lim=None):
x,y,z = (xyz.T[subset]).T
m = remnant.data[subset]['m']
return remnant.ellipsoid_axes(m, x, y, z, r_lim)
```
Subset the data by origin, galaxy/particle type:
```
MWh = get_where('MW', 1)
MWd = get_where('MW', 2)
MWb = get_where('MW', 3)
M31h = get_where('M31', 1)
M31d = get_where('M31', 2)
M31b = get_where('M31', 3)
x,y,z = xyz
m = remnant.data['m']
abc = {}
axes = {}
```
Get inertia tensor and eigenvalues
```
r_lim = 40
r = norm(xyz, axis=0)
central = np.where(r < r_lim)
m_c = m[central]
x_c = x[central]
y_c = y[central]
z_c = z[central]
xyz_c = np.array([x_c, y_c, z_c])
I = remnant.I_tensor(m_c, x_c, y_c, z_c)
w, v = eigh(I)
# moments of intertia around principal axes:
A, B, C = w / np.max(w)
A, B, C
min_axis = I[0]/norm(I[0])
mid_axis = I[1]/norm(I[1])
maj_axis = I[2]/norm(I[2])
min_axis, mid_axis, maj_axis
xyz_c.shape, xyz_c.shape[1]/xyz.shape[1]
R = rotation_matrix_to_vector(maj_axis)
R
x_r, y_r, z_r = R @ xyz_c
R @ min_axis, R @ mid_axis
np.arccos(np.dot(R @ min_axis, np.array([1,0,0]))) * 180/np.pi
fig = plt.figure(figsize=(18,6))
lim = 30
fontsize = 22 # for labels
bins = 100
# left plot x-y
ax0 = fig.add_subplot(1, 3, 1, projection='scatter_density')
ax0.scatter_density(x_r, y_r, norm=im_norm)
ax0.set_xlim(-lim, lim)
ax0.set_ylim(-lim, lim)
ax0.set_xlabel('x (kpc)', fontsize=fontsize)
ax0.set_ylabel('y (kpc)', fontsize=fontsize)
level_vals = [0.5, 0.75, 0.9]
colors = ['red','orange', 'yellow', 'orange', 'yellow']
p.density_contour(x_r, y_r, bins, bins, level_vals, ax=ax0, colors=colors)
# mid plot x-z
ax1 = fig.add_subplot(1, 3, 2, projection='scatter_density')
ax1.scatter_density(x_r, z_r, norm=im_norm)
ax1.set_xlim(-lim, lim)
ax1.set_ylim(-lim, lim)
ax1.set_xlabel('x (kpc)', fontsize=fontsize)
ax1.set_ylabel('z (kpc)', fontsize=fontsize)
c = p.density_contour(x_r, z_r, bins, bins, level_vals, ax=ax1, colors=colors)
# right plot y-z
ax2 = fig.add_subplot(1, 3, 3, projection='scatter_density')
ax2.scatter_density(y_r, z_r, norm=im_norm)
ax2.set_xlim(-lim, lim)
ax2.set_ylim(-lim, lim)
ax2.set_xlabel('y (kpc)', fontsize=fontsize)
ax2.set_ylabel('z (kpc)', fontsize=fontsize);
p.density_contour(y_r, z_r, bins, bins, level_vals, ax=ax2, colors=colors)
#adjust tick label font size
label_size = 16
matplotlib.rcParams['xtick.labelsize'] = label_size
matplotlib.rcParams['ytick.labelsize'] = label_size
plt.tight_layout()
plt.savefig('remnant_shape_tensor.pdf', rasterized=True, dpi=350);
segs = c.allsegs
len(segs), len(segs[2]), len(segs[2][0])
(segs[2][0].T[0].min() + segs[2][0].T[0].max())/2
(segs[2][0].T[1].min() + segs[2][0].T[1].max())/2
np.mean(segs[2][0].T[0]), np.mean(segs[2][0].T[1])
lim=8
fig, ax = plt.subplots(figsize=(8,8))
x = segs[2][0].T[0] - np.mean(segs[2][0].T[0])
y = segs[2][0].T[1] - np.mean(segs[2][0].T[1])
ax.plot(x, y, 'r.', ms=20)
ax.set_xlim(-lim, lim)
ax.set_ylim(-lim, lim)
ax.add_artist(Ellipse((0,0), 15, 9, 60, fill=False, lw=3));
# r = np.sqrt(x**2 + y**2)
# theta = np.arctan2(y, x) * 180/np.pi
# fig, ax = plt.subplots(figsize=(8,8))
# ax.plot(theta, r, 'r.', ms=20)
# lim=30
# fig, ax = plt.subplots(figsize=(8,8))
# x = segs[0][0].T[0] - np.mean(segs[0][0].T[0])
# y = segs[0][0].T[1] - np.mean(segs[0][0].T[1])
# ax.plot(x, y, 'r.', ms=10)
# # ax.set_xlim(-lim, lim)
# # ax.set_ylim(-lim, lim)
# ax.add_artist(Ellipse((0.5,-0.5), 50, 38, 80, fill=False, lw=3));
```
Repeat on subgroups:
```
# abc['total'], axes['total'] = remnant.ellipsoid_axes(m, x, y, z, r_lim)
# abc['MWd'], axes['MWd'] = sub_ellipsoid_axes(MWd, r_lim)
# abc['MWb'], axes['MWb'] = sub_ellipsoid_axes(MWb, r_lim)
# abc['M31d'], axes['M31d'] = sub_ellipsoid_axes(M31d, r_lim)
# abc['M31b'], axes['M31b'] = sub_ellipsoid_axes(M31b, r_lim)
# abc, axes
```
Use pandas to organize and output the results:
```
# df = pd.DataFrame()
# df['set'] = np.array([k for k, _ in abc.items()])
# df['a'] = np.array([np.round(v[0], 2) for _, v in abc.items()])
# df['b'] = np.array([np.round(v[1], 2) for _, v in abc.items()])
# df['c'] = np.array([np.round(v[2], 2) for _, v in abc.items()])
# df
# print(df.to_latex(index=False))
```
What is the relative orientation of component ellipsoids?
```
def tilt_angle(group1, group2):
# these should already be unit vectors, but normalise for safety
vec1 = axes[group1][0]/la.norm(axes[group1][0])
vec2 = axes[group2][0]/la.norm(axes[group2][0])
dot = np.dot(vec1, vec2)
return np.round(np.arccos(np.abs(dot)) * 180 / np.pi, 1)
# groups = [k for k,_ in abc.items()]
# tilts = pd.DataFrame([[tilt_angle(g1, g2) for g1 in groups] for g2 in groups], columns=groups)
# print(groups)
# tilts
# print(tilts.to_latex())
```
| github_jupyter |
```
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data
```
# Load dataset
```
mnist = input_data.read_data_sets("MNIST_data")
train_data = mnist.train.images
train_labels = mnist.train.labels
validation_data = mnist.validation.images
validation_labels = mnist.validation.labels
test_data = mnist.test.images
test_labels = mnist.test.labels
print("train_data", train_data.shape, "train_labels", train_labels.shape)
print("validation_data", validation_data.shape, "validation_labels", validation_labels.shape)
print("test_data", test_data.shape, "test_labels", test_labels.shape)
```
# Create input functions for training, validation and testing
```
train_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"x": train_data},
y=train_labels,
batch_size=64,
num_epochs=10,
shuffle=True
)
validation_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"x": validation_data},
y=validation_labels,
num_epochs=1,
shuffle=False)
test_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"x": test_data},
y=test_labels,
num_epochs=1,
shuffle=False)
```
# Create model function
```
def my_model_fn(features, labels, mode):
"""Model function for our CNN"""
net = tf.reshape(features['x'], [-1, 28, 28, 1])
for _ in range(3):
net = tf.layers.conv2d(
inputs=net,
filters=32,
kernel_size=[3, 3],
padding="same",
activation=tf.nn.relu
)
net = tf.layers.max_pooling2d(
inputs=net,
pool_size=[2, 2],
strides=[2,2]
)
net = tf.layers.flatten(net)
net = tf.layers.dense(inputs=net, units=64)
logits = tf.layers.dense(inputs=net, units=10)
predictions = {
'classes': tf.argmax(input=logits, axis=1),
'probabilities': tf.nn.softmax(logits, name='softmax_tensor')
}
if mode == tf.estimator.ModeKeys.PREDICT:
export_outputs = {
'predictions': tf.estimator.export.PredictOutput(predictions)
}
return tf.estimator.EstimatorSpec(mode=mode, predictions=predictions, export_outputs=export_outputs)
onehot_labels = tf.one_hot(indices=tf.cast(labels, tf.int32), depth=10)
loss = tf.losses.softmax_cross_entropy(onehot_labels=onehot_labels,
logits=logits)
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.001)
train_op = optimizer.minimize(loss=loss, global_step=tf.train.get_global_step())
eval_metric_ops = {
"accuracy": tf.metrics.accuracy(labels=labels, predictions=predictions['classes'])
}
return tf.estimator.EstimatorSpec(mode=mode,
loss=loss,
train_op=train_op,
eval_metric_ops=eval_metric_ops)
```
# Create estimator
```
mnist_estimator = tf.estimator.Estimator(
model_fn=my_model_fn,
model_dir="E:\\temp\\mnist_estimator"
)
```
# Train and evaluate
```
train_spec = tf.estimator.TrainSpec(
input_fn=train_input_fn,
max_steps=20000,
)
validation_spec = tf.estimator.EvalSpec(
input_fn=validation_input_fn,
steps=50, throttle_secs=60
)
tf.estimator.train_and_evaluate(mnist_estimator, train_spec, validation_spec)
```
# Perform evaluation on the test set
```
mnist_estimator.evaluate(input_fn=test_input_fn)
```
# Getting the result of each prediction
```
test_results = mnist_estimator.predict(input_fn=test_input_fn)
print(test_results)
import itertools
list(itertools.islice(test_results, 3))
```
| github_jupyter |
# Largest Rectangle in Histogram
## Approach
### Complexity `O(n)`
For any bar i the maximum rectangle is of width `r - l - 1` where r - is the last coordinate of the bar to the right with height `h[r] >= h[i]` and l - is the last coordinate of the bar to the left which height `h[l] >= h[i]`
The meaning of `r` and `l` is somewhat confusing, to put them more accurately:
`l`: the first coordinate of the bar to the left with height `h[l] < h[i]`.
`r`: the first coordinate of the bar to the right with height `h[r] < h[i]`.
[image](https://i.loli.net/2018/10/29/5bd65b33c2798.png)
So if for any i coordinate we know his utmost higher (or of the same height) neighbors to the right and to the left, we can
easily find the largest rectangle:
```java
int maxArea = 0;
for (int i = 0; i < height.length; i++) {
maxArea = Math.max(maxArea, height[i] * (lessFromRight[i] - lessFromLeft[i] - 1));
}
```
The main trick is how to effectively calculate lessFromRight and lessFromLeft arrays. The trivial solution is to use `O(n^2) `solution and for each i element first find his left/right heighbour in the second inner loop just iterating back or forward:
```java
for (int i = 1; i < height.length; i++) {
int p = i - 1;
while (p >= 0 && height[p] >= height[i]) {
p--;
}
lessFromLeft[i] = p;
}
```
The only line change shifts this algorithm from `O(n^2) -> O(n)` complexity: we don't need to rescan each item to the left - we can reuse results of previous calculations and "jump" through indices in quick manner:
```java
while (p >= 0 && height[p] >= height[i]) {
p = lessFromLeft[p];
}
```
Here is the whole solution:
```java
public static int largestRectangleArea(int[] height) {
if (height == null || height.length == 0) {
return 0;
}
int[] lessFromLeft = new int[height.length]; // idx of the first bar the left that is lower than current
int[] lessFromRight = new int[height.length]; // idx of the first bar the right that is lower than current
lessFromRight[height.length - 1] = height.length;
lessFromLeft[0] = -1;
for (int i = 1; i < height.length; i++) {
int p = i - 1;
while (p >= 0 && height[p] >= height[i]) {
p = lessFromLeft[p];
}
lessFromLeft[i] = p;
}
for (int i = height.length - 2; i >= 0; i--) {
int p = i + 1;
while (p < height.length && height[p] >= height[i]) {
p = lessFromRight[p];
}
lessFromRight[i] = p;
}
int maxArea = 0;
for (int i = 0; i < height.length; i++) {
maxArea = Math.max(maxArea, height[i] * (lessFromRight[i] - lessFromLeft[i] - 1));
}
return maxArea;
}
```
[Question Link](https://leetcode.com/problems/largest-rectangle-in-histogram/)
| github_jupyter |
# This notebook creates the stopwords csvs needed for word vectorizations
```
import csv
cooking_stop_words = list(set([
'canned', 'cans', 'drained', 'and', 'halved', 'cup', 'cups',
'teaspoon', 'tablespoon', 'teaspoons', 'tablespoons',
'finely', 'freshly', 'fresh', 'thickcut', 'to', 'taste',
'grated', 'cut', 'into', 'wedges', 'pounds', 'unpeeled', 'large',
'minced', 'slice', 'slices', 'sliced', 'thick-cut', 'cut',
'crosswise', 'pieces', 'toothpicks', 'low-fat', 'chopped', 'or',
'taste', 'cooked', 'dry', 'shredded', 'beaten', 'dried', 'melted',
'stems', 'removed', 'diced', 'ounce', 'ounces', 'packages',
'softened', 'such', 'RedHot®', 'RedHot', 'Franks', "Frank's",
'crumbled', 'Old', 'Bay®', 'Bay', 'pinch', 'for', 'garnish', 'slice',
'slices', 'needed', 'inch', 'cubes', 'cooking', 'spray', 'ground',
'rotisserie', 'lowfat', 'as', 'quarteres', 'cloves', 'more', 'can',
'package', 'frozen', 'thawed', 'packet', 'reducedfat', 'Knorr',
'container', 'pound', 'peeled', 'deveined', 'seeded', 'ripe',
'English', 'juiced', 'plus', 'more', 'Hass', 'cubed', 'Mexicanstyle',
'hearts', 'prepared', 'party', 'pitted', 'mashed',
'roma', 'optional', 'chunk', 'Hot', 'bunch', 'cleaned', 'box',
'chickenflavored', 'Golden', 'delicious', 'cored', 'any', 'flavor',
'flavored', 'whole', 'allpurpose', 'all', 'purpose', 'deep', 'frying',
'dash', 'packed', 'in', 'French', 'jar', 'small', 'head', 'little',
'smokie', 'seasoned', 'Boston', 'Bibb', 'leaves', 'lean', 'pickled',
'Asian', 'dark', 'flaked', 'rolled', 'packed', 'jellied',
'thirds', 'with', 'attached', 'skewers', 'skinless', 'boneless',
'half', 'kernels', 'rinsed', 'quart', 'quarts', 'kernel',
'Italianstyle', 'unpopped', 'lightly', 'coating', 'SAUCE',
'lengthwise', 'miniature', 'semisweet', 'rinsed', 'round',
'squeezed', 'stewed', 'raw', 'the', 'liquid', 'reserved', 'medium',
'instant', 'solid', 'pack', 'refrigerated', 'halves', 'distilled',
'loaf', 'extra', 'virgin', 'crushed', 'kosher', 'toasted', 'buttery',
'TM', 'panko', 'Japanese', 'regular', 'bottle', 'bottles', 'thin',
'peel', 'paper', 'thick', 'circles', 'unbleached',
'breast', 'breasts', 'wings', 'strips', 'jumbo', 'giant', 'chunks',
'quickcooking', 'sweetened', 'flakes', 'Ranchstyle', 'snipped',
'food', 'ROTEL', 'Italian', 'sticks', 'stick', 'crescent', 'thinly',
'boiled', 'Genoa', 'roasted', 'thin', 'extrasharp', 'pressed',
'sifted', 'split', 'tips', 'discarded', 'mini', 'deli', 'drain',
'reserve', 'diameter', 'Greek', 'Thai', 'drops', 'square', 'crusty',
'American', 'selfrising', 'imitation', 'Wings', 'apart', 'at',
'joints', 'wing', 'tips', 'discarded', 'parts',
'tops', 'seperated', 'blend', 'coarsely', 'sweet', 'stalk', 'heads',
'husked', 'divided', 'pats', 'unsalted', 'active', 'warm', 'sea',
'separated', 'herb', 'overripe', 'degrees', 'F', 'C', 'room',
'temperature', 'machine', 'very', 'pint', 'puree', 'coarse',
'envelopes', 'lukewarm', 'creamstyle', 'unsweetened',
'lite', 'of', 'chilled', 'freezer', 'cold', 'brushing', 'nonfat',
'squares', 'tails', 'thigh', 'quarters', 'Masterpiece', 'KC', 'from',
'El', 'Paso', 'bulk', 'Hunts', 'Roma', 'light', 'fluid', 'lagerstyle',
'stalks', 'quartered', 'undrained', 'drained', 'Tony', 'Chacheres',
'lump', 'uncooked', 'cube', 'bits', 'hair', 'angel', 'trimmed',
'stew', 'brisket', 'bitesized', 'matchstick', 'Chobani',
'unbaked', 'crust', 'torn', 'bonein', 'pounded', 'bitesize',
'granules', 'boiling', 'yolk', 'coloring', 'pinch', 'a', 'blender',
'fine', 'which', 'extralarge', 'use', 'will', 'make', 'garnish',
'barely', 'moistened', 'about', 'right', 'before', 'serving', 'mix',
'thinly-sliced', 'thinly-sliced', 'etc', 'excess', 'granulated', 'peeled',
'extra-virgin', 'pinch'
]))
unhelpful = list(set(['fresh', 'ripe', 'cracked', 'cooking', 'coarse', 'light',
'mild', 'hot', 'minced', 'dark roast', 'unsifted', 'canned',
'cans', 'drained', 'halved', 'finely', 'freshly', 'thickcut',
'grated', 'cut', 'unpeeled', 'large', 'minced', 'slice',
'slices', 'sliced', 'chopped','shredded', 'beaten', 'dried',
'melted', 'stems', 'softened', 'packages', 'crumbled', 'ground',
'low-fat', 'rotisserie', 'lowfat', 'can', 'thawed', 'packet',
'reducedfat', 'small', 'pats', 'regular', 'lukewarm', 'mashed',
'stalk', 'breast', 'breasts', 'juiced', 'halves', 'extrasharp',
'sharp', 'extra sharp', 'frozen', 'raw', 'warm', 'divided',
'little', 'squares', 'thinly', 'thick', 'rinsed', 'toasted',
'bitesize', 'chunks', 'refrigerated', 'kernel', 'kernels',
'jar', 'lengthwise', 'unpeeled', 'cleaned', 'paper', 'melted',
'separated', 'seperated', 'deveined', 'party', 'bunch', 'overripe',
'boiled', 'chunk', 'container', 'bitesized', 'sweet', 'strips',
'sifted', 'roma', 'very', 'undrained', 'stewed', 'thawed', 'lean',
'roasted', 'extra', 'lite', 'coarsely', 'pressed', 'square',
'jumbo', 'yolk', 'yolks', 'barely', 'pitted', 'cored', 'puree',
'cubes', 'angel', 'hair', 'angelhair', 'giant', 'husked', 'chilled',
'thigh', 'trimmed', 'thin', 'lightly', 'cubed', 'drops', 'grated',
'boneless', 'unsalted', 'pieces', 'skinless', 'pounded',
'chickenflavored', 'extralarge', 'medium', 'reserve', 'unbaked',
'crushed', 'wings', 'crosswise', 'cold', 'bonein', 'bone in',
'squeezed', 'kosher', 'miniature', 'tails', 'quarters', 'attached',
'loaf', 'dry', 'more', 'head', 'removed', 'packed', 'hearts',
'matchstick', 'unbleached', 'heads', 'stems', 'sea', 'diced',
'mini', 'cut', 'unpopped', 'box', 'uncooked', 'freezer', 'stalks',
'shredded', 'halved', 'snipped', 'thick-cut', 'split', 'seeded',
'sweetened', 'discarded', 'lump', 'boiling', 'whole', 'semisweet',
'semi-sweet', 'quartered', 'moistened', 'reserved', 'prepared',
'fresh', 'ripe', 'cracked', 'cooking', 'coarse', 'light', 'mild',
'hot', 'minced', 'dark roast', 'unsifted', 'quaker', 'raw', 'frozen',
'calore-wise', 'ziploc bag', 'real', 'lite', 'crisp', 'decaffeinated',
'canned', 'processed', 'cooked', 'unpeeled', 'also', 'store-bought',
'accepted', 'accommodate', 'accompaniment', 'accompaniments',
'according', 'across', 'acting', 'active', 'add', 'added', 'additional',
'additive', 'adds', 'adjoining', 'adjustable', 'adjusting', 'aged',
'ahead', 'aisle', 'aka', 'almost', 'along', 'alternatively', 'aluminum',
'amazon', 'amount', 'amounts', 'angle', 'angled', 'animal', 'another',
'apart', 'appetizer', 'appropriate', 'approx', 'approximately', 'area',
'aroma', 'aromatic', 'around', 'aside', 'ask', 'assembling', 'assorted',
'attachment', 'attachments', 'authentic', 'available', 'avoid', 'away',
'bag', 'bags', 'baked', 'baker', 'bakery', 'bakeware', 'ball', 'balls',
'bands', 'based', 'basket', 'baskets', 'baster', 'basting', 'bearded',
'beards', 'beating', 'becomes', 'bench', 'best', 'better', 'big', 'bit',
'bite', 'bits', 'blade', 'blend', 'blended', 'blender', 'blotted', 'body',
'boil', 'boning', 'bottom', 'bottomed', 'bottoms', 'bought', 'bowl', 'bowls',
'boxes', 'braised', 'braising','branch','branches','brand','brands','brewed',
'briefly','bright','broil','broiled','broken','brought','brush','brushed',
'brushes','brushing','bulk','burn','burner','burning','buy','call','called',
'calorie','canal','canner','canning','canvas','capacity','carbon','cardboard',
'cards','careful','carefully','carton','case','casing','casings','cast',
'catalogue','cavity','centimeter','ceramic','changes','charcoal','charlotte',
'charred','cheap','chef','chewy','chiffonade','choice','chopping','chopstick',
'chopsticks','cl','clarified','clark','classic','clean','clear','cleaver','click',
'cloth','club','cm','coarser','coat','coated','coating','coiled','coils','coin','coins',
'colander','coleman','collapsible','colman','color','colored','com','combination',
'combine','combined','come','comes','commercial','commercially','commonly','company',
'completely','con','condiment','condiments','connective','consistency','containers',
'containing','contains','content','converted','cook','cooker','copper','core','cores',
'corkscrew','count','counter','couple','course','court','cracks','craft','credit',
'crisply','crock','cross','crosscut','cultivated','cupmirin','cupsfresh','cupspanko',
'cupstomato','cupsturkey','cure','cured','curing','curled','curly','cutinto','damaged',
'damp','dampened','damson','dark','dash','dashes','de','debearded','deboned','decorate',
'decorating','decoration','decorations','decorative','deep','defatted','defrosted','del',
'dense','depending','desert','desiccated','desired','dessert','dessicated','diagonal',
'diagonally','diameter','diamond','dice','diet','different','difficult','digital',
'diluted','dim','dinner','dipper','dipping','directions','discard','discarding',
'discolored','dish','dishes','disk','disks','disposable','dissolved','distilled',
'domestic','done','doone','double','dover','dowel','dozen','drain','draining',
'dredging','drink','drips','drizzle','drizzling','drop','dust','dusting','duty',
'dyed','easier','east','easter','eastern','easy','edge','edged','edges','edible',
'edward','eight','eighteen','eighth','eighths','either','el','electric','elixir',
'em','empty','en','enameled','end','ends','english','enough','epi','equipment',
'equivalent','etc','ethnic','european','even','ever','exceed','excellence',
'excellent','excess','excluding','exotic','expose','exposed','extremely','eyed',
'fairly','family','famous','fan','fancy','farm','farmed','farmer','farmers',
'farms','fashioned','fast','faster','favorite','feathers','fed','feet','fermented',
'fiery','filled','filling','fillings','filter','filtered','find','fine','finishing',
'fire','firm','firmly','first','fishmonger','fit','fitted','fitting','five','fl',
'flameproof','flattened','flavor','flavored','flavorful','flavoring','flavors',
'flexible','floral','fluid','flying','foam','foamy','foil','following','follows',
'food','foods','forced','form','formed','forming','forms','found','four','fourths',
'fragments','fragrant','free','freeze','freshepazote','freshest','freshwater','fried',
'fries','frosting','full','fully','fun','ga','gallon','gallons','garnish','garnishes',
'garnishing','gas','gem','generous','generously','gently','girl','give','gives','glass',
'glasses','glitter','globe','glove','gloves','go','goes','good','gourmet','grade',
'grain','grained','grains','grainy','gram','grams','granulated','grate','grater',
'grating','gratings','greased','greasing','great','griddle','grill','grilled',
'grilling','grind','grinder','grinding','grindings','grinds','groceries','grocery',
'gummy','gutted','ha','half','hammer','hand','handful','handfuls','handheld','handle',
'handles','handling','hands','hard','headnote','health','heaped','heaping','hearty',
'heat','heated','heatproof','heavily','heavy','hellmann','herb','herbed','herbes',
'herbs','high','hinged','hispanic','hold','hole','holes','holiday','holland',
'hollow','hollowed','holy','home','homemade','hon','honeyed','hook','hoop',
'horizontally','hour','hours','house','hulled','icing','id','igourmet',
'imitation','imported','inch','inches','include','included','including',
'increase','india','indian','individual','indonesian','info','ingredient',
'ingredients','inner','insert','inside','insist','insta','instacure','instant',
'instead','instructions','intact','interior','intervals','inverted','irish',
'iron','island','israeli','italian','jarred','jars','jell','john','joint',
'joints','juice','juices','juicy','keep','keeping','kept','kettle','key',
'kg','kha','kikkoman','kilogram','kind','king','kirby','kitchen','kneading',
'knife','knob','knot','known','knuckle','kum','la','label','labeled','ladle',
'larger','last','late','latin','latino','latter','laver','layer','layers',
'leafed','leafy','least','leathery','leave','leaved','leaving','lebanese',
'left','leftover','length','lengths','less','let','level','leveled','leveling',
'lexington','lid','lids','like','lined','liner','liners','lining','link','links',
'lip','liquefy','liquid','list','liter','liters','litres','live','lo','local',
'log','logs','long','loofah','look','loose','loosely','loosen','lot','lots',
'low','lower','machine','made','mae','mail','major','make','maker','makes',
'making','maldon','malted','mam','manischewitz','manual','many','marbled',
'marked','market','markets','mash','mason','master','mat','may','maytag',
'measure','measured','measuring','meaty','medallions','mediterranean','melt','melting','members','metal','method','mexican','mexicana','mexico','mg','micro','microplane','middle','milder','mildly','mill','milled','milliliter','milliliters','minces','minus','minute','minutes','mission','mitts','mix','mixed','mixer','mixing','mixture','ml','mm','moderately','moist','moistening','moisture','mold','molds','moon','moons','mortar','mountain','mrs','mt','much','mullet','multi','multigrain','nationwide','native','natural','naturally','near','necessary','need','needed','needle','needlenose','needles','nests','neutral','nine','non','nonreactive','nonstick','nontoxic','north','northern','notes','number','nutritional','oblong','offset','often','old','ole','omit','omitting','one','ones','online','oounces','open','opened','opening','opposite','optional','order','ordered','organic','oriental','original','ounceahi','outer','outside','oven','ovenproof','overnight','pack','package','packaged','packets','pad','page','paintbrush','pair','pan','pans','papery','parboiled','parchment','pareve','paring','park','part','partially','parts','parve','passed','passing','passion','passover','pasteurized','pat','patted','pattern','patties','peasant','peeled','peeler','peeling','pencil','people','peppered','pepperidge','per','percent','perfect','perfectly','perforated','perilla','pernod','persian','person','pesticide','pestle','petals','petit','petite','petits','philadelphia','pi','pick','picked','pickle','pickled','pickling','picks','picnic','piece','pierced','pin','pinch','pinches','pint','pints','piping','pit','pits','pkg','place','places','plain','plane','plant','plastic','plate','plates','platter','platters','pleated','plenty','plump','plumped','plunged','plus','poached','poaching','pocket','pocketless','pockets','pod','pods','point','points','polish','polished','pollen','pop','popped','portion','portions','portuguese','possible','pot','pounder','pour','poured','powder','powdered','powdery','power','pre','prebaked','precooked','prefer','preferable','preferably','preferred','preheat','premade','premium','preparation','prepare','preserved','preserves','presliced','press','pressure','pretty','prevent','prickly','prime','prince','problem','procedure','process','processor','produce','product','products','pt','pulled','pullman','pulp','pulsed','pulverized','purchase','purchased','pure','purpose','qt','qts','quality','quart','quarter','quarts','quatre','quick','quill','quills','rack','racks','raised','ram','ramekin','ramekins','range','rapid','rashers','rather','razor','read','ready','really','recipe','recipelink','recipes','recommend','recommended','reconstituted','rectangle','rectangles','rectangular','reduce','reduced','referred','refried','refrigerator','reheated','release','remainder','remaining','remains','removable','remove','rendered','request','resealable','resembles','rest','restaurant','results','retain','rib','ribbon','ribbons','ricer','rich','rick','rim','rimmed','rinse','ripened','river','roast','roasting','roasts','robust','rock','roll','rolled','rolling','rolls','room','rough','roughly','round','rounded','rounds','royal','rub','rubbed','rubber','rubbing','ruler','runny','rustic','rustique','sack','safe','safety','saucepan','save','sawdust','sawed','scalded','scale','scaled','scales','scant','scharffen','scissors','scoop','scooped','scooping','scoops','scored','scout','scrambled','scraped','scraper','scraps','screw','scrubbed','sealable','sealing','seasonal','seasonally','seasoned','seasoning','seasonings','second','seconds','section','sectioned','sections','securely','sediment','segment','segmented','segments','self','semi','semicircles','semidried','semisoft','separate','separately','separator','serve','serving','set','seven','several','shape','shaped','shapes','shaping','shards','shaved','shaving','shavings','shears','sheet','sheets','shipping','shop','shopping','shops','shortcut','shred','shreds','shucked','side','sides','sieve','sift','sifter','sifting','silpat','silver','similar','simmer','simmering','simple','simply','single','six','sixteen','sixths','size','sized','sizes','skewer','skewers','skillet','skillets','skim','skin','skinned','skins','skirt','slather','sleeve','slender','slicer','slicing','slightly','slim','slit','slivered','slivers','slotted','slow','slowly','smaller','smashed','snack','snacks','snapped','snow','soak','soaked','soaking','soft','soften','softer','softly','sold','solid','solids','something','sometimes','somewhat','sorted','sources','spare','sparkling','spatula','spatulas','spear','spears','special','specialty','spiced','spices','spicy','spider','spiked','splash','sponge','spongy','spoon','spooned','spoons','spots','spotted','spray','sprig','sprigs','spring','springform','springs','squeeze','st','stacked','stainless','stale','stand','standing','starchy','starter','states','steamer','steaming','steel','steen','stem','stemmed','step','sticky','stiff','stiffly','still','stir','stirred','stirring','stone','stop','storage','store','storebought','stores','stovetop','strained','strainer','strands','straw','streaky','streaming','street','strength','string','stringed','strings','strip','striped','stripped','strong','stronger','stuck','stuffed','stuffing','sturdy','style','sub','submerged','substitute','substituted','substitutes','sum','super','superfine','superior','supermarket','supermarkets','superpremium','supply','sure','surface','swanson','sweep','sweetener','sweetness','table','tablepoons','tablet','tall','tart','taste','tasting','tbsp','tear','teardrop','teaspoonfenugreek','teaspooon','tel','temperature','ten','tender','tends','tepid','terra','terrine','terry','test','tester','texture','textured','thai','thaw','thermometer','thickest','thickly','thickness','think','thinned','thinning','third','thirds','thirty','thoroughly','thousand','thread','threads','three','tie','tied','tight','tightly','time','times','tin','tins','tiny','tip','tissue','toaster','toasting','together','ton','tongs','toothpicks','toothpicks','top','topping','toppings','tops','torch','torn','toss','tossed','tossing','total','tough','towel','towels','traditional','tradizionale','trans','transfer','transparent','tray','treated','tree','tri','triangles','triangular','trimmings','triple','tropical','true','trumpet','try','tub','tube','tubes','tubular','turning','tweezers','twelve','twenty','twice','twine','two','tying','type','ultra','ultrapasteurized','unavailable','unblanched','uncle','unconverted','uncured','uncut','underripe','unfiltered','unflavored','unglazed','ungreased','unhulled','unhusked','uniform','uniformly','united','unless','unpitted','unrefined','unrinsed','unripe','unrolled','unscented','unseasoned','unshelled','unskinned','unsliced','unsmoked','unsprayed','unsulfured','unsweetened','unthawed','untoasted','untreated','untrimmed','unwaxed','unwrapped','upper','use','used','using','usually','uwajimaya','vacuum','valrhona','varieties','variety','various','vary','version','vertically','video','visible','vitamin','warmed','wash','washed','watch','watercolor','watery','waxed','waxy','way','wear','weave','weighed','weighing','weight','weighting','weights','well','west','westphalian','wet','whatever','wheel','whip','whipped','whipping','whisk','whisked','wide','width','wild','william','wilted','wilton','wineglass','winter','wipe','wiped','wire','without','wok','wood','wooden','woody','work','works','would','wrapped','wrappers','wrapping','www'
'year', 'yield',
'zester'
]))
brands = list(set(['rotel', 'absolut', 'betty crocker', 'jello', 'diana', 'ener-g',
'del-monte', "hunt's", 'martha', 'goya', 'cracker barrel',
'hamburger helper', "mccormick's", 'pepperidge farm', 'knorr',
'godiva', 'hidden valley', 'tabasco', 'branston', "kellogg's",
'hodgson mill', 'kraft', 'johnsonville', 'jim beam', 'mccormick',
'equal', 'jell-o', 'jimmy dean', 'country bob', "smucker's",
'toblerone', 'gerber', 'nestle', 'nestl', 'malt-o-meal', 'triscuit',
'ragu', 'campbell', 'hormel', 'earth balance', 'pillsbury',
"bird's eye", "campbell's", "betty crocker's", 'gold medal',
'crystal light', 'milnot', "land o' lakes", 'herb-ox', 'quaker',
'coffee-mate', 'contadina', 'j&d', 'fantastic foods', 'bacardi',
'eckrich', 'little smokies', 'snickers', 'ortega', 'bayou blast',
"annie's", 'mrs. dash', 'mori-nu', 'old el paso', 'original supreme',
'morton', 'nabisco', 'rice-a-roni', 'stolichnaya', "lawry's",
'st. germain', "eggland's best", 'club house "lagrille"', 'hostess',
'giada de laurentiis genovese', '*available in most target stores',
'jarlsberg', 'pillsbury plus', 'ro-tel', 'pillsbury grands',
'shilling', 'hershey', 'hershey carb alternatives', 'pasta roni',
'pastaroni', 'torani', 'v8', 'v8 fusion', 'ghiradelli', 'oscar mayer',
"bird's", 'smithfield', 'cadbury', 'sun-maid', 'karo',
'wishbone deluxe', 'vochelle', 'laughing cow', 'omega', 'stirrings',
'duncan hines', 'barilla', 'carnation', 'bertolli', 'bertolli®',
'breyers®', "1/2", "14.5", "''", "'s", "best®", "bell®", "bragg",
"crystal®", "crock®", "foods®", "frank's®", "gold®", "hellmann's®",
"johnsonville®", "knorr®", "lipton®", "n't", "n", "pillsbury™",
"progresso™", "progresso", "ranch®", "secrets®", "valley®", "vay®",
"yoplait®", "®", "’", 'boboli', 'bobolis', 'classico'
]))
measures = list(set(['cup', 'cups', 'pound', 'pounds', 'teaspoon', 'tsp', 'teaspoons',
'tablespoon', 'tablespoons', 'tbs', 'ounce', 'ounces', 'oz', 'see',
'note', 'lb', 'lbs', '¼', '½', '⅓', '⅔']))
all_list = list(set(cooking_stop_words + unhelpful + brands + measures))
with open("../write_data/food_stopwords.csv", "w") as myfile:
wr = csv.writer(myfile, quoting=csv.QUOTE_ALL)
wr.writerow(all_list)
```
| github_jupyter |
## Example of using the Google API Client to access BigQuery
Note that this is <b>not</b> the recommended approach. You should use the BigQuery client library because that is idiomatic Python.
See [the bigquery_client notebook](bigquery_client.ipynb) for examples.
### Authenticate and build stubs
```
PROJECT='cloud-training-demos' # CHANGE THIS
from googleapiclient.discovery import build
service = build('bigquery', 'v2')
```
### Get info about a dataset
```
# information about the ch04 dataset
dsinfo = service.datasets().get(datasetId="ch04", projectId=PROJECT).execute()
for info in dsinfo.items():
print(info)
```
### List tables and creation times
```
# list tables in dataset
tables = service.tables().list(datasetId="ch04", projectId=PROJECT).execute()
for t in tables['tables']:
print(t['tableReference']['tableId'] + ' was created at ' + t['creationTime'])
```
### Query and get result
```
# send a query request
request={
"useLegacySql": False,
"query": "SELECT start_station_name , AVG(duration) as duration , COUNT(duration) as num_trips FROM `bigquery-public-data`.london_bicycles.cycle_hire GROUP BY start_station_name ORDER BY num_trips DESC LIMIT 5"
}
print(request)
response = service.jobs().query(projectId=PROJECT, body=request).execute()
print('----' * 10)
for r in response['rows']:
print(r['f'][0]['v'])
```
### Asynchronous query and paging through results
```
# send a query request that will not terminate within the timeout specified and will require paging
request={
"useLegacySql": False,
"timeoutMs": 0,
"useQueryCache": False,
"query": "SELECT start_station_name , AVG(duration) as duration , COUNT(duration) as num_trips FROM `bigquery-public-data`.london_bicycles.cycle_hire GROUP BY start_station_name ORDER BY num_trips DESC LIMIT 5"
}
response = service.jobs().query(projectId=PROJECT, body=request).execute()
print(response)
jobId = response['jobReference']['jobId']
print(jobId)
# get query results
while (not response['jobComplete']):
response = service.jobs().getQueryResults(projectId=PROJECT,
jobId=jobId,
maxResults=2,
timeoutMs=5).execute()
while (True):
# print responses
for row in response['rows']:
print(row['f'][0]['v']) # station name
print('--' * 5)
# page through responses
if 'pageToken' in response:
pageToken = response['pageToken']
# get next page
response = service.jobs().getQueryResults(projectId=PROJECT,
jobId=jobId,
maxResults=2,
pageToken=pageToken,
timeoutMs=5).execute()
else:
break
```
Copyright 2019 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| github_jupyter |
# User comparison tests
# Table of Contents
[Preparation](#preparation)
[User data vectors](#userdatavectors)
[User lists](#userlists)
[Sessions' checkpoints](#sessionscheckpoints)
[Assembly](#assembly)
[Time](#time)
# Preparation
<a id=preparation />
```
%run "../Functions/1. Google form analysis.ipynb"
%run "../Functions/4. User comparison.ipynb"
```
# Data vectors of users
<a id=userdatavectors />
```
#getAllResponders()
setAnswerTemporalities(gform)
```
# getAllUserVectorData
```
# small sample
#allData = getAllUserVectorData( getAllUsers( rmdf152 )[:10] )
# complete set
#allData = getAllUserVectorData( getAllUsers( rmdf152 ) )
# subjects who answered the gform
allData = getAllUserVectorData( getAllResponders() )
# 10 subjects who answered the gform
#allData = getAllUserVectorData( getAllResponders()[:10] )
efficiencies = allData.loc['efficiency'].sort_values()
efficiencies.index = range(0, len(allData.columns))
efficiencies.plot(title = 'efficiency')
efficiencies2 = allData.loc['efficiency'].sort_values()
efficiencies2 = efficiencies2[efficiencies2 != 0]
efficiencies2.index = range(0, len(efficiencies2))
efficiencies2 = np.log(efficiencies2)
efficiencies2.plot(title = 'efficiency log')
maxChapter = allData.loc['maxChapter'].sort_values()
maxChapter.index = range(0, len(allData.columns))
maxChapter.plot(title = 'maxChapter')
len(allData.columns)
userIds = getAllResponders()
_source = correctAnswers
# _source is used as correction source, if we want to include answers to these questions
#def getAllUserVectorData( userIds, _source = [] ):
# result
isInitialized = False
allData = []
f = FloatProgress(min=0, max=len(userIds))
display(f)
for userId in userIds:
#print(str(userId))
f.value += 1
if not isInitialized:
isInitialized = True
allData = getUserDataVector(userId, _source = _source)
else:
allData = pd.concat([allData, getUserDataVector(userId, _source = _source)], axis=1)
#print('done')
allData
userId
```
# Correlation Matrix
```
methods = ['pearson', 'kendall', 'spearman']
_allUserVectorData = allData.T
_method = methods[0]
_title='RedMetrics Correlations'
_abs=True
_clustered=False
_figsize = (20,20)
#def plotAllUserVectorDataCorrelationMatrix(
# _allUserVectorData,
# _method = methods[0],
# _title='RedMetrics Correlations',
# _abs=False,
# _clustered=False,
# _figsize = (20,20)
#):
_progress = FloatProgress(min=0, max=3)
display(_progress)
# computation of correlation matrix
_m = _method
if(not (_method in methods)):
_m = methods[0]
_correlation = _allUserVectorData.astype(float).corr(_m)
_progress.value += 1
if(_abs):
_correlation = _correlation.abs()
_progress.value += 1
# plot
if(_clustered):
sns.clustermap(_correlation,cmap=plt.cm.jet,square=True,figsize=_figsize)
else:
_fig = plt.figure(figsize=_figsize)
_ax = plt.subplot(111)
_ax.set_title(_title)
sns.heatmap(_correlation,ax=_ax,cmap=plt.cm.jet,square=True)
_progress.value += 1
gform['Temporality'].unique()
allData.loc['scoreundefined'].dropna()
getAllUsers(rmdf152)[:10]
len(getAllUsers(rmdf152))
```
# List of users and their sessions
<a id=userlists />
```
userSessionsRelevantColumns = ['customData.localplayerguid', 'sessionId']
userSessions = rmdf152[rmdf152['type']=='start'].loc[:,userSessionsRelevantColumns]
userSessions = userSessions.rename(index=str, columns={'customData.localplayerguid': 'userId'})
userSessions.head()
#groupedUserSessions = userSessions.groupby('customData.localplayerguid')
#groupedUserSessions.head()
#groupedUserSessions.describe().head()
```
# List of sessions with their checkpoints achievements
<a id=sessionscheckpoints />
```
checkpointsRelevantColumns = ['sessionId', 'customData.localplayerguid', 'type', 'section', 'userTime']
checkpoints = rmdf152.loc[:, checkpointsRelevantColumns]
checkpoints = checkpoints[checkpoints['type']=='reach'].loc[:,['section','sessionId','userTime']]
checkpoints = checkpoints[checkpoints['section'].str.startswith('tutorial', na=False)]
#checkpoints = checkpoints.groupby("sessionId")
#checkpoints = checkpoints.max()
checkpoints.head()
```
# Assembly of both
<a id=assembly />
```
#assembled = userSessions.combine_first(checkpoints)
assembled = pd.merge(userSessions, checkpoints, on='sessionId', how='outer')
assembled.head()
userSections = assembled.drop('sessionId', 1)
userSections.head()
userSections = userSections.dropna()
userSections.head()
checkpoints = userSections.groupby("userId")
checkpoints = checkpoints.max()
checkpoints.head()
```
# Time analysis
<a id=time />
```
#userTimedSections = userSections.groupby("userId").agg({ "userTime": np.min })
#userTimedSections = userSections.groupby("userId")
userTimes = userSections.groupby("userId").agg({ "userTime": [np.min, np.max] })
userTimes["duration"] = pd.to_datetime(userTimes["userTime"]["amax"]) - pd.to_datetime(userTimes["userTime"]["amin"])
userTimes["duration"] = userTimes["duration"].map(lambda x: np.timedelta64(x, 's'))
userTimes = userTimes.sort_values(by=['duration'], ascending=[False])
userTimes.head()
```
# TODO
userTimes.loc[:,'duration']
userTimes = userTimes[4:]
userTimes["duration_seconds"] = userTimes["duration"].map(lambda x: pd.Timedelta(x).seconds)
maxDuration = np.max(userTimes["duration_seconds"])
userTimes["duration_rank"] = userTimes["duration_seconds"].rank(ascending=False)
userTimes.plot(x="duration_rank", y="duration_seconds")
plt.xlabel("game session")
plt.ylabel("time played (s)")
plt.legend('')
plt.xlim(0, 139)
plt.ylim(0, maxDuration)
userTimedSections = userSections.groupby("section").agg({ "userTime": np.min })
userTimedSections
userTimedSections["firstReached"] = pd.to_datetime(userTimedSections["userTime"])
userTimedSections.head()
userTimedSections.drop('userTime', 1)
userTimedSections.head()
userTimedSections["firstCompletionDuration"] = userTimedSections["firstReached"].diff()
userTimedSections.head()
```
sessionCount = 1
_rmDF = rmdf152
sample = gform
before = False
after = True
gfMode = False
rmMode = True
#def getAllUserVectorDataCustom(before, after, gfMode = False, rmMode = True, sessionCount = 1, _rmDF = rmdf152)
userIds = []
if (before and after):
userIds = getSurveysOfUsersWhoAnsweredBoth(sample, gfMode = gfMode, rmMode = rmMode)
elif before:
if rmMode:
userIds = getRMBefores(sample)
else:
userIds = getGFBefores(sample)
elif after:
if rmMode:
userIds = getRMAfters(sample)
else:
userIds = getGFormAfters(sample)
if(len(userIds) > 0):
userIds = userIds[localplayerguidkey]
allUserVectorData = getAllUserVectorData(userIds, _rmDF = _rmDF)
allUserVectorData = allUserVectorData.T
result = allUserVectorData[allUserVectorData['sessionsCount'] == sessionCount].T
else:
print("no matching user")
result = []
result
getAllUserVectorDataCustom(False, True)
userIdsBoth = getSurveysOfUsersWhoAnsweredBoth(gform, gfMode = True, rmMode = True)[localplayerguidkey]
allUserVectorData = getAllUserVectorData(userIdsBoth)
allUserVectorData = allUserVectorData.T
allUserVectorData[allUserVectorData['sessionsCount'] == 1]
```
#### user progress classification
##### tinkering
```
testUser = "3685a015-fa97-4457-ad73-da1c50210fe1"
def getScoreFromBinarized(binarizedAnswers):
gformIndices = binarizedAnswers.index.map(lambda s: int(s.split(correctionsColumnNameStem)[1]))
return pd.Series(np.dot(binarizedAnswers, np.ones(binarizedAnswers.shape[1])), index=gform.loc[gformIndices, localplayerguidkey])
#allResponders = getAllResponders()
#gf_both = getSurveysOfUsersWhoAnsweredBoth(gform, gfMode = True, rmMode = False)
rm_both = getSurveysOfUsersWhoAnsweredBoth(gform, gfMode = False, rmMode = True)
#gfrm_both = getSurveysOfUsersWhoAnsweredBoth(gform, gfMode = True, rmMode = True)
sciBinarizedBefore = getAllBinarized(_form = getRMBefores(rm_both))
sciBinarizedAfter = getAllBinarized(_form = getRMAfters(rm_both))
scoresBefore = getScoreFromBinarized(sciBinarizedBefore)
scoresAfter = getScoreFromBinarized(sciBinarizedAfter)
medianBefore = np.median(scoresBefore)
medianAfter = np.median(scoresAfter)
maxScore = sciBinarizedBefore.shape[1]
indicators = pd.DataFrame()
indicators['before'] = scoresBefore
indicators['after'] = scoresAfter
indicators['delta'] = scoresAfter - scoresBefore
indicators['maxPotentialDelta'] = maxScore - scoresBefore
for index in indicators['maxPotentialDelta'].index:
if (indicators.loc[index, 'maxPotentialDelta'] == 0):
indicators.loc[index, 'maxPotentialDelta'] = 1
indicators['relativeBefore'] = scoresBefore / medianBefore
indicators['relativeAfter'] = scoresAfter / medianBefore
indicators['relativeDelta'] = indicators['delta'] / medianBefore
indicators['realizedPotential'] = indicators['delta'] / indicators['maxPotentialDelta']
indicators['increaseRatio'] = indicators['before']
for index in indicators['increaseRatio'].index:
if (indicators.loc[index, 'increaseRatio'] == 0):
indicators.loc[index, 'increaseRatio'] = 1
indicators['increaseRatio'] = indicators['delta'] / indicators['increaseRatio']
indicators
(min(indicators['relativeBefore']), max(indicators['relativeBefore'])),\
(min(indicators['relativeDelta']), max(indicators['relativeDelta'])),\
medianBefore,\
np.median(indicators['relativeBefore']),\
np.median(indicators['relativeDelta'])\
indicatorX = 'relativeBefore'
indicatorY = 'relativeDelta'
def scatterPlotIndicators(indicatorX, indicatorY):
print(indicatorX + ' range: ' + str((min(indicators[indicatorX]), max(indicators[indicatorX]))))
print(indicatorY + ' range: ' + str((min(indicators[indicatorY]), max(indicators[indicatorY]))))
print(indicatorX + ' median: ' + str(np.median(indicators[indicatorX])))
print(indicatorY + ' median: ' + str(np.median(indicators[indicatorY])))
fig = plt.figure()
ax1 = fig.add_subplot(111)
ax1.scatter(indicators[indicatorX], indicators[indicatorY])
plt.xlabel(indicatorX)
plt.ylabel(indicatorY)
# vertical line
plt.plot( [np.median(indicators[indicatorX]), np.median(indicators[indicatorX])],\
[min(indicators[indicatorY]), max(indicators[indicatorY])],\
'k-', lw=2)
# horizontal line
plt.plot( [min(indicators[indicatorX]), max(indicators[indicatorX])],\
[np.median(indicators[indicatorY]), np.median(indicators[indicatorY])],\
'k-', lw=2)
indicators.columns
scatterPlotIndicators('relativeBefore', 'relativeDelta')
scatterPlotIndicators('relativeBefore', 'realizedPotential')
scatterPlotIndicators('relativeBefore', 'increaseRatio')
scatterPlotIndicators('relativeBefore', 'relativeAfter')
scatterPlotIndicators('maxPotentialDelta', 'realizedPotential')
```
| github_jupyter |

<a href="https://hub.callysto.ca/jupyter/hub/user-redirect/git-pull?repo=https%3A%2F%2Fgithub.com%2Fcallysto%2Fcurriculum-notebooks&branch=master&subPath=Mathematics/BinaryDecimalConversion/binary-decimal-conversion.ipynb&depth=1" target="_parent"><img src="https://raw.githubusercontent.com/callysto/curriculum-notebooks/master/open-in-callysto-button.svg?sanitize=true" width="123" height="24" alt="Open in Callysto"/></a>
```
%%html
<script>
function code_toggle() {
if (code_shown){
$('div.input').hide('500');
$('#toggleButton').val('Show Code')
} else {
$('div.input').show('500');
$('#toggleButton').val('Hide Code')
}
code_shown = !code_shown
}
$( document ).ready(function(){
code_shown=false;
$('div.input').hide()
});
</script>
<form action="javascript:code_toggle()"><input type="submit" id="toggleButton" value="Show Code"></form>
#!pip install --upgrade --force-reinstall --user git+git://github.com/callysto/nbplus.git#egg=geogebra\&subdirectory=geogebra
try:
from geogebra.ggb import *
except ImportError:
!pip install --upgrade --force-reinstall --user git+git://github.com/callysto/nbplus.git#egg=geogebra\&subdirectory=geogebra
from importlib import reload
import site
reload(site)
from geogebra.ggb import *
ggb = GGB()
from __future__ import print_function
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
import IPython
from IPython.display import HTML,display, Math, Latex, clear_output
import math
import random
import numpy as np
import plotly.offline as py
from plotly.offline import init_notebook_mode, iplot
import plotly.graph_objs as go
```
# Binary representation of numbers
## Introduction
A *positional numeral system* is a way of writing numbers using place value.<br>
Our usual way of writing numbers is an example:<br>
In a number like 1234, the 4 is in the 'ones' spot, the 3 in the 'tens' spot, and so on.<br>
Each position represents a power of 10, so we call this the *decimal*, or base-10 system.
This notebook will look at positional numeral systems in general, but a focus on two familiar examples:<br>
The decimal (base-10) system we all know and love, and the binary (base-2) system.
Topics include:
- the *base* of the numeral system
- how the choice of base effects the *digits* we use to write a number
- how to convert between decimal and binary representations of a number.
<img src="images/binary.png" alt="drawing" style="width:500px;"/>
### Big Ideas
Here are some key takeaways you should have in mind as you progress through this notebook:
* Computers understand numbers in binary, while humans understand decimal.
* Conversion from decimal to binary is fundamental to human-computer interaction.
* Understanding this process requires only basic arithmetic: addition, subtraction, and powers.
## Background information
### Numeral systems
In mathematical terminology, a **_number system_** refers to the type of number being used.<br>
Number systems include the integers, the rational numbers (fractions), and real numbers.<br>
A [**_numeral system_**](https://en.wikipedia.org/wiki/Numeral_system) refers to the way we write those numbers down.
The numeral system we use when we write a number like 3127 is called the *decimal system*.<br>
You've gotten so used to using the decimal system that you understand the number 3127 without thinking!<br>
Take a moment to remind yourself that the *position* of each digit represents a value: 1, 10, 100, etc.<br>
The digit in that position then tells us how many of that quantity we have.<br>
In 3127, there are 3 thousands, one hundred, two tens, and seven ones.
But our method of expressing numbers is by no means universal.<br>
Another numeral system you've probably encountered is that of Roman numerals.<br>
The Roman system uses symbols instead of position to represent certain quantities.<br>
We count how many times each symbol appears to know how many of that quantity we have.
For example, the Roman numeral system uses I for 1, V for 5, X for 10, C for 100, and D for 1000, among others.<br>
In the Roman system, the number 3127 is written as MMMCXXVII:
$$\begin{aligned}
3127&=3000+100+20+7\\
& = \underbrace{1000+1000+1000}_{\text{M M M}}+\underbrace{100}_{\text{C}}+\underbrace{10+10}_{\text{X X}}+\underbrace{5}_{\text{V}}+\underbrace{1+1}_{\text{I I}}.
\end{aligned}$$
Note that in the Roman system, there is no symbol for zero: a number like 210 has no ones, so we don't write any.<br>
The number 210 is written CCX. Many early numeral systems did not have a zero. <br>
The invention (discovery?) of zero is usually credited to India, around the 4th or 5th century.<br>
However, zero was also used in [Mayan calendars](https://en.wikipedia.org/wiki/0#History).<br>
The number zero is an interesting topic in its own right, but it's not the subject of this notebook.<br>
If you're curious, you might enjoy this [Scientific American article](https://www.scientificamerican.com/article/history-of-zero/).
Binary and decimal numbers are two examples of *positional numeral systems*, where the relative placement of digits tells us about the values they represent.<br>
One of the earliest examples of a positional system is the remarkable Babylonian [sexagesimal (base-60) system](https://en.wikipedia.org/wiki/Babylonian_numerals).
Our numeral system traces its origins back to India and the Middle East, and is known as the [Hindu-Arabic numeral system](https://en.wikipedia.org/wiki/Hindu%E2%80%93Arabic_numeral_system).<br>
This is a decimal system, where positions represent powers of 10, and symbols represent numbers from 0 to 9.<br>
For example, as we mentioned above, the expression 3127 is convenient shorthand for
$$3\times 1000 + 1\times 1000 + 2\times 10 + 7\times 1.$$
Notice that 1000, 100, 10, and 1 are all powers of 10: $1=10^0, 10=10^1, 100=10^2$, and $1000=10^3$.<br>
When each position represents successive powers of some number, we call that number the *base* of the system.<br>
The decimal system uses base 10. The binary system, which is essential to computing, uses base 2.
### Why Binary?
Why do computers use binary? First, we might ask: why do we use decimal?<br>
We’ve been using decimal so long, you may not have wondered why we chose the base-10 number system for our everyday number needs.<br>
(There's a pretty good chance that it has something to do with humans having 10 fingers.)
Regardless of what led to it, tricks we’ve learned along the way have solidified base-10’s place in our heart.<br>
Everyone can count by 10s. We even round large numbers to the nearest multiple of 10. We’re obsessed with 10!
Computers, being electronic machines, only understand two things:<br>
'ON' or '1': electricity is flowing (through circuit, gate, transistor, whatever).<br>
'OFF' or '0': electricity is not flowing.
In a computer, everything must be represented as collections of ONs and OFFs.
Unfortunately, this scheme is difficult for humans to process.<br>
To make things easier on ourselves, humans represent "computerian" as binary numbers:
ON is represented by a 1, and OFF is represented by a 0.<br>
So instead of "OFF ON ON OFF", we humans can read or write this as "0110".
|
### Base and Digits in a Positional System
The **base** in a positional numeral system like Hindu-Arabic is the number whose powers we represent by the position of the digits in a number.<br>
Note that the value of the base is also the number of different symbols we need for digits.<br>
For example, the decimal (base-10) system uses 10 digits: 0, 1, 2, 3, 4, 5, 6, 7, 8, and 9.<br>
In some computer applications (like many WiFi passwords) a hexadecimal (base-16) system is used, with digits
> 0, 1, 2, 3, 4, 5, 6, 7, 8, 9, a, b, c, d, e, f.
Of course, we also use "base" to refer to the number that gets multiplied when applying an exponent.
Examples:
* In $8^{2}$, 8 is the base, and the result is $8 × 8 = 64$.
* In $5^{3}$, 5 is the base, and the result is $5 × 5 × 5 = 125$.
This isn't really a coincidence, since we use powers of the base to reconstruct numbers from positional notation.<br>
The way our notation works is as follows:
- Position of a digit (read from right to left) tells us which power of the base to use.
- The digit in that position tells us what to multiply that power by.
Any integer greater than 1 can be used as a base. (Why can't we use 1?)<br>
For a hypothetical "base-$r$" system, we would have $r$ symbols to represent the digits, and each position represents a power of $r$.<br>
For example, we could imagine a base-26 system with each letter of the alphabet representing a digit.<br>
(It's doubtful anyone would want to *use* this system, but we can imagine it!)<br>
In this system, the word "number" would actually *be* a number!<br>
If we assign each letter its place value in the alphabet, we have
$$
n = 14, u = 21, m = 13, b = 2, e = 5, r = 18,
$$
and

This doesn't seem very practical, however, so let's sum up the key points and move on to numbers in decimal and binary.
* The base of a positional numeral system is equal to the number of digits (symbols) needed to represent numbers in that system.
* The position of a digit tells us what the corresponding power of the base should be.
* We multiply that power by the value of the digit, for each digit present.
* Finally, we add everything up to get our number.
For a number $a_na_{n-1}\cdots a_2a_1a_0$ with digits $a_0, a_1, \ldots, a_n$ written in base $r$, we get the number
$$a_nr^n+a_{n-1}r^{n-1}+\cdots a_2r^2+a_1r+a_0.$$
We won't do it here, but see if you can convince yourself any number can be written *uniquely* in any base. We never miss a number, and we never have two different ways of writing down the same number.
While working with numbers of different bases, its best to put the base as a subscript to the number to avoid confusion like $(Number)_{Base}$. For example in order to represent a binary number we place 2 as a subscript like $(11)_{2}$ and 10 to denote decimal numbers like $(11)_{10}$.
#### An answer to an impractical problem.
In the example above, were you left wondering what (base-10) number is "number" in (base-26)?<br>
We can ask the computer to calculate it for us:
```
14*26**5 + 21*26**4 + 13*26**3 + 2*26**2 + 5*26 + 18
```
### Counting in binary
We can count in decimal endlessly, even in our sleep, but how can we count in binary?<br>
Zero and one in base-two should look pretty familiar: 0 and 1. From there things get decidedly binary.
In decimal, we start in the 'ones' position, and count until we run out of digits:
$$0, 1, 2, 3, 4, 5, 6, 7, 8, 9.$$
After 9, we hit the next power of 10, so we add 1 to the 'tens' position, and reset the'ones' to 0.<br>
Then we go back to counting in the 'ones' position until we reach 9 again:
$$10, 11, 12, 13, 14, 15, 16, 17, 18, 19.$$
Now, we increase the 'tens' digit by 1 again, and reset the 'ones', getting 20.<br>
We continue like this until we reach 99, and then we're ready to add a 1 in the 'hundreds' position.<br>
When we do, the digits in both the 'tens' and 'ones' positions reset to zero, and then we continue the process.
Counting in binary is similar, with two important differences:
1. Positions represent powers of 2, not 10: $ 2^0=1, 2^1 = 2, 2^2 = 4, 2^3 = 8, 2^4 = 16, 2^5 = 32, 2^6=64$, etc.
2. Digits are only 0 and 1, so the 'change position and reset to zero' step is a lot more frequent!
Remember that we’ve only got those two digits, so as we do in decimal, when we run out of symbols we shift one column to the left, add a 1, and turn all of the digits to the right of that column to 0. <br>
So after 1 we get 10, then 11, then 100. Let’s start counting…
| Decimal | Binary |
|:-------:|:------:|
| 0 | 0000 |
| 1 | 0001 |
| 2 | 0010 |
| 3 | 0011 |
| 4 | 0100 |
| 5 | 0101 |
| 6 | 0110 |
| 7 | 0111 |
| 8 | 1000 |
| 9 | 1001 |
| 10 | 1010 |
| 11 | 1011 |
| 12 | 1100 |
| 13 | 1101 |
| 14 | 1110 |
| 15 | 1111 |
## Converting between decimal and binary notation.
### Converting binary to decimal
Mathematically, this is straightforward. The digits in binary are called **_bits_**.<br>
Each position represents a power of 2. We multiply each power of 2 by the bit in that position, and then add it all up.
#### Example:
To convert the number 1101 from binary to decimal, we compute as follows:
$$\begin{aligned}
1101 &= 1 * 2^{3} + 1 * 2^{2} + 0 * 2^{1} + 1 * 2^{0} \\
&= 1*8+1*4+0*2+1*1\\
&= 8+4+1\\&=13
\end{aligned}$$
### Interactive Method
We will now demonstrate an interactive approach to doing this conversion.<br>
Our implementation uses buttons on a computer screen, but you can do this "in real life" as well!<br>
For demonstration we start off with a 5 bit number; that is, a 5 digit binary number, which can represent up to 31 in decimal.
1. Represent each binary digit with a single card, so there will be 5 cards.
2. Mark one side of each card with a power of 2: our five cards will be 1, 2, 4, 8, and 16.<br>
Mark the other side with a 0.
3. Arrange the cards to reflect our positional system: Our order is 16, 8, 4, 2, 1.
4. The side with the power of 2 represents 1 in binary and the other side represents 0.
5. Start with the 0 on each card. Choose a binary number, then flip the cards whose digit is 1.<br>
For example, if we chose the number 11001, we flip the 16, the 8, and the 1.
6. Now to convert to decimal we just add up the numbers showing in the cards!<br>
For 11001 we have $16+8+0+0+1=25$, so 11001 in binary is equivalent to 25 in decimal.
#### Applet Instructions:
1. Create a number in binary: clicking on a button with a 0 turns it into a 1.
2. The corresponding power of 2 below will change colour, and the 0 in front will change to a 1.
3. The decimal equivalent will appear at the bottom.
```
%%HTML
<!DOCTYPE html>
<html>
<head>
<meta name="viewport" content="width=device-width, initial-scale=1">
<style>
.button {
padding: 15px 25px;
font-size: 24px;
text-align: center;
cursor: pointer;
outline: none;
color: #fff;
background-color: #4CAF50;
border: none;
border-radius: 15px;
box-shadow: 0 9px #999;
}
.button:hover {background-color: #3e8e41}
.disabled {
opacity: 0.6;
cursor: not-allowed;
}
.button:active {
background-color: #3e8e41;
box-shadow: 0 5px #666;
transform: translateY(4px);
}
.txtbox{
display: block;
float: left;
height: 50px;
width: 50px;
font-size: 24px;
}
.new-meows{
padding: 10px;
border: 1px solid grey;
margin : 10px;
width: 97%;
display:flex;
}
.divider{
width:10px;
height:auto;
display:inline-block;
}
</style>
</head>
<body>
<h3>Enter a binary number</h3><br>
<button id="card1" onclick="myFunction(this.id);myFunction1(this.id, 'Decimal_card_16', 'box1');" class= "button">0</button>
<button id="card2" onclick="myFunction(this.id);myFunction1(this.id,'Decimal_card_8', 'box2' );" class= "button">0</button>
<button id="card3" onclick="myFunction(this.id);myFunction1(this.id,'Decimal_card_4', 'box3');" class= "button">0</button>
<button id="card4" onclick="myFunction(this.id);myFunction1(this.id,'Decimal_card_2', 'box4');" class= "button">0</button>
<button id="card5" onclick="myFunction(this.id);myFunction1(this.id,'Decimal_card_1', 'box5');" class= "button">0</button>
<br>
<br>
<h3>Decimal Equivalent of the given binary number</h3>
<div class = "new-meows">
<input type="text" id = "box1" value="0" class ="txtbox" /> <font size="24" style= vertical-align: middle;> × </font>
<div class="divider"/>
<button id="Decimal_card_16" class= "button ">16</button><font size="24"> + </font>
<div class="divider"/>
<input type="text" id = "box2" value="0" class ="txtbox" /> <font size="24"> × </font>
<div class="divider"/>
<button id="Decimal_card_8" class= "button ">8</button> <font size="24">+</font>
<div class="divider"/>
<input type="text" id = "box3" value="0" class ="txtbox" /> <font size="24"> × </font>
<div class="divider"/>
<button id="Decimal_card_4" class= "button ">4</button><font size="24">+</font>
<div class="divider"/>
<input type="text" id = "box4" value="0" class ="txtbox" /> <font size="24"> × </font>
<div class="divider"/>
<button id="Decimal_card_2" class= "button ">2</button><font size="24">+</font>
<div class="divider"/>
<input type="text" id = "box5" value="0" class ="txtbox" /> <font size="24"> × </font>
<div class="divider"/>
<button id="Decimal_card_1" class= "button ">1</button><font size="24"> =</font>
<div class="divider"/>
<input type="text" id = "box6" value="" class ="txtbox" />
</div>
<script>
function myFunction(clicked_id) {
var Button_id = document.getElementById(clicked_id);
if (Button_id.innerHTML === "0") {
Button_id.innerHTML = "1";
} else {
Button_id.innerHTML = "0";
}
}
</script>
<script>
function setColor(btn, color_id) {
var Button_id = document.getElementById(btn);
if (color_id == 1) {
Button_id.style.backgroundColor = "#FF0000"
}
else {
Button_id.style.backgroundColor = "#4CAF50"
color_id = 0;
}
}
</script>
<script>
function setValue(box_id, txt_id) {
document.getElementById(box_id).value = "1";
if (txt_id == 1) {
document.getElementById(box_id).value = "1";
}
else {
document.getElementById(box_id).value = "0";
}
}
</script>
<script>
var value1, value2, value3, value4, value5;
var result = 0;
function myFunction1(clicked_id, decimal_card, txt_box_no) {
//value1 = 0;
//value2 = 0;
//value3 = 0;
//value4 = 0;
//value5 = 0;
var Button_id = document.getElementById(clicked_id);
if(clicked_id ==="card1" && Button_id.innerHTML ==="1"){
value1 = 16;
result = result + value1;
setColor(decimal_card, 1);
setValue(txt_box_no, 1);
}
else if(clicked_id ==="card1" && Button_id.innerHTML ==="0"){
result = result - 16;
value1 = 0;
setColor(decimal_card, 0);
setValue(txt_box_no, 0);
}
else if(clicked_id ==="card2" && Button_id.innerHTML ==="1"){
value2 = 8;
result = result + value2;
setColor(decimal_card, 1);
setValue(txt_box_no, 1);
}
else if(clicked_id ==="card2" && Button_id.innerHTML ==="0"){
result = result - 8;
value2 = 0;
setColor(decimal_card, 0);
setValue(txt_box_no, 0);
}
else if(clicked_id ==="card3" && Button_id.innerHTML ==="1"){
value3 = 4;
result = result + value3;
setColor(decimal_card, 1);
setValue(txt_box_no, 1);
}
else if(clicked_id ==="card3" && Button_id.innerHTML ==="0"){
value3 = 0;
result = result - 4;
setColor(decimal_card, 0);
setValue(txt_box_no, 0);
}
else if(clicked_id ==="card4" && Button_id.innerHTML ==="1"){
value4 = 2;
result = result + value4;
setColor(decimal_card, 1);
setValue(txt_box_no, 1);
}
else if(clicked_id ==="card4" && Button_id.innerHTML ==="0"){
value4 = 0;
result = result - 2;
setColor(decimal_card, 0);
setValue(txt_box_no, 0);
}
else if(clicked_id ==="card5" && Button_id.innerHTML ==="1"){
value5 = 1;
result = result + value5;
setColor(decimal_card, 1);
setValue(txt_box_no, 1);
}
else if(clicked_id ==="card5" && Button_id.innerHTML ==="0"){
value5 = 0;
result = result - 1;
setColor(decimal_card, 0);
setValue(txt_box_no, 0);
}
document.getElementById("Result_of_binary").innerHTML = result;
document.getElementById("box6").value = result;
}
</script>
<h3>The decimal equivalent is: <span id="Result_of_binary"></span></h3>
</body>
</html>
```
### A remark on notation
When we're working with more than one base system at the same time (for example, when converting from decimal to binary) we need a way of specifying what base we're using.
A common notation is to place the digits in parentheses, with the base as a subscript.<br>
For example, we can write $(10110)_2$ to indicate that we're expressing a number in binary.<br>
This lets us write equations, like $(10110)_2 = (22)_{10}$, to indicate that we've expressed the same number in two different bases.
Some people use notation like $\mathbf{Bin}\,10110 = \mathbf{Dec}\, 22$ instead.<br>
This notation is a little clumsier, but it can be used for a [great joke](https://armchairdissident.wordpress.com/2009/04/15/a-long-explanation-of-a-damned-fine-joke/).
### Converting Decimal to Binary
In the other direction, our goal is to flip the right cards to add up to a given number.<br>
Let's say we want the binary representation for 13.
We could simply employ trial and error, flipping cards and checking each time to see if we have it right.<br>
This isn't very efficient though. Instead, let's think like a computer scientist, and devise an algorithm.
1. First, we definitely don't want to add any numbers *greater* than our number. So 16 is out: we set that card to 0.
2. Next, we're going to want the biggest power of 2 that's *less* than our number. So 8 is in: we set that card to 1.
3. Now we ask ourselves, what do we have left to add? This is a subtraction problem! We want 13 and we have 8, so we still have $13-8=5$ left to add.
4. To continue, we take the value that's left -- 5 -- and repeat steps 2 and 3. Since 4 is less than 5, we flip that card to 1, and find the difference: $5-4=1$.
5. We keep repeating, until there's nothing left to add. But all we have left is 1, so we flip that card, and we're done!
**Note:** the steps we just described give an example of what's called a *recursive algorithm*.<br>
In a recursive algorithm, we repeat a series of steps until a desired goal is reached.<br>
Many computer programs employ recursive algorithms to accomplish their task.
### Visualizing the Algorithm
#### Instructions:
1. Enter a value for n between 1 and 31 in the input box.
2. A graph will be generated to show the original numbers as a blue dot.
3. The red dots will represent the powers of 2 we need to make the remainder 0.
4. Below the graph, the steps are also shown for further clarification. For Example if the input number is 25, then the steps will be
the first step : 25 - 16 = 9
the next step : 9 - 8 = 1
the final step : 1 - 1 = 0
From the steps we know that we need 16, 8, 1 as powers of 2 to make the remainder 0.
* So The binary for $(25)_{10}$ is $(11001)_{2}$.
```
py.init_notebook_mode(connected=True)
def plot_stored_values(store_plot_value, value):
value_list = []
value_list.append(value)
N = len(store_plot_value)
random_y = np.zeros(N)
# Create a trace
trace = go.Scatter(
x = store_plot_value,
y = random_y,
marker = dict(color = 'rgb(128, 0, 0)',),
mode = 'markers',
name = "Derived Numbers of power of 2"
)
trace_value = go.Scatter(
x = value_list,
y = [0],
marker = dict(color = 'rgb(0, 0, 128)',),
mode = 'markers',
name = "Original number"
)
data = [trace , trace_value]
layout = go.Layout(
title ='Decimal Value numbers',
xaxis =dict(
title='Decimal Values',
titlefont=dict(
family='Courier New, monospace',
size=18,
color='#7f7f7f'
)
),
yaxis=dict(
title='',
titlefont=dict(
family='Courier New, monospace',
size=18,
color='#7f7f7f'
)
)
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='basic-line')
#function for finding the power of 2
def power_two_1(n):
return int(math.log(n, 2))
def power_two(x):
return x.bit_length()-1
#function for calculating the algorithm value
store_plot_value = []
def Calculate_Value(x):
store_plot_value = []
power_of_2 = power_two(x)
#print(power_of_2)
first_value = 2 ** power_of_2
store_plot_value.append(first_value)
plot_value = x - 2 ** power_of_2
#store_plot_value.append(plot_value)
while(plot_value > 0):
power_of_2 = power_two(plot_value)
#print(power_of_2)
store_value = 2 ** power_of_2
store_plot_value.append(store_value)
plot_value = plot_value - 2 ** power_of_2
#plot_stored_values(store_plot_value)
return store_plot_value
Algorithm_value = widgets.BoundedIntText(
value = 0,
min = 0,
max = 256,
step = 1,
description = 'Enter value:',
disabled = False
)
def on_value_change(change):
clear_output(wait= False)
display(Algorithm_value)
Plot_values = []
#Calculate_Value(change['new'])
Plot_values = Calculate_Value(change['new'])
plot_stored_values(Plot_values, change['new'])
visual_value = change['new'] - Plot_values[0]
interim_value = visual_value
print("the first step : {} - {} = {}".format(change['new'],Plot_values[0], interim_value ))
for i in Plot_values[1:]:
temp = interim_value
interim_value = interim_value - i
print("the next step : {} - {} = {}".format(temp, i , interim_value ))
print("From the steps we see that we need", Plot_values, " as powers of 2 to make the remainder 0.")
display(Algorithm_value)
Algorithm_value.observe(on_value_change, names='value')
```
### Try doing the algorithm yourself!
#### Instructions:
1. Enter a value for n between 1 and 31 in the input box.
2. When you click on a power of 2, it will be subtracted from your number.
3. Keep working until the amount r that remains is zero.
4. Your number in binary will be shown at the bottom.
```
ggb.material('VJJpYMQ2').draw()
```
## Practice Questions
Now that you've seen a few examples, it's time to try some on your own.<br>
But remember: two problems worth of practice is not enough to master this skill.<br>
Make sure you take the time to work through additional problems from your teacher or textbook.
### Convert the following Decimal Numbers to binary
##### Question 1
$(57)_{10} = (A)_2$
```
a = widgets.IntText(value=0, description='A =', disabled=False)
IPython.display.display(a)
def check(q):
IPython.display.clear_output(wait=False)
IPython.display.display(a)
if a.value == 111001:
print("That's right! Great work!")
else:
print("Sorry, try again or practice some more using the tools above")
a.observe(check, 'value')
%%html
<html>
<head>
<script type="text/javascript">
<!--
function toggle(id) {
var e = document.getElementById(id);
if(e.style.display == 'none')
e.style.display = 'block';
else
e.style.display = 'none';
}
//-->
</script>
</head>
<body>
<div id="question1">
Solution Question 1. <button id = "A"
onclick="toggle('answer1');">Click Here</button> to see the answer.
</div>
<div style="display:none" id="answer1">
As $64(2^6)$ is closest power of 2 for 57, so there will be atleast 6 digits for binary. <br />
the first step : 57 - 32 = 25 <br />
the next step : 25 - 16 = 9 <br />
the next step : 9 - 8 = 1 <br />
the last step : 1 - 1 = 0 <br />
so, the solution is: <br /> $1*32+ 1*16+ 1*8+ 0*4 + 0*2 + 1*1 $= $(111001)_2$
</div>
```
##### Question 2
$(79)_{10} = (B)_2$
```
b = widgets.IntText(value=0, description='B =', disabled=False)
IPython.display.display(b)
def check(q):
IPython.display.clear_output(wait=False)
IPython.display.display(b)
if b.value == 1001111:
print("That's right! Great work!")
else:
print("Sorry, try again or practice some more using the tools above")
b.observe(check, 'value')
%%html
<html>
<body>
<div id="question2">
Solution Question 2. <button id = "B"
onclick="toggle('answer2');">Click Here</button> to see the answer.
</div>
<div style="display:none" id="answer2">
As $128(2^7)$ is closest power of 2 for 79, so there will be atleast 7 digits for binary. <br />
the first step : 79 - 64 = 15 <br />
the next step : 15 - 8 = 7 <br />
the next step : 7 - 4 = 3 <br />
the next step : 3 - 2 = 1 <br />
the last step : 1 - 1 = 0 <br />
so, the solution is: <br /> $1*64+ 0*32+ 0*16+ 1*8+ 1*4 + 1*2 + 1*1 $= $(1001111)_2$
</div>
</body>
</html>
```
##### Question 3
$(39)_{10} = (C)_2$
```
c = widgets.IntText(value=0, description='C =', disabled=False)
IPython.display.display(c)
def check(q):
IPython.display.clear_output(wait=False)
IPython.display.display(c)
if c.value == 100111:
print("That's right! Great work!")
else:
print("Sorry, try again or practice some more using the tools above")
c.observe(check, 'value')
%%html
<html>
<body>
<div id="question3">
Solution Question 3. <button id = "C"
onclick="toggle('answer3');">Click Here</button> to see the answer.
</div>
<div style="display:none" id="answer3">
As $64(2^6)$ is closest power of 2 for 39, so there will be atleast 6 digits for binary. <br />
the first step : 39 - 32 = 7 <br />
the next step : 7 - 4 = 3 <br />
the next step : 3 - 2 = 1 <br />
the last step : 1 - 1 = 0 <br />
so, the solution is: <br /> $1*32+ 0*16+ 0*8+ 1*4 + 1*2 + 1*1 $= $(100111)_2$
</div>
</body>
</html>
```
### Convert the following Binary Numbers to Decimal
##### Question 4
$(10101)_{2} = (D)_{10}$
```
d = widgets.IntText(value=0, description='D =', disabled=False)
IPython.display.display(d)
def check(q):
IPython.display.clear_output(wait=False)
IPython.display.display(d)
if d.value == 21:
print("That's right! Great work!")
else:
print("Sorry, try again or practice some more using the tools above")
d.observe(check, 'value')
%%html
<html>
<body>
<div id="question4">
Solution Question 4. <button id = "C"
onclick="toggle('answer4');">Click Here</button> to see the answer.
</div>
<div style="display:none" id="answer4">
As $64(2^6)$ is closest power of 2 for 39, so there will be atleast 6 digits for binary. <br />
the first step : 39 - 32 = 7 <br />
the next step : 7 - 4 = 3 <br />
the next step : 3 - 2 = 1 <br />
the last step : 1 - 1 = 0 <br />
so, the solution is: <br /> $1*32+ 0*16+ 0*8+ 1*4 + 1*2 + 1*1 $= $(100111)_2$
</div>
</body>
</html>
```
##### Question 5
$(01111)_{2} = (E)_{10}$
```
e = widgets.IntText(value=0, description='E =', disabled=False)
IPython.display.display(e)
def check(q):
IPython.display.clear_output(wait=False)
IPython.display.display(e)
if e.value == 15:
print("That's right! Great work!")
else:
print("Sorry, try again or practice some more using the tools above")
e.observe(check, 'value')
%%html
<html>
<body>
<div id="question5">
Solution Question 5. <button id = "D"
onclick="toggle('answer5');">Click Here</button> to see the answer.
</div>
<div style="display:none" id="answer5">
$(10101)_2 = 1 * 2^{4} + 0 * 2^{3} + 1 * 2^{2} + 0* 2{1} + 1*2{0}$ <br />        
= $1*16+ 0*8+ 1*4 + 0*2 + 1*1$ <br />        
= $(21)_{10}$ <br />
</div>
</body>
</html>
```
##### Question 6
$(100100)_{2} = (F)_{10}$
```
f = widgets.IntText(value=0, description='F =', disabled=False)
IPython.display.display(f)
def check(q):
IPython.display.clear_output(wait=False)
IPython.display.display(f)
if f.value == 36:
print("That's right! Great work!")
else:
print("Sorry, try again or practice some more using the tools above")
f.observe(check, 'value')
%%html
<html>
<body>
<div id="question6">
Solution Question 6. <button id = "D"
onclick="toggle('answer6');">Click Here</button> to see the answer.
</div>
<div style="display:none" id="answer6">
$(100100)_2 = 1 * 2^{5} + 0 * 2^{4} + 0 * 2^{3} + 1 * 2^{2} + 0* 2{1} + 1*2{0}$ <br />        
= $1*32+ 0*16+ 0*8+ 1*4 + 0*2 + 0*1$ <br />        
= $(36)_{10}$
</div>
</body>
</html>
```
## Conclusion
There are many different ways to represent the numbers we use every day.<br>
The best way of representing a number can depend on what we're trying to do.
Digital computers typically use two states rather than more because:
* It is usually easy to distinguish two states;
* Two is enough to represent anything you like as a sequence of bits; and
* Computers don't mind long sequences.
Since we live in a digital era surrounded by machines, we must learn how machines understand the living world.
[](https://github.com/callysto/curriculum-notebooks/blob/master/LICENSE.md)
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from numpy.random import default_rng
# Numpy financial functions have been removed to their own package
# See https://numpy.org/numpy-financial/
import numpy_financial as npf
from whatif import Model
from whatif import get_sim_results_df
%matplotlib inline
class SingleProductSPF(Model):
def __init__(self, fixed_cost=60e+3, var_cost=90, selling_price=120,
spf_scale=10e+6, spf_exponent=-1.8):
self.fixed_cost = fixed_cost
self.var_cost = var_cost
self.selling_price = selling_price
self.spf_scale = spf_scale
self.spf_exponent = spf_exponent
def demand(self):
"""Demand based on SPF power function
D = a * P ** b
"""
_demand = self.spf_scale * self.selling_price ** self.spf_exponent
return _demand
def tot_revenue(self):
"""Total revenue
D * P
"""
_tot_revenue = self.demand() * self.selling_price
return _tot_revenue
def tot_cost(self):
"""Total cost
Fixed Cost + Var Cost * P
"""
_tot_cost = self.fixed_cost + self.var_cost * self.demand()
return _tot_cost
def profit(self):
"""Profit
Total revenue - total cost
"""
_profit = self.tot_revenue() - self.tot_cost()
return _profit
model = SingleProductSPF()
print(model)
print(model.demand())
print(model.tot_revenue())
print(model.tot_cost())
print(model.profit())
model.goal_seek?
print(f"Break even selling price: {model.goal_seek('profit', 0, 'selling_price', 90, 250):.2f}")
selling_prices = {'selling_price': np.arange(90, 501, 10)}
dt_selling_price = model.data_table(selling_prices, ['demand', 'tot_revenue', 'tot_cost', 'profit'])
dt_selling_price
plt.title("Sensitivity of profit to selling price")
plt.xlabel("Selling price")
plt.ylabel("Profit")
plt.plot(dt_selling_price['selling_price'], dt_selling_price['profit'])
plt.hlines(0, dt_selling_price['selling_price'].min(), dt_selling_price['selling_price'].max(),
linestyles='--')
plt.show();
```
## Simulation
Let's add uncertainty around the spf exponent.
```
rg = default_rng(4)
num_sim_reps = 1000
random_inputs = {'spf_exponent': rg.uniform(-2.0, -1.6, size=num_sim_reps)}
sim_results = model.simulate(random_inputs, ['profit'])
sim_results_df = get_sim_results_df(sim_results)
plt.title("Profit distribution")
plt.xlabel("Profit")
plt.ylabel("density")
plt.hist(sim_results_df['profit'], density=True);
plt.show();
```
## Extended model
```
class SingleProductSPF_extended(Model):
def __init__(self, fixed_cost=60e+3, var_cost=90, selling_price=120,
spf_scale=10e+6, spf_exponent=-1.8,
ot_threshold=1000, ot_var_cost=110, lost_demand_threshold=1200):
self.fixed_cost = fixed_cost
self.var_cost = var_cost
self.selling_price = selling_price
self.spf_scale = spf_scale
self.spf_exponent = spf_exponent
self.ot_threshold = ot_threshold
self.ot_var_cost = ot_var_cost
self.lost_demand_threshold = lost_demand_threshold
def demand(self):
"""Demand based on SPF power function
D = a * P ** b
"""
_demand = self.spf_scale * self.selling_price ** self.spf_exponent
return _demand
def units_sold(self):
"""Number of units sold
Min of D and lost_demand_threshold
"""
demand = self.demand()
_units_sold = np.minimum(demand, self.lost_demand_threshold)
return _units_sold
def tot_revenue(self):
"""Total revenue
D * P
"""
_tot_revenue = self.units_sold() * self.selling_price
return _tot_revenue
def tot_var_cost(self):
units_sold = self.units_sold()
units_at_base_cost = np.minimum(units_sold, self.ot_threshold)
units_at_ot_cost = np.maximum(0, units_sold - self.ot_threshold)
_tot_var_cost = self.var_cost * units_at_base_cost + self.ot_var_cost * units_at_ot_cost
return _tot_var_cost
def tot_cost(self):
"""Total cost
Fixed Cost + Var Cost * P
"""
_tot_cost = self.fixed_cost + self.tot_var_cost()
return _tot_cost
def profit(self):
"""Profit
Total revenue - total cost
"""
_profit = self.tot_revenue() - self.tot_cost()
return _profit
model_x = SingleProductSPF_extended()
print(model_x.units_sold())
print(model_x.tot_var_cost())
print(model_x.profit())
dt_selling_price_x = model_x.data_table(selling_prices, ['profit', 'demand', 'units_sold', ])
dt_selling_price_x
plt.title("Sensitivity of profit to selling price")
plt.xlabel("Selling price")
plt.ylabel("Profit")
plt.plot(dt_selling_price_x['selling_price'], dt_selling_price_x['profit'])
plt.hlines(0, dt_selling_price_x['selling_price'].min(), dt_selling_price_x['selling_price'].max(),
linestyles='--')
plt.show();
print(f"Break even selling price: {model_x.goal_seek('profit', 0, 'selling_price', 90, 250):.2f}")
print(f"Break even selling price: {model_x.goal_seek('profit', 0, 'selling_price', 200, 500):.2f}")
```
| github_jupyter |
# Executing Squonk services
This notebook is an example of executing Squonk services using Python's requests module.
It assumes you are executing against the JobExector service running in an OpenShift environment.
```
import requests
import json
# requests_toolbelt module is used to handle the multipart responses.
# Need to `pip install requests-toolbelt` from a terminal to install. This might need doing each time the Notebook pod starts
from requests_toolbelt.multipart import decoder
# Define some URLs and params
base_url = 'https://jobexecutor.prod.openrisknet.org/jobexecutor/rest'
services_url = base_url + '/v1/services'
jobexecutor_url = base_url + '/v1/jobs'
keycloak_url = 'https://sso.prod.openrisknet.org/auth/realms/openrisknet/protocol/openid-connect/token'
# set to False if self signed certificates are being used
tls_verify=True
```
## Check basic operation
```
# Test the PING service. Should give a 200 response and return 'OK'.
# If not then nothing else is going to work.
url = base_url + '/ping'
print("Requesting GET " + url)
resp = requests.get(url, verify=tls_verify)
print('Response Code: ' + str(resp.status_code))
print(resp.text)
```
## Authentication
```
# Need to specify your Keycloak SSO username and password so that we can get a token
import getpass
username = input('Username')
password = getpass.getpass('Password')
# Get token from Keycloak. This will have a finite lifetime.
# If your requests are getting a 401 error your token has probably expired.
data = {'grant_type': 'password', 'client_id': 'squonk-jobexecutor', 'username': username, 'password': password}
kresp = requests.post(keycloak_url, data = data)
j = kresp.json()
token = j['access_token']
token
```
## List all services
```
# Get a list of all the Squonk services that can be executed.
#
print("Requesting GET " + services_url)
jobs_resp = requests.get(services_url, headers={'Authorization': 'bearer ' + token}, verify=tls_verify)
print('Response Code: ' + str(jobs_resp.status_code))
json = jobs_resp.json()
print(str(len(json)) + " services found")
print(json)
```
## Getting details of a particular service
```
# find the service ID from the list in the list services cell
#service_id = 'core.dataset.filter.slice.v1'
#service_id = 'pipelines.rdkit.conformer.basic'
service_id = 'pipelines.rdkit.o3da.basic'
url = services_url + '/' + service_id
print("Requesting GET " + url)
jobs_resp = requests.get(url, headers={'Authorization': 'bearer ' + token}, verify=tls_verify)
print('Response Code: ' + str(jobs_resp.status_code))
json = jobs_resp.json()
print(json)
```
## List all jobs
```
# Result of the request is an array of JobStatus objects.
# The job ID and status are listed
print("Requesting GET " + jobexecutor_url)
jobs_resp = requests.get(jobexecutor_url, headers={'Authorization': 'bearer ' + token}, verify=tls_verify)
print('Response Code: ' + str(jobs_resp.status_code))
json = jobs_resp.json()
print(str(len(json)) + " jobs found")
for status in json:
print(status['jobId'] + ' ' + status['status'])
```
## Execute the 'Dataset Slice' service
```
# The 'Datast slice' takes a slice through a dataset specified by the number of records to skip and then the number to include.
# This is one of Squonk's 'internal' services.
# The job ID is stored in the job_id variable.
url = jobexecutor_url + '/core.dataset.filter.slice.v1'
data = {
'options': '{"skip":2,"count":3}',
'input_data': ('input_data', open('nci10.data', 'rb'), 'application/x-squonk-molecule-object+json'),
'input_metadata': ('input_metadata', open('nci10.metadata', 'rb'), 'application/x-squonk-dataset-metadata+json')
}
print("Requesting POST " + jobexecutor_url)
jobs_resp = requests.post(url, files=data, headers = {'Authorization': 'bearer ' + token, 'Content-Type': 'multipart/form'}, verify=tls_verify)
print('Response Code: ' + str(jobs_resp.status_code))
job_status = jobs_resp.json()
job_id = job_status['jobId']
print(job_status)
print("\nJobID: " + job_id)
```
## Get the status of the current job
```
# The job is defined by the job_id variable and is probably the last job executed
url = jobexecutor_url + '/' + job_id + '/status'
print("Requesting GET " + url )
jobs_resp = requests.get(url, headers={'Authorization': 'bearer ' + token}, verify=tls_verify)
print('Response Code: ' + str(jobs_resp.status_code))
json = jobs_resp.json()
json
```
## Get the results of a job.
```
# The job is defined by the job_id variable and is probably the last job executed.
# The status of the job needs to be 'RESULTS_READY'
# The response is a multipart response, typically containing the job status, the results metadata and the results data.
# This method can be called for a job any number of times until the job is deleted.
url = jobexecutor_url + '/' + job_id + '/results'
print("Requesting GET " + url )
jobs_resp = requests.get(url, headers={'Authorization': 'bearer ' + token}, verify=tls_verify)
print('Response Code: ' + str(jobs_resp.status_code))
multipart_data = decoder.MultipartDecoder.from_response(jobs_resp)
for part in multipart_data.parts:
print(part.content)
print(part.headers)
```
## Delete the job
```
# Once you have fetched the results you MUST delete the job.
# The job is defined by the job_id variable and is probably the last job executed.
url = jobexecutor_url + '/' + job_id
print("Requesting DELETE " + url)
jobs_resp = requests.delete(url, headers={'Authorization': 'bearer ' + token}, verify=tls_verify)
print('Response Code: ' + str(jobs_resp.status_code))
json = jobs_resp.json()
if 'status' in json and json['status'] == 'COMPLETED':
print('Job deleted')
else:
print('Problem deleting job')
```
## Delete all jobs
This is to help clean up if you get into a mess!
```
# Delete all jobs
# First get the current jobs
jobs_resp = requests.get(jobexecutor_url, headers={'Authorization': 'bearer ' + token}, verify=tls_verify)
print('Response Code: ' + str(jobs_resp.status_code))
json = jobs_resp.json()
print('Found ' + str(len(json)) + ' jobs')
# Now go through them and delete
# If successful the status of the job will then be COMPLETED.
for job in json:
id = job['jobId']
url = jobexecutor_url + '/' + id
print("Deleting " + url)
jobs_resp = requests.delete(url, headers={'Authorization': 'bearer ' + token}, verify=tls_verify)
j = jobs_resp.json()
print("Status: " + j['status'])
```
## Other services
In addition to the simple 'dataset slice' service many more meaningful ones are available.
Here are some examples illustrating the different categories of Squonk services:
1. Built in services running within the job executor Java process. These are limited to very simple and very fast operations
1. HTTP services running in the chemservices module that stream results and are designed for relatively short term execution (seconds or at most a few minutes) with the results being streamed immediately back to the requester.
1. Services running in a Docker container given the input data as files and writing the results as files. These are designed for more flexible implementation of services that can take longer to execute.
1. Nextflow services. Similar to Docker services, but defined as a Nextflow workflow that typically allows parallel execution on the K8S cluster or potentionally on an external cluster.
Execute one of these instead of the dataset slice one above.
```
# The 'Lipinski filter' takes calculates the classical rule of five properties and allows to filter based on these.
# We have implementations for ChemAxon and RDKit. Here we use the RDKit one.
# The default filter is the classical drug-likeness one defined by Lipinski but you can specify your owwn criteria instaead.
# This is one of Squonk's 'HTTP' services.
# The job ID is stored in the job_id variable.
url = jobexecutor_url + '/rdkit.calculators.lipinski'
data = {
'options': '{"filterMode":"INCLUDE_PASS"}',
'input_data': ('input_data', open('nci10.data', 'rb'), 'application/x-squonk-molecule-object+json'),
'input_metadata': ('input_metadata', open('nci10.metadata', 'rb'), 'application/x-squonk-dataset-metadata+json')
}
print("Requesting POST " + url)
jobs_resp = requests.post(url, files=data, headers = {'Authorization': 'bearer ' + token, 'Content-Type': 'multipart/form'}, verify=tls_verify)
print('Response Code: ' + str(jobs_resp.status_code))
job_status = jobs_resp.json()
job_id = job_status['jobId']
print(job_status)
print("\nJobID: " + job_id)
# passing data as SDF
url = jobexecutor_url + '/rdkit.calculators.lipinski'
data = {
'options': '{"filterMode":"INCLUDE_PASS"}',
'input': ('input', open('Kinase_inhibs.sdf', 'rb'), 'chemical/x-mdl-sdfile')
}
print("Requesting POST " + url)
jobs_resp = requests.post(url, files=data, headers = {'Authorization': 'bearer ' + token, 'Content-Type': 'multipart/form'}, verify=tls_verify)
print('Response Code: ' + str(jobs_resp.status_code))
job_status = jobs_resp.json()
job_id = job_status['jobId']
print(job_status)
print("\nJobID: " + job_id)
# sucos scoring passing 2 inputs as SDF
url = jobexecutor_url + '/pipelines.rdkit.sucos.basic'
data = {
'options': '{}',
'input': ('input', open('mols.sdf', 'rb'), 'chemical/x-mdl-sdfile'),
'target': ('target', open('benzene.sdf', 'rb'), 'chemical/x-mdl-sdfile')
}
print("Requesting POST " + url)
jobs_resp = requests.post(url, files=data, headers = {'Authorization': 'bearer ' + token, 'Content-Type': 'multipart/form'}, verify=tls_verify)
print('Response Code: ' + str(jobs_resp.status_code))
job_status = jobs_resp.json()
job_id = job_status['jobId']
print(job_status)
print("\nJobID: " + job_id)
# open3dAlign scoring passing 2 inputs as SDF
# passing the queryMol as pyrimethamine.mol does not work - it needs tob e converted to SDF
url = jobexecutor_url + '/pipelines.rdkit.o3da.basic'
data = {
'options': '{"arg.crippen":"false"}',
'input': ('input', open('dhfr_3d.sdf', 'rb'), 'chemical/x-mdl-sdfile'),
'queryMol': ('queryMol', open('pyrimethamine.sdf', 'rb'), 'chemical/x-mdl-sdfile')
}
print("Requesting POST " + url)
jobs_resp = requests.post(url, files=data, headers = {'Authorization': 'bearer ' + token, 'Content-Type': 'multipart/form'}, verify=tls_verify)
print('Response Code: ' + str(jobs_resp.status_code))
job_status = jobs_resp.json()
job_id = job_status['jobId']
print(job_status)
print("\nJobID: " + job_id)
# open3dAlign scoring passing inputs as dataset and query as SDF
url = jobexecutor_url + '/pipelines.rdkit.o3da.basic'
data = {
'options': '{"arg.crippen":"false"}',
'input_data': ('input_data', open('dhfr_3d.data.gz', 'rb'), 'application/x-squonk-molecule-object+json'),
'input_metadata': ('input_metadata', open('dhfr_3d.metadata', 'rb'), 'application/x-squonk-dataset-metadata+json'),
'queryMol': ('queryMol', open('pyrimethamine.sdf', 'rb'), 'chemical/x-mdl-sdfile')
}
print("Requesting POST " + url)
jobs_resp = requests.post(url, files=data, headers = {'Authorization': 'bearer ' + token, 'Content-Type': 'multipart/form'}, verify=tls_verify)
print('Response Code: ' + str(jobs_resp.status_code))
job_status = jobs_resp.json()
job_id = job_status['jobId']
print(job_status)
print("\nJobID: " + job_id)
# The 'Conformer generator' used RDKit ETKDG conformer generation tool to generate a number of conformers for the input structures.
# This is one of Squonk's 'Docker' services.
# The job ID is stored in the job_id variable.
service_id = 'pipelines.rdkit.conformer.basic'
data = {
'options': '{"arg.num":10,"arg.method":"RMSD"}',
'input_data': ('input_data', open('nci10.data', 'rb'), 'application/x-squonk-molecule-object+json'),
'input_metadata': ('input_metadata', open('nci10.metadata', 'rb'), 'application/x-squonk-dataset-metadata+json')
}
jobs_resp = requests.post(jobexecutor_url + '/' + service_id, files=data, headers = {'Authorization': 'bearer ' + token, 'Content-Type': 'multipart/form'}, verify=tls_verify)
print('Response Code: ' + str(jobs_resp.status_code))
job_status = jobs_resp.json()
job_id = job_status['jobId']
print(job_status)
print("\nJobID: " + job_id)
# Similarity screening using RDKit.
# This is one of Squonk's 'Nextflow' services.
# The job ID is stored in the job_id variable.
# NOTE: THIS IS NOT WORKING AS THE QUERY STRUCTURE IS NOT BEING PASSED CORRECTLY
service_id = 'pipelines.rdkit.screen.basic'
data = {
'options': '{"arg.query":{"source":"CC1=CC(=O)C=CC1=O","format":"smiles"},"arg.sim":{"minValue":0.5,"maxValue":1.0}}',
'input_data': ('input_data', open('nci10_data.json', 'rb'), 'application/x-squonk-molecule-object+json'),
'input_metadata': ('input_metadata', open('nci10_meta.json', 'rb'), 'application/x-squonk-dataset-metadata+json')
}
jobs_resp = requests.post(jobexecutor_url + '/' + service_id, files=data, headers = {'Authorization': 'bearer ' + token, 'Content-Type': 'multipart/form'}, verify=tls_verify)
print('Response Code: ' + str(jobs_resp.status_code))
job_status = jobs_resp.json()
job_id = job_status['jobId']
print(job_status)
print("\nJobID: " + job_id)
```
| github_jupyter |
# Determining Shoppers Online Purchasing Intent
<img align="center" width="600" height="500" src="https://nmgprod.s3.amazonaws.com/media/files/7c/58/7c58ed58f9ed85184957c34b7124e0b0/cover_image.jpg.640x360_q85_crop.jpg">
## Table of Contents <a name="t"></a>
1. [EDA](#eda)
- [Finding Errors](#errors)
- [Data Assumptions](#assump)
- [Interesting Relationships](#relat)
- [Outlier Significance](#out)
- [Interesting Facts](#facts)
- [Correlation Analysis](#corr)
- [Chi-Square Test](#chi)
2. [Benchmark Models](#bench)
3. [Data Pre-processing](#proc)
4. [Feature Selection](#feat)
5. [Modeling with Pre-processed Data](#aftpip)
6. [Hyperparameter Optimization](#hyopt)
7. [Modeling with Optimized Hyperparameters](#modhypopt)
8. [Ensamble Modeling](#ensamble)
9. [Exploring Feature Importance with Our Best Model](#featimp)
10. [Results & Reccomendations](#resrec)
11. [How about if we exclude page values?](#nopageval)
12. [What you can get better results](#futurework)
Online e-commerce applications are becoming a primary vehicle for people to find, compare, and ultimately purchase products. One of the fundamental questions that arises in e-commerce is to characterize, understand, and model user long-term purchasing intent, which is important as it allows for personalized and context relevant e-commerce services.
Understanding online purchase intent and its buildup over time is important because individuals spend large amounts of time and resources on online shopping—in the U.S. alone, e-commerce sales have reached over 350 billion USD per year and are expected to grow at around 15% annually.
Visit [Business Understanding Notebook](https://github.com/fairfield-university-ba545/project2-mem/blob/master/1_Business_Understanding/business_understanding.ipynb) for more detail.
In this project we aim to answer the following analytical questions:
- How accurately are we able to predict that a customer is going to make an online purchase on a website?
- What and what is the range of driving factors that lead to a purchase?
- Can the probability of online purchase produced through data mining represent the "real" probability of customer online purchase?
- Do those who purchase products tend to be of a certain region? returning or new visitor? buying near a special day, particular month or weekend? or spending more time on a specific page or click-thorugh (bounce) freuqently? What pages do they visit and time spent on them before exit?
## Data Dictionary
<img align="center" width="600" height="500" src="data_dictionary.jpg">
The origial dataset can be found here: [UCI Data](http://archive.ics.uci.edu/ml/datasets/Online+Shoppers+Purchasing+Intention+Dataset)
## Project Outline
- **Our project is divided into 3 main parts**
- **Steps and important findings will be shared in this report. If you want to see how the code was executed links to full Notebooks are provided at each step.**
<img align="left" width="500" height="500" src="Data_Process_Diagram.jpeg">
<span style="font-family:Papyrus; font-size:2em;">Now that we got the formalities out of the way, let's begin!</span>
- if you're wondering how I changed the font size and type: https://support.squarespace.com/hc/en-us/articles/206543587-Markdown-cheat-sheet
```
# Import packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
# setting to display max columns
pd.set_option('display.max_columns', 500)
# read in data
data = pd.read_csv('online_shoppers_intention.csv')
print('Number of rows, columns: ', data.shape)
```
## STEP 1: Exploratory Data Analysis (EDA) <a name="eda"></a>
The link to whole EDA notebook can be found here: **[Data Understanding](https://github.com/fairfield-university-ba545/project2-mem/blob/master/Project2FinalReport/EDA/EDA_data_understanding.ipynb)**
<font size="3">- Explore the dataset</font><br>
<font size="3">- Find errors to clean </font><br>
<font size="3">- Find interesting relationships that could answer/half-answer our analytical questions </font><br>
<font size="3">- Create new features out of those relationships </font><br>
Back to [Table of Contents](#t)
Month and VisitorType needs to be encoded
```
data.dtypes
```
- we have no missing values BUT that does not mean we do not have any logical errors
```
data.isnull().sum()
```
- There are many positive (4th quantile) outliers in our data **BUT** that is not neccessarally a bad thing
- Page durations have most evident outliers in their 4th quantile
- The following values occur the **most frequently**: Month of May, returning visitor, not the weekend, and revenue false
- **Missing 2 months** (only 10)
- **Target Feature (Revenue) Imbalanced** (10,422 False, 1908 True)
```
# Looking at the descriptive statisticss for each column
# verifies no missing values with count
data.describe(include='all').T
```
Let's look more closely.
## Finding Errors <a name="errors"></a>
- We will bin and encode Month, SpecialDay, and VisitorType
**Months**<br>
- Website had **most traffic** in 2nd and 4th quarter
- January and april are missing
- We decide to **bin it into quarters** and then use **One-Hot encoding** becasue we think we can capture more seasonality/trend which would be more meaningfull to the business /
```
print('Nbr of months: ',len(data.Month.value_counts()))
data.Month.value_counts()
```
- bin and one hot encode visitor type (i.e. Returning_Visitor = 1, New_Visitor = 2, Other = 3, then One-Hot)
```
data.VisitorType.value_counts()
```
### Encoding
```
# we have 10 months, missing Jan, Apr
# encode into quarters
def division_func_month(div) :
if div == 'Feb':
return 1
elif div == 'Mar':
return 1
elif div == 'May':
return 2
elif div == 'June':
return 2
elif div == 'Jul':
return 3
elif div == 'Aug':
return 3
elif div == 'Sep':
return 3
elif div == 'Oct':
return 4
elif div == 'Nov':
return 4
elif div == 'Dec':
return 4
else:
return 0
# from object to integers
data['quarter'] = data['Month']
# apply function
data['quarter'] = data['quarter'].apply(division_func_month)
# encoding for revenue and weekend
def div_fun_visType(div) :
if div == 'Returning_Visitor':
return 1
if div == 'New_Visitor':
return 2
else:
return 0
# from boolean to integers
data['VisitorType_encode'] = data['VisitorType']
# apply function
data['VisitorType_encode'] = data['VisitorType_encode'].apply(div_fun_visType)
data1=data.copy()
# One-hot encode
data1 = pd.get_dummies(data1, columns=['VisitorType_encode','quarter','SpecialDay'])
```
## Data Assumptions <a name="assump"></a>
**We assume the following to be the logical data values**
- No NA values allowed
- Administrative, Informational, Product related page and their corresponding duration can be zero (e.g. don't visit and spend time on that page) but cannot be a negative value visit
- Similarly you cannot have negative values for Bounce Rates, Exit rates, Page values, Operating system, Browser, Region
- You cannot have zero values for Exit rates as at some point user will exit
- Browser, region, traffic type and operating system represent segment a user based on their usage and zero category was not mentioned in data description
- Special Day similarly only contains certain values from 0 to 1 in increments of 0.2 (cannot have other values)
**We assume the following to be the correct data types**
- Count: Administrative, Informational, ProductRelated, Browser, OperatingSystem
- Binary: Revenue (Y), Weekend
- Categorical: TrafficType, Region, VisitorType, SpecialDay
- Continous: Administrative_Duration,Informational_Duration, ProductRelated_Duration, BounceRates, ExitRates, PageValues
- Date/time: Month
**NOTE:** Need to convert counts into ratios (Administrative, Informational, and Product Related)
**We only found one logical error**
- Where exit rates == 0 impute with median (becasue there is only 76 values we believe imputing with median is sufficient)
- We assume that exit rate cannot be zero for a user who logs into a website session
```
# we have 76 values to impute
data[(data['ExitRates'] == 0) & (data['BounceRates'] == 0)]
```
### Dealing with imbalanced dataset
You need to deal with **imbalanced data set** when the value of finding the minority class is much higher than that of finding the majority.
Let say that 1% of the population have that rare disease. Suppose that you assign the same cost to saying that a healthy man is sick or saying that a sick man is healthy. Provide a model that say that everybody are healthy, get 99% accuracy and go home early. The problem with such a model is that though it has high accuracy, it will probably not what you are looking for. Most of the time you want to find the people with the disease (giving high weigh to false negatives) much more than you are afraid to send an healthy person to unneeded test (low weight to false positives). In a real world health problem the ratio between the weight can easily be 1 to 1,000.
The imbalance in the distribution fails most algorithms from finding a proper solution.
We used **SMOTE over-sampling technique** to handle this<br>
- Visit our **[Baseline Modeling Notebook]()** to see the implementation.<br>
```
data.Revenue.value_counts().plot(kind='bar')
plt.title('Target Variable (Revenue) Imbalance')
```
## Interesting relationships - Outlier Significance <a name="relat"></a>
Through our EDA below we confirm our suspicion that outliers are important and decide to **skip the outlier treatment step** in our pipeline. More importantly it gives us insight in what **new features we can engineer**.
### Linear Relationship
- Bounce and Exit Rates have a good linear relationship
- Outliers could indicate Revenue false
- Revenue True seems more concentrated between 0 - 0.1
- Revenue false has much more observations
- Higher exit and bounce rates might mean a higher probability that the revenue will be false
- **Combining the following 2 features might give us better modeling results**
- we will use weighted average
```
ax = sns.scatterplot(x="BounceRates", y="ExitRates",hue = 'Revenue', data=data1)
```
### Negative Relationship
- We found significant negative relationships in many features and decide to create new ones with **ratios**
- We discover **3 interesting patterns**:
- Higher page values and lower duration is more likely to lead to a user purchase
- The higher the bounce rate the more likely revenue will be False
#### High Page Value and Lower Page Duration
- Higher page values and lower duration is more likely to lead to a user purchase
- Or in general higher page calue can lead to higher revenue
```
ax = sns.scatterplot(x="ProductRelated_Duration", y="PageValues",hue = 'Revenue', data=data1)
ax = sns.scatterplot(x="Informational_Duration", y="PageValues",hue = 'Revenue', data=data)
ax = sns.scatterplot(x="Administrative_Duration", y="PageValues",hue = 'Revenue', data=data)
```
**High Bounce Rate = higher probability of revenue being false (and vice versa)**
- Here we have a flipped relationship
- The higher the bounce rate the more likely revenue will be False
```
ax = sns.scatterplot(x="PageValues", y="BounceRates",hue = 'Revenue', data=data1)
ax = sns.scatterplot(x="Informational_Duration", y="BounceRates",hue = 'Revenue', data=data)
ax = sns.scatterplot(x="Administrative_Duration", y="BounceRates",hue = 'Revenue', data=data)
ax = sns.scatterplot(x="ProductRelated_Duration", y="BounceRates",hue = 'Revenue', data=data)
```
**Higher Exit rate = higher probability of revenue being false (and vice versa)!!**
```
ax = sns.scatterplot(x="PageValues", y="ExitRates",hue = 'Revenue', data=data1)
ax = sns.scatterplot(x="Informational_Duration", y="ExitRates",hue = 'Revenue', data=data)
ax = sns.scatterplot(x="Administrative_Duration", y="ExitRates",hue = 'Revenue', data=data)
ax = sns.scatterplot(x="ProductRelated_Duration", y="ExitRates",hue = 'Revenue', data=data)
```
### Confirming Outlier Significance (Box-plots & IQR) <a name="out"></a>
There are many outliers in out dataset and by removing them (as seen above) we could introduce a lot of error into our dataset. More importatnly we can benefit from these relationships to build better models.
```
for el in ['BounceRates', 'ExitRates','PageValues', 'Administrative_Duration','Informational_Duration',
'ProductRelated_Duration']:
sns.boxplot(x = 'Revenue', y = el, data = data1)
plt.show()
# select
outlier = data1.select_dtypes(include=['float64'])
outlier = outlier.drop('SpecialDay', axis = 1 )
# Calculate IQR range
def iqr_calc(df):
Q1 = df.quantile(0.25) # Q1 range
Q3 = df.quantile(0.75) # Q3 range
IQR = Q3 - Q1 # IQR range
outliers = ((df < (Q1 - 1.5 * IQR)) | (df > (Q3 + 1.5 * IQR))).sum()
return outliers
```
- About 10-20% of our data consists of outliers. We assume it would not be reccomended to impute so much in this case.
```
outlier.apply(iqr_calc)
```
### Interesting Facts <a name="facts"></a>
- most common number for page visits is 0 (could indicate that user does not visit all three pages in a session)
- a day before and after a special day (holiday) has the most volume
- 2 operating systems and browsers are used the most when on website
- most visitors come from region 2 (we do not know what region this actually is)
- traffic type 2 is most common
- 4th and then 2nd quarter have the most volume on the site
- most volume is during the week
- Returning visitors are the most common on the website
```
col_names = ['Administrative', 'Informational', 'SpecialDay','OperatingSystems', 'Browser', 'Region',
'TrafficType','quarter', 'Weekend', 'VisitorType_encode']
df = data[col_names]
for i, col in enumerate(df.columns):
plt.figure(i)
sns.countplot(x=col, data=df)
```
### Correlation Analysis <a name="corr"></a>
- High negative correlation between Exit Rates, Page durations,and page value makes sense. If the user spends more time on the page it will exit less and vice versa.
- Exit and Bounce rates might have a linear relationship becasue the more the user "bounces" from one page to another, more chances there are he/she will exit.
- Now interesting question is whether bounce rates cause more users to exit or is it something else?
- Page values correlation is highest for product related duration
- this means product related page makes the most revenue for the website
```
#Created a heatmap to see the correlation between the different variables
#Created a figure and are able to adjust the size of the graph
plt.figure(figsize = (20,13))
# select continous feature columns
continous_data = data1.select_dtypes(include=['float64']).drop('SpecialDay', axis = 1)
#Created a heatmap using Seaborn to display the correlation based on 'Spearman' method
##Added the annotation so that the numbers appear in each box
display = sns.heatmap(data1[continous_data.columns].corr(method = 'spearman'), annot=True, fmt='.2f', cmap='BuPu',
linewidth=.75, cbar_kws={'shrink': .75})
#Created a title
plt.title('Correlation Heatmap')
plt.savefig('corr_raw.png')
```
## Chi-Square (for categorical) <a name="chi"></a>
```
from sklearn.feature_selection import chi2
# features
X = data[['Administrative', 'Informational', 'SpecialDay','OperatingSystems', 'Browser', 'Region',
'TrafficType','quarter', 'Weekend', 'VisitorType_encode']]
# target
y = data['Revenue']
# chi-square test
chi_scores = chi2(X,y)
# here first array represents chi square values and second array represnts p-values
chi_scores
```
Since Operating system, traffic type, and region has higher the p-value, it says that this variables is independent of the responce and it **may not be viable for model training** (or have poorer performance). This was actually accurate as we will find out in our feature selection.
```
p_values = pd.Series(chi_scores[1],index = X.columns)
p_values.sort_values(ascending = False , inplace = True)
p_values.plot.bar()
plt.title('Significance (p-value) of Categorical Features')
```
### Benchmark Modeling Results <a name="bench"></a>
Before we began our pipelines we wanted to know what would be our baseline results. For this step we encoded our data (as above) and performed min-max standardization as it does not assume normality. Here are our results:
Code for the following results can be found here: **[Baseline Modeling Notebook](https://github.com/fairfield-university-ba545/project2-mem/blob/master/5_Modeling/Baseline_Modeling.ipynb)**
<img align="center" width="500" height="500" src="Baseline_models_results.jpeg">
We also ran **TPOT** (Genetic Algorithm). The result gave us the hint that **random forest might be the best model** and its parameters. **The best AUC score was 93.06%.** As this was not our best result we will only link the **[full notebook HERE](https://github.com/fairfield-university-ba545/project2-mem/blob/master/5_Modeling/tpot.ipynb).**
Back to [Table of Contents.](#t)
### Step 2: Data Pre-processing <a name="proc"></a>
We will not spend to much time here as these steps are quite standard. Visit the notebooks if you want to see how each step in the pipeline was implemented.
<img align="center" width="500" height="500" src="data_preprocessing_pic.jpeg">
**Pipeline A**
- [Feature Engineering](https://github.com/fairfield-university-ba545/project2-mem/blob/master/3_Data_Preprocessing/pipelineA/PipelineA_full.ipynb)
- [MinMax](https://github.com/fairfield-university-ba545/project2-mem/blob/master/3_Data_Preprocessing/pipelineA/PipelineA-MinMax.ipynb)
- [Normlize](https://github.com/fairfield-university-ba545/project2-mem/blob/master/3_Data_Preprocessing/pipelineA/PipelineA-Normalize_Skew.ipynb)
**Pipeline B**
- [Feature Engineering](https://github.com/fairfield-university-ba545/project2-mem/blob/master/3_Data_Preprocessing/PipelineB/PipelineB_full.ipynb)
- [Zscore](https://github.com/fairfield-university-ba545/project2-mem/blob/master/3_Data_Preprocessing/PipelineB/PipelineB-Zscore.ipynb)
- [Normalize](https://github.com/fairfield-university-ba545/project2-mem/blob/master/Project2FinalReport/Pipelines/PipelineB-Normalize_Skew.ipynb)
Back to [Table of Contents.](#t)
### Data Shape after data merge
- after our 2 pipelines we have 108 features that can be used for training
- our advice is to drop original features for those you engineered new ones becasue it can become tricky as you have to manually select features for modeling. **Only one mix of feature can be in a model** (i.e. PageValues_scales and PageValuesExitRate cannot be in the same model).
```
data_merge = pd.read_csv('all_model_data.csv', index_col=0)
print('New dataframe shape: ', data_merge.shape)
```
## 10-way Voting Feature Selection <a name="feat"></a>
All features were used in the model selection becasue we wanted to see whether our new engineered features beat our standard pipeline features. The most votes the feature receives the higher it ranks or more important it could be.
**Full notebook code implementation can be found [HERE](https://github.com/fairfield-university-ba545/project2-mem/blob/master/4_Feature_Selection/feature_selection_code.ipynb).**
### Methods:
- **Filter:** Pearson, f_classif (Anova F value)
- **Wrapper:** RFE with Logistic regression and XGBoost
- **Embeded:** Logistic Regression, Random Forest, XGBoost, LassoCV, RidgeClassifierCV
**Findings:**
We do not get the definitive answer whether our engineered features are better than "standard" ones. However, a wide range of choice can be good depending on the model. The most important features and variations are the relationships we highlighted above:
- Page durations, page values, bounce rates, and exit rates
- 4th quarter ends up in 20th place as it has the most website volume
Back to [Table of Contents.](#t)
```
feature_selection = pd.read_csv('feature_selection.csv', index_col=0)
feature_selection.head(20)
```
## Modeling After Pipelines and Feature Selection <a name="aftpip"></a>
- After our data was preprocessed from our 2 pipelines we ran many models to get the best features
- Models we ran were still mainly on defualt settings
- We ommited decision tree model as results did not improve significantly
- There were 10 iterations for each model (average or standard deviation was taken of all results)
Our model is indeed Random Forest with Average AUC CV 97%. We also see that standard pipeline features worked best for this model.
Full modeling code can be found here **[preprocessed_data_modeling-Copy1.ipynb](https://github.com/fairfield-university-ba545/project2-mem/blob/master/5_Modeling/preprocessed_data_modeling.ipynb)**
<img align="center" width="600" height="500" src="modeling_preprocessed_results.jpeg">
Back to [Table of Contents.](#t)
## Hyperparameter Optimization (GridSearchCV & RandomSearchCV) <a name="hypopt"></a>
- We ran gridsearchCV and randomsearchCV to get the best parameters possible. We still ran the models on the same features we got after our first modeling results. We received better results for almost all models however, it is important to note that these 2 methods are not always too reliable. A better yet more complicated method to use would be Bayesian Optimization.
**Full code notebook can be found [HERE](https://github.com/fairfield-university-ba545/project2-mem/blob/master/6_Model_Optimization/hyperparameter_tuning.ipynb)**
Back to [Table of Contents.](#t)
### Modeling After Hyperparameter Optimization <a name="modhypopt"></a>
Full notebook code can be found **[HERE](https://github.com/fairfield-university-ba545/project2-mem/blob/master/6_Model_Optimization/hypermerameter_optimized_modeling.ipynb)**
- After hyperperameter optimization we ran models again (10-iterations) with the same features and best parameters we got from gridsearchCV and randomsearchCV.
- We found out that for many models we got poorer results. However, it improved AUC result from XGBoost considerably by more than 3% (and KNN by 1%)
- We assume we could've gotten better results if we included more parameters in the grid search
<img align="center" width="600" height="500" src="modeling_hyp_optimization.jpeg">
Back to [Table of Contents.](#t)
## Ensamble modeling <a name="ensamble"></a>
As an exercise we tried ensamble modeling for which you can view the code **[HERE](https://github.com/fairfield-university-ba545/project2-mem/blob/master/7_Ensamble_Modeling/Ensamble_Modeling.ipynb).**
Back to [Table of Contents.](#t)
## EXPLORING FEATURE IMPORTANCE OF RANDOM FOREST (BEST MODEL) <a name="featimp"></a>
- We will outline the best feature below, however, you can find the **[code for top 3 features here](https://github.com/fairfield-university-ba545/project2-mem/blob/master/5_Modeling/best_model_feature_importance.ipynb)**. The findings will of course be covered in the results.
Back to [Table of Contents.](#t)
```
# read in data
import pandas as pd
data = pd.read_csv('all_model_data.csv', index_col = 0)
# split data into labels and target
X = data.drop('Revenue', axis = 1)
y = data.Revenue
# Split dataset into training set and test set
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2) # 80% training and 20% test
## Import the random forest model.
from sklearn.ensemble import RandomForestClassifier
## This line instantiates the model.
rf = RandomForestClassifier()
## Fit the model on your training data.
rf.fit(X_train, y_train)
feature_importances = pd.DataFrame(rf.feature_importances_,index = X_train.columns,
columns=['importance']).sort_values('importance', ascending=False)
```
Here for instance we see that a lot of page value variations are at the top. We need to choose only one variation of the feature.
```
feature_importances.head() # select one from each variation
```
## Top 5 Features Based on Importance
- it is worth mentioning exit rates that are quite to the top but are a mix feature with page values (PageValues_Scaled more important)
```
feature_importances.iloc[[0,13,18,21,29],:]
import matplotlib.pyplot as plt
import seaborn as sns
# subsetting dataframe
rev_true = data[data['Revenue']==True]
rev_false = data[data['Revenue']==False]
print(rev_true.shape)
print(rev_false.shape)
```
## Top Feature - PageValues_Scaled
```
feat1_1 = rev_true.PageValues_Scaled
sns.distplot(feat1_1, kde=False, label='Revenue True')
# Plot formatting
plt.legend(prop={'size': 12})
plt.title('PageValues_Scaled Revenue True')
plt.xlabel('Page Values')
plt.ylabel('Density')
```
- Higher page value (outliers) lead to revenue
- Page value gives an idea of which page in your site contributed more to your site's revenue
- To increase revenue attention must be given to quality of the webpage
```
feat1_0 = rev_false.PageValues_Scaled
sns.distplot(feat1_0, kde=False, label='Revenue False')
# Plot formatting
plt.legend(prop={'size': 12})
plt.title('PageValues_Scaled Revenue False')
plt.xlabel('Page Values')
plt.ylabel('Density')
```
- Lover the page value, less likely the purchase will occur
- Less outliers (values concentrated around 0)
```
sns.boxplot(x=feat1_1)
plt.title('Boxplot Page Values Revenue True')
```
- we see better here that page values is bigger and outliers are more to the right when revenue is true
```
sns.boxplot(x=feat1_0)
plt.title('Boxplot Page Values Revenue False')
```
- mostly 0 values with majority of outliers around 0-0.3 for revenue false
# RESULTS & RECCOMENDATIONS <a name="resrec"></a>
Back to [Table of Contents.](#t)
### Answering our analytical questions
Our best model was Random Forest (not optimized):
<img align="center" width="600" height="500" src="random_forest_pic.jpeg">
We can say with 97% certainty whether the user will make a online purchase or not. Moreover, the model correctly classified revenue false with 96% precision, and instances of Revenue true with 56% precision. Meainng we a have rather low false positive rate for Revenue false class and rather high false positive rate for revenue true class. Our recal is 0.88 and 0.76 which means the algorythm returned most of the relavent results and the model is good with identifyin false negatives.
We reccomend using our **KNN** model (with no parameter tuning) if your goal is classifying positive classes of revenue true, as it has highest precision - 79%.
<img align="center" width="600" height="500" src="knn_best_precision.jpeg">
## Here are our reccomendations:
1. Make the website as easy to use a possible
- the more time the user spends on product related page the more likely it will make a purchase
2. Product related page and duration is the most important as it has the highest positive correlation with page value
- page value tells us from which page we get the most revenue
3. If you keep the user on the product related page for 0.08 or more there is a higher chance he will make a purchase
- Attention needs to be grabbed here (i.e. promotions, variaty of choice, specific call to action, discounts etc.)
4. The more pages the user visits less likely he will make a purchase
- one explanation for these extremes could be developers websrapping the website, however, we do not believe that is the case
- we think if the user has to go to different websites the information and features are not well organized which in turn leads for users not purchasing and exiting
- the more quality information the user can get without leaving a page the better (mazon 1-click order is a good example)
5. The majority of users do not make a purchase
- we reccomend focusing on a great call to action that would lead users to make a purchase
6. the website has the most returning users which means **good retention rate**
- we reccomend reaching out to new customer segments to reach new users (e.g. promotions, engagement, surveys to learn what to improve...)
- User with traffic type 2 and from region 1 account for the majority of traffic (we do not know from data documentation what these numbers represent). We reccomend expanding to other channels while still taking care of this segment which you clearly know how to attract. A different strategy might need to be used when expanding in the new segment or region. Our reccomendation is reaching out to a segment that has market revevance (likes to buy) and is identified with what you are selling.
7. Focus marketing efforts for when the most website traffic happens
- before and after a special day (holiday)
- 4th and 2nd quarter
- during the week
8. Users generally use 2 operating systems and 2 browsers. We would reccomend improve responsiveness to these 2 most popular types. When expanding these 2 factors will need to be researched if they need to be expanded.
## How about if we exclude page values? <a name="nopageval"></a>
We conducted this analysis as we thought would be interesting to see how the model behaves and what features it considers the most important. Below are a few highlights but if you want to see the whole analysis just click **[HERE](https://github.com/fairfield-university-ba545/project2-mem/blob/master/5_Modeling/modeling_without_pagevalues.ipynb).**
Back to [Table of Contents.](#t)
#### Few take-aways
- For revenue true we see low distribution when duration is lower (0-0.06). Makes sense if they spend less time on the page less likely users will purchase. However, those who do must be familar with the website and know what they want. The distribution then increases exponentially until 0.10 - 0.14 and then spikes again for 0.15. 0.10 - 0.15 duration could be considered the hisghes chance the user will purchase comparing to revenue false. So it would be wise to devise the website in such a way users will spend that amount of time on the page (specific call to action, discounts etc.)
- For revenue false there are many 0 values which signify the user did not make it to the page = no revenue. It would be wise to get some answers as to why that happened.
- The closer the the exit rate is to zero and less outliers it has to the right, the more likely user will make a purchase.
- It would be interesting see what are the factors on the website that customers do not like that prompts them to exit
<img align="center" width="400" height="500" src="no_pagevalues_feat_importance.jpeg">
#### Modeling results
As wee see our modeling results are far worse comparing to precision. However, recall does not seem that bad meaning we received we got revavent results (lower number of false negatives).
<img align="center" width="400" height="500" src="no_pagevalues_modeling_results.jpeg">
### Here are our reccomendations of what could lead to better results <a name="futurework"></a>
Back to [Table of Contents.](#t)
There are limitations with this dataset as it is focused on short-term user activity on whether a given user session will result in a purchase. But its important to note that the purchase intent of a consumer may slowly build up over time, and may not instantaneously lead to a purchase. Furthermore, traditional studies often examine user behavior on a single e-commerce platform, while users may use several different services and move across e-commerce platforms when deciding which product to purchase and where. Thus, what is missing from the picture is a **cross-platform analysis of how user purchase intent varies over time.** To this end, it is important to contrast the population of purchasing users with the population of non- purchasing users, and then also identify how purchasers’ online behavior changes over time from the norm as a result of impending purchases.
- make use of ensambles models especially the voting ensamble
| github_jupyter |
### Project :: Evolution Strategies

Remember the idea behind Evolution Strategies? Here's a neat [blog post](https://blog.openai.com/evolution-strategies/) about 'em.
Can you reproduce their success? You will have to implement evolutionary strategies and see how they work.
This project is optional; has several milestones each worth a number of points [and swag].
__Milestones:__
* [10pts] Basic prototype of evolutionary strategies that works in one thread on CartPole
* [+5pts] Modify the code to make them work in parallel
* [+5pts] if you can run ES distributedly on at least two PCs
* [+10pts] Apply ES to play Atari Pong at least better than random
* [++] Additional points for all kinds of cool stuff besides milestones
__Rules:__
* This is __not a mandatory assignment__, but it's a way to learn some cool things if you're getting bored with default assignments.
* Once you decided to take on this project, please tell any of course staff members so that we can help ypu if you get stuck.
* There's a default implementation of ES in this [openai repo](https://github.com/openai/evolution-strategies-starter). It's okay to look there if you get stuck or want to compare your solutions, but each copy-pasted chunk of code should be understood thoroughly. We'll test that with questions.
### Tips on implementation
* It would be very convenient later if you implemented a function that takes policy weights, generates a session and returns policy changes -- so that you could then run a bunch of them in parallel.
* The simplest way you can do multiprocessing is to use [joblib](https://www.google.com/search?client=ubuntu&channel=fs&q=joblib&ie=utf-8&oe=utf-8)
* For joblib, make sure random variables are independent in each job. Simply add `np.random.seed()` at the beginning of your "job" function.
Later once you got distributed, you may need a storage that gathers gradients from all workers. In such case we recommend [Redis](https://redis.io/) due to it's simplicity.
Here's a speed-optimized saver/loader to store numpy arrays in Redis as strings.
```
import joblib
from six import BytesIO
def dumps(data):
"""converts whatever to string"""
s = BytesIO()
joblib.dump(data, s)
return s.getvalue()
def loads(self, string):
"""converts string to whatever was dumps'ed in it"""
return joblib.load(BytesIO(string))
```
### Tips on atari games
* There's all the pre-processing and tuning done for you in the code below
* Images rescaled to 42x42 to speed up computation
* We use last 4 frames as observations to account for ball velocity
* The code below requires ```pip install Image``` and ```pip install gym[atari]```
* You may also need some dependencies for gym[atari] - google "gym install all" dependencies or use our pre-built environment.
* The recommended agent architecture is a convolutional neural network. Dense network will also do.
May the force be with you!
```
from pong import make_pong
import numpy as np
env = make_pong()
print(env.action_space)
# get the initial state
s = env.reset()
print(s.shape)
import matplotlib.pyplot as plt
%matplotlib inline
# plot first observation. Only one frame
plt.imshow(s.swapaxes(1, 2).reshape(-1, s.shape[-1]).T)
# next frame
new_s, r, done, _ = env.step(env.action_space.sample())
plt.imshow(new_s.swapaxes(1, 2).reshape(-1, s.shape[-1]).T)
# after 10 frames
for _ in range(10):
new_s, r, done, _ = env.step(env.action_space.sample())
plt.imshow(new_s.swapaxes(1, 2).reshape(-1, s.shape[-1]).T, vmin=0)
< tons of your code here or elsewhere >
```
| github_jupyter |
```
import numpy as np
import pymc3 as pm
import theano.tensor as tt
import statsmodels
import patsy
import pystan
import matplotlib.pylab as plt
%matplotlib inline
dataset = statsmodels.datasets.get_rdataset(package='survival', dataname='flchain' )
d = dataset.data.query('futime > 7').sample(500)
d.reset_index(level=0, inplace=True)
d.rename(columns={'futime': 't', 'death': 'event'}, inplace=True)
d.head()
y, x_df = patsy.dmatrices("event ~ age + sex", d, return_type='dataframe')
x_df = x_df.iloc[:, x_df.columns != 'Intercept']
N, M = x_df.shape
x = x_df.as_matrix()
y = d['t'].values
event = d['event'].values.astype(int)
```
https://github.com/hammerlab/survivalstan/blob/master/survivalstan/stan/weibull_survival_model.stan
```
datadict = dict(N=N, M=M, x=x, y=y, event=event)
stan_m = """
functions {
vector sqrt_vec(vector x) {
vector[dims(x)[1]] res;
for (m in 1:dims(x)[1]){
res[m] = sqrt(x[m]);
}
return res;
}
vector bg_prior_lp(real r_global, vector r_local) {
r_global ~ normal(0.0, 10.0);
r_local ~ inv_chi_square(1.0);
return r_global * sqrt_vec(r_local);
}
}
data {
// dimensions
int<lower=0> N; // number of observations
int<lower=1> M; // number of predictors
// observations
matrix[N, M] x; // predictors for observation n
vector[N] y; // time for observation n
vector[N] event; // event status (1:event, 0:censor) for obs n
}
transformed data {
real<lower=0> tau_mu;
real<lower=0> tau_al;
tau_mu = 10.0;
tau_al = 10.0;
}
parameters {
real<lower=0> tau_s_raw;
vector<lower=0>[M] tau_raw;
real alpha_raw;
vector[M] beta_raw;
real mu;
}
transformed parameters {
vector[M] beta;
real<lower=0> alpha;
vector[N] lp;
beta = bg_prior_lp(tau_s_raw, tau_raw) .* beta_raw;
alpha = exp(tau_al * alpha_raw);
for (n in 1:N) {
lp[n] = mu + dot_product(x[n], beta);
}
}
model {
// priors
beta_raw ~ normal(0.0, 1.0);
alpha_raw ~ normal(0.0, 0.1);
mu ~ normal(0.0, tau_mu);
// likelihood
for (n in 1:N) {
if (event[n]==1)
y[n] ~ weibull(alpha, exp(-(lp[n])/alpha));
else
target += weibull_lccdf(y[n] | alpha, exp(-(lp[n])/alpha));
}
}
"""
tr_stan = pystan.stan(model_code=stan_m, data=datadict,
iter=2000, warmup=1000,
chains=4, n_jobs=4, verbose=False)
tr_stan.plot(pars=['mu','alpha', 'beta']);
sampler_params = tr_stan.get_sampler_params(inc_warmup=False)
divergent = [x for y in sampler_params for x in y['divergent__']]
n = sum(divergent)
N = len(divergent)
print('{} of {} iterations ended with a divergence ({}%)'.format(n, N,
100 * n / N))
mu_stan = tr_stan.extract(permuted=True)['mu'].mean(axis=0)
alpha_stan = tr_stan.extract(permuted=True)['alpha'].mean(axis=0)
beta_stan = tr_stan.extract(permuted=True)['beta'].mean(axis=0)
sd_mu = 10.0
sd_al = 10.0
def weibull_lccdf(value, alpha, beta):
return - (value / beta)**alpha
with pm.Model() as weibull:
beta_raw = pm.Normal('b0', mu=0., sd=1., shape=M)
alpha_raw = pm.Normal('a0', mu=0., sd=.1)
mu = pm.Normal('mu', mu=0., sd=sd_mu)
tau_s_raw = pm.HalfNormal('tau_s_raw', 10.)
tau_raw = pm.ChiSquared('tau_raw', 1., shape=M)
beta = pm.Deterministic('beta', tau_s_raw * tt.sqrt(1. / tau_raw) * beta_raw)
alpha = pm.Deterministic('alpha', tt.exp(sd_al * alpha_raw))
lp = mu + tt.dot(x, beta)
y1 = pm.Weibull(
'y1', alpha, tt.exp(-(lp[event == 1]) / alpha), observed=y[event == 1])
y0 = pm.Potential(
'y0', weibull_lccdf(y[event == 0], alpha,
tt.exp(-(lp[event == 0]) / alpha)))
with weibull:
trace = pm.sample(1000, tune=1000, init='adapt_diag')
pm.traceplot(trace,
varnames=['mu','alpha', 'beta'],
lines=dict(mu=mu_stan, alpha=alpha_stan, beta=beta_stan));
```
[Weibull Survival Regression](https://discourse.pymc.io/t/weibull-survival-regression-aft/1107/3)
```
time <-c(59, 115, 156, 421, 431, 448, 464, 475, 477, 563, 638, 744, 769, 770, 803, 855, 1040, 1106, 1129, 1206, 1227, 268, 329, 353, 365, 377)
event <- c(1, 1, 1, 0, 1, 0, 1, 1, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 0)
library(survival)
r <- survreg(Surv(time, event) ~ 1, dist="weibull")
beta <- 1/r$scale
eta <- exp(r$coefficients[1])
> beta
[1] 1.10806
> eta
(Intercept)
1225.419
```
```
time = np.asarray([59, 115, 156, 421, 431, 448, 464, 475, 477, 563, 638, 744,
769, 770, 803, 855, 1040, 1106, 1129, 1206, 1227, 268, 329, 353, 365, 377], dtype=np.float64)
event = np.asarray([1, 1, 1, 0, 1, 0, 1, 1, 0, 1, 1, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 1, 1, 0])
sd_mu = 100.0
sd_al = 10.0
def weibull_lccdf(value, alpha, beta):
return - (value / beta)**alpha
with pm.Model() as weibull:
alpha_raw = pm.Normal('a0', mu=0., sd=.1)
mu = pm.Normal('mu', mu=0., sd=sd_mu)
alpha = pm.Deterministic('alpha', tt.exp(sd_al * alpha_raw))
beta = pm.Deterministic('beta', tt.exp(mu / alpha))
y1 = pm.Weibull(
'y1', alpha, beta, observed=time[event == 1])
y0 = pm.Potential(
'y0', weibull_lccdf(time[event == 0], alpha, beta))
trace = pm.sample(1000, tune=1000, init='adapt_diag')
pm.traceplot(trace);
df = pm.summary(trace)
df
```
https://github.com/stan-dev/example-models/blob/master/bugs_examples/vol1/kidney/kidney.stan
```
with pm.Model() as m:
alpha = pm.Normal('alpha', 0., 10.)
r = pm.Gamma('r', 1., .001, testval=.25)
beta = pm.Deterministic('beta', tt.exp(-alpha / r))
y1 = pm.Weibull(
'y1', r, beta, observed=time[event == 1])
y0 = pm.Bernoulli(
'y0', tt.exp(-tt.pow(time[event == 0] / beta, r)),
observed=np.ones(np.sum(event == 0)))
# y0 = pm.Potential(
# 'y0', weibull_lccdf(time[event == 0], r, beta))
for var in m.basic_RVs:
print(var.name, var.logp(m.test_point))
with m:
tr = pm.sample(1000, tune=1000, init='adapt_diag')
pm.traceplot(tr);
pm.summary(tr)
```
http://austinrochford.com/posts/2017-10-02-bayes-param-survival.html
```
logtime = np.log(time)
def gumbel_sf(y, mu, sigma):
return 1.0 - tt.exp(-tt.exp(-(y - mu) / sigma))
# Build Bayesian model
with pm.Model() as model:
# Hyperprior
s = pm.HalfNormal("s", tau=5.0)
# Priors
gamma = pm.Normal("gamma", 0., 5.0)
# Likelihood for uncensored and censored survival times
y_obs = pm.Gumbel("y_obs", mu=gamma, beta=s, observed=logtime[event == 1])
y_cens = pm.Bernoulli("y_cens", p=gumbel_sf(
logtime[event == 0], mu=gamma, sigma=s), observed=np.ones(np.sum(event == 0)))
trace = pm.sample(1000, tune=1000, init='adapt_diag')
pm.traceplot(trace);
df = pm.summary(trace)
df
```
| github_jupyter |
<h1>Video Sliding Windows</h1>
<p>
So far we restricted ourselves to 1D time series, but the idea of recovering periodic dynamics with geometry can just as easily apply to multivariate signals. In this module, we will examine sliding windows of videos as an exmaple. Many natural videos also have periodicity, such as this video of a woman doing jumping jacks
</p>
```
import io
import base64
from IPython.display import HTML
video = io.open('jumpingjacks.ogg', 'r+b').read()
encoded = base64.b64encode(video)
HTML(data='''<video alt="test" controls>
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded.decode('ascii')))
```
<p>
Video can be decomposed into a 3D array, which has dimensions width x height x time. To tease out periodicity in geometric form, we will do the exact same thing as with sliding window 1D signal embeddings, but instead of just one sample per time shift, we need to take every pixel in every frame in the time window. The figure below depicts this
</p>
<img src = "VideoStackTime.svg"><BR><BR>
To see this visually in the video next to PCA of the embedding, look at the following video
```
video = io.open('jumpingjackssliding.ogg', 'r+b').read()
encoded = base64.b64encode(video)
HTML(data='''<video alt="test" controls>
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded.decode('ascii')))
```
<h2>PCA Preprocessing for Efficiency</h2><BR>
One issue we have swept under the rug so far is memory consumption and computational efficiency. Doing a raw sliding window of every pixel of every frame in the video would blow up in memory. However, even though there are <code>WH</code> pixels in each frame, there are only <code>N</code> frames in the video. This means that each frame in the video can be represented in an <code>(N-1)</code> dimensional subspace of the pixel space, and the coordinates of this subspace can be used in lieu of the pixels in the sliding window embedding. This can be done efficiently with a PCA step before the sliding window embedding. Run the cell below to load code that does PCA efficiently
```
#Do all of the imports and setup inline plotting
%matplotlib notebook
import numpy as np
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA
from mpl_toolkits.mplot3d import Axes3D
import scipy.interpolate
from ripser import ripser
from persim import plot_diagrams
from VideoTools import *
##Here is the actual PCA code
def getPCAVideo(I):
ICov = I.dot(I.T)
[lam, V] = linalg.eigh(ICov)
V = V*np.sqrt(lam[None, :])
return V
```
<h2>Jumping Jacks Example Live Demo</h2><BR>
Let's now load in code that does sliding window embeddings of videos. The code is very similar to the 1D case, and it has the exact same parameters. The only difference is that each sliding window lives in a Euclidean space of dimension the number of pixels times <code>dim</code>. We're also using linear interpolation instead of spline interpolation to keep things fast
```
def getSlidingWindowVideo(I, dim, Tau, dT):
N = I.shape[0] #Number of frames
P = I.shape[1] #Number of pixels (possibly after PCA)
pix = np.arange(P)
NWindows = int(np.floor((N-dim*Tau)/dT))
X = np.zeros((NWindows, dim*P))
idx = np.arange(N)
for i in range(NWindows):
idxx = dT*i + Tau*np.arange(dim)
start = int(np.floor(idxx[0]))
end = int(np.ceil(idxx[-1]))+2
if end >= I.shape[0]:
X = X[0:i, :]
break
f = scipy.interpolate.interp2d(pix, idx[start:end+1], I[idx[start:end+1], :], kind='linear')
X[i, :] = f(pix, idxx).flatten()
return X
```
Finally, let's load in the jumping jacks video and perform PCA to reduce the number of effective pixels. <BR>
<i>Note that loading the video may take a few seconds on the virtual image</i>
```
#Load in video and do PCA to compress dimension
(X, FrameDims) = loadImageIOVideo("jumpingjacks.ogg")
X = getPCAVideo(X)
```
Now let's do a sliding window embedding and examine the sliding window embedding using TDA. As before, you should tweak the parameters of the sliding window embedding and study the effect on the geometry.
```
#Given that the period is 30 frames per cycle, choose a dimension and a Tau that capture
#this motion in the roundest possible way
#Plot persistence diagram and PCA
dim = 30
Tau = 1
dT = 1
#Get sliding window video
XS = getSlidingWindowVideo(X, dim, Tau, dT)
#Mean-center and normalize sliding window
XS = XS - np.mean(XS, 1)[:, None]
XS = XS/np.sqrt(np.sum(XS**2, 1))[:, None]
#Get persistence diagrams
dgms = ripser(XS)['dgms']
#Do PCA for visualization
pca = PCA(n_components = 3)
Y = pca.fit_transform(XS)
fig = plt.figure(figsize=(12, 6))
plt.subplot(121)
plot_diagrams(dgms)
plt.title("1D Persistence Diagram")
c = plt.get_cmap('nipy_spectral')
C = c(np.array(np.round(np.linspace(0, 255, Y.shape[0])), dtype=np.int32))
C = C[:, 0:3]
ax2 = fig.add_subplot(122, projection = '3d')
ax2.set_title("PCA of Sliding Window Embedding")
ax2.scatter(Y[:, 0], Y[:, 1], Y[:, 2], c=C)
ax2.set_aspect('equal', 'datalim')
plt.show()
```
<h1>Periodicities in The KTH Dataset</h1><BR>
We will now examine videos from the <a href = "http://www.nada.kth.se/cvap/actions/">KTH dataset</a>, which is a repository of black and white videos of human activities. It consists of 25 subjects performing 6 different actions in each of 4 scenarios. We will use the algorithms developed in this section to measure and rank the periodicity of the different video clips.
<h2>Varying Window Length</h2><BR>
For our first experiment, we will be showing some precomputed results of varying the sliding window length, while choosing Tau and dT appropriately to keep the dimension and the number of points, respectively, the same in the sliding window embedding. As an example, we will apply it to one of the videos of a subject waving his hands back and forth, as shown below
```
video = io.open('KTH/handwaving/person01_handwaving_d1_uncomp.ogg', 'r+b').read()
encoded = base64.b64encode(video)
HTML(data='''<video alt="test" controls>
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded.decode('ascii')))
```
We have done some additional preprocessing, including applying a bandpass filter to each PCA pixel to cut down on drift in the video. Below we show a video varying the window size of the embedding and plotting the persistence diagram, "self-similarity matrix" (distance matrix), and PCA of the embedding, as well as an evolving plot of the maximum persistence versus window size:
```
video = io.open('Handwaving_Deriv10_Block160_PCA10.ogg', 'r+b').read()
encoded = base64.b64encode(video)
HTML(data='''<video alt="test" controls>
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded.decode('ascii')))
```
As you can see, the maximum persistence peaks at around 40 frames, which is the period of each hand wave. This is what the theory we developed for 1D time series would have predicted as the roundest window.<BR>
<h1>Quasiperiodicity Quantification in Video</h1><BR>
<p>
We now examine how this pipeline can be used to detect quasiperiodicity in videos. As an example, we examine videos from high-speed glottography, or high speed videos (4000 fps) of the left and right vocal folds in the human vocal tract. When a person has a normal voice, the vocal folds oscillate in a periodic fashion. On the other hand, if they have certain types of paralysis or near chaotic dynamics, they can exhibit biphonation just as the horse whinnies did. More info can be found in <a href = "https://arxiv.org/abs/1704.08382">this paper</a>.
</p>
<h2>Healthy Subject</h2>
<p>
Let's begin by analyzing a video of a healthy person. In this example and in the following example, we will be computing both persistent H1 and persistent H2, so the code may take a bit longer to run.
</p>
#### Questions
* What can we say about the vocal folds of a healthy subject based on the persistence diagram?
```
video = io.open('NormalPeriodicCrop.ogg', 'r+b').read()
encoded = base64.b64encode(video)
HTML(data='''<video alt="test" controls>
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded.decode('ascii')))
(X, FrameDims) = loadVideo("NormalPeriodicCrop.ogg")
X = getPCAVideo(X)
dim = 70
Tau = 0.5
dT = 1
derivWin = 10
#Take a bandpass filter in time at each pixel to smooth out noise
[X, validIdx] = getTimeDerivative(X, derivWin)
#Do the sliding window
XS = getSlidingWindowVideo(X, dim, Tau, dT)
#Mean-center and normalize sliding window
XS = XS - np.mean(XS, 1)[:, None]
XS = XS/np.sqrt(np.sum(XS**2, 1))[:, None]
#Compute and plot persistence diagrams
print("Computing persistence diagrams...")
dgms = ripser(XS, maxdim=2)['dgms']
print("Finished computing persistence diagrams")
plt.figure()
plot_diagrams(dgms)
plt.title("Persistence Diagrams$")
plt.show()
```
<h2>Subject with Biphonation</h2>
<p>
Let's now examine a video of someone with a vocal pathology. This video may still appear periodic, but if you look closely there's a subtle shift going on over time
</p>
```
video = io.open('ClinicalAsymmetry.mp4', 'r+b').read()
encoded = base64.b64encode(video)
HTML(data='''<video alt="test" controls>
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded.decode('ascii')))
(X, FrameDims) = loadVideo("ClinicalAsymmetry.mp4")
X = getPCAVideo(X)
X = X[0:200, :]
#'dim':32, 'Tau':0.25, 'dT':0.25, 'derivWin':2
dim = 100
Tau = 0.25
dT = 0.5
derivWin = 5
#Take a bandpass filter in time at each pixel to smooth out noise
[X, validIdx] = getTimeDerivative(X, derivWin)
#Do the sliding window
XS = getSlidingWindowVideo(X, dim, Tau, dT)
print("XS.shape = ", XS.shape)
#Mean-center and normalize sliding window
XS = XS - np.mean(XS, 1)[:, None]
XS = XS/np.sqrt(np.sum(XS**2, 1))[:, None]
#Compute and plot persistence diagrams
print("Computing persistence diagrams...")
dgms = ripser(XS, maxdim=2)['dgms']
print("Finished computing persistence diagrams")
plt.figure()
plt.title("Persistence Diagrams$")
plot_diagrams(dgms)
plt.show()
```
#### Question:
* What shape is this? What does this say about the underlying frequencies involved?
<h2>Another Subject with Biphonation</h2>
<p>
Let's now examine another person with a vocal pathology, this time due to mucus that is pushed out of the vocal folds every other oscillation. This time, we will look at both $\mathbb{Z} / 2\mathbb{Z}$ coefficients and $\mathbb{Z} / 3 \mathbb{Z}$ coefficients.
</p>
#### Questions
* Can you see any changes between $\mathbb{Z} / 2\mathbb{Z}$ coefficients and $\mathbb{Z} / 3 \mathbb{Z}$ coefficients? the What shape is this? Can you relate this to something we've seen before?
```
video = io.open('LTR_ED_MucusBiphonCrop.ogg', 'r+b').read()
encoded = base64.b64encode(video)
HTML(data='''<video alt="test" controls>
<source src="data:video/mp4;base64,{0}" type="video/mp4" />
</video>'''.format(encoded.decode('ascii')))
(X, FrameDims) = loadVideo("LTR_ED_MucusBiphonCrop.ogg")
X = getPCAVideo(X)
X = X[0:200, :]
#'dim':32, 'Tau':0.25, 'dT':0.25, 'derivWin':2
dim = 100
Tau = 1
dT = 0.25
derivWin = 5
#Take a bandpass filter in time at each pixel to smooth out noise
[X, validIdx] = getTimeDerivative(X, derivWin)
#Do the sliding window
XS = getSlidingWindowVideo(X, dim, Tau, dT)
print("XS.shape = ", XS.shape)
#Mean-center and normalize sliding window
XS = XS - np.mean(XS, 1)[:, None]
XS = XS/np.sqrt(np.sum(XS**2, 1))[:, None]
#Compute and plot persistence diagrams
print("Computing persistence diagrams...")
dgms2 = ripser(XS, maxdim=2, coeff=2)['dgms']
dgms3 = ripser(XS, maxdim=2, coeff=3)['dgms']
print("Finished computing persistence diagrams")
plt.figure(figsize=(8, 4))
plt.subplot(121)
plot_diagrams(dgms2)
plt.title("Persistence Diagrams $\mathbb{Z}2$")
plt.subplot(122)
plot_diagrams(dgms3)
plt.title("Persistence Diagrams $\mathbb{Z}3$")
plt.show()
```
<h1>Summary</h1>
<ul>
<li>Periodicity can be studied on general time series data, including multivariate time series such as video</li>
<li>Computational tricks, such as PCA, can be employed to make sliding window videos computationally tractable</li>
<li>It is even possible to pick up on quasiperiodicity/biphonation in videos without doing any tracking.</li>
</ul>
| github_jupyter |
[](https://colab.research.google.com/github/ksachdeva/rethinking-tensorflow-probability/blob/master/notebooks/12_monsters_and_mixtures.ipynb)
# Chapter 12 - Monsters and Mixtures
## Imports and utility functions
```
# Install packages that are not installed in colab
try:
import google.colab
IN_COLAB = True
except:
IN_COLAB = False
if IN_COLAB:
%tensorflow_version 2.X
print("Installing watermark & arviz ...")
!pip install -q watermark
!pip install -q arviz
%load_ext watermark
# Core
import collections
import numpy as np
import arviz as az
import pandas as pd
import tensorflow as tf
import tensorflow_probability as tfp
# visualization
import matplotlib.pyplot as plt
# aliases
tfd = tfp.distributions
tfb = tfp.bijectors
Root = tfd.JointDistributionCoroutine.Root
%watermark -p numpy,tensorflow,tensorflow_probability,arviz,scipy,pandas
# config of various plotting libraries
%config InlineBackend.figure_format = 'retina'
az.style.use('arviz-darkgrid')
```
### Tensorflow MCMC sampling helpers
```
USE_XLA = False #@param
NUMBER_OF_CHAINS = 2 #@param
NUMBER_OF_BURNIN = 500 #@param
NUMBER_OF_SAMPLES = 500 #@param
NUMBER_OF_LEAPFROG_STEPS = 4 #@param
def _trace_to_arviz(trace=None,
sample_stats=None,
observed_data=None,
prior_predictive=None,
posterior_predictive=None,
inplace=True):
if trace is not None and isinstance(trace, dict):
trace = {k: v.numpy()
for k, v in trace.items()}
if sample_stats is not None and isinstance(sample_stats, dict):
sample_stats = {k: v.numpy().T for k, v in sample_stats.items()}
if prior_predictive is not None and isinstance(prior_predictive, dict):
prior_predictive = {k: v[np.newaxis]
for k, v in prior_predictive.items()}
if posterior_predictive is not None and isinstance(posterior_predictive, dict):
if isinstance(trace, az.InferenceData) and inplace == True:
return trace + az.from_dict(posterior_predictive=posterior_predictive)
else:
trace = None
return az.from_dict(
posterior=trace,
sample_stats=sample_stats,
prior_predictive=prior_predictive,
posterior_predictive=posterior_predictive,
observed_data=observed_data,
)
@tf.function(autograph=False, experimental_compile=USE_XLA)
def run_hmc_chain(init_state,
bijectors,
step_size,
target_log_prob_fn,
num_leapfrog_steps=NUMBER_OF_LEAPFROG_STEPS,
num_samples=NUMBER_OF_SAMPLES,
burnin=NUMBER_OF_BURNIN,
):
def _trace_fn_transitioned(_, pkr):
return (
pkr.inner_results.inner_results.log_accept_ratio
)
hmc_kernel = tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn,
num_leapfrog_steps=num_leapfrog_steps,
step_size=step_size)
inner_kernel = tfp.mcmc.TransformedTransitionKernel(
inner_kernel=hmc_kernel,
bijector=bijectors)
kernel = tfp.mcmc.SimpleStepSizeAdaptation(
inner_kernel=inner_kernel,
target_accept_prob=.8,
num_adaptation_steps=int(0.8*burnin),
log_accept_prob_getter_fn=lambda pkr: pkr.inner_results.log_accept_ratio
)
results, sampler_stat = tfp.mcmc.sample_chain(
num_results=num_samples,
num_burnin_steps=burnin,
current_state=init_state,
kernel=kernel,
trace_fn=_trace_fn_transitioned)
return results, sampler_stat
def sample_posterior(jdc,
observed_data,
params,
init_state=None,
bijectors=None,
step_size = 0.1,
num_chains=NUMBER_OF_CHAINS,
num_samples=NUMBER_OF_SAMPLES,
burnin=NUMBER_OF_BURNIN):
if init_state is None:
init_state = list(jdc.sample(num_chains)[:-1])
if bijectors is None:
bijectors = [tfb.Identity() for i in init_state]
target_log_prob_fn = lambda *x: jdc.log_prob(x + observed_data)
results, sample_stats = run_hmc_chain(init_state,
bijectors,
step_size=step_size,
target_log_prob_fn=target_log_prob_fn,
num_samples=num_samples,
burnin=burnin)
stat_names = ['mean_tree_accept']
sampler_stats = dict(zip(stat_names, [sample_stats]))
transposed_results = []
for r in results:
if len(r.shape) == 2:
transposed_shape = [1,0]
else:
transposed_shape = [1,0,2]
transposed_results.append(tf.transpose(r, transposed_shape))
posterior = dict(zip(params, transposed_results))
az_trace = _trace_to_arviz(trace=posterior,
sample_stats=sampler_stats)
return posterior, az_trace
```
### Dataset URLs & Utils
```
# You could change base url to local dir or a remoate raw github content
_BASE_URL = "https://raw.githubusercontent.com/rmcelreath/rethinking/master/data"
UCBADMIT_DATASET_PATH = f"{_BASE_URL}/UCBadmit.csv"
KLINE_DATASET_PATH = f"{_BASE_URL}/Kline.csv"
# A utility method to convert data (columns) from pandas dataframe
# into tensors with appropriate type
def df_to_tensors(name, df, columns, default_type=tf.float32):
""" name : Name of the dataset
df : pandas dataframe
colums : a list of names that have the same type or
a dictionary where keys are the column names and values are the tensorflow type (e.g. tf.float32)
"""
if isinstance(columns,dict):
column_names = columns.keys()
fields = [tf.cast(df[k].values, dtype=v) for k,v in columns.items()]
else:
column_names = columns
fields = [tf.cast(df[k].values, dtype=default_type) for k in column_names]
# build the cls
tuple_cls = collections.namedtuple(name, column_names)
# build the obj
return tuple_cls._make(fields)
```
# Introduction
## 12.1 Over-dispersed counts
### 12.1.1 Beta-binomial
##### Code 12.1
A beta distribution is a probability distribution for probabilities !
```
pbar = 0.5 # mean
theta = 5 # total concentration
alpha = pbar * theta
beta = (1 - pbar) * theta
x = np.linspace(0, 1, 101)
plt.plot(x, tf.exp(tfd.Beta(alpha, beta).log_prob(x)))
plt.gca().set(xlabel="probability", ylabel="Density");
```
##### Code 12.2
```
d = pd.read_csv(UCBADMIT_DATASET_PATH, sep=";")
d["gid"] = (d["applicant.gender"] != "male").astype(int)
tdf = df_to_tensors("UCBAdmit", d, {
"gid" : tf.int32,
"applications" : tf.float32,
"admit" : tf.float32,
"reject" : tf.float32
})
def model_12_1(gid, N):
def _generator():
alpha = yield Root(tfd.Sample(tfd.Normal(loc=0., scale=1.5), sample_shape=2))
phi = yield Root(tfd.Sample(tfd.Exponential(rate=1.), sample_shape=1))
theta = phi + 2
pbar = tf.sigmoid(tf.squeeze(tf.gather(alpha, gid, axis=-1)))
# prepare the concentration vector
concentration1 = pbar * theta[...,tf.newaxis]
concentration0 = (1 - pbar) * theta[...,tf.newaxis]
concentration = tf.stack([concentration1, concentration0], axis=-1)
# outcome A i.e. admit
# since it is a multinomial we will have K = 2
# note - this does not really behave like Binomial in terms of the sample shape
A = yield tfd.Independent(tfd.DirichletMultinomial(total_count=N, concentration=concentration), reinterpreted_batch_ndims=1)
return tfd.JointDistributionCoroutine(_generator, validate_args=True)
jdc_12_1 = model_12_1(tdf.gid, tdf.applications)
```
##### Code 12.3
```
# Prepare the expected shape by the DirichletMultinomial
obs_values = tf.stack([tdf.admit, tdf.reject], axis=-1)
NUMBER_OF_CHAINS_12_1 = 2
init_state = [
tf.zeros([NUMBER_OF_CHAINS_12_1, 2]),
tf.ones([NUMBER_OF_CHAINS_12_1])
]
bijectors = [
tfb.Identity(),
tfb.Identity()
]
posterior_12_1, trace_12_1 = sample_posterior(
jdc_12_1,
num_samples=1000,
observed_data=(obs_values,),
init_state=init_state,
bijectors=bijectors,
params=['alpha', 'phi'])
# compute the difference between alphas
trace_12_1.posterior["da"] = trace_12_1.posterior["alpha"][:,:,0] - \
trace_12_1.posterior["alpha"][:,:,1]
# compute theta
trace_12_1.posterior["theta"] = trace_12_1.posterior["phi"] + 2
az.summary(trace_12_1, hdi_prob=0.89)
```
##### Code 12.4
```
# Since we have two chains and data is stored in InferenceData format
# we have to manually extract it
#
# Here I am using the data from chain 0
sample_alpha = trace_12_1.posterior["alpha"][0,:].values
sample_theta = trace_12_1.posterior["theta"][0,:].values
gid = 1
# draw posterior mean beta distribution
x = np.linspace(0, 1, 101)
pbar = tf.reduce_mean(tf.sigmoid(sample_alpha[:, gid]))
theta = tf.reduce_mean(sample_theta)
plt.plot(x, tf.exp(tfd.Beta(pbar * theta, (1 - pbar) * theta).log_prob(x)))
plt.gca().set(ylabel="Density", xlabel="probability admit", ylim=(0, 3));
# draw 50 beta distributions sampled from posterior
for i in range(50):
p = tf.sigmoid(sample_alpha[i, gid])
theta = sample_theta[i]
plt.plot(x, tf.exp(tfd.Beta(p * theta, (1 - p) * theta).log_prob(x)),
"k", alpha=0.2)
plt.title("distribution of female admission rates");
```
##### Code 12.5
```
# get samples given the posterior distribution
N = tf.cast(d.applications.values, dtype=tf.float32)
gid = d.gid.values
sample_pbar = tf.sigmoid(tf.squeeze(tf.gather(sample_alpha, gid, axis=-1)))
# need to reshape it to make it happy
st = tf.reshape(sample_theta, shape=(1000,1))
# prepare the concentration vector
concentration1 = sample_pbar * st
concentration0 = (1 - sample_pbar) * st
concentration = tf.stack([concentration1, concentration0], axis=-1)
dist = tfd.DirichletMultinomial(total_count=N, concentration=concentration)
predictive_samples = dist.sample()
# numpy style indexing magic ! .. hate it !
admit_rate = predictive_samples[::,::,0] / N
plt.scatter(range(1, 13), d.admit.values /N)
plt.errorbar(range(1, 13), np.mean(admit_rate, 0), np.std(admit_rate, 0) / 2,
fmt="o", c="k", mfc="none", ms=7, elinewidth=1)
plt.plot(range(1, 13), np.percentile(admit_rate, 5.5, 0), "k+")
plt.plot(range(1, 13), np.percentile(admit_rate, 94.5, 0), "k+");
```
In the above plot, the vertifical axis shows the predicted proportion admitted, for each case on the horizontal.
Blue points show the empirical proportion admitted on each row of data
Open circles are the posterior mean pbar and + symbols mark the 89% interval of predicted counts of admission
### 12.1.2 Negative-binomial or gamma-Poisson
##### Code 12.6 (Not working !)
Start to use **Gamma-Poisson** (also known as **Negative Binomial**) models.
Essentially Gamm-Poisson is about associating a rate to each Posisson count observation. Estimates the shape of gamma distribution to describe the Poisson rates across cases.
Gamma-Poisson also expects more variation around the mean rate.
The negative binomial distribution arises naturally from a probability experiment of performing a series of independent Bernoulli trials until the occurrence of the rth success where r is a positive integer.
```
d = pd.read_csv(KLINE_DATASET_PATH, sep=";")
d["P"] = d.population.pipe(np.log).pipe(lambda x: (x - x.mean()) / x.std())
d["cid"] = (d.contact == "high").astype(int)
d.head()
def model_12_2(cid, P):
def _generator():
alpha = yield Root(tfd.Sample(tfd.Normal(loc=1., scale=1.), sample_shape=2))
beta = yield Root(tfd.Sample(tfd.Exponential(rate=1.), sample_shape=2))
gamma = yield Root(tfd.Sample(tfd.Exponential(rate=1.), sample_shape=1))
phi = yield Root(tfd.Sample(tfd.Exponential(rate=1.), sample_shape=1))
lambda_ = tf.exp(tf.squeeze(tf.gather(alpha, cid, axis=-1))) * \
tf.math.pow(P, tf.squeeze(tf.gather(beta, cid, axis=-1))) / gamma
g_concentration = lambda_ / phi
g_rate = 1 / phi
t1 = yield tfd.Independent(
tfd.Gamma(concentration=g_concentration, rate=g_rate), reinterpreted_batch_ndims=1)
T = yield tfd.Independent(tfd.Poisson(rate=t1), reinterpreted_batch_ndims=1)
return tfd.JointDistributionCoroutine(_generator, validate_args=True)
jdc_12_2 = model_12_2(d.cid.values, tf.cast(d.P.values, dtype=tf.float32))
# Issue -
# Even prior sampling is not coming back !
# jdc_12_2.sample()
# NUMBER_OF_CHAINS_12_2 = 1
# alpha_init, beta_init, gamma_init, phi_init, t1_init, _ = jdc_12_2.sample(2)
# init_state = [
# alpha_init,
# beta_init,
# gamma_init,
# phi_init,
# t1_init
# ]
# bijectors = [
# tfb.Identity(),
# tfb.Exp(),
# tfb.Exp(),
# tfb.Exp(),
# tfb.Identity(),
# ]
# posterior_12_2, trace_12_2 = sample_posterior(jdc_12_2,
# observed_data=(d.total_tools.values,),
# init_state=init_state,
# bijectors=bijectors,
# params=['alpha', 'beta', 'gamma' 'phi', 't1'])
# az.summary(trace_12_2, credible_interval=0.89)
```
## 12.2 Zero-inflated outcomes (TODO)
## 12.3 Ordered categorical outcomes (TODO)
## 12.4 Ordered categorical predictors (TODO)
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from PIL import Image
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix, accuracy_score, recall_score, f1_score
import seaborn as sns
import torch
import torch.nn as nn
from torch.utils.data import Dataset, DataLoader
from torchvision import models, transforms, utils
import torch.optim as optim
from torch.optim import lr_scheduler
import time
import os
import zipfile
from copy import deepcopy
%matplotlib inline
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
# vision transformer
from linformer import Linformer
from vit_pytorch.efficient import ViT
torch.manual_seed(0)
dir_cat = "/home/hhg/Research/galaxyClassify/catalog/galaxyZoo/zoo2/"
file_csv = dir_cat+'gz2_all_v2.csv'
f_train = 0.64
f_valid = 0.16
f_test = 0.20
seed = 3
import pandas as pd
from sklearn.model_selection import train_test_split
def data_split(file_csv, f_train=0.64, f_valid=0.16, f_test=0.20, random_state=None, stats=False, label_tag='label_8'):
'''train-valid-test splits
Args:
file_csv (str) : path to the full catalog csv file
f_train, f_valid, f_test : fractions of training, validation, test samples
stats (bool): display splitting statistics if True
Returns:
df_train (pd.dataframes) : splitted training sample
df_valid (pd.dataframes) : validation
df_test (pd.dataframes) : test
'''
assert f_train + f_valid + f_test == 1, 'fractions have to sum to 1.'
df = pd.read_csv(file_csv)
df_train, df_temp = train_test_split(df, train_size=f_train, random_state=random_state)
relative_f_valid = f_valid/(f_valid+f_test)
df_valid, df_test = train_test_split(df_temp, train_size=relative_f_valid, random_state=random_state)
if stats:
df_stats=df.groupby([label_tag])[label_tag].agg('count').to_frame('count').reset_index()
df_stats['full'] = df_stats['count']/df_stats['count'].sum()
df_stats['train'] = df_train.groupby([label_tag]).size()/df_train.groupby([label_tag]).size().sum()
df_stats['valid'] = df_valid.groupby([label_tag]).size()/df_valid.groupby([label_tag]).size().sum()
df_stats['test'] = df_test.groupby([label_tag]).size()/df_test.groupby([label_tag]).size().sum()
ax = df_stats.plot.bar(x=label_tag, y=['full', 'train', 'valid', 'test'], rot=0)
ax.set_ylabel('class fraction')
return df_train.reset_index(drop=True), df_valid.reset_index(drop=True), df_test.reset_index(drop=True)
df_train, df_valid, df_test = data_split(file_csv, f_train, f_valid, f_test, random_state=seed, stats=True)
```
# Datasets
```
import os
from PIL import Image
from torch.utils.data import Dataset
class GalaxyZooDataset(Dataset):
'''Galaxy Zoo 2 image dataset
Args:
dataframe : pd.dataframe, outputs from the data_split function
e.g. df_train / df_valid / df_test
dir_image : str, path where galaxy images are located
label_tag : str, class label system to be used for training
e.g. label_tag = 'label_8' / 'label_3'
'''
def __init__(self, dataframe, dir_image, label_tag='label_8', transform=None):
self.df = dataframe
self.transform = transform
self.dir_image = dir_image
self.label_tag = label_tag
def __getitem__(self, index):
galaxyID = self.df.iloc[[index]].galaxyID.values[0]
file_img = os.path.join(self.dir_image, str(galaxyID) + '.jpg')
image = Image.open(file_img)
if self.transform:
image = self.transform(image)
label = self.df.iloc[[index]][self.label_tag].values[0]
return image, label, int(galaxyID)
def __len__(self):
return len(self.df)
```
# Transforms
```
def create_data_transforms(is_for_inception=False):
"""
Create Pytorch data transforms for the GalaxyZoo datasets.
Args:
is_for_inception (bool): True for inception neural networks
Outputs:
train_transform: transform for the training data
test_transform: transform for the testing data
"""
if is_for_inception:
input_size = 299
else:
input_size = 224
# transforms for training data
train_transform = transforms.Compose([transforms.CenterCrop(input_size),
transforms.RandomRotation(90),
transforms.RandomHorizontalFlip(),
transforms.RandomVerticalFlip(),
transforms.RandomResizedCrop(input_size, scale=(0.8, 1.0), ratio=(0.99, 1.01)),
transforms.ToTensor(),
transforms.Normalize([0.094, 0.0815, 0.063], [0.1303, 0.11, 0.0913])])
# transforms for validation data
valid_transform = transforms.Compose([transforms.CenterCrop(input_size),
transforms.ToTensor(),
transforms.Normalize([0.094, 0.0815, 0.063], [0.1303, 0.11, 0.0913])])
# transforms for test data
test_transform = transforms.Compose([transforms.CenterCrop(input_size),
transforms.ToTensor(),
transforms.Normalize([0.094, 0.0815, 0.063], [0.1303, 0.11, 0.0913])])
return train_transform, valid_transform, test_transform
```
# Train function
```
def train_model(model, num_epochs, criterion, optimizer, scheduler, print_every=1, early_stop_epochs=10):
"""
Train the model
Args:
model: Pytorch neural model
num_epochs: number of epochs to train
criterion: the loss function object
optimizer: the optimizer
scheduler: the learning rate decay scheduler
print_every: print the information every X epochs
early_stop_epochs: early stopping if the model doesn't improve after X epochs
"""
# cache the best model
best_model_weights = deepcopy(model.state_dict())
# best train acc
best_train_acc = 0.0
# best valid acc
best_valid_acc = 0.0
# best epoch
best_epoch = -1
# intiate dict to records the history of loss and acc
history_dic = dict()
history_dic['train_loss'] = []
history_dic['train_acc'] = []
history_dic['valid_loss'] = []
history_dic['valid_acc'] = []
history_dic['lr'] = []
for epoch in range(num_epochs):
# time of start
epoch_start_time = time.time()
"""
Train
"""
model.train()
epoch_train_cum_loss = 0.0
epoch_train_cum_corrects = 0
for images, labels, _ in train_loader:
images = images.to(device)
labels = labels.long().to(device)
optimizer.zero_grad()
pred_logits = model(images)
loss = criterion(pred_logits, labels)
_, pred_classes = torch.max(pred_logits.detach(), dim=1)
pred_classes = pred_classes.long()
epoch_train_cum_loss += loss.item() * images.size(0)
epoch_train_cum_corrects += torch.sum(pred_classes==labels.data).detach().to('cpu').item()
loss.backward()
optimizer.step()
"""
Eval
"""
model.eval()
epoch_valid_cum_loss = 0.0
epoch_valid_cum_corrects = 0
for images, labels, _ in valid_loader:
images = images.to(device)
labels = labels.long().to(device)
with torch.no_grad():
pred_logits = model(images)
_, pred_classes = torch.max(pred_logits.detach(), dim=1)
loss = criterion(pred_logits, labels)
epoch_valid_cum_loss += loss.item() * images.size(0)
epoch_valid_cum_corrects += torch.sum(pred_classes==labels.data).detach().to('cpu').item()
## Calculate metrics
train_loss = epoch_train_cum_loss / len(data_train)
train_acc = epoch_train_cum_corrects / len(data_train)
valid_loss = epoch_valid_cum_loss / len(data_valid)
valid_acc = epoch_valid_cum_corrects / len(data_valid)
# update history_dic
history_dic['train_loss'].append(train_loss)
history_dic['train_acc'].append(train_acc)
history_dic['valid_loss'].append(valid_loss)
history_dic['valid_acc'].append(valid_acc)
history_dic['lr'].append(scheduler.get_last_lr()[0])
# check if is the best acc ever
if valid_acc > best_valid_acc:
best_train_acc = train_acc
best_valid_acc = valid_acc
best_epoch = epoch + 1
# update the best model weights
best_model_weights = deepcopy(model.state_dict())
# save the best model weights to Google drive
torch.save(model.state_dict(), model_name+"_cache.pth")
epoch_end_time = time.time()
epoch_time_used = epoch_end_time - epoch_start_time
# convert epoch_time_used into mm:ss
mm = epoch_time_used // 60
ss = epoch_time_used % 60
## Print metrics
if (epoch+1) % print_every == 0:
# if is best valid acc
if epoch == (best_epoch - 1):
print("Epoch {}/{}\tTrain loss: {:.4f}\tTrain acc: {:.4f}\tValid loss: {:.4f}\tValid acc: {:.4f}\tTime: {:.0f}m {:.0f}s\t<--".format(
epoch+1, num_epochs, train_loss, train_acc, valid_loss, valid_acc, mm, ss))
# not a better model
else:
print("Epoch {}/{}\tTrain loss: {:.4f}\tTrain acc: {:.4f}\tValid loss: {:.4f}\tValid acc: {:.4f}\tTime: {:.0f}m {:.0f}s".format(
epoch+1, num_epochs, train_loss, train_acc, valid_loss, valid_acc, mm, ss))
## Early stopping
if (epoch+1) - best_epoch >= early_stop_epochs:
print("Early stopping... (Model did not improve after {} epochs)".format(early_stop_epochs))
break
scheduler.step()
# load the best weights into the model
model.load_state_dict(best_model_weights)
# print the best epoch
print("Best epoch = {}, with training accuracy = {:.4f} and validation accuracy = {:.4f}".format(best_epoch, best_train_acc, best_valid_acc))
# return the best model
return model, history_dic
```
# Create Dataloaders
```
dir_image = '/home/hhg/Research/galaxyClassify/catalog/galaxyZoo_kaggle/gz2_images/images'
# the batch size
BATCH_SIZE = 64
# create transforms
train_transform, valid_transform, test_transform = create_data_transforms(is_for_inception=False)
# create datasets
data_train = GalaxyZooDataset(df_train, dir_image, label_tag='label_8', transform=train_transform)
data_valid = GalaxyZooDataset(df_valid, dir_image, label_tag='label_8', transform=valid_transform)
data_test = GalaxyZooDataset(df_test , dir_image, label_tag='label_8', transform=test_transform)
# dataloaders
train_loader = DataLoader(data_train, batch_size=BATCH_SIZE, shuffle=True)
valid_loader = DataLoader(data_valid, batch_size=BATCH_SIZE, shuffle=True)
test_loader = DataLoader(data_test, batch_size=BATCH_SIZE, shuffle=True)
# check the sizes
print("**Dataloaders**")
print("Number of training data: {} ({} batches)".format(len(data_train), len(train_loader)))
print("Number of validation data: {} ({} batches)".format(len(data_valid), len(valid_loader)))
print("Number of test data: {} ({} batches)".format(len(data_test), len(test_loader)))
print("===============================")
df_train.groupby(['label_8']).size()
Ngals = np.array([21690., 25064., 4727., 10502., 13118., 19490., 3830., 1387.])
invNgals = 1./Ngals
invNgals /= invNgals[0]
print(invNgals)
#print(invNgals*4.6)
"""
Parameters
"""
PATCH_SIZE = 28
DEPTH = 12
HIDDEN_DIM = 128
K_DIM = 64
NUM_HEADS = 8
LR = 5e-4
STEP_SIZE = 200
GAMMA = 1
MAX_EPOCH = 200
LIN_DROPOUT = 0.1
# loss calculation for each class
class_weights = torch.FloatTensor([ 0.83, 1., 4.6, 1.77, 2.5, 3.1, 8.6, 15.2]).to(device)
#class_weights = torch.FloatTensor([ 0.9, 1., 4.6, 1.8, 2.5, 3.1, 8.5, 15.2]).to(device)
#class_weights = torch.FloatTensor([ 0.9, 1., 4.6, 1.8, 2.5, 3.1, 8.2, 15.2]).to(device)
#class_weights = torch.FloatTensor([ 1., 1., 4.6, 2.0, 2.5, 3., 8., 15.]).to(device)
#class_weights = torch.FloatTensor([ 1., 0.865, 4.589, 2.065, 1.653, 1.112, 5.663, 15.638]).to(device)
#class_weights = torch.FloatTensor([1., 1., 2., 1., 1., 1., 4., 3.]).to(device)
#class_weights = torch.FloatTensor([4.6, 4.0, 20.9, 9.6, 7.7, 5.1, 26.5, 73.3]).to(device)
#class_weights = torch.FloatTensor([1., 1., 1., 1., 1., 1., 1., 1.]).to(device)
## file name
model_name = "gz2_vit_09212021_0200"
# calculate seq_len
seq_len = int((224/PATCH_SIZE)**2) + 1
## Linformer
lin = Linformer(dim=HIDDEN_DIM, seq_len=seq_len, depth=DEPTH, k=K_DIM, heads=NUM_HEADS,
dim_head=None, one_kv_head=False, share_kv=False, reversible=False, dropout=LIN_DROPOUT)
## Vision Transformer
model = ViT(image_size=224, patch_size=PATCH_SIZE, num_classes=8, dim=HIDDEN_DIM, transformer=lin, pool='cls', channels=3)
# print out model details
print("*******[ " + model_name + " ]*******")
print("===============================")
print("patch_size = {}".format(PATCH_SIZE))
print("depth = {}".format(DEPTH))
print("hidden_dim = {}".format(HIDDEN_DIM))
print("k_dim = {}".format(K_DIM))
print("num_heads = {}".format(NUM_HEADS))
print("dropout = {}".format(LIN_DROPOUT))
print("batch_size = {}".format(BATCH_SIZE))
print("lr = {}".format(LR))
print("step_size = {}".format(STEP_SIZE))
print("gamma = {}".format(GAMMA))
print("max_epoch = {}".format(MAX_EPOCH))
print("class weights = {}".format(class_weights))
print("===============================")
print("Number of trainable parameters: {}".format(sum(param.numel() for param in model.parameters() if param.requires_grad)))
print("===============================")
```
# Train the model
```
# move to gpu
model = model.to(device)
# loss function
criterion = nn.CrossEntropyLoss(weight=class_weights)
# optimizer
optimizer = optim.Adam(model.parameters(), lr=LR)
# scheduler
scheduler = lr_scheduler.StepLR(optimizer, step_size=STEP_SIZE, gamma=GAMMA)
## train and return the best model
model, history_dic = train_model(model, MAX_EPOCH, criterion, optimizer, scheduler, print_every=1, early_stop_epochs=10)
## Save the best weights
#torch.save(model.state_dict(), model_name + '.pth')
torch.save(model.state_dict(), model_name + '.pth')
## Convert history to dataframe
history_df = pd.DataFrame(history_dic)
## Save the history
#history_df.to_csv(model_name + '_history.csv')
history_df.to_csv(model_name + '_history.csv')
```
# Plot loss and accuracy history
```
# read csv
history_df = pd.read_csv(model_name + '_history.csv')
# number of epochs
epochs = len(history_df['valid_acc'])
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(15, 5))
ax1.plot(range(epochs), history_df['train_loss'], 'b', label='train')
ax1.plot(range(epochs), history_df['valid_loss'], 'r', label='val')
ax1.set_title("Loss")
ax1.set_xlabel("Epochs")
ax1.set_ylabel("Loss")
ax1.legend()
ax2.plot(range(epochs), history_df['train_acc'], 'b', label='train')
ax2.plot(range(epochs), history_df['valid_acc'], 'r', label='val')
ax2.set_title("Accuracy")
ax2.set_xlabel("Epochs")
ax2.set_ylabel("Acc")
ax2.legend()
plt.show()
```
# Predict test data
```
def predict_model(model):
"""
Predict test data
"""
# evaluation
model.eval()
# empty lists for results
y_true = []
y_pred = []
y_label = []
for images, labels, galaxy_id in test_loader:
images = images.to(device)
labels = labels.long().to(device)
with torch.no_grad():
pred_logits = model(images)
_, pred_classes = torch.max(pred_logits.detach(), dim=1)
loss = criterion(pred_logits, labels)
y_true += torch.squeeze(labels.cpu()).tolist()
y_pred += torch.squeeze(pred_classes).tolist()
y_label += torch.squeeze(galaxy_id.cpu()).tolist()
# create a DataFrame with columns 'GalaxyID', 'class', 'predicted labels'
predict_df = pd.DataFrame(data={'GalaxyID': y_label, 'class': y_true, 'pred': y_pred})
return y_true, y_pred, predict_df
# move to gpu
model = model.to(device)
# model evaluation
y_true, y_pred, predict_df = predict_model(model)
# save predict_df
predict_df.to_csv(model_name + '_predictions.csv')
predict_df.head()
```
# Evaluation matrix
```
# galaxy classes
gxy_labels = ['Round Elliptical',
'In-between Elliptical',
'Cigar-shaped Elliptical',
'Edge-on Spiral',
'Barred Spiral',
'Unbarred Spiral',
'Irregular',
'Merger']
# confusion matrix
cm = confusion_matrix(y_true, y_pred, normalize='true')
cm_df = pd.DataFrame(cm, index=gxy_labels, columns=gxy_labels)
# accuracy of each class
for c in range(8):
print("Class {}: accuracy = {:.4f} ({})".format(c, cm[c,c]/sum(cm[c,:]), gxy_labels[c]))
print("================")
# accuracy
acc = accuracy_score(y_true, y_pred)
print("Total Accuracy = {:.4f}\n".format(acc))
# recall
recall = recall_score(y_true, y_pred, average='macro')
print("Recall = {:.4f}\n".format(recall))
# f1 score
F1 = f1_score(y_true, y_pred, average='macro')
print("F1 score = {:.4f}\n".format(F1))
# plot confusion matrix
sns.set(font_scale=1.6)
fig = plt.figure(figsize=(10, 10))
sns.heatmap(cm_df, annot=True, fmt=".1%", cmap="YlGnBu", cbar=False, annot_kws={"size": 16})
plt.show()
```
| github_jupyter |
### NivLink Demo: Multidimensional Decision-Making Task
This demo uses data from a multidimensional decision-making paradigm similar to the one described in:
> Leong, Y. C., Radulescu, A., Daniel, R., Dewoskin, V., Niv, Y. (2017). Dynamic Interaction between Reinforcement Learning and Attention in Multidimensional Environments. _Neuron_, 93(2), 451–463. http://doi.org/10.1016/j.neuron.2016.12.040
On every trial of the task, participants choose between three column stimuli which vary along three dimensions: faces, houses and tools. Upon choosing a column, they either receive a reward or not, presented as a green or red rectangle around the chosen column. After a variable delay, the features in each column reshuffle along each dimension, and the next trial begins.
```
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import scipy.io as io
import pandas as pd
import matplotlib.pyplot as plt
from pandas import DataFrame, read_csv
from nivlink import ScreenInfo
from nivlink import epoching_fht, align_to_aoi, compute_fixations
sns.set_style('white')
sns.set_context('notebook', font_scale=1.5)
%matplotlib inline
```
#### Load eye-tracking data.
Refer to [DataViewer documentation](https://github.com/nivlab/NivLink/blob/master/docs/DataViewer.Md) for details on how to generate raw text files for your experiment.
```
## Load single subject data from text file.
et_data_path = 'sample_eyetrack_data.txt'
raw_data = pd.read_table(et_data_path);
## Filter out columns of interest, convert to array and replace '.' with NaNs.
raw_data_lean = raw_data.filter(['SAMPLE_MESSAGE','RIGHT_GAZE_X','RIGHT_GAZE_Y'], axis=1)
raw_data_lean = raw_data_lean.replace('.',np.nan)
raw_data_array = raw_data_lean.values
```
#### Load behavioral data.
Here, we're loading the corresponding RTs for each trial.
```
behavior = read_csv('sample_behav_data.csv',index_col=0)
behavior.head(5)
```
#### Load pre-generated events file.
The events file is a 3-column `.mat` file with line per trial which defines:
- The run number (here, "run" refers to a continuous chunk of time during which the eye-tracker was recording)
- The trial onset relative to when the eye-tracker started recording on this run
- The trial duration
```
events_file_path = 'sample_events.mat'
events_mat = io.loadmat(events_file_path)
events = np.array(events_mat["events_array"])
```
#### Load pre-generated feature map file.
The feature map file is a n_trial by n_aoi with one line per trial which defines the task feature corresponding to each Area of Interest (AoI).
```
featmap_file_path = 'sample_featmap.mat'
featmap_mat = io.loadmat(featmap_file_path)
featmap = np.array(featmap_mat["features_aoi_map"])
```
#### Define screen parameters.
```
## Define metadata.
xdim, ydim, sfreq = 1600, 1200, 500
## Initialize ScreenInfo object.
info = ScreenInfo(xdim, ydim, sfreq, )
## Define and add areas of interest.
n_aois = 9
aois = np.empty((n_aois,2,2))
# Left column
aois[0] = np.matrix([[425, 325], [575, 475]])
aois[1] = np.matrix([[425, 525], [575, 675]])
aois[2] = np.matrix([[425, 725], [575, 875]])
# Middle column
aois[3] = np.matrix([[725, 325], [875, 475]])
aois[4] = np.matrix([[725, 525], [875, 675]])
aois[5] = np.matrix([[725, 725], [875, 875]])
# Right column
aois[6] = np.matrix([[1025, 325], [1175, 475]])
aois[7] = np.matrix([[1025, 525], [1175, 675]])
aois[8] = np.matrix([[1025, 725], [1175, 875]])
for a in range(0, n_aois):
info.add_rectangle_aoi(aois[a,0,0], aois[a,1,0], aois[a,0,1], aois[a,1,1])
## Visually inspect AoIs.
fig, ax = info.plot_aoi(1,6,False,)
```
#### Preprocess.
```
## Run epoching.
epochs = epoching_fht(raw_data_array, info, events)
## Define screenidx array.
n_trials = max(behavior.Trial)
# In this demo, AoIs are identically distributed across trials
screenidx = np.ones((n_trials,1))
## Align to AoI.
aligned = align_to_aoi(epochs, info, screenidx)
## Compute fixations.
fixations = compute_fixations(aligned, info)
## Add screen column.
fixations['Screen'] = screenidx[fixations.Trial.values.astype(int) - 1].astype(int)
## Map to task features.
fixations['Feature'] = featmap[fixations.Trial.values.astype(int) - 1,fixations.AoI.values.astype(int) - 1]
fixations.head(10)
```
#### Run some basic QA checks.
```
fixations.hist('Trial',bins=120,figsize=[8,6])
plt.xlabel('Trial',fontsize=15)
plt.ylabel('Count',fontsize=15)
plt.title('Distribution of fixations per trial',fontsize=20);
fixations.hist('AoI',bins=9,figsize=[8,6])
plt.xlabel('AoI',fontsize=15)
plt.ylabel('Count',fontsize=15)
plt.title('Distribution of AoIs',fontsize=20);
fixations.hist('Feature',bins=9,figsize=[8,6])
plt.xlabel('Feature',fontsize=15)
plt.ylabel('Count',fontsize=15)
plt.title('Distribution of Features',fontsize=20);
```
#### Visualize mapping for a single trial.
```
## Select trial.
t = 1
fixations_this_trial = fixations.loc[fixations['Trial'] == t]
[n_events, dummy] = fixations_this_trial.shape
## Plot trajectory in feature space.
plt.plot(np.arange(1, n_events+1, step=1), fixations_this_trial.Feature.values, '-o',markersize=12)
plt.xlabel('Time within trial',fontsize=15)
plt.ylabel('Feature',fontsize=15)
plt.ylim((0,10))
plt.xticks(np.arange(1, n_events+1, step=1))
plt.yticks(np.arange(1, 10, step=1))
plt.title('Fixation trajectory in feature space',fontsize=20);
```
#### Pandas Example 1: Merge fixations and behavior dataframes along trial number.
```
data = pd.merge(fixations, behavior, on ='Trial')
data.head(10)
```
#### Pandas Example 2: Remove fixations lasting less than 100ms.
```
data = data[data.Duration >= 0.10].reset_index(drop=True)
```
#### Pandas Example 3: Remove fixations after response.
```
## Drop missing responses, fast responses.
data = data.dropna()
data = data[data.RT > 0.2]
## Realign fixations to response.
data.Onset -= data.RT
data.Offset -= data.RT
## Remove fixations beginning after response.
data = data[data.Onset < 0]
data.head(10)
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# Explain binary classification model predictions
_**This notebook showcases how to use the Azure Machine Learning Interpretability SDK to explain and visualize a binary classification model predictions.**_
## Table of Contents
1. [Introduction](#Introduction)
1. [Setup](#Setup)
1. [Run model explainer locally at training time](#Explain)
1. Train a binary classification model
1. Explain the model
1. Generate global explanations
1. Generate local explanations
1. [Visualize results](#Visualize)
1. [Next steps](#Next)
## Introduction
This notebook illustrates how to explain a binary classification model predictions locally at training time without contacting any Azure services.
It demonstrates the API calls that you need to make to get the global and local explanations and a visualization dashboard that provides an interactive way of discovering patterns in data and explanations.
We will showcase three tabular data explainers: TabularExplainer (SHAP), MimicExplainer (global surrogate), and PFIExplainer.
|  |
|:--:|
| *Interpretability Toolkit Architecture* |
Problem: Breast cancer diagnosis classification with scikit-learn (run model explainer locally)
1. Train a SVM classification model using Scikit-learn
2. Run 'explain_model' globally and locally with full dataset in local mode, which doesn't contact any Azure services.
3. Visualize the global and local explanations with the visualization dashboard.
---
Setup: If you are using Jupyter notebooks, the extensions should be installed automatically with the package.
If you are using Jupyter Labs run the following command:
```
(myenv) $ jupyter labextension install @jupyter-widgets/jupyterlab-manager
```
## Explain
### Run model explainer locally at training time
```
from sklearn.datasets import load_breast_cancer
from sklearn import svm
# Explainers:
# 1. SHAP Tabular Explainer
from interpret.ext.blackbox import TabularExplainer
# OR
# 2. Mimic Explainer
from interpret.ext.blackbox import MimicExplainer
# You can use one of the following four interpretable models as a global surrogate to the black box model
from interpret.ext.glassbox import LGBMExplainableModel
from interpret.ext.glassbox import LinearExplainableModel
from interpret.ext.glassbox import SGDExplainableModel
from interpret.ext.glassbox import DecisionTreeExplainableModel
# OR
# 3. PFI Explainer
from interpret.ext.blackbox import PFIExplainer
```
### Load the breast cancer diagnosis data
```
breast_cancer_data = load_breast_cancer()
classes = breast_cancer_data.target_names.tolist()
# Split data into train and test
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(breast_cancer_data.data, breast_cancer_data.target, test_size=0.2, random_state=0)
```
### Train a SVM classification model, which you want to explain
```
clf = svm.SVC(gamma=0.001, C=100., probability=True)
model = clf.fit(x_train, y_train)
```
### Explain predictions on your local machine
```
# 1. Using SHAP TabularExplainer
explainer = TabularExplainer(model,
x_train,
features=breast_cancer_data.feature_names,
classes=classes)
# 2. Using MimicExplainer
# augment_data is optional and if true, oversamples the initialization examples to improve surrogate model accuracy to fit original model. Useful for high-dimensional data where the number of rows is less than the number of columns.
# max_num_of_augmentations is optional and defines max number of times we can increase the input data size.
# LGBMExplainableModel can be replaced with LinearExplainableModel, SGDExplainableModel, or DecisionTreeExplainableModel
# explainer = MimicExplainer(model,
# x_train,
# LGBMExplainableModel,
# augment_data=True,
# max_num_of_augmentations=10,
# features=breast_cancer_data.feature_names,
# classes=classes)
# 3. Using PFIExplainer
# Use the parameter "metric" to pass a metric name or function to evaluate the permutation.
# Note that if a metric function is provided a higher value must be better.
# Otherwise, take the negative of the function or set the parameter "is_error_metric" to True.
# Default metrics:
# F1 Score for binary classification, F1 Score with micro average for multiclass classification and
# Mean absolute error for regression
# explainer = PFIExplainer(model,
# features=breast_cancer_data.feature_names,
# classes=classes)
```
### Generate global explanations
Explain overall model predictions (global explanation)
```
# Passing in test dataset for evaluation examples - note it must be a representative sample of the original data
# x_train can be passed as well, but with more examples explanations will take longer although they may be more accurate
global_explanation = explainer.explain_global(x_test)
# Note: if you used the PFIExplainer in the previous step, use the next line of code instead
# global_explanation = explainer.explain_global(x_test, true_labels=y_test)
# Sorted SHAP values
print('ranked global importance values: {}'.format(global_explanation.get_ranked_global_values()))
# Corresponding feature names
print('ranked global importance names: {}'.format(global_explanation.get_ranked_global_names()))
# Feature ranks (based on original order of features)
print('global importance rank: {}'.format(global_explanation.global_importance_rank))
# Note: PFIExplainer does not support per class explanations
# Per class feature names
print('ranked per class feature names: {}'.format(global_explanation.get_ranked_per_class_names()))
# Per class feature importance values
print('ranked per class feature values: {}'.format(global_explanation.get_ranked_per_class_values()))
# Print out a dictionary that holds the sorted feature importance names and values
print('global importance rank: {}'.format(global_explanation.get_feature_importance_dict()))
```
### Explain overall model predictions as a collection of local (instance-level) explanations
```
# feature shap values for all features and all data points in the training data
print('local importance values: {}'.format(global_explanation.local_importance_values))
```
### Generate local explanations
Explain local data points (individual instances)
```
# Note: PFIExplainer does not support local explanations
# You can pass a specific data point or a group of data points to the explain_local function
# E.g., Explain the first data point in the test set
instance_num = 0
local_explanation = explainer.explain_local(x_test[instance_num,:])
# Get the prediction for the first member of the test set and explain why model made that prediction
prediction_value = clf.predict(x_test)[instance_num]
sorted_local_importance_values = local_explanation.get_ranked_local_values()[prediction_value]
sorted_local_importance_names = local_explanation.get_ranked_local_names()[prediction_value]
print('local importance values: {}'.format(sorted_local_importance_values))
print('local importance names: {}'.format(sorted_local_importance_names))
```
## Visualize
Load the visualization dashboard
```
from azureml.contrib.interpret.visualize import ExplanationDashboard
ExplanationDashboard(global_explanation, model, x_test)
```
## Next
Learn about other use cases of the explain package on a:
1. [Training time: regression problem](./explain-regression-local.ipynb)
1. [Training time: multiclass classification problem](./explain-multiclass-classification-local.ipynb)
1. Explain models with engineered features:
1. [Simple feature transformations](./simple-feature-transformations-explain-local.ipynb)
1. [Advanced feature transformations](./advanced-feature-transformations-explain-local.ipynb)
1. [Save model explanations via Azure Machine Learning Run History](../azure-integration/run-history/save-retrieve-explanations-run-history.ipynb)
1. [Run explainers remotely on Azure Machine Learning Compute (AMLCompute)](../azure-integration/remote-explanation/explain-model-on-amlcompute.ipynb)
1. Inferencing time: deploy a classification model and explainer:
1. [Deploy a locally-trained model and explainer](../azure-integration/scoring-time/train-explain-model-locally-and-deploy.ipynb)
1. [Deploy a remotely-trained model and explainer](../azure-integration/scoring-time/train-explain-model-on-amlcompute-and-deploy.ipynb)
| github_jupyter |
```
import pandas as pd
import collections
import matplotlib.pyplot as plt
df=pd.read_csv('../data/preprocessed/Final_uwe.csv')
df.drop(['Unnamed: 0'],inplace=True,axis=1)
#Preprocessing
for i,row in df.iterrows():
if row['Course'].startswith('CCC'):
df.drop(i,inplace=True)
for i,row in df.iterrows():
if row['Grade']=='AF' or row['Grade']=='R':
df.drop(i,inplace=True)
#Creating the dictionary from the most frequent grades entered as list.
def f(l):
d=dict()
for i in l:
d[i[0]] = i[1]
return d
def avgGrade(d):
listOfGrades = ["A","A-","B","B-","C","C-","D"]
gradesVal = range(10,3,-1)
MappedGrades = dict()
for i in range(len(listOfGrades)):
MappedGrades[listOfGrades[i]] = gradesVal[i]
MappedGrades["F"]=0
MappedGrades["W"] = 6
MappedGrades["AP"] = 10
s=0
for key,val in d.items():
s+= MappedGrades[key]*val
return s/sum(d.values())
df['Course'].unique()
df['Course'].nunique()
df['Grade'].unique()
df.drop(['Sem_year'],inplace=True,axis=1)
course_grades=df.groupby('Course')['Grade'].apply(list).to_dict()
uwe_grades=pd.DataFrame(list(course_grades.items()))
uwe_grades=uwe_grades.rename( columns={0: "Course", 1: "Grade"})
uwe_grades['grades_dict']=""
uwe_grades['5_largest']=''
uwe_grades['total_students']=''
uwe_grades['avg_grade']=''
for i,row in uwe_grades.iterrows():
uwe_grades.loc[i][2]=collections.Counter(uwe_grades.loc[i][1])
for i,row in uwe_grades.iterrows():
uwe_grades.loc[i][3]=f(collections.Counter(uwe_grades.loc[i][2]).most_common(5))
length_course_grades = {key: len(value) for key, value in course_grades.items()}
total_student_list=list(length_course_grades.values())
uwe_grades['total_students']=total_student_list
for i,row in uwe_grades.iterrows():
if row['total_students']<=5:
uwe_grades.drop(i,inplace=True)
avgGradesList = []
for i,row in uwe_grades.iterrows():
avgGradesList.append(avgGrade(uwe_grades.loc[i]["5_largest"]))
#appending the avgGrades as columns
uwe_grades["avg_grade"] = avgGradesList
# print('9-10')
AminustoA=[]
for i,row in uwe_grades.iterrows():
if row['avg_grade']>=9:
# print(row['Course'])
AminustoA.append(row['Course'])
# print('8-9')
BtoAminus=[]
for i,row in uwe_grades.iterrows():
if row['avg_grade']>=8 and row['avg_grade']<9:
# print(row['Course'])
BtoAminus.append(row['Course'])
# print('7-8')
BminustoB=[]
for i,row in uwe_grades.iterrows():
if row['avg_grade']>=7 and row['avg_grade']<8:
# print(row['Course'])
BminustoB.append(row['Course'])
# print('6-7')
CtoBminus=[]
for i,row in uwe_grades.iterrows():
if row['avg_grade']>=6 and row['avg_grade']<7:
# print(row['Course'])
CtoBminus.append(row['Course'])
# print('<6')
Below_C=[]
for i,row in uwe_grades.iterrows():
if row['avg_grade']<6:
# print(row['Course'])
Below_C.append(row['Course'])
#-------------------------------------------------------------
course_dept=df.groupby('Major')['Course'].apply(list).to_dict()
#dept wise analysis1
uwe_dept=pd.DataFrame(list(course_dept.items()))
uwe_dept=uwe_dept.rename( columns={0: "Major", 1: "Course"})
uwe_dept['dept_dict']=''
uwe_dept['10_largest']=''
for i,row in uwe_dept.iterrows():
uwe_dept.loc[i][2]=collections.Counter(uwe_dept.loc[i][1])
for i,row in uwe_dept.iterrows():
uwe_dept.loc[i][3]=f(collections.Counter(uwe_dept.loc[i][2]).most_common(10))
#Major dept wise analysis of uwes's
#for i,row in uwe_dept.iterrows(10):
uwe_df=uwe_dept.head(15)
for i,row in uwe_df.iterrows():
plt.ylabel('Number of students')
plt.title(uwe_dept['Major'].iloc[i])
plt.bar(range(len(uwe_dept['10_largest'].iloc[i])), list(uwe_dept['10_largest'].iloc[i].values()), align='center')
plt.xticks(range(len(uwe_dept['10_largest'].iloc[i])), list(uwe_dept['10_largest'].iloc[i].keys()),rotation=60)
plt.figure(figsize=(5, 3))
# UWE wise find the Major backgrounds of students who opt that UWE
course_dept2=df.groupby('Course')['Major'].apply(list).to_dict()
uwe_dept2=pd.DataFrame(list(course_dept2.items()))
uwe_dept2=uwe_dept2.rename( columns={0: "Major", 1: "Course"})
uwe_dept2['dept_dict']=''
uwe_dept2['10_largest']=''
for i,row in uwe_dept2.iterrows():
uwe_dept2.loc[i][2]=collections.Counter(uwe_dept2.loc[i][1])
for i,row in uwe_dept2.iterrows():
uwe_dept2.loc[i][3]=f(collections.Counter(uwe_dept2.loc[i][2]).most_common(10))
uwe_df2=uwe_dept2.head(15)
for i,row in uwe_df2.iterrows():
plt.ylabel('Number of students')
plt.title(uwe_dept2['Major'].iloc[i])
plt.bar(range(len(uwe_dept2['10_largest'].iloc[i])), list(uwe_dept2['10_largest'].iloc[i].values()), align='center')
plt.xticks(range(len(uwe_dept2['10_largest'].iloc[i])), list(uwe_dept2['10_largest'].iloc[i].keys()),rotation=60)
plt.figure(figsize=(5, 3))
```
# CONCLUSION
- We wanted to find out from what background, students are likely to opt for UWEs in each department, based on factors like their interests or similarity of background of UWEs with their major course.
- Like, above we observe that `BDA690` UWE which is a course offered in the department of Big Data Analytics is opted by students belonging from 2 backgrounds: `Computer Science` (majority) and `Physics`, which is quite reasonable also, as students studying `Computer Science` are ore likely to explore the field of `Big Data analytics`
- Also, we have found out, for each Major department, what all UWEs people tend to opt for. For example, the first plot shows that the students who are from `BIO` Major tend to choose UWEs like `CHY213`, `CHY122`, `CHY352`, etc which are all `Chemistry` related courses, which might indicate that they are trying to combine the degree of `Chemistry` and want to pursue it as a `Minor` along with their major `BIO`. such kinds of other patterns can also be mined from other plots above
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
plt.style.use('classic')
%matplotlib inline
```
# Class 6: Pandas and Statsmodels
## Background
In their article "The New Kaldor Facts: Ideas, Institutions, Population, and Human Capital" in the January 2010 issue of the *American Economic Journal: Macroeconomics*, Charles Jones and Paul Romer document six empirical features about macroeconomics that are yet to be adequately explained by a general theory of economic growth (link to article: [https://www.jstor.org/stable/25760291](https://www.jstor.org/stable/25760291)). Among these facts is the observation that total factor productivity is positively correlated with a country's GDP per worker.
Recall the Cobb-Douglas production function with human capital:
\begin{align}
Y & = A K^{\alpha}\left(hL\right)^{1-\alpha}, \label{eqn:production}
\end{align}
where $Y$ denotes the production of final goods and services, $K$ is the stock of physical capital, $L$ is the labor force (either number of workers or number of worker hours), $h$ is human capital per worker, and $A$ is *total factor productivity* or TFP which represents all other factors that might affect production that are not captured by $K$, $L$, or $h$.
For most countries, we can measure $Y$, $K$, $L$ and $h$. Measurements may not be perfect; for example, we often associate $h$ with years of schooling in a country even though, strictly speaking, on-the-job experience and training also contribute to human capital accumulation. Regardless, the point is that the production function *implies* a value for TFP:
\begin{align}
A & = \frac{Y}{K^{\alpha}(hL)^{1-\alpha}}
\end{align}
TFP is like a residual in a linear regression: it is the part of production that is not explained by observable inputs for production. Remarkably, Jones and Romer show that TFP explains a lot of the variation in GDP per worker across countries. This fact is captured in Figure 1.
<br>
<div align="center">
<figure style="display: table">
<img src="https://raw.githubusercontent.com/letsgoexploring/econ126/master/Img/external_fig_08_New_Kaldor_Facts_Jones_Romer_Fig4.png" width="75%" height="75%">
<figcaption style="display: table-caption; caption-side: top; text-align: center;" >Figure 1. <strong>TFP and GDP per capita across countries </strong> relative to the US. Source: Jones and Romer (2010)</figcaption>
</figure>
</div>
<br>
In this Notebook, we will replicate Figure 1 and use a linear regression procedure to estimate the relationship between TFP and GDP per worker. The data required for the exercise is named `cross_country_production.csv` and is available here: https://github.com/letsgoexploring/econ126/raw/master/Data/Csv/cross_country_production.csv
The file contains production data in 2014 for 144 countries. In addition to the name and country code, the file contains production data for 144 countries in 2014:
| Name | Description | Units |
|--------------------|---------------------------|-----------------------------|
| `gdp` | real GDP | Millions of 2001 US dollars |
| `labor` | Average number of workers | Millions |
| `physical_capital` | Physical capital stock | Millions of 2001 US dollars |
| `human_capital` | Human capital per worker | Index (unitless) |
We can use the data to compute TFP for each country in the sample and then to replicate Figure 1.
## Replicate Figure from Jones and Romer (2010)
Import the production data. Add new columns containing GDP per worker, physical capital per worker, and TFP. Construct three scatter plots:
1. GDP per worker against physical capital per worker
2. GDP per worker against human capital per worker
3. TFP against GDP per worker
```
# Import data into a DataFrame called 'df'. Use the leftmost column as the index
# Print the first five rows of the imported data
# Add a column to df called 'gdp_pw' equal to GDP divided by labor divided by 1000 so units are in thousands of $
# Add a column to df called 'physical_capital_pw' equal to physical capital divided by labor divided by 1000 so units are in thousands of $
# Set alpha to 1/3
# Add a column to df called 'tfp' equal to TFP computed using the production function
# Sort the rows of df so that the GDP per worker column is in ascending order
# Construct a scatter plot of real GDP per worker against human capital per worker with:
# 1. All countries as blue circles, size 75, opacity (alpha) 0.25, label: 'All countries'
# 2. USA as red square, size 75, opacity (alpha) 0.5, label='USA'
# 3. Add a legend in the upper left corner with scatterpoints=1
# 4. y-axis limits: [-10,250]
# 5. x-axis label: 'Human capital per worker\n(Index)'
# 6. y-axis label: 'Real GDP per worker\n(Thousands of 2001 $)'
# 7. Add a grid if you'd like
# Construct a scatter plot of real GDP per worker against physical capital per worker with:
# 1. All countries as blue circles, size 75, opacity (alpha) 0.25, label: 'All countries'
# 2. USA as red square, size 75, opacity (alpha) 0.5, label='USA'
# 3. Add a legend in the upper left corner with scatterpoints=1
# 4. y-axis limits: [-10,250]
# 5. x-axis label: 'Physical capital per worker\n(Thousands of 2001 $)'
# 6. y-axis label: 'Real GDP per worker\n(Thousands of 2001 $)'
# 7. Add a grid if you'd like
# Construct a scatter plot of TFP against real GDP per worker with:
# 1. All countries as blue circles, size 75, opacity (alpha) 0.25, label: 'All countries'
# 2. USA as red square, size 75, opacity (alpha) 0.5, label='USA'
# 3. Add a legend in the upper left corner with scatterpoints=1
# 4. x-axis limits: [-10,250]
# 4. y-axis limits: [-10,1600]
# 6. x-axis label: 'Real GDP capital per worker\n(Thousands of 2001 $)'
# 7. y-axis label: 'TFP (Unitless)'
# 8. Add a grid if you'd like
```
## Estimate Relationship Between TFP and GDP per Worker
The Statsmodels module is an excellent tool for a broad range of statistical applications. Learn more about Statsmodels here: https://www.statsmodels.org/stable/index.html. We can use the module to estimate the relationship between TFP and GDP per worker in our data.
Since the scatter plot reveals some curvature in the relationship between TFP and GDP per worker, let's specifiy the linear regression model with the variables in logs:
\begin{align}
\log \text{TFP}_i & = \beta_0 + \beta_1 \log \text{GDP}_i + \epsilon_i
\end{align}
Once we've obtained the estimated coefficients $\hat{\beta}_0$ and $\hat{\beta}_1$, we can compute predicted values for TFP $\widehat{TFP}_i$ using by setting $\epsilon_i=0$ and solving the regression equation for TFP:
\begin{align}
\widehat{TFP}_i & = \exp\left(\hat{\beta}_0 + \hat{\beta}_1 \log \text{GDP}_i\right)
\end{align}
```
# Import the Statsmodels module
# Create a variable called 'y' that will be to the dependent variable in the regression
# Create a variable called 'x' that will be to the independent variable in the regression
# Use the function sm.add_constant() to add a constant to x
# Print the first five rows of x
# Create an OLS model using Statsmodels
# Fit the OLS model to data
# Print the summary2() method of results to print summary of regression results:
```
Estimated coefficients are stored in the `params` attribute of `results`. Residuals of the regression are stored in the attribute `resid`. And predicted values of the dependent variable are stored in `fittedvalues`.
```
# Print estimated coefficients of linear regression
# Print R-squared of the regression
```
**Question**
1. What is the interpretation of the coefficient on log GDP per worker?
2. What is the R-squared of the regression? What does it mean?
**Answer**
1.
2.
```
# Print the first five rows of residuals.
# Print the mean of the residuals of the regression.
# Create a variable called 'yhat' equal to the fitted values of y:
# Print the first five rows of yhat
```
Recall that we can use estimated parameters can be used to compute out-of-sample predictions:
\begin{align}
\widehat{TFP}_i & = \exp\left(\hat{\beta}_0 + \hat{\beta}_1 \log \text{GDP}_i\right)
\end{align}
```
# Create variable called 'gdp_values' that is an array from 0.1 to 300
# Create variable called 'beta0' equal to the constant of the regression
# Create variable called 'beta1' equal to the coefficient on log GDP per worker from the regression
# Create a variable called 'ahat' that is equal to the predicted values of TFP corresponding to gdp_values
# Recreate the scatter plot of TFP against real GDP per worker with:
# 1. All features and settings from the previous plot
# 2. A plot of ahat against gdp_values; linewidth: 3, opacity: 0.5, label: 'OLS', color: magenta (c='m')
```
| github_jupyter |
# BGS Archetypes
The goal of this notebook is to derive a set of spectral archetypes from the BGS template set using Guangtun Zhu's [SetCoverPy](https://github.com/guangtunbenzhu/SetCoverPy) algorithm.
### Preliminaries.
```
import os
import numpy as np
import matplotlib.pyplot as plt
from desispec.io.util import write_bintable, makepath
from desisim.io import write_templates
from desisim.archetypes import compute_chi2, ArcheTypes
import multiprocessing
nproc = multiprocessing.cpu_count() // 2
plt.style.use('seaborn-talk')
%matplotlib inline
```
#### Initialize the random seed so the results are reproducible, below.
```
seed = 123
rand = np.random.RandomState(seed)
```
#### Output path and filenames.
```
version = 'v1.0'
outdir = os.path.join(os.getenv('DESI_ROOT'), 'spectro', 'templates', 'archetypes', 'bgs', version)
print('Setting output directory to {}'.format(outdir))
os.makedirs(outdir, exist_ok=True)
chi2file = os.path.join(outdir, 'bgs_archetypes_chi2_{}.fits'.format(version))
archfile = os.path.join(outdir, 'bgs_archetypes_{}.fits'.format(version))
```
### Read the BGS basis templates.
Read both a set of lower-resolution (1 A/pix) templates sampled over a restricted wavelength range (roughly 3500-7000 A) and the same set at higher resolution (0.2 A/pix) and over the wavelength range 0.12-2 micron. The lower-resolution templates will be used to determine the archetypes (since speed is an issue) while the full-resolution templates is what we actually write out.
In both cases we (arbitrarily) normalize every template to *r=18* and adopt a nominal velocity dispersion of 100 km/s.
```
def _build_templates(args):
"""Filler function for the multiprocessing."""
build_templates(*args)
def build_templates(bgs, input_meta, verbose=False):
flux, _, meta = bgs.make_templates(input_meta=input_meta, novdisp=True,
nocolorcuts=True, verbose=verbose)
return [flux.astype('f4'), meta]
def read_and_normalize(verbose=False, nproc=1, minwave=1200, maxwave=2e4,
cdelt=0.2, nominal_rmag=18.0, nominal_vdisp=1000,
subset=False):
"""Read and normalize the full set of basis templates.
"""
from astropy.table import vstack
from desisim.templates import BGS
from desisim.io import empty_metatable
bgs = BGS(minwave=minwave, maxwave=maxwave, cdelt=cdelt)
bgs.normline = None # no emission line
nspec = len(bgs.basemeta)
if subset:
nspec = 1000
these = rand.choice(len(bgs.basemeta), nspec)
print('Selecting a subset of {} / {} templates!'.format(nspec, len(bgs.basemeta)))
else:
these = np.arange(nspec)
input_meta = empty_metatable(nmodel=nspec, objtype='BGS')
input_meta['TEMPLATEID'] = these
input_meta['REDSHIFT'] = 0.0
input_meta['MAG'] = nominal_rmag
input_meta['VDISP'] = nominal_vdisp
input_meta['SEED'] = rand.randint(2**32, size=nspec)
# Not sure why multiprocessing isn't working in this case.
if nproc > 1:
chunk = np.array_split(these, nproc)
tempargs = list()
for ii in range(nproc):
tempargs.append((bgs, input_meta[chunk[ii]], verbose))
pool = multiprocessing.Pool(nproc)
out = pool.map(_build_templates, tempargs)
flux = np.vstack(out[0])
meta = vstack(out[1])
else:
flux, meta = build_templates(bgs, input_meta, verbose)
nspec, npix = flux.shape
print('Generated {} rest-frame BGS spectra with {} pixels.'.format(nspec, npix))
return flux, bgs.wave, meta, bgs.basemeta[these]
%time flux, wave, meta, basemeta = read_and_normalize(nproc=1, cdelt=1.0, minwave=3600, maxwave=7000, subset=True)
nspec, npix = flux.shape
%time hiresflux, hireswave, _, _ = read_and_normalize(nproc=1, cdelt=0.2, minwave=1200, maxwave=2e4, subset=True)
_, hiresnpix = hiresflux.shape
def plot_subset(nplot=25, ncol=5):
"""Plot a random sampling of the basis templates."""
nspec, npix = flux.shape
nrow = np.ceil(nplot / ncol).astype('int')
these = rand.choice(nspec, nplot, replace=False)
fig, ax = plt.subplots(nrow, ncol, figsize=(2.2*ncol, 2.2*nrow), sharey=True, sharex=True)
for thisax, indx in zip(ax.flat, these):
thisax.plot(wave, flux[indx, :])
thisax.text(0.95, 0.93, '{:0d}'.format(indx), ha='right',
va='top', transform=thisax.transAxes, fontsize=11)
thisax.xaxis.set_major_locator(plt.MaxNLocator(3))
fig.subplots_adjust(wspace=0.05, hspace=0.05)
plot_subset()
```
### Compute the NxN chi2 matrix.
We use chi2 as the "distance" matrix for the Set Cover problem.
Then, we need to determine what threshold chi2 value differentiates "different" templates.
Note that the threshold chi^2 value can be tuned until the desired number of archetypes is achieved. However, If we want the archetypes to describe each spectrum in the parent sample to a precision of prec=0.1 (10%) then we we should set chi2min to be approximately npix*prec^2.
```
def write_chi2(chi2):
from astropy.io import fits
print('Writing {}'.format(chi2file))
hdu = fits.PrimaryHDU(chi2)
hdu.writeto(chi2file, overwrite=True)
%time chi2, amp = compute_chi2(flux)
write_chi2(chi2)
prec = 0.1
chi2min_nominal = npix*prec**2
print(chi2min_nominal, np.log10(chi2min_nominal)) # seems high...
with np.errstate(divide='ignore'):
logchi2 = np.log10(chi2)
logchi2[chi2 == 0] = -1
fig, ax = plt.subplots(1, 2, figsize=(12, 4))
im = ax[0].imshow(logchi2, origin='lower', interpolation='nearest',
vmin=-1.0, cmap='viridis')
ax[0].set_xlabel('Spectrum Number')
ax[0].set_ylabel('Spectrum Number')
plt.colorbar(im, label='$log_{10}(\chi^{2})$', ax=ax[0])
_ = ax[1].hist(logchi2.reshape(nspec * nspec), bins=30, range=(-1.2, np.max(logchi2)))
ax[1].set_ylabel('Number')
ax[1].set_xlabel('$log_{10}(\chi^{2}$)')
```
### Compute and plot the number of archetypes vs chi2 threshold.
```
def narch_vs_chi2min(Arch):
"""Determine the number of archtypes vs chi2 threshold.
"""
cost = np.ones(nspec) # uniform cost
chi2min = np.logspace(1, 5, 10)
print(chi2min)
narch = np.zeros_like(chi2min)
for ii, cmin in enumerate(chi2min):
iarch = Arch.get_archetypes(chi2_thresh=cmin)
narch[ii] = len(iarch)
return narch, chi2min
def qa_narch_vs_chi2min():
fig, ax = plt.subplots()
ax.scatter(np.log10(chi2min), narch)
ax.set_xlabel('$log_{10}(\chi^{2})$ Threshold')
ax.set_ylabel('Number of Archetypes')
ax.axvline(x=np.log10(chi2min_nominal), color='red', ls='-')
ax.grid(True)
Arch = ArcheTypes(chi2)
narch, chi2min = narch_vs_chi2min(Arch)
qa_narch_vs_chi2min()
```
### Choose a chi2 threshold value then get the final set of archetypes.
```
def write_archetypes():
"""ToDo: Write out the responsibility indices for each archetype."""
from astropy.table import Column
outmeta = meta[iarch]
outmeta.add_column(Column(name='RESPONSIBILITY', length=len(iarch), dtype='int8'))
outmeta['RESPONSIBILITY'] = resp
print('Writing {}'.format(archfile))
write_templates(archfile, hiresflux[iarch, :], hireswave, outmeta, objtype='BGS Archetypes')
chi2_thresh = 10**2.5
print('Choosing a log10(chi2) threshold value of {:.1f}.'.format(np.log10(chi2_thresh)))
_iarch, _resp, _respindx = Arch.get_archetypes(chi2_thresh=chi2_thresh, responsibility=True)
print('Generated {} archetypes.'.format(len(_iarch)))
```
#### Sort by Dn(4000).
```
srt = np.argsort(meta['D4000'][_iarch])
iarch = _iarch[srt]
resp = _resp[srt]
respindx = []
for ss in srt:
respindx.append(_respindx[ss])
write_archetypes()
```
### Generate some QAplots.
```
def _markers():
d4000 = meta['D4000']
size = 110 * (1+(resp - resp.min()) / resp.ptp())
shade = (d4000[iarch] - d4000[iarch].min()) / d4000[iarch].ptp()
col = plt.cm.coolwarm(shade)
return size, col
def qa_responsibility():
"""Generate a color-color plot with the symbol size scaled by the responsibility.
"""
rz = -2.5 * np.log10(meta['FLUX_R'] / meta['FLUX_Z'])
d4000 = meta['D4000']
size, col = _markers()
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(11, 4), sharey=True)
ax1.scatter(rz[iarch], resp, c=col, marker='o', s=size,
edgecolor='k')
ax1.set_xlabel('r - z')
ax1.set_ylabel('Responsibility')
ax1.grid(True)
ax2.scatter(d4000[iarch], resp, c=col, marker='o', s=size,
edgecolor='k')
ax2.set_xlabel('$D_{n}(4000)$')
ax2.grid(True)
fig.subplots_adjust(wspace=0.05)
qa_responsibility()
def qa_colorcolor():
"""Generate a color-color plot with the symbol size scaled by the responsibility.
"""
gr = -2.5 * np.log10(meta['FLUX_G'] / meta['FLUX_R'])
rz = -2.5 * np.log10(meta['FLUX_R'] / meta['FLUX_Z'])
d4000 = meta['D4000']
size, col = _markers()
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 4))
ax1.scatter(rz, gr, s=30, c='lightgray', edgecolor='k')
ax1.scatter(rz[iarch], gr[iarch], c=col, marker='o', s=size,
alpha=1, edgecolor='k')
ax1.set_xlabel(r'$r - z$')
ax1.set_ylabel(r'$g - r$')
ax1.grid(True)
ax2.scatter(d4000, rz, s=30, c='lightgray', edgecolor='k')
ax2.scatter(d4000[iarch], rz[iarch], c=col, marker='o', s=size,
alpha=1, edgecolor='k')
ax2.set_xlabel('$D_{n}(4000)$')
ax2.set_ylabel(r'$g - r$')
ax2.grid(True)
fig.subplots_adjust(wspace=0.3)
qa_colorcolor()
def qa_archetypes(ncol=5, nfilter=11):
"""Plot the archetypes and the spectra for which they're responsible."""
from scipy.signal import medfilt
_, col = _markers()
narch = len(iarch)
nrow = np.ceil(narch / ncol).astype('int')
fig, ax = plt.subplots(nrow, ncol, figsize=(2.5*ncol, 2.5*nrow), sharey=True, sharex=True)
ww = (hireswave > 3000) * (hireswave < 1e4)
for jj, (thisax, indx, rindx, rr) in enumerate(zip(ax.flat, iarch, respindx, resp)):
if rr > 1:
for ii in rindx:
thisax.plot(hireswave[ww], hiresflux[ii, ww], color='lightgrey')
smoothflux = medfilt(hiresflux[indx, ww], 11)
thisax.plot(hireswave[ww], smoothflux, color=col[jj])
thisax.xaxis.set_major_locator(plt.MaxNLocator(2))
thisax.text(0.95, 0.93, '{:04d}\nResp={}'.format(indx, rr), ha='right',
va='top', transform=thisax.transAxes, fontsize=11)
fig.subplots_adjust(wspace=0.05, hspace=0.05)
qa_archetypes()
def qa_ages_colormag():
"""Generate a color-magnitude plot for the original AGES sample.
"""
Mr = basemeta['SDSS_UGRIZ_ABSMAG_Z01'][:, 2]
gr = basemeta['SDSS_UGRIZ_ABSMAG_Z01'][:, 1]-basemeta['SDSS_UGRIZ_ABSMAG_Z01'][:, 2]
size, col = _markers()
fig, ax = plt.subplots(figsize=(6, 4))
ax.scatter(Mr, gr, s=30, c='lightgray', edgecolor='k')
ax.scatter(Mr[iarch], gr[iarch], c=col, marker='o', s=size,
alpha=1, edgecolor='k')
ax.set_xlabel(r'$M_{0.1r}$')
ax.set_ylabel(r'$^{0.1}(g - r)$')
ax.set_xlim(-16, -23)
ax.set_ylim(0, 1.3)
ax.grid(True)
qa_ages_colormag()
```
| github_jupyter |
```
import h5py
import os
import cPickle as pickle
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import pandas as pd
import seaborn as sns
import collections
import Queue
from collections import Counter
%matplotlib inline
%load_ext autoreload
%autoreload 2
(image_metadata, book_metadata, image_to_idx) = pickle.load(open("/data/all_metadata_1M_tags.pkl", 'r'))
image_hdf5 = h5py.File('/data/image_data.hdf5','r')
ocr_hdf5 = h5py.File('/data/ocr_data.hdf5','r')
```
## Set up natural language processing
```
# coding=UTF-8
import nltk
from nltk.corpus import brown
# This is a fast and simple noun phrase extractor (based on NLTK)
# Feel free to use it, just keep a link back to this post
# http://thetokenizer.com/2013/05/09/efficient-way-to-extract-the-main-topics-of-a-sentence/
# Create by Shlomi Babluki
# May, 2013
# This is our fast Part of Speech tagger
#############################################################################
brown_train = brown.tagged_sents(categories='news')
regexp_tagger = nltk.RegexpTagger(
[(r'^-?[0-9]+(.[0-9]+)?$', 'CD'),
(r'(-|:|;)$', ':'),
(r'\'*$', 'MD'),
(r'(The|the|A|a|An|an)$', 'AT'),
(r'.*able$', 'JJ'),
(r'^[A-Z].*$', 'NNP'),
(r'.*ness$', 'NN'),
(r'.*ly$', 'RB'),
(r'.*s$', 'NNS'),
(r'.*ing$', 'VBG'),
(r'.*ed$', 'VBD'),
(r'.*', 'NN')
])
unigram_tagger = nltk.UnigramTagger(brown_train, backoff=regexp_tagger)
bigram_tagger = nltk.BigramTagger(brown_train, backoff=unigram_tagger)
#############################################################################
# This is our semi-CFG; Extend it according to your own needs
#############################################################################
cfg = {}
cfg["NNP+NNP"] = "NNP"
cfg["NN+NN"] = "NNI"
cfg["NNI+NN"] = "NNI"
cfg["JJ+JJ"] = "JJ"
cfg["JJ+NN"] = "NNI"
#############################################################################
class NPExtractor(object):
def __init__(self, sentence):
self.sentence = sentence
# Split the sentence into singlw words/tokens
def tokenize_sentence(self, sentence):
tokens = nltk.word_tokenize(sentence)
return tokens
# Normalize brown corpus' tags ("NN", "NN-PL", "NNS" > "NN")
def normalize_tags(self, tagged):
n_tagged = []
for t in tagged:
if t[1] == "NP-TL" or t[1] == "NP":
n_tagged.append((t[0], "NNP"))
continue
if t[1].endswith("-TL"):
n_tagged.append((t[0], t[1][:-3]))
continue
if t[1].endswith("S"):
n_tagged.append((t[0], t[1][:-1]))
continue
n_tagged.append((t[0], t[1]))
return n_tagged
# Extract the main topics from the sentence
def extract(self):
tokens = self.tokenize_sentence(self.sentence)
tags = self.normalize_tags(bigram_tagger.tag(tokens))
merge = True
while merge:
merge = False
for x in range(0, len(tags) - 1):
t1 = tags[x]
t2 = tags[x + 1]
key = "%s+%s" % (t1[1], t2[1])
value = cfg.get(key, '')
if value:
merge = True
tags.pop(x)
tags.pop(x)
match = "%s %s" % (t1[0], t2[0])
pos = value
tags.insert(x, (match, pos))
break
matches = []
for t in tags:
if t[1] == "NNP" or t[1] == "NNI":
#if t[1] == "NNP" or t[1] == "NNI" or t[1] == "NN":
matches.append(t[0])
return matches
```
## Do TF-IDF
we are sampling 10K images
```
def get_noun_phrases_from_img(chunk, img):
chunk = int(chunk)
img = int(img)
phrase = ' '.join(ocr_hdf5['Chunk{}'.format(chunk)][img][1:]).decode("ascii", errors="ignore")
np_extractor = NPExtractor(phrase)
multiword_res = [x.lower() for x in np_extractor.extract()]
res = []
for word in multiword_res:
res.extend(word.split(' '))
# number of items in the set divided by total length; a marker of English or not-English
pct_np = float(len(set(res)))/len(phrase)
# get the words that appear most often
#print Counter(res).most_common(10)
return pct_np, res
def get_noun_phrases_from_text(txt):
np_extractor = NPExtractor(txt)
multiword_res = [x.lower() for x in np_extractor.extract()]
res = []
for word in multiword_res:
res.extend(word.split(' '))
res = [x for x in res if len(x) > 3]
return res
sampled_images = []
sampled_text = []
for_lang = ['avec', 'sich', 'eine', 'nach', 'auch', 'nicht', 'wurde', 'alle']
for chunk in range(200):
if chunk % 50 == 0:
print(chunk)
for index in np.random.choice(range(5000), 50):
try:
ocr = ' '.join(ocr_hdf5['Chunk{}'.format(chunk)][index][1:]).decode("ascii", errors="ignore")
if ocr == ' ': continue
if any(word in ocr for word in for_lang): continue
# pct, words = get_noun_phrases_from_img(chunk, index)
# if pct > 0.03: continue
sampled_images.append((chunk, index))
sampled_text.append(ocr)
except:
pass
np.random.shuffle(sampled_text)
from sklearn.feature_extraction.text import TfidfVectorizer
#define vectorizer parameters
tfidf_vectorizer = TfidfVectorizer(max_df=0.8, max_features=200000,
min_df=0.05, stop_words='english',
use_idf=True, tokenizer=get_noun_phrases_from_text, ngram_range=(1,1))
%time tfidf_matrix = tfidf_vectorizer.fit_transform(sampled_text[:1000]) #fit the vectorizer to synopses
print(tfidf_matrix.shape)
terms = tfidf_vectorizer.get_feature_names()
terms
tfidf_matrix.
```
## K-means
```
from sklearn.cluster import KMeans
num_clusters = 5
km = KMeans(n_clusters=num_clusters)
%time km.fit(tfidf_matrix)
clusters = km.labels_.tolist()
from sklearn.externals import joblib
#uncomment the below to save your model
#since I've already run my model I am loading from the pickle
joblib.dump(km, '/data/ocr_clustering/doc_cluster.pkl')
km = joblib.load('/data/ocr_clustering/doc_cluster.pkl')
clusters = km.labels_.tolist()
from __future__ import print_function
terms = tfidf_vectorizer.get_feature_names()
print("Top terms per cluster:")
print()
#sort cluster centers by proximity to centroid
order_centroids = km.cluster_centers_.argsort()[:, ::-1]
for i in range(num_clusters):
print("Cluster %d words:" % i, end='')
for ind in order_centroids[i, :6]: #replace 6 with n words per cluster
print(' %s' % terms[ind], end=',')
print() #add whitespace
print("Number of images: {}".format(clusters.count(i)))
print() #add whitespace
print()
print()
```
## PCA
```
import sklearn
tsne = sklearn.manifold.TSNE(2)
tfidf_matrix_pca = tsne.fit_transform(tfidf_matrix.toarray())
sns.pairplot(pd.DataFrame(tfidf_matrix_pca))
```
## Hierarchical
```
from scipy.cluster.hierarchy import ward, dendrogram
from sklearn.metrics.pairwise import cosine_similarity
dist = 1 - cosine_similarity(tfidf_matrix)
linkage_matrix = ward(dist) #define the linkage_matrix using ward clustering pre-computed distances
fig, ax = plt.subplots(figsize=(5,5)) # set size
ax = dendrogram(linkage_matrix, orientation="right");
plt.tick_params(\
axis= 'x', # changes apply to the x-axis
which='both', # both major and minor ticks are affected
bottom='off', # ticks along the bottom edge are off
top='off', # ticks along the top edge are off
labelbottom='off')
plt.tight_layout() #show plot with tight layout
#uncomment below to save figure
#plt.savefig('ward_clusters.png', dpi=200) #save figure as ward_clusters
```
## Latent Dirichlet Association
```
from gensim import corpora, models, similarities
stopwords = nltk.corpus.stopwords.words('english')
%time tokenized_text = [get_noun_phrases_from_text(text) for text in sampled_text[:1000]]
%time texts = [[word for word in text if word not in stopwords] for text in tokenized_text]
%time a = [get_noun_phrases_from_text(text) for text in sampled_text[:1000]]
#create a Gensim dictionary from the texts
dictionary = corpora.Dictionary(texts)
#remove extremes (similar to the min/max df step used when creating the tf-idf matrix)
dictionary.filter_extremes(no_below=0.05, no_above=0.8)
#convert the dictionary to a bag of words corpus for reference
corpus = [dictionary.doc2bow(text) for text in texts]
%time lda = models.LdaModel(corpus, num_topics=5, id2word=dictionary, update_every=5, chunksize=500, passes=20)
lda.show_topics()
topics_matrix = lda.show_topics(formatted=False, num_words=20)
for i in topics_matrix:
print([str(word[0]) for word in i[1]][:5])
print()
```
| github_jupyter |
# Housing Regression
This notebook predicts housing prices in the Kaggle pratice competition "House Prices: Advanced Regression Techniques."
## Get Data
```
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import seaborn as sns
from scipy import stats
PATH = '/kaggle/input/house-prices-advanced-regression-techniques/'
data = pd.read_csv(PATH + 'train.csv')
test = pd.read_csv(PATH + 'test.csv')
data = data.sample(frac=1).reset_index(drop=True)
data.head()
```
## Feature Handling
```
data.drop('Id', axis = 1, inplace=True)
def combineFeatures(df):
#Total Floor area of entire house
df['TotalSF'] = df['TotalBsmtSF'] + df['1stFlrSF'] + df['2ndFlrSF']
#Total number of baths
df['TotalBath'] = df['FullBath'] + df['HalfBath'] + df['BsmtFullBath'] + df['BsmtHalfBath']
#Total porch area
df['TotalPorchSF'] = df['OpenPorchSF'] + df['3SsnPorch'] + df['EnclosedPorch'] + df['ScreenPorch'] + df['WoodDeckSF']
combineFeatures(data)
combineFeatures(test)
```
## Analyze Correlations
```
corr_matrix = data.corr()
plt.subplots(figsize=(15,15))
plt.title('Correlation Matrix for All Numeric Features')
sns.heatmap(corr_matrix, vmin=-1, vmax=1, center= 0, linewidth = 1, linecolor='black', cmap='coolwarm')
plt.subplots(figsize=(15,5))
plt.title('Correlations Of Features to SalePrice')
corr_matrix["SalePrice"].sort_values(ascending=False).plot.bar(x='Feature', y='Correlation')
```
## Analyze Distributions
```
print('SalePrice Skew : ' + str(data['SalePrice'].skew()))
print(data['SalePrice'].describe())
plt.subplots(figsize=(10,5))
sns.distplot(data['SalePrice'])
# Top ten features correlated to SalePrice
plt.subplots(figsize=(10,40))
plt.subplot(10, 1, 1)
ax = sns.distplot(data['OverallQual'])
plt.subplot(10, 1, 2)
ax = sns.distplot(data['TotalSF'])
plt.subplot(10, 1, 3)
ax = sns.distplot(data['GrLivArea'])
plt.subplot(10, 1, 4)
ax = sns.distplot(data['GarageCars'])
plt.subplot(10, 1, 5)
ax = sns.distplot(data['GarageArea'])
plt.subplot(10, 1, 6)
ax = sns.distplot(data['TotalBsmtSF'])
plt.subplot(10, 1, 7)
ax = sns.distplot(data['TotalBath'])
plt.subplot(10, 1, 8)
ax = sns.distplot(data['1stFlrSF'])
plt.subplot(10, 1, 9)
ax = sns.distplot(data['FullBath'])
plt.subplot(10, 1, 10)
ax = sns.distplot(data['TotRmsAbvGrd'])
```
## Transform Skewed Data
```
MIN_SKEW = 1
boxcox_dict = {}
data['SalePrice'] = data['SalePrice'] + 1
data['SalePrice'], boxcox_dict['SalePrice'] = stats.boxcox(data['SalePrice'] + 1)
print('SalePrice Skew : ' + str(data['SalePrice'].skew()))
print(data['SalePrice'].describe())
plt.subplots(figsize=(10,5))
sns.distplot(data['SalePrice'])
# Split features into types based on how we will preprocess them
# Refer to data_description.txt for more info on features
# Numeric features
# Note: The description is incorrect Kitchen and Bedroom are labeled KitchenAbvGr and BedroomAbvGr
num_features = ["LotFrontage", "LotArea", "YearBuilt", "OverallQual", "OverallCond", "MasVnrArea", "BsmtFinSF1", "BsmtFinSF2", "BsmtUnfSF", "TotalBsmtSF", "1stFlrSF", "2ndFlrSF",
"LowQualFinSF", "GrLivArea", "BsmtFullBath", "BsmtHalfBath", "FullBath", "HalfBath", "BedroomAbvGr", "KitchenAbvGr", "TotRmsAbvGrd", "Fireplaces", "GarageYrBlt",
"GarageCars", "GarageArea", "WoodDeckSF", "OpenPorchSF", "EnclosedPorch", "3SsnPorch", "ScreenPorch", "PoolArea", "MiscVal", "YrSold","YearRemodAdd"]
# Categoric features that have simple, (a>b>c), correlations and can be ordinally encoded, NOTE: Be careful about order
cat_features_ordcorr = ["ExterQual", "ExterCond", "BsmtQual", "BsmtCond", "BsmtExposure", "BsmtFinType1", "BsmtFinType2", "HeatingQC", "KitchenQual", "Functional", "FireplaceQu",
"GarageFinish", "GarageQual", "GarageCond", "PavedDrive", "PoolQC", "Fence", "LotShape", "LandContour", "LandSlope"]
# Categoric features that likely don't have any correlation for use with one hot encoders
cat_features_uncorr = ["Street", "Alley", "RoofStyle", "RoofMatl", "Exterior1st", "Exterior2nd", "MasVnrType", "Foundation", "Heating", "Electrical", "CentralAir", "GarageType",
"MiscFeature", "SaleType", "SaleCondition", "MSSubClass", "MSZoning", "Utilities", "LotConfig", "Neighborhood", "Condition1", "Condition2", "BldgType", "HouseStyle"]
# Features that may require different preprocessing steps than the above
# Dropping MoSold, month sold
odd_features = ["MoSold"]
data.drop(odd_features, axis=1, inplace=True)
# MSSubClass is a number but it's a categoric feature
data['MSSubClass'] = data['MSSubClass'].apply(str)
for feature in num_features:
data[feature].fillna(data[feature].median(), inplace = True)
test[feature].fillna(data[feature].median(), inplace = True)
data_skews = data[num_features].skew(axis = 0, skipna = True).sort_values(ascending=False)
data_skews
for feature, skew in zip(data_skews.index, data_skews.values):
if skew >= MIN_SKEW or skew <= -(MIN_SKEW):
data[feature], boxcox_dict[feature] = stats.boxcox(data[feature] + 1)
test[feature] = stats.boxcox(test[feature] + 1, lmbda = boxcox_dict[feature])
data.skew(axis = 0, skipna = True).sort_values(ascending=False)
## Keep in mind that all box cox transformations have the +1 shift parameter
```
## Data Preprocessing
```
from sklearn.preprocessing import OrdinalEncoder
from sklearn.preprocessing import MinMaxScaler
target_scaler = MinMaxScaler()
target_data = target_scaler.fit_transform(data['SalePrice'].to_numpy().reshape(-1, 1))
scaler = MinMaxScaler()
scaler.fit(data[num_features])
num_data = scaler.transform(data[num_features])
num_test = scaler.transform(test[num_features])
# For some reason the description lists 'NA' as a category but they're actual NaNs not the string 'NA'
for feature in cat_features_uncorr:
data[feature].fillna('NA', inplace = True)
test[feature].fillna('NA', inplace = True)
uncorr_data = pd.get_dummies(data=data[cat_features_uncorr])
uncorr_test = pd.get_dummies(data=test[cat_features_uncorr])
# Remove categories we haven't seen in the training data
uncorr_test = uncorr_test.reindex(columns = uncorr_data.columns, fill_value=0)
# Same as uncorr categoric features
for feature in cat_features_ordcorr:
# These ones have NaNs but they don't have an NA category
if feature in ['KitchenQual', 'Functional']:
data[feature].fillna(data[feature].mode()[0], inplace = True)
test[feature].fillna(test[feature].mode()[0], inplace = True)
else:
data[feature].fillna('NA', inplace = True)
test[feature].fillna('NA', inplace = True)
# From the description.txt file
feature_cat_pairs = [('ExterQual', ['Po', 'Fa', 'TA', 'Gd', 'Ex']),
('ExterCond', ['Po', 'Fa', 'TA', 'Gd', 'Ex']),
('BsmtQual', ['NA', 'Po', 'Fa', 'TA', 'Gd', 'Ex']),
('BsmtCond', ['NA', 'Po', 'Fa', 'TA', 'Gd', 'Ex']),
('BsmtExposure', ['NA', 'No', 'Mn', 'Av', 'Gd']),
('BsmtFinType1', ['NA', 'Unf', 'LwQ', 'Rec', 'BLQ', 'ALQ', 'GLQ']),
('BsmtFinType2', ['NA', 'Unf', 'LwQ', 'Rec', 'BLQ', 'ALQ', 'GLQ']),
('HeatingQC', ['Po', 'Fa', 'TA', 'Gd', 'Ex']),
('KitchenQual', ['Po', 'Fa', 'TA', 'Gd', 'Ex']),
('Functional', ['Sal', 'Sev', 'Maj2', 'Maj1', 'Mod', 'Min2', 'Min1', 'Typ']),
('FireplaceQu', ['NA', 'Po', 'Fa', 'TA', 'Gd', 'Ex']),
('GarageFinish', ['NA', 'Unf', 'RFn', 'Fin']),
('GarageQual', ['NA', 'Po', 'Fa', 'TA', 'Gd', 'Ex']),
('GarageCond', ['NA', 'Po', 'Fa', 'TA', 'Gd', 'Ex']),
('PavedDrive', ['N', 'P', 'Y']),
('PoolQC', ['NA', 'Po', 'Fa', 'TA', 'Gd', 'Ex']),
('Fence', ['NA', 'MnWw', 'GdWo', 'MnPrv', 'GdPrv']),
('LotShape', ['IR3', 'IR2', 'IR1', 'Reg']),
('LandContour', ['Low', 'HLS', 'Bnk', 'Lvl']),
('LandSlope', ['Sev', 'Mod', 'Gtl'])
]
scaler = MinMaxScaler()
ordcorr_data = np.zeros((len(data), len(feature_cat_pairs)))
ordcorr_test = np.zeros((len(test), len(feature_cat_pairs)))
for i, pair in enumerate(feature_cat_pairs):
enc = OrdinalEncoder(categories=[pair[1]])
transformed_data = scaler.fit_transform(enc.fit_transform(data[pair[0]].to_numpy().reshape(-1, 1)))
transformed_test = scaler.transform(enc.transform(test[pair[0]].to_numpy().reshape(-1, 1)))
# Probably a better way to do this
for j, value in enumerate(transformed_data):
ordcorr_data[j][i] = value
for j, value in enumerate(transformed_test):
ordcorr_test[j][i] = value
```
## Models
```
from sklearn.metrics import mean_squared_error
from sklearn import linear_model
from sklearn import ensemble
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeRegressor
from sklearn.experimental import enable_hist_gradient_boosting
from sklearn.ensemble import HistGradientBoostingRegressor
from xgboost import XGBRegressor
from sklearn.ensemble import VotingRegressor
def showPredInfo(train, train_pred, test, test_pred):
print('Training RMSE : ' + str(np.sqrt(mean_squared_error(train, train_pred))))
print('Validation RMSE : ' + str(np.sqrt(mean_squared_error(test, test_pred))))
plt.subplots(figsize=(5,10))
ax = plt.subplot(2, 1, 1)
ax.plot([0,1], [0,1], color='cyan')
ax.plot(train, train_pred, 'o', color='green')
ax = plt.subplot(2, 1, 2)
ax.plot([0,1], [0,1], color='cyan')
ax.plot(test, test_pred, 'o', color='green')
plt.show()
test_data = np.hstack((num_test, uncorr_test.to_numpy(), ordcorr_test))
full_data = np.hstack((num_data, uncorr_data.to_numpy(), ordcorr_data))
X_train, X_test, y_train, y_test = train_test_split(full_data, target_data, test_size=0.28, random_state=8)
y_train = np.ravel(y_train)
```
### Ridge Regression
```
ridge = linear_model.Ridge(alpha=1.0)
ridge.fit(X_train, y_train)
ridge_train_pred = ridge.predict(X_train)
ridge_test_pred = ridge.predict(X_test)
showPredInfo(y_train, ridge_train_pred, y_test, ridge_test_pred)
```
### Lasso
```
lasso = linear_model.LassoCV(cv=5)
lasso.fit(X_train, y_train)
lasso_train_pred = lasso.predict(X_train)
lasso_test_pred = lasso.predict(X_test)
showPredInfo(y_train, lasso_train_pred, y_test, lasso_test_pred)
```
### Elastic Net
```
elastic = linear_model.ElasticNetCV(cv=5)
elastic.fit(X_train, y_train)
elastic_train_pred = elastic.predict(X_train)
elastic_test_pred = elastic.predict(X_test)
showPredInfo(y_train, elastic_train_pred, y_test, elastic_test_pred)
```
### Lars
```
lars = linear_model.Lars(eps=1.2)
lars.fit(X_train, y_train)
lars_train_pred = lars.predict(X_train)
lars_test_pred = lars.predict(X_test)
showPredInfo(y_train, lars_train_pred, y_test, lars_test_pred)
```
### Bayesion Ridge
```
bRidge = linear_model.BayesianRidge()
bRidge.fit(X_train, y_train)
bRidge_train_pred = bRidge.predict(X_train)
bRidge_test_pred = bRidge.predict(X_test)
showPredInfo(y_train, bRidge_train_pred, y_test, bRidge_test_pred)
```
### ARD
```
ard = linear_model.ARDRegression()
ard.fit(X_train, y_train)
ard_train_pred = ard.predict(X_train)
ard_test_pred = ard.predict(X_test)
showPredInfo(y_train, ard_train_pred, y_test, ard_test_pred)
```
### OMP
```
omp = linear_model.OrthogonalMatchingPursuitCV(cv=5)
omp.fit(X_train, y_train)
omp_train_pred = omp.predict(X_train)
omp_test_pred = omp.predict(X_test)
showPredInfo(y_train, omp_train_pred, y_test, omp_test_pred)
```
### Random Forest
```
forest = ensemble.RandomForestRegressor(max_depth=10, n_estimators=1000, random_state=8)
forest.fit(X_train, y_train)
forest_train_pred = forest.predict(X_train)
forest_test_pred = forest.predict(X_test)
showPredInfo(y_train, forest_train_pred, y_test, forest_test_pred)
```
### Extra Trees
```
extra = ensemble.ExtraTreesRegressor(max_depth=10, n_estimators=1000, random_state=8)
extra.fit(X_train, y_train)
extra_train_pred = extra.predict(X_train)
extra_test_pred = extra.predict(X_test)
showPredInfo(y_train, extra_train_pred, y_test, extra_test_pred)
```
### Adaptive Boosting
```
ada = ensemble.AdaBoostRegressor(base_estimator=DecisionTreeRegressor(max_depth=12),
n_estimators=1000, random_state=8)
ada.fit(X_train, y_train)
ada_train_pred = ada.predict(X_train)
ada_test_pred = ada.predict(X_test)
showPredInfo(y_train, ada_train_pred, y_test, ada_test_pred)
```
### Gradient Boosting
```
gbc = ensemble.GradientBoostingRegressor(n_estimators=1000, learning_rate=0.3, max_depth=1, random_state=8)
gbc.fit(X_train, y_train)
gbc_train_pred = gbc.predict(X_train)
gbc_test_pred = gbc.predict(X_test)
showPredInfo(y_train, gbc_train_pred, y_test, gbc_test_pred)
```
### Histogram Gradient Boosting
```
hgb = HistGradientBoostingRegressor(learning_rate=0.1)
hgb.fit(X_train, y_train)
hgb_train_pred = hgb.predict(X_train)
hgb_test_pred = hgb.predict(X_test)
showPredInfo(y_train, hgb_train_pred, y_test, hgb_test_pred)
```
### XGBoost
```
xgb = XGBRegressor(n_estimators=1000, learning_rate=0.2, max_depth=1, random_state=8)
xgb.fit(X_train, y_train)
xgb_train_pred = xgb.predict(X_train)
xgb_test_pred = xgb.predict(X_test)
showPredInfo(y_train, xgb_train_pred, y_test, xgb_test_pred)
plt.subplots(figsize=(5,10))
ax = plt.subplot(2, 1, 1)
ax.plot([0,1], [0,1], color='cyan')
plt.plot(y_train, ridge_train_pred, 'o', color=(0.0, 0.0, 0.0), alpha=0.2)
plt.plot(y_train, lasso_train_pred, 'o', color=(0.4, 0.0, 0.0), alpha=0.2)
plt.plot(y_train, elastic_train_pred, 'o', color=(0.0, 0.4, 0.0), alpha=0.2)
plt.plot(y_train, lars_train_pred, 'o', color=(0.0, 0.0, 0.4), alpha=0.2)
plt.plot(y_train, bRidge_train_pred, 'o', color=(0.4, 0.4, 0.0), alpha=0.2)
plt.plot(y_train, ard_train_pred, 'o', color=(0.4, 0.0, 0.4), alpha=0.2)
plt.plot(y_train, omp_train_pred, 'o', color=(0.0, 0.4, 0.4), alpha=0.2)
plt.plot(y_train, forest_train_pred, 'o', color=(0.4, 0.4, 0.4), alpha=0.2)
plt.plot(y_train, extra_train_pred, 'o', color=(0.8, 0.0, 0.0), alpha=0.2)
plt.plot(y_train, ada_train_pred, 'o', color=(0.0, 0.8, 0.0), alpha=0.2)
plt.plot(y_train, gbc_train_pred, 'o', color=(0.0, 0.0, 0.8), alpha=0.2)
plt.plot(y_train, hgb_train_pred, 'o', color=(0.8, 0.8, 0.0), alpha=0.2)
plt.plot(y_train, xgb_train_pred, 'o', color=(0.0, 0.8, 0.8), alpha=0.2)
ax = plt.subplot(2, 1, 2)
ax.plot([0,1], [0,1], color='cyan')
plt.plot(y_test, ridge_test_pred, 'o', color=(0.0, 0.0, 0.0), alpha=0.2)
plt.plot(y_test, lasso_test_pred, 'o', color=(0.4, 0.0, 0.0), alpha=0.2)
plt.plot(y_test, elastic_test_pred, 'o', color=(0.0, 0.4, 0.0), alpha=0.2)
plt.plot(y_test, lars_test_pred, 'o', color=(0.0, 0.0, 0.4), alpha=0.2)
plt.plot(y_test, bRidge_test_pred, 'o', color=(0.4, 0.4, 0.0), alpha=0.2)
plt.plot(y_test, ard_test_pred, 'o', color=(0.4, 0.0, 0.4), alpha=0.2)
plt.plot(y_test, omp_test_pred, 'o', color=(0.0, 0.4, 0.4), alpha=0.2)
plt.plot(y_test, forest_test_pred, 'o', color=(0.4, 0.4, 0.4), alpha=0.2)
plt.plot(y_test, extra_test_pred, 'o', color=(0.8, 0.0, 0.0), alpha=0.2)
plt.plot(y_test, ada_test_pred, 'o', color=(0.0, 0.8, 0.0), alpha=0.2)
plt.plot(y_test, gbc_test_pred, 'o', color=(0.0, 0.0, 0.8), alpha=0.2)
plt.plot(y_test, hgb_test_pred, 'o', color=(0.8, 0.8, 0.0), alpha=0.2)
plt.plot(y_test, xgb_test_pred, 'o', color=(0.0, 0.8, 0.8), alpha=0.2)
plt.show()
```
### Voting Regressor
```
# (bRidge, 'bRidge'), doesn't like this one for some reason
vote = VotingRegressor([('ridge', ridge), ('lasso', lasso), ('elastic', elastic),
('lars', lars), ('ard', ard), ('omp', omp), ('gbc', gbc),
('forest', forest), ('extra', extra), ('ada', ada),
('hgb', hgb), ('xgb', xgb)])
vote.fit(X_train, y_train)
vote_train_pred = vote.predict(X_train)
vote_test_pred = vote.predict(X_test)
showPredInfo(y_train, vote_train_pred, y_test, vote_test_pred)
```
## Submission
```
from scipy.special import inv_boxcox
vote.fit(full_data, np.ravel(target_data))
sub_pred = vote.predict(test_data)
sub_pred = target_scaler.inverse_transform(sub_pred.reshape(-1, 1))
sub_pred = inv_boxcox(sub_pred, boxcox_dict['SalePrice']) + 1
submission = pd.DataFrame({'Id' : test['Id'], 'SalePrice' : np.ravel(sub_pred)})
submission.to_csv('submission.csv', index=False)
```
| github_jupyter |
## Imports and Libaries
```
import cv2
import os
import numpy as np
import matplotlib.pyplot as plt
import random
```
## Data preperation and pre-processing
```
!unzip Face\ Image\ Dataset_2020.zip -d ./face_dataset
folder_path = 'face_dataset/'
train_data = []
test_date = []
X_train = []
Y_train = []
X_test = []
Y_test = []
train_ratio = 0.6
unprocessed_img_dim = None
for folder in os.listdir(folder_path):
if os.path.isfile(folder_path + folder):
continue
D = []
for file in os.listdir( folder_path + folder + '/'):
img = cv2.imread( folder_path + folder + '/' + file)
img = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
if unprocessed_img_dim is None:
unprocessed_img_dim = img.shape
img = img.flatten()
D.append(img)
random.shuffle(D)
train_boundary = train_ratio * len(D)
label = folder
for i in range(len(D)):
if i < train_boundary:
X_train.append(D[i])
Y_train.append(label)
else:
X_test.append(D[i])
Y_test.append(label)
X_train = np.asarray(X_train)
Y_train = np.asarray(Y_train)
X_test = np.asarray(X_test)
Y_test = np.asarray(Y_test)
```
## a) PCA Implementation and analysis
```
class PCA(object):
match_threshold = 40000000
def __init__(self, X_train, Y_train, k = 100):
self.k = k
self.X_train = X_train
self.Y_train = Y_train
self.mean_image = None
self.weights = None
self.eigen_faces = None
self.W = None
self.fit(X_train)
def fit(self, X_train):
self.mean_image = np.mean(X_train, axis = 0)
P = len(X_train)
self.W = (X_train - self.mean_image)
C = np.matmul(self.W , self.W.T) / (P - 1)
eigen_vectors, _, _ = np.linalg.svd(C)
final_eigen_vectors = eigen_vectors[:,:self.k]
self.eigen_faces = np.matmul(final_eigen_vectors.T, self.W )
self.weights = np.matmul( self.W , self.eigen_faces.T)
def predict(self, X):
X_centred = (X - self.mean_image)
weight_img = np.matmul( X_centred, self.eigen_faces.T)
min_dist = 1e19
ind = 0
for i in range( len(self.weights) ):
dist = np.linalg.norm(weight_img - self.weights[i])
if dist < min_dist:
min_dist, ind = dist, i
if min_dist >= PCA.match_threshold:
return "Not Enrolled"
return self.Y_train[ind]
def getAccuracy(self, X_test, Y_test):
freq = 0
n = len(X_test)
for i in range(n):
if self.predict(X_test[i]) == Y_test[i]:
freq += 1
return freq * 100 / n
def displayMeanImage(self):
plt.imshow(self.mean_image.reshape(unprocessed_img_dim), cmap = 'gray')
plt.title('Mean Image')
k = 100
PCA_model = PCA(X_train, Y_train, k)
accuracy = PCA_model.getAccuracy(X_test, Y_test)
print(f'Accuracy : {round(accuracy, 2)}%')
```
### Visualizations and Plots
#### k vs Accuracy plot
```
k_list = [10, 20, 30, 40, 50, 60, 70, 80, 90, 100]
acc_list = []
for k in k_list:
PCA_model = PCA(X_train, Y_train, k)
accuracy = PCA_model.getAccuracy(X_test, Y_test)
acc_list.append(accuracy)
plt.plot(k_list, acc_list)
plt.title('K vs Accuracy Plot')
plt.xlabel('K')
plt.ylabel('Accuracy')
plt.show()
```
#### Input data before centering
```
PCA_model.displayMeanImage()
fig, ax = plt.subplots(nrows = 2, ncols = 3)
sub_data_ind_list = random.sample(range(0, len(X_train)), 6)
ind = 0
for i, a in enumerate(ax.ravel()):
img = X_train[sub_data_ind_list[ind]]
a.imshow(img.reshape(unprocessed_img_dim), cmap='gray')
ind += 1
```
#### Input data after centering
```
fig, ax = plt.subplots(nrows = 2, ncols = 3)
ind = 0
for i, a in enumerate(ax.ravel()):
img = PCA_model.W[sub_data_ind_list[ind]]
a.imshow(img.reshape(unprocessed_img_dim), cmap='gray')
ind += 1
```
#### Examples of correctly classified and mis-classified images
```
sub_data_ind_list = random.sample(range(0, len(X_test)), 6)
for ind in sub_data_ind_list:
fig,ax = plt.subplots(nrows = 1, ncols = 1)
ax.set_title('Actual Label: ' + Y_test[ind] + ', Predicted Label: ' + PCA_model.predict(X_test[ind]) )
ax.imshow(X_test[ind].reshape(unprocessed_img_dim), cmap='gray')
```
## b) Add imposters into the test set and then recognize it as the not enrolled person.
```
img = cv2.imread('lena_img.png')
img = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
img = img.flatten()
plt.imshow(img.reshape(unprocessed_img_dim), cmap = 'gray')
print(f'Class label for the above image : {PCA_model.predict(img)}')
```
| github_jupyter |
# Multi-Layer Perceptron, MNIST
---
In this notebook, we will train an MLP to classify images from the [MNIST database](http://yann.lecun.com/exdb/mnist/) hand-written digit database.
The process will be broken down into the following steps:
>1. Load and visualize the data
2. Define a neural network
3. Train the model
4. Evaluate the performance of our trained model on a test dataset!
Before we begin, we have to import the necessary libraries for working with data and PyTorch.
```
# import libraries
import torch
import numpy as np
```
---
## Load and Visualize the [Data](http://pytorch.org/docs/stable/torchvision/datasets.html)
Downloading may take a few moments, and you should see your progress as the data is loading. You may also choose to change the `batch_size` if you want to load more data at a time.
This cell will create DataLoaders for each of our datasets.
```
from torchvision import datasets
import torchvision.transforms as transforms
# number of subprocesses to use for data loading
num_workers = 0
# how many samples per batch to load
batch_size = 20
# convert data to torch.FloatTensor
transform = transforms.ToTensor()
# choose the training and test datasets
train_data = datasets.MNIST(root='data', train=True,
download=True, transform=transform)
test_data = datasets.MNIST(root='data', train=False,
download=True, transform=transform)
# prepare data loaders
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
num_workers=num_workers)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=batch_size,
num_workers=num_workers)
```
### Visualize a Batch of Training Data
The first step in a classification task is to take a look at the data, make sure it is loaded in correctly, then make any initial observations about patterns in that data.
```
import matplotlib.pyplot as plt
%matplotlib inline
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy()
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
# print out the correct label for each image
# .item() gets the value contained in a Tensor
ax.set_title(str(labels[idx].item()))
```
### View an Image in More Detail
```
img = np.squeeze(images[1])
fig = plt.figure(figsize = (12,12))
ax = fig.add_subplot(111)
ax.imshow(img, cmap='gray')
width, height = img.shape
thresh = img.max()/2.5
for x in range(width):
for y in range(height):
val = round(img[x][y],2) if img[x][y] !=0 else 0
ax.annotate(str(val), xy=(y,x),
horizontalalignment='center',
verticalalignment='center',
color='white' if img[x][y]<thresh else 'black')
```
---
## Define the Network [Architecture](http://pytorch.org/docs/stable/nn.html)
The architecture will be responsible for seeing as input a 784-dim Tensor of pixel values for each image, and producing a Tensor of length 10 (our number of classes) that indicates the class scores for an input image. This particular example uses two hidden layers and dropout to avoid overfitting.
```
import torch.nn as nn
import torch.nn.functional as F
## TODO: Define the NN architecture
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.fc1 = nn.Linear(28 * 28, 256)
self.fc2 = nn.Linear(256, 128)
self.fc3 = nn.Linear(128, 10)
def forward(self, x):
# flatten image input
x = x.flatten(start_dim=1)
# add hidden layer, with relu activation function
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = F.log_softmax(self.fc3(x))
return x
# initialize the NN
model = Net()
print(model)
```
### Specify [Loss Function](http://pytorch.org/docs/stable/nn.html#loss-functions) and [Optimizer](http://pytorch.org/docs/stable/optim.html)
It's recommended that you use cross-entropy loss for classification. If you look at the documentation (linked above), you can see that PyTorch's cross entropy function applies a softmax funtion to the output layer *and* then calculates the log loss.
```
## TODO: Specify loss and optimization functions
import torch.optim as optim
# specify loss function
criterion = nn.NLLLoss()
# specify optimizer
optimizer = optim.SGD(model.parameters(), lr=0.01, momentum=0.9)
```
---
## Train the Network
The steps for training/learning from a batch of data are described in the comments below:
1. Clear the gradients of all optimized variables
2. Forward pass: compute predicted outputs by passing inputs to the model
3. Calculate the loss
4. Backward pass: compute gradient of the loss with respect to model parameters
5. Perform a single optimization step (parameter update)
6. Update average training loss
The following loop trains for 30 epochs; feel free to change this number. For now, we suggest somewhere between 20-50 epochs. As you train, take a look at how the values for the training loss decrease over time. We want it to decrease while also avoiding overfitting the training data.
```
# number of epochs to train the model
n_epochs = 30 # suggest training between 20-50 epochs
model.train() # prep model for training
for epoch in range(n_epochs):
# monitor training loss
train_loss = 0.0
###################
# train the model #
###################
for data, target in train_loader:
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update running training loss
train_loss += loss.item()*data.size(0)
# print training statistics
# calculate average loss over an epoch
train_loss = train_loss/len(train_loader.sampler)
print('Epoch: {} \tTraining Loss: {:.6f}'.format(
epoch+1,
train_loss
))
```
---
## Test the Trained Network
Finally, we test our best model on previously unseen **test data** and evaluate it's performance. Testing on unseen data is a good way to check that our model generalizes well. It may also be useful to be granular in this analysis and take a look at how this model performs on each class as well as looking at its overall loss and accuracy.
#### `model.eval()`
`model.eval(`) will set all the layers in your model to evaluation mode. This affects layers like dropout layers that turn "off" nodes during training with some probability, but should allow every node to be "on" for evaluation!
```
# initialize lists to monitor test loss and accuracy
test_loss = 0.0
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
model.eval() # prep model for *evaluation*
for data, target in test_loader:
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# update test loss
test_loss += loss.item()*data.size(0)
# convert output probabilities to predicted class
_, pred = torch.max(output, 1)
# compare predictions to true label
correct = np.squeeze(pred.eq(target.data.view_as(pred)))
# calculate test accuracy for each object class
for i in range(len(target)):
label = target.data[i]
class_correct[label] += correct[i].item()
class_total[label] += 1
# calculate and print avg test loss
test_loss = test_loss/len(test_loader.sampler)
print('Test Loss: {:.6f}\n'.format(test_loss))
for i in range(10):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
str(i), 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
```
### Visualize Sample Test Results
This cell displays test images and their labels in this format: `predicted (ground-truth)`. The text will be green for accurately classified examples and red for incorrect predictions.
```
# obtain one batch of test images
dataiter = iter(test_loader)
images, labels = dataiter.next()
# get sample outputs
output = model(images)
# convert output probabilities to predicted class
_, preds = torch.max(output, 1)
# prep images for display
images = images.numpy()
# plot the images in the batch, along with predicted and true labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
ax.set_title("{} ({})".format(str(preds[idx].item()), str(labels[idx].item())),
color=("green" if preds[idx]==labels[idx] else "red"))
```
| github_jupyter |
# PDBe API Training
This interactive Python notebook will guide you through various ways of programmatically accessing Protein Data Bank in Europe (PDBe) data using REST API
The REST API is a programmatic way to obtain information from the PDB and EMDB. You can access details about:
* sample
* experiment
* models
* compounds
* cross-references
* publications
* quality
* assemblies
and more...
For more information, visit https://www.ebi.ac.uk/pdbe/pdbe-rest-api
# Notebook #7
This notebook is the seventh in the training material series, and focuses on searching for sequences in the PDBe search API
## 1) Making imports and setting variables
First, we import some packages that we will use, and set some variables.
Note: Full list of valid URLs is available from https://www.ebi.ac.uk/pdbe/api/doc/
```
from pprint import pprint # used for pretty printing
import requests # used to get data from the a URL
import pandas as pd # used to analyse the results
search_url = "https://www.ebi.ac.uk/pdbe/search/pdb/select?" # the rest of the URL used for PDBe's search API.
```
We will now define some functions which will use to get the data from PDBe's search API
```
def make_request_post(search_dict, number_of_rows=10):
"""
makes a post request to the PDBe API
:param dict search_dict: the terms used to search
:param number_of_rows: number or rows to return - initially limited to 10
:return dict: response JSON
"""
# make sure we get the number of rows we need
if 'rows' not in search_dict:
search_dict['rows'] = number_of_rows
# set the return type to JSON
search_dict['wt'] = 'json'
# do the query
response = requests.post(search_url, data=search_dict)
if response.status_code == 200:
return response.json()
else:
print("[No data retrieved - %s] %s" % (response.status_code, response.text))
return {}
def format_sequence_search_terms(sequence, filter_terms=None):
"""
Format parameters for a sequence search
:param str sequence: one letter sequence
:param lst filter_terms: Terms to filter the results by
:return str: search string
"""
# first we set the parameters which we will pass to PDBe's search
params = {
'json.nl': 'map',
'start': '0',
'sort': 'fasta(e_value) asc',
'xjoin_fasta': 'true',
'bf': 'fasta(percentIdentity)',
'xjoin_fasta.external.expupperlim': '0.1',
'xjoin_fasta.external.sequence': sequence,
'q': '*:*',
'fq': '{!xjoin}xjoin_fasta'
}
# we make sure that we add required filter terms if they aren't present
if filter_terms:
for term in ['pdb_id', 'entity_id', 'entry_entity', 'chain_id']:
filter_terms.append(term)
filter_terms = list(set(filter_terms))
params['fl'] = ','.join(filter_terms)
# returns the parameter dictionary
return params
def run_sequence_search(sequence, filter_terms=None, number_of_rows=10):
"""
Runs a sequence search and results the results
:param str sequence: sequence in one letter code
:param lst filter_terms: terms to filter the results by
:param int number_of_rows: number of results to return
:return lst: List of results
"""
search_dict = format_sequence_search_terms(sequence=sequence, filter_terms=filter_terms)
response = make_request_post(search_dict=search_dict, number_of_rows=number_of_rows)
results = response.get('response', {}).get('docs', [])
print('Number of results {}'.format(len(results)))
# we now have to go through the FASTA results and join them with the main results
raw_fasta_results = response.get('xjoin_fasta').get('external')
fasta_results = {} # results from FASTA will be stored here - key'd by PDB ID and Chain ID
# go through each FASTA result and get the E value, percentage identity and sequence from the result
for fasta_row in raw_fasta_results:
# join_id = fasta_row.get('joinId')
fasta_doc = fasta_row.get('doc', {})
percent_identity = fasta_doc.get('percent_identity')
e_value = fasta_doc.get('e_value')
return_sequence = fasta_row.get('return_sequence_string')
pdb_id_chain = fasta_doc.get('pdb_id_chain').split('_')
pdb_id = pdb_id_chain[0].lower()
chain_id = pdb_id_chain[-1]
join_id = '{}_{}'.format(pdb_id, chain_id)
fasta_results[join_id] = {'e_value': e_value,
'percentage_identity': percent_identity,
'return_sequence': return_sequence}
# now we go through the main results and add the FASTA results
ret = [] # final results will be stored here.
for row in results:
pdb_id = row.get('pdb_id').lower()
chain_ids = row.get('chain_id')
for chain_id in chain_ids:
search_id = '{}_{}'.format(pdb_id, chain_id)
entry_fasta_results = fasta_results.get(search_id, {})
# we will only keep results that match the search ID
if entry_fasta_results:
row['e_value'] = entry_fasta_results.get('e_value')
row['percentage_identity'] = entry_fasta_results.get('percentage_identity')
row['result_sequence'] = entry_fasta_results.get('return_sequence_string')
ret.append(row)
return ret
```
We will search for a sequence with an example sequence from UniProt P24941 -
Cyclin-dependent kinase 2
```
sequence_to_search = """
MENFQKVEKIGEGTYGVVYKARNKLTGEVVALKKIRLDTETEGVPSTAIREISLLKELNH
PNIVKLLDVIHTENKLYLVFEFLHQDLKKFMDASALTGIPLPLIKSYLFQLLQGLAFCHS
HRVLHRDLKPQNLLINTEGAIKLADFGLARAFGVPVRTYTHEVVTLWYRAPEILLGCKYY
STAVDIWSLGCIFAEMVTRRALFPGDSEIDQLFRIFRTLGTPDEVVWPGVTSMPDYKPSF
PKWARQDFSKVVPPLDEDGRSLLSQMLHYDPNKRISAKAALAHPFFQDVTKPVPHLRL"""
filter_list = ['pfam_accession', 'pdb_id', 'molecule_name', 'ec_number',
'uniprot_accession_best', 'tax_id']
first_results = run_sequence_search(sequence_to_search, filter_terms=filter_list)
```
Print the first result to see what we have
```
pprint(first_results[0])
```
Notice that some of the results are lists
Before we do any further analysis we should get a few more results so we can see some patterns.
We are going to increase the number of results to 1000
```
first_results = run_sequence_search(sequence_to_search,
filter_terms=filter_list,
number_of_rows=1000
)
```
Load the results into a Pandas Dataframe so we can query them
Before we do this we have to do a bit of house keeping. We are going to change the lists (results with [] around them)
into comma separated values
```
def change_lists_to_strings(results):
"""
updates lists to strings for loading into Pandas
:param dict results: dictionary of results to process
:return dict: dictionary of results
"""
for row in results:
for data in row:
if type(row[data]) == list:
# if there are any numbers in the list change them into strings
row[data] = [str(a) for a in row[data]]
# unique and sort the list and then change the list into a string
row[data] = ','.join(sorted(list(set(row[data]))))
return results
def pandas_dataset(list_of_results):
results = change_lists_to_strings(list_of_results) # we have added our function to change lists to strings
df = pd.DataFrame(results)
return df
df = pandas_dataset(first_results)
```
Lets see what we have - you'll see it looks a bit like a spreadsheet or a database
```
print(df.head())
```
We can save the results to a CSV file which we can load into excel
```
df.to_csv("search_results.csv")
```
There isn't a cut off of eValue or percentage identity in our search
so we should look what the values go to
we can select the column and find the minimum value with .min() or maximum value with .max()
```
df['percentage_identity'].max()
df['percentage_identity'].min()
```
same for e value - here we want the min and max
```
df['e_value'].min()
df['e_value'].max()
```
We can see that percentage identity drops to as low as 36%
Lets say we want to restrict it to 50%
```
df2 = df.query('percentage_identity > 50')
```
We stored the results in a new Dataframe called "df2"
```
df2.head()
```
Number of entries in the Dataframe
```
len(df2)
```
Max value of percentage identity
```
df2['percentage_identity'].max()
```
Min value of percentage identity
```
df2['percentage_identity'].min()
```
How many unique Pfam domains or UniProts did we get back?
We can group the results by Pfam using "groupby" and then counting the results
```
df.groupby('pfam_accession').count()
```
same for uniprot accession
This time we will sort the values by the number of PDB entries ("pdb_id"'s) they appear in.
```
group_by_uniprot = df.groupby('uniprot_accession_best').count().sort_values('pdb_id', ascending=False)
group_by_uniprot
```
In this case the most common UniProt accession is P24941.
How many UniProt accessions were there?
```
len(group_by_uniprot)
```
How many are enzymes? We can use "ec_number" to work see how many have E.C. numbers
```
uniprot_with_ec = group_by_uniprot.query('ec_number != 0')
len(uniprot_with_ec)
```
| github_jupyter |
## Training
# Example Predictor: Linear Rollout Predictor
This example contains basic functionality for training and evaluating a linear predictor that rolls out predictions day-by-day.
First, a training data set is created from historical case and npi data.
Second, a linear model is trained to predict future cases from prior case data along with prior and future npi data.
The model is an off-the-shelf sklearn Lasso model, that uses a positive weight constraint to enforce the assumption that increased npis has a negative correlation with future cases.
Third, a sample evaluation set is created, and the predictor is applied to this evaluation set to produce prediction results in the correct format.
```
import pickle
import numpy as np
import pandas as pd
from sklearn.linear_model import Lasso
from sklearn.model_selection import train_test_split
```
### Copy the data locally
```
# Main source for the training data
DATA_URL = 'https://raw.githubusercontent.com/OxCGRT/covid-policy-tracker/master/data/OxCGRT_latest.csv'
# Local file
DATA_FILE = 'data/OxCGRT_latest.csv'
import os
import urllib.request
if not os.path.exists('data'):
os.mkdir('data')
# urllib.request.urlretrieve(DATA_URL, DATA_FILE)
# Load historical data from local file
df = pd.read_csv(DATA_FILE,
parse_dates=['Date'],
encoding="ISO-8859-1",
dtype={"RegionName": str,
"RegionCode": str},
error_bad_lines=False)
df.columns
# For testing, restrict training data to that before a hypothetical predictor submission date
HYPOTHETICAL_SUBMISSION_DATE = np.datetime64("2020-07-31")
df = df[df.Date <= HYPOTHETICAL_SUBMISSION_DATE]
# Add RegionID column that combines CountryName and RegionName for easier manipulation of data
# GeoID 作为地区的唯一识别号,用于后续的分组中
df['GeoID'] = df['CountryName'] + '__' + df['RegionName'].astype(str)
# Add new cases column
# ConfirmedCases 确诊病例 NewCases 新增病例
df['NewCases'] = df.groupby('GeoID').ConfirmedCases.diff().fillna(0)
# Keep only columns of interest
id_cols = ['CountryName',
'RegionName',
'GeoID',
'Date']
cases_col = ['NewCases']
npi_cols = ['C1_School closing',
'C2_Workplace closing',
'C3_Cancel public events',
'C4_Restrictions on gatherings',
'C5_Close public transport',
'C6_Stay at home requirements',
'C7_Restrictions on internal movement',
'C8_International travel controls',
'H1_Public information campaigns',
'H2_Testing policy',
'H3_Contact tracing',
'H6_Facial Coverings']
# 12 列为干预措施
print(len(npi_cols))
df = df[id_cols + cases_col + npi_cols]
# Fill any missing case values by interpolation and setting NaNs to 0
df.update(df.groupby('GeoID').NewCases.apply(
lambda group: group.interpolate()).fillna(0))
# Fill any missing NPIs by assuming they are the same as previous day
for npi_col in npi_cols:
df.update(df.groupby('GeoID')[npi_col].ffill().fillna(0))
df
# Set number of past days to use to make predictions
# 回看的天数,可以自行设置
nb_lookback_days = 30
# Create training data across all countries for predicting one day ahead
# 用过去一个月的数据预测当天的值
X_cols = cases_col + npi_cols
y_col = cases_col
X_samples = []
y_samples = []
geo_ids = df.GeoID.unique()
for g in geo_ids:
# 筛选出特定区域的数据
gdf = df[df.GeoID == g]
all_case_data = np.array(gdf[cases_col])
all_npi_data = np.array(gdf[npi_cols])
# Create one sample for each day where we have enough data
# Each sample consists of cases and npis for previous nb_lookback_days
# 每个样本包括过去一个月病例和干预措施,相当于当天的预测只参考于过去一个月的数据, K阶马尔可夫
nb_total_days = len(gdf)
for d in range(nb_lookback_days, nb_total_days - 1):
X_cases = all_case_data[d-nb_lookback_days:d]
# Take negative of npis to support positive
# weight constraint in Lasso.
# 取负值,是为了让权重变成正的
X_npis = -all_npi_data[d - nb_lookback_days:d]
# X_cases 是一个矩阵,需要拉平为一个向量
# Flatten all input data so it fits Lasso input format.
X_sample = np.concatenate([X_cases.flatten(),
X_npis.flatten()])
y_sample = all_case_data[d + 1]
X_samples.append(X_sample)
y_samples.append(y_sample)
X_samples = np.array(X_samples)
y_samples = np.array(y_samples).flatten()
#30 * 1 + 30 *12 前面30列为过去每天的新增病例 后面360列为过去30天的干预措施逐天叠加在一起
print(X_samples, X_samples.shape)
print(y_samples)
# Helpful function to compute mae
def mae(pred, true):
return np.mean(np.abs(pred - true))
# Split data into train and test sets
X_train, X_test, y_train, y_test = train_test_split(X_samples,
y_samples,
test_size=0.2,
random_state=301)
# Create and train Lasso model.
# Set positive=True to enforce assumption that cases are positively correlated
# with future cases and npis are negatively correlated.
model = Lasso(alpha=0.1,
precompute=True,
max_iter=10000,
positive=True,
selection='random')
# Fit model
model.fit(X_train, y_train)
# Evaluate model
train_preds = model.predict(X_train)
train_preds = np.maximum(train_preds, 0) # Don't predict negative cases
print('Train MAE:', mae(train_preds, y_train))
test_preds = model.predict(X_test)
test_preds = np.maximum(test_preds, 0) # Don't predict negative cases
print('Test MAE:', mae(test_preds, y_test))
# Inspect the learned feature coefficients for the model
# to see what features it's paying attention to.
# Give names to the features
x_col_names = []
for d in range(-nb_lookback_days, 0):
x_col_names.append('Day ' + str(d) + ' ' + cases_col[0])
for d in range(-nb_lookback_days, 1):
for col_name in npi_cols:
x_col_names.append('Day ' + str(d) + ' ' + col_name)
# View non-zero coefficients
for (col, coeff) in zip(x_col_names, list(model.coef_)):
if coeff != 0.:
print(col, coeff)
print('Intercept', model.intercept_)
# Save model to file
if not os.path.exists('models'):
os.mkdir('models')
with open('models/model.pkl', 'wb') as model_file:
pickle.dump(model, model_file)
```
## Evaluation
Now that the predictor has been trained and saved, this section contains the functionality for evaluating it on sample evaluation data.
```
# Reload the module to get the latest changes
import predict
from importlib import reload
reload(predict)
from predict import predict_df
# 会碰到历史数据很少而不能满足模型需求的情况(冷启动)
%%time
preds_df = predict_df("2020-08-01", "2020-08-31", path_to_ips_file="../../../validation/data/2020-09-30_historical_ip.csv", verbose=True)
# Check the predictions
preds_df.head()
```
# Validation
This is how the predictor is going to be called during the competition.
!!! PLEASE DO NOT CHANGE THE API !!!
```
!python predict.py -s 2020-08-01 -e 2020-08-04 -ip ../../../validation/data/2020-09-30_historical_ip.csv -o predictions/2020-08-01_2020-08-04.csv
!head predictions/2020-08-01_2020-08-04.csv
```
# Test cases
We can generate a prediction file. Let's validate a few cases...
```
import os
from covid_xprize.validation.predictor_validation import validate_submission
def validate(start_date, end_date, ip_file, output_file):
# First, delete any potential old file
try:
os.remove(output_file)
except OSError:
pass
# Then generate the prediction, calling the official API
!python predict.py -s {start_date} -e {end_date} -ip {ip_file} -o {output_file}
# And validate it
errors = validate_submission(start_date, end_date, ip_file, output_file)
if errors:
for error in errors:
print(error)
else:
print("All good!")
```
## 4 days, no gap
- All countries and regions
- Official number of cases is known up to start_date
- Intervention Plans are the official ones
```
validate(start_date="2020-08-01",
end_date="2020-08-04",
ip_file="../../../validation/data/2020-09-30_historical_ip.csv",
output_file="predictions/val_4_days.csv")
```
## 1 month in the future
- 2 countries only
- there's a gap between date of last known number of cases and start_date
- For future dates, Intervention Plans contains scenarios for which predictions are requested to answer the question: what will happen if we apply these plans?
```
%%time
validate(start_date="2021-01-01",
end_date="2021-01-31",
ip_file="../../../validation/data/future_ip.csv",
output_file="predictions/val_1_month_future.csv")
```
## 180 days, from a future date, all countries and regions
- Prediction start date is 1 week from now. (i.e. assuming submission date is 1 week from now)
- Prediction end date is 6 months after start date.
- Prediction is requested for all available countries and regions.
- Intervention plan scenario: freeze last known intervention plans for each country and region.
As the number of cases is not known yet between today and start date, but the model relies on them, the model has to predict them in order to use them.
This test is the most demanding test. It should take less than 1 hour to generate the prediction file.
### Generate the scenario
```
from datetime import datetime, timedelta
start_date = datetime.now() + timedelta(days=7)
start_date_str = start_date.strftime('%Y-%m-%d')
end_date = start_date + timedelta(days=180)
end_date_str = end_date.strftime('%Y-%m-%d')
print(f"Start date: {start_date_str}")
print(f"End date: {end_date_str}")
from covid_xprize.validation.scenario_generator import get_raw_data, generate_scenario, NPI_COLUMNS
DATA_FILE = 'data/OxCGRT_latest.csv'
latest_df = get_raw_data(DATA_FILE, latest=True)
scenario_df = generate_scenario(start_date_str, end_date_str, latest_df, countries=None, scenario="Freeze")
scenario_file = "predictions/180_days_future_scenario.csv"
scenario_df.to_csv(scenario_file, index=False)
print(f"Saved scenario to {scenario_file}")
```
### Check it
```
%%time
validate(start_date=start_date_str,
end_date=end_date_str,
ip_file=scenario_file,
output_file="predictions/val_6_month_future.csv")
```
| github_jupyter |
# Self-Driving Car Engineer Nanodegree
### Deep Learning
### Project: Build a Traffic Sign Recognition Classifier
The neural networks in this project reference the `LeNet-5` implementation with some layers and parameters fine tuned. The best validation result is **97.8%** and the final test result is **95.4%**.
---
## Step 0: Load The Data
```
# Load pickled data
import pickle
training_file = '../traffic-signs-data/train.p'
validation_file= '../traffic-signs-data/valid.p'
testing_file = '../traffic-signs-data/test.p'
with open(training_file, mode='rb') as f:
train = pickle.load(f)
with open(validation_file, mode='rb') as f:
valid = pickle.load(f)
with open(testing_file, mode='rb') as f:
test = pickle.load(f)
X_train, y_train = train['features'], train['labels']
X_valid, y_valid = valid['features'], valid['labels']
X_test, y_test = test['features'], test['labels']
# Mapping ClassID to traffic sign names
import csv
signs = []
with open('signnames.csv', 'r') as csvfile:
signnames = csv.reader(csvfile, delimiter=',')
next(signnames,None)
for row in signnames:
signs.append(row[1])
csvfile.close()
```
---
## Step 1: Dataset Summary & Exploration
The pickled data is a dictionary with 4 key/value pairs:
- `'features'` is a 4D array containing raw pixel data of the traffic sign images, (num examples, width, height, channels).
- `'labels'` is a 1D array containing the label/class id of the traffic sign. The file `signnames.csv` contains id -> name mappings for each id.
- `'sizes'` is a list containing tuples, (width, height) representing the original width and height the image.
- `'coords'` is a list containing tuples, (x1, y1, x2, y2) representing coordinates of a bounding box around the sign in the image.
**THESE COORDINATES ASSUME THE ORIGINAL IMAGE. THE PICKLED DATA CONTAINS RESIZED VERSIONS (32 by 32) OF THESE IMAGES**
```
# Number of training examples
n_train = X_train.shape[0]
# Number of validation examples
n_validation = X_valid.shape[0]
# Number of testing examples.
n_test = X_test.shape[0]
# The shape of an traffic sign image
image_shape = train['features'].shape[1:]
# Number of unique classes/labels there are in the dataset.
n_classes = len(set(train['labels']))
print("Number of training examples =", n_train)
print("Number of testing examples =", n_test)
print("Image data shape =", image_shape)
print("Number of classes =", n_classes)
```
### Include an exploratory visualization of the dataset
Visualize the German Traffic Signs Dataset using the pickled file(s).
```
# Show an image selected randomly
import random
import numpy as np
import matplotlib.pyplot as plt
# Visualizations will be shown in the notebook.
%matplotlib inline
index_im = random.randint(0, len(X_train))
image = X_train[index_im].squeeze()
plt.figure(figsize = (2,2))
plt.imshow(image)
print('data index = ' + str(index_im))
print('label number = ' + str(y_train[index_im]))
print('Image label = ' + str(signs[y_train[index_im]]))
# plotting the count of each sign in training data
sign_count = []
for index in range(43):
sign_count.append(list(y_train).count(index))
plt.figure(figsize=(12, 4))
plt.bar(range(len(sign_count)), sign_count)
plt.xlabel('sign-label-number')
plt.title('The count of each sign in training data')
plt.show()
```
----
## Step 2: Design and Test a Model Architecture
Design and implement a deep learning model that learns to recognize traffic signs.
### Pre-process the Data Set (normalization, grayscale, etc.)
```
# Converting to grayscale
import cv2
def conv_gray(images):
images_gray = []
for image in images:
# image = cv2.cvtColor(image,cv2.COLOR_HSV2RGB)
image_gray = np.array(cv2.cvtColor(image,cv2.COLOR_RGB2GRAY))
image_gray.resize(32, 32, 1)
images_gray.append(image_gray)
return np.array(images_gray)
X_train = conv_gray(X_train)
X_valid = conv_gray(X_valid)
X_test = conv_gray(X_test)
# Plotting the gray image
image_gray = X_train[index_im].squeeze()
plt.figure(figsize = (5,5))
ax1 = plt.subplot(1,2,1)
ax2 = plt.subplot(1,2,2)
plt.sca(ax1)
plt.xlabel('full color image')
plt.imshow(image)
plt.sca(ax2)
plt.xlabel('grayscale image')
plt.imshow(image_gray, cmap='gray')
plt.show()
# Image normalize
X_train = (X_train / 255.0)
X_valid = (X_valid / 255.0)
X_test = (X_test / 255.0)
#Shuffle the training data.
from sklearn.utils import shuffle
X_train, y_train = shuffle(X_train, y_train)
```
### Model Architecture
```
import tensorflow as tf
EPOCHS = 50
BATCH_SIZE = 128
from tensorflow.contrib.layers import flatten
def LeNet(x):
# Arguments used for tf.truncated_normal, randomly defines variables for the weights and biases for each layer
mu = 0
sigma = 0.1
# Layer 1: Convolutional. Input = 32x32x1. Output = 28x28x48. Pooling. Input = 28x28x48. Output = 14x14x48.
conv1_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 1, 48), mean = mu, stddev = sigma))
conv1_b = tf.Variable(tf.zeros(48))
conv1 = tf.nn.conv2d(x, conv1_W, strides=[1, 1, 1, 1], padding='VALID') + conv1_b
conv1 = tf.nn.relu(conv1)
conv1 = tf.nn.max_pool(conv1, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
# Layer 2: Convolutional. Output = 10x10x96. Pooling. Input = 10x10x96. Output = 5x5x96.
conv2_W = tf.Variable(tf.truncated_normal(shape=(5, 5, 48, 96), mean = mu, stddev = sigma))
conv2_b = tf.Variable(tf.zeros(96))
conv2 = tf.nn.conv2d(conv1, conv2_W, strides=[1, 1, 1, 1], padding='VALID') + conv2_b
conv2 = tf.nn.relu(conv2)
conv2 = tf.nn.max_pool(conv2, ksize=[1, 2, 2, 1], strides=[1, 2, 2, 1], padding='VALID')
# Layer 3: Convolutional. Output = 3x3x172. Pooling. Input = 3x3x172. Output = 2x2x172.
conv3_W = tf.Variable(tf.truncated_normal(shape=(3, 3, 96, 172), mean = mu, stddev = sigma))
conv3_b = tf.Variable(tf.zeros(172))
conv3 = tf.nn.conv2d(conv2, conv3_W, strides=[1, 1, 1, 1], padding='VALID') + conv3_b
conv3 = tf.nn.relu(conv3)
conv3 = tf.nn.max_pool(conv3, ksize=[1, 2, 2, 1], strides=[1, 1, 1, 1], padding='VALID')
# Flatten. Input = 2x2x172. Output = 688.
fc0 = flatten(conv3)
# SOLUTION: Layer 4: Fully Connected. Input = 688. Output = 240.
fc1_W = tf.Variable(tf.truncated_normal(shape=(688, 240), mean = mu, stddev = sigma))
fc1_b = tf.Variable(tf.zeros(240))
fc1 = tf.matmul(fc0, fc1_W) + fc1_b
# SOLUTION: Activation.
fc1 = tf.nn.relu(fc1)
# SOLUTION: Layer 5: Fully Connected. Input = 240. Output = 84.
fc2_W = tf.Variable(tf.truncated_normal(shape=(240, 84), mean = mu, stddev = sigma))
fc2_b = tf.Variable(tf.zeros(84))
fc2 = tf.matmul(fc1, fc2_W) + fc2_b
# SOLUTION: Activation.
fc2 = tf.nn.relu(fc2)
# SOLUTION: Layer 6: Fully Connected. Input = 84. Output = 43.
fc3_W = tf.Variable(tf.truncated_normal(shape=(84, 43), mean = mu, stddev = sigma))
fc3_b = tf.Variable(tf.zeros(43))
logits = tf.matmul(fc2, fc3_W) + fc3_b
return logits
#Features and Labels
x = tf.placeholder(tf.float32, (None, 32, 32, 1))
y = tf.placeholder(tf.int32, (None))
one_hot_y = tf.one_hot(y, 43)
#Training Pipeline
rate = 0.001
logits = LeNet(x)
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(labels=one_hot_y, logits=logits)
loss_operation = tf.reduce_mean(cross_entropy)
optimizer = tf.train.AdamOptimizer(learning_rate = rate)
training_operation = optimizer.minimize(loss_operation)
#Model Evaluation
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1))
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
saver = tf.train.Saver()
def evaluate(X_data, y_data):
num_examples = len(X_data)
total_accuracy = 0
sess = tf.get_default_session()
for offset in range(0, num_examples, BATCH_SIZE):
batch_x, batch_y = X_data[offset:offset+BATCH_SIZE], y_data[offset:offset+BATCH_SIZE]
accuracy = sess.run(accuracy_operation, feed_dict={x: batch_x, y: batch_y})
total_accuracy += (accuracy * len(batch_x))
return total_accuracy / num_examples
```
### Train, Validate and Test the Model
A validation set can be used to assess how well the model is performing. A low accuracy on the training and validation
sets imply underfitting. A high accuracy on the training set but low accuracy on the validation set implies overfitting.
```
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
num_examples = len(X_train)
print("Training...")
print()
for i in range(EPOCHS):
X_train, y_train = shuffle(X_train, y_train)
for offset in range(0, num_examples, BATCH_SIZE):
end = offset + BATCH_SIZE
batch_x, batch_y = X_train[offset:end], y_train[offset:end]
sess.run(training_operation, feed_dict={x: batch_x, y: batch_y})
validation_accuracy = evaluate(X_valid, y_valid)
print("EPOCH {} : ".format(i+1) + "Validation Accuracy = {:.3f}".format(validation_accuracy))
saver.save(sess, './lenet')
print("Model saved")
#Evaluate the Model
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
test_accuracy = evaluate(X_test, y_test)
print("Test Accuracy = {:.3f}".format(test_accuracy))
```
---
## Step 3: Test a Model on New Images
To give yourself more insight into how your model is working, download at least five pictures of German traffic signs from the web and use your model to predict the traffic sign type.
You may find `signnames.csv` useful as it contains mappings from the class id (integer) to the actual sign name.
### Load and Output the Images
```
# Load 5 new test images
import os
new_test_images = []
path = '../traffic-signs-data/new_test_images/'
print('The new test images are :')
for image in os.listdir(path):
print(image)
img = cv2.imread(path + image)
img = cv2.resize(img, (32,32))
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
new_test_images.append(img)
# The new test images' labels
new_test_signs = [40, 20, 4, 18, 14]
# Plotting the new test images
plt.figure(figsize=(10, 10))
for i in range(5):
plt.subplot(2, 5, i+1)
plt.imshow(new_test_images[i])
plt.xlabel(signs[new_test_signs[i]])
plt.xticks([])
plt.yticks([])
plt.tight_layout(pad=0, h_pad=0, w_pad=0)
plt.show()
```
### Predict the Sign Type for Each Image
```
# Pre-process the new test images
new_test_gray = conv_gray(new_test_images)
new_test_norm = new_test_gray / 255.0
# Predict the Sign Type for Each Image
def new_test(X_data, sess):
pred_sign = sess.run(tf.argmax(logits, 1), feed_dict={x: X_data, y: 1.0})
return pred_sign
with tf.Session() as sess:
saver.restore(sess, './lenet')
signs_classes=new_test(new_test_norm, sess)
print(str(new_test_signs) + ' <== Actural Sign')
print(str(list(signs_classes)) + ' <== Predict Sign')
plt.figure(figsize=(20, 20))
for i in range(5):
plt.subplot(3, 5, i+1)
plt.imshow(new_test_images[i])
plt.title(signs[signs_classes[i]], color = 'blue')
plt.axis('off')
plt.show()
```
### Analyze Performance
```
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
test_accuracy = evaluate(new_test_norm, new_test_signs)
print("Test Accuracy = {:.3f}".format(test_accuracy))
```
### Output Top 5 Softmax Probabilities For Each Image Found on the Web
```
def new_test_top(X_data, sess):
prob = sess.run(tf.nn.softmax(logits), feed_dict={x: X_data, y: 1.0})
top_5 = tf.nn.top_k(prob, k=5)
return sess.run(top_5)
with tf.Session() as sess:
saver.restore(sess, './lenet')
signs_top_5=new_test_top(new_test_norm, sess)
print(signs_top_5)
plt.figure(figsize=(16, 21))
for i in range(5):
plt.subplot(12, 2, 2*i+1)
plt.imshow(new_test_images[i])
plt.axis('off')
plt.subplot(12, 2, 2*i+2)
plt.barh(np.arange(1, 6, 1), signs_top_5.values[i, :])
labs=[signs[j] for j in signs_top_5.indices[i]]
plt.yticks(np.arange(1, 6, 1), labs)
plt.show()
```
| github_jupyter |
## Scalar differentiation
```
import sympy
a = sympy.Symbol('a')
b = sympy.Symbol('b')
e = (a + 2*b)**5
print("\nExpression : ")
pprint(e)
print("\n\nDifferentiating w.r.t. a:")
pprint(e.diff(a))
print("\n\nDifferentiating w.r.t. b:")
pprint(e.diff(b))
print("\n\nSecond derivative of the above result w.r.t. a:")
pprint(e.diff(b).diff(a, 2))
print("\n\nExpanding the above result:")
pprint(e.expand().diff(b).diff(a, 2))
```
## Approximate differentiation
```
from sympy import *
from sympy.printing.str import StrPrinter
from sympy.printing.latex import LatexPrinter
##### M E T H O D S
def matrices(names):
''' Call with A,B,C = matrix('A B C') '''
return symbols(names,commutative=False)
# Transformations
d = Function("d",commutative=False)
inv = Function("inv",commutative=False)
class t(Function):
''' The transposition, with special rules
t(A+B) = t(A) + t(B) and t(AB) = t(B)t(A) '''
is_commutative = False
def __new__(cls,arg):
if arg.is_Add:
return Add(*[t(A) for A in arg.args])
elif arg.is_Mul:
L = len(arg.args)
return Mul(*[t(arg.args[L-i-1]) for i in range(L)])
else:
return Function.__new__(cls,arg)
# Differentiation
MATRIX_DIFF_RULES = {
# e =expression, s = a list of symbols respsect to which
# we want to differentiate
Symbol : lambda e,s : d(e) if s.has(e) else 0,
Add : lambda e,s : Add(*[matDiff(arg,s) for arg in e.args]),
Mul : lambda e,s : Mul(matDiff(e.args[0],s),Mul(*e.args[1:]))
+ Mul(e.args[0],matDiff(Mul(*e.args[1:]),s)) ,
t : lambda e,s : t( matDiff(e.args[0],s) ),
inv : lambda e,s : - e * matDiff(e.args[0],s) * e
}
def matDiff(expr,symbols):
if expr.__class__ in MATRIX_DIFF_RULES.keys():
return MATRIX_DIFF_RULES[expr.__class__](expr,symbols)
else:
return 0
##### C O S M E T I C S
# Console mode
class matStrPrinter(StrPrinter):
''' Nice printing for console mode : X¯¹, X', ∂X '''
def _print_inv(self, expr):
if expr.args[0].is_Symbol:
return self._print(expr.args[0]) +'¯¹'
else:
return '(' + self._print(expr.args[0]) + ')¯¹'
def _print_t(self, expr):
return self._print(expr.args[0]) +"'"
def _print_d(self, expr):
if expr.args[0].is_Symbol:
return '∂'+ self._print(expr.args[0])
else:
return '∂('+ self._print(expr.args[0]) +')'
def matPrint(m):
mem = Basic.__str__
Basic.__str__ = lambda self: matStrPrinter().doprint(self)
print(str(m).replace('*',''))
Basic.__str__ = mem
# Latex mode
class matLatPrinter(LatexPrinter):
''' Printing instructions for latex : X^{-1}, X^T, \partial X '''
def _print_inv(self, expr):
if expr.args[0].is_Symbol:
return self._print(expr.args[0]) +'^{-1}'
else:
return '(' + self._print(expr.args[0]) + ')^{-1}'
def _print_t(self, expr):
return self._print(expr.args[0]) +'^T'
def _print_d(self, expr):
if expr.args[0].is_Symbol:
return '\partial '+ self._print(expr.args[0])
else:
return '\partial ('+ self._print(expr.args[0]) +')'
def matLatex(expr, profile=None, **kargs):
if profile is not None:
profile.update(kargs)
else:
profile = kargs
return matLatPrinter(profile).doprint(expr)
```
$ H = X {(X^T S^{-1} X)}^{-1} X^T S^{-1} $
How much is $\displaystyle \frac{\partial H}{\partial X}$?
```
X, S = matrices("X S")
H = X * inv(t(X) * inv(S) * X) * t(X) * inv(S)
expr = expand(expand(matDiff(H,X)))
expr
matPrint(expr)
latex = matLatex(matDiff(H,X))
latex
from IPython.display import Math
Math(latex)
```
## Matrix differentiation
If $E = x^T A x$ then:
$ \displaystyle \frac{\partial E}{\partial x} = x^T (A + A^T) $
```
x = MatrixSymbol('x', 3, 1)
A = MatrixSymbol('A', 3, 3)
alpha = x.T * A * x
print(alpha)
print(Matrix(alpha))
print(Matrix(alpha).diff(x[0, 0]))
for expr in derive_by_array(Matrix(alpha), Matrix(x)):
print(expr)
expr1 = Matrix(derive_by_array(Matrix(alpha), Matrix(x))).T
expr2 = Matrix(x.T * (A + A.T))
simplify(expr1 - expr2)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/mayankshouche/DSLabFinalProject/blob/main/Bert_Fine_Tuning" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
!pip install transformers datasets tweet-preprocessor
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import wordcloud
import preprocessor as p # tweet-preprocessor
import nltk
import re
import seaborn as sns
import torch
from transformers import BertTokenizer, BertForSequenceClassification, Trainer, TrainingArguments, AdamW, get_linear_schedule_with_warmup
from sklearn.metrics import accuracy_score, roc_auc_score
from datasets import Dataset
from sklearn.model_selection import train_test_split
from scipy.special import softmax
from torch.utils.data import TensorDataset, DataLoader, RandomSampler, SequentialSampler
from tqdm.notebook import tqdm
dataset = pd.read_csv('/content/drive/MyDrive/Copy of tweets_3k.csv', index_col=0)
dataset.head()
print(f"States: {dataset['State'].nunique()}")
dataset["State"].value_counts().plot.bar()
```
### Build dataset for classification
```
dataset_final = dataset.copy()
LABEL_MAP = {
"AntiDemocrats": 0,
"ProDemocrats": 1,
"AntiRepublicans": 2,
"ProRepublicans": 3
}
def buildLabels(row):
if row["Score"] < 0:
return LABEL_MAP.get("Anti" + row["PartyName"])
else:
return LABEL_MAP.get("Pro" + row["PartyName"])
def cleanTweet(row):
tweet = row["Text"]
tweet = str(p.clean(tweet))
tweet = re.sub(r'[^\w\s]', '', tweet) # punctuation
tweet = re.sub("^\d+\s|\s\d+\s|\s\d+$", " ", tweet) # numbers
return tweet
dataset_final["label"] = dataset_final.apply(lambda row: buildLabels(row), axis=1)
dataset_final["clean_text"] = dataset_final.apply(lambda row: cleanTweet(row),
axis=1)
dataset_final.drop(columns=['Created-At', 'From-User-Id', 'To-User-Id', 'Language',
'Retweet-Count', 'PartyName', 'Id', 'Score', 'Scoring String',
'Negativity', 'Positivity', 'Uncovered Tokens', 'Total Tokens'],
inplace=True)
X_train, X_val, y_train, y_val = train_test_split(dataset_final.index.values,
dataset_final.label.values,
test_size=0.15,
random_state=42,
stratify=dataset_final.label.values)
dataset_final['data_type'] = ['not_set']*dataset_final.shape[0]
dataset_final.loc[X_train, 'data_type'] = 'train'
dataset_final.loc[X_val, 'data_type'] = 'test'
dataset_final.groupby(['label', 'data_type']).count()
```
### Try out BERTForSequenceClassification to see how it goes
```
tokenizer = BertTokenizer.from_pretrained('bert-base-uncased',
do_lower_case=True)
# find distribution of length of tokenized tweets
lens = []
for tweet in dataset_final.clean_text:
tokens = tokenizer.encode(tweet, max_length=512)
lens.append(len(tokens))
sns.displot(lens)
plt.xlim([0, 128])
# tokenize train and test data so BERT can understand it
encoded_data_train = tokenizer.batch_encode_plus(
dataset_final[dataset_final.data_type=='train'].clean_text.values,
add_special_tokens=True,
return_attention_mask=True,
padding=True,
max_length=64,
return_tensors='pt'
)
encoded_data_test = tokenizer.batch_encode_plus(
dataset_final[dataset_final.data_type=='test'].clean_text.values,
add_special_tokens=True,
return_attention_mask=True,
padding=True,
max_length=64,
return_tensors='pt'
)
# destructure out the input_ids, attention masks, and labels from tokenizer & encoder output
input_ids_train = encoded_data_train['input_ids']
attention_masks_train = encoded_data_train['attention_mask']
labels_train = torch.tensor(dataset_final[dataset_final.data_type=='train'].label.values)
input_ids_test = encoded_data_test['input_ids']
attention_masks_test = encoded_data_test['attention_mask']
labels_test = torch.tensor(dataset_final[dataset_final.data_type=='test'].label.values)
train_data = TensorDataset(input_ids_train, attention_masks_train, labels_train)
test_data = TensorDataset(input_ids_test, attention_masks_test, labels_test)
BATCH_SIZE = 8
train_dataloader = DataLoader(train_data,
sampler=RandomSampler(train_data),
batch_size=BATCH_SIZE)
test_dataloader = DataLoader(test_data,
sampler=SequentialSampler(test_data),
batch_size=BATCH_SIZE)
model = BertForSequenceClassification.from_pretrained("bert-base-uncased",
num_labels=4,
output_attentions=False,
output_hidden_states=False)
EPOCHS = 5
optimizer = AdamW(model.parameters(),
lr=1e-5,
eps=1e-8)
scheduler = get_linear_schedule_with_warmup(optimizer,
num_warmup_steps=0,
num_training_steps=len(train_dataloader)*EPOCHS)
def auc_score(preds, labels):
soft_preds = softmax(preds, axis=1) # logit -> probability
return roc_auc_score(labels, soft_preds, multi_class='ovr')
def acc_score_by_class(preds, labels):
label_dict_inverse = {v: k for k, v in LABEL_MAP.items()}
preds_flat = np.argmax(preds, axis=1).flatten()
labels_flat = labels.flatten()
for label in np.unique(labels_flat):
y_preds = preds_flat[labels_flat==label]
y_true = labels_flat[labels_flat==label]
print(f'Class: {label_dict_inverse[label]}')
print(f'Accuracy: {len(y_preds[y_preds==label])}/{len(y_true)}\n')
```
following is mostly taken from [this medium article](https://towardsdatascience.com/multi-class-text-classification-with-deep-learning-using-bert-b59ca2f5c613)
```
def evaluate(dataloader):
model.eval()
loss_val_total = 0
predictions, true_vals = [], []
for batch in dataloader:
# convert data to CUDA
batch = tuple(b.to(device) for b in batch)
inputs = {
'input_ids': batch[0],
'attention_mask': batch[1],
'labels': batch[2],
}
with torch.no_grad():
outputs = model(**inputs) # get predictions
loss = outputs[0]
logits = outputs[1]
loss_val_total += loss.item()
logits = logits.detach().cpu().numpy()
label_ids = inputs['labels'].cpu().numpy()
predictions.append(logits)
true_vals.append(label_ids)
loss_val_avg = loss_val_total/len(dataloader)
predictions = np.concatenate(predictions, axis=0)
true_vals = np.concatenate(true_vals, axis=0)
return loss_val_avg, predictions, true_vals
import random
seed_val = 42
random.seed(seed_val)
np.random.seed(seed_val)
torch.manual_seed(seed_val)
torch.cuda.manual_seed_all(seed_val)
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
model.to(device)
print(device)
torch.cuda.get_device_name(0)
for epoch in tqdm(range(1, EPOCHS+1)): # use tqdm for a progress bar
model.train() # enter training mode
loss_train_total = 0
progress_bar = tqdm(train_dataloader, desc=f'Epoch {epoch}', leave=False, disable=False)
for batch in progress_bar:
model.zero_grad()
# get CUDA data
batch = tuple(b.to(device) for b in batch)
inputs = {
'input_ids': batch[0],
'attention_mask': batch[1],
'labels': batch[2],
}
outputs = model(**inputs) # evaluate
# for reference, we are using cross-entropy loss here,
# as implemented in https://huggingface.co/transformers/_modules/transformers/modeling_bert.html
loss = outputs[0]
loss_train_total += loss.item()
loss.backward() # do backprop
torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
optimizer.step()
scheduler.step()
progress_bar.set_postfix({'training_loss': '{:.3f}'.format(loss.item()/len(batch))})
torch.save(model.state_dict(), f'/content/drive/MyDrive/finetuned_BERT_epoch_{epoch}.model')
tqdm.write(f'\nEpoch {epoch}')
loss_train_avg = loss_train_total/len(train_dataloader)
tqdm.write(f'Training loss: {loss_train_avg}')
val_loss, predictions, true_vals = evaluate(test_dataloader)
auc = auc_score(predictions, true_vals)
tqdm.write(f'Testing loss: {val_loss}')
tqdm.write(f'AUC: {auc}')
_, preds, true_vals = evaluate(test_dataloader)
acc_score_by_class(preds, true_vals)
```
| github_jupyter |
```
import pandas as pd
import orca
import os; os.chdir('../')
from scripts import datasources
trips = pd.read_csv('./data/persons_HWtrips.csv')
trips.head()
job_coords = orca.merge_tables('jobs', ['jobs', 'buildings', 'parcels'])
job_coords = job_coords[['x', 'y']]
hh_coords = orca.merge_tables('households', ['households', 'units', 'buildings', 'parcels'])
hh_coords = hh_coords[['x', 'y']]
trips = trips[['person_id', 'household_id', 'job_id', 'MODE', 'HW_departure_time', 'WH_departure_time']].rename(
columns={'HW_departure_time': 'Home', 'WH_departure_time': 'Work'})
trip_data = trips.merge(
hh_coords, left_on='household_id', right_index=True).merge(
job_coords, left_on='job_id', right_index=True, suffixes=('_home', '_work'))
trip_data = trip_data[['person_id', 'MODE', 'Home', 'Work', 'x_home', 'y_home', 'x_work', 'y_work']]
melted = trip_data.melt(
id_vars=['person_id', 'MODE', 'x_home', 'y_home', 'x_work', 'y_work'],
var_name='activityType', value_name='endTime')
melted['x'] = None
melted['y'] = None
melted.loc[melted['activityType'] == 'Home', 'x'] = melted.loc[melted['activityType'] == 'Home', 'x_home']
melted.loc[melted['activityType'] == 'Home', 'y'] = melted.loc[melted['activityType'] == 'Home', 'y_home']
melted.loc[melted['activityType'] == 'Work', 'x'] = melted.loc[melted['activityType'] == 'Work', 'x_work']
melted.loc[melted['activityType'] == 'Work', 'y'] = melted.loc[melted['activityType'] == 'Work', 'y_work']
plans = melted.sort_values(['person_id', 'endTime'])[['person_id', 'MODE', 'activityType', 'endTime', 'x', 'y']].reset_index(drop=True)
plans['planElement'] = 'activity'
plans['planElementIndex'] = plans.groupby('person_id').cumcount() * 2 + 1
returnActivity = plans[plans['planElementIndex'] == 1]
returnActivity.loc[:, 'planElementIndex'] = 5
returnActivity.loc[:, 'endTime'] = ''
plans = plans.append(returnActivity, ignore_index=True).sort_values(['person_id', 'planElementIndex'])
legs = plans[plans['planElementIndex'].isin([1,3])]
legs.loc[:, 'planElementIndex'] = legs.loc[:, 'planElementIndex'] + 1
legs.loc[:, 'activityType'] = ''
legs.loc[:, 'endTime'] = ''
legs.loc[:, 'x'] = ''
legs.loc[:, 'y'] = ''
legs.loc[:, 'planElement'] = 'leg'
plans = plans.append(legs, ignore_index=True).sort_values(['person_id', 'planElementIndex']).rename(
columns={'person_id': 'personId', 'MODE': 'mode'}).reset_index(drop=True)
plans = plans[['personId', 'planElement', 'planElementIndex', 'activityType', 'x', 'y', 'endTime', 'mode']]
plans.loc[plans['planElement'] == 'activity', 'mode'] = ''
plans.to_csv('./data/urbansim_beam_plans.csv', index=False)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as m
df = pd.read_csv('summer.csv')
df.head()
len(df)
tv = len(df) * 9
print(tv)
```
DataFrame
```
df
df.isnull().sum()
```
NAME OF CITIES WHERE SUMMER OLYMPICS IS HELD
```
c = []
c = df['Country'].unique()
c
```
NUMBER OF CITIES WHERE SUMMER OLYMPICS IS HELD
```
n = len(c) - 1
print("The Number of Cities Where Summer Olympics is held is \n", n)
```
Sport which is having most number of Gold Medals so far (Top 5)
```
x = df[df['Medal'] == 'Gold']
gold = []
for i in x['Sport'].unique():
gold.append([i, len(x[x['Sport'] == i])])
gold = pd.DataFrame(gold, columns = ['Sport', 'Medals'])
gold = gold.sort_values(by = 'Medals', ascending = False).head()
gold
gold.plot(x = 'Sport', y = 'Medals', kind = 'bar', color = 'gold', figsize = (6,6))
```
Sport which is having most number of medals so far (Top 5)
```
tm = []
for m in df['Sport'].unique():
tm.append([m, len(df[df['Sport'] == m])])
tm = pd.DataFrame(tm, columns = ['Sport', 'Total Medals'])
tm = tm.sort_values(by = 'Total Medals', ascending = False).head()
tm
tm.plot(x = 'Sport', y = 'Total Medals', kind = 'bar', color = 'red', figsize = (6,6))
```
Players who have won most number of medals (Top 5)
```
at = []
for ap in df['Athlete'].unique():
at.append([ap, len(df[df['Athlete'] == ap])])
at = pd.DataFrame(at, columns = ['Player', 'Total Medals'])
at = at.sort_values(by = 'Total Medals', ascending = False).head()
at
at.plot(x = 'Player', y = 'Total Medals', kind = 'bar', color = 'green', figsize = (6,6))
```
Players who have won most number Gold Medals of medals (Top 5)
```
x = df[df['Medal'] == 'Gold']
plgold = []
for i in x['Athlete'].unique():
plgold.append([i, len(x[x['Athlete'] == i])])
plgold = pd.DataFrame(plgold, columns = ['Player', 'Gold Medals'])
plgold = plgold.sort_values(by = 'Gold Medals', ascending = False).head()
plgold
plgold.plot(x = 'Player', y = 'Gold Medals', kind = 'bar', color = 'gold', figsize = (6,6))
```
The year where India won first Gold Medal in Summer Olympics
```
x = df[df['Medal'] == 'Gold']
y = x.loc[x['Country'] == 'IND']
y.iloc[0]
print("The first Gold Medal in Summer Olympics won by India was in the year")
y['Year'].iloc[0]
```
Most popular event in terms on number of players (Top 5)
```
eve = []
for i in df['Event'].unique():
eve.append([i, len(df[df['Event'] == i])])
eve = pd.DataFrame(eve, columns = ['Event', 'Total Players'])
eve = eve.sort_values(by = 'Total Players', ascending = False).head()
eve
eve.plot(x = 'Event', y = 'Total Players', kind = 'bar', color = 'black', figsize = (6,6))
```
Sport which is having most female Gold Medalists (Top 5)
```
x = df[df['Medal'] == 'Gold']
f = x[x['Gender'] == 'Women']
wgold = []
for i in f['Sport'].unique():
wgold.append([i, len(f[f['Sport'] == i])])
wgold = pd.DataFrame(wgold, columns = ['Sport', 'Female Gold Medalists'])
wgold = wgold.sort_values(by = 'Female Gold Medalists', ascending = False).head()
wgold
wgold.plot(x = 'Sport', y = 'Female Gold Medalists', kind = 'bar', color = 'pink', figsize = (6,6))
```
Done by Sushanth UV
| github_jupyter |
```
import boto3
s3 = boto3.resource('s3')
bucket_name = 'canopy-production-ml'
pc_bucket = s3.Bucket(bucket_name)
all_objects = pc_bucket.objects.all()
yes_chips = []
for obj in all_objects:
if 'cloudfree-merge-polygons/yes' in obj.key:
yes_chips.append(obj)
len(yes_chips)
yes_chips[0].key
key = yes_chips[0].key
key.split('/')
split = key.split('/')
split[2] = 'dataset_v2'
'/'.join(split)
total = len(yes_chips)
for i, chip in enumerate(yes_chips):
print(f'Copying chip {i+1} of {total}', end='\r', flush=True)
key = chip.key
copy_source = {
'Bucket': bucket_name,
'Key': key
}
key_split = key.split('/')
key_split[2] = 'dataset_v2'
new_key = '/'.join(key_split)
pc_bucket.copy(copy_source, new_key)
print(end - start)
all_chips = []
for obj in all_objects:
if 'cloudfree-merge-polygons/full' in obj.key:
all_chips.append(obj)
len(all_chips)
yes_filenames = [chip.key.split('/')[-1] for chip in yes_chips]
yes_filenames[:10]
all_chips[0].key
all_chips[0].key[-3:]
no_chips = []
total = len(all_chips)
for i, chip in enumerate(all_chips):
filename = chip.key.split('/')[-1]
if filename not in yes_filenames:
no_chips.append(chip)
len(no_chips)
import collections
count = collections.Counter(yes_filenames)
duplicates = [i for i in count if count[i] >= 2]
duplicates
odd_keys
assert len(yes_chips) + len(no_chips) == len(all_chips)
len(yes_chips) + len(no_chips)
len(all_chips)
chip_filenames = [chip.key.split('/')[-1] for chip in all_chips]
missing = []
total = len(yes_chips)
for i, chip in enumerate(yes_chips):
print(f'Chip {i+1} of {total}', end='\r', flush=True)
if chip.key.split('/')[-1] not in chip_filenames:
missing.append(chip.key)
len(missing)
missing
len(no_chips)
def boto3_copy_objs(keys, old_bucket_name, new_bucket,
folder_change_dict=None, folder_add_dict=None, folder_swap_dict=None):
total = len(keys)
for i, key in enumerate(keys):
print(f'Copying object {i+1} of {total}', end='\r', flush=True)
copy_source = {
'Bucket': old_bucket_name,
'Key': key
}
key_split = key.split('/')
if folder_change_dict:
for k in folder_change_dict:
try:
key_split[k] = folder_change_dict[k]
except:
raise ValueError('Error when applying the folder_change_dict')
if folder_add_dict:
for k in folder_add_dict:
try:
key_split.insert(k, folder_add_dict[k])
except:
raise ValueError('Error when applying the folder_add_dict')
if folder_swap_dict:
for k in folder_swap_dict:
i = folder_swap_dict[k]
try:
key_split[k], key_split[i] = key_split[i], key_split[k]
except:
raise ValueError('Error when applying the folder_swap_dict')
new_key = '/'.join(key_split)
#print(new_key)
new_bucket.copy(copy_source, new_key)
no_chips[0].key
boto3_copy_objs(no_chips, bucket_name, pc_bucket, {2: 'dataset_v2'}, {3: 'misc'}, {4: 5})
keep_cats = ['ISL', 'Roads', 'Industrial_agriculture', 'Shifting_cultivation', 'Mining']
keep_filenames = []
for obj in all_objects:
for category in keep_cats:
if f'cloudfree-merge-polygons/dataset_v2/{category}' in obj.key:
filename = obj.key.split('/')[-1]
keep_filenames.append(filename)
len(keep_filenames)
keep_filenames[0]
remove_cats = ['River', 'Rainforest', 'Water', 'Savanna', 'Habitation']
move_chips = []
delete_chips = []
for obj in all_objects:
for category in remove_cats:
if f'cloudfree-merge-polygons/dataset_v2/{category}' in obj.key:
filename = obj.key.split('/')[-1]
if filename not in keep_filenames:
move_chips.append(obj)
else:
delete_chips.append(obj)
len(move_chips) + len(delete_chips)
import pandas as pd
data = {'Keys': [chip.key for chip in move_chips]}
df = pd.DataFrame(data=data)
df.head()
df['Filenames'] = df['Keys'].apply(lambda x:x.split('/')[-1])
df.head()
print(df.shape)
df2 = df.drop_duplicates(subset=['Filenames'])
print(df2.shape)
move_chips_no_dupes = df2['Keys'].tolist()
len(move_chips_no_dupes)
move_chips_no_dupes[0]
boto3_copy_objs(move_chips_no_dupes, bucket_name, pc_bucket, {3: 'misc'})
delete_chips_full = move_chips + delete_chips
len(delete_chips_full)
# delete_objs_list = [{'Key': chip.key for chip in delete_chips_full}]
#client = boto3.client('s3')
#response = client.delete_objects(
# Bucket=bucket_name,
# Delete=delete_objs_list
#)
```
### train/test split based off polygons
```
all_objects = pc_bucket.objects.all()
all_chips = []
for obj in all_objects:
if 'cloudfree-merge-polygons/dataset_v2/' in obj.key:
all_chips.append(obj)
print(len(all_chips))
print(all_chips[0].key)
all_chips[1].key
assert 'chips/cloudfree-merge-polygons/dataset_v2/ISL/' in all_chips
chip_count = {}
for chip in all_chips:
key = chip.key
try:
poly_id = key.split('/')[5]
if poly_id in chip_count.keys():
chip_count[poly_id] += 1
else:
chip_count[poly_id] = 1
except:
print(key)
sum(chip_count.values())
chip_count
import json
json_object = json.dumps(chip_count)
with open('chip_count.json', 'w') as fp:
json.dump(chip_count, fp)
bucket_name = 'canopy-production-ml-output'
my_bucket = s3.Bucket(bucket_name)
path = 'ckpt/pc-tf-custom-container-2021-04-14-15-15-52-166/'
h5_files = [obj.key for obj in my_bucket.objects.filter(Prefix=path) if obj.key[-2:] == 'h5']
h5_files
h5_files = h5_files[1:]
h5_files_dict = {}
for file in h5_files:
key = int(file.split('_')[-1].split('.')[0])
h5_files_dict[key] = file
h5_files_dict
max_epoch = max(h5_files_dict)
h5_files_dict[max_epoch]
max(h5_files_dict.keys())
```
| github_jupyter |
# [Numpy](https://numpy.org/devdocs/user/quickstart.html)
В 2019 году между народная группа ученых на основании данных, которые были получены с 8 обсерваторий по всему миру сделали то, что никто ранее не делал, фотографию черной дыры:
<img src="https://leonardo.osnova.io/4d8e3b6f-4d76-698c-f4b1-d1e39610e5be/-/scale_crop/2100x1223/center/-/format/webp/" width=300/>
Исследователи сообщили, что в получении такого результата им помогли следующие библиотеки Python:
<ul>
<li>astropy</li>
<li>matplotlib</li>
<li>networkx</li>
<li>numpy</li>
<li>pandas</li>
<li>scipy</li>
<li>и другие</li>
</ul>
В нашем курсе мы рассмотрим библиотеки __numpy__, __pandas__ и __matplotlib__ с целью исследовать машинное обучение. Но, как вы видите, их использование не ограничивается машинным обучением, они могут быть применены и в высокой науке.
## Numpy. Операции над векторами и тензорами
Ядром пакета NumPy является объект [ndarray](https://docs.scipy.org/doc/numpy/reference/generated/numpy.ndarray.html)
Библеотека NumPy предоставляет следующие **возможности**:
* работать с многомерными массивами (включая матрицы)
* производить быстрое вычисление математических функций на многомерных массивах
Сегодня нам потребуются следующие библиотеки:
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
## рисование графиков внутри ноутбука
```
### Справка
```
?np.ndarray
#можно еще ??nd.array, а вот ???np.ndarray уже нельзя
```
### Поиск в библиотеке
```
np.lookfor('create array')
```
## Генерация Numpy arrays
Numpy позволяет просто и быстро работать с векторами, матрицами и структурами более высоких порядков: тензорами. Давайте объявим несколько простых numpy объектов:
```
a = np.array([4, 1, 6])
b = np.array([
[0, 1, 2],
[4, 5, 6]
])
c = np.array([
[
[3, 1, 0],
[2, 2, 0]
],
[
[5, 55, 5],
[5, 55, 5]
]
])
```
Теперь случае __a__, - вектор, __b__ - матрица, __c__ - тензор 3 размерности
```
a
b
c
```
### Тензоры
<img src = "https://www.cc.gatech.edu/~san37/img/dl/tensor.png" width=500>
### Генерация массивов из коробки
* [arange](https://docs.scipy.org/doc/numpy/reference/generated/numpy.arange.html) — аналог range из Python, которому можно передать нецелочисленный шаг
```
np.arange(0, 5, 0.5)
```
* [linspace](https://docs.scipy.org/doc/numpy/reference/generated/numpy.linspace.html) — способ равномерно разбить отрезок на n-1 интервал
```
np.linspace(0, 5, 13)
```
* [logspace](https://docs.scipy.org/doc/numpy/reference/generated/numpy.logspace.html) — способ разбить отрезок по логарифмической шкале
```
print(np.logspace(start=1, stop=5, num=6, base=2))
print(2 ** np.linspace(start=1, stop=5, num=6))
#делаем linspace, а потом возводим base в степень каждого элемента linspace
```
* [zeros](https://docs.scipy.org/doc/numpy/reference/generated/numpy.zeros.html) — создаёт массив заполненный нулями заданной размерности
```
np.zeros((2, 2, 5))
```
* [ones](https://docs.scipy.org/doc/numpy/reference/generated/numpy.ones.html) — создаёт массив заполненный единицами заданной размерности
```
np.ones((2, 2, 7))
```
* [eye](https://docs.scipy.org/doc/numpy/reference/generated/numpy.eye.html) - создаёт единичную матрицу заданного размера
```
np.eye(5)
```
Если не знаем какую-то функцию, можно сделать поиск прямо внутри numpy
## Преобразования размерностей над матрицами
Pазмеры массива хранятся в поле **shape**, а количество размерностей - в **ndim**
```
array = np.ones([2, 4, 3])
print(array)
array.shape, array.ndim
```
Метод [reshape](https://docs.scipy.org/doc/numpy/reference/generated/numpy.reshape.html) позволяет преобразовать размеры массива без изменения данных
```
array = np.arange(0, 6, 0.5)
print(array)
array = array.reshape((2, 2, 3))
array
```
Для того, что бы развернуть многомерный массив в вектор, можно воспользоваться функцией [ravel](https://docs.scipy.org/doc/numpy-1.15.0/reference/generated/numpy.ravel.html)
```
array.ravel()
```
### Транспонирование матрицы
```
A = np.arange(10).reshape(2, 5)
print(A)
A.T
```
Заметим, что сама матрица А не изменилась. A.T - новый объект, который возвращает функция транспонирования.
```
A
```
Однако транспонировать вектор мы не умеем
```
a = np.array([1, 2, 3, 4])
a.T
```
### Добавление размерностей
Чтобы транспонировать вектор нам необходимо превратить его в матрицу (n,1). Это можно сделать двумя способами:
* [np.expand_dims](https://docs.scipy.org/doc/numpy/reference/generated/numpy.expand_dims.html)
* [np.newaxis](https://stackoverflow.com/questions/29241056/how-does-numpy-newaxis-work-and-when-to-use-it)
```
a = np.array([1, 2, 3, 4])
a1 = np.expand_dims(a, 1) #добавить размерность по след. координате
a2 = a[:, np.newaxis] # : - все элементы по 1 коорд.(см. индексацию ниже), np.newaxis - новая координата
a1
a2
a.shape, a1.shape, a2.shape
```
Заметьте, получается, что изначальный numpy.ndarray - столбец, а вовсе не строка!
## Операции над матрицами
```
A = np.arange(9).reshape(3, 3)
B = np.arange(2, 11).reshape(3, 3)
print(A)
print(B)
```
Операции над матрицами одинаковых размерностей осуществляются поэлементно
```
A + B
A / B
```
Отдельно обратим внимание на то, что умножение массивов также является **поэлементным**, а не матричным:
```
A * B
```
Для выполнения матричного умножения необходимо использовать функцию dot:
```
A.dot(B)
```
А что будет если произвести оперцию между матрицей и скаляром?
```
4 * A
A + 10
A ** B
```
А если добавим к матрице вектор?
```
np.zeros([4, 3]) + np.array([0, 1, 2])
```
А если добавим к столбцу строку?
```
np.array([[ 0],
[10],
[20],
[30]]) + np.array([0, 1, 2])
```
Почему так происходит?
### [Broadcasting](https://docs.scipy.org/doc/numpy/user/basics.broadcasting.html)
**Замечание:** Все арифметические операции над массивами одинаковой размерности производятся поэлементно
Broadcasting снимает правило одной размерности и позволяет производить арифметические операции над массивами разных, но всётаки согласованных размерностей.
Если количество размерностей не совпадают, то к массиву меньшей размерности добавляются фиктивные размерности "слева".
Прозе всего показать на картинке:
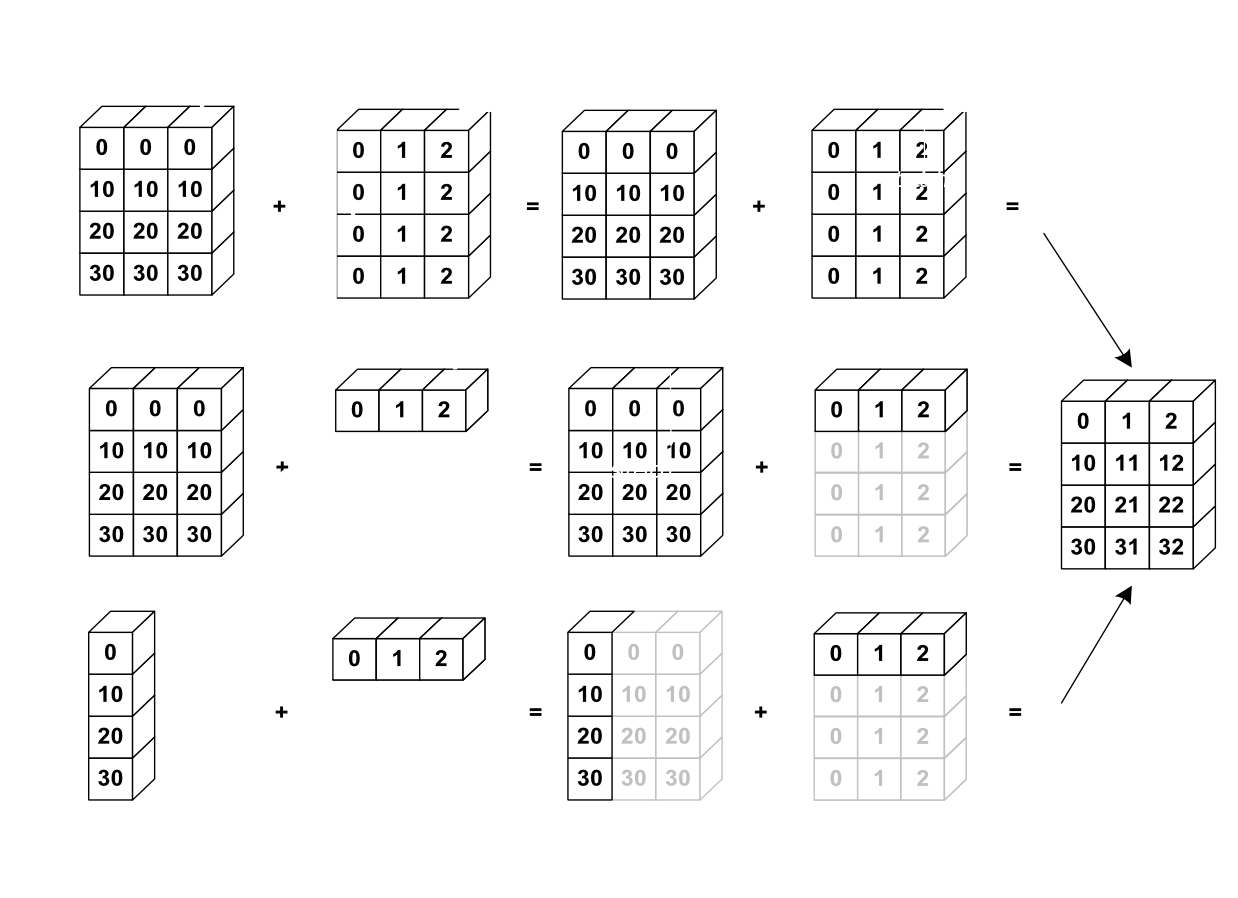
Если размерности не согласуются, то операция невозможна.

```
np.zeros([4, 3]) + np.array([0, 1,2, 3])
```
Очевидно, что broadcasting работает не только для сложения, но и для любой операции.
```
np.array([[ 0],
[10],
[20],
[30]]) ** np.array([0, 1, 2])
np.arange(27).reshape(3, 3, 3) + np.arange(3)
```
## Конкатенация многомерных массивов
Конкатенировать несколько массивом можно с помощью функций [**np.concatenate, np.hstack, np.vstack**](https://stackoverflow.com/questions/33356442/when-should-i-use-hstack-vstack-vs-append-vs-concatenate-vs-column-stack)
```
A = np.arange(9).reshape(3, 3)
B = np.arange(1, 10).reshape(3, 3)
np.hstack([A, B])
np.vstack([A, B])
```
## Агрегирующие операции
Numpy поддерживает множество аггрегирующих операций, такие как sum(), prod(), max(), min(). Эти операции применяются на выбранные оси тензора и размерность тенхора уменьшается. Давайте сгенерируем тензор размерности 3 и попробуем применить эти операции:
```
tensor = np.random.choice(a=[0,1,2,3], size=(2,3,4)) #рандомайз
tensor
```
np.random дает возможность использовать различные функции генерации данных. np.random.choice генерирует тензор данного размера size, который состоит из элементов множества a.
Давайте просуммируем все значения тензора:
```
tensor.sum()
```
Давайте просуммируем все значения тензора оп оси 0:
```
tensor_sum_0 = tensor.sum(axis = (0,1))
tensor_sum_0.shape, tensor_sum_0
```
Как видите, размерность тензора уменьшилась. Давайте выберем другую агрегирующую операцию и другую ось:
```
tensor_max_2 = tensor.max(axis = 2)
tensor_max_2.shape, tensor_max_2
```
Универсальные функции (sin, cos, exp и т.д.) также применяются поэлементно:
```
A = np.arange(9).reshape(3, 3)
np.exp(A)
```
Некоторые операции над массивами (например, вычисления минимума, максимума, суммы элементов) выполняются над всеми элементами вне зависимости от формы массива, однако при указании оси выполняются вдоль нее (например, для нахождения максимума каждой строки или каждого столбца):
```
A.min()
A.max(axis = 0)
A.sum(axis = 1)
```
Так же в NumPy реализованно много полезных операций для работы с массивами: [np.min](https://docs.scipy.org/doc/numpy/reference/generated/numpy.ndarray.min.html), [np.max](https://docs.scipy.org/doc/numpy/reference/generated/numpy.ndarray.max.html), [np.sum](https://docs.scipy.org/doc/numpy/reference/generated/numpy.sum.html), [np.mean](https://docs.scipy.org/doc/numpy/reference/generated/numpy.mean.html) и т.д.
**Замечание:** В каждой из перечисленных функций есть параметр **axis**, который указывает по какому измерению производить данную операцию. По умолчанию операция производится по всем значениям массива
## [Типы Данных](https://docs.scipy.org/doc/numpy-1.13.0/reference/arrays.dtypes.html)
Мы много поигрались с простыми числами `int`, `float`. Но можно использовать комплексные числа и очень **длинные** числа
```
a = np.array([2 + 3j, 4 + 5j])
b = np.array([np.longdouble(1/10**(1000))])
a.dtype, b.dtype
```
Можно и строки хранить.
```
s = np.array(['abra', 'zxcv', 'щаоштпавтищав'])
s.dtype, type(s[0]), s[0].dtype
```
Читаем Unicode длины меньше 13 (максимальное слово)
Да и вообще можно кастовать любой лист к массиву. Тип приводится к наследнику. Сложные объекты также хранятся по ссылке.
```
def f():
pass
class A(object):
def __init__(self):
pass
a = A()
np.array([2, 'asd', [1,2], {'d':'a'}, a, f])
```
Однако есть некоторые странности. Например при присутствии типа `str` в листе, где все остальные типы стандартные - все остальные элементы кастуются к стрингу
```
a = np.array([2, False, 'str', 'aaaaa'])
a.dtype, type(a[1])
```
Для всех кто хочет глубже погрузиться в систему типов numpy - [вот сюда](https://docs.scipy.org/doc/numpy-1.13.0/reference/arrays.dtypes.html).
И в само устройство массивов - [сюда](https://docs.scipy.org/doc/numpy/reference/arrays.html)
## Зачем Numpy?
Посмотрим с какой скоростью выполняется скалярное произведение для list и ndarrays
```
import time
A_quick = np.random.normal(size = (1000000,))
B_quick = np.random.normal(size = (1000000,))
A_slow, B_slow = list(A_quick), list(B_quick)
%%time
ans = sum([A_slow[i] * B_slow[i] for i in range(1000000)])
%%time
ans = np.sum(A_quick * B_quick)
```
Встроенная функция скалярного произведения
Почему же так происходит?
* NumPy array имеет фиксированную длину, которая определяется в момент его создания (в отличие от Python lists, которые могут расти динамически).
* Тип массива приводится к минимальному возможному для определения всех объектов. Поэтому мы четко понимаем сколько памяти у нас занимает массив и как оперативно с ним работать.
* Большинство `математическиx` операций реализовано на С в обход питоновских циклов.
Для более детального разбора:
* [тред на stackoverflow](https://stackoverflow.com/questions/8385602/why-are-numpy-arrays-so-fast)
* [Locality of reference](https://en.wikipedia.org/wiki/Locality_of_reference)
* [Внутреннее устройство массива numpy](https://docs.scipy.org/doc/numpy/reference/arrays.html)

## Индексация
В NumPy работает привычная индексация Python, включая использование отрицательных индексов и срезов
```
array = np.arange(0,5)
print(array[0])
print(array[-1])
print(array[1:-1])
print(array[1:-1:2])
print(array[::-1])
```
**Замечание**: Индексы и срезы в многомерных массивах не нужно разделять квадратными скобками
т.е. вместо ```matrix[i][j]``` нужно использовать ```matrix[i, j]```
Чтобы взять срезы по матрице можно использовать `:`
```
a = np.arange(27).reshape(3,3,3)
a[:,:,2], a[:,2,2]
```
Или еще более странный синтаксический сахар: `...` - `все остальное`
```
a[...,2], a[..., 2, 2]
```
### Индексирование масками
```
a = np.arange(0, 10)
(a % 3 == 0)
mask = (a % 3 == 0)
extract_from_a = a[mask]
extract_from_a
```
Индексирование маской может быть очень полезным для присваивания значений части элементов массива:
```
a[a % 3 == 0] = -1
a
```
### Индексирование массивом целых чисел
```
a = np.arange(10, 27)
a
a[[2, 3, 2, 4]]
a[[2, 4]] = -1000
a
a = np.arange(10, 20)
idx = np.array([[3, 4],
[9, 7]])
idx.shape
a[idx]
b = a.reshape((2, 5))
idx = np.array([(1, 1)])
b[idx]
```
### Собственные функции в Numpy
Чтобы сделать собственную `массовую` функцию в numpy недостаточно просто сделать ее, нужно ее еще `векторизовать`. Создадим функцю проверяющую массив на палиндром.
```
pal = lambda x: str(x) == str(x)[::-1]
a = np.array([11, 21, 343, 40])
pal(a) #воспринимает за x весь массив, str(x) - строковое представление массива
vpal = np.vectorize(pal)
vpal(a)
```
# Пример использования на чуть более реальной задаче.
Есть несколько способов индексирования тензоров. Мы их рассмотрим подробнее в этом разделе.
Но с маленькими тензорами работать не интересно. Давайте для начала создадим большой тензор. Сделаем матрицу размерности (1000, 2) проинициализируем ее элементы случайными числами из простых нормальных распределений:
```
coordinates_1 = np.random.normal(loc=[10, 10], scale = [30, 20], size = (500,2))
coordinates_2 = np.random.normal(loc=[50, 20], scale = [10, 70], size = (500,2))
coordinates = np.concatenate([coordinates_1, coordinates_2])
```
Эта матрица будет соответствовать координатам 1000 точек в пространстве. Давайте для наглядности их отобразим на плоскости:
Давайте раскрасим наши распределения:
```
plt.scatter(coordinates[:,0], coordinates[:,1], s=1, color=np.where(np.arange(0, 1000) < 500, 'blue', 'green'));
plt.grid()
```
Здесь мы встретились с функцией **np.where**, которая является некоторым аналогом if-then-else. В качестве первого аргумента она принимает condiniton - вектор булевых значений, в качестве двух других - вектор соответствующего размера или константу. Для тех позиций, где condiniton == True будут выбраны элементы первого вектора, в остальных случаях будут выбраны элементы второго вектора.
Для индексации в каждой компоненте мы можем использовать синтаксическую структуру slice. В этом случае размерность результата падать не будет, но будет осуществляться выборка в матрице по каждой координате.
Давайте оставим только те точки, в которых X принимает значения от 25 до 75:
```
X_bordered_coordinates = coordinates[(coordinates[:,0] > 25) & (coordinates[:,0] < 75)]
```
Здесь мы использовали индексирование булевым вектором. Этот трюк позволяет фильтровать numpy-объекты по некоторому условию.
Давайте отрисуем получившиеся точки поверх всех точек из coordinates:
```
plt.scatter(coordinates[:, 0], coordinates[:, 1], s=1, color='red')
plt.scatter(X_bordered_coordinates[:, 0], X_bordered_coordinates[:, 1], s=2, color='black')
plt.grid()
```
Для фильтрации мы можем использовать более сложные условия.
```
plt.scatter(coordinates[:, 0], coordinates[:, 1], s=1)
bordered_coordinates = coordinates[(coordinates[:,0] > 25) & (coordinates[:,0] < 75) &\
(coordinates[:,1] > -50) & (coordinates[:,1] < 50)]
plt.scatter(bordered_coordinates[:, 0], bordered_coordinates[:, 1], s=10, color='black');
plt.grid()
```
И еще более сложные:
```
new_coordinates = np.random.uniform(-20, 20, (100000, 2))
plt.scatter(new_coordinates[:, 0], new_coordinates[:, 1], s=1)
xs = new_coordinates[:, 0]
ys = new_coordinates[:, 1]
bordered_coordinates = new_coordinates[np.sin(np.sqrt(xs*xs + ys*ys) + np.arctan2(xs,ys)*5) < 0]
plt.scatter(bordered_coordinates[:, 0], bordered_coordinates[:, 1], s=1);
```
Как вы видите, в numpy также реализованы основные функции над векторами, аналогичные функциям над скалярами в библиотеке math
## Дополнительные материалы
* [Numpy documentation](https://numpy.org/devdocs/user/quickstart.html)
* [100 numpy execises](https://github.com/rougier/numpy-100/blob/master/100_Numpy_exercises.ipynb)
## Источники материалов:
* [mlcourse.ai](https://github.com/Yorko/mlcourse.ai) - курс Машинного обучения с OpenDataScience
* [AI Seminars](https://github.com/AICommunityInno/Seminars) - семинары по Машинному обучению в Иннополисе
* [HSE-ML course](https://github.com/esokolov/ml-course-hse) - курс Машинного обучения ФКН ВШЭ
| github_jupyter |
## Setup
```
%matplotlib qt
import matplotlib.pyplot as plt
import tensorflow as tf
import tensorflow_gan as tfgan
import numpy as np
import os, sys
from tqdm.notebook import tqdm
from pathlib import Path
sys.path.append( os.path.abspath('..') )
import utils
Path('MNIST').mkdir(exist_ok=True)
os.chdir('MNIST')
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.mnist.load_data()
data = np.concatenate((x_train, x_test))
data = (data.astype('float32') - 127.5) / 127.5 # normalize to [-1, 1]
data = np.expand_dims(data, axis=-1) # add channels dimension
assert data.shape == (70000, 28, 28, 1) # (batch, height, width, channel)
NUM_CLASSES = 10
labels = np.concatenate((y_train, y_test))
labels = np.expand_dims(labels, -1)
assert labels.shape == (70000, 1)
```
## 1 Models
### 1.1 Architecture
```
def generator_model(latent_dims):
## Label input
label = tf.keras.Input(shape=(1,), name='label', dtype=tf.int32)
embedding = tf.keras.layers.Embedding(input_dim=NUM_CLASSES, output_dim=36)(label)
label_channel = tf.keras.layers.Dense(7*7)(embedding)
label_channel = tf.keras.layers.Reshape((7, 7, 1))(label_channel)
## Latent input
seed = tf.keras.Input(shape=(latent_dims,), name='seed')
seed_channels = tf.keras.layers.Dense(7*7*255, input_shape=(latent_dims,))(seed)
seed_channels = tf.keras.layers.Reshape((7, 7, 255))(seed_channels)
channels = tf.keras.layers.Concatenate(axis=-1)([label_channel, seed_channels])
channels = tf.keras.layers.LeakyReLU()(channels)
channels = tf.keras.layers.BatchNormalization()(channels)
channels = tf.keras.layers.Conv2D(128, kernel_size=5, strides=1, padding='same')(channels)
channels = tf.keras.layers.LeakyReLU()(channels)
channels = tf.keras.layers.BatchNormalization()(channels)
channels = tf.keras.layers.UpSampling2D(size=2, interpolation='bilinear')(channels)
channels = tf.keras.layers.Conv2D(64, kernel_size=5, strides=1, padding='same')(channels)
channels = tf.keras.layers.LeakyReLU()(channels)
channels = tf.keras.layers.BatchNormalization()(channels)
channels = tf.keras.layers.UpSampling2D(size=2, interpolation='bilinear')(channels)
img = tf.keras.layers.Conv2D(1, kernel_size=5, strides=1, padding='same', activation='tanh')(channels)
return tf.keras.Model(inputs=[seed, label], outputs=img, name='generator')
def discriminator_model():
## Label input
label = tf.keras.Input(shape=(1,), name='label', dtype=tf.int32)
embedding = tf.keras.layers.Embedding(input_dim=NUM_CLASSES, output_dim=36)(label)
label_channel = tf.keras.layers.Dense(28*28)(embedding)
label_channel = tf.keras.layers.Reshape((28, 28, 1))(label_channel)
## Image input
image = tf.keras.Input(shape=(28, 28, 1), name='image')
channels = tf.keras.layers.Concatenate(axis=-1)([label_channel, image])
channels = tf.keras.layers.Conv2D(64, kernel_size=5, strides=2, padding='same', input_shape=(28,28,1))(channels)
channels = tf.keras.layers.LeakyReLU()(channels)
channels = tf.keras.layers.Dropout(0.3)(channels)
channels = tf.keras.layers.Conv2D(128, kernel_size=5, strides=2, padding='same')(channels)
channels = tf.keras.layers.LeakyReLU()(channels)
channels = tf.keras.layers.Dropout(0.3)(channels)
channels = tf.keras.layers.Flatten()(channels)
logit = tf.keras.layers.Dense(1)(channels)
return tf.keras.Model(inputs=[image, label], outputs=logit)
```
### 1.2 Losses
The binary cross entropy (BCE) between $y$ and $\hat{y}$ is calculated as:
$$
\mathrm{BCE}(y, \hat{y}) = - y \log\left(\hat{y}\right) - (1-y) \log\left(1 - \hat{y}\right)
$$
```
cross_entropy = tf.keras.losses.BinaryCrossentropy(from_logits=True)
```
The generator tries to maximize the chance of the discriminator being wrong. This is equivalent of trying to minimize the following loss function:
$$
J^{(G)} = -\log\bigl(D\bigl(G(z)\bigr)\bigr)
$$
```
def generator_loss(fake_output):
return cross_entropy(tf.ones_like(fake_output), fake_output)
```
The discriminator tries to correctly classify real data as real and fake data as fake. This is equivalent to minimizing the following loss function:
$$
J^{(D)} = -\log\bigr(D(x)\bigl) - \log\bigl(1 - D\bigl(G(z)\bigr)\bigr)
$$
Here we scale down the loss by a factor of $\;0.5$
```
def discriminator_loss_normal(real_output, fake_output):
real_loss = cross_entropy(tf.ones_like(real_output), real_output)
fake_loss = cross_entropy(tf.zeros_like(fake_output), fake_output)
return 0.5 * (real_loss + fake_loss)
```
This function applies one sided label smoothing of $\:0.9\:$ to the discriminator loss.
```
def discriminator_loss_smooth(real_output, fake_output):
real_loss = cross_entropy(0.9 * tf.ones_like(real_output), real_output)
fake_loss = cross_entropy(tf.zeros_like(fake_output), fake_output)
return 0.5 * (real_loss + fake_loss)
```
## 2 Training
### 2.1 Main Functions
```
def discriminator_train_step(generator, discriminator, images, labels, latent_dims):
noise = tf.random.normal([images.shape[0], latent_dims])
with tf.GradientTape() as disc_tape:
generated_imgs = generator([noise, labels], training=True)
real_output = discriminator([images, labels], training=True)
fake_output = discriminator([generated_imgs, labels], training=True)
loss_D = discriminator_loss(real_output, fake_output)
grads_D = disc_tape.gradient(loss_D, discriminator.trainable_variables)
discriminator.optimizer.apply_gradients(zip(grads_D, discriminator.trainable_variables))
def generator_train_step(generator, discriminator, y, latent_dims):
noise = tf.random.normal([y.shape[0], latent_dims])
with tf.GradientTape() as gen_tape:
generated_imgs = generator([noise, y], training=True)
fake_output = discriminator([generated_imgs, y], training=True)
loss_G = generator_loss(fake_output)
grads_G = gen_tape.gradient(loss_G, generator.trainable_variables)
generator.optimizer.apply_gradients(zip(grads_G, generator.trainable_variables))
def train(generator, discriminator, data, labels, epochs, batch_size=None, callbacks=None):
latent_dims = generator.input_shape[0][1]
batch_size = batch_size if batch_size is not None else 32
num_batches = 1 + (data.shape[0] - 1) // batch_size
X = tf.data.Dataset.from_tensor_slices(data)
Y = tf.data.Dataset.from_tensor_slices(labels)
dataset = tf.data.Dataset.zip((X, Y)).shuffle(data.shape[0]).batch(batch_size)
generator_step = tf.function(generator_train_step)
discriminator_step = tf.function(discriminator_train_step)
callbacks = callbacks or []
for epoch in tqdm(range(epochs)):
for c in callbacks:
c.on_epoch_begin(epoch=epoch + 1, generator=generator, discriminator=discriminator)
for images, labels in tqdm(dataset, leave=False, total=num_batches):
discriminator_step(generator, discriminator, images, labels, latent_dims)
generator_step(generator, discriminator, labels, latent_dims)
for c in callbacks:
c.on_epoch_end(epoch=epoch + 1, generator=generator, discriminator=discriminator)
```
### 2.2 Metrics classifier
Loading the classifier that will be used to calculate the *Classifier Score* (CS) and *Fréchet Classifier Distance* (FCD). \
The features of the real data are also precalculated to avoid doing that for each epoch.
```
classifier = tf.keras.models.load_model('../../Classifiers/mnist.h5')
feature_layer = classifier.get_layer('features')
logits_layer = classifier.get_layer('logits')
precalculated_features = utils.fn.calculate_features(classifier, feature_layer, data)
```
### 2.3 Hyperparameter Testing
This function will overload the function of the same name in the MetricsCallback instance, this is because the default for this class does not generate the labels as input.
```
def get_random_inputs(n_samples):
seeds = tf.random.normal((n_samples, LATENT_DIMS))
labels = tf.random.uniform(
shape=(n_samples, 1),
minval=0, maxval=NUM_CLASSES,
dtype=tf.int32
)
return [seeds, labels]
```
These were the hyperparameters tested for the final document. Training all of them simultaneously may take a long time, consider commenting out some options to run the tests individually.
```
LATENT_DIMS = 24
hparams_list = [
{'batch_size': 16, 'smooth_labels': False},
{'batch_size': 32, 'smooth_labels': False},
{'batch_size': 16, 'smooth_labels': True},
{'batch_size': 32, 'smooth_labels': True}
]
for hparams in hparams_list:
dirname = 'BS{}{}'.format(
hparams['batch_size'],
'_SMOOTH' if hparams['smooth_labels'] else ''
)
Path(dirname).mkdir(exist_ok=True)
generator = generator_model(LATENT_DIMS)
generator.optimizer = tf.keras.optimizers.Adam(1e-4)
discriminator = discriminator_model()
discriminator.optimizer = tf.keras.optimizers.Adam(1e-4)
discriminator_loss = discriminator_loss_smooth if hparams['smooth_labels'] else discriminator_loss_normal
## Callbacks
timer = utils.callback.TimerCallback()
save_samples = utils.callback.SaveSamplesCallback(
path_format=os.path.join(dirname, 'epoch-{}'),
inputs= [
tf.random.normal((10*10, LATENT_DIMS)),
np.expand_dims(np.repeat(np.arange(10), 10, axis=0), -1)
],
n_cols=10,
savefig_kwargs={'bbox_inches': 'tight', 'pad_inches': 0, 'dpi': 192},
imshow_kwargs={'cmap': 'gray_r', 'vmin': -1, 'vmax': 1}
)
metrics = utils.callback.MetricsCallback(
generator=generator,
classifier=classifier,
latent_dims=LATENT_DIMS,
feature_layer=feature_layer,
logits_layer=logits_layer,
precalculated_features=precalculated_features,
save_after=5, save_to=os.path.join(dirname, 'best.h5')
)
metrics.get_random_inputs = get_random_inputs #overloading default function
## Train and save results
train(
generator, discriminator, data, labels, epochs=30,
batch_size=hparams['batch_size'],
callbacks=[timer, save_samples, metrics]
)
metrics_obj = metrics.get_metrics()
metrics_obj['time'] = timer.get_time()
utils.fn.update_json_log(os.path.join(dirname, 'log.json'), metrics_obj)
generator.save (os.path.join(dirname, 'generator.h5' ), overwrite=True, save_format='h5')
discriminator.save(os.path.join(dirname, 'discriminator.h5'), overwrite=True, save_format='h5')
```
\
In windows the command bellow is used to turn down the machine after the training finishes, very useful if you wanna let the computer running while you go to sleep :)
```
# !shutdown /s /t 60
```
| github_jupyter |
### Imports
```
import matplotlib.pyplot as plt
import tensorflow as tf
import numpy as np
from sklearn.metrics import confusion_matrix
%matplotlib inline
%pylab inline
pylab.rcParams['figure.figsize'] = (15, 10)
```
### Load Data
```
from tensorflow.examples.tutorials.mnist import input_data
data = input_data.read_data_sets("data/MNIST/", one_hot=True)
print("Shape of:")
print("- Training-set:\t\t{}".format(data.train.labels.shape))
print("- Test-set:\t\t{}".format(data.test.labels.shape))
print("- Validation-set:\t{}".format(data.validation.labels.shape))
```
### One-Hot Encoding
```
data.test.labels[:5]
data.test.cls = np.array([label.argmax() for label in data.test.labels])
data.test.cls[:5]
```
### Data Dimensions
```
img_size = 28
img_size_flat = img_size * img_size
img_shape = (img_size, img_size)
num_classes = 10
```
### Helper function for plotting images
```
def plot_images(images, generated_images=None):
assert len(images) == 5
# Create figure with 3x3 subplots
if generated_images is None:
fig, axes = plt.subplots(1, 5)
else:
fig, axes = plt.subplots(2, 5)
for i, ax in enumerate(axes.flat):
if i < 5:
ax.imshow(images[i].reshape(img_shape), cmap='gray')
else:
if generated_images is None:
break
ax.imshow(generated_images[i - 5].reshape(img_shape), cmap='gray')
ax.set_xticks([])
ax.set_yticks([])
```
### Plot a few images to see if data is correct
```
plot_images(data.test.images[:5])
```
### Define Computational Graph
#### Helper function for adding layer
```
def add_layer(_input_layer, num_neurons, use_relu=True):
# Assume _input_layer is of shape (None, prev_neurons)
prev_neurons = _input_layer.shape[1]
weights = tf.Variable(np.float32(np.random.randn(prev_neurons, num_neurons) * 1e-3))
biases = tf.Variable(tf.zeros((num_neurons)))
layer = tf.matmul(_input_layer, weights) + biases
if use_relu:
layer = tf.nn.relu(layer)
return layer, weights, biases
```
#### Placeholders
```
batch_size = 100
x = tf.placeholder(tf.float32, shape=(None, num_classes))
y_true = tf.placeholder(tf.float32, shape=(None, img_size_flat))
#from tensorflow.contrib.image import rotate
#rotation_angles = (np.random.rand(batch_size) * 2 - 1) * 20
#x = rotate(tf.reshape(x, (-1, img_size, img_size, 1)), rotation_angles)
#x = tf.reshape(x, (-1, img_size_flat))
```
#### Model
```
layer_1, weights, biases = add_layer(x, 25)
logits, _, _ = add_layer(layer_1, img_size_flat)
y_pred = logits
```
#### Cost function to be optimized
```
error = (y_pred - y_true)**2
loss = tf.reduce_mean(error)
```
#### Optimization Method
```
optimizer = tf.train.AdamOptimizer(learning_rate=0.05).minimize(loss)
```
### Tensorflow Run
#### Create Tensorflow Session
```
session = tf.Session()
```
#### Initialize Variables
```
session.run(tf.global_variables_initializer())
```
#### Helper Function to perform optimization iterations
```
def plot_performance():
image_idx = np.random.randint(0, data.test.num_examples, 5)
original_images = data.test.images[image_idx]
generated_images = session.run(y_pred, {x: data.test.labels[image_idx], y_true: data.test.images[image_idx]})
plot_images(original_images, generated_images)
def optimize(num_iterations=1):
plot_performance()
for i in range(num_iterations):
# Get a batch of training examples
y_true_batch, x_batch = data.train.next_batch(batch_size)
y_true_batch_cls = np.array([label.argmax() for label in y_true_batch])
# Create feed-dict to send it to the graph run
feed_dict_train = {x: x_batch, y_true: y_true_batch}
# Run the optimizer
session.run(optimizer, feed_dict_train)
if i % int(55000 / batch_size) == 0:
plot_performance()
cost = session.run(loss, feed_dict_train)
print("Epoch: {}, loss=".format(int(i / int(55000 / batch_size)) + 1, cost))
optimize(10000)
vectors = np.eye(10)
generated_images = session.run(y_pred, {x: vectors})
plot_images(generated_images[:5], generated_images[5:])
hybrid_vectors = np.zeros((10, 10))
hybrid_vectors[range(10), np.random.randint(0, 10, 10)] += 0.5
hybrid_vectors[range(10), np.random.randint(0, 10, 10)] += 0.5
hybrid_images = session.run(y_pred, {x: hybrid_vectors})
plot_images(hybrid_images[:5], hybrid_images[5:])
print(hybrid_vectors)
random_vectors = np.random.randn(10, 10)
random_images = session.run(y_pred, {x: random_vectors})
plot_images(random_images[:5], random_images[5:])
print(np.round(random_vectors, 2))
session.close()
```
| github_jupyter |
Mount my google drive, where I stored the dataset.
```
from google.colab import drive
drive.mount('/content/drive')
```
**Download dependencies**
```
!pip3 install sklearn matplotlib GPUtil
```
**Download Data**
In order to acquire the dataset please navigate to:
https://ieee-dataport.org/documents/cervigram-image-dataset
Unzip the dataset into the folder "dataset".
For your environment, please adjust the paths accordingly.
```
!rm -vrf "dataset"
!mkdir "dataset"
!cp -r "/content/drive/My Drive/Studiu doctorat leziuni cervicale/cervigram-image-dataset-v2.zip" "dataset/cervigram-image-dataset-v2.zip"
# !cp -r "cervigram-image-dataset-v2.zip" "dataset/cervigram-image-dataset-v2.zip"
!unzip "dataset/cervigram-image-dataset-v2.zip" -d "dataset"
```
**Constants**
For your environment, please modify the paths accordingly.
```
TRAIN_PATH = '/content/dataset/data/train/'
TEST_PATH = '/content/dataset/data/test/'
CROP_SIZE = 260
IMAGE_SIZE = 224
BATCH_SIZE = 150
```
**Imports**
```
import torch as t
import torchvision as tv
import numpy as np
import PIL as pil
import matplotlib.pyplot as plt
from torchvision.datasets import ImageFolder
from torch.utils.data import DataLoader
from torch.nn import Linear, BCEWithLogitsLoss
import sklearn as sk
import sklearn.metrics
from os import listdir
import time
import random
import GPUtil
```
**Memory Stats**
```
import GPUtil
def memory_stats():
for gpu in GPUtil.getGPUs():
print("GPU RAM Free: {0:.0f}MB | Used: {1:.0f}MB | Util {2:3.0f}% | Total {3:.0f}MB".format(gpu.memoryFree, gpu.memoryUsed, gpu.memoryUtil*100, gpu.memoryTotal))
memory_stats()
```
**Deterministic Measurements**
This statements help making the experiments reproducible by fixing the random seeds. Despite fixing the random seeds, experiments are usually not reproducible using different PyTorch releases, commits, platforms or between CPU and GPU executions. Please find more details in the PyTorch documentation:
https://pytorch.org/docs/stable/notes/randomness.html
```
SEED = 0
t.manual_seed(SEED)
t.cuda.manual_seed(SEED)
t.backends.cudnn.deterministic = True
t.backends.cudnn.benchmark = False
np.random.seed(SEED)
random.seed(SEED)
```
**Loading Data**
The dataset is structured in multiple small folders of 7 images each. This generator iterates through the folders and returns the category and 7 paths: one for each image in the folder. The paths are ordered; the order is important since each folder contains 3 types of images, first 5 are with acetic acid solution and the last two are through a green lens and having iodine solution(a solution of a dark red color).
```
def sortByLastDigits(elem):
chars = [c for c in elem if c.isdigit()]
return 0 if len(chars) == 0 else int(''.join(chars))
def getImagesPaths(root_path):
for class_folder in [root_path + f for f in listdir(root_path)]:
category = int(class_folder[-1])
for case_folder in listdir(class_folder):
case_folder_path = class_folder + '/' + case_folder + '/'
img_files = [case_folder_path + file_name for file_name in listdir(case_folder_path)]
yield category, sorted(img_files, key = sortByLastDigits)
```
We define 3 datasets, which load 3 kinds of images: natural images, images taken through a green lens and images where the doctor applied iodine solution (which gives a dark red color). Each dataset has dynamic and static transformations which could be applied to the data. The static transformations are applied on the initialization of the dataset, while the dynamic ones are applied when loading each batch of data.
```
class SimpleImagesDataset(t.utils.data.Dataset):
def __init__(self, root_path, transforms_x_static = None, transforms_x_dynamic = None, transforms_y_static = None, transforms_y_dynamic = None):
self.dataset = []
self.transforms_x = transforms_x_dynamic
self.transforms_y = transforms_y_dynamic
for category, img_files in getImagesPaths(root_path):
for i in range(1, 5):
img = pil.Image.open(img_files[i])
if transforms_x_static != None:
img = transforms_x_static(img)
if transforms_y_static != None:
category = transforms_y_static(category)
self.dataset.append((img, category))
def __getitem__(self, i):
x, y = self.dataset[i]
if self.transforms_x != None:
x = self.transforms_x(x)
if self.transforms_y != None:
y = self.transforms_y(y)
return x, y
def __len__(self):
return len(self.dataset)
class GreenLensImagesDataset(SimpleImagesDataset):
def __init__(self, root_path, transforms_x_static = None, transforms_x_dynamic = None, transforms_y_static = None, transforms_y_dynamic = None):
self.dataset = []
self.transforms_x = transforms_x_dynamic
self.transforms_y = transforms_y_dynamic
for category, img_files in getImagesPaths(root_path):
# Only the green lens image
img = pil.Image.open(img_files[-2])
if transforms_x_static != None:
img = transforms_x_static(img)
if transforms_y_static != None:
category = transforms_y_static(category)
self.dataset.append((img, category))
class RedImagesDataset(SimpleImagesDataset):
def __init__(self, root_path, transforms_x_static = None, transforms_x_dynamic = None, transforms_y_static = None, transforms_y_dynamic = None):
self.dataset = []
self.transforms_x = transforms_x_dynamic
self.transforms_y = transforms_y_dynamic
for category, img_files in getImagesPaths(root_path):
# Only the green lens image
img = pil.Image.open(img_files[-1])
if transforms_x_static != None:
img = transforms_x_static(img)
if transforms_y_static != None:
category = transforms_y_static(category)
self.dataset.append((img, category))
```
**Preprocess Data**
Convert pytorch tensor to numpy array.
```
def to_numpy(x):
return x.cpu().detach().numpy()
```
Data transformations for the test and training sets.
```
norm_mean = [0.485, 0.456, 0.406]
norm_std = [0.229, 0.224, 0.225]
transforms_train = tv.transforms.Compose([
tv.transforms.RandomAffine(degrees = 45, translate = None, scale = (1., 2.), shear = 30),
# tv.transforms.CenterCrop(CROP_SIZE),
tv.transforms.Resize(IMAGE_SIZE),
tv.transforms.RandomHorizontalFlip(),
tv.transforms.ToTensor(),
tv.transforms.Lambda(lambda t: t.cuda()),
tv.transforms.Normalize(mean=norm_mean, std=norm_std)
])
transforms_test = tv.transforms.Compose([
# tv.transforms.CenterCrop(CROP_SIZE),
tv.transforms.Resize(IMAGE_SIZE),
tv.transforms.ToTensor(),
tv.transforms.Normalize(mean=norm_mean, std=norm_std)
])
y_transform = tv.transforms.Lambda(lambda y: t.tensor(y, dtype=t.long, device = 'cuda:0'))
```
Initialize pytorch datasets and loaders for training and test.
```
def create_loaders(dataset_class):
dataset_train = dataset_class(TRAIN_PATH, transforms_x_dynamic = transforms_train, transforms_y_dynamic = y_transform)
dataset_test = dataset_class(TEST_PATH, transforms_x_static = transforms_test,
transforms_x_dynamic = tv.transforms.Lambda(lambda t: t.cuda()), transforms_y_dynamic = y_transform)
loader_train = DataLoader(dataset_train, BATCH_SIZE, shuffle = True, num_workers = 0)
loader_test = DataLoader(dataset_test, BATCH_SIZE, shuffle = False, num_workers = 0)
return loader_train, loader_test, len(dataset_train), len(dataset_test)
loader_train_simple_img, loader_test_simple_img, len_train, len_test = create_loaders(GreenLensImagesDataset)
```
**Visualize Data**
Load a few images so that we can see the effects of the data augmentation on the training set.
```
def plot_one_prediction(x, label, pred):
x, label, pred = to_numpy(x), to_numpy(label), to_numpy(pred)
x = np.transpose(x, [1, 2, 0])
if x.shape[-1] == 1:
x = x.squeeze()
x = x * np.array(norm_std) + np.array(norm_mean)
plt.title(label, color = 'green' if label == pred else 'red')
plt.imshow(x)
def plot_predictions(imgs, labels, preds):
fig = plt.figure(figsize = (20, 5))
for i in range(20):
fig.add_subplot(2, 10, i + 1, xticks = [], yticks = [])
plot_one_prediction(imgs[i], labels[i], preds[i])
# x, y = next(iter(loader_train_simple_img))
# plot_predictions(x, y, y)
```
**Model**
Define a few models to experiment with.
```
def get_mobilenet_v2():
model = t.hub.load('pytorch/vision', 'mobilenet_v2', pretrained=True)
# model.classifier[0] = t.nn.Dropout(p=0.9, inplace=False)
# model.classifier[1] = Linear(in_features=1280, out_features=4, bias=True)
# model.features[18].add_module('cnn_drop_18', t.nn.Dropout2d(p = .3))
# model.features[17]._modules['conv'][1].add_module('cnn_drop_17', t.nn.Dropout2d(p = .2))
# model.features[16]._modules['conv'][1].add_module('cnn_drop_16', t.nn.Dropout2d(p = .1))
model = model.cuda()
return model
def get_vgg_19():
model = tv.models.vgg19(pretrained = True)
model = model.cuda()
model.classifier[2].p = .2
model.classifier[6].out_features = 4
return model
def get_res_next_101():
model = t.hub.load('facebookresearch/WSL-Images', 'resnext101_32x8d_wsl')
model.fc = t.nn.Sequential(
t.nn.Dropout(p = .9),
t.nn.Linear(in_features=2048, out_features=4)
)
model = model.cuda()
return model
def get_resnet_18():
model = tv.models.resnet18(pretrained = True)
model.fc = t.nn.Sequential(
t.nn.Dropout(p = .9),
t.nn.Linear(in_features=512, out_features=4)
)
model = model.cuda()
return model
def get_dense_net():
model = tv.models.densenet121(pretrained = True)
model.classifier = t.nn.Sequential(
t.nn.Dropout(p = .9),
t.nn.Linear(in_features = 1024, out_features = 4)
)
model = model.cuda()
return model
class MobileNetV2_FullConv(t.nn.Module):
def __init__(self):
super().__init__()
self.cnn = get_mobilenet_v2().features
self.cnn[18] = t.nn.Sequential(
tv.models.mobilenet.ConvBNReLU(320, 64, kernel_size=1),
t.nn.Dropout2d(p = .7)
)
# self.fc = t.nn.Sequential(
# t.nn.Flatten(),
# t.nn.Dropout(0.4),
# t.nn.Linear(8 * 7 * 10, 4),
# )
self.fc = t.nn.Linear(64, 4)
def forward(self, x):
x = self.cnn(x)
x = x.mean([2, 3])
x = self.fc(x);
return x
model_simple = MobileNetV2_FullConv().cuda()
```
**Train & Evaluate**
Timer utility function. This is used to measure the execution speed.
```
time_start = 0
def timer_start():
global time_start
time_start = time.time()
def timer_end():
return time.time() - time_start
```
This function trains the network and evaluates it at the same time. It outputs the metrics recorded during the training for both train and test. We are measuring accuracy and the loss. The function also saves a checkpoint of the model every time the accuracy is improved. In the end we will have a checkpoint of the model which gave the best accuracy.
```
def train_eval(optimizer, model, loader_train, loader_test, chekpoint_name, epochs):
metrics = {
'losses_train': [],
'losses_test': [],
'acc_train': [],
'acc_test': [],
'prec_train': [],
'prec_test': [],
'rec_train': [],
'rec_test': [],
'f_score_train': [],
'f_score_test': []
}
best_acc = 0
loss_fn = t.nn.CrossEntropyLoss()
for epoch in range(epochs):
timer_start()
train_epoch_loss, train_epoch_acc, train_epoch_precision, train_epoch_recall, train_epoch_f_score = 0, 0, 0, 0, 0
test_epoch_loss, test_epoch_acc, test_epoch_precision, test_epoch_recall, test_epoch_f_score = 0, 0, 0, 0, 0
# Train
model.train()
for x, y in loader_train:
y_pred = model.forward(x)
loss = loss_fn(y_pred, y)
loss.backward()
optimizer.step()
# memory_stats()
optimizer.zero_grad()
y_pred, y = to_numpy(y_pred), to_numpy(y)
pred = y_pred.argmax(axis = 1)
ratio = len(y) / len_train
train_epoch_loss += (loss.item() * ratio)
train_epoch_acc += (sk.metrics.classification.accuracy_score(y, pred) * ratio)
precision, recall, f_score, _ = sk.metrics.classification.precision_recall_fscore_support(y, pred, average = 'macro')
train_epoch_precision += (precision * ratio)
train_epoch_recall += (recall * ratio)
train_epoch_f_score += (f_score * ratio)
metrics['losses_train'].append(train_epoch_loss)
metrics['acc_train'].append(train_epoch_acc)
metrics['prec_train'].append(train_epoch_precision)
metrics['rec_train'].append(train_epoch_recall)
metrics['f_score_train'].append(train_epoch_f_score)
# Evaluate
model.eval()
with t.no_grad():
for x, y in loader_test:
y_pred = model.forward(x)
loss = loss_fn(y_pred, y)
y_pred, y = to_numpy(y_pred), to_numpy(y)
pred = y_pred.argmax(axis = 1)
ratio = len(y) / len_test
test_epoch_loss += (loss * ratio)
test_epoch_acc += (sk.metrics.classification.accuracy_score(y, pred) * ratio )
precision, recall, f_score, _ = sk.metrics.classification.precision_recall_fscore_support(y, pred, average = 'macro')
test_epoch_precision += (precision * ratio)
test_epoch_recall += (recall * ratio)
test_epoch_f_score += (f_score * ratio)
metrics['losses_test'].append(test_epoch_loss)
metrics['acc_test'].append(test_epoch_acc)
metrics['prec_test'].append(test_epoch_precision)
metrics['rec_test'].append(test_epoch_recall)
metrics['f_score_test'].append(test_epoch_f_score)
if metrics['acc_test'][-1] > best_acc:
best_acc = metrics['acc_test'][-1]
t.save({'model': model.state_dict()}, 'checkpint {}.tar'.format(chekpoint_name))
print('Epoch {} acc {} prec {} rec {} f {} minutes {}'.format(
epoch + 1, metrics['acc_test'][-1], metrics['prec_test'][-1], metrics['rec_test'][-1], metrics['f_score_test'][-1], timer_end() / 60))
print('Ended training')
return metrics
```
Plot a metric for both train and test.
```
def plot_train_test(train, test, title, y_title):
plt.plot(range(len(train)), train, label = 'train')
plt.plot(range(len(test)), test, label = 'test')
plt.xlabel('Epochs')
plt.ylabel(y_title)
plt.title(title)
plt.legend()
plt.show()
```
Plot precision - recall curve
```
def plot_precision_recall(metrics):
plt.scatter(metrics['prec_train'], metrics['rec_train'], label = 'train')
plt.scatter(metrics['prec_test'], metrics['rec_test'], label = 'test')
plt.legend()
plt.title('Precision-Recall')
plt.xlabel('Precision')
plt.ylabel('Recall')
```
Train a model for several epochs. The steps_learning parameter is a list of tuples. Each tuple specifies the steps and the learning rate.
```
def do_train(model, loader_train, loader_test, checkpoint_name, steps_learning):
for steps, learn_rate in steps_learning:
metrics = train_eval(t.optim.Adam(model.parameters(), lr = learn_rate, weight_decay = 0), model, loader_train, loader_test, checkpoint_name, steps)
print('Best test accuracy :', max(metrics['acc_test']))
plot_train_test(metrics['losses_train'], metrics['losses_test'], 'Loss (lr = {})'.format(learn_rate))
plot_train_test(metrics['acc_train'], metrics['acc_test'], 'Accuracy (lr = {})'.format(learn_rate))
```
Perform actual training.
```
def do_train(model, loader_train, loader_test, checkpoint_name, steps_learning):
t.cuda.empty_cache()
for steps, learn_rate in steps_learning:
metrics = train_eval(t.optim.Adam(model.parameters(), lr = learn_rate, weight_decay = 0), model, loader_train, loader_test, checkpoint_name, steps)
index_max = np.array(metrics['acc_test']).argmax()
print('Best test accuracy :', metrics['acc_test'][index_max])
print('Corresponding precision :', metrics['prec_test'][index_max])
print('Corresponding recall :', metrics['rec_test'][index_max])
print('Corresponding f1 score :', metrics['f_score_test'][index_max])
plot_train_test(metrics['losses_train'], metrics['losses_test'], 'Loss (lr = {})'.format(learn_rate), 'Loss')
plot_train_test(metrics['acc_train'], metrics['acc_test'], 'Accuracy (lr = {})'.format(learn_rate), 'Accuracy')
plot_train_test(metrics['prec_train'], metrics['prec_test'], 'Precision (lr = {})'.format(learn_rate), 'Precision')
plot_train_test(metrics['rec_train'], metrics['rec_test'], 'Recall (lr = {})'.format(learn_rate), 'Recall')
plot_train_test(metrics['f_score_train'], metrics['f_score_test'], 'F1 Score (lr = {})'.format(learn_rate), 'F1 Score')
plot_precision_recall(metrics)
do_train(model_simple, loader_train_simple_img, loader_test_simple_img, 'simple_1', [(300, 1e-4)])
```
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W2D3_BiologicalNeuronModels/student/W2D3_Tutorial2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Tutorial 2: Effects of Input Correlation
**Week 2, Day 3: Biological Neuron Models**
**By Neuromatch Academy**
__Content creators:__ Qinglong Gu, Songtin Li, John Murray, Richard Naud, Arvind Kumar
__Content reviewers:__ Maryam Vaziri-Pashkam, Ella Batty, Lorenzo Fontolan, Richard Gao, Matthew Krause, Spiros Chavlis, Michael Waskom
**Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
<p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
---
# Tutorial Objectives
In this tutorial, we will use the leaky integrate-and-fire (LIF) neuron model (see Tutorial 1) to study how they transform input correlations to output properties (transfer of correlations). In particular, we are going to write a few lines of code to:
- inject correlated GWN in a pair of neurons
- measure correlations between the spiking activity of the two neurons
- study how the transfer of correlation depends on the statistics of the input, i.e. mean and standard deviation.
---
# Setup
```
# Import libraries
import matplotlib.pyplot as plt
import numpy as np
import time
# @title Figure Settings
import ipywidgets as widgets # interactive display
%config InlineBackend.figure_format = 'retina'
# use NMA plot style
plt.style.use("https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/nma.mplstyle")
my_layout = widgets.Layout()
# @title Helper functions
def default_pars(**kwargs):
pars = {}
### typical neuron parameters###
pars['V_th'] = -55. # spike threshold [mV]
pars['V_reset'] = -75. # reset potential [mV]
pars['tau_m'] = 10. # membrane time constant [ms]
pars['g_L'] = 10. # leak conductance [nS]
pars['V_init'] = -75. # initial potential [mV]
pars['V_L'] = -75. # leak reversal potential [mV]
pars['tref'] = 2. # refractory time (ms)
### simulation parameters ###
pars['T'] = 400. # Total duration of simulation [ms]
pars['dt'] = .1 # Simulation time step [ms]
### external parameters if any ###
for k in kwargs:
pars[k] = kwargs[k]
pars['range_t'] = np.arange(0, pars['T'], pars['dt']) # Vector of discretized
# time points [ms]
return pars
def run_LIF(pars, Iinj):
"""
Simulate the LIF dynamics with external input current
Args:
pars : parameter dictionary
Iinj : input current [pA]. The injected current here can be a value or an array
Returns:
rec_spikes : spike times
rec_v : mebrane potential
"""
# Set parameters
V_th, V_reset = pars['V_th'], pars['V_reset']
tau_m, g_L = pars['tau_m'], pars['g_L']
V_init, V_L = pars['V_init'], pars['V_L']
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
tref = pars['tref']
# Initialize voltage and current
v = np.zeros(Lt)
v[0] = V_init
Iinj = Iinj * np.ones(Lt)
tr = 0.
# simulate the LIF dynamics
rec_spikes = [] # record spike times
for it in range(Lt - 1):
if tr > 0:
v[it] = V_reset
tr = tr - 1
elif v[it] >= V_th: # reset voltage and record spike event
rec_spikes.append(it)
v[it] = V_reset
tr = tref / dt
# calculate the increment of the membrane potential
dv = (-(v[it] - V_L) + Iinj[it] / g_L) * (dt / tau_m)
# update the membrane potential
v[it + 1] = v[it] + dv
rec_spikes = np.array(rec_spikes) * dt
return v, rec_spikes
def my_GWN(pars, sig, myseed=False):
"""
Function that calculates Gaussian white noise inputs
Args:
pars : parameter dictionary
mu : noise baseline (mean)
sig : noise amplitute (standard deviation)
myseed : random seed. int or boolean
the same seed will give the same random number sequence
Returns:
I : Gaussian white noise input
"""
# Retrieve simulation parameters
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
# Set random seed. You can fix the seed of the random number generator so
# that the results are reliable however, when you want to generate multiple
# realization make sure that you change the seed for each new realization
if myseed:
np.random.seed(seed=myseed)
else:
np.random.seed()
# generate GWN
# we divide here by 1000 to convert units to sec.
I_GWN = sig * np.random.randn(Lt) * np.sqrt(pars['tau_m'] / dt)
return I_GWN
def Poisson_generator(pars, rate, n, myseed=False):
"""
Generates poisson trains
Args:
pars : parameter dictionary
rate : noise amplitute [Hz]
n : number of Poisson trains
myseed : random seed. int or boolean
Returns:
pre_spike_train : spike train matrix, ith row represents whether
there is a spike in ith spike train over time
(1 if spike, 0 otherwise)
"""
# Retrieve simulation parameters
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
# set random seed
if myseed:
np.random.seed(seed=myseed)
else:
np.random.seed()
# generate uniformly distributed random variables
u_rand = np.random.rand(n, Lt)
# generate Poisson train
poisson_train = 1. * (u_rand < rate * (dt / 1000.))
return poisson_train
def example_plot_myCC():
pars = default_pars(T=50000, dt=.1)
c = np.arange(10) * 0.1
r12 = np.zeros(10)
for i in range(10):
I1gL, I2gL = correlate_input(pars, mu=20.0, sig=7.5, c=c[i])
r12[i] = my_CC(I1gL, I2gL)
plt.figure()
plt.plot(c, r12, 'bo', alpha=0.7, label='Simulation', zorder=2)
plt.plot([-0.05, 0.95], [-0.05, 0.95], 'k--', label='y=x',
dashes=(2, 2), zorder=1)
plt.xlabel('True CC')
plt.ylabel('Sample CC')
plt.legend(loc='best')
def LIF_output_cc(pars, mu, sig, c, bin_size, n_trials=20):
""" Simulates two LIF neurons with correlated input and computes output correlation
Args:
pars : parameter dictionary
mu : noise baseline (mean)
sig : noise amplitute (standard deviation)
c : correlation coefficient ~[0, 1]
bin_size : bin size used for time series
n_trials : total simulation trials
Returns:
r : output corr. coe.
sp_rate : spike rate
sp1 : spike times of neuron 1 in the last trial
sp2 : spike times of neuron 2 in the last trial
"""
r12 = np.zeros(n_trials)
sp_rate = np.zeros(n_trials)
for i_trial in range(n_trials):
I1gL, I2gL = correlate_input(pars, mu, sig, c)
_, sp1 = run_LIF(pars, pars['g_L'] * I1gL)
_, sp2 = run_LIF(pars, pars['g_L'] * I2gL)
my_bin = np.arange(0, pars['T'], bin_size)
sp1_count, _ = np.histogram(sp1, bins=my_bin)
sp2_count, _ = np.histogram(sp2, bins=my_bin)
r12[i_trial] = my_CC(sp1_count[::20], sp2_count[::20])
sp_rate[i_trial] = len(sp1) / pars['T'] * 1000.
return r12.mean(), sp_rate.mean(), sp1, sp2
def plot_c_r_LIF(c, r, mycolor, mylabel):
z = np.polyfit(c, r, deg=1)
c_range = np.array([c.min() - 0.05, c.max() + 0.05])
plt.plot(c, r, 'o', color=mycolor, alpha=0.7, label=mylabel, zorder=2)
plt.plot(c_range, z[0] * c_range + z[1], color=mycolor, zorder=1)
```
The helper function contains the:
- Parameter dictionary: `default_pars( **kwargs)`
- LIF simulator: `run_LIF`
- Gaussian white noise generator: `my_GWN(pars, sig, myseed=False)`
- Poisson type spike train generator: `Poisson_generator(pars, rate, n, myseed=False)`
- Two LIF neurons with correlated inputs simulator: `LIF_output_cc(pars, mu, sig, c, bin_size, n_trials=20)`
- Some additional plotting utilities
---
# Section 1: Correlations (Synchrony)
Correlation or synchrony in neuronal activity can be described for any readout of brain activity. Here, we are concerned with the spiking activity of neurons.
In the simplest way, correlation/synchrony refers to coincident spiking of neurons, i.e., when two neurons spike together, they are firing in **synchrony** or are **correlated**. Neurons can be synchronous in their instantaneous activity, i.e., they spike together with some probability. However, it is also possible that spiking of a neuron at time $t$ is correlated with the spikes of another neuron with a delay (time-delayed synchrony).
## Origin of synchronous neuronal activity:
- Common inputs, i.e., two neurons are receiving input from the same sources. The degree of correlation of the shared inputs is proportional to their output correlation.
- Pooling from the same sources. Neurons do not share the same input neurons but are receiving inputs from neurons which themselves are correlated.
- Neurons are connected to each other (uni- or bi-directionally): This will only give rise to time-delayed synchrony. Neurons could also be connected via gap-junctions.
- Neurons have similar parameters and initial conditions.
## Implications of synchrony
When neurons spike together, they can have a stronger impact on downstream neurons. Synapses in the brain are sensitive to the temporal correlations (i.e., delay) between pre- and postsynaptic activity, and this, in turn, can lead to the formation of functional neuronal networks - the basis of unsupervised learning (we will study some of these concepts in a forthcoming tutorial).
Synchrony implies a reduction in the dimensionality of the system. In addition, correlations, in many cases, can impair the decoding of neuronal activity.
```
# @title Video 1: Input & output correlations
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="nsAYFBcAkes", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
## How to study the emergence of correlations
A simple model to study the emergence of correlations is to inject common inputs to a pair of neurons and measure the output correlation as a function of the fraction of common inputs.
Here, we are going to investigate the transfer of correlations by computing the correlation coefficient of spike trains recorded from two unconnected LIF neurons, which received correlated inputs.
The input current to LIF neuron $i$ $(i=1,2)$ is:
\begin{equation}
\frac{I_i}{g_L} =\mu_i + \sigma_i (\sqrt{1-c}\xi_i + \sqrt{c}\xi_c) \quad (1)
\end{equation}
where $\mu_i$ is the temporal average of the current. The Gaussian white noise $\xi_i$ is independent for each neuron, while $\xi_c$ is common to all neurons. The variable $c$ ($0\le c\le1$) controls the fraction of common and independent inputs. $\sigma_i$ shows the variance of the total input.
So, first, we will generate correlated inputs.
```
# @title
#@markdown Execute this cell to get a function for generating correlated GWN inputs
def correlate_input(pars, mu=20., sig=7.5, c=0.3):
"""
Args:
pars : parameter dictionary
mu : noise baseline (mean)
sig : noise amplitute (standard deviation)
c. : correlation coefficient ~[0, 1]
Returns:
I1gL, I2gL : two correlated inputs with corr. coe. c
"""
# generate Gaussian whute noise xi_1, xi_2, xi_c
xi_1 = my_GWN(pars, sig)
xi_2 = my_GWN(pars, sig)
xi_c = my_GWN(pars, sig)
# Generate two correlated inputs by Equation. (1)
I1gL = mu + np.sqrt(1. - c) * xi_1 + np.sqrt(c) * xi_c
I2gL = mu + np.sqrt(1. - c) * xi_2 + np.sqrt(c) * xi_c
return I1gL, I2gL
print(help(correlate_input))
```
### Exercise 1: Compute the correlation
The _sample correlation coefficient_ between two input currents $I_i$ and $I_j$ is defined as the sample covariance of $I_i$ and $I_j$ divided by the square root of the sample variance of $I_i$ multiplied with the square root of the sample variance of $I_j$. In equation form:
\begin{align}
r_{ij} &= \frac{cov(I_i, I_j)}{\sqrt{var(I_i)} \sqrt{var(I_j)}}\\
cov(I_i, I_j) &= \sum_{k=1}^L (I_i^k -\bar{I}_i)(I_j^k -\bar{I}_j) \\
var(I_i) &= \sum_{k=1}^L (I_i^k -\bar{I}_i)^2
\end{align}
where $\bar{I}_i$ is the sample mean, k is the time bin, and L is the length of $I$. This means that $I_i^k$ is current i at time $k\cdot dt$. Note that the equations above are not accurate for sample covariances and variances as they should be additionally divided by L-1 - we have dropped this term because it cancels out in the sample correlation coefficient formula.
The _sample correlation coefficient_ may also be referred to as the _sample Pearson correlation coefficient_. Here, is a beautiful paper that explains multiple ways to calculate and understand correlations [Rodgers and Nicewander 1988](https://www.stat.berkeley.edu/~rabbee/correlation.pdf).
In this exercise, we will create a function, `my_CC` to compute the sample correlation coefficient between two time series. Note that while we introduced this computation here in the context of input currents, the sample correlation coefficient is used to compute the correlation between any two time series - we will use it later on binned spike trains.
```
def my_CC(i, j):
"""
Args:
i, j : two time series with the same length
Returns:
rij : correlation coefficient
"""
########################################################################
## TODO for students: compute rxy, then remove the NotImplementedError #
# Tip1: array([a1, a2, a3])*array([b1, b2, b3]) = array([a1*b1, a2*b2, a3*b3])
# Tip2: np.sum(array([a1, a2, a3])) = a1+a2+a3
# Tip3: square root, np.sqrt()
# Fill out function and remove
raise NotImplementedError("Student exercise: compute the sample correlation coefficient")
########################################################################
# Calculate the covariance of i and j
cov = ...
# Calculate the variance of i
var_i = ...
# Calculate the variance of j
var_j = ...
# Calculate the correlation coefficient
rij = ...
return rij
# Uncomment the line after completing the my_CC function
# example_plot_myCC()
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D3_BiologicalNeuronModels/solutions/W2D3_Tutorial2_Solution_03e44bdc.py)
*Example output:*
<img alt='Solution hint' align='left' width=558 height=413 src=https://raw.githubusercontent.com/NeuromatchAcademy/course-content/master/tutorials/W2D3_BiologicalNeuronModels/static/W2D3_Tutorial2_Solution_03e44bdc_0.png>
### Exercise 2: Measure the correlation between spike trains
After recording the spike times of the two neurons, how can we estimate their correlation coefficient?
In order to find this, we need to bin the spike times and obtain two time series. Each data point in the time series is the number of spikes in the corresponding time bin. You can use `np.histogram()` to bin the spike times.
Complete the code below to bin the spike times and calculate the correlation coefficient for two Poisson spike trains. Note that `c` here is the ground-truth correlation coefficient that we define.
```
# @title
# @markdown Execute this cell to get a function for generating correlated Poisson inputs (generate_corr_Poisson)
def generate_corr_Poisson(pars, poi_rate, c, myseed=False):
"""
function to generate correlated Poisson type spike trains
Args:
pars : parameter dictionary
poi_rate : rate of the Poisson train
c. : correlation coefficient ~[0, 1]
Returns:
sp1, sp2 : two correlated spike time trains with corr. coe. c
"""
range_t = pars['range_t']
mother_rate = poi_rate / c
mother_spike_train = Poisson_generator(pars, rate=mother_rate,
n=1, myseed=myseed)[0]
sp_mother = range_t[mother_spike_train > 0]
L_sp_mother = len(sp_mother)
sp_mother_id = np.arange(L_sp_mother)
L_sp_corr = int(L_sp_mother * c)
np.random.shuffle(sp_mother_id)
sp1 = np.sort(sp_mother[sp_mother_id[:L_sp_corr]])
np.random.shuffle(sp_mother_id)
sp2 = np.sort(sp_mother[sp_mother_id[:L_sp_corr]])
return sp1, sp2
print(help(generate_corr_Poisson))
def corr_coeff_pairs(pars, rate, c, trials, bins):
"""
Calculate the correlation coefficient of two spike trains, for different
realizations
Args:
pars : parameter dictionary
rate : rate of poisson inputs
c : correlation coefficient ~ [0, 1]
trials : number of realizations
bins : vector with bins for time discretization
Returns:
r12 : correlation coefficient of a pair of inputs
"""
r12 = np.zeros(n_trials)
for i in range(n_trials):
##############################################################
## TODO for students: Use np.histogram to bin the spike time #
## e.g., sp1_count, _= np.histogram(...)
# Use my_CC() compute corr coe, compare with c
# Note that you can run multiple realizations and compute their r_12(diff_trials)
# with the defined function above. The average r_12 over trials can get close to c.
# Note: change seed to generate different input per trial
# Fill out function and remove
raise NotImplementedError("Student exercise: compute the correlation coefficient")
##############################################################
# Generate correlated Poisson inputs
sp1, sp2 = generate_corr_Poisson(pars, ..., ..., myseed=2020+i)
# Bin the spike times of the first input
sp1_count, _ = np.histogram(..., bins=...)
# Bin the spike times of the second input
sp2_count, _ = np.histogram(..., bins=...)
# Calculate the correlation coefficient
r12[i] = my_CC(..., ...)
return r12
poi_rate = 20.
c = 0.2 # set true correlation
pars = default_pars(T=10000)
# bin the spike time
bin_size = 20 # [ms]
my_bin = np.arange(0, pars['T'], bin_size)
n_trials = 100 # 100 realizations
# Uncomment to test your function
# r12 = corr_coeff_pairs(pars, rate=poi_rate, c=c, trials=n_trials, bins=my_bin)
# print(f'True corr coe = {c:.3f}')
# print(f'Simu corr coe = {r12.mean():.3f}')
```
Sample output
```
True corr coe = 0.200
Simu corr coe = 0.197
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D3_BiologicalNeuronModels/solutions/W2D3_Tutorial2_Solution_e5eaac3e.py)
---
# Section 2: Investigate the effect of input correlation on the output correlation
Now let's combine the aforementioned two procedures. We first generate the correlated inputs by Equation (1). Then we inject the correlated inputs $I_1, I_2$ into a pair of neurons and record their output spike times. We continue measuring the correlation between the output and
investigate the relationship between the input correlation and the output correlation.
## Drive a neuron with correlated inputs and visualize its output
In the following, you will inject correlated GWN in two neurons. You need to define the mean (`gwn_mean`), standard deviation (`gwn_std`), and input correlations (`c_in`).
We will simulate $10$ trials to get a better estimate of the output correlation. Change the values in the following cell for the above variables (and then run the next cell) to explore how they impact the output correlation.
```
# Play around with these parameters
pars = default_pars(T=80000, dt=1.) # get the parameters
c_in = 0.3 # set input correlation value
gwn_mean = 10.
gwn_std = 10.
# @title
# @markdown Do not forget to execute this cell to simulate the LIF
bin_size = 10. # ms
starttime = time.perf_counter() # time clock
r12_ss, sp_ss, sp1, sp2 = LIF_output_cc(pars, mu=gwn_mean, sig=gwn_std, c=c_in,
bin_size=bin_size, n_trials=10)
# just the time counter
endtime = time.perf_counter()
timecost = (endtime - starttime) / 60.
print(f"Simulation time = {timecost:.2f} min")
print(f"Input correlation = {c_in}")
print(f"Output correlation = {r12_ss}")
plt.figure(figsize=(12, 6))
plt.plot(sp1, np.ones(len(sp1)) * 1, '|', ms=20, label='neuron 1')
plt.plot(sp2, np.ones(len(sp2)) * 1.1, '|', ms=20, label='neuron 2')
plt.xlabel('time (ms)')
plt.ylabel('neuron id.')
plt.xlim(1000, 8000)
plt.ylim(0.9, 1.2)
plt.legend()
plt.show()
```
## Think!
- Is the output correlation always smaller than the input correlation? If yes, why?
- Should there be a systematic relationship between input and output correlations?
You will explore these questions in the next figure but try to develop your own intuitions first!
Lets vary `c_in` and plot the relationship between the `c_in` and output correlation. This might take some time depending on the number of trials.
```
#@title
#@markdown Don't forget to execute this cell!
pars = default_pars(T=80000, dt=1.) # get the parameters
bin_size = 10.
c_in = np.arange(0, 1.0, 0.1) # set the range for input CC
r12_ss = np.zeros(len(c_in)) # small mu, small sigma
starttime = time.perf_counter() # time clock
for ic in range(len(c_in)):
r12_ss[ic], sp_ss, sp1, sp2 = LIF_output_cc(pars, mu=10.0, sig=10.,
c=c_in[ic], bin_size=bin_size,
n_trials=10)
endtime = time.perf_counter()
timecost = (endtime - starttime) / 60.
print(f"Simulation time = {timecost:.2f} min")
plt.figure(figsize=(7, 6))
plot_c_r_LIF(c_in, r12_ss, mycolor='b', mylabel='Output CC')
plt.plot([c_in.min() - 0.05, c_in.max() + 0.05],
[c_in.min() - 0.05, c_in.max() + 0.05],
'k--', dashes=(2, 2), label='y=x')
plt.xlabel('Input CC')
plt.ylabel('Output CC')
plt.legend(loc='best', fontsize=16)
plt.show()
```
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D3_BiologicalNeuronModels/solutions/W2D3_Tutorial2_Solution_71e76f4d.py)
---
# Section 3: Correlation transfer function
The above plot of input correlation vs. output correlation is called the __correlation transfer function__ of the neurons.
## Section 3.1: How do the mean and standard deviation of the GWN affect the correlation transfer function?
The correlations transfer function appears to be linear. The above can be taken as the input/output transfer function of LIF neurons for correlations, instead of the transfer function for input/output firing rates as we had discussed in the previous tutorial (i.e., F-I curve).
What would you expect to happen to the slope of the correlation transfer function if you vary the mean and/or the standard deviation of the GWN?
```
#@markdown Execute this cell to visualize correlation transfer functions
pars = default_pars(T=80000, dt=1.) # get the parameters
no_trial = 10
bin_size = 10.
c_in = np.arange(0., 1., 0.2) # set the range for input CC
r12_ss = np.zeros(len(c_in)) # small mu, small sigma
r12_ls = np.zeros(len(c_in)) # large mu, small sigma
r12_sl = np.zeros(len(c_in)) # small mu, large sigma
starttime = time.perf_counter() # time clock
for ic in range(len(c_in)):
r12_ss[ic], sp_ss, sp1, sp2 = LIF_output_cc(pars, mu=10.0, sig=10.,
c=c_in[ic], bin_size=bin_size,
n_trials=no_trial)
r12_ls[ic], sp_ls, sp1, sp2 = LIF_output_cc(pars, mu=18.0, sig=10.,
c=c_in[ic], bin_size=bin_size,
n_trials=no_trial)
r12_sl[ic], sp_sl, sp1, sp2 = LIF_output_cc(pars, mu=10.0, sig=20.,
c=c_in[ic], bin_size=bin_size,
n_trials=no_trial)
endtime = time.perf_counter()
timecost = (endtime - starttime) / 60.
print(f"Simulation time = {timecost:.2f} min")
plt.figure(figsize=(7, 6))
plot_c_r_LIF(c_in, r12_ss, mycolor='b', mylabel=r'Small $\mu$, small $\sigma$')
plot_c_r_LIF(c_in, r12_ls, mycolor='y', mylabel=r'Large $\mu$, small $\sigma$')
plot_c_r_LIF(c_in, r12_sl, mycolor='r', mylabel=r'Small $\mu$, large $\sigma$')
plt.plot([c_in.min() - 0.05, c_in.max() + 0.05],
[c_in.min() - 0.05, c_in.max() + 0.05],
'k--', dashes=(2, 2), label='y=x')
plt.xlabel('Input CC')
plt.ylabel('Output CC')
plt.legend(loc='best', fontsize=14)
plt.show()
```
### Think!
Why do both the mean and the standard deviation of the GWN affect the slope of the correlation transfer function?
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D3_BiologicalNeuronModels/solutions/W2D3_Tutorial2_Solution_2deb4ccb.py)
## Section 3.2: What is the rationale behind varying $\mu$ and $\sigma$?
The mean and the variance of the synaptic current depends on the spike rate of a Poisson process. We can use [Campbell's theorem](https://en.wikipedia.org/wiki/Campbell%27s_theorem_(probability)) to estimate the mean and the variance of the synaptic current:
\begin{align}
\mu_{\rm syn} = \lambda J \int P(t) \\
\sigma_{\rm syn} = \lambda J \int P(t)^2 dt\\
\end{align}
where $\lambda$ is the firing rate of the Poisson input, $J$ the amplitude of the postsynaptic current and $P(t)$ is the shape of the postsynaptic current as a function of time.
Therefore, when we varied $\mu$ and/or $\sigma$ of the GWN, we mimicked a change in the input firing rate. Note that, if we change the firing rate, both $\mu$ and $\sigma$ will change simultaneously, not independently.
Here, since we observe an effect of $\mu$ and $\sigma$ on correlation transfer, this implies that the input rate has an impact on the correlation transfer function.
### Think!
- What are the factors that would make output correlations smaller than input correlations? (Notice that the colored lines are below the black dashed line)
- What does it mean for the correlation in the network?
- Here we have studied the transfer of correlations by injecting GWN. But in the previous tutorial, we mentioned that GWN is unphysiological. Indeed, neurons receive colored noise (i.e., Shot noise or OU process). How do these results obtained from injection of GWN apply to the case where correlated spiking inputs are injected in the two LIFs? Will the results be the same or different?
Reference
- De La Rocha, Jaime, et al. "Correlation between neural spike trains increases with firing rate." Nature (2007) (https://www.nature.com/articles/nature06028/)
- Bujan AF, Aertsen A, Kumar A. Role of input correlations in shaping the variability and noise correlations of evoked activity in the neocortex. Journal of Neuroscience. 2015 Jun 3;35(22):8611-25. (https://www.jneurosci.org/content/35/22/8611)
[*Click for solution*](https://github.com/NeuromatchAcademy/course-content/tree/master//tutorials/W2D3_BiologicalNeuronModels/solutions/W2D3_Tutorial2_Solution_39d29f52.py)
---
# Summary
In this tutorial, we studied how the input correlation of two LIF neurons is mapped to their output correlation. Specifically, we:
- injected correlated GWN in a pair of neurons,
- measured correlations between the spiking activity of the two neurons, and
- studied how the transfer of correlation depends on the statistics of the input, i.e., mean and standard deviation.
Here, we were concerned with zero time lag correlation. For this reason, we restricted estimation of correlation to instantaneous correlations. If you are interested in time-lagged correlation, then we should estimate the cross-correlogram of the spike trains and find out the dominant peak and area under the peak to get an estimate of output correlations.
We leave this as a future to-do for you if you are interested.
---
# Bonus 1: Example of a conductance-based LIF model
Above, we have written code to generate correlated Poisson spike trains. You can write code to stimulate the LIF neuron with such correlated spike trains and study the correlation transfer function for spiking input and compare it to the correlation transfer function obtained by injecting correlated GWNs.
```
# @title Function to simulate conductance-based LIF
def run_LIF_cond(pars, I_inj, pre_spike_train_ex, pre_spike_train_in):
"""
conductance-based LIF dynamics
Args:
pars : parameter dictionary
I_inj : injected current [pA]. The injected current here can
be a value or an array
pre_spike_train_ex : spike train input from presynaptic excitatory neuron
pre_spike_train_in : spike train input from presynaptic inhibitory neuron
Returns:
rec_spikes : spike times
rec_v : mebrane potential
gE : postsynaptic excitatory conductance
gI : postsynaptic inhibitory conductance
"""
# Retrieve parameters
V_th, V_reset = pars['V_th'], pars['V_reset']
tau_m, g_L = pars['tau_m'], pars['g_L']
V_init, E_L = pars['V_init'], pars['E_L']
gE_bar, gI_bar = pars['gE_bar'], pars['gI_bar']
VE, VI = pars['VE'], pars['VI']
tau_syn_E, tau_syn_I = pars['tau_syn_E'], pars['tau_syn_I']
tref = pars['tref']
dt, range_t = pars['dt'], pars['range_t']
Lt = range_t.size
# Initialize
tr = 0.
v = np.zeros(Lt)
v[0] = V_init
gE = np.zeros(Lt)
gI = np.zeros(Lt)
Iinj = I_inj * np.ones(Lt) # ensure I has length Lt
if pre_spike_train_ex.max() == 0:
pre_spike_train_ex_total = np.zeros(Lt)
else:
pre_spike_train_ex_total = pre_spike_train_ex * np.ones(Lt)
if pre_spike_train_in.max() == 0:
pre_spike_train_in_total = np.zeros(Lt)
else:
pre_spike_train_in_total = pre_spike_train_in * np.ones(Lt)
# simulation
rec_spikes = [] # recording spike times
for it in range(Lt - 1):
if tr > 0:
v[it] = V_reset
tr = tr - 1
elif v[it] >= V_th: # reset voltage and record spike event
rec_spikes.append(it)
v[it] = V_reset
tr = tref / dt
# update the synaptic conductance
gE[it+1] = gE[it] - (dt / tau_syn_E) * gE[it] + gE_bar * pre_spike_train_ex_total[it + 1]
gI[it+1] = gI[it] - (dt / tau_syn_I) * gI[it] + gI_bar * pre_spike_train_in_total[it + 1]
# calculate the increment of the membrane potential
dv = (-(v[it] - E_L) - (gE[it + 1] / g_L) * (v[it] - VE) - \
(gI[it + 1] / g_L) * (v[it] - VI) + Iinj[it] / g_L) * (dt / tau_m)
# update membrane potential
v[it + 1] = v[it] + dv
rec_spikes = np.array(rec_spikes) * dt
return v, rec_spikes, gE, gI
print(help(run_LIF_cond))
```
## Interactive Demo: Correlated spike input to an LIF neuron
In the following you can explore what happens when the neurons receive correlated spiking input.
You can vary the correlation between excitatory input spike trains. For simplicity, the correlation between inhibitory spike trains is set to 0.01.
Vary both excitatory rate and correlation and see how the output correlation changes. Check if the results are qualitatively similar to what you observed previously when you varied the $\mu$ and $\sigma$.
```
# @title
# @markdown Make sure you execute this cell to enable the widget!
my_layout.width = '450px'
@widgets.interact(
pwc_ee=widgets.FloatSlider(0.3, min=0.05, max=0.99, step=0.01,
layout=my_layout),
exc_rate=widgets.FloatSlider(1e3, min=500., max=5e3, step=50.,
layout=my_layout),
inh_rate=widgets.FloatSlider(500., min=300., max=5e3, step=5.,
layout=my_layout),
)
def EI_isi_regularity(pwc_ee, exc_rate, inh_rate):
pars = default_pars(T=1000.)
# Add parameters
pars['V_th'] = -55. # spike threshold [mV]
pars['V_reset'] = -75. # reset potential [mV]
pars['tau_m'] = 10. # membrane time constant [ms]
pars['g_L'] = 10. # leak conductance [nS]
pars['V_init'] = -65. # initial potential [mV]
pars['E_L'] = -75. # leak reversal potential [mV]
pars['tref'] = 2. # refractory time (ms)
pars['gE_bar'] = 4.0 # [nS]
pars['VE'] = 0. # [mV] excitatory reversal potential
pars['tau_syn_E'] = 2. # [ms]
pars['gI_bar'] = 2.4 # [nS]
pars['VI'] = -80. # [mV] inhibitory reversal potential
pars['tau_syn_I'] = 5. # [ms]
my_bin = np.arange(0, pars['T']+pars['dt'], .1) # 20 [ms] bin-size
# exc_rate = 1e3
# inh_rate = 0.4e3
# pwc_ee = 0.3
pwc_ii = 0.01
# generate two correlated spike trains for excitatory input
sp1e, sp2e = generate_corr_Poisson(pars, exc_rate, pwc_ee)
sp1_spike_train_ex, _ = np.histogram(sp1e, bins=my_bin)
sp2_spike_train_ex, _ = np.histogram(sp2e, bins=my_bin)
# generate two uncorrelated spike trains for inhibitory input
sp1i, sp2i = generate_corr_Poisson(pars, inh_rate, pwc_ii)
sp1_spike_train_in, _ = np.histogram(sp1i, bins=my_bin)
sp2_spike_train_in, _ = np.histogram(sp2i, bins=my_bin)
v1, rec_spikes1, gE, gI = run_LIF_cond(pars, 0, sp1_spike_train_ex, sp1_spike_train_in)
v2, rec_spikes2, gE, gI = run_LIF_cond(pars, 0, sp2_spike_train_ex, sp2_spike_train_in)
# bin the spike time
bin_size = 20 # [ms]
my_bin = np.arange(0, pars['T'], bin_size)
spk_1, _ = np.histogram(rec_spikes1, bins=my_bin)
spk_2, _ = np.histogram(rec_spikes2, bins=my_bin)
r12 = my_CC(spk_1, spk_2)
print(f"Input correlation = {pwc_ee}")
print(f"Output correlation = {r12}")
plt.figure(figsize=(14, 7))
plt.subplot(211)
plt.plot(sp1e, np.ones(len(sp1e)) * 1, '|', ms=20,
label='Exc. input 1')
plt.plot(sp2e, np.ones(len(sp2e)) * 1.1, '|', ms=20,
label='Exc. input 2')
plt.plot(sp1i, np.ones(len(sp1i)) * 1.3, '|k', ms=20,
label='Inh. input 1')
plt.plot(sp2i, np.ones(len(sp2i)) * 1.4, '|k', ms=20,
label='Inh. input 2')
plt.ylim(0.9, 1.5)
plt.legend()
plt.ylabel('neuron id.')
plt.subplot(212)
plt.plot(pars['range_t'], v1, label='neuron 1')
plt.plot(pars['range_t'], v2, label='neuron 2')
plt.xlabel('time (ms)')
plt.ylabel('membrane voltage $V_{m}$')
plt.tight_layout()
plt.show()
```
Above, we are estimating the output correlation for one trial. You can modify the code to get a trial average of output correlations.
---
# Bonus 2: Ensemble Response
Finally, there is a short BONUS lecture video on the firing response of an ensemble of neurons to time-varying input. There are no associated coding exercises - just enjoy.
```
# @title Video 2 (Bonus): Response of ensemble of neurons to time-varying input
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = 'https://player.bilibili.com/player.html?bvid={0}&page={1}'.format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id="", width=854, height=480, fs=1)
print('Video available at https://www.bilibili.com/video/{0}'.format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id="78_dWa4VOIo", width=854, height=480, fs=1, rel=0)
print('Video available at https://youtube.com/watch?v=' + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
| github_jupyter |
```
%matplotlib inline
import os
import time
import datetime
import warnings
import collections
import matplotlib
matplotlib.rcParams.update({'font.size': 18})
import pandas
import numpy
import scipy.stats
import abcutils
```
## Load and Synthesize Data from CSV
This process loads each summary CSV file, creates a few derived metrics, and then merges each system's CSV into a single global dataset that can be sliced and diced by system, benchmark, or any other way. We are now caching the processed CSV in HDF5 format to speed up initial data ingest at the beginning of each analysis. Delete the `CACHE_FILE` to re-generate this cache (e.g., when the contents of the CSV are updated).
```
filtered_df = abcutils.sc18paper.load_dataset()
```
## Demonstrate a Single Test Platform
Look at one combination of (compute system, file system, benchmark) to show what this UMAMI analysis can do.
### Define Input Parameters
```
plot_metric = 'darshan_normalized_perf_by_max'
group_by = ['_test_platform', '_benchmark_id']
print("plot_metric =", abcutils.CONFIG['metric_labels'].get(plot_metric, plot_metric))
print("date_start =", abcutils.sc18paper.DATE_START.isoformat())
print("date_end =", abcutils.sc18paper.DATE_END.isoformat())
# Width of simple moving average (SMA) short/long windows
short_window = pandas.Timedelta(days=14)
long_window = pandas.Timedelta(days=49)
print("Short window will average over %s measurements at a time" % short_window)
print("Long window will average over %s measurements at a time" % long_window)
```
## Classification
We actually only attempt to correlate against a fixed subset of the features. This is a time-saving measure; the features in `umami_row_order` were determined by performing an unguided correlation against everything and only selecting those features which showed some degree of correlation _and_ were not degenerate of other features.
```
CORRELATION_METRICS = abcutils.CONFIG['umami_row_order']
CORRELATION_METRICS
```
### Generate or load calculated contributors
This can take an inconvenient amount of time, so we cache the results to `contributors.hdf`
```
all_contributors = None
# Load the cached contributors list if available
try:
all_contributors = pandas.read_hdf('contributors.hdf5', 'contributors')
except IOError:
pass
# Otherwise recalculate the contributors list
if all_contributors is None:
grouped_df = filtered_df.groupby(by=group_by)
for group in grouped_df.groups:
example_df = grouped_df.get_group(group)
intercepts = abcutils.features.sma_intercepts(example_df,
plot_metric,
short_window=short_window,
long_window=long_window)
loci = abcutils.features.generate_loci_sma(example_df,
plot_metric,
mins=True,
maxes=False,
short_window=short_window,
long_window=long_window)
regions = list(abcutils.features.intercepts_to_region(example_df, intercepts))
for region in regions:
contributors = abcutils.classify.identify_contributors(region=region,
target_column=plot_metric,
target_index=region[plot_metric].idxmin(),
correlate_columns=CORRELATION_METRICS,
want_good=False,
classifier='minmax')
if all_contributors is None:
all_contributors = contributors
else:
all_contributors = pandas.concat((all_contributors, contributors))
# Cache the contributors list for the next time
all_contributors.index = numpy.arange(len(all_contributors))
all_contributors.to_hdf('contributors.hdf5', key='contributors', mode='w', format='fixed', complevel=9, complib='zlib')
print("Number of target indices:", len(all_contributors['target_index'].unique()))
```
Apply filters to remove very high p-value measurements from the dataset. These cause problems when performing significance testing later on, since they dilute the significance of the results.
```
# Also drop any contributors who lack statistical confidence because of duplicate values
significant_contributors = all_contributors[all_contributors['pvalue'] < 0.10]
print("Discarding %d contributors with p-values < 0.10" % (len(all_contributors) - len(significant_contributors)))
# Keep all data and let p-values speak for themselves
#significant_contributors = all_contributors
print("Number of contributors remaining:", len(significant_contributors))
print("Number of target indices ('bad' jobs):", len(significant_contributors['target_index'].unique()))
print("Number of unclassified jobs:", (all_contributors.groupby(['target_index']).sum()['target_metric_matches'] < 1.0).sum())
```
The following table is the total number of observations broken down by file system. For example, `fs_ave_mds_cpu` for `cscratch@cori-knl` = `102.0` means that there were 102 cases where poor performance was observed on `cscratch@cori-knl` while at the same time the `fs_ave_mds_cpu` metric was available.
```
metric_observation_counts = pandas.pivot_table(significant_contributors,
values='target_metric_matches',
index=['metric_name'],
columns=['_test_platform'],
aggfunc=lambda x: (~numpy.isnan(x)).sum()).fillna(0.0)
metric_observation_counts
```
The following table shows how many of the observations in the previous table were actually implicated (tagged) as being correlated with poor performance.
```
tagged_metric_counts = pandas.pivot_table(significant_contributors,
values='target_metric_matches',
index=['metric_name'],
columns=['_test_platform'],
aggfunc=numpy.sum).fillna(0.0)
tagged_metric_counts
```
We then divide the number of times each metric was implicated (tagged) as correlating with poor performance by the total number of times that metric was observed on each file system. The result is the fraction of times each metric was observed to correlate with poor performance on a per-file system basis.
```
contributor_distribution = tagged_metric_counts.div(metric_observation_counts.sum(axis=1), axis=0)
contributor_distribution = contributor_distribution.reindex(index=contributor_distribution.sum(axis=1).sort_values(ascending=False).index)
try:
del contributor_distribution.columns.name
except AttributeError:
pass
contributor_distribution
```
We then take the total number of times each metric was tagged across _all file systems_ (across an entire row in the previous table) and divide it by the total number of observations of that metric to calculate the fraction of observations where each metric was tagged as being correlated with poor performance.
```
def draw_stacked_bars(contributor_distribution, legendprops=None):
_legendprops = {}
if legendprops:
_legendprops.update(legendprops)
row_sums = contributor_distribution.sum(axis=1)
fig, ax = matplotlib.pyplot.subplots(figsize=(8,4))
contributor_distribution.plot.bar(stacked=True, ax=ax, width=0.90)
ax.grid()
ax.set_ylim(0, 0.5)
ax.set_axisbelow(True)
xticklabels = [abcutils.CONFIG['umami_rows'].get(x.get_text(), x.get_text()) for x in ax.get_xticklabels()]
ax.set_xticklabels(xticklabels, rotation=30, ha='right')
ax.set_xlabel("")
ax.set_ylabel("Fraction of tests")
handles, labels = ax.get_legend_handles_labels()
ax.legend(handles, [abcutils.CONFIG['platform_labels'].get(x, x) for x in labels], **_legendprops)
for index, x_value in enumerate(ax.get_xticks()):
ax.annotate("%d%%" % (100.0 * row_sums[index]), xy=(x_value, row_sums[index] + 0.02),
ha='center',
backgroundcolor='#FFFFFFAA')
return ax
```
## Assert confidence
We use the binomial test to calculate the p-values of each fraction of tests asserted above.
```
results = []
for metric in contributor_distribution.index.values: # ['fs_ave_oss_cpu']: #
prob_success = 1.0
prob_failure = 1.0
successes = []
failures = []
num_matches = 0
metric_measures = significant_contributors[significant_contributors['metric_name'] == metric]
for row in metric_measures.itertuples():
if row.target_metric_matches:
num_matches += 1
prob_success *= row.pvalue
successes.append(row.pvalue)
else:
prob_failure *= (1.0 - row.pvalue)
failures.append(row.pvalue)
pick_n = num_matches
out_of = len(metric_measures)
if not successes:
continue
# what is the probability that we observe pick_n / out_of jobs with this
# tagged metric given the probability of encountering a tagged metric
# if there's no relationship between this metric being tagged and each
# job's performance?
#
# binomial test: assume the null hypothesis is TRUE
# 1. pick the highest p-value observed for this metric - that is the
# case where the null hypothesis is most likely to be true
# 2. perform the binomial test to see what the odds are of observing
# pick_n **or more** tagged metrics if the null hypothesis is true?
probability = numpy.max(successes)
pvalue = scipy.stats.binom_test(pick_n,
out_of,
probability,
alternative='greater')
result = collections.OrderedDict({})
result['metric'] = metric
result['pick_n'] = pick_n
result['out_of'] = out_of
result['probability_used'] = probability
result['calculated_pvalue'] = pvalue
results.append(result)
binomial_results = pandas.DataFrame.from_dict(results).set_index('metric')
binomial_results
ax = draw_stacked_bars(contributor_distribution.loc[binomial_results.index])
# relabel legend
for text in ax.legend().get_texts():
text_str = text.get_text()
text.set_text(abcutils.config.CONFIG['platform_labels_public'].get(text_str, text_str))
ax.legend(fontsize="small")
```
Shade off the statistically insignificant metrics
```
# Build a mapping from metrics to rectangles
reverse_metric_map = {}
for key, val in abcutils.CONFIG['umami_rows'].items():
reverse_metric_map[val] = key
if 'CF' in val:
reverse_metric_map[val.replace(' CF', ' Contention')] = key
# Find all rectangles corresponding to each metric
rectangle_map = {}
xticks = ax.get_xticks()
xticklabels = [x.get_text() for x in ax.get_xticklabels()]
for child in ax.get_children():
if isinstance(child, matplotlib.patches.Rectangle) and child.get_width() == 0.9:
child_x = int(round(child.xy[0] + child.get_width() / 2))
key = reverse_metric_map[xticklabels[child_x]]
if key not in rectangle_map:
rectangle_map[key] = []
rectangle_map[key].append(child)
# Actually apply a grey box over the box of each metric that is not statistically significant
for row in binomial_results.itertuples():
if row.calculated_pvalue > 0.10:
for rectangle in rectangle_map[row.Index]:
rectangle.set_color("#DDDDDD")
rectangle.set_edgecolor('#DDDDDD')
# relabel legend
new_labels = []
for text in ax.legend().get_texts():
text_str = text.get_text()
new_text_str = abcutils.config.CONFIG['platform_labels_public'].get(text_str, text_str)
print("Replacing %s with %s" % (text_str, new_text_str))
new_labels.append(new_text_str)
ax.legend(labels=new_labels, fontsize="small")
ax.xaxis.grid(False)
ax.get_figure().savefig('figs/contributors-bad-by-system-grey.pdf', bbox_inches='tight', transparent=True)
ax.get_figure()
```
Also provide a less confusing version of the plot without the per-file system resolution.
```
for row in binomial_results.itertuples():
if row.calculated_pvalue > 0.10:
for rectangle in rectangle_map[row.Index]:
rectangle.set_color("#DDDDDD")
rectangle.set_edgecolor('#DDDDDD')
else:
for rectangle in rectangle_map[row.Index]:
rectangle.set_color("C0")
rectangle.set_edgecolor('C0')
ax.get_legend().set_visible(False)
ax.get_figure().savefig('figs/contributors-bad-grey.pdf', bbox_inches='tight')
ax.get_figure()
```
Then only show the metrics that are statistically significant at all.
```
ax = draw_stacked_bars(contributor_distribution.loc[(binomial_results['calculated_pvalue'] < 0.10).values],
legendprops={
'loc': 'upper right',
'bbox_to_anchor': (1.01, 1.03),
'labelspacing': 0.4
})
ax.set_xticklabels([x.get_text().replace(' CF', '\nContention') for x in ax.get_xticklabels()], rotation=30)
ax.xaxis.grid(False)
ax.get_figure().savefig('figs/contributors-bad-by-system.pdf', bbox_inches='tight')
binomial_results.sort_values('calculated_pvalue')
```
| github_jupyter |
```
import plotting_functions as pf
import pandas as pd
import numpy as np
clk = "5.00"
freq = "1GHz" if clk == "1.00" else "200MHz"
export = f"../results/breakdown"
prism_prune = [
"#5F4690",
"#38A6A5",
"#0F8554",
"#EDAD08",
"#CC503E",
"#94346E",
"#d68bd2",
"#666666",
]
clock = float(clk)
DIR_FU = f"../results/breakdown/{clk}/FU"
DIR_SWU = f"../results/breakdown/{clk}/SWU"
df_FU = pd.read_csv(f"{DIR_FU}/power.csv", index_col=[0, 1])
df_SWU = pd.read_csv(f"{DIR_SWU}/power.csv", index_col=[0, 1])
df_FU.loc["8x8"] = df_FU.loc["8x8"].mul(clock * (10 ** 3) / 512).values
df_FU.loc["8x4"] = df_FU.loc["8x4"].mul(clock * (10 ** 3) / 1024).values
df_FU.loc["8x2"] = df_FU.loc["8x2"].mul(clock * (10 ** 3) / 2048).values
df_FU.loc["4x4"] = df_FU.loc["4x4"].mul(clock * (10 ** 3) / 2048).values
df_FU.loc["2x2"] = df_FU.loc["2x2"].mul(clock * (10 ** 3) / 8192).values
df_SWU.loc["8x8"] = df_SWU.loc["8x8"].mul(clock * (10 ** 3) / 512).values
df_SWU.loc["4x4"] = df_SWU.loc["4x4"].mul(clock * (10 ** 3) / 1024).values
df_SWU.loc["2x2"] = df_SWU.loc["2x2"].mul(clock * (10 ** 3) / 2048).values
for prec in ["8x4", "8x2"]:
for row in df_SWU.index.levels[1]:
df_SWU.loc[(prec, row), :] = 0
df_area_FU = pd.read_csv(f"{DIR_FU}/area.csv", index_col=0)
df_area_SWU = pd.read_csv(f"{DIR_SWU}/area.csv", index_col=0)
df_area = pd.concat([df_area_FU, df_area_SWU])
df_area.rename(
index={
"BITFUSION": "BG_L2_L4_00_L3_11_L2_11_DVAFS_0",
"BITBLADE": "BG_L3_L4_00_L3_11_L2_11_DVAFS_0",
"LOOM": "BG_BS_L4_00_L3_00_L2_11_DVAFS_0",
},
columns={"comb": "Combinational", "seq": "Sequential"},
inplace=True,
)
df_area.index.set_names("design", inplace=True)
df_area = df_area / 1e6
df_energy = pd.concat([df_FU, df_SWU])
df_energy.index.set_names(("prec", "design"), inplace=True)
ordered_list = [
"FU\nBG: L2\nL2: IS",
"FU\nBG: L2\nL2: HS",
"FU\nBG: L2\nL2: OS",
"FU\nBG: L3\nL2: HS",
"FU\nBG: L3\nL2: OS",
"FU\nBG: BS\nL2: OS",
"SWU\nBG: L2\nL2: NO",
"SWU\nBG: L2\nL2: OS",
]
col_dict = {
"mult_2x2": "L1 Multipliers",
"L2_tree": "L2 Adder Tree",
"L3_tree": "L3 Adder Tree",
"L4_tree": "L4 Adder Tree",
"out_reg": "Output Registers",
"pipe_reg": "Internal Registers",
"in_reg": "Input Registers",
"accum": "Output Accumulators",
}
df_energy.rename(
index={
"BITFUSION": "BG_L2_L4_00_L3_11_L2_11_DVAFS_0",
"BITBLADE": "BG_L3_L4_00_L3_11_L2_11_DVAFS_0",
"LOOM": "BG_BS_L4_00_L3_00_L2_11_DVAFS_0",
},
columns=col_dict,
inplace=True,
)
df_scatter = pd.DataFrame("0", index=df_energy.index, columns=["Energy/Op", "Area"])
for prec in ["8x8", "8x4", "8x2", "4x4", "2x2"]:
df_scatter.loc[prec] = pd.concat(
[df_energy.loc[prec]["top"], df_area["top"]], axis=1
).values
df_scatter.reset_index(level="design", inplace=True)
df_scatter["Config / BG / L2"] = df_scatter["design"].apply(pf.SWPBGL2_noline)
df_scatter["L4 / L3 Modes"] = df_scatter["design"].apply(pf.L4L3)
df_scatter["SWP"] = df_scatter["design"].apply(pf.SWP)
df_scatter.set_index([df_scatter.index, "design"], inplace=True)
df_scatter.rename(
index={
"8x8": "8b x 8b",
"8x4": "8b x 4b",
"8x2": "8b x 2b",
"4x4": "4b x 4b",
"2x2": "2b x 2b",
},
inplace=True,
)
palette = {
"FU / BG: L2 / L2: IS": "tab:blue",
"FU / BG: L2 / L2: HS": "tab:green",
"FU / BG: L2 / L2: OS": "tab:red",
"FU / BG: L3 / L2: HS": "tab:brown",
"FU / BG: L3 / L2: OS": "tab:orange",
"FU / BG: BS / L2: OS": "tab:cyan",
"SWU / BG: L2 / L2: NO": "tab:pink",
"SWU / BG: L2 / L2: OS": "tab:purple",
}
df_energy.reset_index(level="design", inplace=True)
df_energy["BG Unrolling / L2 Mode"] = df_energy["design"].apply(pf.SWPBGL2)
df_energy["L4 / L3 Modes"] = df_energy["design"].apply(pf.L4L3)
df_energy.drop(labels="design", axis="columns", inplace=True)
df_energy.set_index(
[df_energy.index, "BG Unrolling / L2 Mode", "L4 / L3 Modes"], inplace=True
)
df_energy = df_energy.reindex(ordered_list, level=1)
df_area.reset_index(level="design", inplace=True)
df_area["BG Unrolling / L2 Mode"] = df_area["design"].apply(pf.SWPBGL2)
df_area["L4 / L3 Modes"] = df_area["design"].apply(pf.L4L3)
df_area.drop(labels="design", axis="columns", inplace=True)
df_area.set_index(["BG Unrolling / L2 Mode", "L4 / L3 Modes"], inplace=True)
df_area = df_area.reindex(ordered_list, level=0)
BAR_AREA = ["Combinational", "Sequential"]
pf.plot_clustered_stacked(
df_area[BAR_AREA],
title=f"Area_Breakdown",
save=True,
export=f"{export}/{clk}",
ext="png",
cmap="tab20b",
width=1,
)
# In the paper, there was no room to show all 5 tested precisions
# We opted to drop 8b x 2b precision, and show the rest
# This is optional, you can still plot 8x2 precision
pf.scatter_extract(
df_scatter.drop("8b x 2b", level="prec").replace(0, np.nan).dropna(),
name="alpha",
ext="png",
export=f"{export}/{clk}",
legend=(freq == "1GHz"),
palette=palette,
)
BAR_POWER = [
"L1 Multipliers",
"L2 Adder Tree",
"L3 Adder Tree",
"L4 Adder Tree",
"Output Registers",
"Input Registers",
"Internal Registers",
"Output Accumulators",
]
for prec in [
"8x8",
"8x4",
"8x2",
"4x4",
"2x2",
]:
pf.plot_clustered_stacked(
df_energy.xs(prec)[BAR_POWER],
title=f"Energy_{prec}".replace(" ", "").replace("b", ""),
save=True,
export=f"{export}/{clk}",
ext="png",
color=prism_prune,
width=1,
sep_legend=True,
)
df_sq_5_FU = pf.energy_extract("5.00", False)
df_sq_1_FU = pf.energy_extract("1.00", False)
df_sq_5_SWU = pf.energy_extract("5.00", True)
df_sq_1_SWU = pf.energy_extract("1.00", True)
df_sq_5_total = pd.concat([df_sq_5_FU, df_sq_5_SWU], axis=1, join="inner")
df_sq_1_total = pd.concat([df_sq_1_FU, df_sq_1_SWU], axis=1, join="inner")
df_sq_5_total.rename(
index={
"8x8": "8b x 8b",
"8x4": "8b x 4b",
"8x2": "8b x 2b",
"4x4": "4b x 4b",
"2x2": "2b x 2b",
},
inplace=True
)
df_sq_1_total.rename(
index={
"8x8": "8b x 8b",
"8x4": "8b x 4b",
"8x2": "8b x 2b",
"4x4": "4b x 4b",
"2x2": "2b x 2b",
},
inplace=True
)
# ideal_loops = {"B": 1, "K": 64, "C": 256, "OY": 8, "OX": 8, "FY": 4, "FX": 4}
# realistic_loops = {"B": 1, "K": 64, "C": 256, "OY": 7, "OX": 7, "FY": 3, "FX": 3}
# df_util = pf.square_util(realistic_loops)
# df_util_5 = df_sq_5 * df_util
# df_util_1 = df_sq_1 * df_util
# df_mix_5 = (
# df_sq_5.xs("8x8") * 0.2
# + df_sq_5.xs("8x4") * 0.15
# + df_sq_5.xs("8x2") * 0.15
# + df_sq_5.xs("4x4") * 0.3
# + df_sq_5.xs("2x2") * 0.2
# )
# df_mix_1 = (
# df_sq_1.xs("8x8") * 0.2
# + df_sq_1.xs("8x4") * 0.15
# + df_sq_1.xs("8x2") * 0.15
# + df_sq_1.xs("4x4") * 0.3
# + df_sq_1.xs("2x2") * 0.2
# )
# df_mix = pd.concat([df_mix_5, df_mix_1], keys=["200 MHz", "1 GHz"], names=["freq"])
DVAFS = False
pf.heatmap_extract(
df_sq=df_sq_5_total.drop("8b x 2b", level="prec"),
name="total",
cmap="viridis_r",
ext="png",
export=f"{export}/5.00",
DVAFS=DVAFS,
)
pf.heatmap_extract(
df_sq=df_sq_1_total.drop("8b x 2b", level="prec"),
name="total",
cmap="viridis_r",
ext="png",
export=f"{export}/1.00",
DVAFS=DVAFS,
ylabels=False,
)
```
| github_jupyter |
# Apache Arrow shared between Python and R with rpy2
The Python package shown here is available at:
https://github.com/rpy2/rpy2-arrow
```
import pyarrow
import pyarrow.dataset as ds
import rpy2_arrow.pyarrow_rarrow as pyra
```
Our dataset in the NYC taxi one made available to download in the Parquet format by Ursa Labs.
```
DATA_PATH = 'nyc-taxi'
```
## Setup: Get a dataset
Most will not have that data already available locally in directories. We use a Python
translation of the code as https://ursalabs.org/arrow-r-nightly/articles/dataset.html
to fetch the data.
```
# This allows download an incomplete dataset
# in the interest of time. Set it to None or -1
# to download the complete dataset.
MAX_NMONTHS = 10
import os
import urllib.parse
import urllib.request
import shutil
BUCKET = 'https://ursa-labs-taxi-data.s3.us-east-2.amazonaws.com'
paths = []
print(' | |')
for year in range(2009, 2020):
if len(paths) == MAX_NMONTHS:
print()
break
print(f'{year} ', end='', flush=True)
if year == 2019:
# We only have through June 2019 there
months = range(1, 7)
else:
months = range(1, 13)
for month in months:
if len(paths) == MAX_NMONTHS:
print()
break
month_str = f'{month:02d}'
year_str = str(year)
url = urllib.parse.urljoin(BUCKET, '/'.join((year_str, month_str, 'data.parquet')))
filename = os.path.join(DATA_PATH, year_str, month_str, 'data.parquet')
if os.path.exists(filename):
print('s', end='', flush=True)
paths.append(filename)
continue
print('D', end='', flush=True)
os.makedirs(os.path.join(DATA_PATH, year_str, month_str))
with urllib.request.urlopen(url) as response, open(filename, 'wb') as output_file:
shutil.copyfileobj(response, output_file)
paths.append(filename)
print()
```
## We are ready, let's load a dataset
The dataset is read using `pyarrow.dataset`:
```
dataset = ds.dataset(paths, format='parquet')
dataset
```
The following line reads the dataset into an Arrow table. Depending on the size
of the data this may require a lot of memory so we apply a filter (check the relevant documentation
for more details: https://arrow.apache.org/docs/python/dataset.html).
```
tbl = dataset.to_table(filter=ds.field('tip_amount') > 10, batch_size=5E6)
tbl.shape
```
The table has a schema that describes types in its content.
```
tbl.schema.types
```
Each column is an array (this is a column-oriented format), or more specically a chunked array.
```
import timeit
N = 3
for col_i in (0, 1, 5):
print(f'Column: {tbl.schema.types[col_i]}')
t_getitem = timeit.timeit(lambda: tbl[col_i], number=N) / N
print(f' getitem: {t_getitem:.2e}s', end='', flush=True)
array = tbl[col_i]
t_convert = timeit.timeit(lambda: pyra.converter.py2rpy(array), number=N) / N
print(f' to R: {t_convert:.2e}s')
```
We can also convert the Arrow table directly. Note that it takes the cumulated time required to convert each one of the columns.
```
%%time
r_tbl = pyra.converter.py2rpy(tbl)
```
This is disappointingly slow. We can go **much** faster by first combining the chunks.
```
%%time
cb_tbl = tbl.combine_chunks()
r_tbl = pyra.converter.py2rpy(cb_tbl)
```
The table can then used in R, allowing to take advantage of individual strengths in a data science team
(some prefer writing R code) or libraries in R for which there is arguably no matching equivalent in Python.
```
%load_ext rpy2.ipython
%%R -i r_tbl
print(class(r_tbl))
%%R -i r_tbl
library(dplyr, warn.conflicts = FALSE)
r_tbl %>%
mutate(tip_group = round(tip_amount / 5) * 5) %>%
count(tip_group)
%%R -w 800 -h 600 --type cairo-png
library(ggplot2, warn.conflicts = FALSE)
library(viridis)
options(bitmapType="cairo")
X11.options(antialias = "subpixel")
p <- ggplot(r_tbl %>% collect()) +
geom_hex(aes(x = fare_amount, y = tip_amount/fare_amount),
bins = 75) +
scale_fill_viridis(trans="log10") +
scale_y_continuous("tip", labels = scales::percent, trans="log10") +
ggtitle("Tip as a percentage of the fare") +
theme_gray(base_size=19)
print(p)
```
| github_jupyter |
# Sandboxing some pytorch and pyro stuff
```
import torch
import numpy as np
import pyro
%matplotlib inline
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
```
## Pytorch Blitz
### What is pytorch?
let's omit this shit...
### AUTOGRAD: Automatic Differentiation
scalar function grad
```
x = torch.ones(2, 2, requires_grad=True)
y = x + 2
y.grad_fn
z = y*y*3
out=z.mean()
out
out.backward(torch.tensor(2.))
print(x.grad)
```
vector function grad...
```
x=torch.tensor([1., 2., 3.], requires_grad=True)
y=2*x*x
v = torch.tensor([1., 1., 2.])
y.backward(v)
x.grad
x = torch.tensor([[0., 0.], [1., 0]], requires_grad=True)
beta=1.
u = beta * ((x[0,:] - x[1,:]).norm(p=2) - 1.1)**2
u.backward()
x.grad
```
for simplicity's sake, let's create a 1-dimensional potential...
```
x = torch.tensor([[0., 0.], [1., 0.], [0., 1.]], requires_grad=True)
u = ()
```
## Intro to Pyro
```
import pyro
pyro.set_rng_seed(101)
#draw a sample from a standard normal
loc=0; scale=1.
normal=torch.distributions.Normal(loc, scale)
x=normal.rsample()
energy=-normal.log_prob(x)
print(x, energy)
def weather():
cloudy=pyro.sample('cloudy', pyro.distributions.Bernoulli(0.3))
cloudy='cloudy' if cloudy.item()==1. else 'sunny'
mean_temp={'cloudy':55., 'sunny': 75.}['cloudy']
scale_temp = {'cloudy': 10.0, 'sunny': 15.0}[cloudy]
temp = pyro.sample('temp', pyro.distributions.Normal(mean_temp, scale_temp))
return cloudy, temp.item()
weather()
def normal_product(loc, scale):
z1 = pyro.sample("z1", pyro.distributions.Normal(loc, scale))
z2 = pyro.sample("z2", pyro.distributions.Normal(loc, scale))
y = z1 * z2
return y
def make_normal_normal():
mu_latent = pyro.sample("mu_latent", pyro.distributions.Normal(0, 1))
fn = lambda scale: normal_product(mu_latent, scale)
return fn
print(make_normal_normal()(1.))
```
### SVI part 1: an intro to stochastic variational inference in pyro
we define a joint distribution $$p(x, z | \theta) = p(x|z, \theta)p(z|\theta)$$
where we want to maximize the log evidence (i.e. $\log p(x|\theta)$
`model` is the posterior, whereas `guide` is a family of distributions $q_{\phi}(z)$ that can be optimized w.r.t. $\phi$ (i.e. the variational parameters). To be concrete, we need to be able to compute the distribution of the latent variables given the data (aka the posterior)
$$p(z|x, \theta) = \frac {p(x, z | \theta)} {\int dz p(x, z | \theta)}$$
#### ELBO (Evidence Lower Bound)
$$
ELBO =
\langle
\log p(x, z | \theta) - \log q_{\phi}(z)
\rangle
_{q_{\phi}(z)}
$$
### Class
in order to do VI, we use `SVI`, which supports ELBO optimization.<br>
The user needs to provide the model, the guide, and the optimizer...
```
import pyro
from pyro.infer import SVI, Trace_ELBO
svi = SVI(model, guide, optimizer, loss=Trace_ELBO())
```
### Simple Coin Example
you have a 2 sided coin and you want to determine whether it's fair; really, you are trying to determine the latent bias parameter;<br>
- encode heads and tails as 1 ,0 <br>
- the fairness is $f$, where $f \in \left[ 0.0, 1.0 \right]$
- our prior $p(f) = \text{Beta}(10, 10)$
```
xs = torch.linspace(0,1)
beta = pyro.distributions.Beta(10, 10)
log_probs = torch.tensor([beta.log_prob(x) for x in xs[1:-1]])
xs = torch.linspace(0,1)
beta = pyro.distributions.Beta(10, 10)
log_probs = torch.tensor([beta.log_prob(x) for x in xs[1:-1]])
plt.plot(xs[1:-1], np.exp(log_probs))
plt.xlabel('f')
plt.ylabel("p(f)")
plt.title(f"prior")
```
- let's say we have collected the data in a list; let's write the corresponding model..
```
import pyro.distributions as dist
import torch.distributions.constraints as constraints
def model(data):
"""
the model is the product of the likelihood and the prior;
define samples of the likelihood with `obs` arg for all of the data
"""
alpha0 = torch.tensor(10.) #alpha hyperparam
beta0 = torch.tensor(10.) #beta hyperparam
#sample an f from the prior given prior hyperparams
f = pyro.sample('latent fairness', dist.Beta(alpha0, beta0))
for i in range(len(data)):
pyro.sample(f"obs_{i}", dist.Bernoulli(f), obs=data[i])
def guide(data):
"""
define a guide, which is a family of distributions on the latent variables
that is parameterized by `\phi` variables
"""
#in this case, we are going to optimize 2 parameters: alpha_q and beta_q
#and the parameters will parameterize a family of distributions of the beta form
alpha_q = pyro.param('alpha_q', torch.tensor(15.), constraint=constraints.positive)
beta_q = pyro.param('beta_q', torch.tensor(15.), constraint=constraints.positive)
#and just sample the latent fairness from the distribution
pyro.sample('latent_fairness', dist.Beta(alpha_q, beta_q))
```
Note:
- the model and the guide have the same latent variable names
- the model is not an explicit function of the latent variable, but instead set by data
- the guide also holds `data` arguments
- the variational parameters are torch tensors, and `pyro.param` automatically sets `requires_grad` to `True`
- we put positive constraints on the variational parameters
now we can to Stochastic Variational Inference (SVI)
```
from pyro.optim import Adam
from pyro.infer import SVI, Trace_ELBO
import tqdm
```
we just need to add some data first...
```
data = []
for _ in range(6):
data.append(torch.tensor(1.))
for _ in range(4):
data.append(torch.tensor(0.))
adam_params = {'lr': 5e-4, 'betas': (0.9, 0.999)}
optimizer=Adam(adam_params)
svi=SVI(model, guide, optimizer, loss=Trace_ELBO())
n_steps=5000
for step in tqdm.trange(n_steps):
svi.step(data)
alpha_q = pyro.param('alpha_q').item()
beta_q = pyro.param('beta_q').item()
# here we use some facts about the beta distribution
# compute the inferred mean of the coin's fairness
inferred_mean = alpha_q / (alpha_q + beta_q)
# compute inferred standard deviation
factor = beta_q / (alpha_q * (1.0 + alpha_q + beta_q))
inferred_std = inferred_mean * np.sqrt(factor)
print("\nbased on the data and our prior belief, the fairness " +
"of the coin is %.3f +- %.3f" % (inferred_mean, inferred_std))
```
### SVI part 2: conditional independence, subsampling, and amortization
for models with N observables, running `model` and `guide` requires `ELBO` estimates that scales poorly with N; but we can subsample in batches provided that there are some conditional independencies that we can take advantage of. <br>
For example, if the observations are conditionally independent given latent variables, the log likelihood can be approximated as
$$
\sum_{i=1}^{N}\ln p(x_i|z) \approx \frac{N}{M} \sum_{i \in I_M} \ln p(x_i|z)
$$
where $I_M$ is a minibatch of the N data points
for some reason in pyro, it is necessary to explcitly replace the `for` loop in `model` (the one that adds the data as `obs`)
```
def model(data):
"""
the model is the product of the likelihood and the prior;
define samples of the likelihood with `obs` arg for all of the data
"""
alpha0 = torch.tensor(10.) #alpha hyperparam
beta0 = torch.tensor(10.) #beta hyperparam
#sample an f from the prior given prior hyperparams
f = pyro.sample('latent fairness', dist.Beta(alpha0, beta0))
# for i in range(len(data)):
# pyro.sample(f"obs_{i}", dist.Bernoulli(f), obs=data[i])
#the above code has to be replaced with
for i in pyro.plate('data_loop', len(data)):
pyro.sample(f"obs_{i}", dist.Bernoulli(f), obs=data[i])
```
as a note, you can also vectorize `plate` like such:<br>
```
data=torch.zeros(10)
data[:6]=torch.ones()
with plate('observed_data'):
pyro.sample('obs', dist.Bernoulli(f), obs=data)
```
#### subsamping
subsample to run on large datasets...<br>
simply put, we can just subsample in the plate...<br>
```
for i in pyro.plate("data_loop", len(data), subsample_size=5):
pyro.sample("obs_{}".format(i), dist.Bernoulli(f), obs=data[i])
```
which,by the way, does automatic scaling;<br>
this can also be vectorized...<br>
```
with plate('observe_data', size=10, subsample_size=5) as ind:
pyro.sample('obs', dist.Bernoulli(f),
obs=data.index_select(0, ind))
```
However, if the data size is really big, theres a nonnegligible probability tat some dat apoints will never have been subsampled; you can use the `subsample` argument in `plate`
| github_jupyter |
```
import pickle
import numpy as np
import load
import matplotlib.pyplot as plt
import tree.halomodule as hmo
import utils.match as mtc
def bcg_prop(cat, verbose=False):
sort_mstar = cat['mstar'].argsort()
bcg = cat[sort_mstar[-1]] # BCG
sbcg = cat[sort_mstar[-2]] # 2nd BCG
dominance = 2.5* (np.log10(bcg.mstar) - np.log10(sbcg.mstar))
if verbose:
pass
#print(bcg.mstar, sbcg.mstar)
return bcg, dominance
def assign_recarr(recarr, ind, data, drop=None):
"""
If dtype varies among clusters (different version of lambda_gal),
available
"""
names = list(data.dtype.names)
if drop is not None:
for dd in drop:
names.remove(dd)
for fieldname in names:
recarr[fieldname][ind] = data[fieldname]
def halo_of_gal(halo, catalog, galid, dim6=False):
gal = catalog[catalog.id == galid]
center = np.array([gal.xc, gal.yc, gal.zc, gal.rhalo, gal.vx, gal.vy, gal.vz])
if dim6 :
norm = np.sqrt(np.square(center[0] - halo.x) +
np.square(center[1] - halo.y) +
np.square(center[2] - halo.z) +
np.square(center[3] - halo.rvir) +
np.square(center[4] - halo.vx) +
np.square(center[5] - halo.vy) +
np.square(center[6] - halo.vz))
else:
norm = np.sqrt(np.square(center[0] - halo.x) +
np.square(center[1] - halo.y) +
np.square(center[2] - halo.z) +
np.square(center[3] - halo.rvir))
i_match = norm.argmin()
return halo[i_match]
nout = 187
clusters = ['05427', '01605', '29172', '28928']
cdir = 'catalog_GM/'
# check if the clusters have relevant data
check_file=False
if check_file:
from glob import glob
for i, cluster in enumerate(clusters):
wdir = '/home/hoseung/Work/data/' + cluster + '/'
cat_list = glob("")
for file in glob(wdir + cdir + 'catalog' + str(nout) + '.pickle'):
print(file)
for file in glob(wdir + 'halo/DM/tree_bricks' + str(nout)):
print(file)
bcgs = np.zeros(len(clusters),
dtype=[('index', '<i8'), ('boxtokpc', '<f8'), ('id', '<i8'),
('idx', '<i8'), ('lambda_r', '<f8'),
('mgas', '<f8'), ('mstar', '<f8'), ('nstar', '<i8'),
('rgal', '<f8'), ('rhalo', '<f8'),
('vx', '<f8'), ('vy', '<f8'), ('vz', '<f8'),
('xc', '<f8'), ('yc', '<f8'), ('zc', '<f8'),
('mhalo', '<f8'), ('dominance', '<f8'), ('cluster', '<i8'),
('haloid', '<i8')])
# lambda_arr is removed from bcgs and also will be ignored in assign_recarr
# by drop=['lambda_arr'] option.
dominance =[]
rgal = []
mhalo = []
rhalo =[]
lambdar = []
mstar = []
for i, cluster in enumerate(clusters):
wdir = '/home/hoseung/Work/data/' + cluster + '/' #'05427/'
cat = pickle.load(open(wdir + cdir + 'catalog' + str(nout) + '.pickle', 'rb'))
bcg, dom = bcg_prop(cat, verbose=True)
# plot_gal_merger_history(cluster, bcg)
# exclude lambda_arr
assign_recarr(bcgs, i, bcg, drop=['lambda_arr'])
info = load.info.Info(base = wdir, nout=nout, load=True)
hh = hmo.Halo(nout=nout, base=wdir, info=info, halofinder="HM", is_gal=False, load=True)
hh.data.x *= info.pboxsize
hh.data.y *= info.pboxsize
hh.data.z *= info.pboxsize
hh.data.rvir *= info.pboxsize
halo = halo_of_gal(hh.data, cat, bcg.id) # halo data
halo = hh.data[hh.data.np.argmax()]
print("{:.4f} {:.4f} {:.4f} {:.4e}".format(halo['x'], halo['y'], halo['z'], halo.mvir))
print("{:.4f} {:.4f} {:.4f} {:.4e}".format(bcg['xc'], bcg['yc'], bcg['zc'], bcg.mstar))
#print(halo.mvir, hh.data.mvir[hh.data.np.argmax()])
rgal.append(np.log10(bcg['rgal'])) # in kpc
rhalo.append(np.log10(bcg['rhalo'] * info.pboxsize * 1000)) # in kpc (/h?)
mhalo.append(np.log10(halo['mvir']))
lambdar.append(bcg['lambda_r'])
mstar.append(np.log10(bcg['mstar']))
dominance.append(dom)
bcgs[i]['mhalo'] = halo['mvir']
bcgs[i]['dominance'] = dom
bcgs[i]['cluster'] = cluster
bcgs[i]['haloid'] = halo.id
#np.savetxt("ss.txt", bcgs)
np.save("Oliva_data.npy", bcgs)
```
# confirm halos matching
fig, axs = plt.subplots(2)
axs[0].plot(halos.id, cat.id)
axs[0].set_title("id vs id")
axs[1].plot(halos.rvir, cat.rhalo)
axs[1].set_title("rvir vs rvir")
plt.show()
```
#samples = bcgs
fig, axs = plt.subplots(3,3)
axs = axs.ravel()
#rgal = np.log10(samples['rgal']) # in kpc
#rhalo = np.log10(samples['rhalo'] * info.pboxsize * 1000) # in kpc (/h?)
#mhalo = np.log10(samples['mvir'])
#lambdar = samples['lambda_r']
#mstar = np.log10(samples['mstar'])
axs[0].scatter(mstar, lambdar, c = lambdar)
axs[0].set_title("rotation vs Mstar, fig3")
axs[1].scatter(rgal, lambdar, c = lambdar)
axs[1].set_title("rotation vs Rgal, fig7")
axs[2].scatter(mhalo, lambdar, c = lambdar)
axs[2].set_title("rotation vs Mhalo, fig8")
axs[3].scatter(dominance, lambdar, c = lambdar)
axs[3].set_title("rotation vs dominance, fig9")
axs[4].scatter(mstar, rgal, c = lambdar)
axs[4].set_title("Rgal vs Mstar, fig10")
axs[5].scatter(mhalo, rhalo, c = lambdar)
axs[5].set_title("Mhalo vs Mstar, fig11")
plt.suptitle("nout = {}, z= {:.3f}".format(str(nout), info.zred))
plt.tight_layout()
#plt.show()
plt.savefig('Oliva_fig.png', dpi=200)
```
| github_jupyter |
## Data Types & Variable
### Syntax [ var/val identifier: Type = initialization ]
```
var company: String = "PT Company"
company = "PT Mahkota"
company
val company: String = "PT Company"
company = "PT Mahkota"
company
val firstWord = "PT "
val lastWord = "Mahkota"
firstWord + lastWord
val valueA: Int = 10
val valueB = 20
valueA + valueB
```
### Char
```
val character = 'A'
character
val character = 'ABC' // Incorrect character literal
var vocal = 'A'
println("Vocal " + vocal++)
println("Vocal " + vocal++)
println("Vocal " + vocal++)
println("Vocal " + vocal--)
println("Vocal " + vocal--)
println("Vocal " + vocal--)
println("Vocal " + vocal--)
```
### String
```
val textString = "Kotlin"
textString
val text = "Kotlin"
val firstChar = text[0]
print("First character of $text is $firstChar")
val text = "Kotlin"
for (char in text){
print("$char ")
}
```
### Escaped String
```
val statement = "Kotlin is "Awesome!""
statement
```
\t: menambah tab ke dalam teks.
\n: membuat baris baru di dalam teks.
\’: menambah karakter single quote kedalam teks.
\”: menambah karakter double quote kedalam teks.
\\: menambah karakter backslash kedalam teks.
```
val statement = "Kotlin is \"Awesome!\""
statement
```
### Unicode
```
val name = "Unicode test: \u00A9"
print(name)
```
### Raw String
```
val line = """
Line 1
Line 2
Line 3
Line 4
""".trimIndent()
print(line)
```
### Boolean
#### Conjunction atau AND (&&)
```
val officeOpen = 7
val officeClosed = 16
val now = 20
val isOpen = now >= officeOpen && now <= officeClosed
print("Office is open : $isOpen")
```
#### Disjunction atau OR (||)
```
val officeOpen = 7
val officeClosed = 16
val now = 20
val isClose = now < officeOpen || now > officeClosed
print("Office is closed : $isClose")
```
#### Negation atau NOT (!)
```
val officeOpen = 7
val now = 10
val isOpen = now > officeOpen
if (!isOpen) {
print("Office is closed")
} else {
print("Office is open")
}
```
### Number (Integer)
#### Byte (8 Bit)
```
val maxByte = Byte.MAX_VALUE
val minByte = Byte.MIN_VALUE
println(maxByte)
println(minByte)
val byteNumber = 0b11010010
byteNumber
```
#### Short (16 Bit)
```
val maxShort = Short.MAX_VALUE
val minShort = Short.MIN_VALUE
println(maxShort)
println(minShort)
val shortNumber: Short = 10
shortNumber
```
#### Int (32 Bit) range -2^31 sampai +2^31-1
```
val maxInt = Int.MAX_VALUE
val minInt = Int.MIN_VALUE
println(maxInt)
println(minInt)
val intNumber = 100
intNumber
val byteNumber: Byte = 1
val intNumber: Int = byteNumber // compile error
val byteNumber: Byte = 10
val intNumber: Int = byteNumber.toInt() // ready to go
intNumber
```
#### Long (64 Bit) range -2^63 sampai +2^63-1
```
val maxLong = Long.MAX_VALUE
val minLong = Long.MIN_VALUE
println(maxLong)
println(minLong)
val longNumber: Long = 100
longNumber
val longNumber = 100L
longNumber
```
### Number (Floating Point)
### Float (32 Bit)
```
val maxFloat = Float.MAX_VALUE
val minFloat = Float.MIN_VALUE
println(maxFloat)
println(minFloat)
val floatNumber: Float = 0.123456789f
floatNumber
```
### Double (64 Bit)
```
val maxDouble = Double.MAX_VALUE
val minDouble = Double.MIN_VALUE
println(maxDouble)
println(minDouble)
val doubleNumber: Double = 1.3
doubleNumber
```
### Arrays
```
val array = arrayOf(1, 3, 5, 7)
array.contentToString()
val mixArray = arrayOf(1, 3, 5, 7 , "Indra" , true)
mixArray.contentDeepToString()
val intArray = Array(4, { i -> i * i }) // [0, 1, 4, 9]
intArray.contentToString()
```
| github_jupyter |
# Transforms and Resampling <a href="https://mybinder.org/v2/gh/InsightSoftwareConsortium/SimpleITK-Notebooks/master?filepath=Python%2F21_Transforms_and_Resampling.ipynb"><img style="float: right;" src="https://mybinder.org/badge_logo.svg"></a>
This notebook explains how to apply transforms to images, and how to perform image resampling.
```
import SimpleITK as sitk
import numpy as np
%matplotlib inline
import gui
from matplotlib import pyplot as plt
from ipywidgets import interact, fixed
# Utility method that either downloads data from the Girder repository or
# if already downloaded returns the file name for reading from disk (cached data).
%run update_path_to_download_script
from downloaddata import fetch_data as fdata
```
## Creating and Manipulating Transforms
A number of different spatial transforms are available in SimpleITK.
The simplest is the Identity Transform. This transform simply returns input points unaltered.
```
dimension = 2
print("*Identity Transform*")
identity = sitk.Transform(dimension, sitk.sitkIdentity)
print("Dimension: " + str(identity.GetDimension()))
# Points are always defined in physical space
point = (1.0, 1.0)
def transform_point(transform, point):
transformed_point = transform.TransformPoint(point)
print("Point " + str(point) + " transformed is " + str(transformed_point))
transform_point(identity, point)
```
Transform are defined by two sets of parameters, the *Parameters* and *FixedParameters*. *FixedParameters* are not changed during the optimization process when performing registration. For the TranslationTransform, the Parameters are the values of the translation Offset.
```
print("*Translation Transform*")
translation = sitk.TranslationTransform(dimension)
print("Parameters: " + str(translation.GetParameters()))
print("Offset: " + str(translation.GetOffset()))
print("FixedParameters: " + str(translation.GetFixedParameters()))
transform_point(translation, point)
print("")
translation.SetParameters((3.1, 4.4))
print("Parameters: " + str(translation.GetParameters()))
transform_point(translation, point)
```
The affine transform is capable of representing translations, rotations, shearing, and scaling.
```
print("*Affine Transform*")
affine = sitk.AffineTransform(dimension)
print("Parameters: " + str(affine.GetParameters()))
print("FixedParameters: " + str(affine.GetFixedParameters()))
transform_point(affine, point)
print("")
affine.SetTranslation((3.1, 4.4))
print("Parameters: " + str(affine.GetParameters()))
transform_point(affine, point)
```
A number of other transforms exist to represent non-affine deformations, well-behaved rotation in 3D, etc. See the [Transforms](22_Transforms.ipynb) tutorial for more information.
## Applying Transforms to Images
Create a function to display the images that is aware of image spacing.
```
def myshow(img, title=None, margin=0.05, dpi=80):
nda = sitk.GetArrayViewFromImage(img)
spacing = img.GetSpacing()
ysize = nda.shape[0]
xsize = nda.shape[1]
figsize = (1 + margin) * ysize / dpi, (1 + margin) * xsize / dpi
fig = plt.figure(title, figsize=figsize, dpi=dpi)
ax = fig.add_axes([margin, margin, 1 - 2 * margin, 1 - 2 * margin])
extent = (0, xsize * spacing[1], 0, ysize * spacing[0])
t = ax.imshow(
nda, extent=extent, interpolation="hamming", cmap="gray", origin="lower"
)
if title:
plt.title(title)
```
Create a grid image.
```
grid = sitk.GridSource(
outputPixelType=sitk.sitkUInt16,
size=(250, 250),
sigma=(0.5, 0.5),
gridSpacing=(5.0, 5.0),
gridOffset=(0.0, 0.0),
spacing=(0.2, 0.2),
)
myshow(grid, "Grid Input")
```
To apply the transform, a resampling operation is required.
```
def resample(image, transform):
# Output image Origin, Spacing, Size, Direction are taken from the reference
# image in this call to Resample
reference_image = image
interpolator = sitk.sitkCosineWindowedSinc
default_value = 100.0
return sitk.Resample(image, reference_image, transform, interpolator, default_value)
translation.SetOffset((3.1, 4.6))
transform_point(translation, point)
resampled = resample(grid, translation)
myshow(resampled, "Resampled Translation")
```
What happened? The translation is positive in both directions. Why does the output image move down and to the left? It important to keep in mind that a transform in a resampling operation defines *the transform from the output space to the input space*.
```
translation.SetOffset(-1 * np.array(translation.GetParameters()))
transform_point(translation, point)
resampled = resample(grid, translation)
myshow(resampled, "Inverse Resampled")
```
An affine (line preserving) transformation, can perform translation:
```
def affine_translate(transform, x_translation=3.1, y_translation=4.6):
new_transform = sitk.AffineTransform(transform)
new_transform.SetTranslation((x_translation, y_translation))
resampled = resample(grid, new_transform)
myshow(resampled, "Translated")
return new_transform
affine = sitk.AffineTransform(dimension)
interact(
affine_translate,
transform=fixed(affine),
x_translation=(-5.0, 5.0),
y_translation=(-5.0, 5.0),
);
```
or scaling:
```
def affine_scale(transform, x_scale=3.0, y_scale=0.7):
new_transform = sitk.AffineTransform(transform)
matrix = np.array(transform.GetMatrix()).reshape((dimension, dimension))
matrix[0, 0] = x_scale
matrix[1, 1] = y_scale
new_transform.SetMatrix(matrix.ravel())
resampled = resample(grid, new_transform)
myshow(resampled, "Scaled")
print(matrix)
return new_transform
affine = sitk.AffineTransform(dimension)
interact(affine_scale, transform=fixed(affine), x_scale=(0.2, 5.0), y_scale=(0.2, 5.0));
```
or rotation:
```
def affine_rotate(transform, degrees=15.0):
parameters = np.array(transform.GetParameters())
new_transform = sitk.AffineTransform(transform)
matrix = np.array(transform.GetMatrix()).reshape((dimension, dimension))
radians = -np.pi * degrees / 180.0
rotation = np.array(
[[np.cos(radians), -np.sin(radians)], [np.sin(radians), np.cos(radians)]]
)
new_matrix = np.dot(rotation, matrix)
new_transform.SetMatrix(new_matrix.ravel())
resampled = resample(grid, new_transform)
print(new_matrix)
myshow(resampled, "Rotated")
return new_transform
affine = sitk.AffineTransform(dimension)
interact(affine_rotate, transform=fixed(affine), degrees=(-90.0, 90.0));
```
or shearing:
```
def affine_shear(transform, x_shear=0.3, y_shear=0.1):
new_transform = sitk.AffineTransform(transform)
matrix = np.array(transform.GetMatrix()).reshape((dimension, dimension))
matrix[0, 1] = -x_shear
matrix[1, 0] = -y_shear
new_transform.SetMatrix(matrix.ravel())
resampled = resample(grid, new_transform)
myshow(resampled, "Sheared")
print(matrix)
return new_transform
affine = sitk.AffineTransform(dimension)
interact(affine_shear, transform=fixed(affine), x_shear=(0.1, 2.0), y_shear=(0.1, 2.0));
```
## Composite Transform
It is possible to compose multiple transform together into a single transform object. With a composite transform, multiple resampling operations are prevented, so interpolation errors are not accumulated. For example, an affine transformation that consists of a translation and rotation,
```
translate = (8.0, 16.0)
rotate = 20.0
affine = sitk.AffineTransform(dimension)
affine = affine_translate(affine, translate[0], translate[1])
affine = affine_rotate(affine, rotate)
resampled = resample(grid, affine)
myshow(resampled, "Single Transform")
```
can also be represented with two Transform objects applied in sequence with a Composite Transform,
```
composite = sitk.CompositeTransform(dimension)
translation = sitk.TranslationTransform(dimension)
translation.SetOffset(-1 * np.array(translate))
composite.AddTransform(translation)
affine = sitk.AffineTransform(dimension)
affine = affine_rotate(affine, rotate)
composite.AddTransform(translation)
composite = sitk.CompositeTransform(dimension)
composite.AddTransform(affine)
resampled = resample(grid, composite)
myshow(resampled, "Two Transforms")
```
*Beware*, tranforms are non-commutative -- order matters!
```
composite = sitk.CompositeTransform(dimension)
composite.AddTransform(affine)
composite.AddTransform(translation)
resampled = resample(grid, composite)
myshow(resampled, "Composite transform in reverse order")
```
## Resampling
<img src="resampling.svg"/><br><br>
Resampling as the verb implies is the action of sampling an image, which itself is a sampling of an original continuous signal.
Generally speaking, resampling in SimpleITK involves four components:
1. Image - the image we resample, given in coordinate system $m$.
2. Resampling grid - a regular grid of points given in coordinate system $f$ which will be mapped to coordinate system $m$.
2. Transformation $T_f^m$ - maps points from coordinate system $f$ to coordinate system $m$, $^mp = T_f^m(^fp)$.
3. Interpolator - method for obtaining the intensity values at arbitrary points in coordinate system $m$ from the values of the points defined by the Image.
While SimpleITK provides a large number of interpolation methods, the two most commonly used are ```sitkLinear``` and ```sitkNearestNeighbor```. The former is used for most interpolation tasks, a compromise between accuracy and computational efficiency. The later is used to interpolate labeled images representing a segmentation, it is the only interpolation approach which will not introduce new labels into the result.
SimpleITK's procedural API provides three methods for performing resampling, with the difference being the way you specify the resampling grid:
1. ```Resample(const Image &image1, Transform transform, InterpolatorEnum interpolator, double defaultPixelValue, PixelIDValueEnum outputPixelType)```
2. ```Resample(const Image &image1, const Image &referenceImage, Transform transform, InterpolatorEnum interpolator, double defaultPixelValue, PixelIDValueEnum outputPixelType)```
3. ```Resample(const Image &image1, std::vector< uint32_t > size, Transform transform, InterpolatorEnum interpolator, std::vector< double > outputOrigin, std::vector< double > outputSpacing, std::vector< double > outputDirection, double defaultPixelValue, PixelIDValueEnum outputPixelType)```
```
def resample_display(image, euler2d_transform, tx, ty, theta):
euler2d_transform.SetTranslation((tx, ty))
euler2d_transform.SetAngle(theta)
resampled_image = sitk.Resample(image, euler2d_transform)
plt.imshow(sitk.GetArrayFromImage(resampled_image))
plt.axis("off")
plt.show()
logo = sitk.ReadImage(fdata("SimpleITK.jpg"))
euler2d = sitk.Euler2DTransform()
# Why do we set the center?
euler2d.SetCenter(
logo.TransformContinuousIndexToPhysicalPoint(np.array(logo.GetSize()) / 2.0)
)
interact(
resample_display,
image=fixed(logo),
euler2d_transform=fixed(euler2d),
tx=(-128.0, 128.0, 2.5),
ty=(-64.0, 64.0),
theta=(-np.pi / 4.0, np.pi / 4.0),
);
```
### Common Errors
It is not uncommon to end up with an empty (all black) image after resampling. This is due to:
1. Using wrong settings for the resampling grid, not too common, but does happen.
2. Using the inverse of the transformation $T_f^m$. This is a relatively common error, which is readily addressed by invoking the transformations ```GetInverse``` method.
### Defining the Resampling Grid
In the example above we arbitrarily used the original image grid as the resampling grid. As a result, for many of the transformations the resulting image contained black pixels, pixels which were mapped outside the spatial domain of the original image and a partial view of the original image.
If we want the resulting image to contain all of the original image no matter the transformation, we will need to define the resampling grid using our knowledge of the original image's spatial domain and the **inverse** of the given transformation.
Computing the bounds of the resampling grid when dealing with an affine transformation is straightforward. An affine transformation preserves convexity with extreme points mapped to extreme points. Thus we only need to apply the **inverse** transformation to the corners of the original image to obtain the bounds of the resampling grid.
Computing the bounds of the resampling grid when dealing with a BSplineTransform or DisplacementFieldTransform is more involved as we are not guaranteed that extreme points are mapped to extreme points. This requires that we apply the **inverse** transformation to all points in the original image to obtain the bounds of the resampling grid.
```
euler2d = sitk.Euler2DTransform()
# Why do we set the center?
euler2d.SetCenter(
logo.TransformContinuousIndexToPhysicalPoint(np.array(logo.GetSize()) / 2.0)
)
tx = 64
ty = 32
euler2d.SetTranslation((tx, ty))
extreme_points = [
logo.TransformIndexToPhysicalPoint((0, 0)),
logo.TransformIndexToPhysicalPoint((logo.GetWidth(), 0)),
logo.TransformIndexToPhysicalPoint((logo.GetWidth(), logo.GetHeight())),
logo.TransformIndexToPhysicalPoint((0, logo.GetHeight())),
]
inv_euler2d = euler2d.GetInverse()
extreme_points_transformed = [inv_euler2d.TransformPoint(pnt) for pnt in extreme_points]
min_x = min(extreme_points_transformed)[0]
min_y = min(extreme_points_transformed, key=lambda p: p[1])[1]
max_x = max(extreme_points_transformed)[0]
max_y = max(extreme_points_transformed, key=lambda p: p[1])[1]
# Use the original spacing (arbitrary decision).
output_spacing = logo.GetSpacing()
# Identity cosine matrix (arbitrary decision).
output_direction = [1.0, 0.0, 0.0, 1.0]
# Minimal x,y coordinates are the new origin.
output_origin = [min_x, min_y]
# Compute grid size based on the physical size and spacing.
output_size = [
int((max_x - min_x) / output_spacing[0]),
int((max_y - min_y) / output_spacing[1]),
]
resampled_image = sitk.Resample(
logo,
output_size,
euler2d,
sitk.sitkLinear,
output_origin,
output_spacing,
output_direction,
)
plt.imshow(sitk.GetArrayViewFromImage(resampled_image))
plt.axis("off")
plt.show()
```
Are you puzzled by the result? Is the output just a copy of the input? Add a rotation to the code above and see what happens (```euler2d.SetAngle(0.79)```).
### Resampling at a set of locations
In some cases you may be interested in obtaining the intensity values at a set of points (e.g. coloring the vertices of a mesh model segmented from an image).
The code below generates a random point set in the image and resamples the intensity values at these locations. It is written so that it works for all image-dimensions and types (scalar or vector pixels).
```
img = logo
# Generate random samples inside the image, we will obtain the intensity/color values at these points.
num_samples = 10
physical_points = []
for pnt in zip(*[list(np.random.random(num_samples) * sz) for sz in img.GetSize()]):
physical_points.append(img.TransformContinuousIndexToPhysicalPoint(pnt))
# Create an image of size [num_samples,1...1], actual size is dependent on the image dimensionality. The pixel
# type is irrelevant, as the image is just defining the interpolation grid (sitkUInt8 has minimal memory footprint).
interp_grid_img = sitk.Image(
[num_samples] + [1] * (img.GetDimension() - 1), sitk.sitkUInt8
)
# Define the displacement field transformation, maps the points in the interp_grid_img to the points in the actual
# image.
displacement_img = sitk.Image(
[num_samples] + [1] * (img.GetDimension() - 1),
sitk.sitkVectorFloat64,
img.GetDimension(),
)
for i, pnt in enumerate(physical_points):
displacement_img[[i] + [0] * (img.GetDimension() - 1)] = np.array(pnt) - np.array(
interp_grid_img.TransformIndexToPhysicalPoint(
[i] + [0] * (img.GetDimension() - 1)
)
)
# Actually perform the resampling. The only relevant choice here is the interpolator. The default_output_pixel_value
# is set to 0.0, but the resampling should never use it because we expect all points to be inside the image and this
# value is only used if the point is outside the image extent.
interpolator_enum = sitk.sitkLinear
default_output_pixel_value = 0.0
output_pixel_type = (
sitk.sitkFloat32
if img.GetNumberOfComponentsPerPixel() == 1
else sitk.sitkVectorFloat32
)
resampled_points = sitk.Resample(
img,
interp_grid_img,
sitk.DisplacementFieldTransform(displacement_img),
interpolator_enum,
default_output_pixel_value,
output_pixel_type,
)
# Print the interpolated values per point
for i in range(resampled_points.GetWidth()):
print(
str(physical_points[i])
+ ": "
+ str(resampled_points[[i] + [0] * (img.GetDimension() - 1)])
+ "\n"
)
```
## <font color="red">Homework:</font> creating a color mesh
You will now use the code for resampling at arbitrary locations to create a colored mesh.
Using the color image of the [visible human](https://en.wikipedia.org/wiki/Visible_Human_Project) head [`img = sitk.ReadImage(fdata('vm_head_rgb.mha'))`]:
1. Implement the [marching cubes algorithm](https://en.wikipedia.org/wiki/Marching_cubes) to obtain the set of triangles corresponding to the iso-surface of structures of interest (skin, white matter,...).
2. Find the color associated with each of the triangle vertices using the code above.
3. Save the data using the ASCII version of the [PLY](https://en.wikipedia.org/wiki/PLY_(file_format)), Polygon File Format (a.k.a. Stanford Triangle Format).
4. Use [meshlab](http://www.meshlab.net/) to view your creation.
### Creating thumbnails - changing image size, spacing and intensity range
As bio-medical images are most often an-isotropic, have a non uniform size (number of pixels), with a high dynamic range of intensities, some caution is required when converting them to an arbitrary desired size with isotropic spacing and the more common low dynamic intensity range.
The code in the following cells illustrates how to take an arbitrary set of images with various sizes, spacings and intensities and resize all of them to a common arbitrary size, isotropic spacing, and low dynamic intensity range.
```
file_names = ["cxr.dcm", "photo.dcm", "POPI/meta/00-P.mhd", "training_001_ct.mha"]
images = []
image_file_reader = sitk.ImageFileReader()
for fname in file_names:
image_file_reader.SetFileName(fdata(fname))
image_file_reader.ReadImageInformation()
image_size = list(image_file_reader.GetSize())
# 2D image posing as a 3D one
if len(image_size) == 3 and image_size[2] == 1:
image_size[2] = 0
image_file_reader.SetExtractSize(image_size)
images.append(image_file_reader.Execute())
# 2D image
elif len(image_size) == 2:
images.append(image_file_reader.Execute())
# 3D image grab middle x-z slice
elif len(image_size) == 3:
start_index = [0, image_size[1] // 2, 0]
image_size[1] = 0
image_file_reader.SetExtractSize(image_size)
image_file_reader.SetExtractIndex(start_index)
images.append(image_file_reader.Execute())
# 4/5D image
else:
raise ValueError(f"{image.GetDimension()}D image not supported.")
# Notice that in the display the coronal slices are flipped. As we are
# using matplotlib for display, it is not aware of radiological conventions
# and treats the image as an isotropic array of pixels.
gui.multi_image_display2D(images);
```
## <font color="red">Homework:</font> Why do some of the images displayed above look different from others?
What are the differences between the various images in the `images` list? Write code to query them and check their intensity ranges, sizes and spacings.
The next cell illustrates how to resize all images to an arbitrary size, using isotropic spacing while maintaining the original aspect ratio.
```
def resize_and_scale_uint8(image, new_size, outside_pixel_value=0):
"""
Resize the given image to the given size, with isotropic pixel spacing
and scale the intensities to [0,255].
Resizing retains the original aspect ratio, with the original image centered
in the new image. Padding is added outside the original image extent using the
provided value.
:param image: A SimpleITK image.
:param new_size: List of ints specifying the new image size.
:param outside_pixel_value: Value in [0,255] used for padding.
:return: a 2D SimpleITK image with desired size and a pixel type of sitkUInt8
"""
# Rescale intensities if scalar image with pixel type that isn't sitkUInt8.
# We rescale first, so that the zero padding makes sense for all original image
# ranges. If we resized first, a value of zero in a high dynamic range image may
# be somewhere in the middle of the intensity range and the outer border has a
# constant but arbitrary value.
if (
image.GetNumberOfComponentsPerPixel() == 1
and image.GetPixelID() != sitk.sitkUInt8
):
final_image = sitk.Cast(sitk.RescaleIntensity(image), sitk.sitkUInt8)
else:
final_image = image
new_spacing = [
((osz - 1) * ospc) / (nsz - 1)
for ospc, osz, nsz in zip(
final_image.GetSpacing(), final_image.GetSize(), new_size
)
]
new_spacing = [max(new_spacing)] * final_image.GetDimension()
center = final_image.TransformContinuousIndexToPhysicalPoint(
[sz / 2.0 for sz in final_image.GetSize()]
)
new_origin = [
c - c_index * nspc
for c, c_index, nspc in zip(center, [sz / 2.0 for sz in new_size], new_spacing)
]
final_image = sitk.Resample(
final_image,
size=new_size,
outputOrigin=new_origin,
outputSpacing=new_spacing,
defaultPixelValue=outside_pixel_value,
)
return final_image
# Select the arbitrary new size
new_size = [128, 128]
resized_images = [resize_and_scale_uint8(image, new_size, 50) for image in images]
gui.multi_image_display2D(resized_images);
```
| github_jupyter |
# Compute Sci-Hub coverage of journals and publishers in Scopus
See all Crossref API types at https://api.crossref.org/types.
```
import collections
import json
import io
import lzma
import os
import pandas
import requests
with open('00.configuration.json') as read_file:
config = json.load(read_file)
config
```
## Read DOIs
```
# Read catalog of Crossref DOIs
path = os.path.join('data', 'doi.tsv.xz')
doi_df = pandas.read_table(path, parse_dates=['issued'])
doi_df['year'] = doi_df.issued.dt.year
def compute_coverage(df):
row = collections.OrderedDict()
row['scihub'] = sum(df.in_scihub_dois)
row['crossref'] = len(df)
row['coverage'] = df.in_scihub_dois.mean()
return row
```
# Compute coverage by article type
Compute type coverage before restricting to Scopus mapping DOIs.
```
results = doi_df.groupby(['type']).apply(compute_coverage)
type_coverage_df = pandas.DataFrame(list(results), index=results.index).reset_index()
type_coverage_df
path = os.path.join('data', 'type-coverage.tsv')
type_coverage_df.to_csv(path, index=False, sep='\t', float_format='%.5g')
```
# Compute coverage by year
Compute year coverage before restricting to Scopus mapping DOIs.
```
results = (
doi_df
.query("year <= 2017")
.groupby(['year'])
.apply(compute_coverage)
)
year_coverage_df = pandas.DataFrame(list(results), index=results.index).reset_index()
year_coverage_df.year = year_coverage_df.year.astype(int)
year_coverage_df.head(3)
path = os.path.join('data', 'year-coverage.tsv')
year_coverage_df.to_csv(path, index=False, sep='\t', float_format='%.5g')
```
## Compute journal coverage
The remainder of the coverage datasets refer to only articles attributed to a Scopus journal.
```
# One DOI can map to multiple journals, so these values are best calcuated before merging with the journal mapping
doi_df = doi_df.query("in_scopus == 1")
print(f'''\
{len(doi_df):,} articles attributed to a Scopus journal
Sci-Hub contains {doi_df.in_scihub_dois.sum():,} ({doi_df.in_scihub_dois.mean():.1%})
''')
# Read Scopus title names and attributes
url = config['scopus_url'] + 'data/titles.tsv'
journal_df = pandas.read_table(url)
url = config['scopus_url'] + 'data/title-attributes.tsv'
journal_attributes_df = pandas.read_table(url)
journal_df = (
journal_df
.merge(journal_attributes_df)
[['scopus_id', 'title_name', 'active', 'open_access']]
)
journal_df.head(2)
# Map DOIs to scopus titles
path = os.path.join('data', 'scopus-title-to-doi-map.tsv.xz')
doi_df = (
pandas.read_table(path)
.merge(doi_df)
)
doi_df.head(3)
results = doi_df.groupby(['scopus_id']).apply(compute_coverage)
journal_coverage_df = pandas.DataFrame(list(results), index=results.index).reset_index()
journal_coverage_df = journal_df.merge(journal_coverage_df)
journal_coverage_df.sort_values('crossref', ascending=False).head(4)
path = os.path.join('data', 'journal-coverage.tsv')
journal_coverage_df.to_csv(path, index=False, sep='\t', float_format='%.5g')
```
## Compute journal-year coverage
```
results = (
doi_df
.query("year <= 2017")
.groupby(['scopus_id', 'year'])
.apply(compute_coverage)
)
journal_year_coverage_df = pandas.DataFrame(list(results), index=results.index).reset_index()
journal_year_coverage_df.year = journal_year_coverage_df.year.astype(int)
journal_year_coverage_df.head(3)
path = os.path.join('data', 'journal-year-coverage.tsv.xz')
journal_year_coverage_df.to_csv(path, index=False, sep='\t', float_format='%.5g', compression='xz')
```
## Compute year by access-status coverage
```
def compute_coverage_by_access(df):
row = compute_coverage(df)
row['scihub_journals'] = df.query("in_scihub_dois == 1").scopus_id.nunique()
row['crossref_journals'] = df.scopus_id.nunique()
return row
results = (
doi_df
.merge(journal_df)
.query("year <= 2017")
.groupby(['year', 'open_access'])
.apply(compute_coverage_by_access)
)
year_access_coverage_df = pandas.DataFrame(list(results), index=results.index).reset_index()
year_access_coverage_df.year = year_access_coverage_df.year.astype(int)
year_access_coverage_df.tail(4)
path = os.path.join('data', 'year-coverage-by-access.tsv')
year_access_coverage_df.to_csv(path, index=False, sep='\t', float_format='%.5g')
```
## Compute publisher-year by access-status coverage
```
results = (
journal_attributes_df
[['scopus_id', 'open_access', 'main_publisher']]
.merge(doi_df)
.query("year <= 2017")
.groupby(['main_publisher', 'open_access', 'year'])
.apply(compute_coverage_by_access)
)
year_publisher_coverage_df = pandas.DataFrame(list(results), index=results.index).reset_index()
year_publisher_coverage_df.year = year_publisher_coverage_df.year.astype(int)
year_publisher_coverage_df.tail(4)
path = os.path.join('data', 'year-publisher-coverage-by-access.tsv.xz')
year_publisher_coverage_df.to_csv(path, index=False, sep='\t', float_format='%.5g', compression='xz')
```
| github_jupyter |
```
from sklearn.datasets import make_classification
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, f1_score
import random
import numpy as np
import pandas as pd
import os
try:
import google.colab
IN_COLAB = True
except:
IN_COLAB = False
if not IN_COLAB:
os.chdir("..")
%load_ext autoreload
%autoreload 2
```
# Utility Functions
```
def make_imbalanced_mixed_classification(n_samples, n_features, n_categories):
X,y = make_classification(n_samples=n_samples, n_features=n_features, random_state=42, n_informative=5, weights=[0.7], flip_y=0.3)
cat_cols = random.choices(list(range(X.shape[-1])),k=n_categories)
num_cols = [i for i in range(X.shape[-1]) if i not in cat_cols]
for col in cat_cols:
X[:,col] = pd.qcut(X[:,col], q=4).codes.astype(int)
col_names = []
num_col_names=[]
cat_col_names=[]
for i in range(X.shape[-1]):
if i in cat_cols:
col_names.append(f"cat_col_{i}")
cat_col_names.append(f"cat_col_{i}")
if i in num_cols:
col_names.append(f"num_col_{i}")
num_col_names.append(f"num_col_{i}")
X = pd.DataFrame(X, columns=col_names)
y = pd.Series(y, name="target")
data = X.join(y)
return data, cat_col_names, num_col_names
def print_metrics(y_true, y_pred, tag):
if isinstance(y_true, pd.DataFrame) or isinstance(y_true, pd.Series):
y_true = y_true.values
if isinstance(y_pred, pd.DataFrame) or isinstance(y_pred, pd.Series):
y_pred = y_pred.values
if y_true.ndim>1:
y_true=y_true.ravel()
if y_pred.ndim>1:
y_pred=y_pred.ravel()
val_acc = accuracy_score(y_true, y_pred)
val_f1 = f1_score(y_true, y_pred)
print(f"{tag} Acc: {val_acc} | {tag} F1: {val_f1}")
```
# Generate Synthetic Data
First of all, let's create a synthetic data which is a mix of numerical and categorical features
```
data, cat_col_names, num_col_names = make_imbalanced_mixed_classification(n_samples=10000, n_features=20, n_categories=4)
train, test = train_test_split(data, random_state=42)
train, val = train_test_split(train, random_state=42)
```
# Importing the Library
```
from pytorch_tabular import TabularModel
from pytorch_tabular.models import CategoryEmbeddingModelConfig
from pytorch_tabular.config import DataConfig, OptimizerConfig, TrainerConfig, ExperimentConfig
```
## Define the Configs
```
data_config = DataConfig(
target=['target'], #target should always be a list. Multi-targets are only supported for regression. Multi-Task Classification is not implemented
continuous_cols=num_col_names,
categorical_cols=cat_col_names,
)
trainer_config = TrainerConfig(
auto_lr_find=True, # Runs the LRFinder to automatically derive a learning rate
batch_size=1024,
max_epochs=100,
gpus=-1, #index of the GPU to use. -1 means all available GPUs, None, means CPU
)
optimizer_config = OptimizerConfig()
model_config = CategoryEmbeddingModelConfig(
task="classification",
layers="1024-512-512", # Number of nodes in each layer
activation="LeakyReLU", # Activation between each layers
learning_rate = 1e-3,
metrics=["f1","accuracy"],
metrics_params=[{"num_classes":2},{}]
)
```
## Training the Model
```
tabular_model = TabularModel(
data_config=data_config,
model_config=model_config,
optimizer_config=optimizer_config,
trainer_config=trainer_config,
)
tabular_model.fit(train=train, validation=val)
result = tabular_model.evaluate(test)
```
## Custom Sampler
PyTorch Tabular also allows custom batching strategy through Custom Samplers which comes in handy when working with imbalanced data.
Although you can use any sampler, Pytorch Tabular has a few handy utility functions which takes in the target array and implements WeightedRandomSampler using inverse frequency sampling to combat imbalance. This is analogous to preprocessing techniques like Under or OverSampling in traditional ML systems.
```
from pytorch_tabular.utils import get_balanced_sampler, get_class_weighted_cross_entropy
tabular_model = TabularModel(
data_config=data_config,
model_config=model_config,
optimizer_config=optimizer_config,
trainer_config=trainer_config,
)
sampler = get_balanced_sampler(train['target'].values.ravel())
tabular_model.fit(train=train, validation=val, train_sampler=sampler)
result = tabular_model.evaluate(test)
```
## Custom Weighted Loss
If Samplers were like Over/Under Sampling, Custom Weighted Loss is similar to `class_weights`. Depending on the problem, one of these might help you with imbalance. You can easily make calculate the class_weights and provide them to the CrossEntropyLoss using the parameter `weight`. To make this easier, PyTorch Tabular has a handy utility method which calculates smoothed class weights and initializes a weighted loss. Once you have that loss, it's just a matter of passing it to the 1fit1 method using the `loss` parameter.
```
tabular_model = TabularModel(
data_config=data_config,
model_config=model_config,
optimizer_config=optimizer_config,
trainer_config=trainer_config,
)
weighted_loss = get_class_weighted_cross_entropy(train["target"].values.ravel(), mu=0.1)
tabular_model.fit(train=train, validation=val, loss=weighted_loss)
result = tabular_model.evaluate(test)
```
| github_jupyter |

# Introduction to python3
## Python Features
### 1-Easy-to-learn
### 2-Easy-to-read
### 3-Easy-to-maintain
### 4-A broad standard library − Python's bulk of the library is very portable
### 5-and cross-platform compatible on UNIX, Windows, and Macintosh.
### 6-Portable − Python can run on a wide variety of hardware platforms and has the same interface on all platforms.
# python vs nodejs
Math.pow(2,1000)
2**1000
# Variable Types
```
counter = 100 # An integer assignment
miles = 1000.0 # A floating point
name = "John" # A string
print ('counter->',counter)
print ('miles-->',miles)
print ('name-->',name)
print(type(name))
```
# Multiple Assignment
```
a=b=c=1
print(a)
c=d=e='reza',19.5,18
print(c[1])
```
# Standard Data Types
## Python has five standard data types −
### 1-Numbers
### 2-String
### 3-List
### 4-Tuple
### 5-Dictionary
# Python Numbers
```
a = 25
c=2564899756565
print('--------numbers---------->',c)
print(type(c))
```
# Python Strings
```
str = "Hello World!" #or '2564899756565'
print('-----------strings------------->',str)
print(type(str))
#Python Strings
str = 'Hello World!'
print(str[2]) # Prints first character of the string
print (str[2:7]) # Prints characters starting from 3rd to 5th
print (str[2:]) # Prints string starting from 3rd character
```
# Python Lists
```
list = [ 'abcd', 786 , 2.23, 'john', 70.2 ]
tinylist = [123, 'john']
print('-----------------------------')
print (list)
print('-----------------------------')
list[0]=2
print('------------>>',list)
print(list[2])
print('-----------------------------')
print(list[1:3])
print('-----------------------------')
print (tinylist * 3) # Prints list two times
```

# Python Tuples
```
tuple = ( 'abcd', 786 , 2.23, 'john', 70.2 )
print(tuple)
print(tuple)
print(tuple[0])
print (tuple[1:4])
```
# Python Dictionary
```
dict = {}
dict['one'] = "This is one"
dict[2]= "This is two"
print('dict--------->',dict)
```

```
test ={"name":["a",b],"lname":"bojnordi","country":"iran"}
print("---------------------------------------------------")
print("test------->",test)
print("---------------------------------------------------")
print (test.keys()) # Prints all the keys
print("---------------------------------------------------")
print (test.values()) # Prints all the values
```

```
# documentation
dataset =[{
"name":"reza",
"lname":"bojnordi",
"age":27,
"job":"web developer"
},
{
"name":"mahmmod",
"lname":"mahmodi",
"age":27,
"job":"web developer"
},
{
"name":"hashem",
"lname":"jahangiri",
"age":30,
"job":"android developer"
}
]
print('--------------------------------------------')
print('***********>',dataset[0])
print('dataset:',dataset[1]["lname"])
print('--------------------------------------------')
print('dataset:',dataset)
```
# Data Type Conversion
```
phone = "963258"
print("phone",type(phone))
print("type",type(phone))
print("phone------>",int(phone))
print("type",type(int(phone)))
```
# Decision Making
```
age = input("لطفا سن خود را وارد کنید:")
if(int(age) > 20):
print("-----1>",int(age) +2)
elif(int(age) < 20):
print(int(age) -2)
else:
print("noting")
```
# while Loop Statements
```
counte =0
while(counte < 9):
print ('The count is:', counte)
counte= counte + 1
print("Good Bye")
```
# Loops
```
list = [2,5,10,25]
for i in list:
print('list:',i)
print("------------------------------------------------")
```
# Nested loops
```
for i in range(1,11):
for j in range(1,11):
k = i * j
print (k, end=' ')
print()
```
# mean
```
list = [2,4,6,8]
a =0
for i in list:
a = a + i
print("mean:",a / len(list))
```
# Lists
```
list = ['physics', 'chemistry', 1997, 2000]
print ("Value available at index 2 : ", list[2])
list[2] = 2001
print ("New value available at index 2 : ", list[2])
```

# Basic List Operations
```
list = ['physics', 'chemistry', 1997, 2000]
print("Length:",len(list))
print("concat:",[2,5,8,4]+[87,45,6])
print("Membership:", 3 in [3,5,9,58,45])
print("max:",max([55,12,4,69,4,12,1,50,80,4]))
print("min:",min([55,12,4,69,4,12,1,50,80,4]))
```
# append
```
list1 = ['python','ruby','node','c++']
list1.append(2)
print('list1:',list1)
print('pop list:',list1.pop(0))
print('pop list--->:',list1)
```
# sort
```
list2 = [50,6,25,45,7,78,23,5]
list3 = [50,6,25,45,7,78,23,5]
list2.sort()
list2.sort(reverse=True)
print(list2)
```
# Dictionary
```
dict = {'Name': 'Zara', 'Age': 7, 'Class': 'First'}
print ("dict['Name']: ", dict['Name'])
print ("dict['Age']: ", dict['Age'])
```
# Error Keys
```
dict = {'Name': 'Zara', 'Age': 7, 'Class': 'First'};
print ("dict['Alice']: ", dict['Age'])
```
# Updating Dictionary
```
dict = {'Name': 'Zara', 'Age': 7, 'Class': 'First'}
print(dict)
dict['Age'] = 8; # update existing entry
dict['School'] = "DPS School" # Add new entry
print ("dict['Age']: ", dict['Age'])
print ("dict['School']: ", dict['School'])
print('---------------------------->',dict)
print(dict.keys())
print(dict.values())
```
# len
```
dict = {'Name': 'Manni', 'Age': 7, 'Class': 'First'}
print ("Length : %d" % len (dict))
```
# dictionary copy()
```
dict1 = {'Name': 'Manni', 'Age': 7, 'Class': 'First'}
dict2 = dict1.copy()
print ("New Dictionary : ",dict2)
```
# Functions
```
# Function definition is here
def printme(str):
"This prints a passed string into this function"
print ('------>',str)
# Now you can call printme function
printme("This is first call to the user defined function!")
printme("Again second call to the same function")
def Multiplication (x,b):
return x * b
print(Multiplication(6,70))
```
# random in numpy
```
import numpy as np
print("Normal distribution",np.random.randn(1,10))
print('-------------------------------------------------------------------')
print("Uniform distribution",np.random.uniform(0 ,1 , 10))
import numpy as np
estimated_action_value = np.zeros(10)
print(estimated_action_value)
```
# numpy.matmul
```
a = [[2, 0],
[5, 1]
]
b = [
[4, 1],
[2, 2]
]
print(np.matmul(a, b))
```
# source:https://www.tutorialspoint.com/python3/
| github_jupyter |
```
from collections import defaultdict, deque
class Jump(Exception):
def __init__(self, offset):
self.offset = offset
super().__init__(offset)
def opcode(operands):
def decorator(f):
class Opcode:
def __set_name__(self, owner, name):
self.opcode = name[3:]
owner.opcodes[self.opcode] = self
def __repr__(self):
return f'<opcode {self.opcode} {operands!r}>'
def value(self, operand, type_):
if type_ == 'r':
return operand
try:
return int(operand)
except ValueError:
return self.registers[operand]
def __call__(self, cpu, *ops):
self.registers = cpu.registers
try:
result = f(cpu, *map(self.value, ops, operands))
cpu.pos += 1
except Jump as j:
cpu.pos += j.offset
result = None
return self.opcode, result
return Opcode()
return decorator
class Proc:
opcodes = {}
def __init__(self):
self.reset()
def reset(self):
self.registers = defaultdict(int)
self.sound_freq = 0
self.pos = 0
def run(self, instructions):
while True:
opcode, *ops = instructions[self.pos]
yield self.opcodes[opcode](self, *ops)
def run_until_rcv(self, instructions):
return next(
val
for op, val in self.run(instructions)
if op == 'rcv' and val is not None)
@opcode('v')
def op_snd(self, x):
self.sound_freq = x
@opcode('rv')
def op_set(self, x, y):
self.registers[x] = y
@opcode('rv')
def op_add(self, x, y):
self.registers[x] += y
@opcode('rv')
def op_mul(self, x, y):
self.registers[x] *= y
@opcode('rv')
def op_mod(self, x, y):
self.registers[x] %= y
@opcode('r')
def op_rcv(self, x):
if self.registers[x]:
return self.sound_freq
@opcode('vv')
def op_jgz(self, x, y):
if x > 0:
raise Jump(y)
class SendingProc(Proc):
opcodes = Proc.opcodes.copy()
def __init__(self, cpu_id):
self.cpu_id = cpu_id
super().__init__()
def set_pair(self, cpu):
self.paired = cpu
def reset(self):
super().reset()
self.message_queue = deque()
self.registers['p'] = self.cpu_id
@opcode('v')
def op_snd(self, x):
self.paired.message_queue.append(x)
@opcode('r')
def op_rcv(self, x):
if not self.message_queue:
raise Jump(0)
value = self.message_queue.popleft()
self.registers[x] = value
return value
def parallel_run(instructions):
cpu0 = SendingProc(0)
cpu1 = SendingProc(1)
cpu0.set_pair(cpu1)
cpu1.set_pair(cpu0)
sendcount = 0
for (op0, res0), (op1, res1) in zip(cpu0.run(instructions), cpu1.run(instructions)):
if op1 == 'snd':
sendcount += 1
if (op0, res0, op1, res1) == ('rcv', None, 'rcv', None):
# deadlock
return sendcount
proc = Proc()
test_instr = [instr.split() for instr in '''\
set a 1
add a 2
mul a a
mod a 5
snd a
set a 0
rcv a
jgz a -1
set a 1
jgz a -2
'''.splitlines()]
assert proc.run_until_rcv(test_instr) == 4
test_instr = [instr.split() for instr in '''\
snd 1
snd 2
snd p
rcv a
rcv b
rcv c
rcv d
'''.splitlines()]
assert parallel_run(test_instr) == 3
import aocd
data = aocd.get_data(day=18, year=2017)
instructions = [line.split() for line in data.splitlines()]
proc = Proc()
print('Part 1:', proc.run_until_rcv(instructions))
print('Part 2:', parallel_run(instructions))
```
| github_jupyter |
```
%matplotlib inline
import gym
import matplotlib.pyplot as plt
import torch
import torch.nn.functional as F
from torch.optim import Adam
from torch import nn
from torch.autograd import Variable
from utils import grid_from_state, process_state
from tutorial import select_action, calculate_epsilon
from models import DQN
from memory import ReplayMemory
from atari_wrappers import wrap_deepmind
```
## Double Q learning
- https://arxiv.org/pdf/1509.06461.pdf
- Stabilizes training and sometimes improves performance.
- ~5 lines of code
```
env = gym.make('Breakout-v0')
env = wrap_deepmind(env, frame_stack=True)
_ = env.reset()
action = env.action_space.sample()
state, reward, _, _ = env.step(action) # carry out action/observe reward
next_state, _, _, _ = env.step(env.action_space.sample()) # carry out action/observe reward
plt.imshow(grid_from_state(state));
```
use a second DQN to compute `next_q_values` The target network,
with parameters θ−, is the same as the online network except
that its parameters are copied every τ steps from the online network,
so that then θ−t = θt, and kept fixed on all other steps.
- it is common to copy params every 1k-10k frames
## q network and target network are the same
```
q_network = DQN(env.action_space.n) # initialize action-value function Q with random weights
target_network = DQN(env.action_space.n)
q_network
target_network
```
## New target calculation process

```
s = Variable(process_state(state))
r = Variable(torch.Tensor([reward]))
s2 = Variable(process_state(next_state))
q_values = q_network(s)[:, action] # q vals for action we took
target_actions = q_network(s2).max(dim=1)[1]
next_q_values = target_network(s2)[range(len(target_actions)), target_actions]
targets = r + (0.99 * next_q_values)
```
## sync process, every 1-10k steps
```
dqn_params = q_network.state_dict()
target_network.load_state_dict(dqn_params)
```
## new training loop (single episode)
```
# hyper params
batch_size = 32
gamma = 0.99
learning_rate =1e-4
capacity=10000
sync_interval=1000 # new parameter t from paper
memory = ReplayMemory(capacity) # initialize replay memory
q_network = DQN(env.action_space.n).cuda() # initialize action-value function Q with random weights
target_network = DQN(env.action_space.n).cuda() # init target network
optimizer = Adam(q_network.parameters(), lr=learning_rate)
criterion = nn.SmoothL1Loss()
state = env.reset() # observe initial state
current_step = 0
while True:
env.render() # so we can watch!
action = select_action(q_network, state, env, calculate_epsilon(current_step)) # select action
next_state, reward, done, info = env.step(action) # carry out action/observe reward
# store experience s, a, r, s' in replay memory
memory.add(state, action, reward, next_state, done)
# sample random transitions
states, actions, rewards, next_states, done_mask = memory.sample(batch_size)
# prepare batch
states = Variable(states).cuda()
next_states = Variable(next_states).cuda()
rewards = Variable(rewards).cuda()
done_mask = Variable(done_mask).cuda()
# calculate target
# find next Q values and set Q values for done states to 0
### DOUBLE Q LEARNING
target_actions = q_network(next_states).max(dim=1)[1]
next_q_values = target_network(next_states)[range(len(target_actions)), target_actions].detach() * done_mask
### END DOUBLE Q LEARNING
# calculate targets = rewards + (gamma * next_Q_values)
targets = rewards + (gamma * next_q_values)
q_values = q_network(states)[range(len(actions)), actions] # select only Q values for actions we took
# train network
loss = criterion(q_values, targets) # smooth l1 loss
optimizer.zero_grad()
loss.backward()
# gradient clipping to prevent exploding gradient
for param in q_network.parameters():
param.grad.data.clamp_(-1, 1)
optimizer.step()
state = next_state # move to next state
current_step += 1
### DDQN sync
if current_step % sync_interval == 0:
dqn_params = q_network.state_dict()
target_network.load_state_dict(dqn_params)
if done:
break
env.close()
```
| github_jupyter |
```
# def warper(img, src, dst):
# # Compute and apply perpective transform
# img_size = (img.shape[1], img.shape[0])
# M = cv2.getPerspectiveTransform(src, dst)
# warped = cv2.warpPerspective(img, M, img_size, flags=cv2.INTER_NEAREST) # keep same size as input image
# return warped
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import cv2
# %matplotlib qt
%matplotlib inline
fileName = '../test_images/test5.jpg'
image = mpimg.imread(fileName)
gray = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
R = image[:, :, 0]
G = image[:, :, 1]
B = image[:, :, 2]
# HLS color
hls = cv2.cvtColor(image, cv2.COLOR_RGB2HLS)
H = hls[:,:,0]
L = hls[:,:,1]
S = hls[:,:,2]
thresh = (218, 255)
binary = np.zeros_like(S)
binary[(R > thresh[0]) & (R <= thresh[1])] = 1
a2 = binary
# Define a function that applies Sobel x and y,
# then computes the direction of the gradient
# and applies a threshold.
def dir_threshold(img, sobel_kernel=3, thresh=(0, np.pi/2)):
# Apply the following steps to img
# 1) Convert to grayscale
# gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
gray = img
# 2) Take the gradient in x and y separately
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# 3) Take the absolute value of the x and y gradients
abs_sobelx = np.sqrt(np.square(sobelx))
abs_sobely = np.sqrt(np.square(sobely))
abs_sobelxy = np.sqrt(np.square(sobelx)+np.square(sobely))
# 4) Use np.arctan2(abs_sobely, abs_sobelx) to calculate the direction of the gradient
theta = np.arctan2(abs_sobely, abs_sobelx)
# normalize
# theta = np.uint8(theta/(np.max(theta)/255))
scaled_theta = np.uint8(theta/np.max(theta/255))
# 5) Create a binary mask where direction thresholds are met
# binary_output = np.zeros_like(theta)
binary_output = np.zeros_like(scaled_theta)
binary_output[(theta >= thresh[0]) & (theta <= thresh[1])] = 1
# 6) Return this mask as your binary_output image
# binary_output = np.copy(img) # Remove this line
return binary_output
def abs_sobel_thresh(gray, orient='x', sobel_kernel=3, thresh=(0, 255)):
# gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# gray =img
# Calculate directional gradient
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# 3) Take the absolute value of the x and y gradients
abs_sobelx = np.sqrt(np.square(sobelx))
abs_sobely = np.sqrt(np.square(sobely))
abs_sobelxy = np.sqrt(np.square(sobelx)+np.square(sobely))
scaled_sobel = np.uint8(255*abs_sobelx/np.max(abs_sobelx))
# Apply threshold
sxbinary = np.zeros_like(scaled_sobel)
sxbinary[(scaled_sobel >= thresh[0]) & (scaled_sobel <= thresh[1])] = 1
return sxbinary
# Run the function
sx_L = abs_sobel_thresh(L, orient='x', sobel_kernel=3, thresh=(20, 100))
a1 = sx_L
# # dir_binary = dir_threshold(binary, sobel_kernel=15, thresh=(0.7, 1.3))
# # Plot the result
# f, (ax1, ax2) = plt.subplots(1, 2, figsize=(24, 9))
# f.tight_layout()
# ax1.imshow(L, cmap='gray')
# ax1.set_title('Original Image', fontsize=50)
# ax2.imshow(sx_L, cmap='gray')
# ax2.set_title('Thresholded Grad. Dir.', fontsize=50)
# plt.subplots_adjust(left=0., right=1, top=0.9, bottom=0.)
def abs_sobel_thresh(img, orient='x', sobel_kernel=3, thresh=(0, 255)):
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# gray =img
# Calculate directional gradient
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# 3) Take the absolute value of the x and y gradients
abs_sobelx = np.sqrt(np.square(sobelx))
abs_sobely = np.sqrt(np.square(sobely))
abs_sobelxy = np.sqrt(np.square(sobelx)+np.square(sobely))
if orient == 'x':
scaled_sobel = np.uint8(255*abs_sobelx/np.max(abs_sobelx))
else:
scaled_sobel = np.uint8(255*abs_sobely/np.max(abs_sobely))
# Apply threshold
grad_binary = np.zeros_like(scaled_sobel)
grad_binary[(scaled_sobel >= thresh[0]) & (scaled_sobel <= thresh[1])] = 1
return grad_binary
# Define a function to return the magnitude of the gradient
# for a given sobel kernel size and threshold values
def mag_thresh(img, sobel_kernel=3, mag_thresh=(0, 255)):
# Calculate gradient magnitude
# Apply threshold
# Convert to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# Take both Sobel x and y gradients
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# Calculate the gradient magnitude
gradmag = np.sqrt(sobelx**2 + sobely**2)
# Rescale to 8 bit
scale_factor = np.max(gradmag)/255
gradmag = (gradmag/scale_factor).astype(np.uint8)
# Create a binary image of ones where threshold is met, zeros otherwise
mag_binary = np.zeros_like(gradmag)
mag_binary[(gradmag >= mag_thresh[0]) & (gradmag <= mag_thresh[1])] = 1
# Return the binary image
return mag_binary
def dir_threshold(img, sobel_kernel=3, thresh=(0, np.pi/2)):
# Calculate gradient direction
# Apply threshold
# Apply the following steps to img
# 1) Convert to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# gray = img
# 2) Take the gradient in x and y separately
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# 3) Take the absolute value of the x and y gradients
abs_sobelx = np.sqrt(np.square(sobelx))
abs_sobely = np.sqrt(np.square(sobely))
abs_sobelxy = np.sqrt(np.square(sobelx)+np.square(sobely))
# 4) Use np.arctan2(abs_sobely, abs_sobelx) to calculate the direction of the gradient
theta = np.arctan2(abs_sobely, abs_sobelx)
# normalize
# theta = np.uint8(theta/(np.max(theta)/255))
scaled_theta = np.uint8(theta/np.max(theta/255))
# 5) Create a binary mask where direction thresholds are met
# binary_output = np.zeros_like(theta)
dir_binary = np.zeros_like(scaled_theta)
dir_binary[(theta >= thresh[0]) & (theta <= thresh[1])] = 1
return dir_binary
# Choose a Sobel kernel size
ksize = 3 # Choose a larger odd number to smooth gradient measurements
# Apply each of the thresholding functions
gradx = abs_sobel_thresh(image, orient='x', sobel_kernel=ksize, thresh=(14, 130))
grady = abs_sobel_thresh(image, orient='y', sobel_kernel=ksize, thresh=(44, 115))
mag_binary = mag_thresh(image, sobel_kernel=ksize, mag_thresh=(50, 255))
dir_binary = dir_threshold(image, sobel_kernel=ksize, thresh=(np.pi/5, np.pi/3))
combined = np.zeros_like(dir_binary)
combined[((gradx == 1) & (grady == 1)) | ((mag_binary == 1) & (dir_binary == 1))] = 1
best_binary2 = combined
# # Plotting thresholded images
# f, (ax1, ax2) = plt.subplots(1, 2, figsize=(20,10))
# ax1.set_title('sobel mag dir')
# ax1.imshow(mag_binary, cmap ='gray')
# ax2.set_title('sobel mag dir')
# ax2.imshow(combined, cmap='gray')
# %matplotlib qt
# Stack each channel to view their individual contributions in green and blue respectively
# This returns a stack of the two binary images, whose components you can see as different colors
color_binary = np.dstack(( np.zeros_like(a1), a1, a2)) * 255
# Combine the two binary thresholds
combined_binary = np.zeros_like(a1)
combined_binary[(a2 == 1) | (a1 == 1)] = 1
best_binary1 = combined_binary
# Stack each channel to view their individual contributions in green and blue respectively
# This returns a stack of the two binary images, whose components you can see as different colors
color_binary = np.dstack(( np.zeros_like(best_binary1), best_binary1, best_binary2)) * 255
# Combine the two binary thresholds
combined_binary = np.zeros_like(best_binary1)
combined_binary[(best_binary2 == 1) | (best_binary1 == 1)] = 1
# best_binary1 = combined_binary
best_final = combined_binary
img_size = (image.shape[1], image.shape[0])
src = np.float32(
[[(img_size[0] / 2) - 55, img_size[1] / 2 + 100],
[((img_size[0] / 6) - 10), img_size[1]],
[(img_size[0] * 5 / 6) + 60, img_size[1]],
[(img_size[0] / 2 + 55), img_size[1] / 2 + 100]])
dst = np.float32(
[[(img_size[0] / 4), 0],
[(img_size[0] / 4), img_size[1]],
[(img_size[0] * 3 / 4), img_size[1]],
[(img_size[0] * 3 / 4), 0]])
M = cv2.getPerspectiveTransform(src, dst)
warped = cv2.warpPerspective(image, M, img_size, flags=cv2.INTER_LINEAR)
# trythis = warped.copy()
trythis2 = image.copy()
# img_size = (image.shape[1], image.shape[0])
# src = np.float32(
# [[(img_size[0] / 2) - 55, img_size[1] / 2 + 100],
# [((img_size[0] / 6) - 10), img_size[1]],
# [(img_size[0] * 5 / 6) + 60, img_size[1]],
# [(img_size[0] / 2 + 55), img_size[1] / 2 + 100]])
# dst = np.float32(
# [[(img_size[0] / 4), 0],
# [(img_size[0] / 4), img_size[1]],
# [(img_size[0] * 3 / 4), img_size[1]],
# [(img_size[0] * 3 / 4), 0]])
# M = cv2.getPerspectiveTransform(src, dst)
warped = cv2.warpPerspective(best_final, M, img_size, flags=cv2.INTER_LINEAR)
def hist(image):
bottom_half = img[img.shape[0]//2:,:]
histogram = np.sum(bottom_half, axis=0)
return histogram
img = warped/255
histogram = hist(img)
# plt.plot(histogram)
# Iterate through nwindows to track culvature
# 1. Loop through each windows in nwindows
# 2. Find the boundaries of our current windows. (leftx_current and rightx_current) as well as the margin you set in the hyperparameters
# 3. Use cv2.rectangle to draw these windows boundares onto our visualization image out_img.
# 4. Now that we know the boundaries of our window. find out which activated pixels from nonzeroy, and nonzerox above actually fall into the window.
# 5. Append these to our lists left_lane_inds and right_lanes_inds.
# 6. If the number of pixels you found in step 4 are geater than you hyperparameter minpix, re-center our window(i.e. leftx_current or rightx_current) based on the mean position of these pixels
# Load our image
binary_warped = warped.copy()
def find_lane_pixels(binary_warped):
# Take a histogram of the bottom half of the image
histogram = np.sum(binary_warped[binary_warped.shape[0]//2:,:], axis=0)
# Create an output image to draw on and visualize the result
out_img = np.dstack((binary_warped, binary_warped, binary_warped))
# Find the peak of the left and right halves of the histogram
# These will be the starting point for the left and right lines
midpoint = np.int(histogram.shape[0]//2)
leftx_base = np.argmax(histogram[:midpoint])
rightx_base = np.argmax(histogram[midpoint:]) + midpoint
# HYPERPARAMETERS
# Choose the number of sliding windows
nwindows = 9
# Set the width of the windows +/- margin
margin = 100
# Set minimum number of pixels found to recenter window
minpix = 50
# Set height of windows - based on nwindows above and image shape
window_height = np.int(binary_warped.shape[0]//nwindows)
# Identify the x and y positions of all nonzero pixels in the image
nonzero = binary_warped.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
# Current positions to be updated later for each window in nwindows
leftx_current = leftx_base
rightx_current = rightx_base
# Create empty lists to receive left and right lane pixel indices
left_lane_inds = []
right_lane_inds = []
# Step through the windows one by one
for window in range(nwindows):
# Identify window boundaries in x and y (and right and left)
win_y_low = binary_warped.shape[0] - (window+1)*window_height
win_y_high = binary_warped.shape[0] - window*window_height
win_xleft_low = leftx_current - margin
win_xleft_high = leftx_current + margin
win_xright_low = rightx_current - margin
win_xright_high = rightx_current + margin
# Draw the windows on the visualization image
cv2.rectangle(out_img,(win_xleft_low,win_y_low),
(win_xleft_high,win_y_high),(0,255,0), 2)
cv2.rectangle(out_img,(win_xright_low,win_y_low),
(win_xright_high,win_y_high),(0,255,0), 2)
# Identify the nonzero pixels in x and y within the window #
good_left_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_xleft_low) & (nonzerox < win_xleft_high)).nonzero()[0]
good_right_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_xright_low) & (nonzerox < win_xright_high)).nonzero()[0]
# Append these indices to the lists
left_lane_inds.append(good_left_inds)
right_lane_inds.append(good_right_inds)
# If you found > minpix pixels, recenter next window on their mean position
if len(good_left_inds) > minpix:
leftx_current = np.int(np.mean(nonzerox[good_left_inds]))
if len(good_right_inds) > minpix:
rightx_current = np.int(np.mean(nonzerox[good_right_inds]))
# Concatenate the arrays of indices (previously was a list of lists of pixels)
try:
left_lane_inds = np.concatenate(left_lane_inds)
right_lane_inds = np.concatenate(right_lane_inds)
except ValueError:
# Avoids an error if the above is not implemented fully
pass
# Extract left and right line pixel positions
leftx = nonzerox[left_lane_inds]
lefty = nonzeroy[left_lane_inds]
rightx = nonzerox[right_lane_inds]
righty = nonzeroy[right_lane_inds]
return leftx, lefty, rightx, righty, out_img
def fit_polynomial(binary_warped):
# Find our lane pixels first
leftx, lefty, rightx, righty, out_img = find_lane_pixels(binary_warped)
# Fit a second order polynomial to each using `np.polyfit`
left_fit = np.polyfit(lefty, leftx, 2)
right_fit = np.polyfit(righty, rightx, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, binary_warped.shape[0]-1, binary_warped.shape[0] )
try:
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
except TypeError:
# Avoids an error if `left` and `right_fit` are still none or incorrect
print('The function failed to fit a line!')
left_fitx = 1*ploty**2 + 1*ploty
right_fitx = 1*ploty**2 + 1*ploty
## Visualization ##
# Colors in the left and right lane regions
out_img[lefty, leftx] = [255, 0, 0]
out_img[righty, rightx] = [0, 0, 255]
#
# Plots the left and right polynomials on the lane lines
# plt.plot(left_fitx, ploty, color='yellow')
# plt.plot(right_fitx, ploty, color='yellow')
return out_img, ploty, left_fit, right_fit
out_img, ploty,left_fit, right_fit = fit_polynomial(binary_warped)
def fit_poly(img_shape, leftx, lefty, rightx, righty):
### TO-DO: Fit a second order polynomial to each with np.polyfit() ###
left_fit = np.polyfit(lefty, leftx, 2)
right_fit = np.polyfit(righty, rightx, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, img_shape[0]-1, img_shape[0])
### TO-DO: Calc both polynomials using ploty, left_fit and right_fit ###
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
return left_fitx, right_fitx, ploty
def search_around_poly(binary_warped):
# HYPERPARAMETER
# Choose the width of the margin around the previous polynomial to search
# The quiz grader expects 100 here, but feel free to tune on your own!
margin = 100
# Grab activated pixels
nonzero = binary_warped.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
### TO-DO: Set the area of search based on activated x-values ###
### within the +/- margin of our polynomial function ###
### Hint: consider the window areas for the similarly named variables ###
### in the previous quiz, but change the windows to our new search area ###
left_lane_inds = ((nonzerox > (left_fit[0]*(nonzeroy**2) + left_fit[1]*nonzeroy +
left_fit[2] - margin)) & (nonzerox < (left_fit[0]*(nonzeroy**2) +
left_fit[1]*nonzeroy + left_fit[2] + margin)))
right_lane_inds = ((nonzerox > (right_fit[0]*(nonzeroy**2) + right_fit[1]*nonzeroy +
right_fit[2] - margin)) & (nonzerox < (right_fit[0]*(nonzeroy**2) +
right_fit[1]*nonzeroy + right_fit[2] + margin)))
# Again, extract left and right line pixel positions
leftx = nonzerox[left_lane_inds]
lefty = nonzeroy[left_lane_inds]
rightx = nonzerox[right_lane_inds]
righty = nonzeroy[right_lane_inds]
# Fit new polynomials
left_fitx, right_fitx, ploty = fit_poly(binary_warped.shape, leftx, lefty, rightx, righty)
## Visualization ##
# Create an image to draw on and an image to show the selection window
out_img = np.dstack((binary_warped, binary_warped, binary_warped))*255
window_img = np.zeros_like(out_img)
# Color in left and right line pixels
out_img[nonzeroy[left_lane_inds], nonzerox[left_lane_inds]] = [255, 0, 0]
out_img[nonzeroy[right_lane_inds], nonzerox[right_lane_inds]] = [0, 0, 255]
# Generate a polygon to illustrate the search window area
# And recast the x and y points into usable format for cv2.fillPoly()
left_line_window1 = np.array([np.transpose(np.vstack([left_fitx-margin, ploty]))])
left_line_window2 = np.array([np.flipud(np.transpose(np.vstack([left_fitx+margin,
ploty])))])
left_line_pts = np.hstack((left_line_window1, left_line_window2))
right_line_window1 = np.array([np.transpose(np.vstack([right_fitx-margin, ploty]))])
right_line_window2 = np.array([np.flipud(np.transpose(np.vstack([right_fitx+margin,
ploty])))])
right_line_pts = np.hstack((right_line_window1, right_line_window2))
# Draw the lane onto the warped blank image
cv2.fillPoly(window_img, np.int_([left_line_pts]), (0,255, 0))
cv2.fillPoly(window_img, np.int_([right_line_pts]), (0,255, 0))
result = cv2.addWeighted(out_img, 1, window_img, 0.3, 0)
# Plot the polynomial lines onto the image
# plt.plot(left_fitx, ploty, color='yellow')
# plt.plot(right_fitx, ploty, color='yellow')
## End visualization steps ##
return result, left_fitx, right_fitx, ploty, out_img
# Run image through the pipeline
# Note that in your project, you'll also want to feed in the previous fits
result, left_fitx, right_fitx, ploty, out_img = search_around_poly(binary_warped)
# %matplotlib qt
warped = binary_warped
image = binary_warped
# print(type(warped))
# Create an image to draw the lines on
warp_zero = np.zeros_like(warped).astype(np.uint8)
color_warp = np.dstack((warp_zero, warp_zero, warp_zero))
# Recast the x and y points into usable format for cv2.fillPoly()
pts_left = np.array([np.transpose(np.vstack([left_fitx, ploty]))])
pts_right = np.array([np.flipud(np.transpose(np.vstack([right_fitx, ploty])))])
pts = np.hstack((pts_left, pts_right))
# Draw the lane onto the warped blank image
cv2.fillPoly(color_warp, np.int_([pts]), (0,255, 0))
img_size = (warped.shape[1], warped.shape[0])
src = np.float32(
[[(img_size[0] / 2) - 55, img_size[1] / 2 + 100],
[((img_size[0] / 6) - 10), img_size[1]],
[(img_size[0] * 5 / 6) + 60, img_size[1]],
[(img_size[0] / 2 + 55), img_size[1] / 2 + 100]])
dst = np.float32(
[[(img_size[0] / 4), 0],
[(img_size[0] / 4), img_size[1]],
[(img_size[0] * 3 / 4), img_size[1]],
[(img_size[0] * 3 / 4), 0]])
Minv = cv2.getPerspectiveTransform(dst, src)
# Warp the blank back to original image space using inverse perspective matrix (Minv)
newwarp = cv2.warpPerspective(color_warp, Minv, (image.shape[1], image.shape[0]))
# Warp the blank back to original image space using inverse perspective matrix (Minv)
# result = cv2.warpPerspective(trythis2, Minv, (image.shape[1], image.shape[0]))
# Combine the result with the original image
result = cv2.addWeighted(trythis2, 1, newwarp, 0.3, 0)
plt.imshow(result)
# plt.imshow(result)
# Import everything needed to edit/save/watch video clips
from moviepy.editor import VideoFileClip
from IPython.display import HTML
# def warper(img, src, dst):
# # Compute and apply perpective transform
# img_size = (img.shape[1], img.shape[0])
# M = cv2.getPerspectiveTransform(src, dst)
# warped = cv2.warpPerspective(img, M, img_size, flags=cv2.INTER_NEAREST) # keep same size as input image
# return warped
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import cv2
# %matplotlib qt
%matplotlib inline
def process_image(image):
# fileName = '../test_images/test5.jpg'
# image = mpimg.imread(fileName)
gray = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
R = image[:, :, 0]
G = image[:, :, 1]
B = image[:, :, 2]
# HLS color
hls = cv2.cvtColor(image, cv2.COLOR_RGB2HLS)
H = hls[:,:,0]
L = hls[:,:,1]
S = hls[:,:,2]
thresh = (218, 255)
binary = np.zeros_like(S)
binary[(R > thresh[0]) & (R <= thresh[1])] = 1
a2 = binary
# Define a function that applies Sobel x and y,
# then computes the direction of the gradient
# and applies a threshold.
def dir_threshold(img, sobel_kernel=3, thresh=(0, np.pi/2)):
# Apply the following steps to img
# 1) Convert to grayscale
# gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
gray = img
# 2) Take the gradient in x and y separately
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# 3) Take the absolute value of the x and y gradients
abs_sobelx = np.sqrt(np.square(sobelx))
abs_sobely = np.sqrt(np.square(sobely))
abs_sobelxy = np.sqrt(np.square(sobelx)+np.square(sobely))
# 4) Use np.arctan2(abs_sobely, abs_sobelx) to calculate the direction of the gradient
theta = np.arctan2(abs_sobely, abs_sobelx)
# normalize
# theta = np.uint8(theta/(np.max(theta)/255))
scaled_theta = np.uint8(theta/np.max(theta/255))
# 5) Create a binary mask where direction thresholds are met
# binary_output = np.zeros_like(theta)
binary_output = np.zeros_like(scaled_theta)
binary_output[(theta >= thresh[0]) & (theta <= thresh[1])] = 1
# 6) Return this mask as your binary_output image
# binary_output = np.copy(img) # Remove this line
return binary_output
def abs_sobel_thresh(gray, orient='x', sobel_kernel=3, thresh=(0, 255)):
# gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# gray =img
# Calculate directional gradient
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# 3) Take the absolute value of the x and y gradients
abs_sobelx = np.sqrt(np.square(sobelx))
abs_sobely = np.sqrt(np.square(sobely))
abs_sobelxy = np.sqrt(np.square(sobelx)+np.square(sobely))
scaled_sobel = np.uint8(255*abs_sobelx/np.max(abs_sobelx))
# Apply threshold
sxbinary = np.zeros_like(scaled_sobel)
sxbinary[(scaled_sobel >= thresh[0]) & (scaled_sobel <= thresh[1])] = 1
return sxbinary
# Run the function
sx_L = abs_sobel_thresh(L, orient='x', sobel_kernel=3, thresh=(20, 100))
a1 = sx_L
# # dir_binary = dir_threshold(binary, sobel_kernel=15, thresh=(0.7, 1.3))
# # Plot the result
# f, (ax1, ax2) = plt.subplots(1, 2, figsize=(24, 9))
# f.tight_layout()
# ax1.imshow(L, cmap='gray')
# ax1.set_title('Original Image', fontsize=50)
# ax2.imshow(sx_L, cmap='gray')
# ax2.set_title('Thresholded Grad. Dir.', fontsize=50)
# plt.subplots_adjust(left=0., right=1, top=0.9, bottom=0.)
def abs_sobel_thresh(img, orient='x', sobel_kernel=3, thresh=(0, 255)):
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# gray =img
# Calculate directional gradient
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# 3) Take the absolute value of the x and y gradients
abs_sobelx = np.sqrt(np.square(sobelx))
abs_sobely = np.sqrt(np.square(sobely))
abs_sobelxy = np.sqrt(np.square(sobelx)+np.square(sobely))
if orient == 'x':
scaled_sobel = np.uint8(255*abs_sobelx/np.max(abs_sobelx))
else:
scaled_sobel = np.uint8(255*abs_sobely/np.max(abs_sobely))
# Apply threshold
grad_binary = np.zeros_like(scaled_sobel)
grad_binary[(scaled_sobel >= thresh[0]) & (scaled_sobel <= thresh[1])] = 1
return grad_binary
# Define a function to return the magnitude of the gradient
# for a given sobel kernel size and threshold values
def mag_thresh(img, sobel_kernel=3, mag_thresh=(0, 255)):
# Calculate gradient magnitude
# Apply threshold
# Convert to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# Take both Sobel x and y gradients
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# Calculate the gradient magnitude
gradmag = np.sqrt(sobelx**2 + sobely**2)
# Rescale to 8 bit
scale_factor = np.max(gradmag)/255
gradmag = (gradmag/scale_factor).astype(np.uint8)
# Create a binary image of ones where threshold is met, zeros otherwise
mag_binary = np.zeros_like(gradmag)
mag_binary[(gradmag >= mag_thresh[0]) & (gradmag <= mag_thresh[1])] = 1
# Return the binary image
return mag_binary
def dir_threshold(img, sobel_kernel=3, thresh=(0, np.pi/2)):
# Calculate gradient direction
# Apply threshold
# Apply the following steps to img
# 1) Convert to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# gray = img
# 2) Take the gradient in x and y separately
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# 3) Take the absolute value of the x and y gradients
abs_sobelx = np.sqrt(np.square(sobelx))
abs_sobely = np.sqrt(np.square(sobely))
abs_sobelxy = np.sqrt(np.square(sobelx)+np.square(sobely))
# 4) Use np.arctan2(abs_sobely, abs_sobelx) to calculate the direction of the gradient
theta = np.arctan2(abs_sobely, abs_sobelx)
# normalize
# theta = np.uint8(theta/(np.max(theta)/255))
scaled_theta = np.uint8(theta/np.max(theta/255))
# 5) Create a binary mask where direction thresholds are met
# binary_output = np.zeros_like(theta)
dir_binary = np.zeros_like(scaled_theta)
dir_binary[(theta >= thresh[0]) & (theta <= thresh[1])] = 1
return dir_binary
# Choose a Sobel kernel size
ksize = 3 # Choose a larger odd number to smooth gradient measurements
# Apply each of the thresholding functions
gradx = abs_sobel_thresh(image, orient='x', sobel_kernel=ksize, thresh=(14, 130))
grady = abs_sobel_thresh(image, orient='y', sobel_kernel=ksize, thresh=(44, 115))
mag_binary = mag_thresh(image, sobel_kernel=ksize, mag_thresh=(50, 255))
dir_binary = dir_threshold(image, sobel_kernel=ksize, thresh=(np.pi/5, np.pi/3))
combined = np.zeros_like(dir_binary)
combined[((gradx == 1) & (grady == 1)) | ((mag_binary == 1) & (dir_binary == 1))] = 1
best_binary2 = combined
# # Plotting thresholded images
# f, (ax1, ax2) = plt.subplots(1, 2, figsize=(20,10))
# ax1.set_title('sobel mag dir')
# ax1.imshow(mag_binary, cmap ='gray')
# ax2.set_title('sobel mag dir')
# ax2.imshow(combined, cmap='gray')
# %matplotlib qt
# Stack each channel to view their individual contributions in green and blue respectively
# This returns a stack of the two binary images, whose components you can see as different colors
color_binary = np.dstack(( np.zeros_like(a1), a1, a2)) * 255
# Combine the two binary thresholds
combined_binary = np.zeros_like(a1)
combined_binary[(a2 == 1) | (a1 == 1)] = 1
best_binary1 = combined_binary
# Stack each channel to view their individual contributions in green and blue respectively
# This returns a stack of the two binary images, whose components you can see as different colors
color_binary = np.dstack(( np.zeros_like(best_binary1), best_binary1, best_binary2)) * 255
# Combine the two binary thresholds
combined_binary = np.zeros_like(best_binary1)
combined_binary[(best_binary2 == 1) | (best_binary1 == 1)] = 1
# best_binary1 = combined_binary
best_final = combined_binary
img_size = (image.shape[1], image.shape[0])
src = np.float32(
[[(img_size[0] / 2) - 55, img_size[1] / 2 + 100],
[((img_size[0] / 6) - 10), img_size[1]],
[(img_size[0] * 5 / 6) + 60, img_size[1]],
[(img_size[0] / 2 + 55), img_size[1] / 2 + 100]])
dst = np.float32(
[[(img_size[0] / 4), 0],
[(img_size[0] / 4), img_size[1]],
[(img_size[0] * 3 / 4), img_size[1]],
[(img_size[0] * 3 / 4), 0]])
M = cv2.getPerspectiveTransform(src, dst)
warped2 = cv2.warpPerspective(image, M, img_size, flags=cv2.INTER_LINEAR)
# trythis = warped.copy()
trythis2 = image
# img_size = (image.shape[1], image.shape[0])
# src = np.float32(
# [[(img_size[0] / 2) - 55, img_size[1] / 2 + 100],
# [((img_size[0] / 6) - 10), img_size[1]],
# [(img_size[0] * 5 / 6) + 60, img_size[1]],
# [(img_size[0] / 2 + 55), img_size[1] / 2 + 100]])
# dst = np.float32(
# [[(img_size[0] / 4), 0],
# [(img_size[0] / 4), img_size[1]],
# [(img_size[0] * 3 / 4), img_size[1]],
# [(img_size[0] * 3 / 4), 0]])
# M = cv2.getPerspectiveTransform(src, dst)
warped = cv2.warpPerspective(best_final, M, img_size, flags=cv2.INTER_LINEAR)
def hist(image):
bottom_half = img[img.shape[0]//2:,:]
histogram = np.sum(bottom_half, axis=0)
return histogram
img = warped/255
histogram = hist(img)
# plt.plot(histogram)
# Iterate through nwindows to track culvature
# 1. Loop through each windows in nwindows
# 2. Find the boundaries of our current windows. (leftx_current and rightx_current) as well as the margin you set in the hyperparameters
# 3. Use cv2.rectangle to draw these windows boundares onto our visualization image out_img.
# 4. Now that we know the boundaries of our window. find out which activated pixels from nonzeroy, and nonzerox above actually fall into the window.
# 5. Append these to our lists left_lane_inds and right_lanes_inds.
# 6. If the number of pixels you found in step 4 are geater than you hyperparameter minpix, re-center our window(i.e. leftx_current or rightx_current) based on the mean position of these pixels
# Load our image
binary_warped = warped.copy()
def find_lane_pixels(binary_warped):
# Take a histogram of the bottom half of the image
histogram = np.sum(binary_warped[binary_warped.shape[0]//2:,:], axis=0)
# Create an output image to draw on and visualize the result
out_img = np.dstack((binary_warped, binary_warped, binary_warped))
# Find the peak of the left and right halves of the histogram
# These will be the starting point for the left and right lines
midpoint = np.int(histogram.shape[0]//2)
leftx_base = np.argmax(histogram[:midpoint])
rightx_base = np.argmax(histogram[midpoint:]) + midpoint
# HYPERPARAMETERS
# Choose the number of sliding windows
nwindows = 9
# Set the width of the windows +/- margin
margin = 100
# Set minimum number of pixels found to recenter window
minpix = 50
# Set height of windows - based on nwindows above and image shape
window_height = np.int(binary_warped.shape[0]//nwindows)
# Identify the x and y positions of all nonzero pixels in the image
nonzero = binary_warped.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
# Current positions to be updated later for each window in nwindows
leftx_current = leftx_base
rightx_current = rightx_base
# Create empty lists to receive left and right lane pixel indices
left_lane_inds = []
right_lane_inds = []
# Step through the windows one by one
for window in range(nwindows):
# Identify window boundaries in x and y (and right and left)
win_y_low = binary_warped.shape[0] - (window+1)*window_height
win_y_high = binary_warped.shape[0] - window*window_height
win_xleft_low = leftx_current - margin
win_xleft_high = leftx_current + margin
win_xright_low = rightx_current - margin
win_xright_high = rightx_current + margin
# Draw the windows on the visualization image
cv2.rectangle(out_img,(win_xleft_low,win_y_low),
(win_xleft_high,win_y_high),(0,255,0), 2)
cv2.rectangle(out_img,(win_xright_low,win_y_low),
(win_xright_high,win_y_high),(0,255,0), 2)
# Identify the nonzero pixels in x and y within the window #
good_left_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_xleft_low) & (nonzerox < win_xleft_high)).nonzero()[0]
good_right_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_xright_low) & (nonzerox < win_xright_high)).nonzero()[0]
# Append these indices to the lists
left_lane_inds.append(good_left_inds)
right_lane_inds.append(good_right_inds)
# If you found > minpix pixels, recenter next window on their mean position
if len(good_left_inds) > minpix:
leftx_current = np.int(np.mean(nonzerox[good_left_inds]))
if len(good_right_inds) > minpix:
rightx_current = np.int(np.mean(nonzerox[good_right_inds]))
# Concatenate the arrays of indices (previously was a list of lists of pixels)
try:
left_lane_inds = np.concatenate(left_lane_inds)
right_lane_inds = np.concatenate(right_lane_inds)
except ValueError:
# Avoids an error if the above is not implemented fully
pass
# Extract left and right line pixel positions
leftx = nonzerox[left_lane_inds]
lefty = nonzeroy[left_lane_inds]
rightx = nonzerox[right_lane_inds]
righty = nonzeroy[right_lane_inds]
return leftx, lefty, rightx, righty, out_img
def fit_polynomial(binary_warped):
# Find our lane pixels first
leftx, lefty, rightx, righty, out_img = find_lane_pixels(binary_warped)
# Fit a second order polynomial to each using `np.polyfit`
left_fit = np.polyfit(lefty, leftx, 2)
right_fit = np.polyfit(righty, rightx, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, binary_warped.shape[0]-1, binary_warped.shape[0] )
try:
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
except TypeError:
# Avoids an error if `left` and `right_fit` are still none or incorrect
print('The function failed to fit a line!')
left_fitx = 1*ploty**2 + 1*ploty
right_fitx = 1*ploty**2 + 1*ploty
## Visualization ##
# Colors in the left and right lane regions
out_img[lefty, leftx] = [255, 0, 0]
out_img[righty, rightx] = [0, 0, 255]
#
# Plots the left and right polynomials on the lane lines
# plt.plot(left_fitx, ploty, color='yellow')
# plt.plot(right_fitx, ploty, color='yellow')
return out_img, ploty, left_fit, right_fit
out_img, ploty,left_fit, right_fit = fit_polynomial(binary_warped)
def fit_poly(img_shape, leftx, lefty, rightx, righty):
### TO-DO: Fit a second order polynomial to each with np.polyfit() ###
left_fit = np.polyfit(lefty, leftx, 2)
right_fit = np.polyfit(righty, rightx, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, img_shape[0]-1, img_shape[0])
### TO-DO: Calc both polynomials using ploty, left_fit and right_fit ###
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
return left_fitx, right_fitx, ploty
def search_around_poly(binary_warped):
# HYPERPARAMETER
# Choose the width of the margin around the previous polynomial to search
# The quiz grader expects 100 here, but feel free to tune on your own!
margin = 100
# Grab activated pixels
nonzero = binary_warped.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
### TO-DO: Set the area of search based on activated x-values ###
### within the +/- margin of our polynomial function ###
### Hint: consider the window areas for the similarly named variables ###
### in the previous quiz, but change the windows to our new search area ###
left_lane_inds = ((nonzerox > (left_fit[0]*(nonzeroy**2) + left_fit[1]*nonzeroy +
left_fit[2] - margin)) & (nonzerox < (left_fit[0]*(nonzeroy**2) +
left_fit[1]*nonzeroy + left_fit[2] + margin)))
right_lane_inds = ((nonzerox > (right_fit[0]*(nonzeroy**2) + right_fit[1]*nonzeroy +
right_fit[2] - margin)) & (nonzerox < (right_fit[0]*(nonzeroy**2) +
right_fit[1]*nonzeroy + right_fit[2] + margin)))
# Again, extract left and right line pixel positions
leftx = nonzerox[left_lane_inds]
lefty = nonzeroy[left_lane_inds]
rightx = nonzerox[right_lane_inds]
righty = nonzeroy[right_lane_inds]
# Fit new polynomials
left_fitx, right_fitx, ploty = fit_poly(binary_warped.shape, leftx, lefty, rightx, righty)
## Visualization ##
# Create an image to draw on and an image to show the selection window
out_img = np.dstack((binary_warped, binary_warped, binary_warped))*255
window_img = np.zeros_like(out_img)
# Color in left and right line pixels
out_img[nonzeroy[left_lane_inds], nonzerox[left_lane_inds]] = [255, 0, 0]
out_img[nonzeroy[right_lane_inds], nonzerox[right_lane_inds]] = [0, 0, 255]
# Generate a polygon to illustrate the search window area
# And recast the x and y points into usable format for cv2.fillPoly()
left_line_window1 = np.array([np.transpose(np.vstack([left_fitx-margin, ploty]))])
left_line_window2 = np.array([np.flipud(np.transpose(np.vstack([left_fitx+margin,
ploty])))])
left_line_pts = np.hstack((left_line_window1, left_line_window2))
right_line_window1 = np.array([np.transpose(np.vstack([right_fitx-margin, ploty]))])
right_line_window2 = np.array([np.flipud(np.transpose(np.vstack([right_fitx+margin,
ploty])))])
right_line_pts = np.hstack((right_line_window1, right_line_window2))
# Draw the lane onto the warped blank image
cv2.fillPoly(window_img, np.int_([left_line_pts]), (0,255, 0))
cv2.fillPoly(window_img, np.int_([right_line_pts]), (0,255, 0))
result = cv2.addWeighted(out_img, 1, window_img, 0.3, 0)
# Plot the polynomial lines onto the image
# plt.plot(left_fitx, ploty, color='yellow')
# plt.plot(right_fitx, ploty, color='yellow')
## End visualization steps ##
return result, left_fitx, right_fitx, ploty, out_img
# Run image through the pipeline
# Note that in your project, you'll also want to feed in the previous fits
result, left_fitx, right_fitx, ploty, out_img = search_around_poly(binary_warped)
# %matplotlib qt
warped = binary_warped
image = binary_warped
# print(type(warped))
# Create an image to draw the lines on
warp_zero = np.zeros_like(warped).astype(np.uint8)
color_warp = np.dstack((warp_zero, warp_zero, warp_zero))
# Recast the x and y points into usable format for cv2.fillPoly()
pts_left = np.array([np.transpose(np.vstack([left_fitx, ploty]))])
pts_right = np.array([np.flipud(np.transpose(np.vstack([right_fitx, ploty])))])
pts = np.hstack((pts_left, pts_right))
# Draw the lane onto the warped blank image
cv2.fillPoly(color_warp, np.int_([pts]), (0,255, 0))
img_size = (warped.shape[1], warped.shape[0])
src = np.float32(
[[(img_size[0] / 2) - 55, img_size[1] / 2 + 100],
[((img_size[0] / 6) - 10), img_size[1]],
[(img_size[0] * 5 / 6) + 60, img_size[1]],
[(img_size[0] / 2 + 55), img_size[1] / 2 + 100]])
dst = np.float32(
[[(img_size[0] / 4), 0],
[(img_size[0] / 4), img_size[1]],
[(img_size[0] * 3 / 4), img_size[1]],
[(img_size[0] * 3 / 4), 0]])
Minv = cv2.getPerspectiveTransform(dst, src)
# Warp the blank back to original image space using inverse perspective matrix (Minv)
newwarp = cv2.warpPerspective(color_warp, Minv, (image.shape[1], image.shape[0]))
# Warp the blank back to original image space using inverse perspective matrix (Minv)
# result = cv2.warpPerspective(trythis2, Minv, (image.shape[1], image.shape[0]))
# Combine the result with the original image
result = cv2.addWeighted(trythis2, 1, newwarp, 0.3, 0)
# plt.imshow(result)
return result
# process_image()
white_output = '../HC_video_output/project_video.mp4'
## To speed up the testing process you may want to try your pipeline on a shorter subclip of the video
## To do so add .subclip(start_second,end_second) to the end of the line below
## Where start_second and end_second are integer values representing the start and end of the subclip
## You may also uncomment the following line for a subclip of the first 5 seconds
clip1 = VideoFileClip("../project_video.mp4").subclip(0,5)
# clip1 = VideoFileClip("../project_video.mp4")
white_clip = clip1.fl_image(process_image_end) #NOTE: this function expects color images!!
%time white_clip.write_videofile(white_output, audio=False)
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(white_output))
# Import everything needed to edit/save/watch video clips
from moviepy.editor import VideoFileClip
from IPython.display import HTML
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import cv2
# %matplotlib qt
%matplotlib inline
def abs_sobel_thresh(img, orient='x', sobel_kernel=3, thresh=(0, 255)):
# gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
gray =img
# Calculate directional gradient
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# 3) Take the absolute value of the x and y gradients
abs_sobelx = np.sqrt(np.square(sobelx))
abs_sobely = np.sqrt(np.square(sobely))
abs_sobelxy = np.sqrt(np.square(sobelx)+np.square(sobely))
if orient == 'x':
scaled_sobel = np.uint8(255*abs_sobelx/np.max(abs_sobelx))
else:
scaled_sobel = np.uint8(255*abs_sobely/np.max(abs_sobely))
# Apply threshold
grad_binary = np.zeros_like(scaled_sobel)
grad_binary[(scaled_sobel >= thresh[0]) & (scaled_sobel <= thresh[1])] = 1
return grad_binary
def mag_thresh(img, sobel_kernel=3, mag_thresh=(0, 255)):
# Calculate gradient magnitude
# Apply threshold
# Convert to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# Take both Sobel x and y gradients
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# Calculate the gradient magnitude
gradmag = np.sqrt(sobelx**2 + sobely**2)
# Rescale to 8 bit
scale_factor = np.max(gradmag)/255
gradmag = (gradmag/scale_factor).astype(np.uint8)
# Create a binary image of ones where threshold is met, zeros otherwise
mag_binary = np.zeros_like(gradmag)
mag_binary[(gradmag >= mag_thresh[0]) & (gradmag <= mag_thresh[1])] = 1
# Return the binary image
return mag_binary
def dir_threshold(img, sobel_kernel=3, thresh=(0, np.pi/2)):
# Calculate gradient direction
# Apply threshold
# Apply the following steps to img
# 1) Convert to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# gray = img
# 2) Take the gradient in x and y separately
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# 3) Take the absolute value of the x and y gradients
abs_sobelx = np.sqrt(np.square(sobelx))
abs_sobely = np.sqrt(np.square(sobely))
abs_sobelxy = np.sqrt(np.square(sobelx)+np.square(sobely))
# 4) Use np.arctan2(abs_sobely, abs_sobelx) to calculate the direction of the gradient
theta = np.arctan2(abs_sobely, abs_sobelx)
# normalize
# theta = np.uint8(theta/(np.max(theta)/255))
scaled_theta = np.uint8(theta/np.max(theta/255))
# 5) Create a binary mask where direction thresholds are met
# binary_output = np.zeros_like(theta)
dir_binary = np.zeros_like(scaled_theta)
dir_binary[(theta >= thresh[0]) & (theta <= thresh[1])] = 1
return dir_binary
def hist(image):
bottom_half = image[image.shape[0]//2:,:]
histogram = np.sum(bottom_half, axis=0)
return histogram
def find_lane_pixels(binary_warped):
# Take a histogram of the bottom half of the image
histogram = np.sum(binary_warped[binary_warped.shape[0]//2:,:], axis=0)
# Create an output image to draw on and visualize the result
out_img = np.dstack((binary_warped, binary_warped, binary_warped))
# Find the peak of the left and right halves of the histogram
# These will be the starting point for the left and right lines
midpoint = np.int(histogram.shape[0]//2)
leftx_base = np.argmax(histogram[:midpoint])
rightx_base = np.argmax(histogram[midpoint:]) + midpoint
# HYPERPARAMETERS
# Choose the number of sliding windows
nwindows = 9
# Set the width of the windows +/- margin
margin = 100
# Set minimum number of pixels found to recenter window
minpix = 50
# Set height of windows - based on nwindows above and image shape
window_height = np.int(binary_warped.shape[0]//nwindows)
# Identify the x and y positions of all nonzero pixels in the image
nonzero = binary_warped.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
# Current positions to be updated later for each window in nwindows
leftx_current = leftx_base
rightx_current = rightx_base
# Create empty lists to receive left and right lane pixel indices
left_lane_inds = []
right_lane_inds = []
# Step through the windows one by one
for window in range(nwindows):
# Identify window boundaries in x and y (and right and left)
win_y_low = binary_warped.shape[0] - (window+1)*window_height
win_y_high = binary_warped.shape[0] - window*window_height
win_xleft_low = leftx_current - margin
win_xleft_high = leftx_current + margin
win_xright_low = rightx_current - margin
win_xright_high = rightx_current + margin
# Draw the windows on the visualization image
cv2.rectangle(out_img,(win_xleft_low,win_y_low),
(win_xleft_high,win_y_high),(0,255,0), 2)
cv2.rectangle(out_img,(win_xright_low,win_y_low),
(win_xright_high,win_y_high),(0,255,0), 2)
# Identify the nonzero pixels in x and y within the window #
good_left_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_xleft_low) & (nonzerox < win_xleft_high)).nonzero()[0]
good_right_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_xright_low) & (nonzerox < win_xright_high)).nonzero()[0]
# Append these indices to the lists
left_lane_inds.append(good_left_inds)
right_lane_inds.append(good_right_inds)
# If you found > minpix pixels, recenter next window on their mean position
if len(good_left_inds) > minpix:
leftx_current = np.int(np.mean(nonzerox[good_left_inds]))
if len(good_right_inds) > minpix:
rightx_current = np.int(np.mean(nonzerox[good_right_inds]))
# Concatenate the arrays of indices (previously was a list of lists of pixels)
try:
left_lane_inds = np.concatenate(left_lane_inds)
right_lane_inds = np.concatenate(right_lane_inds)
except ValueError:
# Avoids an error if the above is not implemented fully
pass
# Extract left and right line pixel positions
leftx = nonzerox[left_lane_inds]
lefty = nonzeroy[left_lane_inds]
rightx = nonzerox[right_lane_inds]
righty = nonzeroy[right_lane_inds]
return leftx, lefty, rightx, righty, out_img
def fit_polynomial(binary_warped):
# Find our lane pixels first
leftx, lefty, rightx, righty, out_img = find_lane_pixels(binary_warped)
# Fit a second order polynomial to each using `np.polyfit`
left_fit = np.polyfit(lefty, leftx, 2)
right_fit = np.polyfit(righty, rightx, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, binary_warped.shape[0]-1, binary_warped.shape[0] )
try:
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
except TypeError:
# Avoids an error if `left` and `right_fit` are still none or incorrect
print('The function failed to fit a line!')
left_fitx = 1*ploty**2 + 1*ploty
right_fitx = 1*ploty**2 + 1*ploty
## Visualization ##
# Colors in the left and right lane regions
out_img[lefty, leftx] = [255, 0, 0]
out_img[righty, rightx] = [0, 0, 255]
#
# Plots the left and right polynomials on the lane lines
# plt.plot(left_fitx, ploty, color='yellow')
# plt.plot(right_fitx, ploty, color='yellow')
return out_img, ploty, left_fit, right_fit
def fit_poly(img_shape, leftx, lefty, rightx, righty):
### TO-DO: Fit a second order polynomial to each with np.polyfit() ###
left_fit = np.polyfit(lefty, leftx, 2)
right_fit = np.polyfit(righty, rightx, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, img_shape[0]-1, img_shape[0])
### TO-DO: Calc both polynomials using ploty, left_fit and right_fit ###
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
return left_fitx, right_fitx, ploty
def search_around_poly(binary_warped,left_fit, right_fit):
# HYPERPARAMETER
# Choose the width of the margin around the previous polynomial to search
# The quiz grader expects 100 here, but feel free to tune on your own!
margin = 100
# Grab activated pixels
nonzero = binary_warped.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
### TO-DO: Set the area of search based on activated x-values ###
### within the +/- margin of our polynomial function ###
### Hint: consider the window areas for the similarly named variables ###
### in the previous quiz, but change the windows to our new search area ###
left_lane_inds = ((nonzerox > (left_fit[0]*(nonzeroy**2) + left_fit[1]*nonzeroy +
left_fit[2] - margin)) & (nonzerox < (left_fit[0]*(nonzeroy**2) +
left_fit[1]*nonzeroy + left_fit[2] + margin)))
right_lane_inds = ((nonzerox > (right_fit[0]*(nonzeroy**2) + right_fit[1]*nonzeroy +
right_fit[2] - margin)) & (nonzerox < (right_fit[0]*(nonzeroy**2) +
right_fit[1]*nonzeroy + right_fit[2] + margin)))
# Again, extract left and right line pixel positions
leftx = nonzerox[left_lane_inds]
lefty = nonzeroy[left_lane_inds]
rightx = nonzerox[right_lane_inds]
righty = nonzeroy[right_lane_inds]
# Fit new polynomials
left_fitx, right_fitx, ploty = fit_poly(binary_warped.shape, leftx, lefty, rightx, righty)
## Visualization ##
# Create an image to draw on and an image to show the selection window
out_img = np.dstack((binary_warped, binary_warped, binary_warped))*255
window_img = np.zeros_like(out_img)
# Color in left and right line pixels
out_img[nonzeroy[left_lane_inds], nonzerox[left_lane_inds]] = [255, 0, 0]
out_img[nonzeroy[right_lane_inds], nonzerox[right_lane_inds]] = [0, 0, 255]
# Generate a polygon to illustrate the search window area
# And recast the x and y points into usable format for cv2.fillPoly()
left_line_window1 = np.array([np.transpose(np.vstack([left_fitx-margin, ploty]))])
left_line_window2 = np.array([np.flipud(np.transpose(np.vstack([left_fitx+margin,
ploty])))])
left_line_pts = np.hstack((left_line_window1, left_line_window2))
right_line_window1 = np.array([np.transpose(np.vstack([right_fitx-margin, ploty]))])
right_line_window2 = np.array([np.flipud(np.transpose(np.vstack([right_fitx+margin,
ploty])))])
right_line_pts = np.hstack((right_line_window1, right_line_window2))
# Draw the lane onto the warped blank image
cv2.fillPoly(window_img, np.int_([left_line_pts]), (0,255, 0))
cv2.fillPoly(window_img, np.int_([right_line_pts]), (0,255, 0))
result = cv2.addWeighted(out_img, 1, window_img, 0.3, 0)
# Plot the polynomial lines onto the image
# plt.plot(left_fitx, ploty, color='yellow')
# plt.plot(right_fitx, ploty, color='yellow')
## End visualization steps ##
return result, left_fitx, right_fitx, ploty, out_img
def process_image(image):
# fileName = '../test_images/test5.jpg'
# image = mpimg.imread(fileName)
gray = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
R = image[:, :, 0]
G = image[:, :, 1]
B = image[:, :, 2]
# HLS color
hls = cv2.cvtColor(image, cv2.COLOR_RGB2HLS)
H = hls[:,:,0]
L = hls[:,:,1]
S = hls[:,:,2]
thresh = (218, 255)
binary = np.zeros_like(R)
binary[(R > thresh[0]) & (R <= thresh[1])] = 1
R_binary = binary
sx_L = abs_sobel_thresh(L, orient='x', sobel_kernel=3, thresh=(20, 100))
Sx_binary = sx_L
# Define a function to return the magnitude of the gradient
# for a given sobel kernel size and threshold values
# Choose a Sobel kernel size
ksize = 3 # Choose a larger odd number to smooth gradient measurements
# Apply each of the thresholding functions
gradx = abs_sobel_thresh(gray, orient='x', sobel_kernel=ksize, thresh=(14, 130))
grady = abs_sobel_thresh(gray, orient='y', sobel_kernel=ksize, thresh=(44, 115))
mag_binary = mag_thresh(image, sobel_kernel=ksize, mag_thresh=(50, 255))
dir_binary = dir_threshold(image, sobel_kernel=ksize, thresh=(np.pi/5, np.pi/3))
combined = np.zeros_like(dir_binary)
combined[((gradx == 1) & (grady == 1)) | ((mag_binary == 1) & (dir_binary == 1))] = 1
best_binary2 = combined
# Stack each channel to view their individual contributions in green and blue respectively
# This returns a stack of the two binary images, whose components you can see as different colors
color_binary = np.dstack(( np.zeros_like(Sx_binary), Sx_binary, R_binary)) * 255
# Combine the two binary thresholds
combined_binary = np.zeros_like(Sx_binary)
combined_binary[(R_binary == 1) | (Sx_binary == 1)] = 1
best_binary1 = combined_binary
# Stack each channel to view their individual contributions in green and blue respectively
# This returns a stack of the two binary images, whose components you can see as different colors
color_binary = np.dstack(( np.zeros_like(best_binary1), best_binary1, best_binary2)) * 255
# Combine the two binary thresholds
combined_binary = np.zeros_like(best_binary1)
combined_binary[(best_binary2 == 1) | (best_binary1 == 1)] = 1
# best_binary1 = combined_binary
best_final = combined_binary
img_size = (image.shape[1], image.shape[0])
src = np.float32(
[[(img_size[0] / 2) - 55, img_size[1] / 2 + 100],
[((img_size[0] / 6) - 10), img_size[1]],
[(img_size[0] * 5 / 6) + 60, img_size[1]],
[(img_size[0] / 2 + 55), img_size[1] / 2 + 100]])
dst = np.float32(
[[(img_size[0] / 4), 0],
[(img_size[0] / 4), img_size[1]],
[(img_size[0] * 3 / 4), img_size[1]],
[(img_size[0] * 3 / 4), 0]])
M = cv2.getPerspectiveTransform(src, dst)
warped2 = cv2.warpPerspective(image, M, img_size, flags=cv2.INTER_LINEAR)
# trythis = warped.copy()
trythis2 = image
warped = cv2.warpPerspective(best_final, M, img_size, flags=cv2.INTER_LINEAR)
img = warped/255
histogram = hist(img)
# plt.plot(histogram)
binary_warped = warped
out_img, ploty,left_fit, right_fit = fit_polynomial(binary_warped)
# Run image through the pipeline
# Note that in your project, you'll also want to feed in the previous fits
result, left_fitx, right_fitx, ploty, out_img = search_around_poly(binary_warped, left_fit, right_fit)
# %matplotlib qt
warped = binary_warped
image = binary_warped
# print(type(warped))
# Create an image to draw the lines on
warp_zero = np.zeros_like(warped).astype(np.uint8)
color_warp = np.dstack((warp_zero, warp_zero, warp_zero))
# Recast the x and y points into usable format for cv2.fillPoly()
pts_left = np.array([np.transpose(np.vstack([left_fitx, ploty]))])
pts_right = np.array([np.flipud(np.transpose(np.vstack([right_fitx, ploty])))])
pts = np.hstack((pts_left, pts_right))
# Draw the lane onto the warped blank image
cv2.fillPoly(color_warp, np.int_([pts]), (0,255, 0))
img_size = (warped.shape[1], warped.shape[0])
src = np.float32(
[[(img_size[0] / 2) - 55, img_size[1] / 2 + 100],
[((img_size[0] / 6) - 10), img_size[1]],
[(img_size[0] * 5 / 6) + 60, img_size[1]],
[(img_size[0] / 2 + 55), img_size[1] / 2 + 100]])
dst = np.float32(
[[(img_size[0] / 4), 0],
[(img_size[0] / 4), img_size[1]],
[(img_size[0] * 3 / 4), img_size[1]],
[(img_size[0] * 3 / 4), 0]])
Minv = cv2.getPerspectiveTransform(dst, src)
# Warp the blank back to original image space using inverse perspective matrix (Minv)
newwarp = cv2.warpPerspective(color_warp, Minv, (image.shape[1], image.shape[0]))
warp2 = cv2.warpPerspective(warped2, Minv, (image.shape[1], image.shape[0]))
# Warp the blank back to original image space using inverse perspective matrix (Minv)
# result = cv2.warpPerspective(trythis2, Minv, (image.shape[1], image.shape[0]))
# Combine the result with the original image
result = cv2.addWeighted(warp2, 1, newwarp, 0.3, 0)
# plt.imshow(result)
return newwarp
fileName = '../test_images/test5.jpg'
image = mpimg.imread(fileName)
result_a = process_image(image)
result = cv2.addWeighted(image, 1, result_a, 0.3, 0)
plt.imshow(result)
# Import everything needed to edit/save/watch video clips
from moviepy.editor import VideoFileClip
from IPython.display import HTML
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import cv2
# %matplotlib qt
%matplotlib inline
def abs_sobel_thresh(img, orient='x', sobel_kernel=3, thresh=(0, 255)):
# gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
gray =img
# Calculate directional gradient
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# 3) Take the absolute value of the x and y gradients
abs_sobelx = np.sqrt(np.square(sobelx))
abs_sobely = np.sqrt(np.square(sobely))
abs_sobelxy = np.sqrt(np.square(sobelx)+np.square(sobely))
if orient == 'x':
scaled_sobel = np.uint8(255*abs_sobelx/np.max(abs_sobelx))
else:
scaled_sobel = np.uint8(255*abs_sobely/np.max(abs_sobely))
# Apply threshold
grad_binary = np.zeros_like(scaled_sobel)
grad_binary[(scaled_sobel >= thresh[0]) & (scaled_sobel <= thresh[1])] = 1
return grad_binary
def mag_thresh(img, sobel_kernel=3, mag_thresh=(0, 255)):
# Calculate gradient magnitude
# Apply threshold
# Convert to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# Take both Sobel x and y gradients
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# Calculate the gradient magnitude
gradmag = np.sqrt(sobelx**2 + sobely**2)
# Rescale to 8 bit
scale_factor = np.max(gradmag)/255
gradmag = (gradmag/scale_factor).astype(np.uint8)
# Create a binary image of ones where threshold is met, zeros otherwise
mag_binary = np.zeros_like(gradmag)
mag_binary[(gradmag >= mag_thresh[0]) & (gradmag <= mag_thresh[1])] = 1
# Return the binary image
return mag_binary
def dir_threshold(img, sobel_kernel=3, thresh=(0, np.pi/2)):
# Calculate gradient direction
# Apply threshold
# Apply the following steps to img
# 1) Convert to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_RGB2GRAY)
# gray = img
# 2) Take the gradient in x and y separately
sobelx = cv2.Sobel(gray, cv2.CV_64F, 1, 0, ksize=sobel_kernel)
sobely = cv2.Sobel(gray, cv2.CV_64F, 0, 1, ksize=sobel_kernel)
# 3) Take the absolute value of the x and y gradients
abs_sobelx = np.sqrt(np.square(sobelx))
abs_sobely = np.sqrt(np.square(sobely))
abs_sobelxy = np.sqrt(np.square(sobelx)+np.square(sobely))
# 4) Use np.arctan2(abs_sobely, abs_sobelx) to calculate the direction of the gradient
theta = np.arctan2(abs_sobely, abs_sobelx)
# normalize
# theta = np.uint8(theta/(np.max(theta)/255))
scaled_theta = np.uint8(theta/np.max(theta/255))
# 5) Create a binary mask where direction thresholds are met
# binary_output = np.zeros_like(theta)
dir_binary = np.zeros_like(scaled_theta)
dir_binary[(theta >= thresh[0]) & (theta <= thresh[1])] = 1
return dir_binary
def hist(image):
bottom_half = image[image.shape[0]//2:,:]
histogram = np.sum(bottom_half, axis=0)
return histogram
def find_lane_pixels(binary_warped):
# Take a histogram of the bottom half of the image
histogram = np.sum(binary_warped[binary_warped.shape[0]//2:,:], axis=0)
# Create an output image to draw on and visualize the result
out_img = np.dstack((binary_warped, binary_warped, binary_warped))
# Find the peak of the left and right halves of the histogram
# These will be the starting point for the left and right lines
midpoint = np.int(histogram.shape[0]//2)
leftx_base = np.argmax(histogram[:midpoint])
rightx_base = np.argmax(histogram[midpoint:]) + midpoint
# HYPERPARAMETERS
# Choose the number of sliding windows
nwindows = 9
# Set the width of the windows +/- margin
margin = 100
# Set minimum number of pixels found to recenter window
minpix = 50
# Set height of windows - based on nwindows above and image shape
window_height = np.int(binary_warped.shape[0]//nwindows)
# Identify the x and y positions of all nonzero pixels in the image
nonzero = binary_warped.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
# Current positions to be updated later for each window in nwindows
leftx_current = leftx_base
rightx_current = rightx_base
# Create empty lists to receive left and right lane pixel indices
left_lane_inds = []
right_lane_inds = []
# Step through the windows one by one
for window in range(nwindows):
# Identify window boundaries in x and y (and right and left)
win_y_low = binary_warped.shape[0] - (window+1)*window_height
win_y_high = binary_warped.shape[0] - window*window_height
win_xleft_low = leftx_current - margin
win_xleft_high = leftx_current + margin
win_xright_low = rightx_current - margin
win_xright_high = rightx_current + margin
# Draw the windows on the visualization image
cv2.rectangle(out_img,(win_xleft_low,win_y_low),
(win_xleft_high,win_y_high),(0,255,0), 2)
cv2.rectangle(out_img,(win_xright_low,win_y_low),
(win_xright_high,win_y_high),(0,255,0), 2)
# Identify the nonzero pixels in x and y within the window #
good_left_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_xleft_low) & (nonzerox < win_xleft_high)).nonzero()[0]
good_right_inds = ((nonzeroy >= win_y_low) & (nonzeroy < win_y_high) &
(nonzerox >= win_xright_low) & (nonzerox < win_xright_high)).nonzero()[0]
# Append these indices to the lists
left_lane_inds.append(good_left_inds)
right_lane_inds.append(good_right_inds)
# If you found > minpix pixels, recenter next window on their mean position
if len(good_left_inds) > minpix:
leftx_current = np.int(np.mean(nonzerox[good_left_inds]))
if len(good_right_inds) > minpix:
rightx_current = np.int(np.mean(nonzerox[good_right_inds]))
# Concatenate the arrays of indices (previously was a list of lists of pixels)
try:
left_lane_inds = np.concatenate(left_lane_inds)
right_lane_inds = np.concatenate(right_lane_inds)
except ValueError:
# Avoids an error if the above is not implemented fully
pass
# Extract left and right line pixel positions
leftx = nonzerox[left_lane_inds]
lefty = nonzeroy[left_lane_inds]
rightx = nonzerox[right_lane_inds]
righty = nonzeroy[right_lane_inds]
return leftx, lefty, rightx, righty, out_img
def fit_polynomial(binary_warped):
# Find our lane pixels first
leftx, lefty, rightx, righty, out_img = find_lane_pixels(binary_warped)
# Fit a second order polynomial to each using `np.polyfit`
left_fit = np.polyfit(lefty, leftx, 2)
right_fit = np.polyfit(righty, rightx, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, binary_warped.shape[0]-1, binary_warped.shape[0] )
try:
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
except TypeError:
# Avoids an error if `left` and `right_fit` are still none or incorrect
print('The function failed to fit a line!')
left_fitx = 1*ploty**2 + 1*ploty
right_fitx = 1*ploty**2 + 1*ploty
## Visualization ##
# Colors in the left and right lane regions
out_img[lefty, leftx] = [255, 0, 0]
out_img[righty, rightx] = [0, 0, 255]
#
# Plots the left and right polynomials on the lane lines
# plt.plot(left_fitx, ploty, color='yellow')
# plt.plot(right_fitx, ploty, color='yellow')
return out_img, ploty, left_fit, right_fit
def fit_poly(img_shape, leftx, lefty, rightx, righty):
### TO-DO: Fit a second order polynomial to each with np.polyfit() ###
left_fit = np.polyfit(lefty, leftx, 2)
right_fit = np.polyfit(righty, rightx, 2)
# Generate x and y values for plotting
ploty = np.linspace(0, img_shape[0]-1, img_shape[0])
### TO-DO: Calc both polynomials using ploty, left_fit and right_fit ###
left_fitx = left_fit[0]*ploty**2 + left_fit[1]*ploty + left_fit[2]
right_fitx = right_fit[0]*ploty**2 + right_fit[1]*ploty + right_fit[2]
return left_fitx, right_fitx, ploty
def search_around_poly(binary_warped,left_fit, right_fit):
# HYPERPARAMETER
# Choose the width of the margin around the previous polynomial to search
# The quiz grader expects 100 here, but feel free to tune on your own!
margin = 100
# Grab activated pixels
nonzero = binary_warped.nonzero()
nonzeroy = np.array(nonzero[0])
nonzerox = np.array(nonzero[1])
### TO-DO: Set the area of search based on activated x-values ###
### within the +/- margin of our polynomial function ###
### Hint: consider the window areas for the similarly named variables ###
### in the previous quiz, but change the windows to our new search area ###
left_lane_inds = ((nonzerox > (left_fit[0]*(nonzeroy**2) + left_fit[1]*nonzeroy +
left_fit[2] - margin)) & (nonzerox < (left_fit[0]*(nonzeroy**2) +
left_fit[1]*nonzeroy + left_fit[2] + margin)))
right_lane_inds = ((nonzerox > (right_fit[0]*(nonzeroy**2) + right_fit[1]*nonzeroy +
right_fit[2] - margin)) & (nonzerox < (right_fit[0]*(nonzeroy**2) +
right_fit[1]*nonzeroy + right_fit[2] + margin)))
# Again, extract left and right line pixel positions
leftx = nonzerox[left_lane_inds]
lefty = nonzeroy[left_lane_inds]
rightx = nonzerox[right_lane_inds]
righty = nonzeroy[right_lane_inds]
# Fit new polynomials
left_fitx, right_fitx, ploty = fit_poly(binary_warped.shape, leftx, lefty, rightx, righty)
## Visualization ##
# Create an image to draw on and an image to show the selection window
out_img = np.dstack((binary_warped, binary_warped, binary_warped))*255
window_img = np.zeros_like(out_img)
# Color in left and right line pixels
out_img[nonzeroy[left_lane_inds], nonzerox[left_lane_inds]] = [255, 0, 0]
out_img[nonzeroy[right_lane_inds], nonzerox[right_lane_inds]] = [0, 0, 255]
# Generate a polygon to illustrate the search window area
# And recast the x and y points into usable format for cv2.fillPoly()
left_line_window1 = np.array([np.transpose(np.vstack([left_fitx-margin, ploty]))])
left_line_window2 = np.array([np.flipud(np.transpose(np.vstack([left_fitx+margin,
ploty])))])
left_line_pts = np.hstack((left_line_window1, left_line_window2))
right_line_window1 = np.array([np.transpose(np.vstack([right_fitx-margin, ploty]))])
right_line_window2 = np.array([np.flipud(np.transpose(np.vstack([right_fitx+margin,
ploty])))])
right_line_pts = np.hstack((right_line_window1, right_line_window2))
# Draw the lane onto the warped blank image
cv2.fillPoly(window_img, np.int_([left_line_pts]), (0,255, 0))
cv2.fillPoly(window_img, np.int_([right_line_pts]), (0,255, 0))
result = cv2.addWeighted(out_img, 1, window_img, 0.3, 0)
# Plot the polynomial lines onto the image
# plt.plot(left_fitx, ploty, color='yellow')
# plt.plot(right_fitx, ploty, color='yellow')
## End visualization steps ##
return result, left_fitx, right_fitx, ploty, out_img
def process_image(image):
# fileName = '../test_images/test5.jpg'
# image = mpimg.imread(fileName)
gray = cv2.cvtColor(image, cv2.COLOR_RGB2GRAY)
R = image[:, :, 0]
G = image[:, :, 1]
B = image[:, :, 2]
# HLS color
hls = cv2.cvtColor(image, cv2.COLOR_RGB2HLS)
H = hls[:,:,0]
L = hls[:,:,1]
S = hls[:,:,2]
thresh = (218, 255)
binary = np.zeros_like(R)
binary[(R > thresh[0]) & (R <= thresh[1])] = 1
R_binary = binary
sx_L = abs_sobel_thresh(L, orient='x', sobel_kernel=3, thresh=(20, 100))
Sx_binary = sx_L
# Define a function to return the magnitude of the gradient
# for a given sobel kernel size and threshold values
# Choose a Sobel kernel size
ksize = 3 # Choose a larger odd number to smooth gradient measurements
# Apply each of the thresholding functions
gradx = abs_sobel_thresh(gray, orient='x', sobel_kernel=ksize, thresh=(14, 130))
grady = abs_sobel_thresh(gray, orient='y', sobel_kernel=ksize, thresh=(44, 115))
mag_binary = mag_thresh(image, sobel_kernel=ksize, mag_thresh=(50, 255))
dir_binary = dir_threshold(image, sobel_kernel=ksize, thresh=(np.pi/5, np.pi/3))
combined = np.zeros_like(dir_binary)
combined[((gradx == 1) & (grady == 1)) | ((mag_binary == 1) & (dir_binary == 1))] = 1
best_binary2 = combined
# Stack each channel to view their individual contributions in green and blue respectively
# This returns a stack of the two binary images, whose components you can see as different colors
color_binary = np.dstack(( np.zeros_like(Sx_binary), Sx_binary, R_binary)) * 255
# Combine the two binary thresholds
combined_binary = np.zeros_like(Sx_binary)
combined_binary[(R_binary == 1) | (Sx_binary == 1)] = 1
best_binary1 = combined_binary
# Stack each channel to view their individual contributions in green and blue respectively
# This returns a stack of the two binary images, whose components you can see as different colors
color_binary = np.dstack(( np.zeros_like(best_binary1), best_binary1, best_binary2)) * 255
# Combine the two binary thresholds
combined_binary = np.zeros_like(best_binary1)
combined_binary[(best_binary2 == 1) | (best_binary1 == 1)] = 1
# best_binary1 = combined_binary
best_final = combined_binary
img_size = (image.shape[1], image.shape[0])
src = np.float32(
[[(img_size[0] / 2) - 55, img_size[1] / 2 + 100],
[((img_size[0] / 6) - 10), img_size[1]],
[(img_size[0] * 5 / 6) + 60, img_size[1]],
[(img_size[0] / 2 + 55), img_size[1] / 2 + 100]])
dst = np.float32(
[[(img_size[0] / 4), 0],
[(img_size[0] / 4), img_size[1]],
[(img_size[0] * 3 / 4), img_size[1]],
[(img_size[0] * 3 / 4), 0]])
M = cv2.getPerspectiveTransform(src, dst)
warped2 = cv2.warpPerspective(image, M, img_size, flags=cv2.INTER_LINEAR)
# trythis = warped.copy()
trythis2 = image
warped = cv2.warpPerspective(best_final, M, img_size, flags=cv2.INTER_LINEAR)
img = warped/255
histogram = hist(img)
# plt.plot(histogram)
binary_warped = warped
out_img, ploty,left_fit, right_fit = fit_polynomial(binary_warped)
# Run image through the pipeline
# Note that in your project, you'll also want to feed in the previous fits
result, left_fitx, right_fitx, ploty, out_img = search_around_poly(binary_warped, left_fit, right_fit)
# %matplotlib qt
warped = binary_warped
image = binary_warped
# print(type(warped))
# Create an image to draw the lines on
warp_zero = np.zeros_like(warped).astype(np.uint8)
color_warp = np.dstack((warp_zero, warp_zero, warp_zero))
# Recast the x and y points into usable format for cv2.fillPoly()
pts_left = np.array([np.transpose(np.vstack([left_fitx, ploty]))])
pts_right = np.array([np.flipud(np.transpose(np.vstack([right_fitx, ploty])))])
pts = np.hstack((pts_left, pts_right))
# Draw the lane onto the warped blank image
cv2.fillPoly(color_warp, np.int_([pts]), (0,255, 0))
# img_size = (warped.shape[1], warped.shape[0])
Minv = cv2.getPerspectiveTransform(dst, src)
# Warp the blank back to original image space using inverse perspective matrix (Minv)
newwarp = cv2.warpPerspective(color_warp, Minv, (image.shape[1], image.shape[0]))
warp2 = cv2.warpPerspective(warped2, Minv, (image.shape[1], image.shape[0]))
# Warp the blank back to original image space using inverse perspective matrix (Minv)
# result = cv2.warpPerspective(trythis2, Minv, (image.shape[1], image.shape[0]))
# Combine the result with the original image
# result = cv2.addWeighted(warp2, 1, newwarp, 0.3, 0)
# plt.imshow(result)
return newwarp
def process_image_end(img):
image = img
result_a = process_image(image)
result = cv2.addWeighted(image, 1, result_a, 0.3, 0)
return result
fileName = '../test_images/test3.jpg'
image = mpimg.imread(fileName)
result =process_image_end(image)
# result_a = process_image(image)
# result = cv2.addWeighted(image, 1, result_a, 0.3, 0)
plt.imshow(result)
# process_image_end
white_output = '../HC_video_output/project_video.mp4'
## To speed up the testing process you may want to try your pipeline on a shorter subclip of the video
## To do so add .subclip(start_second,end_second) to the end of the line below
## Where start_second and end_second are integer values representing the start and end of the subclip
## You may also uncomment the following line for a subclip of the first 5 seconds
clip1 = VideoFileClip("../project_video.mp4").subclip(22,24)
# clip1 = VideoFileClip("../project_video.mp4")
white_clip = clip1.fl_image(process_image_end) #NOTE: this function expects color images!!
%time white_clip.write_videofile(white_output, audio=False)
HTML("""
<video width="960" height="540" controls>
<source src="{0}">
</video>
""".format(white_output))
```
| github_jupyter |
```
from keras.models import Model, load_model
from keras.layers import Input, Convolution1D, Dot, Dense, Activation, Concatenate
from keras.utils import Sequence
import numpy as np
import random
import json
from typing import List, Dict, Tuple
```
## Load the model
```
with open('options.json', encoding='utf-8') as f:
options = json.load(f)
print(options)
latent_dim = 256
def create_model():
# Define an input sequence and process it.
encoder_inputs = Input(shape=(None, options['num_encoder_tokens']))
# Encoder
x_encoder = Convolution1D(256, kernel_size=3, activation='relu',
padding='causal')(encoder_inputs)
x_encoder = Convolution1D(256, kernel_size=3, activation='relu',
padding='causal', dilation_rate=2)(x_encoder)
x_encoder = Convolution1D(256, kernel_size=3, activation='relu',
padding='causal', dilation_rate=4)(x_encoder)
decoder_inputs = Input(shape=(None, options['num_decoder_tokens']))
# Decoder
x_decoder = Convolution1D(256, kernel_size=3, activation='relu',
padding='causal')(decoder_inputs)
x_decoder = Convolution1D(256, kernel_size=3, activation='relu',
padding='causal', dilation_rate=2)(x_decoder)
x_decoder = Convolution1D(256, kernel_size=3, activation='relu',
padding='causal', dilation_rate=4)(x_decoder)
# Attention
attention = Dot(axes=[2, 2])([x_decoder, x_encoder])
attention = Activation('softmax')(attention)
context = Dot(axes=[2, 1])([attention, x_encoder])
decoder_combined_context = Concatenate(axis=-1)([context, x_decoder])
decoder_outputs = Convolution1D(64, kernel_size=3, activation='relu',
padding='causal')(decoder_combined_context)
decoder_outputs = Convolution1D(64, kernel_size=3, activation='relu',
padding='causal')(decoder_outputs)
# Output
decoder_dense = Dense(options['num_decoder_tokens'], activation='softmax')
decoder_outputs = decoder_dense(decoder_outputs)
# Define the model that will turn
# `encoder_input_data` & `decoder_input_data` into `decoder_target_data`
model = Model([encoder_inputs, decoder_inputs], decoder_outputs)
model.summary()
return model
#model = create_model()
#model.load_weights('s2s.h5')
model = load_model('s2s.h5')
#model.compile(optimizer='adam', loss='categorical_crossentropy')
user_inputs = [
"There are numerous weaknesses with the bag of words model especially when applied to natural language processing tasks that graph ranking algorithms such as TextRank are able to address.",
"Since purple yams happen to be starchy root vegetables, they also happen to be a great source of carbs, potassium, and vitamin C.",
"Recurrent Neural Networks (RNNs) have been used successfully for many tasks involving sequential data such as machine translation, sentiment analysis, image captioning, time-series prediction etc.",
"Improved RNN models such as Long Short-Term Memory networks (LSTMs) enable training on long sequences overcoming problems like vanishing gradients.",
"However, even the more advanced models have their limitations and researchers had a hard time developing high-quality models when working with long data sequences.",
"In machine translation, for example, the RNN has to find connections between long input and output sentences composed of dozens of words.",
"It seemed that the existing RNN architectures needed to be changed and adapted to better deal with such tasks.",
"Wenger ended his 22-year Gunners reign after the 2017-18 season and previously stated he intended to take charge of a new club in early 2019.",
"It will not prevent the Frenchman from resuming his career in football.",
"However 12 months out of work has given him a different outlook and may influence his next move.",
]
def user_input_to_inputs(ui: List[str]):
max_encoder_seq_length = options['max_encoder_seq_length']
num_encoder_tokens = options['num_encoder_tokens']
input_token_index = options['input_token_index']
encoder_input_data = np.zeros(
(len(ui), max_encoder_seq_length, num_encoder_tokens),
dtype='float32')
for i, input_text in enumerate(ui):
for t, char in enumerate(input_text):
encoder_input_data[i, t, input_token_index[char]] = 1.
return encoder_input_data
inputs = user_input_to_inputs(user_inputs)
def print_predictions(inputs:np.array, user_inputs: List[str]):
max_decoder_seq_length = options['max_decoder_seq_length']
num_decoder_tokens = options['num_decoder_tokens']
input_token_index = options['input_token_index']
target_token_index = options['target_token_index']
# Define sampling models
reverse_input_char_index = dict(
(i, char) for char, i in input_token_index.items())
reverse_target_char_index = dict(
(i, char) for char, i in target_token_index.items())
in_encoder = inputs
in_decoder = np.zeros(
(len(in_encoder), max_decoder_seq_length, num_decoder_tokens),
dtype='float32')
in_decoder[:, 0, target_token_index["\t"]] = 1
predict = np.zeros(
(len(in_encoder), max_decoder_seq_length),
dtype='float32')
for i in range(max_decoder_seq_length - 1):
predict = model.predict([in_encoder, in_decoder])
predict = predict.argmax(axis=-1)
predict_ = predict[:, i].ravel().tolist()
for j, x in enumerate(predict_):
in_decoder[j, i + 1, x] = 1
for seq_index in range(len(in_encoder)):
# Take one sequence (part of the training set)
# for trying out decoding.
output_seq = predict[seq_index, :].ravel().tolist()
decoded = []
for x in output_seq:
if reverse_target_char_index[x] == "\n":
break
else:
decoded.append(reverse_target_char_index[x])
decoded_sentence = "".join(decoded)
print('-')
print('Input sentence:', user_inputs[seq_index])
print('Decoded sentence:', decoded_sentence)
print_predictions(inputs, user_inputs)
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Tutorials/GlobalSurfaceWater/2_water_occurrence_change_intensity.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Tutorials/GlobalSurfaceWater/2_water_occurrence_change_intensity.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=Tutorials/GlobalSurfaceWater/2_water_occurrence_change_intensity.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Tutorials/GlobalSurfaceWater/2_water_occurrence_change_intensity.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as emap
except:
import geemap as emap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google Satellite`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py#L13) can be added using the `Map.add_basemap()` function.
```
Map = emap.Map(center=[40,-100], zoom=4)
Map.add_basemap('ROADMAP') # Add Google Map
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
###############################
# Asset List
###############################
gsw = ee.Image('JRC/GSW1_1/GlobalSurfaceWater')
occurrence = gsw.select('occurrence')
change = gsw.select("change_abs")
roi = ee.Geometry.Polygon(
[[[-74.17213, -8.65569],
[-74.17419, -8.39222],
[-74.38362, -8.36980],
[-74.43031, -8.61293]]])
###############################
# Constants
###############################
VIS_OCCURRENCE = {
'min':0,
'max':100,
'palette': ['red', 'blue']
}
VIS_CHANGE = {
'min':-50,
'max':50,
'palette': ['red', 'black', 'limegreen']
}
VIS_WATER_MASK = {
'palette': ['white', 'black']
}
###############################
# Calculations
###############################
# Create a water mask layer, and set the image mask so that non-water areas are transparent.
water_mask = occurrence.gt(90).mask(1)
# # Generate a histogram object and print it to the console tab.
# histogram = ui.Chart.image.histogram({
# 'image': change,
# 'region': roi,
# 'scale': 30,
# 'minBucketWidth': 10
# })
# histogram.setOptions({
# title: 'Histogram of surface water change intensity.'
# })
# print(histogram)
###############################
# Initialize Map Location
###############################
# Uncomment one of the following statements to center the map on
# a particular location.
# Map.setCenter(-90.162, 29.8597, 10) # New Orleans, USA
# Map.setCenter(-114.9774, 31.9254, 10) # Mouth of the Colorado River, Mexico
# Map.setCenter(-111.1871, 37.0963, 11) # Lake Powell, USA
# Map.setCenter(149.412, -35.0789, 11) # Lake George, Australia
# Map.setCenter(105.26, 11.2134, 9) # Mekong River Basin, SouthEast Asia
# Map.setCenter(90.6743, 22.7382, 10) # Meghna River, Bangladesh
# Map.setCenter(81.2714, 16.5079, 11) # Godavari River Basin Irrigation Project, India
# Map.setCenter(14.7035, 52.0985, 12) # River Oder, Germany & Poland
# Map.setCenter(-59.1696, -33.8111, 9) # Buenos Aires, Argentina\
Map.setCenter(-74.4557, -8.4289, 11) # Ucayali River, Peru
###############################
# Map Layers
###############################
Map.addLayer(water_mask, VIS_WATER_MASK, '90% occurrence water mask', False)
Map.addLayer(occurrence.updateMask(occurrence.divide(100)), VIS_OCCURRENCE, "Water Occurrence (1984-2015)")
Map.addLayer(change, VIS_CHANGE,'occurrence change intensity')
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
```
from keras.models import Sequential
from keras.layers import Dense
from keras import optimizers
import numpy as np
import pandas as pd
import mlflow
import mlflow.keras
import warnings
import mlflow.pyfunc
# source: https://androidkt.com/linear-regression-model-in-keras/
# Modified and extended
# Generate X, y data
X_fahrenheit = np.array(
[-140, -136, -124, -112, -105, -96, -88, -75, -63, -60,
-58, -40, -20, -10, 0, 30, 35, 48, 55, 69, 81, 89, 95,
99,105, 110, 120, 135, 145, 158, 160, 165, 170, 175, 180,
185, 187, 190, 195, 198, 202, 205, 207, 210, 215, 220], dtype=float)
y_celsius = np.array(
[-95.55, -93.33, -86.66, -80, -76.11, -71.11, -66.66, -59.44, -52.77, -51.11,
-50, -40, -28.88, -23.33, -17.77, -1.11, 1.66, 8.88, 12, 20,
27.22, 31.66, 35, 37.22, 40.55, 43.33, 48.88, 57.22, 62.77, 70,
71.11, 73.88, 76.66, 79.44, 82.22, 85, 86.11,87.77,90.55, 92.22,
94.44, 96.11, 97.22, 98.88, 101.66, 104.44], dtype=float)
# Define the model
def baseline_model():
model = Sequential([
Dense(64, activation='relu', input_shape=[1]),
Dense(64, activation='relu'),
Dense(1)
])
optimizer = optimizers.RMSprop(0.001)
# Compile the model
model.compile(loss='mean_squared_error',
optimizer=optimizer,
metrics=['mean_absolute_error', 'mean_squared_error'])
return model
def mlflow_run(params, run_name="Keras Linear Regression"):
# Start MLflow run and log everyting...
with mlflow.start_run(run_name=run_name) as run:
model = baseline_model()
# single line of MLflow Fluent API obviates the need to log
# individual parameters, metrics, model, artifacts etc...
# https://mlflow.org/docs/latest/python_api/mlflow.keras.html#mlflow.keras.autolog
mlflow.keras.autolog()
model.fit(X_fahrenheit, y_celsius, batch_size=params['batch_size'], epochs=params['epochs'])
run_id = run.info.run_uuid
exp_id = run.info.experiment_id
for f in [200, 206]:
print(f"F={f}; C={model.predict([f])}")
return (exp_id, run_id)
# suppress any deprecated warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
params = {'batch_size': 5, 'epochs': 1000}
(exp_id, run_id) = mlflow_run(params)
print(f"Finished Experiment id={exp_id} and run id = {run_id}")
# Load this Keras Model as a pyfunc model and make a prediction
pyfunc_uri = f"runs:/{run_id}/model"
pyfunc_model = mlflow.pyfunc.load_model(pyfunc_uri)
# Given Fahernheight -> Predict Celcius
df = pd.DataFrame(np.array([32, 212, 200, 206]))
pred = pyfunc_model.predict(df)
print(pred)
! mlflow ui
```
| github_jupyter |
# Train the StarNet Model
This notebook takes you through the steps of how to train a StarNet Model
- Required Python packages: `numpy h5py keras`
- Required data files: training_data.h5, mean_and_std.npy
Note: We use tensorflow for the keras backend.
```
import numpy as np
import h5py
import random
from keras.models import Model
from keras.layers import Input, Dense, InputLayer, Flatten, Reshape
from keras.layers.convolutional import Conv1D
from keras.layers.convolutional import MaxPooling1D
from keras.optimizers import Adam
from keras.callbacks import EarlyStopping, ReduceLROnPlateau
from keras.utils import HDF5Matrix
datadir = ""
training_set = datadir + 'training_data.h5'
normalization_data = datadir + 'mean_and_std.npy'
```
** Normalization **
Write a function to normalize the output labels. Each label will be normalized to have approximately have a mean of zero and unit variance.
NOTE: This is necessary to put output labels on a similar scale in order for the model to train properly, this process is reversed in the test stage to give the output labels their proper units
```
mean_and_std = np.load(normalization_data)
mean_labels = mean_and_std[0]
std_labels = mean_and_std[1]
def normalize(labels):
# Normalize labels
return (labels-mean_labels) / std_labels
```
** Obtain training data **
Here we will collect the output labels for the training and cross-validation sets, then normalize each.
Next we will create an HDF5Matrix for the training and cross-validation input spectra rather than loading them all into memory. This is useful to save RAM when training the model.
```
# Define the number of output labels
num_labels = np.load(datadir+'mean_and_std.npy').shape[1]
# Define the number of training spectra
num_train = 41000
# Load labels
with h5py.File(training_set, 'r') as F:
y_train = np.hstack((F['TEFF'][0:num_train], F['LOGG'][0:num_train], F['FE_H'][0:num_train]))
y_cv = np.hstack((F['TEFF'][num_train:], F['LOGG'][num_train:], F['FE_H'][num_train:]))
# Normalize labels
y_train = normalize(y_train)
y_cv = normalize(y_cv)
# Create the spectra training and cv datasets
x_train = HDF5Matrix(training_set, 'spectrum',
start=0, end=num_train)
x_cv = HDF5Matrix(training_set, 'spectrum',
start=num_train, end=None)
# Define the number of output labels
num_labels = y_train.shape[1]
num_fluxes = x_train.shape[1]
print('Each spectrum contains ' + str(num_fluxes) + ' wavelength bins')
print('Training set includes ' + str(x_train.shape[0]) +
' spectra and the cross-validation set includes ' + str(x_cv.shape[0])+' spectra')
```
**Build the StarNet model architecture**
The StarNet architecture is built with:
- input layer
- 2 convolutional layers
- 1 maxpooling layer followed by flattening for the fully connected layer
- 2 fully connected layers
- output layer
First, let's define some model variables.
```
# activation function used following every layer except for the output layers
activation = 'relu'
# model weight initializer
initializer = 'he_normal'
# number of filters used in the convolutional layers
num_filters = [4,16]
# length of the filters in the convolutional layers
filter_length = 8
# length of the maxpooling window
pool_length = 4
# number of nodes in each of the hidden fully connected layers
num_hidden = [256,128]
# number of spectra fed into model at once during training
batch_size = 64
# maximum number of interations for model training
max_epochs = 30
# initial learning rate for optimization algorithm
lr = 0.0007
# exponential decay rate for the 1st moment estimates for optimization algorithm
beta_1 = 0.9
# exponential decay rate for the 2nd moment estimates for optimization algorithm
beta_2 = 0.999
# a small constant for numerical stability for optimization algorithm
optimizer_epsilon = 1e-08
# Input spectra
input_spec = Input(shape=(num_fluxes,), name='starnet_input_x')
# Reshape spectra for CNN layers
cur_in = Reshape((num_fluxes, 1))(input_spec)
# CNN layers
cur_in = Conv1D(kernel_initializer=initializer, activation=activation,
padding="same", filters=num_filters[0], kernel_size=filter_length)(cur_in)
cur_in = Conv1D(kernel_initializer=initializer, activation=activation,
padding="same", filters=num_filters[1], kernel_size=filter_length)(cur_in)
# Max pooling layer
cur_in = MaxPooling1D(pool_size=pool_length)(cur_in)
# Flatten the current input for the fully-connected layers
cur_in = Flatten()(cur_in)
# Fully-connected layers
cur_in = Dense(units=num_hidden[0], kernel_initializer=initializer,
activation=activation)(cur_in)
cur_in = Dense(units=num_hidden[1], kernel_initializer=initializer,
activation=activation)(cur_in)
# Output nodes
output_label = Dense(units=num_labels, activation="linear",
input_dim=num_hidden[1], name='starnet_output_y')(cur_in)
model = Model(input_spec, output_label, name='StarNet')
```
**More model techniques**
* The `Adam` optimizer is the gradient descent algorithm used for minimizing the loss function
* `EarlyStopping` uses the cross-validation set to test the model following every iteration and stops the training if the cv loss does not decrease by `min_delta` after `patience` iterations
* `ReduceLROnPlateau` is a form of learning rate decay where the learning rate is decreased by a factor of `factor` if the training loss does not decrease by `epsilon` after `patience` iterations unless the learning rate has reached `min_lr`
```
# Default loss function parameters
early_stopping_min_delta = 0.0001
early_stopping_patience = 4
reduce_lr_factor = 0.5
reuce_lr_epsilon = 0.0009
reduce_lr_patience = 2
reduce_lr_min = 0.00008
# loss function to minimize
loss_function = 'mean_squared_error'
# compute mean absolute deviation
metrics = ['mae']
optimizer = Adam(lr=lr, beta_1=beta_1, beta_2=beta_2, epsilon=optimizer_epsilon, decay=0.0)
early_stopping = EarlyStopping(monitor='val_loss', min_delta=early_stopping_min_delta,
patience=early_stopping_patience, verbose=2, mode='min')
reduce_lr = ReduceLROnPlateau(monitor='loss', factor=0.5, epsilon=reuce_lr_epsilon,
patience=reduce_lr_patience, min_lr=reduce_lr_min, mode='min', verbose=2)
```
**Compile model**
```
model.compile(optimizer=optimizer, loss=loss_function, metrics=metrics)
model.summary()
```
**Train model**
```
model.fit(x_train, y_train, validation_data=(x_cv, y_cv),
epochs=max_epochs, verbose=1, shuffle='batch',
callbacks=[early_stopping, reduce_lr])
```
**Save model**
```
starnet_model = 'starnet_cnn.h5'
model.save(datadir + starnet_model)
print(starnet_model+' saved.')
```
| github_jupyter |
# Calculate ensemble mean
This notebook shows an example of working with ensemble weather forecast data. This is based on the [ensemble example from the iris documentation](https://scitools.org.uk/iris/docs/v0.9.1/examples/graphics/lagged_ensemble.html).
In this notebook we will:
- Load multiple files from the MOGREPS dataset on AWS Earth
- Add missing metadata to cubes from the object key
- Plot multiple cubes in a 'postage stamp' style plot
- Calculate and plot the ensemble mean
```
import iris
import numpy as np
import matplotlib.pyplot as plt
import iris.plot as iplt
import warnings # Useful for supressing some of the chatty iris warnings
```
## Files to load
As the files in the MOGREPS dataset contain metadata information in the object key we can load twelve files, each of which contain a different ensemble member. Let's start by generating the file paths based on our prior knowledge of the dataset.
```
MEMBERS = range(12)
FILE_LIST = ["/s3/mogreps-uk/prods_op_mogreps-uk_20130102_09_{member:02d}_003.nc".format(member=member) for member in MEMBERS]
FILE_LIST
```
## Loading the data
If we inspect a single cube within our dataset we will sadly discover that the ensemble member information is missing.
```
with warnings.catch_warnings():
warnings.simplefilter('ignore')
cube = iris.load_cube(FILE_LIST[0], 'surface_air_pressure')
cube
```
Thankfully when calling `iris.load` you can provide a callback function which will be called on each cube that has been loaded before attempting to merge them into a single cube. We will use this callback to extract the `realization` number from the object key metadata and add it to the cube as a coordinate. We will also pass a constraint telling iris that we are only interested in the surface air pressure data.
```
def realization_metadata(cube, field, fname):
"""
A function which modifies the cube's metadata to add a "realization" (ensemble member) coordinate from the filename if one
doesn't already exist in the cube.
"""
# add an ensemble member coordinate if one doesn't already exist
if not cube.coords('realization'):
realization_number = fname[-9:-7]
realization_coord = iris.coords.AuxCoord(np.int32(realization_number), 'realization')
cube.add_aux_coord(realization_coord)
with warnings.catch_warnings():
warnings.simplefilter('ignore')
air_pressure = iris.load_cube(
FILE_LIST,
iris.Constraint(name='surface_air_pressure'),
callback=realization_metadata)
air_pressure
```
## Extract the first timestep
The data we have loaded has multiple timesteps, however we are only interested in a single timestep so we will extract it. We could potentially plot all timesteps and create a video of the data as a further exercise.
```
first_timestep = air_pressure[:, -1, :, :]
first_timestep
```
## Postage stamp plots
In the following code cell we will create a plot, iterate over our cube sliced by `realization` and create a subplot for each one displayed as a grid in our overall plot. We will also add a colourbar, title, axis ticks, etc.
```
# Create a wider than normal figure to support our many plots
plt.figure(figsize=(12, 6), dpi=100)
# Also manually adjust the spacings which are used when creating subplots
plt.gcf().subplots_adjust(hspace=0.05, wspace=0.05, top=0.95, bottom=0.05, left=0.075, right=0.925)
# iterate over all possible latitude longitude slices
for cube in first_timestep.slices(['grid_latitude', 'grid_longitude']):
# get the ensemble member number from the ensemble coordinate
ens_member = cube.coord('realization').points[0]
# plot the data in a 4x3 grid, with each plot's position in the grid being determined by ensemble member number
plt.subplot(3, 6, ens_member+1)
cf = iplt.pcolormesh(cube)
# add coastlines
plt.gca().coastlines('50m')
# make an axes to put the shared colorbar in
colorbar_axes = plt.gcf().add_axes([0.35, 0.1, 0.3, 0.05])
colorbar = plt.colorbar(cf, colorbar_axes, orientation='horizontal')
colorbar.set_label('%s' % first_timestep.units)
# limit the colorbar to 8 tick marks
import matplotlib.ticker
colorbar.locator = matplotlib.ticker.MaxNLocator(8)
colorbar.update_ticks()
# get the time for the entire plot
time_coord = first_timestep.coord('time')
time = time_coord.units.num2date(time_coord.points[0])
# set a global title for the postage stamps with the date formated by "monthname year"
plt.suptitle('Surface temperature ensemble forecasts for %s' % time.strftime('%B %Y'))
iplt.show()
```
## Calculating the ensemble mean
We may also wish to calculate the mean of the ensemble. We can do this by collapsing our cube along the `realization` dimension using the `MEAN` analysis operator.
```
ensemble_mean = first_timestep.collapsed('realization', iris.analysis.MEAN)
ensemble_mean
```
Now that we have calculated the mean we can plot it. This is an excellent example of the laziness in iris, the mean isn't actually calculated at this point, we have just created a cube which represents the mean of the data. When we ask matplotlib to draw the data it will then go away and run the calculation.
```
fig = plt.figure(figsize=(10, 15))
qplt.pcolormesh(ensemble_mean)
# Add coastlines to the map created by contourf.
plt.gca().coastlines('50m')
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/adityabingi/cs330-meta-learning/blob/main/CS330_Homework2_Stencil.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
##Setup
You will need to make a copy of this Colab notebook in your Google Drive before you can edit the homework files. You can do so with **File → Save a copy in Drive**.
```
import os
from google_drive_downloader import GoogleDriveDownloader as gdd
# Need to download the Omniglot dataset -- DON'T MODIFY THIS CELL
if not os.path.isdir('./omniglot_resized'):
gdd.download_file_from_google_drive(file_id='1iaSFXIYC3AB8q9K_M-oVMa4pmB7yKMtI',
dest_path='./omniglot_resized.zip',
unzip=True)
assert os.path.isdir('./omniglot_resized')
from google.colab import drive
drive.mount('/content/drive')
""" Utility functions. """
## NOTE: You do not need to modify this block but you will need to use it.
import numpy as np
import os
import random
import tensorflow as tf
## Loss utilities
def cross_entropy_loss(pred, label, k_shot):
return tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits=pred, labels=tf.stop_gradient(label)) / k_shot)
def accuracy(labels, predictions):
return tf.reduce_mean(tf.cast(tf.equal(labels, predictions), dtype=tf.float32))
"""Convolutional layers used by MAML model."""
## NOTE: You do not need to modify this block but you will need to use it.
seed = 123
def conv_block(inp, cweight, bweight, bn, activation=tf.nn.relu, residual=False, training=True):
""" Perform, conv, batch norm, nonlinearity, and max pool """
stride, no_stride = [1,2,2,1], [1,1,1,1]
conv_output = tf.nn.conv2d(input=inp, filters=cweight, strides=no_stride, padding='SAME') + bweight
normed = bn(conv_output)
normed = activation(normed)
normed = tf.nn.max_pool(normed, stride, stride, padding='VALID')
return normed
class ConvLayers(tf.keras.layers.Layer):
def __init__(self, channels, dim_hidden, dim_output, img_size):
super(ConvLayers, self).__init__()
self.channels = channels
self.dim_hidden = dim_hidden
self.dim_output = dim_output
self.img_size = img_size
weights = {}
dtype = tf.float32
weight_initializer = tf.keras.initializers.GlorotUniform()
k = 3
weights['conv1'] = tf.Variable(weight_initializer(shape=[k, k, self.channels, self.dim_hidden]), name='conv1', dtype=dtype)
weights['b1'] = tf.Variable(tf.zeros([self.dim_hidden]), name='b1')
self.bn1 = tf.keras.layers.BatchNormalization(name='bn1')
weights['conv2'] = tf.Variable(weight_initializer(shape=[k, k, self.dim_hidden, self.dim_hidden]), name='conv2', dtype=dtype)
weights['b2'] = tf.Variable(tf.zeros([self.dim_hidden]), name='b2')
self.bn2 = tf.keras.layers.BatchNormalization(name='bn2')
weights['conv3'] = tf.Variable(weight_initializer(shape=[k, k, self.dim_hidden, self.dim_hidden]), name='conv3', dtype=dtype)
weights['b3'] = tf.Variable(tf.zeros([self.dim_hidden]), name='b3')
self.bn3 = tf.keras.layers.BatchNormalization(name='bn3')
weights['conv4'] = tf.Variable(weight_initializer([k, k, self.dim_hidden, self.dim_hidden]), name='conv4', dtype=dtype)
weights['b4'] = tf.Variable(tf.zeros([self.dim_hidden]), name='b4')
self.bn4 = tf.keras.layers.BatchNormalization(name='bn4')
weights['w5'] = tf.Variable(weight_initializer(shape=[self.dim_hidden, self.dim_output]), name='w5', dtype=dtype)
weights['b5'] = tf.Variable(tf.zeros([self.dim_output]), name='b5')
self.conv_weights = weights
def call(self, inp, weights, training=True):
channels = self.channels
inp = tf.reshape(inp, [-1, self.img_size, self.img_size, channels])
hidden1 = conv_block(inp, weights['conv1'], weights['b1'], self.bn1, training = training)
hidden2 = conv_block(hidden1, weights['conv2'], weights['b2'], self.bn2, training = training)
hidden3 = conv_block(hidden2, weights['conv3'], weights['b3'], self.bn3, training = training)
hidden4 = conv_block(hidden3, weights['conv4'], weights['b4'], self.bn4, training = training)
hidden4 = tf.reduce_mean(input_tensor=hidden4, axis=[1, 2])
return tf.matmul(hidden4, weights['w5']) + weights['b5']
"""Data loading scripts"""
## NOTE: You do not need to modify this block but you will need to use it.
import numpy as np
import os
import random
import tensorflow as tf
from scipy import misc
import imageio
def get_images(paths, labels, n_samples=None, shuffle=True):
"""
Takes a set of character folders and labels and returns paths to image files
paired with labels.
Args:
paths: A list of character folders
labels: List or numpy array of same length as paths
n_samples: Number of images to retrieve per character
Returns:
List of (label, image_path) tuples
"""
if n_samples is not None:
sampler = lambda x: random.sample(x, n_samples)
else:
sampler = lambda x: x
images_labels = [(i, os.path.join(path, image))
for i, path in zip(labels, paths)
for image in sampler(os.listdir(path))]
if shuffle:
random.shuffle(images_labels)
return images_labels
def image_file_to_array(filename, dim_input):
"""
Takes an image path and returns numpy array
Args:
filename: Image filename
dim_input: Flattened shape of image
Returns:
1 channel image
"""
image = imageio.imread(filename)
image = image.reshape([dim_input])
image = image.astype(np.float32) / 255.0
image = 1.0 - image
return image
class DataGenerator(object):
"""
Data Generator capable of generating batches of Omniglot data.
A "class" is considered a class of omniglot digits.
"""
def __init__(self, num_classes, num_samples_per_class, num_meta_test_classes, num_meta_test_samples_per_class, config={}):
"""
Args:
num_classes: Number of classes for classification (K-way)
num_samples_per_class: num samples to generate per class in one batch
num_meta_test_classes: Number of classes for classification (K-way) at meta-test time
num_meta_test_samples_per_class: num samples to generate per class in one batch at meta-test time
batch_size: size of meta batch size (e.g. number of functions)
"""
self.num_samples_per_class = num_samples_per_class
self.num_classes = num_classes
self.num_meta_test_samples_per_class = num_meta_test_samples_per_class
self.num_meta_test_classes = num_meta_test_classes
data_folder = config.get('data_folder', './omniglot_resized')
self.img_size = config.get('img_size', (28, 28))
self.dim_input = np.prod(self.img_size)
self.dim_output = self.num_classes
character_folders = [os.path.join(data_folder, family, character)
for family in os.listdir(data_folder)
if os.path.isdir(os.path.join(data_folder, family))
for character in os.listdir(os.path.join(data_folder, family))
if os.path.isdir(os.path.join(data_folder, family, character))]
random.seed(123)
random.shuffle(character_folders)
num_val = 100
num_train = 1100
self.metatrain_character_folders = character_folders[: num_train]
self.metaval_character_folders = character_folders[
num_train:num_train + num_val]
self.metatest_character_folders = character_folders[
num_train + num_val:]
def sample_batch(self, batch_type, batch_size, shuffle=True, swap=False):
"""
Samples a batch for training, validation, or testing
Args:
batch_type: meta_train/meta_val/meta_test
shuffle: randomly shuffle classes or not
swap: swap number of classes (N) and number of samples per class (K) or not
Returns:
A a tuple of (1) Image batch and (2) Label batch where
image batch has shape [B, N, K, 784] and label batch has shape [B, N, K, N] if swap is False
where B is batch size, K is number of samples per class, N is number of classes
"""
if batch_type == "meta_train":
folders = self.metatrain_character_folders
num_classes = self.num_classes
num_samples_per_class = self.num_samples_per_class
elif batch_type == "meta_val":
folders = self.metaval_character_folders
num_classes = self.num_classes
num_samples_per_class = self.num_samples_per_class
else:
folders = self.metatest_character_folders
num_classes = self.num_meta_test_classes
num_samples_per_class = self.num_meta_test_samples_per_class
all_image_batches, all_label_batches = [], []
for i in range(batch_size):
sampled_character_folders = random.sample(
folders, num_classes)
labels_and_images = get_images(sampled_character_folders, range(
num_classes), n_samples=num_samples_per_class, shuffle=False)
labels = [li[0] for li in labels_and_images]
images = [image_file_to_array(
li[1], self.dim_input) for li in labels_and_images]
images = np.stack(images)
labels = np.array(labels).astype(np.int32)
labels = np.reshape(
labels, (num_classes, num_samples_per_class))
labels = np.eye(num_classes, dtype=np.float32)[labels]
images = np.reshape(
images, (num_classes, num_samples_per_class, -1))
batch = np.concatenate([labels, images], 2)
if shuffle:
for p in range(num_samples_per_class):
np.random.shuffle(batch[:, p])
labels = batch[:, :, :num_classes]
images = batch[:, :, num_classes:]
if swap:
labels = np.swapaxes(labels, 0, 1)
images = np.swapaxes(images, 0, 1)
all_image_batches.append(images)
all_label_batches.append(labels)
all_image_batches = np.stack(all_image_batches)
all_label_batches = np.stack(all_label_batches)
return all_image_batches, all_label_batches
```
# MAML (Training and Evaluation Code)
```
"""MAML model code"""
import numpy as np
import sys
import tensorflow as tf
from functools import partial
class MAML(tf.keras.Model):
def __init__(self, dim_input=1, dim_output=1,
num_inner_updates=1,
inner_update_lr=0.4, num_filters=32, k_shot=5, learn_inner_update_lr=False):
super(MAML, self).__init__()
self.dim_input = dim_input
self.dim_output = dim_output
self.inner_update_lr = inner_update_lr
self.loss_func = partial(cross_entropy_loss, k_shot=k_shot)
self.dim_hidden = num_filters
self.channels = 1
self.img_size = int(np.sqrt(self.dim_input/self.channels))
# outputs_ts[i] and losses_ts_post[i] are the output and loss after i+1 inner gradient updates
losses_tr_pre, outputs_tr, losses_ts_post, outputs_ts = [], [], [], []
accuracies_tr_pre, accuracies_ts = [], []
# for each loop in the inner training loop
outputs_ts = [[]]*num_inner_updates
losses_ts_post = [[]]*num_inner_updates
accuracies_ts = [[]]*num_inner_updates
# Define the weights - these should NOT be directly modified by the
# inner training loop
tf.random.set_seed(seed)
self.conv_layers = ConvLayers(self.channels, self.dim_hidden, self.dim_output, self.img_size)
self.learn_inner_update_lr = learn_inner_update_lr
if self.learn_inner_update_lr:
self.inner_update_lr_dict = {}
for key in self.conv_layers.conv_weights.keys():
self.inner_update_lr_dict[key] = [tf.Variable(self.inner_update_lr, name='inner_update_lr_%s_%d' % (key, j)) for j in range(num_inner_updates)]
def call(self, inp, meta_batch_size=25, num_inner_updates=1):
def task_inner_loop(inp, reuse=True,
meta_batch_size=25, num_inner_updates=1):
"""
Perform gradient descent for one task in the meta-batch (i.e. inner-loop).
Args:
inp: a tuple (input_tr, input_ts, label_tr, label_ts), where input_tr and label_tr are the inputs and
labels used for calculating inner loop gradients and input_ts and label_ts are the inputs and
labels used for evaluating the model after inner updates.
Should be shapes:
input_tr: [N*K, 784]
input_ts: [N*K, 784]
label_tr: [N*K, N]
label_ts: [N*K, N]
Returns:
task_output: a list of outputs, losses and accuracies at each inner update
"""
# the inner and outer loop data
input_tr, input_ts, label_tr, label_ts = inp
# weights corresponds to the initial weights in MAML (i.e. the meta-parameters)
weights = self.conv_layers.conv_weights
# the predicted outputs, loss values, and accuracy for the pre-update model (with the initial weights)
# evaluated on the inner loop training data
task_output_tr_pre, task_loss_tr_pre, task_accuracy_tr_pre = None, None, None
# lists to keep track of outputs, losses, and accuracies of test data for each inner_update
# where task_outputs_ts[i], task_losses_ts[i], task_accuracies_ts[i] are the output, loss, and accuracy
# after i+1 inner gradient updates
task_outputs_ts, task_losses_ts, task_accuracies_ts = [], [], []
#############################
#### YOUR CODE GOES HERE ####
# perform num_inner_updates to get modified weights
# modified weights should be used to evaluate performance
# Note that at each inner update, always use input_tr and label_tr for calculating gradients
# and use input_ts and labels for evaluating performance
# HINTS: You will need to use tf.GradientTape().
# Read through the tf.GradientTape() documentation to see how 'persistent' should be set.
# Here is some documentation that may be useful:
# https://www.tensorflow.org/guide/advanced_autodiff#higher-order_gradients
# https://www.tensorflow.org/api_docs/python/tf/GradientTape
with tf.GradientTape(persistent=True) as tape1:
task_output_tr_pre = self.conv_layers(input_tr, weights)
task_loss_tr_pre = self.loss_func(pred = task_output_tr_pre, label=label_tr)
grads = tape1.gradient(task_loss_tr_pre, list(weights.values()))
gradients = dict(zip(weights.keys(), grads))
if self.learn_inner_update_lr:
fast_weights = dict(zip(weights.keys(), [weights[key]-self.inner_update_lr_dict[key][0]*gradients[key] for key in weights.keys()]))
else:
fast_weights = dict(zip(weights.keys(), [weights[key] - self.inner_update_lr*gradients[key] for key in weights.keys()]))
for i in range(1, num_inner_updates):
test_output = self.conv_layers(input_ts, fast_weights, training=False)
task_outputs_ts.append(tf.stop_gradient(test_output))
ts_loss = self.loss_func(pred = tf.stop_gradient(test_output), label=label_ts)
task_losses_ts.append(ts_loss)
with tf.GradientTape(persistent=True) as tape2:
preds = self.conv_layers(input_tr, fast_weights)
loss = self.loss_func(pred = preds, label=label_tr)
grads = tape2.gradient(loss, list(fast_weights.values()))
gradients = dict(zip(fast_weights.keys(), grads))
if self.learn_inner_update_lr:
fast_weights = dict(zip(fast_weights.keys(), [fast_weights[key] - self.inner_update_lr_dict[key][i]*gradients[key] for key in fast_weights.keys()]))
else:
fast_weights = dict(zip(fast_weights.keys(), [fast_weights[key] - self.inner_update_lr*gradients[key] for key in fast_weights.keys()]))
test_output = self.conv_layers(input_ts, fast_weights, training=False)
task_outputs_ts.append(test_output)
ts_loss = self.loss_func(pred = test_output, label=label_ts)
task_losses_ts.append(ts_loss)
#############################
# Compute accuracies from output predictions
task_accuracy_tr_pre = accuracy(tf.argmax(input=label_tr, axis=1), tf.argmax(input=tf.nn.softmax(task_output_tr_pre), axis=1))
for j in range(num_inner_updates):
task_accuracies_ts.append(accuracy(tf.argmax(input=label_ts, axis=1), tf.argmax(input=tf.nn.softmax(task_outputs_ts[j]), axis=1)))
task_output = [task_output_tr_pre, task_outputs_ts, task_loss_tr_pre, task_losses_ts, task_accuracy_tr_pre, task_accuracies_ts]
return task_output
input_tr, input_ts, label_tr, label_ts = inp
# to initialize the batch norm vars, might want to combine this, and not run idx 0 twice.
unused = task_inner_loop((input_tr[0], input_ts[0], label_tr[0], label_ts[0]),
False,
meta_batch_size,
num_inner_updates)
out_dtype = [tf.float32, [tf.float32]*num_inner_updates, tf.float32, [tf.float32]*num_inner_updates]
out_dtype.extend([tf.float32, [tf.float32]*num_inner_updates])
task_inner_loop_partial = partial(task_inner_loop, meta_batch_size=meta_batch_size, num_inner_updates=num_inner_updates)
result = tf.map_fn(task_inner_loop_partial,
elems=(input_tr, input_ts, label_tr, label_ts),
dtype=out_dtype,
parallel_iterations=meta_batch_size)
return result
"""Model training code"""
"""
Usage Instructions:
5-way, 1-shot omniglot:
python main.py --meta_train_iterations=15000 --meta_batch_size=25 --k_shot=1 --inner_update_lr=0.4 --num_inner_updates=1 --logdir=logs/omniglot5way/
20-way, 1-shot omniglot:
python main.py --meta_train_iterations=15000 --meta_batch_size=16 --k_shot=1 --n_way=20 --inner_update_lr=0.1 --num_inner_updates=5 --logdir=logs/omniglot20way/
To run evaluation, use the '--meta_train=False' flag and the '--meta_test_set=True' flag to use the meta-test set.
"""
import csv
import numpy as np
import pickle
import random
import tensorflow as tf
from collections import OrderedDict
@tf.function
def outer_train_step(inp, model, optim, meta_batch_size=25, num_inner_updates=1):
with tf.GradientTape(persistent=False) as outer_tape:
result = model(inp, meta_batch_size=meta_batch_size, num_inner_updates=num_inner_updates)
outputs_tr, outputs_ts, losses_tr_pre, losses_ts, accuracies_tr_pre, accuracies_ts = result
total_losses_ts = [tf.reduce_mean(loss_ts) for loss_ts in losses_ts]
gradients = outer_tape.gradient(total_losses_ts[-1], model.trainable_variables)
optim.apply_gradients(zip(gradients, model.trainable_variables))
total_loss_tr_pre = tf.reduce_mean(losses_tr_pre)
total_accuracy_tr_pre = tf.reduce_mean(accuracies_tr_pre)
total_accuracies_ts = [tf.reduce_mean(accuracy_ts) for accuracy_ts in accuracies_ts]
return outputs_tr, outputs_ts, total_loss_tr_pre, total_losses_ts, total_accuracy_tr_pre, total_accuracies_ts
@tf.function
def outer_eval_step(inp, model, meta_batch_size=25, num_inner_updates=1):
result = model(inp, meta_batch_size=meta_batch_size, num_inner_updates=num_inner_updates)
outputs_tr, outputs_ts, losses_tr_pre, losses_ts, accuracies_tr_pre, accuracies_ts = result
total_loss_tr_pre = tf.reduce_mean(losses_tr_pre)
total_losses_ts = [tf.reduce_mean(loss_ts) for loss_ts in losses_ts]
total_accuracy_tr_pre = tf.reduce_mean(accuracies_tr_pre)
total_accuracies_ts = [tf.reduce_mean(accuracy_ts) for accuracy_ts in accuracies_ts]
return outputs_tr, outputs_ts, total_loss_tr_pre, total_losses_ts, total_accuracy_tr_pre, total_accuracies_ts
def meta_train_fn(model, exp_string, data_generator,
n_way=5, meta_train_iterations=15000, meta_batch_size=25,
log=True, logdir='/tmp/data', k_shot=1, num_inner_updates=1, meta_lr=0.001):
SUMMARY_INTERVAL = 10
SAVE_INTERVAL = 100
PRINT_INTERVAL = 10
TEST_PRINT_INTERVAL = PRINT_INTERVAL *5
pre_accuracies, post_accuracies = [], []
num_classes = data_generator.num_classes
optimizer = tf.keras.optimizers.Adam(learning_rate=meta_lr)
meta_val_accs = OrderedDict()
for itr in range(meta_train_iterations):
#############################
#### YOUR CODE GOES HERE ####
# sample a batch of training data and partition into
# the support/training set (input_tr, label_tr) and the query/test set (input_ts, label_ts)
# NOTE: The code assumes that the support and query sets have the same number of examples.
image_batches, label_batches = data_generator.sample_batch('meta_train', meta_batch_size)
input_tr = image_batches[:, :, :k_shot, :].reshape(meta_batch_size, n_way*k_shot, -1)
label_tr = label_batches[:, :, :k_shot, :].reshape(meta_batch_size, n_way*k_shot, -1)
input_ts = image_batches[:, :, k_shot:, :].reshape(meta_batch_size, n_way*k_shot, -1)
label_ts = label_batches[:, :, k_shot:, :].reshape(meta_batch_size, n_way*k_shot, -1)
#############################
inp = (input_tr, input_ts, label_tr, label_ts)
result = outer_train_step(inp, model, optimizer, meta_batch_size=meta_batch_size, num_inner_updates=num_inner_updates)
if (itr+1) % SUMMARY_INTERVAL == 0:
pre_accuracies.append(result[-2])
post_accuracies.append(result[-1][-1])
if (itr!=0) and (itr+1) % PRINT_INTERVAL == 0:
print_str = 'Iteration %d: pre-inner-loop train accuracy: %.5f, post-inner-loop test accuracy: %.5f' % (itr+1, np.mean(pre_accuracies), np.mean(post_accuracies))
print(print_str)
pre_accuracies, post_accuracies = [], []
if (itr!=0) and (itr+1) % TEST_PRINT_INTERVAL == 0:
#############################
#### YOUR CODE GOES HERE ####
# sample a batch of validation data and partition it into
# the support/training set (input_tr, label_tr) and the query/test set (input_ts, label_ts)
# NOTE: The code assumes that the support and query sets have the same number of examples.
image_batches, label_batches = data_generator.sample_batch('meta_val', meta_batch_size)
input_tr = image_batches[:, :, :k_shot, :].reshape(meta_batch_size, n_way*k_shot, -1)
label_tr = label_batches[:, :, :k_shot, :].reshape(meta_batch_size, n_way*k_shot, -1)
input_ts = image_batches[:, :, k_shot:, :].reshape(meta_batch_size, n_way*k_shot, -1)
label_ts = label_batches[:, :, k_shot:, :].reshape(meta_batch_size, n_way*k_shot, -1)
#############################
inp = (input_tr, input_ts, label_tr, label_ts)
result = outer_eval_step(inp, model, meta_batch_size=meta_batch_size, num_inner_updates=num_inner_updates)
print('Meta-validation pre-inner-loop train accuracy: %.5f, meta-validation post-inner-loop test accuracy: %.5f' % (result[-2], result[-1][-1]))
meta_val_accs[itr+1] = result[-1][-1].numpy()
model_file = logdir + exp_string + '/model' + str(itr)
print("Saving to ", model_file)
model.save_weights(model_file)
log_file = logdir + 'meta_train' + exp_string + '.pkl'
with open(log_file, 'wb') as f:
pickle.dump(meta_val_accs, f)
# calculated for omniglot
NUM_META_TEST_POINTS = 600
def meta_test_fn(model, data_generator, exp_string, n_way=5, meta_batch_size=25, k_shot=1,
num_inner_updates=1):
num_classes = data_generator.num_classes
np.random.seed(1)
random.seed(1)
meta_test_accuracies = []
for _ in range(NUM_META_TEST_POINTS):
#############################
#### YOUR CODE GOES HERE ####
# sample a batch of test data and partition it into
# the support/training set (input_tr, label_tr) and the query/test set (input_ts, label_ts)
# NOTE: The code assumes that the support and query sets have the same number of examples.
image_batches, label_batches = data_generator.sample_batch('meta_test', meta_batch_size)
input_tr = image_batches[:, :, :k_shot, :].reshape(meta_batch_size, n_way*k_shot, -1)
label_tr = label_batches[:, :, :k_shot, :].reshape(meta_batch_size, n_way*k_shot, -1)
input_ts = image_batches[:, :, k_shot:, :].reshape(meta_batch_size, n_way*k_shot, -1)
label_ts = label_batches[:, :, k_shot:, :].reshape(meta_batch_size, n_way*k_shot, -1)
#############################
inp = (input_tr, input_ts, label_tr, label_ts)
result = outer_eval_step(inp, model, meta_batch_size=meta_batch_size, num_inner_updates=num_inner_updates)
meta_test_accuracies.append(result[-1][-1])
meta_test_accuracies = np.array(meta_test_accuracies)
means = np.mean(meta_test_accuracies)
stds = np.std(meta_test_accuracies)
ci95 = 1.96*stds/np.sqrt(NUM_META_TEST_POINTS)
print('Mean meta-test accuracy/loss, stddev, and confidence intervals')
print((means, stds, ci95))
meta_test_results = {'meta_test_acc': means, 'meta_test_acc_std': stds,
'CI95': ci95, 'num_meta_test_points': NUM_META_TEST_POINTS}
log_file = logdir + 'meta_test' + exp_string + '.pkl'
with open(log_file, 'wb') as f:
pickle.dump(meta_test_results, f)
def run_maml(n_way=5, k_shot=1, meta_batch_size=25, meta_lr=0.001,
inner_update_lr=0.4, num_filters=32, num_inner_updates=1,
learn_inner_update_lr=False,
resume=False, resume_itr=0, log=True, logdir='/tmp/data/',
data_path='./omniglot_resized',meta_train=True,
meta_train_iterations=15000, meta_train_k_shot=-1,
meta_train_inner_update_lr=-1):
# call data_generator and get data with k_shot*2 samples per class
data_generator = DataGenerator(n_way, k_shot*2, n_way, k_shot*2, config={'data_folder': data_path})
# set up MAML model
dim_output = data_generator.dim_output
dim_input = data_generator.dim_input
model = MAML(dim_input,
dim_output,
num_inner_updates=num_inner_updates,
inner_update_lr=inner_update_lr,
k_shot=k_shot,
num_filters=num_filters,
learn_inner_update_lr=learn_inner_update_lr)
if meta_train_k_shot == -1:
meta_train_k_shot = k_shot
if meta_train_inner_update_lr == -1:
meta_train_inner_update_lr = inner_update_lr
exp_string = '.n_way_'+str(n_way)+'.mbs_'+str(meta_batch_size) + '.k_shot_' + str(meta_train_k_shot) + '.inner_numstep_' + str(num_inner_updates) + '.inner_updatelr_' + str(meta_train_inner_update_lr) + '.learn_inner_update_lr_' + str(learn_inner_update_lr)
if meta_train:
meta_train_fn(model, exp_string, data_generator,
n_way, meta_train_iterations, meta_batch_size, log, logdir,
k_shot, num_inner_updates, meta_lr)
print("Evaluating trained MAML on held-out test-classes")
meta_test_fn(model, data_generator, exp_string, n_way=n_way,
meta_batch_size=1,k_shot=k_shot,
num_inner_updates=num_inner_updates)
else:
meta_batch_size = 1
model_file = tf.train.latest_checkpoint(logdir + exp_string)
print("Restoring model weights from ", model_file)
model.load_weights(model_file)
if meta_train_k_shot != k_shot:
exp_string = '.n_way_'+str(n_way)+'.mbs_'+str(meta_batch_size) + '.k_shot_' + str(k_shot) + '.inner_numstep_' + str(num_inner_updates) + '.inner_updatelr_' + str(meta_train_inner_update_lr) + '.learn_inner_update_lr_' + str(learn_inner_update_lr)
meta_test_fn(model, data_generator, exp_string, n_way, meta_batch_size, k_shot, num_inner_updates)
logdir = '/content/drive/MyDrive/CS330-MetaLearning/Hw2/maml/'
run_maml(n_way=5, k_shot=1, inner_update_lr= 4.0, num_inner_updates=1, learn_inner_update_lr=True, logdir=logdir)
```
#Plotting Utility Functions
```
def load_metrics(pkl_file):
with open(pkl_file, 'rb') as f:
metrics = pickle.load(f)
return metrics
import matplotlib.pyplot as plt
import seaborn as sns
def plot_metrics(ax, x, y, stds=None, label=None, marker=None):
"""
Plots y vs x normal and fill_between type for the ax
"""
x = np.array(x)
y = np.array(y)
if marker is not None:
#ax.errorbar(x, y, yerr)
ax.plot(x, y, label=label, marker=marker)
else:
ax.plot(x, y, label=label)
if stds is not None:
ax.fill_between(x, y-stds, y+stds, alpha=0.2, facecolor=c)
return ax
def plot_results(hp_values, hp_x, hp_y, label_strings, x_label, y_label, title, save_path, marker=None):
"""
Plots results of metrics with respect to hyperparameters
Args:
hp_values : list of hyperparameter values for which plot is drawn
hp_x : x_axis values
hp_y: stacked array of y_axis values for each value in hp_values
label_stings: hyperparameter strings considered for plotting
x_label: label for x_axis
y_label: label for y-axis
title : plot title
save_path: location to save the plot
Returns:
Saves plot at specified 'save_path' location
"""
sns.set(style='darkgrid')
fig, ax = plt.subplots(figsize = (10,7))
for i, value in enumerate(hp_values):
ax = plot_metrics(ax, hp_x, hp_y[i]*100, label= label_strings[i] +f'{value}', marker=marker)
ax.set_xlabel(x_label)
ax.set_ylabel(y_label)
#ax.set_xticks(iterations)
#ax.set_yticks(np.arange(0.1, 1.1, 0.05))
ax.legend(loc='best')
ax.set_title(title)
fig.savefig(save_path)
```
# Plot Results For MAML
Plotting Meta Validation Accuracies for MAML training with respect to learning rates [0.04, 0.4, 4.0]
```
logdir = '/content/drive/MyDrive/CS330-MetaLearning/Hw2/maml/'
log_file1 = logdir + 'meta_train.n_way_5.mbs_25.k_shot_1.inner_numstep_1.inner_updatelr_0.04.learn_inner_update_lr_False' + '.pkl'
metrics1 = load_metrics(log_file1)
print(metrics1)
logdir = '/content/drive/MyDrive/CS330-MetaLearning/Hw2/maml/'
learning_rates=[0.04, 0.4, 4.0]
maml_metrics = []
for lr in learning_rates:
file_name = f'meta_train.n_way_5.mbs_25.k_shot_1.inner_numstep_1.inner_updatelr_{lr}.learn_inner_update_lr_False.pkl'
metrics = load_metrics(logdir +file_name)
meta_train_iterations = list(metrics.keys())
maml_metrics.append(list(metrics.values()))
y_stacked = np.stack(maml_metrics, axis=0)
x_label = 'Number of Meta-Training Iterations'
y_label = 'Meta-Validation Accuracies'
title = 'MAML 5-way 1-shot validation accuracies'
hp_values= learning_rates
label_strings = ['lr']*3
save_path = logdir + 'maml-meta-val-plot-manual-lr.jpg'
plot_results(hp_values, meta_train_iterations, y_stacked, label_strings, x_label, y_label, title, save_path)
```
# Inner Loop Learning Rate Plot
Let's now compare MAML training with fixed learning rate in inner loop vs learning learning rate in inner loop
```
logdir = '/content/drive/MyDrive/CS330-MetaLearning/Hw2/maml/'
learning_rates=[0.04, 0.4, 4.0]
fixed_inner_lr = []
for lr in learning_rates:
file_name = f'meta_train.n_way_5.mbs_25.k_shot_1.inner_numstep_1.inner_updatelr_{lr}.learn_inner_update_lr_False.pkl'
metrics = load_metrics(logdir +file_name)
meta_train_iterations = list(metrics.keys())
fixed_inner_lr.append(list(metrics.values()))
learned_inner_lr = []
for lr in learning_rates:
file_name = f'meta_train.n_way_5.mbs_25.k_shot_1.inner_numstep_1.inner_updatelr_{lr}.learn_inner_update_lr_True.pkl'
metrics = load_metrics(logdir +file_name)
meta_train_iterations = list(metrics.keys())
learned_inner_lr.append(list(metrics.values()))
y_stacked = np.stack(fixed_inner_lr+learned_inner_lr, axis=0)
x_label = 'Number of Meta-Training Iterations'
y_label = 'Meta-Validation Accuracies'
title = 'MAML 5-way 1-shot fixed inner-loop lr vs learning inner-loop lr for each variable'
hp_values= learning_rates*2
label_strings = ['fixed_lr']*3 +['learning_lr_with_initial_lr'] *3
save_path = logdir + 'maml-meta-val-plot-manual-lr-vs-learned-lr.jpg'
plot_results(hp_values, meta_train_iterations, y_stacked, label_strings, x_label, y_label, title, save_path)
```
# Prototypical Networks (Training and Evaluation Code)
```
# models/ProtoNet
import numpy as np
import tensorflow as tf
from tensorflow.keras import layers
class ProtoNet(tf.keras.Model):
def __init__(self, num_filters, latent_dim):
super(ProtoNet, self).__init__()
self.num_filters = num_filters
self.latent_dim = latent_dim
num_filter_list = self.num_filters + [latent_dim]
self.convs = []
for i, num_filter in enumerate(num_filter_list):
block_parts = [
layers.Conv2D(
filters=num_filter,
kernel_size=3,
padding='SAME',
activation='linear'),
]
block_parts += [layers.BatchNormalization()]
block_parts += [layers.Activation('relu')]
block_parts += [layers.MaxPool2D()]
block = tf.keras.Sequential(block_parts, name='conv_block_%d' % i)
self.__setattr__("conv%d" % i, block)
self.convs.append(block)
self.flatten = tf.keras.layers.Flatten()
def call(self, inp):
out = inp
for conv in self.convs:
out = conv(out)
out = self.flatten(out)
return out
def ProtoLoss(x_latent, q_latent, labels_onehot, num_classes, num_support, num_queries):
"""
calculates the prototype network loss using the latent representation of x
and the latent representation of the query set
Args:
x_latent: latent representation of supports with shape [N*S, D], where D is the latent dimension
q_latent: latent representation of queries with shape [N*Q, D], where D is the latent dimension
labels_onehot: one-hot encodings of the labels of the queries with shape [N, Q, N]
num_classes: number of classes (N) for classification
num_support: number of examples (S) in the support set
num_queries: number of examples (Q) in the query set
Returns:
ce_loss: the cross entropy loss between the predicted labels and true labels
acc: the accuracy of classification on the queries
"""
#############################
#### YOUR CODE GOES HERE ####
# compute the prototypes
# compute the distance from the prototypes
# compute cross entropy loss
# note - additional steps are needed!
# return the cross-entropy loss and accuracy
prototypes = tf.reduce_mean(tf.reshape(x_latent,[num_classes, num_support, -1]), axis=1)
tiled_proto = tf.tile(tf.expand_dims(prototypes, axis=0), (num_classes*num_queries, 1, 1))
tiled_queries = tf.tile(tf.expand_dims(q_latent, axis=1), (1, num_classes, 1))
distances = tf.reduce_mean(tf.square(tiled_proto - tiled_queries), axis=2)
log_probs = tf.reshape(tf.nn.log_softmax(-distances), [num_classes, num_queries, -1])
ce_loss = -tf.reduce_mean(tf.reshape(tf.reduce_sum(tf.multiply(labels_onehot, log_probs), axis=-1), [-1]))
acc = tf.reduce_mean(tf.cast(tf.equal(tf.argmax(log_probs, axis=-1), tf.argmax(labels_onehot, axis=-1)), tf.float32))
#############################
return ce_loss, acc
# run_ProtoNet
from PIL import Image
import numpy as np
import tensorflow as tf
from tensorflow.keras import layers
import os
import glob
import matplotlib.pyplot as plt
@tf.function
def proto_net_train_step(model, optim, x, q, labels_ph):
num_classes, num_support, im_height, im_width, channels = x.shape
num_queries = q.shape[1]
x = tf.reshape(x, [-1, im_height, im_width, channels])
q = tf.reshape(q, [-1, im_height, im_width, channels])
with tf.GradientTape() as tape:
x_latent = model(x)
q_latent = model(q)
ce_loss, acc = ProtoLoss(x_latent, q_latent, labels_ph, num_classes, num_support, num_queries)
gradients = tape.gradient(ce_loss, model.trainable_variables)
optim.apply_gradients(zip(gradients, model.trainable_variables))
return ce_loss, acc
@tf.function
def proto_net_eval(model, x, q, labels_ph):
num_classes, num_support, im_height, im_width, channels = x.shape
num_queries = q.shape[1]
x = tf.reshape(x, [-1, im_height, im_width, channels])
q = tf.reshape(q, [-1, im_height, im_width, channels])
x_latent = model(x)
q_latent = model(q)
ce_loss, acc = ProtoLoss(x_latent, q_latent, labels_ph, num_classes, num_support, num_queries)
return ce_loss, acc
def run_protonet(data_path='./omniglot_resized', logdir= '/tmp/data/',n_way=20,
k_shot=1, n_query=5, n_meta_test_way=20, k_meta_test_shot=5, n_meta_test_query=5):
n_epochs = 20
n_episodes = 100
im_width, im_height, channels = 28, 28, 1
num_filters = 32
latent_dim = 16
num_conv_layers = 3
n_meta_test_episodes = 1000
model = ProtoNet([num_filters]*num_conv_layers, latent_dim)
optimizer = tf.keras.optimizers.Adam(learning_rate=0.001)
# call DataGenerator with k_shot+n_query samples per class
data_generator = DataGenerator(n_way, k_shot+n_query, n_meta_test_way, k_meta_test_shot+n_meta_test_query)
exp_string = '.n_way_'+str(n_way) + '.k_shot_' + str(k_shot) + '.n_query_' + str(n_query) + '.n_meta_test_way_' + str(n_meta_test_way) + '.k_meta_test_shot_' + str(k_meta_test_shot) + '.n_meta_test_query_' + str(n_meta_test_query)
val_accs = OrderedDict()
for ep in range(n_epochs):
for epi in range(n_episodes):
#############################
#### YOUR CODE GOES HERE ####
# sample a batch of training data and partition it into
# support and query sets
image_batches, label_batches = data_generator.sample_batch('meta_train', 1, shuffle=False)
support = image_batches[:, :, :k_shot, :].reshape(n_way, k_shot, im_height, im_width, channels)
query = image_batches[:, :, k_shot:, :].reshape(n_way, n_query, im_height, im_width, channels)
labels = label_batches[:, :, k_shot:, :].reshape(n_way, n_query, n_way)
#############################
ls, ac = proto_net_train_step(model, optimizer, x=support, q=query, labels_ph=labels)
if (epi+1) % 50 == 0:
#############################
#### YOUR CODE GOES HERE ####
# sample a batch of validation data and partition it into
# support and query sets
image_batches, label_batches = data_generator.sample_batch('meta_val', 1, shuffle=False)
support = image_batches[:, :, :k_shot, :].reshape(n_way, k_shot, im_height, im_width, channels)
query = image_batches[:, :, k_shot:, :].reshape(n_way, n_query, im_height, im_width, channels)
labels = label_batches[:, :, k_shot:, :].reshape(n_way, n_query, n_way)
#############################
val_ls, val_ac = proto_net_eval(model, x=support, q=query, labels_ph=labels)
print('[epoch {}/{}, episode {}/{}] => meta-training loss: {:.5f}, meta-training acc: {:.5f}, meta-val loss: {:.5f}, meta-val acc: {:.5f}'.format(ep+1,
n_epochs,
epi+1,
n_episodes,
ls,
ac,
val_ls,
val_ac))
val_accs[(ep * n_episodes) + (epi+1)] = val_ac.numpy()
log_file = logdir + 'meta_train' + exp_string + '.pkl'
with open(log_file, 'wb') as f:
pickle.dump(val_accs, f)
print('Testing...')
meta_test_accuracies = []
for epi in range(n_meta_test_episodes):
#############################
#### YOUR CODE GOES HERE ####
# sample a batch of test data and partition it into
# support and query sets
image_batches, label_batches = data_generator.sample_batch('meta_test', 1, shuffle=False)
support = image_batches[:, :, :k_meta_test_shot, :].reshape(n_meta_test_way, k_meta_test_shot, im_height, im_width, channels)
query = image_batches[:, :, k_meta_test_shot:, :].reshape(n_meta_test_way, n_meta_test_query, im_height, im_width, channels)
labels = label_batches[:, :, k_meta_test_shot:, :].reshape(n_meta_test_way, n_meta_test_query, n_meta_test_way)
#############################
ls, ac = proto_net_eval(model, x=support, q=query, labels_ph=labels)
meta_test_accuracies.append(ac)
if (epi+1) % 50 == 0:
print('[meta-test episode {}/{}] => loss: {:.5f}, acc: {:.5f}'.format(epi+1, n_meta_test_episodes, ls, ac))
avg_acc = np.mean(meta_test_accuracies)
stds = np.std(meta_test_accuracies)
print('Average Meta-Test Accuracy: {:.5f}, Meta-Test Accuracy Std: {:.5f}'.format(avg_acc, stds))
meta_test_results = {'meta_test_acc': avg_acc, 'meta_test_acc_std': stds,
'num_meta_episodes':n_meta_test_episodes}
log_file = logdir + 'meta_test' + exp_string + '.pkl'
with open(log_file, 'wb') as f:
pickle.dump(meta_test_results, f)
logdir = '/content/drive/MyDrive/CS330-MetaLearning/Hw2/proto_nets/'
run_protonet('./omniglot_resized/', logdir = logdir, n_way=5, k_shot=1, n_query=5, n_meta_test_way=5, k_meta_test_shot=10, n_meta_test_query=10)
```
# Plot Results for ProtoNets
```
logdir = '/content/drive/MyDrive/CS330-MetaLearning/Hw2/proto_nets/'
log_file = logdir + 'meta_train.n_way_5.k_shot_1.n_query_5.n_meta_test_way_5.k_meta_test_shot_4.n_meta_test_query_4' + '.pkl'
metrics1 = load_metrics(log_file1)
print(metrics1)
meta_train_iterations = list(metrics1.keys())
y_1 = list(metrics1.values())
y_stacked = np.stack([y_1], axis=0)
x_label = 'Number of Meta-Training Iterations'
y_label = 'Meta-Validation Accuracies'
title = 'ProtoNet 5-way 1-shot validation accuracies'
hp_values=['0.001']
label_strings = ['lr']
save_path = logdir + 'protonet-meta-val-plot-manual-lr.jpg'
plot_results(hp_values, meta_train_iterations, y_stacked, label_strings, x_label, y_label, title, save_path)
```
# Comparision MAML vs ProtoNet
MAMl vs ProtoNet meta-test accuracy comparision;
Both the algorithms are trained with 5-way 1-shot setting at meta-train time however at meta-test time comparision is made with varying k-shot settings with k= [4, 6, 8, 10]
```
logdir = '/content/drive/MyDrive/CS330-MetaLearning/Hw2/maml/'
run_maml(n_way=5, k_shot=10, inner_update_lr= 0.04, num_inner_updates=1, logdir=logdir, learn_inner_update_lr=True, meta_train=False, meta_train_k_shot=1)
# MAML Meta-Test Metrics
logdir = '/content/drive/MyDrive/CS330-MetaLearning/Hw2/maml/'
logfile1 = logdir + 'meta_test.n_way_5.mbs_1.k_shot_4.inner_numstep_1.inner_updatelr_0.04.learn_inner_update_lr_True' +'.pkl'
metrics1 = load_metrics(logfile1)
# ProtoNet Meta-Test Metrics
logdir = '/content/drive/MyDrive/CS330-MetaLearning/Hw2/proto_nets/'
logfile2 = logdir + 'meta_test.n_way_5.k_shot_1.n_query_5.n_meta_test_way_5.k_meta_test_shot_4.n_meta_test_query_4' +'.pkl'
metrics2 = load_metrics(logfile2)
print(metrics1)
print(metrics2)
K_shots= [4, 6, 8, 10]
logdir = '/content/drive/MyDrive/CS330-MetaLearning/Hw2/maml/'
maml_accuracies = []
for k in K_shots:
file_name = f'meta_test.n_way_5.mbs_1.k_shot_{k}.inner_numstep_1.inner_updatelr_0.04.learn_inner_update_lr_True.pkl'
metrics = load_metrics(logdir +file_name)
maml_accuracies.append(metrics['meta_test_acc'])
logdir = '/content/drive/MyDrive/CS330-MetaLearning/Hw2/proto_nets/'
proto_accuracies=[]
for k in K_shots:
file_name = f'meta_test.n_way_5.k_shot_1.n_query_5.n_meta_test_way_5.k_meta_test_shot_{k}.n_meta_test_query_{k}.pkl'
metrics = load_metrics(logdir +file_name)
proto_accuracies.append(metrics['meta_test_acc'])
y_stacked= np.stack([maml_accuracies, proto_accuracies], axis=0)
x_label = 'K-shot at Meta-Test time'
y_label = 'Meta-Test Accuracies'
title = '5-way 1-shot Meta-test accs with varying k-shot at meta-test time'
hp_values= ['', '']
label_strings = ['maml', 'proto-net']
save_path = logdir + 'maml-vs-proto.jpg'
plot_results(hp_values, K_shots, y_stacked, label_strings, x_label, y_label, title, save_path, marker='o')
```
| github_jupyter |
# Illustration of Various Kernels
---------------------------------
This function wll illustrate how to implement various kernels in TensorFlow.
Linear Kernel:
K(x1, x2) = t(x1) * x2
Gaussian Kernel (RBF):
K(x1, x2) = exp(-gamma * abs(x1 - x2)^2)
We start by loading the necessary libraries
```
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
from sklearn import datasets
from tensorflow.python.framework import ops
ops.reset_default_graph()
```
Start a computational graph session:
```
sess = tf.Session()
```
For this example, we will generate fake non-linear data. The data we will generate is concentric ring data.
```
# Generate non-lnear data
(x_vals, y_vals) = datasets.make_circles(n_samples=350, factor=.5, noise=.1)
y_vals = np.array([1 if y==1 else -1 for y in y_vals])
class1_x = [x[0] for i,x in enumerate(x_vals) if y_vals[i]==1]
class1_y = [x[1] for i,x in enumerate(x_vals) if y_vals[i]==1]
class2_x = [x[0] for i,x in enumerate(x_vals) if y_vals[i]==-1]
class2_y = [x[1] for i,x in enumerate(x_vals) if y_vals[i]==-1]
```
We declare the batch size (large for SVMs), create the placeholders, and declare the $b$ variable for the SVM model.
```
# Declare batch size
batch_size = 350
# Initialize placeholders
x_data = tf.placeholder(shape=[None, 2], dtype=tf.float32)
y_target = tf.placeholder(shape=[None, 1], dtype=tf.float32)
prediction_grid = tf.placeholder(shape=[None, 2], dtype=tf.float32)
# Create variables for svm
b = tf.Variable(tf.random_normal(shape=[1,batch_size]))
```
Here we will apply the kernel. Note that the `Linear Kernel` is commented out. If you choose to use the linear kernel, then uncomment the linear `my_kernel` variable, and comment out the five RBF kernel lines.
```
# Apply kernel
# Linear Kernel
# my_kernel = tf.matmul(x_data, tf.transpose(x_data))
# Gaussian (RBF) kernel
gamma = tf.constant(-50.0)
dist = tf.reduce_sum(tf.square(x_data), 1)
dist = tf.reshape(dist, [-1,1])
sq_dists = tf.add(tf.subtract(dist, tf.multiply(2., tf.matmul(x_data, tf.transpose(x_data)))), tf.transpose(dist))
my_kernel = tf.exp(tf.multiply(gamma, tf.abs(sq_dists)))
```
Next we compute the SVM model and create a loss function.
```
# Compute SVM Model
first_term = tf.reduce_sum(b)
b_vec_cross = tf.matmul(tf.transpose(b), b)
y_target_cross = tf.matmul(y_target, tf.transpose(y_target))
second_term = tf.reduce_sum(tf.multiply(my_kernel, tf.multiply(b_vec_cross, y_target_cross)))
loss = tf.negative(tf.subtract(first_term, second_term))
```
Just like we created the kernel for the training points, we need to create the kernel for the test/prediction points.
Again, comment/uncomment the appropriate lines for using the linear or RBF kernel.
```
# Create Prediction Kernel
# Linear prediction kernel
# my_kernel = tf.matmul(x_data, tf.transpose(prediction_grid))
# Gaussian (RBF) prediction kernel
rA = tf.reshape(tf.reduce_sum(tf.square(x_data), 1),[-1,1])
rB = tf.reshape(tf.reduce_sum(tf.square(prediction_grid), 1),[-1,1])
pred_sq_dist = tf.add(tf.subtract(rA, tf.multiply(2., tf.matmul(x_data, tf.transpose(prediction_grid)))), tf.transpose(rB))
pred_kernel = tf.exp(tf.multiply(gamma, tf.abs(pred_sq_dist)))
```
In order to use the kernel to classify points, we create a prediction operation. This prediction operation will be the sign ( positive or negative ) of the model outputs. The accuracy can then be computed if we know the actual target labels.
```
prediction_output = tf.matmul(tf.multiply(tf.transpose(y_target),b), pred_kernel)
prediction = tf.sign(prediction_output-tf.reduce_mean(prediction_output))
accuracy = tf.reduce_mean(tf.cast(tf.equal(tf.squeeze(prediction), tf.squeeze(y_target)), tf.float32))
```
We now declare the optimizer and variable initialization operations.
```
# Declare optimizer
my_opt = tf.train.GradientDescentOptimizer(0.002)
train_step = my_opt.minimize(loss)
# Initialize variables
init = tf.global_variables_initializer()
sess.run(init)
```
We start the training loop for the SVM. We will randomly choose a batch of points and run the train step. Then we calculate the loss and accuracy.
```
# Training loop
loss_vec = []
batch_accuracy = []
for i in range(1000):
rand_index = np.random.choice(len(x_vals), size=batch_size)
rand_x = x_vals[rand_index]
rand_y = np.transpose([y_vals[rand_index]])
sess.run(train_step, feed_dict={x_data: rand_x, y_target: rand_y})
temp_loss = sess.run(loss, feed_dict={x_data: rand_x, y_target: rand_y})
loss_vec.append(temp_loss)
acc_temp = sess.run(accuracy, feed_dict={x_data: rand_x,
y_target: rand_y,
prediction_grid:rand_x})
batch_accuracy.append(acc_temp)
if (i+1)%250==0:
print('Step #' + str(i+1))
print('Loss = ' + str(temp_loss))
```
To plot a pretty picture of the regions we fit, we create a fine mesh to run through our model and get the predictions. (This is very similar to the SVM plotting code from sci-kit learn).
```
# Create a mesh to plot points in
x_min, x_max = x_vals[:, 0].min() - 1, x_vals[:, 0].max() + 1
y_min, y_max = x_vals[:, 1].min() - 1, x_vals[:, 1].max() + 1
xx, yy = np.meshgrid(np.arange(x_min, x_max, 0.02),
np.arange(y_min, y_max, 0.02))
grid_points = np.c_[xx.ravel(), yy.ravel()]
[grid_predictions] = sess.run(prediction, feed_dict={x_data: rand_x,
y_target: rand_y,
prediction_grid: grid_points})
grid_predictions = grid_predictions.reshape(xx.shape)
```
Plot the results
```
# Plot points and grid
plt.contourf(xx, yy, grid_predictions, cmap=plt.cm.Paired, alpha=0.8)
plt.plot(class1_x, class1_y, 'ro', label='Class 1')
plt.plot(class2_x, class2_y, 'kx', label='Class -1')
plt.title('Gaussian SVM Results')
plt.xlabel('x')
plt.ylabel('y')
plt.legend(loc='lower right')
plt.ylim([-1.5, 1.5])
plt.xlim([-1.5, 1.5])
plt.show()
# Plot batch accuracy
plt.plot(batch_accuracy, 'k-', label='Accuracy')
plt.title('Batch Accuracy')
plt.xlabel('Generation')
plt.ylabel('Accuracy')
plt.legend(loc='lower right')
plt.show()
# Plot loss over time
plt.plot(loss_vec, 'k-')
plt.title('Loss per Generation')
plt.xlabel('Generation')
plt.ylabel('Loss')
plt.show()
```
| github_jupyter |
Versjon 08.01.2020
# Introduksjon
Denne notebooken er ment som en relativt enkel illustrasjon av analyse av sensordata med maskinlæring. Se slides fra introduksjonen for motivasjon.
# Setup
```
%matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import sklearn
from pathlib import Path
import subprocess
```
# Data
Vi bruker et sensor-datasett fra UCI Machine Learning, tilgjenglig via Kaggle: https://www.kaggle.com/uciml/human-activity-recognition-with-smartphones. Fra 30 personer ble det samlet målinger fra en smart-telefon mens de utførte dagligdagse oppgaver. Vår oppgave er å predikere hvilken oppgave som ble utført direkte fra sensor-målingene.
Video som beskriver innsamlingen av data: https://www.youtube.com/watch?v=XOEN9W05_4A
```
import IPython
IPython.display.IFrame(width="560", height="315", src="https://www.youtube.com/embed/XOEN9W05_4A")
```
Her er en beskrivelse av datasettet, sakset fra Kaggle:
The Human Activity Recognition database was built from the recordings of 30 study participants performing activities of daily living (ADL) while carrying a waist-mounted smartphone with embedded inertial sensors. The objective is to classify activities into one of the six activities performed.
> **Description of experiment**<br>
The experiments have been carried out with a group of 30 volunteers within an age bracket of 19-48 years. Each person performed six activities (WALKING, WALKING_UPSTAIRS, WALKING_DOWNSTAIRS, SITTING, STANDING, LAYING) wearing a smartphone (Samsung Galaxy S II) on the waist. Using its embedded accelerometer and gyroscope, we captured 3-axial linear acceleration and 3-axial angular velocity at a constant rate of 50Hz. The experiments have been video-recorded to label the data manually. The obtained dataset has been randomly partitioned into two sets, where **70% of the volunteers was selected for generating the training data and 30% the test data**.
> The sensor signals (accelerometer and gyroscope) were pre-processed by applying noise filters and then sampled in fixed-width sliding windows of 2.56 sec and 50% overlap (128 readings/window). The sensor acceleration signal, which has gravitational and body motion components, was separated using a Butterworth low-pass filter into body acceleration and gravity. The gravitational force is assumed to have only low frequency components, therefore a filter with 0.3 Hz cutoff frequency was used. From each window, a vector of features was obtained by calculating variables from the time and frequency domain.
> **Attribute information**<br>
> For each record in the dataset the following is provided:
> * Triaxial acceleration from the accelerometer (total acceleration) and the estimated body acceleration.
* Triaxial Angular velocity from the gyroscope.
* A 561-feature vector with time and frequency domain variables.
* Its activity label.
* An identifier of the subject who carried out the experiment.
## Last inn og utforsk data
Vi har allerede hentet data fra Kaggle. Plassert i katalogen `../data`
```
DATA = Path('../data/sensor')
train = pd.read_csv(DATA/'train.csv')
test = pd.read_csv(DATA/'test.csv')
```
Vi har fått to dataframes bestående av en lang rekke sensormålinger, markert med tilhørende aktiviteter:
```
train.info()
test.info()
```
Vi ser at det er 7352 treningsdata og 2947 testdata.
Hvordan ser data ut?
```
# For å vise alle søylene i data frames:
pd.set_option('display.max_columns', 600)
train.head()
test.head()
```
Her er noen labels:
```
np.random.choice(train['Activity'], size=50)
```
De seks ulike aktivitetene vi skal detektere er:
```
np.unique(train['Activity'])
```
Fordelingen av disse i treningsdata er:
```
train['Activity'].value_counts().plot(kind='bar')
plt.show()
```
## Ekstra: korrelasjoner
Det er helt sikkert stor korrelasjon mellom mange av features i dette datasettet (akselerasjon og gyroskop-features, for eksempel). Vi kan avdekke dette ved å bruke korrelasjonsmatrisen, og så trekke ut egenskapene som er mest korrelert:
```
correlation_matrix = train.corr()
correlation_matrix.info()
```
Korrelasjonsmatrisen er en 562x562-matrise (alle numeriske features korrelert med alle numeriske features). Her er de føste 10 søyler og 10 rader:
```
correlation_matrix.iloc[0:10, 0:10]
```
Vi ønsker å plukke ut parene av features som har høyest korrelasjon. Vi kan gjøre dette med `unstack`, som gjør alle verdiene i søylen helt til venstre (index-søylen) til søyler:
```
correlation_matrix.unstack().shape
correlation_matrix.unstack()[:5]
```
Vi får altså 315844 entries
```
562*562
```
Nå kan vi plukke ut de 15 minste og største tallene:
```
correlation_matrix.unstack().drop_duplicates().sort_values()[:15]
correlation_matrix.unstack().drop_duplicates().sort_values()[-15:]
```
Som vi trodde: det er mange features som er veldig høyt korrelert.
> En kan (ofte med fordel) fjerne features som er veldig høyt korrelert fra data. Det kan øke ytelsen til modellene. Forsøk gjerne dette her!
# Splitt opp data
Vi deler opp data i input X og output y:
```
X_train = train.drop('Activity', axis=1)
y_train = train['Activity']
X_test = test.drop('Activity', axis=1)
y_test = test['Activity']
```
# Modell
Vi bruker vår venn `RandomForestClassifier`.
> Senere skal du få vite nøyaktig hvordan denne fungerer!
```
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier(n_estimators=100, n_jobs=-1, random_state=42)
rf.fit(X_train, y_train)
```
Hvor bra accuracy får vi?
```
y_pred = rf.predict(X_test)
from sklearn.metrics import accuracy_score
accuracy_score(y_test, y_pred)
```
Over 92%!
Er dette bra? For å svare på det må vi bruke verktøyene vi har lært om for evaluering av klassifikatorer:
# Evaluer resultatet
## Forvirringsmatrise
```
from utils import plot_confusion_matrix, plot_confusion_matrix_with_colorbar
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
plot_confusion_matrix_with_colorbar(cm, classes=np.unique(y_test), figsize=(10,10))
```
## Feature importance
Her er de 10 features som ble vektet høyest av vår modell:
```
importances = rf.feature_importances_
# Find index of those with highest importance, sorted from largest to smallest:
indices = np.argsort(importances)[::-1]
for f in range(10):
print(f'{X_test.columns[indices[f]]}: {np.round(importances[indices[f]],2)}')
```
## Permutation importance
Som kjent er feature importance for random forests en veldig ustabil og lite gunstig måte å måle faktisk viktighet av features (ta en titt tilbake på Lab 0 for mer om dette). Permutation importance gir mer stabile, data-drevne estimat av feature importance.
**Advarsel:** dette tar litt tid siden vi har såpass mange features-søyler som skal shuffles...
```
import eli5
from eli5.sklearn import PermutationImportance
#?PermutationImportance
perm = PermutationImportance(rf, random_state=42)
perm.fit(X_test, y_test)
eli5.show_weights(perm, feature_names = X_test.columns.tolist())
```
**Resultat:**
<img width=30% src="assets/permimportance_sensor.png">
# Fin-tuning
Som diskutert i notebooken fra Lab 1 er det en rekke ting en kan gjøre dersom en ikke er fornøyd med ytelsen til en maskinlæringsmodell. En av disse er å justere på såkalte **hyperparametre** i modellen (dvs parametre som ikke settes under trening, men velges av oss).
Som vi skal se senere (når vi kommer til hvordan random forests fungerer) er det en rekke hyperparametre i random forests som kan influere ytelsen.
En mye brukt strategi for å finne gode valg av parametre er å *søke* gjennom et bestemt *grid* av potensielle parameterkombinasjoner. Enten ved å forsøke alle (dette kalles **grid search**) eller ved å forsøke et tilfeldig antall valg (dette kalles **randomized search**). Det finnes også andre, mer avanserte former for *hyperparameteroptimalisering*, for eksempel **bayesian search**. Vi skal se på grid search og randomized search. For bayesiansk søk, se for eksempel `scikit-optimize`: https://scikit-optimize.github.io/#skopt.BayesSearchCV.
(Det er også mulig å søke gjennom ulike *modeller* i tillegg til deres hyperparemetre, men det skal vi ikke gå inn på her)
Vi forsøker:
## Et mulig parametergrid å søke gjennom
Her er to parametergrid som ofte vil fungere bra for random forest. Nøyaktig hvilke parametre som gir mening å forsøke i en gitt situasjon avhenger blant annet av datasettet en har. For å velge klokt her kreves det en del erfaring, samt forståelse av modellen.
La oss bare velge noe relativt trygt, og så ikke bry oss om dette er det *beste* valget.
Vi lager to grids, et lite; et stort:
```
param_grid_small = {
'max_depth': [5, 10, 15, 20, 30, 100, None],
'n_estimators': [50, 100, 500, 1000]
}
param_grid_large = {
'bootstrap': [True, False],
'max_depth': [5, 10, 20, 30, 40, 50, 60, 70, 80, 90, 100, None],
'min_samples_leaf': [1, 2, 4],
'min_samples_split': [2, 5, 10],
'n_estimators': [50, 100, 500, 1000]
}
```
Når vi senere kjører grid search, randomized search og bayesian search, blir disse grids konvertert til en type matriser, og alle kombinasjoner av parametre blir potensielle kandidater.
Det betyr 7x4 = 28 kombinasjoner for `param_grid_small` og 2x12x3x3x4 = 864 kombinasjoner for `param_grid_large`.
## Grid search
Å søke gjennom absolutt alle kombinasjoner i `param_grid_large` blir for kostbart tidsmessig. Vi bruker derfor `param_grid_small`:
```
from sklearn.model_selection import GridSearchCV
rf_gs = GridSearchCV(estimator=rf, param_grid=param_grid_small, cv=3, n_jobs=-1)
rf_gs.fit(X_train, y_train)
best_gs_model = rf_gs.best_estimator_
best_gs_model
best_gs_model.score(X_test, y_test)
```
Søket klarte i dette tilfellet ikke å finne en bedre parameterkombinasjon enn den vi allerede hadde.
## Randomized search
Med randomized search er de vi som bestemmer antall (tilfeldig valgte) kombinasjoner som skal forsøkes. Vi kan derfor tillate oss å bruke `param_grid_large`:
```
from sklearn.model_selection import RandomizedSearchCV
rf_rs = RandomizedSearchCV(estimator=rf, param_distributions=param_grid_large, n_iter=50, cv=3, n_jobs=-1, random_state=42)
rf_rs.fit(X_train, y_train)
best_rs_model = rf_rs.best_estimator_
best_rs_model
best_rs_model.score(X_test, y_test)
```
Fortsatt ikke vesentlig bedre enn vår første modell.
Med et større søk (det vil si, `n_iter` satt til et større tall) kan det hende at vi oppdager bedre parametre. Men jo større antall forsøk, jo lenger beregningstid...
# Noen oppgaver
> **Din tur!** Klarer du å lage en modell som kan predikere hvilken person som genererte hver sensormåling? <em>Gi meg din mobiltelefon så skal jeg fortelle deg hvem du er</em>
> **Din tur!** Undersøk hvilke bevegelser som best skiller personer fra hverandre.
# Ekstra
La oss forsøke en annen modell: **logistisk regresjon**:
```
from sklearn.linear_model import LogisticRegression
from sklearn.preprocessing import StandardScaler
std_sc = StandardScaler()
X_train_std = std_sc.fit_transform(X_train)
X_test_std = std_sc.transform(X_test)
log_reg = LogisticRegression(solver='lbfgs', multi_class='auto', max_iter=1000, C=0.2, random_state=42)
log_reg.fit(X_train_std, y_train)
log_reg.score(X_test_std, y_test)
```
Denne modellen scorer 96%, altså langt bedre enn de random forest-variantene vi forsøkte over. Dette illusterer viktigheten av hensiktsmessig modellvalg, tilpasset data og problemstilling man står ovenfor.
## Forvirringsmatrise
```
y_pred_logreg = log_reg.predict(X_test)
cm_logreg = confusion_matrix(y_test, y_pred_logreg)
plot_confusion_matrix_with_colorbar(cm_logreg, classes=np.unique(y_test), figsize=(10,10))
```
Denne kan sammenlignes fra vår forvirringsmatrise fra random forest:
```
plot_confusion_matrix_with_colorbar(cm, classes=np.unique(y_test), figsize=(10,10))
```
# Ekstra ekstra
For moro skyld, la oss også forsøke noe kraftigere: en gradient boosting-basert modell.
```
from sklearn.ensemble import GradientBoostingClassifier
gb = GradientBoostingClassifier(random_state=42, n_estimators=500)
gb.fit(X_train, y_train)
gb.score(X_test, y_test)
```
Ca. 94%.
## Forvirringsmatrise
```
y_pred_gs = gb.predict(X_test)
cm_gs = confusion_matrix(y_test, y_pred_gs)
fig, ax = plt.subplots(figsize=(12,12))
_ = plot_confusion_matrix(cm_gs, classes=np.unique(y_test), ax=ax)
```
## Ensembling
En (av mange) mulige måter å forbedre modeller på er å bruke **ensembling**: slå sammen prediksjonene fra flere modeller. Dette er spesielt nyttig dersom man har flere, svært ulike modeller, som hver for seg scorer høyt. Sammen kan de da ofte bli enda bedre (*wisdom of the crowd*).
La oss forsøke med de vi har til nå:
```
from sklearn.ensemble import VotingClassifier
eclf = VotingClassifier(estimators=[('rf', best_rs_model), ('logreg', log_reg), ('gnb', gb)], voting='soft')
eclf.fit(X_train, y_train)
eclf.score(X_test, y_test)
```
I vårt tilfelle gav ikke dette en bedre modell enn logistisk regresjon alene.
```
y_pred_eclf = eclf.predict(X_test)
cm_eclf = confusion_matrix(y_test, y_pred_eclf)
fig, ax = plt.subplots(figsize=(12,12))
_ = plot_confusion_matrix(cm_eclf, classes=np.unique(y_test), ax=ax)
# Merk: For å slippe å trene modellene på nytt når de ensembles kunne vi brukt mlextend
# sin EnsembleVoteClassifier. Det vil spare mye tid:
#!pip install mlextend
#from mlxtend.classifier import EnsembleVoteClassifier
#import copy
#eclf = EnsembleVoteClassifier(clfs=[best_rs_model, gnb], weights=[1,1], refit=False)
```
| github_jupyter |
# Getting Started with Amazon SageMaker Studio Lab
## Welcome to your SageMaker Studio Lab project
Your SageMaker Studio Lab project is a notebook development environment with 25 GB of persistent storage and access to a CPU or GPU runtime. Everything about your project is automatically saved (notebooks, source code files, datasets, Git repos you have cloned, Conda environments, JupyterLab extensions, etc.) so each time you launch your project you can pick up where you left off. SageMaker Studio Lab is based on the open-source JupyterLab, so you can take advantage of open-source Jupyter extensions in your project.
## Running Python code
This Getting Started document is a [Jupyter notebook](https://jupyter.org/). Notebooks enable you to combine live code, equations, Markdown, images, visualizations, and other content into a single document that you can share with other people.
To run the following Python code, select the cell with a click, and then type `Shift-Enter` on your keyboard or click the play button in the notebook toolbar at the top of the document.
```
a = 10
b = 20
c = a + b
print(a, b, c)
```
To learn more about Python see [The Python Tutorial](https://docs.python.org/3/tutorial/).
## Creating notebooks, source code files and accessing the Terminal
SageMaker Studio Lab lets you create notebooks, source code files, and access the built-in Terminal. You can do this by clicking on the "+" button at the top of the file browser in the left panel to open the Launcher:

In the Launcher, there are a set of cards that allow you to launch notebooks in different environments, create source code files, or access the Terminal:

All of the notebooks, files, and datasets that you create are saved in your persistent project directory and are available when you open your project. To get help or access documentation, click on the **Help** menu in the menu bar at the top of the page.
## Installing Python packages
The simplest way of installing Python packages is to use either of the following magic commands in a code cell of a notebook:
`%conda install <package>`
`%pip install <package>`
These magic commands will always install packages into the environment used by that notebook and any packages you install are saved in your persistent project directory. Note: we don't recommend using `!pip` or `!conda` as those can behave in unexpected ways when you have multiple environments.
Here is an example that shows how to install NumPy into the environment used by this notebook:
```
%conda install numpy
```
Now you can use NumPy:
```
import numpy as np
np.random.rand(10)
```
## SageMaker Studio Lab example notebooks
SageMaker Studio Lab works with familiar open-source data science and machine learning libraries, such as [NumPy](https://numpy.org/), [pandas](https://pandas.pydata.org/), [scikit-learn](https://scikit-learn.org/stable/), [PyTorch](https://pytorch.org/), and [TensorFlow](https://www.tensorflow.org/).
To help you take the next steps, we have a GitHub repository with a set of example notebooks that cover a wide range of data science and machine learning topics, from importing and cleaning data to data visualization and training machine learning models.
<button class="jp-mod-styled" data-commandlinker-command="git:clone" data-commandlinker-args="{"URL": "https://github.com/aws/studio-lab-examples.git"}">Clone SageMaker Studio Lab Example Notebooks</button>
## AWS Machine Learning University
[Machine Learning University (MLU)](https://aws.amazon.com/machine-learning/mlu/) provides anybody, anywhere, at any time access to the same machine learning courses used to train Amazon’s own developers on machine learning. Learn how to use ML with the learn-at-your-own-pace MLU Accelerator learning series.
<button class="jp-mod-styled" data-commandlinker-command="git:clone" data-commandlinker-args="{"URL": "https://github.com/aws-samples/aws-machine-learning-university-accelerated-tab.git"}">Clone MLU Notebooks</button>
## Dive into Deep Learning (D2L)
[Dive into Deep Learning (D2L)](https://www.d2l.ai/) is an open-source, interactive book that teaches the ideas, the mathematical theory, and the code that powers deep learning. With over 150 Jupyter notebooks, D2L provides a comprehensive overview of deep learning principles and a state-of-the-art introduction to deep learning in computer vision and natural language processing. With tens of millions of online page views, D2L has been adopted for teaching by over 300 universities from 55 countries, including Stanford, MIT, Harvard, and Cambridge.
<button class="jp-mod-styled" data-commandlinker-command="git:clone" data-commandlinker-args="{"URL": "https://github.com/d2l-ai/d2l-pytorch-sagemaker-studio-lab.git"}">Clone D2L Notebooks</button>
## Hugging Face
[Hugging Face](http://huggingface.co/) is the home of the [Transformers](https://huggingface.co/transformers/) library and state-of-the-art natural language processing, speech, and computer vision models.
<button class="jp-mod-styled" data-commandlinker-command="git:clone" data-commandlinker-args="{"URL": "https://github.com/huggingface/notebooks.git"}">Clone Hugging Face Notebooks</button>
## Switching to a GPU runtime
Depending on the kinds of algorithms you are using, you may want to switch to a GPU or a CPU runtime for faster computation. First, save your work and then navigate back to your project overview page to select the instance type you want. You can navigate back to your project page by selecting the **Open Project Overview Page** in the **Amazon SageMaker Studio Lab** menu. Switching the runtime will stop all your kernels, but all of your notebooks, files, and datasets will be saved in your persistent project directory.
Note that a GPU runtime session is limited to 4 hours and a CPU runtime session is limited to 12 hours of continuous use.
## Managing packages and Conda environments
### Your default environment
SageMaker Studio Lab uses Conda environments to encapsulate the software (Python, R, etc.) packages needed to run notebooks. Your project contains a default Conda environment, named `default`, with the [IPython kernel](https://ipython.readthedocs.io/en/stable/) and that is about it. There are a couple of ways to install additional packages into this environment.
As described above, you can use the following magic commands in any notebook:
`%conda install <package>`
`%pip install <package>`
These magic commands will always install packages into the environment used by that notebook and any packages you install are saved in your persistent project directory. Note: we don't recommend using `!pip` or `!conda` as those can behave in unexpected ways when you have multiple environments.
Alternatively, you can open the Terminal and activate the environment using:
`$ conda activate default`
Once the environment is activated, you can install packages using the [Conda](https://docs.conda.io/en/latest/) or [pip](https://pip.pypa.io/en/stable/) command lines:
`$ conda install <package>`
`$ pip install <package>`
The conda installation for SageMaker Studio Lab uses a default channel of [conda-forge](https://conda-forge.org/), so you don't need to add the `-c conda-forge` argument when calling `conda install`.
### Creating and using new Conda environments
There are a couple of ways of creating new Conda environments.
**First**, you can open the Terminal and directly create a new environment using the Conda command line:
`$ conda env create --name my_environment python=3.9`
This example creates an new environment named `my_environment` with Python 3.9.
**Alternatively**, if you have a Conda environment file, can right click on the file in the JupyterLab file browser, and select the "Build Conda Environment" item:

To activate any Conda environment in the Terminal, run:
`$ conda activate my_environment`
Once you do this, any pakcages installed using Conda or pip will be installed in that environment.
To use your new Conda environments with notebooks, make sure the `ipykernel` package is installed into that environment:
`$ conda install ipykernel`
Once installed `ipykernel`, you should see a card in the launcher for that environment and kernel after about a minute.
<div class="alert alert-info"> <b>Note:</b> It may take about one minute for the new environment to appear as a kernel option.</div>
## Installing JupyterLab and Jupyter Server extensions
SageMaker Studio Lab enables you to install open-source JupyterLab and Jupyter Server extensions. These extensions are typically Python packages that can be installed using `conda` or `pip`. To install these extensions, open the Terminal and activate the `studiolab` environment:
`$ conda activate studiolab`
Then you can install the relevant JupyterLab or Jupyter Server extension:
`$ conda install <jupyter_extension>`
You will need to refresh your page to pickup any JupyterLab extensions you have installed, or power cycle your project runtime to pickup any Jupyter server extensions.
## Adding *Open in Studio Lab* links to your GitHub repositories
If you have public GitHub repositories with Jupyter Notebooks, you can make it easy for other users to open these notebooks in SageMaker Studio Lab by adding an *Open in Studio Lab* link to a README.md or notebook. This allows anyone to quickly preview the notebook and import it into their SageMaker Studio Lab project.
To add an *Open in Studio Lab* badge to your README.md file use the following markdown
```
[](https://studiolab.sagemaker.aws/import/github/org/repo/blob/master/path/to/notebook.ipynb)
```
and replace `org`, `repo`, the path and the notebook filename with those for your repo. Or in HTML:
```
<a href="https://studiolab.sagemaker.aws/import/github/org/repo/blob/master/path/to/notebook.ipynb">
<img src="https://studiolab.sagemaker.aws/studiolab.svg" alt="Open In SageMaker Studio Lab"/>
</a>
```
This will creates a badge like:
[](https://studiolab.sagemaker.aws/import/github/d2l-ai/d2l-pytorch-sagemaker-studio-lab/blob/161e45f1055654c547ffe3c81bd5f06310e96cff/GettingStarted-D2L.ipynb)
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
Licensed under the Apache License, Version 2.0 (the "License").
# Neural Machine Translation with Attention
<table class="tfo-notebook-buttons" align="left"><td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/tensorflow/blob/master/tensorflow/contrib/eager/python/examples/nmt_with_attention/nmt_with_attention.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td><td>
<a target="_blank" href="https://github.com/tensorflow/tensorflow/tree/master/tensorflow/contrib/eager/python/examples/nmt_with_attention/nmt_with_attention.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a></td></table>
This notebook trains a sequence to sequence (seq2seq) model for Spanish to English translation using [tf.keras](https://www.tensorflow.org/programmers_guide/keras) and [eager execution](https://www.tensorflow.org/programmers_guide/eager). This is an advanced example that assumes some knowledge of sequence to sequence models.
After training the model in this notebook, you will be able to input a Spanish sentence, such as *"¿todavia estan en casa?"*, and return the English translation: *"are you still at home?"*
The translation quality is reasonable for a toy example, but the generated attention plot is perhaps more interesting. This shows which parts of the input sentence has the model's attention while translating:
<img src="https://tensorflow.org/images/spanish-english.png" alt="spanish-english attention plot">
Note: This example takes approximately 10 mintues to run on a single P100 GPU.
```
from __future__ import absolute_import, division, print_function
# Import TensorFlow >= 1.10 and enable eager execution
import tensorflow as tf
tf.enable_eager_execution()
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
import unicodedata
import re
import numpy as np
import os
import time
print(tf.__version__)
```
## Download and prepare the dataset
We'll use a language dataset provided by http://www.manythings.org/anki/. This dataset contains language translation pairs in the format:
```
May I borrow this book? ¿Puedo tomar prestado este libro?
```
There are a variety of languages available, but we'll use the English-Spanish dataset. For convenience, we've hosted a copy of this dataset on Google Cloud, but you can also download your own copy. After downloading the dataset, here are the steps we'll take to prepare the data:
1. Add a *start* and *end* token to each sentence.
2. Clean the sentences by removing special characters.
3. Create a word index and reverse word index (dictionaries mapping from word → id and id → word).
4. Pad each sentence to a maximum length.
```
# Download the file
path_to_zip = tf.keras.utils.get_file(
'spa-eng.zip', origin='http://download.tensorflow.org/data/spa-eng.zip',
extract=True)
path_to_file = os.path.dirname(path_to_zip)+"/spa-eng/spa.txt"
# Converts the unicode file to ascii
def unicode_to_ascii(s):
return ''.join(c for c in unicodedata.normalize('NFD', s)
if unicodedata.category(c) != 'Mn')
def preprocess_sentence(w):
w = unicode_to_ascii(w.lower().strip())
# creating a space between a word and the punctuation following it
# eg: "he is a boy." => "he is a boy ."
# Reference:- https://stackoverflow.com/questions/3645931/python-padding-punctuation-with-white-spaces-keeping-punctuation
w = re.sub(r"([?.!,¿])", r" \1 ", w)
w = re.sub(r'[" "]+', " ", w)
# replacing everything with space except (a-z, A-Z, ".", "?", "!", ",")
w = re.sub(r"[^a-zA-Z?.!,¿]+", " ", w)
w = w.rstrip().strip()
# adding a start and an end token to the sentence
# so that the model know when to start and stop predicting.
w = '<start> ' + w + ' <end>'
return w
# 1. Remove the accents
# 2. Clean the sentences
# 3. Return word pairs in the format: [ENGLISH, SPANISH]
def create_dataset(path, num_examples):
lines = open(path, encoding='UTF-8').read().strip().split('\n')
word_pairs = [[preprocess_sentence(w) for w in l.split('\t')] for l in lines[:num_examples]]
return word_pairs
# This class creates a word -> index mapping (e.g,. "dad" -> 5) and vice-versa
# (e.g., 5 -> "dad") for each language,
class LanguageIndex():
def __init__(self, lang):
self.lang = lang
self.word2idx = {}
self.idx2word = {}
self.vocab = set()
self.create_index()
def create_index(self):
for phrase in self.lang:
self.vocab.update(phrase.split(' '))
self.vocab = sorted(self.vocab)
self.word2idx['<pad>'] = 0
for index, word in enumerate(self.vocab):
self.word2idx[word] = index + 1
for word, index in self.word2idx.items():
self.idx2word[index] = word
def max_length(tensor):
return max(len(t) for t in tensor)
def load_dataset(path, num_examples):
# creating cleaned input, output pairs
pairs = create_dataset(path, num_examples)
# index language using the class defined above
inp_lang = LanguageIndex(sp for en, sp in pairs)
targ_lang = LanguageIndex(en for en, sp in pairs)
# Vectorize the input and target languages
# Spanish sentences
input_tensor = [[inp_lang.word2idx[s] for s in sp.split(' ')] for en, sp in pairs]
# English sentences
target_tensor = [[targ_lang.word2idx[s] for s in en.split(' ')] for en, sp in pairs]
# Calculate max_length of input and output tensor
# Here, we'll set those to the longest sentence in the dataset
max_length_inp, max_length_tar = max_length(input_tensor), max_length(target_tensor)
# Padding the input and output tensor to the maximum length
input_tensor = tf.keras.preprocessing.sequence.pad_sequences(input_tensor,
maxlen=max_length_inp,
padding='post')
target_tensor = tf.keras.preprocessing.sequence.pad_sequences(target_tensor,
maxlen=max_length_tar,
padding='post')
return input_tensor, target_tensor, inp_lang, targ_lang, max_length_inp, max_length_tar
```
### Limit the size of the dataset to experiment faster (optional)
Training on the complete dataset of >100,000 sentences will take a long time. To train faster, we can limit the size of the dataset to 30,000 sentences (of course, translation quality degrades with less data):
```
# Try experimenting with the size of that dataset
num_examples = 30000
input_tensor, target_tensor, inp_lang, targ_lang, max_length_inp, max_length_targ = load_dataset(path_to_file, num_examples)
# Creating training and validation sets using an 80-20 split
input_tensor_train, input_tensor_val, target_tensor_train, target_tensor_val = train_test_split(input_tensor, target_tensor, test_size=0.2)
# Show length
len(input_tensor_train), len(target_tensor_train), len(input_tensor_val), len(target_tensor_val)
```
### Create a tf.data dataset
```
BUFFER_SIZE = len(input_tensor_train)
BATCH_SIZE = 64
N_BATCH = BUFFER_SIZE//BATCH_SIZE
embedding_dim = 256
units = 1024
vocab_inp_size = len(inp_lang.word2idx)
vocab_tar_size = len(targ_lang.word2idx)
dataset = tf.data.Dataset.from_tensor_slices((input_tensor_train, target_tensor_train)).shuffle(BUFFER_SIZE)
dataset = dataset.batch(BATCH_SIZE, drop_remainder=True)
```
## Write the encoder and decoder model
Here, we'll implement an encoder-decoder model with attention which you can read about in the TensorFlow [Neural Machine Translation (seq2seq) tutorial](https://www.tensorflow.org/tutorials/seq2seq). This example uses a more recent set of APIs. This notebook implements the [attention equations](https://www.tensorflow.org/tutorials/seq2seq#background_on_the_attention_mechanism) from the seq2seq tutorial. The following diagram shows that each input words is assigned a weight by the attention mechanism which is then used by the decoder to predict the next word in the sentence.
<img src="https://www.tensorflow.org/images/seq2seq/attention_mechanism.jpg" width="500" alt="attention mechanism">
The input is put through an encoder model which gives us the encoder output of shape *(batch_size, max_length, hidden_size)* and the encoder hidden state of shape *(batch_size, hidden_size)*.
Here are the equations that are implemented:
<img src="https://www.tensorflow.org/images/seq2seq/attention_equation_0.jpg" alt="attention equation 0" width="800">
<img src="https://www.tensorflow.org/images/seq2seq/attention_equation_1.jpg" alt="attention equation 1" width="800">
We're using *Bahdanau attention*. Lets decide on notation before writing the simplified form:
* FC = Fully connected (dense) layer
* EO = Encoder output
* H = hidden state
* X = input to the decoder
And the pseudo-code:
* `score = FC(tanh(FC(EO) + FC(H)))`
* `attention weights = softmax(score, axis = 1)`. Softmax by default is applied on the last axis but here we want to apply it on the *1st axis*, since the shape of score is *(batch_size, max_length, hidden_size)*. `Max_length` is the length of our input. Since we are trying to assign a weight to each input, softmax should be applied on that axis.
* `context vector = sum(attention weights * EO, axis = 1)`. Same reason as above for choosing axis as 1.
* `embedding output` = The input to the decoder X is passed through an embedding layer.
* `merged vector = concat(embedding output, context vector)`
* This merged vector is then given to the GRU
The shapes of all the vectors at each step have been specified in the comments in the code:
```
def gru(units):
# If you have a GPU, we recommend using CuDNNGRU(provides a 3x speedup than GRU)
# the code automatically does that.
if tf.test.is_gpu_available():
return tf.keras.layers.CuDNNGRU(units,
return_sequences=True,
return_state=True,
recurrent_initializer='glorot_uniform')
else:
return tf.keras.layers.GRU(units,
return_sequences=True,
return_state=True,
recurrent_activation='sigmoid',
recurrent_initializer='glorot_uniform')
class Encoder(tf.keras.Model):
def __init__(self, vocab_size, embedding_dim, enc_units, batch_sz):
super(Encoder, self).__init__()
self.batch_sz = batch_sz
self.enc_units = enc_units
self.embedding = tf.keras.layers.Embedding(vocab_size, embedding_dim)
self.gru = gru(self.enc_units)
def call(self, x, hidden):
x = self.embedding(x)
output, state = self.gru(x, initial_state = hidden)
return output, state
def initialize_hidden_state(self):
return tf.zeros((self.batch_sz, self.enc_units))
class Decoder(tf.keras.Model):
def __init__(self, vocab_size, embedding_dim, dec_units, batch_sz):
super(Decoder, self).__init__()
self.batch_sz = batch_sz
self.dec_units = dec_units
self.embedding = tf.keras.layers.Embedding(vocab_size, embedding_dim)
self.gru = gru(self.dec_units)
self.fc = tf.keras.layers.Dense(vocab_size)
# used for attention
self.W1 = tf.keras.layers.Dense(self.dec_units)
self.W2 = tf.keras.layers.Dense(self.dec_units)
self.V = tf.keras.layers.Dense(1)
def call(self, x, hidden, enc_output):
# enc_output shape == (batch_size, max_length, hidden_size)
# hidden shape == (batch_size, hidden size)
# hidden_with_time_axis shape == (batch_size, 1, hidden size)
# we are doing this to perform addition to calculate the score
hidden_with_time_axis = tf.expand_dims(hidden, 1)
# score shape == (batch_size, max_length, hidden_size)
score = tf.nn.tanh(self.W1(enc_output) + self.W2(hidden_with_time_axis))
# attention_weights shape == (batch_size, max_length, 1)
# we get 1 at the last axis because we are applying score to self.V
attention_weights = tf.nn.softmax(self.V(score), axis=1)
# context_vector shape after sum == (batch_size, hidden_size)
context_vector = attention_weights * enc_output
context_vector = tf.reduce_sum(context_vector, axis=1)
# x shape after passing through embedding == (batch_size, 1, embedding_dim)
x = self.embedding(x)
# x shape after concatenation == (batch_size, 1, embedding_dim + hidden_size)
x = tf.concat([tf.expand_dims(context_vector, 1), x], axis=-1)
# passing the concatenated vector to the GRU
output, state = self.gru(x)
# output shape == (batch_size * 1, hidden_size)
output = tf.reshape(output, (-1, output.shape[2]))
# output shape == (batch_size * 1, vocab)
x = self.fc(output)
return x, state, attention_weights
def initialize_hidden_state(self):
return tf.zeros((self.batch_sz, self.dec_units))
encoder = Encoder(vocab_inp_size, embedding_dim, units, BATCH_SIZE)
decoder = Decoder(vocab_tar_size, embedding_dim, units, BATCH_SIZE)
```
## Define the optimizer and the loss function
```
optimizer = tf.train.AdamOptimizer()
def loss_function(real, pred):
mask = 1 - np.equal(real, 0)
loss_ = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=real, logits=pred) * mask
return tf.reduce_mean(loss_)
```
## Checkpoints (Object-based saving)
```
checkpoint_dir = './training_checkpoints'
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
checkpoint = tf.train.Checkpoint(optimizer=optimizer,
encoder=encoder,
decoder=decoder)
```
## Training
1. Pass the *input* through the *encoder* which return *encoder output* and the *encoder hidden state*.
2. The encoder output, encoder hidden state and the decoder input (which is the *start token*) is passed to the decoder.
3. The decoder returns the *predictions* and the *decoder hidden state*.
4. The decoder hidden state is then passed back into the model and the predictions are used to calculate the loss.
5. Use *teacher forcing* to decide the next input to the decoder.
6. *Teacher forcing* is the technique where the *target word* is passed as the *next input* to the decoder.
7. The final step is to calculate the gradients and apply it to the optimizer and backpropagate.
```
EPOCHS = 10
for epoch in range(EPOCHS):
start = time.time()
hidden = encoder.initialize_hidden_state()
total_loss = 0
for (batch, (inp, targ)) in enumerate(dataset):
loss = 0
with tf.GradientTape() as tape:
enc_output, enc_hidden = encoder(inp, hidden)
dec_hidden = enc_hidden
dec_input = tf.expand_dims([targ_lang.word2idx['<start>']] * BATCH_SIZE, 1)
# Teacher forcing - feeding the target as the next input
for t in range(1, targ.shape[1]):
# passing enc_output to the decoder
predictions, dec_hidden, _ = decoder(dec_input, dec_hidden, enc_output)
loss += loss_function(targ[:, t], predictions)
# using teacher forcing
dec_input = tf.expand_dims(targ[:, t], 1)
batch_loss = (loss / int(targ.shape[1]))
total_loss += batch_loss
variables = encoder.variables + decoder.variables
gradients = tape.gradient(loss, variables)
optimizer.apply_gradients(zip(gradients, variables))
if batch % 100 == 0:
print('Epoch {} Batch {} Loss {:.4f}'.format(epoch + 1,
batch,
batch_loss.numpy()))
# saving (checkpoint) the model every 2 epochs
if (epoch + 1) % 2 == 0:
checkpoint.save(file_prefix = checkpoint_prefix)
print('Epoch {} Loss {:.4f}'.format(epoch + 1,
total_loss / N_BATCH))
print('Time taken for 1 epoch {} sec\n'.format(time.time() - start))
```
## Translate
* The evaluate function is similar to the training loop, except we don't use *teacher forcing* here. The input to the decoder at each time step is its previous predictions along with the hidden state and the encoder output.
* Stop predicting when the model predicts the *end token*.
* And store the *attention weights for every time step*.
Note: The encoder output is calculated only once for one input.
```
def evaluate(sentence, encoder, decoder, inp_lang, targ_lang, max_length_inp, max_length_targ):
attention_plot = np.zeros((max_length_targ, max_length_inp))
sentence = preprocess_sentence(sentence)
inputs = [inp_lang.word2idx[i] for i in sentence.split(' ')]
inputs = tf.keras.preprocessing.sequence.pad_sequences([inputs], maxlen=max_length_inp, padding='post')
inputs = tf.convert_to_tensor(inputs)
result = ''
hidden = [tf.zeros((1, units))]
enc_out, enc_hidden = encoder(inputs, hidden)
dec_hidden = enc_hidden
dec_input = tf.expand_dims([targ_lang.word2idx['<start>']], 0)
for t in range(max_length_targ):
predictions, dec_hidden, attention_weights = decoder(dec_input, dec_hidden, enc_out)
# storing the attention weigths to plot later on
attention_weights = tf.reshape(attention_weights, (-1, ))
attention_plot[t] = attention_weights.numpy()
predicted_id = tf.argmax(predictions[0]).numpy()
result += targ_lang.idx2word[predicted_id] + ' '
if targ_lang.idx2word[predicted_id] == '<end>':
return result, sentence, attention_plot
# the predicted ID is fed back into the model
dec_input = tf.expand_dims([predicted_id], 0)
return result, sentence, attention_plot
# function for plotting the attention weights
def plot_attention(attention, sentence, predicted_sentence):
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(1, 1, 1)
ax.matshow(attention, cmap='viridis')
fontdict = {'fontsize': 14}
ax.set_xticklabels([''] + sentence, fontdict=fontdict, rotation=90)
ax.set_yticklabels([''] + predicted_sentence, fontdict=fontdict)
plt.show()
def translate(sentence, encoder, decoder, inp_lang, targ_lang, max_length_inp, max_length_targ):
result, sentence, attention_plot = evaluate(sentence, encoder, decoder, inp_lang, targ_lang, max_length_inp, max_length_targ)
print('Input: {}'.format(sentence))
print('Predicted translation: {}'.format(result))
attention_plot = attention_plot[:len(result.split(' ')), :len(sentence.split(' '))]
plot_attention(attention_plot, sentence.split(' '), result.split(' '))
```
## Restore the latest checkpoint and test
```
# restoring the latest checkpoint in checkpoint_dir
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
translate('hace mucho frio aqui.', encoder, decoder, inp_lang, targ_lang, max_length_inp, max_length_targ)
translate('esta es mi vida.', encoder, decoder, inp_lang, targ_lang, max_length_inp, max_length_targ)
translate('¿todavia estan en casa?', encoder, decoder, inp_lang, targ_lang, max_length_inp, max_length_targ)
# wrong translation
translate('trata de averiguarlo.', encoder, decoder, inp_lang, targ_lang, max_length_inp, max_length_targ)
```
## Next steps
* [Download a different dataset](http://www.manythings.org/anki/) to experiment with translations, for example, English to German, or English to French.
* Experiment with training on a larger dataset, or using more epochs
| github_jupyter |
<h1>Using pre-trained embeddings with TensorFlow Hub</h1>
This notebook illustrates:
<ol>
<li>How to instantiate a TensorFlow Hub module</li>
<li>How to find pre-trained TensorFlow Hub modules for a variety of purposes</li>
<li>How to examine the embeddings of a Hub module</li>
<li>How one Hub module composes representations of sentences from individual words</li>
<li>How to assess word embeddings using a semantic similarity test</li>
</ol>
```
# change these to try this notebook out# chang
BUCKET = 'cloud-training-demos-ml'
PROJECT = 'cloud-training-demos'
REGION = 'us-central1'
```
Install the TensorFlow Hub library
```
!pip install -q tensorflow-hub
import tensorflow as tf
import tensorflow_hub as hub
import matplotlib.pyplot as plt
import numpy as np
import os
import pandas as pd
import re
import seaborn as sns
import scipy
import math
os.environ['BUCKET'] = BUCKET
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
os.environ['TFVERSION'] = '1.8'
import tensorflow as tf
print(tf.__version__)
```
<h2>TensorFlow Hub Concepts</h2>
TensorFlow Hub is a library for the publication, discovery, and consumption of reusable parts of machine learning models. A module is a self-contained piece of a TensorFlow graph, along with its weights and assets, that can be reused across different tasks in a process known as transfer learning, which we covered as part of the course on Image Models.
To download and use a module, it's as easy as:
However, because modules are self-contained parts of a TensorFlow graph, in order to actually collect values from a module, you'll need to evaluate it in the context of a session.
First, let's explore what hub modules there are. Go to [the documentation page](https://www.tensorflow.org/hub/modules) and explore a bit.
Note that TensorFlow Hub has modules for Images, Text, and Other. In this case, we're interested in a Text module, so navigate to the Text section.
Within the Text section, there are a number of modules. If you click on a link, you'll be taken to a page that describes the module and links to the original paper where the model was proposed. Click on a model in the Word2Vec section of the page.
Note the details section, which describes what the module expects as input, how it preprocesses data, what it does when it encounters a word it hasn't seen before (OOV means "out of vocabulary") and in this case, how word embeddings can be composed to form sentence embeddings.
Finally, note the URL of the page. This is the URL you can copy to instantiate your module.
<h2>Task 1: Create an embedding using the NNLM model</h2>
To complete this task:
<ol>
<li>Find the module URL for the NNLM 50 dimensional English model</li>
<li>Use it to instantiate a module as 'embed'</li>
<li>Print the embedded representation of "cat"</li>
</ol>
NOTE: downloading hub modules requires downloading a lot of data. Instantiating the module will take a few minutes.
```
# Task 1
embed = ...
```
When I completed this exercise, I got a vector that looked like:
[[ 0.11233182 -0.3176392 -0.01661182...]]
<h2>Task 2: Assess the Embeddings Informally</h2>
<ol>
<li>Identify some words to test</li>
<li>Retrieve the embeddings for each word</li>
<li>Determine what method to use to compare each pair of embeddings</li>
</ol>
So, now we have some vectors but the question is, are they any good? One way of testing whether they are any good is to try them for your task. But, first, let's just take a peak.
For our test, we'll need three common words such that two of the words are much closer in meaning than the third.
```
word_1 = #
word_2 = #
word_3 = #
```
Now, we'll use the same process of using our Hub module to generate embeddings but instead of printing the embeddings, capture them in a variable called 'my_embeddings'.
```
# Task 2b
```
Now, we'll use Seaborn's heatmap function to see how the vectors compare to each other. I've written the shell of a function that you'll need to complete that will generate a heatmap. The one piece that's missing is how we'll compare each pair of vectors. Note that because we are computing a score for every pair of vectors, we should have len(my_embeddings)^2 scores. There are many valid ways of comparing vectors. Generality, similarity scores are symmetric. The simplest is to take their dot product. For extra credit, implement a more complicated vector comparison function.
```
def plot_similarity(labels, embeddings):
corr = # ... TODO: fill out a len(embeddings) x len(embeddings) array
sns.set(font_scale=1.2)
g = sns.heatmap(
corr,
xticklabels=labels,
yticklabels=labels,
vmin=0,
vmax=1,
cmap="YlOrRd")
g.set_xticklabels(labels, rotation=90)
g.set_title("Semantic Textual Similarity")
plot_similarity([word_1, word_2, word_3], my_embeddings)
```
What you should observe is that, trivially, all words are identical to themselves, and, more interestingly, that the two more similar words have more similar embeddings than the third word.
<h2>Task 3: From Words to Sentences</h2>
Up until now, we've used our module to produce representations of words. But, in fact, if we want to, we can also use it to construct representations of sentences. The methods used by the module to compose a representation of a sentence won't be as nuanced as what an RNN might do, but they are still worth examining because they are so convenient.
<ol>
<li> Examine the documentation for our hub module and determine how to ask it to construct a representation of a sentence</li>
<li> Figure out how the module takes word embeddings and uses them to construct sentence embeddings </li>
<li> Construct a embeddings of a "cat", "The cat sat on the mat", "dog" and "The cat sat on the dog" and plot their similarity
</ol>
```
# Task 3
```
Which is cat more similar to, "The cat sat on the mat" or "dog"? Is this desireable?
Think back to how an RNN scans a sequence and maintains its state. Naive methods of embedding composition (mapping many to one) can't possibly compete with a network trained for this very purpose!
<h2>Task 4: Assessing the Embeddings Formally</h2>
Of course, it's great to know that our embeddings match our intuitions to an extent, but it'd be better to have a formal, data-driven measure of the quality of the representation.
Researchers have
The [STS Benchmark](http://ixa2.si.ehu.es/stswiki/index.php/STSbenchmark) provides an intristic evaluation of the degree to which similarity scores computed using sentence embeddings align with human judgements. The benchmark requires systems to return similarity scores for a diverse selection of sentence pairs. Pearson correlation is then used to evaluate the quality of the machine similarity scores against human judgements.
```
def load_sts_dataset(filename):
# Loads a subset of the STS dataset into a DataFrame. In particular both
# sentences and their human rated similarity score.
sent_pairs = []
with tf.gfile.GFile(filename, "r") as f:
for line in f:
ts = line.strip().split("\t")
# (sent_1, sent_2, similarity_score)
sent_pairs.append((ts[5], ts[6], float(ts[4])))
return pd.DataFrame(sent_pairs, columns=["sent_1", "sent_2", "sim"])
def download_and_load_sts_data():
sts_dataset = tf.keras.utils.get_file(
fname="Stsbenchmark.tar.gz",
origin="http://ixa2.si.ehu.es/stswiki/images/4/48/Stsbenchmark.tar.gz",
extract=True)
sts_dev = load_sts_dataset(
os.path.join(os.path.dirname(sts_dataset), "stsbenchmark", "sts-dev.csv"))
sts_test = load_sts_dataset(
os.path.join(
os.path.dirname(sts_dataset), "stsbenchmark", "sts-test.csv"))
return sts_dev, sts_test
sts_dev, sts_test = download_and_load_sts_data()
sts_dev.head()
```
<h3>Build the Evaluation Graph</h3>
Next, we need to build the evaluation graph.
```
sts_input1 = tf.placeholder(tf.string, shape=(None))
sts_input2 = tf.placeholder(tf.string, shape=(None))
# For evaluation we use exactly normalized rather than
# approximately normalized.
sts_encode1 = tf.nn.l2_normalize(embed(sts_input1), axis=1)
sts_encode2 = tf.nn.l2_normalize(embed(sts_input2), axis=1)
cosine_similarities = tf.reduce_sum(tf.multiply(sts_encode1, sts_encode2), axis=1)
clip_cosine_similarities = tf.clip_by_value(cosine_similarities, -1.0, 1.0)
sim_scores = 1.0 - tf.acos(clip_cosine_similarities)
```
<h3>Evaluate Sentence Embeddings</h3>
Finally, we need to create a session and run our evaluation.
```
sts_data = sts_dev #@param ["sts_dev", "sts_test"] {type:"raw"}
text_a = sts_data['sent_1'].tolist()
text_b = sts_data['sent_2'].tolist()
dev_scores = sts_data['sim'].tolist()
def run_sts_benchmark(session):
"""Returns the similarity scores"""
emba, embb, scores = session.run(
[sts_encode1, sts_encode2, sim_scores],
feed_dict={
sts_input1: text_a,
sts_input2: text_b
})
return scores
with tf.Session() as session:
session.run(tf.global_variables_initializer())
session.run(tf.tables_initializer())
scores = run_sts_benchmark(session)
pearson_correlation = scipy.stats.pearsonr(scores, dev_scores)
print('Pearson correlation coefficient = {0}\np-value = {1}'.format(
pearson_correlation[0], pearson_correlation[1]))
```
<h3>Extra Credit</h3>
For extra credit, re-run this analysis with a different Hub module. Are the results different? If so, how?
<h2>Further Reading</h2>
We published a [blog post](https://developers.googleblog.com/2018/04/text-embedding-models-contain-bias.html) on how bias can affect text embeddings. It's worth a read!
| github_jupyter |
# some of the common NLP tasks reference notebook
__(using NLTK)__
## Text transformation tasks
```
with open("sample_text.txt") as file:
raw_text = file.read()
from nltk.tokenize import word_tokenize, sent_tokenize
some_text = "hello world! Its Aman Singh from India. This is a reference notebook for some of the common Natural Language Processing (NLP) tasks."
# word tokenization
tokenized_words = word_tokenize(some_text)
print("Text:\n", some_text)
print("\ntokenized words:")
print(tokenized_words)
# sentence tokenization
tokenized_sents = sent_tokenize(some_text)
print("\ntokenized sentences:")
print(tokenized_sents)
# sentence/word tokenization with other laguages
french_text = "Ceci est 1 première phrase. Puis j'en écris une seconde. pour finir en voilà une troisième sans mettre de majuscule"
tokenized_sents = sent_tokenize(french_text, language = 'french')
print("\nFrench Text:\n", french_text)
print("\ntokenized french sentences:")
print(tokenized_sents)
tokenized_words = word_tokenize(french_text, language = 'french')
print("\ntokenized french words:")
print(tokenized_words)
# nltk's wordnet package includes groups of synonyms, antonyms and also a brief definition for each
# here only examplifying synonyms and antonyms
from nltk.corpus import wordnet
synonyms, antonyms = [], []
for syn in wordnet.synsets("big"):
for lemma in syn.lemmas():
synonyms.append(lemma.name())
if lemma.antonyms():
antonyms.append(lemma.antonyms()[0].name())
print("SYNONYMS:\n",synonyms)
print("\nANTONYMS:\n",antonyms)
# stemming vs lemmatization
from nltk.stem import PorterStemmer, WordNetLemmatizer
# stemming
stemmer = PorterStemmer()
print("stemming:")
print(stemmer.stem("increases"))
# lemmatization
lemmatizer = WordNetLemmatizer()
print("\nlemmatization:")
print(lemmatizer.lemmatize("increases"))
# generating n-grams from tokens
from nltk.tokenize import word_tokenize
from nltk.util import ngrams
text = "In the fields of computational linguistics and probability, an n-gram is a contiguous sequence of n items from a given sample of text or speech."
word_tokens = word_tokenize(text)
bigrams = ngrams(word_tokens, 2)
trigrams = ngrams(word_tokens, 3)
print("Text:\n", text)
print("\nbi-grams:\n", list(bigrams))
print("\ntri-grams:\n", list(trigrams))
```
## Information extraction tasks
```
# Part-Of-Speech (POS) tagging
from nltk.tag import pos_tag
text = "I like to go to the park with my dog"
tokens = word_tokenize(text)
tags = pos_tag(tokens)
print("POS Tags:\n", tags)
```
#### POS tags References:
---------------------------------
CC | Coordinating conjunction |
CD | Cardinal number |
DT | Determiner |
EX | Existential there |
FW | Foreign word |
IN | Preposition or subordinating conjunction |
JJ | Adjective |
JJR | Adjective, comparative |
JJS | Adjective, superlative |
LS | List item marker |
MD | Modal |
NN | Noun, singular or mass |
NNS | Noun, plural |
NNP | Proper noun, singular |
NNPS| Proper noun, plural |
PDT | Predeterminer |
POS | Possessive ending |
PRP | Personal pronoun |
PRP$| Possessive pronoun |
RB | Adverb |
RBR | Adverb, comparative |
RBS | Adverb, superlative |
RP | Particle |
SYM | Symbol |
TO | to |
UH | Interjection |
VB | Verb, base form |
VBD | Verb, past tense |
VBG | Verb, gerund or present participle |
VBN | Verb, past participle |
VBP | Verb, non-3rd person singular present |
VBZ | Verb, 3rd person singular present |
WDT | Wh-determiner |
WP | Wh-pronoun |
WP$ | Possessive wh-pronoun |
WRB | Wh-adverb |
```
# Named-Entity-Recognition (NER)
from nltk.tokenize import word_tokenize
from nltk.tag import pos_tag
from nltk.chunk import ne_chunk
text = "Mark Elliot Zuckerberg (born May 14, 1984) is a co-founder of Facebook."
# tokenize sentence into words
tokens = word_tokenize(text)
# POS tagging of the tokens
tags = pos_tag(tokens)
# using the ner function
ner = ne_chunk(tags)
print("Named-Entity-Recognition:\n", ner)
#todo: make an abstractive summarizer like smmry.com
#PAPER: https://arxiv.org/pdf/1602.06023.pdf
```
| github_jupyter |
## Training knowledge graph embedding by using the Deep Graph Library with MXNet backend
The **Amazon SageMaker Python SDK** makes it easy to train Deep Graph Library (DGL) models. In this example, you generate knowledge graph embedding using the [DMLC DGL API](https://github.com/dmlc/dgl.git) and FB15k dataset.
For more information about knowledge graph embedding and this example, see https://github.com/dmlc/dgl/tree/master/apps/kg
### Setup
Define a few variables that are needed later in the example.
```
import sagemaker
from sagemaker import get_execution_role
from sagemaker.session import Session
# Setup session
sess = sagemaker.Session()
# S3 bucket for saving code and model artifacts.
# Feel free to specify a different bucket here if you wish.
bucket = sess.default_bucket()
# Location to put your custom code.
custom_code_upload_location = "customcode"
# IAM execution role that gives Amazon SageMaker access to resources in your AWS account.
# You can use the Amazon SageMaker Python SDK to get the role from the notebook environment.
role = get_execution_role()
```
### Amazon SageMaker estimator class
The Amazon SageMaker estimator allows you to run a single machine in Amazon SageMaker, using CPU or GPU-based instances.
When you create the estimator, pass in the file name of the training script and the name of the IAM execution role. Also provide a few other parameters. train_instance_count and train_instance_type determine the number and type of Amazon SageMaker instances that are used for the training job. The hyperparameters parameter is a dictionary of values that is passed to your training script as parameters that you can use argparse to parse.
Here, you can directly use the DL Container provided by Amazon SageMaker for training DGL models by specifying the MXNet framework version (>= 1.6.0) and the python version (only py3). You can also add a task_tag with value 'DGL' to help tracking the task.
```
from sagemaker.mxnet.estimator import MXNet
ENTRY_POINT = "train.py"
CODE_PATH = "./"
account = sess.boto_session.client("sts").get_caller_identity()["Account"]
region = sess.boto_session.region_name
params = {}
params["dataset"] = "FB15k"
params["model"] = "DistMult"
params["batch_size"] = 1024
params["neg_sample_size"] = 256
params["hidden_dim"] = 2000
params["gamma"] = 500.0
params["lr"] = 0.1
params["max_step"] = 100000
params["batch_size_eval"] = 16
params["valid"] = True
params["test"] = True
params["neg_adversarial_sampling"] = True
task_tags = [{"Key": "ML Task", "Value": "DGL"}]
estimator = MXNet(
entry_point=ENTRY_POINT,
source_dir=CODE_PATH,
role=role,
train_instance_count=1,
train_instance_type="ml.p3.2xlarge",
framework_version="1.6.0",
py_version="py3",
hyperparameters=params,
tags=task_tags,
sagemaker_session=sess,
)
```
### Running the Training Job
After you construct the Estimator object, you can fit it by using Amazon SageMaker. The dataset is automatically downloaded.
```
estimator.fit()
```
## Output
You can get the resulting embedding output from the Amazon SageMaker console by searching for the training task and looking for the address of 'S3 model artifact'
| github_jupyter |
```
import os
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import torch.utils.data
import torchvision
from torchvision import datasets
from torchvision import transforms
from torchsummary import summary
!pip install optuna
import optuna
DEVICE = torch.device("cuda") ##'cuda' or 'cpu'
BATCHSIZE = 128
CLASSES = 10 #CLASSES = 10 for cifar10 and 100 for cifar100
DIR = os.getcwd()
EPOCHS = 10
LOG_INTERVAL = 10
N_TRAIN_EXAMPLES = BATCHSIZE * 30
N_VALID_EXAMPLES = BATCHSIZE * 10
def define_model(trial):
layers = []
layers.append(nn.Conv2d(in_channels=3, out_channels=32, kernel_size=3, stride=1))
layers.append(nn.BatchNorm2d(32))
layers.append(nn.ReLU())
layers.append(nn.Dropout(0.2))
layers.append(nn.Conv2d(in_channels=32, out_channels=64, kernel_size=3, stride=2))
layers.append(nn.BatchNorm2d(64))
layers.append(nn.ReLU())
layers.append(nn.Dropout(0.2))
layers.append(nn.Conv2d(in_channels=64, out_channels=128, kernel_size=3, stride=2))
layers.append(nn.BatchNorm2d(128))
layers.append(nn.ReLU())
layers.append(nn.Dropout(0.2))
layers.append(nn.Conv2d(in_channels=128, out_channels=256, kernel_size=3, stride=2))
layers.append(nn.BatchNorm2d(256))
layers.append(nn.ReLU())
layers.append(nn.Flatten())
layers.append(nn.Linear(256*2*2, 500)) #output size found by printing the model detail using summary in torchsummary
layers.append(nn.Dropout(0.2))
layers.append(nn.Linear(500, CLASSES)) #CLASSES = 10 for cifar10 and 100 for cifar100
#cross entropy loss used as loss function, therefore no softmax layer here
return nn.Sequential(*layers)
def get_cifar10():
# Load cifar10 dataset.
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root=DIR, train=True,
download=True, transform=transform)
#split training data into training-80% and validation-20%
train_set, val_set = torch.utils.data.random_split(trainset, [int(0.8*len(trainset)), int(0.2*len(trainset))])
train_loader = torch.utils.data.DataLoader(train_set, batch_size=BATCHSIZE,
shuffle=True, num_workers=2)
valid_loader = torch.utils.data.DataLoader(val_set, batch_size=BATCHSIZE,
shuffle=False, num_workers=2)
"""
testset = torchvision.datasets.CIFAR10(root=DIR, train=False,
download=True, transform=transform)
test_loader = torch.utils.data.DataLoader(testset, batch_size=BATCHSIZE,
shuffle=False, num_workers=2)
"""
return train_loader, valid_loader
def objective(trial):
# Generate the model.
model = define_model(trial).to(DEVICE)
# Generate the optimizers.
#optimizer_name = trial.suggest_categorical("optimizer", ["Adam", "RMSprop", "SGD"]) #for hp tuning
optimizer_name = "Adam"
lr = trial.suggest_float("lr", 1e-5, 1e-1, log=True) #for hp tuning
#lr = 0.001
optimizer = getattr(optim, optimizer_name)(model.parameters(), lr=lr)
CEloss = nn.CrossEntropyLoss() ## this loss object must be used the loop. Directly using nn.CrossEntropyLoss() gives error
# Get the MNIST dataset.
train_loader, valid_loader = get_cifar10()
# Training of the model.
for epoch in range(EPOCHS):
model.train()
for batch_idx, (data, target) in enumerate(train_loader):
# Limiting training data for faster epochs.
if batch_idx * BATCHSIZE >= N_TRAIN_EXAMPLES:
break
#data, target = data.view(data.size(0), -1).to(DEVICE), target.to(DEVICE) ## for mnist
data, target = data.to(DEVICE), target.to(DEVICE) ## for cifar 10 and 100
optimizer.zero_grad()
output = model(data)
loss = CEloss(output, target) ## used cross entropy loss
loss.backward()
optimizer.step()
# Validation of the model.
model.eval()
correct = 0
with torch.no_grad():
val_loss_batch = 0
for batch_idx, (data, target) in enumerate(valid_loader):
# Limiting validation data.
if batch_idx * BATCHSIZE >= N_VALID_EXAMPLES:
break
#data, target = data.view(data.size(0), -1).to(DEVICE), target.to(DEVICE) ## for mnist
data, target = data.to(DEVICE), target.to(DEVICE) ## for cifar 10 and 100
output = model(data)
# Get the index of the max log-probability.
pred = output.argmax(dim=1, keepdim=True)
correct += pred.eq(target.view_as(pred)).sum().item()
val_loss_batch += CEloss(output, target).item() ## used cross entropy loss
#accuracy = correct / min(len(valid_loader.dataset), N_VALID_EXAMPLES)
val_loss_epoch = val_loss_batch / min(len(valid_loader.dataset), N_VALID_EXAMPLES)
#trial.report(accuracy, epoch)
trial.report(val_loss_epoch, epoch)
# Handle pruning based on the intermediate value.
if trial.should_prune():
raise optuna.exceptions.TrialPruned()
return val_loss_epoch #accuracy
if __name__ == "__main__":
study = optuna.create_study(direction="minimize") # 'maximize' because objective function is returning accuracy
study.optimize(objective, n_trials=50, timeout=600) ## link: https://optuna.readthedocs.io/en/stable/reference/generated/optuna.study.Study.html#optuna.study.Study.optimize
pruned_trials = [t for t in study.trials if t.state == optuna.trial.TrialState.PRUNED]
complete_trials = [t for t in study.trials if t.state == optuna.trial.TrialState.COMPLETE]
print("Study statistics: ")
print(" Number of finished trials: ", len(study.trials))
print(" Number of pruned trials: ", len(pruned_trials))
print(" Number of complete trials: ", len(complete_trials))
print("Best trial:")
trial = study.best_trial
print(" Value: ", trial.value)
print(" Params: ")
for key, value in trial.params.items():
print(" {}: {}".format(key, value))
study.best_trial
optuna.visualization.plot_optimization_history(study)
optuna.visualization.plot_param_importances(study) ## this is important to figure out which hp is important
optuna.visualization.plot_slice(study) ## this gives a clear picture
optuna.visualization.plot_parallel_coordinate(study)
# SKIP THIS
#### used for testing output sizes of layers in the model
#****important: only change the input filter to maintain the output size of each layer
"""
model = nn.Sequential(nn.Conv2d(in_channels=3, out_channels=32, kernel_size=3, stride=1)
,nn.BatchNorm2d(32)
,nn.ReLU()
,nn.Dropout(0.2)
,nn.Conv2d(in_channels=32, out_channels=128, kernel_size=3, stride=2)
,nn.BatchNorm2d(128) #this must be same as the out_channel of the previous layer
,nn.ReLU()
,nn.Dropout(0.2)
,nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, stride=2)
,nn.BatchNorm2d(128)
,nn.ReLU()
,nn.Dropout(0.2)
,nn.Conv2d(in_channels=128, out_channels=256, kernel_size=3, stride=2)
,nn.BatchNorm2d(256)
,nn.ReLU()
,nn.Flatten()
,nn.Linear(256*2*2, 500) #output size found by printing the model detail using summary in torchsummary
,nn.Dropout(0.2)
,nn.Linear(500, CLASSES)) #CLASSES = 10 for cifar10 and 100 for cifar100
print(summary(model,(3,32,32)))
"""
```
| github_jupyter |
# RMSprop from scratch
```
from mxnet import ndarray as nd
# RMSProp.
def rmsprop(params, sqrs, lr, gamma, batch_size):
eps_stable = 1e-8
for param, sqr in zip(params, sqrs):
g = param.grad / batch_size
sqr[:] = gamma * sqr + (1. - gamma) * nd.square(g)
div = lr * g / nd.sqrt(sqr + eps_stable)
param[:] -= div
import mxnet as mx
from mxnet import autograd
from mxnet import gluon
import random
mx.random.seed(1)
random.seed(1)
# Generate data.
num_inputs = 2
num_examples = 1000
true_w = [2, -3.4]
true_b = 4.2
X = nd.random_normal(scale=1, shape=(num_examples, num_inputs))
y = true_w[0] * X[:, 0] + true_w[1] * X[:, 1] + true_b
y += .01 * nd.random_normal(scale=1, shape=y.shape)
dataset = gluon.data.ArrayDataset(X, y)
# Construct data iterator.
def data_iter(batch_size):
idx = list(range(num_examples))
random.shuffle(idx)
for batch_i, i in enumerate(range(0, num_examples, batch_size)):
j = nd.array(idx[i: min(i + batch_size, num_examples)])
yield batch_i, X.take(j), y.take(j)
# Initialize model parameters.
def init_params():
w = nd.random_normal(scale=1, shape=(num_inputs, 1))
b = nd.zeros(shape=(1,))
params = [w, b]
sqrs = []
for param in params:
param.attach_grad()
sqrs.append(param.zeros_like())
return params, sqrs
# Linear regression.
def net(X, w, b):
return nd.dot(X, w) + b
# Loss function.
def square_loss(yhat, y):
return (yhat - y.reshape(yhat.shape)) ** 2 / 2
%matplotlib inline
import matplotlib as mpl
mpl.rcParams['figure.dpi']= 120
import matplotlib.pyplot as plt
import numpy as np
def train(batch_size, lr, gamma, epochs, period):
assert period >= batch_size and period % batch_size == 0
[w, b], sqrs = init_params()
total_loss = [np.mean(square_loss(net(X, w, b), y).asnumpy())]
# Epoch starts from 1.
for epoch in range(1, epochs + 1):
for batch_i, data, label in data_iter(batch_size):
with autograd.record():
output = net(data, w, b)
loss = square_loss(output, label)
loss.backward()
rmsprop([w, b], sqrs, lr, gamma, batch_size)
if batch_i * batch_size % period == 0:
total_loss.append(np.mean(square_loss(net(X, w, b), y).asnumpy()))
print("Batch size %d, Learning rate %f, Epoch %d, loss %.4e" %
(batch_size, lr, epoch, total_loss[-1]))
print('w:', np.reshape(w.asnumpy(), (1, -1)),
'b:', b.asnumpy()[0], '\n')
x_axis = np.linspace(0, epochs, len(total_loss), endpoint=True)
plt.semilogy(x_axis, total_loss)
plt.xlabel('epoch')
plt.ylabel('loss')
plt.show()
train(batch_size=10, lr=0.03, gamma=0.9, epochs=3, period=10)
```
## Next
[RMSProp with Gluon](../chapter06_optimization/rmsprop-gluon.ipynb)
For whinges or inquiries, [open an issue on GitHub.](https://github.com/zackchase/mxnet-the-straight-dope)
| github_jupyter |
```
import tensorflow as tf
import numpy as np
import pandas as pd
import matplotlib.gridspec as gridspec
import matplotlib.pyplot as plot
from tensorflow.examples.tutorials.mnist import input_data
batch_size = 32
X_dim = 784
z_dim = 16
c_dim = 10
h_dim = 256
mnist = input_data.read_data_sets('../../MNIST_data', one_hot=True)
def plot_images(samples):
fig = plot.figure(figsize=(4, 4))
gs = gridspec.GridSpec(4, 4)
gs.update(wspace=0.05, hspace=0.05)
for i, sample in enumerate(samples):
ax = plot.subplot(gs[i])
plot.axis('off')
ax.set_xticklabels([])
ax.set_yticklabels([])
ax.set_aspect('equal')
plot.imshow(sample.reshape(28, 28), cmap='Greys_r')
return fig
def xavier_init(size):
input_dim = size[0]
xavier_stddev = 1. / tf.sqrt(input_dim / 2.)
return tf.random_normal(shape=size, stddev=xavier_stddev)
X = tf.placeholder(tf.float32, shape=[None, X_dim])
D_W1 = tf.Variable(xavier_init([X_dim, h_dim]))
D_b1 = tf.Variable(tf.zeros(shape=[h_dim]))
D_W2 = tf.Variable(xavier_init([h_dim, 1]))
D_b2 = tf.Variable(tf.zeros(shape=[1]))
theta_D = [D_W1, D_W2, D_b1, D_b2]
z = tf.placeholder(tf.float32, shape=[None, z_dim])
c = tf.placeholder(tf.float32, shape=[None, c_dim])
G_W1 = tf.Variable(xavier_init([z_dim + c_dim, h_dim // 2]))
G_b1 = tf.Variable(tf.zeros(shape=[h_dim // 2]))
G_W2 = tf.Variable(xavier_init([h_dim // 2, X_dim]))
G_b2 = tf.Variable(tf.zeros(shape=[X_dim]))
theta_G = [G_W1, G_W2, G_b1, G_b2]
Q_W1 = tf.Variable(xavier_init([X_dim, h_dim // 2]))
Q_b1 = tf.Variable(tf.zeros(shape=[h_dim // 2]))
Q_W2 = tf.Variable(xavier_init([h_dim // 2, 10]))
Q_b2 = tf.Variable(tf.zeros(shape=[10]))
theta_Q = [Q_W1, Q_W2, Q_b1, Q_b2]
def get_sample_z(size):
return np.random.uniform(-1., 1., size=size)
def get_sample_c(m):
return np.random.multinomial(1, 10*[0.1], size=m)
def generator(z, c):
inputs = tf.concat(axis=1, values=[z, c])
G_h1 = tf.nn.relu(tf.matmul(inputs, G_W1) + G_b1)
G_out = tf.sigmoid(tf.matmul(G_h1, G_W2) + G_b2)
return G_out
def discriminator(x):
D_h1 = tf.nn.relu(tf.matmul(x, D_W1) + D_b1)
D_out = tf.sigmoid(tf.matmul(D_h1, D_W2) + D_b2)
return D_out
def Q(x):
Q_h1 = tf.nn.relu(tf.matmul(x, Q_W1) + Q_b1)
Q_out = tf.nn.softmax(tf.matmul(Q_h1, Q_W2) + Q_b2)
return Q_out
generator_sample = generator(z, c)
discriminator_real = discriminator(X)
discriminator_fake = discriminator(generator_sample)
q_c_x = Q(generator_sample)
d_loss = -tf.reduce_mean(tf.log(discriminator_real + 1e-8) + tf.log(1 - discriminator_fake + 1e-8))
g_loss = -tf.reduce_mean(tf.log(discriminator_fake + 1e-8))
q_loss = tf.reduce_mean(-tf.reduce_sum(tf.log(q_c_x + 1e-8) * c, 1)) + \
tf.reduce_mean(-tf.reduce_sum(tf.log(c + 1e-8) * c, 1))
d_step = tf.train.AdamOptimizer(0.0001).minimize(d_loss, var_list=theta_D)
g_step = tf.train.AdamOptimizer(0.0001).minimize(g_loss, var_list=theta_G)
q_step = tf.train.AdamOptimizer(0.0001).minimize(q_loss, var_list=theta_G + theta_Q)
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
for i in range(100000):
x_batch, _ = mnist.train.next_batch(batch_size)
z_batch = get_sample_z((batch_size, z_dim))
c_batch = get_sample_c(batch_size)
_, d_loss_val = sess.run(
[d_step, d_loss],
feed_dict={X: x_batch, z: z_batch, c: c_batch}
)
_, g_loss_val = sess.run(
[g_step, g_loss],
feed_dict={z: z_batch, c: c_batch}
)
sess.run([q_step], feed_dict={z: z_batch, c: c_batch})
if i % 100 == 0:
print('Iteration: {} - Discriminator Loss: {:.4}, Generator Loss: {:.4}'
.format(i, d_loss_val, g_loss_val))
if i % 1000 == 0:
samples = sess.run(generator_sample, feed_dict={z: get_sample_z(size=(16, z_dim)),
c: get_sample_c(16)})
fig = plot_images(samples)
plot.show()
plot.close(fig)
```
| github_jupyter |
# Basic Usage
This notebook is one small example of what is possible with Jupyter notebooks, but there are many more out there. For a quick idea about what Jupyter notebooks can do, check out the live demos available at:
* http://www.nature.com/news/ipython-interactive-demo-7.21492
* https://try.jupyter.org/
The IPython/Jupyter project maintains a curated example of amazing notebooks on a variety of topics:
https://github.com/ipython/ipython/wiki/A-gallery-of-interesting-IPython-Notebooks#entire-books-or-other-large-collections-of-notebooks-on-a-topic
## Demo
For this demonstration, I will use the example of Planck's Law describing the radiation intensity of a blackbody at a certain temperature for a given wavelength of light. This example is trivial, but it will show the various aspects of Jupyter notebooks, including:
1. Writing prose with Markdown
2. Linking in rich media
3. Writing equations in Latex
4. Writing Python code
5. Visualization
## Planck's Law
### 1. Prose in Markdown
Planck's Law describes the electromagnetic radiation emited by a black body at a given temperature for a given wavelength.
This description is made possible by using a "Markdown" cell block. Markdown is an incredibly simple yet powerful markup language for formatting text. Documentation for Markdown is readily available, including at the following links:
* https://daringfireball.net/projects/markdown/
* https://help.github.com/articles/markdown-basics/
* https://github.com/adam-p/markdown-here/wiki/Markdown-Cheatsheet
* https://guides.github.com/features/mastering-markdown/
Get to know Markdown -- it is used in Jupyter notebooks, on Github, and in many other places.
To use Markdown in a cell, first create a new cell by clicking the "Insert" menu on your toolbar and then the "Insert Cell Below" menu item (shortcut: CTRL + M, b).
By default, new cells are code cells. Change the type of cell by clicking the "Cell" menu on your toolbar and then clicking "Cell Type -> Markdown" (shortcut: CTRL + M + m).
This is now a new code cell that is parsed as Markdown. I can now continue my description of Planck's Law.
Some quick facts about Planck's Law in the form of an unordered list. I am using asterisks "*" for my bullet points, but you can also use dashes "-" and plus signs "+". See [Github's documentation](https://help.github.com/articles/markdown-basics/#unordered-lists)) for more information.
* Named after scientist Max Planck
* Originall propsed in 1900
* There were two competing descriptions of the spectrum of radiation that were not entirely accurate
To list these two competing descriptions, I can use an ordered list (see [Github's documentation](https://help.github.com/articles/markdown-basics/#ordered-lists)):
Competing descriptions:
1. Wien approximation
2. Rayleigh–Jeans law
To add more information, I could use a nested list ([Github's documentation](https://help.github.com/articles/markdown-basics/#nested-lists)):
1. Wien approximation
* Inaccurate for long wavelengths
2. Rayleigh-Jeans law
* Inaccurate for short wavelengths
For more information, I can refer the reader to the [Wikipedia page on Planck's Law](https://en.wikipedia.org/wiki/Planck%27s_law). To make a hyperlink, you can write the link text within square brackets "[TEXT]" and immediatelly follow the text with the URL within parentheses "(URL)" (see [Github's documentation](https://help.github.com/articles/markdown-basics/#links)).
### 2. Rich media
I can even pull in images from the web to include in my notebook. To create a link to an image, write the URL to the image as a normal link, but pre-pend an exclamation point. For example, "\!\[link text\](URL)":

Linked image by Darth Kule (Own work) [Public domain], via Wikimedia Commons.
Using some code from the IPython project, I can even embed a YouTube video by passing the video ID to the `YouTubeVideo` function [documented here](http://ipython.readthedocs.org/en/stable/api/generated/IPython.display.html?highlight=youtube#IPython.display.YouTubeVideo):
```
from IPython.display import YouTubeVideo
# Title: Max Planck Solves the Ultraviolet Catastrophe for Blackbody Radiation | Doc Physics
# Author: Doc Schuster
YouTubeVideo('H-7f-3OAXm0')
```
### 3. Equations
Markdown allows you to write mathematical equations using [LaTeX](https://www.latex-project.org/) that will be rendered using [MathJax](https://www.mathjax.org/). The simplest way of including an equations is to wrap the LaTeX code in sets of double dollar signs, "$$":
Planck Equation:
$$ B_{\lambda}(\lambda, T) = \frac{2hc^2}{\lambda^5} \frac{1}{e^{\frac{hc}{\lambda k_B T}} - 1}$$
Another, more difficult, method of writing equations is to instead use a cell block as a LaTeX block. This may be accomplished using "cell magics", specifically "%%latex":
```
%%latex
\begin{aligned}
B_{\lambda}(\lambda, T) = \frac{2hc^2}{\lambda^5} \frac{1}{e^{\frac{hc}{\lambda k_B T}} - 1}
\end{aligned}
```
You can also use "line magics" to write LaTeX inline in Markdown cells:
%latex \begin{aligned} B_{\lambda}(\lambda, T) = \frac{2hc^2}{\lambda^5} \frac{1}{e^{\frac{hc}{\lambda k_B T}} - 1} \end{aligned}
### 4. Code
You can, of course, also include code in the notebook as code is the default cell type.
Let's create a function for Planck's Law:
```
import numpy as np
def planck(wavelength, temp):
""" Return the emitted radiation from a blackbody of a given temp and wavelength
Args:
wavelength (float): wavelength (m)
temp (float): temperature of black body (Kelvin)
Returns:
float: spectral radiance (W / (sr m^3))
"""
k_b = 1.3806488e-23 # J/K Boltzmann constant
h = 6.626070040e-34 # J s - Planck's constant
c = 3e8 # m/s - speed of light
return ((2 * h * c ** 2) / wavelength ** 5 *
1 / (np.exp(h * c / (wavelength * k_b * temp)) - 1))
for temp in (3000, 4000, 5000):
rad = planck(0.5e-6, temp)
rad_kW_per_sr_m2_nm = rad / 1e3 / 1e9 # convert from W to kW and m to nm
print('%.3f K: %.5f kW/(sr m^2 nm)' % (temp, rad_kW_per_sr_m2_nm))
```
### 5. Visualization
Not only can the notebooks display console style text outputs from the code, but it can also display and save very detaild plots.
Below I use the Python plotting library, [matplotlib](http://matplotlib.org/), to reproduce the plot displayed in section 2.
```
# Import and alias to "plt"
import matplotlib.pyplot as plt
# Calculate
wavelength = np.linspace(1e-7, 3e-6, 1000)
temp = np.array([3000, 4000, 5000])
rad = np.zeros((wavelength.size, temp.size), dtype=np.float)
for i, t in enumerate(temp):
rad[:, i] = planck(wavelength, t)
% matplotlib nbagg
# Plot
text_x = wavelength[rad.argmax(axis=0)] * 1e6
text_y = rad.max(axis=0) / 1e3 / 1e9
temp_str = ['%.2f K' % t for t in temp]
fig, ax = plt.subplots()
ax.plot(wavelength * 1e6, rad / 1e3 / 1e9)
for _x, _y, _temp in zip(text_x, text_y, temp_str):
ax.text(_x, _y, _temp, ha='center')
plt.legend(labels=['%.2f K' % t for t in temp])
plt.xlabel(r'Wavelength ($\mu m$)')
plt.ylabel(r'Spectral radiance ($kW \cdot sr^{-1} \cdot m^{-2} \cdot nm^{-1}$)')
```
| github_jupyter |
<center><h1>Python Loops - Break or Conditionals?</h1></center>
<center><h3>Written 22/11/2020, by Tyler J. Russell</h3></center>
***
#### Abstract
I was working on my [Fox and Hen Game] project for computer science when I found myself questioning whether `break` statements or conditionals are more efficient for `for` and `while` loops. Now, obviously, there is a difference: one will always run at least one iteration, the other depends on the condition. However, in this paper, I will only be comparing them with the same use case; id est, at least one iteration with a break check.
As such, in this test, superiority is measured in performance - both time to execute and operational expense.
To ensure fairness of testing, I will be using the same number of tests to average for each (1 million) and also for the conditionals I will include the initial assignment in the setup rather than in the actual test block to avoid adding to the time for initial assignment. However, it is worth noting that initial assignment and cleanup afterwards would count into the operational expense and time difference in any real circumstance, so that should be taken into consideration. My aim in excluding it is simply to measure raw performance of both methods alone.
[Fox and Hen Game]: https://github.com/Nytelife26/transparency/tree/main/works/academic/2020/computer-science/assignments/fox-and-hen
#### Hypothesis
It is difficult to draw a logically concluded hypothesis for such a test, as one would assume reasonably that the two operations should be relatively similar. However, on the simple basis that conditionals require an assignment to memory, a loop back, and then another fetch to check the condition, whereas `break` requires only a single instruction to jump out of the loop, I believe `break` should be at least somewhat faster, if not significantly.
#### Testing
Setting up our test suite:
```
from timeit import timeit
def test(tests):
for (key, value) in tests.items():
time = timeit(value[0], value[2], number=value[1])
print(f"{key}: {time}")
```
Running our tests:
```
tests = {}
tests["while_break"] = [
"""
while True:
break
""",
1000000,
""
]
tests["while_check"] = [
"""
while not over:
over = True
""",
1000000,
"over = False"
]
tests["forin_break"] = [
"""
for x in p:
break
""",
1000000,
"p = [0, 1, 2]"
]
# It is not actually possible to directly build a
# conditional into a for loop in Python, as a
# consequence of how Python is built. As such, we
# will test 2 methods: erasing the rest of the list,
# and an `if...break`.
tests["forin_check_del"] = [
"""
for x in p:
if over:
p.clear()
over = True
""",
1000000,
"over = False;p = [0, 1, 2]"
]
tests["forin_check_brk"] = [
"""
for x in p:
if over:
break
over = True
""",
1000000,
"over = False;p = [0, 1, 2]"
]
test(tests)
```
Of course, I always run my test suites more than once. However, upon doing so, I noticed an inconsistency in the primary results. They seemed to change back and forth quite often, so I'll now be constructing a table to measure results and see if the discrepancies warrant a change in approach or an inconclusive mark.
As the reader you are welcome at any time to run the suite on your own machine to verify my claims.
```
import numpy as np
from matplotlib import pyplot as plt
from matplotlib import colors as col
def plot(checks):
"""Plots test results from checks. `checks` must be dict, where the key is the name and the value is an instance of `np.array`"""
# calculates plot values
low = min([checks[x].min() for x in checks])
lim = max([checks[x].max() for x in checks])
tests = range(1, len(checks[list(checks.keys())[0]])+1)
# plots graph
plt.xlabel("test no.")
plt.ylabel("time (s)")
for name, x in checks.items():
colour = list(col.BASE_COLORS.keys())[list(checks.keys()).index(name)]
plt.plot(tests, x, linestyle="-", c=colour, label=name)
plt.xticks(tests)
plt.ylim([low-(0.1*low), lim+(0.1*low)])
plt.legend(loc="upper left")
plt.show()
```
| Test | `while_break` | `while_check` |
|:----:|:--------------------:|:--------------------:|
| 1 | 0.016855678999945667 | 0.014753502000075969 |
| 2 | 0.017085809000036534 | 0.012262844999895606 |
| 3 | 0.01806852899972 | 0.021265754000069137 |
| 4 | 0.015186278999863134 | 0.01617280499976914 |
| 5 | 0.01349683799980994 | 0.020387914999446366 |
| 6 | 0.022207265000361076 | 0.01812310300010722 |
| 7 | 0.01449698000033095 | 0.016063858000052278 |
| 8 | 0.019192011000086495 | 0.013797925999824656 |
| 9 | 0.02037990900043951 | 0.014765822000299522 |
| 10 | 0.018137518000003183 | 0.013981764999698498 |
```
%matplotlib inline
res_brk = np.array([0.016855678999945667, 0.017085809000036534, 0.01806852899972, 0.015186278999863134, 0.01349683799980994, 0.022207265000361076, 0.01449698000033095, 0.019192011000086495, 0.02037990900043951, 0.018137518000003183])
res_chk = np.array([0.014753502000075969, 0.012262844999895606, 0.021265754000069137, 0.01617280499976914, 0.020387914999446366, 0.01812310300010722, 0.016063858000052278, 0.013797925999824656, 0.014765822000299522, 0.013981764999698498])
plot({"while_break": res_brk, "while_check": res_chk})
```
The results prove to be inconsistent a lot of the time. First, I will try increasing the base number of tests to average out, in order to see if the problem is actually that I am measuring too small a number of tests for the perhaps miniscule difference in execution time between the two constructs. This time, I will run for 1 billion tests, in order to magnify the results enough to see any genuinely major differences.
```
tests = {}
tests["while_break"] = [
"""
while True:
break
""",
1000000000,
""
]
tests["while_check"] = [
"""
while not over:
over = True
""",
1000000000,
"over = False"
]
test(tests)
```
| Test | `while_break` | `while_check` |
|:----:|:------------------:|:-----------------:|
| 1 | 7.545864007000091 | 8.893152812000153 |
| 2 | 6.9994925880000665 | 8.789875424999991 |
| 3 | 6.99228112200035 | 8.772128739999971 |
| 4 | 7.016520256000149 | 8.774441176999972 |
| 5 | 7.008395603999816 | 9.082717027000399 |
| 6 | 7.012604857000042 | 8.75714720399992 |
| 7 | 7.0721877389996735 | 8.862611889000618 |
| 8 | 7.0326520609996805 | 8.788487822999741 |
| 9 | 7.169503927000733 | 9.04424666399973 |
| 10 | 7.203111390999766 | 9.042622316000234 |
Now, to compare our new results:
```
%matplotlib inline
res_brk = np.array([7.545864007000091, 6.9994925880000665, 6.99228112200035, 7.016520256000149, 7.008395603999816, 7.012604857000042, 7.0721877389996735, 7.0326520609996805, 7.169503927000733, 7.203111390999766])
res_chk = np.array([8.893152812000153, 8.789875424999991, 8.772128739999971, 8.774441176999972, 9.082717027000399, 8.75714720399992, 8.862611889000618, 8.788487822999741, 9.04424666399973, 9.042622316000234])
plot({"while_break": res_brk, "while_check": res_chk})
```
As the graph shows, these results are significantly more consistent. Furthermore, they produce a conclusive result.
Now that the inconclusivity is resolved, we can draw the conclusions for our hypothesis.
#### Conclusion
Our hypothesis holds true for `while` loops: the `break` statement proves to be faster than conditionals in our test cases.
However, another interesting result can be seen from our testing; id est, in terms of `for` loops, conditionals are actually faster.
Not just conditionals, though - interestingly enough, using `:list:.clear()` to erase the rest of the elements on a conditional is faster than running `break` conditionally.
In summary:
- `break` is faster than conditionals for `while` loops.
- Conditionals are faster than `break` for `for` loops.
* Conditionally reserved `break` is faster than pure `break`.
* Conditionally reserved `:list:.clear()` is faster than conditionally reserved `break`.
Therefore, I conclude that `break` should be preferred to end `while` loops, and conditionals should be preferred to end `for` loops, with `:list:.clear()` being preferred in instances where the list does not need to remain intact and conditionally reserved `break` being preferred otherwise.
May this document serve useful to you, its reader, as this research did to me.
| github_jupyter |
```
import simulate
import withdrawal
import market
import harvesting
import metrics
import lens
from decimal import Decimal
import decimal
import itertools
import pandas
import numpy
import math
%matplotlib inline
#%pdb on
import seaborn
from matplotlib import pyplot as plt
import matplotlib
seaborn.set(style="whitegrid")
seaborn.set_context('poster')
stock_er = Decimal('.05')
bond_er = Decimal('.018')
inflation = Decimal('.02')
stock_pct = Decimal('.6')
discount_rate = (stock_pct * stock_er) + ((1-stock_pct) * bond_er)
#discount_rate = Decimal('.0054')
max_age = 92 # 95th percentile SOA2012 for 65/65 male/female
expenses = 40_000
def pv_expenses(age):
years = max_age - age + 1
e = [expenses] * years
npv = numpy.npv(discount_rate, e)
return npv
print(discount_rate)
pv_expenses(70)
def run(age, funded_ratio):
npv = pv_expenses(age)
portfolio_value = npv * funded_ratio
portfolio = (portfolio_value, 0) # this is a 100/0 portfolio but it'll get rebalanced before we start.
iwd = expenses / portfolio_value
wd_s = lambda p, h: withdrawal.ConstantDollar(p, h, rate=iwd)
df = simulate.calc_lens(lambda p: harvesting.AgeBased(p, 100, starting_age=age), wd_s, max_age-age, lens.calc_years_sustained, portfolio=portfolio)
return df
#run(70, Decimal('1.0')).head()
def long_sim():
data = pandas.DataFrame(columns=['age', 'funded_ratio', 'failure_rate'])
for age in range(40, 90):
for funded in range(80, 251, 1):
f = Decimal(funded) / 100
s = run(age, f)
failures = len(s[s<0])
count = len(s)
failure_rate = failures / count
data = data.append({'age': age, 'funded_ratio': f, 'failure_rate': failure_rate}, ignore_index=True)
print(data.head())
return data
data = long_sim()
#data.to_csv('funded_failures.csv')
#data = pandas.read_csv('funded_failures.csv')
plt.figure(figsize=(12,8))
plt.title('Failure Rates')
seaborn.scatterplot(data=data, x='age', y='funded_ratio', hue='failure_rate', legend=False)
plt.figure(figsize=(12,8))
plt.title('Failure Rates')
seaborn.scatterplot(data=data[data['failure_rate']>0.05], x='age', y='funded_ratio', hue='failure_rate', legend=False)
plt.figure(figsize=(8,6))
plt.title('Funded Ratio 0.8')
seaborn.lineplot(data=data[data['funded_ratio'] == Decimal('0.8')], x='age', y='failure_rate')
plt.figure(figsize=(8,6))
plt.title('Funded Ratio 0.9')
seaborn.lineplot(data=data[data['funded_ratio'] == Decimal('0.9')], x='age', y='failure_rate')
plt.figure(figsize=(8,6))
plt.title('Funded Ratio 1.0')
seaborn.lineplot(data=data[data['funded_ratio'] == Decimal('1.0')], x='age', y='failure_rate')
plt.figure(figsize=(8,6))
plt.title('Funded Ratio 1.1')
seaborn.lineplot(data=data[data['funded_ratio'] == Decimal('1.1')], x='age', y='failure_rate')
plt.figure(figsize=(8,6))
plt.title('Funded Ratio 1.2')
seaborn.lineplot(data=data[data['funded_ratio'] == Decimal('1.2')], x='age', y='failure_rate')
plt.figure(figsize=(8,6))
plt.title('Funded Ratio 1.3')
seaborn.lineplot(data=data[data['funded_ratio'] == Decimal('1.3')], x='age', y='failure_rate')
plt.figure(figsize=(8,6))
plt.title('Funded Ratio 1.4')
seaborn.lineplot(data=data[data['funded_ratio'] == Decimal('1.4')], x='age', y='failure_rate')
plt.figure(figsize=(8,6))
plt.title('Funded Ratio 1.5')
seaborn.lineplot(data=data[data['funded_ratio'] == Decimal('1.5')], x='age', y='failure_rate')
f_by_age = data[data['failure_rate'] <= Decimal('.05')].groupby('age').min()
print(f_by_age.head())
plt.figure(figsize=(8,6))
plt.title('Funded Ratio by Age: 5% failure rate')
seaborn.lineplot(data=f_by_age['funded_ratio'].astype(float))
f_by_age = data[data['failure_rate'] <= Decimal('0')].groupby('age').min()
print(f_by_age.head())
plt.figure(figsize=(8,6))
plt.title('Funded Ratio by Age: 0% failure rate')
seaborn.lineplot(data=f_by_age['funded_ratio'].astype(float))
import scipy.stats
for age in [50, 55, 60, 65, 70]:
f = data[data['age'] == age]
f = f[f['funded_ratio'] <= Decimal('1.5')]
print(f.head())
plt.figure(figsize=(8,6))
plt.title(f'Age {age}')
seaborn.lineplot(data=f, x='funded_ratio', y='failure_rate')
print(age, scipy.stats.linregress(f['funded_ratio'], f['failure_rate']))
f = data[data['age'] == 50]
f[f['funded_ratio'] <= Decimal('1.11')]
def sim():
data = pandas.DataFrame(columns=['age', 'funded_ratio', 'failure_rate'])
for age in range(40, 90):
for funded in range(80, 151, 1):
f = Decimal(funded) / 100
s = run(age, f)
failures = len(s[s<0])
count = len(s)
failure_rate = failures / count
data = data.append({'age': age, 'funded_ratio': f, 'failure_rate': failure_rate}, ignore_index=True)
print(data.head())
return data
data_new = sim()
#data_new.to_csv('funded_failures_tips.csv')
#data_new = pandas.read_csv('funded_failures_tips.csv')
plt.figure(figsize=(8,6))
plt.title('Funded Ratio 1.0')
seaborn.lineplot(data=data_new[data_new['funded_ratio'] == Decimal('1.0')], x='age', y='failure_rate')
new = pandas.DataFrame(data_new[data_new['funded_ratio'] == Decimal('1.0')])
new['discount'] = '0.54%'
old = pandas.DataFrame(data[data['funded_ratio'] == Decimal('1.0')])
old['discount'] = '3.73%'
joint = old.append(new)
plt.figure(figsize=(8,6))
plt.title('Funded Ratio 1.0')
seaborn.lineplot(data=joint, x='age', y='failure_rate', hue='discount')
f_by_age = data_new[data_new['failure_rate'] <= Decimal('0')].groupby('age').min()
print(f_by_age.head())
plt.figure(figsize=(8,6))
plt.title('Funded Ratio (0.54% discount rate) by Age: 0% failure rate')
seaborn.lineplot(data=f_by_age['funded_ratio'].astype(float))
```
| github_jupyter |
# Coursework 1: Data loading, visualisation and simple analysis using Python
In this coursework, we will deal with a dataset stored in the ".csv" format, which describes the housing price in Boston. This is a small dataset with only 506 cases. But it would be a good illustration how Python can be used for loading, visualising and analysing a dataset. The dataset was originally published at
\[1\] Harrison, D. and Rubinfeld, D.L. Hedonic prices and the demand for clean air, J. Environ. Economics & Management, vol.5, 81-102, 1978.
## Dataset
A copy of the .csv data is already here after you git clone this repository. The .csv format is a format for spreadsheet, which can be opened using Microsoft Excel or Libreoffice.
## Import libraries
The code importing the libraries is already provided, which includes
* pandas: a library for loading .csv datasets
* numpy: a library for manipulating numbers and arrays
* matplotlib: for data visualisation
* seaborn: for data visualisation as well
* sklearn: for linear regression and machine learing
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
from sklearn.metrics import r2_score
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
```
## 1. Load data and print the first few lines using pandas (10 points).
## Dataset description
Each row is a case of the housing price. There are 506 cases in total. Each column is an attribute, there are 14 attributes, including:
**crim**: per capita crime rate by town
**zn**: proportion of residential land zoned for lots over 25,000 sq.ft.
**indus**: proportion of non-retail business acres per town
**chas**: Charles River dummy variable (1 if tract bounds river; 0 otherwise)
**nox**: nitric oxides concentration (parts per 10 million)
**rm**: average number of rooms per dwelling
**age**: proportion of owner-occupied units built prior to 1940
**dis**: weighted distances to five Boston employment centres
**rad**: index of accessibility to radial highways
**tax**: full-value property-tax rate per \$10,000
**ptratio**: pupil-teacher ratio by town
**b**: 1000(Bk - 0.63)^2 where Bk is the proportion of blacks by town
**lstat**: lower status of the population
**medv**: Median value of owner-occupied homes in $1000's
## 2. Simple statistics (10 points).
Print the basic statistics (mean and standard deviation) for the crime rate, nitric oxides concentration, pupil-teacher ratio and median value of owner-occupied homes.
## 3. Data visualisation (30 points).
### 3.1 Plot the histogram distribution for each data column (10 points).
### 3.2 Plot the correlation matrix between the data columns (10 points).
### 3.3 Plot the house price (the last data column) against each feature (each of the first 13 data columns) (10 points).
## 4. Linear regression (30 points).
### 4.1. Regress the house price against all the features (15 points).
* First, split the whole dataset into a training set and a test set using a pre-defined ratio (80:20 in this case).
* Then, train the linear regression model using the training set.
* Finally, plot the predicted house price on the training set.
The dataset split is provided for consistent evaluation. Please do not change the random_state seed.
```
X = df.iloc[:, :13]
y = df.iloc[:, 13]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
```
### 4.2 Quantitatively evaluate the linear model using the root of mean squared error (RMSE) and the R squared (R2) on both the training set and test set (15 points).
## 5. Challenge yourself (20 points)
Previously, we use 13 features to predict the house price. Perhaps some of the features are more relevant to the price, whereas some are less.
### 5.1 Explore the features and develop a linear model for house price prediction using only 5 features as input (10 points).
Hint: either using feature selection or dimensionality reduction.
### 5.2 Evaluate the quantitative performance of the new model in terms of RMSE and R2 on the test set (10 points).
## 6. Survey
How long did it take you to complete the coursework? What is your background and how you feel?
| github_jupyter |

[](https://colab.research.google.com/github/JohnSnowLabs/nlu/blob/master/examples/colab/component_examples/named_entity_recognition_(NER)/NLU_ner_ONTO_18class_example.ipynb.ipynb)
# Named-entity recognition with Deep Learning ONTO NOTES
Named-Entity recognition is a well-known technique in information extraction it is also known as entity identification, entity chunking and entity extraction. Knowing the relevant tags for each article help in automatically categorizing the articles in defined hierarchies and enable smooth content discovery. This pipeline is based on NerDLApproach annotator with Char CNN - BiLSTM and GloVe Embeddings on the OntoNotes corpus and supports the identification of 18 entities.
Following NER classes can be detected by this model
|Type | Description |
|------|--------------|
| PERSON | People, including fictional like **Harry Potter** |
| NORP | Nationalities or religious or political groups like the **Germans** |
| FAC | Buildings, airports, highways, bridges, etc. like **New York Airport** |
| ORG | Companies, agencies, institutions, etc. like **Microsoft** |
| GPE | Countries, cities, states. like **Germany** |
| LOC | Non-GPE locations, mountain ranges, bodies of water. Like the **Sahara desert**|
| PRODUCT | Objects, vehicles, foods, etc. (Not services.) like **playstation** |
| EVENT | Named hurricanes, battles, wars, sports events, etc. like **hurricane Katrina**|
| WORK_OF_ART | Titles of books, songs, etc. Like **Mona Lisa** |
| LAW | Named documents made into laws. Like : **Declaration of Independence** |
| LANGUAGE | Any named language. Like **Turkish**|
| DATE | Absolute or relative dates or periods. Like every second **friday**|
| TIME | Times smaller than a day. Like **every minute**|
| PERCENT | Percentage, including ”%“. Like **55%** of workers enjoy their work |
| MONEY | Monetary values, including unit. Like **50$** for those pants |
| QUANTITY | Measurements, as of weight or distance. Like this person weights **50kg** |
| ORDINAL | “first”, “second”, etc. Like David placed **first** in the tournament |
| CARDINAL | Numerals that do not fall under another type. Like **hundreds** of models are avaiable in NLU |
```
import os
! apt-get update -qq > /dev/null
# Install java
! apt-get install -y openjdk-8-jdk-headless -qq > /dev/null
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["PATH"] = os.environ["JAVA_HOME"] + "/bin:" + os.environ["PATH"]
! pip install nlu pyspark==2.4.7 > /dev/null
```
# NLU makes NER easy.
You just need to load the NER model via ner.load() and predict on some dataset.
It could be a pandas dataframe with a column named text or just an array of strings.
```
import nlu
example_text = ['People, including fictional like Harry Potter.',
'Nationalities or religious or political groups like Germans.',
'Buildings, airports, highways, bridges, etc. like New York Airport',
'Companies, agencies, institutions, etc. like Microsoft',
'Countries, cities, states. like Germany',
'Non-GPE locations, mountain ranges, bodies of water. Like Sahara Destert',
'Objects, vehicles, foods, etc. (Not services.) Like the a or playstation or Playstation',
'Named hurricanes, battles, wars, sports events, etc. like hurricane Katrina',
'Titles of books, songs, etc. Like the Mona Lisa',
'Named documents made into laws. Like the Declaration of Independence',
'Any named language. Like English',
'Absolute or relative dates or periods. Like every second friday',
'Times smaller than a day. Like every minute',
'Percentage, including ”%“. Like 55% of workers enjoy their work',
'Monetary values, including unit. Like 50$ for those pants',
'Measurements, as of weight or distance. Like this person weights 50kg',
'“first”, “second”, etc. Like David place first in the tournament',
'Numerals that do not fall under another type. Like hundreds of models are avaiable in NLU',]
nlu.load('ner.onto').predict(example_text)
text = ["Barclays misled shareholders and the public about one of the biggest investments in the bank's history, a BBC Panorama investigation has found.",
"The bank announced in 2008 that Manchester City owner Sheikh Mansour had agreed to invest more than £3bn.",
"But the BBC found that the money, which helped Barclays avoid a bailout by British taxpayers, actually came from the Abu Dhabi government.",
"Barclays said the mistake in its accounts was 'a drafting error'.",
"Unlike RBS and Lloyds TSB, Barclays narrowly avoided having to request a government bailout late in 2008 after it was rescued by £7bn worth of new investment, most of which came from the Gulf states of Qatar and Abu Dhabi.",
"The S&P 500's price to earnings multiple is 71% higher than Apple's, and if Apple were simply valued at the same multiple, its share price would be $840, which is 52% higher than its current price.",
"Alice has a cat named Alice and also a dog named Alice and also a parrot named Alice, it is her favorite name!"
] + example_text
ner_df = nlu.load('ner.onto').predict(text, output_level='chunk')
```
## Lets explore our data which the predicted NER tags and visalize them!
We specify [1:] so we dont se the count for the O-tag wich is the most common, since most words in a sentence are not named entities and thus not part of a chunk
```
ner_df['entities'].value_counts()[1:].plot.bar(title='Occurence of Named Entity tokens in dataset')
ner_type_to_viz = 'ORG'
ner_df[ner_df.entities_confidence == ner_type_to_viz]['entities'].value_counts().plot.bar(title='Most often occuring ORG labeled tokens in the dataset')
ner_type_to_viz = 'LOC'
ner_df[ner_df.entities_confidence == ner_type_to_viz]['entities'].value_counts().plot.bar(title='Most often occuring LOC labeled tokens in the dataset')
```
| github_jupyter |
# Logistic Regression with a Neural Network mindset
Welcome to your first (required) programming assignment! You will build a logistic regression classifier to recognize cats. This assignment will step you through how to do this with a Neural Network mindset, and so will also hone your intuitions about deep learning.
**Instructions:**
- Do not use loops (for/while) in your code, unless the instructions explicitly ask you to do so.
**You will learn to:**
- Build the general architecture of a learning algorithm, including:
- Initializing parameters
- Calculating the cost function and its gradient
- Using an optimization algorithm (gradient descent)
- Gather all three functions above into a main model function, in the right order.
## <font color='darkblue'>Updates</font>
This notebook has been updated over the past few months. The prior version was named "v5", and the current versionis now named '6a'
#### If you were working on a previous version:
* You can find your prior work by looking in the file directory for the older files (named by version name).
* To view the file directory, click on the "Coursera" icon in the top left corner of this notebook.
* Please copy your work from the older versions to the new version, in order to submit your work for grading.
#### List of Updates
* Forward propagation formula, indexing now starts at 1 instead of 0.
* Optimization function comment now says "print cost every 100 training iterations" instead of "examples".
* Fixed grammar in the comments.
* Y_prediction_test variable name is used consistently.
* Plot's axis label now says "iterations (hundred)" instead of "iterations".
* When testing the model, the test image is normalized by dividing by 255.
## 1 - Packages ##
First, let's run the cell below to import all the packages that you will need during this assignment.
- [numpy](www.numpy.org) is the fundamental package for scientific computing with Python.
- [h5py](http://www.h5py.org) is a common package to interact with a dataset that is stored on an H5 file.
- [matplotlib](http://matplotlib.org) is a famous library to plot graphs in Python.
- [PIL](http://www.pythonware.com/products/pil/) and [scipy](https://www.scipy.org/) are used here to test your model with your own picture at the end.
```
import numpy as np
import matplotlib.pyplot as plt
import h5py
import scipy
from PIL import Image
from scipy import ndimage
from lr_utils import load_dataset
%matplotlib inline
```
## 2 - Overview of the Problem set ##
**Problem Statement**: You are given a dataset ("data.h5") containing:
- a training set of m_train images labeled as cat (y=1) or non-cat (y=0)
- a test set of m_test images labeled as cat or non-cat
- each image is of shape (num_px, num_px, 3) where 3 is for the 3 channels (RGB). Thus, each image is square (height = num_px) and (width = num_px).
You will build a simple image-recognition algorithm that can correctly classify pictures as cat or non-cat.
Let's get more familiar with the dataset. Load the data by running the following code.
```
# Loading the data (cat/non-cat)
train_set_x_orig, train_set_y, test_set_x_orig, test_set_y, classes = load_dataset()
```
We added "_orig" at the end of image datasets (train and test) because we are going to preprocess them. After preprocessing, we will end up with train_set_x and test_set_x (the labels train_set_y and test_set_y don't need any preprocessing).
Each line of your train_set_x_orig and test_set_x_orig is an array representing an image. You can visualize an example by running the following code. Feel free also to change the `index` value and re-run to see other images.
```
# Example of a picture
index = 25
plt.imshow(train_set_x_orig[index])
print ("y = " + str(train_set_y[:, index]) + ", it's a '" + classes[np.squeeze(train_set_y[:, index])].decode("utf-8") + "' picture.")
```
Many software bugs in deep learning come from having matrix/vector dimensions that don't fit. If you can keep your matrix/vector dimensions straight you will go a long way toward eliminating many bugs.
**Exercise:** Find the values for:
- m_train (number of training examples)
- m_test (number of test examples)
- num_px (= height = width of a training image)
Remember that `train_set_x_orig` is a numpy-array of shape (m_train, num_px, num_px, 3). For instance, you can access `m_train` by writing `train_set_x_orig.shape[0]`.
```
### START CODE HERE ### (≈ 3 lines of code)
m_train = len(train_set_x_orig)
m_test = len(test_set_x_orig)
num_px = train_set_x_orig[25].shape[0]
### END CODE HERE ###
print ("Number of training examples: m_train = " + str(m_train))
print ("Number of testing examples: m_test = " + str(m_test))
print ("Height/Width of each image: num_px = " + str(num_px))
print ("Each image is of size: (" + str(num_px) + ", " + str(num_px) + ", 3)")
print ("train_set_x shape: " + str(train_set_x_orig.shape))
print ("train_set_y shape: " + str(train_set_y.shape))
print ("test_set_x shape: " + str(test_set_x_orig.shape))
print ("test_set_y shape: " + str(test_set_y.shape))
```
**Expected Output for m_train, m_test and num_px**:
<table style="width:15%">
<tr>
<td>**m_train**</td>
<td> 209 </td>
</tr>
<tr>
<td>**m_test**</td>
<td> 50 </td>
</tr>
<tr>
<td>**num_px**</td>
<td> 64 </td>
</tr>
</table>
For convenience, you should now reshape images of shape (num_px, num_px, 3) in a numpy-array of shape (num_px $*$ num_px $*$ 3, 1). After this, our training (and test) dataset is a numpy-array where each column represents a flattened image. There should be m_train (respectively m_test) columns.
**Exercise:** Reshape the training and test data sets so that images of size (num_px, num_px, 3) are flattened into single vectors of shape (num\_px $*$ num\_px $*$ 3, 1).
A trick when you want to flatten a matrix X of shape (a,b,c,d) to a matrix X_flatten of shape (b$*$c$*$d, a) is to use:
```python
X_flatten = X.reshape(X.shape[0], -1).T # X.T is the transpose of X
```
```
# Reshape the training and test examples
### START CODE HERE ### (≈ 2 lines of code)
train_set_x_flatten = train_set_x_orig.reshape(train_set_x_orig.shape[0], -1).T
test_set_x_flatten = test_set_x_orig.reshape(test_set_x_orig.shape[0], -1).T
### END CODE HERE ###
print ("train_set_x_flatten shape: " + str(train_set_x_flatten.shape))
print ("train_set_y shape: " + str(train_set_y.shape))
print ("test_set_x_flatten shape: " + str(test_set_x_flatten.shape))
print ("test_set_y shape: " + str(test_set_y.shape))
print ("sanity check after reshaping: " + str(train_set_x_flatten[0:5,0]))
```
**Expected Output**:
<table style="width:35%">
<tr>
<td>**train_set_x_flatten shape**</td>
<td> (12288, 209)</td>
</tr>
<tr>
<td>**train_set_y shape**</td>
<td>(1, 209)</td>
</tr>
<tr>
<td>**test_set_x_flatten shape**</td>
<td>(12288, 50)</td>
</tr>
<tr>
<td>**test_set_y shape**</td>
<td>(1, 50)</td>
</tr>
<tr>
<td>**sanity check after reshaping**</td>
<td>[17 31 56 22 33]</td>
</tr>
</table>
To represent color images, the red, green and blue channels (RGB) must be specified for each pixel, and so the pixel value is actually a vector of three numbers ranging from 0 to 255.
One common preprocessing step in machine learning is to center and standardize your dataset, meaning that you substract the mean of the whole numpy array from each example, and then divide each example by the standard deviation of the whole numpy array. But for picture datasets, it is simpler and more convenient and works almost as well to just divide every row of the dataset by 255 (the maximum value of a pixel channel).
<!-- During the training of your model, you're going to multiply weights and add biases to some initial inputs in order to observe neuron activations. Then you backpropogate with the gradients to train the model. But, it is extremely important for each feature to have a similar range such that our gradients don't explode. You will see that more in detail later in the lectures. !-->
Let's standardize our dataset.
```
train_set_x = train_set_x_flatten/255.
test_set_x = test_set_x_flatten/255.
```
<font color='blue'>
**What you need to remember:**
Common steps for pre-processing a new dataset are:
- Figure out the dimensions and shapes of the problem (m_train, m_test, num_px, ...)
- Reshape the datasets such that each example is now a vector of size (num_px \* num_px \* 3, 1)
- "Standardize" the data
## 3 - General Architecture of the learning algorithm ##
It's time to design a simple algorithm to distinguish cat images from non-cat images.
You will build a Logistic Regression, using a Neural Network mindset. The following Figure explains why **Logistic Regression is actually a very simple Neural Network!**
<img src="images/LogReg_kiank.png" style="width:650px;height:400px;">
**Mathematical expression of the algorithm**:
For one example $x^{(i)}$:
$$z^{(i)} = w^T x^{(i)} + b \tag{1}$$
$$\hat{y}^{(i)} = a^{(i)} = sigmoid(z^{(i)})\tag{2}$$
$$ \mathcal{L}(a^{(i)}, y^{(i)}) = - y^{(i)} \log(a^{(i)}) - (1-y^{(i)} ) \log(1-a^{(i)})\tag{3}$$
The cost is then computed by summing over all training examples:
$$ J = \frac{1}{m} \sum_{i=1}^m \mathcal{L}(a^{(i)}, y^{(i)})\tag{6}$$
**Key steps**:
In this exercise, you will carry out the following steps:
- Initialize the parameters of the model
- Learn the parameters for the model by minimizing the cost
- Use the learned parameters to make predictions (on the test set)
- Analyse the results and conclude
## 4 - Building the parts of our algorithm ##
The main steps for building a Neural Network are:
1. Define the model structure (such as number of input features)
2. Initialize the model's parameters
3. Loop:
- Calculate current loss (forward propagation)
- Calculate current gradient (backward propagation)
- Update parameters (gradient descent)
You often build 1-3 separately and integrate them into one function we call `model()`.
### 4.1 - Helper functions
**Exercise**: Using your code from "Python Basics", implement `sigmoid()`. As you've seen in the figure above, you need to compute $sigmoid( w^T x + b) = \frac{1}{1 + e^{-(w^T x + b)}}$ to make predictions. Use np.exp().
```
# GRADED FUNCTION: sigmoid
def sigmoid(z):
"""
Compute the sigmoid of z
Arguments:
z -- A scalar or numpy array of any size.
Return:
s -- sigmoid(z)
"""
### START CODE HERE ### (≈ 1 line of code)
s = 1 / (1 + np.exp(-z))
### END CODE HERE ###
return s
print ("sigmoid([0, 2]) = " + str(sigmoid(np.array([0,2]))))
```
**Expected Output**:
<table>
<tr>
<td>**sigmoid([0, 2])**</td>
<td> [ 0.5 0.88079708]</td>
</tr>
</table>
### 4.2 - Initializing parameters
**Exercise:** Implement parameter initialization in the cell below. You have to initialize w as a vector of zeros. If you don't know what numpy function to use, look up np.zeros() in the Numpy library's documentation.
```
# GRADED FUNCTION: initialize_with_zeros
def initialize_with_zeros(dim):
"""
This function creates a vector of zeros of shape (dim, 1) for w and initializes b to 0.
Argument:
dim -- size of the w vector we want (or number of parameters in this case)
Returns:
w -- initialized vector of shape (dim, 1)
b -- initialized scalar (corresponds to the bias)
"""
### START CODE HERE ### (≈ 1 line of code)
w = np.zeros((dim, 1))
b = 0
### END CODE HERE ###
assert(w.shape == (dim, 1))
assert(isinstance(b, float) or isinstance(b, int))
return w, b
dim = 2
w, b = initialize_with_zeros(dim)
print ("w = " + str(w))
print ("b = " + str(b))
```
**Expected Output**:
<table style="width:15%">
<tr>
<td> ** w ** </td>
<td> [[ 0.]
[ 0.]] </td>
</tr>
<tr>
<td> ** b ** </td>
<td> 0 </td>
</tr>
</table>
For image inputs, w will be of shape (num_px $\times$ num_px $\times$ 3, 1).
### 4.3 - Forward and Backward propagation
Now that your parameters are initialized, you can do the "forward" and "backward" propagation steps for learning the parameters.
**Exercise:** Implement a function `propagate()` that computes the cost function and its gradient.
**Hints**:
Forward Propagation:
- You get X
- You compute $A = \sigma(w^T X + b) = (a^{(1)}, a^{(2)}, ..., a^{(m-1)}, a^{(m)})$
- You calculate the cost function: $J = -\frac{1}{m}\sum_{i=1}^{m}y^{(i)}\log(a^{(i)})+(1-y^{(i)})\log(1-a^{(i)})$
Here are the two formulas you will be using:
$$ \frac{\partial J}{\partial w} = \frac{1}{m}X(A-Y)^T\tag{7}$$
$$ \frac{\partial J}{\partial b} = \frac{1}{m} \sum_{i=1}^m (a^{(i)}-y^{(i)})\tag{8}$$
```
# GRADED FUNCTION: propagate
def propagate(w, b, X, Y):
"""
Implement the cost function and its gradient for the propagation explained above
Arguments:
w -- weights, a numpy array of size (num_px * num_px * 3, 1)
b -- bias, a scalar
X -- data of size (num_px * num_px * 3, number of examples)
Y -- true "label" vector (containing 0 if non-cat, 1 if cat) of size (1, number of examples)
Return:
cost -- negative log-likelihood cost for logistic regression
dw -- gradient of the loss with respect to w, thus same shape as w
db -- gradient of the loss with respect to b, thus same shape as b
Tips:
- Write your code step by step for the propagation. np.log(), np.dot()
"""
m = X.shape[1]
# FORWARD PROPAGATION (FROM X TO COST)
### START CODE HERE ### (≈ 2 lines of code)
A = sigmoid(np.dot(w.T, X) + b) # compute activation
cost = -1/m * np.sum( (Y*np.log(A) + (1-Y)*np.log(1-A)), axis=1, keepdims=True) # compute cost
### END CODE HERE ###
# BACKWARD PROPAGATION (TO FIND GRAD)
### START CODE HERE ### (≈ 2 lines of code)
dw = 1/m * np.dot(X, (A-Y).T)
db = 1/m * np.sum((A-Y).T)
### END CODE HERE ###
assert(dw.shape == w.shape)
assert(db.dtype == float)
cost = np.squeeze(cost)
assert(cost.shape == ())
grads = {"dw": dw,
"db": db}
return grads, cost
w, b, X, Y = np.array([[1.],[2.]]), 2., np.array([[1.,2.,-1.],[3.,4.,-3.2]]), np.array([[1,0,1]])
grads, cost = propagate(w, b, X, Y)
print ("dw = " + str(grads["dw"]))
print ("db = " + str(grads["db"]))
print ("cost = " + str(cost))
```
**Expected Output**:
<table style="width:50%">
<tr>
<td> ** dw ** </td>
<td> [[ 0.99845601]
[ 2.39507239]]</td>
</tr>
<tr>
<td> ** db ** </td>
<td> 0.00145557813678 </td>
</tr>
<tr>
<td> ** cost ** </td>
<td> 5.801545319394553 </td>
</tr>
</table>
### 4.4 - Optimization
- You have initialized your parameters.
- You are also able to compute a cost function and its gradient.
- Now, you want to update the parameters using gradient descent.
**Exercise:** Write down the optimization function. The goal is to learn $w$ and $b$ by minimizing the cost function $J$. For a parameter $\theta$, the update rule is $ \theta = \theta - \alpha \text{ } d\theta$, where $\alpha$ is the learning rate.
```
# GRADED FUNCTION: optimize
def optimize(w, b, X, Y, num_iterations, learning_rate, print_cost = False):
"""
This function optimizes w and b by running a gradient descent algorithm
Arguments:
w -- weights, a numpy array of size (num_px * num_px * 3, 1)
b -- bias, a scalar
X -- data of shape (num_px * num_px * 3, number of examples)
Y -- true "label" vector (containing 0 if non-cat, 1 if cat), of shape (1, number of examples)
num_iterations -- number of iterations of the optimization loop
learning_rate -- learning rate of the gradient descent update rule
print_cost -- True to print the loss every 100 steps
Returns:
params -- dictionary containing the weights w and bias b
grads -- dictionary containing the gradients of the weights and bias with respect to the cost function
costs -- list of all the costs computed during the optimization, this will be used to plot the learning curve.
Tips:
You basically need to write down two steps and iterate through them:
1) Calculate the cost and the gradient for the current parameters. Use propagate().
2) Update the parameters using gradient descent rule for w and b.
"""
costs = []
for i in range(num_iterations):
# Cost and gradient calculation (≈ 1-4 lines of code)
### START CODE HERE ###
grads, cost = propagate(w, b, X, Y)
### END CODE HERE ###
# Retrieve derivatives from grads
dw = grads["dw"]
db = grads["db"]
# update rule (≈ 2 lines of code)
### START CODE HERE ###
w = w - learning_rate*dw
b = b - learning_rate*db
### END CODE HERE ###
# Record the costs
if i % 100 == 0:
costs.append(cost)
# Print the cost every 100 training iterations
if print_cost and i % 100 == 0:
print ("Cost after iteration %i: %f" %(i, cost))
params = {"w": w,
"b": b}
grads = {"dw": dw,
"db": db}
return params, grads, costs
params, grads, costs = optimize(w, b, X, Y, num_iterations= 100, learning_rate = 0.009, print_cost = False)
print ("w = " + str(params["w"]))
print ("b = " + str(params["b"]))
print ("dw = " + str(grads["dw"]))
print ("db = " + str(grads["db"]))
```
**Expected Output**:
<table style="width:40%">
<tr>
<td> **w** </td>
<td>[[ 0.19033591]
[ 0.12259159]] </td>
</tr>
<tr>
<td> **b** </td>
<td> 1.92535983008 </td>
</tr>
<tr>
<td> **dw** </td>
<td> [[ 0.67752042]
[ 1.41625495]] </td>
</tr>
<tr>
<td> **db** </td>
<td> 0.219194504541 </td>
</tr>
</table>
**Exercise:** The previous function will output the learned w and b. We are able to use w and b to predict the labels for a dataset X. Implement the `predict()` function. There are two steps to computing predictions:
1. Calculate $\hat{Y} = A = \sigma(w^T X + b)$
2. Convert the entries of a into 0 (if activation <= 0.5) or 1 (if activation > 0.5), stores the predictions in a vector `Y_prediction`. If you wish, you can use an `if`/`else` statement in a `for` loop (though there is also a way to vectorize this).
```
# GRADED FUNCTION: predict
def predict(w, b, X):
'''
Predict whether the label is 0 or 1 using learned logistic regression parameters (w, b)
Arguments:
w -- weights, a numpy array of size (num_px * num_px * 3, 1)
b -- bias, a scalar
X -- data of size (num_px * num_px * 3, number of examples)
Returns:
Y_prediction -- a numpy array (vector) containing all predictions (0/1) for the examples in X
'''
m = X.shape[1]
Y_prediction = np.zeros((1, m))
w = w.reshape(X.shape[0], 1)
# Compute vector "A" predicting the probabilities of a cat being present in the picture
### START CODE HERE ### (≈ 1 line of code)
A = sigmoid(np.dot(w.T, X) + b)
### END CODE HERE ###
for i in range(A.shape[1]):
# Convert probabilities a[0,i] to actual predictions p[0,i]
### START CODE HERE ### (≈ 4 lines of code)
if A[0, i] >=0.5:
Y_prediction[0, i] = 1
elif A[0, i] <0.5:
Y_prediction[0, i] = 0
### END CODE HERE ###
assert(Y_prediction.shape == (1, m))
return Y_prediction
w = np.array([[0.1124579],[0.23106775]])
b = -0.3
X = np.array([[1.,-1.1,-3.2],[1.2,2.,0.1]])
print ("predictions = " + str(predict(w, b, X)))
```
**Expected Output**:
<table style="width:30%">
<tr>
<td>
**predictions**
</td>
<td>
[[ 1. 1. 0.]]
</td>
</tr>
</table>
<font color='blue'>
**What to remember:**
You've implemented several functions that:
- Initialize (w,b)
- Optimize the loss iteratively to learn parameters (w,b):
- computing the cost and its gradient
- updating the parameters using gradient descent
- Use the learned (w,b) to predict the labels for a given set of examples
## 5 - Merge all functions into a model ##
You will now see how the overall model is structured by putting together all the building blocks (functions implemented in the previous parts) together, in the right order.
**Exercise:** Implement the model function. Use the following notation:
- Y_prediction_test for your predictions on the test set
- Y_prediction_train for your predictions on the train set
- w, costs, grads for the outputs of optimize()
```
# GRADED FUNCTION: model
def model(X_train, Y_train, X_test, Y_test, num_iterations = 2000, learning_rate = 0.5, print_cost = False):
"""
Builds the logistic regression model by calling the function you've implemented previously
Arguments:
X_train -- training set represented by a numpy array of shape (num_px * num_px * 3, m_train)
Y_train -- training labels represented by a numpy array (vector) of shape (1, m_train)
X_test -- test set represented by a numpy array of shape (num_px * num_px * 3, m_test)
Y_test -- test labels represented by a numpy array (vector) of shape (1, m_test)
num_iterations -- hyperparameter representing the number of iterations to optimize the parameters
learning_rate -- hyperparameter representing the learning rate used in the update rule of optimize()
print_cost -- Set to true to print the cost every 100 iterations
Returns:
d -- dictionary containing information about the model.
"""
### START CODE HERE ###
# initialize parameters with zeros (≈ 1 line of code)
w, b = initialize_with_zeros(dim = X_train.shape[0])
# Gradient descent (≈ 1 line of code)
parameters, grads, costs = optimize(w, b, X_train, Y_train, num_iterations, learning_rate, print_cost)
# Retrieve parameters w and b from dictionary "parameters"
w = parameters["w"]
b = parameters["b"]
# Predict test/train set examples (≈ 2 lines of code)
Y_prediction_test = predict(w, b, X_test)
Y_prediction_train = predict(w, b, X_train)
### END CODE HERE ###
# Print train/test Errors
print("train accuracy: {} %".format(100 - np.mean(np.abs(Y_prediction_train - Y_train)) * 100))
print("test accuracy: {} %".format(100 - np.mean(np.abs(Y_prediction_test - Y_test)) * 100))
d = {"costs": costs,
"Y_prediction_test": Y_prediction_test,
"Y_prediction_train" : Y_prediction_train,
"w" : w,
"b" : b,
"learning_rate" : learning_rate,
"num_iterations": num_iterations}
return d
```
Run the following cell to train your model.
```
d = model(train_set_x, train_set_y, test_set_x, test_set_y, num_iterations = 2000, learning_rate = 0.005, print_cost = True)
```
**Expected Output**:
<table style="width:40%">
<tr>
<td> **Cost after iteration 0 ** </td>
<td> 0.693147 </td>
</tr>
<tr>
<td> <center> $\vdots$ </center> </td>
<td> <center> $\vdots$ </center> </td>
</tr>
<tr>
<td> **Train Accuracy** </td>
<td> 99.04306220095694 % </td>
</tr>
<tr>
<td>**Test Accuracy** </td>
<td> 70.0 % </td>
</tr>
</table>
**Comment**: Training accuracy is close to 100%. This is a good sanity check: your model is working and has high enough capacity to fit the training data. Test accuracy is 68%. It is actually not bad for this simple model, given the small dataset we used and that logistic regression is a linear classifier. But no worries, you'll build an even better classifier next week!
Also, you see that the model is clearly overfitting the training data. Later in this specialization you will learn how to reduce overfitting, for example by using regularization. Using the code below (and changing the `index` variable) you can look at predictions on pictures of the test set.
```
# Example of a picture that was wrongly classified.
index = 1
plt.imshow(test_set_x[:,index].reshape((num_px, num_px, 3)))
print ("y = " + str(test_set_y[0,index]) + ", you predicted that it is a \"" + classes[d["Y_prediction_test"][0,index]].decode("utf-8") + "\" picture.")
```
Let's also plot the cost function and the gradients.
```
# Plot learning curve (with costs)
costs = np.squeeze(d['costs'])
plt.plot(costs)
plt.ylabel('cost')
plt.xlabel('iterations (per hundreds)')
plt.title("Learning rate =" + str(d["learning_rate"]))
plt.show()
```
**Interpretation**:
You can see the cost decreasing. It shows that the parameters are being learned. However, you see that you could train the model even more on the training set. Try to increase the number of iterations in the cell above and rerun the cells. You might see that the training set accuracy goes up, but the test set accuracy goes down. This is called overfitting.
## 6 - Further analysis (optional/ungraded exercise) ##
Congratulations on building your first image classification model. Let's analyze it further, and examine possible choices for the learning rate $\alpha$.
#### Choice of learning rate ####
**Reminder**:
In order for Gradient Descent to work you must choose the learning rate wisely. The learning rate $\alpha$ determines how rapidly we update the parameters. If the learning rate is too large we may "overshoot" the optimal value. Similarly, if it is too small we will need too many iterations to converge to the best values. That's why it is crucial to use a well-tuned learning rate.
Let's compare the learning curve of our model with several choices of learning rates. Run the cell below. This should take about 1 minute. Feel free also to try different values than the three we have initialized the `learning_rates` variable to contain, and see what happens.
```
learning_rates = [0.01, 0.001, 0.0001]
models = {}
for i in learning_rates:
print ("learning rate is: " + str(i))
models[str(i)] = model(train_set_x, train_set_y, test_set_x, test_set_y, num_iterations = 1500, learning_rate = i, print_cost = False)
print ('\n' + "-------------------------------------------------------" + '\n')
for i in learning_rates:
plt.plot(np.squeeze(models[str(i)]["costs"]), label= str(models[str(i)]["learning_rate"]))
plt.ylabel('cost')
plt.xlabel('iterations (hundreds)')
legend = plt.legend(loc='upper center', shadow=True)
frame = legend.get_frame()
frame.set_facecolor('0.90')
plt.show()
```
**Interpretation**:
- Different learning rates give different costs and thus different predictions results.
- If the learning rate is too large (0.01), the cost may oscillate up and down. It may even diverge (though in this example, using 0.01 still eventually ends up at a good value for the cost).
- A lower cost doesn't mean a better model. You have to check if there is possibly overfitting. It happens when the training accuracy is a lot higher than the test accuracy.
- In deep learning, we usually recommend that you:
- Choose the learning rate that better minimizes the cost function.
- If your model overfits, use other techniques to reduce overfitting. (We'll talk about this in later videos.)
## 7 - Test with your own image (optional/ungraded exercise) ##
Congratulations on finishing this assignment. You can use your own image and see the output of your model. To do that:
1. Click on "File" in the upper bar of this notebook, then click "Open" to go on your Coursera Hub.
2. Add your image to this Jupyter Notebook's directory, in the "images" folder
3. Change your image's name in the following code
4. Run the code and check if the algorithm is right (1 = cat, 0 = non-cat)!
```
## START CODE HERE ## (PUT YOUR IMAGE NAME)
my_image = "my_image.jpg" # change this to the name of your image file
## END CODE HERE ##
# We preprocess the image to fit your algorithm.
fname = "images/" + my_image
image = np.array(ndimage.imread(fname, flatten=False))
image = image/255.
my_image = scipy.misc.imresize(image, size=(num_px,num_px)).reshape((1, num_px*num_px*3)).T
my_predicted_image = predict(d["w"], d["b"], my_image)
plt.imshow(image)
print("y = " + str(np.squeeze(my_predicted_image)) + ", your algorithm predicts a \"" + classes[int(np.squeeze(my_predicted_image)),].decode("utf-8") + "\" picture.")
```
<font color='blue'>
**What to remember from this assignment:**
1. Preprocessing the dataset is important.
2. You implemented each function separately: initialize(), propagate(), optimize(). Then you built a model().
3. Tuning the learning rate (which is an example of a "hyperparameter") can make a big difference to the algorithm. You will see more examples of this later in this course!
Finally, if you'd like, we invite you to try different things on this Notebook. Make sure you submit before trying anything. Once you submit, things you can play with include:
- Play with the learning rate and the number of iterations
- Try different initialization methods and compare the results
- Test other preprocessings (center the data, or divide each row by its standard deviation)
Bibliography:
- http://www.wildml.com/2015/09/implementing-a-neural-network-from-scratch/
- https://stats.stackexchange.com/questions/211436/why-do-we-normalize-images-by-subtracting-the-datasets-image-mean-and-not-the-c
| github_jupyter |
# Machine Learning Workshop
Here we will walk through an example of a machine learning workflow following five steps:
<img src="../_img/ml_workflow.png" alt="ML Workflow" width="800"/>
For more detailed information on the Shiu Lab's ML pipeline, including explanations of all output files,
check out the [README](https://github.com/ShiuLab/ML-Pipeline).
***
## Step 0. Set up Jupyter notebook & software
Check out this [**guide**](https://github.com/ShiuLab/ML-Pipeline/tree/master/Workshop) to learn how to set up Jupyter notebook and the software needed to run the Shiu Lab's ML pipeline.
***

**What do we want to predict?**
If a gene is annotated as being involved in specialized or general metabolism.
**What are the labeled instances?**
Tomato genes annotated as being involved in specialized or general metabolism by TomatoCyc.
**What are the predictive features?**
- duplication information (e.g. number of paralogs, gene family size)
- sequence conservation (e.g. nonsynonymous/synonymouse substitution rates between homologs)
- gene expression (e.g. breadth, stress specific, co-expression)
- protein domain conent (e.g. p450, Aldedh)
- epigenetic modification (e.g. H3K23ac histone marks)
- network properties (# protein-protein interactions, network connectivity).
**What data do we have?**
- 532 tomato genes with specialized metabolism annotation by TomatoCyc
- 2,318 tomato genes with general metabolism annotation by TomatoCyc
- 4,197 features (we are only using a subset of **564** for this workshop)
***

```
## A. Lets look at the data (note, you can do this in excel or R!)
import pandas as pd
d = pd.read_table('data.txt', sep='\t', index_col = 0)
print('Shape of data (rows, cols):')
print(d.shape)
print('\nSnapshot of data:')
print(d.iloc[:6,:5]) # prints first 6 rows and 5 columns
print('\nList of class labels')
print(d['Class'].value_counts())
```
**Things to notice:**
- Our data has NAs. ML algorithms cannot handel NAs. We either needs to drop or impute NA values!
- We have binary, continuous, and categorical features in this dataset. A perk of ML models is that they can integrate multiple datatypes in a single model.
- However, before being used as input, a categorical feature needs to be converted into set binary features using an approach called [one-hot-encoding](https://www.kaggle.com/dansbecker/using-categorical-data-with-one-hot-encoding).
*Before One-Hot Encoding:*
| ID | Class | Weather |
|--- |--- |--- |
| instance_A | 1 | sunny |
| instance_B | 0 | overcast |
| instance_C | 0 | rain |
| instance_D | 1 | sunny |
*After One-Hot Encoding:*
| ID | Class | Weather_sunny | Weather_overcast | Weather_rain |
|--- |--- |--- |--- |--- |
| instance_A | 1 | 1 | 0 | 0 |
| instance_B | 0 | 0 | 1 | 0 |
| instance_C | 0 | 0 | 0 | 1 |
| instance_D | 1 | 1 | 0 | 0 |
***
### Automated data cleaning: ML_preprocess.py
Input
```
-df: your data table
-na_method: how you want to impute NAs (options: drop, mean, median, mode)
-h: show more options
```
```
# B. Drop/Impute NAs and one-hot-encode categorical features
%run ../ML_preprocess.py -df data.txt -na_method median
```
***
## Set aside instances for testing
We want to set aside a subset of our data to use to test how well our model performed. Note that this is done before feature engineering, parameter selection, or model training. This will ensure our performance metric is entirely independent from our modeling!
### Automated selection of test set: test_set.py
Input:
```
-df: your data table
-use: what class labels to include in the test set (we don't want to include unknowns!)
-type: (c) classification or (r) regression
-p: What percent of instances from each class to select for test (0.1 = 10%)
-save: save name for test set
```
```
# C. Define test set
%run ../test_set.py -df data_mod.txt \
-use gen,special \
-type c \
-p 0.1 \
-save test_genes.txt
```
***

While one major advantage of ML approaches is that they are robust when the number of features is very large, there are cases where removing unuseful features or selecting only the best features may help you better answer your question. One common issue we see with using feature selection for machine learning is using the whole dataset to select the best features, which results in overfitting! **Be sure you specify your test set so that this data is not used for feature selection!**
### Automated feature selection: Feature_Selection.py
Input
```
-df: your data table
-test: what instances to hold out (i.e. test instances!)
-cl_train: labels to include in training the feature selection algorithm
-type: (c) classification or (r) regression
-alg: what feature selection algorithm to use (e.g. lasso, elastic net, random forest)
-p: Parameter specific to different algorithms (use -h for more information)
-n: Number of feature to select (unless algorithm does this automatically)
-save: save name for list of selected features
```
Here we will use one of the most common feature selection algorithms: LASSO. LASSO requires the user to select the level of sparcity (-p) they want to induce during feature selection, where a larger value will result in more features being selected and a smaller value resulting in fewer features being selected. You can play around with this value to see what it does for your data.
```
%run ../Feature_Selection.py -df data_mod.txt \
-test test_genes.txt \
-cl_train special,gen \
-type c \
-alg lasso \
-p 0.01 \
-save top_feat_lasso.txt
%run ../Feature_Selection.py -df data_mod.txt \
-test test_genes.txt \
-cl_train special,gen \
-type c \
-alg random \
-n 10 \
-save rand_feat.txt
```
***

Next we want to determine which ML algorithm we should use and what combination of hyperparameters for those algorithms work best. Importantly, at this stage we **only assess our model performance on the validation data** in order to assure we aren't just selecting the algorithm that works best on our held out testing data. The pipeline will automatically withhold the testing data from the parameter selection (i.e. grid search) step.
Note, the pipeline **automatically "balances" your data**, meaning it pulls the same number of instances of each class for training. This avoids biasing the model to just predict everything as the more common class. This is a major reason why we want to run multiple replicates of the model!
### Algorithm Selection
The machine learning algorithms in the ML_Pipeline are implement from [SciKit-Learn](https://scikit-learn.org/stable/), which has excellent resources to learn more about the ins and outs of these algorithms.
**Why is algorithm selection useful?** ML models are able to learn patterns from data without the being explictely programmed to look for those patterns. ML algorithms differ in what patterns they excel at finding. For example, SVM is limited to linear relationships between feature and labels, while RF, because of its heiarchical structure, is able to model interactive patterns between your features. Furthermore, algorithms vary in their complexity and the amount of training data that is needed in order to
### [Hyper]-Parameter Selection
Most ML algorithms have internal parameters that need to be set by the user. For example:

There are two general strategies for parameter selection: the grid search (default option: left) and the random search (use "-gs_type random": right):

*Image: Bergstra & Bengio 2012; used under CC-BY license*
### Automated Training and Validation
Training and validation is done using a [cross-validation (CV)](https://towardsdatascience.com/cross-validation-70289113a072) scheme. CV is useful because it makes good use of our data (i.e. uses all non-test data for training at some point) but also makes sure we are selecting the best parameters/algorithms on models that aren't overfit to the training data. Here is a visual to demonstrate how CV works (with 10-cv folds in this example):

### ML_classification.py (similar to ML_regression.py)
**Input:**
```
-df: your data table
-test: what instances to hold out (i.e. test instances)
-cl_train: labels to include in training the feature selection algorithm
-alg: what ML algorithm to use (e.g. SVM, RF, LogReg (classification only), LR (regression only))
-cv: Number of cross-validation folds (default = 10, use fewer if data set is small)
-n: Number of replicates of the balanced cross-validation scheme to run (default = 100)
-save: Name to save output to (will over-write old files)
```
*There are many functions available within the pipeline that are not described in this workshop. For more options run:*
```
python ML_classification.py -h
```
```
%run ../ML_classification.py -df data_mod.txt \
-test test_genes.txt \
-cl_train special,gen \
-alg SVM \
-cv 5 \
-n 10 \
-save metab_SVM
```
#### Results Breakdown
There are dozens of [performance metrics](https://scikit-learn.org/stable/modules/model_evaluation.html) that can be used to assess how well a ML model works. While the best metric for you depends on the type of question you are asking, some of the most generally useful metrics include the area under the Receiver Operator Characteristic curve (AUC-ROC), the area under the Precision-Recall curve (AUC_PRc), and the F-measure (F1).

Running the same script (only changing **-alg XXX**), average performance on the validation data using other algorithms:
| Alg | F1 | AUC-ROC |
|--- |--- |--- |
| RF | 0.787 | 0.824 |
| SVMpoly | 0.833 | 0.897 |
| SVMrbf | 0.855 | 0.905 |
| SVM | 0.856 | 0.911 |
***SVM performed best on the validation data so we will continue with that algorithm!***

Now that we have our best performing algorithm, we will run the pipeline one more time, but with more replicates (note, I still just use 10 here for time!) and we will use it to predict our unknown genes.
**Additional input:**
```
- apply: List of lable names to apply trained model to (i.e. all, or 'unknown')
- plots: True/False if you want the pipeline to generate performance metric plots (default = F)
```
```
%run ../ML_classification.py -df data_mod.txt \
-test test_genes.txt \
-cl_train special,gen \
-alg SVM \
-cv 5 \
-n 10 \
-apply unknown \
-plots T \
-save metab_SVM
```
**Let's check out our results...**
Here are the files that are output from the model:
- **data.txt_results:** A detailed look at the model that was run and its performance.
- **data.txt_scores:** The probability score for each gene (i.e. how confidently it was predicted) and the final classification for each gene, including the unknowns the model was applied to.
- **data.txt_imp:** The importance of each feature in your model.
- **data.txt_GridSearch:** Detailed results from the parameter grid search.
- **data.txt_BalancedID:** A list of the genes that were included in each replicate after downsampling to balance the model.
*For a detailed description of the content of the pipeline output see the [README](../README.md)*
***
## What if we use fewer features?
Additional input:
```
- feat: List of features to use.
```
```
%run ../ML_classification.py -df data_mod.txt \
-test test_genes.txt \
-cl_train special,gen \
-alg SVM \
-cv 5 \
-n 10 \
-feat top_feat_lasso.txt \
-save metab_SVM_lasso10
%run ../ML_classification.py -df data_mod.txt \
-test test_genes.txt \
-cl_train special,gen \
-alg SVM \
-cv 5 \
-n 10 \
-feat rand_feat.txt_11 \
-save metab_SVM_rand
```
### Visualizing Your Results
There are a number of vizualization tools available in the ML-Pipeline (see ML_Postprocessing). Here we will use ML_plots.
**ML_plots.py input:**
```
-save: Name to save output figures
-cl_train: positive and negative classes
-names: short names to call each model being included
-scores: path to name_scores.txt files to include
```
```
%run ../scripts_PostAnalysis/ML_plots.py -save compare_SVM \
-cl_train special gen \
-names All LASSO Random \
-scores metab_SVM_scores.txt metab_SVM_lasso10_scores.txt metab_SVM_rand_scores.txt
```
***
## Final Thoughts
Here we went through one example of how the ML pipeline can be used to automate a machine learning experiment. There are numerous advanced features included into the pipeline that were not covered. Run any script in the pipeline with no arguments (i.e. -arg) to see more options.
**Some advanced options include:**
- multi-class classification
- transfer learning
- comparing instance classification across models
- single feature ML models
| github_jupyter |
# Classification modeling
---
Working with interpolated data!!!
```
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import random
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split, GridSearchCV, cross_val_score
from sklearn.preprocessing import StandardScaler, PolynomialFeatures
from sklearn.pipeline import Pipeline
from sklearn.metrics import confusion_matrix, ConfusionMatrixDisplay, plot_confusion_matrix, classification_report, plot_roc_curve
from sklearn.tree import DecisionTreeClassifier, plot_tree, export_text
random.seed(42)
```
---
### Load the data
```
df = pd.read_csv('../../coastal_upwelling_output/interpolated.csv')
df.rename({'Unnamed: 0':'time'},inplace=True, axis=1)
# df.set_index('time', inplace=True)
df
df['upwelling'].value_counts(normalize=True)
df.isna().sum()
```
---
### Checking feature correlation
One of the big assumptions we make when building logistic regression models is that our independent features are independent of each other. We can print out a heatmap to check whether our features are correlated to each other or not.
```
plt.figure(figsize=(12,12))
sns.heatmap(df.corr()[['CUTI']].sort_values(by='CUTI', ascending=False),
annot=True);
```
Are the deeper depths correlated here because they change so little?
---
### Using PolynomialFeatures
Now let's add those features back in and use feature interactions combined with regularization to try upping the accuracy while accounting for the multicollinearity.
An easy way to get a variety of feature interactions is using sklearn's PolynomialFeatures function. There are four features in this model, so I'll set the degree to 4 so that there will be an engineered feature that includes all 4 of the original features.
```
X = df.drop(columns=['upwelling', 'time', 'CUTI'])
y = df['upwelling']
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y, random_state=42)
pipe = Pipeline([
('sc', StandardScaler()),
('logreg', LogisticRegression(max_iter=1000, solver='liblinear'))
])
pipe_params = {
'logreg__penalty':['l1', 'l2'],
'logreg__C': np.linspace(0.001, 1, 10)
}
gs_lr = GridSearchCV(pipe, pipe_params, cv=5, verbose=1, return_train_score=True)
gs_lr.fit(X_train, y_train)
print(f'Best parameters: {gs_lr.best_params_}')
print(f'Best score: {gs_lr.best_score_}')
```
Now that we have the best parameters, we can create a logistic regression model with these parameters and see what the coefficients are for our poly features.
```
print(gs_lr.cv_results_['mean_train_score'].mean())
print(gs_lr.cv_results_['mean_test_score'].mean())
print(f'Train accuracy: {gs_lr.score(X_train, y_train)}')
print(f'Test accuracy: {gs_lr.score(X_test, y_test)}')
```
Question to self: do the coefficients need to be exponentiated to get their actual values, since the logistic regression model uses the logit function to transform the data?
```
gs_lr.predict(X_train)
gs_lr.best_estimator_['logreg'].coef_
coefs = gs_lr.best_estimator_['logreg'].coef_[0]
coefs = np.exp(coefs)
sorted(list(zip(X.columns, coefs)), key=lambda x: x[1], reverse=True)
gs_lr_train_preds = gs_lr.predict(X_train)
gs_lr_test_preds = gs_lr.predict(X_test)
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 8))
cm = confusion_matrix(y_train, gs_lr_train_preds)
ConfusionMatrixDisplay(cm).plot(ax=ax1)
ax1.set_title('Confusion Matrix: Train Data')
cm = confusion_matrix(y_test, gs_lr_test_preds)
ConfusionMatrixDisplay(cm).plot(ax=ax2)
ax2.set_title('Confusion Matrix: Test Data');
```
Looks like our false negatives outnumber our false positives
```
print(classification_report(y_test, gs_lr_test_preds))
# ROC curve
plot_roc_curve(gs_lr, X_test, y_test)
# add worst case scenario line
plt.plot([0,1],[0,1], label='baseline', linestyle='--')
# add a legend
plt.legend();
# want AUC (area under curve) to be as close to 1 as possible
```
#### Explore misclassified data
```
# Get indices of misclassified data source: https://stackoverflow.com/questions/25551977/retrieve-misclassified-documents-using-scikitlearn
misclass_ind_lr = np.where(y_test != gs_lr_test_preds)
misclass_ind_lr
X_test
df
df.iloc[X_test.index]['time']
X_test_times = df.iloc[X_test.index]['time']
X_test_times.iloc[misclass_ind_lr]
pd.DataFrame(X_test_times.iloc[misclass_ind_lr]).reset_index(drop=True)
frames_lr = [pd.DataFrame(X_test_times.iloc[misclass_ind_lr]), X_test.iloc[misclass_ind_lr], pd.DataFrame(y_test.iloc[misclass_ind_lr])]
misclass_df_lr = pd.concat(frames_lr, axis=1)
misclass_df_lr
```
---
### Decision tree classifiers
Decision trees come in a lot of different shapes, so it'd be best to use GridSearchCV to find the best parameters for a tree for upwelling classification.
```
X = df.drop(columns=['time', 'upwelling', 'CUTI'])
y = df['upwelling']
X_train, X_test, y_train, y_test = train_test_split(X, y, stratify=y, random_state=42)
param_grid = {
'max_depth': [5, 7, 9],
'min_samples_split': [5, 10, 15, 20],
'min_samples_leaf': [2, 3, 4, 5, 6],
'ccp_alpha': [0, 0.01, 0.1, 1, 10]
}
gs_dt = GridSearchCV(estimator=DecisionTreeClassifier(),
param_grid=param_grid,
verbose=1,
cv=5)
%time gs_dt.fit(X_train, y_train)
gs_dt.best_estimator_
print(f'Score on training set: {gs_dt.score(X_train, y_train)}')
print(f'Score on testing set: {gs_dt.score(X_test, y_test)}')
gs_dt_train_preds = gs_dt.predict(X_train)
gs_dt_test_preds = gs_dt.predict(X_test)
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 8))
cm = confusion_matrix(y_train, gs_dt_train_preds)
ConfusionMatrixDisplay(cm).plot(ax=ax1)
ax1.set_title('Confusion Matrix: Train Data')
cm = confusion_matrix(y_test, gs_dt_test_preds)
ConfusionMatrixDisplay(cm).plot(ax=ax2)
ax2.set_title('Confusion Matrix: Test Data');
sorted(list(zip(X_train.columns, gs_dt.best_estimator_.feature_importances_)), key=lambda x: x[1], reverse=True)
print(gs_dt.best_estimator_.feature_importances_)
list(X_train.columns)
```
We saw that `seawater_temperature` was the most strongly correlated feature to upwelling, but `sea_surface_temperature` ended up having the greatest feature importance. I want to trust the model on this, but I'm wondering why this happened.
```
# Establish size of figure.
plt.figure(figsize = (50, 30))
# Plot our tree.
plot_tree(gs_dt.best_estimator_,
feature_names = X_train.columns,
class_names = ['Not upwelling', 'Upwelling'],
filled = True);
print(export_text(gs_dt.best_estimator_,
list(X_train.columns)));
print(classification_report(y_test, gs_dt.predict(X_test)))
# ROC curve
fig, ax = plt.subplots(figsize=(8,8))
plot_roc_curve(gs_dt, X_test, y_test, ax=ax, name='Decision Tree')
plot_roc_curve(gs_lr, X_test, y_test, ax=ax, name='Logistic Regression')
# add worst case scenario line
plt.plot([0,1],[0,1], label='baseline', linestyle='--')
# add a legend
plt.legend();
# want AUC (area under curve) to be as close to 1 as possible
```
Interpretation goes here
#### Explore misclassified data
```
# Get indices of misclassified data
misclass_ind_dt = np.where(y_test != gs_dt_test_preds)
# misclass_ind_dt
X_test_times = df.iloc[X_test.index]['time']
X_test_times.iloc[misclass_ind_dt]
pd.DataFrame(X_test_times.iloc[misclass_ind_dt]).reset_index(drop=True)
frames_dt = [pd.DataFrame(X_test_times.iloc[misclass_ind_dt]), X_test.iloc[misclass_ind_dt], pd.DataFrame(y_test.iloc[misclass_ind_dt])]
misclass_df_dt = pd.concat(frames_dt, axis=1)
misclass_df_dt
df['CUTI'] = df['CUTI']
df['CUTI']
```
| github_jupyter |
# langages de script – Python
## Introduction et remise à niveau
### M2 Ingénierie Multilingue – INaLCO
loic.grobol@gmail.com
yoa.dupont@gmail.com
# Vue générale
## Les cours
* Le mercredi de 9h à 12h en 7.03, sauf indication contraire
* une page d'accueil vous donnera accès aux cours déjà effectués, aux corrections d'exercices, etc: https://loicgrobol.github.io/python-im-2/
* la page sera souvent mise à jour, pensez à y jeter un œil régulièrement !
## L'évaluation
* Un examen qui vaudra pour 50% de la note finale
* Un projet qui vaudra pour 50% de la note finale
# L'examen
Un examen de 2h sur ordinateur où il faudra rendre un **script python** (extension `.py`) qui renvoie les résultats attendus à chaque question.
**Si vous rendez un notebook, on vous retirera des points.**
# Le projet
Vous allez construire une [bibliothèque logicielle](https://fr.wikipedia.org/wiki/Biblioth%C3%A8que_logicielle) (ou librairie). Le sujet sera à votre choix.
# Rappel
* Nous travaillerons avec Python3 (`3.6` ou supérieur de préférence)
* Pour le travail avec la console vous utiliserez `python3` (ou `python` si vous avez une installation récente) ou `ipython`
<small>Tapez `python3` ou `python3 -m IPython` dans un terminal pour accéder à la console</small>
* Choisissez l'éditeur de texte que vous préférez (emacs, vi, atom, visual studio, sublime text, …)
* Vos scripts devront être encodés en utf-8, indiquez-le dans vos scripts avec le commentaire suivant en première ligne :
`# -*- coding: utf-8 -*-` ou `# coding=utf-8`
* pensez à aller sur la doc en ligne : https://docs.python.org/3/
# Rappel (2)
Pour exécuter vos scripts, deux options :
`> python3 mon_script.py`
ou <small>(mais pourquoi faire compliqué ?)</small>
`> chmod u+x`
`> ./mon_script.py` en ajoutant ce shebang en première ligne :
`#!/usr/bin/env python3`
# À vos marques, prêts, ...
* avant d'entrer dans le vif du sujet, nous allons passer en revue les notions vues l'année dernière (pour qui a suivi l'intro en M1 à l'Inalco).
# Opérateurs mathématiques
`+` addition (`+` est aussi l'opérateur de concaténation de chaînes de caractères)
`-` soustraction
`*` multiplication
`/` division
`//` la division entière
`%` modulo (reste de la division)
`**` puissance
* L'ordre des opérations est l'ordre classique en mathématiques (puissance passe avant les opérations).
* vous pouvez utiliser des parenthèses pour définir des priorités.
### Opérateurs de comparaison
`<` inférieur / `<=` inférieur ou égal
`>` supérieur / `>=` supérieur ou égal
`==` égal / `!=` non égal
`is` identité (pour les objets surtout) / `is not` non identité
# Les variables
* L'affectation des variables se fait à l'aide du symbole `=`
* Si la variable est placée à droite du symbole `=`, sa *valeur* est affectée à la variable placée à gauche.
* Les noms de variable sont composés de car. alphabétiques (min ou maj), des chiffres et de l'underscore. C'est tout.
* Les noms de variable sont choisis par le programmeur, ils doivent être le plus clair possible. Il est conseillé de suivre la [PEP 8](https://www.python.org/dev/peps/pep-0008/).
```
var = 3 + 2
print(var)
another_var = var
print(another_var)
je-ne-suis-pas-une-variable = 2
3_moi_non_plus = 2 + 3
```
* Seuls les mots réservés sont interdits.
```
import keyword
print(keyword.kwlist)
```
# Les types
* Python est un langage à typage *dynamique* fort : le type d'une variable est déterminé par l'interpréteur.
* Python est un langage à typage dynamique *fort* : pas de conversion implicite, certaines actions sont interdites.
```
"Hello" + 1
```
* La fonction `type()` retourne le type de la variable donnée en argument.
```
type("Hello")
```
* `str()`, `int()`, `float()` convertit l'argument dans le type désiré.
* `bool()` retourne `True` si l'argument est vrai, `False` sinon (ce qui est rarement utile)
```
int(3.14159265359)
"Hello" + str(1)
```
# Les fonctions
* Une fonction, pour simplifier, va être un bout de code qu'on pourra réutiliser et qui fait un tâche bien précise
```python
def ma_fonction(arg1, arg2):
""" description de la fonction """
instruction1
instruction2
return resultat
truc
```
* L'ordre des arguments est déterminant
* Il est possible d'outrepasser l'ordre en nommant les arguments lors de l'appel de fonction
```
def soustraction(arg1, arg2):
"""On va soustraire arg2 à arg1"""
res = arg1 - arg2
return res
diff = soustraction(5, 3)
diff
diff = soustraction(3, 5)
diff
diff = soustraction(arg2=5, arg1=3)
diff
```
# Rappel
Avant d'utiliser une fonction il est indispensable de savoir : ce que fait la fonction, quels sont les paramètres attendus, quelle est la valeur de retour.
Pour accéder à la documentation d'une fonction :
* dans la console : `help(int)` <small>(d'une fonction ou d'une classe ou d'un module)</small>
<small>`q` pour sortir de la doc</small>
* sur le web : [https://docs.python.org/3/library/functions.html](https://docs.python.org/3/library/functions.html)
* ou bien : [DevDocs](http://devdocs.io/)
# Les chaînes de caractères
* Les chaînes de caractères sont entourées soit de quotes simples `'`, soit de guillemets `"`
* Si votre mot contient une apostrophe, entourez-le de guillemets `"`
```
'Ça donne une erreur t'as vu'
"Ça donne une erreur t'as vu"
```
Les chaînes sont des *sequences*, on peut leur appliquer les opérations suivantes propres à la catégorie d'objets *sequence* :
* longueur, minimum, maximum
```
var = "bonjour"
# longueur, minimum, maximum
print(len(var))
print(max(var))
```
* _indexing_
```
# indexing
var = "bonjour"
print(var[2])
print(var[-1])
```
* _slicing_
```
# slicing
print(var[0:3]) # 3 premiers éléments
print(var[-3:]) # 3 dernier éléments
```
* _membership_
```
if 'u' in var:
print("Il y a un u dans {}".format(var))
```
Les chaînes ont aussi des fonctions qui leur sont propres
Voir la liste complète dans la doc python
`lower()` transforme la chaine en minuscules
`upper()` transforme la chaine en majuscules
`replace(old, new)` remplace les occurrences de `old` par `new`
`strip(chars=None)` appelé sans arguments supprime le ou les espaces en tête et en fin de chaîne
`rstrip(chars=None)` fait la même chose en fin de chaîne uniquement
`lstrip(chars=None)` idem en début de chaîne
`split(sep=None)` découpe une chaîne en fonction de `sep` et renvoie une liste. Si `sep` n'est pas donné, coupe sur tous les caractères d'espace
`join(iterable)` est l'inverse de `split`, il permet de joindre les éléments d'une liste de chaînes pour former une seule chaîne de caractères
[`format()`](https://docs.python.org/3.5/library/string.html#formatstrings) depuis python3 (et python2.7) pour effectuer l'[interpolation de chaîne](https://en.wikipedia.org/wiki/String_interpolation)
## Formatage de chaînes
« There should be one-- and preferably only one --obvious way to do it. » _Zen of Python_
Sauf que :
* concaténation avec `+`
* interpolation avec `format()`
* [f-string](https://docs.python.org/3.6/reference/lexical_analysis.html#f-strings) depuis python3.6
```
name = ""
coffee_price = 0.6
print("Tiens salut " + name + ". T'aurais pas " + str(coffee_price*2) + " euros pour 2 cafés ?")
print("Tiens salut {}. T'aurais pas {} euros pour 2 cafés ?".format(name, coffee_price*2))
print(f"Tiens salut {name}. T'aurais pas {coffee_price*2} euros pour 2 cafés ?")
```
# Les structures de donnée
## Les listes
* Une liste une structure de données ordonnée
* Une liste peut contenir plusieurs valeurs, variables, listes, objets, ... le tout de types différents
* On accède à un élément par son indice (de 0 à n-1, n étant le nombre d'éléments)
```
voyelles = []
voyelles = ['a', 'e', 'i', 'o', 'u']
print(voyelles[0])
chaines = ['abc', 'def', 'ghi']
print(chaines[2])
print(chaines[-1])
chiffres = [0, 1, 2, 3, 4, 5, 6, 7, 8, 9]
print(chiffres[5:])
```
## Les dictionnaires
* Un dictionnaire est une structure de données associative de type 'clé' → 'valeur'
* Les données ne sont pas ordonnées comme dans les listes
* On accède à une valeur par sa clé
* Les clés sont uniques : on ne peut pas associer deux valeurs à une même clé
* `keys()` renvoie la liste des clés, `values()` la liste des valeurs
```
couleurs = {'a':'noir', 'e':'blanc', 'i':'rouge', 'u':'vert', 'o':'bleu'}
couleurs['i'] = "pourpre"
couleurs
couleurs.keys()
couleurs.values()
couleurs.items()
```
# Les structures conditionnelles
```python
if condition:
[...]
elif condition: # si besoin
[...]
else: # si besoin
[...]
```
### Opérateurs booléens
``not`` négation
``and`` conjonction (True si les deux opérandes sont vraies, False sinon)
``or`` disjonction (True si une des deux opérandes est vraie)
* Les valeurs ci-dessous sont toutes évaluées par l'interpréteur comme ayant la valeur booléenne *false*
`False` `None` `0` `""` `()` `[]` `{}`
* Tout le reste<sup>1</sup> sera évalué comme _true_
Vous pouvez écrire :
```python
>>> if var: ou while my_list:
```
plutôt que :
```python
>>> if var != "": ou while my_list != []:
```
<sup>1</sup> <small>Sauf les objets dont vous avez construit les classes. Voir les diapos à venir sur Classes et objets.</small>
```
x = 4
if x > 3:
print("x a grandi")
if x > 3 and x <= 5:
print("x a grandi, un peu")
elif x > 5:
print("x a grandi")
else:
print("x n'a pas grandi")
if x is not None:
print("x n'est pas rien")
if not x is None: # Moins élégant et plus lent
print("x n'est toujours pas rien")
```
# Les boucles
* Les boucles `while` nécessitent que la valeur utilisée dans la condition d'arrêt soit modifiée dans le corps de la boucle.
```
i = 1
while i < 5:
print(i)
i = i + 1
```
* Les boucles `for` s'appliquent sur les *séquences* (`list`, `str`, `tuple`) et plus généralement sur les *iterables* [voir doc](https://docs.python.org/3/glossary.html#term-iterable)
* Les *iterables* sont des objets issus de classes qui implémentent la méthode `__iter__()` et/ou `__getitem__()`
* L'instruction `continue` permet de passer à l'itération suivante
* L'instruction `break` permet de quitter la boucle en cours
```
for item in voyelles:
print(item)
for item in couleurs.keys():
if item == 'i':
continue
print(item)
for key, value in couleurs.items():
print(key, value)
if key == 'i':
break
```
* `zip` permet de boucler sur plusieurs séquences
* Si les séquences sont de tailles différentes `zip` s'arrête à la longueur la plus petite
```
noms = ['einstein', 'planck', 'turing', 'curie', 'bohr', 'shannon']
facs = ['inalco', 'p3', 'p10', 'inalco', 'p3', 'inalco']
parcours = ['pro', 'r&d', 'r&d', 'pro', 'pro', 'r&d']
for nom, fac, parcours in zip(noms, facs, parcours):
print("{} est inscrit en {} à {}".format(nom, parcours, fac))
```
# Exercices série 1 ! Des chiffres et des lettres
1. Des triangles
1. écrire une fonction `la_plus_grande(longueur1, longueur2, longueur3)` qui renvoie la longueur du plus grand côté (une fonction de python fait peut-être déjà cela...).
2. écrire une fonction `est_equilateral(longueur1, longueur2, longueur3)` qui détermine si un triangle est équilatéral ou non (les trois côtés ont la même longueur).
2. écrire une fonction `est_isocele(longueur1, longueur2, longueur3)` qui détermine si un triangle est isocèle (deux côtés de même longueur mais pas trois) ou non.
3. Écrire une fonction `caracteristiques(longueur1, longueur2, longueur3)` qui affiche à l'écran la nature et la taille du plus grand côté d'un triangle. On dira qu'un triangle est `quelconque` s'il n'est ni équilatéral ni isocèle. Affiche `pas un triangle` si les longueurs données ne font pas un triangle (la longueur du plus grand côté est supérieure à celle des deux autres).
```
def la_plus_grande(longueur1, longueur2, longueur3):
"""Renvoie la plus grande longueur."""
# TODO: codez !
def est_equilateral(longueur1, longueur2, longueur3):
"""Renvoie si un triangle est équilatéral."""
# TODO: codez !
def est_isocele(longueur1, longueur2, longueur3):
"""Renvoie si un triangle est isocele."""
# TODO: codez !
def est_triangle(longueur1, longueur2, longueur3):
"""Renvoie si les longueurs données font bien un triangle."""
# TODO: codez !
def caracteristiques(longueur1, longueur2, longueur3):
"""Affiche les caractéristiques d'un triangle.
Les caractéristiques d'un triangle sont :
- sa nature
- la taille de son plus grand côté.
On dira qu'un triangle est `quelconque` s'il n'est ni équilatéral ni isocèle.
Affiche `pas un triangle` si les longueurs données ne font pas un triangle
(la longueur du plus grand côté est supérieure à celle des deux autres).
"""
# TODO: codez !
caracteristiques(1, 1, 1) # equilatéral 1
caracteristiques(1, 1, 2) # isocèle 2
caracteristiques(1, 2, 1) # isocèle 2
caracteristiques(2, 1, 1) # isocèle 2
caracteristiques(2, 3, 1) # quelconque 3
caracteristiques(2, 3, 6) # pas un triangle
caracteristiques(6, 3, 2) # pas un triangle
caracteristiques(2, 6, 3) # pas un triangle
```
2. Des heures
1. écrire une fonction `heures(secondes)` qui prend un nombre de secondes (entier) et le convertit en heures, minutes et secondes sous le format `H:M:S` où `H` est le nombre d'heures, `M` le nombre de minutes et `S` le nombre de secondes.
2. écrire une fonction `secondes(heure)` qui prend une heure au format `H:M:S` et renvoie le nombre de secondes correspondantes (entier).
On ne gèrera ici pas les cas incohérents comme un nombre de secondes négatif ou une heure mal formatée.
```
def heures(secondes):
"""Prend un nombre de secondes (entier) et le convertit en heures, minutes
et secondes sous le format `H:M:S` où `H` est le nombre d'heures,
`M` le nombre de minutes et `S` le nombre de secondes.
On suppose que secondes est positif ou nul (secondes >= 0).
"""
# TODO: codez !
def secondes(heure):
"""Prend une heure au format `H:M:S` et renvoie le nombre de secondes
correspondantes (entier).
On suppose que l'heure est bien formattée. On aura toujours un nombre
d'heures valide, un nombre de minutes valide et un nombre de secondes valide.
"""
# TODO: codez !
print(heures(0)) # 0:0:0
print(heures(30)) # 0:0:30
print(heures(60)) # 0:1:0
print(heures(66)) # 0:1:6
print(heures(3600)) # 1:0:0
print(heures(86466)) # 24:1:6
print(secondes('0:0:0')) # 0
print(secondes('6:6:6')) # 21966
print(secondes(heures(86466))) # 86466
print(heures(secondes('24:1:1'))) # 24:1:1
```
3. Des cartes
Nous jouons aux cartes à quatre personnes. On appelle un pli l'ensemble des cartes jouées dans un tour (ici, quatre cartes). Chaque carte a une valeur (un entier de 1 à 13). Chaque carte a également une couleur : carreau, trèfle, cœur ou pic. Ces couleurs sont notés avec une lettre: carreau=`D`, trèfle=`C`, cœur=`H` et pic=`S`. Une carte est alors une chaîne avec sa couleur et sa valeur, par exemple l'as de pic est noté `S1`, la dame de cœur `H12`. La carte du premier joueur `carte1` donne la couleur attendue. Une carte qui n'est pas à la bonne couleur perd automatiquement. Écrire une fonction `gagne_couleur(carte1, carte2, carte3, carte4)` qui affiche la carte qui remporte le pli en faisant attention aux couleurs.
On ne gèrera pas certains cas incohérents comme une carte ou un pli invalide.
```
def gagne_couleur(carte1, carte2, carte3, carte4):
"""Affiche la carte qui remporte le pli en faisant attention aux couleurs :
- la carte du premier joueur `carte1` donne la couleur attendue.
- une carte qui n'est pas à la bonne couleur perd automatiquement.
On ne gèrera pas certains cas incohérents comme une carte ou un pli invalide.
"""
# TODO: codez !
gagne_couleur('S1', 'S2', 'S3', 'S4') # S4
gagne_couleur('S4', 'S3', 'S2', 'S1') # S4
gagne_couleur('S1', 'D2', 'C3', 'H4') # S1
gagne_couleur('S1', 'D2', 'S13', 'S10') # S13
```
Faire l'exercice ASCII ART de codingame: https://www.codingame.com/ide/puzzle/ascii-art
| github_jupyter |
<a href="https://colab.research.google.com/github/martin-fabbri/colab-notebooks/blob/master/xlm_r/xlm_r_explore_vocabulary.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Setup
--------------------------
```
!pip install transformers -Uqq
!pip install sentencepiece -Uqq
!wget -q http://www.gutenberg.org/files/3201/files/NAMES.TXT
import torch
import random
import pandas as pd
import sentencepiece as spm
import matplotlib.pyplot as plt
import seaborn as sns
from transformers import XLMRobertaTokenizer
from transformers import BertTokenizer
# download the tokenizer for the XLM-Robert base model
tokenizer = XLMRobertaTokenizer.from_pretrained('xlm-roberta-base')
```
## Inspect XLM-R Vocabulary
--------------------------
### Vocabulary Dump
```
# retrieve the full list of tokens
all_tokens = list(tokenizer.get_vocab().keys())
print(f'The vocabulary contains {len(all_tokens):,} tokens.')
all_tokens[:5], all_tokens[-5:]
```
## SentencePiece vs. WordPiece
------------------------------
XLM-R uses a `SentencePiece` model, which is a little different than BERT's WordPiece model.
As an example, I'll use a misspelling of the word "philosophy": "philosphy". This will cause the tokenizer to break the word into subwords.
In WordPiece, subwords are denoted by two hash characters, except the *first* subword in a word.
```
BERT / WordPiece: phil ##os ##phy
XLM-R / SentencePiece: ▁phil os phy
```
In SentencePiece, all spacing is captured.
> Note: The marking character used by SentencePiece looks like an underscore, but it as actually unicode symbol U+2581, named "Lower One Eighth Block". I'll refer to it as an underscore for simplicity, though.
* Underscore: _
* U+2581: ▁
With SentencePiece, you can interpret the underscore as a space, because that's exactly what it represents!
```
bert_tokenizer = BertTokenizer.from_pretrained('bert-base-cased')
example_sentence = 'The reactions were unexpectedly uplifting and spontaneous. Just welcoming and inviting!'
xlmr_tokens = tokenizer.tokenize(example_sentence)
bert_tokens = bert_tokenizer.tokenize(example_sentence)
def print_as_rows(list_a, list_b):
'''
Prints two lists as rows, with padding to make them line up neatly.
'''
row_a = ""
row_b = ""
# Pad the lists to the same length.
while len(list_a) < len(list_b):
list_a.append("")
while len(list_b) < len(list_a):
list_b.append("")
for i in range(0, len(list_a)):
str_a = list_a[i]
str_b = list_b[i]
pad_len = max(len(str_a), len(str_b)) + 2
row_a += ("{:>{width}}").format(str_a, width=pad_len)
row_b += ("{:>{width}}").format(str_b, width=pad_len)
print(row_a)
print(row_b)
print_as_rows(xlmr_tokens, bert_tokens)
pd.DataFrame([xlmr_tokens, bert_tokens])
```
### Token Lengths
Let's gather some statistics on the vocabulary.
```
sns.set(style='darkgrid')
# Increase the plot size and font size.
sns.set(font_scale=1.5)
plt.rcParams['figure.figsize'] = (10,5)
# Measure the length of every token in the vocab.
token_lengths = [len(token) for token in all_tokens]
# Plot the number of tokens of each length.
sns.countplot(x=token_lengths)
plt.title('Vocab Token Lengths')
plt.xlabel('Token Length')
plt.ylabel('# of Tokens');
print('Maximum token length:', max(token_lengths), '\n\n')
long_tokens = []
for token in all_tokens:
if len(token) == 16:
long_tokens.append(token)
long_tokens[:5]
# Read them in.
with open('NAMES.TXT', 'rb') as f:
names_encoded = f.readlines()
names = []
# Decode the names, convert to lowercase, and strip newlines.
for name in names_encoded:
try:
names.append(name.rstrip().lower().decode('utf-8'))
except:
continue
print('Number of names: {:,}'.format(len(names)))
print('Example:', names[:5], names[-5:])
# Count how many numbers are in the vocabulary.
numbers_tokens = []
# For each token in the vocabulary...
for token in all_tokens:
# If it's a whole word, and it's all digits...
if (token[0] == '▁') and token[1:].isdigit():
# Grab it.
numbers_tokens.append(token[1:])
# Any numbers > ▁9999?
if len(token) > 5:
print(token)
# Count how many dates between 1600 and 2021 are included.
count = 0
for i in range(1600, 2021):
if '▁' + str(i) in all_tokens:
count += 1
print('Vocab includes {:,} of 421 dates from 1600 - 2021'.format(count))
```
| github_jupyter |
```
# !wget https://f000.backblazeb2.com/file/malaya-model/bert-bahasa/bert-base-2020-10-08.tar.gz
# !tar -zxf bert-base-2020-10-08.tar.gz
import json
import re
import sentencepiece as spm
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '3'
from xlnet.prepro_utils import preprocess_text, encode_ids, encode_pieces
sp_model = spm.SentencePieceProcessor()
sp_model.Load('pretrained-model/preprocess/sp10m.cased.bert.model')
with open('pretrained-model/preprocess/sp10m.cased.bert.vocab') as fopen:
v = fopen.read().split('\n')[:-1]
v = [i.split('\t') for i in v]
v = {i[0]: i[1] for i in v}
class Tokenizer:
def __init__(self, v):
self.vocab = v
pass
def tokenize(self, string):
return encode_pieces(sp_model, string, return_unicode=False, sample=False)
def convert_tokens_to_ids(self, tokens):
return [sp_model.PieceToId(piece) for piece in tokens]
def convert_ids_to_tokens(self, ids):
return [sp_model.IdToPiece(i) for i in ids]
tokenizer = Tokenizer(v)
from malaya.train.model.bigbird import modeling, optimization
import numpy as np
import json
import tensorflow as tf
import itertools
from unidecode import unidecode
import re
import random
emotion_label = ['anger', 'fear', 'happy', 'love', 'sadness', 'surprise']
with open('/home/husein/sentiment/emotion-twitter-lexicon.json') as fopen:
emotion = json.load(fopen)
emotion.keys()
texts, labels = [], []
for k, v in emotion.items():
if len(v) > 30000:
emotion[k] = random.sample(v, 30000)
print(k, len(emotion[k]))
texts.extend(emotion[k])
labels.extend([emotion_label.index(k)] * len(emotion[k]))
from malaya.text.rules import normalized_chars
import random
laughing = {
'huhu',
'haha',
'gagaga',
'hihi',
'wkawka',
'wkwk',
'kiki',
'keke',
'huehue',
'hshs',
'hoho',
'hewhew',
'uwu',
'sksk',
'ksks',
'gituu',
'gitu',
'mmeeooww',
'meow',
'alhamdulillah',
'muah',
'mmuahh',
'hehe',
'salamramadhan',
'happywomensday',
'jahagaha',
'ahakss',
'ahksk'
}
def make_cleaning(s, c_dict):
s = s.translate(c_dict)
return s
def cleaning(string):
"""
use by any transformer model before tokenization
"""
string = unidecode(string)
string = ' '.join(
[make_cleaning(w, normalized_chars) for w in string.split()]
)
string = re.sub('\(dot\)', '.', string)
string = (
re.sub(re.findall(r'\<a(.*?)\>', string)[0], '', string)
if (len(re.findall(r'\<a (.*?)\>', string)) > 0)
and ('href' in re.findall(r'\<a (.*?)\>', string)[0])
else string
)
string = re.sub(
r'\w+:\/{2}[\d\w-]+(\.[\d\w-]+)*(?:(?:\/[^\s/]*))*', ' ', string
)
chars = '.,/'
for c in chars:
string = string.replace(c, f' {c} ')
string = re.sub(r'[ ]+', ' ', string).strip().split()
string = [w for w in string if w[0] != '@']
x = []
for word in string:
word = word.lower()
if any([laugh in word for laugh in laughing]):
if random.random() >= 0.5:
x.append(word)
else:
x.append(word)
string = [w.title() if w[0].isupper() else w for w in x]
return ' '.join(string)
from tqdm import tqdm
for i in tqdm(range(len(texts))):
texts[i] = cleaning(texts[i])
actual_t, actual_l = [], []
for i in tqdm(range(len(texts))):
if len(texts[i]) > 2:
actual_t.append(texts[i])
actual_l.append(labels[i])
from tqdm import tqdm
input_ids, input_masks = [], []
for text in tqdm(actual_t):
tokens_a = tokenizer.tokenize(text)
tokens = ["[CLS]"] + tokens_a + ["[SEP]"]
input_id = tokenizer.convert_tokens_to_ids(tokens)
input_mask = [1] * len(input_id)
input_ids.append(input_id)
input_masks.append(input_mask)
maxlen = 512
bert_config = {
'attention_probs_dropout_prob': 0.2,
'hidden_act': 'gelu',
'hidden_dropout_prob': 0.2,
'hidden_size': 768,
'initializer_range': 0.02,
'intermediate_size': 3072,
'max_position_embeddings': 4096,
'max_encoder_length': maxlen,
'num_attention_heads': 12,
'num_hidden_layers': 12,
'type_vocab_size': 2,
'scope': 'bert',
'use_bias': True,
'rescale_embedding': False,
'vocab_model_file': None,
'attention_type': 'block_sparse',
'norm_type': 'postnorm',
'block_size': 16,
'num_rand_blocks': 3,
'vocab_size': 32000,
}
epoch = 3
batch_size = 16
warmup_proportion = 0.1
num_train_steps = int(len(texts) / batch_size * epoch)
num_warmup_steps = int(num_train_steps * warmup_proportion)
dimension_output = 6
def create_initializer(initializer_range=0.02):
return tf.truncated_normal_initializer(stddev=initializer_range)
class Model:
def __init__(
self,
dimension_output,
learning_rate = 2e-5,
training = True,
):
self.X = tf.placeholder(tf.int32, [None, None])
self.Y = tf.placeholder(tf.int32, [None])
model = modeling.BertModel(bert_config)
sequence_output, pooled_output = model(self.X)
output_layer = sequence_output
output_layer = tf.layers.dense(
output_layer,
bert_config['hidden_size'],
activation=tf.tanh,
kernel_initializer=create_initializer())
self.logits_seq = tf.layers.dense(output_layer, dimension_output,
kernel_initializer=create_initializer())
self.logits_seq = tf.identity(self.logits_seq, name = 'logits_seq')
self.logits = self.logits_seq[:, 0]
self.logits = tf.identity(self.logits, name = 'logits')
self.cost = tf.reduce_mean(
tf.nn.sparse_softmax_cross_entropy_with_logits(
logits = self.logits, labels = self.Y
)
)
self.optimizer = optimization.create_optimizer(self.cost, learning_rate,
num_train_steps, num_warmup_steps, False)
correct_pred = tf.equal(
tf.argmax(self.logits, 1, output_type = tf.int32), self.Y
)
self.accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
dimension_output = 6
learning_rate = 2e-5
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Model(
dimension_output,
learning_rate
)
sess.run(tf.global_variables_initializer())
import collections
import re
def get_assignment_map_from_checkpoint(tvars, init_checkpoint):
"""Compute the union of the current variables and checkpoint variables."""
assignment_map = {}
initialized_variable_names = {}
name_to_variable = collections.OrderedDict()
for var in tvars:
name = var.name
m = re.match('^(.*):\\d+$', name)
if m is not None:
name = m.group(1)
name_to_variable[name] = var
init_vars = tf.train.list_variables(init_checkpoint)
assignment_map = collections.OrderedDict()
for x in init_vars:
(name, var) = (x[0], x[1])
name_r = name.replace('bert/embeddings/LayerNorm', 'bert/encoder/LayerNorm')
if name_r not in name_to_variable:
continue
if 'embeddings/position_embeddings' in name_r:
continue
assignment_map[name] = name_to_variable[name_r]
initialized_variable_names[name_r] = 1
initialized_variable_names[name_r + ':0'] = 1
return (assignment_map, initialized_variable_names)
tvars = tf.trainable_variables()
checkpoint = 'bert-base/model.ckpt-1000000'
assignment_map, initialized_variable_names = get_assignment_map_from_checkpoint(tvars,
checkpoint)
saver = tf.train.Saver(var_list = assignment_map)
saver.restore(sess, checkpoint)
from sklearn.model_selection import train_test_split
train_input_ids, test_input_ids, train_Y, test_Y, train_mask, test_mask = train_test_split(
input_ids, actual_l, input_masks, test_size = 0.2
)
pad_sequences = tf.keras.preprocessing.sequence.pad_sequences
from tqdm import tqdm
import time
for EPOCH in range(epoch):
train_acc, train_loss, test_acc, test_loss = [], [], [], []
pbar = tqdm(
range(0, len(train_input_ids), batch_size), desc = 'train minibatch loop'
)
for i in pbar:
index = min(i + batch_size, len(train_input_ids))
batch_x = train_input_ids[i: index]
batch_x = pad_sequences(batch_x, padding='post', maxlen = maxlen)
batch_y = train_Y[i: index]
acc, cost, _ = sess.run(
[model.accuracy, model.cost, model.optimizer],
feed_dict = {
model.Y: batch_y,
model.X: batch_x,
},
)
train_loss.append(cost)
train_acc.append(acc)
pbar.set_postfix(cost = cost, accuracy = acc)
pbar = tqdm(range(0, len(test_input_ids), batch_size), desc = 'test minibatch loop')
for i in pbar:
index = min(i + batch_size, len(test_input_ids))
batch_x = test_input_ids[i: index]
batch_x = pad_sequences(batch_x, padding='post', maxlen = maxlen)
batch_y = test_Y[i: index]
acc, cost = sess.run(
[model.accuracy, model.cost],
feed_dict = {
model.Y: batch_y,
model.X: batch_x,
},
)
test_loss.append(cost)
test_acc.append(acc)
pbar.set_postfix(cost = cost, accuracy = acc)
train_loss = np.mean(train_loss)
train_acc = np.mean(train_acc)
test_loss = np.mean(test_loss)
test_acc = np.mean(test_acc)
print(
'epoch: %d, training loss: %f, training acc: %f, valid loss: %f, valid acc: %f\n'
% (EPOCH, train_loss, train_acc, test_loss, test_acc)
)
saver = tf.train.Saver(tf.trainable_variables())
saver.save(sess, 'bigbird-base-emotion/model.ckpt')
strings = ','.join(
[
n.name
for n in tf.get_default_graph().as_graph_def().node
if ('Variable' in n.op
or 'Placeholder' in n.name
or 'logits' in n.name
or 'alphas' in n.name
or 'self/Softmax' in n.name)
and 'adam' not in n.name
and 'beta' not in n.name
and 'global_step' not in n.name
]
)
strings.split(',')
real_Y, predict_Y = [], []
pbar = tqdm(
range(0, len(test_input_ids), batch_size), desc = 'validation minibatch loop'
)
for i in pbar:
index = min(i + batch_size, len(test_input_ids))
batch_x = test_input_ids[i: index]
batch_x = pad_sequences(batch_x, padding='post', maxlen=maxlen)
batch_y = test_Y[i: index]
predict_Y += np.argmax(sess.run(model.logits,
feed_dict = {
model.X: batch_x,
},
), 1, ).tolist()
real_Y += batch_y
from sklearn import metrics
print(
metrics.classification_report(
real_Y, predict_Y, target_names = ['anger', 'fear', 'happy', 'love', 'sadness', 'surprise'],
digits = 5
)
)
def freeze_graph(model_dir, output_node_names):
if not tf.gfile.Exists(model_dir):
raise AssertionError(
"Export directory doesn't exists. Please specify an export "
'directory: %s' % model_dir
)
checkpoint = tf.train.get_checkpoint_state(model_dir)
input_checkpoint = checkpoint.model_checkpoint_path
absolute_model_dir = '/'.join(input_checkpoint.split('/')[:-1])
output_graph = absolute_model_dir + '/frozen_model.pb'
clear_devices = True
with tf.Session(graph = tf.Graph()) as sess:
saver = tf.train.import_meta_graph(
input_checkpoint + '.meta', clear_devices = clear_devices
)
saver.restore(sess, input_checkpoint)
output_graph_def = tf.graph_util.convert_variables_to_constants(
sess,
tf.get_default_graph().as_graph_def(),
output_node_names.split(','),
)
with tf.gfile.GFile(output_graph, 'wb') as f:
f.write(output_graph_def.SerializeToString())
print('%d ops in the final graph.' % len(output_graph_def.node))
freeze_graph('bigbird-base-emotion', strings)
def load_graph(frozen_graph_filename):
with tf.gfile.GFile(frozen_graph_filename, 'rb') as f:
graph_def = tf.GraphDef()
graph_def.ParseFromString(f.read())
with tf.Graph().as_default() as graph:
tf.import_graph_def(graph_def)
return graph
# g = load_graph('bigbird-base-emotion/frozen_model.pb')
# x = g.get_tensor_by_name('import/Placeholder:0')
# logits = g.get_tensor_by_name('import/logits:0')
# test_sess = tf.InteractiveSession(graph = g)
# result = test_sess.run(tf.nn.softmax(logits), feed_dict = {x: [input_id]})
# result
```
| github_jupyter |
# Event-related fields
Event-related fields (ERFs) as well as event-related potentials are generated by neuronal activity elicited by a given events. They are often used in cognitive and clinical neurosciene to quantify the brain activity associated with a given task (Luck 2014, Woodman 2010). The aim of this section is generate event-related fields (ERFs) in response to the visual input and to do some simple plotting of time course and topographies. Event-related fields are generated by neuronal activity phase-locked to a given events.
## Preparation
Import the relevant modules.
```
import os.path as op
import os
import sys
import matplotlib.pyplot as plt
import mne
import numpy as np
```
See the local paths of the data:
```
data_path = r'C:\Users\JensenO\Dropbox\FLUX\Development\dataRaw'
result_path = r'C:\Users\JensenO\Dropbox\FLUX\Development\dataResults'
file_name = 'training_epo.fif'
path_file = op.join(result_path,file_name)
```
Read the epochs:
```
epochs = mne.read_epochs(path_file,
preload=True,
verbose=True)
```
## Averaging the trial data
Identify the epochs for the left condition with respect to the onset of the moving gratings and then average over trials. Subsequently apply a lowpass filter at 30 Hz and crop the data to the time interval from -100 to 400 ms from stimulus onset. The 30 Hz low-pass filter is typically used for cognitive type event-related responses.
```
evoked_left= epochs['left'].copy().average(method='mean').filter(0.0, 30).crop(-0.1,0.4)
```
## Plotting event-related fields
To plot the single trials and the averaged event-related fields for a single magnetometer write:
```
epochs['left'].copy().filter(0.0,30).crop(-0.1,0.4).plot_image(picks=['MEG1911'],vmin=-500,vmax=500);
evoked_left.copy().apply_baseline(baseline=(-0.1, 0))
evoked_left.copy().pick_types(meg='mag').plot_topo(title = 'Magnetometers');
```
Above shows the event-related fields from -100 to 400 ms over each magnetometer. Note the strong response at ~110 ms over posterior sensors. This is often referred to as N100m.
To plot a topographic map of the response at 110 ms write:
```
evoked_left.plot_topomap(0.110, ch_type='mag', time_unit='s');
```
**Question 1:** Explain how an equivalent current dipole (ECD) can account for the posterior event-related field by drawing the ECD on top of the topographic plot (hint: consider the 'right-hand rule')
**Question 2:** Plot a sequence of topographic maps for the time-points ranging from 0 to 300 ms in steps of 20 ms.
Now we plot the same event-related fields over each planar gradiometer. Baseline correction is applyed beforehand
```
evoked_left.copy().apply_baseline(baseline=(-0.1, 0))
evoked_left.copy().pick_types(meg='grad').plot_topo(title = 'Gradiometers');
```
The planar gradiometers are arranged in pairs. Each pair is composed of two orthogonal planar sensors (ending in XXX2 and XXX3). It is not straightforward to interpret the field maps for planar gradiometers. Nevertheless, the magnitude of the planar gradiometers is typically the largest directly above a given current dipole.
**Question 3:** Why is the magnitude of the planar gradient typically strongest directly above a current dipole (hint: explain on the basis of the topographic map of the N100m)
To better interpret the planar gradiometers one can apply the root-mean-square operation in which the magnitude of the field for two orthogonal gradiometers are combined:
```
evoked_left.copy().pick_types(meg='grad').plot_topo(title='Gradiometers', merge_grads=True);
```
To plot a topographic map of the combined planar gradient at 110 ms write:
```
evoked_left.plot_topomap(0.11, ch_type='grad', time_unit='s');
```
**Question 3:** Interpret the topograpgy of the combined planar gradient in relation to the dipolar map (see Question 1).
## Preregistration and publications
Publication, example:
"Prior to calculating the event-related fields, the data were lowpass filtered at 30 Hz (non-causal finite impulse response filter implemented using a Hamming window and a 441 sample filter length). After averaging, a 100 ms baseline was subtracted."
## References
Woodman G.F. (2010) A brief introduction to the use of event-related potentials in studies of perception and attention. Atten Percept Psychophys. 72(8):2031-46.
Luck, S.J. (2014) An Introduction to the Event-Related Potential Technique, Second Edition. Bradford Books
(many of the recommendation for calculated event-related potentials also apply to calculating event-related fields)
| github_jupyter |
# Introduction
In this notebook, we will provide information about key concepts in Machine Learning as well as discuss some of the most useful tools for predictive data analysis in Python.
## Table of Content
- [0. Packages](#0)
- [1. Scikit Learn](#1)
- [1.1 Alternatives](#1-1)
- [2. Supervised Learning](#2)
- [2.1 Classification](#2-1)
- [Jupyter Notebook Widgets (Data Visualisation)](#2-1-1)
- [Preprocessing](#2-1-2)
- [Classify Digits](#2-1-3)
- [Categorical Variables Encoding](#2-1-4)
- [2.2 Regression](#2-2)
- [Other Linear Regression Models](#2-2-1)
- [Making Predictions](#2-2-2)
- [3.Unsupervised Learning](#3)
- [3.2 Clustering](#3-1)
- [3.1 PCA](#3-2)
# 0. Packages <a name="0"></a>
In this session, we will make use of the following packages:
- [numpy](https://docs.scipy.org/doc/numpy/) is a popular library for scientific computing.
- [matplotlib](https://matplotlib.org/3.1.1/contents.html) is a plotting library compatible with numpy.
- [pandas](https://pandas.pydata.org/docs/) is what we'll use to manipulate our data.
- [sklearn](https://scikit-learn.org/stable/index.html) will be used to measure the performance of our model.
Run the next cell to import the necessary packages mentioned before. Besides, we will add more packages as needed while progressing in this session.
```
# Good practice to use short but clear aliases for the imported libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import sklearn
from sklearn import preprocessing
# Set a seed
np.random.seed(2020)
# Magic Function
%matplotlib inline
# Hide all warnings
import warnings
warnings.filterwarnings('ignore') # warnings.filterwarnings(action='once')
```
# 1. Scikit-learn <a name="1"></a>
Scikit-learn is one of the most popular libraries for ML and predictive data analysis in python. Scikit-learn was built on top of other popular libraries such as NumPy, SciPy and matplotlib [1].
Some of the benefits of using Scikit-learn:
- Simple and consistent API
- It covers most of the machine learning tasks
- Great documentation
- You can finetune many parameters and still have good default values
- Efficient models
- Open source and commercially usable
According to the creators of Scikit-learn:
> Often the hardest part of solving a machine learning problem can be finding the right estimator (algorithm or model) for the job. Different estimators are better suited for different types of data and different problems. The flowchart below is designed to give users a bit of a rough guide on how to approach problems concerning which estimators to try on your data [2].
In the official scikit-learn documentation, a useful map showing the landscape of machine learning is provided [2].
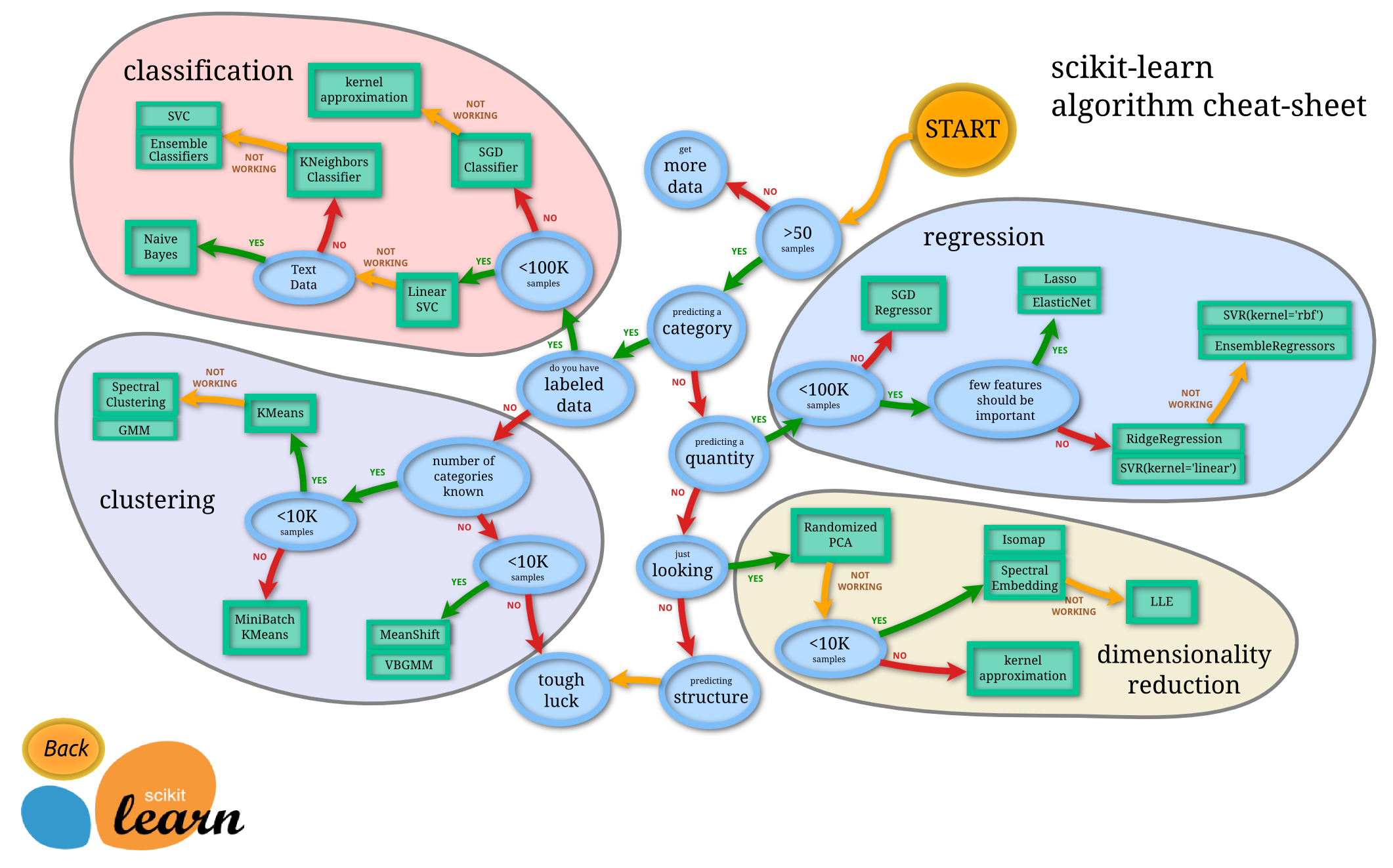
Sources:
- 1. [Scikit-learn official website](https://scikit-learn.org/stable/)
- 2. [Machine Learning Map](https://scikit-learn.org/stable/tutorial/machine_learning_map/index.html)
# 1.1 Alternatives <a name="1-1"></a>
Scikit-learn is one of the most popular ML libraries in Python that provide out-of-the-box implementations of the most popular ML algorithms. However, given some circumstances, you might find want to use other libraries or complement your pipelines with other ML libraries.
Other popular ML libraries in Python are:
**SciPy library:**
> The SciPy library is one of the core packages that make up the SciPy stack. It provides many user-friendly and efficient numerical routines, such as routines for numerical integration, interpolation, optimization, linear algebra, and statistics.
Source: [SciPy](https://www.scipy.org/)
**RAPIDS:**
> The RAPIDS data science framework includes a collection of libraries for executing end-to-end data science pipelines completely in the GPU. It is designed to have a familiar look and feel to data scientists working in Python.
RAPIDS uses optimized NVIDIA CUDA® primitives and high-bandwidth GPU memory to accelerate data preparation and machine learning. The goal of RAPIDS is not only to accelerate the individual parts of the typical data science workflow, but to accelerate the complete end-to-end workflow.
One of the current limitations of the Scikit-learn library is that it is implemented to be used in CPU. This might be ok for many use cases, however, as more complicated and larger you get the longer will take to train the ML algorithms.
RAPIDS has a specific module for ML (cuML) in GPU which accelerates the performance significantly compared with CPU-based implementation libraries such as Scikit-learn.
> cuML is a suite of fast, GPU-accelerated machine learning algorithms designed for data science and analytical tasks. Our API mirrors Sklearn’s, and we provide practitioners with the easy fit-predict-transform paradigm without ever having to program on a GPU.
Source: [RAPIDS](https://rapids.ai/start.html)
**PyTorch/Tensorflow**
Deep learning can use more complex data, but needs more data. This approaches typically use deep learning frame works like PyTorch which we will use in the next course.
# 2. Supervised Learning <a name="2"></a>
Supervised learning is the machine learning (ML) task of learning a function that maps an input to an output based on example input-output pairs [3]. It infers a function from labelled training data consisting of a set of training examples [4].
Consider the example below, where we have a classification problem. The MNIST dataset uses images corresponding to handwritten digits from 0 to 9 for a total of 10 different classes. Suppose that we already have a trained model.

In the example above, the image represents the digit 1 which is passed to the Machine Learning classifier, then the output generates an array with 10 positions (every position represents a class), being the class 0 and 9, the first and last position of the array. So we can define the ML classifier as a function that is going to map the input (in our case an image of the digit 1), to a set of probabilities (values ranging from 0 to 1) for the 10 different classes.
In our example, $p(1|x, \theta)$ = 0.84. In ML is common to use the <code>argmax()</code> function to get the position of the array with the maximum probability. This index would correspond to the most likely label. If we apply <code>argmax()</code> to the output array, we will get the value 1 in return given that 0.84 is the maximum probability value.
To train an ML algorithm, we have to modify the internal learnable parameters $\theta$. Those values will allow the algorithm to map correctly its input to the desired output.
In the supervised learning approach, we need to provide many examples of inputs with their correspondent label or class to the algorithm for the training phase. In the image below, we can observe a representation of this process.

In this iterative process, a loss or cost function is used to measure the distance between the real label and the prediction. The aim of the training process is then to get a closer distance between the prediction and real label on unseen images.
Find more information about the Supervised Learning algorithms available in scikit-learn [here](https://scikit-learn.org/stable/supervised_learning.html#supervised-learning)
Sources:
- [Wikipedia](https://en.wikipedia.org/wiki/Supervised_learning#:~:text=Supervised%20learning%20is%20the%20machine,a%20set%20of%20training%20examples.)
- 3. Stuart J. Russell, Peter Norvig (2010) Artificial Intelligence: A Modern Approach, Third Edition, Prentice Hall ISBN 9780136042594.
- 4. Mehryar Mohri, Afshin Rostamizadeh, Ameet Talwalkar (2012) Foundations of Machine Learning, The MIT Press ISBN 9780262018258.
Consider the example below where we have the ground truth label `y` and 3 predictions made by different machine learning algorithm. We will determine what is the best prediction using `Mean Absolute Error` (MAE).
*Note:* `MAE` is not usually used for classification problems. Instead, there are better metrics for classification that will be discussed in the next session. However, `MAE` is one of the simplest error functions.
```
y = np.array([0,0,0,0,1,0,0,0,0,0])
print('Correct Answer:', y.argmax())
pred_1 = np.array([0,0,0,0,0,0,0,0,0,1])
pred_2 = np.array([0.01,0,0,0,0.89,0,0,0,0,0.1])
pred_3 = np.array([0,0,0,0,0.95,0,0,0,0,0.05])
error1 = np.abs(y-pred_1).mean()
error2 = np.abs(y-pred_2).mean()
error3 = np.abs(y-pred_3).mean()
print(error1, error2, error3)
```
<div class="alert alert-success">
<h2>Exercise 1</h2>
Description:
1. Given `y` calculate the error using `Mean Absolute Error` (MAE) for every prediction.
```python
y = np.array([0.35,0.05,0.05,0,0,0,0.55,0,0,0])
pred_1 = np.array([0.75,0,0,0,0,0,0,0.2,0,0.05])
pred_2 = np.array([0.01,0,0,0,0.89,0,0,0,0,0.1])
pred_3 = np.array([0,0,0,0,0,0,0,0.95,0,0.05])
```
2. Another popular error is `Mean Square Error` or `MSE`. Calculate `MSE` for every prediction using `np.square` instead of `np.abs`
<details>
<summary><b>→ Hints</b></summary>
- Use `np.abs` and `.mean()` to calculate the `Mean Absolute Error`.
- To calculate `Mean Squared Error` use `np.square(y-pred).mean()`
</details>
<br/>
<br/>
<details>
<summary>
<b>→ Solution</b>
</summary>
Mean Absolute Error:
```python
error1 = np.abs(y-pred_1).mean()
error2 = np.abs(y-pred_2).mean()
error3 = np.abs(y-pred_3).mean()
print(error1, error2, error3)
```
Mean Squared Error:
```python
error1 = np.square(y-pred_1).mean()
error2 = np.square(y-pred_2).mean()
error3 = np.square(y-pred_3).mean()
print(error1, error2, error3)
```
</details>
</div>
# 2.1 Classification <a name="2-1"></a>
While regression methods map inputs to a continuous dependent variable from a number of independent variables. In classification problems, the inputs will be mapped to a defined set of classes. There are many ML algorithms for regression and classification.
For this session, we will explore the most basic ML algorithm for binary classification, the <code>Logistic Regression</code>.
> ... the logistic model (or logit model) is used to model the probability of a certain class or event existing such as pass/fail, win/lose, alive/dead or healthy/sick. This can be extended to model several classes of events such as determining whether an image contains a cat, dog, lion, etc. Each object detected in the image would be assigned a probability between 0 and 1, with a sum of one.
In the next session, we will explore more complex methods for classification.
## Classifying Images of Digits
Scikit-learn provides some datasets that can be used to test different machine learning techniques. For this example, we will use the Digits dataset provided by Scikit-learn which consists of 1,797 images of 8x8 pixels. Each image, like the one shown below, is of a hand-written digit. In order to utilize an 8x8 figure like this, we would have to first transform it into a feature vector with length 64.
Source: [The Digit Dataset](https://scikit-learn.org/stable/auto_examples/datasets/plot_digits_last_image.html)
More information about the dataset [here](https://archive.ics.uci.edu/ml/datasets/Pen-Based+Recognition+of+Handwritten+Digits)
```
from sklearn import datasets
# Load the digits dataset
digits = datasets.load_digits()
# Get keys from dictionary
digits.keys()
```
# Let's see the targets
```
print(digits['target'], len(digits['target']))
idx = 100
# Let's first see one of the images
plt.figure(1, figsize=(3, 3))
print('Target: {}'.format(digits.target[0]))
plt.imshow(digits.images[idx], cmap=plt.cm.gray_r)
plt.show()
```
<a name="2-1-1"></a>
# Jupyter Notebook Widgets
Let's explore the dataset in a different way using widgets !
```
from ipywidgets import interact, widgets
from IPython.display import display
fig = plt.figure(figsize=(10, 10))
total_images = digits.target.shape[0]
def f(index):
print('Target: {}'.format(digits.target[index]))
plt.imshow(digits.images[index], cmap=plt.cm.gray_r)
fig.canvas.draw()
display(fig)
plt.show()
interact(f, index=widgets.IntSlider(min=0, max=total_images-1))
```
<a name="2-1-2"></a>
# Preprocessing
Before training any ML model, we have to prepare the raw data and process it to make it suitable for a machine learning model.
Usually, in the preprocessing step, we have to deal with some problems like:
1. Finding Missing Data and dealing with it
2. Encoding categorical features
3. Splitting the dataset into training and test set.
4. Data Normalisation
In this session and futures ones, we will provide examples of those problems and how to deal with them.
To apply a classifier on this data, we need to flatten the image, to turn the data in a (samples, features) matrix:
```
n_samples = len(digits.images)
X = digits.images.reshape((n_samples, 64))
y = digits.target
print(digits.images.shape)
print(X.shape)
```
We will further divide the dataset into the <code>train</code> and <code>test</code> set. To do that, scikit-learn provides the <code>sklearn.model_selection.train_test_split</code> function.
In ML is important to separate the dataset and have one subset for training only and another one for testing and validation. In order to evaluate the performance of ML algorithms, it is important that we test in unseen data. There are some exceptions to this rule when using much advance evaluation techniques such as cross-validation, but we will talk about it in future sessions.
<a name="2-1-3"></a>
# Train, Predict and Evaluate
```
from sklearn.linear_model import LogisticRegression
# Let's import the train_test_split function
from sklearn.model_selection import train_test_split
# Create a classifier: a Logistic Regression with All vs Rest multiclass strategy
classifier = LogisticRegression(multi_class="ovr", max_iter=1000, random_state=2020)
# Split data into train and test subsets
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.5, shuffle=False)
# We learn the digits on the first half of the digits
classifier.fit(X_train, y_train)
# Now predict the value of the digit
predicted = classifier.predict(X_test)
# Let's check the accuracy achieved by our model
print('Accuracy: {}'.format(sklearn.metrics.accuracy_score(y_test, predicted)))
```
We achieved an accuracy of 92% using one of the most basic classification algorithms, the Logistic Regression, for small images.
**Note:** For more complex images with higher dimensionality is recommended to use Deep Learning techniques such as Convolutional Neural Networks or CNNs.
<a name="2-1-4"></a>
## Encoding Categorical Features
Encoding is a preprocessing step where we convert categorical features to representations that an ML algorithm can process.
There are two popular types of encoding:
- One Hot Encoding: [`OneHotEncoder`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OneHotEncoder.html) is a utility function in scikit-learn that helps to encode categorical features as a one-hot numeric array.
- Integer/Ordinal Encoding:
In this type of encoding we will map a label or class to an integer representation. Scikit-learn provides two functions to do this. <code>[OrdinalEncoder](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OrdinalEncoder.html)</code> and <code>[LabelEncoder](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.LabelEncoder.html)</code>.
## One Hot Encoding
For this example, imagine that you have an output sequence of the following 3 labels corresponding to the number of medals a country got in the last Olympic Games.
- "bronze"
- "silver"
- "gold"
This sequence might be represented using an integer representation using `LabelEncoder` from scikit-learn. Using integer encoding the labels might have this representations:
- "bronze" -> 1
- "silver" -> 2
- "gold" -> 3
One-hot encoding uses binary vectors instead to represent the labels.
The example below shows an intuitive one-hot representation for the labels where `1` represents the presence of the label and `0` the abscence of it.
- "bronze" -> [1,0,0]
- "silver" -> [0,1,0]
- "gold" -> [0,0,1]
We are currently using 3-length vectors to represent the labels. However, there are more efficient ways of representing the same data using different representations. For example, consider the one-hot prepresentations below:
- "bronze" -> [0,0]
- "silver" -> [1,0]
- "gold" -> [0,1]
If we had one more label we could use [1,1] to represent it. This is the main idea behind the one-hot encoding process. You could create your own code to create these representations, however, there are different variations and in some cases it might be easier to use `OneHotEncoder` from scikit-learn, which handles special cases.
Consider the example below which encodes two variables (two cagegorical and three numerical) into a single one-hot representation using `OneHotEncoder` in scikit-learn:
```
from sklearn.preprocessing import OneHotEncoder
# OneHotEncoder will ignore classes that were not seen during fit
enc = OneHotEncoder(handle_unknown="ignore")
X = [["bronze"], ["silver"], ["gold"]]
# Learn Categories from seen data
enc.fit(X)
```
Now let's see what categories were learnt by the encoder
```
enc.categories_
# Transform Cagegorical values to one-hot encoding
enc.transform([["silver"], ["gold"], ["bronze"],["bronze"]]).toarray()
# You can reverse the operation as well
enc.inverse_transform([[0, 1, 0], [1.0, 0.0, 0.0]])
```
Now let's try with two encoded categorical values
```
# Data
X_2 = [["bronze","archery"],["bronze","weightlifting"],["silver","fencing"],["gold","table tennis"]]
enc_2 = OneHotEncoder(handle_unknown="ignore")
enc_2.fit(X_2)
enc_2.get_feature_names(["medal", "sport"])
```
<a name="ex-2"></a>
<div class="alert alert-success">
<h2>Exercise 2</h2>
Have a look at the result of the one-hot encoding in the example below.
1. Analyse the one-hot encoding results and compare it with the input. Can you spot any pattern?
2. Change the values of the input with valid data and try to determine if there is any pattern.
</div>
```
test = ["silver", "archery"], ["gold", "archery"], ["gold", "fencing"], ["bronze","fencing"]
enc_2.transform(test).toarray()
```
One can always drop the first column for each feature:
```
drop_enc = OneHotEncoder(drop='first').fit(X_2)
drop_enc.categories_
```
<a name="ex-3"></a>
<div class="alert alert-success">
<h2>Exercise 3</h2>
- Now that we dropped the first column, analyse how the encoding `drop_enc` changed compared to the first one-hot encoding.
</div>
```
test
drop_enc.transform(test).toarray()
```
## Label Encoding
Example:
```
le = preprocessing.LabelEncoder()
le.fit(["apples", "blueberries", "pineapple", "apples", "apples", "coconut"])
print(le.classes_)
print(le.transform(["coconut", "apples", "apples", "blueberries"]))
# Create a random list with 10 elements in the range from 0 to 4
data_example = np.random.randint(0, 4, 10)
print(data_example)
le.inverse_transform(data_example)
```
# 2.2 Regression <a name="2-2"></a>
According to Wikipedia:
> In statistical modelling, regression analysis is a set of statistical processes for estimating the relationships between a dependent variable (often called the 'outcome variable') and one or more independent variables (often called 'predictors', 'covariates', or 'features'). The most common form of regression analysis is linear regression, in which a researcher finds the line (or a more complex linear combination) that most closely fits the data according to a specific mathematical criterion.
> Most regression models propose that $Y_{i}$ is a function of $X_{i}$ and $\beta$ , with $e_{i}$ representing an additive error term that may stand in for un-modelled determinants of $Y_{i}$ or random statistical noise:
$Y_{i}=f(X_{i},\beta )+e_{i}$
**Linear regression:** It is one of the most basic regression methods. <code>LinearRegression</code> fits a linear model with coefficients to minimize the residual sum of squares between the observed targets in the dataset, and the targets predicted by the linear approximation. Mathematically it solves a problem of the form:
$\min_{w} || X w - y||_2^2$
Formula: $ f\left(x\right)=mx+b $
Sources:
- [Regression Analysis](https://en.wikipedia.org/wiki/Regression_analysis)
```
# Generate 30 random numbers
x = np.linspace(0, 50, 30)
# Random Delta
delta = np.random.uniform(-10, 10, x.size)
y = 0.5 * x + 5 + delta
plt.scatter(x, y)
from sklearn import linear_model
reg = linear_model.LinearRegression()
# Fit data
reg.fit(x.reshape(-1, 1), y.reshape(-1, 1))
# Check Regression coefficients and intercept
m = reg.coef_[0][0]
b = reg.intercept_[0]
print('m:{} b:{}'.format(m, b))
```
Let's plot the line that minimise the distance to the points
```
predictions = [(m * x[i]) + b for i in range(len(x))]
plt.scatter(x, y)
plt.plot(x, predictions, color="r")
plt.xlabel('input')
plt.ylabel('output')
```
Let's create some test data:
```
# Generate 30 random numbers
x_2 = np.linspace(0, 50, 30)
# Random Delta
delta_2 = np.random.uniform(-10, 10, x.size)
test_x = 0.5 * x + 5 + delta
test_y = reg.predict(test_x.reshape(-1,1))
```
Let's plot the new predictions:
```
predictions = [(m * x[i]) + b for i in range(len(x))]
plt.scatter(x, y)
plt.scatter(test_x, test_y, color='g')
plt.plot(x, predictions, color="r")
plt.xlabel('input')
plt.ylabel('output')
```
In this case, our features will be the pixels of the images. However, to use the image input we will need to convert it into a 1-dimensional array with length 64.
<a name="2-2-1"></a>
# Other Linear Regression Models
Let's see how different linear regression methods finds solutions to the same problem.
## Bayesian Ridge Regression
```
bayesian_ridge = linear_model.BayesianRidge()
bayesian_ridge.fit(x.reshape(-1, 1), y.reshape(-1, 1))
m2 = bayesian_ridge.coef_[0]
b2 = bayesian_ridge.intercept_
print('m:{} b:{}'.format(m2, b2))
# Let's plot the line that minimise the distance
predictions2 = [(m2 * x[i]) + b2 for i in range(len(x))]
plt.scatter(x, y)
plt.plot(x, predictions2, color="g")
plt.show()
```
# Lasso Regression
```
lasso = linear_model.Lasso(alpha=0.1)
lasso.fit(x.reshape(-1, 1), y.reshape(-1, 1))
m3 = lasso.coef_[0]
b3 = lasso.intercept_
print('m:{} b:{}'.format(m3, b3))
# Let's plot the line that minimised the dis
predictions3 = [(m3 * x[i]) + b3 for i in range(len(x))]
plt.scatter(x, y)
plt.plot(x, predictions3, color="b")
plt.show()
# Let's plot the lines created by the 3 regression models
plt.scatter(x, y)
plt.plot(x, predictions3, color="b")
plt.plot(x, predictions, color="r")
plt.plot(x, predictions2, color="g")
plt.show()
```
Let's try a regression problem with another example.
Imagine that a teacher in high school is interested to know if there is any correlation in regards to the number of hours their students studied for the final exam and the score they got. For this example, we will assume that the number of hours is the only factor that was collected. In this dataset, every row represents a student with the corresponding number of study hours and the score.
```
# Let's use pandas to explore the dataset
# Load the dataset
students = pd.read_csv('hours_vs_scores.csv')
students
```
## Splitting the dataset
Let's split the student's dataset. First, let's split <code>X</code> and <code>y</code>.
```
# Being X the features (hours)
X = students.iloc[:, 0]
# Being y, what we want to predict (score)
y = students.iloc[:, 1]
# We can specify the percentage for the splitting for test_size. This means that 20% of the data will be used for testing and the rest for training.
# random_state is set for reproducibility
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=2020
)
X_train.shape
```
We need to reshape <code>x</code> and <code>y</code>. The `LinearRegression` API from scikit-learn requires the values for `x` and `y` to be shaped like (num_features, 1).
```
# Reshape X_train
X_train.values.reshape(-1, 1).shape
# Train LinearRegression
regressor = linear_model.LinearRegression()
regressor.fit(X_train.values.reshape(-1, 1), y_train.values.reshape(-1, 1))
print(regressor.coef_[0][0])
print(regressor.intercept_[0])
```
<a name="2-1-2"></a>
# Making Predictions
Now let's make some predictions.
**Important Note:** Always use the test set for predictions. You should never use the train data for predictions.
```
# Use the test dataset for predictions
y_pred = regressor.predict(X_test.values.reshape(-1, 1)).flatten()
y_pred
# Let's compare the actual values with the predictions
df = pd.DataFrame({"Actual": y_test, "Predicted": y_pred})
df
```
<div class="alert alert-success">
<h2>Exercise 4</h2>
Description:
- Given `actual` and `predicted` calculate the error using `Mean Absolute Error` and `Mean Squared Error`.
<br/>
<details>
<summary><b>→ Hints</b></summary>
- Use `np.abs` and `.mean()` to calculate the `Mean Absolute Error`.
- Use `np.squared` and `.mean()` to calculate the `Mean Squared Error`. Alternatively you can calculate it like: `((actual-predicted)**2).mean()`
</details>
<br/>
<br/>
<details>
<summary>
<b>→ Solution</b>
</summary>
```python
actual = df.iloc[:,0]
predicted = df.iloc[:,1]
mae = np.abs(actual-predicted).mean()
mse = np.square(actual-predicted).mean()
```
</details>
</div>
<a name="3"></a>
# 3. Unsupervised Learning
According to [Wikipedia](https://en.wikipedia.org/wiki/Unsupervised_learning):
> Unsupervised learning is a type of machine learning that looks for previously undetected patterns in a data set with no pre-existing labels and with a minimum of human supervision. In contrast to supervised learning that usually makes use of human-labelled data, unsupervised learning, also known as self-organization allows for modelling of probability densities over inputs.[1] It forms one of the three main categories of machine learning, along with supervised and reinforcement learning. Semi-supervised learning, a related variant, makes use of supervised and unsupervised techniques.
... Two of the main methods used in unsupervised learning are principal component and cluster analysis.
<a name="3-1"></a>
## 3.1 Clustering <a name="clustering"></a>
Cluster analysis or clustering is the task of grouping a set of objects in such a way that objects in the same group (called a cluster) are more similar (in some sense) to each other than to those in other groups (clusters).
Sources: [Wikipedia](https://en.wikipedia.org/wiki/Cluster_analysis)
<a name="3-2"></a>
## 3.2 Principal Component Analysis (PCA) <a name="pca"></a>
PCA is a common tool used in exploratory data analysis and in machine learning for predictive models.
The main idea of principal component analysis (PCA) is to reduce the dimensionality of a data set consisting of many variables correlated with each other, either heavily or lightly, while retaining the most representative features in the dataset.
Sources: [Wikipedia](https://en.wikipedia.org/wiki/Principal_component_analysis)
There is a session dedicated to unsupervised learning, we will provide examples of the different techniques mentioned here for clustering and dimensionality reduction.
## References and further reading
The following sources have been used in the creation of this notebook:
- [One Hot Encoding Examples](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OneHotEncoder.html)
- [Label Encoding](https://scikit-learn.org/stable/modules/preprocessing_targets.html#preprocessing-targets)
- [Supervised Learning](https://en.wikipedia.org/wiki/Supervised_learning)
- [Unsupervised Learning](https://en.wikipedia.org/wiki/Unsupervised_learning)
- [Normalisation](https://en.wikipedia.org/wiki/Normalization_(statistics))
- [Overfitting](https://en.wikipedia.org/wiki/Overfitting)
- [SciPy](https://www.scipy.org/)
- [RAPIDS](https://rapids.ai/start.html)
| github_jupyter |
# DataSet class walkthrough
In this notebook we are going to go through the mighty `DataSet` class, and get an overview of the most of it's methods and properties. In this case, the more you know, the more you can achieve.
## Note on DataSet object creation
Users are expected to obtain `DataSet` objects in the following ways:
* run a `Measurement` and get a `DataSet` via a `DataSaver` after exiting the measurement context, as shown here:
```python
...
meas = Measurement(...)
...
with meas.run() as datasaver:
...
dataset = datasaver.dataset # <- this one
...
```
* load a `DataSet` using one of the `load_*` functions from `qcodes.dataset.data_set` module, for example, `load_by_run_spec`:
```python
dataset = load_by_run_spec(
experiment_name='pinchoff',
sample_name='SDH-X-13c',
captured_run_id=12
)
```
Users should **NOT** instantiate a `DataSet` object via its constructor, or via the `new_data_set` function.
## Preparation: a DataSet from a dummy Measurement
In order to obtain a `DataSet` object, we are going to run a `Measurement` storing some dummy data (see [Dataset Context Manager](Dataset%20Context%20Manager.ipynb) notebook for more details).
```
import tempfile
import os
import time
import numpy as np
import qcodes
from qcodes import initialise_or_create_database_at, \
load_or_create_experiment, Measurement, Parameter, \
Station
from qcodes.dataset.plotting import plot_dataset
db_path = os.path.join(tempfile.gettempdir(),
'data_access_example.db')
initialise_or_create_database_at(db_path)
experiment = load_or_create_experiment(
experiment_name='greco',
sample_name='draco')
# This parameter is created in order to
# justify creation of a `Station` object,
# which, in turn, is needed to remind the
# reader about the importance of adding
# all instruments to the station.
fs = Parameter(name='force_side', label='Side of the Force',
initial_value='dark',
set_cmd=None, get_cmd=None)
station = Station()
station.add_component(fs)
x = Parameter(name='x', label='Voltage', unit='V',
set_cmd=None, get_cmd=None)
t = Parameter(name='t', label='Time', unit='s',
set_cmd=None, get_cmd=None)
y = Parameter(name='y', label='Voltage', unit='V',
set_cmd=None, get_cmd=None)
y2 = Parameter(name='y2', label='Current', unit='A',
set_cmd=None, get_cmd=None)
meas = Measurement(exp=experiment,
name='first_run',
station=station)
meas.register_parameter(x)
meas.register_parameter(t)
meas.register_parameter(y, setpoints=(x, t))
meas.register_parameter(y2, setpoints=(x, t))
x_vals = np.linspace(-4, 5, 50)
t_vals = np.linspace(-500, 1500, 25)
with meas.run() as datasaver:
for xv in x_vals:
for tv in t_vals:
yv = np.sin(2*np.pi*xv)*np.cos(2*np.pi*0.001*tv) + 0.001*tv
y2v = np.sin(2*np.pi*xv)*np.cos(2*np.pi*0.001*tv + 0.5*np.pi) - 0.001*tv
datasaver.add_result((x, xv), (t, tv), (y, yv), (y2, y2v))
time.sleep(1.0)
dataset = datasaver.dataset
```
For the sake of demonstrating what kind of data we've produced, let's use `plot_dataset` to make some default plots of the data.
```
plot_dataset(dataset)
```
## Note on immutability of DataSet
Users are NOT expected to use `DataSet`s methods that modify its content, for example, `add_result`. Only `Measurement` and `DataSaver` objects use those methods. This is due to the philosophy of the `DataSet` - once created within a `Measurement` context, it should not change so that the user don't accidentally modify/remove the important measured data. In the future versions of QCoDeS the design of the `DataSet` object will be improved to make the described philosophy clear.
## DataSet indentity
Before we dive into what's in the `DataSet`, let's briefly note how a `DataSet` is identified.
TL;DR:
> * Both `experiment name` and `sample name` are great means for identifying a dataset, especially when coupled with the `captured_run_id`.
> * `guid` is the only globally unique identifier for datasets.
### GUID
Globally, "in this universe", a dataset is identified by a globally unique identifier:
```
dataset.guid
```
`guid` is generated when a dataset is created, and is globally unique.
Due to the nature of the `guid` and its size, it is very user-friendly. For example, it is convenient to use it as a reference in a conversation between users. However, the `guid` is crucial for use in automated/computer systems, for example, when moving a dataset from one database file to a different one.
### Captured run ID
Within a given single database file (!) a dataset can be identified by its `captured_run_id`, an integer index:
```
dataset.captured_run_id
```
`captured_run_id` gets assigned to a `DataSet` at its creation time such that the new `captured_run_id` is always one larger than the current number of `DataSet`s in the database file:
```
captured_run_id_for_new_dataset = 1 + current_total_number_of_datasets_in_the_database
```
Note that `captured_run_id` is preserved when exporting the `DataSet` to another database file but may not be unique if combined with datasets from other database files. For example, it is possible to export 2 datasets with the same `captured_run_id` to the same database file.
We will see below how to disambiguate multiple datasets that share the same `captured_run_id`.
#### Note on run ID
`DataSet` object also has a `run_id`, also an integer index:
```
dataset.run_id
```
`run_id` also gets assigned to a dataset upon its creation, and due to its implementation is unique only within that database file. Unlike `captured_run_id`, `run_id` is **NOT** preserved when exporting a dataset from one database file to another, hence it is **NOT** recommended for any use.
### Experiment and sample names
Every dataset contains the name of the experiment, and the name of the sample - those come from the `Experiment` object that was passed to the `Measurement` object with which the dataset has been created.
So here is the experiment name that has been used above:
```
dataset.exp_name
```
And here is the sample name that has been used above:
```
dataset.sample_name
```
Both experiment name and sample name are great means for identifying a dataset, especially when coupled with the `captured_run_id`.
### DataSet name
Last but not least, a dataset may have a `name`. This name gets assigned to the dataset upon its creation, and is taken from the `name` attribute of the `Measurement` object that was used to create the dataset. For example, above the `name` of the `Measurement` was set by passing it as an argument to its constructor, hence the produced `DataSet` also has this name:
```
dataset.name
```
If the `name` of the `Measurement` is not explicitly defined, the created dataset gets a default name.
### Note on experiments
Conceptually experiments are meant to group datasets under the same experiment name and sample name. This is why every dataset also contains the following `Experiment`-related identifications.
Experiments have integer ids within a database file. A dataset knows about the id of the experiment it belongs to via `exp_id` property:
```
dataset.exp_id
```
Because an `Experiment` is a group of datasets, every dataset also has a `captured_counter` within an `Experiment`:
```
dataset.captured_counter
```
Similar to `captured_run_id`, `captured_counter` gets assigned to a `DataSet` upon its creation, and is preserved when exporting a dataset to another database file. The formula for `captured_counter` is:
```
captured_counter_for_new_dataset = 1 + current_total_number_of_datasets_within_the_given_experiment_in_the_database
```
#### Note on counter
Upon creation, a `DataSet` also gets a `counter` index within an `Experiment`:
```
dataset.counter
```
However, similar to `run_id`, `counter` is **NOT** preserved when exporting a dataset to a different database file, hence it is **NOT** recommended for any use.
## DataSet info
In this section we are going to look at different minor pieces of information about the dataset.
### Database file
Dataset is obviously aware of which database file it is contained in:
```
dataset.path_to_db
```
### Timestamps
A dataset has two associated timestamps: `run_timestamp` of the moment when it has been started (say, when the measurement was started), and `completed_timestamp` of the moment when it has been completed (say, when the measurement was finished). Both of the timestamps are available as `*_raw` properties of the `DataSet`, and as convenience methods of `DataSet` which allow formatting of the raw timestamps.
```
dataset.run_timestamp_raw
dataset.run_timestamp()
dataset.completed_timestamp_raw
dataset.completed_timestamp()
```
### Snapshot
The snapshot of the `Station` at the moment the measurement started is stored in the `DataSet`, and is available via `snapshot` property that returns the snapshot as a Python dictionary:
```
dataset.snapshot
```
If needed, `snapshot_raw` property of the `DataSet` can be used to retrieve the snapshot in the way it is persisted - it's a JSON-formatted string:
```
dataset.snapshot_raw
```
### Parent dataset links
The feature of linking `DataSet`s is described in detail in a separate [Linking to parent datasets](Linking to parent datasets.ipynb) notebook, hence here we will just mention the `parent_dataset_links` property:
```
dataset.parent_dataset_links
```
### Note on DataSet states
The `DataSet` object publicly exposes properties which give information about its state. However, users are expected to always get `DataSet` objects in their "final", last, unmodifiable state. This means that the "state" properties of the dataset will have the following values:
```
dataset.completed
dataset.started
dataset.running
dataset.pristine
```
### Note on metadata
`DataSet` object also supports storing arbitrary "metadata" inside it, however, this feature is not complete and may be flaky and buggy at the moment. Yet, for completeness of this article we still mention the `metadata` property of the `DataSet`:
```
dataset.metadata
```
## Parameters in DataSet
In this section we are getting information about the parameters stored in the given `DataSet`.
### Interdependencies
`DataSet` object has a `description` property that returns a `RunDescriber` object. The `RunDescriber` object in turn contains information about the parameters under its `interdeps` attribute:
```
interdeps = dataset.description.interdeps
interdeps
```
This is `InterDependencies_` object, it stores information about every parameter in the form of `ParamSpecBase` objects, and the releationship between parameters via `dependencies`, `inferences`, and `standalones` attributes.
For example, the dataset that we are inspecting contains no inferences, and no standalone parameters, but it contains two dependent parameters `y` and `y2`, which both depend on independent `x` and `t` parameters:
```
interdeps.inferences
interdeps.standalones
interdeps.dependencies
ps = list(interdeps.dependencies.keys())[0]
print(f'Parameter {ps} depends on:')
for p in interdeps.dependencies[ps]:
print(f'- {p}')
```
Note that `ParamSpecBase` objects contain all the necessary information about a parameter, including its `name` and `unit`:
```
ps = list(interdeps.dependencies.keys())[0]
print(f'Parameter {ps.name!r} is in {ps.unit!r}')
```
More information on how to work with this object is provided in a separate example notebook, [Accessing data in DataSet](Accessing-data-in-DataSet.ipynb).
### Shortcuts to important parameters
For the frequently needed groups of parameters, `DataSet` object itself provides convenient methods.
For example, use `dependent_parameters` property to get a only dependent parameters of a given `DataSet`:
```
dataset.dependent_parameters
```
### Note on ParamSpec's
> `ParamSpec`s originate from QCoDeS versions prior to `0.2.0` and for now are kept for backwards compatibility. `ParamSpec`s are completely superseded by `InterDependencies_`/`ParamSpecBase` bundle and will likely be deprecated in future versions of QCoDeS together with the `DataSet` methods/properties that return `ParamSpec`s objects.
In addition to the `Interdependencies_` object, `DataSet` also holds `ParamSpec` objects (not to be confused with `ParamSpecBase` objects from above). Similar to `Interdependencies_` object, the `ParamSpec` objects hold information about parameters and their interdependencies but in a different way: for a given parameter, `ParamSpec` object itself contains information on names of parameters that it depends on, while for the `InterDependencies_`/`ParamSpecBase`s this information is stored only in the `InterDependencies_` object.
`DataSet` exposes `paramspecs` property and `get_parameters()` method, both of which return `ParamSpec` objects of all the parameters of the dataset, and are not recommended for use:
```
dataset.paramspecs
dataset.get_parameters()
```
Lastly, `DataSet` has `parameters` that returns a string with comma-separated names of all the dataset parameters (will likely be deprecated soon):
```
dataset.parameters
```
## Data access in DataSet
`DataSet` provides one main method of accessing data - `get_parameter_data`. It returns data for groups of dependent-parameter-and-its-independent-parameters in a form of a nested dictionary of `numpy` arrays:
```
dataset.get_parameter_data()
```
Additionally, `DataSet` provides the following convenient methods:
* `DataSet.get_data_as_pandas_dataframe` ([more info in Working with pandas and xarray article](Working-With-Pandas-and-XArray.ipynb))
* `DataSet.write_data_to_text_file`
More information on how to access data in a dataset is provided in a separate example notebook, [Accessing data in DataSet](Accessing-data-in-DataSet.ipynb).
### Not recommended data access methods
The following tree methods of accessing data in a dataset are not recommended for use, and will be deprecated soon:
* `DataSet.get_data`
* `DataSet.get_values`
* `DataSet.get_setpoints`
`DataSet` also provides a property `number_of_results` which reflects the size of the data the dataset holds. For our example dataset, this number is equal to the number of `x` values multiplied by the number of `t` values multiplied by the number of dependent parameters (`y` and `y2`) -- `50 * 25 * 2`:
```
dataset.number_of_results
```
## What about the rest?
An interested user might discover other methods and properties of the `DataSet` class that are not mentioned here. Those methods and properties are likely to be made private or deprecated in future QCoDeS releases... or be documented and presented for use :)
| github_jupyter |
# Enable Spot Training with Amazon SageMaker Debugger
Amazon SageMaker Debugger is a new capability of Amazon SageMaker that allows debugging machine learning training.
It lets you go beyond just looking at scalars like losses and accuracies during training and gives you full visibility into all tensors 'flowing through the graph' during training. Amazon SageMaker Debugger helps you to monitor your training in near real time using rules and would provide you alerts, once it has detected inconsistency in training flow.
Using Amazon SageMaker Debugger is a two step process: Saving tensors and Analysis.
### Saving tensors
Tensors define the state of the training job at any particular instant in its lifecycle. Debugger exposes a library which allows you to capture these tensors and save them for analysis.
### Analysis
There are two ways to get to tensors and run analysis on them. One way is to use concept called ***Rules***. For more information about a rules-based approach to analysis, see [Rules](https://github.com/awslabs/sagemaker-debugger/blob/master/docs/analysis.md#Rules). You can also perform interactive analysis in a notebook. Please refer to our other notebooks on how to do that.
## Spot Training
This notebook talks about how Amazon SageMaker Debugger feature can also be used with Spot Training. For more information related to spot training in Amazon SageMaker please see [Spot Training](https://docs.aws.amazon.com/sagemaker/latest/dg/model-managed-spot-training.html).
The examples uses a small gluon CNN model and trains it on the FashionMNIST dataset. If during the training spot instance terminates, the training and analysis of tensors will continue from the last saved checkpoint.
```
import sagemaker
import boto3
import os
from sagemaker.mxnet import MXNet
from sagemaker.debugger import Rule, rule_configs
```
### Configuring the inputs for the training job
Now call the Amazon SageMaker MXNet Estimator to kick off a training job along with enabling Debugger functionality.
- `entrypoint_script` points to the simple MXNet training script that is ran by training job
- `hyperparameters` are the parameters that will be passed to the training script.
```
# Set the SageMaker Session
sagemaker_session = sagemaker.Session()
# Define the entrypoint script
entrypoint_script='mxnet_gluon_spot_training.py'
hyperparameters = {'batch-size' : 100, 'epochs' : 5, 'checkpoint-path' : '/opt/ml/checkpoints' }
```
## Training MXNet models in Amazon SageMaker with Amazon SageMaker Debugger
Train a small MXNet CNN model with the FashonMNIST dataset in this notebook, with Amazon SageMaker Debugger enabled. This is done using an Amazon SageMaker MXNet 1.6.0 container with script mode. Amazon SageMaker Debugger currently works with Python3, so be sure to set `py_version='py3'` when creating the Amazon SageMaker Estimator.
## Enable Amazon SageMaker Debugger and Spot Training in Estimator object
Enabling Amazon SageMaker Debugger in training job can be accomplished by adding its configuration into Estimator object constructor:
```python
sagemaker_simple_estimator = MXNet(...,
# Parameters required to enable spot training.
train_use_spot_instances=True, #Set it to True to enable spot training.
train_max_wait = 10000 # This should be equal to or greater than train_max_run in seconds'
checkpoint_local_path = '/opt/ml/checkpoints/' # This is local path where checkpoints will be stored during training. Default path is /opt/ml/checkpoints'.The training script should generate the checkpoints.
checkpoint_s3_uri = 's3://bucket/prefix' # Uri to S3 bucket where the checkpoints captured by the model will be stored.
## Rule Parameter
rules = [Rule.sagemaker(rule_configs.vanishing_gradient())]
)
```
In this section, we will focus on parameters that are needed to enable Spot Training.
- `train_use_spot_instance` : This parameter should be set to 'True' to enable the spot training.
- `train_max_wait` : This parameter (in seconds) should be set equal to or greater than 'train_max_run'.
- `checkpoint_s3_uri` : This is URI to S3 bucket where the checkpoints will be stored before the spot instance terminated. Once the training is resumed, the checkpoints from this S3 bucket will be restored to 'checkpoint_local_path' in the new instance. Ensure that the S3 bucket is created in the same region as that of current session.
- `checkpoint_local_path`: This is the local path where the model will save the checkpoints perodically. The default path is set to '/opt/ml/checkpoints'. Ensure that the model under training is saving the checkpoints in this path. Note that in hyperparameters we are setting 'checkpoint-path' so that the training script will save the checkpoints in that directory.
### Rule Parameter
We are going to run the *vanishing_gradient* rule during this training. By specifying this parameter, we are enabling the Amazon SageMaker Debugger functionality to collect the *gradients* during this training. The *gradients* will be collected every 500th step as part of the default configurations for this Rule.
## How Spot Training works with Amazon SageMaker Debugger
Amazon SageMaker Debugger can be enabled even for training with Spot Instances. Spot instances can be interrupted, causing jobs to take longer to start or finish. To leverage the managed spot instance support that Amazon SageMaker provides, you need to configure your training job to save checkpoints. Amazon SageMaker copies checkpoint data from a local path to Amazon S3. When the job is restarted on a different instance, Amazon SageMaker copies the data from Amazon S3 back into the local path. The training can then resume from the last checkpoint instead of restarting.
Amazon SageMaker Debugger relies on the checkpoints mechanism to continue emitting tensors from the last saved checkpoint. The Amazon SageMaker Debugger saves the metadata containing last saved state whenver user creates a checkpoint in *checkpoint_local_path*. Along with the checkpoints, this metadata also gets saved to Amazon S3 when the instance is interrupted. Upon restart, along with the checkpoints, this metadata is also copied back to the instance. The Amazon SageMaker Debugger reads the last saved state from the metadata and continues to emit the tensors from that step. This minimizes the emission of duplicate tensors. Note that currently, the rule job continues to wait till even if the training job is interrupted.
```
# Make sure to set this to your bucket and location
# Ensure that the bucket exists in the same region as that of current region.
BUCKET_NAME = sagemaker_session.default_bucket()
LOCATION_IN_BUCKET = 'smdebug-checkpoints'
checkpoint_s3_bucket = 's3://{BUCKET_NAME}/{LOCATION_IN_BUCKET}'.format(BUCKET_NAME=BUCKET_NAME, LOCATION_IN_BUCKET=LOCATION_IN_BUCKET)
# Local path where the model will save its checkpoints.
checkpoint_local_path = '/opt/ml/checkpoints'
estimator = MXNet(
role=sagemaker.get_execution_role(),
base_job_name='smdebugger-spot-training-demo-mxnet',
train_instance_count=1,
train_instance_type='ml.m4.xlarge',
train_volume_size = 400,
entry_point=entrypoint_script,
hyperparameters = hyperparameters,
framework_version='1.6.0',
py_version='py3',
train_max_run=3600,
sagemaker_session=sagemaker_session,
# Parameters required to enable spot training.
train_use_spot_instances=True, #Set it to True to enable spot training.
train_max_wait = 3600, #This should be equal to or greater than train_max_run in seconds
checkpoint_s3_uri = checkpoint_s3_bucket, #Set the S3 URI to store the checkpoints.
checkpoint_local_path = checkpoint_local_path, #This is default path where checkpoints will be stored. The training script should generate the checkpoints.
## Rule parameter
rules = [Rule.sagemaker(rule_configs.vanishing_gradient())]
)
estimator.fit()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
pd.set_option('display.max_columns', None)
%matplotlib inline
# Loading training data
training = pd.read_csv("data/kaggle-house-prices/train.csv")
training.info()
training.sample(10)
SalesPrice = training.SalePrice
# Loading testing data
testing = pd.read_csv("data/kaggle-house-prices/test.csv")
testing.info()
training.shape, testing.shape
# Combining training and testing data into a single dataframe
df = pd.concat([training.iloc[:, 0:80], testing], ignore_index = True)
df.info()
print("Training: ", training.shape)
print("Testing: ", testing.shape)
print("Combined: ", df.shape)
```
Loaded both datasets and combined them into a single dataset for data preprocessing. When we are ready for modeling prepearion, we will split them into 2 datasets.
Amount of NA values
```
df.isnull().sum().sum() / (df.shape[0] * df.shape[1])
```
Close 6% of data points across entire dataset is NA. Find out which columns have null in them
```
def find_null_columns():
null_columns = df.isnull().sum().sort_values(ascending = False)
null_columns = null_columns[null_columns > 0]
print("Total number of records: ", df.shape[0])
print("No of columns with null values: ", len(null_columns))
print("\nMissing values per column:")
print(null_columns)
find_null_columns()
```
There are 34 columns that contain null values. Let's analyze each of these columns one by one in the order of null occurrences.
### PoolQC
How many column names contain "pool" - likely that these are pool data related.
```
def find_columns(df, search):
matched = [c for c in df.columns if search.lower() in c.lower()]
return matched
find_columns(df, "pool")
```
From the field types described above, we can see PoolArea is numeric and PoolQC is categorical. Find out where are any records where PoolArea is mentioned but PoolQC is null.
```
df.query("PoolArea >= 0 and PoolQC != PoolQC").shape
```
Out of these, how many have >0 PoolArea?
```
df.query("PoolArea > 0 and PoolQC != PoolQC")[['PoolArea', 'PoolQC']]
```
We can impute the values of PoolQC, by looking at PoolArea value. Find out how many diff values are there for PoolQC and their freq and mean PoolArea
```
df.groupby("PoolQC").PoolArea.agg([np.mean, len])
```
So, we can input Ex, Ex, Fa respectively so that PoolArea values are closer the mean values for the respective PoolQC. Replace the rest of NA values under PoolQC with "None".
```
df.loc[2420,'PoolQC'] = "Ex"
df.loc[2503,'PoolQC'] = "Ex"
df.loc[2599,'PoolQC'] = "Fa"
idx = df.query("PoolArea == 0 and PoolQC != PoolQC").index
df.loc[idx, 'PoolQC'] = "None"
df.query("PoolArea >= 0 and PoolQC != PoolQC").shape
```
### Garage Fields
Find garage fields
```
garage_cols = find_columns(df, "garage")
df[garage_cols].info()
```
Lets see what the deal is with GarageYrBlt. It seems reasonable that most houses would build a garage when the house itself was built. We can check this by seeing how many houses were built the same year their garage was built.
```
(df.GarageYrBlt == df.YearBuilt).value_counts()
```
2216 of the 2919 houses have same year for for GarageYrBlt and YearBuilt. Lets replace any of the NA’s for GarageYrBlt with the year from YearBuilt.
```
df.query("GarageYrBlt != GarageYrBlt").shape
idx = df.query("GarageYrBlt != GarageYrBlt").index
df.loc[idx, "GarageYrBlt"] = df.loc[idx, "YearBuilt"]
(df.GarageYrBlt == df.YearBuilt).value_counts()
pd.isnull(df.GarageYrBlt).value_counts()
```
So, there is no more NA in the GarageYrBlt field.
Let's look at top 10 values of GarageYrBlt and top 10 YearBuilt values to to find any anamolies
```
print(df.GarageYrBlt.sort_values(ascending = False)[:10])
print(df.YearBuilt.sort_values(ascending = False)[:10])
df[df.GarageYrBlt == 2207][["GarageYrBlt", "YearBuilt"]]
```
Replace GarageYrBlt with YearBuild for record 2593
```
df.loc[2592, "GarageYrBlt"] = 2006
```
Plot histogram of GarageYrBlt
```
df.GarageYrBlt.plot.hist(bins = 100)
plt.title("Histogram by GarageYrBlt")
plt.xlabel("GarageYrBlt")
```
Plot histogram of YearBuilt
```
df.YearBuilt.plot.hist(bins = 100)
plt.title("Histogram by YearBuilt")
plt.xlabel("YearBuilt")
```
Let's see values at lower end to find any anamolies
```
print(df.GarageYrBlt.sort_values(ascending = True)[:10])
print(df.YearBuilt.sort_values(ascending = True)[:10])
```
At the lower values there is no obvious anamoly.
That leaves 6 garage features in our dataset and 4 of them have at least 157 missing values while GarageArea and GarageCars both only have 1, thus we can assume this particular house does not have a garage at all. For the rest of the houses we can check to see that if the NA’s recorded also have 0 GarageArea and 0 GarageCars. If they do we can fill in their missing values with ‘None’ since having 0 area and 0 cars in their garage will imply that they do not have any at all.
```
df[df.GarageCond.isnull()][garage_cols].query("GarageArea > 0")
```
Only one house who had NA’s in their garage columns had an area graeteer than 0. We can fill this house in manually and set the rest of the houses NA’s to 0.
For the house with GarageArea = 360 and GarageCars = 1, but NA’s in the other columns, we can use the most frequent values for each columns from houses with a similar area and car count.
```
idx = df.query("GarageArea >= 350 and GarageArea <= 370").index
df2 = df.loc[idx, garage_cols]
df2
```
Find most common values for each column
```
def mostFrequent(col):
counts = col.value_counts()
counts = counts.sort_values(ascending = False)
return counts.index[0]
df2.apply(mostFrequent, axis = 0)
df.loc[2126, "GarageQual"] = "TA"
df.loc[2126, "GarageFinish"] = "Unf"
df.loc[2126, "GarageCond"] = "TA"
```
Now we can fill in any missing numeric values with 0 and categoric with ‘None’ since these houses recorded having 0 area and 0 cars in their garage.
```
pd.isnull(df[garage_cols]).sum()
garage_cols
def replace_with(df, col, fill_with):
idx = df[df[col].isnull()].index
df.loc[idx, col] = fill_with
for c in ["GarageType", "GarageFinish", "GarageQual", "GarageCond"]:
replace_with(df, c, "None")
for c in ["GarageCars", "GarageArea"]:
replace_with(df, c, 0.0)
pd.isnull(df[garage_cols]).sum()
```
### KitchenQual and Electrical
With only 1 missing value for KitchenQual and Electrical each we can fill in the missing value with the most frequent value from each column.
```
df[find_columns(df, "kitchen")].info()
df.KitchenQual.value_counts()
replace_with(df, "KitchenQual", "TA")
df[find_columns(df, "electrical")].info()
df.Electrical.value_counts()
replace_with(df, "Electrical", "SBrkr")
df[find_columns(df, "electrical")].info()
```
### Basement
```
basement_columns = find_columns(df, "bsmt")
df[basement_columns].info()
```
From the documetation below are the description of these fields.
- BsmtQual: Height of the basement
- BsmtCond: General condition of the basement
- BsmtExposure: Walkout or garden level basement walls
- BsmtFinType1: Quality of basement finished area
- BsmtFinSF1: Type 1 finished square feet
- BsmtFinType2: Quality of second finished area (if present)
- BsmtFinSF2: Type 2 finished square feet
- BsmtUnfSF: Unfinished square feet of basement area
- TotalBsmtSF: Total square feet of basement area
- BsmtFullBath: Basement full bathrooms
- BsmtHalfBath: Basement half bathrooms
There are 11 basement features each with at least 1 missing value. We can take a look at the subset of just these columns from our data.
```
df[df.BsmtExposure.isnull()][basement_columns]
df.BsmtExposure.value_counts()
```
Almost all of the missing values for each categoric basement feature comes from houses with 0 on each features corresponding to area. We can fill in these values with ‘None’ since these houses certainly don’t have basements. Rows 949, 1488 and 2349 are the only missing values from BsmtExposure, we can fill this with No as that is the most frequent value and these houses most likely don’t have any exposure for their basements. The rest of the basement columns corresponding to area will be filled with 0 since they likely don’t have a basement and the categoric missing values will be filled with NoBsmt.
```
df.loc[[949, 1488, 2349], 'BsmtExposure'] = 'No'
df[basement_columns].info()
for c in ["BsmtQual", "BsmtCond", "BsmtExposure", "BsmtFinType1", "BsmtFinType2"]:
replace_with(df, c, "None")
for c in ["BsmtFinSF1", "BsmtFinSF2", "BsmtUnfSF",
"TotalBsmtSF", "BsmtFullBath", "BsmtHalfBath"]:
replace_with(df, c, 0.0)
df[basement_columns].info()
find_null_columns()
```
### Exterior
```
exterior_columns = find_columns(df, "exterior")
exterior_columns
df.query("Exterior1st != Exterior1st or Exterior2nd != Exterior2nd")[exterior_columns]
df.loc[2151, "Exterior1st"] = "Other"
df.loc[2151, "Exterior2nd"] = "Other"
find_null_columns()
```
### Sale Type
SaleType, Functional and Utilities have less than 3 missing values. For SaleType we can see what the SaleCondition of the house was and use a contingency table to see which SaleType and SaleCondition overlap together the most.
```
sale_columns = find_columns(df, "sale")
sale_columns
df[df.SaleType.isnull()][sale_columns]
df.groupby(["SaleType", "SaleCondition"]).Id.count().unstack()
df.loc[2489, "SaleType"] = "WD"
find_null_columns()
```
### Functional
There is no field that could help us replace the NA value for Functional. So let replace NA with most freq value.
```
functional_columns = find_columns(df, "func")
functional_columns
df.Functional.value_counts()
replace_with(df, "Functional", "Typ")
find_null_columns()
```
### Utilities
Utilities only has 1 value for NoSeWa and the rest AllPub. We can drop this feature from our dataset as the house with ‘NoSeWa’ is from our training set and will have won’t help with any predictive modelling
```
df.Utilities.value_counts()
del df["Utilities"]
find_null_columns()
```
### MSZoning
MSZoning: The general zoning classification
MSSubClass: The building class
There are only 4 missing values for MSZoning. We can see what the subclass is for the houses with missing values for Zoning.
```
zoning_columns = ["MSZoning", "MSSubClass"]
df[df.MSZoning.isnull()][zoning_columns]
```
Find the most frequent corresponding subclass
```
df.groupby(["MSSubClass", "MSZoning"]).Id.count().unstack()
df.loc[1915, "MSZoning"] = "RM"
df.loc[2216, "MSZoning"] = "RL"
df.loc[2250, "MSZoning"] = "RM"
df.loc[2904, "MSZoning"] = "RL"
find_null_columns()
```
### Masonry
MasVnrType: Masonry veneer type
MasVnrArea: Masonry veneer area in square feet
There are 23 missing values for MasVnrArea and 24 for MasVnrType. We can see if both missing values come from the same houses
```
masonary_columns = find_columns(df, "mas")
masonary_columns
df[df.MasVnrType.isnull() | df.MasVnrArea.isnull()][masonary_columns]
```
All but one house has missing values for both columns. For houses with NA’s on both columns we can fill 0 for the area and None for the type since they likely do not have a masonry veneer. For the house with a MasVnrArea of 198 but NA for MasVnrType we can record the median areas for each type and see which type is closest to 198.
```
df.groupby("MasVnrType").MasVnrArea.agg([len, np.median])
df.loc[2610, "MasVnrType"] = "Stone"
replace_with(df, "MasVnrType", "None")
replace_with(df, "MasVnrArea", 0)
find_null_columns()
```
### LotFrontage: Linear feet of street connected to property
There are 486 missing values for LotFrontage, which is quite a lot of values to fill and we can’t just replace these with 0. We’re given that “LotFrontage: Linear feet of street connected to property.” The area of each street connected to the house property is most likely going to have a similar area to other houses in its neighborhood. We can group by each neighborhood and take the median of each LotFrontage and fill the missing values of each LotFrontage based on what neighborhood the house comes from.
```
lot_columns = find_columns(df, "lot")
lot_columns.append("Neighborhood")
lot_columns
neighborhood = df.groupby("Neighborhood").LotFrontage.agg([len, np.median])
neighborhood
df2 = df[lot_columns].merge(neighborhood,
left_on="Neighborhood", right_index=True)
df2
idx = df[df.LotFrontage.isnull()].index
df.loc[idx, "LotFrontage"] = df2.loc[idx, "median"]
df[lot_columns].merge(neighborhood,
left_on="Neighborhood", right_index=True)
find_null_columns()
```
### Fence: Fence quality
We can replace any missing vlues for Fence and MiscFeature with ‘None’ as they probably don’t have this feature with their property.
```
df.Fence.value_counts()
replace_with(df, "Fence", "None")
find_null_columns()
```
### MiscFeature
```
replace_with(df, "MiscFeature", "None")
find_null_columns()
```
### Fireplace
- Fireplaces: Number of fireplaces
- FireplaceQu: Fireplace quality. FireplaceQu denotes the fireplace quality.
We can check to see if any of the missing values for FireplaceQu come from houses that recorded having at least 1 fireplace.
```
fire_columns = find_columns(df, "fire")
fire_columns
df.groupby(["FireplaceQu", "Fireplaces"]).Id.count().unstack()
df[df.FireplaceQu.isnull()][fire_columns]
```
All the houses that have missing values did not record having any fireplaces. We can replace the NA’s with ‘None’ since these houses don’t have any fireplaces at all.
```
replace_with(df, "FireplaceQu", "None")
find_null_columns()
```
### Alley: Type of alley access
There are 2721 missing values for Alley and only 2 potential options - Grvl and Pave. We can fill ‘None’ for any of the houses with NA’s as these houses must not have any type of alley access.
```
replace_with(df, "Alley", "None")
find_null_columns()
```
# Congratulations! There is no more missing values
Let's add back the SalePrice column to the dataset. Note for the testing data, there is not SalePrice. So for those, the field will be null.
```
df.head()
df["SalesPrice"] = np.nan
df.loc[0:training.shape[0], "SalesPrice"] = SalesPrice
df.head()
df.to_csv("data/kaggle-house-prices/data_combined_cleaned.csv", index = False)
```
# Visualization
Plot the following for train.csv
- Saleprice - histogram or boxplot
- YearBuilt - histogram (better) or frequency plot
- Saleprice vs YearBuilt - horizontal bar chart
- Saleprice vs GarageArea - scatter chart
- Median Saleprice by SaleCondition and BldgType - heatmap
```
xlims = training.SalePrice.min(), training.SalePrice.max()
plt.subplot(2, 1, 1)
training.SalePrice.plot.hist(bins = 50, xlim = xlims, sharex = True, title = "Histogram of SalePrice")
plt.subplot(2, 1, 2)
training.SalePrice.plot.box(vert = False, xlim = xlims, sharex = True, title = "Boxplot of SalePrice")
plt.tight_layout()
plt.xlabel("SalePrice")
training.YearBuilt.plot.hist(bins = 50, title = "Frequency by YearBuilt")
grouped = training.groupby("YearBuilt").SalePrice.median().sort_index()
plt.figure(figsize=(15, 5))
plt.bar(grouped.index, grouped)
plt.xlabel("YearBuilt")
plt.ylabel("Median SalePrice")
plt.title("Median SalePrice by YearBuilt")
training.plot.scatter("GarageArea", "SalePrice")
plt.title("SalePrice by GarageArea")
import seaborn as sns
grouped = training.groupby(["SaleCondition","BldgType"]).SalePrice.median().unstack()
blues = sns.color_palette("Blues", n_colors=10)
sns.heatmap(grouped, cmap = blues)
plt.title("Median Saleprice \n by SaleCondition and BldgType")
```
| github_jupyter |
<a href="https://colab.research.google.com/github/jantic/DeOldify/blob/master/VideoColorizerColab.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
### **<font color='blue'> Video Colorizer </font>**
#◢ DeOldify - Colorize your own videos!
_FYI: This notebook is intended as a tool to colorize gifs and short videos, if you are trying to convert longer video you may hit the limit on processing space. Running the Jupyter notebook on your own machine is recommended (and faster) for larger video sizes._
####**Credits:**
Big special thanks to:
Robert Bell for all his work on the video Colab notebook, and paving the way to video in DeOldify!
Dana Kelley for doing things, breaking stuff & having an opinion on everything.
---
#◢ Verify Correct Runtime Settings
**<font color='#FF000'> IMPORTANT </font>**
In the "Runtime" menu for the notebook window, select "Change runtime type." Ensure that the following are selected:
* Runtime Type = Python 3
* Hardware Accelerator = GPU
#◢ Git clone and install DeOldify
```
!git clone https://github.com/jantic/DeOldify.git DeOldify
cd DeOldify
```
#◢ Setup
```
#NOTE: This must be the first call in order to work properly!
from deoldify import device
from deoldify.device_id import DeviceId
#choices: CPU, GPU0...GPU7
device.set(device=DeviceId.GPU0)
import torch
if not torch.cuda.is_available():
print('GPU not available.')
from os import path
!pip install -r colab_requirements.txt
import fastai
from deoldify.visualize import *
from pathlib import Path
torch.backends.cudnn.benchmark=True
!mkdir 'models'
!wget https://www.dropbox.com/s/336vn9y4qwyg9yz/ColorizeVideo_gen.pth?dl=0 -O ./models/ColorizeVideo_gen.pth
!wget https://media.githubusercontent.com/media/jantic/DeOldify/master/resource_images/watermark.png -O ./resource_images/watermark.png
colorizer = get_video_colorizer()
```
#◢ Instructions
### source_url
Type in a url hosting a video from YouTube, Imgur, Twitter, Reddit, Vimeo, etc. Many sources work! GIFs also work. Full list here: https://ytdl-org.github.io/youtube-dl/supportedsites.html NOTE: If you want to use your own video, upload it first to a site like YouTube.
### render_factor
The default value of 21 has been carefully chosen and should work -ok- for most scenarios (but probably won't be the -best-). This determines resolution at which the color portion of the video is rendered. Lower resolution will render faster, and colors also tend to look more vibrant. Older and lower quality film in particular will generally benefit by lowering the render factor. Higher render factors are often better for higher quality videos and inconsistencies (flashy render) will generally be reduced, but the colors may get slightly washed out.
### watermarked
Selected by default, this places a watermark icon of a palette at the bottom left corner of the image. This is intended to be a standard way to convey to others viewing the image that it is colorized by AI. We want to help promote this as a standard, especially as the technology continues to improve and the distinction between real and fake becomes harder to discern. This palette watermark practice was initiated and lead by the company MyHeritage in the MyHeritage In Color feature (which uses a newer version of DeOldify than what you're using here).
### How to Download a Copy
Simply right click on the displayed video and click "Save video as..."!
## Pro Tips
1. If a video takes a long time to render and you're wondering how well the frames will actually be colorized, you can preview how well the frames will be rendered at each render_factor by using the code at the bottom. Just stop the video rendering by hitting the stop button on the cell, then run that bottom cell under "See how well render_factor values perform on a frame here". It's not perfect and you may still need to experiment a bit especially when it comes to figuring out how to reduce frame inconsistency. But it'll go a long way in narrowing down what actually works.
2. If videos are taking way too much time for your liking, running the Jupyter notebook VideoColorizer.ipynb on your own machine (with DeOldify installed) will generally be much faster (as long as you have the hardware for it).
## Troubleshooting
The video player may wind up not showing up, in which case- make sure to wait for the Jupyter cell to complete processing first (the play button will stop spinning). Then follow these alternative download instructions
1. In the menu to the left, click Files
2. If you don't see the 'DeOldify' folder, click "Refresh"
3. By default, rendered video will be in /DeOldify/video/result/
If a video you downloaded doesn't play, it's probably because the cell didn't complete processing and the video is in a half-finished state.
#◢ Colorize!!
```
source_url = '' #@param {type:"string"}
render_factor = 21 #@param {type: "slider", min: 5, max: 40}
watermarked = True #@param {type:"boolean"}
if source_url is not None and source_url !='':
video_path = colorizer.colorize_from_url(source_url, 'video.mp4', render_factor, watermarked=watermarked)
show_video_in_notebook(video_path)
else:
print('Provide a video url and try again.')
```
## See how well render_factor values perform on a frame here
```
for i in range(10,40,2):
colorizer.vis.plot_transformed_image('video/bwframes/video/00001.jpg', render_factor=i, display_render_factor=True, figsize=(8,8))
```
---
#⚙ Recommended video and gif sources
* [/r/Nickelodeons/](https://www.reddit.com/r/Nickelodeons/)
* [r/silentmoviegifs](https://www.reddit.com/r/silentmoviegifs/)
* https://twitter.com/silentmoviegifs
| github_jupyter |
```
# This notebook calculates the returns over 1 through 15 bars for trading signals.
# Every signal is evaluated so this return does not reflect the returns that would have been realized
# by entering trades. Data is pulled from Yahoo finance for any given symbol. Other data sources may
# be substituted in this cell. The key fields are "Open", "High", "Low", and "Close".
# Import Yahoo finance and talib (for ATR calculations)
import yfinance as yf
import talib
import numpy as np
import random
from matplotlib.ticker import PercentFormatter
import matplotlib.pyplot as plt
# Get the desired ticker symbol
sym = " "
sym = input("Enter Symbol: ")
# Get test interval
print("valid intervals: 1m,2m,5m,15m,30m,60m,90m,1h,1d,5d,1wk,1mo,3mo")
print("Intraday if period < 60 days)")
test_interval = ""
test_interval = input("Test Interval")
raise_error = 0
if test_interval == "1d" or test_interval == "5d" or test_interval == "1wk" or test_interval == "1mo" or test_interval == "3mo":
# Set the desired Start and End Dates
start_date = "2000-06-01"
end_date = "2021-06-15"
elif test_interval == "1m" or test_interval == "2m" or test_interval == "5m" or test_interval == "15m" or test_interval == "30m" or test_interval == "60m" or test_interval == "90m":
# Set the desired Start and End Dates
start_date = "2021-05-01"
end_date = "2021-06-15"
else:
raise_error = 1
print("Invalid test interval")
if raise_error == 0:
# Get data from Yahoo Finance
# fetch data by interval (including intraday if period < 60 days)
# valid intervals: 1m,2m,5m,15m,30m,60m,90m,1h,1d,5d,1wk,1mo,3mo
# (optional, default is '1d')
# For more data an alternative to Yahoo may be required
ticker = yf.Ticker(sym)
df = ticker.history(start=start_date, end=end_date, interval=test_interval)
df = df.dropna()
print(df.head(5))
# TRADING SIGNAL GENERATION ALGORITHM
# The result of this cell is a trading signal AND a support level. The output of this cell
# must be a df['SIGNAL'] AND a df['SUPPORT'] column added to the dataframe.
# Any signal generation can be substituted as long as the result is a df['SIGNAL'] and
# df['SUPPORT'] value.
# Awesome Osc
# This signal is based on Bill Williams momentum oscillator.
# There are multiple trade signal methods used for this indicator.
# This current implementation uses the most basic.
# AWESOME
# Fast AverageFC
df['MedianPrice'] = (df['High'] + df['Low']) * 0.5
df['FC_5'] = df['MedianPrice'].rolling(5).sum()
df['AvgFast'] = df['FC_5']/5
# Slow AverageFC
df['FC_34'] = df['MedianPrice'].rolling(34).sum()
df['AvgSlow'] = df['FC_34']/34
df['AO'] = df['AvgFast'] - df['AvgSlow']
# BUY SETUPS
# TSEL: AOPivotLow = CurrentBar > 2 and AO > AO[1] and AO[1] < AO[2] ;
df['AOPivotLow'] = 0
df.loc[(df['AO'] > df['AO'].shift(1)) & (df['AO'].shift(1) < df['AO'].shift(2)), 'AOPivotLow'] = 1
# TSEL: AOSaucerBuy = AO > 0 and AO[1] > 0 and AO[2] > 0 and AOPivotLow ;
df['AOSaucerBuy'] = 0
df.loc[(df['AO'] > 0) & (df['AO'].shift(1) > 0) & (df['AO'].shift(2) > 0) &
(df['AOPivotLow'] == 1), 'AOSaucerBuy'] = 1
# TSEL: AOCrossBuy = AO > 0 and AO[1] < 0 ;
df['AOCrossBuy'] = 0
df.loc[(df['AO'] > 0) & (df['AO'].shift(1) < 0), 'AOCrossBuy'] = 1
df['Awesome'] = 0
df.loc[(df['AOSaucerBuy'] + df['AOCrossBuy'] > 0), 'SIGNAL'] = 1
# Set the support leve to the low of the signal bar - Other values can be tested.
df.loc[(df['SIGNAL'] == 1), 'SUPPORT'] = df['Low']
print(df.head(5))
# Determine the frequency of the trading signal
# How many bars are in the file?
bars = len(df)
# How many signals were generated?
signals = df['SIGNAL'].sum()
# Signal Frequency
cl_freq = 0
if bars != 0:
cl_freq = signals/bars
print("There were ", signals, " signals")
print("Probablity of Trading Signal: ", '{:2.2%}'.format(cl_freq))
# CLOSING PRICE CALCULATIONS - Calculate the returns based upon the closing price of the signal bar.
# This assumes that we are watching the market and could possibly enter on the close of the signal bar.
# Calculate the returns for the signals. Since I am using the zero element of the array I want to adjust the
# shift value to start at 1 and the name of the RetCL column to reflect the bar number in the analysis.
for i in range(0,15):
df.loc[(df['SIGNAL'] == 1), 'RetCL' + str(i + 1)] = np.log(df['Close'].shift(-(i+1))/df['Close'])
# Find the mean and slope of the returns. The mean is calcaluted over the full data population (versus a moving avearge)
ret_close = [0]*15
ret_slope_close = [0]*15
for i in range(0,15):
ret_close[i] = np.nanmean(df['RetCL' + str(i + 1)])
if i > 1:
ret_slope_close[i] = (ret_close[i] - ret_close[0])/i
# Set up the plot x axis and zero reference for future plots
bar_count = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15]
zeros = np.zeros_like(ret_close)
# Plot the return results
plt.title("RETURNS BASED ON CLOSING PRICE")
plt.gca().yaxis.set_major_formatter(PercentFormatter(1))
plt.plot(bar_count, ret_close)
plt.plot(bar_count, zeros)
plt.xlabel('Bars After Signal')
plt.ylabel('Returns (Close)')
print("")
# Plot the slope of the returns over the 15 bar period. Slope doesn't become interesing until the 3rd bar...
plt.title("SLOPE OF THE RETURNS BASED ON CLOSING PRICE")
plt.plot(bar_count[2:14], ret_slope_close[2:14])
plt.plot(bar_count[2:14], zeros[2:14])
plt.xlabel('Bars After Signal')
plt.ylabel('Slope of Returns (Close)')
print("")
# RANDOM SIGNAL COMPARISON WITH SIGNAL BASED ON CLOSING PRICE
# Calculate returns based on a random signals. Calculation is repeated 20 times
# and compared to the returns generated by the signal. We are only going to
# compare the returns and the slope.
# Set the number of trials
cl_trials = 20
# Set up signal versus random performance comparison array
svr_close = [0]*15
svr_slope_close = [0]*15
print("Frequency of Signal under evaluation (Close): ", '{:2.2%}'.format(cl_freq))
# Do "x" iterations of random experiments
for i in range(0,cl_trials):
# Generate a random number for each line/bar in the file
df['CLX'] = 0
for j, row in df.iterrows():
df.at[j,'cl_x'] = random.random()
df.loc[(df['cl_x'] <= cl_freq), 'CLX'] = 1
# How many random signals were generated?
r_cl_signals = df['CLX'].sum()
# Signal Frequency
r_cl_freq = 0
if bars != 0:
r_cl_freq = r_cl_signals/bars
print("Frequency of Random Signal: ", '{:2.2%}'.format(r_cl_freq))
# Clean the random returns column
for j in range(0,15):
df['RandomCL' + str(j + 1)] = np.nan
# Calculate the random returns
for j in range(0,15):
df.loc[(df['CLX'] == 1), 'RandomCL' + str(j + 1)] = np.log(df['Close'].shift(-(j+1))/df['Close'])
# Calculate the average returns per bar
random_close = [0]*15
random_slope_close = [0]*15
for j in range(0,15):
random_close[j] = np.nanmean(df['RandomCL' + str(j + 1)])
if j > 1:
random_slope_close[j] = (random_close[j] - random_close[0])/j
# Compare the random trial returns to the trading signal returns
for j in range(0,15):
if ret_close[j] > random_close[j]:
svr_close[j] += 1
# Compare the random trial returns slope to the trading signal returns slope
for j in range(2,15):
if ret_slope_close[j] > random_slope_close[j]:
svr_slope_close[j] += 1
# Get the average number of times the signal was better than random
for i in range(0,15):
svr_close[i] /= cl_trials
svr_slope_close[i] /= cl_trials
plt.title("SIGNAL VERSUS RANDOM TRIALS - RETURNS (CLOSE)")
plt.gca().yaxis.set_major_formatter(PercentFormatter(1))
plt.plot(bar_count, svr_close)
plt.xlabel('Bars After Signal')
plt.ylabel('Percentage Signal Outperforms')
print("")
plt.title("SIGNAL VERSUS RANDOM TRIALS - SLOPE OF RETURNS (CLOSE)")
plt.gca().yaxis.set_major_formatter(PercentFormatter(1))
plt.plot(bar_count[2:14], svr_slope_close[2:14])
plt.xlabel('Bars After Signal')
plt.ylabel('Percentage Signal Outperforms')
print("")
# BUY STOP SCENARIO - Assume we wait until a buy stop order would have filled.
# Calculate returns on the high price of the signal bar - assuming the next bar takes out the high
# This is a buy stop scenario.
df.loc[(df['SIGNAL'] == 1) & (df['High'].shift(-1) > df['High']), 'BS_fill'] = 1
# How many fills for the buy stop?
bs_fill = df['BS_fill'].sum()
# Signal Frequency
bs_freq = 0
if bars != 0:
bs_freq = bs_fill/bars
print("There were ", int(bs_fill), " trades")
print("Probability of Trading Signal: ", '{:2.2%}'.format(bs_freq))
# Calculate "Buy Stop" returns
# Again, I want to use "+1" with the shift and column name. The return is now based on the high price of
# the signal bar
for i in range(0,15):
df.loc[(df['BS_fill'] == 1), 'RetBS' + str(i + 1)] = np.log(df['Close'].shift(-(i+1))/df['High'])
ret_buystop = [0]*15
ret_slope_buystop = [0]*15
for i in range(0,15):
ret_buystop[i] = np.nanmean(df['RetBS' + str(i + 1)])
if i > 0:
ret_slope_buystop[i] = (ret_buystop[i] - ret_buystop[0])/i
plt.title("RETURNS BASED ON BUY STOP")
plt.gca().yaxis.set_major_formatter(PercentFormatter(1))
plt.plot(bar_count, ret_buystop)
plt.plot(bar_count, zeros)
plt.xlabel('Bars After Signal')
plt.ylabel('Returns (Buy Stop)')
print("")
# Plot the slope of the returns over the 15 bar period
plt.title("SLOPE OF RETURNS BASED ON BUY STOP")
plt.plot(bar_count[2:14], ret_slope_buystop[2:14])
plt.plot(bar_count[2:14], zeros[2:14])
plt.xlabel('Bars After Signal')
plt.ylabel('Slope of Returns (Buy Stop)')
print("")
# RANDOM SIGNAL COMPARISON WITH SIGNAL BASED ON BUY STOP
# Calculate returns based on a random signals. Calculation is repeated 20 times
# and compared to the returns generated by the signal.
# Set the number of trials
bs_trials = 20
# Set up signal versus random performance comparison array
svr_buystop = [0]*15
svr_slope_buystop = [0]*15
print("Frequency of Signal under evaluation (Buy Stop): ", '{:2.2%}'.format(bs_freq))
# Do "x" iterations of random experiments
for i in range(0,bs_trials):
# Generate a random number for each line/bar in the file
df['BSX'] = 0
for j, row in df.iterrows():
df.at[j,'bs_x'] = random.random()
df.loc[(df['bs_x'] <= bs_freq), 'BSX'] = 1
# How many random signals were generated?
r_bs_signals = df['BSX'].sum()
# Signal Frequency
r_bs_freq = 0
if bars != 0:
r_bs_freq = r_bs_signals/bars
print("Frequency of Random Signal: ", '{:2.2%}'.format(r_bs_freq))
# Clean the random returns column
for j in range(0,15):
df['RandomBS' + str(j + 1)] = np.nan
# Calculate the random returns
for j in range(0,15):
df.loc[(df['BSX'] == 1), 'RandomBS' + str(j + 1)] = np.log(df['Close'].shift(-(j+1))/df['Close'])
# Calculate the average returns per bar
random_buystop = [0]*15
random_slope_buystop = [0]*15
for j in range(0,15):
random_buystop[j] = np.nanmean(df['RandomBS' + str(j + 1)])
if j > 0:
random_slope_buystop[j] = (random_buystop[j] - random_buystop[0])/j
# Compare the random trial returns to the trading signal returns
for j in range(0,15):
if ret_buystop[j] > random_buystop[j]:
svr_buystop[j] += 1
# Compare the random trial returns slope to the trading signal returns slope
for j in range(2,15):
if ret_slope_buystop[j] > random_slope_buystop[j]:
svr_slope_buystop[j] += 1
# Get the average number of times the signal was better than random
for i in range(0,15):
svr_buystop[i] /= bs_trials
svr_slope_buystop[i] /= bs_trials
plt.title("SIGNAL VERSUS RANDOM TRIALS - RETURNS (BUY STOP)")
plt.gca().yaxis.set_major_formatter(PercentFormatter(1))
plt.plot(bar_count, svr_close)
plt.xlabel('Bars After Signal')
plt.ylabel('Percentage Signal Outperforms')
print("")
plt.title("SIGNAL VERSUS RANDOM TRIALS - SLOPE OF RETURNS (BUY STOP)")
plt.gca().yaxis.set_major_formatter(PercentFormatter(1))
plt.plot(bar_count[2:14], svr_slope_buystop[2:14])
plt.xlabel('Bars After Signal')
plt.ylabel('Percentage Signal Outperforms')
print("")
```
| github_jupyter |
```
from PIL import Image
import numpy as np
import matplotlib.pyplot as plt
```
## Useful to separate an image into 'semantic' components: Things are different
<img src="images4/segmentation.png">
### Let's start with a simple example. How would you do this?
```
img = Image.open('images4/apple.png')
img
data = np.array(img)
data.shape
mask = data[:,:,0] > 120
mask.shape
plt.imshow(mask, cmap='gray')
mask = np.tile(mask.reshape(514, 496, 1), (1,1,4))
mask.shape
plt.imshow(data * mask)
```
### Takeaway: An object (or a portion of the object) is similarly coloured across its surface
### Something we can do now is to look at neighbouring regions of these pixels, since the pixels of an object are also *Close together*
# How do we automatically find "similar" pixels (close together in RGB and distance)?
### Clustering - Finding groups of things (eg. people or pixels), that have similar values (eg. height, or color)
```
X = np.array([1,2,2,5,7,6,4,6,8,7,8])
Y = np.array([2,1,2,8,7,8,4,6,1,1,3])
plt.scatter(X,Y)
plt.xlim(0, 10)
plt.ylim(0, 10)
joint = np.array([[x,y] for x,y in zip(X,Y)])
joint
```
### Complete the below functions (answers filled in)
```
def label(x, y):
#decide which cluster (among y) x should be assigned to
x_repeated = np.tile(x,(y.shape[0], 1))
distances = np.sum((y-x)**2, axis=1)
label = np.argmin(distances)
return label
def pairwise_distances_argmin(X, centers):
#for every point decide which cluster it should belong to
labels = []
for i in range(X.shape[0]):
labels.append(label(X[i], centers))
return np.array(labels)
def find_clusters(X, n_clusters, rseed=2):
rng = np.random.RandomState(rseed)
i = rng.permutation(X.shape[0])[:n_clusters]
centers = X[i]
all_labels = []
all_centers = []
while True:
labels = pairwise_distances_argmin(X, centers)
new_centers = np.array([X[labels == i].mean(0)
for i in range(n_clusters)])
if np.all(centers == new_centers):
break
centers = new_centers
all_labels.append(labels)
all_centers.append(centers)
return all_centers, all_labels
```
### Let's run it on the toy dataset
```
all_centers, all_labels = find_clusters(joint, 3)
print(len(all_centers), len(all_labels))
print(all_labels)
print(all_centers)
step = 1
centers = all_centers[-step]
labels = all_labels[-step]
plt.scatter(X[labels == 0],Y[labels == 0], color='yellow')
plt.scatter(X[labels == 1],Y[labels == 1])
plt.scatter(X[labels == 2],Y[labels == 2])
plt.scatter(centers[:,0], centers[:,1], marker='x', s=100)
plt.xlim(0, 10)
plt.ylim(0, 10)
```
### This is to test the performance of k-Means on a sample
```
raw_img = Image.open('images4/apple.png')
raw_img
img = np.array(raw_img)
from sklearn.cluster import KMeans
alpha = 0.1
normalized = (img/255.0)
values = []
for x in range(normalized.shape[0]):
for y in range(normalized.shape[1]):
x_norm = float(x)/normalized.shape[0]
y_norm = float(y)/normalized.shape[1]
values.append([alpha * x_norm, alpha * y_norm,normalized[x,y,0], normalized[x,y,1], normalized[x,y,2]])
#values.append([normalized[x,y,0], normalized[x,y,1], normalized[x,y,2]])
kmeans = KMeans(n_clusters=4, random_state=0).fit(values)
kmeans.cluster_centers_
segments = kmeans.labels_.reshape(normalized.shape[0], normalized.shape[1])
plt.imshow(segments)
```
| github_jupyter |
# Disjunctive Programming
TODO: Rewrite following new style guide
TODO: MS ... contrast bigm vs gdp
## Television advertising
A business manager for a local television station needs to select advertisements for a 120 second slot during an evening news broadcast. The station's sales department has prepared a list of candidate advertisements detailing the client, ad titile, length in seconds, and the revenue eared by running the ad in that time slot.
| Client | Title | Length | Revenue |
| :----- | :---- | :----: | :-----: |
| Super Cola | "Best Cola Ever" | 30 | 520 |
| Cheap Cola | "Good Cola, Low Price" | 15 | 250 |
| Corner Mart | "20% off sale" | 15 | 300 |
| Pat Smith for Mayo | "Vote Smith for Mayor" | 30 | 600 |
| Pat Smith for Mayor | "Vote Smith for Mayor" | 60 | 1300 |
| Pat Smith for Mayor | "Vote Smith for Mayor" | 15 | 400 |
| Central Grocers | "Great coffee" | 10 | 250 |
| Quick Auto | "Good cars at low prices" | 15 | 280 |
| Fancy Auto | "Fancy cars at high prices" | 30 | 480 |
| Hollywood Studios | "Professor MegaMath saves the World" | 30 | 540 |
| Childrens Book Store | "Funny Stories" | 15 | 10 | 200 |
Any unfilled time 5 seconds or longer would be use to promote other programming with an assumed value of 10 € per seconnd.
There are some additional considerations in selecting ads for thi
```
import pandas as pd
ads = pd.DataFrame([
["Super Cola", "Best Cola Ever", 30, 520],
["Cheap Cola", "Good Cola, Low Price", 15, 250],
["Corner Mart", "20% Sale", 15, 300],
["Pat Smith for Mayor", "Vote Smith for Mayor", 30, 600],
["Pat Smith for Mayor", "Vote Smith for Mayor", 60, 1300],
["Pat Smith for Mayor", "Vote Smith for Mayor", 15, 400],
["Central Grocers", "Great Coffee", 10, 250],
["Quick Auto", "Good Cars at Low Prices", 15, 200],
["Fancy Auto", "Better Cars", 30, 400],
["Hollywood Studios", "Professor MegaMeth Saves the World", 30, 540],
["Children's Bookstore", "Funny Story Books", 15, 200],
],
columns = ["client", "title", "length", "revenue"])
# these are competitors that should not appear in the same time slot
conflicts = [
["Corner Mart", "Central Grocers"],
["Super Cola", "Cheap Cola"],
["Quick Auto", "Fancy Auto"]
]
# these are clients that wish to cross promotion
joint_promotions = [
["Corner Mart", "Cheap Cola"],
["Central Grocers", "Super Cola"]
]
display(ads)
import pyomo.environ as pyo
def build_ad_model(ads, time_slot, conflicts):
m = pyo.ConcreteModel()
m.ADS = pyo.Set(initialize=ads.index)
m.CLIENTS = pyo.Set(initialize=list(set(ads["client"])))
m.CLIENT_ADS = pyo.Set(m.CLIENTS,
initialize=lambda m, client: ads[ads["client"]==client].index)
for client in m.CLIENT_ADS:
print(client, [ad for ad in m.CLIENT_ADS[client]])
m.x = pyo.Var(m.ADS, domain=pyo.Binary)
@m.Objective(sense=pyo.maximize)
def maximize_revenue(m):
return sum(m.x[i] * ads.loc[i, "revenue"] for i in m.ADS)
@m.Constraint()
def max_time(m):
return sum(m.x[i] * ads.loc[i, "length"] for i in m.ADS) <= time_slot
# @m.Constraint(m.CLIENTS)
# def no_repeat_clients(m, client):
# client_ads = ads[ads["client"]==client].index
# return sum(m.x[i] for i in client_ads) <= 1
@m.Constraint(m.CLIENT_ADS)
def no_repeat_client(m, client):
print(m.CLIENT_ADS[client])
return sum([m.x[ad] for ad in m.CLIENT_ADS[client]]) <= 1
@m.Disjunction(xor=True)
def test(m):
return [[m.x[2] == 1, m.x[4]==0], [m.x[2] == 0, m.x[4]==1]]
pyo.TransformationFactory('gdp.hull').apply_to(m)
solver = pyo.SolverFactory('cbc')
solver.solve(m)
soln = ads.copy()
soln["run"] = [int(m.x[i]()) for i in m.ADS]
return soln[soln["run"] == 1]
run_ads = build_ad_model(ads, 120, conflicts)
display(run_ads)
print(run_ads["length"].sum())
print(run_ads["revenue"].sum())
```
## Logical Relationships
MIT OpenCourseWare example: https://ocw.mit.edu/courses/sloan-school-of-management/15-053-optimization-methods-in-management-science-spring-2013/tutorials/MIT15_053S13_tut09.pdf
Example: Assign advertisers to television ad slots
```
import pyomo.environ as pyo
import pyomo.gdp as gdp
m = pyo.ConcreteModel()
pyo.TransformationFactory("gdp.bigm").apply_to(m)
pyo.SolverFactory("gurobi_direct").solve(m)
```
Disjunctive programming describes a class of optimization problems that include disjunctive ("or") constraints. These are encountered when there is a choice to be made, such as whether to do perform job A before job B on a machine, or use raw supplier X rather than Y in a supply chain, or open a retail store at location A, B, C, or D in a metropolitan area.
```
import random
import matplotlib.pyplot as plt
n_boxes = 10
boxes = dict()
for n in range(n_boxes):
w = random.randint(10, 30)
h = random.randint(10, 30)
boxes[n] = {"W": w, "H": h, "D": }
print(boxes)
fig, ax = plt.subplots(1, 1, figsize=(12, 3))
x = 0
for box in boxes.keys():
ax.rectangle(
import pyomo.environ as pyo
import pyomo.gdp as gdp
m = pyo.ConcreteModel()
m.BOXES = pyo.Set(initialize=boxes.keys())
m.width_on_shelf = pyo.Var(m.BOXES, domain=pyo.NonNegativeReals, bounds=(0, 50))
m.orientation = pyo.Var(m.BOXES, domain=pyo.NonNegativeIntegers, bounds=(1, 3))
@m.Objective(sense=pyo.minimize)
def shelf_width(m):
return sum(m.width_on_shelf[box] for box in m.BOXES)
@m.Disjunction(m.BOXES, xor=True)
def orient_box(m, box):
return [[m.orientation[box]==1,
m.width_on_shelf[box]==boxes[box]["W"]],
[m.orientation[box]==2,
m.width_on_shelf[box]==boxes[box]["H"]],
[m.orientation[box]==3,
m.width_on_shelf[box]==boxes[box]["D"]],
]
pyo.TransformationFactory("gdp.bigm").apply_to(m)
pyo.SolverFactory("cbc").solve(m)
for box in m.BOXES:
print(f"Box {box:3d} Orientation {m.orientation[box]()} Width on Shelf {m.width_on_shelf[box]():4.1f}")
```
Suppose we have a set of items to deliver to customer that need to bo packaged into boxes for express shipping. Express shipping requires each package to weigh less than 30 kg.
```
import pyomo.environ as pyo
import pyomo.gdp as gdp
# data dictionary
weights = {"A": 12, "B": 8, "C": 20, "D": 10, "E": 25}
m = pyo.ConcreteModel()
m.BOXES = pyo.RangeSet(3)
m.ITEMS = pyo.Set(initialize=weights.keys())
m.assign = pyo.Var(m.ITEMS, m.BOXES, domain=pyo.Binary)
m.use_box = pyo.Var(m.BOXES, domain=pyo.Binary)
@m.Disjunction(m.ITEMS, xor=True)
def assign_item_to_one_box(model, item):
return [[m.assign[item, box] == 1, m.use_box[box] == 1] for box in m.BOXES]
@m.Disjunction(m.BOXES, xor=True)
def assign_all_items(model, box):
return [m.assign[item, box] == 1 for item in m.ITEMS]
@m.Constraint(m.BOXES)
def weight_limit(model, box):
return sum(m.assign[item, box]*weights[item] for item in m.ITEMS) <= 30
pyo.TransformationFactory("gdp.hull").apply_to(m)
pyo.SolverFactory('gurobi_direct').solve(m)
for box in m.BOXES:
print(f"\nPacking list for Box {box}")
for item in m.ITEMS:
if m.assign[item, box]():
print(f" Item {item}: {weights[item]:5.2f} kg")
print(f" TOTAL: {m.weight_limit[box]():5.2f} kg")
```
## Logical
```
import pyomo.environ as pyo
import pyomo.gdp as gdp
m = pyo.ConcreteModel()
m.x = pyo.Var(bounds=(0, 1000))
m.y = pyo.Var(bounds=(0, 1000))
```
## Installations and imports
```
import pyomo.environ as pyo
import pyomo.gdp as gdp
import pandas as pd
```
## Problem statement
### Component data
```
# load data as dictionary of components
# component data consists of cost and composition
comp_data = {
"A": {"cost": 2.0, "Vit A": 0.5, "Vit B": 0.2},
"B": {"cost": 2.0, "Vit A": 0.4, "Vit B": 0.1},
"C": {"cost": 5.0, "Vit A": 0.3, "Vit B": 0.3},
}
# use pandas to create a nice display
pd.DataFrame.from_dict(comp_data, orient='index')
```
### Product Composition Requirements
Find the lowest cost blend
* Vit A: less than 0.4
* Vit B: greater than 0.2
Your code should be able to accept alternative specification for data and product requirements.
```
prod_req = {
"Vit A": {"lb": 0.0, "ub": 0.4},
"Vit B": {"lb": 0.2, "ub": 1.0},
}
pd.DataFrame.from_dict(prod_req, orient='index')
```
### Component Compatibility
For this application, we consider an additional type of constraint specifying the incompatability of certain blends of components. For example, suppose we have a constraint:
* A and B cannot be mixed together in the final product
The constraint is specified by creating a list of incompatabile pairs.
```
excl_pairs = [("A", "B")]
```
## Version 0: Neglecting the compatibility requirments
```
m = pyo.ConcreteModel()
# define sets that will be used to index decision variables and constraints
m.COMPONENTS = pyo.Set(initialize=comp_data.keys())
m.REQUIREMENTS = pyo.Set(initialize=prod_req.keys())
# decision variables
m.x = pyo.Var(m.COMPONENTS, domain=pyo.NonNegativeReals)
@m.Objective(sense=pyo.minimize)
def cost(m):
return sum(m.x[c]*comp_data[c]["cost"] for c in m.COMPONENTS)
@m.Constraint(m.COMPONENTS)
def mass_fraction(m, c):
return sum(m.x[c] for c in m.COMPONENTS)==1)
# com
m.lb = pyo.Constraint(m.req, rule=lambda m, r: sum(m.x[c]*comp_data[c][r] for c in m.comp) >= prod_req[r]["lb"])
m.ub = pyo.Constraint(m.req, rule=lambda m, r: sum(m.x[c]*comp_data[c][r] for c in m.comp) <= prod_req[r]["ub"])
solver = pyo.SolverFactory('cbc')
solver.solve(m)
for c in m.comp:
print(f"{c} = {m.x[c]()}")
```
## Version 1: Including compatibility requirements with Big-M
The challenge of this problem are the disjunctive constraints associated with the component incompatability data. Here we associated a boolean variable for each pair, then use the boolean variable to determine which member of the pair to keep in the blend.
```
m = ConcreteModel()
# define sets that will be used to index decision variables and constraints
# remember to use initialize keyword
m.comp = Set(initialize=comp_data.keys())
m.req = Set(initialize=prod_req.keys())
# define a set to that includes the excluded pairs
m.pairs = Set(initialize=excl_pairs)
# decision variables
m.x = Var(m.comp, domain=NonNegativeReals)
# for each excluded pair, create a boolean variable. The value of the boolean
# variable will determine which member of the pair is allowed in the product
m.y = Var(m.pairs, domain=Boolean)
# objective function
m.cost = Objective(expr=sum(m.x[c]*comp_data[c]["cost"] for c in m.comp), sense=minimize)
# structural constraints
m.massfraction = Constraint(expr=sum(m.x[c] for c in m.comp)==1)
# composition constraints
m.lb = Constraint(m.req, rule=lambda m, r: sum(m.x[c]*comp_data[c][r] for c in m.comp) >= prod_req[r]["lb"])
m.ub = Constraint(m.req, rule=lambda m, r: sum(m.x[c]*comp_data[c][r] for c in m.comp) <= prod_req[r]["ub"])
# component incompatability constraints
M = 100
m.disj = ConstraintList()
for pair in m.pairs:
a, b = pair
m.disj.add(m.x[a] <= M*m.y[pair])
m.disj.add(m.x[b] <= M*(1-m.y[pair]))
solver = SolverFactory('cbc')
solver.solve(m)
for c in m.comp:
print(f"{c} = {m.x[c]()}")
```
## Version 2. Disjunctive Constraints
```
m = ConcreteModel()
# define sets that will be used to index decision variables and constraints
# remember to use initialize keyword
m.comp = Set(initialize=comp_data.keys())
m.req = Set(initialize=prod_req.keys())
# define a set to that includes the excluded pairs
m.pairs = Set(initialize=excl_pairs)
# decision variables
m.x = Var(m.comp, domain=NonNegativeReals, bounds=(0, 1))
# objective function
m.cost = Objective(expr=sum(m.x[c]*comp_data[c]["cost"] for c in m.comp), sense=minimize)
# structural constraints
m.massfraction = Constraint(expr=sum(m.x[c] for c in m.comp)==1)
# composition constraints
m.lb = Constraint(m.req, rule=lambda m, r: sum(m.x[c]*comp_data[c][r] for c in m.comp) >= prod_req[r]["lb"])
m.ub = Constraint(m.req, rule=lambda m, r: sum(m.x[c]*comp_data[c][r] for c in m.comp) <= prod_req[r]["ub"])
# component incompatability constraints
m.disj = Disjunction(m.pairs, rule=lambda m, a, b: [m.x[a] == 0, m.x[b] == 0])
# apply transformations
TransformationFactory('gdp.hull').apply_to(m)
# solve
solver = SolverFactory('cbc')
solver.solve(m)
for c in m.comp:
print(f"{c} = {m.x[c]()}")
```
## Analysis
```
comp_data = {
"A": {"cost": 2.0, "Vit A": 0.5, "Vit B": 0.2},
"B": {"cost": 2.0, "Vit A": 0.4, "Vit B": 0.1},
"C": {"cost": 4.0, "Vit A": 0.3, "Vit B": 0.3},
}
prod_req = {
"Vit A": {"lb": 0.0, "ub": 0.4},
"Vit B": {"lb": 0.2, "ub": 1.0},
}
excl_pairs = [("A", "B")]
```
\begin{align*}
x_A + x_B + x_C & = 1 \\
0.5 x_A + 0.4 x_B + 0.3 x_C & \leq 0.4 \\
0.2 x_A + 0.1 x_B + 0.3 x_C & \geq 0.2 \\
\end{align*}
Solving for x_C
\begin{align*}
x_C & = 1 - x_A - x_B
\end{align*}
Substitution
\begin{align*}
0.2 x_A + 0.1 x_B & \leq 0.1 \\
-0.1 x_A - 0.2 x_B & \geq -0.1 \\
\end{align*}
```
TransformationFactory
```
| github_jupyter |
Up until know, you should now intuitively that linear regression is the least squares line that minimizes the sum of squared residuals. We can check the conditions for linear regression, by looking at **linearity, nearly normal residuals, and constant variability.** We also looking for linear regression of categorical variables, and how to make inference on it.
<!--TEASER_END-->
## Linearity
The conditions for linearity, is whether you have linear relationship between explanatory and response variables. This is to be expected. Non-linearity algorithm is exist, but beyond the scope of this blog. You might want to check my other [blog posts](http://napitupulu-jon.appspot.com/categories/logistic-regression.html).
We can check the relationship based on scatter plot or **residual plots**. Take a look at the graph below.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/147) 01:37*
The first plot you have good linearity, second plot will give you a more curvy plot, and the third one has highest residual. Not obvious linearity in the third plot.We see residual plots below for the the respective plot. So, how do we make it?

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/147) 04:56*
We can make the residual plot by plotting the residuals between observed and predicted points. If we predict based on % HS grad, we will have 14.46% for Rhode Island, and 11.36% for DC. We calculate the residuals and have -4.16% for RI (overestimate) and 5.44% for DC(underestimate). Calculate the rest of the data points, and incorporating the linear regression, we will have residual plots.
The ideal residuals is zero, means plot right at the projected line. This is almost zero chance, but nevertheless we want the residuals to be small, scattered around zero. We want the linearity to capture all the pattern in the data, and let scattered around zero would means that it happen by random chance.
## Nearly normal residuals

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/147) 05:35*
The conditions can also be looked at the residuals plot. We can convert all of the predicted and observed to residuals, plot the histogram and see whether it's normally distributed, centered around zero. By looking at plot above, there's few **unusual observations**, you can also look at the residuals plot, it slightly away from the trend. But this only few observations, so the condition is still met.
## Constant Variability

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/147) 06:25*
The variability for points that scattered around zero in residuals plot is constant. This condition is often called **homoscedasticity**. By looking at the residuals plot, we can see the variability area (shaded by grey color), constant along x-axis. So you see that as explanatory increases, the variability doesn't get affected.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/147) 10:05*
Take a look at this link http://bitly.com/slr_diag. Dr Mine Cetinkaya has been kind enough to provide us with this link to play around with kinds of scatter plot and the resulting residuals. You can see that based on the plot in the upper right corner, we can see whether it's constant variability(Plot 1), nearly normal residuals, centered around zero(Plot 2), and linearity(Plot 3).
# R Squared
After we checked the conditions of linear regression, we want to meassure how fit our prediction to the observed value. This is where R squared comes into play. This will tell how fit our linear model is. This can be calculated by **square the correlation coeffecient**. Mathematically speaking because correlation coeffecient is between 0 to 1 and can be negative/positive, R squared is always positive and ranging from 0 to 1.
R squared will tell us the percentage variability explained by the model, and the rest percentage is what's not explained by the model.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/149) 02:53*
Take a look at the example. We can have R squared by achieving +/- 0.75 in correlation coeffecient.How can we interpret the results in linear regression?
**Example A**. 56.25% of the time, the prediction will be correct. This would means that 56.25% of the predicted value is fall perfectly in the regression line. This is incorrect. The interpretation is not about the predicting correctly.
**Example B**. This is talking about the complementary value of R squared and variability of response variable.Although it's more intuitive, this is also incorrect. The complementary value of R Squared is the variability that can't be explained by the model, and not the other way around.
**Example C**. This explained about the R Squared value and the explanatory variable. This is once again incorrect. We know all about the explanatory variable, and it's meaningless to tell the variability explained by the model.
**Example D**. This is the one that's correct. We know the R-Squared value is the variability of the response variable that's explained by the model.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/149) 04:03*
In determining the value of correlation coeffecient, you can rely the on the computation software to compute the square root of the given R squared, but then you have to look at the plot to see whether it has positive/negative relationship.
# Regression with Categorical Explanatory Variables

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/151) 01:44*
Turns out linear regression can also work with categorical variable as the explanatory. We can have level encoded in the explanatory, and plug in the encoded value, in thise case 0 is east and 1 is west, to the regression formula. An indicator variable is binary categorical variable with two levels(explanatory).

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/151) 02:50*
So for the the intercept alone, we can predict that the model predicts 11.17% average poverty percentage in eastern states. That is if we plug zero, which the category of west. If we plug one, then the slope will be live, and the model predicts 0.38% higher average than easterns states.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/151) 04:03*
Let's take a look at this example. We ahve 4 levels of categorical variables. We can then make a formula with one intercept and multiple slope, and then just try to make binary 1 level that we focus on otherwise zero.So then what's the reference level? The level that we're trying to set as base comparison? You've seen in the image that midwest,west and south has been set for the slope. So all that's left is northeast, as our intercept.So when we're focusing on the northeast, we set all zero for the slope.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/151) 05:40*
So if we want to focus on west, we can eliminate all the pther slopes so we only have west weight parameters, and the resulting will be 11.29. To interpret this we say, the model predicted 11.29% poverty on average of western state.
# Outliers

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/153) 01:03*
In this section we're going to discuss about outliers. Here without the outliers, the regression line can be horizontal flat. But because outliers, the regression line then change to provide linear regression with outliers included. So you see single outlier will affect our regression line greatly.
**Outliers** are points that distance away from cluster of points. There are many types of ourliers. **Leverage points** is points that fall horizontall away from the cluster, so it doesn't affect the slope of the regression, while **influential points** are high level leverage pints, are points that doesn't fall horizontally, and affect the slope of the line.Usually a better way to test this is plot the points with and without the outliers, and see if the slope line have considerable change.If the slope doesn't change, then the points are leverage, but if it does change, then the points are influential. After detecting the outliers, one must be careful whether to include or exclude the outliers.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/153) 04:41*
This is an example of points between light intensity and temperature, by log scale. We can see that with/without outliers, the slope of the line vary greatly. It's depend on what type of finding that you want to focus. If outliers are more interesting thing, then you should include the outliers. But one can not simply join all the trend and outliers. Divide them into two groups, trend and outliers, and make regression line for each of the group. Don't join the model altogether, because it will hurt the model.
So is R squared always get reduced by influential points? No. In some cases, it will affect R squared greatly. So it's not enough to just see R squared and make a decision. You also have make a scatter plot, and detect any influential points that occured. Event one influential points will leverage the slope line.
So leverage points that lies away in horizontal directions, it doesn't change the slope direction. Influential points is the points that change the direction. Determining whether the outlier is leverage/influential, is try to imagine the slope when the outlier is there vs not there.There are have to be a good reason to remove outliers, and if there's level in categorical explanatory that contains few observations but all of it are outliers, this may act as influential points.
# Inference
In this section we want to talk about Hypothesis Testing significance, confidence interval, and additional conditions for inference linear regression.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/155) 01:30*
Here the study is about different in intelligence when twins raised by one foster and one biological. We can see the from the plot we have positive strong relationship, with correlation coeffecient 0.882

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/155) 02:17*
In this example, we can say " 78% of foster IQ can be explained by biological IQ". For each 10 points increase in biological twins IQ, we would expect the foster twins IQ to be increase as well by average 9 points. The explanatory is bioIQ and response variable is fosterIQ. twins foster IQ with higher than average are predicted to have biological twins IQ to be higher than average as well.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/155) 03:22*
For Hypothesis Testing, we want to test that the data provide convincing evidence that biological twin IQ is significant predictor for IQ foster twin.That is hypothesis testing to test whether or not explanatory variable affected the response variable. The skeptical would say no, and the slope is zero, alternative stated otherwise. Recall that B1 is the slope, whereas B0 is null hypothesis.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/155) 04:13*
Recall that the formulation of t-statistic, we have the calculation for z critical and degree of freedom. So we can plug the formula with slope as point estimate.So why n-2? Remember that for each parameter we calculated, we lose a degree of freedom. Even if you just focus on the slope, it will end up for intercept as well. So we lose two degree of freedom, hence n-2.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/155) 06:52*
Calculating standard error is tedious and very error prone. So here we have already computed table score. Given this we can calculate t-score and get 9.36. As we know this is very huge, and rounding to 4 digits we still get approximate zero. Remember! Even if the computation results to zero, it doesn't mean it equals zero, because p-value can never be zero, it simply just too small even for rounding 4 digits. We can also calculate the intercep and we have almost 1 standard deviation. Remember that hypothesis testing is about two-sided value, so 16% x 2 equals 0.33 like in the computation. So indeed the data provide convincing evidence that biological twin IQ is a significant predictor for foster twin IQ.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/155) 08:54*
```
qt(0.025, df = 25)
```
For confidence interval, we can subtitute the slope and using t-critical for z-score. 95% will results in a little above 1.96, to count t-star for wider interval. Using qt in R we can get our t-critical, which should always be positive. Then the point estimate +/- t_score times standard error, we get the confidence interval. How do we interpret CI? We can say, "We are 95% confident that for each additional point on biological twin IQ, foster twin IQ are expected to be higher by 0.7 to 1.1 points.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/155) 10:43*
Null value is often zero. Since here we have to catch any relationship between explanatory and response variable. That's also why the null value is zero. On computation, regression output b1,SEb1, two-tailed p-value given null value zero, for the slope. We rarely talk about intercept, because what we often want to infer is the relationship between both variables, which is done by observing the slope.
So conditions must be checked to make inference for linear regression as well. We want's to have random sample (if not often unreliable), less than 10% population, and else to make sure that observations are independent of one another. And if you're already have population data, it's useless to make inference, we only work of what based on the sample data.
# Variability Partitioning

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/155) 00:54*
Recall that R squared is about variability of y explained by x. But could it be makes to other way around? Why not unexplained variability dictates y? This requires us to use **ANOVA**.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/155) 01:57*
Recall that sum of squares total can be calculated by the difference between actual output and the average outputs. So we can sum of squares and have the total difference. We also can calculate sum of squares residuals, with the difference between predicted and the actual output. So we can take explained variability by complements of the residuals.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/155) 02:34*
For the degree of freedom, we just observing the slope (one parameter) and have 27 samples, so we have 26 total. And because 1 predictor, we get regression df 1. Residuals is just the complement.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/155) 03:19*
Means of squares can be get by SS/df for each regression and residuals. Recall that F-statistic is ratio of explained to unexplained variability. Which means that as we get higher F score, we get a good estimate.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/155) 04:02*
For hypothesis testing, the p-value is approximate zero. Hence we reject the null hypothesis and conclude the data provide convincing evidence that the slope is significantly different than zero(we said significantly, if it pass the significance level), i.e explanatory variable is a significant predictor of the response variable.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/155) 05:03*
R squared is the proportion of variability y explained by the model. Larger value will yield larger proportion of y variability y explained by the model. Small will means the evidence is not convincing to be significant predictor. Two ways to calculate R squared, square the correlation coeffecient, or proportion of explained ratio to total variability, as calculated by ANOVA. We can validate R squared using both methods.

*Screenshot taken from [Coursera](https://class.coursera.org/statistics-003/lecture/155) 05:46*
Using both models we achieved roughly 78%. So 78% of twin foster IQ (response variable) can be explained by the model, in other words, twin biological IQ (explanatory variable).We want to keep R squared to be approach 1, because that is the ideal model, which can predict all the y output.
In summary, we can use t-test and the resulting p-value to determine whether the indicator variable, the predictor, is sifnificant predictor or not. Regression output computed by the software always yields two-sided p-value given null hypothesis zero. We lose two degree freedom to account for intercept and slope parameters.
Why we rarely have HT for intercept? Because the intercept point may outside the realm of data and usually an extrapolation. In other words, **extrapolation** means predicting for any given value of x outside the realm of data.
> **REFERENCES**:
> Dr. Mine Çetinkaya-Rundel, [Cousera](https://class.coursera.org/statistics-003/lecture)
| github_jupyter |
The first notebook to test FSIC idea. Likely to be deprecated in the near future. Created on 16 June 2016.
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import fsic.util as util
import fsic.data as data
import fsic.kernel as kernel
import fsic.indtest as it
import fsic.glo as glo
import scipy.stats as stats
# font options
font = {
#'family' : 'normal',
#'weight' : 'bold',
'size' : 15
}
plt.rc('font', **font)
plt.rc('lines', linewidth=2)
def get_quad_psfunc():
"""
Return a PairedSource to generate y = x^2 + Gaussian noise.
"""
px = lambda n: np.random.rand(n, 1)*8 - 4
f = lambda x: 0.2*x**2 + np.random.randn(x.shape[0], 1)
return data.PSFunc(f, px)
def kl_kgauss_median(pdata, med_factor=1):
"""
Get two Gaussian kernels constructed with the median heuristic.
"""
xtr, ytr = pdata.xy()
dx = xtr.shape[1]
dy = ytr.shape[1]
medx2 = util.meddistance(xtr, subsample=1000)**2
medy2 = util.meddistance(ytr, subsample=1000)**2
k = kernel.KGauss(medx2*med_factor)
l = kernel.KGauss(medy2*med_factor)
return k, l
# paired source
alpha = 0.01
n = 1000
dx = 100
dy = 100
seed = 393
#ps = data.PSIndSameGauss(dx, dy)
#ps = get_quad_psfunc()
ps = data.PS2DSinFreq(freq=5)
#ps = data.PSIndUnif(xlb=[0, 3], xub=[1, 10], ylb=[-5, 5], yub=[8, 10])
pdata = ps.sample(n, seed=seed)
X, Y = pdata.xy()
#tr, te = pdata.split_tr_te(tr_proportion=0.5, seed=10)
# data and parameters
#xtr, ytr = tr.xy()
#xte, yte = te.xy()
k, l = kl_kgauss_median(pdata)
# number of test locations
J = 5
dx = pdata.dx()
dy = pdata.dy()
#V = np.random.randn(J, dx)
#W = np.random.randn(J, dy)
V, W = it.GaussNFSIC.init_locs_joint_randn(pdata, J, seed=seed+2)
#V, W = it.GaussNFSIC.init_locs_2randn(pdata, J, seed=seed+2)
# perform test
nfsic = it.NFSIC(k, l, V, W, alpha=alpha, reg=1e-5)
nfsic_result = nfsic.perform_test(pdata)
nfsic_result
```
## Null distribution
```
# permute and compute the statistic many times
n_permute = 1000
with util.ContextTimer() as t1:
test_stats = nfsic.list_permute(X, Y, k, l, V, W, n_permute=n_permute, reg='auto')
#test_stats = nfsic._list_permute_naive(X, Y, k, l, V, W, n_permute=n_permute)
sim_pval = np.mean(test_stats > nfsic_result['test_stat'])
asymp_pval = nfsic_result['pvalue']
print 'p-value by permutations: %.3g'%sim_pval
print 'p-value by asymptotic chi-square: %.3g'%asymp_pval
print 'permutation took: %.3f s'%t1.secs
dom = np.linspace(1e-1, np.max(test_stats), 500)
chi2den = stats.chi2.pdf(dom, df=J)
nc2den = stats.ncx2.pdf(dom, df=J, nc=np.mean(test_stats))
# histogram
plt.figure(figsize=(10,4))
plt.hist(test_stats, alpha=0.5, bins=20, normed=True, label='Stats under $H_0$')
plt.plot(dom, chi2den, label=r'$\chi^2(J)$')
#plt.plot(dom, nc2den, label=r'$\chi^2(J, \lambda)$')
plt.legend()
plt.title('FSIC. $J=%d$'%J)
# plot empirical cdf
sorted_h0_stats = np.sort(test_stats)
normed_ranks = np.arange(len(sorted_h0_stats))/float(len(sorted_h0_stats))
plt.plot(sorted_h0_stats, normed_ranks, label='Simulated ECDF')
cdfs = stats.chi2.cdf(dom, df=J)
plt.plot(dom, cdfs, label='$\chi^2(J)$ CDF')
plt.legend()
# diff CDF
diff_cdf = normed_ranks - stats.chi2.cdf(sorted_h0_stats, df=J)
plt.plot(sorted_h0_stats, diff_cdf)
plt.title('Diff in the CDFs')
plt.grid(True)
```
## Permutation test
```
#nfsic_perm = it.NFSIC(k, l, V, W, alpha=alpha, reg=1e-6, n_permute=500)
st = nfsic.compute_stat(pdata)
pval = np.mean(test_stats > st)
print 'stat: %.3f'%st
print 'p-value: %.3f'%pval
```
## Test power
Simulate from a toy example and try to compute the test power.
```
reps = 1000
n = 1000
J = 10
alpha = 0.05
# None = use aymptotics
n_permute = None
#n_permute = 200
ps = data.PSIndSameGauss(dx=20, dy=20)
k, l = kl_kgauss_median(ps.sample(1000, seed=2198), med_factor=1.0)
with util.NumpySeedContext(seed=23):
V = np.random.randn(J, ps.dx())
W = np.random.randn(J, ps.dy())
test_results = []
for r in range(reps):
pdata = ps.sample(n, seed=r)
pdata2 = ps.sample(300, seed=r+66)
#with util.NumpySeedContext(seed=23):
# V = np.random.randn(J, ps.dx())
# W = np.random.randn(J, ps.dy())
V, W = it.GaussNFSIC.init_locs_joint_subset(pdata2, J, seed=r+1)
#V, W = it.GaussNFSIC.init_locs_2randn(pdata, J, seed=r+3)
#k, l = kl_kgauss_median(pdata, med_factor=1.0)
nfsic = it.NFSIC(k, l, V, W, alpha=alpha, reg='auto', n_permute=n_permute, seed=89)
result = nfsic.perform_test(pdata)
test_results.append(result)
rejs = [r['h0_rejected'] for r in test_results]
rep_stats = [r['test_stat'] for r in test_results]
thresh = stats.chi2.isf(alpha, df=J)
power = np.mean(rejs)
print 'power: %g'%power
# histogram
dom = np.linspace(max(1e-1, np.min(rep_stats)), np.max(rep_stats), 600)
chi2_den = stats.chi2.pdf(dom, df=J)
plt.figure(figsize=(10, 5))
plt.hist(rep_stats, bins=20, alpha=0.5, label='Repeated trials', normed=True)
plt.plot(dom, chi2_den, '-', label=r'$\chi^2(%d)$'%J)
plt.legend()
```
## When two locations are very close
```
# 2d data
#ps = data.PSIndSameGauss(dx=1, dy=1)
ps = data.PS2DSinFreq(freq=1)
pdata = ps.sample(n=700, seed=8)
X, Y = pdata.xy()
k, l = kl_kgauss_median(pdata, med_factor=0.85)
reg = 1e-6
W = np.array([[-1.7], [-1.7]])
v0_cand = np.hstack( (np.linspace(-4, 4, 500), [0, -1] ))
v0_cand.sort()
nfsics = np.zeros(len(v0_cand))
for i, v0 in enumerate(v0_cand):
V = np.array([[v0], [-1]])
nfsic = it.NFSIC(k, l, V, W, alpha=0.05, reg=reg)
nfsics[i] = nfsic.compute_stat(pdata)
# plot
#plt.figure(figsize=(7, 2))
ax1 = plt.subplot(2, 1, 1)
plt.locator_params(nbins=5)
plt.plot(X[:, 0], Y[:, 0], 'k.', label='Sample', alpha=0.8)
plt.plot(v0_cand, np.ones(len(v0_cand))*W[0], 'g-', linewidth=3, label='$\mathbf{t}_2$ trajectory')
plt.plot(V[1], W[1], 'r*', markersize=23, label=r'$\mathbf{t}_1$')
plt.xlabel('$x$')
plt.ylabel('$y$')
plt.xlim([-np.pi, np.pi])
plt.ylim([-np.pi, np.pi])
plt.legend(numpoints=1, ncol=3, fontsize=16,
# bbox_to_anchor=(1.02, 1.9)
bbox_to_anchor=(1.02, 1.5)
)
plt.setp(ax1.get_xticklabels(), visible=False)
# values
plt.subplot(2, 1, 2, sharex=ax1)
plt.locator_params(nbins=5)
plt.plot(v0_cand, nfsics, 'b-', label='$\hat{\lambda}_n(\mathbf{t}_1, \mathbf{t}_2)$')
#plt.title('V: [[v0], [%.1f]], W: %s'%(V[1], W))
plt.xlabel(r'$\mathbf{t}_2$')
plt.ylabel('$\hat{\lambda}_n(\mathbf{t}_1, \mathbf{t}_2)$')
plt.ylim([np.min(nfsics)-10, np.max(nfsics)+10])
plt.xlim([-np.pi, np.pi])
#plt.legend(numpoints=1, ncol=3, loc='lower right')
#plt.gca().get_yaxis().set_visible(False)
plt.savefig('redundant_locs.pdf', bbox_inches='tight')
```
--------------
## Test power when J is high
```
# paired source
alpha = 0.01
n = 1000
dx = 100
dy = 100
seed = 393
#ps = data.PSIndSameGauss(dx, dy)
#ps = get_quad_psfunc()
ps = data.PS2DSinFreq(freq=2)
#ps = data.PSIndUnif(xlb=[0, 3], xub=[1, 10], ylb=[-5, 5], yub=[8, 10])
pdata = ps.sample(n, seed=seed)
X, Y = pdata.xy()
k, l = kl_kgauss_median(pdata)
tr, te = pdata.split_tr_te(tr_proportion=0.5, seed=10)
def test_power(ps, nte, J, reps):
rejs = np.zeros(reps)
for r in range(reps):
te = ps.sample(nte, seed=r+9827)
tr = ps.sample(nte, seed=r+27)
V, W = it.GaussNFSIC.init_locs_2randn(tr, J, seed=r+2)
#V, W = it.GaussNFSIC.init_locs_joint_randn(tr, J, seed=r+2)
#V, W = it.GaussNFSIC.init_locs_marginals_subset(tr, J, seed=r+2)
#V, W = it.GaussNFSIC.init_locs_joint_subset(tr, J, seed=r+2)
nfsic = it.NFSIC(k, l, V, W, alpha=alpha, reg='auto')
try:
test_result = nfsic.perform_test(te)
rejs[r] = test_result['h0_rejected']
except:
rejs[r] = False
return np.mean(rejs)
nte = 800
reps = 500
Js = range(1, 600+3, 50) + [10, 20, 30, 40]
Js = np.sort(np.array(Js))
#Js = np.logspace(0, 2.6, 10).astype(np.int64)
Js_pow = np.zeros(len(Js))
test_results = np.zeros(len(Js), dtype=np.object)
for i, J in enumerate(Js):
tpow = test_power(ps, nte, J, reps)
Js_pow[i] = tpow
#plt.semilogx(Js, Js_pow, 'bo-')
plt.plot(Js, Js_pow, 'bo-')
plt.xlim([np.min(Js), np.max(Js)])
plt.ylim([np.min(Js_pow), 1])
plt.xlabel('J')
plt.ylabel('Test power')
plt.grid()
fname = 'pow_vs_J.pdf'
plt.savefig(fname, bbox_inches='tight')
```
| github_jupyter |
```
import theme
theme.load_style()
```
# Lesson 17: Thermal Convection Element
<a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Creative Commons License" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a>
This lecture by Tim Fuller is licensed under the
<a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Creative Commons Attribution 4.0 International License</a>. All code examples are also licensed under the [MIT license](http://opensource.org/licenses/MIT).
# Introduction
Using simple axial bar elements and transformation equations we were able to solve for the deformation of a 3D space truss under prescribed load/support conditions.
Using the same connectivity table we were able to solve for the temperature distribution assuming a prescribed temperature at the support and loaded nodes, and neglecting the effect of convection to the surrounds.
In reality, the heat loss to the surrounding air would be significant and should be considered in the design process.
Thermal expansion and self weight may significantly affect the structural response.
<div class='msg'>
How can we capture these important thermal and mechanical mechanisms without the cost and complexity of a 3D solid model?
</div>
# Axial Bar with Surface Convection
For the bar shown, two heat transfer mechanisms occur at each point along the bar
<table id='mytable'>
<tr><td>
<img src='./Images/thermal-convection.d/therm-conv-1.png' style='width:80%'/>
</td>
<td>
<img src='./Images/thermal-convection.d/therm-conv-2.png' style='width:50%'/>
</td>
</tr>
</table>
**Axial conduction**, governed by Fourier’s law; the area‐specific heat flux trhough the bar is equal to the negative product of the thermal conductivity and the temperature gradient.
$$
q_{\text{cond}}=-\kappa\frac{dT}{dx}
$$
**Surface convection**: The area specific heat to the surroundings is equal to the product of the heat transfer coefficient and the difference between the surface temperature and free stream temperature.
$$
q_{\text{conv}} = h\left(T_x - T_{\infty}\right)
$$
For free convection to air, typical values for are 2 - 25 w/m$^2$ K and can be estimated based on model geometry and gas properties.
## Convection Coefficient
<table id='mytable'>
<tr>
<td><img src='./Images/thermal-convection.d/buoyancy-flow.png' style='width:70%'/></td>
<td><img src='./Images/thermal-convection.d/heat-trans-coef.png' style='width:70%'/></td>
</tr>
</table>
Free convection over a long horizontal cylinder is a buoyancy-driven flow and has been studied extensively. Churchill and Chu (1975) present the following correlation:
Average heat transfer coefficient
$$
\overline{h} = \frac{\overline{Nu_D}k}{D}
$$
Nusselt number
$$
\overline{Nu_D} = \left(.6+
\frac{.387Ra_D^{1/6}}{\left(1+\left(.599/Pr\right)^{9/16}\right)^{8/27}}\right)^2
$$
Rayleigh number:
$$
Ra_D = \frac{g\beta\left(T_s-T_{\infty}\right)D^3}{\nu\alpha}
$$
Ideal gas compressibility:
$$
\beta = \frac{1}{\left(T_s-T_{\infty}\right)/2}
$$
Evaluating this with properties for air at 300K, and allowing that the surface temperature may vary with position along the bar, we obtain the following:
$$
h(x) = \frac{.026}{d}\left(19.8\left(\lvert T(x)-T_{\infty}\rvert \right)^{1/6} \left(\frac{d^3}{T(x)+T_{\infty}}\right)^{1/6}+.6\right)^2
$$
<div class='msg'>
If large surface temperature variations occur over the structure, this correlation could be used to develop a more accurate model by allowing $h(x)$ to vary with position over an element, otherwise some constant $h_e$ could be computed for each element.
</div>
## Strong Form
### Balance Law
At each point, the axial conduction is equal to the conduction out plus the convection to the surroundings.
For some differential length:
$$
q(x)A(x) - \beta h\left(T(x) - T_\infty(x)\right)\Delta x -
q(x+\Delta x)A\left(x+\Delta x\right)=0
$$
Taking the limit as $\Delta x\rightarrow 0$, we obtain
$$
\beta h\left(T(x) - T_{\infty}\right)+A(x)\frac{dq}{dx}=0
$$
### Flux-Potential Relationship
For heat conduction, the flux-potential relationship is modeled by Fourier's law:
$$
T(x) - T(x+\Delta x) = q(x)\frac{\Delta x}{k_{th}A}
$$
or, in differential form
$$
q(x) = -k_{th}\frac{dT}{dx}
$$
Substituting the flux potential relationship into the balance law we obtain the **strong form** for axial conduction in a bar with surface convection
$$
k_{th}A(x)\frac{d^2T}{dx^2}-\beta h T\left(T(x)-T_{\infty}\right)=0, \quad 0 \leq x \leq L
$$
with Dirichlet boundary conditions
$$
T(0) = T_h, \quad T(L) = T_c
$$
## Weak Form
Assuming the problem data (what are all the problem data?) are constant, the strong form is
$$
kAT''-\beta h\left(T-T_{\infty}\right)=0, \quad 0 \leq x \leq L
$$
$$
T(0) = T_h, \quad T(L) = T_c
$$
Following the 3 step procedure
1. Multiply by a weight function and integrate over the domain
$$
\int_0^L w\left(
kAT''-\beta h\left(T-T_{\infty}\right)\right)dx = 0 \quad \forall w
$$
2. Integrate by parts
$$
-\int_0^L w'kAT'-\beta h\left(T-T_{\infty}\right)dx
+kAwT'\Bigg|_0^L
$$
3. Enforce essential boundary conditions
$$
kAwT'\Bigg|_0^L=kAw(L)T'(L)-kAw(0)T'(0)=0, \quad \forall w
$$
So, the weak form for axial conduction in a bar with surface convection and Dirichlet boundaries is
$$
\int_0^L w'kAT'dx +
\int_0^L\beta hTdx -
\int_0^L\beta hT_{\infty}dx=0, \quad
\forall w, \ w(0) = w(L) = 0
$$
# Approximate Solution
Express the integral over the domain as a sum of integrals over elements
$$ \begin{multline}
\sum_e \left(
\int_{x_0^{(e)}}^{x_L^{(e)}} {w^{(e)}}' k^{(e)}A^{(e)} {T^{(e)}}' dx +
\int_{x_0^{(e)}}^{x_L^{(e)}} w^{(e)} \beta^{(e)} h^{(e)} T^{(e)} dx - \\
\int_{x_0^{(e)}}^{x_L^{(e)}} w^{(e)} \beta^{(e)} h^{(e)} T_{\infty} dx
\right)=0, \quad
\forall w, \ w(0) = w(L) = 0
\end{multline} $$
Introduce an approximation for the "trial" solution and weight "test" function
$$
T^e = N_i^e T_i^e, \quad
\frac{dT^e}{dx} = \frac{dN_i^e}{dx}T_i^e = B_i^eT_i^e, \quad
w^e = w_i^eN_i^e, \quad
\frac{dw^e}{dx} = w_i^e\frac{dN_i^e}{dx}
$$
Substituting in to the weak form gives
$$
\begin{multline}
\sum_e\left(\int_{x_0^{(e)}}^{x_L^{(e)}} w_i^{(e)}\frac{dN_i^{(e)}}{dx}k^{(e)}A^{(e)}\frac{dN_j^{(e)}}{dx} T_j^{(e)} +
\int_{x_0^{(e)}}^{x_L^{(e)}} w_i^{(e)}N_i^{(e)}\beta^{(e)} h^{(e)} N_j^{(e)}T_j^{(e)}dx \right. \\ \left. -
\int_{x_0^{(e)}}^{x_L^{(e)}} w_i^{(e)}N_i^{(e)} \beta^{(e)} h^{(e)} T_{\infty}dx\right)=0, \quad
\forall w, \ w_1 = w_n = 0
\end{multline}
$$
We factor out the $w_i^{(e)}$ of each term and rearrange
$$
\begin{multline}
\sum_ew_i^{(e)}\left[
\left(
\int_{x_0^{(e)}}^{x_L^{(e)}} k^{(e)}A^{(e)}\frac{dN_i^{(e)}}{dx}\frac{dN_j^{(e)}}{dx} +
\int_{x_0^{(e)}}^{x_L^{(e)}} \beta^{(e)} h^{(e)} N_i^{(e)} N_j^{(e)}dx\right)T_j^{(e)} \right.\\
\left.-
\int_{x_0^{(e)}}^{x_L^{(e)}} \beta^{(e)} h^{(e)} N_i^{(e)} T_{\infty}dx\right] = 0, \quad
\forall w, \ w_1 = w_n = 0
\end{multline}
$$
Writing this in a more compact form
$$
\sum_e w_i^{(e)} \left(k_{ij}^{(e)}T_j^{(e)} - f_i^{(e)}\right) = 0
$$
where the stiffness $k_{ij}^{(e)}$ is
$$
k_{ij}^{(e)}=\int_{x_0^{(e)}}^{x_L^{(e)}} k^{(e)}A^{(e)}\frac{dN_i^{(e)}}{dx}\frac{dN_j^{(e)}}{dx} +
\int_{x_0^{(e)}}^{x_L^{(e)}} \beta^{(e)} h^{(e)} N_i^{(e)} N_j^{(e)}dx
$$
and the **element external flux array**, (equivalent to the force array for our structural problem), which is a sum of boundary and body forces
$$
f_i^{(e)} = f_{\Gamma i}^{(e)} + f_{\Omega i}^{(e)} =
\int_{x_0^{(e)}}^{x_L^{(e)}} \beta^{(e)} h^{(e)} N_i^{(e)} T_{\infty}dx
$$
For this problem the boundary source array is zero, as we have specified Dirichlet boundaries at each end of the domain. This is often done in discussion of FE when the focus is the development of an element stiffness matrix. We have not yet specified the function $T_{\infty}$ that determines how the free stream temperature varies along the bar.
# Thermal Stresses
From the temperature distribution in the truss members, we can compute the thermal expansion strains to see how these affect the stresses and displacements in the structure.
**Strong Form**
$$
\frac{d}{dx}\left(EA(x)\frac{du}{dx}-E\alpha A(x) T(x)\right) = 0
$$
**Weak Form**
$$
\int_0^Lw'(x) EA u'(x) dx + \int_0^L w(x) EA\alpha T'(x) dx = 0, \quad
\forall w(x), w(x)_{x=0,L}=0
$$
**Finite Element Equations**
$$
k_{ij}^{(e)}u_j^{(e)} - f_i^{(e)} = 0
$$
where
$$
k_{ij}^{(e)} = \int_{x_0^{(e)}}^{x_L^{(e)}}
A^{(e)}E^{(e)}\frac{dN_i}{dx}\frac{dN_j}{dx} dx
$$
$$
f_i^{(e)} = -\int_{x_0^{(e)}}^{x_L^{(e)}}N_i^{(e)}(x)EA\alpha\frac{dT^{(e)}}{dx}dx
$$
The thermal stresses appear in the form of a body force, adding a force that counteracts that normally required to achieve some amount of thermal expansion.
<div class='msg'>
Thus, we see that thermal effects can be readily added to an existing model, through the addition of force terms. This approach can be used to model all types of zero‐stress deformation mechanics such as curing or drying of composites, water absorption, etc.
</div>
# Exercises
## Exercise 1 [60]
Modified from Fish & Belytschko, Problem 5.1
For this problem do NOT use the direct stiffness method.
Consider a heat conduction problem in the domain [0, 20]m. The bar has a unit cross section, constant thermal conductivity $k=5$W/$^{\text{o}}C$$\cdot$m and a uniform heat source $s=100$ W/m. The boundary conditions are $T(0)=0^{\text{o}}C$ and $\overline{q}(20)=0$W/m$^2$. Solve the problem with two equal linear elements. Plot the finite element solution $T_N(x)$ and $dT_N(x)/dx$ and compare to the exact solution which is given by $T(x)=-10x^2+400x$.
## Exercise 2
<img src='./Images/thermal-convection.d/ex-2.png' style='width:70%'/>
For the bar shown,
a) State the strong form representing heat flow and solve analytically. Give the symbolic solution $T(x)$ and total heat flux $Q(x)$ along the length of the bar.
b) Construct the element body source arrays, the boundary force arrays, and assemble the global external force array.
c) Construct the element stiffness matrices and assemble the global stiffness matrix.
d) Solve for the temperature distribution $T(x)$. Generate a plot of the global approximate solution $T_N(x)$ along the length of the bar overlaid with the analytic solution.
| github_jupyter |
# Logistic Regression with a Neural Network mindset
Welcome to your first (required) programming assignment! You will build a logistic regression classifier to recognize cats. This assignment will step you through how to do this with a Neural Network mindset, and so will also hone your intuitions about deep learning.
**Instructions:**
- Do not use loops (for/while) in your code, unless the instructions explicitly ask you to do so.
**You will learn to:**
- Build the general architecture of a learning algorithm, including:
- Initializing parameters
- Calculating the cost function and its gradient
- Using an optimization algorithm (gradient descent)
- Gather all three functions above into a main model function, in the right order.
## 1 - Packages ##
First, let's run the cell below to import all the packages that you will need during this assignment.
- [numpy](www.numpy.org) is the fundamental package for scientific computing with Python.
- [h5py](http://www.h5py.org) is a common package to interact with a dataset that is stored on an H5 file.
- [matplotlib](http://matplotlib.org) is a famous library to plot graphs in Python.
- [PIL](http://www.pythonware.com/products/pil/) and [scipy](https://www.scipy.org/) are used here to test your model with your own picture at the end.
```
import numpy as np
import matplotlib.pyplot as plt
import h5py
import scipy
from PIL import Image
from scipy import ndimage
from lr_utils import load_dataset
%matplotlib inline
```
## 2 - Overview of the Problem set ##
**Problem Statement**: You are given a dataset ("data.h5") containing:
- a training set of m_train images labeled as cat (y=1) or non-cat (y=0)
- a test set of m_test images labeled as cat or non-cat
- each image is of shape (num_px, num_px, 3) where 3 is for the 3 channels (RGB). Thus, each image is square (height = num_px) and (width = num_px).
You will build a simple image-recognition algorithm that can correctly classify pictures as cat or non-cat.
Let's get more familiar with the dataset. Load the data by running the following code.
```
# Loading the data (cat/non-cat)
train_set_x_orig, train_set_y, test_set_x_orig, test_set_y, classes = load_dataset()
```
We added "_orig" at the end of image datasets (train and test) because we are going to preprocess them. After preprocessing, we will end up with train_set_x and test_set_x (the labels train_set_y and test_set_y don't need any preprocessing).
Each line of your train_set_x_orig and test_set_x_orig is an array representing an image. You can visualize an example by running the following code. Feel free also to change the `index` value and re-run to see other images.
```
# Example of a picture
index = 25
plt.imshow(train_set_x_orig[index])
print ("y = " + str(train_set_y[:, index]) + ", it's a '" + classes[np.squeeze(train_set_y[:, index])].decode("utf-8") + "' picture.")
```
Many software bugs in deep learning come from having matrix/vector dimensions that don't fit. If you can keep your matrix/vector dimensions straight you will go a long way toward eliminating many bugs.
**Exercise:** Find the values for:
- m_train (number of training examples)
- m_test (number of test examples)
- num_px (= height = width of a training image)
Remember that `train_set_x_orig` is a numpy-array of shape (m_train, num_px, num_px, 3). For instance, you can access `m_train` by writing `train_set_x_orig.shape[0]`.
```
### START CODE HERE ### (≈ 3 lines of code)
m_train = train_set_x_orig.shape[0]
m_test = test_set_x_orig.shape[0]
num_px = train_set_x_orig.shape[1]
### END CODE HERE ###
print ("Number of training examples: m_train = " + str(m_train))
print ("Number of testing examples: m_test = " + str(m_test))
print ("Height/Width of each image: num_px = " + str(num_px))
print ("Each image is of size: (" + str(num_px) + ", " + str(num_px) + ", 3)")
print ("train_set_x shape: " + str(train_set_x_orig.shape))
print ("train_set_y shape: " + str(train_set_y.shape))
print ("test_set_x shape: " + str(test_set_x_orig.shape))
print ("test_set_y shape: " + str(test_set_y.shape))
```
**Expected Output for m_train, m_test and num_px**:
<table style="width:15%">
<tr>
<td>**m_train**</td>
<td> 209 </td>
</tr>
<tr>
<td>**m_test**</td>
<td> 50 </td>
</tr>
<tr>
<td>**num_px**</td>
<td> 64 </td>
</tr>
</table>
For convenience, you should now reshape images of shape (num_px, num_px, 3) in a numpy-array of shape (num_px $*$ num_px $*$ 3, 1). After this, our training (and test) dataset is a numpy-array where each column represents a flattened image. There should be m_train (respectively m_test) columns.
**Exercise:** Reshape the training and test data sets so that images of size (num_px, num_px, 3) are flattened into single vectors of shape (num\_px $*$ num\_px $*$ 3, 1).
A trick when you want to flatten a matrix X of shape (a,b,c,d) to a matrix X_flatten of shape (b$*$c$*$d, a) is to use:
```python
X_flatten = X.reshape(X.shape[0], -1).T # X.T is the transpose of X
```
```
# Reshape the training and test examples
### START CODE HERE ### (≈ 2 lines of code)
train_set_x_flatten = train_set_x_orig.reshape(train_set_x_orig.shape[0], -1).T
test_set_x_flatten = test_set_x_orig.reshape(test_set_x_orig.shape[0], -1).T
### END CODE HERE ###
print ("train_set_x_flatten shape: " + str(train_set_x_flatten.shape))
print ("train_set_y shape: " + str(train_set_y.shape))
print ("test_set_x_flatten shape: " + str(test_set_x_flatten.shape))
print ("test_set_y shape: " + str(test_set_y.shape))
print ("sanity check after reshaping: " + str(train_set_x_flatten[0:5,0]))
```
**Expected Output**:
<table style="width:35%">
<tr>
<td>**train_set_x_flatten shape**</td>
<td> (12288, 209)</td>
</tr>
<tr>
<td>**train_set_y shape**</td>
<td>(1, 209)</td>
</tr>
<tr>
<td>**test_set_x_flatten shape**</td>
<td>(12288, 50)</td>
</tr>
<tr>
<td>**test_set_y shape**</td>
<td>(1, 50)</td>
</tr>
<tr>
<td>**sanity check after reshaping**</td>
<td>[17 31 56 22 33]</td>
</tr>
</table>
To represent color images, the red, green and blue channels (RGB) must be specified for each pixel, and so the pixel value is actually a vector of three numbers ranging from 0 to 255.
One common preprocessing step in machine learning is to center and standardize your dataset, meaning that you substract the mean of the whole numpy array from each example, and then divide each example by the standard deviation of the whole numpy array. But for picture datasets, it is simpler and more convenient and works almost as well to just divide every row of the dataset by 255 (the maximum value of a pixel channel).
<!-- During the training of your model, you're going to multiply weights and add biases to some initial inputs in order to observe neuron activations. Then you backpropogate with the gradients to train the model. But, it is extremely important for each feature to have a similar range such that our gradients don't explode. You will see that more in detail later in the lectures. !-->
Let's standardize our dataset.
```
train_set_x = train_set_x_flatten/255.
test_set_x = test_set_x_flatten/255.
```
<font color='blue'>
**What you need to remember:**
Common steps for pre-processing a new dataset are:
- Figure out the dimensions and shapes of the problem (m_train, m_test, num_px, ...)
- Reshape the datasets such that each example is now a vector of size (num_px \* num_px \* 3, 1)
- "Standardize" the data
## 3 - General Architecture of the learning algorithm ##
It's time to design a simple algorithm to distinguish cat images from non-cat images.
You will build a Logistic Regression, using a Neural Network mindset. The following Figure explains why **Logistic Regression is actually a very simple Neural Network!**
<img src="images/LogReg_kiank.png" style="width:650px;height:400px;">
**Mathematical expression of the algorithm**:
For one example $x^{(i)}$:
$$z^{(i)} = w^T x^{(i)} + b \tag{1}$$
$$\hat{y}^{(i)} = a^{(i)} = sigmoid(z^{(i)})\tag{2}$$
$$ \mathcal{L}(a^{(i)}, y^{(i)}) = - y^{(i)} \log(a^{(i)}) - (1-y^{(i)} ) \log(1-a^{(i)})\tag{3}$$
The cost is then computed by summing over all training examples:
$$ J = \frac{1}{m} \sum_{i=1}^m \mathcal{L}(a^{(i)}, y^{(i)})\tag{6}$$
**Key steps**:
In this exercise, you will carry out the following steps:
- Initialize the parameters of the model
- Learn the parameters for the model by minimizing the cost
- Use the learned parameters to make predictions (on the test set)
- Analyse the results and conclude
## 4 - Building the parts of our algorithm ##
The main steps for building a Neural Network are:
1. Define the model structure (such as number of input features)
2. Initialize the model's parameters
3. Loop:
- Calculate current loss (forward propagation)
- Calculate current gradient (backward propagation)
- Update parameters (gradient descent)
You often build 1-3 separately and integrate them into one function we call `model()`.
### 4.1 - Helper functions
**Exercise**: Using your code from "Python Basics", implement `sigmoid()`. As you've seen in the figure above, you need to compute $sigmoid( w^T x + b) = \frac{1}{1 + e^{-(w^T x + b)}}$ to make predictions. Use np.exp().
```
# GRADED FUNCTION: sigmoid
def sigmoid(z):
"""
Compute the sigmoid of z
Arguments:
z -- A scalar or numpy array of any size.
Return:
s -- sigmoid(z)
"""
### START CODE HERE ### (≈ 1 line of code)
s = 1./(1+np.exp(-z))
### END CODE HERE ###
return s
print ("sigmoid([0, 2]) = " + str(sigmoid(np.array([0,2]))))
```
**Expected Output**:
<table>
<tr>
<td>**sigmoid([0, 2])**</td>
<td> [ 0.5 0.88079708]</td>
</tr>
</table>
### 4.2 - Initializing parameters
**Exercise:** Implement parameter initialization in the cell below. You have to initialize w as a vector of zeros. If you don't know what numpy function to use, look up np.zeros() in the Numpy library's documentation.
```
# GRADED FUNCTION: initialize_with_zeros
def initialize_with_zeros(dim):
"""
This function creates a vector of zeros of shape (dim, 1) for w and initializes b to 0.
Argument:
dim -- size of the w vector we want (or number of parameters in this case)
Returns:
w -- initialized vector of shape (dim, 1)
b -- initialized scalar (corresponds to the bias)
"""
### START CODE HERE ### (≈ 1 line of code)
w = np.zeros((dim, 1))
b = 0
### END CODE HERE ###
assert(w.shape == (dim, 1))
assert(isinstance(b, float) or isinstance(b, int))
return w, b
dim = 2
w, b = initialize_with_zeros(dim)
print ("w = " + str(w))
print ("b = " + str(b))
```
**Expected Output**:
<table style="width:15%">
<tr>
<td> ** w ** </td>
<td> [[ 0.]
[ 0.]] </td>
</tr>
<tr>
<td> ** b ** </td>
<td> 0 </td>
</tr>
</table>
For image inputs, w will be of shape (num_px $\times$ num_px $\times$ 3, 1).
### 4.3 - Forward and Backward propagation
Now that your parameters are initialized, you can do the "forward" and "backward" propagation steps for learning the parameters.
**Exercise:** Implement a function `propagate()` that computes the cost function and its gradient.
**Hints**:
Forward Propagation:
- You get X
- You compute $A = \sigma(w^T X + b) = (a^{(0)}, a^{(1)}, ..., a^{(m-1)}, a^{(m)})$
- You calculate the cost function: $J = -\frac{1}{m}\sum_{i=1}^{m}y^{(i)}\log(a^{(i)})+(1-y^{(i)})\log(1-a^{(i)})$
Here are the two formulas you will be using:
$$ \frac{\partial J}{\partial w} = \frac{1}{m}X(A-Y)^T\tag{7}$$
$$ \frac{\partial J}{\partial b} = \frac{1}{m} \sum_{i=1}^m (a^{(i)}-y^{(i)})\tag{8}$$
```
# GRADED FUNCTION: propagate
def propagate(w, b, X, Y):
"""
Implement the cost function and its gradient for the propagation explained above
Arguments:
w -- weights, a numpy array of size (num_px * num_px * 3, 1)
b -- bias, a scalar
X -- data of size (num_px * num_px * 3, number of examples)
Y -- true "label" vector (containing 0 if non-cat, 1 if cat) of size (1, number of examples)
Return:
cost -- negative log-likelihood cost for logistic regression
dw -- gradient of the loss with respect to w, thus same shape as w
db -- gradient of the loss with respect to b, thus same shape as b
Tips:
- Write your code step by step for the propagation. np.log(), np.dot()
"""
m = X.shape[1] # number of examples
# FORWARD PROPAGATION (FROM X TO COST)
### START CODE HERE ### (≈ 2 lines of code)
A = sigmoid(np.dot(w.T, X) + b) # compute activation
cost = np.sum(Y * np.log(A) + (1 - Y) * np.log(1 - A)) * 1. / (- m) # compute cost
### END CODE HERE ###
# BACKWARD PROPAGATION (TO FIND GRAD)
### START CODE HERE ### (≈ 2 lines of code)
dw = np.dot(X, (A - Y).T) * 1. / m
db = np.sum(A - Y) * 1. / m
### END CODE HERE ###
assert(dw.shape == w.shape)
assert(db.dtype == float)
cost = np.squeeze(cost)
assert(cost.shape == ())
grads = {"dw": dw,
"db": db}
return grads, cost
w, b, X, Y = np.array([[1],[2]]), 2, np.array([[1,2],[3,4]]), np.array([[1,0]])
grads, cost = propagate(w, b, X, Y)
print ("dw = " + str(grads["dw"]))
print ("db = " + str(grads["db"]))
print ("cost = " + str(cost))
```
**Expected Output**:
<table style="width:50%">
<tr>
<td> ** dw ** </td>
<td> [[ 0.99993216]
[ 1.99980262]]</td>
</tr>
<tr>
<td> ** db ** </td>
<td> 0.499935230625 </td>
</tr>
<tr>
<td> ** cost ** </td>
<td> 6.000064773192205</td>
</tr>
</table>
### d) Optimization
- You have initialized your parameters.
- You are also able to compute a cost function and its gradient.
- Now, you want to update the parameters using gradient descent.
**Exercise:** Write down the optimization function. The goal is to learn $w$ and $b$ by minimizing the cost function $J$. For a parameter $\theta$, the update rule is $ \theta = \theta - \alpha \text{ } d\theta$, where $\alpha$ is the learning rate.
```
# GRADED FUNCTION: optimize
def optimize(w, b, X, Y, num_iterations, learning_rate, print_cost = False):
"""
This function optimizes w and b by running a gradient descent algorithm
Arguments:
w -- weights, a numpy array of size (num_px * num_px * 3, 1)
b -- bias, a scalar
X -- data of shape (num_px * num_px * 3, number of examples)
Y -- true "label" vector (containing 0 if non-cat, 1 if cat), of shape (1, number of examples)
num_iterations -- number of iterations of the optimization loop
learning_rate -- learning rate of the gradient descent update rule
print_cost -- True to print the loss every 100 steps
Returns:
params -- dictionary containing the weights w and bias b
grads -- dictionary containing the gradients of the weights and bias with respect to the cost function
costs -- list of all the costs computed during the optimization, this will be used to plot the learning curve.
Tips:
You basically need to write down two steps and iterate through them:
1) Calculate the cost and the gradient for the current parameters. Use propagate().
2) Update the parameters using gradient descent rule for w and b.
"""
costs = []
for i in range(num_iterations):
# Cost and gradient calculation (≈ 1-4 lines of code)
### START CODE HERE ###
grads, cost = propagate(w, b, X, Y)
### END CODE HERE ###
# Retrieve derivatives from grads
dw = grads["dw"]
db = grads["db"]
# update rule (≈ 2 lines of code)
### START CODE HERE ###
w = w - learning_rate * dw
b = b - learning_rate * db
### END CODE HERE ###
# Record the costs
if i % 100 == 0:
costs.append(cost)
# Print the cost every 100 training examples
if print_cost and i % 100 == 0:
print ("Cost after iteration %i: %f" %(i, cost))
params = {"w": w,
"b": b}
grads = {"dw": dw,
"db": db}
return params, grads, costs
params, grads, costs = optimize(w, b, X, Y, num_iterations= 100, learning_rate = 0.009, print_cost = False)
print ("w = " + str(params["w"]))
print ("b = " + str(params["b"]))
print ("dw = " + str(grads["dw"]))
print ("db = " + str(grads["db"]))
```
**Expected Output**:
<table style="width:40%">
<tr>
<td> **w** </td>
<td>[[ 0.1124579 ]
[ 0.23106775]] </td>
</tr>
<tr>
<td> **b** </td>
<td> 1.55930492484 </td>
</tr>
<tr>
<td> **dw** </td>
<td> [[ 0.90158428]
[ 1.76250842]] </td>
</tr>
<tr>
<td> **db** </td>
<td> 0.430462071679 </td>
</tr>
</table>
**Exercise:** The previous function will output the learned w and b. We are able to use w and b to predict the labels for a dataset X. Implement the `predict()` function. There is two steps to computing predictions:
1. Calculate $\hat{Y} = A = \sigma(w^T X + b)$
2. Convert the entries of a into 0 (if activation <= 0.5) or 1 (if activation > 0.5), stores the predictions in a vector `Y_prediction`. If you wish, you can use an `if`/`else` statement in a `for` loop (though there is also a way to vectorize this).
```
# GRADED FUNCTION: predict
def predict(w, b, X):
'''
Predict whether the label is 0 or 1 using learned logistic regression parameters (w, b)
Arguments:
w -- weights, a numpy array of size (num_px * num_px * 3, 1)
b -- bias, a scalar
X -- data of size (num_px * num_px * 3, number of examples)
Returns:
Y_prediction -- a numpy array (vector) containing all predictions (0/1) for the examples in X
'''
m = X.shape[1]
Y_prediction = np.zeros((1,m))
w = w.reshape(X.shape[0], 1)
# Compute vector "A" predicting the probabilities of a cat being present in the picture
### START CODE HERE ### (≈ 1 line of code)
A = sigmoid(np.dot(w.T, X) + b)
### END CODE HERE ###
for i in range(A.shape[1]):
# Convert probabilities A[0,i] to actual predictions p[0,i]
### START CODE HERE ### (≈ 4 lines of code)
Y_prediction[0,i] = 1 if A[0,i] > 0.5 else 0
### END CODE HERE ###
assert(Y_prediction.shape == (1, m))
return Y_prediction
print ("predictions = " + str(predict(w, b, X)))
```
**Expected Output**:
<table style="width:30%">
<tr>
<td>
**predictions**
</td>
<td>
[[ 1. 1.]]
</td>
</tr>
</table>
<font color='blue'>
**What to remember:**
You've implemented several functions that:
- Initialize (w,b)
- Optimize the loss iteratively to learn parameters (w,b):
- computing the cost and its gradient
- updating the parameters using gradient descent
- Use the learned (w,b) to predict the labels for a given set of examples
## 5 - Merge all functions into a model ##
You will now see how the overall model is structured by putting together all the building blocks (functions implemented in the previous parts) together, in the right order.
**Exercise:** Implement the model function. Use the following notation:
- Y_prediction for your predictions on the test set
- Y_prediction_train for your predictions on the train set
- w, costs, grads for the outputs of optimize()
```
# GRADED FUNCTION: model
def model(X_train, Y_train, X_test, Y_test, num_iterations = 2000, learning_rate = 0.5, print_cost = False):
"""
Builds the logistic regression model by calling the function you've implemented previously
Arguments:
X_train -- training set represented by a numpy array of shape (num_px * num_px * 3, m_train)
Y_train -- training labels represented by a numpy array (vector) of shape (1, m_train)
X_test -- test set represented by a numpy array of shape (num_px * num_px * 3, m_test)
Y_test -- test labels represented by a numpy array (vector) of shape (1, m_test)
num_iterations -- hyperparameter representing the number of iterations to optimize the parameters
learning_rate -- hyperparameter representing the learning rate used in the update rule of optimize()
print_cost -- Set to true to print the cost every 100 iterations
Returns:
d -- dictionary containing information about the model.
"""
### START CODE HERE ###
# initialize parameters with zeros (≈ 1 line of code)
w, b = initialize_with_zeros(X_train.shape[0])
# Gradient descent (≈ 1 line of code)
parameters, grads, costs = optimize(w, b, X_train, Y_train, num_iterations, learning_rate, print_cost)
# Retrieve parameters w and b from dictionary "parameters"
w = parameters["w"]
b = parameters["b"]
# Predict test/train set examples (≈ 2 lines of code)
Y_prediction_test = predict(w, b, X_test)
Y_prediction_train = predict(w, b, X_train)
### END CODE HERE ###
# Print train/test Errors
print("train accuracy: {} %".format(100 - np.mean(np.abs(Y_prediction_train - Y_train)) * 100))
print("test accuracy: {} %".format(100 - np.mean(np.abs(Y_prediction_test - Y_test)) * 100))
d = {"costs": costs,
"Y_prediction_test": Y_prediction_test,
"Y_prediction_train" : Y_prediction_train,
"w" : w,
"b" : b,
"learning_rate" : learning_rate,
"num_iterations": num_iterations}
return d
```
Run the following cell to train your model.
```
d = model(train_set_x, train_set_y, test_set_x, test_set_y, num_iterations = 2000, learning_rate = 0.005, print_cost = True)
```
**Expected Output**:
<table style="width:40%">
<tr>
<td> **Train Accuracy** </td>
<td> 99.04306220095694 % </td>
</tr>
<tr>
<td>**Test Accuracy** </td>
<td> 70.0 % </td>
</tr>
</table>
**Comment**: Training accuracy is close to 100%. This is a good sanity check: your model is working and has high enough capacity to fit the training data. Test error is 68%. It is actually not bad for this simple model, given the small dataset we used and that logistic regression is a linear classifier. But no worries, you'll build an even better classifier next week!
Also, you see that the model is clearly overfitting the training data. Later in this specialization you will learn how to reduce overfitting, for example by using regularization. Using the code below (and changing the `index` variable) you can look at predictions on pictures of the test set.
```
# Example of a picture that was wrongly classified.
index = 1
plt.imshow(test_set_x[:,index].reshape((num_px, num_px, 3)))
print ("y = " + str(test_set_y[0,index]) + ", you predicted that it is a \"" + classes[d["Y_prediction_test"][0,index]].decode("utf-8") + "\" picture.")
```
Let's also plot the cost function and the gradients.
```
# Plot learning curve (with costs)
costs = np.squeeze(d['costs'])
plt.plot(costs)
plt.ylabel('cost')
plt.xlabel('iterations (per hundreds)')
plt.title("Learning rate =" + str(d["learning_rate"]))
plt.show()
```
**Interpretation**:
You can see the cost decreasing. It shows that the parameters are being learned. However, you see that you could train the model even more on the training set. Try to increase the number of iterations in the cell above and rerun the cells. You might see that the training set accuracy goes up, but the test set accuracy goes down. This is called overfitting.
## 6 - Further analysis (optional/ungraded exercise) ##
Congratulations on building your first image classification model. Let's analyze it further, and examine possible choices for the learning rate $\alpha$.
#### Choice of learning rate ####
**Reminder**:
In order for Gradient Descent to work you must choose the learning rate wisely. The learning rate $\alpha$ determines how rapidly we update the parameters. If the learning rate is too large we may "overshoot" the optimal value. Similarly, if it is too small we will need too many iterations to converge to the best values. That's why it is crucial to use a well-tuned learning rate.
Let's compare the learning curve of our model with several choices of learning rates. Run the cell below. This should take about 1 minute. Feel free also to try different values than the three we have initialized the `learning_rates` variable to contain, and see what happens.
```
learning_rates = [0.01, 0.001, 0.0001]
models = {}
for i in learning_rates:
print ("learning rate is: " + str(i))
models[str(i)] = model(train_set_x, train_set_y, test_set_x, test_set_y, num_iterations = 1500, learning_rate = i, print_cost = False)
print ('\n' + "-------------------------------------------------------" + '\n')
for i in learning_rates:
plt.plot(np.squeeze(models[str(i)]["costs"]), label= str(models[str(i)]["learning_rate"]))
plt.ylabel('cost')
plt.xlabel('iterations')
legend = plt.legend(loc='upper center', shadow=True)
frame = legend.get_frame()
frame.set_facecolor('0.90')
plt.show()
```
**Interpretation**:
- Different learning rates give different costs and thus different predictions results.
- If the learning rate is too large (0.01), the cost may oscillate up and down. It may even diverge (though in this example, using 0.01 still eventually ends up at a good value for the cost).
- A lower cost doesn't mean a better model. You have to check if there is possibly overfitting. It happens when the training accuracy is a lot higher than the test accuracy.
- In deep learning, we usually recommend that you:
- Choose the learning rate that better minimizes the cost function.
- If your model overfits, use other techniques to reduce overfitting. (We'll talk about this in later videos.)
## 7 - Test with your own image (optional/ungraded exercise) ##
Congratulations on finishing this assignment. You can use your own image and see the output of your model. To do that:
1. Click on "File" in the upper bar of this notebook, then click "Open" to go on your Coursera Hub.
2. Add your image to this Jupyter Notebook's directory, in the "images" folder
3. Change your image's name in the following code
4. Run the code and check if the algorithm is right (1 = cat, 0 = non-cat)!
```
## START CODE HERE ## (PUT YOUR IMAGE NAME)
my_image = "roller_cat.jpg" # change this to the name of your image file
## END CODE HERE ##
# We preprocess the image to fit your algorithm.
fname = "images/" + my_image
image = np.array(ndimage.imread(fname, flatten=False))
my_image = scipy.misc.imresize(image, size=(num_px,num_px)).reshape((1, num_px*num_px*3)).T
my_predicted_image = predict(d["w"], d["b"], my_image)
plt.imshow(image)
print("y = " + str(np.squeeze(my_predicted_image)) + ", your algorithm predicts a \"" + classes[int(np.squeeze(my_predicted_image)),].decode("utf-8") + "\" picture.")
```
<font color='blue'>
**What to remember from this assignment:**
1. Preprocessing the dataset is important.
2. You implemented each function separately: initialize(), propagate(), optimize(). Then you built a model().
3. Tuning the learning rate (which is an example of a "hyperparameter") can make a big difference to the algorithm. You will see more examples of this later in this course!
Finally, if you'd like, we invite you to try different things on this Notebook. Make sure you submit before trying anything. Once you submit, things you can play with include:
- Play with the learning rate and the number of iterations
- Try different initialization methods and compare the results
- Test other preprocessings (center the data, or divide each row by its standard deviation)
Bibliography:
- http://www.wildml.com/2015/09/implementing-a-neural-network-from-scratch/
- https://stats.stackexchange.com/questions/211436/why-do-we-normalize-images-by-subtracting-the-datasets-image-mean-and-not-the-c
| github_jupyter |
```
# 安装pandas
# pip install Pandas
# 运行测试套件
# 运行前需要安装: hypothesis和pytest
import pandas as pd
# pd.test()
# 对象创建
# 传入一些值的列表来创建一个Series,pandas会自动创建一个默认的整数索引.
import pandas as pd
import numpy as np
import pprint
s = pd.Series([1,3,5,np.nan,6,8])
print(s)
print('-'*30)
# 传递带有日期时间索引和带标签列的NumPy数组来创建DataFrame
dates = pd.date_range('20190815',periods=6)
pprint.pprint(dates)
df = pd.DataFrame(np.random.randn(6,4),index=dates,columns=list('ABCD'))
# df.to_excel('./output.xlsx')
pprint.pprint(df)
# 转化为类似Series的dict对象来创建DataFrame
df2 = pd.DataFrame({'A': 1.,
'B': pd.Timestamp('20190820'),
'C': pd.Series(1,index=list(range(4)),dtype='float32'),
'D': np.array([3] * 4,dtype='int32'),
'E': pd.Categorical(["test","train","test1","train2"]),
'F': 'foo'})
print(df2)
# DataFrame的列具有不同的数据类型
print(df2.dtypes)
### 查看数据
# 查看DataFrame顶部数据
print(df.head(3))
print('+='*30)
# 查看DataFrame尾部数据
print(df.tail(3))
print('--+'*30)
# 显示索引,列和底层NumPy数据.
print(df.index)
print('-='*30)
print(df.columns)
print('-|'*30)
# DataFrame.to_numpy() 会给出Numpy对象. 输出时不包含行索引和列索引.
print(df.to_numpy())
# describe()方法显示数据的快速统计摘要
print(df.describe())
print('--'*30)
# 转置数据
print(df.T)
print('=='*30)
# 按轴排序
print(df.sort_index(axis=1,ascending=False))
print('-='*30)
# 按值排序
print(df.sort_values(by='B'))
### 获取
print(df['A'])
# 对行进行切片
print(df[0:])
print(df[0:2])
print('-=='*30)
print(df['20190816':'20190818'])
### 按标签选择
# 通过标签获取一行数据
print(df.loc[dates[0]])
print(df.loc[dates[1]])
print('=='*30)
# 通过标签在多个轴上选择数据
print('通过标签在多个轴上选择数据')
print(df.loc[:,['A','B']])
print('--'*30)
print(df.loc[:,['C']])
# 通过标签同时在两个轴上切片
print('通过标签同时在两个轴上切片')
print(df.loc['20190817':'20190819',['A','B']])
# 减小返回对象的大小
print(df.loc['20190820',['A','B']])
# 获取标量值
print(df.loc[dates[0],'A'])
# 快速访问标量
print(df.at[dates[0],'A'])
### 布尔索引
# 使用单个列的值来选择数据
print(df[df.A > 0]) # 会输出为True的内容.
print(df.A > 0) # 将True和False都打印出来.
# 从满足布尔条件的DataFrame中选择值:
print(df[df > 0])
# 使用isin()方法过滤
df3 = df.copy()
# print(df3)
# df3['E'] = ['one','one','two','three','four','three']
df3['E'] = ['one','two','three','four','five','six']
# print(df3)
print('-='*30)
print(df3[df3['E'].isin(['two','four'])])
### 赋值
# 添加新列将自动根据索引对齐数据.
s1 = pd.Series([1,2,3,4,5,6],index=pd.date_range('20190818',periods=6))
print(s1)
df3['F'] = s1
print(df3['F'])
# 通过标签赋值
df3.at[dates[0],'A'] = 0
print(df3)
### 通过位置赋值
df.iat[0,1] = 0
print(df)
# 使用NumPy数组赋值
df3.loc[:,'D'] = np.array([5] * len(df))
print(df3) # 前面一系列赋值操作的结果.
# 带有where条件的赋值操作.
df2 = df.copy()
df2[df2 > 0] = -df2
print(df2)
### 缺失值
# pandas主要使用值np.nan来表示缺失的数据.
# 重建索引允许更改/添加/删除指定轴上的索引.这个操作会返回一个副本.
df5 = df.reindex(index=dates[0:4],columns=list(df.columns) + ['E'])
df5.loc[dates[0]:dates[1],'E'] = 1
print(df5)
# 删除任何带有缺失值的行
print(df5.dropna(how='any'))
# 填充缺失值
print(df5.fillna(value=5))
# 获取值为nan的掩码
print(pd.isna(df5))
### 统计
# 进行描述性统计
print(df5.mean())
print('-='*30)
# 在其它轴上进行同样的操作:
print(df5.mean(1))
# 使用具有不同维度且需要对齐的对象进行操作. pandas会自动沿指定维度进行广播.
s = pd.Series([1,3,5,np.nan,6,8],index=dates).shift(2)
print(s)
print(df5.sub(s,axis='index'))
### 应用
# 将函数应用于数据
print(df5.apply(np.cumsum))
print(df5.apply(lambda x: x.max() - x.min()))
### 直方图化
s1 = pd.Series(np.random.randint(0,7,size=10))
print(s1)
print('=+'*30)
print(s1.value_counts())
### 字符串方法
# Series在str属性中有一组字符串处理方法,可对数组的每个元素进行操作.
s2 = pd.Series(['A','B','C','Aaba','Baca',np.nan,'CABA','dog','cat'])
print(s2.str.lower())
## 合并
### 连接
# 使用concat()连接pandas对象.
df6 = pd.DataFrame(np.random.randn(10,4))
print(df6)
print('---'*30)
pieces = [df6[:3],df6[3:7],df6[7:]]
print(pd.concat(pieces))
### Join
# SQL风格的合并
left = pd.DataFrame({'key': ['foo','foo'],'lval':[1,2]})
right = pd.DataFrame({'key': ['foo','foo'],'rval': [4,5]})
print(left)
print('='*30)
print(right)
print('='*30)
print(pd.merge(left,right,on='key'))
# 另一个例子
left = pd.DataFrame({'key': ['foo','bar'],'lval': [1,2]})
right = pd.DataFrame({'key': ['foo','bar'],'rval':[4,5]})
print(left)
print('-'*30)
print(right)
print(pd.merge(left,right,on='key'))
### 追加
df7 = pd.DataFrame(np.random.randn(8,4),columns=['A','B','C','D'])
print(df7)
print('=='*30)
s3 = df7.iloc[3]
print(df7.append(s3,ignore_index=True))
### 分组
"""
group by包括:
分割: 根据一些标准将数据分解成组.
应用: 将函数独立地应用于每个组.
组合: 将结果组合成数据结构.
"""
df8 = pd.DataFrame({'A': ['foo','bar','foo','bar',
'foo','bar','foo','foo'],
'B': ['one','one','two','three',
'two','two','one','three'],
'C': np.random.randn(8),
'D': np.random.randn(8)})
print(df8)
# 分组,然后将sum()函数应用于分组结果.
print(df8.groupby('A').sum())
print('=-'*30)
# 按多列分组形成层次索引,用sum函数
print(df8.groupby(['A','B']).sum())
### 堆叠(Stack)
tuples = list(zip(*[['bar','bar','baz','baz',
'foo','foo','qux','qux'],
['one','two','one','two',
'one','two','one','two']]))
index = pd.MultiIndex.from_tuples(tuples,names=['first','second'])
df = pd.DataFrame(np.random.randn(8,2),index=index,columns=['A','B'])
df9 = df[:4]
print(df9)
### stack()方法压缩DataFrame的列
stacked = df9.stack()
print(stacked)
# stack()的逆操作是unstack(),默认情况下取消最后压缩的哪个级别.
print(stacked.unstack())
print('=='*30)
print(stacked.unstack(1))
print('-='*30)
print(stacked.unstack(0))
### 数据透视表
df10 = pd.DataFrame({'A': ['one','one','two','three'] * 3,
'B': ['A','B','C'] * 4,
'C': ['foo','foo','foo','bar','bar','bar'] * 2,
'D': np.random.randn(12),
'E': np.random.randn(12)})
print(df10)
print('-='*30)
# 从这些数据生成数据透视表
pd.pivot_table(df10,values='D',index=['A','B'],columns=['C'])
### 时间序列(TimeSeries)
# 用于在频率转换期间执行重采样操作.
rng = pd.date_range('22/08/2019',periods=100,freq='S')
ts = pd.Series(np.random.randint(0,500,len(rng)),index=rng)
print(ts.resample('5Min').sum())
# 时区代表
rng = pd.date_range('21/08/2019 21:29:30',periods=5,freq='D')
ts = pd.Series(np.random.randn(len(rng)),rng)
print(ts)
print('-='*30)
ts_utc = ts.tz_localize('UTC')
print(ts_utc)
print('-='*30)
# 转换为另一个时区
print(ts_utc.tz_convert('US/Eastern'))
# 在时间跨度表示之间转换
rng = pd.date_range('22/08/2019',periods=5,freq='M')
ts = pd.Series(np.random.randn(len(rng)),index=rng)
print(ts)
print('-='*30)
ps = ts.to_period()
print(ps)
print('-='*30)
print(ps.to_timestamp())
# 周期和时间戳之间的转换可以用算术函数.
# 示例: 以11月为结束年份的季度频率转换为季度结束后一个月末的上午9点.
prng = pd.period_range('2010Q1','2019Q4',freq='Q-NOV')
ts = pd.Series(np.random.randn(len(prng)),prng)
ts.index = (prng.asfreq('M','e') + 1).asfreq('H','s') + 9
print(ts.head())
### 分类(Categoricals)
# pandas可以在DataFrame中包含分类数据.
df11 = pd.DataFrame({"id": [1,2,3,4,5,6],
"raw_grade": ['a','b','b','a','a','e']})
# 将原始成绩转换为category数据类型
df11["grade"] = df11["raw_grade"].astype("category")
print(df11["grade"])
print('-='*30)
# 将类别重命名为更有意义的名称(通过调用Series.cat.categories来替换)
df11["grade"].cat.categories = ["very good","good","very bad"]
print(df11["grade"].cat.categories)
# 对categories重新排序并同时添加缺少的category(Series.cat下的方法默认返回一个新的Series)
df11["grade"] = df11["grade"].cat.set_categories(["very bad","bad","medium",
"good","very good"])
print(df11["grade"])
# 排序时按categories中的顺序排序,不是按照词汇顺序排序.
print(df11.sort_values(by="grade"))
# 按分好类的列分组(groupby)可以显示空categories。
print(df11.groupby("grade").size())
### 绘图
ts = pd.Series(np.random.randn(1000),
index=pd.date_range('22/08/2019',periods=1000))
ts = ts.cumsum()
ts.plot()
import matplotlib.pyplot as plt
# 在一个DataFrame中,plot方法绘制带有label的所有列.
df12 = pd.DataFrame(np.random.randn(1000,4),index=ts.index,
columns=['A','B','C','D'])
df13 = df12.cumsum()
plt.figure()
df13.plot()
plt.legend(loc='best')
### 数据输入/输出
# 写入csv文件
df13.to_csv('./best.csv')
# 从csv文件读数据
pd.read_csv('./best.csv')
### HDF5
# pip install tables
# 写入HDF5
df13.to_hdf('./best.h5','df')
# 从HDF5读数据
pd.read_hdf('./best.h5','df')
### Excel
# 写入excel文件
df13.to_excel('./best.xlsx',sheet_name='best')
# 从excel文件读取数据
pd.read_excel('./best.xlsx','best',index_col=None,na_values=['NA'])
# Gotchas 坑
# 异常
if pd.Series([False,True,False]):
print("I was true")
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.